Authors: Luis U. Aguilera, William S. Raymond, Rhiannon M. Sears, Nathan L. Nowling, Brian Munsky, Ning Zhao
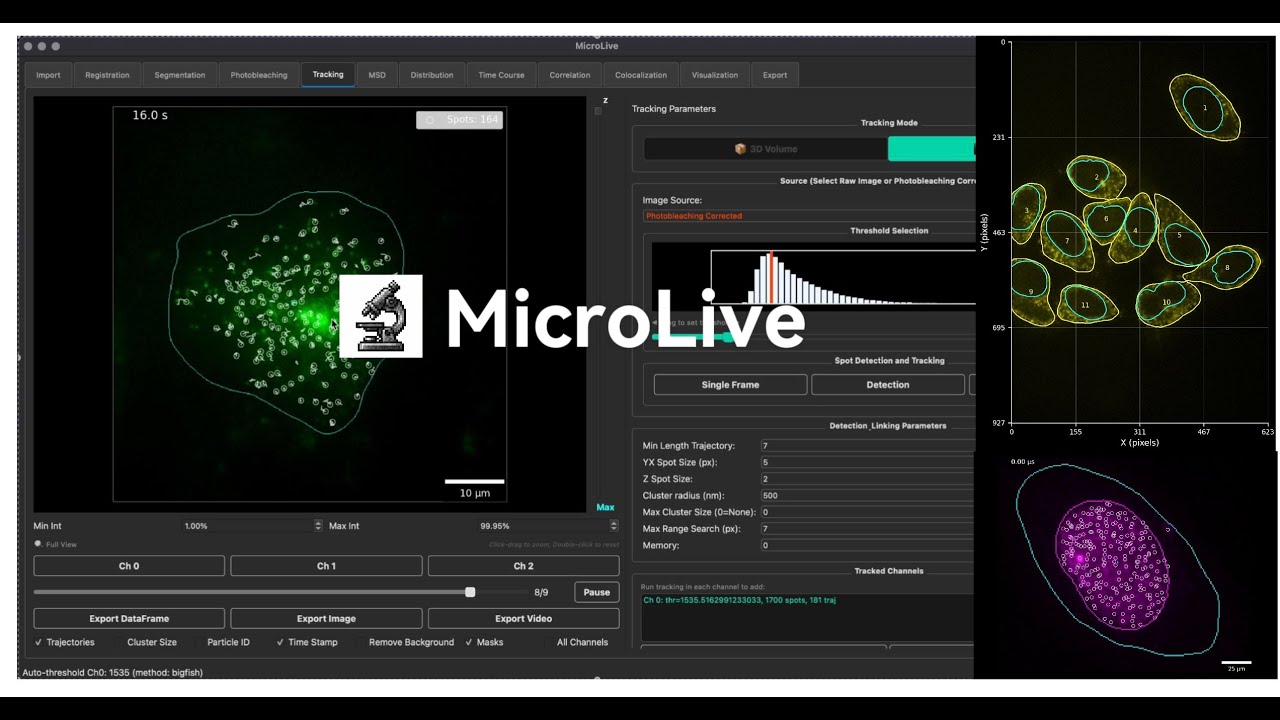
MicroLive is a Python-based GUI application for live-cell microscopy image analysis and single-molecule measurements. It provides an end-to-end workflow from image loading through particle tracking, colocalization analysis, and statistical analysis.
Click the image above to watch the demo video on YouTube
- Image I/O: Load .lif, .tif, .ome.tif with metadata extraction and dimension mapping
- Registration: Drift correction via phase correlation
- Segmentation: Cellpose (GPU), watershed, manual ROI, or external mask import
- Photobleaching correction: Exponential decay modeling
- Particle tracking: 2D (TrackPy) and 3D (Big-FISH) detection with multi-channel support
- Automated threshold detection: Hybrid Big-FISH/TrueSpot method with fixed-threshold mode for inhibitor experiments
- Trajectory linking: Nearest-neighbor with memory and cluster analysis
- Intensity quantification: Background subtraction, PSF fitting, SNR calculation
- Colocalization: CNN-based, distance-based, and manual verification
- MSD analysis: Per-cell diffusion coefficient calculation
- Correlation: Auto- and cross-correlation with exponential/linear fitting
- Export: PNG, TIFF, CSV, MP4/GIF with full metadata logging
- User Guide — Complete guide to using MicroLive
- Tutorial — Step-by-step tutorials for all workflows
- API Reference — Technical documentation for developers
# Create and activate conda environment
conda create -n microlive python=3.10 -y
conda activate microlive
# Install MicroLive
pip install microlive
# Launch the GUI
microliveThat's it! MicroLive will launch with GPU acceleration automatically enabled:
- macOS (Apple Silicon): MPS GPU acceleration works automatically
- Linux/Windows (CPU): Works out of the box
For NVIDIA GPU acceleration, install PyTorch with CUDA support before installing MicroLive:
# Create environment
conda create -n microlive python=3.10 -y
conda activate microlive
# Install PyTorch with CUDA 12.4 (adjust version as needed)
pip install torch torchvision --index-url https://download.pytorch.org/whl/cu124
# Then install MicroLive
pip install microlive
# Launch
microlivePre-configured environment files are available:
# macOS / CPU
conda env create -f installation/microlive.yml
conda activate microlive
# Windows/Linux with NVIDIA GPU
conda env create -f installation/microlive_cuda.yml
conda activate microlivefrom microlive.utils.device import check_gpu_status
check_gpu_status()Expected output:
PyTorch version: 2.x.x
✅ CUDA available: NVIDIA GeForce RTX ...
Memory: 8.0 GB
Or for Apple Silicon:
PyTorch version: 2.x.x
✅ MPS available: Apple Silicon GPU (MPS)
For developers who want to modify the source code:
# Clone the repository
git clone --depth 1 https://github.com/ningzhaoAnschutz/microlive.git
cd microlive
# Create environment
conda create -n microlive python=3.10 -y
conda activate microlive
# Install in editable mode
pip install -e .
# Launch
microliveIf you encounter errors during installation (especially on Python 3.12):
# Option 1: Upgrade pip and pre-install numpy
pip install --upgrade pip setuptools wheel
pip install numpy
pip install microlive
# Option 2: Use Python 3.10 (most compatible)
conda create -n microlive python=3.10 -y
conda activate microlive
pip install microliveCommon error: ModuleNotFoundError: No module named 'numpy' during pystackreg build
- This occurs when pip tries to build older package versions from source
- Solution: Pre-install NumPy before installing MicroLive
conda activate microlive
microliveimport microlive.microscopy as mi
# Load images from a Leica .lif file
reader = mi.ReadLif("experiment.lif")
images = reader.get_images()
# Access image data
image = reader.list_images[0]
print(f"Image shape: {image.shape}") # (T, Z, Y, X, C)
# Run Cellpose segmentation
cellpose = mi.Cellpose(image_for_segmentation=image[0, 0, :, :, 0])
masks = cellpose.calculate_masks()
# Spot detection
spots = mi.SpotDetection(
image=image,
spot_size_yx=5,
threshold=100
)
detected = spots.detect_spots()microlive/
├── microlive/ # Core package (pip installable)
│ ├── microscopy.py # Main analysis classes
│ ├── imports.py # Central import management
│ ├── ml_spot_detection.py # ML-based spot detection
│ ├── gui/ # GUI application
│ │ ├── app.py # Main GUI window
│ │ └── main.py # CLI entry point
│ ├── utils/ # Utility modules
│ │ ├── device.py # GPU detection
│ │ └── resources.py # Resource paths
│ ├── data/ # ML models and resources
│ └── pipelines/ # Analysis pipeline modules
├── simulations/ # Spot simulation & validation (not in pip)
│ ├── spot_simulator.py # Multi-cell simulation engine
│ ├── run_simulation.py # CLI runner
│ ├── visualize_results.py # Visualization tools
│ ├── config_*.yaml # Configuration examples
│ └── tests/ # 8-test validation suite (100% pass)
├── docs/ # Documentation
│ ├── user_guide.md # User manual
│ ├── tutorial.md # Step-by-step tutorials
│ └── api_reference.md # API documentation
├── modeling/ # Research/development (not in pip package)
├── notebooks/ # Example Jupyter notebooks
├── installation/ # Environment files
│ ├── microlive.yml # Conda env (macOS / CPU)
│ └── microlive_cuda.yml # Conda env (NVIDIA GPU)
├── tests/ # Test suite
├── pyproject.toml # Package configuration
└── LICENSE # GPL v3 License
This project is licensed under the GNU General Public License v3 (GPLv3). See LICENSE for details.
If you use MicroLive in your research, please cite:
Aguilera LU, Raymond WS, Sears RM, Nowling NL, Munsky B, Zhao N. MicroLive: An Image Processing Toolkit for Quantifying Live-cell Single-Molecule Microscopy. bioRxiv, 2025.
DOI: 10.1101/2025.09.25.678587
@article{aguilera2025microlive,
title={MicroLive: An Image Processing Toolkit for Quantifying Live-cell Single-Molecule Microscopy},
author={Aguilera, Luis U and Raymond, William S and Sears, Rhiannon M and Nowling, Nathan L and Munsky, Brian and Zhao, Ning},
journal={bioRxiv},
pages={2025--09},
year={2025},
publisher={Cold Spring Harbor Laboratory},
doi={10.1101/2025.09.25.678587}
}- Documentation: User Guide | Tutorial | API Reference
- Issues & Contributions: GitHub