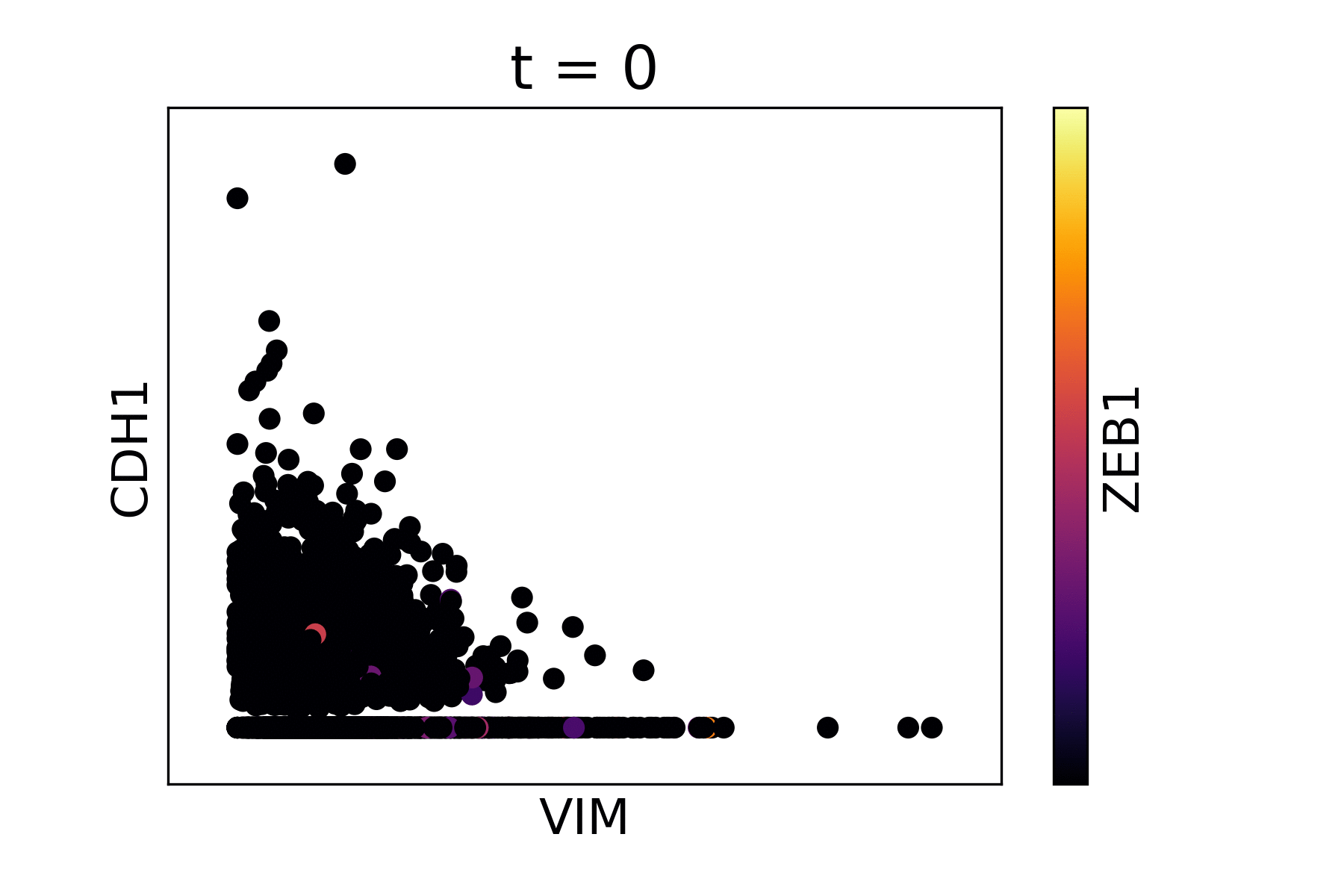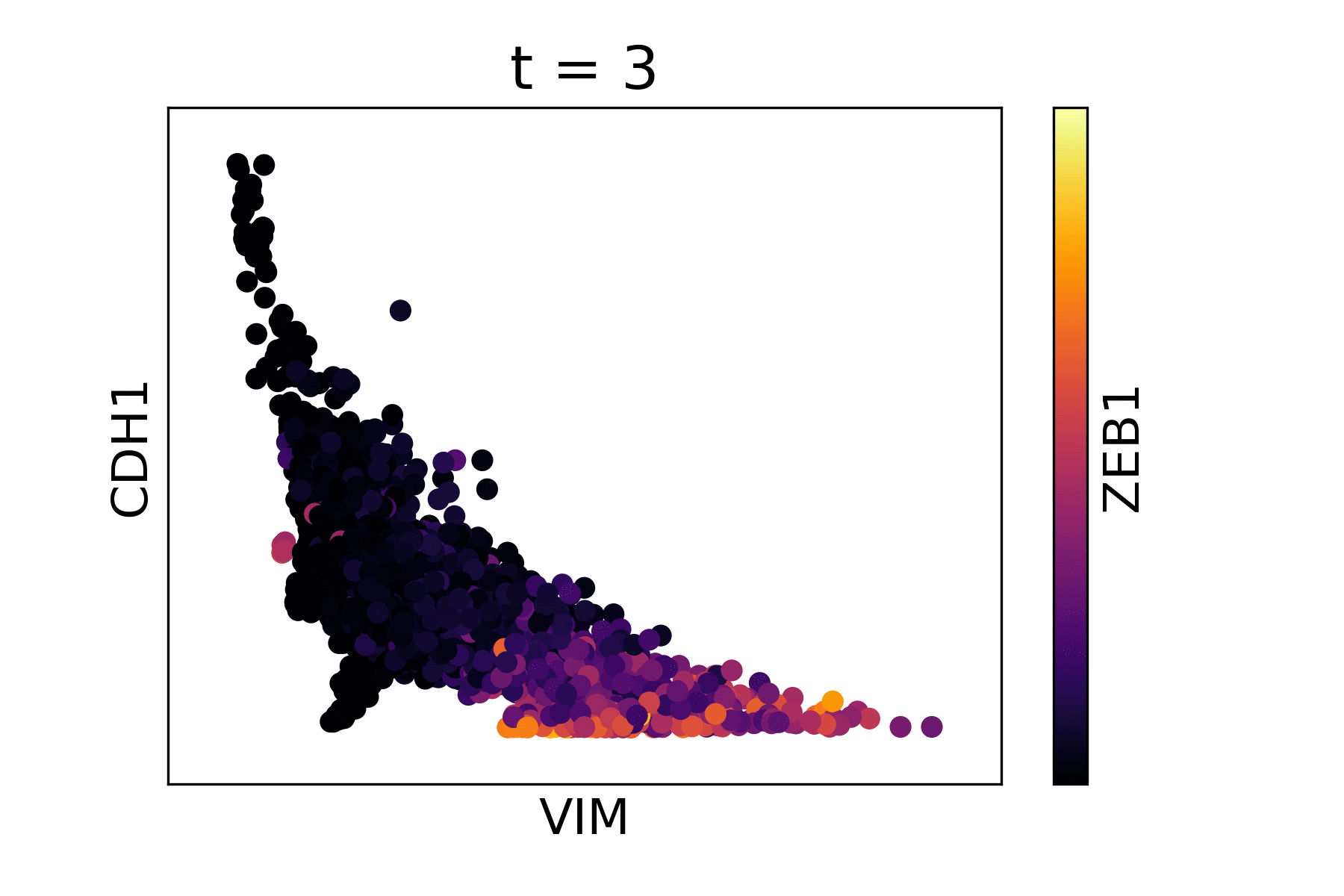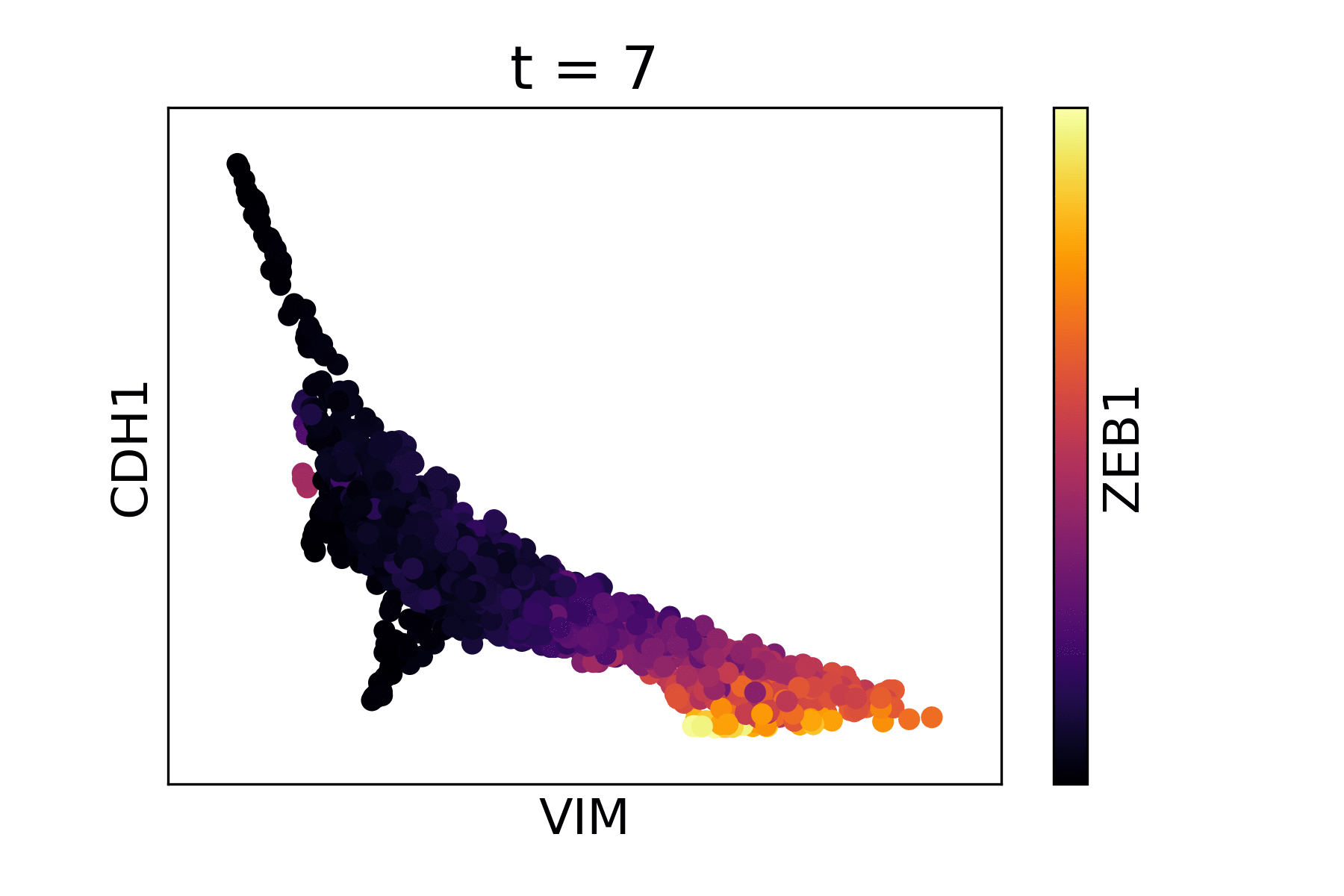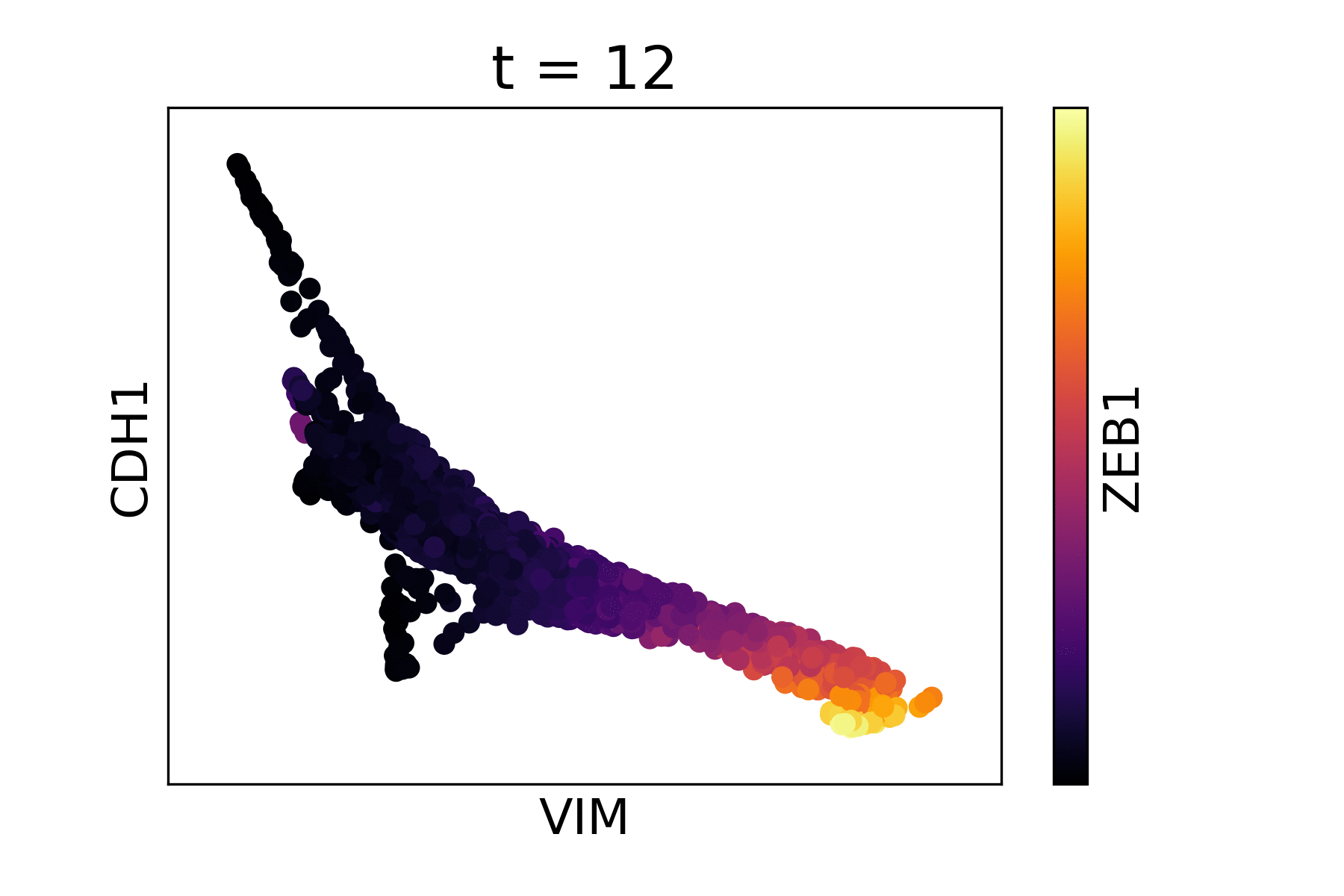
Markov Affinity-based Graph Imputation of Cells (MAGIC) is an algorithm for denoising high-dimensional data most commonly applied to single-cell RNA sequencing data. MAGIC learns the manifold data, using the resultant graph to smooth the features and restore the structure of the data.
To see how MAGIC can be applied to single-cell RNA-seq, elucidating the epithelial-to-mesenchymal transition, read our publication in Cell.
For R and MATLAB implementations of MAGIC, see https://github.com/KrishnaswamyLab/MAGIC.
Magic reveals the interaction between Vimentin (VIM), Cadherin-1 (CDH1), and Zinc finger E-box-binding homeobox 1 (ZEB1, encoded by colors).
To install with pip, run the following from a terminal:
pip install --user magic-impute
To clone the repository and install manually, run the following from a terminal:
git clone git://github.com/KrishnaswamyLab/MAGIC.git cd MAGIC/python python setup.py install --user
The following code runs MAGIC on test data located in the MAGIC repository:
import magic
import pandas as pd
import matplotlib.pyplot as plt
X = pd.read_csv("MAGIC/data/test_data.csv")
magic_operator = magic.MAGIC()
X_magic = magic_operator.fit_transform(X, genes=['VIM', 'CDH1', 'ZEB1'])
plt.scatter(X_magic['VIM'], X_magic['CDH1'], c=X_magic['ZEB1'], s=1, cmap='inferno')
plt.show()
magic.plot.animate_magic(X, gene_x='VIM', gene_y='CDH1', gene_color='ZEB1', operator=magic_operator)
We have included two tutorial notebooks on MAGIC usage and results visualization for single cell RNA-seq data.
EMT data notebook: http://nbviewer.jupyter.org/github/KrishnaswamyLab/magic/blob/master/python/tutorial_notebooks/emt_tutorial.ipynb
Bone Marrow data notebook: http://nbviewer.jupyter.org/github/KrishnaswamyLab/magic/blob/master/python/tutorial_notebooks/bonemarrow_tutorial.ipynb
If you have any questions or require assistance using MAGIC, please contact us at https://krishnaswamylab.org/get-help.