Requirements:
- Prepare ligand and receptors in pdb format.
- Add hydrogens and charges.
- Correct ligand coordinations.
- SwissDock server: http://www.swissdock.ch/docking
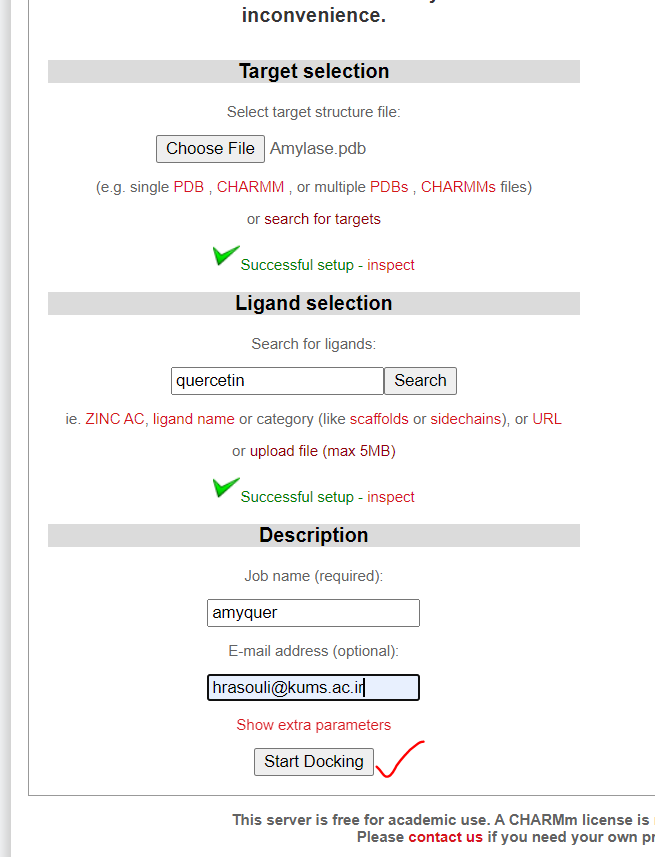
- Upload your receptor or fetch it by pdb id.
- Upload ligand or fetch it by name.
- Go to SwissDock homepage:
Note: In this tutorial, we will dock human alpha amylase and quercetin ligand.
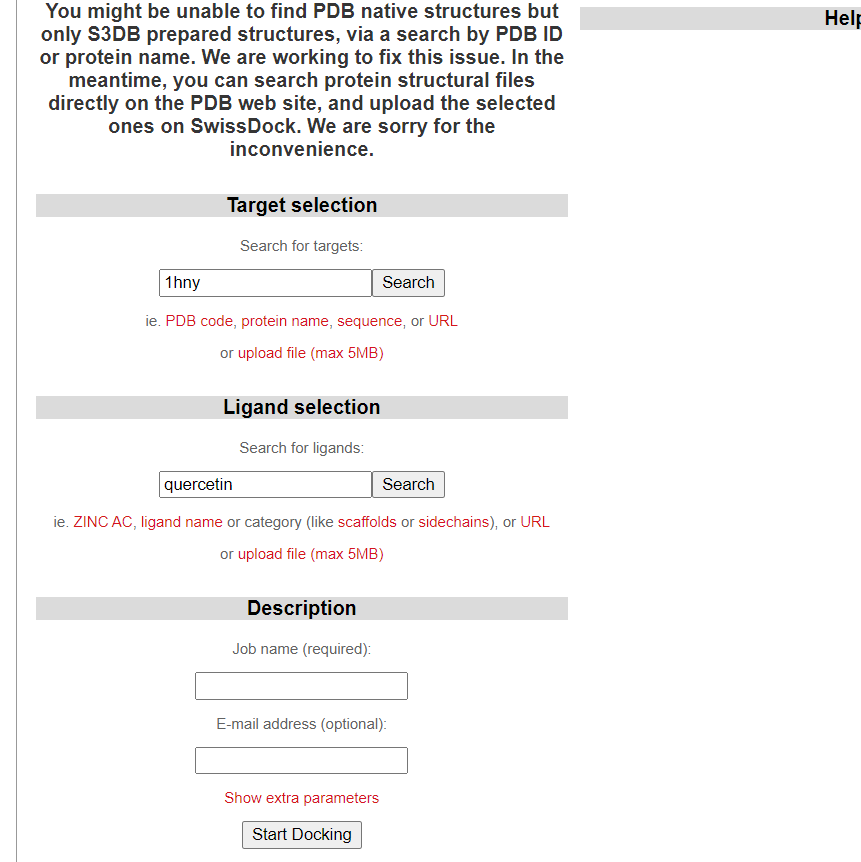
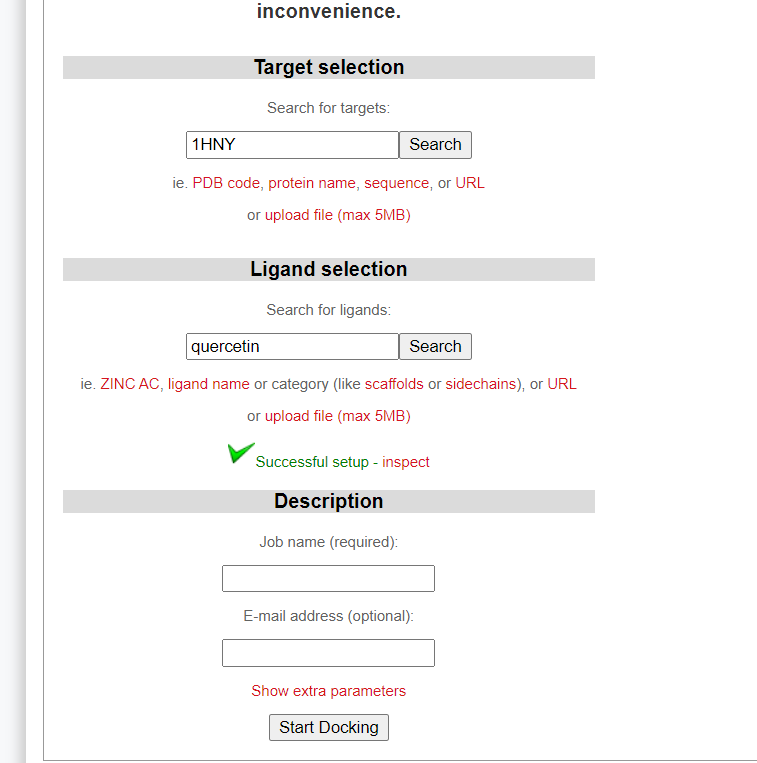
- Enter receptor pdb Id in target box and then enter ligand name and search it in SwissDock.
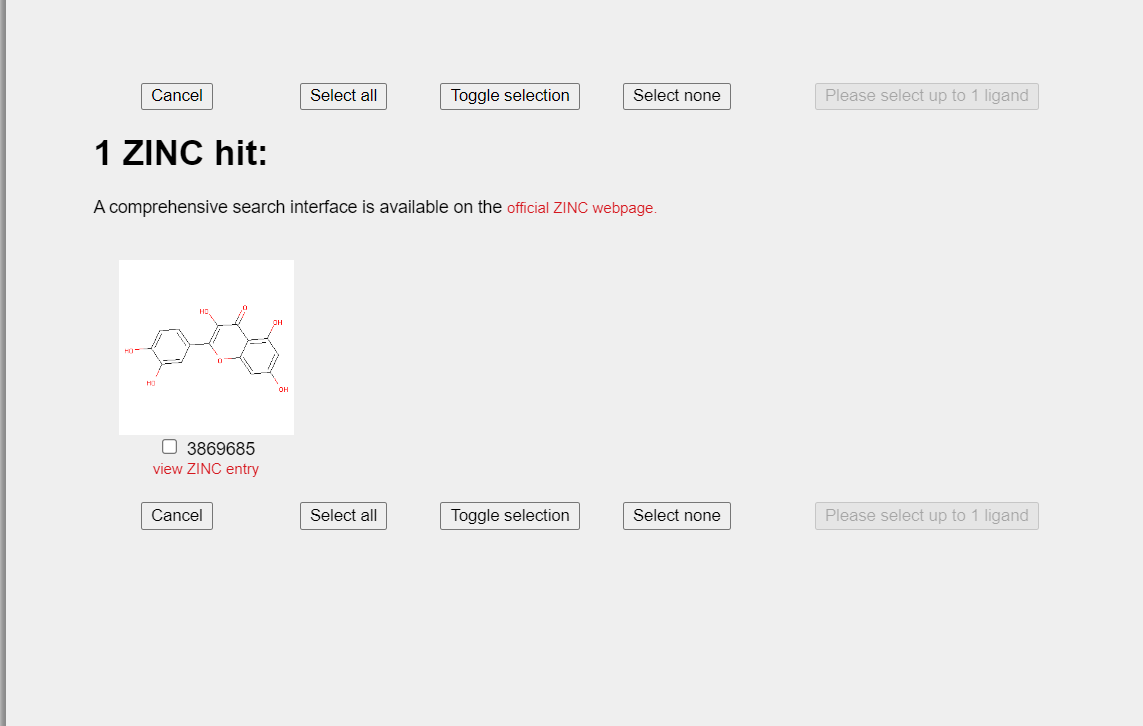
- Select the target ligand and wait to fetch its structure.
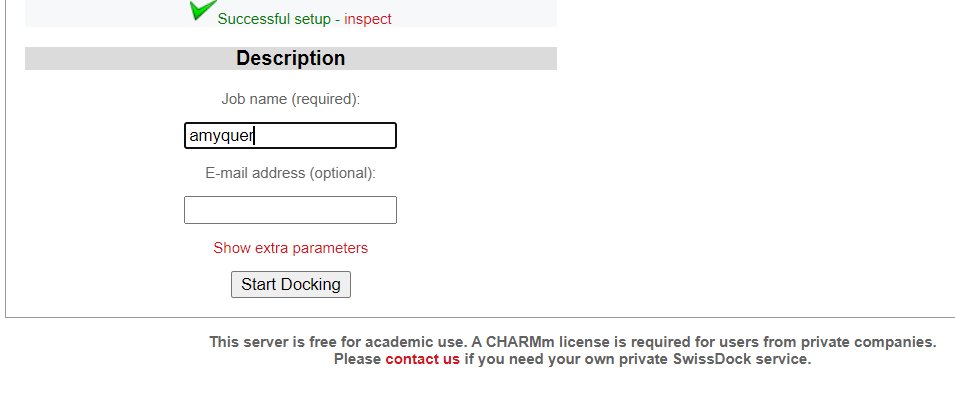
- Assign a job name for your docking and add your email address.
- Click on Start Docking bottom to set off your first docking project.
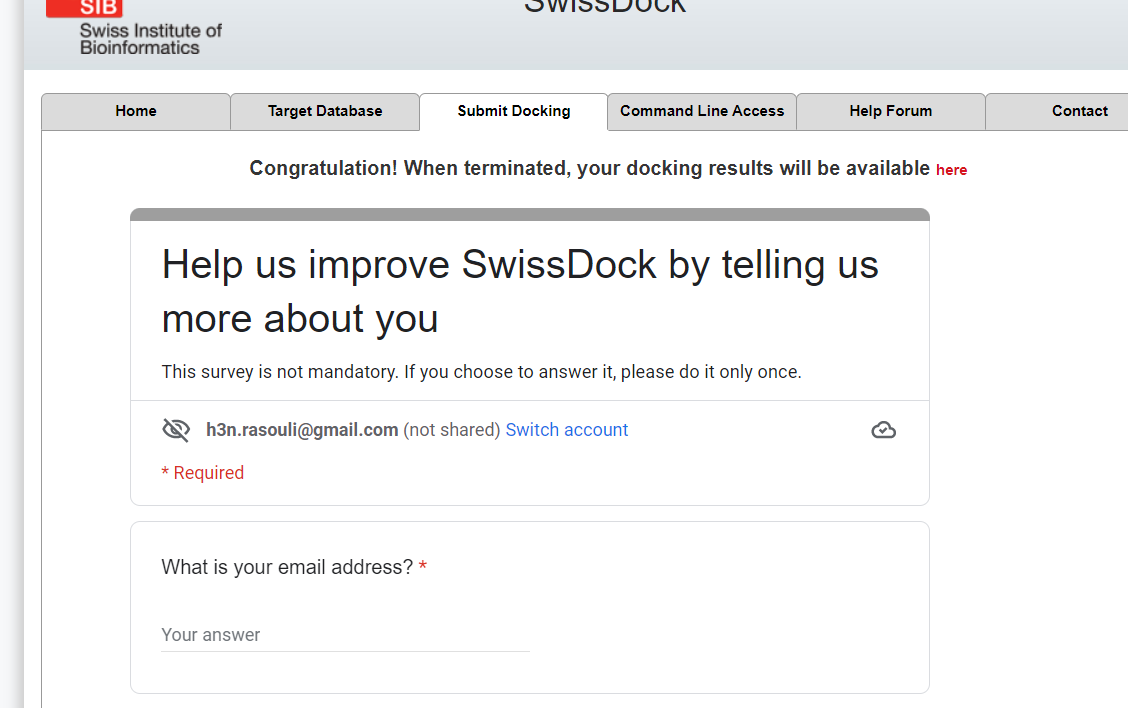
- Wait for results.
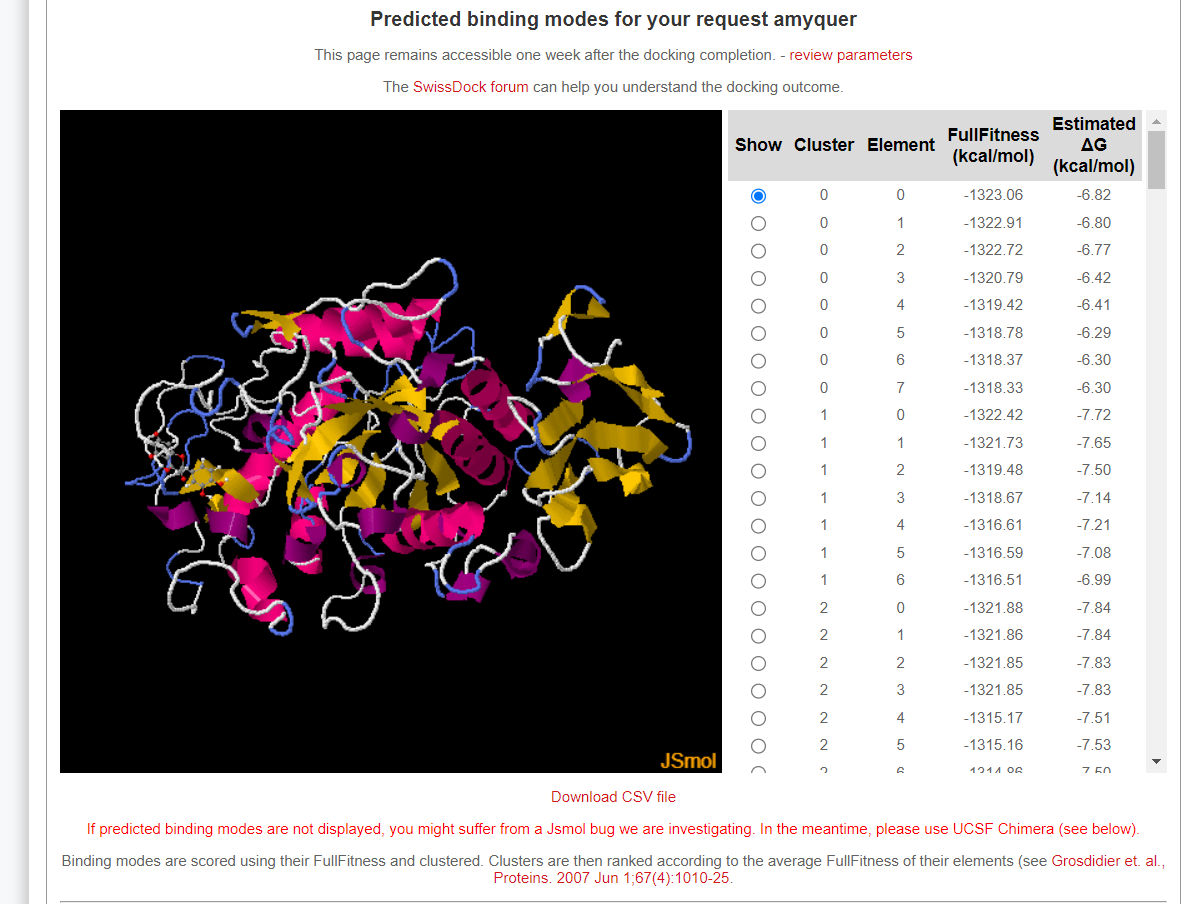
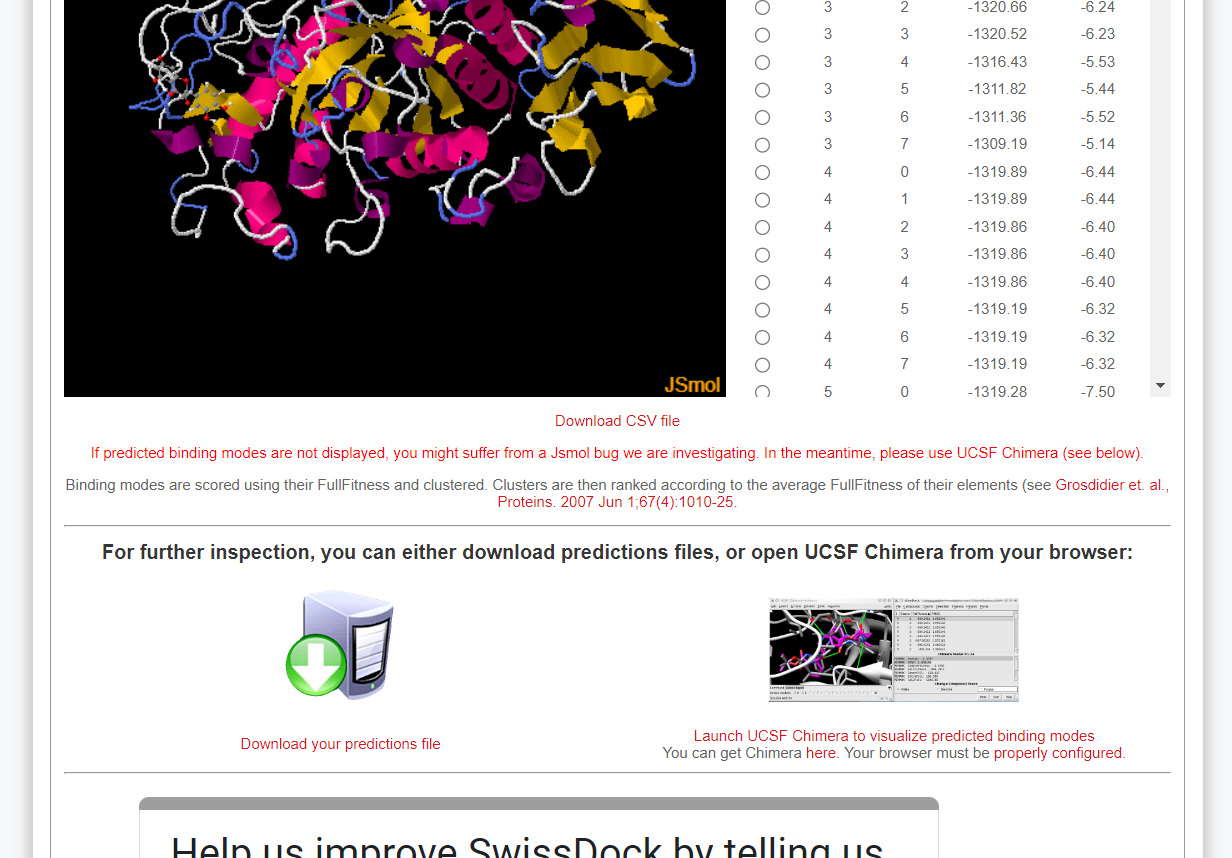
- After finishing your docking job, you should download the outputs and moniotr the produced poses to find the best interaction profile.
Make sure that the top docking pose should be selected for post-docking analyses.