MIT License Addendum:
(c) To all the cancer patients, their courage and love that fuel this work and countless others fighting to defeat cancer.
I work on machine learning (ML) and it's applications in computational biology and bioengineering, with a singular focus: adapting emerging AI advances to precision medicine in cancer and nuero-degeneration. My formal background is in applied math and theoretical machine learning, but my mission is in engaging cancer.
I seek problems that focus on precise translational inflection points, where fundamental biological insights and data can be computationally harnessed to engineer game changing breakthroughs. This is where I look for leaps toward a cure, at the intersection of scientific discovery and actionable intervention for patients.
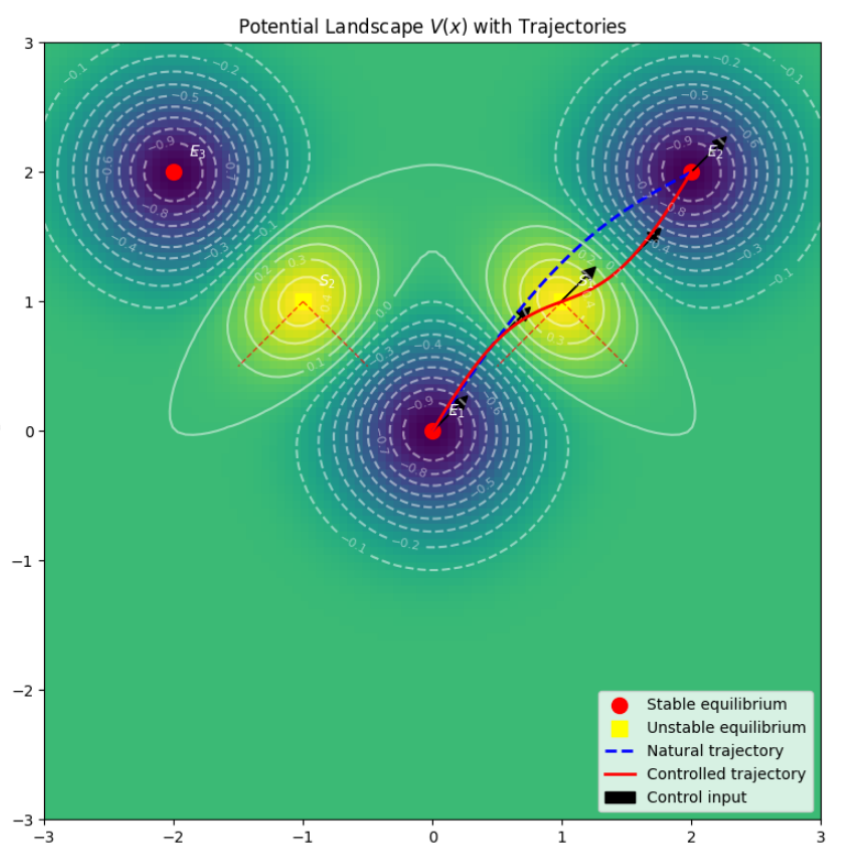
I see the immune system as the key and most promising frontier in medicine and battle against cancer. Each immune cell is a programmable molecular dynamical system, that working collectively give rise to the emergent functional complexity and success of our immune system. I develop predictive and generative models at genomic, cellular and system levels, aiming to decipher the genomic programming language that governs this system. Growing evidence suggests that the key to solving cancer lies within the regulatory programs of our cells. Can we re-program our cells or steer their states to modulate the immune system and/or the tumor microenvironments to stop solid tumor progressions? These are the questions that drive my work.
On a more theoretical front, a current priority is uncovering the mathematical underpinnings of the Transformer models needed for their adaptation to genomic data. I am working toward modiying the Transformer formulations for building better predictive ML models in biology.
Cancer requires interdisciplinary team work. To that end, I work on creating accelerated pathways for mathematicians and engineers to transition into computational biology, while helping biologists gain formal computational fluencies, including strategic and guided proliferation of AI-assisted programming.
If you're working directly or indirectly on cancer research, feel free to reach out. I reserve time to volunteer and support cancer researchers.
Below are a sample of tools and programs to guide you into the brand of computational biology problems I'm working on. Completed tools on GitHub are free to use under academic licensing, in any way you can, to make progress in your work. May the force be with you!
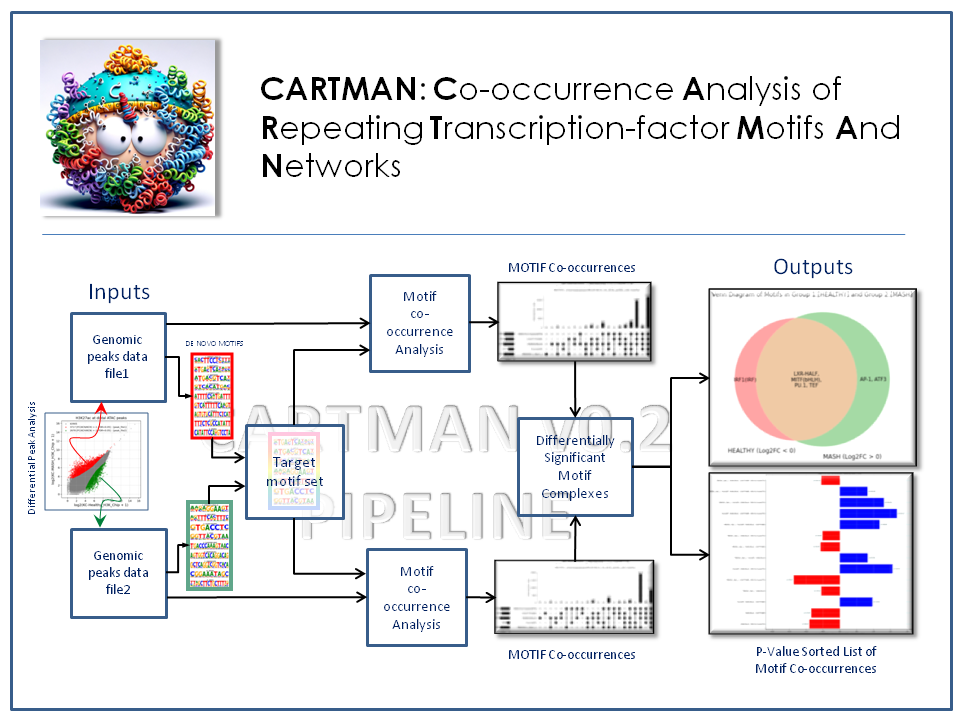 |
🔥 CARTMAN
Co-occurrence Analysis of Repeating Transcription-factor Motifs and Networks. A computational tool for motif discovery and transcription factor co-occurrence analysis in regulatory genomics. 🔗 GitHub Repository |
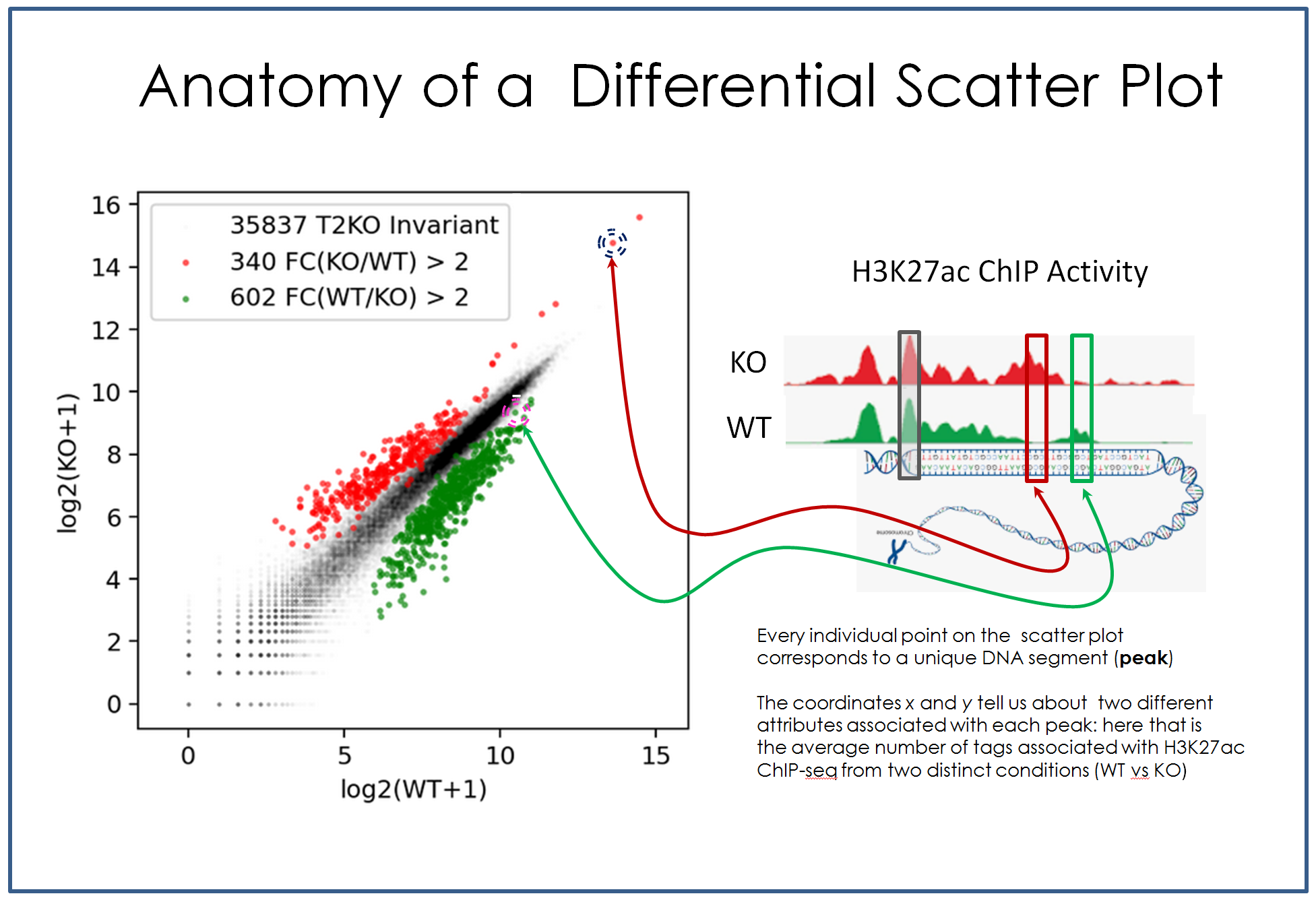 |
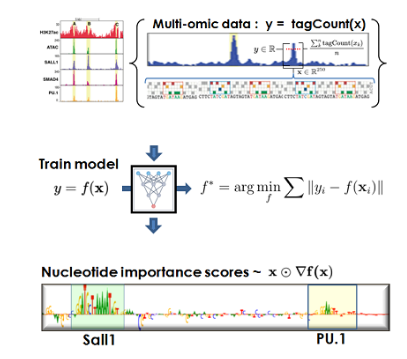
🔥 PEAKDIFF
Differential peak analysis for identifying distinct genomic regions between experimental conditions. Includes integration with *ChIP-seq, ATAC-seq*, and *HOMER-based enhancer prediction*. 🔗 GitHub Repository |
 |
🔥 PIPSCOUT
PIPseeker-based Single-Cell Output & UMAP Typing A tool for deciphering PIPseeker's single-cell outputs for downstream analytical pipelines. 🔗 GitHub Repository |
 |
🔥 SPATIOME
Synthetic Platform for Advanced Transcriptomics Integrating Omics and Multidimensional Exploration. Computational framework for integrating high-dimensional transcriptomics data. GitHub Repository |

|
|

|
|

|
| Kaggle is a Google Subsidiary |