Interactive visualization tool for (meta)genome assembly graphs.
MetagenomeScope decomposes the graph into structural patterns and highlights these as annotations on the graph. By default it lays out the graph hierarchically, using Graphviz' dot algorithm.
MetagenomeScope also contains various functionalities for visualizing assembly graphs at larger scales -- for example, highlighting scaffold paths on the graph and drawing summary plots of the graph's structure.
MetagenomeScope supports the outputs of most modern assemblers, can handle large graphs including tens of thousands of nodes, and is backed by over four hundred automatic software tests.
The tool is under active development, so please let us know if you have any feedback!
| Stool metagenome assembly (MetaCarvel) | Yeast genome assembly (Flye) |
|---|---|
 |
 |
| Data source: SRS049959 | Data source: AGB |
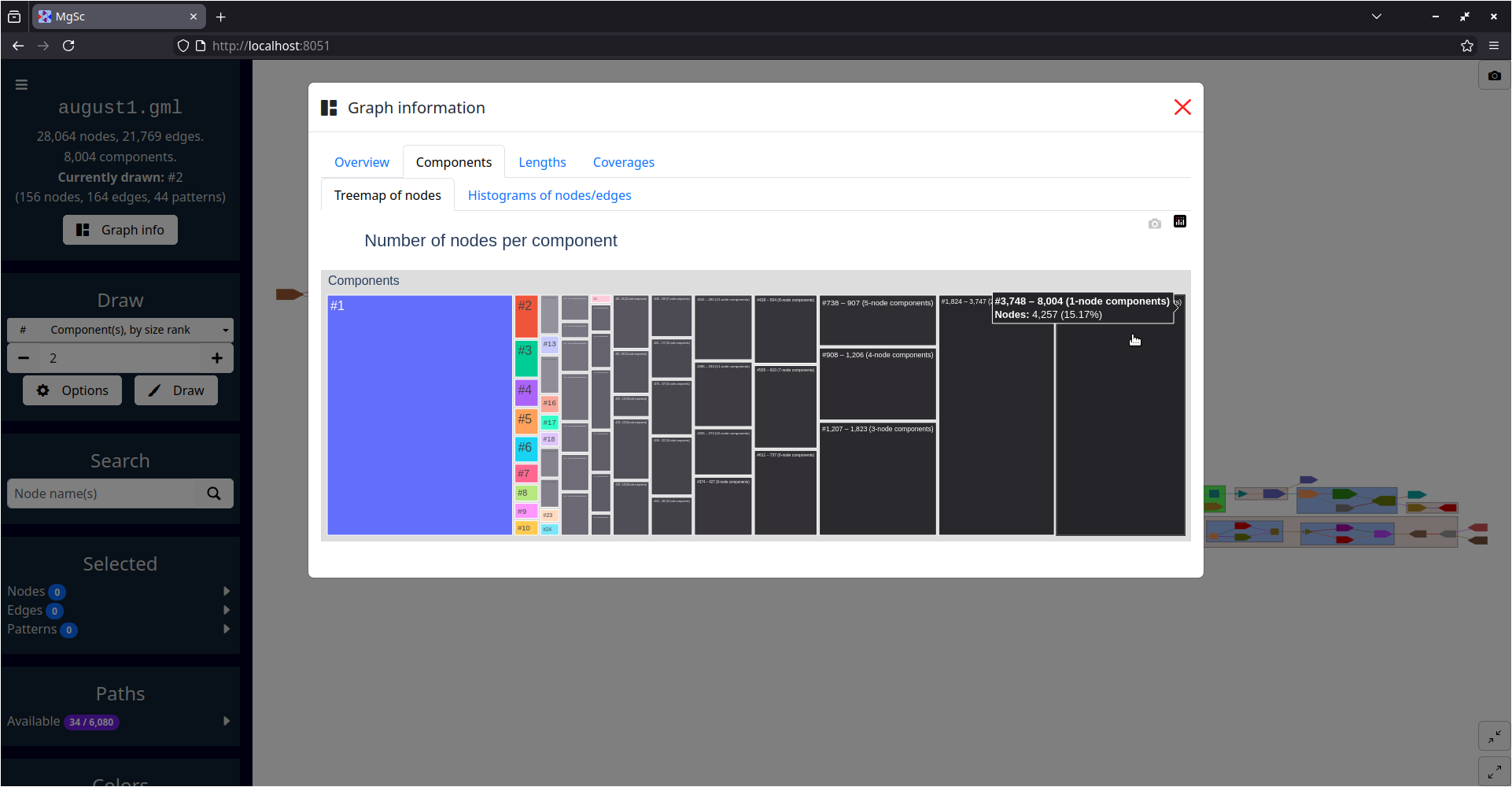
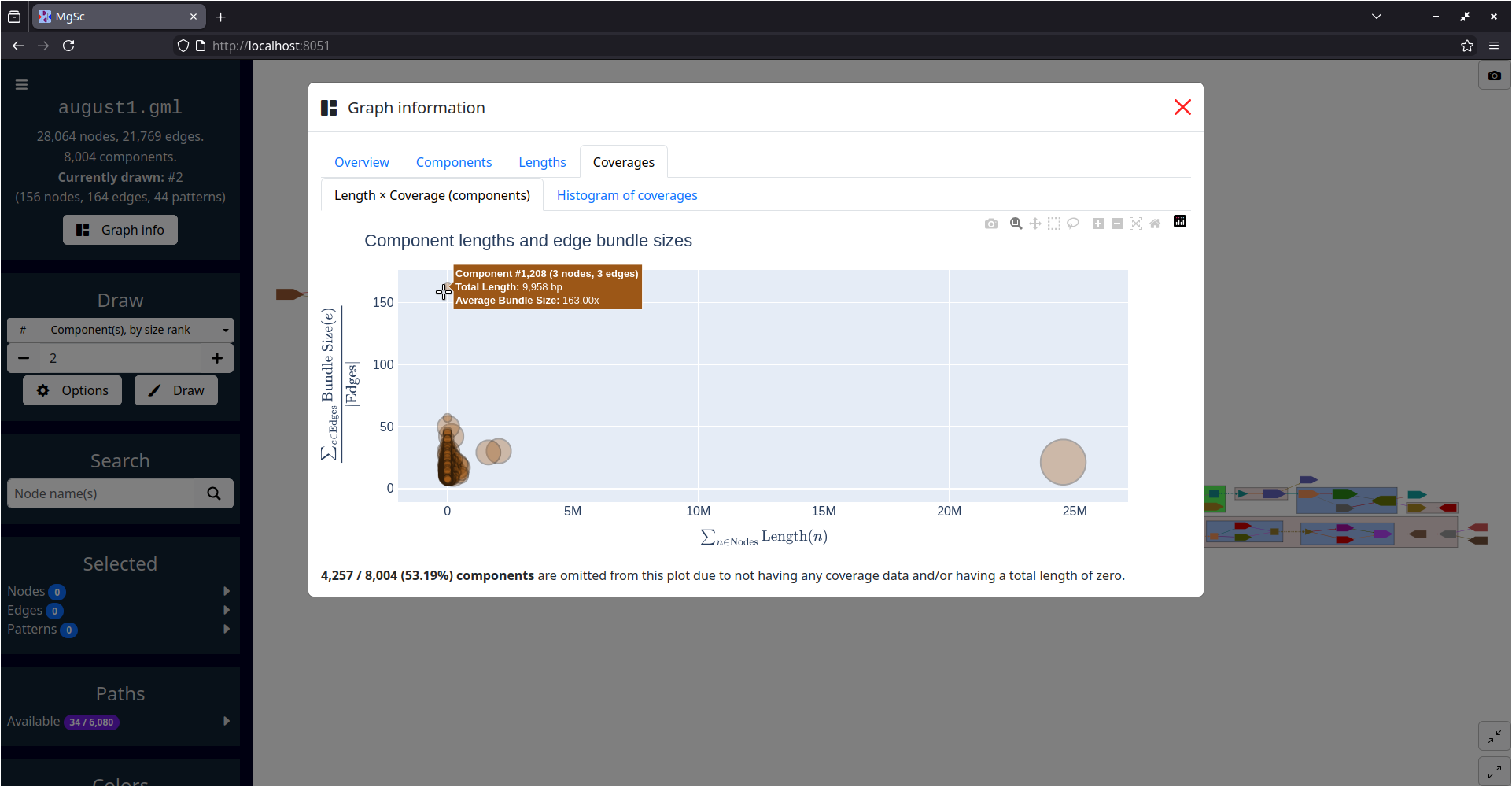
| Summarizing graph structure in a treemap | Interactive charts of graph statistics |
|---|---|
 |
 |
| Data source: SRS049959 | |
mamba install -c bioconda -c conda-forge metagenomescopeUsing pip
First, you need to make sure that Graphviz
and Pygraphviz are installed
properly (so that Pygraphviz knows where to find Graphviz).
See Pygraphviz'
INSTALL.txt
for details.
(Probably the most consistent way to do this is just installing Graphviz and Pygraphviz from conda-forge, but at that point you might as well do the entire installation from within conda...)
Anyway, once Graphviz and Pygraphviz are installed, you should be able to just run:
pip install metagenomescopemgsc -g graph.gfa... where graph.gfa is a path to the assembly graph you want to visualize
(see information below on supported graph filetypes).
This will start a server using Dash.
The port number of the server defaults to 8050, so navigate
to localhost:8050 in a web browser to access the visualization.
Usage: mgsc [OPTIONS]
Visualizes an assembly graph.
Please visit https://github.com/marbl/MetagenomeScope for more information.
Options:
-g, --graph FILE In GFA, FASTG, DOT, GML, or LastGraph format. [required]
-a, --agp FILE AGP file describing paths (e.g. scaffolds) in the graph.
-i, --info FILE Flye assembly_info.txt file describing contigs/scaffolds.
-p, --port INTEGER RANGE Server port number. [default: 8050; 1024<=x<=65535]
--verbose / --no-verbose Log extra details. [default: no-verbose]
--debug / --no-debug Use Dash's debug mode. [default: no-debug]
-v, --version Show the version and exit.
-h, --help Show this message and exit.
| Filetype | Generated by | Notes |
|---|---|---|
GFA (.gfa) |
Flye, LJA, hifiasm, verkko, ... | Both GFA 1 and GFA 2 files are accepted. Currently we visualize segments, links (GFA 1), non-containment edges (GFA 2), and paths of segments. |
FASTG (.fastg) |
SPAdes, MEGAHIT | Expects FASTG files produced by SPAdes or MEGAHIT. |
DOT (.dot, .gv) |
Flye, LJA | Expects DOT files produced by Flye or LJA. See "What filetype should I use for de Bruijn graphs?" in the FAQs below. |
GML (.gml) |
MetaCarvel | Expects GML files produced by MetaCarvel. |
LastGraph (.LastGraph) |
Velvet | Currently we just visualize the raw structure (nodes and arcs). |
Should you run into additional assembly graph filetypes you'd like us to support, feel free to open a GitHub issue.
Paths can optionally be specified through any of the following inputs:
AGP files (-a)
See the AGP specification for details.
If your graph is in DOT format:
- We assume the
component_ids in column 6a of the AGP file correspond to edge IDs.
Otherwise:
- We assume the
component_ids correspond to node IDs.
Flye assembly_info.txt files (-i)
See Flye's documentation for details.
If your graph is in DOT format:
- We will visualize the edge-paths described in the
.txtfile.
If your graph is in GFA format:
-
The contigs in the GFA file should correspond to collapsed edge-paths in the
.txtfile, so we can't really visualize these edge-paths. -
However, we will extract contig information from the
.txtfile (e.g. coverage) and show it in the interface as node data.
If your graph is not in DOT or GFA format:
- We will ignore the
.txtfile. Flye should only generate DOT or GFA files, so like... where did you even get this data from 💀
P-lines in GFA 1 files, or O-lines in GFA 2 files (-g)
See the GFA 1 and GFA 2 specifications for details.
Currently, we only show segments on GFA paths (not edges, gaps, etc.)
MetagenomeScope detects and highlights five types of structural patterns on the graph:
Bubbles (Miller et al., 2010; Nijkamp et al., 2013) follow a diverge-coverge pattern. They generally indicate variation -- either real (e.g. an alternate path is caused by a SNP) or erroneous (e.g. an alternate path is caused by a sequencing error). We identify bubbles using a modified version of the algorithm given in Onodera et al., 2013.
Similarly, bulges (Pevzner et al., 2004; Vasilinetc et al., 2015) are pairs of nodes where there exist multiple parallel edges from one node to another.
Bulges can typically be interpreted the same way as bubbles -- you generally see bulges in "edge-centric" (e.g. de Bruijn) graphs, and bubbles in "node-centric" (e.g. overlap) graphs. So, we label both bubbles and bulges identically.
Frayed ropes (Miller et al., 2010) follow a converge-diverge pattern; they have the opposite structure as bubbles. They generally indicate interspersed repeats in the middle region.
Chains are just non-branching paths of at least two nodes. Cyclic chains are chains where the end node has an outgoing edge to the start node. Cyclic chains represent a simpler form of what are known in edge-centric graphs as whirl structures (Pevzner et al., 2004).
Bipartites are regions of the graph that can be partitioned into two layers of nodes (let's call them Left and Right), such that all of the nodes in Left have outgoing edges to all of the nodes in Right. We require that both Left and Right contain at least two nodes each. (Such a pattern is essentially a stricter version of a complete bipartite graph.)
Surprisingly, bipartites pop up a lot in certain assembly graphs! These are less well-documented in the literature than the above types of patterns, but our suspicion is that these are another indication (like frayed ropes) of repeats -- and that a lot of these patterns in succession might indicate things like strain heterogeneity. See Figure 5 of Li et al., 2012 for an example of how a bipartite (or, viewed another way, a frayed rope) could be caused by a repeat.
Sometimes, it is best to consider a node as the child of two patterns. A common example of this is a bubble chain (Dabbaghie et al., 2022), where multiple bubbles occur one after another. In a bubble chain, the "end node" of one bubble is also the "start node" of another bubble!
To accommodate these cases, MetagenomeScope splits the boundary nodes of a pattern.
Splitting a node A transforms it into two nodes: A-L and A-R, which are connected by a single "fake edge" A-L -> A-R.
Because this allows a node to be in two patterns simultaneously, this makes it possible for us to identify a much richer set of patterns and describe the graph structure more accurately.
"Split nodes" and "fake edges" are drawn with distinct visual styles, in order to make them clearer -- split nodes are drawn in a way that looks like the node has been split in half, and fake edges are drawn as thick dashed lines.
| Examples of split nodes | |
|---|---|
 |
 |
| MetaCarvel stool metagenome scaffold graph (the "large graph" shown below), component #17 | jumboDBG de Bruijn graph of human chromosome 15 (available as chr15_full.gv in metagenome/tests/input/) |
Note that node splitting is not always necessary -- as the figures above show, sometimes a boundary node of a pattern doesn't need to be the boundary node of any other pattern. To limit the amount of split nodes we need to show in the visualization, we detect and remove unnecessary split nodes after finishing the decomposition procedure.
Here are three graphs of various sizes, each produced by a different assembly program.
This data is from AGB's GitHub repository.
wget https://raw.githubusercontent.com/marbl/MetagenomeScope/refs/heads/main/metagenomescope/tests/input/flye_yeast.gv
wget https://raw.githubusercontent.com/marbl/MetagenomeScope/refs/heads/main/metagenomescope/tests/input/flye_yeast_assembly_info.txt
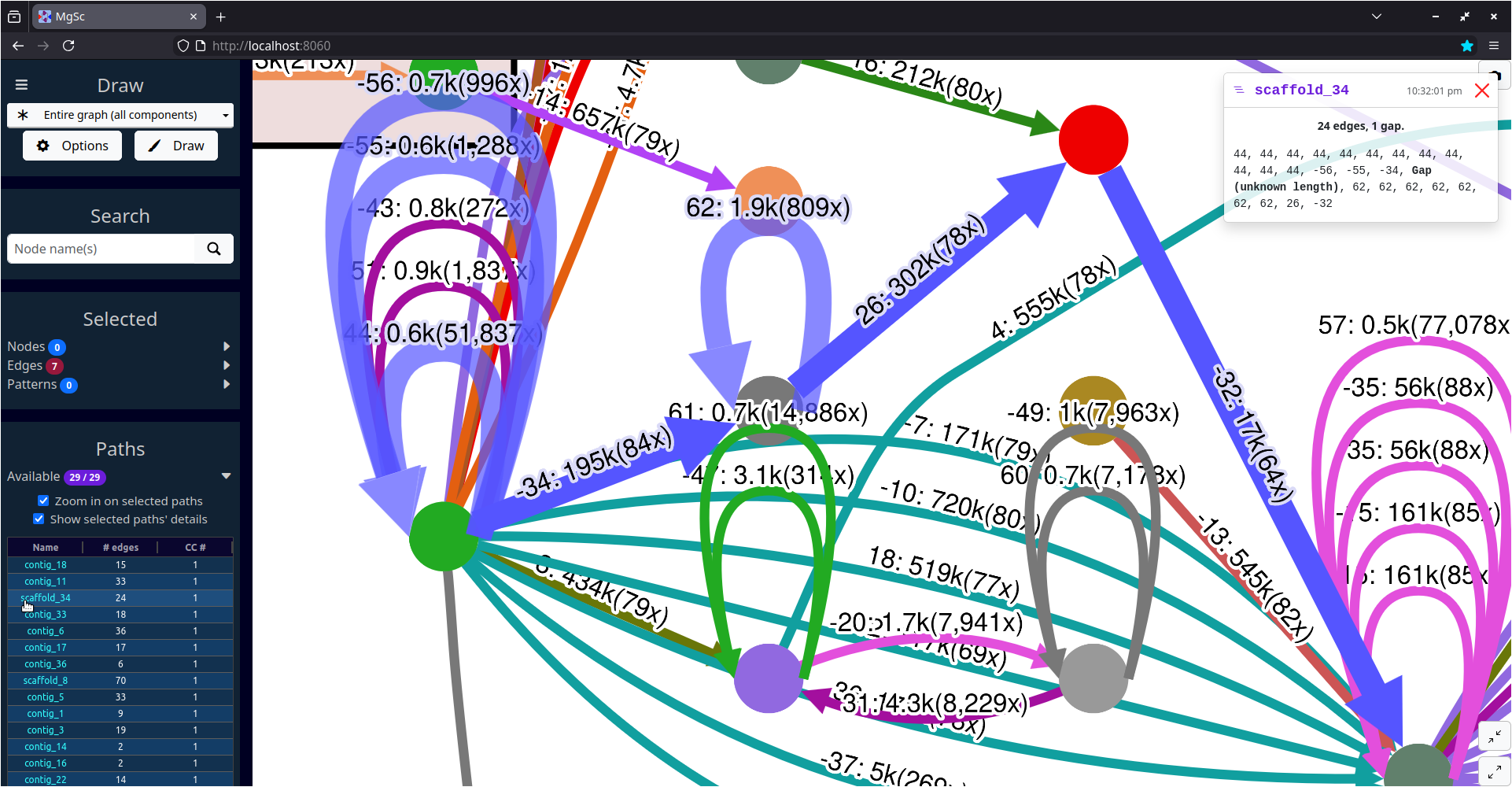
mgsc -g flye_yeast.gv -i flye_yeast_assembly_info.txt| Yeast assembly graph screenshots | |
|---|---|
 |
 |
| Entire graph | Zoomed in on scaffold_34 in component #1 |
Tip
The "Labels" section of the interface has some settings that help make labels prettier. I produced the screenshot on the right above using the Offset, Outline, and Rotate edge label settings.
Note
We draw DOT files from Flye and LJA using the typical conventions for drawing de Bruijn graphs -- with nodes represented as circles, and edges given labels with their length and coverage. This resembles the styles from various papers that show visualizations of these kinds of graphs, including Pevzner et al., 2004; Mikheenko & Kolmogorov 2019; and the DOT outputs of Flye and LJA.
This is an example graph from Bandage.
wget https://github.com/rrwick/Bandage/raw/refs/heads/gh-pages/samples/E_coli_LastGraph.zip
unzip E_coli_LastGraph.zip
mgsc -g E_coli_LastGraph| E. coli assembly graph screenshots | |
|---|---|
 |
 |
| Hierarchical layout with dot | Force-directed layout with sfdp (using an overlap scaling factor of -15) |
Note
As discussed in the FAQs below on "Reverse-complementary sequences," we represent each pair of nodes (A, -A) separately in the graph. This makes it easier to lay out the graph nicely.
Notice that components #2 and #3 in these screenshots are reverse-complements of each other, as are components #4 and #5, etc.
However, component #1 is "strand-mixed" -- it contains both nodes 1 and -1!
This is a scaffold graph created by MetaCarvel. Note that this graph is fairly old (it dates back to August 2017!); MetaCarvel has been updated a decent amount since then.
wget https://zenodo.org/records/18316065/files/august1.gml
wget https://zenodo.org/records/18316065/files/scaffolds_august1_fixed.agp
# Use --verbose to show more information in the terminal about how long each step takes
mgsc -g august1.gml -a scaffolds_august1_fixed.agp --verbose| Stool metagenome scaffold graph screenshots | |
|---|---|
 |
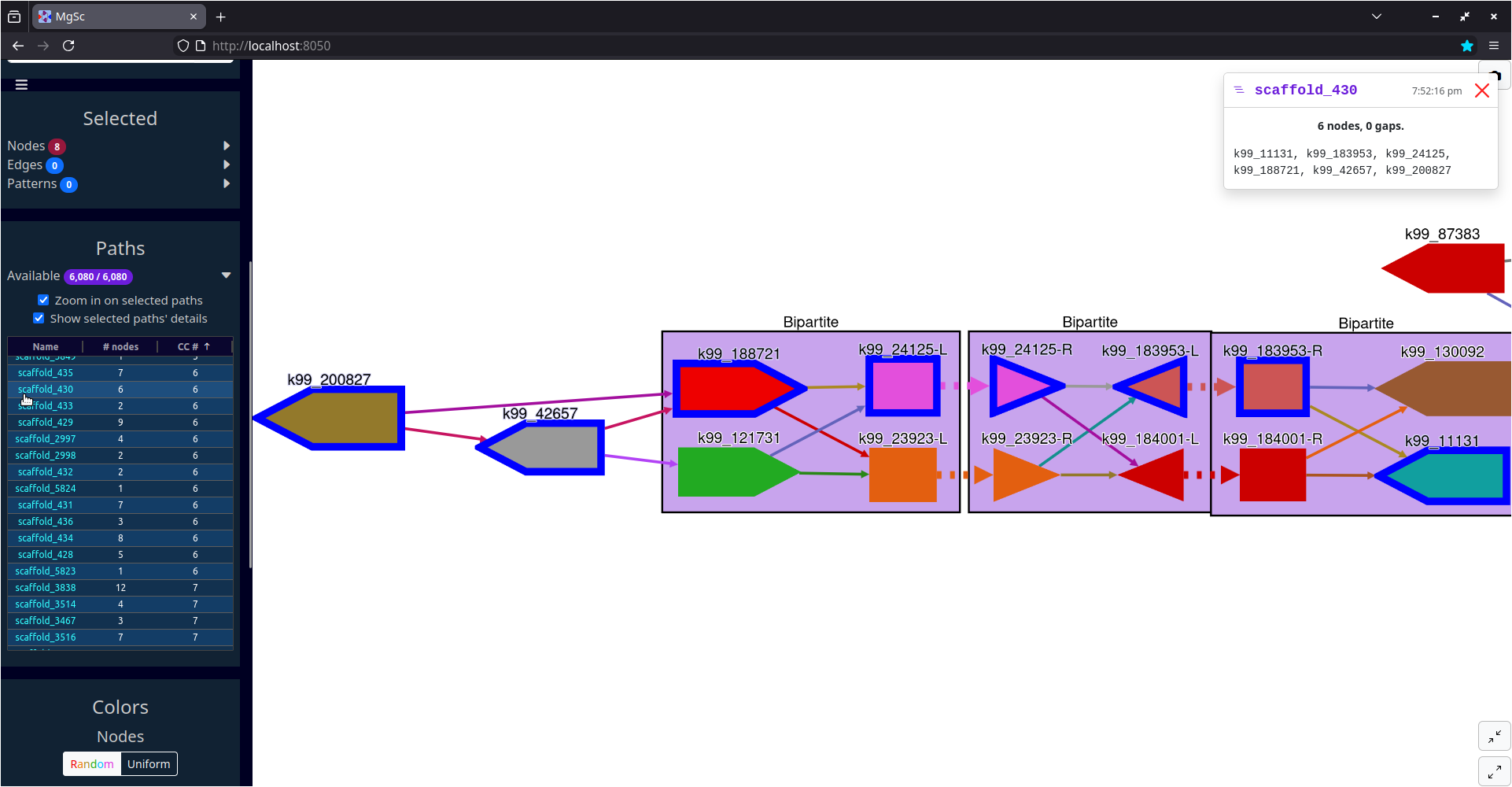 |
| Entire graph (on my 2018 laptop this takes about 2.5 minutes to lay out and draw; see tip below) | Zoomed in on scaffold_430 in component #6 |
Tip
You can get a sense for the sizes of this graph's connected components by clicking the "Graph info" button in the left sidebar, then examining the charts in the "Components" tab.
This shows that the largest connected component in this graph (i.e. the one with size rank #1) contains over six thousand nodes. You can certainly draw this component in MetagenomeScope, but it will take a few seconds to lay out and draw. The resulting interface may also become a bit sluggish due to the size of the graph. (You could even draw the entire graph if you wanted to, as shown above! But it will be even more sluggish.)
If you would like to examine the smaller parts of this graph, you can start by drawing component #2 -- or even a range of components, for example #2 - 10. (Try pasting 2-10 into the "Component(s), by size rank" input to draw all of these components at once!)
You can also draw only a subregion of a larger component, using functionality inspired by Bandage. In the "Draw" section, change the "Component(s), by size rank" dropdown to the "Around certain node(s)" option, and then type in k99_38. Try increasing the distance and redrawing to see more and more of component #1!
See the metagenomescope/tests/input/
directory.
FAQ 1. How do you handle reverse-complementary nodes/edges?
The answer to this depends on the filetype of the graph you are using.
When MetagenomeScope reads in FASTG, DOT, and GML files, it assumes that these files explicitly describe all of the nodes and edges in the graph. So, let's say you give MetagenomeScope the following DOT file:
digraph g {
1 -> 2 [label="edge1 A99(2.4)"];
}We will interpret this as a graph with two nodes (1, 2) and one edge
(1 -> 2).
However, for GFA and LastGraph files, MetagenomeScope cannot make the assumption that these files explicitly describe all of the nodes and edges in the graph. In these files, each declaration of a node / edge (in GFA parlance, "segment" / "link"; in LastGraph parlance, "node" / "arc") also declares this node / edge's reverse complement.
So, let's say you give MetagenomeScope the following GFA file (based on this example):
H VN:Z:1.0
S 1 CGATGCAA
S 2 TGCAAAGTAC
L 1 + 2 + 5M
We will interpret this as a graph with four nodes (1, -1, 2, -2)
and two edges (1 -> 2, -2 -> -1). The presence of node X
"implies"
the existence of the reverse complement node -X, and the presence of edge
X -> Y "implies" the existence of the reverse complement edge -Y -> -X.
Interpreting the graph file in this way is analogous to
how "double mode" works in Bandage.
Based on the FASTG specification, shouldn't FASTG be an "implicit" instead of an "explicit" filetype?
It's complicated. The way I interpret the FASTG specification, each declaration of an edge sequence implicitly also declares this edge sequence's reverse complement; however, this is not the case for "adjacencies" between edge sequences.
In any case, the "dialect" of FASTG files produced by SPAdes and MEGAHIT lists edge sequences and their reverse complements (as well as adjacencies between edge sequences and their reverse complements) separately. Because of this, we consider FASTG to be an "explicit" filetype. (See pyfastg's documentation for details on how we handle reverse complements in FASTG files.)
FAQ 2. Why does my graph have node X and -X in the same component?
One common reason this happens is the presence of palindromic sequences: these can cause both a sequence and its reverse-complement to be connected to each other.
This often occurs with the big ("hairball") component in an assembly graph.
FAQ 3. What happens if an edge is its own reverse complement?
(This assumes that you have read FAQ 1.)
This can happen if an edge exists from X -> -X or from -X -> X in an
"implicit" graph file (GFA / LastGraph). Consider
this GFA file:
H VN:Z:1.0
S 1 AAA
S 2 ACG
S 3 CAT
S 4 TTT
L 1 + 1 + 2M
L 2 + 2 - 2M
L 3 - 3 + 2M
L 4 - 4 - 2M
Since this GFA file contains four "link" lines, we might think at first that the corresponding graph
contains 4 × 2 = 8 edges. However, the graph only contains 6 unique
edges. This is because the reverse complement of 2 -> -2 is itself:
we know from above that X -> Y implies -Y -> -X, but
-(-2) -> -(2) is equal to 2 -> -2! The same goes for -3 -> 3:
-(3) -> -(-3) is equal to -3 -> 3.
Both of these edges "imply" themselves as their own reverse complements!
How do we handle this situation? As of writing, when MetagenomeScope visualizes these graphs it will only draw one copy of these "self-implying" edges. This matches the original visualization of this graph, and also matches Bandage's visualization of this GFA file.
Notably, since we assume that "explicit" graph files (FASTG / DOT / GML)
explicitly define all of the nodes and edges in their graph, MetagenomeScope doesn't do anything
special for this case for these files. (If your DOT file describes one edge
from X -> -X, then that's fine; if it describes two or more edges from X -> -X,
then that's also fine, and we'll visualize all of them.)
FAQ 4. What do you mean by a component's "size rank"?
Given a graph with N connected components: we sort these components by the number of nodes they contain, from high to low. We then assign each of these components a size rank, a number from 1 to N: the component with size rank #1 corresponds to the largest component, and the component with size rank #N corresponds to the smallest component.
Often, we only care about looking at individual components in a graph -- laying out and drawing the entire graph is not always a good idea when the graph is massive. Component size ranks are a nice way of formalizing this.
Some details about component size ranks, if you are interested:
-
The numbers shown in the treemap (accessible in the "Graph info" dialog) correspond exactly to component size ranks. So, the rectangle labelled #1 in the treemap corresponds to the largest component, the rectangle labelled #2 corresponds to the second-largest component, etc.
-
The exact component sorting functionality accounts for ties by using four different sorting criteria, in the following order. Ties at one level cause later levels to be considered for breaking ties.
- the number of "full" nodes in the component (treating a pair of split nodes 40-L → 40-R as a single node)
- the number of "total" nodes in the component (treating a pair of split nodes 40-L → 40-R as two nodes)
- the number of "total" edges in the component (including both real edges and "fake" edges between pairs of split nodes like 40-L → 40-R)
- the number of patterns in the component
FAQ 5. Can my graphs have parallel edges?
Yes! MetagenomeScope supports
multigraphs. If your assembly graph
file describes more than one edge from X -> Y, then MetagenomeScope will
visualize all of these "parallel" edges. (This is mostly useful when visualizing
de Bruijn graphs.)
Notably, parallel edges not supported right now for FASTG files. I don't think I've ever seen any FASTG files that have parallel edges, so I don't think this is a big priority, but please let me know if you would like us to add support for it.
FAQ 6. What filetype should I use for de Bruijn graphs?
If you are visualizing output from LJA or Flye, you may want to use a DOT file instead of a GFA / FASTG file as input.
This is because GFA and FASTG are not ideal for representing graphs in which sequences are stored on edges rather than nodes (i.e. de Bruijn / repeat graphs). The DOT files output by Flye and LJA should contain the original structure of these graphs (in which edges and nodes in the visualization actually correspond to edges and nodes in the original graph, respectively); the GFA / FASTG files usually represent altered versions in which nodes and edges have been swapped, which is not always an ideal representation (especially if you are doing something where you really care about the structure of the original graph).
That being said, please note that -- if you are using an assembler that outputs graphs in different filetypes -- these files may have additional differences beyond the usual filetype differences. For example, Flye's GFA and DOT files can have slightly different coverages, since Flye produces them at different times in its pipeline.
FAQ 7. I got an error saying Custom record types are not supported in GFA1?
Some assemblers include additional kinds of lines in their output GFA files. For example,
hifiasm and hifiasm-meta include A-lines describing alignments.
These "custom" lines can cause problems when parsing these graphs, because they may not be technically allowed in certain GFA versions.
The simplest way around this is just deleting or commenting out these custom lines. Here is an examle of commenting out the A-lines in hifiasm GFA files using sed:
sed -i -e 's/^A/#A/' hifiasm-out.p_ctg.gfaEventually I'd like to implement a better solution for this...
FAQ 8. How can I run the pattern decomposition process programmatically?
Creating a metagenomescope.graph.AssemblyGraph object will automatically run the decomposition process:
>>> from metagenomescope.graph import AssemblyGraph
>>> ag = AssemblyGraph("graph.gfa") # replace with your graph's filepathAt this point:
-
The "decomposed graph" (where patterns are collapsed into nodes) is represented by
ag.decomposed_graph(a NetworkXMultiDiGraph). -
The "true graph" (i.e. with all patterns fully uncollapsed, revealing all "original" nodes and edges) is represented by
ag.graph(also a NetworkXMultiDiGraph)- Note that this graph will still include split nodes and fake edges, if any remain after the decomposition process.
-
All nodes, edges, and patterns will have unique integer IDs. These IDs can be used to look up information about nodes, edges, and patterns in the
ag.nodeid2obj,ag.edgeid2obj, andag.pattid2objdictionaries, respectively.
Some examples of analyzing the decomposition results:
>>> from metagenomescope.graph import AssemblyGraph
>>> ag = AssemblyGraph("metagenomescope/tests/input/E_coli_LastGraph")
>>> # Inspect nodes, edges, and patterns
>>> ag.nodeid2obj
{0: Node 0 (name: 1),
1: Node 1 (name: -1),
2: Node 2 (name: 2),
...}
>>> ag.edgeid2obj
{558: Edge 558 (orig: 0 -> 244; new: 0 -> 244; dec: 0 -> 1421),
559: Edge 559 (orig: 1 -> 342; new: 1 -> 342; dec: 1527 -> 342),
560: Edge 560 (orig: 2 -> 477; new: 2 -> 477; dec: 2 -> 477),
...}
>>> ag.pattid2obj
{1222: bubble1222 containing nodes [33, 283, 395, 39] from [33] to [39],
1227: bubble1227 containing nodes [34, 76, 382, 303] from [34] to [76],
1232: bubble1232 containing nodes [40, 43, 35, 501] from [35] to [43],
...}
>>> # Go through just the bubble patterns
>>> ag.bubbles
[bubble1222 containing nodes [33, 283, 395, 39] from [33] to [39],
bubble1227 containing nodes [34, 76, 382, 303] from [34] to [76],
bubble1232 containing nodes [40, 43, 35, 501] from [35] to [43],
...]
>>> # Look up a node by name (if a node was split, this will list both halves)
>>> ag.nodename2objs
defaultdict(<class 'list'>,
{'1': [Node 0 (name: 1)],
'-1': [Node 1 (name: -1)],
'2': [Node 2 (name: 2)],
...
'40-R': [Node 78 (name: 40-R)],
'40': [Node 78 (name: 40-R), Node 1259 (name: 40-L)],
'40-L': [Node 1259 (name: 40-L)],
...})
>>> # Examine split nodes
>>> for n in ag.nodeid2obj.values():
... if n.split is not None:
... print(n)
Node 32 (name: 17-L)
Node 33 (name: -17-R)
Node 34 (name: 18-R)
...
>>> # Distinguish fake from real edges
>>> for e in ag.edgeid2obj.values():
... print(e, e.is_fake)
Edge 558 (orig: 0 -> 244; new: 0 -> 244; dec: 0 -> 1421) False
Edge 559 (orig: 1 -> 342; new: 1 -> 342; dec: 1527 -> 342) False
...
Edge*1634 (orig: 348 -> 1633; new: 348 -> 1633; dec: 1628 -> 1666) True
Edge*1639 (orig: 1638 -> 451; new: 1638 -> 451; dec: 1671 -> 1635) TrueThis interface should remain relatively stable, although I may change things slightly as development continues. If you have any questions, please reach out.
FAQ 9. What's the biggest possible graph I can visualize?
We're still figuring that out. There are two main bottlenecks I am aware of:
-
Laying out the graph.
-
We usually only lay out one component at a time, so generally the problem comes with laying out the large "hairball" component(s) of the graph, if any.
-
When you get to the order of, say, thousands of nodes, laying out a component will probably become somewhat slow (especially if you select the
Lay out patterns recursivelyoption in the draw options dialog). -
To my understanding, a big factor here is the ratio of nodes to edges: when there are many more edges than nodes in a component (indicating a very densely connected structure), Graphviz has to do a lot of work to position things properly.
-
-
Drawing the graph's elements.
-
Cytoscape.js has a lot of optimizations built-in, but I think there are some inherent limitations of drawing using a HTML canvas.
-
With graphs containing thousands of nodes, interaction (e.g. zooming, panning) starts to feel a bit sluggish.
-
See the "Large graph" section under "Example datasets" above for some tips for working with large graphs.
-
Edge flattening: Cytoscape.js (the library we use to visualize graphs in the browser) can sometimes determine that the control points used to draw certain complex edges are invalid. Mainly, this can happen as you adjust the visualization after drawing (e.g. by selecting nodes or moving them around).
To prevent these "invalid" edges from being hidden, MetagenomeScope will detect them and "flatten" them into simple Bezier edges (usually straight lines). This way, we can at least draw something for each edge in the graph.
See CONTRIBUTING.md.
MetagenomeScope is licensed under the GNU GPL, version 3.
MetagenomeScope's code is distributed with
Bootstrap,
Bootstrap Icons,
Cytoscape.js,
cytoscape-dagre,
cytoscape-fcose,
and
cytoscape-svg.
Please see the metagenomescope/assets/vendor/licenses/ directory for copies of these tools' licenses.
Thanks to various people in the Pop, Knight, and Pevzner Labs over the years for their kind feedback and helpful suggestions.
Thanks also to the developers of the many excellent open-source software packages used by MetagenomeScope. In particular, Graphviz (graph layout), Cytoscape.js (interactive graph drawing), and Dash (application framework) have been extremely helpful tools throughout the development of this project.
Please open a GitHub issue if you have any questions or suggestions.


