diff --git a/coreforecast/.nojekyll b/coreforecast/.nojekyll
new file mode 100644
index 00000000..e69de29b
diff --git a/coreforecast/dark.png b/coreforecast/dark.png
new file mode 100644
index 00000000..4142a0bb
Binary files /dev/null and b/coreforecast/dark.png differ
diff --git a/coreforecast/differences.mdx b/coreforecast/differences.mdx
new file mode 100644
index 00000000..ea03e909
--- /dev/null
+++ b/coreforecast/differences.mdx
@@ -0,0 +1,94 @@
+
+
+
+
+# module `coreforecast.differences`
+
+
+
+
+
+---
+
+
+
+## function `num_diffs`
+
+```python
+num_diffs(x: ndarray, max_d: int = 1) → int
+```
+
+Find the optimal number of differences
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Array with the time series.
+ - `max_d` (int, optional): Maximum number of differences to consider. Defaults to 1.
+
+
+
+**Returns:**
+
+ - `int`: Optimal number of differences.
+
+
+---
+
+
+
+## function `num_seas_diffs`
+
+```python
+num_seas_diffs(x: ndarray, season_length: int, max_d: int = 1) → int
+```
+
+Find the optimal number of seasonal differences
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Array with the time series.
+ - `season_length` (int): Length of the seasonal pattern.
+ - `max_d` (int, optional): Maximum number of differences to consider. Defaults to 1.
+
+
+
+**Returns:**
+
+ - `int`: Optimal number of seasonal differences.
+
+
+---
+
+
+
+## function `diff`
+
+```python
+diff(x: ndarray, d: int) → ndarray
+```
+
+Subtract previous values of the series
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Array with the time series.
+ - `d` (int): Lag to subtract
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Differenced time series.
+
+
+
+
+---
+
+_This file was automatically generated via [lazydocs](https://github.com/ml-tooling/lazydocs)._
diff --git a/coreforecast/expanding.mdx b/coreforecast/expanding.mdx
new file mode 100644
index 00000000..3213fc8a
--- /dev/null
+++ b/coreforecast/expanding.mdx
@@ -0,0 +1,141 @@
+
+
+
+
+# module `coreforecast.expanding`
+
+
+
+
+
+---
+
+
+
+## function `expanding_mean`
+
+```python
+expanding_mean(x: ndarray) → ndarray
+```
+
+Compute the expanding_mean of the input array.
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Input array.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the expanding statistic
+
+
+---
+
+
+
+## function `expanding_std`
+
+```python
+expanding_std(x: ndarray) → ndarray
+```
+
+Compute the expanding_std of the input array.
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Input array.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the expanding statistic
+
+
+---
+
+
+
+## function `expanding_min`
+
+```python
+expanding_min(x: ndarray) → ndarray
+```
+
+Compute the expanding_min of the input array.
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Input array.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the expanding statistic
+
+
+---
+
+
+
+## function `expanding_max`
+
+```python
+expanding_max(x: ndarray) → ndarray
+```
+
+Compute the expanding_max of the input array.
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Input array.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the expanding statistic
+
+
+---
+
+
+
+## function `expanding_quantile`
+
+```python
+expanding_quantile(x: ndarray, p: float) → ndarray
+```
+
+Compute the expanding_quantile of the input array.
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Input array.
+ - `p` (float): Quantile to compute.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the expanding statistic
+
+
+
+
+---
+
+_This file was automatically generated via [lazydocs](https://github.com/ml-tooling/lazydocs)._
diff --git a/coreforecast/exponentially_weighted.mdx b/coreforecast/exponentially_weighted.mdx
new file mode 100644
index 00000000..5b1916f4
--- /dev/null
+++ b/coreforecast/exponentially_weighted.mdx
@@ -0,0 +1,41 @@
+
+
+
+
+# module `coreforecast.exponentially_weighted`
+
+
+
+
+
+---
+
+
+
+## function `exponentially_weighted_mean`
+
+```python
+exponentially_weighted_mean(x: ndarray, alpha: float) → ndarray
+```
+
+Compute the exponentially weighted mean of the input array.
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Input array.
+ - `alpha` (float): Weight parameter.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the exponentially weighted mean.
+
+
+
+
+---
+
+_This file was automatically generated via [lazydocs](https://github.com/ml-tooling/lazydocs)._
diff --git a/coreforecast/favicon.svg b/coreforecast/favicon.svg
new file mode 100644
index 00000000..e5f33342
--- /dev/null
+++ b/coreforecast/favicon.svg
@@ -0,0 +1,5 @@
+
diff --git a/coreforecast/grouped_array.mdx b/coreforecast/grouped_array.mdx
new file mode 100644
index 00000000..2526b876
--- /dev/null
+++ b/coreforecast/grouped_array.mdx
@@ -0,0 +1,16 @@
+
+
+
+
+# module `coreforecast.grouped_array`
+
+
+
+
+
+
+
+
+---
+
+_This file was automatically generated via [lazydocs](https://github.com/ml-tooling/lazydocs)._
diff --git a/coreforecast/index.mdx b/coreforecast/index.mdx
new file mode 100644
index 00000000..b7a5532f
--- /dev/null
+++ b/coreforecast/index.mdx
@@ -0,0 +1,58 @@
+---
+description: Fast implementations of common forecasting routines
+title: "coreforecast"
+---
+## Motivation
+At Nixtla we have implemented several libraries to deal with time series data. We often have to apply some transformation over all of the series, which can prove time consuming even for simple operations like performing some kind of scaling.
+
+We've used [numba](https://numba.pydata.org/) to speed up our expensive computations, however that comes with other issues such as cold starts and more dependencies (LLVM). That's why we developed this library, which implements several operators in C++ to transform time series data (or other kind of data that can be thought of as independent groups), with the possibility to use multithreading to get the best performance possible.
+
+You probably won't need to use this library directly but rather use one of our higher level libraries like [mlforecast](https://nixtlaverse.nixtla.io/mlforecast/docs/how-to-guides/lag_transforms_guide.html#built-in-transformations-experimental), which will use this library under the hood. If you're interested on using this library directly (only depends on numpy) you should continue reading.
+
+## Installation
+
+### PyPI
+```python
+pip install coreforecast
+```
+
+### conda-forge
+```python
+conda install -c conda-forge coreforecast
+```
+
+## Minimal example
+The base data structure is the "grouped array" which holds two numpy 1d arrays:
+
+* **data**: values of the series.
+* **indptr**: series boundaries such that `data[indptr[i] : indptr[i + 1]]` returns the `i-th` series. For example, if you have two series of sizes 5 and 10 the indptr would be [0, 5, 15].
+
+```python
+import numpy as np
+from coreforecast.grouped_array import GroupedArray
+
+data = np.arange(10)
+indptr = np.array([0, 3, 10])
+ga = GroupedArray(data, indptr)
+```
+
+Once you have this structure you can run any of the provided transformations, for example:
+
+```python
+from coreforecast.lag_transforms import ExpandingMean
+from coreforecast.scalers import LocalStandardScaler
+
+exp_mean = ExpandingMean(lag=1).transform(ga)
+scaler = LocalStandardScaler().fit(ga)
+standardized = scaler.transform(ga)
+```
+
+## Single-array functions
+We've also implemented some functions that work on single arrays, you can refer to the following pages:
+
+* [differences](https://nixtlaverse.nixtla.io/coreforecast/differences)
+* [scalers](https://nixtlaverse.nixtla.io/coreforecast/scalers)
+* [seasonal](https://nixtlaverse.nixtla.io/coreforecast/seasonal)
+* [rolling](https://nixtlaverse.nixtla.io/coreforecast/rolling)
+* [expanding](https://nixtlaverse.nixtla.io/coreforecast/expanding)
+* [exponentially weighted](https://nixtlaverse.nixtla.io/coreforecast/exponentially_weighted)
diff --git a/coreforecast/lag_transforms.mdx b/coreforecast/lag_transforms.mdx
new file mode 100644
index 00000000..644fb252
--- /dev/null
+++ b/coreforecast/lag_transforms.mdx
@@ -0,0 +1,1522 @@
+
+
+
+
+# module `coreforecast.lag_transforms`
+
+
+
+
+**Global Variables**
+---------------
+- **TYPE_CHECKING**
+
+
+---
+
+
+
+## class `Lag`
+Simple lag operator
+
+
+
+**Args:**
+
+ - `lag` (int): Number of periods to offset
+
+
+
+### method `__init__`
+
+```python
+__init__(lag: int)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(transforms: Sequence[ForwardRef('_BaseLagTransform')]) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(_idxs: ndarray) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+
+---
+
+
+
+## class `RollingMean`
+Rolling Mean
+
+
+
+**Args:**
+
+ - `lag` (int): Number of periods to offset by before applying the transformation.
+ - `window_size` (int): Length of the rolling window.
+ - `min_samples` (int, optional): Minimum number of samples required to compute the statistic. If None, defaults to window_size.
+
+
+
+### method `__init__`
+
+```python
+__init__(lag: int, window_size: int, min_samples: Optional[int] = None)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(transforms: Sequence[ForwardRef('_BaseLagTransform')]) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(_idxs: ndarray) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+
+---
+
+
+
+## class `RollingStd`
+Rolling Standard Deviation
+
+
+
+**Args:**
+
+ - `lag` (int): Number of periods to offset by before applying the transformation.
+ - `window_size` (int): Length of the rolling window.
+ - `min_samples` (int, optional): Minimum number of samples required to compute the statistic. If None, defaults to window_size.
+
+
+
+### method `__init__`
+
+```python
+__init__(lag: int, window_size: int, min_samples: Optional[int] = None)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(transforms: Sequence[ForwardRef('_BaseLagTransform')]) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(_idxs: ndarray) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+
+---
+
+
+
+## class `RollingMin`
+Rolling Minimum
+
+
+
+**Args:**
+
+ - `lag` (int): Number of periods to offset by before applying the transformation.
+ - `window_size` (int): Length of the rolling window.
+ - `min_samples` (int, optional): Minimum number of samples required to compute the statistic. If None, defaults to window_size.
+
+
+
+### method `__init__`
+
+```python
+__init__(lag: int, window_size: int, min_samples: Optional[int] = None)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(transforms: Sequence[ForwardRef('_BaseLagTransform')]) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(_idxs: ndarray) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+
+---
+
+
+
+## class `RollingMax`
+Rolling Maximum
+
+
+
+**Args:**
+
+ - `lag` (int): Number of periods to offset by before applying the transformation.
+ - `window_size` (int): Length of the rolling window.
+ - `min_samples` (int, optional): Minimum number of samples required to compute the statistic. If None, defaults to window_size.
+
+
+
+### method `__init__`
+
+```python
+__init__(lag: int, window_size: int, min_samples: Optional[int] = None)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(transforms: Sequence[ForwardRef('_BaseLagTransform')]) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(_idxs: ndarray) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+
+---
+
+
+
+## class `RollingQuantile`
+Rolling quantile
+
+
+
+**Args:**
+
+ - `lag` (int): Number of periods to offset by before applying the transformation
+ - `p` (float): Quantile to compute
+ - `window_size` (int): Length of the rolling window
+ - `min_samples` (int, optional): Minimum number of samples required to compute the statistic. If None, defaults to window_size.
+
+
+
+### method `__init__`
+
+```python
+__init__(
+ lag: int,
+ p: float,
+ window_size: int,
+ min_samples: Optional[int] = None
+)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(transforms: Sequence[ForwardRef('_BaseLagTransform')]) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(_idxs: ndarray) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+
+---
+
+
+
+## class `SeasonalRollingMean`
+Seasonal rolling Mean
+
+
+
+**Args:**
+
+ - `lag` (int): Number of periods to offset by before applying the transformation
+ - `season_length` (int): Length of the seasonal period, e.g. 7 for weekly data
+ - `window_size` (int): Length of the rolling window
+ - `min_samples` (int, optional): Minimum number of samples required to compute the statistic. If None, defaults to window_size.
+
+
+
+### method `__init__`
+
+```python
+__init__(
+ lag: int,
+ season_length: int,
+ window_size: int,
+ min_samples: Optional[int] = None
+)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(transforms: Sequence[ForwardRef('_BaseLagTransform')]) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(_idxs: ndarray) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+
+---
+
+
+
+## class `SeasonalRollingStd`
+Seasonal rolling Standard Deviation
+
+
+
+**Args:**
+
+ - `lag` (int): Number of periods to offset by before applying the transformation
+ - `season_length` (int): Length of the seasonal period, e.g. 7 for weekly data
+ - `window_size` (int): Length of the rolling window
+ - `min_samples` (int, optional): Minimum number of samples required to compute the statistic. If None, defaults to window_size.
+
+
+
+### method `__init__`
+
+```python
+__init__(
+ lag: int,
+ season_length: int,
+ window_size: int,
+ min_samples: Optional[int] = None
+)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(transforms: Sequence[ForwardRef('_BaseLagTransform')]) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(_idxs: ndarray) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+
+---
+
+
+
+## class `SeasonalRollingMin`
+Seasonal rolling Minimum
+
+
+
+**Args:**
+
+ - `lag` (int): Number of periods to offset by before applying the transformation
+ - `season_length` (int): Length of the seasonal period, e.g. 7 for weekly data
+ - `window_size` (int): Length of the rolling window
+ - `min_samples` (int, optional): Minimum number of samples required to compute the statistic. If None, defaults to window_size.
+
+
+
+### method `__init__`
+
+```python
+__init__(
+ lag: int,
+ season_length: int,
+ window_size: int,
+ min_samples: Optional[int] = None
+)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(transforms: Sequence[ForwardRef('_BaseLagTransform')]) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(_idxs: ndarray) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+
+---
+
+
+
+## class `SeasonalRollingMax`
+Seasonal rolling Maximum
+
+
+
+**Args:**
+
+ - `lag` (int): Number of periods to offset by before applying the transformation
+ - `season_length` (int): Length of the seasonal period, e.g. 7 for weekly data
+ - `window_size` (int): Length of the rolling window
+ - `min_samples` (int, optional): Minimum number of samples required to compute the statistic. If None, defaults to window_size.
+
+
+
+### method `__init__`
+
+```python
+__init__(
+ lag: int,
+ season_length: int,
+ window_size: int,
+ min_samples: Optional[int] = None
+)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(transforms: Sequence[ForwardRef('_BaseLagTransform')]) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(_idxs: ndarray) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+
+---
+
+
+
+## class `SeasonalRollingQuantile`
+Seasonal rolling statistic
+
+
+
+**Args:**
+
+ - `lag` (int): Number of periods to offset by before applying the transformation
+ - `p` (float): Quantile to compute
+ - `season_length` (int): Length of the seasonal period, e.g. 7 for weekly data
+ - `window_size` (int): Length of the rolling window
+ - `min_samples` (int, optional): Minimum number of samples required to compute the statistic. If None, defaults to window_size.
+
+
+
+### method `__init__`
+
+```python
+__init__(
+ lag: int,
+ p: float,
+ season_length: int,
+ window_size: int,
+ min_samples: Optional[int] = None
+)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(transforms: Sequence[ForwardRef('_BaseLagTransform')]) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(_idxs: ndarray) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+
+---
+
+
+
+## class `ExpandingMean`
+Expanding Mean
+
+
+
+**Args:**
+
+ - `lag` (int): Number of periods to offset by before applying the transformation
+
+
+
+### method `__init__`
+
+```python
+__init__(lag: int)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(transforms: Sequence[ForwardRef('_BaseLagTransform')]) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(idxs: ndarray) → _ExpandingBase
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+
+---
+
+
+
+## class `ExpandingStd`
+Expanding Standard Deviation
+
+
+
+**Args:**
+
+ - `lag` (int): Number of periods to offset by before applying the transformation
+
+
+
+### method `__init__`
+
+```python
+__init__(lag: int)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(transforms: Sequence[ForwardRef('_BaseLagTransform')]) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(idxs: ndarray) → _ExpandingBase
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+
+---
+
+
+
+## class `ExpandingMin`
+Expanding Minimum
+
+
+
+**Args:**
+
+ - `lag` (int): Number of periods to offset by before applying the transformation
+
+
+
+### method `__init__`
+
+```python
+__init__(lag: int)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(transforms: Sequence[ForwardRef('_BaseLagTransform')]) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(idxs: ndarray) → _ExpandingBase
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+
+---
+
+
+
+## class `ExpandingMax`
+Expanding Maximum
+
+
+
+**Args:**
+
+ - `lag` (int): Number of periods to offset by before applying the transformation
+
+
+
+### method `__init__`
+
+```python
+__init__(lag: int)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(transforms: Sequence[ForwardRef('_BaseLagTransform')]) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(idxs: ndarray) → _ExpandingBase
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+
+---
+
+
+
+## class `ExpandingQuantile`
+Expanding quantile
+
+
+
+**Args:**
+ lag (int): Number of periods to offset by before applying the transformation p (float): Quantile to compute
+
+
+
+### method `__init__`
+
+```python
+__init__(lag: int, p: float)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(transforms: Sequence[ForwardRef('_BaseLagTransform')]) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(_idxs: ndarray) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+
+---
+
+
+
+## class `ExponentiallyWeightedMean`
+Exponentially weighted mean
+
+
+
+**Args:**
+
+ - `lag` (int): Number of periods to offset by before applying the transformation
+ - `alpha` (float): Smoothing factor
+
+
+
+### method `__init__`
+
+```python
+__init__(lag: int, alpha: float)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(transforms: Sequence[ForwardRef('_BaseLagTransform')]) → _BaseLagTransform
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(idxs: ndarray) → ExponentiallyWeightedMean
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArray') → ndarray
+```
+
+
+
+
+
+
+
+
+---
+
+_This file was automatically generated via [lazydocs](https://github.com/ml-tooling/lazydocs)._
diff --git a/coreforecast/light.png b/coreforecast/light.png
new file mode 100644
index 00000000..bbb99b54
Binary files /dev/null and b/coreforecast/light.png differ
diff --git a/coreforecast/mint.json b/coreforecast/mint.json
new file mode 100644
index 00000000..e3fdd385
--- /dev/null
+++ b/coreforecast/mint.json
@@ -0,0 +1,41 @@
+{
+ "$schema": "https://mintlify.com/schema.json",
+ "name": "Nixtla",
+ "logo": {
+ "light": "/light.png",
+ "dark": "/dark.png"
+ },
+ "favicon": "/favicon.svg",
+ "colors": {
+ "primary": "#0E0E0E",
+ "light": "#FAFAFA",
+ "dark": "#0E0E0E",
+ "anchors": {
+ "from": "#2AD0CA",
+ "to": "#0E00F8"
+ }
+ },
+ "topbarCtaButton": {
+ "type": "github",
+ "url": "https://github.com/Nixtla/coreforecast"
+ },
+ "navigation": [
+ {
+ "group": "",
+ "pages": ["index"]
+ },
+ {
+ "group": "API Reference",
+ "pages": [
+ "grouped_array",
+ "lag_transforms",
+ "scalers",
+ "differences",
+ "seasonal",
+ "rolling",
+ "expanding",
+ "exponentially_weighted"
+ ]
+ }
+ ]
+}
diff --git a/coreforecast/rolling.mdx b/coreforecast/rolling.mdx
new file mode 100644
index 00000000..f084a355
--- /dev/null
+++ b/coreforecast/rolling.mdx
@@ -0,0 +1,339 @@
+
+
+
+
+# module `coreforecast.rolling`
+
+
+
+
+
+---
+
+
+
+## function `rolling_mean`
+
+```python
+rolling_mean(
+ x: ndarray,
+ window_size: int,
+ min_samples: Optional[int] = None
+) → ndarray
+```
+
+Compute the rolling_mean of the input array.
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Input array.
+ - `window_size` (int): The size of the rolling window.
+ - `min_samples` (int, optional): The minimum number of samples required to compute the statistic. If None, it is set to `window_size`.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the rolling statistic
+
+
+---
+
+
+
+## function `rolling_std`
+
+```python
+rolling_std(
+ x: ndarray,
+ window_size: int,
+ min_samples: Optional[int] = None
+) → ndarray
+```
+
+Compute the rolling_std of the input array.
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Input array.
+ - `window_size` (int): The size of the rolling window.
+ - `min_samples` (int, optional): The minimum number of samples required to compute the statistic. If None, it is set to `window_size`.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the rolling statistic
+
+
+---
+
+
+
+## function `rolling_min`
+
+```python
+rolling_min(
+ x: ndarray,
+ window_size: int,
+ min_samples: Optional[int] = None
+) → ndarray
+```
+
+Compute the rolling_min of the input array.
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Input array.
+ - `window_size` (int): The size of the rolling window.
+ - `min_samples` (int, optional): The minimum number of samples required to compute the statistic. If None, it is set to `window_size`.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the rolling statistic
+
+
+---
+
+
+
+## function `rolling_max`
+
+```python
+rolling_max(
+ x: ndarray,
+ window_size: int,
+ min_samples: Optional[int] = None
+) → ndarray
+```
+
+Compute the rolling_max of the input array.
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Input array.
+ - `window_size` (int): The size of the rolling window.
+ - `min_samples` (int, optional): The minimum number of samples required to compute the statistic. If None, it is set to `window_size`.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the rolling statistic
+
+
+---
+
+
+
+## function `rolling_quantile`
+
+```python
+rolling_quantile(
+ x: ndarray,
+ p: float,
+ window_size: int,
+ min_samples: Optional[int] = None
+) → ndarray
+```
+
+Compute the rolling_quantile of the input array.
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Input array.
+ - `q` (float): Quantile to compute.
+ - `window_size` (int): The size of the rolling window.
+ - `min_samples` (int, optional): The minimum number of samples required to compute the statistic. If None, it is set to `window_size`.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with rolling statistic
+
+
+---
+
+
+
+## function `seasonal_rolling_mean`
+
+```python
+seasonal_rolling_mean(
+ x: ndarray,
+ season_length: int,
+ window_size: int,
+ min_samples: Optional[int] = None
+) → ndarray
+```
+
+Compute the seasonal_rolling_mean of the input array
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Input array.
+ - `season_length` (int): The length of the seasonal period.
+ - `window_size` (int): The size of the rolling window.
+ - `min_samples` (int, optional): The minimum number of samples required to compute the statistic. If None, it is set to `window_size`.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the seasonal rolling statistic
+
+
+---
+
+
+
+## function `seasonal_rolling_std`
+
+```python
+seasonal_rolling_std(
+ x: ndarray,
+ season_length: int,
+ window_size: int,
+ min_samples: Optional[int] = None
+) → ndarray
+```
+
+Compute the seasonal_rolling_std of the input array
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Input array.
+ - `season_length` (int): The length of the seasonal period.
+ - `window_size` (int): The size of the rolling window.
+ - `min_samples` (int, optional): The minimum number of samples required to compute the statistic. If None, it is set to `window_size`.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the seasonal rolling statistic
+
+
+---
+
+
+
+## function `seasonal_rolling_min`
+
+```python
+seasonal_rolling_min(
+ x: ndarray,
+ season_length: int,
+ window_size: int,
+ min_samples: Optional[int] = None
+) → ndarray
+```
+
+Compute the seasonal_rolling_min of the input array
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Input array.
+ - `season_length` (int): The length of the seasonal period.
+ - `window_size` (int): The size of the rolling window.
+ - `min_samples` (int, optional): The minimum number of samples required to compute the statistic. If None, it is set to `window_size`.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the seasonal rolling statistic
+
+
+---
+
+
+
+## function `seasonal_rolling_max`
+
+```python
+seasonal_rolling_max(
+ x: ndarray,
+ season_length: int,
+ window_size: int,
+ min_samples: Optional[int] = None
+) → ndarray
+```
+
+Compute the seasonal_rolling_max of the input array
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Input array.
+ - `season_length` (int): The length of the seasonal period.
+ - `window_size` (int): The size of the rolling window.
+ - `min_samples` (int, optional): The minimum number of samples required to compute the statistic. If None, it is set to `window_size`.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the seasonal rolling statistic
+
+
+---
+
+
+
+## function `seasonal_rolling_quantile`
+
+```python
+seasonal_rolling_quantile(
+ x: ndarray,
+ p: float,
+ season_length: int,
+ window_size: int,
+ min_samples: Optional[int] = None
+) → ndarray
+```
+
+Compute the seasonal_rolling_quantile of the input array.
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Input array.
+ - `q` (float): Quantile to compute.
+ - `season_length` (int): The length of the seasonal period.
+ - `window_size` (int): The size of the rolling window.
+ - `min_samples` (int, optional): The minimum number of samples required to compute the statistic. If None, it is set to `window_size`.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with rolling statistic
+
+
+
+
+---
+
+_This file was automatically generated via [lazydocs](https://github.com/ml-tooling/lazydocs)._
diff --git a/coreforecast/scalers.mdx b/coreforecast/scalers.mdx
new file mode 100644
index 00000000..abe68848
--- /dev/null
+++ b/coreforecast/scalers.mdx
@@ -0,0 +1,1201 @@
+
+
+
+
+# module `coreforecast.scalers`
+
+
+
+
+**Global Variables**
+---------------
+- **TYPE_CHECKING**
+
+---
+
+
+
+## function `boxcox_lambda`
+
+```python
+boxcox_lambda(
+ x: ndarray,
+ method: str,
+ season_length: Optional[int] = None,
+ lower: float = -0.9,
+ upper: float = 2.0
+) → float
+```
+
+Find optimum lambda for the Box-Cox transformation
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Array with data to transform.
+ - `method` (str): Method to use. Valid options are 'guerrero' and 'loglik'. 'guerrero' minimizes the coefficient of variation for subseries of `x` and supports negative values. 'loglik' maximizes the log-likelihood function.
+ - `season_length` (int, optional): Length of the seasonal period. Only required if method='guerrero'.
+ - `lower` (float): Lower bound for the lambda.
+ - `upper` (float): Upper bound for the lambda.
+
+
+
+**Returns:**
+
+ - `float`: Optimum lambda.
+
+
+---
+
+
+
+## function `boxcox`
+
+```python
+boxcox(x: ndarray, lmbda: float) → ndarray
+```
+
+Apply the Box-Cox transformation
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Array with data to transform.
+ - `lmbda` (float): Lambda value to use.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the transformed data.
+
+
+---
+
+
+
+## function `inv_boxcox`
+
+```python
+inv_boxcox(x: ndarray, lmbda: float) → ndarray
+```
+
+Invert the Box-Cox transformation
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Array with data to transform.
+ - `lmbda` (float): Lambda value to use.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the inverted transformation.
+
+
+---
+
+
+
+## class `LocalMinMaxScaler`
+Scale each group to the [0, 1] interval
+
+
+
+
+---
+
+
+
+### method `fit`
+
+```python
+fit(ga: 'GroupedArrayT') → _BaseLocalScaler
+```
+
+Compute the statistics for each group.
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `self`: The fitted scaler object.
+
+---
+
+
+
+### method `fit_transform`
+
+```python
+fit_transform(ga: 'GroupedArrayT') → ndarray
+```
+
+"Compute the statistics for each group and apply the transformation.
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the transformed data.
+
+---
+
+
+
+### method `inverse_transform`
+
+```python
+inverse_transform(ga: 'GroupedArrayT') → ndarray
+```
+
+Use the computed statistics to invert the transformation.
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the inverted transformation.
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(scalers: Sequence[ForwardRef('_BaseLocalScaler')]) → _BaseLocalScaler
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(idxs: ndarray) → _BaseLocalScaler
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArrayT') → ndarray
+```
+
+Use the computed statistics to apply the transformation.
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the transformed data.
+
+
+---
+
+
+
+## class `LocalStandardScaler`
+Scale each group to have zero mean and unit variance
+
+
+
+
+---
+
+
+
+### method `fit`
+
+```python
+fit(ga: 'GroupedArrayT') → _BaseLocalScaler
+```
+
+Compute the statistics for each group.
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `self`: The fitted scaler object.
+
+---
+
+
+
+### method `fit_transform`
+
+```python
+fit_transform(ga: 'GroupedArrayT') → ndarray
+```
+
+"Compute the statistics for each group and apply the transformation.
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the transformed data.
+
+---
+
+
+
+### method `inverse_transform`
+
+```python
+inverse_transform(ga: 'GroupedArrayT') → ndarray
+```
+
+Use the computed statistics to invert the transformation.
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the inverted transformation.
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(scalers: Sequence[ForwardRef('_BaseLocalScaler')]) → _BaseLocalScaler
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(idxs: ndarray) → _BaseLocalScaler
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArrayT') → ndarray
+```
+
+Use the computed statistics to apply the transformation.
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the transformed data.
+
+
+---
+
+
+
+## class `LocalRobustScaler`
+Scale each group using robust statistics
+
+
+
+**Args:**
+
+ - `scale` (str): Type of robust scaling to use. Valid options are 'iqr' and 'mad'. If 'iqr' will use the inter quartile range as the scale. If 'mad' will use median absolute deviation as the scale.
+
+
+
+### method `__init__`
+
+```python
+__init__(scale: str)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `fit`
+
+```python
+fit(ga: 'GroupedArrayT') → _BaseLocalScaler
+```
+
+Compute the statistics for each group.
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `self`: The fitted scaler object.
+
+---
+
+
+
+### method `fit_transform`
+
+```python
+fit_transform(ga: 'GroupedArrayT') → ndarray
+```
+
+"Compute the statistics for each group and apply the transformation.
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the transformed data.
+
+---
+
+
+
+### method `inverse_transform`
+
+```python
+inverse_transform(ga: 'GroupedArrayT') → ndarray
+```
+
+Use the computed statistics to invert the transformation.
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the inverted transformation.
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(scalers: Sequence[ForwardRef('_BaseLocalScaler')]) → _BaseLocalScaler
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(idxs: ndarray) → _BaseLocalScaler
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArrayT') → ndarray
+```
+
+Use the computed statistics to apply the transformation.
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the transformed data.
+
+
+---
+
+
+
+## class `LocalBoxCoxScaler`
+Find the optimum lambda for the Box-Cox transformation by group and apply it
+
+
+
+**Args:**
+
+ - `season_length` (int, optional): Length of the seasonal period. Only required if method='guerrero'.
+ - `lower` (float): Lower bound for the lambda.
+ - `upper` (float): Upper bound for the lambda.
+ - `method` (str): Method to use. Valid options are 'guerrero' and 'loglik'. 'guerrero' minimizes the coefficient of variation for subseries of `x` and supports negative values. 'loglik' maximizes the log-likelihood function.
+
+
+
+### method `__init__`
+
+```python
+__init__(
+ method: str,
+ season_length: Optional[int] = None,
+ lower: float = -0.9,
+ upper: float = 2.0
+)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `fit`
+
+```python
+fit(ga: 'GroupedArrayT') → _BaseLocalScaler
+```
+
+Compute the statistics for each group.
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `self`: The fitted scaler object.
+
+---
+
+
+
+### method `fit_transform`
+
+```python
+fit_transform(ga: 'GroupedArrayT') → ndarray
+```
+
+"Compute the statistics for each group and apply the transformation.
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the transformed data.
+
+---
+
+
+
+### method `inverse_transform`
+
+```python
+inverse_transform(ga: 'GroupedArrayT') → ndarray
+```
+
+Use the computed lambdas to invert the transformation.
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the inverted transformation.
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(scalers: Sequence[ForwardRef('_BaseLocalScaler')]) → _BaseLocalScaler
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(idxs: ndarray) → _BaseLocalScaler
+```
+
+
+
+
+
+---
+
+
+
+### method `transform`
+
+```python
+transform(ga: 'GroupedArrayT') → ndarray
+```
+
+Use the computed lambdas to apply the transformation.
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the transformed data.
+
+
+---
+
+
+
+## class `Difference`
+Subtract a lag to each group
+
+
+
+**Args:**
+
+ - `d` (int): Lag to subtract.
+
+
+
+### method `__init__`
+
+```python
+__init__(d: int)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `fit_transform`
+
+```python
+fit_transform(ga: 'GroupedArrayT') → ndarray
+```
+
+Apply the transformation
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the transformed data.
+
+---
+
+
+
+### method `inverse_transform`
+
+```python
+inverse_transform(ga: 'GroupedArrayT') → ndarray
+```
+
+Invert the transformation
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the inverted transformation.
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(scalers: Sequence[ForwardRef('Difference')]) → Difference
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(idxs: ndarray) → Difference
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArrayT') → ndarray
+```
+
+Update the last observations from each serie
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the updated data.
+
+
+---
+
+
+
+## class `AutoDifferences`
+Find and apply the optimal number of differences to each group.
+
+
+
+**Args:**
+
+ - `max_diffs` (int): Maximum number of differences to apply.
+
+
+
+### method `__init__`
+
+```python
+__init__(max_diffs: int)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `fit_transform`
+
+```python
+fit_transform(ga: 'GroupedArrayT') → ndarray
+```
+
+Compute and apply the optimal number of differences for each group
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the transformed data.
+
+---
+
+
+
+### method `inverse_transform`
+
+```python
+inverse_transform(ga: 'GroupedArrayT') → ndarray
+```
+
+Invert the differences
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the inverted transformation.
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(scalers: Sequence[ForwardRef('AutoDifferences')]) → AutoDifferences
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(idxs: ndarray) → AutoDifferences
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArrayT') → ndarray
+```
+
+Update the last observations from each serie
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the updated data.
+
+
+---
+
+
+
+## class `AutoSeasonalDifferences`
+Find and apply the optimal number of seasonal differences to each group.
+
+
+
+**Args:**
+
+ - `season_length` (int): Length of the seasonal period.
+ - `max_diffs` (int): Maximum number of differences to apply.
+ - `n_seasons` (int | None): Number of seasons to use to determine the number of differences. Defaults to 10. If `None` will use all samples, otherwise `season_length` * `n_seasons` samples will be used for the test. Smaller values will be faster but could be less accurate.
+
+
+
+### method `__init__`
+
+```python
+__init__(season_length: int, max_diffs: int, n_seasons: Optional[int] = 10)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `fit_transform`
+
+```python
+fit_transform(ga: 'GroupedArrayT') → ndarray
+```
+
+Compute and apply the optimal number of seasonal differences for each group
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the transformed data.
+
+---
+
+
+
+### method `inverse_transform`
+
+```python
+inverse_transform(ga: 'GroupedArrayT') → ndarray
+```
+
+Invert the seasonal differences
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the inverted transformation.
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(scalers: Sequence[ForwardRef('AutoDifferences')]) → AutoDifferences
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(idxs: ndarray) → AutoDifferences
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArrayT') → ndarray
+```
+
+Update the last observations from each serie
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the updated data.
+
+
+---
+
+
+
+## class `AutoSeasonalityAndDifferences`
+Find the length of the seasonal period and apply the optimal number of differences to each group.
+
+
+
+**Args:**
+
+ - `max_season_length` (int): Maximum length of the seasonal period.
+ - `max_diffs` (int): Maximum number of differences to apply.
+ - `n_seasons` (int | None): Number of seasons to use to determine the number of differences. Defaults to 10. If `None` will use all samples, otherwise `max_season_length` * `n_seasons` samples will be used for the test. Smaller values will be faster but could be less accurate.
+
+
+
+### method `__init__`
+
+```python
+__init__(max_season_length: int, max_diffs: int, n_seasons: Optional[int] = 10)
+```
+
+
+
+
+
+
+
+
+---
+
+
+
+### method `fit_transform`
+
+```python
+fit_transform(ga: 'GroupedArrayT') → ndarray
+```
+
+Compute the optimal length of the seasonal period and apply the optimal number of differences for each group
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the transformed data.
+
+---
+
+
+
+### method `inverse_transform`
+
+```python
+inverse_transform(ga: 'GroupedArrayT') → ndarray
+```
+
+Invert the seasonal differences
+
+
+
+**Args:**
+
+ - `ga` (GroupedArray): Array with grouped data.
+
+
+
+**Returns:**
+
+ - `np.ndarray`: Array with the inverted transformation.
+
+---
+
+
+
+### method `stack`
+
+```python
+stack(
+ scalers: Sequence[ForwardRef('AutoSeasonalityAndDifferences')]
+) → AutoSeasonalityAndDifferences
+```
+
+
+
+
+
+---
+
+
+
+### method `take`
+
+```python
+take(idxs: ndarray) → AutoSeasonalityAndDifferences
+```
+
+
+
+
+
+---
+
+
+
+### method `update`
+
+```python
+update(ga: 'GroupedArrayT') → ndarray
+```
+
+
+
+
+
+
+
+
+---
+
+_This file was automatically generated via [lazydocs](https://github.com/ml-tooling/lazydocs)._
diff --git a/coreforecast/seasonal.mdx b/coreforecast/seasonal.mdx
new file mode 100644
index 00000000..7aaf79ec
--- /dev/null
+++ b/coreforecast/seasonal.mdx
@@ -0,0 +1,40 @@
+
+
+
+
+# module `coreforecast.seasonal`
+
+
+
+
+
+---
+
+
+
+## function `find_season_length`
+
+```python
+find_season_length(x: ndarray, max_season_length: int) → int
+```
+
+Find the length of the seasonal period of the time series. Returns 0 if no seasonality is found.
+
+
+
+**Args:**
+
+ - `x` (np.ndarray): Array with the time series.
+
+
+
+**Returns:**
+
+ - `int`: Season period.
+
+
+
+
+---
+
+_This file was automatically generated via [lazydocs](https://github.com/ml-tooling/lazydocs)._
diff --git a/coreforecast/utils.mdx b/coreforecast/utils.mdx
new file mode 100644
index 00000000..b8259835
--- /dev/null
+++ b/coreforecast/utils.mdx
@@ -0,0 +1,16 @@
+
+
+
+
+# module `coreforecast.utils`
+
+
+
+
+
+
+
+
+---
+
+_This file was automatically generated via [lazydocs](https://github.com/ml-tooling/lazydocs)._
diff --git a/datasetsforecast/.nojekyll b/datasetsforecast/.nojekyll
new file mode 100644
index 00000000..e69de29b
diff --git a/datasetsforecast/dark.png b/datasetsforecast/dark.png
new file mode 100644
index 00000000..4142a0bb
Binary files /dev/null and b/datasetsforecast/dark.png differ
diff --git a/datasetsforecast/favicon.svg b/datasetsforecast/favicon.svg
new file mode 100644
index 00000000..e5f33342
--- /dev/null
+++ b/datasetsforecast/favicon.svg
@@ -0,0 +1,5 @@
+
diff --git a/datasetsforecast/favorita.html.mdx b/datasetsforecast/favorita.html.mdx
new file mode 100644
index 00000000..aeec307c
--- /dev/null
+++ b/datasetsforecast/favorita.html.mdx
@@ -0,0 +1,373 @@
+---
+output-file: favorita.html
+title: Favorita
+---
+
+
+## Auxiliary Functions
+
+This auxiliary functions are used to efficiently create and wrangle
+Favorita’s series.
+
+### Numpy Wrangling
+
+------------------------------------------------------------------------
+
+source
+
+#### numpy_balance
+
+> ``` text
+> numpy_balance (*arrs)
+> ```
+
+\*Fast NumPy implementation of ‘balance’ operation, useful to create a
+balanced panel dataset, ie a dataset with all the interactions of
+‘unique_id’ and ‘ds’.
+
+**Parameters:** `arrs`: NumPy arrays.
+
+**Returns:** `out`: NumPy array.\*
+
+------------------------------------------------------------------------
+
+source
+
+#### numpy_ffill
+
+> ``` text
+> numpy_ffill (arr)
+> ```
+
+\*Fast NumPy implementation of `ffill` that fills missing values in an
+array by propagating the last non-missing value forward.
+
+For example, if the array has the following values: 0 1 2 3 1 2
+NaN 4
+
+The `ffill` method would fill the missing values as follows: 0 1 2
+3 1 2 2 4
+
+**Parameters:** `arr`: NumPy array.
+
+**Returns:** `out`: NumPy array.\*
+
+------------------------------------------------------------------------
+
+source
+
+#### numpy_bfill
+
+> ``` text
+> numpy_bfill (arr)
+> ```
+
+\*Fast NumPy implementation of `bfill` that fills missing values in an
+array by propagating the last non-missing value backwards.
+
+For example, if the array has the following values: 0 1 2 3 1 2
+NaN 4
+
+The `bfill` method would fill the missing values as follows: 0 1 2
+3 1 2 4 4
+
+**Parameters:** `arr`: NumPy array.
+
+**Returns:** `out`: NumPy array.\*
+
+### Pandas Wrangling
+
+------------------------------------------------------------------------
+
+source
+
+#### one_hot_encoding
+
+> ``` text
+> one_hot_encoding (df, index_col)
+> ```
+
+\*Encodes dataFrame `df`’s categorical variables skipping `index_col`.
+
+**Parameters:** `df`: pd.DataFrame with categorical columns.
+`index_col`: str, the index column to avoid encoding.
+
+**Returns:** `one_hot_concat_df`: pd.DataFrame with one hot encoded
+categorical columns. \*
+
+------------------------------------------------------------------------
+
+source
+
+#### nested_one_hot_encoding
+
+> ``` text
+> nested_one_hot_encoding (df, index_col)
+> ```
+
+\*Encodes dataFrame `df`’s hierarchically-nested categorical variables
+skipping `index_col`.
+
+Nested categorical variables (example geographic levels country\>state),
+require the dummy features to preserve encoding order, to reflect the
+hierarchy of the categorical variables.
+
+**Parameters:** `df`: pd.DataFrame with hierarchically-nested
+categorical columns. `index_col`: str, the index column to avoid
+encoding.
+
+**Returns:** `one_hot_concat_df`: pd.DataFrame with one hot encoded
+hierarchically-nested categorical columns. \*
+
+------------------------------------------------------------------------
+
+source
+
+#### get_levels_from_S_df
+
+> ``` text
+> get_levels_from_S_df (S_df)
+> ```
+
+\*Get hierarchical index levels implied by aggregation constraints
+dataframe `S_df`.
+
+Create levels from summation matrix (base, bottom). Goes through the
+rows until all the bottom level series are ‘covered’ by the aggregation
+constraints to discover blocks/hierarchy levels.
+
+**Parameters:** `S_df`: pd.DataFrame with summing matrix of size
+`(base, bottom)`, see [aggregate
+method](https://nixtlaverse.nixtla.io/hierarchicalforecast/src/utils.html#aggregate).
+
+**Returns:** `levels`: list, with hierarchical aggregation indexes,
+where each entry is a level.\*
+
+## Favorita Dataset
+
+### Favorita Raw
+
+------------------------------------------------------------------------
+
+source
+
+#### FavoritaRawData
+
+> ``` text
+> FavoritaRawData ()
+> ```
+
+\*Favorita Raw Data
+
+Raw subset datasets from the Favorita 2018 Kaggle competition. This
+class contains utilities to download, load and filter portions of the
+dataset.
+
+If you prefer, you can also download original dataset available from
+Kaggle directly. `pip install kaggle --upgrade`
+`kaggle competitions download -c favorita-grocery-sales-forecasting`\*
+
+------------------------------------------------------------------------
+
+source
+
+#### FavoritaRawData.\_load_raw_group_data
+
+> ``` text
+> FavoritaRawData._load_raw_group_data (directory, group, verbose=False)
+> ```
+
+\*Load raw group data.
+
+Reads, filters and sorts Favorita subset dataset.
+
+**Parameters:** `directory`: str, Directory where data will be
+downloaded. `group`: str, dataset group name in ‘Favorita200’,
+‘Favorita500’, ‘FavoritaComplete’. `verbose`: bool=False, wether or
+not print partial outputs.
+
+**Returns:** `filter_items`: ordered list with unique items
+identifiers in the Favorita subset. `filter_stores`: ordered list
+with unique store identifiers in the Favorita subset.
+`filter_dates`: ordered list with dates in the Favorita subset.
+`raw_group_data`: dictionary with original raw Favorita pd.DataFrames,
+temporal, oil, items, store_info, holidays, transactions. \*
+
+#### Favorita Raw Usage example
+
+```python
+from datasetsforecast.favorita import FavoritaRawData
+
+verbose = True
+group = 'Favorita200' # 'Favorita500', 'FavoritaComplete'
+directory = './data/favorita' # directory = f's3://favorita'
+
+filter_items, filter_stores, filter_dates, raw_group_data = \
+ FavoritaRawData._load_raw_group_data(directory=directory, group=group, verbose=verbose)
+n_items = len(filter_items)
+n_stores = len(filter_stores)
+n_dates = len(filter_dates)
+
+print('\n')
+print('n_stores: \t', n_stores)
+print('n_items: \t', n_items)
+print('n_dates: \t', n_dates)
+print('n_items * n_dates: \t\t',n_items * n_dates)
+print('n_items * n_stores: \t\t',n_items * n_stores)
+print('n_items * n_dates * n_stores: \t', n_items * n_dates * n_stores)
+```
+
+### FavoritaData
+
+------------------------------------------------------------------------
+
+source
+
+#### FavoritaData
+
+> ``` text
+> FavoritaData ()
+> ```
+
+\*Favorita Data
+
+The processed Favorita dataset of grocery contains item sales daily
+history with additional information on promotions, items, stores, and
+holidays, containing 371,312 series from January 2013 to August 2017,
+with a geographic hierarchy of states, cities, and stores. This
+wrangling matches that of the DPMN paper.
+
+- [Kin G. Olivares, O. Nganba Meetei, Ruijun Ma, Rohan Reddy, Mengfei
+ Cao, Lee Dicker (2022).”Probabilistic Hierarchical Forecasting with
+ Deep Poisson Mixtures”. International Journal Forecasting, special
+ issue.](https://doi.org/10.1016/j.ijforecast.2023.04.007)\*
+
+------------------------------------------------------------------------
+
+source
+
+#### FavoritaData.load_preprocessed
+
+> ``` text
+> FavoritaData.load_preprocessed (directory:str, group:str,
+> cache:bool=True, verbose:bool=False)
+> ```
+
+\*Load Favorita group datasets.
+
+For the exploration of more complex models, we make available the entire
+information including data at the bottom level of the items sold in
+Favorita stores, in addition to the aggregate/national level information
+for the items.
+
+**Parameters:** `directory`: str, directory where data will be
+downloaded and saved. `group`: str, dataset group name in
+‘Favorita200’, ‘Favorita500’, ‘FavoritaComplete’. `cache`:
+bool=False, If `True` saves and loads. `verbose`: bool=False, wether
+or not print partial outputs.
+
+**Returns:** `static_bottom`: pd.DataFrame, with static variables of
+bottom level series. `static_agg`: pd.DataFrame, with static
+variables of aggregate level series. `temporal_bottom`:
+pd.DataFrame, with temporal variables of bottom level series.
+`temporal_agg`: pd.DataFrame, with temporal variables of aggregate level
+series. \*
+
+------------------------------------------------------------------------
+
+source
+
+#### FavoritaData.load
+
+> ``` text
+> FavoritaData.load (directory:str, group:str, cache:bool=True,
+> verbose:bool=False)
+> ```
+
+\*Load Favorita forecasting benchmark dataset.
+
+In contrast with other hierarchical datasets, this dataset contains a
+geographic hierarchy for each individual grocery item series, identified
+with ‘item_id’ column. The geographic hierarchy is captured by the
+‘hier_id’ column.
+
+For this reason minor wrangling is needed to adapt it for use with
+[`HierarchicalForecast`](https://github.com/Nixtla/hierarchicalforecast),
+and [`StatsForecast`](https://github.com/Nixtla/statsforecast)
+libraries.
+
+**Parameters:** `directory`: str, directory where data will be
+downloaded and saved. `group`: str, dataset group name in
+‘Favorita200’, ‘Favorita500’, ‘FavoritaComplete’. `cache`:
+bool=False, If `True` saves and loads. `verbose`: bool=False, wether
+or not print partial outputs.
+
+**Returns:** `Y_df`: pd.DataFrame, target base time series with
+columns \[‘item_id’, ‘hier_id’, ‘ds’, ‘y’\]. `S_df`: pd.DataFrame,
+hierarchical constraints dataframe of size (base, bottom). \*
+
+```python
+# #| hide
+# #| eval: false
+# # Test the equality of created and loaded datasets columns and rows
+# static_agg1, static_bottom1, temporal_agg1, temporal_bottom1, S_df1 = \
+# FavoritaData.load_preprocessed(directory=directory, group=group, cache=False)
+
+# static_agg2, static_bottom2, temporal_agg2, temporal_bottom2, S_df2 = \
+# FavoritaData.load_preprocessed(directory=directory, group=group)
+
+# test_eq(len(static_agg1)+len(static_agg1.columns),
+# len(static_agg2)+len(static_agg2.columns))
+# test_eq(len(static_bottom1)+len(static_bottom1.columns),
+# len(static_bottom2)+len(static_bottom2.columns))
+
+# test_eq(len(temporal_agg1)+len(temporal_agg1.columns),
+# len(temporal_agg2)+len(temporal_agg2.columns))
+# test_eq(len(temporal_bottom1)+len(temporal_bottom1.columns),
+# len(temporal_bottom2)+len(temporal_bottom2.columns))
+```
+
+#### Favorita Usage Example
+
+```python
+# Qualitative evaluation of hierarchical data
+from datasetsforecast.favorita import FavoritaData
+from hierarchicalforecast.utils import HierarchicalPlot
+
+group = 'Favorita200' # 'Favorita500', 'FavoritaComplete'
+directory = './data/favorita'
+Y_df, S_df, tags = FavoritaData.load(directory=directory, group=group)
+
+Y_item_df = Y_df[Y_df.item_id==1916577] # 112830, 1501570, 1916577
+Y_item_df = Y_item_df.rename(columns={'hier_id': 'unique_id'})
+Y_item_df = Y_item_df.set_index('unique_id')
+del Y_item_df['item_id']
+
+hplots = HierarchicalPlot(S=S_df, tags=tags)
+hplots.plot_hierarchically_linked_series(
+ Y_df=Y_item_df, bottom_series='store_[40]',
+)
+```
+
diff --git a/datasetsforecast/hierarchical.html.mdx b/datasetsforecast/hierarchical.html.mdx
new file mode 100644
index 00000000..8bd08937
--- /dev/null
+++ b/datasetsforecast/hierarchical.html.mdx
@@ -0,0 +1,168 @@
+---
+output-file: hierarchical.html
+title: Hierarchical Datasets
+---
+
+
+Here we host a collection of datasets used in previous hierarchical
+research by Rangapuram et al. \[2021\], Olivares et al. \[2023\], and
+Kamarthi et al. \[2022\]. The benchmark datasets utilized include
+Australian Monthly Labour
+([`Labour`](https://Nixtla.github.io/datasetsforecast/hierarchical.html#labour)),
+SF Bay Area daily Traffic
+([`Traffic`](https://Nixtla.github.io/datasetsforecast/hierarchical.html#traffic),
+[`OldTraffic`](https://Nixtla.github.io/datasetsforecast/hierarchical.html#oldtraffic)),
+Quarterly Australian Tourism Visits
+([`TourismSmall`](https://Nixtla.github.io/datasetsforecast/hierarchical.html#tourismsmall)),
+Monthly Australian Tourism visits
+([`TourismLarge`](https://Nixtla.github.io/datasetsforecast/hierarchical.html#tourismlarge),
+[`OldTourismLarge`](https://Nixtla.github.io/datasetsforecast/hierarchical.html#oldtourismlarge)),
+and daily Wikipedia article views
+([`Wiki2`](https://Nixtla.github.io/datasetsforecast/hierarchical.html#wiki2)).
+Old datasets favor the original datasets with minimal target variable
+preprocessing (Rangapuram et al. \[2021\], Olivares et al. \[2023\]),
+while the remaining datasets follow PROFHIT experimental settings.
+
+## References
+
+- [Syama Sundar Rangapuram, Lucien D Werner, Konstantinos Benidis,
+ Pedro Mercado, Jan Gasthaus, Tim Januschowski. (2021). “End-to-End
+ Learning of Coherent Probabilistic Forecasts for Hierarchical Time
+ Series”. Proceedings of the 38th International Conference on Machine
+ Learning
+ (ICML).](https://proceedings.mlr.press/v139/rangapuram21a.html)
+- [Kin G. Olivares, O. Nganba Meetei, Ruijun Ma, Rohan Reddy, Mengfei
+ Cao, Lee Dicker (2022).”Probabilistic Hierarchical Forecasting with
+ Deep Poisson Mixtures”. International Journal Forecasting, special
+ issue.](https://doi.org/10.1016/j.ijforecast.2023.04.007)
+- [Harshavardhan Kamarthi, Lingkai Kong, Alexander Rodriguez, Chao
+ Zhang, and B. Prakash. PROFHIT: Probabilistic robust forecasting for
+ hierarchical time-series. Computing Research Repository.URL
+ https://arxiv.org/abs/2206.07940.](https://arxiv.org/abs/2206.07940)
+
+------------------------------------------------------------------------
+
+source
+
+### Labour
+
+> ``` text
+> Labour (freq:str='MS', horizon:int=8, papers_horizon:int=12,
+> seasonality:int=12, test_size:int=125,
+> tags_names:Tuple[str]=('Country', 'Country/Region',
+> 'Country/Gender/Region', 'Country/Employment/Gender/Region'))
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### TourismLarge
+
+> ``` text
+> TourismLarge (freq:str='MS', horizon:int=12, papers_horizon:int=12,
+> seasonality:int=12, test_size:int=57,
+> tags_names:Tuple[str]=('Country', 'Country/State',
+> 'Country/State/Zone', 'Country/State/Zone/Region',
+> 'Country/Purpose', 'Country/State/Purpose',
+> 'Country/State/Zone/Purpose',
+> 'Country/State/Zone/Region/Purpose'))
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### TourismSmall
+
+> ``` text
+> TourismSmall (freq:str='Q', horizon:int=4, papers_horizon:int=4,
+> seasonality:int=4, test_size:int=9,
+> tags_names:Tuple[str]=('Country', 'Country/Purpose',
+> 'Country/Purpose/State',
+> 'Country/Purpose/State/CityNonCity'))
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### Traffic
+
+> ``` text
+> Traffic (freq:str='D', horizon:int=14, papers_horizon:int=7,
+> seasonality:int=7, test_size:int=91,
+> tags_names:Tuple[str]=('Level1', 'Level2', 'Level3', 'Level4'))
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### Wiki2
+
+> ``` text
+> Wiki2 (freq:str='D', horizon:int=14, papers_horizon:int=7,
+> seasonality:int=7, test_size:int=91,
+> tags_names:Tuple[str]=('Views', 'Views/Country',
+> 'Views/Country/Access', 'Views/Country/Access/Agent',
+> 'Views/Country/Access/Agent/Topic'))
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### OldTraffic
+
+> ``` text
+> OldTraffic (freq:str='D', horizon:int=1, papers_horizon:int=1,
+> seasonality:int=7, test_size:int=91,
+> tags_names:Tuple[str]=('Level1', 'Level2', 'Level3',
+> 'Level4'))
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### OldTourismLarge
+
+> ``` text
+> OldTourismLarge (freq:str='MS', horizon:int=12, papers_horizon:int=12,
+> seasonality:int=12, test_size:int=57,
+> tags_names:Tuple[str]=('Country', 'Country/State',
+> 'Country/State/Zone', 'Country/State/Zone/Region',
+> 'Country/Purpose', 'Country/State/Purpose',
+> 'Country/State/Zone/Purpose',
+> 'Country/State/Zone/Region/Purpose'))
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### HierarchicalData
+
+> ``` text
+> HierarchicalData ()
+> ```
+
+*Initialize self. See help(type(self)) for accurate signature.*
+
diff --git a/datasetsforecast/index.html.mdx b/datasetsforecast/index.html.mdx
new file mode 100644
index 00000000..e15ce8a3
--- /dev/null
+++ b/datasetsforecast/index.html.mdx
@@ -0,0 +1,45 @@
+---
+output-file: index.html
+title: datasetsforecast
+---
+
+
+Datasets for time series forecasting
+
+## Install
+
+```sh
+pip install datasetsforecast
+```
+
+## Datasets
+
+- [Favorita](https://nixtlaverse.nixtla.io/datasetsforecast/favorita.html)
+- [Hierarchical](https://nixtlaverse.nixtla.io/datasetsforecast/hierarchical.html)
+- [Long
+ horizon](https://nixtlaverse.nixtla.io/datasetsforecast/long_horizon.html)
+- [M3](https://nixtlaverse.nixtla.io/datasetsforecast/m3.html)
+- [M4](https://nixtlaverse.nixtla.io/datasetsforecast/m4.html)
+- [M5](https://nixtlaverse.nixtla.io/datasetsforecast/m5.html)
+- [PHM2008](https://nixtlaverse.nixtla.io/datasetsforecast/phm2008.html)
+
+## How to use
+
+All the modules have a `load` method which you can use to load the
+dataset for a specific group. If you don’t have the data locally it will
+be downloaded for you.
+
+```python
+from datasetsforecast.phm2008 import PHM2008
+```
+
+
+```python
+train_df, test_df = PHM2008.load(directory='data', group='FD001')
+train_df.shape, test_df.shape
+```
+
+``` text
+((20631, 17), (13096, 17))
+```
+
diff --git a/datasetsforecast/light.png b/datasetsforecast/light.png
new file mode 100644
index 00000000..bbb99b54
Binary files /dev/null and b/datasetsforecast/light.png differ
diff --git a/datasetsforecast/long_horizon.html.mdx b/datasetsforecast/long_horizon.html.mdx
new file mode 100644
index 00000000..02c8028b
--- /dev/null
+++ b/datasetsforecast/long_horizon.html.mdx
@@ -0,0 +1,218 @@
+---
+description: Download and wrangling utility for long-horizon datasets.
+output-file: long_horizon.html
+title: Long-Horizon Datasets
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### ETTm2
+
+> ``` text
+> ETTm2 (freq:str='15T', name:str='ETTm2', n_ts:int=7, test_size:int=11520,
+> val_size:int=11520, horizons:Tuple[int]=(96, 192, 336, 720))
+> ```
+
+The ETTm2 dataset monitors an electricity transformer from a region of
+a province of China including oil temperature and variants of load (such
+as high useful load and high useless load) from July 2016 to July 2018
+at a fifteen minute frequency.
+
+Reference:
+- [Zhou, et al. Informer: Beyond Efficient Transformer for Long Sequence Time-Series Forecasting. AAAI 2021.](https://arxiv.org/abs/2012.07436)
+
+------------------------------------------------------------------------
+
+source
+
+### ETTm1
+
+> ``` text
+> ETTm1 (freq:str='15T', name:str='ETTm1', n_ts:int=7, test_size:int=11520,
+> val_size:int=11520, horizons:Tuple[int]=(96, 192, 336, 720))
+> ```
+
+*The ETTm1 dataset monitors an electricity transformer from a region of
+a province of China including oil temperature and variants of load (such
+as high useful load and high useless load) from July 2016 to July 2018
+at a fifteen minute frequency.*
+
+------------------------------------------------------------------------
+
+source
+
+### ETTh2
+
+> ``` text
+> ETTh2 (freq:str='H', name:str='ETTh2', n_ts:int=1, test_size:int=11520,
+> val_size:int=11520, horizons:Tuple[int]=(96, 192, 336, 720))
+> ```
+
+*The ETTh2 dataset monitors an electricity transformer from a region of
+a province of China including oil temperature and variants of load (such
+as high useful load and high useless load) from July 2016 to July 2018
+at an hourly frequency.*
+
+------------------------------------------------------------------------
+
+source
+
+### ETTh1
+
+> ``` text
+> ETTh1 (freq:str='H', name:str='ETTh1', n_ts:int=1, test_size:int=11520,
+> val_size:int=11520, horizons:Tuple[int]=(96, 192, 336, 720))
+> ```
+
+*The ETTh1 dataset monitors an electricity transformer from a region of
+a province of China including oil temperature and variants of load (such
+as high useful load and high useless load) from July 2016 to July 2018
+at an hourly frequency.*
+
+------------------------------------------------------------------------
+
+source
+
+### ECL
+
+> ``` text
+> ECL (freq:str='15T', name:str='ECL', n_ts:int=321, test_size:int=5260,
+> val_size:int=2632, horizons:Tuple[int]=(96, 192, 336, 720))
+> ```
+
+The Electricity dataset reports the fifteen minute electricity
+consumption (KWh) of 321 customers from 2012 to 2014. For comparability,
+we aggregate it hourly.
+
+Reference:
+
+- [Li, S et al. Enhancing the locality and breaking the memory bottleneck of Transformer on time series forecasting. NeurIPS 2019.](http://arxiv.org/abs/1907.00235)
+
+
+
+------------------------------------------------------------------------
+
+source
+
+### Exchange
+
+> ``` text
+> Exchange (freq:str='D', name:str='Exchange', n_ts:int=8,
+> test_size:int=1517, val_size:int=760, horizons:Tuple[int]=(96,
+> 192, 336, 720))
+> ```
+
+The Exchange dataset is a collection of daily exchange rates of eight
+countries relative to the US dollar. The countries include Australia,
+UK, Canada, Switzerland, China, Japan, New Zealand and Singapore from
+1990 to 2016.
+
+Reference:
+
+- [Lai, G., Chang, W., Yang, Y., and Liu, H. Modeling Long and Short-Term Temporal Patterns with Deep Neural Networks. SIGIR 2018.](http://arxiv.org/abs/1703.07015)
+
+------------------------------------------------------------------------
+
+source
+
+### TrafficL
+
+> ``` text
+> TrafficL (freq:str='H', name:str='traffic', n_ts:int=862,
+> test_size:int=3508, val_size:int=1756, horizons:Tuple[int]=(96,
+> 192, 336, 720))
+> ```
+
+This large Traffic dataset was collected by the California Department
+of Transportation, it reports road hourly occupancy rates of 862
+sensors, from January 2015 to December 2016.
+
+Reference:
+
+- [Lai, G., Chang, W., Yang, Y., and Liu, H. Modeling Long and Short-Term Temporal Patterns with Deep Neural Networks. SIGIR 2018.](http://arxiv.org/abs/1703.07015)
+- [Wu, H., Xu, J., Wang, J., and Long, M. Autoformer: Decomposition Transformers with auto-correlation for long-term series forecasting. NeurIPS 2021](https://arxiv.org/abs/2106.13008)
+
+------------------------------------------------------------------------
+
+source
+
+### ILI
+
+> ``` text
+> ILI (freq:str='W', name:str='ili', n_ts:int=7, test_size:int=193,
+> val_size:int=97, horizons:Tuple[int]=(24, 36, 48, 60))
+> ```
+
+This dataset reports weekly recorded influenza-like illness (ILI)
+patients from Centers for Disease Control and Prevention of the United
+States from 2002 to 2021. It is measured as a ratio of ILI patients
+versus the total patients in the week.
+
+Reference:
+
+- [Wu, H., Xu, J., Wang, J., and Long, M. Autoformer: Decomposition Transformers with auto-correlation for long-term series forecasting. NeurIPS 2021.](https://arxiv.org/abs/2106.13008)
+
+------------------------------------------------------------------------
+
+source
+
+### Weather
+
+> ``` text
+> Weather (freq:str='10M', name:str='weather', n_ts:int=21,
+> test_size:int=10539, val_size:int=5270, horizons:Tuple[int]=(96,
+> 192, 336, 720))
+> ```
+
+This Weather dataset contains the 2020 year of 21 meteorological
+measurements recorded every 10 minutes from the Weather Station of the
+Max Planck Biogeochemistry Institute in Jena, Germany.
+
+Reference:
+
+- [Wu, H., Xu, J., Wang, J., and Long, M. Autoformer: Decomposition Transformers with auto-correlation for long-term series forecasting. NeurIPS 2021. ](https://arxiv.org/abs/2106.13008)
+
+------------------------------------------------------------------------
+
+source
+
+### LongHorizon
+
+> ``` text
+> LongHorizon (source_url:str='https://nhits-
+> experiments.s3.amazonaws.com/datasets.zip')
+> ```
+
+This Long-Horizon datasets wrapper class, provides with utility to
+download and wrangle the following datasets:
+ETT, ECL, Exchange, Traffic, ILI and Weather.
+
+- Each set is normalized with the train data mean and standard
+ deviation.
+- Datasets are partitioned into train, validation and test splits.
+- For all datasets: 70%, 10%, and 20% of observations are train,
+ validation, test, except ETT that uses 20% validation.
+
diff --git a/datasetsforecast/long_horizon2.html.mdx b/datasetsforecast/long_horizon2.html.mdx
new file mode 100644
index 00000000..b57b20d7
--- /dev/null
+++ b/datasetsforecast/long_horizon2.html.mdx
@@ -0,0 +1,185 @@
+---
+description: >-
+ Download and wrangling utility for long-horizon datasets. These datasets have
+ been used by `NHITS, AutoFormer, Informer, PatchTST, TiDE` among many other
+ neural forecasting methods. The datasets include the original [ETTh1, ETTh2,
+ ETTm1, ETTm2, Weather, ILI, TrafficL](https://github.com/zhouhaoyi/ETDataset)
+ benchmark datasets.
+output-file: long_horizon2.html
+title: Long-Horizon Original Datasets
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### Weather
+
+> ``` text
+> Weather (freq:str='10M', name:str='weather', n_ts:int=21,
+> test_size:int=10539, val_size:int=5270, horizons:Tuple[int]=(96,
+> 192, 336, 720))
+> ```
+
+\*This Weather dataset contains the 2020 year of 21 meteorological
+measurements recorded every 10 minutes from the Weather Station of the
+Max Planck Biogeochemistry Institute in Jena, Germany.
+
+Reference: Wu, H., Xu, J., Wang, J., and Long, M. Autoformer:
+Decomposition Transformers with auto-correlation for long-term series
+forecasting. NeurIPS 2021. https://arxiv.org/abs/2106.13008.\*
+
+------------------------------------------------------------------------
+
+source
+
+### TrafficL
+
+> ``` text
+> TrafficL (freq:str='H', name:str='traffic', n_ts:int=862,
+> test_size:int=3508, val_size:int=1756, horizons:Tuple[int]=(96,
+> 192, 336, 720))
+> ```
+
+\*This large Traffic dataset was collected by the California Department
+of Transportation, it reports road hourly occupancy rates of 862
+sensors, from January 2015 to December 2016.
+
+Reference: Lai, G., Chang, W., Yang, Y., and Liu, H. Modeling Long and
+Short-Term Temporal Patterns with Deep Neural Networks. SIGIR 2018.
+http://arxiv.org/abs/1703.07015.
+
+Wu, H., Xu, J., Wang, J., and Long, M. Autoformer: Decomposition
+Transformers with auto-correlation for long-term series forecasting.
+NeurIPS 2021. https://arxiv.org/abs/2106.13008.\*
+
+------------------------------------------------------------------------
+
+source
+
+### ECL
+
+> ``` text
+> ECL (freq:str='15T', name:str='ECL', n_ts:int=321, n_time:int=26304,
+> test_size:int=5260, val_size:int=2632, horizons:Tuple[int]=(96, 192,
+> 336, 720))
+> ```
+
+\*The Electricity dataset reports the fifteen minute electricity
+consumption (KWh) of 321 customers from 2012 to 2014. For comparability,
+we aggregate it hourly.
+
+Reference: Li, S et al. Enhancing the locality and breaking the memory
+bottleneck of Transformer on time series forecasting. NeurIPS 2019.
+http://arxiv.org/abs/1907.00235.\*
+
+------------------------------------------------------------------------
+
+source
+
+### ETTm2
+
+> ``` text
+> ETTm2 (freq:str='15T', name:str='ETTm2', n_ts:int=7, n_time:int=57600,
+> test_size:int=11520, val_size:int=11520, horizons:Tuple[int]=(96,
+> 192, 336, 720))
+> ```
+
+\*The ETTm2 dataset monitors an electricity transformer from a region of
+a province of China including oil temperature and variants of load (such
+as high useful load and high useless load) from July 2016 to July 2018
+at a fifteen minute frequency.
+
+Reference: Zhou, et al. Informer: Beyond Efficient Transformer for Long
+Sequence Time-Series Forecasting. AAAI 2021.
+https://arxiv.org/abs/2012.07436\*
+
+------------------------------------------------------------------------
+
+source
+
+### ETTm1
+
+> ``` text
+> ETTm1 (freq:str='15T', name:str='ETTm1', n_ts:int=7, n_time:int=57600,
+> test_size:int=11520, val_size:int=11520, horizons:Tuple[int]=(96,
+> 192, 336, 720))
+> ```
+
+*The ETTm1 dataset monitors an electricity transformer from a region of
+a province of China including oil temperature and variants of load (such
+as high useful load and high useless load) from July 2016 to July 2018
+at a fifteen minute frequency.*
+
+------------------------------------------------------------------------
+
+source
+
+### ETTh2
+
+> ``` text
+> ETTh2 (freq:str='H', name:str='ETTh2', n_ts:int=7, n_time:int=14400,
+> test_size:int=2880, val_size:int=2880, horizons:Tuple[int]=(96,
+> 192, 336, 720))
+> ```
+
+*The ETTh2 dataset monitors an electricity transformer from a region of
+a province of China including oil temperature and variants of load (such
+as high useful load and high useless load) from July 2016 to July 2018
+at an hourly frequency.*
+
+------------------------------------------------------------------------
+
+source
+
+### ETTh1
+
+> ``` text
+> ETTh1 (freq:str='H', name:str='ETTh1', n_ts:int=7, n_time:int=14400,
+> test_size:int=2880, val_size:int=2880, horizons:Tuple[int]=(96,
+> 192, 336, 720))
+> ```
+
+*The ETTh1 dataset monitors an electricity transformer from a region of
+a province of China including oil temperature and variants of load (such
+as high useful load and high useless load) from July 2016 to July 2018
+at an hourly frequency.*
+
+------------------------------------------------------------------------
+
+source
+
+### LongHorizon2
+
+> ``` text
+> LongHorizon2 (source_url:str='https://www.dropbox.com/s/rlc1qmprpvuqrsv/a
+> ll_six_datasets.zip?dl=1')
+> ```
+
+\*This Long-Horizon datasets wrapper class, provides with utility to
+download and wrangle the following datasets:
+ETT, ECL, Exchange, Traffic, ILI and Weather.
+
+- Each set is normalized with the train data mean and standard
+ deviation.
+- Datasets are partitioned into train, validation and test splits.
+- For all datasets: 70%, 10%, and 20% of observations are train,
+ validation, test, except ETT that uses 20% validation.\*
+
diff --git a/datasetsforecast/m3.html.mdx b/datasetsforecast/m3.html.mdx
new file mode 100644
index 00000000..30acbe07
--- /dev/null
+++ b/datasetsforecast/m3.html.mdx
@@ -0,0 +1,73 @@
+---
+description: Download the M3 dataset.
+output-file: m3.html
+title: M3 dataset
+---
+
+
+> M3 meta information
+
+------------------------------------------------------------------------
+
+source
+
+### Other
+
+> ``` text
+> Other (seasonality:int=1, horizon:int=8, freq:str='D',
+> sheet_name:str='M3Other', name:str='Other', n_ts:int=174)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### Monthly
+
+> ``` text
+> Monthly (seasonality:int=12, horizon:int=18, freq:str='M',
+> sheet_name:str='M3Month', name:str='Monthly', n_ts:int=1428)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### Quarterly
+
+> ``` text
+> Quarterly (seasonality:int=4, horizon:int=8, freq:str='Q',
+> sheet_name:str='M3Quart', name:str='Quarterly', n_ts:int=756)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### Yearly
+
+> ``` text
+> Yearly (seasonality:int=1, horizon:int=6, freq:str='Y',
+> sheet_name:str='M3Year', name:str='Yearly', n_ts:int=645)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### M3
+
+> ``` text
+> M3 ()
+> ```
+
diff --git a/datasetsforecast/m4.html.mdx b/datasetsforecast/m4.html.mdx
new file mode 100644
index 00000000..bf3881da
--- /dev/null
+++ b/datasetsforecast/m4.html.mdx
@@ -0,0 +1,184 @@
+---
+description: Download and evaluate the M4 dataset.
+output-file: m4.html
+title: M4 dataset
+---
+
+
+> M4 meta information
+
+------------------------------------------------------------------------
+
+source
+
+### Other
+
+> ``` text
+> Other (seasonality:int=1, horizon:int=8, freq:str='D', name:str='Other',
+> n_ts:int=5000, included_groups:Tuple=('Weekly', 'Daily',
+> 'Hourly'))
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### Hourly
+
+> ``` text
+> Hourly (seasonality:int=24, horizon:int=48, freq:str='H',
+> name:str='Hourly', n_ts:int=414)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### Daily
+
+> ``` text
+> Daily (seasonality:int=1, horizon:int=14, freq:str='D', name:str='Daily',
+> n_ts:int=4227)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### Weekly
+
+> ``` text
+> Weekly (seasonality:int=1, horizon:int=13, freq:str='W',
+> name:str='Weekly', n_ts:int=359)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### Monthly
+
+> ``` text
+> Monthly (seasonality:int=12, horizon:int=18, freq:str='M',
+> name:str='Monthly', n_ts:int=48000)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### Quarterly
+
+> ``` text
+> Quarterly (seasonality:int=4, horizon:int=8, freq:str='Q',
+> name:str='Quarterly', n_ts:int=24000)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### Yearly
+
+> ``` text
+> Yearly (seasonality:int=1, horizon:int=6, freq:str='Y',
+> name:str='Yearly', n_ts:int=23000)
+> ```
+
+## Download data class
+
+------------------------------------------------------------------------
+
+source
+
+### M4
+
+> ``` text
+> M4 (source_url:str='https://raw.githubusercontent.com/Mcompetitions/M4-
+> methods/master/Dataset/', naive2_forecast_url:str='https://github.com
+> /Nixtla/m4-forecasts/raw/master/forecasts/submission-Naive2.zip')
+> ```
+
+```python
+group = 'Hourly'
+await M4.async_download('data', group=group)
+df, *_ = M4.load(directory='data', group=group)
+n_series = len(np.unique(df.unique_id.values))
+display_str = f'Group: {group} '
+display_str += f'n_series: {n_series}'
+print(display_str)
+```
+
+## Evaluation class
+
+------------------------------------------------------------------------
+
+source
+
+### M4Evaluation
+
+> ``` text
+> M4Evaluation ()
+> ```
+
+*Initialize self. See help(type(self)) for accurate signature.*
+
+### URL-based evaluation
+
+The method `evaluate` from the class
+[`M4Evaluation`](https://Nixtla.github.io/datasetsforecast/m4.html#m4evaluation)
+can receive a url of a [benchmark uploaded to the M4
+competiton](https://github.com/Mcompetitions/M4-methods/tree/master/Point%20Forecasts).
+
+The results compared to the on-the-fly evaluation were obtained from the
+[official
+evaluation](https://github.com/Mcompetitions/M4-methods/blob/master/Evaluation%20and%20Ranks.xlsx).
+
+```python
+from fastcore.test import test_close
+```
+
+
+```python
+esrnn_url = 'https://github.com/Nixtla/m4-forecasts/raw/master/forecasts/submission-118.zip'
+esrnn_evaluation = M4Evaluation.evaluate('data', 'Hourly', esrnn_url)
+# Test of the same evaluation as the original one
+test_close(esrnn_evaluation['SMAPE'].item(), 9.328, eps=1e-3)
+test_close(esrnn_evaluation['MASE'].item(), 0.893, eps=1e-3)
+test_close(esrnn_evaluation['OWA'].item(), 0.440, eps=1e-3)
+esrnn_evaluation
+```
+
+### Numpy-based evaluation
+
+Also the method `evaluate` can recevie a numpy array of forecasts.
+
+```python
+fforma_url = 'https://github.com/Nixtla/m4-forecasts/raw/master/forecasts/submission-245.zip'
+fforma_forecasts = M4Evaluation.load_benchmark('data', 'Hourly', fforma_url)
+fforma_evaluation = M4Evaluation.evaluate('data', 'Hourly', fforma_forecasts)
+# Test of the same evaluation as the original one
+test_close(fforma_evaluation['SMAPE'].item(), 11.506, eps=1e-3)
+test_close(fforma_evaluation['MASE'].item(), 0.819, eps=1e-3)
+test_close(fforma_evaluation['OWA'].item(), 0.484, eps=1e-3)
+fforma_evaluation
+```
+
diff --git a/datasetsforecast/m5.html.mdx b/datasetsforecast/m5.html.mdx
new file mode 100644
index 00000000..04f5506c
--- /dev/null
+++ b/datasetsforecast/m5.html.mdx
@@ -0,0 +1,114 @@
+---
+description: Download and evaluate the M5 dataset.
+output-file: m5.html
+title: M5 dataset
+---
+
+
+## Download data class
+
+------------------------------------------------------------------------
+
+source
+
+### M5
+
+> ``` text
+> M5 (source_url:str='https://github.com/Nixtla/m5-
+> forecasts/raw/main/datasets/m5.zip')
+> ```
+
+## Test number of series
+
+## Evaluation class
+
+------------------------------------------------------------------------
+
+source
+
+### M5Evaluation
+
+> ``` text
+> M5Evaluation ()
+> ```
+
+*Initialize self. See help(type(self)) for accurate signature.*
+
+### URL-based evaluation
+
+The method `evaluate` from the class
+[`M5Evaluation`](https://Nixtla.github.io/datasetsforecast/m5.html#m5evaluation)
+can receive a url of a [submission to the M5
+competiton](https://github.com/Nixtla/m5-forecasts/tree/main/forecasts).
+
+The results compared to the on-the-fly evaluation were obtained from the
+[official
+evaluation](https://github.com/Mcompetitions/M5-methods/blob/master/Scores%20and%20Ranks.xlsx).
+
+```python
+m5_winner_url = 'https://github.com/Nixtla/m5-forecasts/raw/main/forecasts/0001 YJ_STU.zip'
+winner_evaluation = M5Evaluation.evaluate('data', m5_winner_url)
+# Test of the same evaluation as the original one
+test_close(winner_evaluation.loc['Total'].item(), 0.520, eps=1e-3)
+winner_evaluation
+```
+
+### Pandas-based evaluation
+
+Also the method `evaluate` can recevie a pandas DataFrame of forecasts.
+
+```python
+m5_second_place_url = 'https://github.com/Nixtla/m5-forecasts/raw/main/forecasts/0002 Matthias.zip'
+m5_second_place_forecasts = M5Evaluation.load_benchmark('data', m5_second_place_url)
+second_place_evaluation = M5Evaluation.evaluate('data', m5_second_place_forecasts)
+# Test of the same evaluation as the original one
+test_close(second_place_evaluation.loc['Total'].item(), 0.528, eps=1e-3)
+second_place_evaluation
+```
+
+By default you can load the winner benchmark using the following.
+
+```python
+winner_benchmark = M5Evaluation.load_benchmark('data')
+winner_evaluation = M5Evaluation.evaluate('data', winner_benchmark)
+# Test of the same evaluation as the original one
+test_close(winner_evaluation.loc['Total'].item(), 0.520, eps=1e-3)
+winner_evaluation
+```
+
+### Validation evaluation
+
+You can also evaluate the official validation set.
+
+```python
+winner_benchmark_val = M5Evaluation.load_benchmark('data', validation=True)
+winner_evaluation_val = M5Evaluation.evaluate('data', winner_benchmark_val, validation=True)
+winner_evaluation_val
+```
+
+## Kaggle-Competition-M5 References
+
+The evaluation metric of the Favorita Kaggle competition was the
+normalized weighted root mean squared logarithmic error (NWRMSLE).
+Perishable items have a score weight of 1.25; otherwise, the weight is
+1.0.
+
+$$ NWRMSLE = \sqrt{\frac{\sum^{n}_{i=1} w_{i}\left(log(\hat{y}_{i}+1) - log(y_{i}+1)\right)^{2}}{\sum^{n}_{i=1} w_{i}}}$$
+
+| Kaggle Competition Forecasting Methods | 16D ahead NWRMSLE |
+|:---------------------------------------------------------------:|:-----:|
+| [LGBM](https://www.kaggle.com/shixw125/1st-place-lgb-model-public-0-506-private-0-511/comments) \[1\] | 0.5091 |
+| [Seq2Seq WaveNet](https://arxiv.org/abs/1803.04037) \[2\] | 0.5129 |
+
+1. [Corporación Favorita. Corporación favorita grocery sales
+ forecasting. Kaggle Competition Leaderboard,
+ 2018.](https://www.kaggle.com/c/favorita-grocery-sales-forecasting/leaderboard)
+2. [Glib Kechyn, Lucius Yu, Yangguang Zang, and Svyatoslav Kechyn.
+ Sales forecasting using wavenet within the framework of the Favorita
+ Kaggle competition. Computing Research Repository, abs/1803.04037,
+ 2018](https://arxiv.org/abs/1803.04037).
+
diff --git a/datasetsforecast/mint.json b/datasetsforecast/mint.json
new file mode 100644
index 00000000..e0a88151
--- /dev/null
+++ b/datasetsforecast/mint.json
@@ -0,0 +1,42 @@
+{
+ "$schema": "https://mintlify.com/schema.json",
+ "name": "Nixtla",
+ "logo": {
+ "light": "/light.png",
+ "dark": "/dark.png"
+ },
+ "favicon": "/favicon.svg",
+ "colors": {
+ "primary": "#0E0E0E",
+ "light": "#FAFAFA",
+ "dark": "#0E0E0E",
+ "anchors": {
+ "from": "#2AD0CA",
+ "to": "#0E00F8"
+ }
+ },
+ "topbarCtaButton": {
+ "type": "github",
+ "url": "https://github.com/Nixtla/datasetsforecast"
+ },
+ "navigation": [
+ {
+ "group": "",
+ "pages": ["index.html"]
+ },
+ {
+ "group": "Datasets",
+ "pages": [
+ "favorita.html",
+ "hierarchical.html",
+ "long_horizon.html",
+ "long_horizon2.html",
+ "m3.html",
+ "m4.html",
+ "m5.html",
+ "phm2008.html",
+ "utils.html"
+ ]
+ }
+ ]
+}
diff --git a/datasetsforecast/phm2008.html.mdx b/datasetsforecast/phm2008.html.mdx
new file mode 100644
index 00000000..e11c249e
--- /dev/null
+++ b/datasetsforecast/phm2008.html.mdx
@@ -0,0 +1,75 @@
+---
+description: Download the PHM2008 dataset.
+output-file: phm2008.html
+title: PHM2008 dataset
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### FD004
+
+> ``` text
+> FD004 (seasonality:int=1, horizon:int=8, freq:str='None',
+> train_file:str='train_FD004.txt', test_file:str='test_FD004.txt',
+> rul_file:str='RUL_FD004.txt', n_ts:int=249, n_test:int=248)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### FD003
+
+> ``` text
+> FD003 (seasonality:int=1, horizon:int=1, freq:str='None',
+> train_file:str='train_FD003.txt', test_file:str='test_FD003.txt',
+> rul_file:str='RUL_FD003.txt', n_ts:int=100, n_test:int=100)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### FD002
+
+> ``` text
+> FD002 (seasonality:int=1, horizon:int=1, freq:str='None',
+> train_file:str='train_FD002.txt', test_file:str='test_FD002.txt',
+> rul_file:str='RUL_FD002.txt', n_ts:int=260, n_test:int=259)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### FD001
+
+> ``` text
+> FD001 (seasonality:int=1, horizon:int=1, freq:str='None',
+> train_file:str='train_FD001.txt', test_file:str='test_FD001.txt',
+> rul_file:str='RUL_FD001.txt', n_ts:int=100, n_test:int=100)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### PHM2008
+
+> ``` text
+> PHM2008 ()
+> ```
+
diff --git a/datasetsforecast/utils.html.mdx b/datasetsforecast/utils.html.mdx
new file mode 100644
index 00000000..187843b6
--- /dev/null
+++ b/datasetsforecast/utils.html.mdx
@@ -0,0 +1,113 @@
+---
+output-file: utils.html
+title: Datasets Utils
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### download_file
+
+> ``` text
+> download_file (directory:str, source_url:str, decompress:bool=False)
+> ```
+
+*Download data from source_ulr inside directory.*
+
+------------------------------------------------------------------------
+
+source
+
+### extract_file
+
+> ``` text
+> extract_file (filepath, directory)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### async_download_files
+
+> ``` text
+> async_download_files (path:Union[str,pathlib.Path], urls:Iterable[str])
+> ```
+
+```python
+import os
+import tempfile
+
+import requests
+```
+
+
+```python
+gh_url = 'https://api.github.com/repos/Nixtla/datasetsforecast/contents/'
+base_url = 'https://raw.githubusercontent.com/Nixtla/datasetsforecast/main'
+
+headers = {}
+gh_token = os.getenv('GITHUB_TOKEN')
+if gh_token is not None:
+ headers = {'Authorization': f'Bearer: {gh_token}'}
+resp = requests.get(gh_url, headers=headers)
+if resp.status_code != 200:
+ raise Exception(resp.text)
+urls = [f'{base_url}/{e["path"]}' for e in resp.json() if e['type'] == 'file']
+with tempfile.TemporaryDirectory() as tmp:
+ tmp = Path(tmp)
+ await async_download_files(tmp, urls)
+ files = list(tmp.iterdir())
+ assert len(files) == len(urls)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### download_files
+
+> ``` text
+> download_files (directory:Union[str,pathlib.Path], urls:Iterable[str])
+> ```
+
+```python
+with tempfile.TemporaryDirectory() as tmp:
+ tmp = Path(tmp)
+ fname = tmp / 'script.py'
+ fname.write_text(f"""
+from datasetsforecast.utils import download_files
+
+download_files('{tmp.as_posix()}', {urls})
+ """)
+ !python {fname}
+ fname.unlink()
+ files = list(tmp.iterdir())
+ assert len(files) == len(urls)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### Info
+
+> ``` text
+> Info (class_groups:Tuple[dataclass])
+> ```
+
+*Info Dataclass of datasets. Args: groups (Tuple): Tuple of str groups
+class_groups (Tuple): Tuple of dataclasses.*
+
diff --git a/docs.json b/docs.json
new file mode 100644
index 00000000..83bf38ff
--- /dev/null
+++ b/docs.json
@@ -0,0 +1,564 @@
+{
+ "$schema": "https://mintlify.com/docs.json",
+
+ "banner": {
+ "content": "[Announcement](https://www.nixtla.io/blog/genai-announcement): Nixtla Enterprise now offers top foundation models, MCP, and agentic capabilities: [join the waitlist](https://www.nixtla.io?waitlistSheet=true)",
+ "dismissible": true
+ },
+ "colors": {
+ "primary": "#161616",
+ "light": "#FFF",
+ "dark": "#161616"
+ },
+ "favicon": "/favicon.svg",
+ "footer": {
+ "socials": {
+ "github": "https://github.com/Nixtla",
+ "slack": "https://join.slack.com/t/nixtlaworkspace/shared_invite/zt-135dssye9-fWTzMpv2WBthq8NK0Yvu6A",
+ "twitter": "https://twitter.com/nixtlainc"
+ }
+ },
+ "integrations": {
+ "gtm": {
+ "tagId": "GTM-TBJ64S3X"
+ },
+ "intercom": {
+ "appId": "j7y9c2ep"
+ }
+ },
+ "logo": {
+ "light": "/light.png",
+ "dark": "/dark.png"
+ },
+ "name": "Nixtla",
+ "navbar": {
+ "primary": {
+ "type": "github",
+ "href": "https://github.com/Nixtla"
+ }
+ },
+ "navigation": {
+ "anchors": [
+ {
+ "anchor": "TimeGPT",
+ "icon": "clock-nine",
+ "href": "https://nixtla.io/docs"
+ },
+ {
+ "anchor": "StatsForecast",
+ "icon": "bolt",
+ "groups": [
+ {
+ "group": " ",
+ "pages": ["statsforecast/index.html"]
+ },
+ {
+ "group": "Getting Started",
+ "pages": [
+ "statsforecast/docs/getting-started/installation.html",
+ "statsforecast/docs/getting-started/getting_started_short.html",
+ "statsforecast/docs/getting-started/getting_started_complete.html",
+ "statsforecast/docs/getting-started/getting_started_complete_polars.html"
+ ]
+ },
+ {
+ "group": "Tutorials",
+ "pages": [
+ "statsforecast/docs/tutorials/anomalydetection.html",
+ "statsforecast/docs/tutorials/conformalprediction.html",
+ "statsforecast/docs/tutorials/crossvalidation.html",
+ "statsforecast/docs/tutorials/electricityloadforecasting.html",
+ "statsforecast/docs/tutorials/electricitypeakforecasting.html",
+ "statsforecast/docs/tutorials/garch_tutorial.html",
+ "statsforecast/docs/tutorials/intermittentdata.html",
+ "statsforecast/docs/tutorials/mlflow.html",
+ "statsforecast/docs/tutorials/multipleseasonalities.html",
+ "statsforecast/docs/tutorials/statisticalneuralmethods.html",
+ "statsforecast/docs/tutorials/uncertaintyintervals.html"
+ ]
+ },
+ {
+ "group": "How to Guides",
+ "pages": [
+ "statsforecast/docs/how-to-guides/automatic_forecasting.html",
+ "statsforecast/docs/how-to-guides/exogenous.html",
+ "statsforecast/docs/how-to-guides/generating_features.html",
+ "statsforecast/docs/how-to-guides/sklearn_models.html",
+ "statsforecast/docs/how-to-guides/migrating_R",
+ "statsforecast/docs/how-to-guides/numba_cache.html"
+ ]
+ },
+ {
+ "group": "Distributed",
+ "pages": [
+ "statsforecast/docs/distributed/dask.html",
+ "statsforecast/docs/distributed/ray.html",
+ "statsforecast/docs/distributed/spark.html"
+ ]
+ },
+ {
+ "group": "Experiments",
+ "pages": [
+ "statsforecast/docs/experiments/amazonstatsforecast.html",
+ "statsforecast/docs/experiments/autoarima_vs_prophet.html",
+ "statsforecast/docs/experiments/ets_ray_m5.html",
+ "statsforecast/docs/experiments/prophet_spark_m5.html"
+ ]
+ },
+ {
+ "group": "Model References",
+ "pages": [
+ "statsforecast/docs/models/adida.html",
+ "statsforecast/docs/models/arch.html",
+ "statsforecast/docs/models/arima.html",
+ "statsforecast/docs/models/autoarima.html",
+ "statsforecast/docs/models/autoces.html",
+ "statsforecast/docs/models/autoets.html",
+ "statsforecast/docs/models/autoregressive.html",
+ "statsforecast/docs/models/autotheta.html",
+ "statsforecast/docs/models/crostonclassic.html",
+ "statsforecast/docs/models/crostonoptimized.html",
+ "statsforecast/docs/models/crostonsba.html",
+ "statsforecast/docs/models/dynamicoptimizedtheta.html",
+ "statsforecast/docs/models/dynamicstandardtheta.html",
+ "statsforecast/docs/models/garch.html",
+ "statsforecast/docs/models/holt.html",
+ "statsforecast/docs/models/holtwinters.html",
+ "statsforecast/docs/models/imapa.html",
+ "statsforecast/docs/models/mfles.html",
+ "statsforecast/docs/models/multipleseasonaltrend.html",
+ "statsforecast/docs/models/optimizedtheta.html",
+ "statsforecast/docs/models/seasonalexponentialsmoothing.html",
+ "statsforecast/docs/models/seasonalexponentialsmoothingoptimized.html",
+ "statsforecast/docs/models/simpleexponentialoptimized.html",
+ "statsforecast/docs/models/simpleexponentialsmoothing.html",
+ "statsforecast/docs/models/standardtheta.html",
+ "statsforecast/docs/models/tsb.html"
+ ]
+ },
+ {
+ "group": "API Reference",
+ "pages": [
+ "statsforecast/src/core/core.html",
+ "statsforecast/src/core/distributed.fugue.html",
+ "statsforecast/src/core/models.html",
+ "statsforecast/src/core/models_intro",
+ "statsforecast/src/feature_engineering.html"
+ ]
+ },
+ {
+ "group": "Contributing",
+ "pages": [
+ "statsforecast/docs/contribute/contribute",
+ "statsforecast/docs/contribute/docs",
+ "statsforecast/docs/contribute/issue-labels",
+ "statsforecast/docs/contribute/issues",
+ "statsforecast/docs/contribute/step-by-step",
+ "statsforecast/docs/contribute/techstack"
+ ]
+ }
+ ]
+ },
+ {
+ "anchor": "MLForecast",
+ "icon": "robot",
+ "groups": [
+ {
+ "group": " ",
+ "pages": ["mlforecast/index.html"]
+ },
+ {
+ "group": "Getting Started",
+ "pages": [
+ "mlforecast/docs/getting-started/install.html",
+ "mlforecast/docs/getting-started/quick_start_local.html",
+ "mlforecast/docs/getting-started/quick_start_distributed.html",
+ "mlforecast/docs/getting-started/end_to_end_walkthrough.html"
+ ]
+ },
+ {
+ "group": "How-to guides",
+ "pages": [
+ "mlforecast/docs/how-to-guides/exogenous_features.html",
+ "mlforecast/docs/how-to-guides/lag_transforms_guide.html",
+ "mlforecast/docs/how-to-guides/hyperparameter_optimization.html",
+ "mlforecast/docs/how-to-guides/sklearn_pipelines.html",
+ "mlforecast/docs/how-to-guides/sample_weights.html",
+ "mlforecast/docs/how-to-guides/cross_validation.html",
+ "mlforecast/docs/how-to-guides/prediction_intervals.html",
+ "mlforecast/docs/how-to-guides/target_transforms_guide.html",
+ "mlforecast/docs/how-to-guides/analyzing_models.html",
+ "mlforecast/docs/how-to-guides/mlflow.html",
+ "mlforecast/docs/how-to-guides/transforming_exog.html",
+ "mlforecast/docs/how-to-guides/custom_training.html",
+ "mlforecast/docs/how-to-guides/training_with_numpy.html",
+ "mlforecast/docs/how-to-guides/one_model_per_horizon.html",
+ "mlforecast/docs/how-to-guides/custom_date_features.html",
+ "mlforecast/docs/how-to-guides/predict_callbacks.html",
+ "mlforecast/docs/how-to-guides/predict_subset.html",
+ "mlforecast/docs/how-to-guides/transfer_learning.html"
+ ]
+ },
+ {
+ "group": "Tutorials",
+ "pages": [
+ "mlforecast/docs/tutorials/electricity_load_forecasting.html",
+ "mlforecast/docs/tutorials/electricity_peak_forecasting.html",
+ "mlforecast/docs/tutorials/prediction_intervals_in_forecasting_models.html"
+ ]
+ },
+ {
+ "group": "API Reference",
+ "pages": [
+ {
+ "group": "Local",
+ "pages": [
+ "mlforecast/forecast.html",
+ "mlforecast/auto.html",
+ "mlforecast/lgb_cv.html",
+ "mlforecast/optimization.html",
+ "mlforecast/utils.html",
+ "mlforecast/core.html",
+ "mlforecast/target_transforms.html",
+ "mlforecast/lag_transforms.html",
+ "mlforecast/feature_engineering.html",
+ "mlforecast/callbacks.html"
+ ]
+ },
+ {
+ "group": "Distributed",
+ "pages": [
+ "mlforecast/distributed.forecast.html",
+ {
+ "group": "Models",
+ "pages": [
+ "mlforecast/distributed.models.dask.lgb.html",
+ "mlforecast/distributed.models.dask.xgb.html",
+ "mlforecast/distributed.models.ray.lgb.html",
+ "mlforecast/distributed.models.ray.xgb.html",
+ "mlforecast/distributed.models.spark.lgb.html",
+ "mlforecast/distributed.models.spark.xgb.html"
+ ]
+ }
+ ]
+ }
+ ]
+ }
+ ]
+ },
+ {
+ "anchor": "NeuralForecast",
+ "icon": "brain-circuit",
+ "groups": [
+ {
+ "group": "Getting Started",
+ "pages": [
+ "neuralforecast/docs/getting-started/introduction.html",
+ "neuralforecast/docs/getting-started/quickstart.html",
+ "neuralforecast/docs/getting-started/installation.html",
+ "neuralforecast/docs/getting-started/datarequirements.html"
+ ]
+ },
+ {
+ "group": "Capabilities",
+ "pages": [
+ "neuralforecast/docs/capabilities/overview.html",
+ "neuralforecast/docs/capabilities/objectives.html",
+ "neuralforecast/docs/capabilities/exogenous_variables.html",
+ "neuralforecast/docs/capabilities/cross_validation.html",
+ "neuralforecast/docs/capabilities/hyperparameter_tuning.html",
+ "neuralforecast/docs/capabilities/predictinsample.html",
+ "neuralforecast/docs/capabilities/save_load_models.html",
+ "neuralforecast/docs/capabilities/time_series_scaling.html"
+ ]
+ },
+ {
+ "group": "Tutorials",
+ "pages": [
+ {
+ "group": "Forecasting",
+ "pages": [
+ "neuralforecast/docs/tutorials/getting_started_complete.html",
+ "neuralforecast/docs/tutorials/cross_validation.html",
+ "neuralforecast/docs/tutorials/longhorizon_nhits.html",
+ "neuralforecast/docs/tutorials/longhorizon_transformers.html",
+ "neuralforecast/docs/tutorials/forecasting_tft.html",
+ "neuralforecast/docs/tutorials/multivariate_tsmixer.html"
+ ]
+ },
+ {
+ "group": "Probabilistic Forecasting",
+ "pages": [
+ "neuralforecast/docs/tutorials/uncertainty_quantification.html",
+ "neuralforecast/docs/tutorials/longhorizon_probabilistic.html",
+ "neuralforecast/docs/tutorials/conformal_prediction.html"
+ ]
+ },
+ {
+ "group": "Special Topics",
+ "pages": [
+ "neuralforecast/docs/tutorials/explainability",
+ "neuralforecast/docs/tutorials/hierarchical_forecasting.html",
+ "neuralforecast/docs/tutorials/distributed_neuralforecast.html",
+ "neuralforecast/docs/tutorials/intermittent_data.html",
+ "neuralforecast/docs/tutorials/using_mlflow.html",
+ "neuralforecast/docs/tutorials/robust_forecasting.html",
+ "neuralforecast/docs/tutorials/interpretable_decompositions.html",
+ "neuralforecast/docs/tutorials/comparing_methods.html",
+ "neuralforecast/docs/tutorials/temporal_classification.html",
+ "neuralforecast/docs/tutorials/transfer_learning.html",
+ "neuralforecast/docs/tutorials/adding_models.html",
+ "neuralforecast/docs/tutorials/large_datasets.html"
+ ]
+ }
+ ]
+ },
+ {
+ "group": "Use cases",
+ "pages": [
+ "neuralforecast/docs/use-cases/electricity_peak_forecasting.html",
+ "neuralforecast/docs/use-cases/predictive_maintenance.html"
+ ]
+ },
+ {
+ "group": "API Reference",
+ "pages": [
+ "neuralforecast/core.html",
+ {
+ "group": "Models",
+ "pages": [
+ "neuralforecast/models.autoformer.html",
+ "neuralforecast/models.bitcn.html",
+ "neuralforecast/models.deepar.html",
+ "neuralforecast/models.deepnpts.html",
+ "neuralforecast/models.dilated_rnn.html",
+ "neuralforecast/models.dlinear.html",
+ "neuralforecast/models.fedformer.html",
+ "neuralforecast/models.gru.html",
+ "neuralforecast/models.hint.html",
+ "neuralforecast/models.informer.html",
+ "neuralforecast/models.itransformer.html",
+ "neuralforecast/models.kan.html",
+ "neuralforecast/models.lstm.html",
+ "neuralforecast/models.mlp.html",
+ "neuralforecast/models.mlpmultivariate.html",
+ "neuralforecast/models.nbeats.html",
+ "neuralforecast/models.nbeatsx.html",
+ "neuralforecast/models.nhits.html",
+ "neuralforecast/models.nlinear.html",
+ "neuralforecast/models.patchtst.html",
+ "neuralforecast/models.rmok.html",
+ "neuralforecast/models.rnn.html",
+ "neuralforecast/models.softs.html",
+ "neuralforecast/models.stemgnn.html",
+ "neuralforecast/models.tcn.html",
+ "neuralforecast/models.tft.html",
+ "neuralforecast/models.tide.html",
+ "neuralforecast/models.timellm.html",
+ "neuralforecast/models.timemixer.html",
+ "neuralforecast/models.timesnet.html",
+ "neuralforecast/models.timexer.html",
+ "neuralforecast/models.tsmixer.html",
+ "neuralforecast/models.tsmixerx.html",
+ "neuralforecast/models.vanillatransformer.html",
+ "neuralforecast/models.xlstm"
+ ]
+ },
+ "neuralforecast/models.html",
+ {
+ "group": "Train/Evaluation",
+ "pages": [
+ "neuralforecast/losses.pytorch.html",
+ "neuralforecast/losses.numpy.html"
+ ]
+ },
+ {
+ "group": "Common Components",
+ "pages": [
+ "neuralforecast/common.base_auto.html",
+ "neuralforecast/common.scalers.html",
+ "neuralforecast/common.modules.html"
+ ]
+ },
+ {
+ "group": "Utils",
+ "pages": [
+ "neuralforecast/tsdataset.html",
+ "neuralforecast/utils.html"
+ ]
+ }
+ ]
+ }
+ ]
+ },
+ {
+ "anchor": "HierarchicalForecast",
+ "icon": "crown",
+ "groups": [
+ {
+ "group": " ",
+ "pages": ["hierarchicalforecast/index.html"]
+ },
+ {
+ "group": "Getting Started",
+ "pages": [
+ "hierarchicalforecast/examples/installation.html",
+ "hierarchicalforecast/examples/tourismsmall.html",
+ "hierarchicalforecast/examples/tourismsmallpolars.html",
+ "hierarchicalforecast/examples/introduction.html"
+ ]
+ },
+ {
+ "group": "Tutorials",
+ "pages": [
+ {
+ "group": "Point Reconciliation",
+ "pages": [
+ "hierarchicalforecast/examples/australiandomestictourism.html",
+ "hierarchicalforecast/examples/australianprisonpopulation.html",
+ "hierarchicalforecast/examples/nonnegativereconciliation.html"
+ ]
+ },
+ {
+ "group": "Probabilistic Reconciliation",
+ "pages": [
+ "hierarchicalforecast/examples/australiandomestictourism-intervals.html",
+ "hierarchicalforecast/examples/australiandomestictourism-bootstraped-intervals.html",
+ "hierarchicalforecast/examples/australiandomestictourism-permbu-intervals.html",
+ "hierarchicalforecast/examples/tourismlarge-evaluation.html"
+ ]
+ },
+ {
+ "group": "Temporal Reconciliation",
+ "pages": [
+ "hierarchicalforecast/examples/australiandomestictourismtemporal.html",
+ "hierarchicalforecast/examples/australiandomestictourismcrosstemporal.html",
+ "hierarchicalforecast/examples/m3withthief.html",
+ "hierarchicalforecast/examples/localglobalaggregation.html"
+ ]
+ },
+ "hierarchicalforecast/examples/mlframeworksexample.html"
+ ]
+ },
+ {
+ "group": "API Reference",
+ "pages": [
+ "hierarchicalforecast/src/core.html",
+ "hierarchicalforecast/src/methods.html",
+ "hierarchicalforecast/src/probabilistic_methods.html",
+ "hierarchicalforecast/src/evaluation.html",
+ "hierarchicalforecast/src/utils.html"
+ ]
+ }
+ ]
+ },
+ {
+ "anchor": "UtilsForecast",
+ "icon": "wrench",
+ "groups": [
+ {
+ "group": " ",
+ "pages": ["utilsforecast/index.html"]
+ },
+ {
+ "group": "API Reference",
+ "pages": [
+ "utilsforecast/preprocessing.html",
+ "utilsforecast/feature_engineering.html",
+ "utilsforecast/evaluation.html",
+ "utilsforecast/losses.html",
+ "utilsforecast/plotting.html",
+ "utilsforecast/data.html"
+ ]
+ }
+ ]
+ },
+ {
+ "anchor": "DatasetsForecast",
+ "icon": "chart-simple",
+ "groups": [
+ {
+ "group": " ",
+ "pages": ["datasetsforecast/index.html"]
+ },
+ {
+ "group": "Datasets",
+ "pages": [
+ "datasetsforecast/favorita.html",
+ "datasetsforecast/hierarchical.html",
+ "datasetsforecast/long_horizon.html",
+ "datasetsforecast/long_horizon2.html",
+ "datasetsforecast/m3.html",
+ "datasetsforecast/m4.html",
+ "datasetsforecast/m5.html",
+ "datasetsforecast/phm2008.html",
+ "datasetsforecast/utils.html"
+ ]
+ }
+ ]
+ },
+ {
+ "anchor": "CoreForecast",
+ "icon": "truck-fast",
+ "groups": [
+ {
+ "group": " ",
+ "pages": ["coreforecast/index"]
+ },
+ {
+ "group": "API Reference",
+ "pages": [
+ "coreforecast/grouped_array",
+ "coreforecast/lag_transforms",
+ "coreforecast/scalers",
+ "coreforecast/differences",
+ "coreforecast/seasonal",
+ "coreforecast/rolling",
+ "coreforecast/expanding",
+ "coreforecast/exponentially_weighted"
+ ]
+ }
+ ]
+ }
+ ]
+ },
+ "redirects": [
+ {
+ "source": "/statsforecast/docs/getting-started/0_Installation",
+ "destination": "/statsforecast/docs/getting-started/installation.html"
+ },
+ {
+ "source": "/statsforecast/docs/getting-started/1_Getting_Started_short",
+ "destination": "/statsforecast/docs/getting-started/getting_started_short.html"
+ },
+ {
+ "source": "/statsforecast/docs/getting-started/2_Getting_Started_complete",
+ "destination": "/statsforecast/docs/getting-started/getting_started_complete.html"
+ },
+ {
+ "source": "/statsforecast/docs/tutorials/AnomalyDetection",
+ "destination": "/statsforecast/docs/tutorials/anomalydetection.html"
+ },
+ {
+ "source": "/statsforecast/docs/tutorials/CrossValidation",
+ "destination": "/statsforecast/docs/tutorials/crossvalidation.html"
+ },
+ {
+ "source": "/statsforecast/docs/tutorials/MultipleSeasonalities",
+ "destination": "/statsforecast/docs/tutorials/multipleseasonalities.html"
+ },
+ {
+ "source": "/statsforecast/docs/tutorials/ElectricityPeakForecasting",
+ "destination": "/statsforecast/docs/tutorials/electricitypeakforecasting.html"
+ },
+ {
+ "source": "/statsforecast/docs/tutorials/IntermittentData",
+ "destination": "/statsforecast/docs/tutorials/intermittentdata.html"
+ },
+ {
+ "source": "/statsforecast/docs/how-to-guides/Exogenous",
+ "destination": "/statsforecast/docs/how-to-guides/exogenous.html"
+ }
+ ],
+ "theme": "mint"
+}
diff --git a/fonts/Geist-VariableFont_wght.ttf b/fonts/Geist-VariableFont_wght.ttf
new file mode 100644
index 00000000..ad6f2c5a
Binary files /dev/null and b/fonts/Geist-VariableFont_wght.ttf differ
diff --git a/fonts/PPNeueMontrealMono-Medium.otf b/fonts/PPNeueMontrealMono-Medium.otf
new file mode 100644
index 00000000..1dd7ecce
Binary files /dev/null and b/fonts/PPNeueMontrealMono-Medium.otf differ
diff --git a/fonts/Supply-Regular.otf b/fonts/Supply-Regular.otf
new file mode 100644
index 00000000..99c6986c
Binary files /dev/null and b/fonts/Supply-Regular.otf differ
diff --git a/fonts/ppneuemontreal-medium.otf b/fonts/ppneuemontreal-medium.otf
new file mode 100644
index 00000000..315ac601
Binary files /dev/null and b/fonts/ppneuemontreal-medium.otf differ
diff --git a/hierarchicalforecast/.nojekyll b/hierarchicalforecast/.nojekyll
new file mode 100644
index 00000000..e69de29b
diff --git a/hierarchicalforecast/dark.png b/hierarchicalforecast/dark.png
new file mode 100644
index 00000000..4142a0bb
Binary files /dev/null and b/hierarchicalforecast/dark.png differ
diff --git a/hierarchicalforecast/examples/AustralianDomesticTourism-Bootstraped-Intervals_files/figure-markdown_strict/cell-11-output-1.png b/hierarchicalforecast/examples/AustralianDomesticTourism-Bootstraped-Intervals_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..254eadae
Binary files /dev/null and b/hierarchicalforecast/examples/AustralianDomesticTourism-Bootstraped-Intervals_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/hierarchicalforecast/examples/AustralianDomesticTourism-Bootstraped-Intervals_files/figure-markdown_strict/cell-12-output-1.png b/hierarchicalforecast/examples/AustralianDomesticTourism-Bootstraped-Intervals_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..5437fb5d
Binary files /dev/null and b/hierarchicalforecast/examples/AustralianDomesticTourism-Bootstraped-Intervals_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/hierarchicalforecast/examples/AustralianDomesticTourism-Bootstraped-Intervals_files/figure-markdown_strict/cell-19-output-1.png b/hierarchicalforecast/examples/AustralianDomesticTourism-Bootstraped-Intervals_files/figure-markdown_strict/cell-19-output-1.png
new file mode 100644
index 00000000..338014c6
Binary files /dev/null and b/hierarchicalforecast/examples/AustralianDomesticTourism-Bootstraped-Intervals_files/figure-markdown_strict/cell-19-output-1.png differ
diff --git a/hierarchicalforecast/examples/AustralianDomesticTourism-Bootstraped-Intervals_files/figure-markdown_strict/cell-20-output-1.png b/hierarchicalforecast/examples/AustralianDomesticTourism-Bootstraped-Intervals_files/figure-markdown_strict/cell-20-output-1.png
new file mode 100644
index 00000000..8d053cac
Binary files /dev/null and b/hierarchicalforecast/examples/AustralianDomesticTourism-Bootstraped-Intervals_files/figure-markdown_strict/cell-20-output-1.png differ
diff --git a/hierarchicalforecast/examples/AustralianDomesticTourism-Bootstraped-Intervals_files/figure-markdown_strict/cell-21-output-1.png b/hierarchicalforecast/examples/AustralianDomesticTourism-Bootstraped-Intervals_files/figure-markdown_strict/cell-21-output-1.png
new file mode 100644
index 00000000..989d7f95
Binary files /dev/null and b/hierarchicalforecast/examples/AustralianDomesticTourism-Bootstraped-Intervals_files/figure-markdown_strict/cell-21-output-1.png differ
diff --git a/hierarchicalforecast/examples/AustralianDomesticTourism-Bootstraped-Intervals_files/figure-markdown_strict/cell-22-output-1.png b/hierarchicalforecast/examples/AustralianDomesticTourism-Bootstraped-Intervals_files/figure-markdown_strict/cell-22-output-1.png
new file mode 100644
index 00000000..6c0c5faa
Binary files /dev/null and b/hierarchicalforecast/examples/AustralianDomesticTourism-Bootstraped-Intervals_files/figure-markdown_strict/cell-22-output-1.png differ
diff --git a/hierarchicalforecast/examples/AustralianDomesticTourism-Intervals_files/figure-markdown_strict/cell-11-output-1.png b/hierarchicalforecast/examples/AustralianDomesticTourism-Intervals_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..994c176c
Binary files /dev/null and b/hierarchicalforecast/examples/AustralianDomesticTourism-Intervals_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/hierarchicalforecast/examples/AustralianDomesticTourism-Intervals_files/figure-markdown_strict/cell-12-output-1.png b/hierarchicalforecast/examples/AustralianDomesticTourism-Intervals_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..9ec36845
Binary files /dev/null and b/hierarchicalforecast/examples/AustralianDomesticTourism-Intervals_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/hierarchicalforecast/examples/AustralianDomesticTourism-Intervals_files/figure-markdown_strict/cell-19-output-1.png b/hierarchicalforecast/examples/AustralianDomesticTourism-Intervals_files/figure-markdown_strict/cell-19-output-1.png
new file mode 100644
index 00000000..04ed62e1
Binary files /dev/null and b/hierarchicalforecast/examples/AustralianDomesticTourism-Intervals_files/figure-markdown_strict/cell-19-output-1.png differ
diff --git a/hierarchicalforecast/examples/AustralianDomesticTourism-Intervals_files/figure-markdown_strict/cell-20-output-1.png b/hierarchicalforecast/examples/AustralianDomesticTourism-Intervals_files/figure-markdown_strict/cell-20-output-1.png
new file mode 100644
index 00000000..c1777bf0
Binary files /dev/null and b/hierarchicalforecast/examples/AustralianDomesticTourism-Intervals_files/figure-markdown_strict/cell-20-output-1.png differ
diff --git a/hierarchicalforecast/examples/AustralianDomesticTourism-Intervals_files/figure-markdown_strict/cell-21-output-1.png b/hierarchicalforecast/examples/AustralianDomesticTourism-Intervals_files/figure-markdown_strict/cell-21-output-1.png
new file mode 100644
index 00000000..6644c172
Binary files /dev/null and b/hierarchicalforecast/examples/AustralianDomesticTourism-Intervals_files/figure-markdown_strict/cell-21-output-1.png differ
diff --git a/hierarchicalforecast/examples/AustralianDomesticTourism-Intervals_files/figure-markdown_strict/cell-22-output-1.png b/hierarchicalforecast/examples/AustralianDomesticTourism-Intervals_files/figure-markdown_strict/cell-22-output-1.png
new file mode 100644
index 00000000..4cb0ab7d
Binary files /dev/null and b/hierarchicalforecast/examples/AustralianDomesticTourism-Intervals_files/figure-markdown_strict/cell-22-output-1.png differ
diff --git a/hierarchicalforecast/examples/AustralianDomesticTourism-Permbu-Intervals_files/figure-markdown_strict/cell-11-output-1.png b/hierarchicalforecast/examples/AustralianDomesticTourism-Permbu-Intervals_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..46c5d116
Binary files /dev/null and b/hierarchicalforecast/examples/AustralianDomesticTourism-Permbu-Intervals_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/hierarchicalforecast/examples/AustralianDomesticTourism-Permbu-Intervals_files/figure-markdown_strict/cell-12-output-1.png b/hierarchicalforecast/examples/AustralianDomesticTourism-Permbu-Intervals_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..8a6ee770
Binary files /dev/null and b/hierarchicalforecast/examples/AustralianDomesticTourism-Permbu-Intervals_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/hierarchicalforecast/examples/AustralianDomesticTourism-Permbu-Intervals_files/figure-markdown_strict/cell-19-output-1.png b/hierarchicalforecast/examples/AustralianDomesticTourism-Permbu-Intervals_files/figure-markdown_strict/cell-19-output-1.png
new file mode 100644
index 00000000..54adf490
Binary files /dev/null and b/hierarchicalforecast/examples/AustralianDomesticTourism-Permbu-Intervals_files/figure-markdown_strict/cell-19-output-1.png differ
diff --git a/hierarchicalforecast/examples/AustralianDomesticTourism-Permbu-Intervals_files/figure-markdown_strict/cell-20-output-1.png b/hierarchicalforecast/examples/AustralianDomesticTourism-Permbu-Intervals_files/figure-markdown_strict/cell-20-output-1.png
new file mode 100644
index 00000000..072bc7c3
Binary files /dev/null and b/hierarchicalforecast/examples/AustralianDomesticTourism-Permbu-Intervals_files/figure-markdown_strict/cell-20-output-1.png differ
diff --git a/hierarchicalforecast/examples/AustralianDomesticTourism-Permbu-Intervals_files/figure-markdown_strict/cell-21-output-1.png b/hierarchicalforecast/examples/AustralianDomesticTourism-Permbu-Intervals_files/figure-markdown_strict/cell-21-output-1.png
new file mode 100644
index 00000000..e23a9645
Binary files /dev/null and b/hierarchicalforecast/examples/AustralianDomesticTourism-Permbu-Intervals_files/figure-markdown_strict/cell-21-output-1.png differ
diff --git a/hierarchicalforecast/examples/AustralianDomesticTourismTemporal_files/figure-markdown_strict/cell-28-output-1.png b/hierarchicalforecast/examples/AustralianDomesticTourismTemporal_files/figure-markdown_strict/cell-28-output-1.png
new file mode 100644
index 00000000..57b46caa
Binary files /dev/null and b/hierarchicalforecast/examples/AustralianDomesticTourismTemporal_files/figure-markdown_strict/cell-28-output-1.png differ
diff --git a/hierarchicalforecast/examples/MLFrameworksExample_files/figure-markdown_strict/cell-18-output-1.png b/hierarchicalforecast/examples/MLFrameworksExample_files/figure-markdown_strict/cell-18-output-1.png
new file mode 100644
index 00000000..41af2619
Binary files /dev/null and b/hierarchicalforecast/examples/MLFrameworksExample_files/figure-markdown_strict/cell-18-output-1.png differ
diff --git a/hierarchicalforecast/examples/MLFrameworksExample_files/figure-markdown_strict/cell-19-output-1.png b/hierarchicalforecast/examples/MLFrameworksExample_files/figure-markdown_strict/cell-19-output-1.png
new file mode 100644
index 00000000..0e79ba16
Binary files /dev/null and b/hierarchicalforecast/examples/MLFrameworksExample_files/figure-markdown_strict/cell-19-output-1.png differ
diff --git a/hierarchicalforecast/examples/MLFrameworksExample_files/figure-markdown_strict/cell-6-output-1.png b/hierarchicalforecast/examples/MLFrameworksExample_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..1366b01f
Binary files /dev/null and b/hierarchicalforecast/examples/MLFrameworksExample_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/hierarchicalforecast/examples/TourismLarge-Evaluation_files/figure-markdown_strict/cell-11-output-1.png b/hierarchicalforecast/examples/TourismLarge-Evaluation_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..cb2d0004
Binary files /dev/null and b/hierarchicalforecast/examples/TourismLarge-Evaluation_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/hierarchicalforecast/examples/TourismLarge-Evaluation_files/figure-markdown_strict/cell-7-output-1.png b/hierarchicalforecast/examples/TourismLarge-Evaluation_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..a959ef7f
Binary files /dev/null and b/hierarchicalforecast/examples/TourismLarge-Evaluation_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/hierarchicalforecast/examples/australiandomestictourism-bootstraped-intervals.html.mdx b/hierarchicalforecast/examples/australiandomestictourism-bootstraped-intervals.html.mdx
new file mode 100644
index 00000000..c3e0247b
--- /dev/null
+++ b/hierarchicalforecast/examples/australiandomestictourism-bootstraped-intervals.html.mdx
@@ -0,0 +1,322 @@
+---
+output-file: australiandomestictourism-bootstraped-intervals.html
+title: Bootstrap
+---
+
+
+
+
+In many cases, only the time series at the lowest level of the
+hierarchies (bottom time series) are available. `HierarchicalForecast`
+has tools to create time series for all hierarchies and also allows you
+to calculate prediction intervals for all hierarchies. In this notebook
+we will see how to do it.
+
+
+```python
+!pip install hierarchicalforecast statsforecast
+```
+
+
+```python
+import pandas as pd
+
+# compute base forecast no coherent
+from statsforecast.models import AutoETS
+from statsforecast.core import StatsForecast
+
+#obtain hierarchical reconciliation methods and evaluation
+from hierarchicalforecast.methods import BottomUp, MinTrace
+from hierarchicalforecast.utils import aggregate, HierarchicalPlot
+from hierarchicalforecast.core import HierarchicalReconciliation
+```
+
+## Aggregate bottom time series
+
+In this example we will use the
+[Tourism](https://otexts.com/fpp3/tourism.html) dataset from the
+[Forecasting: Principles and Practice](https://otexts.com/fpp3/) book.
+The dataset only contains the time series at the lowest level, so we
+need to create the time series for all hierarchies.
+
+
+```python
+Y_df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/tourism.csv')
+Y_df = Y_df.rename({'Trips': 'y', 'Quarter': 'ds'}, axis=1)
+Y_df.insert(0, 'Country', 'Australia')
+Y_df = Y_df[['Country', 'Region', 'State', 'Purpose', 'ds', 'y']]
+Y_df['ds'] = Y_df['ds'].str.replace(r'(\d+) (Q\d)', r'\1\2', regex=True)
+Y_df['ds'] = pd.PeriodIndex(Y_df["ds"], freq='Q').to_timestamp()
+Y_df.head()
+```
+
+| | Country | Region | State | Purpose | ds | y |
+|-----|-----------|----------|-----------------|----------|------------|------------|
+| 0 | Australia | Adelaide | South Australia | Business | 1998-01-01 | 135.077690 |
+| 1 | Australia | Adelaide | South Australia | Business | 1998-04-01 | 109.987316 |
+| 2 | Australia | Adelaide | South Australia | Business | 1998-07-01 | 166.034687 |
+| 3 | Australia | Adelaide | South Australia | Business | 1998-10-01 | 127.160464 |
+| 4 | Australia | Adelaide | South Australia | Business | 1999-01-01 | 137.448533 |
+
+The dataset can be grouped in the following non-strictly hierarchical
+structure.
+
+
+```python
+spec = [
+ ['Country'],
+ ['Country', 'State'],
+ ['Country', 'Purpose'],
+ ['Country', 'State', 'Region'],
+ ['Country', 'State', 'Purpose'],
+ ['Country', 'State', 'Region', 'Purpose']
+]
+```
+
+Using the
+[`aggregate`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#aggregate)
+function from `HierarchicalForecast` we can generate: 1. `Y_df`: the
+hierarchical structured series $\mathbf{y}_{[a,b]\tau}$ 2. `S_df`: the
+aggregation constraings dataframe with $S_{[a,b]}$ 3. `tags`: a list
+with the ‘unique_ids’ conforming each aggregation level.
+
+
+```python
+Y_df, S_df, tags = aggregate(df=Y_df, spec=spec)
+```
+
+
+```python
+Y_df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|--------------|
+| 0 | Australia | 1998-01-01 | 23182.197269 |
+| 1 | Australia | 1998-04-01 | 20323.380067 |
+| 2 | Australia | 1998-07-01 | 19826.640511 |
+| 3 | Australia | 1998-10-01 | 20830.129891 |
+| 4 | Australia | 1999-01-01 | 22087.353380 |
+
+
+```python
+S_df.iloc[:5, :5]
+```
+
+| | unique_id | Australia/ACT/Canberra/Business | Australia/ACT/Canberra/Holiday | Australia/ACT/Canberra/Other | Australia/ACT/Canberra/Visiting |
+|----|----|----|----|----|----|
+| 0 | Australia | 1.0 | 1.0 | 1.0 | 1.0 |
+| 1 | Australia/ACT | 1.0 | 1.0 | 1.0 | 1.0 |
+| 2 | Australia/New South Wales | 0.0 | 0.0 | 0.0 | 0.0 |
+| 3 | Australia/Northern Territory | 0.0 | 0.0 | 0.0 | 0.0 |
+| 4 | Australia/Queensland | 0.0 | 0.0 | 0.0 | 0.0 |
+
+
+```python
+tags['Country/Purpose']
+```
+
+``` text
+array(['Australia/Business', 'Australia/Holiday', 'Australia/Other',
+ 'Australia/Visiting'], dtype=object)
+```
+
+We can visualize the `S_df` dataframe and `Y_df` using the
+[`HierarchicalPlot`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#hierarchicalplot)
+class as follows.
+
+
+```python
+hplot = HierarchicalPlot(S=S_df, tags=tags)
+```
+
+
+```python
+hplot.plot_summing_matrix()
+```
+
+
+
+
+```python
+hplot.plot_hierarchically_linked_series(
+ bottom_series='Australia/ACT/Canberra/Holiday',
+ Y_df=Y_df
+)
+```
+
+
+
+### Split Train/Test sets
+
+We use the final two years (8 quarters) as test set.
+
+
+```python
+Y_test_df = Y_df.groupby('unique_id', as_index=False).tail(8)
+Y_train_df = Y_df.drop(Y_test_df.index)
+```
+
+
+```python
+Y_train_df.groupby('unique_id').size()
+```
+
+``` text
+unique_id
+Australia 72
+Australia/ACT 72
+Australia/ACT/Business 72
+Australia/ACT/Canberra 72
+Australia/ACT/Canberra/Business 72
+ ..
+Australia/Western Australia/Experience Perth/Other 72
+Australia/Western Australia/Experience Perth/Visiting 72
+Australia/Western Australia/Holiday 72
+Australia/Western Australia/Other 72
+Australia/Western Australia/Visiting 72
+Length: 425, dtype: int64
+```
+
+## Computing Base Forecasts
+
+The following cell computes the **base forecasts** for each time series
+in `Y_df` using the `AutoETS` and model. Observe that `Y_hat_df`
+contains the forecasts but they are not coherent. Since we are computing
+prediction intervals using bootstrapping, we only need the fitted values
+of the models.
+
+
+```python
+fcst = StatsForecast(models=[AutoETS(season_length=4, model='ZAA')],
+ freq='QS', n_jobs=-1)
+Y_hat_df = fcst.forecast(df=Y_train_df, h=8, fitted=True)
+Y_fitted_df = fcst.forecast_fitted_values()
+```
+
+## Reconcile Base Forecasts
+
+The following cell makes the previous forecasts coherent using the
+[`HierarchicalReconciliation`](https://Nixtla.github.io/hierarchicalforecast/src/core.html#hierarchicalreconciliation)
+class. Since the hierarchy structure is not strict, we can’t use methods
+such as
+[`TopDown`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#topdown)
+or
+[`MiddleOut`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#middleout).
+In this example we use
+[`BottomUp`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#bottomup)
+and
+[`MinTrace`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#mintrace).
+If you want to calculate prediction intervals, you have to use the
+`level` argument as follows and set `intervals_method='bootstrap'`.
+
+
+```python
+reconcilers = [
+ BottomUp(),
+ MinTrace(method='mint_shrink'),
+ MinTrace(method='ols')
+]
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+Y_rec_df = hrec.reconcile(Y_hat_df=Y_hat_df, Y_df=Y_fitted_df, S_df=S_df,
+ tags=tags, level=[80, 90],
+ intervals_method='bootstrap')
+```
+
+The dataframe `Y_rec_df` contains the reconciled forecasts.
+
+
+```python
+Y_rec_df.head()
+```
+
+| | unique_id | ds | AutoETS | AutoETS/BottomUp | AutoETS/BottomUp-lo-90 | AutoETS/BottomUp-lo-80 | AutoETS/BottomUp-hi-80 | AutoETS/BottomUp-hi-90 | AutoETS/MinTrace_method-mint_shrink | AutoETS/MinTrace_method-mint_shrink-lo-90 | AutoETS/MinTrace_method-mint_shrink-lo-80 | AutoETS/MinTrace_method-mint_shrink-hi-80 | AutoETS/MinTrace_method-mint_shrink-hi-90 | AutoETS/MinTrace_method-ols | AutoETS/MinTrace_method-ols-lo-90 | AutoETS/MinTrace_method-ols-lo-80 | AutoETS/MinTrace_method-ols-hi-80 | AutoETS/MinTrace_method-ols-hi-90 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | Australia | 2016-01-01 | 26080.878488 | 24487.152503 | 23242.757311 | 23332.592968 | 25379.829486 | 25424.139137 | 25521.551706 | 24407.442712 | 24698.931479 | 26357.024354 | 26466.740682 | 26034.132091 | 24914.199038 | 25100.470502 | 27102.746065 | 27176.467048 |
+| 1 | Australia | 2016-04-01 | 24587.012115 | 23068.314292 | 21823.919100 | 21910.615057 | 23945.982949 | 24278.683243 | 24106.522479 | 23185.403634 | 23283.902251 | 25098.332342 | 25473.239949 | 24567.457913 | 23483.983814 | 23640.627126 | 25709.792870 | 25809.220444 |
+| 2 | Australia | 2016-07-01 | 24147.307744 | 22686.983933 | 21293.529449 | 21526.525610 | 23697.859931 | 24150.879789 | 23717.610501 | 22603.501507 | 22802.771308 | 24802.973260 | 25228.795629 | 24150.111246 | 23030.178193 | 23154.972436 | 25359.917993 | 25404.792198 |
+| 3 | Australia | 2016-10-01 | 24794.040779 | 23428.037637 | 22034.583153 | 22273.826957 | 24241.840440 | 24438.913635 | 24472.939115 | 23361.285512 | 23584.825871 | 25338.713995 | 25469.426623 | 24831.540721 | 23725.927463 | 23836.401911 | 25900.154695 | 25977.249268 |
+| 4 | Australia | 2017-01-01 | 26283.998654 | 24939.637616 | 23695.217554 | 23903.395713 | 25815.638682 | 25973.164607 | 26029.322724 | 24948.339795 | 25144.179030 | 26900.068461 | 27119.073160 | 26348.229758 | 25254.682234 | 25487.518098 | 27410.894158 | 27477.330557 |
+
+## Plot Predictions
+
+Then we can plot the probabilist forecasts using the following function.
+
+
+```python
+plot_df = Y_df.merge(Y_rec_df, on=['unique_id', 'ds'], how="outer")
+```
+
+### Plot single time series
+
+
+```python
+hplot.plot_series(
+ series='Australia',
+ Y_df=plot_df,
+ models=['y', 'AutoETS', 'AutoETS/MinTrace_method-ols', 'AutoETS/MinTrace_method-mint_shrink'],
+ level=[80]
+)
+```
+
+
+
+
+```python
+# Since we are plotting a bottom time series
+# the probabilistic and mean forecasts
+# differ due to bootstrapping
+hplot.plot_series(
+ series='Australia/Western Australia/Experience Perth/Visiting',
+ Y_df=plot_df,
+ models=['y', 'AutoETS', 'AutoETS/BottomUp'],
+ level=[80]
+)
+```
+
+
+
+### Plot hierarchichally linked time series
+
+
+```python
+hplot.plot_hierarchically_linked_series(
+ bottom_series='Australia/Western Australia/Experience Perth/Visiting',
+ Y_df=plot_df,
+ models=['y', 'AutoETS', 'AutoETS/MinTrace_method-ols', 'AutoETS/BottomUp'],
+ level=[80]
+)
+```
+
+
+
+
+```python
+# ACT only has Canberra
+hplot.plot_hierarchically_linked_series(
+ bottom_series='Australia/ACT/Canberra/Other',
+ Y_df=plot_df,
+ models=['y', 'AutoETS/MinTrace_method-mint_shrink'],
+ level=[80, 90]
+)
+```
+
+
+
+### References
+
+- [Hyndman, R.J., & Athanasopoulos, G. (2021). “Forecasting:
+ principles and practice, 3rd edition: Chapter 11: Forecasting
+ hierarchical and grouped series.”. OTexts: Melbourne, Australia.
+ OTexts.com/fpp3 Accessed on July
+ 2022.](https://otexts.com/fpp3/hierarchical.html)
+- [Shanika L. Wickramasuriya, George Athanasopoulos, and Rob J.
+ Hyndman. Optimal forecast reconciliation for hierarchical and
+ grouped time series through trace minimization.Journal of the
+ American Statistical Association, 114(526):804–819, 2019. doi:
+ 10.1080/01621459.2018.1448825. URL
+ https://robjhyndman.com/publications/mint/.](https://robjhyndman.com/publications/mint/)
+- [Puwasala Gamakumara Ph. D. dissertation. Monash University,
+ Econometrics and Business Statistics (2020). “Probabilistic Forecast
+ Reconciliation”](https://bridges.monash.edu/articles/thesis/Probabilistic_Forecast_Reconciliation_Theory_and_Applications/11869533)
+
diff --git a/hierarchicalforecast/examples/australiandomestictourism-intervals.html.mdx b/hierarchicalforecast/examples/australiandomestictourism-intervals.html.mdx
new file mode 100644
index 00000000..849bd4ff
--- /dev/null
+++ b/hierarchicalforecast/examples/australiandomestictourism-intervals.html.mdx
@@ -0,0 +1,319 @@
+---
+output-file: australiandomestictourism-intervals.html
+title: Normality
+---
+
+
+
+
+In many cases, only the time series at the lowest level of the
+hierarchies (bottom time series) are available. `HierarchicalForecast`
+has tools to create time series for all hierarchies and also allows you
+to calculate prediction intervals for all hierarchies. In this notebook
+we will see how to do it.
+
+
+```python
+!pip install hierarchicalforecast statsforecast
+```
+
+
+```python
+import pandas as pd
+
+# compute base forecast no coherent
+from statsforecast.models import AutoARIMA
+from statsforecast.core import StatsForecast
+
+#obtain hierarchical reconciliation methods and evaluation
+from hierarchicalforecast.methods import BottomUp, MinTrace
+from hierarchicalforecast.utils import aggregate, HierarchicalPlot
+from hierarchicalforecast.core import HierarchicalReconciliation
+```
+
+## Aggregate bottom time series
+
+In this example we will use the
+[Tourism](https://otexts.com/fpp3/tourism.html) dataset from the
+[Forecasting: Principles and Practice](https://otexts.com/fpp3/) book.
+The dataset only contains the time series at the lowest level, so we
+need to create the time series for all hierarchies.
+
+
+```python
+Y_df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/tourism.csv')
+Y_df = Y_df.rename({'Trips': 'y', 'Quarter': 'ds'}, axis=1)
+Y_df.insert(0, 'Country', 'Australia')
+Y_df = Y_df[['Country', 'Region', 'State', 'Purpose', 'ds', 'y']]
+Y_df['ds'] = Y_df['ds'].str.replace(r'(\d+) (Q\d)', r'\1-\2', regex=True)
+Y_df['ds'] = pd.PeriodIndex(Y_df["ds"], freq='Q').to_timestamp()
+Y_df.head()
+```
+
+| | Country | Region | State | Purpose | ds | y |
+|-----|-----------|----------|-----------------|----------|------------|------------|
+| 0 | Australia | Adelaide | South Australia | Business | 1998-01-01 | 135.077690 |
+| 1 | Australia | Adelaide | South Australia | Business | 1998-04-01 | 109.987316 |
+| 2 | Australia | Adelaide | South Australia | Business | 1998-07-01 | 166.034687 |
+| 3 | Australia | Adelaide | South Australia | Business | 1998-10-01 | 127.160464 |
+| 4 | Australia | Adelaide | South Australia | Business | 1999-01-01 | 137.448533 |
+
+The dataset can be grouped in the following non-strictly hierarchical
+structure.
+
+
+```python
+spec = [
+ ['Country'],
+ ['Country', 'State'],
+ ['Country', 'Purpose'],
+ ['Country', 'State', 'Region'],
+ ['Country', 'State', 'Purpose'],
+ ['Country', 'State', 'Region', 'Purpose']
+]
+```
+
+Using the
+[`aggregate`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#aggregate)
+function from `HierarchicalForecast` we can generate: 1. `Y_df`: the
+hierarchical structured series $\mathbf{y}_{[a,b]\tau}$ 2. `S_df`: the
+aggregation constraings dataframe with $S_{[a,b]}$ 3. `tags`: a list
+with the ‘unique_ids’ conforming each aggregation level.
+
+
+```python
+Y_df, S_df, tags = aggregate(df=Y_df, spec=spec)
+```
+
+
+```python
+Y_df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|--------------|
+| 0 | Australia | 1998-01-01 | 23182.197269 |
+| 1 | Australia | 1998-04-01 | 20323.380067 |
+| 2 | Australia | 1998-07-01 | 19826.640511 |
+| 3 | Australia | 1998-10-01 | 20830.129891 |
+| 4 | Australia | 1999-01-01 | 22087.353380 |
+
+
+```python
+S_df.iloc[:5, :5]
+```
+
+| | unique_id | Australia/ACT/Canberra/Business | Australia/ACT/Canberra/Holiday | Australia/ACT/Canberra/Other | Australia/ACT/Canberra/Visiting |
+|----|----|----|----|----|----|
+| 0 | Australia | 1.0 | 1.0 | 1.0 | 1.0 |
+| 1 | Australia/ACT | 1.0 | 1.0 | 1.0 | 1.0 |
+| 2 | Australia/New South Wales | 0.0 | 0.0 | 0.0 | 0.0 |
+| 3 | Australia/Northern Territory | 0.0 | 0.0 | 0.0 | 0.0 |
+| 4 | Australia/Queensland | 0.0 | 0.0 | 0.0 | 0.0 |
+
+
+```python
+tags['Country/Purpose']
+```
+
+``` text
+array(['Australia/Business', 'Australia/Holiday', 'Australia/Other',
+ 'Australia/Visiting'], dtype=object)
+```
+
+We can visualize the `S` matrix and the data using the
+[`HierarchicalPlot`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#hierarchicalplot)
+class as follows.
+
+
+```python
+hplot = HierarchicalPlot(S=S_df, tags=tags)
+```
+
+
+```python
+hplot.plot_summing_matrix()
+```
+
+
+
+
+```python
+hplot.plot_hierarchically_linked_series(
+ bottom_series='Australia/ACT/Canberra/Holiday',
+ Y_df=Y_df
+)
+```
+
+
+
+### Split Train/Test sets
+
+We use the final two years (8 quarters) as test set.
+
+
+```python
+Y_test_df = Y_df.groupby('unique_id', as_index=False).tail(8)
+Y_train_df = Y_df.drop(Y_test_df.index)
+```
+
+
+```python
+Y_train_df.groupby('unique_id').size()
+```
+
+``` text
+unique_id
+Australia 72
+Australia/ACT 72
+Australia/ACT/Business 72
+Australia/ACT/Canberra 72
+Australia/ACT/Canberra/Business 72
+ ..
+Australia/Western Australia/Experience Perth/Other 72
+Australia/Western Australia/Experience Perth/Visiting 72
+Australia/Western Australia/Holiday 72
+Australia/Western Australia/Other 72
+Australia/Western Australia/Visiting 72
+Length: 425, dtype: int64
+```
+
+## Computing base forecasts
+
+The following cell computes the **base forecasts** for each time series
+in `Y_df` using the `AutoARIMA` and model. Observe that `Y_hat_df`
+contains the forecasts but they are not coherent. To reconcile the
+prediction intervals we need to calculate the uncoherent intervals using
+the `level` argument of `StatsForecast`.
+
+
+```python
+fcst = StatsForecast(models=[AutoARIMA(season_length=4)],
+ freq='QS', n_jobs=-1)
+Y_hat_df = fcst.forecast(df=Y_train_df, h=8, fitted=True, level=[80, 90])
+Y_fitted_df = fcst.forecast_fitted_values()
+```
+
+## Reconcile forecasts
+
+The following cell makes the previous forecasts coherent using the
+[`HierarchicalReconciliation`](https://Nixtla.github.io/hierarchicalforecast/src/core.html#hierarchicalreconciliation)
+class. Since the hierarchy structure is not strict, we can’t use methods
+such as
+[`TopDown`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#topdown)
+or
+[`MiddleOut`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#middleout).
+In this example we use
+[`BottomUp`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#bottomup)
+and
+[`MinTrace`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#mintrace).
+If you want to calculate prediction intervals, you have to use the
+`level` argument as follows.
+
+
+```python
+reconcilers = [
+ BottomUp(),
+ MinTrace(method='mint_shrink'),
+ MinTrace(method='ols')
+]
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+Y_rec_df = hrec.reconcile(Y_hat_df=Y_hat_df, Y_df=Y_fitted_df,
+ S_df=S_df, tags=tags, level=[80, 90])
+```
+
+The dataframe `Y_rec_df` contains the reconciled forecasts.
+
+
+```python
+Y_rec_df.head()
+```
+
+| | unique_id | ds | AutoARIMA | AutoARIMA-lo-90 | AutoARIMA-lo-80 | AutoARIMA-hi-80 | AutoARIMA-hi-90 | AutoARIMA/BottomUp | AutoARIMA/BottomUp-lo-90 | AutoARIMA/BottomUp-lo-80 | ... | AutoARIMA/MinTrace_method-mint_shrink | AutoARIMA/MinTrace_method-mint_shrink-lo-90 | AutoARIMA/MinTrace_method-mint_shrink-lo-80 | AutoARIMA/MinTrace_method-mint_shrink-hi-80 | AutoARIMA/MinTrace_method-mint_shrink-hi-90 | AutoARIMA/MinTrace_method-ols | AutoARIMA/MinTrace_method-ols-lo-90 | AutoARIMA/MinTrace_method-ols-lo-80 | AutoARIMA/MinTrace_method-ols-hi-80 | AutoARIMA/MinTrace_method-ols-hi-90 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | Australia | 2016-01-01 | 26212.553553 | 24705.948180 | 25038.715077 | 27386.392029 | 27719.158927 | 24646.517084 | 23983.656843 | 24130.064091 | ... | 25267.797338 | 24491.630618 | 24663.064091 | 25872.530586 | 26043.964058 | 26082.753488 | 25010.876141 | 25247.623803 | 26917.883174 | 27154.630835 |
+| 1 | Australia | 2016-04-01 | 25033.667125 | 23337.267588 | 23711.954696 | 26355.379554 | 26730.066662 | 22942.957703 | 22229.916838 | 22387.407579 | ... | 23836.804444 | 23002.620214 | 23186.868128 | 24486.740760 | 24670.988674 | 24822.102094 | 23616.734393 | 23882.966332 | 25761.237857 | 26027.469796 |
+| 2 | Australia | 2016-07-01 | 24507.027198 | 22640.028798 | 23052.396413 | 25961.657983 | 26374.025599 | 22568.286488 | 21805.892199 | 21974.283728 | ... | 23294.240908 | 22410.719833 | 22605.864873 | 23982.616942 | 24177.761983 | 24269.578724 | 22944.380043 | 23237.079287 | 25302.078162 | 25594.777406 |
+| 3 | Australia | 2016-10-01 | 25598.928613 | 23575.665243 | 24022.547410 | 27175.309816 | 27622.191983 | 23113.075726 | 22308.671860 | 22486.342127 | ... | 24154.484487 | 23221.706185 | 23427.730766 | 24881.238208 | 25087.262790 | 25340.549923 | 23905.434070 | 24222.410936 | 26458.688911 | 26775.665777 |
+| 4 | Australia | 2017-01-01 | 26982.576796 | 24669.535238 | 25180.421285 | 28784.732308 | 29295.618354 | 23779.264921 | 22874.194227 | 23074.098975 | ... | 25155.001372 | 24125.268915 | 24352.707952 | 25957.294793 | 26184.733830 | 26690.200927 | 25051.352698 | 25413.328335 | 27967.073518 | 28329.049155 |
+
+## Plot forecasts
+
+Then we can plot the probabilistic forecasts using the following
+function.
+
+
+```python
+plot_df = Y_df.merge(Y_rec_df, on=['unique_id', 'ds'], how="outer")
+```
+
+### Plot single time series
+
+
+```python
+hplot.plot_series(
+ series='Australia',
+ Y_df=plot_df,
+ models=['y', 'AutoARIMA', 'AutoARIMA/MinTrace_method-ols'],
+ level=[80]
+)
+```
+
+
+
+
+```python
+# Since we are plotting a bottom time series
+# the probabilistic and mean forecasts
+# are the same
+hplot.plot_series(
+ series='Australia/Western Australia/Experience Perth/Visiting',
+ Y_df=plot_df,
+ models=['y', 'AutoARIMA', 'AutoARIMA/BottomUp'],
+ level=[80]
+)
+```
+
+
+
+### Plot hierarchichally linked time series
+
+
+```python
+hplot.plot_hierarchically_linked_series(
+ bottom_series='Australia/Western Australia/Experience Perth/Visiting',
+ Y_df=plot_df,
+ models=['y', 'AutoARIMA', 'AutoARIMA/MinTrace_method-ols', 'AutoARIMA/BottomUp'],
+ level=[80]
+)
+```
+
+
+
+
+```python
+# ACT only has Canberra
+hplot.plot_hierarchically_linked_series(
+ bottom_series='Australia/ACT/Canberra/Other',
+ Y_df=plot_df,
+ models=['y', 'AutoARIMA/MinTrace_method-mint_shrink'],
+ level=[80, 90]
+)
+```
+
+
+
+### References
+
+- [Hyndman, R.J., & Athanasopoulos, G. (2021). “Forecasting:
+ principles and practice, 3rd edition: Chapter 11: Forecasting
+ hierarchical and grouped series.”. OTexts: Melbourne, Australia.
+ OTexts.com/fpp3 Accessed on July
+ 2022.](https://otexts.com/fpp3/hierarchical.html)
+- [Shanika L. Wickramasuriya, George Athanasopoulos, and Rob J.
+ Hyndman. Optimal forecast reconciliation for hierarchical and
+ grouped time series through trace minimization.Journal of the
+ American Statistical Association, 114(526):804–819, 2019. doi:
+ 10.1080/01621459.2018.1448825. URL
+ https://robjhyndman.com/publications/mint/.](https://robjhyndman.com/publications/mint/)
+
diff --git a/hierarchicalforecast/examples/australiandomestictourism-multimodel.html.mdx b/hierarchicalforecast/examples/australiandomestictourism-multimodel.html.mdx
new file mode 100644
index 00000000..e111eac8
--- /dev/null
+++ b/hierarchicalforecast/examples/australiandomestictourism-multimodel.html.mdx
@@ -0,0 +1,415 @@
+---
+description: >-
+ Geographical Hierarchical Forecasting on Australian Tourism Data using
+ multiple models for each level in the hierarchy.
+output-file: australiandomestictourism-multimodel.html
+title: Multi-model Aggregation
+---
+
+
+This notebook extends the classic Australian Domestic Tourism
+(`Tourism`) geographical aggregation example to showcase how
+`HierarchicalForecast` can be used to produce coherent forecasts when
+**different forecasting models are applied at each level of the
+hierarchy**. We will use the `Tourism` dataset, which contains monthly
+time series of the number of visitors to each state of Australia.
+
+Specifically, we will demonstrate fitting a diverse set of models across
+the hierarchical levels. This includes statistical models like `AutoETS`
+from `StatsForecast`, machine learning models such as
+`HistGradientBoostingRegressor` using `MLForecast`, and neural network
+models like `NBEATS` from `NeuralForecast`. After generating these base
+forecasts, we will reconcile them using
+[`BottomUp`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#bottomup),
+`MinTrace(mint_shrink)`, `TopDown(forecast_proportions)` reconciliators
+from `HierarchicalForecast`.
+
+You can run these experiments using CPU or GPU with Google Colab.
+
+
+
+
+```python
+!pip install hierarchicalforecast statsforecast mlforecast datasetsforecast neuralforecast
+```
+
+## 1. Load and Process Data
+
+In this example we will use the
+[Tourism](https://otexts.com/fpp3/tourism.html) dataset from the
+[Forecasting: Principles and Practice](https://otexts.com/fpp3/) book.
+
+The dataset only contains the time series at the lowest level, so we
+need to create the time series for all hierarchies.
+
+
+```python
+import numpy as np
+import pandas as pd
+```
+
+
+```python
+Y_df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/tourism.csv')
+Y_df = Y_df.rename({'Trips': 'y', 'Quarter': 'ds'}, axis=1)
+Y_df.insert(0, 'Country', 'Australia')
+Y_df = Y_df[['Country', 'Region', 'State', 'ds', 'y']]
+Y_df['ds'] = Y_df['ds'].str.replace(r'(\d+) (Q\d)', r'\1-\2', regex=True)
+Y_df['ds'] = pd.PeriodIndex(Y_df['ds'], freq='Q').to_timestamp()
+Y_df_first = Y_df.groupby(['Country', 'Region', 'State', 'ds'], as_index=False).agg({'y':'sum'})
+Y_df_first.head()
+```
+
+| | Country | Region | State | ds | y |
+|-----|-----------|----------|-----------------|------------|------------|
+| 0 | Australia | Adelaide | South Australia | 1998-01-01 | 658.553895 |
+| 1 | Australia | Adelaide | South Australia | 1998-04-01 | 449.853935 |
+| 2 | Australia | Adelaide | South Australia | 1998-07-01 | 592.904597 |
+| 3 | Australia | Adelaide | South Australia | 1998-10-01 | 524.242760 |
+| 4 | Australia | Adelaide | South Australia | 1999-01-01 | 548.394105 |
+
+The dataset can be grouped in the following hierarchical structure.
+
+
+```python
+spec = [
+ ['Country'],
+ ['Country', 'State'],
+ ['Country', 'State', 'Region']
+]
+```
+
+Using the
+[`aggregate`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#aggregate)
+function from `HierarchicalForecast` we can get the full set of time
+series.
+
+
+```python
+from hierarchicalforecast.utils import aggregate
+```
+
+
+```python
+Y_df, S_df, tags = aggregate(Y_df_first, spec)
+```
+
+### Split Train/Test sets
+
+We use the final two years (8 quarters) as test set.
+
+
+```python
+Y_test_df = Y_df.groupby('unique_id', as_index=False).tail(8)
+Y_train_df = Y_df.drop(Y_test_df.index)
+```
+
+## 2. Computing different models for different hierarchies
+
+In this section, we illustrate how to fit a different type of model for
+each level of the hierarchy. In particular, for each level, we will fit
+the following models:
+
+- **Country**: `AutoETS` model from `StatsForecast`.
+- **Country/State**: `HistGradientBoostingRegressor` model from
+ `scikit-learn` through the `MLForecast` API.
+- **Country/State/Region**: `NBEATS` model from `NeuralForecast`.
+
+
+```python
+from statsforecast.core import StatsForecast
+from statsforecast.models import AutoETS
+
+from mlforecast import MLForecast
+from sklearn.ensemble import HistGradientBoostingRegressor
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import NBEATS
+```
+
+This `fit_predict_any_models` function is a helper function for training
+and forecasting with models from `StatsForecast`, `MLForecast`, and
+`NeuralForecast`.
+
+
+```python
+def fit_predict_any_models(models, df, h):
+ if isinstance(models, StatsForecast):
+ yhat = models.forecast(df=df, h=h, fitted=True)
+ yfitted = models.forecast_fitted_values()
+ elif isinstance(models, MLForecast):
+ models.fit(df, fitted=True)
+ yhat = models.predict(new_df=df, h=h)
+ yfitted = models.forecast_fitted_values()
+
+ elif isinstance(models, NeuralForecast):
+ models.fit(df=df, val_size=h)
+ yhat = models.predict()
+ yfitted = models.predict_insample(step_size=h)
+ yfitted = yfitted.drop(columns=['cutoff'])
+ else:
+ raise ValueError("Model is not a StatsForecast, MLForecast or NeuralForecast object.")
+
+ return yhat, yfitted
+```
+
+We now define the models that we want to use.
+
+
+```python
+h = 8
+stat_models = StatsForecast(models=[AutoETS(season_length=4, model='ZZA')], freq='QS', n_jobs=-1)
+ml_models = MLForecast(models = [HistGradientBoostingRegressor()], freq='QS', lags=[1, 4])
+neural_models = NeuralForecast(models=[NBEATS(h=h, input_size=16)],freq='QS')
+```
+
+We have defined a hierarchy consisting of three levels. We will use the
+different model types for each of the levels in the hierarchy.
+
+
+```python
+models = {
+ 'Country': stat_models,
+ 'Country/State': ml_models,
+ 'Country/State/Region': neural_models
+}
+```
+
+To fit each model and create forecasts with it, we loop over the
+timeseries that are present in each level of the hierarchy, using the
+`tags` we created earlier using the
+[`aggregate`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#aggregate)
+function.
+
+
+```python
+Y_hat = []
+Y_fitted = []
+# We loop through the tags to fit and predict for each level of the hierarchy.
+for key, value in tags.items():
+ # We filter the training dataframe for the current level of the hierarchy.
+ df_level = Y_train_df.query('unique_id.isin(@value)')
+ # We fit and predict using the corresponding model for the current level.
+ yhat_level, yfitted_level = fit_predict_any_models(models[key], df_level, h=h)
+ # We add the predictions for this level
+ Y_hat.append(yhat_level)
+ Y_fitted.append(yfitted_level)
+
+# Concatenate the predictions for all levels into a single DataFrame
+Y_hat_df = pd.concat(Y_hat, ignore_index=True)
+Y_fitted_df = pd.concat(Y_fitted, ignore_index=True)
+```
+
+We have now created forecasts for different levels of the hierarchy,
+using different model types. Let’s look at the forecasts.
+
+
+```python
+Y_hat_df.head(10)
+```
+
+| | unique_id | ds | AutoETS | HistGradientBoostingRegressor | NBEATS |
+|----|----|----|----|----|----|
+| 0 | Australia | 2016-01-01 | 25990.068004 | NaN | NaN |
+| 1 | Australia | 2016-04-01 | 24458.490282 | NaN | NaN |
+| 2 | Australia | 2016-07-01 | 23974.055984 | NaN | NaN |
+| 3 | Australia | 2016-10-01 | 24563.454495 | NaN | NaN |
+| 4 | Australia | 2017-01-01 | 25990.068004 | NaN | NaN |
+| 5 | Australia | 2017-04-01 | 24458.490282 | NaN | NaN |
+| 6 | Australia | 2017-07-01 | 23974.055984 | NaN | NaN |
+| 7 | Australia | 2017-10-01 | 24563.454495 | NaN | NaN |
+| 8 | Australia/ACT | 2016-01-01 | NaN | 571.433902 | NaN |
+| 9 | Australia/ACT | 2016-04-01 | NaN | 548.060532 | NaN |
+
+As you can see, `AutoETS` only has entries for the
+`unique_id=Australia`, which is because we only created forecasts for
+the level `Country` using `AutoETS`.
+
+Secondly, we also only have forecasts using
+`HistGradientBoostingRegressor` for timeseries in the level
+`Country/State`, again as we only created forecasts for the level
+`Country/State` using `HistGradientBoostingRegressor`.
+
+Finally, `NBEATS` shows no forecasts at all in this view, but when we
+look at the tail of the predictions we see that `NBEATS` only has
+forecasts for the level `Country/State/Region`, which was also what we
+intended to create.
+
+
+```python
+Y_hat_df.tail(10)
+```
+
+| | unique_id | ds | AutoETS | HistGradientBoostingRegressor | NBEATS |
+|----|----|----|----|----|----|
+| 670 | Australia/Western Australia/Australia's South ... | 2017-07-01 | NaN | NaN | 416.720154 |
+| 671 | Australia/Western Australia/Australia's South ... | 2017-10-01 | NaN | NaN | 605.681030 |
+| 672 | Australia/Western Australia/Experience Perth | 2016-01-01 | NaN | NaN | 1139.827393 |
+| 673 | Australia/Western Australia/Experience Perth | 2016-04-01 | NaN | NaN | 1017.152527 |
+| 674 | Australia/Western Australia/Experience Perth | 2016-07-01 | NaN | NaN | 917.289673 |
+| 675 | Australia/Western Australia/Experience Perth | 2016-10-01 | NaN | NaN | 1141.263062 |
+| 676 | Australia/Western Australia/Experience Perth | 2017-01-01 | NaN | NaN | 1134.063477 |
+| 677 | Australia/Western Australia/Experience Perth | 2017-04-01 | NaN | NaN | 1021.346558 |
+| 678 | Australia/Western Australia/Experience Perth | 2017-07-01 | NaN | NaN | 839.628418 |
+| 679 | Australia/Western Australia/Experience Perth | 2017-10-01 | NaN | NaN | 972.161499 |
+
+## 3. Reconcile forecasts
+
+First, we need to make sure we have one forecast column containing all
+the forecasts across all the levels, as we want to reconcile the
+forecasts across the levels. We do so by taking the mean across the
+forecast columns. In this case, because there’s only a single entry for
+each unique_id, it would be equivalent to just combine or sum the
+forecast columns. However, you might want to use more than one model
+*per level* in the hierarchy. In that case, you’d need to think about
+how to ensemble the multiple forecasts - a simple mean ensemble
+generally works well in those cases, so you can directly use the below
+code also for the more complex case where you have multiple models for
+each level.
+
+
+```python
+forecast_cols = [col for col in Y_hat_df.columns if col not in ['unique_id', 'ds', 'y']]
+Y_hat_df["all_forecasts"] = Y_hat_df[forecast_cols].mean(axis=1)
+Y_fitted_df["all_forecasts"] = Y_fitted_df[forecast_cols].mean(axis=1)
+```
+
+As we can see, we now have a single column `all_forecasts` that includes
+the forecasts across all the levels:
+
+
+```python
+Y_hat_df.head(10)
+```
+
+| | unique_id | ds | AutoETS | HistGradientBoostingRegressor | NBEATS | all_forecasts |
+|----|----|----|----|----|----|----|
+| 0 | Australia | 2016-01-01 | 25990.068004 | NaN | NaN | 25990.068004 |
+| 1 | Australia | 2016-04-01 | 24458.490282 | NaN | NaN | 24458.490282 |
+| 2 | Australia | 2016-07-01 | 23974.055984 | NaN | NaN | 23974.055984 |
+| 3 | Australia | 2016-10-01 | 24563.454495 | NaN | NaN | 24563.454495 |
+| 4 | Australia | 2017-01-01 | 25990.068004 | NaN | NaN | 25990.068004 |
+| 5 | Australia | 2017-04-01 | 24458.490282 | NaN | NaN | 24458.490282 |
+| 6 | Australia | 2017-07-01 | 23974.055984 | NaN | NaN | 23974.055984 |
+| 7 | Australia | 2017-10-01 | 24563.454495 | NaN | NaN | 24563.454495 |
+| 8 | Australia/ACT | 2016-01-01 | NaN | 571.433902 | NaN | 571.433902 |
+| 9 | Australia/ACT | 2016-04-01 | NaN | 548.060532 | NaN | 548.060532 |
+
+We are now ready to make the forecasts coherent using the
+[`HierarchicalReconciliation`](https://Nixtla.github.io/hierarchicalforecast/src/core.html#hierarchicalreconciliation)
+class. In this example we use
+[`BottomUp`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#bottomup),
+`MinTrace(mint_shrink)`, `TopDown(forecast_proportions)` reconcilers.
+
+
+```python
+from hierarchicalforecast.methods import BottomUp, MinTrace, TopDown
+from hierarchicalforecast.core import HierarchicalReconciliation
+```
+
+
+```python
+reconcilers = [
+ BottomUp(),
+ MinTrace(method='mint_shrink'),
+ TopDown(method='forecast_proportions')
+]
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+Y_rec_df = hrec.reconcile(Y_hat_df=Y_hat_df[["unique_id", "ds", "all_forecasts"]], Y_df=Y_fitted_df[["unique_id", "ds", "y", "all_forecasts"]], S_df=S_df, tags=tags)
+```
+
+The dataframe `Y_rec_df` contains the reconciled forecasts.
+
+
+```python
+Y_rec_df.head()
+```
+
+| | unique_id | ds | all_forecasts | all_forecasts/BottomUp | all_forecasts/MinTrace_method-mint_shrink | all_forecasts/TopDown_method-forecast_proportions |
+|----|----|----|----|----|----|----|
+| 0 | Australia | 2016-01-01 | 25990.068004 | 24916.914513 | 25959.517939 | 25990.068004 |
+| 1 | Australia | 2016-04-01 | 24458.490282 | 22867.133526 | 24656.012177 | 24458.490282 |
+| 2 | Australia | 2016-07-01 | 23974.055984 | 22845.050221 | 24933.182437 | 23974.055984 |
+| 3 | Australia | 2016-10-01 | 24563.454495 | 23901.916314 | 26382.869677 | 24563.454495 |
+| 4 | Australia | 2017-01-01 | 25990.068004 | 25246.089151 | 26923.282464 | 25990.068004 |
+
+## 4. Evaluation
+
+The `HierarchicalForecast` package includes an
+[`evaluate`](https://Nixtla.github.io/hierarchicalforecast/src/evaluation.html#evaluate)
+function to evaluate the different hierarchies. To evaluate models we
+use `mase` metric and compare it to base predictions.
+
+
+```python
+from hierarchicalforecast.evaluation import evaluate
+from utilsforecast.losses import mase
+from functools import partial
+```
+
+
+```python
+eval_tags = {}
+eval_tags['Total'] = tags['Country']
+eval_tags['State'] = tags['Country/State']
+eval_tags['Regions'] = tags['Country/State/Region']
+
+df = Y_rec_df.merge(Y_test_df, on=['unique_id', 'ds'])
+evaluation = evaluate(df = df,
+ tags = eval_tags,
+ train_df = Y_train_df,
+ metrics = [partial(mase, seasonality=4)])
+```
+
+
+```python
+evaluation
+```
+
+| | level | metric | all_forecasts | all_forecasts/BottomUp | all_forecasts/MinTrace_method-mint_shrink | all_forecasts/TopDown_method-forecast_proportions |
+|----|----|----|----|----|----|----|
+| 0 | Total | mase | 1.589074 | 3.002085 | 0.440261 | 1.589074 |
+| 1 | State | mase | 2.166374 | 1.905035 | 1.882345 | 2.361169 |
+| 2 | Regions | mase | 1.342429 | 1.342429 | 1.423867 | 1.458773 |
+| 3 | Overall | mase | 1.422878 | 1.414905 | 1.455446 | 1.545237 |
+
+We find that:
+
+- **No Single Best Method**: The results indicate that there is no
+ universally superior reconciliation method. The optimal choice
+ depends on which level of the hierarchy is most important.
+- **MinTrace for Country and Country/State**: The
+ `MinTrace(mint_shrink)` reconciler shows best performance for the
+ upper levels of the hierarchy, reducing the MASE from 1.59 (base
+ forecast) to just 0.44.
+- **BottomUp for Country/State/Region and Overall**: The
+ [`BottomUp`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#bottomup)
+ method preserves only the NBEATS forecast of the most granular
+ **Country/State/Regions** level, and aggregates those forecasts for
+ the upper levels. It yields the **best Overall MASE score**.
+
+## 6. Recap
+
+This notebook demonstrated the power and flexibility of
+HierarchicalForecast in a multi-model forecasting scenario.
+
+In this example we fitted:
+
+- `StatsForecast` with `AutoETS` model for the **Country** level.
+- `MLForecast` with `HistGradientBoostingRegressor` model for the
+ **Country/State** level.
+- `NeuralForecast` with `NBEATS` model for the
+ **Country/State/Region** level.
+
+We then combined the results into a single prediction.
+
+For the reconciliation of the forecasts, we used
+[`HierarchicalReconciliation`](https://Nixtla.github.io/hierarchicalforecast/src/core.html#hierarchicalreconciliation)
+with three different methods:
+
+- [`BottomUp`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#bottomup)
+- `MinTrace(method='mint_shrink')`
+- `TopDown(method='forecast_proportions')`
+
+Finally, we evaluated the performance of these reconciliation methods.
+
diff --git a/hierarchicalforecast/examples/australiandomestictourism-permbu-intervals.html.mdx b/hierarchicalforecast/examples/australiandomestictourism-permbu-intervals.html.mdx
new file mode 100644
index 00000000..935806ec
--- /dev/null
+++ b/hierarchicalforecast/examples/australiandomestictourism-permbu-intervals.html.mdx
@@ -0,0 +1,299 @@
+---
+output-file: australiandomestictourism-permbu-intervals.html
+title: PERMBU
+---
+
+
+
+
+In many cases, only the time series at the lowest level of the
+hierarchies (bottom time series) are available. `HierarchicalForecast`
+has tools to create time series for all hierarchies and also allows you
+to calculate prediction intervals for all hierarchies. In this notebook
+we will see how to do it.
+
+
+```python
+!pip install hierarchicalforecast statsforecast
+```
+
+
+```python
+import pandas as pd
+
+# compute base forecast no coherent
+from statsforecast.models import AutoARIMA
+from statsforecast.core import StatsForecast
+
+#obtain hierarchical reconciliation methods and evaluation
+from hierarchicalforecast.methods import BottomUp, MinTrace
+from hierarchicalforecast.utils import aggregate, HierarchicalPlot
+from hierarchicalforecast.core import HierarchicalReconciliation
+```
+
+## Aggregate bottom time series
+
+In this example we will use the
+[Tourism](https://otexts.com/fpp3/tourism.html) dataset from the
+[Forecasting: Principles and Practice](https://otexts.com/fpp3/) book.
+The dataset only contains the time series at the lowest level, so we
+need to create the time series for all hierarchies.
+
+
+```python
+Y_df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/tourism.csv')
+Y_df = Y_df.rename({'Trips': 'y', 'Quarter': 'ds'}, axis=1)
+Y_df.insert(0, 'Country', 'Australia')
+Y_df = Y_df[['Country', 'Region', 'State', 'Purpose', 'ds', 'y']]
+Y_df['ds'] = Y_df['ds'].str.replace(r'(\d+) (Q\d)', r'\1-\2', regex=True)
+Y_df['ds'] = pd.PeriodIndex(Y_df["ds"], freq='Q').to_timestamp()
+Y_df.head()
+```
+
+| | Country | Region | State | Purpose | ds | y |
+|-----|-----------|----------|-----------------|----------|------------|------------|
+| 0 | Australia | Adelaide | South Australia | Business | 1998-01-01 | 135.077690 |
+| 1 | Australia | Adelaide | South Australia | Business | 1998-04-01 | 109.987316 |
+| 2 | Australia | Adelaide | South Australia | Business | 1998-07-01 | 166.034687 |
+| 3 | Australia | Adelaide | South Australia | Business | 1998-10-01 | 127.160464 |
+| 4 | Australia | Adelaide | South Australia | Business | 1999-01-01 | 137.448533 |
+
+The dataset can be grouped in the following strictly hierarchical
+structure.
+
+
+```python
+spec = [
+ ['Country'],
+ ['Country', 'State'],
+ ['Country', 'State', 'Region']
+]
+```
+
+Using the
+[`aggregate`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#aggregate)
+function from `HierarchicalForecast` we can get the full set of time
+series.
+
+
+```python
+Y_df, S_df, tags = aggregate(df=Y_df, spec=spec)
+```
+
+
+```python
+Y_df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|--------------|
+| 0 | Australia | 1998-01-01 | 23182.197269 |
+| 1 | Australia | 1998-04-01 | 20323.380067 |
+| 2 | Australia | 1998-07-01 | 19826.640511 |
+| 3 | Australia | 1998-10-01 | 20830.129891 |
+| 4 | Australia | 1999-01-01 | 22087.353380 |
+
+
+```python
+S_df.iloc[:5, :5]
+```
+
+| | unique_id | Australia/ACT/Canberra | Australia/New South Wales/Blue Mountains | Australia/New South Wales/Capital Country | Australia/New South Wales/Central Coast |
+|----|----|----|----|----|----|
+| 0 | Australia | 1.0 | 1.0 | 1.0 | 1.0 |
+| 1 | Australia/ACT | 1.0 | 0.0 | 0.0 | 0.0 |
+| 2 | Australia/New South Wales | 0.0 | 1.0 | 1.0 | 1.0 |
+| 3 | Australia/Northern Territory | 0.0 | 0.0 | 0.0 | 0.0 |
+| 4 | Australia/Queensland | 0.0 | 0.0 | 0.0 | 0.0 |
+
+
+```python
+tags['Country/State']
+```
+
+``` text
+array(['Australia/ACT', 'Australia/New South Wales',
+ 'Australia/Northern Territory', 'Australia/Queensland',
+ 'Australia/South Australia', 'Australia/Tasmania',
+ 'Australia/Victoria', 'Australia/Western Australia'], dtype=object)
+```
+
+We can visualize the `S` matrix and the data using the
+[`HierarchicalPlot`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#hierarchicalplot)
+class as follows.
+
+
+```python
+hplot = HierarchicalPlot(S=S_df, tags=tags)
+```
+
+
+```python
+hplot.plot_summing_matrix()
+```
+
+
+
+
+```python
+hplot.plot_hierarchically_linked_series(
+ bottom_series='Australia/ACT/Canberra',
+ Y_df=Y_df
+)
+```
+
+
+
+### Split Train/Test sets
+
+We use the final two years (8 quarters) as test set.
+
+
+```python
+Y_test_df = Y_df.groupby('unique_id', as_index=False).tail(8)
+Y_train_df = Y_df.drop(Y_test_df.index)
+```
+
+
+```python
+Y_train_df.groupby('unique_id').size()
+```
+
+``` text
+unique_id
+Australia 72
+Australia/ACT 72
+Australia/ACT/Canberra 72
+Australia/New South Wales 72
+Australia/New South Wales/Blue Mountains 72
+ ..
+Australia/Western Australia/Australia's Coral Coast 72
+Australia/Western Australia/Australia's Golden Outback 72
+Australia/Western Australia/Australia's North West 72
+Australia/Western Australia/Australia's South West 72
+Australia/Western Australia/Experience Perth 72
+Length: 85, dtype: int64
+```
+
+## Computing base forecasts
+
+The following cell computes the **base forecasts** for each time series
+in `Y_df` using the `AutoARIMA` and model. Observe that `Y_hat_df`
+contains the forecasts but they are not coherent. To reconcile the
+prediction intervals we need to calculate the uncoherent intervals using
+the `level` argument of `StatsForecast`.
+
+
+```python
+fcst = StatsForecast(models=[AutoARIMA(season_length=4)],
+ freq='QS', n_jobs=-1)
+Y_hat_df = fcst.forecast(df=Y_train_df, h=8, fitted=True, level=[80, 90])
+Y_fitted_df = fcst.forecast_fitted_values()
+```
+
+## Reconcile forecasts and compute prediction intervals using PERMBU
+
+The following cell makes the previous forecasts coherent using the
+[`HierarchicalReconciliation`](https://Nixtla.github.io/hierarchicalforecast/src/core.html#hierarchicalreconciliation)
+class. In this example we use
+[`BottomUp`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#bottomup)
+and
+[`MinTrace`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#mintrace).
+If you want to calculate prediction intervals, you have to use the
+`level` argument as follows and also `intervals_method='permbu'`.
+
+
+```python
+reconcilers = [
+ BottomUp(),
+ MinTrace(method='mint_shrink'),
+ MinTrace(method='ols')
+]
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+Y_rec_df = hrec.reconcile(Y_hat_df=Y_hat_df, Y_df=Y_fitted_df,
+ S_df=S_df, tags=tags,
+ level=[80, 90], intervals_method='permbu')
+```
+
+The dataframe `Y_rec_df` contains the reconciled forecasts.
+
+
+```python
+Y_rec_df.head()
+```
+
+| | unique_id | ds | AutoARIMA | AutoARIMA-lo-90 | AutoARIMA-lo-80 | AutoARIMA-hi-80 | AutoARIMA-hi-90 | AutoARIMA/BottomUp | AutoARIMA/BottomUp-lo-90 | AutoARIMA/BottomUp-lo-80 | ... | AutoARIMA/MinTrace_method-mint_shrink | AutoARIMA/MinTrace_method-mint_shrink-lo-90 | AutoARIMA/MinTrace_method-mint_shrink-lo-80 | AutoARIMA/MinTrace_method-mint_shrink-hi-80 | AutoARIMA/MinTrace_method-mint_shrink-hi-90 | AutoARIMA/MinTrace_method-ols | AutoARIMA/MinTrace_method-ols-lo-90 | AutoARIMA/MinTrace_method-ols-lo-80 | AutoARIMA/MinTrace_method-ols-hi-80 | AutoARIMA/MinTrace_method-ols-hi-90 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | Australia | 2016-01-01 | 26212.553553 | 24705.948180 | 25038.715077 | 27386.392029 | 27719.158927 | 24955.501571 | 24143.056131 | 24387.230200 | ... | 25413.657606 | 24705.682710 | 24905.677772 | 25928.334367 | 26050.232961 | 26142.818016 | 25525.081721 | 25656.537995 | 26606.345032 | 26832.423921 |
+| 1 | Australia | 2016-04-01 | 25033.667125 | 23337.267588 | 23711.954696 | 26355.379554 | 26730.066662 | 23421.312868 | 22762.045247 | 22904.087197 | ... | 24058.906411 | 23486.828548 | 23627.152623 | 24659.405484 | 24847.778503 | 24946.338649 | 24297.061230 | 24434.805048 | 25535.549040 | 25640.659918 |
+| 2 | Australia | 2016-07-01 | 24507.027198 | 22640.028798 | 23052.396413 | 25961.657983 | 26374.025599 | 22807.706826 | 22065.402373 | 22223.120404 | ... | 23438.863893 | 22672.658701 | 22888.299153 | 23971.724733 | 24179.548677 | 24407.245003 | 23712.841797 | 23834.054327 | 25027.073615 | 25189.869286 |
+| 3 | Australia | 2016-10-01 | 25598.928613 | 23575.665243 | 24022.547410 | 27175.309816 | 27622.191983 | 23471.845870 | 22677.593575 | 22892.328939 | ... | 24322.049398 | 23619.419712 | 23682.803746 | 24847.299228 | 25028.345572 | 25496.855604 | 24740.210465 | 24923.560783 | 26094.250414 | 26273.617732 |
+| 4 | Australia | 2017-01-01 | 26982.576796 | 24669.535238 | 25180.421285 | 28784.732308 | 29295.618354 | 24668.735931 | 23760.842072 | 23964.283124 | ... | 25520.163549 | 24720.304392 | 24910.106650 | 26170.552678 | 26347.181903 | 26853.231907 | 26045.213677 | 26149.753374 | 27502.499674 | 27733.985566 |
+
+## Plot forecasts
+
+Then we can plot the probabilist forecasts using the following function.
+
+
+```python
+plot_df = Y_df.merge(Y_rec_df, on=['unique_id', 'ds'], how="outer")
+```
+
+### Plot single time series
+
+
+```python
+hplot.plot_series(
+ series='Australia',
+ Y_df=plot_df,
+ models=['y', 'AutoARIMA',
+ 'AutoARIMA/MinTrace_method-ols',
+ 'AutoARIMA/BottomUp'
+ ],
+ level=[80]
+)
+```
+
+
+
+### Plot hierarchichally linked time series
+
+
+```python
+hplot.plot_hierarchically_linked_series(
+ bottom_series='Australia/Western Australia/Experience Perth',
+ Y_df=plot_df,
+ models=['y', 'AutoARIMA', 'AutoARIMA/MinTrace_method-ols', 'AutoARIMA/BottomUp'],
+ level=[80]
+)
+```
+
+
+
+
+```python
+# ACT only has Canberra
+hplot.plot_hierarchically_linked_series(
+ bottom_series='Australia/ACT/Canberra',
+ Y_df=plot_df,
+ models=['y', 'AutoARIMA/MinTrace_method-mint_shrink'],
+ level=[80, 90]
+)
+```
+
+
+
+### References
+
+- [Hyndman, R.J., & Athanasopoulos, G. (2021). “Forecasting:
+ principles and practice, 3rd edition: Chapter 11: Forecasting
+ hierarchical and grouped series.”. OTexts: Melbourne, Australia.
+ OTexts.com/fpp3 Accessed on July
+ 2022.](https://otexts.com/fpp3/hierarchical.html)
+- [Shanika L. Wickramasuriya, George Athanasopoulos, and Rob J.
+ Hyndman. Optimal forecast reconciliation for hierarchical and
+ grouped time series through trace minimization.Journal of the
+ American Statistical Association, 114(526):804–819, 2019. doi:
+ 10.1080/01621459.2018.1448825. URL
+ https://robjhyndman.com/publications/mint/.](https://robjhyndman.com/publications/mint/)
+
diff --git a/hierarchicalforecast/examples/australiandomestictourism.html.mdx b/hierarchicalforecast/examples/australiandomestictourism.html.mdx
new file mode 100644
index 00000000..afbd2960
--- /dev/null
+++ b/hierarchicalforecast/examples/australiandomestictourism.html.mdx
@@ -0,0 +1,340 @@
+---
+description: Geographical Hierarchical Forecasting on Australian Tourism Data
+output-file: australiandomestictourism.html
+title: Geographical Aggregation (Tourism)
+---
+
+
+In many applications, a set of time series is hierarchically organized.
+Examples include the presence of geographic levels, products, or
+categories that define different types of aggregations. In such
+scenarios, forecasters are often required to provide predictions for all
+disaggregate and aggregate series. A natural desire is for those
+predictions to be **“coherent”**, that is, for the bottom series to add
+up precisely to the forecasts of the aggregated series.
+
+In this notebook we present an example on how to use
+`HierarchicalForecast` to produce coherent forecasts between
+geographical levels. We will use the classic Australian Domestic Tourism
+(`Tourism`) dataset, which contains monthly time series of the number of
+visitors to each state of Australia.
+
+We will first load the `Tourism` data and produce base forecasts using
+an `AutoETS` model from `StatsForecast`, and then reconciliate the
+forecasts with several reconciliation algorithms from
+`HierarchicalForecast`. Finally, we show the performance is comparable
+with the results reported by the [Forecasting: Principles and
+Practice](https://otexts.com/fpp3/tourism.html) which uses the R package
+[fable](https://github.com/tidyverts/fable).
+
+You can run these experiments using CPU or GPU with Google Colab.
+
+
+
+
+```python
+!pip install hierarchicalforecast statsforecast
+```
+
+## 1. Load and Process Data
+
+In this example we will use the
+[Tourism](https://otexts.com/fpp3/tourism.html) dataset from the
+[Forecasting: Principles and Practice](https://otexts.com/fpp3/) book.
+
+The dataset only contains the time series at the lowest level, so we
+need to create the time series for all hierarchies.
+
+
+```python
+import numpy as np
+import pandas as pd
+```
+
+
+```python
+Y_df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/tourism.csv')
+Y_df = Y_df.rename({'Trips': 'y', 'Quarter': 'ds'}, axis=1)
+Y_df.insert(0, 'Country', 'Australia')
+Y_df = Y_df[['Country', 'Region', 'State', 'Purpose', 'ds', 'y']]
+Y_df['ds'] = Y_df['ds'].str.replace(r'(\d+) (Q\d)', r'\1-\2', regex=True)
+Y_df['ds'] = pd.PeriodIndex(Y_df["ds"], freq='Q').to_timestamp()
+Y_df.head()
+```
+
+| | Country | Region | State | Purpose | ds | y |
+|-----|-----------|----------|-----------------|----------|------------|------------|
+| 0 | Australia | Adelaide | South Australia | Business | 1998-01-01 | 135.077690 |
+| 1 | Australia | Adelaide | South Australia | Business | 1998-04-01 | 109.987316 |
+| 2 | Australia | Adelaide | South Australia | Business | 1998-07-01 | 166.034687 |
+| 3 | Australia | Adelaide | South Australia | Business | 1998-10-01 | 127.160464 |
+| 4 | Australia | Adelaide | South Australia | Business | 1999-01-01 | 137.448533 |
+
+The dataset can be grouped in the following non-strictly hierarchical
+structure.
+
+
+```python
+spec = [
+ ['Country'],
+ ['Country', 'State'],
+ ['Country', 'Purpose'],
+ ['Country', 'State', 'Region'],
+ ['Country', 'State', 'Purpose'],
+ ['Country', 'State', 'Region', 'Purpose']
+]
+```
+
+Using the
+[`aggregate`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#aggregate)
+function from `HierarchicalForecast` we can get the full set of time
+series.
+
+
+```python
+from hierarchicalforecast.utils import aggregate
+```
+
+
+```python
+Y_df, S_df, tags = aggregate(Y_df, spec)
+```
+
+
+```python
+Y_df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|--------------|
+| 0 | Australia | 1998-01-01 | 23182.197269 |
+| 1 | Australia | 1998-04-01 | 20323.380067 |
+| 2 | Australia | 1998-07-01 | 19826.640511 |
+| 3 | Australia | 1998-10-01 | 20830.129891 |
+| 4 | Australia | 1999-01-01 | 22087.353380 |
+
+
+```python
+S_df.iloc[:5, :5]
+```
+
+| | unique_id | Australia/ACT/Canberra/Business | Australia/ACT/Canberra/Holiday | Australia/ACT/Canberra/Other | Australia/ACT/Canberra/Visiting |
+|----|----|----|----|----|----|
+| 0 | Australia | 1.0 | 1.0 | 1.0 | 1.0 |
+| 1 | Australia/ACT | 1.0 | 1.0 | 1.0 | 1.0 |
+| 2 | Australia/New South Wales | 0.0 | 0.0 | 0.0 | 0.0 |
+| 3 | Australia/Northern Territory | 0.0 | 0.0 | 0.0 | 0.0 |
+| 4 | Australia/Queensland | 0.0 | 0.0 | 0.0 | 0.0 |
+
+
+```python
+tags['Country/Purpose']
+```
+
+``` text
+array(['Australia/Business', 'Australia/Holiday', 'Australia/Other',
+ 'Australia/Visiting'], dtype=object)
+```
+
+### Split Train/Test sets
+
+We use the final two years (8 quarters) as test set.
+
+
+```python
+Y_test_df = Y_df.groupby('unique_id', as_index=False).tail(8)
+Y_train_df = Y_df.drop(Y_test_df.index)
+```
+
+
+```python
+Y_train_df.groupby('unique_id').size()
+```
+
+``` text
+unique_id
+Australia 72
+Australia/ACT 72
+Australia/ACT/Business 72
+Australia/ACT/Canberra 72
+Australia/ACT/Canberra/Business 72
+ ..
+Australia/Western Australia/Experience Perth/Other 72
+Australia/Western Australia/Experience Perth/Visiting 72
+Australia/Western Australia/Holiday 72
+Australia/Western Australia/Other 72
+Australia/Western Australia/Visiting 72
+Length: 425, dtype: int64
+```
+
+## 2. Computing base forecasts
+
+The following cell computes the **base forecasts** for each time series
+in `Y_df` using the `ETS` model. Observe that `Y_hat_df` contains the
+forecasts but they are not coherent.
+
+
+```python
+from statsforecast.models import AutoETS
+from statsforecast.core import StatsForecast
+```
+
+
+```python
+fcst = StatsForecast(models=[AutoETS(season_length=4, model='ZZA')],
+ freq='QS', n_jobs=-1)
+Y_hat_df = fcst.forecast(df=Y_train_df, h=8, fitted=True)
+Y_fitted_df = fcst.forecast_fitted_values()
+```
+
+## 3. Reconcile forecasts
+
+The following cell makes the previous forecasts coherent using the
+[`HierarchicalReconciliation`](https://Nixtla.github.io/hierarchicalforecast/src/core.html#hierarchicalreconciliation)
+class. Since the hierarchy structure is not strict, we can’t use methods
+such as
+[`TopDown`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#topdown)
+or
+[`MiddleOut`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#middleout).
+In this example we use
+[`BottomUp`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#bottomup)
+and
+[`MinTrace`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#mintrace).
+
+
+```python
+from hierarchicalforecast.methods import BottomUp, MinTrace
+from hierarchicalforecast.core import HierarchicalReconciliation
+```
+
+
+```python
+reconcilers = [
+ BottomUp(),
+ MinTrace(method='mint_shrink'),
+ MinTrace(method='ols')
+]
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+Y_rec_df = hrec.reconcile(Y_hat_df=Y_hat_df, Y_df=Y_fitted_df, S_df=S_df, tags=tags)
+```
+
+The dataframe `Y_rec_df` contains the reconciled forecasts.
+
+
+```python
+Y_rec_df.head()
+```
+
+| | unique_id | ds | AutoETS | AutoETS/BottomUp | AutoETS/MinTrace_method-mint_shrink | AutoETS/MinTrace_method-ols |
+|----|----|----|----|----|----|----|
+| 0 | Australia | 2016-01-01 | 25990.068004 | 24381.911737 | 25428.089783 | 25894.399067 |
+| 1 | Australia | 2016-04-01 | 24458.490282 | 22903.895964 | 23914.271400 | 24357.301898 |
+| 2 | Australia | 2016-07-01 | 23974.055984 | 22412.265739 | 23428.462394 | 23865.910647 |
+| 3 | Australia | 2016-10-01 | 24563.454495 | 23127.349578 | 24089.845955 | 24470.782393 |
+| 4 | Australia | 2017-01-01 | 25990.068004 | 24518.118006 | 25545.358678 | 25901.362283 |
+
+## 4. Evaluation
+
+The `HierarchicalForecast` package includes an
+[`evaluate`](https://Nixtla.github.io/hierarchicalforecast/src/evaluation.html#evaluate)
+function to evaluate the different hierarchies and also is capable of
+compute scaled metrics compared to a benchmark model.
+
+
+```python
+from hierarchicalforecast.evaluation import evaluate
+from utilsforecast.losses import rmse, mase
+from functools import partial
+```
+
+
+```python
+eval_tags = {}
+eval_tags['Total'] = tags['Country']
+eval_tags['Purpose'] = tags['Country/Purpose']
+eval_tags['State'] = tags['Country/State']
+eval_tags['Regions'] = tags['Country/State/Region']
+eval_tags['Bottom'] = tags['Country/State/Region/Purpose']
+
+df = Y_rec_df.merge(Y_test_df, on=['unique_id', 'ds'])
+evaluation = evaluate(df = df,
+ tags = eval_tags,
+ train_df = Y_train_df,
+ metrics = [rmse,
+ partial(mase, seasonality=4)])
+
+evaluation.columns = ['level', 'metric', 'Base', 'BottomUp', 'MinTrace(mint_shrink)', 'MinTrace(ols)']
+numeric_cols = evaluation.select_dtypes(include="number").columns
+evaluation[numeric_cols] = evaluation[numeric_cols].map('{:.2f}'.format).astype(np.float64)
+```
+
+### RMSE
+
+The following table shows the performance measured using RMSE across
+levels for each reconciliation method.
+
+
+```python
+evaluation.query('metric == "rmse"')
+```
+
+| | level | metric | Base | BottomUp | MinTrace(mint_shrink) | MinTrace(ols) |
+|-----|---------|--------|---------|----------|-----------------------|---------------|
+| 0 | Total | rmse | 1743.29 | 3028.62 | 2112.73 | 1818.94 |
+| 2 | Purpose | rmse | 534.75 | 791.19 | 577.14 | 515.53 |
+| 4 | State | rmse | 308.15 | 413.39 | 316.82 | 287.32 |
+| 6 | Regions | rmse | 51.66 | 55.13 | 46.55 | 46.28 |
+| 8 | Bottom | rmse | 19.37 | 19.37 | 17.80 | 18.19 |
+| 10 | Overall | rmse | 41.12 | 49.82 | 40.47 | 38.75 |
+
+### MASE
+
+The following table shows the performance measured using MASE across
+levels for each reconciliation method.
+
+
+```python
+evaluation.query('metric == "mase"')
+```
+
+| | level | metric | Base | BottomUp | MinTrace(mint_shrink) | MinTrace(ols) |
+|-----|---------|--------|------|----------|-----------------------|---------------|
+| 1 | Total | mase | 1.59 | 3.16 | 2.06 | 1.67 |
+| 3 | Purpose | mase | 1.32 | 2.28 | 1.48 | 1.25 |
+| 5 | State | mase | 1.39 | 1.90 | 1.40 | 1.25 |
+| 7 | Regions | mase | 1.12 | 1.19 | 1.01 | 0.99 |
+| 9 | Bottom | mase | 0.98 | 0.98 | 0.94 | 1.01 |
+| 11 | Overall | mase | 1.02 | 1.06 | 0.97 | 1.02 |
+
+### Comparison fable
+
+Observe that we can recover the results reported by the [Forecasting:
+Principles and Practice](https://otexts.com/fpp3/tourism.html). The
+original results were calculated using the R package
+[fable](https://github.com/tidyverts/fable).
+
+
+
+Fable’s reconciliation
+results
+
+
+### References
+
+- [Hyndman, R.J., & Athanasopoulos, G. (2021). “Forecasting:
+ principles and practice, 3rd edition: Chapter 11: Forecasting
+ hierarchical and grouped series.”. OTexts: Melbourne, Australia.
+ OTexts.com/fpp3 Accessed on July
+ 2022.](https://otexts.com/fpp3/hierarchical.html)
+- [Rob Hyndman, Alan Lee, Earo Wang, Shanika Wickramasuriya, and
+ Maintainer Earo Wang (2021). “hts: Hierarchical and Grouped Time
+ Series”. URL https://CRAN.R-project.org/package=hts. R package
+ version
+ 0.3.1.](https://cran.r-project.org/web/packages/hts/index.html)
+- [Mitchell O’Hara-Wild, Rob Hyndman, Earo Wang, Gabriel Caceres,
+ Tim-Gunnar Hensel, and Timothy Hyndman (2021). “fable: Forecasting
+ Models for Tidy Time Series”. URL
+ https://CRAN.R-project.org/package=fable. R package version
+ 6.0.2.](https://CRAN.R-project.org/package=fable)
+
diff --git a/hierarchicalforecast/examples/australiandomestictourismcrosstemporal.html.mdx b/hierarchicalforecast/examples/australiandomestictourismcrosstemporal.html.mdx
new file mode 100644
index 00000000..b89551a2
--- /dev/null
+++ b/hierarchicalforecast/examples/australiandomestictourismcrosstemporal.html.mdx
@@ -0,0 +1,601 @@
+---
+description: Geographical and Temporal Hierarchical Forecasting on Australian Tourism Data
+output-file: australiandomestictourismcrosstemporal.html
+title: Geographical and Temporal Aggregation (Tourism)
+---
+
+
+In many applications, a set of time series is hierarchically organized.
+Examples include the presence of geographic levels, products, or
+categories that define different types of aggregations. In such
+scenarios, forecasters are often required to provide predictions for all
+disaggregate and aggregate series. A natural desire is for those
+predictions to be **“coherent”**, that is, for the bottom series to add
+up precisely to the forecasts of the aggregated series.
+
+In this notebook we present an example on how to use
+`HierarchicalForecast` to produce coherent forecasts between both
+geographical levels and temporal levels. We will use the classic
+Australian Domestic Tourism (`Tourism`) dataset, which contains monthly
+time series of the number of visitors to each state of Australia.
+
+We will first load the `Tourism` data and produce base forecasts using
+an `AutoETS` model from `StatsForecast`. Then, we reconciliate the
+forecasts with several reconciliation algorithms from
+`HierarchicalForecast` according to the cross-sectional geographical
+hierarchies. Finally, we reconciliate the forecasts in the temporal
+dimension according to a temporal hierarchy.
+
+You can run these experiments using CPU or GPU with Google Colab.
+
+
+
+
+```python
+!pip install hierarchicalforecast statsforecast
+```
+
+## 1. Load and Process Data
+
+In this example we will use the
+[Tourism](https://otexts.com/fpp3/tourism.html) dataset from the
+[Forecasting: Principles and Practice](https://otexts.com/fpp3/) book.
+
+The dataset only contains the time series at the lowest level, so we
+need to create the time series for all hierarchies.
+
+
+```python
+import numpy as np
+import pandas as pd
+```
+
+
+```python
+Y_df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/tourism.csv')
+Y_df = Y_df.rename({'Trips': 'y', 'Quarter': 'ds'}, axis=1)
+Y_df.insert(0, 'Country', 'Australia')
+Y_df = Y_df[['Country', 'Region', 'State', 'Purpose', 'ds', 'y']]
+Y_df['ds'] = Y_df['ds'].str.replace(r'(\d+) (Q\d)', r'\1-\2', regex=True)
+Y_df['ds'] = pd.PeriodIndex(Y_df["ds"], freq='Q').to_timestamp()
+Y_df.head()
+```
+
+| | Country | Region | State | Purpose | ds | y |
+|-----|-----------|----------|-----------------|----------|------------|------------|
+| 0 | Australia | Adelaide | South Australia | Business | 1998-01-01 | 135.077690 |
+| 1 | Australia | Adelaide | South Australia | Business | 1998-04-01 | 109.987316 |
+| 2 | Australia | Adelaide | South Australia | Business | 1998-07-01 | 166.034687 |
+| 3 | Australia | Adelaide | South Australia | Business | 1998-10-01 | 127.160464 |
+| 4 | Australia | Adelaide | South Australia | Business | 1999-01-01 | 137.448533 |
+
+## 2. Cross-sectional reconciliation
+
+### 2a. Aggregating the dataset according to cross-sectional hierarchy
+
+The dataset can be grouped in the following non-strictly hierarchical
+structure.
+
+
+```python
+spec = [
+ ['Country'],
+ ['Country', 'State'],
+ ['Country', 'Purpose'],
+ ['Country', 'State', 'Region'],
+ ['Country', 'State', 'Purpose'],
+ ['Country', 'State', 'Region', 'Purpose']
+]
+```
+
+Using the
+[`aggregate`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#aggregate)
+function from `HierarchicalForecast` we can get the full set of time
+series.
+
+
+```python
+from hierarchicalforecast.utils import aggregate
+```
+
+
+```python
+Y_df_cs, S_df_cs, tags_cs = aggregate(Y_df, spec)
+```
+
+
+```python
+Y_df_cs
+```
+
+| | unique_id | ds | y |
+|----|----|----|----|
+| 0 | Australia | 1998-01-01 | 23182.197269 |
+| 1 | Australia | 1998-04-01 | 20323.380067 |
+| 2 | Australia | 1998-07-01 | 19826.640511 |
+| 3 | Australia | 1998-10-01 | 20830.129891 |
+| 4 | Australia | 1999-01-01 | 22087.353380 |
+| ... | ... | ... | ... |
+| 33995 | Australia/Western Australia/Experience Perth/V... | 2016-10-01 | 439.699451 |
+| 33996 | Australia/Western Australia/Experience Perth/V... | 2017-01-01 | 356.867038 |
+| 33997 | Australia/Western Australia/Experience Perth/V... | 2017-04-01 | 302.296119 |
+| 33998 | Australia/Western Australia/Experience Perth/V... | 2017-07-01 | 373.442070 |
+| 33999 | Australia/Western Australia/Experience Perth/V... | 2017-10-01 | 455.316702 |
+
+
+```python
+S_df_cs.iloc[:5, :5]
+```
+
+| | unique_id | Australia/ACT/Canberra/Business | Australia/ACT/Canberra/Holiday | Australia/ACT/Canberra/Other | Australia/ACT/Canberra/Visiting |
+|----|----|----|----|----|----|
+| 0 | Australia | 1.0 | 1.0 | 1.0 | 1.0 |
+| 1 | Australia/ACT | 1.0 | 1.0 | 1.0 | 1.0 |
+| 2 | Australia/New South Wales | 0.0 | 0.0 | 0.0 | 0.0 |
+| 3 | Australia/Northern Territory | 0.0 | 0.0 | 0.0 | 0.0 |
+| 4 | Australia/Queensland | 0.0 | 0.0 | 0.0 | 0.0 |
+
+### 2b. Split Train/Test sets
+
+We use the final two years (8 quarters) as test set. Consequently, our
+forecast horizon=8.
+
+
+```python
+horizon = 8
+```
+
+
+```python
+Y_test_df_cs = Y_df_cs.groupby("unique_id", as_index=False).tail(horizon)
+Y_train_df_cs = Y_df_cs.drop(Y_test_df_cs.index)
+```
+
+### 2c. Computing base forecasts
+
+The following cell computes the **base forecasts** for each time series
+in `Y_df` using the `AutoETS` model. Observe that `Y_hat_df` contains
+the forecasts but they are not coherent.
+
+
+```python
+from statsforecast.models import AutoETS
+from statsforecast.core import StatsForecast
+```
+
+
+```python
+fcst = StatsForecast(models=[AutoETS(season_length=4, model='ZZA')],
+ freq='QS', n_jobs=-1)
+Y_hat_df_cs = fcst.forecast(df=Y_train_df_cs, h=horizon, fitted=True)
+Y_fitted_df_cs = fcst.forecast_fitted_values()
+```
+
+### 2d. Reconcile forecasts
+
+The following cell makes the previous forecasts coherent using the
+[`HierarchicalReconciliation`](https://Nixtla.github.io/hierarchicalforecast/src/core.html#hierarchicalreconciliation)
+class. Since the hierarchy structure is not strict, we can’t use methods
+such as
+[`TopDown`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#topdown)
+or
+[`MiddleOut`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#middleout).
+In this example we use
+[`BottomUp`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#bottomup)
+and
+[`MinTrace`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#mintrace).
+
+
+```python
+from hierarchicalforecast.methods import BottomUp, MinTrace
+from hierarchicalforecast.core import HierarchicalReconciliation
+```
+
+
+```python
+reconcilers = [
+ BottomUp(),
+ MinTrace(method='mint_shrink'),
+ MinTrace(method='ols')
+]
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+Y_rec_df_cs = hrec.reconcile(Y_hat_df=Y_hat_df_cs, Y_df=Y_fitted_df_cs, S_df=S_df_cs, tags=tags_cs)
+```
+
+The dataframe `Y_rec_df` contains the reconciled forecasts.
+
+
+```python
+Y_rec_df_cs.head()
+```
+
+| | unique_id | ds | AutoETS | AutoETS/BottomUp | AutoETS/MinTrace_method-mint_shrink | AutoETS/MinTrace_method-ols |
+|----|----|----|----|----|----|----|
+| 0 | Australia | 2016-01-01 | 25990.068004 | 24381.911737 | 25428.089783 | 25894.399067 |
+| 1 | Australia | 2016-04-01 | 24458.490282 | 22903.895964 | 23914.271400 | 24357.301898 |
+| 2 | Australia | 2016-07-01 | 23974.055984 | 22412.265739 | 23428.462394 | 23865.910647 |
+| 3 | Australia | 2016-10-01 | 24563.454495 | 23127.349578 | 24089.845955 | 24470.782393 |
+| 4 | Australia | 2017-01-01 | 25990.068004 | 24518.118006 | 25545.358678 | 25901.362283 |
+
+## 3. Temporal reconciliation
+
+Next, we aim to reconcile our forecasts also in the temporal domain.
+
+### 3a. Aggregating the dataset according to temporal hierarchy
+
+We first define the temporal aggregation spec. The spec is a dictionary
+in which the keys are the name of the aggregation and the value is the
+amount of bottom-level timesteps that should be aggregated in that
+aggregation. For example, `year` consists of `12` months, so we define a
+key, value pair `"yearly":12`. We can do something similar for other
+aggregations that we are interested in.
+
+In this example, we choose a temporal aggregation of `year`,
+`semiannual` and `quarter`. The bottom level timesteps have a quarterly
+frequency.
+
+
+```python
+spec_temporal = {"year": 4, "semiannual": 2, "quarter": 1}
+```
+
+We next compute the temporally aggregated train- and test sets using the
+[`aggregate_temporal`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#aggregate_temporal)
+function. Note that we have different aggregation matrices `S` for the
+train- and test set, as the test set contains temporal hierarchies that
+are not included in the train set.
+
+
+```python
+from hierarchicalforecast.utils import aggregate_temporal
+```
+
+
+```python
+Y_train_df_te, S_train_df_te, tags_te_train = aggregate_temporal(df=Y_train_df_cs, spec=spec_temporal)
+Y_test_df_te, S_test_df_te, tags_te_test = aggregate_temporal(df=Y_test_df_cs, spec=spec_temporal)
+```
+
+
+```python
+S_train_df_te.iloc[:5, :5]
+```
+
+| | temporal_id | quarter-1 | quarter-2 | quarter-3 | quarter-4 |
+|-----|-------------|-----------|-----------|-----------|-----------|
+| 0 | year-1 | 1.0 | 1.0 | 1.0 | 1.0 |
+| 1 | year-2 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 2 | year-3 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 3 | year-4 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 4 | year-5 | 0.0 | 0.0 | 0.0 | 0.0 |
+
+
+```python
+S_test_df_te.iloc[:5, :5]
+```
+
+| | temporal_id | quarter-1 | quarter-2 | quarter-3 | quarter-4 |
+|-----|--------------|-----------|-----------|-----------|-----------|
+| 0 | year-1 | 1.0 | 1.0 | 1.0 | 1.0 |
+| 1 | year-2 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 2 | semiannual-1 | 1.0 | 1.0 | 0.0 | 0.0 |
+| 3 | semiannual-2 | 0.0 | 0.0 | 1.0 | 1.0 |
+| 4 | semiannual-3 | 0.0 | 0.0 | 0.0 | 0.0 |
+
+If you don’t have a test set available, as is usually the case when
+you’re making forecasts, it is necessary to create a future dataframe
+that holds the correct bottom-level unique_ids and timestamps so that
+they can be temporally aggregated. We can use the
+[`make_future_dataframe`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#make_future_dataframe)
+helper function for that.
+
+
+```python
+from hierarchicalforecast.utils import make_future_dataframe
+```
+
+
+```python
+Y_test_df_te_new = make_future_dataframe(Y_train_df_te, freq="QS", h=horizon)
+```
+
+`Y_test_df_te_new` can be then used in
+[`aggregate_temporal`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#aggregate_temporal)
+to construct the temporally aggregated structures:
+
+
+```python
+Y_test_df_te_new, S_test_df_te_new, tags_te_test_new = aggregate_temporal(df=Y_test_df_te_new, spec=spec_temporal)
+```
+
+And we can verify that we have the same temporally aggregated test set,
+except that `Y_test_df_te_new` doesn’t contain the ground truth values
+`y`.
+
+
+```python
+Y_test_df_te
+```
+
+| | temporal_id | unique_id | ds | y |
+|----|----|----|----|----|
+| 0 | year-1 | Australia | 2016-10-01 | 101484.586551 |
+| 1 | year-2 | Australia | 2017-10-01 | 107709.864650 |
+| 2 | year-1 | Australia/ACT | 2016-10-01 | 2457.401367 |
+| 3 | year-2 | Australia/ACT | 2017-10-01 | 2734.748452 |
+| 4 | year-1 | Australia/ACT/Business | 2016-10-01 | 754.139245 |
+| ... | ... | ... | ... | ... |
+| 5945 | quarter-4 | Australia/Western Australia/Visiting | 2016-10-01 | 787.030391 |
+| 5946 | quarter-5 | Australia/Western Australia/Visiting | 2017-01-01 | 702.777251 |
+| 5947 | quarter-6 | Australia/Western Australia/Visiting | 2017-04-01 | 642.516090 |
+| 5948 | quarter-7 | Australia/Western Australia/Visiting | 2017-07-01 | 646.521395 |
+| 5949 | quarter-8 | Australia/Western Australia/Visiting | 2017-10-01 | 813.184778 |
+
+
+```python
+Y_test_df_te_new
+```
+
+| | temporal_id | unique_id | ds |
+|------|-------------|--------------------------------------|------------|
+| 0 | year-1 | Australia | 2016-10-01 |
+| 1 | year-2 | Australia | 2017-10-01 |
+| 2 | year-1 | Australia/ACT | 2016-10-01 |
+| 3 | year-2 | Australia/ACT | 2017-10-01 |
+| 4 | year-1 | Australia/ACT/Business | 2016-10-01 |
+| ... | ... | ... | ... |
+| 5945 | quarter-4 | Australia/Western Australia/Visiting | 2016-10-01 |
+| 5946 | quarter-5 | Australia/Western Australia/Visiting | 2017-01-01 |
+| 5947 | quarter-6 | Australia/Western Australia/Visiting | 2017-04-01 |
+| 5948 | quarter-7 | Australia/Western Australia/Visiting | 2017-07-01 |
+| 5949 | quarter-8 | Australia/Western Australia/Visiting | 2017-10-01 |
+
+### 3b. Computing base forecasts
+
+Now, we need to compute base forecasts for each temporal aggregation.
+The following cell computes the **base forecasts** for each temporal
+aggregation in `Y_train_df_te` using the `AutoETS` model. Observe that
+`Y_hat_df_te` contains the forecasts but they are not coherent.
+
+Note also that both frequency and horizon are different for each
+temporal aggregation. In this example, the lowest level has a quarterly
+frequency, and a horizon of `8` (constituting `2` years). The `year`
+aggregation thus has a yearly frequency with a horizon of `2`.
+
+It is of course possible to choose a different model for each level in
+the temporal aggregation - you can be as creative as you like!
+
+
+```python
+Y_hat_dfs_te = []
+id_cols = ["unique_id", "temporal_id", "ds", "y"]
+# We will train a model for each temporal level
+for level, temporal_ids_train in tags_te_train.items():
+ # Filter the data for the level
+ Y_level_train = Y_train_df_te.query("temporal_id in @temporal_ids_train")
+ temporal_ids_test = tags_te_test[level]
+ Y_level_test = Y_test_df_te.query("temporal_id in @temporal_ids_test")
+ # For each temporal level we have a different frequency and forecast horizon
+ freq_level = pd.infer_freq(Y_level_train["ds"].unique())
+ horizon_level = Y_level_test["ds"].nunique()
+ # Train a model and create forecasts
+ fcst = StatsForecast(models=[AutoETS(model='ZZZ')], freq=freq_level, n_jobs=-1)
+ Y_hat_df_te_level = fcst.forecast(df=Y_level_train[["ds", "unique_id", "y"]], h=horizon_level)
+ # Add the test set to the forecast
+ Y_hat_df_te_level = Y_hat_df_te_level.merge(Y_level_test, on=["ds", "unique_id"], how="left")
+ # Put cols in the right order (for readability)
+ Y_hat_cols = id_cols + [col for col in Y_hat_df_te_level.columns if col not in id_cols]
+ Y_hat_df_te_level = Y_hat_df_te_level[Y_hat_cols]
+ # Append the forecast to the list
+ Y_hat_dfs_te.append(Y_hat_df_te_level)
+
+Y_hat_df_te = pd.concat(Y_hat_dfs_te, ignore_index=True)
+```
+
+### 3c. Reconcile forecasts
+
+We can again use the
+[`HierarchicalReconciliation`](https://Nixtla.github.io/hierarchicalforecast/src/core.html#hierarchicalreconciliation)
+class to reconcile the forecasts. In this example we use
+[`BottomUp`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#bottomup)
+and
+[`MinTrace`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#mintrace).
+Note that we have to set `temporal=True` in the `reconcile` function.
+
+Note that temporal reconcilation currently isn’t supported for insample
+reconciliation methods, such as `MinTrace(method='mint_shrink')`.
+
+
+```python
+reconcilers = [
+ BottomUp(),
+ MinTrace(method='ols')
+]
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+Y_rec_df_te = hrec.reconcile(Y_hat_df=Y_hat_df_te, S_df=S_test_df_te, tags=tags_te_test, temporal=True)
+```
+
+## 4. Evaluation
+
+The `HierarchicalForecast` package includes the
+[`evaluate`](https://Nixtla.github.io/hierarchicalforecast/src/evaluation.html#evaluate)
+function to evaluate the different hierarchies.
+
+
+```python
+from hierarchicalforecast.evaluation import evaluate
+from utilsforecast.losses import rmse
+```
+
+### 4a. Cross-sectional evaluation
+
+We first evaluate the forecasts *across all cross-sectional
+aggregations*.
+
+
+```python
+eval_tags = {}
+eval_tags['Total'] = tags_cs['Country']
+eval_tags['Purpose'] = tags_cs['Country/Purpose']
+eval_tags['State'] = tags_cs['Country/State']
+eval_tags['Regions'] = tags_cs['Country/State/Region']
+eval_tags['Bottom'] = tags_cs['Country/State/Region/Purpose']
+
+evaluation = evaluate(df = Y_rec_df_te.drop(columns = 'temporal_id'),
+ tags = eval_tags,
+ metrics = [rmse])
+
+evaluation.columns = ['level', 'metric', 'Base', 'BottomUp', 'MinTrace(ols)']
+numeric_cols = evaluation.select_dtypes(include="number").columns
+evaluation[numeric_cols] = evaluation[numeric_cols].map('{:.2f}'.format).astype(np.float64)
+```
+
+
+```python
+evaluation
+```
+
+| | level | metric | Base | BottomUp | MinTrace(ols) |
+|-----|---------|--------|---------|----------|---------------|
+| 0 | Total | rmse | 4249.25 | 4461.95 | 4234.55 |
+| 1 | Purpose | rmse | 1222.57 | 1273.48 | 1137.57 |
+| 2 | State | rmse | 635.78 | 546.02 | 611.32 |
+| 3 | Regions | rmse | 103.67 | 107.00 | 99.23 |
+| 4 | Bottom | rmse | 33.15 | 33.98 | 32.30 |
+| 5 | Overall | rmse | 81.89 | 82.41 | 78.97 |
+
+As can be seen `MinTrace(ols)` seems to be the best forecasting method
+across each cross-sectional aggregation.
+
+### 4b. Temporal evaluation
+
+We then evaluate the temporally aggregated forecasts *across all
+temporal aggregations*.
+
+
+```python
+evaluation = evaluate(df = Y_rec_df_te.drop(columns = 'unique_id'),
+ tags = tags_te_test,
+ metrics = [rmse],
+ id_col="temporal_id")
+
+evaluation.columns = ['level', 'metric', 'Base', 'BottomUp', 'MinTrace(ols)']
+numeric_cols = evaluation.select_dtypes(include="number").columns
+evaluation[numeric_cols] = evaluation[numeric_cols].map('{:.2f}'.format).astype(np.float64)
+```
+
+
+```python
+evaluation
+```
+
+| | level | metric | Base | BottomUp | MinTrace(ols) |
+|-----|------------|--------|--------|----------|---------------|
+| 0 | year | rmse | 480.85 | 581.18 | 515.32 |
+| 1 | semiannual | rmse | 312.33 | 304.98 | 275.30 |
+| 2 | quarter | rmse | 168.02 | 168.02 | 155.61 |
+| 3 | Overall | rmse | 253.94 | 266.17 | 241.19 |
+
+Again, `MinTrace(ols)` is the best overall method, scoring the lowest
+`rmse` on the `quarter` aggregated forecasts, and being slightly worse
+than the `Base` forecasts on the `year` aggregated forecasts.
+
+### 4c. Cross-temporal evaluation
+
+Finally, we evaluate cross-temporally. To do so, we first need to obtain
+the combination of cross-sectional and temporal hierarchies, for which
+we can use the
+[`get_cross_temporal_tags`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#get_cross_temporal_tags)
+helper function.
+
+
+```python
+from hierarchicalforecast.utils import get_cross_temporal_tags
+```
+
+
+```python
+Y_rec_df_te, tags_ct = get_cross_temporal_tags(Y_rec_df_te, tags_cs=tags_cs, tags_te=tags_te_test)
+```
+
+As we can see, we now have a tag `Country//year` that contains
+`Australia//year-1` and `Australia//year-2`, indicating the
+cross-sectional hierarchy `Australia` at the temporal hierarchies `2016`
+and `2017`.
+
+
+```python
+tags_ct["Country//year"]
+```
+
+``` text
+['Australia//year-1', 'Australia//year-2']
+```
+
+We now have our dataset and cross-temporal tags ready for evaluation.
+
+We define a set of eval_tags, and now we split each cross-sectional
+aggregation also by each temporal aggregation. Note that we skip the
+semiannual temporal aggregation in the below overview.
+
+
+```python
+eval_tags = {}
+eval_tags['TotalByYear'] = tags_ct['Country//year']
+eval_tags['RegionsByYear'] = tags_ct['Country/State/Region//year']
+eval_tags['BottomByYear'] = tags_ct['Country/State/Region/Purpose//year']
+eval_tags['TotalByQuarter'] = tags_ct['Country//quarter']
+eval_tags['RegionsByQuarter'] = tags_ct['Country/State/Region//quarter']
+eval_tags['BottomByQuarter'] = tags_ct['Country/State/Region/Purpose//quarter']
+
+
+evaluation = evaluate(df = Y_rec_df_te.drop(columns=['unique_id', 'temporal_id']),
+ tags = eval_tags,
+ id_col = 'cross_temporal_id',
+ metrics = [rmse])
+
+evaluation.columns = ['level', 'metric', 'Base', 'BottomUp', 'MinTrace(ols)']
+numeric_cols = evaluation.select_dtypes(include="number").columns
+evaluation[numeric_cols] = evaluation[numeric_cols].map('{:.2f}'.format).astype(np.float64)
+```
+
+
+```python
+evaluation
+```
+
+| | level | metric | Base | BottomUp | MinTrace(ols) |
+|-----|------------------|--------|---------|----------|---------------|
+| 0 | TotalByYear | rmse | 7148.99 | 8243.06 | 7748.40 |
+| 1 | RegionsByYear | rmse | 151.96 | 175.69 | 158.48 |
+| 2 | BottomByYear | rmse | 46.98 | 50.78 | 46.72 |
+| 3 | TotalByQuarter | rmse | 2060.77 | 2060.77 | 1942.32 |
+| 4 | RegionsByQuarter | rmse | 57.07 | 57.07 | 54.12 |
+| 5 | BottomByQuarter | rmse | 19.42 | 19.42 | 18.69 |
+| 6 | Overall | rmse | 43.14 | 45.27 | 42.49 |
+
+We find that the best method is the cross-temporally reconciled method
+`AutoETS/MinTrace_method-ols`, which achieves overall lowest RMSE.
+
+### References
+
+- [Hyndman, R.J., & Athanasopoulos, G. (2021). “Forecasting:
+ principles and practice, 3rd edition: Chapter 11: Forecasting
+ hierarchical and grouped series.”. OTexts: Melbourne, Australia.
+ OTexts.com/fpp3 Accessed on July
+ 2022.](https://otexts.com/fpp3/hierarchical.html)
+- [Rob Hyndman, Alan Lee, Earo Wang, Shanika Wickramasuriya, and
+ Maintainer Earo Wang (2021). “hts: Hierarchical and Grouped Time
+ Series”. URL https://CRAN.R-project.org/package=hts. R package
+ version
+ 0.3.1.](https://cran.r-project.org/web/packages/hts/index.html)
+- [Mitchell O’Hara-Wild, Rob Hyndman, Earo Wang, Gabriel Caceres,
+ Tim-Gunnar Hensel, and Timothy Hyndman (2021). “fable: Forecasting
+ Models for Tidy Time Series”. URL
+ https://CRAN.R-project.org/package=fable. R package version
+ 6.0.2.](https://CRAN.R-project.org/package=fable)
+- [Athanasopoulos, G, Hyndman, Rob J., Kourentzes, N., Petropoulos,
+ Fotios (2017). Forecasting with temporal hierarchies. European
+ Journal of Operational Research, 262,
+ 60-74](https://www.sciencedirect.com/science/article/pii/S0377221717301911)
+
diff --git a/hierarchicalforecast/examples/australiandomestictourismtemporal.html.mdx b/hierarchicalforecast/examples/australiandomestictourismtemporal.html.mdx
new file mode 100644
index 00000000..61eabecb
--- /dev/null
+++ b/hierarchicalforecast/examples/australiandomestictourismtemporal.html.mdx
@@ -0,0 +1,444 @@
+---
+description: Temporal Hierarchical Forecasting on Australian Tourism Data
+output-file: australiandomestictourismtemporal.html
+title: Temporal Aggregation (Tourism)
+---
+
+
+In many applications, a set of time series is hierarchically organized.
+Examples include the presence of geographic levels, products, or
+categories that define different types of aggregations. In such
+scenarios, forecasters are often required to provide predictions for all
+disaggregate and aggregate series. A natural desire is for those
+predictions to be **“coherent”**, that is, for the bottom series to add
+up precisely to the forecasts of the aggregated series.
+
+In this notebook we present an example on how to use
+`HierarchicalForecast` to produce coherent forecasts between temporal
+levels. We will use the classic Australian Domestic Tourism (`Tourism`)
+dataset, which contains monthly time series of the number of visitors to
+each state of Australia.
+
+We will first load the `Tourism` data and produce base forecasts using
+an `AutoETS` model from `StatsForecast`. Then, we reconciliate the
+forecasts with several reconciliation algorithms from
+`HierarchicalForecast` according to a temporal hierarchy.
+
+You can run these experiments using CPU or GPU with Google Colab.
+
+
+
+
+```python
+!pip install hierarchicalforecast statsforecast
+```
+
+## 1. Load and Process Data
+
+In this example we will use the
+[Tourism](https://otexts.com/fpp3/tourism.html) dataset from the
+[Forecasting: Principles and Practice](https://otexts.com/fpp3/) book.
+
+The dataset only contains the time series at the lowest level, so we
+need to create the time series for all hierarchies.
+
+
+```python
+import numpy as np
+import pandas as pd
+```
+
+
+```python
+Y_df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/tourism.csv')
+Y_df = Y_df.rename({'Trips': 'y', 'Quarter': 'ds'}, axis=1)
+Y_df.insert(0, 'Country', 'Australia')
+Y_df = Y_df[['Country', 'Region', 'State', 'Purpose', 'ds', 'y']]
+Y_df['ds'] = Y_df['ds'].str.replace(r'(\d+) (Q\d)', r'\1-\2', regex=True)
+Y_df['ds'] = pd.PeriodIndex(Y_df["ds"], freq='Q').to_timestamp()
+Y_df.head()
+```
+
+| | Country | Region | State | Purpose | ds | y |
+|-----|-----------|----------|-----------------|----------|------------|------------|
+| 0 | Australia | Adelaide | South Australia | Business | 1998-01-01 | 135.077690 |
+| 1 | Australia | Adelaide | South Australia | Business | 1998-04-01 | 109.987316 |
+| 2 | Australia | Adelaide | South Australia | Business | 1998-07-01 | 166.034687 |
+| 3 | Australia | Adelaide | South Australia | Business | 1998-10-01 | 127.160464 |
+| 4 | Australia | Adelaide | South Australia | Business | 1999-01-01 | 137.448533 |
+
+## 2. Temporal reconciliation
+
+First, we add a `unique_id` to the data.
+
+
+```python
+Y_df["unique_id"] = Y_df["Country"] + "/" + Y_df["State"] + "/" + Y_df["Region"] + "/" + Y_df["Purpose"]
+```
+
+### 2a. Split Train/Test sets
+
+We use the final two years (8 quarters) as test set. Consequently, our
+forecast horizon=8.
+
+
+```python
+horizon = 8
+```
+
+
+```python
+Y_test_df = Y_df.groupby("unique_id", as_index=False).tail(horizon)
+Y_train_df = Y_df.drop(Y_test_df.index)
+```
+
+### 2a. Aggregating the dataset according to temporal hierarchy
+
+We first define the temporal aggregation spec. The spec is a dictionary
+in which the keys are the name of the aggregation and the value is the
+amount of bottom-level timesteps that should be aggregated in that
+aggregation. For example, `year` consists of `12` months, so we define a
+key, value pair `"yearly":12`. We can do something similar for other
+aggregations that we are interested in.
+
+In this example, we choose a temporal aggregation of `year`,
+`semiannual` and `quarter`. The bottom level timesteps have a quarterly
+frequency.
+
+
+```python
+spec_temporal = {"year": 4, "semiannual": 2, "quarter": 1}
+```
+
+We next compute the temporally aggregated train- and test sets using the
+[`aggregate_temporal`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#aggregate_temporal)
+function. Note that we have different aggregation matrices `S` for the
+train- and test set, as the test set contains temporal hierarchies that
+are not included in the train set.
+
+
+```python
+from hierarchicalforecast.utils import aggregate_temporal
+```
+
+
+```python
+Y_train_df, S_train_df, tags_train = aggregate_temporal(df=Y_train_df, spec=spec_temporal)
+Y_test_df, S_test_df, tags_test = aggregate_temporal(df=Y_test_df, spec=spec_temporal)
+```
+
+
+```python
+tags_train
+```
+
+``` text
+{'year': array(['year-1', 'year-2', 'year-3', 'year-4', 'year-5', 'year-6',
+ 'year-7', 'year-8', 'year-9', 'year-10', 'year-11', 'year-12',
+ 'year-13', 'year-14', 'year-15', 'year-16', 'year-17', 'year-18'],
+ dtype=object),
+ 'semiannual': array(['semiannual-1', 'semiannual-2', 'semiannual-3', 'semiannual-4',
+ 'semiannual-5', 'semiannual-6', 'semiannual-7', 'semiannual-8',
+ 'semiannual-9', 'semiannual-10', 'semiannual-11', 'semiannual-12',
+ 'semiannual-13', 'semiannual-14', 'semiannual-15', 'semiannual-16',
+ 'semiannual-17', 'semiannual-18', 'semiannual-19', 'semiannual-20',
+ 'semiannual-21', 'semiannual-22', 'semiannual-23', 'semiannual-24',
+ 'semiannual-25', 'semiannual-26', 'semiannual-27', 'semiannual-28',
+ 'semiannual-29', 'semiannual-30', 'semiannual-31', 'semiannual-32',
+ 'semiannual-33', 'semiannual-34', 'semiannual-35', 'semiannual-36'],
+ dtype=object),
+ 'quarter': array(['quarter-1', 'quarter-2', 'quarter-3', 'quarter-4', 'quarter-5',
+ 'quarter-6', 'quarter-7', 'quarter-8', 'quarter-9', 'quarter-10',
+ 'quarter-11', 'quarter-12', 'quarter-13', 'quarter-14',
+ 'quarter-15', 'quarter-16', 'quarter-17', 'quarter-18',
+ 'quarter-19', 'quarter-20', 'quarter-21', 'quarter-22',
+ 'quarter-23', 'quarter-24', 'quarter-25', 'quarter-26',
+ 'quarter-27', 'quarter-28', 'quarter-29', 'quarter-30',
+ 'quarter-31', 'quarter-32', 'quarter-33', 'quarter-34',
+ 'quarter-35', 'quarter-36', 'quarter-37', 'quarter-38',
+ 'quarter-39', 'quarter-40', 'quarter-41', 'quarter-42',
+ 'quarter-43', 'quarter-44', 'quarter-45', 'quarter-46',
+ 'quarter-47', 'quarter-48', 'quarter-49', 'quarter-50',
+ 'quarter-51', 'quarter-52', 'quarter-53', 'quarter-54',
+ 'quarter-55', 'quarter-56', 'quarter-57', 'quarter-58',
+ 'quarter-59', 'quarter-60', 'quarter-61', 'quarter-62',
+ 'quarter-63', 'quarter-64', 'quarter-65', 'quarter-66',
+ 'quarter-67', 'quarter-68', 'quarter-69', 'quarter-70',
+ 'quarter-71', 'quarter-72'], dtype=object)}
+```
+
+Our aggregation matrices aggregate the lowest temporal granularity
+(quarters) up to years.
+
+
+```python
+S_train_df.iloc[:5, :5]
+```
+
+| | temporal_id | quarter-1 | quarter-2 | quarter-3 | quarter-4 |
+|-----|-------------|-----------|-----------|-----------|-----------|
+| 0 | year-1 | 1.0 | 1.0 | 1.0 | 1.0 |
+| 1 | year-2 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 2 | year-3 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 3 | year-4 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 4 | year-5 | 0.0 | 0.0 | 0.0 | 0.0 |
+
+
+```python
+S_test_df.iloc[:5, :5]
+```
+
+| | temporal_id | quarter-1 | quarter-2 | quarter-3 | quarter-4 |
+|-----|--------------|-----------|-----------|-----------|-----------|
+| 0 | year-1 | 1.0 | 1.0 | 1.0 | 1.0 |
+| 1 | year-2 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 2 | semiannual-1 | 1.0 | 1.0 | 0.0 | 0.0 |
+| 3 | semiannual-2 | 0.0 | 0.0 | 1.0 | 1.0 |
+| 4 | semiannual-3 | 0.0 | 0.0 | 0.0 | 0.0 |
+
+If you don’t have a test set available, as is usually the case when
+you’re making forecasts, it is necessary to create a future dataframe
+that holds the correct bottom-level unique_ids and timestamps so that
+they can be temporally aggregated. We can use the
+[`make_future_dataframe`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#make_future_dataframe)
+helper function for that.
+
+
+```python
+from hierarchicalforecast.utils import make_future_dataframe
+```
+
+
+```python
+Y_test_df_new = make_future_dataframe(Y_train_df, freq="QS", h=horizon)
+```
+
+`Y_test_df_new` can be then used in
+[`aggregate_temporal`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#aggregate_temporal)
+to construct the temporally aggregated structures:
+
+
+```python
+Y_test_df_new, S_test_df_new, tags_test_new = aggregate_temporal(df=Y_test_df_new, spec=spec_temporal)
+```
+
+And we can verify that we have the same temporally aggregated test set,
+except that `Y_test_df_new` doesn’t contain the ground truth values `y`.
+
+
+```python
+S_test_df_new
+```
+
+| | temporal_id | quarter-1 | quarter-2 | quarter-3 | quarter-4 | quarter-5 | quarter-6 | quarter-7 | quarter-8 |
+|----|----|----|----|----|----|----|----|----|----|
+| 0 | year-1 | 1.0 | 1.0 | 1.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 1 | year-2 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 | 1.0 | 1.0 |
+| 2 | semiannual-1 | 1.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 3 | semiannual-2 | 0.0 | 0.0 | 1.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 4 | semiannual-3 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0.0 | 0.0 |
+| 5 | semiannual-4 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 |
+| 6 | quarter-1 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 7 | quarter-2 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 8 | quarter-3 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 9 | quarter-4 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 10 | quarter-5 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 |
+| 11 | quarter-6 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 |
+| 12 | quarter-7 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 13 | quarter-8 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 |
+
+
+```python
+Y_test_df
+```
+
+| | temporal_id | unique_id | ds | y |
+|----|----|----|----|----|
+| 0 | year-1 | Australia/ACT/Canberra/Business | 2016-10-01 | 754.139245 |
+| 1 | year-2 | Australia/ACT/Canberra/Business | 2017-10-01 | 809.950839 |
+| 2 | year-1 | Australia/ACT/Canberra/Holiday | 2016-10-01 | 735.365896 |
+| 3 | year-2 | Australia/ACT/Canberra/Holiday | 2017-10-01 | 834.717900 |
+| 4 | year-1 | Australia/ACT/Canberra/Other | 2016-10-01 | 175.239916 |
+| ... | ... | ... | ... | ... |
+| 4251 | quarter-4 | Australia/Western Australia/Experience Perth/V... | 2016-10-01 | 439.699451 |
+| 4252 | quarter-5 | Australia/Western Australia/Experience Perth/V... | 2017-01-01 | 356.867038 |
+| 4253 | quarter-6 | Australia/Western Australia/Experience Perth/V... | 2017-04-01 | 302.296119 |
+| 4254 | quarter-7 | Australia/Western Australia/Experience Perth/V... | 2017-07-01 | 373.442070 |
+| 4255 | quarter-8 | Australia/Western Australia/Experience Perth/V... | 2017-10-01 | 455.316702 |
+
+
+```python
+Y_test_df_new
+```
+
+| | temporal_id | unique_id | ds |
+|----|----|----|----|
+| 0 | year-1 | Australia/ACT/Canberra/Business | 2016-10-01 |
+| 1 | year-2 | Australia/ACT/Canberra/Business | 2017-10-01 |
+| 2 | year-1 | Australia/ACT/Canberra/Holiday | 2016-10-01 |
+| 3 | year-2 | Australia/ACT/Canberra/Holiday | 2017-10-01 |
+| 4 | year-1 | Australia/ACT/Canberra/Other | 2016-10-01 |
+| ... | ... | ... | ... |
+| 4251 | quarter-4 | Australia/Western Australia/Experience Perth/V... | 2016-10-01 |
+| 4252 | quarter-5 | Australia/Western Australia/Experience Perth/V... | 2017-01-01 |
+| 4253 | quarter-6 | Australia/Western Australia/Experience Perth/V... | 2017-04-01 |
+| 4254 | quarter-7 | Australia/Western Australia/Experience Perth/V... | 2017-07-01 |
+| 4255 | quarter-8 | Australia/Western Australia/Experience Perth/V... | 2017-10-01 |
+
+### 3b. Computing base forecasts
+
+Now, we need to compute base forecasts for each temporal aggregation.
+The following cell computes the **base forecasts** for each temporal
+aggregation in `Y_train_df` using the `AutoETS` model. Observe that
+`Y_hat_df` contains the forecasts but they are not coherent.
+
+Note also that both frequency and horizon are different for each
+temporal aggregation. In this example, the lowest level has a quarterly
+frequency, and a horizon of `8` (constituting `2` years). The `year`
+aggregation thus has a yearly frequency with a horizon of `2`.
+
+It is of course possible to choose a different model for each level in
+the temporal aggregation - you can be as creative as you like!
+
+
+```python
+from statsforecast.models import AutoETS
+from statsforecast.core import StatsForecast
+```
+
+
+```python
+Y_hat_dfs = []
+id_cols = ["unique_id", "temporal_id", "ds", "y"]
+# We will train a model for each temporal level
+for level, temporal_ids_train in tags_train.items():
+ # Filter the data for the level
+ Y_level_train = Y_train_df.query("temporal_id in @temporal_ids_train")
+ temporal_ids_test = tags_test[level]
+ Y_level_test = Y_test_df.query("temporal_id in @temporal_ids_test")
+ # For each temporal level we have a different frequency and forecast horizon
+ freq_level = pd.infer_freq(Y_level_train["ds"].unique())
+ horizon_level = Y_level_test["ds"].nunique()
+ # Train a model and create forecasts
+ fcst = StatsForecast(models=[AutoETS(model='ZZZ')], freq=freq_level, n_jobs=-1)
+ Y_hat_df_level = fcst.forecast(df=Y_level_train[["ds", "unique_id", "y"]], h=horizon_level, level=[80, 90])
+ # Add the test set to the forecast
+ Y_hat_df_level = Y_hat_df_level.merge(Y_level_test, on=["ds", "unique_id"], how="left")
+ # Put cols in the right order (for readability)
+ Y_hat_cols = id_cols + [col for col in Y_hat_df_level.columns if col not in id_cols]
+ Y_hat_df_level = Y_hat_df_level[Y_hat_cols]
+ # Append the forecast to the list
+ Y_hat_dfs.append(Y_hat_df_level)
+
+Y_hat_df = pd.concat(Y_hat_dfs, ignore_index=True)
+```
+
+### 3c. Reconcile forecasts
+
+We can use the
+[`HierarchicalReconciliation`](https://Nixtla.github.io/hierarchicalforecast/src/core.html#hierarchicalreconciliation)
+class to reconcile the forecasts. In this example we use
+[`BottomUp`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#bottomup)
+and
+[`MinTrace`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#mintrace).
+Note that we have to set `temporal=True` in the `reconcile` function.
+
+Note that temporal reconcilation currently isn’t supported for insample
+reconciliation methods, such as `MinTrace(method='mint_shrink')`.
+
+
+```python
+from hierarchicalforecast.methods import BottomUp, MinTrace
+from hierarchicalforecast.core import HierarchicalReconciliation
+```
+
+
+```python
+reconcilers = [
+ BottomUp(),
+ MinTrace(method="ols"),
+]
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+Y_rec_df = hrec.reconcile(Y_hat_df=Y_hat_df,
+ S_df=S_test_df,
+ tags=tags_test,
+ temporal=True,
+ level=[80, 90])
+```
+
+## 4. Evaluation
+
+The `HierarchicalForecast` package includes the
+[`evaluate`](https://Nixtla.github.io/hierarchicalforecast/src/evaluation.html#evaluate)
+function to evaluate the different hierarchies.
+
+We evaluate the temporally aggregated forecasts *across all temporal
+aggregations*.
+
+
+```python
+from hierarchicalforecast.evaluation import evaluate
+from utilsforecast.losses import mae, scaled_crps
+```
+
+
+```python
+evaluation = evaluate(df = Y_rec_df.drop(columns = 'unique_id'),
+ tags = tags_test,
+ metrics = [mae, scaled_crps],
+ level = [80, 90],
+ id_col='temporal_id')
+
+evaluation.columns = ['level', 'metric', 'Base', 'BottomUp', 'MinTrace(ols)']
+numeric_cols = evaluation.select_dtypes(include="number").columns
+evaluation[numeric_cols] = evaluation[numeric_cols].map('{:.3}'.format).astype(np.float64)
+```
+
+
+```python
+evaluation
+```
+
+| | level | metric | Base | BottomUp | MinTrace(ols) |
+|-----|------------|-------------|---------|----------|---------------|
+| 0 | year | mae | 47.0000 | 50.8000 | 46.7000 |
+| 1 | year | scaled_crps | 0.0562 | 0.0620 | 0.0666 |
+| 2 | semiannual | mae | 29.5000 | 30.5000 | 29.1000 |
+| 3 | semiannual | scaled_crps | 0.0643 | 0.0681 | 0.0727 |
+| 4 | quarter | mae | 19.4000 | 19.4000 | 18.7000 |
+| 5 | quarter | scaled_crps | 0.0876 | 0.0876 | 0.0864 |
+| 6 | Overall | mae | 26.2000 | 27.1000 | 25.7000 |
+| 7 | Overall | scaled_crps | 0.0765 | 0.0784 | 0.0797 |
+
+`MinTrace(ols)` is the best overall point method, scoring the lowest
+`mae` on the `year` and `semiannual` aggregated forecasts as well as the
+`quarter` bottom-level aggregated forecasts. However, the `Base` method
+is better overall on the probabilistic measure `crps`, where it scores
+the lowest, indicating that the uncertainty levels predicted with the
+`Base` method are better in this example.
+
+## Appendix: plotting the S matrix
+
+
+```python
+from hierarchicalforecast.utils import HierarchicalPlot
+```
+
+We plot our summing matrix for the test set. It’s fairly
+straightforward: there are two years in the test set, consisting of 4
+quarters each. \* The first row of the `S` matrix shows how the
+aggregation `2016` can be obtained by summing the 4 quarters in 2016. \*
+The second row of the `S` matrix shows how the aggregation `2017` can be
+obtained by summing the 4 quarters in 2017. \* The next 4 rows show how
+the semi-annual aggregations can be obtained. \* The final rows are the
+identity matrix for each quarter, denoting the bottom temporal level
+(each quarter).
+
+
+```python
+hplot = HierarchicalPlot(S=S_test_df, tags=tags_test, S_id_col="temporal_id")
+hplot.plot_summing_matrix()
+```
+
+
+
diff --git a/hierarchicalforecast/examples/australianprisonpopulation.html.mdx b/hierarchicalforecast/examples/australianprisonpopulation.html.mdx
new file mode 100644
index 00000000..69f77855
--- /dev/null
+++ b/hierarchicalforecast/examples/australianprisonpopulation.html.mdx
@@ -0,0 +1,345 @@
+---
+description: Geographical Hierarchical Forecasting on Australian Prison Population Data
+output-file: australianprisonpopulation.html
+title: Geographical Aggregation (Prison Population)
+---
+
+
+In many applications, a set of time series is hierarchically organized.
+Examples include the presence of geographic levels, products, or
+categories that define different types of aggregations. In such
+scenarios, forecasters are often required to provide predictions for all
+disaggregate and aggregate series. A natural desire is for those
+predictions to be **“coherent”**, that is, for the bottom series to add
+up precisely to the forecasts of the aggregated series.
+
+In this notebook we present an example on how to use
+`HierarchicalForecast` to produce coherent forecasts between
+geographical levels. We will use the Australian Prison Population
+dataset.
+
+We will first load the dataset and produce base forecasts using an `ETS`
+model from `StatsForecast`, and then reconciliate the forecasts with
+several reconciliation algorithms from `HierarchicalForecast`. Finally,
+we show the performance is comparable with the results reported by the
+[Forecasting: Principles and
+Practice](https://otexts.com/fpp3/tourism.html) which uses the R package
+[fable](https://github.com/tidyverts/fable).
+
+You can run these experiments using CPU or GPU with Google Colab.
+
+
+
+
+```python
+!pip install hierarchicalforecast statsforecast
+```
+
+## 1. Load and Process Data
+
+The dataset only contains the time series at the lowest level, so we
+need to create the time series for all hierarchies.
+
+
+```python
+import numpy as np
+import pandas as pd
+```
+
+
+```python
+Y_df = pd.read_csv('https://OTexts.com/fpp3/extrafiles/prison_population.csv')
+Y_df = Y_df.rename({'Count': 'y', 'Date': 'ds'}, axis=1)
+Y_df.insert(0, 'Country', 'Australia')
+Y_df = Y_df[['Country', 'State', 'Gender', 'Legal', 'Indigenous', 'ds', 'y']]
+Y_df['ds'] = pd.to_datetime(Y_df['ds']) + pd.DateOffset(months=1)
+Y_df.head()
+```
+
+| | Country | State | Gender | Legal | Indigenous | ds | y |
+|-----|-----------|-------|--------|-----------|------------|------------|-----|
+| 0 | Australia | ACT | Female | Remanded | ATSI | 2005-04-01 | 0 |
+| 1 | Australia | ACT | Female | Remanded | Non-ATSI | 2005-04-01 | 2 |
+| 2 | Australia | ACT | Female | Sentenced | ATSI | 2005-04-01 | 0 |
+| 3 | Australia | ACT | Female | Sentenced | Non-ATSI | 2005-04-01 | 5 |
+| 4 | Australia | ACT | Male | Remanded | ATSI | 2005-04-01 | 7 |
+
+The dataset can be grouped in the following grouped structure.
+
+
+```python
+hiers = [
+ ['Country'],
+ ['Country', 'State'],
+ ['Country', 'Gender'],
+ ['Country', 'Legal'],
+ ['Country', 'State', 'Gender', 'Legal']
+]
+```
+
+Using the
+[`aggregate`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#aggregate)
+function from `HierarchicalForecast` we can get the full set of time
+series.
+
+
+```python
+from hierarchicalforecast.utils import aggregate
+```
+
+
+```python
+Y_df, S_df, tags = aggregate(Y_df, hiers)
+Y_df['y'] = Y_df['y']/1e3
+```
+
+
+```python
+Y_df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|--------|
+| 0 | Australia | 2005-04-01 | 24.296 |
+| 1 | Australia | 2005-07-01 | 24.643 |
+| 2 | Australia | 2005-10-01 | 24.511 |
+| 3 | Australia | 2006-01-01 | 24.393 |
+| 4 | Australia | 2006-04-01 | 24.524 |
+
+
+```python
+S_df.iloc[:5, :5]
+```
+
+| | unique_id | Australia/ACT/Female/Remanded | Australia/ACT/Female/Sentenced | Australia/ACT/Male/Remanded | Australia/ACT/Male/Sentenced |
+|----|----|----|----|----|----|
+| 0 | Australia | 1.0 | 1.0 | 1.0 | 1.0 |
+| 1 | Australia/ACT | 1.0 | 1.0 | 1.0 | 1.0 |
+| 2 | Australia/NSW | 0.0 | 0.0 | 0.0 | 0.0 |
+| 3 | Australia/NT | 0.0 | 0.0 | 0.0 | 0.0 |
+| 4 | Australia/QLD | 0.0 | 0.0 | 0.0 | 0.0 |
+
+
+```python
+tags
+```
+
+``` text
+{'Country': array(['Australia'], dtype=object),
+ 'Country/State': array(['Australia/ACT', 'Australia/NSW', 'Australia/NT', 'Australia/QLD',
+ 'Australia/SA', 'Australia/TAS', 'Australia/VIC', 'Australia/WA'],
+ dtype=object),
+ 'Country/Gender': array(['Australia/Female', 'Australia/Male'], dtype=object),
+ 'Country/Legal': array(['Australia/Remanded', 'Australia/Sentenced'], dtype=object),
+ 'Country/State/Gender/Legal': array(['Australia/ACT/Female/Remanded', 'Australia/ACT/Female/Sentenced',
+ 'Australia/ACT/Male/Remanded', 'Australia/ACT/Male/Sentenced',
+ 'Australia/NSW/Female/Remanded', 'Australia/NSW/Female/Sentenced',
+ 'Australia/NSW/Male/Remanded', 'Australia/NSW/Male/Sentenced',
+ 'Australia/NT/Female/Remanded', 'Australia/NT/Female/Sentenced',
+ 'Australia/NT/Male/Remanded', 'Australia/NT/Male/Sentenced',
+ 'Australia/QLD/Female/Remanded', 'Australia/QLD/Female/Sentenced',
+ 'Australia/QLD/Male/Remanded', 'Australia/QLD/Male/Sentenced',
+ 'Australia/SA/Female/Remanded', 'Australia/SA/Female/Sentenced',
+ 'Australia/SA/Male/Remanded', 'Australia/SA/Male/Sentenced',
+ 'Australia/TAS/Female/Remanded', 'Australia/TAS/Female/Sentenced',
+ 'Australia/TAS/Male/Remanded', 'Australia/TAS/Male/Sentenced',
+ 'Australia/VIC/Female/Remanded', 'Australia/VIC/Female/Sentenced',
+ 'Australia/VIC/Male/Remanded', 'Australia/VIC/Male/Sentenced',
+ 'Australia/WA/Female/Remanded', 'Australia/WA/Female/Sentenced',
+ 'Australia/WA/Male/Remanded', 'Australia/WA/Male/Sentenced'],
+ dtype=object)}
+```
+
+### Split Train/Test sets
+
+We use the final two years (8 quarters) as test set.
+
+
+```python
+Y_test_df = Y_df.groupby('unique_id', as_index=False).tail(8)
+Y_train_df = Y_df.drop(Y_test_df.index)
+```
+
+## 2. Computing base forecasts
+
+The following cell computes the **base forecasts** for each time series
+in `Y_df` using the `ETS` model. Observe that `Y_hat_df` contains the
+forecasts but they are not coherent.
+
+
+```python
+from statsforecast.models import AutoETS
+from statsforecast.core import StatsForecast
+```
+
+
+```python
+fcst = StatsForecast(models=[AutoETS(season_length=4, model='ZMZ')],
+ freq='QS', n_jobs=-1)
+Y_hat_df = fcst.forecast(df=Y_train_df, h=8, fitted=True)
+Y_fitted_df = fcst.forecast_fitted_values()
+```
+
+
+```python
+Y_test_df
+```
+
+| | unique_id | ds | y |
+|------|-----------------------------|------------|--------|
+| 40 | Australia | 2015-04-01 | 35.271 |
+| 41 | Australia | 2015-07-01 | 35.921 |
+| 42 | Australia | 2015-10-01 | 36.067 |
+| 43 | Australia | 2016-01-01 | 36.983 |
+| 44 | Australia | 2016-04-01 | 37.830 |
+| ... | ... | ... | ... |
+| 2155 | Australia/WA/Male/Sentenced | 2016-01-01 | 3.894 |
+| 2156 | Australia/WA/Male/Sentenced | 2016-04-01 | 3.876 |
+| 2157 | Australia/WA/Male/Sentenced | 2016-07-01 | 3.969 |
+| 2158 | Australia/WA/Male/Sentenced | 2016-10-01 | 4.076 |
+| 2159 | Australia/WA/Male/Sentenced | 2017-01-01 | 4.088 |
+
+
+```python
+Y_train_df
+```
+
+| | unique_id | ds | y |
+|------|-----------------------------|------------|--------|
+| 0 | Australia | 2005-04-01 | 24.296 |
+| 1 | Australia | 2005-07-01 | 24.643 |
+| 2 | Australia | 2005-10-01 | 24.511 |
+| 3 | Australia | 2006-01-01 | 24.393 |
+| 4 | Australia | 2006-04-01 | 24.524 |
+| ... | ... | ... | ... |
+| 2147 | Australia/WA/Male/Sentenced | 2014-01-01 | 3.614 |
+| 2148 | Australia/WA/Male/Sentenced | 2014-04-01 | 3.635 |
+| 2149 | Australia/WA/Male/Sentenced | 2014-07-01 | 3.692 |
+| 2150 | Australia/WA/Male/Sentenced | 2014-10-01 | 3.726 |
+| 2151 | Australia/WA/Male/Sentenced | 2015-01-01 | 3.780 |
+
+## 3. Reconcile forecasts
+
+The following cell makes the previous forecasts coherent using the
+[`HierarchicalReconciliation`](https://Nixtla.github.io/hierarchicalforecast/src/core.html#hierarchicalreconciliation)
+class. Since the hierarchy structure is not strict, we can’t use methods
+such as
+[`TopDown`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#topdown)
+or
+[`MiddleOut`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#middleout).
+In this example we use
+[`BottomUp`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#bottomup)
+and
+[`MinTrace`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#mintrace).
+
+
+```python
+from hierarchicalforecast.methods import BottomUp, MinTrace
+from hierarchicalforecast.core import HierarchicalReconciliation
+```
+
+
+```python
+reconcilers = [
+ BottomUp(),
+ MinTrace(method='mint_shrink')
+]
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+Y_rec_df = hrec.reconcile(Y_hat_df=Y_hat_df, Y_df=Y_fitted_df, S_df=S_df, tags=tags)
+```
+
+The dataframe `Y_rec_df` contains the reconciled forecasts.
+
+
+```python
+Y_rec_df.head()
+```
+
+| | unique_id | ds | AutoETS | AutoETS/BottomUp | AutoETS/MinTrace_method-mint_shrink |
+|----|----|----|----|----|----|
+| 0 | Australia | 2015-04-01 | 34.799497 | 34.946476 | 34.923548 |
+| 1 | Australia | 2015-07-01 | 35.192638 | 35.410342 | 35.432421 |
+| 2 | Australia | 2015-10-01 | 35.188216 | 35.580849 | 35.473386 |
+| 3 | Australia | 2016-01-01 | 35.888628 | 35.951878 | 35.939526 |
+| 4 | Australia | 2016-04-01 | 36.045437 | 36.416829 | 36.245158 |
+
+## 4. Evaluation
+
+The `HierarchicalForecast` package includes the
+[`HierarchicalEvaluation`](https://Nixtla.github.io/hierarchicalforecast/src/evaluation.html#hierarchicalevaluation)
+class to evaluate the different hierarchies and also is capable of
+compute scaled metrics compared to a benchmark model.
+
+
+```python
+from hierarchicalforecast.evaluation import evaluate
+from utilsforecast.losses import mase
+from functools import partial
+```
+
+
+```python
+eval_tags = {}
+eval_tags['Total'] = tags['Country']
+eval_tags['State'] = tags['Country/State']
+eval_tags['Legal status'] = tags['Country/Legal']
+eval_tags['Gender'] = tags['Country/Gender']
+eval_tags['Bottom'] = tags['Country/State/Gender/Legal']
+
+df = Y_rec_df.merge(Y_test_df, on=['unique_id', 'ds'])
+evaluation = evaluate(df = df,
+ tags = eval_tags,
+ train_df = Y_train_df,
+ metrics = [partial(mase, seasonality=4)])
+
+numeric_cols = evaluation.select_dtypes(include="number").columns
+evaluation[numeric_cols] = evaluation[numeric_cols].map('{:.2f}'.format).astype(np.float64)
+evaluation.rename(columns={'AutoETS': 'Base'}, inplace=True)
+```
+
+
+```python
+evaluation
+```
+
+| | level | metric | Base | AutoETS/BottomUp | AutoETS/MinTrace_method-mint_shrink |
+|----|----|----|----|----|----|
+| 0 | Total | mase | 1.36 | 1.07 | 1.17 |
+| 1 | State | mase | 1.53 | 1.55 | 1.59 |
+| 2 | Legal status | mase | 2.40 | 2.48 | 2.38 |
+| 3 | Gender | mase | 1.08 | 0.82 | 0.93 |
+| 4 | Bottom | mase | 2.16 | 2.16 | 2.14 |
+| 5 | Overall | mase | 1.99 | 1.98 | 1.98 |
+
+### Fable Comparison
+
+Observe that we can recover the results reported by the [Forecasting:
+Principles and Practice](https://otexts.com/fpp3/prison.html) book. The
+original results were calculated using the R package
+[fable](https://github.com/tidyverts/fable).
+
+
+
+Fable’s reconciliation
+results
+
+
+### References
+
+- [Hyndman, R.J., & Athanasopoulos, G. (2021). “Forecasting:
+ principles and practice, 3rd edition: Chapter 11: Forecasting
+ hierarchical and grouped series.”. OTexts: Melbourne, Australia.
+ OTexts.com/fpp3 Accessed on July
+ 2022.](https://otexts.com/fpp3/hierarchical.html)
+- [Rob Hyndman, Alan Lee, Earo Wang, Shanika Wickramasuriya, and
+ Maintainer Earo Wang (2021). “hts: Hierarchical and Grouped Time
+ Series”. URL https://CRAN.R-project.org/package=hts. R package
+ version
+ 0.3.1.](https://cran.r-project.org/web/packages/hts/index.html)
+- [Mitchell O’Hara-Wild, Rob Hyndman, Earo Wang, Gabriel Caceres,
+ Tim-Gunnar Hensel, and Timothy Hyndman (2021). “fable: Forecasting
+ Models for Tidy Time Series”. URL
+ https://CRAN.R-project.org/package=fable. R package version
+ 6.0.2.](https://CRAN.R-project.org/package=fable)
+
diff --git a/hierarchicalforecast/examples/imgs/AustralianDomesticTourism-results-fable.png b/hierarchicalforecast/examples/imgs/AustralianDomesticTourism-results-fable.png
new file mode 100644
index 00000000..15f437d0
Binary files /dev/null and b/hierarchicalforecast/examples/imgs/AustralianDomesticTourism-results-fable.png differ
diff --git a/hierarchicalforecast/examples/imgs/AustralianPrisonPopulation-results-fable.png b/hierarchicalforecast/examples/imgs/AustralianPrisonPopulation-results-fable.png
new file mode 100644
index 00000000..fc4f988c
Binary files /dev/null and b/hierarchicalforecast/examples/imgs/AustralianPrisonPopulation-results-fable.png differ
diff --git a/hierarchicalforecast/examples/index.mdx b/hierarchicalforecast/examples/index.mdx
new file mode 100644
index 00000000..9d4129a4
--- /dev/null
+++ b/hierarchicalforecast/examples/index.mdx
@@ -0,0 +1,15 @@
+---
+order: 1
+title: Tutorials
+listing:
+ fields:
+ - title
+ type: table
+ sort-ui: false
+ filter-ui: false
+---
+
+
+Click through to any of these tutorials to get started with
+`HierarchicalForecast`’s features.
+
diff --git a/hierarchicalforecast/examples/installation.html.mdx b/hierarchicalforecast/examples/installation.html.mdx
new file mode 100644
index 00000000..ea1b678f
--- /dev/null
+++ b/hierarchicalforecast/examples/installation.html.mdx
@@ -0,0 +1,83 @@
+---
+description: Install HierachicalForecast with pip or conda
+output-file: installation.html
+title: Install
+---
+
+
+We recommend using `uv` as Python package manager, for which you can
+find installation instructions
+[here](https://docs.astral.sh/uv/getting-started/installation/).
+
+You can then install the *released version* of `HierachicalForecast`:
+
+
+```python
+uv pip install hierarchicalforecast
+```
+
+Alternatively, you can directly install from the [Python package
+index](https://pypi.org) with:
+
+
+```python
+pip install hierarchicalforecast
+```
+
+or within a `conda` environment:
+
+
+```python
+conda install -c conda-forge hierarchicalforecast
+```
+
+> **Tip**
+>
+> We recommend installing your libraries inside a python virtual or
+> [conda
+> environment](https://docs.conda.io/projects/conda/en/latest/user-guide/install/macos.html).
+
+#### Installing from source
+
+We recommend using `uv` as Python package manager, for which you can
+find installation instructions
+[here](https://docs.astral.sh/uv/getting-started/installation/).
+
+1. Clone the HierachicalForecast repo:
+
+
+```bash
+$ git clone https://github.com/Nixtla/hierachicalforecast.git && cd hierachicalforecast
+```
+
+1. Create the environment:
+
+
+```bash
+$ uv venv --python 3.10
+```
+
+1. Activate the environment:
+
+- on MacOS / Linux:
+
+
+```bash
+$ source .venv/bin/activate
+```
+
+- on Windows:
+
+
+```bash
+$ .\.venv\Scripts\activate
+```
+
+1. Install the dependencies and the library
+
+
+```bash
+uv pip install -r setup.py
+uv pip install .
+```
+
diff --git a/hierarchicalforecast/examples/introduction.html.mdx b/hierarchicalforecast/examples/introduction.html.mdx
new file mode 100644
index 00000000..7b6fa1da
--- /dev/null
+++ b/hierarchicalforecast/examples/introduction.html.mdx
@@ -0,0 +1,327 @@
+---
+description: Introduction to Hierarchial Forecasting using `HierarchialForecast`
+output-file: introduction.html
+title: Introduction
+---
+
+
+You can run these experiments using CPU or GPU with Google Colab.
+
+
+
+## 1. Hierarchical Series
+
+In many applications, a set of time series is hierarchically organized.
+Examples include the presence of geographic levels, products, or
+categories that define different types of aggregations.
+
+In such scenarios, forecasters are often required to provide predictions
+for all disaggregate and aggregate series. A natural desire is for those
+predictions to be **“coherent”**, that is, for the bottom series to add
+up precisely to the forecasts of the aggregated series.
+
+
+
+The above figure shows a simple hierarchical structure where we have
+four bottom-level series, two middle-level series, and the top level
+representing the total aggregation. Its hierarchical aggregations or
+coherency constraints are:
+
+$$
+y_{\mathrm{Total},\tau} = y_{\beta_{1},\tau}+y_{\beta_{2},\tau}+y_{\beta_{3},\tau}+y_{\beta_{4},\tau}
+ \qquad \qquad \qquad \qquad \qquad \\
+ \mathbf{y}_{[a],\tau}=\left[y_{\mathrm{Total},\tau},\; y_{\beta_{1},\tau}+y_{\beta_{2},\tau},\;y_{\beta_{3},\tau}+y_{\beta_{4},\tau}\right]^{\intercal}
+ \qquad
+ \mathbf{y}_{[b],\tau}=\left[ y_{\beta_{1},\tau},\; y_{\beta_{2},\tau},\; y_{\beta_{3},\tau},\; y_{\beta_{4},\tau} \right]^{\intercal}
+$$
+
+Luckily these constraints can be compactly expressed with the following
+matrices:
+
+$$
+
+\mathbf{S}_{[a,b][b]}
+=
+\begin{bmatrix}
+\mathbf{A}_{\mathrm{[a][b]}} \\
+ \\
+ \\
+\mathbf{I}_{\mathrm{[b][b]}} \\
+ \\
+\end{bmatrix}
+=
+\begin{bmatrix}
+1 & 1 & 1 & 1 \\
+1 & 1 & 0 & 0 \\
+0 & 0 & 1 & 1 \\
+1 & 0 & 0 & 0 \\
+0 & 1 & 0 & 0 \\
+0 & 0 & 1 & 0 \\
+0 & 0 & 0 & 1 \\
+\end{bmatrix}
+
+$$
+
+where $\mathbf{A}_{[a,b][b]}$ aggregates the bottom series to the upper
+levels, and $\mathbf{I}_{\mathrm{[b][b]}}$ is an identity matrix. The
+representation of the hierarchical series is then:
+
+$$
+
+\mathbf{y}_{[a,b],\tau} = \mathbf{S}_{[a,b][b]} \mathbf{y}_{[b],\tau}
+
+$$
+
+To visualize an example, in Figure 2, one can think of the hierarchical
+time series structure levels to represent different geographical
+aggregations. For example, in Figure 2, the top level is the total
+aggregation of series within a country, the middle level being its
+states and the bottom level its regions.
+
+
+
+## 2. Hierarchical Forecast
+
+To achieve **“coherency”**, most statistical solutions to the
+hierarchical forecasting challenge implement a two-stage reconciliation
+process.
+1. First, we obtain a set of the base forecast
+$\mathbf{\hat{y}}_{[a,b],\tau}$
+
+1. Later, we reconcile them into coherent forecasts
+ $\mathbf{\tilde{y}}_{[a,b],\tau}$.
+
+Most hierarchical reconciliation methods can be expressed by the
+following transformations:
+
+$$\tilde{\mathbf{y}}_{[a,b],\tau} = \mathbf{S}_{[a,b][b]} \mathbf{P}_{[b][a,b]} \hat{\mathbf{y}}_{[a,b],\tau}$$
+
+The HierarchicalForecast library offers a Python collection of
+reconciliation methods, datasets, evaluation and visualization tools for
+the task. Among its available reconciliation methods we have
+[`BottomUp`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#bottomup),
+[`TopDown`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#topdown),
+[`MiddleOut`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#middleout),
+[`MinTrace`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#mintrace),
+[`ERM`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#erm).
+Among its probabilistic coherent methods we have
+[`Normality`](https://Nixtla.github.io/hierarchicalforecast/src/probabilistic_methods.html#normality),
+[`Bootstrap`](https://Nixtla.github.io/hierarchicalforecast/src/probabilistic_methods.html#bootstrap),
+[`PERMBU`](https://Nixtla.github.io/hierarchicalforecast/src/probabilistic_methods.html#permbu).
+
+## 3. Minimal Example
+
+
+```python
+!pip install hierarchicalforecast statsforecast datasetsforecast
+```
+
+### Wrangling Data
+
+
+```python
+import numpy as np
+import pandas as pd
+```
+
+We are going to creat a synthetic data set to illustrate a hierarchical
+time series structure like the one in Figure 1.
+
+We will create a two level structure with four bottom series where
+aggregations of the series are self evident.
+
+
+```python
+# Create Figure 1. synthetic bottom data
+ds = pd.date_range(start='2000-01-01', end='2000-08-01', freq='MS')
+y_base = np.arange(1,9)
+r1 = y_base * (10**1)
+r2 = y_base * (10**1)
+r3 = y_base * (10**2)
+r4 = y_base * (10**2)
+
+ys = np.concatenate([r1, r2, r3, r4])
+ds = np.tile(ds, 4)
+unique_ids = ['r1'] * 8 + ['r2'] * 8 + ['r3'] * 8 + ['r4'] * 8
+top_level = 'Australia'
+middle_level = ['State1'] * 16 + ['State2'] * 16
+bottom_level = unique_ids
+
+bottom_df = dict(ds=ds,
+ top_level=top_level,
+ middle_level=middle_level,
+ bottom_level=bottom_level,
+ y=ys)
+bottom_df = pd.DataFrame(bottom_df)
+bottom_df.groupby('bottom_level').head(2)
+```
+
+| | ds | top_level | middle_level | bottom_level | y |
+|-----|------------|-----------|--------------|--------------|-----|
+| 0 | 2000-01-01 | Australia | State1 | r1 | 10 |
+| 1 | 2000-02-01 | Australia | State1 | r1 | 20 |
+| 8 | 2000-01-01 | Australia | State1 | r2 | 10 |
+| 9 | 2000-02-01 | Australia | State1 | r2 | 20 |
+| 16 | 2000-01-01 | Australia | State2 | r3 | 100 |
+| 17 | 2000-02-01 | Australia | State2 | r3 | 200 |
+| 24 | 2000-01-01 | Australia | State2 | r4 | 100 |
+| 25 | 2000-02-01 | Australia | State2 | r4 | 200 |
+
+The previously introduced hierarchical series $\mathbf{y}_{[a,b]\tau}$
+is captured within the `Y_hier_df` dataframe.
+
+The aggregation constraints matrix $\mathbf{S}_{[a][b]}$ is captured
+within the `S_df` dataframe.
+
+Finally the `tags` contains a list within `Y_hier_df` composing each
+hierarchical level, for example the `tags['top_level']` contains
+`Australia`’s aggregated series index.
+
+
+```python
+from hierarchicalforecast.utils import aggregate
+```
+
+
+```python
+# Create hierarchical structure and constraints
+hierarchy_levels = [['top_level'],
+ ['top_level', 'middle_level'],
+ ['top_level', 'middle_level', 'bottom_level']]
+Y_hier_df, S_df, tags = aggregate(df=bottom_df, spec=hierarchy_levels)
+print('S_df.shape', S_df.shape)
+print('Y_hier_df.shape', Y_hier_df.shape)
+print("tags['top_level']", tags['top_level'])
+```
+
+``` text
+S_df.shape (7, 5)
+Y_hier_df.shape (56, 3)
+tags['top_level'] ['Australia']
+```
+
+
+```python
+Y_hier_df.groupby('unique_id').head(2)
+```
+
+| | unique_id | ds | y |
+|-----|---------------------|------------|-----|
+| 0 | Australia | 2000-01-01 | 220 |
+| 1 | Australia | 2000-02-01 | 440 |
+| 8 | Australia/State1 | 2000-01-01 | 20 |
+| 9 | Australia/State1 | 2000-02-01 | 40 |
+| 16 | Australia/State2 | 2000-01-01 | 200 |
+| 17 | Australia/State2 | 2000-02-01 | 400 |
+| 24 | Australia/State1/r1 | 2000-01-01 | 10 |
+| 25 | Australia/State1/r1 | 2000-02-01 | 20 |
+| 32 | Australia/State1/r2 | 2000-01-01 | 10 |
+| 33 | Australia/State1/r2 | 2000-02-01 | 20 |
+| 40 | Australia/State2/r3 | 2000-01-01 | 100 |
+| 41 | Australia/State2/r3 | 2000-02-01 | 200 |
+| 48 | Australia/State2/r4 | 2000-01-01 | 100 |
+| 49 | Australia/State2/r4 | 2000-02-01 | 200 |
+
+
+```python
+S_df
+```
+
+| | unique_id | Australia/State1/r1 | Australia/State1/r2 | Australia/State2/r3 | Australia/State2/r4 |
+|----|----|----|----|----|----|
+| 0 | Australia | 1.0 | 1.0 | 1.0 | 1.0 |
+| 1 | Australia/State1 | 1.0 | 1.0 | 0.0 | 0.0 |
+| 2 | Australia/State2 | 0.0 | 0.0 | 1.0 | 1.0 |
+| 3 | Australia/State1/r1 | 1.0 | 0.0 | 0.0 | 0.0 |
+| 4 | Australia/State1/r2 | 0.0 | 1.0 | 0.0 | 0.0 |
+| 5 | Australia/State2/r3 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 6 | Australia/State2/r4 | 0.0 | 0.0 | 0.0 | 1.0 |
+
+### Base Predictions
+
+Next, we compute the *base forecast* for each time series using the
+`naive` model. Observe that `Y_hat_df` contains the forecasts but they
+are not coherent.
+
+
+```python
+from statsforecast.models import Naive
+from statsforecast.core import StatsForecast
+```
+
+
+```python
+# Split train/test sets
+Y_test_df = Y_hier_df.groupby('unique_id', as_index=False).tail(4)
+Y_train_df = Y_hier_df.drop(Y_test_df.index)
+
+# Compute base Naive predictions
+# Careful identifying correct data freq, this data monthly 'M'
+fcst = StatsForecast(models=[Naive()],
+ freq='MS', n_jobs=-1)
+Y_hat_df = fcst.forecast(df=Y_train_df, h=4, fitted=True)
+Y_fitted_df = fcst.forecast_fitted_values()
+```
+
+### Reconciliation
+
+
+```python
+from hierarchicalforecast.methods import BottomUp
+from hierarchicalforecast.core import HierarchicalReconciliation
+```
+
+
+```python
+# You can select a reconciler from our collection
+reconcilers = [BottomUp()] # MinTrace(method='mint_shrink')
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+
+Y_rec_df = hrec.reconcile(Y_hat_df=Y_hat_df,
+ Y_df=Y_fitted_df,
+ S_df=S_df, tags=tags)
+Y_rec_df.groupby('unique_id').head(2)
+```
+
+| | unique_id | ds | Naive | Naive/BottomUp |
+|-----|---------------------|------------|-------|----------------|
+| 0 | Australia | 2000-05-01 | 880.0 | 880.0 |
+| 1 | Australia | 2000-06-01 | 880.0 | 880.0 |
+| 4 | Australia/State1 | 2000-05-01 | 80.0 | 80.0 |
+| 5 | Australia/State1 | 2000-06-01 | 80.0 | 80.0 |
+| 8 | Australia/State2 | 2000-05-01 | 800.0 | 800.0 |
+| 9 | Australia/State2 | 2000-06-01 | 800.0 | 800.0 |
+| 12 | Australia/State1/r1 | 2000-05-01 | 40.0 | 40.0 |
+| 13 | Australia/State1/r1 | 2000-06-01 | 40.0 | 40.0 |
+| 16 | Australia/State1/r2 | 2000-05-01 | 40.0 | 40.0 |
+| 17 | Australia/State1/r2 | 2000-06-01 | 40.0 | 40.0 |
+| 20 | Australia/State2/r3 | 2000-05-01 | 400.0 | 400.0 |
+| 21 | Australia/State2/r3 | 2000-06-01 | 400.0 | 400.0 |
+| 24 | Australia/State2/r4 | 2000-05-01 | 400.0 | 400.0 |
+| 25 | Australia/State2/r4 | 2000-06-01 | 400.0 | 400.0 |
+
+## References
+
+- [Hyndman, R.J., & Athanasopoulos, G. (2021). “Forecasting:
+ principles and practice, 3rd edition: Chapter 11: Forecasting
+ hierarchical and grouped series.”. OTexts: Melbourne, Australia.
+ OTexts.com/fpp3 Accessed on July
+ 2022.](https://otexts.com/fpp3/hierarchical.html)
+- [Orcutt, G.H., Watts, H.W., & Edwards, J.B.(1968). Data aggregation
+ and information loss. The American Economic Review, 58 ,
+ 773(787).](http://www.jstor.org/stable/1815532)
+- [Disaggregation methods to expedite product line forecasting.
+ Journal of Forecasting, 9 , 233–254.
+ doi:10.1002/for.3980090304.](https://onlinelibrary.wiley.com/doi/abs/10.1002/for.3980090304)
+- [Wickramasuriya, S. L., Athanasopoulos, G., & Hyndman, R. J. (2019).
+ "Optimal forecast reconciliation for hierarchical and grouped time
+ series through trace minimization". Journal of the American
+ Statistical Association, 114 , 804–819.
+ doi:10.1080/01621459.2018.1448825.](https://robjhyndman.com/publications/mint/)
+- [Ben Taieb, S., & Koo, B. (2019). Regularized regression for
+ hierarchical forecasting without unbiasedness conditions. In
+ Proceedings of the 25th ACM SIGKDD International Conference on
+ Knowledge Discovery & Data Mining KDD ’19 (p. 1337(1347). New York,
+ NY, USA: Association for Computing
+ Machinery.](https://doi.org/10.1145/3292500.3330976)
+
diff --git a/hierarchicalforecast/examples/localglobalaggregation.html.mdx b/hierarchicalforecast/examples/localglobalaggregation.html.mdx
new file mode 100644
index 00000000..1fbd946a
--- /dev/null
+++ b/hierarchicalforecast/examples/localglobalaggregation.html.mdx
@@ -0,0 +1,160 @@
+---
+description: Temporal Hierarchical Aggregation on a local or global level.
+output-file: localglobalaggregation.html
+title: Local vs Global Temporal Aggregation
+---
+
+
+In this notebook we explain the difference between temporally
+aggregating timeseries locally and globally.
+
+You can run these experiments using CPU or GPU with Google Colab.
+
+
+
+
+```python
+!pip install hierarchicalforecast utilsforecast
+```
+
+## 1. Generate Data
+
+In this example we will generate synthetic series to explain the
+difference between local- and global temporal aggregation. We will
+generate 2 series with a daily frequency.
+
+
+```python
+from utilsforecast.data import generate_series
+```
+
+
+```python
+freq = "D"
+n_series = 2
+df = generate_series(n_series=n_series,
+ freq=freq,
+ min_length=2 * 365,
+ max_length=4 * 365,
+ equal_ends=True)
+```
+
+Note that our two timeseries do not have the same number of timesteps:
+
+
+```python
+df.groupby('unique_id', observed=True)["ds"].count()
+```
+
+``` text
+unique_id
+0 1414
+1 1289
+Name: ds, dtype: int64
+```
+
+We then define a spec for our temporal aggregations.
+
+
+```python
+spec = {"year": 365, "quarter": 91, "month": 30, "week": 7, "day": 1}
+```
+
+## 2. Local aggregation (default)
+
+In local aggregation, we treat the timestamps of each timeseries
+individually. It means that the temporal aggregation is performed by
+only looking at the timestamps of each series, disregarding the
+timestamps of other series.
+
+
+```python
+from hierarchicalforecast.utils import aggregate_temporal
+```
+
+
+```python
+Y_df_local, S_df_local, tags_local = aggregate_temporal(df, spec)
+```
+
+We have created temporal aggregations *per timeseries*, as the temporal
+aggregation `month-1` doesn’t correspond to the same (year, month) for
+both timeseries. This is because the series with `unique_id=1` is
+shorter and has its first datapoint in July 2000, in contrast to the
+series with `unique_id=0`, which is longer and has its first timestamp
+in March 2000.
+
+
+```python
+Y_df_local.query("temporal_id == 'month-1'")
+```
+
+| | temporal_id | unique_id | ds | y |
+|-----|-------------|-----------|------------|-----------|
+| 39 | month-1 | 0 | 2000-03-16 | 93.574676 |
+| 87 | month-1 | 1 | 2000-07-19 | 91.506421 |
+
+## 2. Global aggregation
+
+In global aggregation, we examine all unique timestamps across all
+timeseries, and base our temporal aggregations on the unique list of
+timestamps across all timeseries. We can specify the aggregation type by
+setting the `aggregation_type` attritbue in
+[`aggregate_temporal`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#aggregate_temporal).
+
+
+```python
+Y_df_global, S_df_global, tags_globval = aggregate_temporal(df, spec, aggregation_type="global")
+```
+
+We have created temporal aggregations *across all timeseries*, as the
+temporal aggregation `month-1` corresponds to the same (year,
+month)-combination for both timeseries. Since `month-1` isn’t present in
+the second timeseries (as it is shorter), we have only one record for
+the aggregation.
+
+
+```python
+Y_df_global.query("temporal_id == 'month-1'")
+```
+
+| | temporal_id | unique_id | ds | y |
+|-----|-------------|-----------|------------|-----------|
+| 39 | month-1 | 0 | 2000-03-16 | 93.574676 |
+
+For `month-5` however, we have a record for both timeseries, as the
+second series has its first datapoint in that month.
+
+
+```python
+Y_df_global.query("temporal_id == 'month-5'")
+```
+
+| | temporal_id | unique_id | ds | y |
+|-----|-------------|-----------|------------|-----------|
+| 43 | month-5 | 0 | 2000-07-14 | 95.169659 |
+| 87 | month-5 | 1 | 2000-07-14 | 74.502584 |
+
+Hence, the global aggregation ensures temporal alignment across all
+series.
+
+## 3. What to choose?
+
+- If all timeseries have the same length and same timestamps, `global`
+ and `local` yield the same results.
+- The default behavior is `local`. This means that temporal
+ aggregations between timeseries can’t be compared unless the series
+ have the same length and timestamp. This behavior is generally
+ safer, and advised to use when time series are not necessarily
+ related, and you are building per-series models using
+ e.g. `StatsForecast`.
+- The `global` behavior can be useful when dealing with timeseries
+ where we expect relationships between the timeseries. For example,
+ in case of forecasting daily product demand individual products may
+ not always have sales for all timesteps, but one is interested in
+ the overall temporal yearly aggregation across all products. The
+ `global` setting has more room for error, so be careful and check
+ the aggregation result carefully. This would typically be the
+ setting used in combination with models from `MLForecast` or
+ `NeuralForecast`.
+
diff --git a/hierarchicalforecast/examples/m3withthief.html.mdx b/hierarchicalforecast/examples/m3withthief.html.mdx
new file mode 100644
index 00000000..8c7a5621
--- /dev/null
+++ b/hierarchicalforecast/examples/m3withthief.html.mdx
@@ -0,0 +1,331 @@
+---
+description: Temporal Hierarchical Forecasting on M3 monthly and quarterly data with THIEF
+output-file: m3withthief.html
+title: Temporal Aggregation with THIEF
+---
+
+
+In this notebook we present an example on how to use
+`HierarchicalForecast` to produce coherent forecasts between temporal
+levels. We will use the monthly and quarterly timeseries of the `M3`
+dataset. We will first load the `M3` data and produce base forecasts
+using an `AutoETS` model from `StatsForecast`. Then, we reconcile the
+forecasts with `THIEF` (Temporal HIerarchical Forecasting) from
+`HierarchicalForecast` according to a specified temporal hierarchy.
+
+### References
+
+[Athanasopoulos, G, Hyndman, Rob J., Kourentzes, N., Petropoulos, Fotios
+(2017). Forecasting with temporal hierarchies. European Journal of
+Operational Research, 262,
+60-74](https://www.sciencedirect.com/science/article/pii/S0377221717301911)
+
+You can run these experiments using CPU or GPU with Google Colab.
+
+
+
+
+```python
+!pip install hierarchicalforecast statsforecast datasetsforecast
+```
+
+## 1. Load and Process Data
+
+
+```python
+import numpy as np
+import pandas as pd
+```
+
+
+```python
+from datasetsforecast.m3 import M3
+```
+
+
+```python
+m3_monthly, _, _ = M3.load(directory='data', group='Monthly')
+m3_quarterly, _, _ = M3.load(directory='data', group='Quarterly')
+```
+
+We will be making aggregations up to yearly levels, so for both monthly
+and quarterly data we make sure each time series has an integer multiple
+of bottom-level timesteps.
+
+For example, the first time series in m3_monthly (with `unique_id='M1'`)
+has 68 timesteps. This is not a multiple of 12 (12 months in one year),
+so we would not be able to aggregate all timesteps into full years.
+Hence, we truncate (remove) the first 8 timesteps, resulting in 60
+timesteps for this series. We do something similar for the quarterly
+data, albeit with a multiple of 4 (4 quarters in one year).
+
+Depending on the highest temporal aggregation in your reconciliation
+problem, you may want to truncate your data differently.
+
+
+```python
+m3_monthly = m3_monthly.groupby("unique_id", group_keys=False)\
+ .apply(lambda x: x.tail(len(x) // 12 * 12))\
+ .reset_index(drop=True)
+
+m3_quarterly = m3_quarterly.groupby("unique_id", group_keys=False)\
+ .apply(lambda x: x.tail(len(x) // 4 * 4))\
+ .reset_index(drop=True)
+```
+
+## 2. Temporal reconciliation
+
+### 2a. Split Train/Test sets
+
+We use as test samples the last 24 observations from the Monthly series
+and the last 8 observations of each quarterly series, following the
+original THIEF paper.
+
+
+```python
+horizon_monthly = 24
+horizon_quarterly = 8
+```
+
+
+```python
+m3_monthly_test = m3_monthly.groupby("unique_id", as_index=False).tail(horizon_monthly)
+m3_monthly_train = m3_monthly.drop(m3_monthly_test.index)
+
+m3_quarterly_test = m3_quarterly.groupby("unique_id", as_index=False).tail(horizon_quarterly)
+m3_quarterly_train = m3_quarterly.drop(m3_quarterly_test.index)
+```
+
+### 2a. Aggregating the dataset according to temporal hierarchy
+
+We first define the temporal aggregation spec. The spec is a dictionary
+in which the keys are the name of the aggregation and the value is the
+amount of bottom-level timesteps that should be aggregated in that
+aggregation. For example, `year` consists of `12` months, so we define a
+key, value pair `"yearly":12`. We can do something similar for other
+aggregations that we are interested in.
+
+
+```python
+spec_temporal_monthly = {"yearly": 12, "semiannually": 6, "fourmonthly": 4, "quarterly": 3, "bimonthly": 2, "monthly": 1}
+spec_temporal_quarterly = {"yearly": 4, "semiannually": 2, "quarterly": 1}
+```
+
+We next compute the temporally aggregated train- and test sets using the
+[`aggregate_temporal`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#aggregate_temporal)
+function. Note that we have different aggregation matrices `S` for the
+train- and test set, as the test set contains temporal hierarchies that
+are not included in the train set.
+
+
+```python
+from hierarchicalforecast.utils import aggregate_temporal
+```
+
+
+```python
+# Monthly
+Y_monthly_train, S_monthly_train, tags_monthly_train = aggregate_temporal(df=m3_monthly_train, spec=spec_temporal_monthly)
+Y_monthly_test, S_monthly_test, tags_monthly_test = aggregate_temporal(df=m3_monthly_test, spec=spec_temporal_monthly)
+
+# Quarterly
+Y_quarterly_train, S_quarterly_train, tags_quarterly_train = aggregate_temporal(df=m3_quarterly_train, spec=spec_temporal_quarterly)
+Y_quarterly_test, S_quarterly_test, tags_quarterly_test = aggregate_temporal(df=m3_quarterly_test, spec=spec_temporal_quarterly)
+```
+
+Our aggregation matrices aggregate the lowest temporal granularity
+(quarters) up to years, for the train- and test set.
+
+
+```python
+S_monthly_train.iloc[:5, :5]
+```
+
+| | temporal_id | monthly-1 | monthly-2 | monthly-3 | monthly-4 |
+|-----|-------------|-----------|-----------|-----------|-----------|
+| 0 | yearly-1 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 1 | yearly-2 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 2 | yearly-3 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 3 | yearly-4 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 4 | yearly-5 | 0.0 | 0.0 | 0.0 | 0.0 |
+
+
+```python
+S_monthly_test.iloc[:5, :5]
+```
+
+| | temporal_id | monthly-1 | monthly-2 | monthly-3 | monthly-4 |
+|-----|----------------|-----------|-----------|-----------|-----------|
+| 0 | yearly-1 | 1.0 | 1.0 | 1.0 | 1.0 |
+| 1 | yearly-2 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 2 | semiannually-1 | 1.0 | 1.0 | 1.0 | 1.0 |
+| 3 | semiannually-2 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 4 | semiannually-3 | 0.0 | 0.0 | 0.0 | 0.0 |
+
+### 2b. Computing base forecasts
+
+Now, we need to compute base forecasts for each temporal aggregation.
+The following cell computes the **base forecasts** for each temporal
+aggregation in `Y_monthly_train` and `Y_quarterly_train` using the
+`AutoARIMA` model. Observe that `Y_hats` contains the forecasts but they
+are not coherent.
+
+Note also that both frequency and horizon are different for each
+temporal aggregation. For the monthly data, the lowest level has a
+monthly frequency, and a horizon of `24` (constituting 2 years).
+However, as example, the `year` aggregation has a yearly frequency with
+a horizon of 2.
+
+It is of course possible to choose a different model for each level in
+the temporal aggregation - you can be as creative as you like!
+
+
+```python
+from statsforecast.models import AutoARIMA
+from statsforecast.core import StatsForecast
+```
+
+
+```python
+Y_hats = []
+id_cols = ["unique_id", "temporal_id", "ds", "y"]
+
+# We loop over the monthly and quarterly data
+for tags_train, tags_test, Y_train, Y_test in zip([tags_monthly_train, tags_quarterly_train],
+ [tags_monthly_test, tags_quarterly_test],
+ [Y_monthly_train, Y_quarterly_train],
+ [Y_monthly_test, Y_quarterly_test]):
+ # We will train a model for each temporal level
+ Y_hats_tags = []
+ for level, temporal_ids_train in tags_train.items():
+ # Filter the data for the level
+ Y_level_train = Y_train.query("temporal_id in @temporal_ids_train")
+ temporal_ids_test = tags_test[level]
+ Y_level_test = Y_test.query("temporal_id in @temporal_ids_test")
+ # For each temporal level we have a different frequency and forecast horizon. We use the timestamps of the first timeseries to automatically derive the frequency & horizon of the temporally aggregated series.
+ unique_id = Y_level_train["unique_id"].iloc[0]
+ freq_level = pd.infer_freq(Y_level_train.query("unique_id == @unique_id")["ds"])
+ horizon_level = Y_level_test.query("unique_id == @unique_id")["ds"].nunique()
+ # Train a model and create forecasts
+ fcst = StatsForecast(models=[AutoARIMA()], freq=freq_level, n_jobs=-1)
+ Y_hat_level = fcst.forecast(df=Y_level_train[["ds", "unique_id", "y"]], h=horizon_level)
+ # Add the test set to the forecast
+ Y_hat_level = pd.concat([Y_level_test.reset_index(drop=True), Y_hat_level.drop(columns=["unique_id", "ds"])], axis=1)
+ # Put cols in the right order (for readability)
+ Y_hat_cols = id_cols + [col for col in Y_hat_level.columns if col not in id_cols]
+ Y_hat_level = Y_hat_level[Y_hat_cols]
+ # Append the forecast to the list
+ Y_hats_tags.append(Y_hat_level)
+
+ Y_hat_tag = pd.concat(Y_hats_tags, ignore_index=True)
+ Y_hats.append(Y_hat_tag)
+```
+
+### 2c. Reconcile forecasts
+
+We can use the
+[`HierarchicalReconciliation`](https://Nixtla.github.io/hierarchicalforecast/src/core.html#hierarchicalreconciliation)
+class to reconcile the forecasts. In this example we use
+[`BottomUp`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#bottomup)
+and `MinTrace(wls_struct)`. The latter is the ‘structural scaling’
+method introduced in [Forecasting with temporal
+hierarchies](https://robjhyndman.com/publications/temporal-hierarchies/).
+
+Note that we have to set `temporal=True` in the `reconcile` function.
+
+
+```python
+from hierarchicalforecast.methods import BottomUp, MinTrace
+from hierarchicalforecast.core import HierarchicalReconciliation
+```
+
+
+```python
+reconcilers = [
+ BottomUp(),
+ MinTrace(method="wls_struct"),
+]
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+Y_recs = []
+# We loop over the monthly and quarterly data
+for Y_hat, S, tags in zip(Y_hats,
+ [S_monthly_test, S_quarterly_test],
+ [tags_monthly_test, tags_quarterly_test]):
+ Y_rec = hrec.reconcile(Y_hat_df=Y_hat, S_df=S, tags=tags, temporal=True)
+ Y_recs.append(Y_rec)
+```
+
+## 3. Evaluation
+
+The `HierarchicalForecast` package includes the
+[`evaluate`](https://Nixtla.github.io/hierarchicalforecast/src/evaluation.html#evaluate)
+function to evaluate the different hierarchies.
+
+We evaluate the temporally aggregated forecasts *across all temporal
+aggregations*.
+
+
+```python
+from hierarchicalforecast.evaluation import evaluate
+from utilsforecast.losses import mae
+```
+
+### 3a. Monthly
+
+
+```python
+Y_rec_monthly = Y_recs[0]
+evaluation = evaluate(df = Y_rec_monthly.drop(columns = 'unique_id'),
+ tags = tags_monthly_test,
+ metrics = [mae],
+ id_col='temporal_id',
+ benchmark="AutoARIMA")
+
+evaluation.columns = ['level', 'metric', 'Base', 'BottomUp', 'MinTrace(wls_struct)']
+numeric_cols = evaluation.select_dtypes(include="number").columns
+evaluation[numeric_cols] = evaluation[numeric_cols].map('{:.2f}'.format).astype(np.float64)
+
+evaluation
+```
+
+| | level | metric | Base | BottomUp | MinTrace(wls_struct) |
+|-----|--------------|------------|------|----------|----------------------|
+| 0 | yearly | mae-scaled | 1.0 | 0.78 | 0.75 |
+| 1 | semiannually | mae-scaled | 1.0 | 0.99 | 0.95 |
+| 2 | fourmonthly | mae-scaled | 1.0 | 0.96 | 0.93 |
+| 3 | quarterly | mae-scaled | 1.0 | 0.95 | 0.93 |
+| 4 | bimonthly | mae-scaled | 1.0 | 0.96 | 0.94 |
+| 5 | monthly | mae-scaled | 1.0 | 1.00 | 0.99 |
+| 6 | Overall | mae-scaled | 1.0 | 0.94 | 0.92 |
+
+`MinTrace(wls_struct)` is the best overall method, scoring the lowest
+`mae` on all levels.
+
+### 3b. Quarterly
+
+
+```python
+Y_rec_quarterly = Y_recs[1]
+evaluation = evaluate(df = Y_rec_quarterly.drop(columns = 'unique_id'),
+ tags = tags_quarterly_test,
+ metrics = [mae],
+ id_col='temporal_id',
+ benchmark="AutoARIMA")
+
+evaluation.columns = ['level', 'metric', 'Base', 'BottomUp', 'MinTrace(wls_struct)']
+numeric_cols = evaluation.select_dtypes(include="number").columns
+evaluation[numeric_cols] = evaluation[numeric_cols].map('{:.2f}'.format).astype(np.float64)
+
+evaluation
+```
+
+| | level | metric | Base | BottomUp | MinTrace(wls_struct) |
+|-----|--------------|------------|------|----------|----------------------|
+| 0 | yearly | mae-scaled | 1.0 | 0.87 | 0.85 |
+| 1 | semiannually | mae-scaled | 1.0 | 1.03 | 1.00 |
+| 2 | quarterly | mae-scaled | 1.0 | 1.00 | 0.97 |
+| 3 | Overall | mae-scaled | 1.0 | 0.97 | 0.94 |
+
+Again, `MinTrace(wls_struct)` is the best overall method, scoring the
+lowest `mae` on all levels.
+
diff --git a/hierarchicalforecast/examples/mlframeworksexample.html.mdx b/hierarchicalforecast/examples/mlframeworksexample.html.mdx
new file mode 100644
index 00000000..c889681f
--- /dev/null
+++ b/hierarchicalforecast/examples/mlframeworksexample.html.mdx
@@ -0,0 +1,313 @@
+---
+output-file: mlframeworksexample.html
+title: Neural/MLForecast
+---
+
+
+This example notebook demonstrates the compatibility of
+HierarchicalForecast’s reconciliation methods with popular
+machine-learning libraries, specifically
+[NeuralForecast](https://github.com/Nixtla/neuralforecast) and
+[MLForecast](https://github.com/Nixtla/mlforecast).
+
+The notebook utilizes NBEATS and XGBRegressor models to create base
+forecasts for the TourismLarge Hierarchical Dataset. After that, we use
+HierarchicalForecast to reconcile the base predictions.
+
+**References** - [Boris N. Oreshkin, Dmitri Carpov, Nicolas
+Chapados, Yoshua Bengio (2019). “N-BEATS: Neural basis expansion
+analysis for interpretable time series forecasting”. url:
+https://arxiv.org/abs/1905.10437](https://arxiv.org/abs/1905.10437) -
+[Tianqi Chen and Carlos Guestrin. “XGBoost: A Scalable Tree Boosting
+System”. In: Proceedings of the 22nd ACM SIGKDD International Conference
+on Knowledge Discovery and Data Mining. KDD ’16. San Francisco,
+California, USA: Association for Computing Machinery, 2016, pp. 785–794.
+isbn: 9781450342322. doi: 10.1145/2939672.2939785. url:
+https://doi.org/10.1145/2939672.2939785 (cit. on
+p. 26).](https://doi.org/10.1145/2939672.2939785)
+
+You can run these experiments using CPU or GPU with Google Colab.
+
+
+
+## 1. Installing packages
+
+
+```python
+!pip install datasetsforecast hierarchicalforecast mlforecast neuralforecast
+```
+
+
+```python
+import numpy as np
+import pandas as pd
+
+from datasetsforecast.hierarchical import HierarchicalData
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import NBEATS
+from neuralforecast.losses.pytorch import GMM
+
+from mlforecast import MLForecast
+from mlforecast.utils import PredictionIntervals
+import xgboost as xgb
+
+#obtain hierarchical reconciliation methods and evaluation
+from hierarchicalforecast.methods import BottomUp, ERM, MinTrace
+from hierarchicalforecast.utils import HierarchicalPlot
+from hierarchicalforecast.core import HierarchicalReconciliation
+from hierarchicalforecast.evaluation import evaluate
+```
+
+## 2. Load hierarchical dataset
+
+This detailed Australian Tourism Dataset comes from the National Visitor
+Survey, managed by the Tourism Research Australia, it is composed of 555
+monthly series from 1998 to 2016, it is organized geographically, and
+purpose of travel. The natural geographical hierarchy comprises seven
+states, divided further in 27 zones and 76 regions. The purpose of
+travel categories are holiday, visiting friends and relatives (VFR),
+business and other. The MinT (Wickramasuriya et al., 2019), among other
+hierarchical forecasting studies has used the dataset it in the past.
+The dataset can be accessed in the [MinT reconciliation
+webpage](https://robjhyndman.com/publications/mint/), although other
+sources are available.
+
+| Geographical Division | Number of series per division | Number of series per purpose | Total |
+|------------------|------------------|------------------|------------------|
+| Australia | 1 | 4 | 5 |
+| States | 7 | 28 | 35 |
+| Zones | 27 | 108 | 135 |
+| Regions | 76 | 304 | 380 |
+| Total | 111 | 444 | 555 |
+
+
+```python
+Y_df, S_df, tags = HierarchicalData.load('./data', 'TourismLarge')
+Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+S_df = S_df.reset_index(names="unique_id")
+```
+
+
+```python
+Y_df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|--------------|
+| 0 | TotalAll | 1998-01-01 | 45151.071280 |
+| 1 | TotalAll | 1998-02-01 | 17294.699551 |
+| 2 | TotalAll | 1998-03-01 | 20725.114184 |
+| 3 | TotalAll | 1998-04-01 | 25388.612353 |
+| 4 | TotalAll | 1998-05-01 | 20330.035211 |
+
+Visualize the aggregation matrix.
+
+
+```python
+hplot = HierarchicalPlot(S=S_df, tags=tags)
+hplot.plot_summing_matrix()
+```
+
+
+
+Split the dataframe in train/test splits.
+
+
+```python
+horizon = 12
+Y_test_df = Y_df.groupby('unique_id', as_index=False).tail(horizon)
+Y_train_df = Y_df.drop(Y_test_df.index)
+```
+
+## 3. Fit and Predict Models
+
+HierarchicalForecast is compatible with many different ML models. Here,
+we show two examples: 1. NBEATS, a MLP-based deep neural
+architecture. 2. XGBRegressor, a tree-based architecture.
+
+
+```python
+level = np.arange(0, 100, 2)
+qs = [[50-lv/2, 50+lv/2] for lv in level]
+quantiles = np.sort(np.concatenate(qs)[1:]/100)
+
+#fit/predict NBEATS from NeuralForecast
+nbeats = NBEATS(h=horizon,
+ input_size=2*horizon,
+ loss=GMM(n_components=10, quantiles=quantiles),
+ scaler_type='robust',
+ max_steps=2000)
+nf = NeuralForecast(models=[nbeats], freq='MS')
+nf.fit(df=Y_train_df)
+Y_hat_nf = nf.predict()
+insample_nf = nf.predict_insample(step_size=horizon)
+
+#fit/predict XGBRegressor from MLForecast
+mf = MLForecast(models=[xgb.XGBRegressor()],
+ freq='MS',
+ lags=[1,2,12,24],
+ date_features=['month'],
+ )
+mf.fit(Y_train_df, fitted=True, prediction_intervals=PredictionIntervals(n_windows=10, h=horizon))
+Y_hat_mf = mf.predict(horizon, level=level)
+insample_mf = mf.forecast_fitted_values()
+```
+
+
+```python
+Y_hat_nf
+```
+
+| | unique_id | ds | NBEATS | NBEATS-lo-98.0 | NBEATS-lo-96.0 | NBEATS-lo-94.0 | NBEATS-lo-92.0 | NBEATS-lo-90.0 | NBEATS-lo-88.0 | NBEATS-lo-86.0 | ... | NBEATS-hi-80.0 | NBEATS-hi-82.0 | NBEATS-hi-84.0 | NBEATS-hi-86.0 | NBEATS-hi-88.0 | NBEATS-hi-90.0 | NBEATS-hi-92.0 | NBEATS-hi-94.0 | NBEATS-hi-96.0 | NBEATS-hi-98.0 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | AAAAll | 2016-01-01 | 2843.298584 | 1764.249023 | 1806.885132 | 1864.019043 | 1906.171021 | 1945.994629 | 1965.081421 | 1998.606812 | ... | 3497.682373 | 3520.107666 | 3561.643799 | 3600.121094 | 3646.954346 | 3703.382324 | 3774.084473 | 3813.719238 | 3902.713867 | 3991.594238 |
+| 1 | AAAAll | 2016-02-01 | 1753.340698 | 1394.245850 | 1414.474976 | 1439.167480 | 1458.228394 | 1474.655640 | 1480.433472 | 1489.651245 | ... | 2024.560791 | 2049.965576 | 2066.480957 | 2090.285156 | 2120.172852 | 2145.964844 | 2201.716064 | 2253.415039 | 2364.905029 | 2441.167480 |
+| 2 | AAAAll | 2016-03-01 | 1878.675171 | 1446.630371 | 1491.637817 | 1513.890137 | 1524.787842 | 1532.539917 | 1547.460205 | 1559.098389 | ... | 2172.270996 | 2189.489990 | 2216.255859 | 2236.661377 | 2286.617676 | 2370.431152 | 2411.910156 | 2477.557373 | 2579.611084 | 2722.415283 |
+| 3 | AAAAll | 2016-04-01 | 2140.948486 | 1661.737793 | 1706.259399 | 1724.914551 | 1736.446045 | 1754.887695 | 1765.482056 | 1772.123901 | ... | 2470.206543 | 2483.571045 | 2493.527588 | 2517.062744 | 2547.355713 | 2577.867676 | 2610.180908 | 2637.010498 | 2700.801758 | 2864.596924 |
+| 4 | AAAAll | 2016-05-01 | 1834.694946 | 1466.314209 | 1485.427002 | 1500.715210 | 1518.462036 | 1535.386475 | 1543.525635 | 1554.429810 | ... | 2093.700684 | 2120.782471 | 2137.882812 | 2154.052002 | 2164.069824 | 2189.309326 | 2234.271973 | 2311.157715 | 2436.267090 | 2659.653809 |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| 6655 | TotalVis | 2016-08-01 | 7362.455078 | 5799.121582 | 5960.676270 | 6073.553223 | 6230.090820 | 6294.191406 | 6365.950684 | 6400.492676 | ... | 8120.279785 | 8144.139648 | 8185.699219 | 8212.809570 | 8255.871094 | 8291.191406 | 8374.907227 | 8435.806641 | 8568.060547 | 8770.566406 |
+| 6656 | TotalVis | 2016-09-01 | 7803.098145 | 6455.050293 | 6612.847168 | 6690.960938 | 6804.897461 | 6848.432617 | 6873.607422 | 6904.770020 | ... | 8562.215820 | 8594.000000 | 8642.083984 | 8715.201172 | 8795.628906 | 8924.573242 | 9053.747070 | 9250.514648 | 9410.338867 | 9818.623047 |
+| 6657 | TotalVis | 2016-10-01 | 8478.570312 | 6592.350098 | 6818.883789 | 7075.323730 | 7223.682129 | 7300.230957 | 7336.740723 | 7391.779785 | ... | 9558.611328 | 9586.333984 | 9658.816406 | 9761.448242 | 9802.087891 | 9870.294922 | 9956.144531 | 10070.672852 | 10195.408203 | 10342.619141 |
+| 6658 | TotalVis | 2016-11-01 | 8251.816406 | 6471.753906 | 6551.861328 | 6621.647461 | 6694.992188 | 6740.827148 | 6798.824707 | 6825.794434 | ... | 9519.825195 | 9557.507812 | 9624.822266 | 9720.269531 | 9811.011719 | 9907.259766 | 10132.628906 | 10362.583984 | 10896.478516 | 11394.652344 |
+| 6659 | TotalVis | 2016-12-01 | 9023.334961 | 6798.515625 | 6978.411621 | 7165.805176 | 7250.106934 | 7333.168457 | 7395.183594 | 7457.470215 | ... | 10221.937500 | 10290.527344 | 10334.883789 | 10399.726562 | 10553.360352 | 10645.852539 | 10806.295898 | 10992.416016 | 11328.151367 | 11933.357422 |
+
+
+```python
+Y_hat_mf
+```
+
+| | unique_id | ds | XGBRegressor | XGBRegressor-lo-98 | XGBRegressor-lo-96 | XGBRegressor-lo-94 | XGBRegressor-lo-92 | XGBRegressor-lo-90 | XGBRegressor-lo-88 | XGBRegressor-lo-86 | ... | XGBRegressor-hi-80 | XGBRegressor-hi-82 | XGBRegressor-hi-84 | XGBRegressor-hi-86 | XGBRegressor-hi-88 | XGBRegressor-hi-90 | XGBRegressor-hi-92 | XGBRegressor-hi-94 | XGBRegressor-hi-96 | XGBRegressor-hi-98 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | AAAAll | 2016-01-01 | 3240.743164 | 2566.404620 | 2638.984995 | 2711.565370 | 2784.145745 | 2856.726120 | 2876.514198 | 2877.447884 | ... | 3601.237386 | 3602.171072 | 3603.104758 | 3604.038444 | 3604.972130 | 3624.760208 | 3697.340583 | 3769.920958 | 3842.501333 | 3915.081708 |
+| 1 | AAAAll | 2016-02-01 | 1583.065063 | 1247.414469 | 1248.895343 | 1250.376217 | 1251.857091 | 1253.337965 | 1263.627340 | 1277.062610 | ... | 1848.761709 | 1862.196978 | 1875.632248 | 1889.067517 | 1902.502787 | 1912.792162 | 1914.273036 | 1915.753910 | 1917.234784 | 1918.715658 |
+| 2 | AAAAll | 2016-03-01 | 2030.168213 | 1345.896497 | 1386.655046 | 1427.413595 | 1468.172144 | 1508.930693 | 1546.207337 | 1582.240444 | ... | 2369.996660 | 2406.029767 | 2442.062874 | 2478.095981 | 2514.129089 | 2551.405733 | 2592.164282 | 2632.922831 | 2673.681380 | 2714.439928 |
+| 3 | AAAAll | 2016-04-01 | 2152.282227 | 1767.276611 | 1772.956049 | 1778.635487 | 1784.314926 | 1789.994364 | 1798.503584 | 1808.023439 | ... | 2467.981448 | 2477.501303 | 2487.021159 | 2496.541014 | 2506.060870 | 2514.570089 | 2520.249527 | 2525.928966 | 2531.608404 | 2537.287842 |
+| 4 | AAAAll | 2016-05-01 | 1970.894775 | 1476.761973 | 1510.667430 | 1544.572887 | 1578.478344 | 1612.383801 | 1625.448072 | 1631.069062 | ... | 2293.857519 | 2299.478509 | 2305.099499 | 2310.720489 | 2316.341479 | 2329.405750 | 2363.311207 | 2397.216664 | 2431.122121 | 2465.027578 |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| 6655 | TotalVis | 2016-08-01 | 7810.465820 | 6251.079674 | 6268.924727 | 6286.769780 | 6304.614833 | 6322.459886 | 6375.977772 | 6442.235956 | ... | 8979.921135 | 9046.179318 | 9112.437501 | 9178.695685 | 9244.953868 | 9298.471754 | 9316.316807 | 9334.161860 | 9352.006913 | 9369.851967 |
+| 6656 | TotalVis | 2016-09-01 | 6887.893555 | 5346.477959 | 5397.795065 | 5449.112170 | 5500.429275 | 5551.746380 | 5604.124112 | 5656.880638 | ... | 7960.636893 | 8013.393419 | 8066.149945 | 8118.906472 | 8171.662998 | 8224.040729 | 8275.357834 | 8326.674940 | 8377.992045 | 8429.309150 |
+| 6657 | TotalVis | 2016-10-01 | 7763.275879 | 6138.534738 | 6267.740281 | 6396.945824 | 6526.151367 | 6655.356910 | 6706.009194 | 6728.606744 | ... | 8730.152366 | 8752.749916 | 8775.347465 | 8797.945014 | 8820.542563 | 8871.194848 | 9000.400391 | 9129.605934 | 9258.811477 | 9388.017020 |
+| 6658 | TotalVis | 2016-11-01 | 7432.722168 | 5703.395148 | 5726.926242 | 5750.457336 | 5773.988430 | 5797.519524 | 5929.164698 | 6099.422043 | ... | 8255.250258 | 8425.507603 | 8595.764948 | 8766.022293 | 8936.279638 | 9067.924811 | 9091.455905 | 9114.986999 | 9138.518093 | 9162.049187 |
+| 6659 | TotalVis | 2016-12-01 | 9624.172852 | 8115.705498 | 8217.381077 | 8319.056655 | 8420.732234 | 8522.407812 | 8566.581883 | 8590.219701 | ... | 10587.212548 | 10610.850366 | 10634.488184 | 10658.126002 | 10681.763820 | 10725.937891 | 10827.613470 | 10929.289048 | 11030.964626 | 11132.640205 |
+
+## 4. Reconcile Predictions
+
+With minimal parsing, we can reconcile the raw output predictions with
+different HierarchicalForecast reconciliation methods.
+
+
+```python
+reconcilers = [
+ ERM(method='closed'),
+ BottomUp(),
+ MinTrace('mint_shrink'),
+]
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+
+Y_rec_nf = hrec.reconcile(Y_hat_df=Y_hat_nf, Y_df=insample_nf, S_df=S_df, tags=tags, level=level)
+Y_rec_mf = hrec.reconcile(Y_hat_df=Y_hat_mf, Y_df=insample_mf, S_df=S_df, tags=tags, level=level)
+```
+
+## 5. Evaluation
+
+To evaluate we use a scaled variation of the CRPS, as proposed by
+Rangapuram (2021), to measure the accuracy of predicted quantiles
+`y_hat` compared to the observation `y`.
+
+$$
+
+\mathrm{sCRPS}(\hat{F}_{\tau}, \mathbf{y}_{\tau}) = \frac{2}{N} \sum_{i}
+\int^{1}_{0}
+\frac{\mathrm{QL}(\hat{F}_{i,\tau}, y_{i,\tau})_{q}}{\sum_{i} | y_{i,\tau} |} dq
+
+$$
+
+We find that XGB with MinTrace(mint_shrink) reconciliation result in the
+lowest CRPS score on the test set, thus giving us the best probabilistic
+forecasts.
+
+
+```python
+from utilsforecast.losses import scaled_crps
+```
+
+
+```python
+rec_model_names_nf = ['NBEATS/BottomUp', 'NBEATS/MinTrace_method-mint_shrink', 'NBEATS/ERM_method-closed_lambda_reg-0.01']
+
+evaluation_nf = evaluate(df = Y_rec_nf.merge(Y_test_df, on=['unique_id', 'ds']),
+ tags = tags,
+ metrics = [scaled_crps],
+ models= rec_model_names_nf,
+ level = list(range(0, 100, 2)),
+ )
+
+rec_model_names_mf = ['XGBRegressor/BottomUp', 'XGBRegressor/MinTrace_method-mint_shrink', 'XGBRegressor/ERM_method-closed_lambda_reg-0.01']
+
+evaluation_mf = evaluate(df = Y_rec_mf.merge(Y_test_df, on=['unique_id', 'ds']),
+ tags = tags,
+ metrics = [scaled_crps],
+ models= rec_model_names_mf,
+ level = list(range(0, 100, 2)),
+ )
+```
+
+
+```python
+name = 'NBEATS/BottomUp'
+quantile_columns = [col for col in Y_rec_mf.columns if (name+'-lo') in col or (name+'-hi') in col]
+```
+
+
+```python
+evaluation_nf.query("level == 'Overall'")
+```
+
+| | level | metric | NBEATS/BottomUp | NBEATS/MinTrace_method-mint_shrink | NBEATS/ERM_method-closed_lambda_reg-0.01 |
+|----|----|----|----|----|----|
+| 8 | Overall | scaled_crps | 2.523212 | 2.43205 | 2.645045 |
+
+
+```python
+evaluation_mf.query("level == 'Overall'")
+```
+
+| | level | metric | XGBRegressor/BottomUp | XGBRegressor/MinTrace_method-mint_shrink | XGBRegressor/ERM_method-closed_lambda_reg-0.01 |
+|----|----|----|----|----|----|
+| 8 | Overall | scaled_crps | 1.98255 | 1.44981 | 1.910014 |
+
+## 6. Visualizations
+
+
+```python
+plot_nf = Y_df.merge(Y_rec_nf, on=['unique_id', 'ds'], how="outer")
+
+plot_mf = Y_df.merge(Y_rec_mf, on=['unique_id', 'ds'], how="outer")
+```
+
+
+```python
+hplot.plot_series(
+ series='TotalVis',
+ Y_df=plot_nf,
+ models=['y', 'NBEATS', 'NBEATS/BottomUp', 'NBEATS/MinTrace_method-mint_shrink', 'NBEATS/ERM_method-closed_lambda_reg-0.01'],
+ level=[80]
+)
+```
+
+
+
+
+```python
+hplot.plot_series(
+ series='TotalVis',
+ Y_df=plot_mf,
+ models=['y', 'XGBRegressor', 'XGBRegressor/BottomUp', 'XGBRegressor/MinTrace_method-mint_shrink', 'XGBRegressor/ERM_method-closed_lambda_reg-0.01'],
+ level=[80]
+)
+```
+
+
+
diff --git a/hierarchicalforecast/examples/nonnegativereconciliation.html.mdx b/hierarchicalforecast/examples/nonnegativereconciliation.html.mdx
new file mode 100644
index 00000000..da20d016
--- /dev/null
+++ b/hierarchicalforecast/examples/nonnegativereconciliation.html.mdx
@@ -0,0 +1,328 @@
+---
+output-file: nonnegativereconciliation.html
+title: Non-Negative MinTrace
+---
+
+
+Large collections of time series organized into structures at different
+aggregation levels often require their forecasts to follow their
+aggregation constraints and to be nonnegative, which poses the challenge
+of creating novel algorithms capable of coherent forecasts.
+
+The `HierarchicalForecast` package provides a wide collection of Python
+implementations of hierarchical forecasting algorithms that follow
+nonnegative hierarchical reconciliation.
+
+In this notebook, we will show how to use the `HierarchicalForecast`
+package to perform nonnegative reconciliation of forecasts on `Wiki2`
+dataset.
+
+You can run these experiments using CPU or GPU with Google Colab.
+
+
+
+
+```python
+!pip install hierarchicalforecast statsforecast datasetsforecast
+```
+
+## 1. Load Data
+
+In this example we will use the `Wiki2` dataset. The following cell gets
+the time series for the different levels in the hierarchy, the summing
+dataframe `S_df` which recovers the full dataset from the bottom level
+hierarchy and the indices of each hierarchy denoted by `tags`.
+
+
+```python
+import numpy as np
+import pandas as pd
+
+from datasetsforecast.hierarchical import HierarchicalData
+```
+
+
+```python
+Y_df, S_df, tags = HierarchicalData.load('./data', 'Wiki2')
+Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+S_df = S_df.reset_index(names="unique_id")
+```
+
+
+```python
+Y_df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|--------|
+| 0 | Total | 2016-01-01 | 156508 |
+| 1 | Total | 2016-01-02 | 129902 |
+| 2 | Total | 2016-01-03 | 138203 |
+| 3 | Total | 2016-01-04 | 115017 |
+| 4 | Total | 2016-01-05 | 126042 |
+
+
+```python
+S_df.iloc[:5, :5]
+```
+
+| | unique_id | de_AAC_AAG_001 | de_AAC_AAG_010 | de_AAC_AAG_014 | de_AAC_AAG_045 |
+|-----|-----------|----------------|----------------|----------------|----------------|
+| 0 | Total | 1 | 1 | 1 | 1 |
+| 1 | de | 1 | 1 | 1 | 1 |
+| 2 | en | 0 | 0 | 0 | 0 |
+| 3 | fr | 0 | 0 | 0 | 0 |
+| 4 | ja | 0 | 0 | 0 | 0 |
+
+
+```python
+tags
+```
+
+``` text
+{'Views': array(['Total'], dtype=object),
+ 'Views/Country': array(['de', 'en', 'fr', 'ja', 'ru', 'zh'], dtype=object),
+ 'Views/Country/Access': array(['de_AAC', 'de_DES', 'de_MOB', 'en_AAC', 'en_DES', 'en_MOB',
+ 'fr_AAC', 'fr_DES', 'fr_MOB', 'ja_AAC', 'ja_DES', 'ja_MOB',
+ 'ru_AAC', 'ru_DES', 'ru_MOB', 'zh_AAC', 'zh_DES', 'zh_MOB'],
+ dtype=object),
+ 'Views/Country/Access/Agent': array(['de_AAC_AAG', 'de_AAC_SPD', 'de_DES_AAG', 'de_MOB_AAG',
+ 'en_AAC_AAG', 'en_AAC_SPD', 'en_DES_AAG', 'en_MOB_AAG',
+ 'fr_AAC_AAG', 'fr_AAC_SPD', 'fr_DES_AAG', 'fr_MOB_AAG',
+ 'ja_AAC_AAG', 'ja_AAC_SPD', 'ja_DES_AAG', 'ja_MOB_AAG',
+ 'ru_AAC_AAG', 'ru_AAC_SPD', 'ru_DES_AAG', 'ru_MOB_AAG',
+ 'zh_AAC_AAG', 'zh_AAC_SPD', 'zh_DES_AAG', 'zh_MOB_AAG'],
+ dtype=object),
+ 'Views/Country/Access/Agent/Topic': array(['de_AAC_AAG_001', 'de_AAC_AAG_010', 'de_AAC_AAG_014',
+ 'de_AAC_AAG_045', 'de_AAC_AAG_063', 'de_AAC_AAG_100',
+ 'de_AAC_AAG_110', 'de_AAC_AAG_123', 'de_AAC_AAG_143',
+ 'de_AAC_SPD_012', 'de_AAC_SPD_074', 'de_AAC_SPD_080',
+ 'de_AAC_SPD_105', 'de_AAC_SPD_115', 'de_AAC_SPD_133',
+ 'de_DES_AAG_064', 'de_DES_AAG_116', 'de_DES_AAG_131',
+ 'de_MOB_AAG_015', 'de_MOB_AAG_020', 'de_MOB_AAG_032',
+ 'de_MOB_AAG_059', 'de_MOB_AAG_062', 'de_MOB_AAG_088',
+ 'de_MOB_AAG_095', 'de_MOB_AAG_109', 'de_MOB_AAG_122',
+ 'de_MOB_AAG_149', 'en_AAC_AAG_044', 'en_AAC_AAG_049',
+ 'en_AAC_AAG_075', 'en_AAC_AAG_114', 'en_AAC_AAG_119',
+ 'en_AAC_AAG_141', 'en_AAC_SPD_004', 'en_AAC_SPD_011',
+ 'en_AAC_SPD_026', 'en_AAC_SPD_048', 'en_AAC_SPD_067',
+ 'en_AAC_SPD_126', 'en_AAC_SPD_140', 'en_DES_AAG_016',
+ 'en_DES_AAG_024', 'en_DES_AAG_042', 'en_DES_AAG_069',
+ 'en_DES_AAG_082', 'en_DES_AAG_102', 'en_MOB_AAG_018',
+ 'en_MOB_AAG_022', 'en_MOB_AAG_101', 'en_MOB_AAG_124',
+ 'fr_AAC_AAG_029', 'fr_AAC_AAG_046', 'fr_AAC_AAG_070',
+ 'fr_AAC_AAG_087', 'fr_AAC_AAG_098', 'fr_AAC_AAG_104',
+ 'fr_AAC_AAG_111', 'fr_AAC_AAG_112', 'fr_AAC_AAG_142',
+ 'fr_AAC_SPD_025', 'fr_AAC_SPD_027', 'fr_AAC_SPD_035',
+ 'fr_AAC_SPD_077', 'fr_AAC_SPD_084', 'fr_AAC_SPD_097',
+ 'fr_AAC_SPD_130', 'fr_DES_AAG_023', 'fr_DES_AAG_043',
+ 'fr_DES_AAG_051', 'fr_DES_AAG_058', 'fr_DES_AAG_061',
+ 'fr_DES_AAG_091', 'fr_DES_AAG_093', 'fr_DES_AAG_094',
+ 'fr_DES_AAG_136', 'fr_MOB_AAG_006', 'fr_MOB_AAG_030',
+ 'fr_MOB_AAG_066', 'fr_MOB_AAG_117', 'fr_MOB_AAG_120',
+ 'fr_MOB_AAG_121', 'fr_MOB_AAG_135', 'fr_MOB_AAG_147',
+ 'ja_AAC_AAG_038', 'ja_AAC_AAG_047', 'ja_AAC_AAG_055',
+ 'ja_AAC_AAG_076', 'ja_AAC_AAG_099', 'ja_AAC_AAG_128',
+ 'ja_AAC_AAG_132', 'ja_AAC_AAG_134', 'ja_AAC_AAG_137',
+ 'ja_AAC_SPD_013', 'ja_AAC_SPD_034', 'ja_AAC_SPD_050',
+ 'ja_AAC_SPD_060', 'ja_AAC_SPD_078', 'ja_AAC_SPD_106',
+ 'ja_DES_AAG_079', 'ja_DES_AAG_081', 'ja_DES_AAG_113',
+ 'ja_MOB_AAG_065', 'ja_MOB_AAG_073', 'ja_MOB_AAG_092',
+ 'ja_MOB_AAG_127', 'ja_MOB_AAG_129', 'ja_MOB_AAG_144',
+ 'ru_AAC_AAG_008', 'ru_AAC_AAG_145', 'ru_AAC_AAG_146',
+ 'ru_AAC_SPD_000', 'ru_AAC_SPD_090', 'ru_AAC_SPD_148',
+ 'ru_DES_AAG_003', 'ru_DES_AAG_007', 'ru_DES_AAG_017',
+ 'ru_DES_AAG_041', 'ru_DES_AAG_071', 'ru_DES_AAG_072',
+ 'ru_MOB_AAG_002', 'ru_MOB_AAG_040', 'ru_MOB_AAG_083',
+ 'ru_MOB_AAG_086', 'ru_MOB_AAG_103', 'ru_MOB_AAG_107',
+ 'ru_MOB_AAG_118', 'ru_MOB_AAG_125', 'zh_AAC_AAG_021',
+ 'zh_AAC_AAG_033', 'zh_AAC_AAG_037', 'zh_AAC_AAG_052',
+ 'zh_AAC_AAG_057', 'zh_AAC_AAG_085', 'zh_AAC_AAG_108',
+ 'zh_AAC_SPD_039', 'zh_AAC_SPD_096', 'zh_DES_AAG_009',
+ 'zh_DES_AAG_019', 'zh_DES_AAG_053', 'zh_DES_AAG_054',
+ 'zh_DES_AAG_056', 'zh_DES_AAG_068', 'zh_DES_AAG_089',
+ 'zh_DES_AAG_139', 'zh_MOB_AAG_005', 'zh_MOB_AAG_028',
+ 'zh_MOB_AAG_031', 'zh_MOB_AAG_036', 'zh_MOB_AAG_138'], dtype=object)}
+```
+
+We split the dataframe in train/test splits.
+
+
+```python
+Y_test_df = Y_df.groupby('unique_id', as_index=False).tail(7)
+Y_train_df = Y_df.drop(Y_test_df.index)
+```
+
+## 2. Base Forecasts
+
+The following cell computes the *base forecast* for each time series
+using the `AutoETS` model. Observe that `Y_hat_df` contains the
+forecasts but they are not coherent.
+
+
+```python
+from statsforecast.models import AutoETS, Naive
+from statsforecast.core import StatsForecast
+```
+
+
+```python
+fcst = StatsForecast(
+ models=[AutoETS(season_length=7, model='ZAA'), Naive()],
+ freq='D',
+ n_jobs=-1
+)
+Y_hat_df = fcst.forecast(df=Y_train_df, h=7)
+```
+
+Observe that the `AutoETS` model computes negative forecasts for some
+series.
+
+
+```python
+Y_hat_df.query('AutoETS < 0')
+```
+
+| | unique_id | ds | AutoETS | Naive |
+|------|----------------|------------|-------------|--------|
+| 28 | de_AAC_AAG_001 | 2016-12-25 | -523.766907 | 340.0 |
+| 29 | de_AAC_AAG_001 | 2016-12-26 | -245.337433 | 340.0 |
+| 30 | de_AAC_AAG_001 | 2016-12-27 | -194.253815 | 340.0 |
+| 33 | de_AAC_AAG_001 | 2016-12-30 | -315.425659 | 340.0 |
+| 34 | de_AAC_AAG_001 | 2016-12-31 | -806.920105 | 340.0 |
+| ... | ... | ... | ... | ... |
+| 1217 | zh_AAC_AAG_033 | 2016-12-31 | -86.466789 | 37.0 |
+| 1345 | zh_MOB | 2016-12-26 | -199.534882 | 1036.0 |
+| 1346 | zh_MOB | 2016-12-27 | -69.527260 | 1036.0 |
+| 1352 | zh_MOB_AAG | 2016-12-26 | -199.534882 | 1036.0 |
+| 1353 | zh_MOB_AAG | 2016-12-27 | -69.527260 | 1036.0 |
+
+## 3. Non-Negative Reconciliation
+
+The following cell makes the previous forecasts coherent and nonnegative
+using the
+[`HierarchicalReconciliation`](https://Nixtla.github.io/hierarchicalforecast/src/core.html#hierarchicalreconciliation)
+class.
+
+
+```python
+from hierarchicalforecast.methods import MinTrace
+from hierarchicalforecast.core import HierarchicalReconciliation
+```
+
+
+```python
+reconcilers = [
+ MinTrace(method='ols'),
+ MinTrace(method='ols', nonnegative=True)
+]
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+Y_rec_df = hrec.reconcile(Y_hat_df=Y_hat_df, Y_df=Y_train_df,
+ S_df=S_df, tags=tags)
+```
+
+Observe that the nonnegative reconciliation method obtains nonnegative
+forecasts.
+
+
+```python
+Y_rec_df
+```
+
+| | unique_id | ds | AutoETS | Naive | AutoETS/MinTrace_method-ols | Naive/MinTrace_method-ols | AutoETS/MinTrace_method-ols_nonnegative-True | Naive/MinTrace_method-ols_nonnegative-True |
+|----|----|----|----|----|----|----|----|----|
+| 0 | Total | 2016-12-25 | 94523.164062 | 95743.0 | 95852.000421 | 95743.0 | 9.664245e+04 | 95743.0 |
+| 1 | Total | 2016-12-26 | 87734.367188 | 95743.0 | 89525.238276 | 95743.0 | 9.028857e+04 | 95743.0 |
+| 2 | Total | 2016-12-27 | 87751.125000 | 95743.0 | 89638.119184 | 95743.0 | 9.056593e+04 | 95743.0 |
+| 3 | Total | 2016-12-28 | 133237.968750 | 95743.0 | 131051.839057 | 95743.0 | 1.314028e+05 | 95743.0 |
+| 4 | Total | 2016-12-29 | 126501.796875 | 95743.0 | 121214.048604 | 95743.0 | 1.218000e+05 | 95743.0 |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| 1388 | zh_MOB_AAG_138 | 2016-12-27 | 62.049744 | 65.0 | -147.399760 | 65.0 | 0.000000e+00 | 65.0 |
+| 1389 | zh_MOB_AAG_138 | 2016-12-28 | 54.934032 | 65.0 | 7.561682 | 65.0 | 4.397229e-15 | 65.0 |
+| 1390 | zh_MOB_AAG_138 | 2016-12-29 | 60.452618 | 65.0 | 114.253489 | 65.0 | 9.321380e+01 | 65.0 |
+| 1391 | zh_MOB_AAG_138 | 2016-12-30 | 50.356693 | 65.0 | 96.446754 | 65.0 | 7.565171e+01 | 65.0 |
+| 1392 | zh_MOB_AAG_138 | 2016-12-31 | 66.735626 | 65.0 | 208.184648 | 65.0 | 1.851130e+02 | 65.0 |
+
+
+```python
+Y_rec_df.query('`AutoETS/MinTrace_method-ols_nonnegative-True` < 0')
+```
+
+| | unique_id | ds | AutoETS | Naive | AutoETS/MinTrace_method-ols | Naive/MinTrace_method-ols | AutoETS/MinTrace_method-ols_nonnegative-True | Naive/MinTrace_method-ols_nonnegative-True |
+|----|----|----|----|----|----|----|----|----|
+
+The free reconciliation method gets negative forecasts.
+
+
+```python
+Y_rec_df.query('`AutoETS/MinTrace_method-ols` < 0')
+```
+
+| | unique_id | ds | AutoETS | Naive | AutoETS/MinTrace_method-ols | Naive/MinTrace_method-ols | AutoETS/MinTrace_method-ols_nonnegative-True | Naive/MinTrace_method-ols_nonnegative-True |
+|----|----|----|----|----|----|----|----|----|
+| 56 | de_DES | 2016-12-25 | -2553.932861 | 495.0 | -3818.990043 | 495.0 | 0.000000e+00 | 495.0 |
+| 57 | de_DES | 2016-12-26 | -2155.228271 | 495.0 | -3309.806933 | 495.0 | 1.909922e-30 | 495.0 |
+| 58 | de_DES | 2016-12-27 | -2720.993896 | 495.0 | -3965.351121 | 495.0 | 1.140223e-13 | 495.0 |
+| 60 | de_DES | 2016-12-29 | -3429.432617 | 495.0 | -3042.502484 | 495.0 | 3.049601e+02 | 495.0 |
+| 61 | de_DES | 2016-12-30 | -3963.202637 | 495.0 | -3476.273292 | 495.0 | 2.877829e+02 | 495.0 |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| 1380 | zh_MOB_AAG_036 | 2016-12-26 | 75.298317 | 115.0 | -166.245228 | 115.0 | 0.000000e+00 | 115.0 |
+| 1381 | zh_MOB_AAG_036 | 2016-12-27 | 72.895554 | 115.0 | -136.553950 | 115.0 | 1.699002e-14 | 115.0 |
+| 1386 | zh_MOB_AAG_138 | 2016-12-25 | 94.796623 | 65.0 | -49.410174 | 65.0 | 0.000000e+00 | 65.0 |
+| 1387 | zh_MOB_AAG_138 | 2016-12-26 | 71.293983 | 65.0 | -170.249562 | 65.0 | 0.000000e+00 | 65.0 |
+| 1388 | zh_MOB_AAG_138 | 2016-12-27 | 62.049744 | 65.0 | -147.399760 | 65.0 | 0.000000e+00 | 65.0 |
+
+## 4. Evaluation
+
+The `HierarchicalForecast` package includes the
+[`evaluate`](https://Nixtla.github.io/hierarchicalforecast/src/evaluation.html#evaluate)
+function to evaluate the different hierarchies. We use `utilsforecast`
+to compute the mean absolute error.
+
+
+```python
+from hierarchicalforecast.evaluation import evaluate
+from utilsforecast.losses import mse
+```
+
+
+```python
+evaluation = evaluate(df = Y_rec_df.merge(Y_test_df, on=['unique_id', 'ds']),
+ tags = tags,
+ train_df = Y_train_df,
+ metrics = [mse],
+ benchmark="Naive")
+
+evaluation.set_index(["level", "metric"]).filter(like='ETS')
+```
+
+| | | AutoETS | AutoETS/MinTrace_method-ols | AutoETS/MinTrace_method-ols_nonnegative-True |
+|----|----|----|----|----|
+| level | metric | | | |
+| Views | mse-scaled | 0.735800 | 0.697371 | 0.675672 |
+| Views/Country | mse-scaled | 1.190354 | 1.053631 | 0.994758 |
+| Views/Country/Access | mse-scaled | 1.086102 | 1.133507 | 1.172270 |
+| Views/Country/Access/Agent | mse-scaled | 1.067394 | 1.100215 | 1.127960 |
+| Views/Country/Access/Agent/Topic | mse-scaled | 1.435105 | 1.381990 | 1.163428 |
+| Overall | mse-scaled | 1.010801 | 0.977667 | 0.939286 |
+
+Observe that the nonnegative reconciliation method performs better
+(lower error) than its unconstrained counterpart.
+
+### References
+
+- [Hyndman, R.J., & Athanasopoulos, G. (2021). “Forecasting:
+ principles and practice, 3rd edition: Chapter 11: Forecasting
+ hierarchical and grouped series.”. OTexts: Melbourne, Australia.
+ OTexts.com/fpp3 Accessed on July
+ 2022.](https://otexts.com/fpp3/hierarchical.html)
+- [Wickramasuriya, S. L., Athanasopoulos, G., & Hyndman, R. J. (2019).
+ "Optimal forecast reconciliation for hierarchical and grouped time
+ series through trace minimization". Journal of the American
+ Statistical Association, 114 , 804–819.
+ doi:10.1080/01621459.2018.1448825.](https://robjhyndman.com/publications/mint/).
+- [Wickramasuriya, S.L., Turlach, B.A. & Hyndman, R.J. (2020).
+ "Optimal non-negative forecast reconciliation”. Stat Comput 30,
+ 1167–1182,
+ https://doi.org/10.1007/s11222-020-09930-0](https://robjhyndman.com/publications/nnmint/).
+
diff --git a/hierarchicalforecast/examples/tourismlarge-evaluation.html.mdx b/hierarchicalforecast/examples/tourismlarge-evaluation.html.mdx
new file mode 100644
index 00000000..d464993e
--- /dev/null
+++ b/hierarchicalforecast/examples/tourismlarge-evaluation.html.mdx
@@ -0,0 +1,277 @@
+---
+description: Hierarchical Forecast's reconciliation and evaluation.
+output-file: tourismlarge-evaluation.html
+title: Probabilistic Forecast Evaluation
+---
+
+
+This notebook offers a step to step guide to create a hierarchical
+forecasting pipeline.
+
+In the pipeline we will use `HierarchicalForecast` and `StatsForecast`
+core class, to create base predictions, reconcile and evaluate them.
+
+We will use the TourismL dataset that summarizes large Australian
+national visitor survey.
+
+Outline 1. Installing Packages 2. Prepare TourismL dataset - Read and
+aggregate - StatsForecast’s Base Predictions 3. Reconciliar 4. Evaluar
+
+
+
+## 1. Installing HierarchicalForecast
+
+We assume you have StatsForecast and HierarchicalForecast already
+installed, if not check this guide for instructions on how to install
+HierarchicalForecast.
+
+
+```python
+!pip install hierarchicalforecast statsforecast datasetsforecast
+```
+
+
+```python
+import os
+import numpy as np
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from statsforecast.core import StatsForecast
+from statsforecast.models import AutoARIMA, Naive
+
+from hierarchicalforecast.core import HierarchicalReconciliation
+from hierarchicalforecast.methods import BottomUp, TopDown, MinTrace, ERM
+
+from hierarchicalforecast.utils import is_strictly_hierarchical
+from hierarchicalforecast.utils import HierarchicalPlot, CodeTimer
+
+from datasetsforecast.hierarchical import HierarchicalData, HierarchicalInfo
+```
+
+## 2. Preparing TourismL Dataset
+
+### 2.1 Read Hierarchical Dataset
+
+
+```python
+# ['Labour', 'Traffic', 'TourismSmall', 'TourismLarge', 'Wiki2']
+dataset = 'TourismSmall' # 'TourismLarge'
+verbose = True
+intervals_method = 'bootstrap'
+LEVEL = np.arange(0, 100, 2)
+```
+
+
+```python
+with CodeTimer('Read and Parse data ', verbose):
+ print(f'{dataset}')
+ if not os.path.exists('./data'):
+ os.makedirs('./data')
+
+ dataset_info = HierarchicalInfo[dataset]
+ Y_df, S_df, tags = HierarchicalData.load(directory=f'./data/{dataset}', group=dataset)
+ Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+
+ # Train/Test Splits
+ horizon = dataset_info.horizon
+ seasonality = dataset_info.seasonality
+ Y_test_df = Y_df.groupby('unique_id', as_index=False).tail(horizon)
+ Y_train_df = Y_df.drop(Y_test_df.index)
+ S_df = S_df.reset_index(names="unique_id")
+```
+
+``` text
+TourismSmall
+Code block 'Read and Parse data ' took: 0.00653 seconds
+```
+
+
+```python
+dataset_info.seasonality
+```
+
+``` text
+4
+```
+
+
+```python
+hplot = HierarchicalPlot(S=S_df, tags=tags)
+hplot.plot_summing_matrix()
+```
+
+
+
+
+```python
+Y_train_df
+```
+
+| | unique_id | ds | y |
+|------|----------------|------------|-------|
+| 0 | total | 1998-03-31 | 84503 |
+| 1 | total | 1998-06-30 | 65312 |
+| 2 | total | 1998-09-30 | 72753 |
+| 3 | total | 1998-12-31 | 70880 |
+| 4 | total | 1999-03-31 | 86893 |
+| ... | ... | ... | ... |
+| 3191 | nt-oth-noncity | 2003-12-31 | 132 |
+| 3192 | nt-oth-noncity | 2004-03-31 | 12 |
+| 3193 | nt-oth-noncity | 2004-06-30 | 40 |
+| 3194 | nt-oth-noncity | 2004-09-30 | 186 |
+| 3195 | nt-oth-noncity | 2004-12-31 | 144 |
+
+### 2.2 StatsForecast’s Base Predictions
+
+This cell computes the base predictions `Y_hat_df` for all the series in
+`Y_df` using StatsForecast’s `AutoARIMA`. Additionally we obtain
+insample predictions `Y_fitted_df` for the methods that require them.
+
+
+```python
+with CodeTimer('Fit/Predict Model ', verbose):
+ # Read to avoid unnecesary AutoARIMA computation
+ yhat_file = f'./data/{dataset}/Y_hat.csv'
+ yfitted_file = f'./data/{dataset}/Y_fitted.csv'
+
+ if os.path.exists(yhat_file):
+ Y_hat_df = pd.read_csv(yhat_file, parse_dates=['ds'])
+ Y_fitted_df = pd.read_csv(yfitted_file, parse_dates=['ds'])
+
+ else:
+ fcst = StatsForecast(
+ models=[AutoARIMA(season_length=seasonality)],
+ fallback_model=[Naive()],
+ freq=dataset_info.freq,
+ n_jobs=-1
+ )
+ Y_hat_df = fcst.forecast(df=Y_train_df, h=horizon, fitted=True, level=LEVEL)
+ Y_fitted_df = fcst.forecast_fitted_values()
+ Y_hat_df.to_csv(yhat_file, index=False)
+ Y_fitted_df.to_csv(yfitted_file, index=False)
+```
+
+## 3. Reconcile Predictions
+
+
+```python
+with CodeTimer('Reconcile Predictions ', verbose):
+ if is_strictly_hierarchical(S=S_df.drop(columns="unique_id").values.astype(np.float32), tags={key: S_df["unique_id"].isin(val).values.nonzero()[0] for key, val in tags.items()}):
+ reconcilers = [
+ BottomUp(),
+ TopDown(method='average_proportions'),
+ TopDown(method='proportion_averages'),
+ MinTrace(method='ols'),
+ MinTrace(method='wls_var'),
+ MinTrace(method='mint_shrink'),
+ ERM(method='closed'),
+ ]
+ else:
+ reconcilers = [
+ BottomUp(),
+ MinTrace(method='ols'),
+ MinTrace(method='wls_var'),
+ MinTrace(method='mint_shrink'),
+ ERM(method='closed'),
+ ]
+
+ hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+ Y_rec_df = hrec.bootstrap_reconcile(Y_hat_df=Y_hat_df,
+ Y_df=Y_fitted_df,
+ S_df=S_df, tags=tags,
+ level=LEVEL,
+ intervals_method=intervals_method,
+ num_samples=10,
+ num_seeds=10)
+
+ Y_rec_df = Y_rec_df.merge(Y_test_df, on=['unique_id', 'ds'], how="left")
+```
+
+``` text
+Code block 'Reconcile Predictions ' took: 7.49314 seconds
+```
+
+Qualitative evaluation, of parsed quantiles
+
+
+```python
+unique_id = "total"
+plot_df = Y_rec_df.query("unique_id == @unique_id").groupby(["unique_id", "ds"], as_index=False).mean()
+for col in hrec.level_names['AutoARIMA/BottomUp']:
+ plt.plot(plot_df["ds"], plot_df[col], color="orange")
+plt.plot(plot_df["ds"], plot_df["y"], label="True")
+plt.title(f"AutoARIMA/BottomUp - {unique_id}")
+plt.legend()
+```
+
+
+
+## 4. Evaluation
+
+
+```python
+from utilsforecast.losses import scaled_crps, msse
+from hierarchicalforecast.evaluation import evaluate
+from functools import partial
+```
+
+
+```python
+with CodeTimer('Evaluate Models CRPS and MSSE ', verbose):
+ metrics_seeds = []
+ for seed in Y_rec_df.seed.unique():
+ df_seed = Y_rec_df.query("seed == @seed")
+ metrics_seed = evaluate(df = df_seed,
+ tags = tags,
+ metrics = [scaled_crps,
+ partial(msse, seasonality=4)],
+ models= hrec.level_names.keys(),
+ level = LEVEL,
+ train_df = Y_train_df,
+ )
+ metrics_seed['seed'] = seed
+ metrics_seeds.append(metrics_seed)
+ metrics_seeds = pd.concat(metrics_seeds)
+
+ metrics_mean = metrics_seeds.groupby(["level", "metric"], as_index=False).mean()
+ metrics_std = metrics_seeds.groupby(["level", "metric"], as_index=False).std()
+
+ results = metrics_mean[hrec.level_names.keys()].round(3).astype(str) + "±" + metrics_std[hrec.level_names.keys()].round(4).astype(str)
+ results.insert(0, "metric", metrics_mean["metric"])
+ results.insert(0, "level", metrics_mean["level"])
+
+results.sort_values(by=["metric", "level"])
+```
+
+``` text
+Code block 'Evaluate Models CRPS and MSSE ' took: 4.25192 seconds
+```
+
+| | level | metric | AutoARIMA/BottomUp | AutoARIMA/TopDown_method-average_proportions | AutoARIMA/TopDown_method-proportion_averages | AutoARIMA/MinTrace_method-ols | AutoARIMA/MinTrace_method-wls_var | AutoARIMA/MinTrace_method-mint_shrink | AutoARIMA/ERM_method-closed_lambda_reg-0.01 |
+|----|----|----|----|----|----|----|----|----|----|
+| 0 | Country | msse | 1.777±0.0 | 2.488±0.0 | 2.488±0.0 | 2.752±0.0 | 2.569±0.0 | 2.775±0.0 | 3.427±0.0 |
+| 2 | Country/Purpose | msse | 1.726±0.0 | 3.181±0.0 | 3.169±0.0 | 2.184±0.0 | 1.876±0.0 | 1.96±0.0 | 3.067±0.0 |
+| 4 | Country/Purpose/State | msse | 0.881±0.0 | 1.657±0.0 | 1.652±0.0 | 0.98±0.0 | 0.857±0.0 | 0.867±0.0 | 1.559±0.0 |
+| 6 | Country/Purpose/State/CityNonCity | msse | 0.95±0.0 | 1.271±0.0 | 1.269±0.0 | 1.033±0.0 | 0.903±0.0 | 0.912±0.0 | 1.635±0.0 |
+| 8 | Overall | msse | 0.973±0.0 | 1.492±0.0 | 1.488±0.0 | 1.087±0.0 | 0.951±0.0 | 0.966±0.0 | 1.695±0.0 |
+| 1 | Country | scaled_crps | 0.043±0.0009 | 0.048±0.0006 | 0.048±0.0006 | 0.05±0.0006 | 0.051±0.0006 | 0.053±0.0006 | 0.054±0.0009 |
+| 3 | Country/Purpose | scaled_crps | 0.077±0.001 | 0.114±0.0003 | 0.112±0.0004 | 0.09±0.0013 | 0.087±0.0009 | 0.089±0.0009 | 0.106±0.0013 |
+| 5 | Country/Purpose/State | scaled_crps | 0.165±0.0009 | 0.249±0.0004 | 0.247±0.0004 | 0.18±0.0018 | 0.169±0.0009 | 0.169±0.0008 | 0.231±0.0021 |
+| 7 | Country/Purpose/State/CityNonCity | scaled_crps | 0.218±0.0013 | 0.289±0.0004 | 0.286±0.0004 | 0.228±0.0018 | 0.217±0.0013 | 0.218±0.0011 | 0.302±0.0033 |
+| 9 | Overall | scaled_crps | 0.193±0.0011 | 0.266±0.0004 | 0.263±0.0004 | 0.205±0.0017 | 0.194±0.0011 | 0.195±0.0009 | 0.268±0.0027 |
+
+## References
+
+- [Syama Sundar Rangapuram, Lucien D Werner, Konstantinos Benidis,
+ Pedro Mercado, Jan Gasthaus, Tim Januschowski. (2021). "End-to-End
+ Learning of Coherent Probabilistic Forecasts for Hierarchical Time
+ Series". Proceedings of the 38th International Conference on Machine
+ Learning
+ (ICML).](https://proceedings.mlr.press/v139/rangapuram21a.html)
+- [Kin G. Olivares, O. Nganba Meetei, Ruijun Ma, Rohan Reddy, Mengfei
+ Cao, Lee Dicker (2022). “Probabilistic Hierarchical Forecasting with
+ Deep Poisson Mixtures”. Submitted to the International Journal
+ Forecasting, Working paper available at
+ arxiv.](https://arxiv.org/pdf/2110.13179.pdf)
+
diff --git a/hierarchicalforecast/examples/tourismsmall.html.mdx b/hierarchicalforecast/examples/tourismsmall.html.mdx
new file mode 100644
index 00000000..ac8e10a6
--- /dev/null
+++ b/hierarchicalforecast/examples/tourismsmall.html.mdx
@@ -0,0 +1,217 @@
+---
+description: Minimal Example of Hierarchical Reconciliation
+output-file: tourismsmall.html
+title: Quick Start
+---
+
+
+Large collections of time series organized into structures at different
+aggregation levels often require their forecasts to follow their
+aggregation constraints, which poses the challenge of creating novel
+algorithms capable of coherent forecasts.
+
+The `HierarchicalForecast` package provides a wide collection of Python
+implementations of hierarchical forecasting algorithms that follow
+classic hierarchical reconciliation.
+
+In this notebook we will show how to use the `StatsForecast` library to
+produce base forecasts, and use `HierarchicalForecast` package to
+perform hierarchical reconciliation.
+
+You can run these experiments using CPU or GPU with Google Colab.
+
+
+
+## 1. Libraries
+
+
+```python
+!pip install hierarchicalforecast statsforecast datasetsforecast
+```
+
+## 2. Load Data
+
+In this example we will use the `TourismSmall` dataset. The following
+cell gets the time series for the different levels in the hierarchy, the
+summing matrix `S` which recovers the full dataset from the bottom level
+hierarchy and the indices of each hierarchy denoted by `tags`.
+
+
+```python
+import pandas as pd
+
+from datasetsforecast.hierarchical import HierarchicalData, HierarchicalInfo
+```
+
+
+```python
+group_name = 'TourismSmall'
+group = HierarchicalInfo.get_group(group_name)
+Y_df, S_df, tags = HierarchicalData.load('./data', group_name)
+S_df = S_df.reset_index(names="unique_id")
+Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+```
+
+
+```python
+S_df.iloc[:6, :6]
+```
+
+| | unique_id | nsw-hol-city | nsw-hol-noncity | vic-hol-city | vic-hol-noncity | qld-hol-city |
+|----|----|----|----|----|----|----|
+| 0 | total | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 |
+| 1 | hol | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 |
+| 2 | vfr | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 3 | bus | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 4 | oth | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 5 | nsw-hol | 1.0 | 1.0 | 0.0 | 0.0 | 0.0 |
+
+
+```python
+tags
+```
+
+``` text
+{'Country': array(['total'], dtype=object),
+ 'Country/Purpose': array(['hol', 'vfr', 'bus', 'oth'], dtype=object),
+ 'Country/Purpose/State': array(['nsw-hol', 'vic-hol', 'qld-hol', 'sa-hol', 'wa-hol', 'tas-hol',
+ 'nt-hol', 'nsw-vfr', 'vic-vfr', 'qld-vfr', 'sa-vfr', 'wa-vfr',
+ 'tas-vfr', 'nt-vfr', 'nsw-bus', 'vic-bus', 'qld-bus', 'sa-bus',
+ 'wa-bus', 'tas-bus', 'nt-bus', 'nsw-oth', 'vic-oth', 'qld-oth',
+ 'sa-oth', 'wa-oth', 'tas-oth', 'nt-oth'], dtype=object),
+ 'Country/Purpose/State/CityNonCity': array(['nsw-hol-city', 'nsw-hol-noncity', 'vic-hol-city',
+ 'vic-hol-noncity', 'qld-hol-city', 'qld-hol-noncity',
+ 'sa-hol-city', 'sa-hol-noncity', 'wa-hol-city', 'wa-hol-noncity',
+ 'tas-hol-city', 'tas-hol-noncity', 'nt-hol-city', 'nt-hol-noncity',
+ 'nsw-vfr-city', 'nsw-vfr-noncity', 'vic-vfr-city',
+ 'vic-vfr-noncity', 'qld-vfr-city', 'qld-vfr-noncity',
+ 'sa-vfr-city', 'sa-vfr-noncity', 'wa-vfr-city', 'wa-vfr-noncity',
+ 'tas-vfr-city', 'tas-vfr-noncity', 'nt-vfr-city', 'nt-vfr-noncity',
+ 'nsw-bus-city', 'nsw-bus-noncity', 'vic-bus-city',
+ 'vic-bus-noncity', 'qld-bus-city', 'qld-bus-noncity',
+ 'sa-bus-city', 'sa-bus-noncity', 'wa-bus-city', 'wa-bus-noncity',
+ 'tas-bus-city', 'tas-bus-noncity', 'nt-bus-city', 'nt-bus-noncity',
+ 'nsw-oth-city', 'nsw-oth-noncity', 'vic-oth-city',
+ 'vic-oth-noncity', 'qld-oth-city', 'qld-oth-noncity',
+ 'sa-oth-city', 'sa-oth-noncity', 'wa-oth-city', 'wa-oth-noncity',
+ 'tas-oth-city', 'tas-oth-noncity', 'nt-oth-city', 'nt-oth-noncity'],
+ dtype=object)}
+```
+
+We split the dataframe in train/test splits.
+
+
+```python
+Y_test_df = Y_df.groupby('unique_id').tail(group.horizon)
+Y_train_df = Y_df.drop(Y_test_df.index)
+```
+
+## 3. Base forecasts
+
+The following cell computes the *base forecast* for each time series
+using the `auto_arima` and `naive` models. Observe that `Y_hat_df`
+contains the forecasts but they are not coherent.
+
+
+```python
+from statsforecast.core import StatsForecast
+from statsforecast.models import AutoARIMA, Naive
+```
+
+
+```python
+fcst = StatsForecast(
+ models=[AutoARIMA(season_length=group.seasonality), Naive()],
+ freq="QE",
+ n_jobs=-1
+)
+Y_hat_df = fcst.forecast(df=Y_train_df, h=group.horizon)
+```
+
+## 4. Hierarchical reconciliation
+
+The following cell makes the previous forecasts coherent using the
+[`HierarchicalReconciliation`](https://Nixtla.github.io/hierarchicalforecast/src/core.html#hierarchicalreconciliation)
+class. The used methods to make the forecasts coherent are:
+
+- [`BottomUp`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#bottomup):
+ The reconciliation of the method is a simple addition to the upper
+ levels.
+- [`TopDown`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#topdown):
+ The second method constrains the base-level predictions to the
+ top-most aggregate-level serie and then distributes it to the
+ disaggregate series through the use of proportions.
+- [`MiddleOut`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#middleout):
+ Anchors the base predictions in a middle level.
+
+
+```python
+from hierarchicalforecast.core import HierarchicalReconciliation
+from hierarchicalforecast.methods import BottomUp, TopDown, MiddleOut
+```
+
+
+```python
+reconcilers = [
+ BottomUp(),
+ TopDown(method='forecast_proportions'),
+ TopDown(method='proportion_averages'),
+ MiddleOut(middle_level="Country/Purpose/State", top_down_method="proportion_averages"),
+]
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+Y_rec_df = hrec.reconcile(Y_hat_df=Y_hat_df, Y_df=Y_train_df, S_df=S_df, tags=tags)
+```
+
+## 5. Evaluation
+
+The `HierarchicalForecast` package includes the
+[`evaluate`](https://Nixtla.github.io/hierarchicalforecast/src/evaluation.html#evaluate)
+function to evaluate the different hierarchies and we can use
+utilsforecast to compute the mean absolute error relative to a baseline
+model.
+
+
+```python
+from hierarchicalforecast.evaluation import evaluate
+from utilsforecast.losses import mse
+```
+
+
+```python
+df = Y_rec_df.merge(Y_test_df, on=['unique_id', 'ds'])
+evaluation = evaluate(df = df,
+ tags = tags,
+ train_df = Y_train_df,
+ metrics = [mse],
+ benchmark="Naive")
+
+evaluation.set_index(["level", "metric"]).filter(like="ARIMA", axis=1)
+```
+
+| | | AutoARIMA | AutoARIMA/BottomUp | AutoARIMA/TopDown_method-forecast_proportions | AutoARIMA/TopDown_method-proportion_averages | AutoARIMA/MiddleOut_middle_level-Country/Purpose/State_top_down_method-proportion_averages |
+|----|----|----|----|----|----|----|
+| level | metric | | | | | |
+| Country | mse-scaled | 0.317897 | 0.367078 | 0.317897 | 0.317897 | 0.305053 |
+| Country/Purpose | mse-scaled | 0.318950 | 0.233606 | 0.262216 | 0.320225 | 0.196062 |
+| Country/Purpose/State | mse-scaled | 0.268057 | 0.281189 | 0.320349 | 0.511356 | 0.268057 |
+| Country/Purpose/State/CityNonCity | mse-scaled | 0.292136 | 0.292136 | 0.323261 | 0.509784 | 0.280599 |
+| Overall | mse-scaled | 0.308942 | 0.295690 | 0.297072 | 0.364775 | 0.255038 |
+
+### References
+
+- [Orcutt, G.H., Watts, H.W., & Edwards, J.B.(1968). Data aggregation
+ and information loss. The American Economic Review, 58 ,
+ 773(787)](http://www.jstor.org/stable/1815532).
+- [Disaggregation methods to expedite product line forecasting.
+ Journal of Forecasting, 9 , 233–254.
+ doi:10.1002/for.3980090304](https://onlinelibrary.wiley.com/doi/abs/10.1002/for.3980090304).
+- [An investigation of aggregate variable time series forecast
+ strategies with specific subaggregate time series statistical
+ correlation. Computers and Operations Research, 26 , 1133–1149.
+ doi:10.1016/S0305-0548(99)00017-9](https://doi.org/10.1016/S0305-0548(99)00017-9).
+- [Hyndman, R.J., & Athanasopoulos, G. (2021). “Forecasting:
+ principles and practice, 3rd edition: Chapter 11: Forecasting
+ hierarchical and grouped series.”. OTexts: Melbourne, Australia.
+ OTexts.com/fpp3 Accessed on July
+ 2022.](https://otexts.com/fpp3/hierarchical.html)
+
diff --git a/hierarchicalforecast/examples/tourismsmallpolars.html.mdx b/hierarchicalforecast/examples/tourismsmallpolars.html.mdx
new file mode 100644
index 00000000..9164aeff
--- /dev/null
+++ b/hierarchicalforecast/examples/tourismsmallpolars.html.mdx
@@ -0,0 +1,221 @@
+---
+description: Minimal Example of Hierarchical Reconciliation using Polars
+output-file: tourismsmallpolars.html
+title: Quick Start (Polars)
+---
+
+
+Large collections of time series organized into structures at different
+aggregation levels often require their forecasts to follow their
+aggregation constraints, which poses the challenge of creating novel
+algorithms capable of coherent forecasts.
+
+The `HierarchicalForecast` package provides a wide collection of Python
+implementations of hierarchical forecasting algorithms that follow
+classic hierarchical reconciliation.
+
+In this notebook we will show how to use the `StatsForecast` library to
+produce base forecasts, and use `HierarchicalForecast` package to
+perform hierarchical reconciliation.
+
+You can run these experiments using CPU or GPU with Google Colab.
+
+
+
+## 1. Libraries
+
+
+```python
+!pip install hierarchicalforecast statsforecast datasetsforecast
+```
+
+## 2. Load Data
+
+In this example we will use the `TourismSmall` dataset. The following
+cell gets the time series for the different levels in the hierarchy, the
+summing matrix `S` which recovers the full dataset from the bottom level
+hierarchy and the indices of each hierarchy denoted by `tags`.
+
+
+```python
+import numpy as np
+import polars as pl
+
+from datasetsforecast.hierarchical import HierarchicalData, HierarchicalInfo
+```
+
+
+```python
+group_name = 'TourismSmall'
+group = HierarchicalInfo.get_group(group_name)
+Y_df, S_df, tags = HierarchicalData.load('./data', group_name)
+
+Y_df = pl.from_pandas(Y_df)
+S_df = pl.from_pandas(S_df.reset_index(names="unique_id"))
+Y_df = Y_df.with_columns(pl.col('ds').cast(pl.Date))
+```
+
+
+```python
+S_df[:6, :6]
+```
+
+| unique_id | nsw-hol-city | nsw-hol-noncity | vic-hol-city | vic-hol-noncity | qld-hol-city |
+|----|----|----|----|----|----|
+| str | f64 | f64 | f64 | f64 | f64 |
+| "total" | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 |
+| "hol" | 1.0 | 1.0 | 1.0 | 1.0 | 1.0 |
+| "vfr" | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
+| "bus" | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
+| "oth" | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
+| "nsw-hol" | 1.0 | 1.0 | 0.0 | 0.0 | 0.0 |
+
+
+```python
+tags
+```
+
+``` text
+{'Country': array(['total'], dtype=object),
+ 'Country/Purpose': array(['hol', 'vfr', 'bus', 'oth'], dtype=object),
+ 'Country/Purpose/State': array(['nsw-hol', 'vic-hol', 'qld-hol', 'sa-hol', 'wa-hol', 'tas-hol',
+ 'nt-hol', 'nsw-vfr', 'vic-vfr', 'qld-vfr', 'sa-vfr', 'wa-vfr',
+ 'tas-vfr', 'nt-vfr', 'nsw-bus', 'vic-bus', 'qld-bus', 'sa-bus',
+ 'wa-bus', 'tas-bus', 'nt-bus', 'nsw-oth', 'vic-oth', 'qld-oth',
+ 'sa-oth', 'wa-oth', 'tas-oth', 'nt-oth'], dtype=object),
+ 'Country/Purpose/State/CityNonCity': array(['nsw-hol-city', 'nsw-hol-noncity', 'vic-hol-city',
+ 'vic-hol-noncity', 'qld-hol-city', 'qld-hol-noncity',
+ 'sa-hol-city', 'sa-hol-noncity', 'wa-hol-city', 'wa-hol-noncity',
+ 'tas-hol-city', 'tas-hol-noncity', 'nt-hol-city', 'nt-hol-noncity',
+ 'nsw-vfr-city', 'nsw-vfr-noncity', 'vic-vfr-city',
+ 'vic-vfr-noncity', 'qld-vfr-city', 'qld-vfr-noncity',
+ 'sa-vfr-city', 'sa-vfr-noncity', 'wa-vfr-city', 'wa-vfr-noncity',
+ 'tas-vfr-city', 'tas-vfr-noncity', 'nt-vfr-city', 'nt-vfr-noncity',
+ 'nsw-bus-city', 'nsw-bus-noncity', 'vic-bus-city',
+ 'vic-bus-noncity', 'qld-bus-city', 'qld-bus-noncity',
+ 'sa-bus-city', 'sa-bus-noncity', 'wa-bus-city', 'wa-bus-noncity',
+ 'tas-bus-city', 'tas-bus-noncity', 'nt-bus-city', 'nt-bus-noncity',
+ 'nsw-oth-city', 'nsw-oth-noncity', 'vic-oth-city',
+ 'vic-oth-noncity', 'qld-oth-city', 'qld-oth-noncity',
+ 'sa-oth-city', 'sa-oth-noncity', 'wa-oth-city', 'wa-oth-noncity',
+ 'tas-oth-city', 'tas-oth-noncity', 'nt-oth-city', 'nt-oth-noncity'],
+ dtype=object)}
+```
+
+We split the dataframe in train/test splits.
+
+
+```python
+Y_test_df = Y_df.group_by('unique_id').tail(group.horizon)
+Y_train_df = Y_df.filter(pl.col('ds') < Y_test_df['ds'].min())
+```
+
+## 3. Base forecasts
+
+The following cell computes the *base forecast* for each time series
+using the `auto_arima` and `naive` models. Observe that `Y_hat_df`
+contains the forecasts but they are not coherent.
+
+
+```python
+from statsforecast.core import StatsForecast
+from statsforecast.models import AutoARIMA, Naive
+```
+
+
+```python
+fcst = StatsForecast(
+ models=[AutoARIMA(season_length=group.seasonality), Naive()],
+ freq="1q",
+ n_jobs=-1
+)
+Y_hat_df = fcst.forecast(df=Y_train_df, h=group.horizon)
+```
+
+## 4. Hierarchical reconciliation
+
+The following cell makes the previous forecasts coherent using the
+[`HierarchicalReconciliation`](https://Nixtla.github.io/hierarchicalforecast/src/core.html#hierarchicalreconciliation)
+class. The used methods to make the forecasts coherent are:
+
+- [`BottomUp`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#bottomup):
+ The reconciliation of the method is a simple addition to the upper
+ levels.
+- [`TopDown`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#topdown):
+ The second method constrains the base-level predictions to the
+ top-most aggregate-level serie and then distributes it to the
+ disaggregate series through the use of proportions.
+- [`MiddleOut`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#middleout):
+ Anchors the base predictions in a middle level.
+
+
+```python
+from hierarchicalforecast.core import HierarchicalReconciliation
+from hierarchicalforecast.methods import BottomUp, TopDown, MiddleOut
+```
+
+
+```python
+reconcilers = [
+ BottomUp(),
+ TopDown(method='forecast_proportions'),
+ MiddleOut(middle_level='Country/Purpose/State',
+ top_down_method='forecast_proportions')
+]
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+Y_rec_df = hrec.reconcile(Y_hat_df=Y_hat_df, Y_df=Y_train_df, S_df=S_df, tags=tags)
+```
+
+## 5. Evaluation
+
+The `HierarchicalForecast` package includes the
+[`evaluate`](https://Nixtla.github.io/hierarchicalforecast/src/evaluation.html#evaluate)
+function to evaluate the different hierarchies and we can use
+utilsforecast to compute the mean absolute error relative to a baseline
+model.
+
+
+```python
+from hierarchicalforecast.evaluation import evaluate
+from utilsforecast.losses import mse
+```
+
+
+```python
+df = Y_rec_df.join(Y_test_df, on=['unique_id', 'ds'])
+evaluation = evaluate(df = df,
+ tags = tags,
+ train_df = Y_train_df,
+ metrics = [mse],
+ benchmark="Naive")
+
+evaluation[["level", "metric", "AutoARIMA", "AutoARIMA/BottomUp", "AutoARIMA/TopDown_method-forecast_proportions"]]
+```
+
+| level | metric | AutoARIMA | AutoARIMA/BottomUp | AutoARIMA/TopDown_method-forecast_proportions |
+|----|----|----|----|----|
+| str | str | f64 | f64 | f64 |
+| "Country" | "mse-scaled" | 0.317897 | 0.226999 | 0.317897 |
+| "Country/Purpose" | "mse-scaled" | 0.323207 | 0.199359 | 0.251368 |
+| "Country/Purpose/State" | "mse-scaled" | 0.266118 | 0.305711 | 0.308241 |
+| "Country/Purpose/State/CityNonC… | "mse-scaled" | 0.305173 | 0.305173 | 0.305913 |
+| "Overall" | "mse-scaled" | 0.311707 | 0.234934 | 0.289406 |
+
+### References
+
+- [Orcutt, G.H., Watts, H.W., & Edwards, J.B.(1968). Data aggregation
+ and information loss. The American Economic Review, 58 ,
+ 773(787)](http://www.jstor.org/stable/1815532).
+- [Disaggregation methods to expedite product line forecasting.
+ Journal of Forecasting, 9 , 233–254.
+ doi:10.1002/for.3980090304](https://onlinelibrary.wiley.com/doi/abs/10.1002/for.3980090304).
+- [An investigation of aggregate variable time series forecast
+ strategies with specific subaggregate time series statistical
+ correlation. Computers and Operations Research, 26 , 1133–1149.
+ doi:10.1016/S0305-0548(99)00017-9](https://doi.org/10.1016/S0305-0548(99)00017-9).
+- [Hyndman, R.J., & Athanasopoulos, G. (2021). “Forecasting:
+ principles and practice, 3rd edition: Chapter 11: Forecasting
+ hierarchical and grouped series.”. OTexts: Melbourne, Australia.
+ OTexts.com/fpp3 Accessed on July
+ 2022.](https://otexts.com/fpp3/hierarchical.html)
+
diff --git a/hierarchicalforecast/favicon.svg b/hierarchicalforecast/favicon.svg
new file mode 100644
index 00000000..e5f33342
--- /dev/null
+++ b/hierarchicalforecast/favicon.svg
@@ -0,0 +1,5 @@
+
diff --git a/hierarchicalforecast/index.html.mdx b/hierarchicalforecast/index.html.mdx
new file mode 100644
index 00000000..e30708ac
--- /dev/null
+++ b/hierarchicalforecast/index.html.mdx
@@ -0,0 +1,206 @@
+---
+output-file: index.html
+title: Hierarchical Forecast 👑
+---
+
+
+Large collections of time series organized into structures at different
+aggregation levels often require their forecasts to follow their
+aggregation constraints, which poses the challenge of creating novel
+algorithms capable of coherent forecasts.
+
+**HierarchicalForecast** offers a collection of cross-sectional and
+temporal reconciliation methods, including
+[`BottomUp`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#bottomup),
+[`TopDown`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#topdown),
+[`MiddleOut`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#middleout),
+[`MinTrace`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#mintrace)
+and
+[`ERM`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#erm),
+as well as probabilistic coherent predictions including
+[`Normality`](https://Nixtla.github.io/hierarchicalforecast/src/probabilistic_methods.html#normality),
+[`Bootstrap`](https://Nixtla.github.io/hierarchicalforecast/src/probabilistic_methods.html#bootstrap),
+and
+[`PERMBU`](https://Nixtla.github.io/hierarchicalforecast/src/probabilistic_methods.html#permbu).
+
+## 🎊 Features
+
+- Classic reconciliation methods:
+ - `BottomUp`: Simple addition to the upper levels.
+ - `TopDown`: Distributes the top levels forecasts trough the
+ hierarchies.
+- Alternative reconciliation methods:
+ - `MiddleOut`: It anchors the base predictions in a middle level.
+ The levels above the base predictions use the bottom-up
+ approach, while the levels below use a top-down.
+ - `MinTrace`: Minimizes the total forecast variance of the space
+ of coherent forecasts, with the Minimum Trace reconciliation.
+ - `ERM`: Optimizes the reconciliation matrix minimizing an L1
+ regularized objective.
+- Probabilistic coherent methods:
+ - `Normality`: Uses MinTrace variance-covariance closed form
+ matrix under a normality assumption.
+ - `Bootstrap`: Generates distribution of hierarchically reconciled
+ predictions using Gamakumara’s bootstrap approach.
+ - `PERMBU`: Reconciles independent sample predictions by
+ reinjecting multivariate dependence with estimated rank
+ permutation copulas, and performing a Bottom-Up aggregation.
+- Temporal reconciliation methods:
+ - All reconciliation methods (except for the insample methods) are
+ available to use with temporal hierarchies too.
+
+Missing something? Please open an issue here or write us in
+[](https://join.slack.com/t/nixtlaworkspace/shared_invite/zt-135dssye9-fWTzMpv2WBthq8NK0Yvu6A)
+
+## 📖 Why?
+
+**Short**: We want to contribute to the ML field by providing reliable
+baselines and benchmarks for hierarchical forecasting task in industry
+and academia. Here’s the complete
+[paper](https://arxiv.org/abs/2207.03517).
+
+**Verbose**: `HierarchicalForecast` integrates publicly available
+processed datasets, evaluation metrics, and a curated set of statistical
+baselines. In this library we provide usage examples and references to
+extensive experiments where we showcase the baseline’s use and evaluate
+the accuracy of their predictions. With this work, we hope to contribute
+to Machine Learning forecasting by bridging the gap to statistical and
+econometric modeling, as well as providing tools for the development of
+novel hierarchical forecasting algorithms rooted in a thorough
+comparison of these well-established models. We intend to continue
+maintaining and increasing the repository, promoting collaboration
+across the forecasting community.
+
+## 💻 Installation
+
+### PyPI
+
+We recommend using `uv` as Python package manager, for which you can
+find installation instructions
+[here](https://docs.astral.sh/uv/getting-started/installation/).
+
+You can then install the *released version* of `HierachicalForecast`:
+
+
+```python
+uv pip install hierarchicalforecast
+```
+
+Alternatively, you can directly install from the [Python package
+index](https://pypi.org) with:
+
+
+```python
+pip install hierarchicalforecast
+```
+
+(Installing inside a python virtualenvironment is recommended.)
+
+### Conda
+
+Also you can install the *released version* of `HierarchicalForecast`
+from [conda](https://anaconda.org) with:
+
+
+```python
+conda install -c conda-forge hierarchicalforecast
+```
+
+(Installing inside a python virtualenvironment or a conda environment is
+recommended.)
+
+### Dev Mode
+
+If you want to make some modifications to the code and see the effects
+in real time (without reinstalling), follow the steps
+[here](https://github.com/Nixtla/hierarchicalforecast/blob/main/CONTRIBUTING.md).
+
+## 🧬 How to use
+
+The following example needs `statsforecast` and `datasetsforecast` as
+additional packages. If not installed, install it via your preferred
+method, e.g. `pip install statsforecast datasetsforecast`. The
+`datasetsforecast` library allows us to download hierarhical datasets
+and we will use `statsforecast` to compute base forecasts to be
+reconciled.
+
+You can open this example in Colab [](https://colab.research.google.com/github/nixtla/hierarchicalforecast/blob/main/nbs/examples/TourismSmall.ipynb)
+
+
+```python
+import pandas as pd
+
+#obtain hierarchical dataset
+from datasetsforecast.hierarchical import HierarchicalData
+
+# compute base forecast no coherent
+from statsforecast.core import StatsForecast
+from statsforecast.models import AutoARIMA, Naive
+
+#obtain hierarchical reconciliation methods and evaluation
+from hierarchicalforecast.core import HierarchicalReconciliation
+from hierarchicalforecast.evaluation import evaluate
+from hierarchicalforecast.methods import BottomUp, TopDown, MiddleOut
+from utilsforecast.losses import mse
+
+# Load TourismSmall dataset
+Y_df, S_df, tags = HierarchicalData.load('./data', 'TourismSmall')
+Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+S_df = S_df.reset_index(names="unique_id")
+
+#split train/test sets
+Y_test_df = Y_df.groupby('unique_id').tail(4)
+Y_train_df = Y_df.drop(Y_test_df.index)
+
+# Compute base auto-ARIMA predictions
+fcst = StatsForecast(models=[AutoARIMA(season_length=4), Naive()],
+ freq='QE', n_jobs=-1)
+Y_hat_df = fcst.forecast(df=Y_train_df, h=4)
+
+# Reconcile the base predictions
+reconcilers = [
+ BottomUp(),
+ TopDown(method='forecast_proportions'),
+ MiddleOut(middle_level='Country/Purpose/State',
+ top_down_method='forecast_proportions')
+]
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+Y_rec_df = hrec.reconcile(Y_hat_df=Y_hat_df, Y_df=Y_train_df,
+ S_df=S_df, tags=tags)
+```
+
+### Evaluation
+
+
+```python
+df = Y_rec_df.merge(Y_test_df, on=['unique_id', 'ds'], how='left')
+
+evaluate(df=df, metrics=[mse],
+ tags=tags, benchmark='Naive')
+```
+
+## How to cite
+
+Here’s the complete [paper](https://arxiv.org/abs/2207.03517).
+
+
+```bibtex
+@article{olivares2022hierarchicalforecast,
+ author = {Kin G. Olivares and
+ Federico Garza and
+ David Luo and
+ Cristian Challú and
+ Max Mergenthaler and
+ Souhaib Ben Taieb and
+ Shanika L. Wickramasuriya and
+ Artur Dubrawski},
+ title = {{HierarchicalForecast}: A Reference Framework for Hierarchical Forecasting in Python},
+ journal = {Work in progress paper, submitted to Journal of Machine Learning Research.},
+ volume = {abs/2207.03517},
+ year = {2022},
+ url = {https://arxiv.org/abs/2207.03517},
+ archivePrefix = {arXiv}
+}
+```
+
diff --git a/hierarchicalforecast/light.png b/hierarchicalforecast/light.png
new file mode 100644
index 00000000..bbb99b54
Binary files /dev/null and b/hierarchicalforecast/light.png differ
diff --git a/hierarchicalforecast/mint.json b/hierarchicalforecast/mint.json
new file mode 100644
index 00000000..7dbd2953
--- /dev/null
+++ b/hierarchicalforecast/mint.json
@@ -0,0 +1,83 @@
+{
+ "$schema": "https://mintlify.com/schema.json",
+ "name": "Nixtla",
+ "logo": {
+ "light": "/light.png",
+ "dark": "/dark.png"
+ },
+ "favicon": "/favicon.svg",
+ "colors": {
+ "primary": "#0E0E0E",
+ "light": "#FAFAFA",
+ "dark": "#0E0E0E",
+ "anchors": {
+ "from": "#2AD0CA",
+ "to": "#0E00F8"
+ }
+ },
+ "topbarCtaButton": {
+ "type": "github",
+ "url": "https://github.com/Nixtla/hierarchicalforecast"
+ },
+ "topAnchor": {
+ "name": "HierarchicalForecast",
+ "icon": "crown"
+ },
+ "navigation": [
+ {
+ "group": "",
+ "pages": ["index.html"]
+ },
+ {
+ "group": "Getting Started",
+ "pages": [
+ "examples/installation.html",
+ "examples/tourismsmall.html",
+ "examples/tourismsmallpolars.html",
+ "examples/introduction.html"
+ ]
+ },
+ {
+ "group": "Tutorials",
+ "pages": [
+ {
+ "group": "Point Reconciliation",
+ "pages": [
+ "examples/australiandomestictourism.html",
+ "examples/australianprisonpopulation.html",
+ "examples/nonnegativereconciliation.html"
+ ]
+ },
+ {
+ "group": "Probabilistic Reconciliation",
+ "pages": [
+ "examples/australiandomestictourism-intervals.html",
+ "examples/australiandomestictourism-bootstraped-intervals.html",
+ "examples/australiandomestictourism-permbu-intervals.html",
+ "examples/tourismlarge-evaluation.html"
+ ]
+ },
+ {
+ "group": "Temporal Reconciliation",
+ "pages": [
+ "examples/australiandomestictourismtemporal.html",
+ "examples/australiandomestictourismcrosstemporal.html",
+ "examples/m3withthief.html",
+ "examples/localglobalaggregation.html"
+ ]
+ },
+ "examples/mlframeworksexample.html"
+ ]
+ },
+ {
+ "group": "API Reference",
+ "pages": [
+ "src/core.html",
+ "src/methods.html",
+ "src/probabilistic_methods.html",
+ "src/evaluation.html",
+ "src/utils.html"
+ ]
+ }
+ ]
+}
diff --git a/hierarchicalforecast/src/core.html.mdx b/hierarchicalforecast/src/core.html.mdx
new file mode 100644
index 00000000..9c28c05c
--- /dev/null
+++ b/hierarchicalforecast/src/core.html.mdx
@@ -0,0 +1,257 @@
+---
+output-file: core.html
+title: Core
+---
+
+
+HierarchicalForecast contains pure Python implementations of
+hierarchical reconciliation methods as well as a
+`core.HierarchicalReconciliation` wrapper class that enables easy
+interaction with these methods through pandas DataFrames containing the
+hierarchical time series and the base predictions.
+
+The `core.HierarchicalReconciliation` reconciliation class operates with
+the hierarchical time series pd.DataFrame `Y_df`, the base predictions
+pd.DataFrame `Y_hat_df`, the aggregation constraints matrix `S`. For
+more information on the creation of aggregation constraints matrix see
+the utils [aggregation
+method](https://nixtlaverse.nixtla.io/hierarchicalforecast/src/utils.html#aggregate).
+
+# HierarchicalReconciliation
+
+------------------------------------------------------------------------
+
+source
+
+### HierarchicalReconciliation
+
+> ``` text
+> HierarchicalReconciliation
+> (reconcilers:list[hierarchicalforecast.method
+> s.HReconciler])
+> ```
+
+\*Hierarchical Reconciliation Class.
+
+The `core.HierarchicalReconciliation` class allows you to efficiently
+fit multiple HierarchicaForecast methods for a collection of time series
+and base predictions stored in pandas DataFrames. The `Y_df` dataframe
+identifies series and datestamps with the unique_id and ds columns while
+the y column denotes the target time series variable. The `Y_h`
+dataframe stores the base predictions, example
+([AutoARIMA](https://Nixtla.github.io/statsforecast/src/core/models.html#autoarima),
+[ETS](https://Nixtla.github.io/statsforecast/src/core/models.html#autoets),
+etc.).
+
+**Parameters:** `reconcilers`: A list of instantiated classes of the
+[reconciliation
+methods](https://nixtla.github.io/hierarchicalforecast/src/methods.html)
+module .
+
+**References:** [Rob J. Hyndman and George Athanasopoulos (2018).
+“Forecasting principles and practice, Hierarchical and Grouped
+Series”.](https://otexts.com/fpp3/hierarchical.html)\*
+
+------------------------------------------------------------------------
+
+source
+
+### reconcile
+
+> ``` text
+> reconcile (Y_hat_df:Union[ForwardRef('DataFrame[Any]'),ForwardRef('LazyFr
+> ame[Any]')], tags:dict[str,numpy.ndarray], S_df:Union[ForwardR
+> ef('DataFrame[Any]'),ForwardRef('LazyFrame[Any]')]=None, Y_df:
+> Union[ForwardRef('DataFrame[Any]'),ForwardRef('LazyFrame[Any]'
+> ),NoneType]=None, level:Optional[list[int]]=None,
+> intervals_method:str='normality', num_samples:int=-1,
+> seed:int=0, is_balanced:bool=False, id_col:str='unique_id',
+> time_col:str='ds', target_col:str='y',
+> id_time_col:str='temporal_id', temporal:bool=False, S:Union[Fo
+> rwardRef('DataFrame[Any]'),ForwardRef('LazyFrame[Any]')]=None)
+> ```
+
+\*Hierarchical Reconciliation Method.
+
+The `reconcile` method is analogous to SKLearn `fit_predict` method, it
+applies different reconciliation techniques instantiated in the
+`reconcilers` list.
+
+Most reconciliation methods can be described by the following convenient
+linear algebra notation:
+
+$$\tilde{\mathbf{y}}_{[a,b],\tau} = \mathbf{S}_{[a,b][b]} \mathbf{P}_{[b][a,b]} \hat{\mathbf{y}}_{[a,b],\tau}$$
+
+where $a, b$ represent the aggregate and bottom levels,
+$\mathbf{S}_{[a,b][b]}$ contains the hierarchical aggregation
+constraints, and $\mathbf{P}_{[b][a,b]}$ varies across reconciliation
+methods. The reconciled predictions are
+$\tilde{\mathbf{y}}_{[a,b],\tau}$, and the base predictions
+$\hat{\mathbf{y}}_{[a,b],\tau}$.
+
+**Parameters:** `Y_hat_df`: DataFrame, base forecasts with columns
+\[‘unique_id’, ‘ds’\] and models to reconcile. `tags`: Each key is a
+level and its value contains tags associated to that level. `S_df`:
+DataFrame with summing matrix of size `(base, bottom)`, see [aggregate
+method](https://nixtlaverse.nixtla.io/hierarchicalforecast/src/utils.html#aggregate).
+`Y_df`: DataFrame, training set of base time series with columns
+`['unique_id', 'ds', 'y']`. If a class of `self.reconciles` receives
+`y_hat_insample`, `Y_df` must include them as columns. `level`:
+positive float list \[0,100), confidence levels for prediction
+intervals. `intervals_method`: str, method used to calculate
+prediction intervals, one of `normality`, `bootstrap`, `permbu`.
+`num_samples`: int=-1, if positive return that many probabilistic
+coherent samples. `seed`: int=0, random seed for numpy generator’s
+replicability. `is_balanced`: bool=False, wether `Y_df` is balanced,
+set it to True to speed things up if `Y_df` is balanced. `id_col` :
+str=‘unique_id’, column that identifies each serie. `time_col` :
+str=‘ds’, column that identifies each timestep, its values can be
+timestamps or integers. `target_col` : str=‘y’, column that contains
+the target.
+
+**Returns:** `Y_tilde_df`: DataFrame, with reconciled predictions.\*
+
+| | **Type** | **Default** | **Details** |
+|----|----|----|----|
+| Y_hat_df | Union | | |
+| tags | dict | | |
+| S_df | Union | None | |
+| Y_df | Union | None | |
+| level | Optional | None | |
+| intervals_method | str | normality | |
+| num_samples | int | -1 | |
+| seed | int | 0 | |
+| is_balanced | bool | False | |
+| id_col | str | unique_id | |
+| time_col | str | ds | |
+| target_col | str | y | |
+| id_time_col | str | temporal_id | |
+| temporal | bool | False | |
+| S | Union | None | For compatibility with the old API, S_df is now S |
+| **Returns** | **FrameT** | | |
+
+------------------------------------------------------------------------
+
+source
+
+### bootstrap_reconcile
+
+> ``` text
+> bootstrap_reconcile (Y_hat_df:Union[ForwardRef('DataFrame[Any]'),ForwardR
+> ef('LazyFrame[Any]')], S_df:Union[ForwardRef('DataFr
+> ame[Any]'),ForwardRef('LazyFrame[Any]')],
+> tags:dict[str,numpy.ndarray], Y_df:Union[ForwardRef(
+> 'DataFrame[Any]'),ForwardRef('LazyFrame[Any]'),NoneT
+> ype]=None, level:Optional[list[int]]=None,
+> intervals_method:str='normality',
+> num_samples:int=-1, num_seeds:int=1,
+> id_col:str='unique_id', time_col:str='ds',
+> target_col:str='y')
+> ```
+
+\*Bootstraped Hierarchical Reconciliation Method.
+
+Applies N times, based on different random seeds, the `reconcile` method
+for the different reconciliation techniques instantiated in the
+`reconcilers` list.
+
+**Parameters:** `Y_hat_df`: DataFrame, base forecasts with columns
+\[‘unique_id’, ‘ds’\] and models to reconcile. `S_df`: DataFrame
+with summing matrix of size `(base, bottom)`, see [aggregate
+method](https://nixtlaverse.nixtla.io/hierarchicalforecast/src/utils.html#aggregate).
+`tags`: Each key is a level and its value contains tags associated to
+that level. `Y_df`: DataFrame, training set of base time series with
+columns `['unique_id', 'ds', 'y']`. If a class of `self.reconciles`
+receives `y_hat_insample`, `Y_df` must include them as columns.
+`level`: positive float list \[0,100), confidence levels for prediction
+intervals. `intervals_method`: str, method used to calculate
+prediction intervals, one of `normality`, `bootstrap`, `permbu`.
+`num_samples`: int=-1, if positive return that many probabilistic
+coherent samples. `num_seeds`: int=1, random seed for numpy generator’s
+replicability. `id_col` : str=‘unique_id’, column that identifies
+each serie. `time_col` : str=‘ds’, column that identifies each
+timestep, its values can be timestamps or integers. `target_col` :
+str=‘y’, column that contains the target.
+
+**Returns:** `Y_bootstrap_df`: DataFrame, with bootstraped
+reconciled predictions.\*
+
+# Example
+
+
+```python
+import pandas as pd
+
+from hierarchicalforecast.core import HierarchicalReconciliation
+from hierarchicalforecast.methods import BottomUp, MinTrace
+from hierarchicalforecast.utils import aggregate
+from hierarchicalforecast.evaluation import evaluate
+from statsforecast.core import StatsForecast
+from statsforecast.models import AutoETS
+from utilsforecast.losses import mase, rmse
+from functools import partial
+
+# Load TourismSmall dataset
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/tourism.csv')
+df = df.rename({'Trips': 'y', 'Quarter': 'ds'}, axis=1)
+df.insert(0, 'Country', 'Australia')
+qs = df['ds'].str.replace(r'(\d+) (Q\d)', r'\1-\2', regex=True)
+df['ds'] = pd.PeriodIndex(qs, freq='Q').to_timestamp()
+
+# Create hierarchical seires based on geographic levels and purpose
+# And Convert quarterly ds string to pd.datetime format
+hierarchy_levels = [['Country'],
+ ['Country', 'State'],
+ ['Country', 'Purpose'],
+ ['Country', 'State', 'Region'],
+ ['Country', 'State', 'Purpose'],
+ ['Country', 'State', 'Region', 'Purpose']]
+
+Y_df, S_df, tags = aggregate(df=df, spec=hierarchy_levels)
+
+# Split train/test sets
+Y_test_df = Y_df.groupby('unique_id').tail(8)
+Y_train_df = Y_df.drop(Y_test_df.index)
+
+# Compute base auto-ETS predictions
+# Careful identifying correct data freq, this data quarterly 'Q'
+fcst = StatsForecast(models=[AutoETS(season_length=4, model='ZZA')], freq='QS', n_jobs=-1)
+Y_hat_df = fcst.forecast(df=Y_train_df, h=8, fitted=True)
+Y_fitted_df = fcst.forecast_fitted_values()
+
+reconcilers = [
+ BottomUp(),
+ MinTrace(method='ols'),
+ MinTrace(method='mint_shrink'),
+ ]
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+Y_rec_df = hrec.reconcile(Y_hat_df=Y_hat_df,
+ Y_df=Y_fitted_df,
+ S_df=S_df, tags=tags)
+
+# Evaluate
+eval_tags = {}
+eval_tags['Total'] = tags['Country']
+eval_tags['Purpose'] = tags['Country/Purpose']
+eval_tags['State'] = tags['Country/State']
+eval_tags['Regions'] = tags['Country/State/Region']
+eval_tags['Bottom'] = tags['Country/State/Region/Purpose']
+
+Y_rec_df_with_y = Y_rec_df.merge(Y_test_df, on=['unique_id', 'ds'], how='left')
+mase_p = partial(mase, seasonality=4)
+
+evaluation = evaluate(Y_rec_df_with_y,
+ metrics=[mase_p, rmse],
+ tags=eval_tags,
+ train_df=Y_train_df)
+
+numeric_cols = evaluation.select_dtypes(include="number").columns
+evaluation[numeric_cols] = evaluation[numeric_cols].map('{:.2f}'.format)
+```
+
diff --git a/hierarchicalforecast/src/evaluation.html.mdx b/hierarchicalforecast/src/evaluation.html.mdx
new file mode 100644
index 00000000..14ec05c8
--- /dev/null
+++ b/hierarchicalforecast/src/evaluation.html.mdx
@@ -0,0 +1,154 @@
+---
+output-file: evaluation.html
+title: Hierarchical Evaluation
+---
+
+
+To assist the evaluation of hierarchical forecasting systems, we make
+available an
+[`evaluate`](https://Nixtla.github.io/hierarchicalforecast/src/evaluation.html#evaluate)
+function that can be used in combination with loss functions from
+`utilsforecast.losses`.
+
+------------------------------------------------------------------------
+
+source
+
+## evaluate
+
+> ``` text
+> evaluate (df:~FrameT, metrics:list[typing.Callable],
+> tags:dict[str,numpy.ndarray], models:Optional[list[str]]=None,
+> train_df:Optional[~FrameT]=None,
+> level:Optional[list[int]]=None, id_col:str='unique_id',
+> time_col:str='ds', target_col:str='y',
+> agg_fn:Optional[str]='mean', benchmark:Optional[str]=None)
+> ```
+
+*Evaluate hierarchical forecast using different metrics.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | FrameT | | Forecasts to evaluate. Must have `id_col`, `time_col`, `target_col` and models’ predictions. |
+| metrics | list | | Functions with arguments `df`, `models`, `id_col`, `target_col` and optionally `train_df`. |
+| tags | dict | | Each key is a level in the hierarchy and its value contains tags associated to that level. |
+| models | Optional | None | Names of the models to evaluate. If `None` will use every column in the dataframe after removing id, time and target. |
+| train_df | Optional | None | Training set. Used to evaluate metrics such as `mase`. |
+| level | Optional | None | Prediction interval levels. Used to compute losses that rely on quantiles. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| agg_fn | Optional | mean | Statistic to compute on the scores by id to reduce them to a single number. |
+| benchmark | Optional | None | If passed, evaluators are scaled by the error of this benchmark model. |
+| **Returns** | **FrameT** | | **Metrics with one row per (id, metric) combination and one column per model. If `agg_fn` is not `None`, there is only one row per metric.** |
+
+# Example
+
+
+```python
+import pandas as pd
+
+from hierarchicalforecast.core import HierarchicalReconciliation
+from hierarchicalforecast.methods import BottomUp, MinTrace
+from hierarchicalforecast.utils import aggregate
+from hierarchicalforecast.evaluation import evaluate
+from statsforecast.core import StatsForecast
+from statsforecast.models import AutoETS
+from utilsforecast.losses import mase, rmse
+from functools import partial
+
+# Load TourismSmall dataset
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/tourism.csv')
+df = df.rename({'Trips': 'y', 'Quarter': 'ds'}, axis=1)
+df.insert(0, 'Country', 'Australia')
+qs = df['ds'].str.replace(r'(\d+) (Q\d)', r'\1-\2', regex=True)
+df['ds'] = pd.PeriodIndex(qs, freq='Q').to_timestamp()
+
+# Create hierarchical seires based on geographic levels and purpose
+# And Convert quarterly ds string to pd.datetime format
+hierarchy_levels = [['Country'],
+ ['Country', 'State'],
+ ['Country', 'Purpose'],
+ ['Country', 'State', 'Region'],
+ ['Country', 'State', 'Purpose'],
+ ['Country', 'State', 'Region', 'Purpose']]
+
+Y_df, S_df, tags = aggregate(df=df, spec=hierarchy_levels)
+
+# Split train/test sets
+Y_test_df = Y_df.groupby('unique_id').tail(8)
+Y_train_df = Y_df.drop(Y_test_df.index)
+
+# Compute base auto-ETS predictions
+# Careful identifying correct data freq, this data quarterly 'Q'
+fcst = StatsForecast(models=[AutoETS(season_length=4, model='ZZA')], freq='QS', n_jobs=-1)
+Y_hat_df = fcst.forecast(df=Y_train_df, h=8, fitted=True)
+Y_fitted_df = fcst.forecast_fitted_values()
+
+reconcilers = [
+ BottomUp(),
+ MinTrace(method='ols'),
+ MinTrace(method='mint_shrink'),
+ ]
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+Y_rec_df = hrec.reconcile(Y_hat_df=Y_hat_df,
+ Y_df=Y_fitted_df,
+ S_df=S_df, tags=tags)
+
+# Evaluate
+eval_tags = {}
+eval_tags['Total'] = tags['Country']
+eval_tags['Purpose'] = tags['Country/Purpose']
+eval_tags['State'] = tags['Country/State']
+eval_tags['Regions'] = tags['Country/State/Region']
+eval_tags['Bottom'] = tags['Country/State/Region/Purpose']
+
+Y_rec_df_with_y = Y_rec_df.merge(Y_test_df, on=['unique_id', 'ds'], how='left')
+mase_p = partial(mase, seasonality=4)
+
+evaluation = evaluate(Y_rec_df_with_y,
+ metrics=[mase_p, rmse],
+ tags=eval_tags,
+ train_df=Y_train_df)
+
+numeric_cols = evaluation.select_dtypes(include="number").columns
+evaluation[numeric_cols] = evaluation[numeric_cols].map('{:.2f}'.format)
+```
+
+# References
+
+- [Gneiting, Tilmann, and Adrian E. Raftery. (2007). "Strictly proper
+ scoring rules, prediction and estimation". Journal of the American
+ Statistical
+ Association.](https://sites.stat.washington.edu/raftery/Research/PDF/Gneiting2007jasa.pdf)
+- [Gneiting, Tilmann. (2011). "Quantiles as optimal point forecasts".
+ International Journal of
+ Forecasting.](https://www.sciencedirect.com/science/article/pii/S0169207010000063)
+- [Spyros Makridakis, Evangelos Spiliotis, Vassilios Assimakopoulos,
+ Zhi Chen, Anil Gaba, Ilia Tsetlin, Robert L. Winkler. (2022). "The
+ M5 uncertainty competition: Results, findings and conclusions".
+ International Journal of
+ Forecasting.](https://www.sciencedirect.com/science/article/pii/S0169207021001722)
+- [Anastasios Panagiotelis, Puwasala Gamakumara, George
+ Athanasopoulos, Rob J. Hyndman. (2022). "Probabilistic forecast
+ reconciliation: Properties, evaluation and score optimisation".
+ European Journal of Operational
+ Research.](https://www.sciencedirect.com/science/article/pii/S0377221722006087)
+- [Syama Sundar Rangapuram, Lucien D Werner, Konstantinos Benidis,
+ Pedro Mercado, Jan Gasthaus, Tim Januschowski. (2021). "End-to-End
+ Learning of Coherent Probabilistic Forecasts for Hierarchical Time
+ Series". Proceedings of the 38th International Conference on Machine
+ Learning
+ (ICML).](https://proceedings.mlr.press/v139/rangapuram21a.html)
+- [Kin G. Olivares, O. Nganba Meetei, Ruijun Ma, Rohan Reddy, Mengfei
+ Cao, Lee Dicker (2022). “Probabilistic Hierarchical Forecasting with
+ Deep Poisson Mixtures”. Submitted to the International Journal
+ Forecasting, Working paper available at
+ arxiv.](https://arxiv.org/pdf/2110.13179.pdf)
+- [Makridakis, S., Spiliotis E., and Assimakopoulos V. (2022). “M5
+ Accuracy Competition: Results, Findings, and Conclusions.”,
+ International Journal of Forecasting, Volume 38, Issue
+ 4.](https://www.sciencedirect.com/science/article/pii/S0169207021001874)
+
diff --git a/hierarchicalforecast/src/methods.html.mdx b/hierarchicalforecast/src/methods.html.mdx
new file mode 100644
index 00000000..d7d54892
--- /dev/null
+++ b/hierarchicalforecast/src/methods.html.mdx
@@ -0,0 +1,1637 @@
+---
+output-file: methods.html
+title: Reconciliation Methods
+---
+
+
+In hierarchical forecasting, we aim to create forecasts for many time
+series concurrently, whilst adhering to pre-specified hierarchical
+relationships that exist between the time series. We can enforce this
+coherence by performing a post-processing reconciliation step on the
+forecasts.
+
+The `HierarchicalForecast` package provides the most comprehensive
+collection of Python implementations of hierarchical forecasting
+algorithms that follow classic hierarchical reconciliation. All the
+methods have a `reconcile` function capable of reconciling base
+forecasts using `numpy` arrays.
+
+## Cross-sectional hierarchies
+
+Traditionally, hierarchical forecasting methods reconcile
+*cross-sectional* aggregations. For example, we may have forecasts for
+individual product demand, but also for the overall product group,
+department and store, and we are interested in making sure these
+forecasts are coherent with each other. This can be formalized as:
+
+$$\tilde{\textbf{Y}} = SP\hat{\textbf{Y}} \;, $$
+
+where $\hat{\textbf{Y}} \in \mathbb{R}^{m \times p}$ denotes the matrix
+of forecasts for all $m$ time series for all $p$ time steps in the
+hierarchy, $S \in \lbrace 0, 1 \rbrace^{m \times n}$ is a matrix that
+defines the hierarchical relationship between the $n$ bottom-level time
+series and the $m^* = m - n$ aggregations,
+$P \in \mathbb{R}^{n \times m}$ is a matrix that encapsulates the
+contribution of each forecast to the final estimate, and
+$\tilde{\textbf{Y}} \in \mathbb{R}^{m \times p}$ is the matrix of
+reconciled forecasts. We can use the matrix $P$ to define various
+forecast contribution scenarios.
+
+Cross-sectional reconciliation methods aim to find the optimal $P$
+matrix.
+
+## Temporal hierarchies
+
+We can also perform *temporal* reconciliation. For example, we may have
+forecasts for daily demand, weekly, and monthly, and we are interested
+in making sure these forecasts are coherent with each other. We
+formalize the temporal hierarchical forecasting problem as:
+
+$$\tilde{\textbf{Y}} = \left( S_{te} P_{te} \hat{\textbf{Y}}^{\intercal} \right)^{\intercal} \;, $$
+
+where $S_{te} \in \lbrace 0, 1 \rbrace^{p \times k}$ is a matrix that
+defines the hierarchical relationship between the $k$ bottom-level time
+steps and the $p^* = p - k$ aggregations and
+$P_{te} \in \mathbb{R}^{k \times p}$ is a matrix that encapsulates the
+contribution of each forecast to the final estimate. We can use the
+matrix $P_{te}$ to define various forecast contribution scenarios.
+
+Temporal reconciliation methods aim to find the optimal $P_{te}$ matrix.
+
+## Cross-temporal reconciliation
+
+We can combine cross-sectional and temporal hierarchical forecasting by
+performing cross-sectional reconciliation and temporal reconciliation in
+a two-step procedure.
+
+**References** -[Hyndman, Rob. Notation for forecast
+reconciliation.](https://robjhyndman.com/hyndsight/reconciliation-notation.html)
+
+# 1. Bottom-Up
+
+------------------------------------------------------------------------
+
+source
+
+### BottomUp
+
+> ``` text
+> BottomUp ()
+> ```
+
+\*Bottom Up Reconciliation Class. The most basic hierarchical
+reconciliation is performed using an Bottom-Up strategy. It was proposed
+for the first time by Orcutt in 1968. The corresponding hierarchical
+“projection” matrix is defined as:
+$$\mathbf{P}_{\text{BU}} = [\mathbf{0}_{\mathrm{[b],[a]}}\;|\;\mathbf{I}_{\mathrm{[b][b]}}]$$
+
+**Parameters:** None
+
+**References:** - [Orcutt, G.H., Watts, H.W., & Edwards, J.B.(1968).
+“Data aggregation and information loss”. The American Economic Review,
+58 , 773(787)](http://www.jstor.org/stable/1815532).\*
+
+------------------------------------------------------------------------
+
+source
+
+### BottomUp.fit
+
+> ``` text
+> BottomUp.fit (S:numpy.ndarray, y_hat:numpy.ndarray,
+> idx_bottom:numpy.ndarray,
+> y_insample:Optional[numpy.ndarray]=None,
+> y_hat_insample:Optional[numpy.ndarray]=None,
+> sigmah:Optional[numpy.ndarray]=None,
+> intervals_method:Optional[str]=None,
+> num_samples:Optional[int]=None, seed:Optional[int]=None,
+> tags:Optional[dict[str,numpy.ndarray]]=None)
+> ```
+
+\*Bottom Up Fit Method.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `idx_bottom`:
+Indices corresponding to the bottom level of `S`, size (`bottom`).
+`y_insample`: In-sample values of size (`base`, `horizon`).
+`y_hat_insample`: In-sample forecast values of size (`base`,
+`horizon`). `sigmah`: Estimated standard deviation of the
+conditional marginal distribution.
+`intervals_method`: Sampler for prediction intervals, one of
+`normality`, `bootstrap`, `permbu`. `num_samples`: Number of samples
+for probabilistic coherent distribution. `seed`: Seed for
+reproducibility. `**sampler_kwargs`: Coherent sampler instantiation
+arguments.
+
+**Returns:** `self`: object, fitted reconciler.\*
+
+------------------------------------------------------------------------
+
+source
+
+### BottomUp.predict
+
+> ``` text
+> BottomUp.predict (S:numpy.ndarray, y_hat:numpy.ndarray,
+> level:Optional[list[int]]=None)
+> ```
+
+\*Predict using reconciler.
+
+Predict using fitted mean and probabilistic reconcilers.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `level`: float
+list 0-100, confidence levels for prediction intervals.
+
+**Returns:** `y_tilde`: Reconciliated predictions.\*
+
+------------------------------------------------------------------------
+
+source
+
+### BottomUp.fit_predict
+
+> ``` text
+> BottomUp.fit_predict (S:numpy.ndarray, y_hat:numpy.ndarray,
+> idx_bottom:numpy.ndarray,
+> y_insample:Optional[numpy.ndarray]=None,
+> y_hat_insample:Optional[numpy.ndarray]=None,
+> sigmah:Optional[numpy.ndarray]=None,
+> level:Optional[list[int]]=None,
+> intervals_method:Optional[str]=None,
+> num_samples:Optional[int]=None,
+> seed:Optional[int]=None,
+> tags:Optional[dict[str,numpy.ndarray]]=None)
+> ```
+
+\*BottomUp Reconciliation Method.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `idx_bottom`:
+Indices corresponding to the bottom level of `S`, size (`bottom`).
+`y_insample`: In-sample values of size (`base`, `insample_size`).
+`y_hat_insample`: In-sample forecast values of size (`base`,
+`insample_size`). `sigmah`: Estimated standard deviation of the
+conditional marginal distribution.
+`level`: float list 0-100, confidence levels for prediction
+intervals. `intervals_method`: Sampler for prediction intervals, one
+of `normality`, `bootstrap`, `permbu`. `num_samples`: Number of
+samples for probabilistic coherent distribution. `seed`: Seed for
+reproducibility. `**sampler_kwargs`: Coherent sampler instantiation
+arguments.
+
+**Returns:** `y_tilde`: Reconciliated y_hat using the Bottom Up
+approach.\*
+
+------------------------------------------------------------------------
+
+source
+
+### BottomUp.sample
+
+> ``` text
+> BottomUp.sample (num_samples:int)
+> ```
+
+\*Sample probabilistic coherent distribution.
+
+Generates n samples from a probabilistic coherent distribution. The
+method uses fitted mean and probabilistic reconcilers, defined by the
+`intervals_method` selected during the reconciler’s instantiation.
+Currently available: `normality`, `bootstrap`, `permbu`.
+
+**Parameters:** `num_samples`: int, number of samples generated from
+coherent distribution.
+
+**Returns:** `samples`: Coherent samples of size (`num_series`,
+`horizon`, `num_samples`).\*
+
+------------------------------------------------------------------------
+
+source
+
+### BottomUpSparse
+
+> ``` text
+> BottomUpSparse ()
+> ```
+
+\*BottomUpSparse Reconciliation Class.
+
+This is the implementation of a Bottom Up reconciliation using the
+sparse matrix approach. It works much more efficient on datasets with
+many time series. \[makoren: At least I hope so, I only checked up until
+~20k time series, and there’s no real improvement, it would be great to
+check for smth like 1M time series, where the dense S matrix really
+stops fitting in memory\]
+
+See the parent class for more details.\*
+
+------------------------------------------------------------------------
+
+source
+
+### BottomUpSparse.fit
+
+> ``` text
+> BottomUpSparse.fit (S:numpy.ndarray, y_hat:numpy.ndarray,
+> idx_bottom:numpy.ndarray,
+> y_insample:Optional[numpy.ndarray]=None,
+> y_hat_insample:Optional[numpy.ndarray]=None,
+> sigmah:Optional[numpy.ndarray]=None,
+> intervals_method:Optional[str]=None,
+> num_samples:Optional[int]=None,
+> seed:Optional[int]=None,
+> tags:Optional[dict[str,numpy.ndarray]]=None)
+> ```
+
+\*Bottom Up Fit Method.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `idx_bottom`:
+Indices corresponding to the bottom level of `S`, size (`bottom`).
+`y_insample`: In-sample values of size (`base`, `horizon`).
+`y_hat_insample`: In-sample forecast values of size (`base`,
+`horizon`). `sigmah`: Estimated standard deviation of the
+conditional marginal distribution.
+`intervals_method`: Sampler for prediction intervals, one of
+`normality`, `bootstrap`, `permbu`. `num_samples`: Number of samples
+for probabilistic coherent distribution. `seed`: Seed for
+reproducibility. `**sampler_kwargs`: Coherent sampler instantiation
+arguments.
+
+**Returns:** `self`: object, fitted reconciler.\*
+
+------------------------------------------------------------------------
+
+source
+
+### BottomUpSparse.predict
+
+> ``` text
+> BottomUpSparse.predict (S:numpy.ndarray, y_hat:numpy.ndarray,
+> level:Optional[list[int]]=None)
+> ```
+
+\*Predict using reconciler.
+
+Predict using fitted mean and probabilistic reconcilers.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `level`: float
+list 0-100, confidence levels for prediction intervals.
+
+**Returns:** `y_tilde`: Reconciliated predictions.\*
+
+------------------------------------------------------------------------
+
+source
+
+### BottomUpSparse.fit_predict
+
+> ``` text
+> BottomUpSparse.fit_predict (S:numpy.ndarray, y_hat:numpy.ndarray,
+> idx_bottom:numpy.ndarray,
+> y_insample:Optional[numpy.ndarray]=None,
+> y_hat_insample:Optional[numpy.ndarray]=None,
+> sigmah:Optional[numpy.ndarray]=None,
+> level:Optional[list[int]]=None,
+> intervals_method:Optional[str]=None,
+> num_samples:Optional[int]=None,
+> seed:Optional[int]=None,
+> tags:Optional[dict[str,numpy.ndarray]]=None)
+> ```
+
+\*BottomUp Reconciliation Method.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `idx_bottom`:
+Indices corresponding to the bottom level of `S`, size (`bottom`).
+`y_insample`: In-sample values of size (`base`, `insample_size`).
+`y_hat_insample`: In-sample forecast values of size (`base`,
+`insample_size`). `sigmah`: Estimated standard deviation of the
+conditional marginal distribution.
+`level`: float list 0-100, confidence levels for prediction
+intervals. `intervals_method`: Sampler for prediction intervals, one
+of `normality`, `bootstrap`, `permbu`. `num_samples`: Number of
+samples for probabilistic coherent distribution. `seed`: Seed for
+reproducibility. `**sampler_kwargs`: Coherent sampler instantiation
+arguments.
+
+**Returns:** `y_tilde`: Reconciliated y_hat using the Bottom Up
+approach.\*
+
+------------------------------------------------------------------------
+
+source
+
+### BottomUpSparse.sample
+
+> ``` text
+> BottomUpSparse.sample (num_samples:int)
+> ```
+
+\*Sample probabilistic coherent distribution.
+
+Generates n samples from a probabilistic coherent distribution. The
+method uses fitted mean and probabilistic reconcilers, defined by the
+`intervals_method` selected during the reconciler’s instantiation.
+Currently available: `normality`, `bootstrap`, `permbu`.
+
+**Parameters:** `num_samples`: int, number of samples generated from
+coherent distribution.
+
+**Returns:** `samples`: Coherent samples of size (`num_series`,
+`horizon`, `num_samples`).\*
+
+# 2. Top-Down
+
+------------------------------------------------------------------------
+
+source
+
+### TopDown
+
+> ``` text
+> TopDown (method:str)
+> ```
+
+\*Top Down Reconciliation Class.
+
+The Top Down hierarchical reconciliation method, distributes the total
+aggregate predictions and decomposes it down the hierarchy using
+proportions $\mathbf{p}_{\mathrm{[b]}}$ that can be actual historical
+values or estimated.
+
+$$\mathbf{P}=[\mathbf{p}_{\mathrm{[b]}}\;|\;\mathbf{0}_{\mathrm{[b][a,b\;-1]}}]$$
+**Parameters:** `method`: One of `forecast_proportions`,
+`average_proportions` and `proportion_averages`.
+
+**References:** - [CW. Gross (1990). “Disaggregation methods to
+expedite product line forecasting”. Journal of Forecasting, 9 , 233–254.
+doi:10.1002/for.3980090304](https://onlinelibrary.wiley.com/doi/abs/10.1002/for.3980090304). -
+[G. Fliedner (1999). “An investigation of aggregate variable time series
+forecast strategies with specific subaggregate time series statistical
+correlation”. Computers and Operations Research, 26 , 1133–1149.
+doi:10.1016/S0305-0548(99)00017-9](https://doi.org/10.1016/S0305-0548(99)00017-9).\*
+
+------------------------------------------------------------------------
+
+source
+
+### TopDown.fit
+
+> ``` text
+> TopDown.fit (S, y_hat, y_insample:numpy.ndarray,
+> y_hat_insample:Optional[numpy.ndarray]=None,
+> sigmah:Optional[numpy.ndarray]=None,
+> intervals_method:Optional[str]=None,
+> num_samples:Optional[int]=None, seed:Optional[int]=None,
+> tags:Optional[dict[str,numpy.ndarray]]=None,
+> idx_bottom:Optional[numpy.ndarray]=None)
+> ```
+
+\*TopDown Fit Method.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `y_insample`:
+Insample values of size (`base`, `insample_size`). Optional for
+`forecast_proportions` method. `y_hat_insample`: Insample forecast
+values of size (`base`, `insample_size`). Optional for
+`forecast_proportions` method. `sigmah`: Estimated standard
+deviation of the conditional marginal distribution.
+`interval_method`: Sampler for prediction intervals, one of `normality`,
+`bootstrap`, `permbu`. `num_samples`: Number of samples for
+probabilistic coherent distribution. `seed`: Seed for
+reproducibility. `tags`: Each key is a level and each value its `S`
+indices. `idx_bottom`: Indices corresponding to the bottom level of
+`S`, size (`bottom`).
+
+**Returns:** `self`: object, fitted reconciler.\*
+
+------------------------------------------------------------------------
+
+source
+
+### TopDown.predict
+
+> ``` text
+> TopDown.predict (S:numpy.ndarray, y_hat:numpy.ndarray,
+> level:Optional[list[int]]=None)
+> ```
+
+\*Predict using reconciler.
+
+Predict using fitted mean and probabilistic reconcilers.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `level`: float
+list 0-100, confidence levels for prediction intervals.
+
+**Returns:** `y_tilde`: Reconciliated predictions.\*
+
+------------------------------------------------------------------------
+
+source
+
+### TopDown.fit_predict
+
+> ``` text
+> TopDown.fit_predict (S:numpy.ndarray, y_hat:numpy.ndarray,
+> tags:dict[str,numpy.ndarray],
+> idx_bottom:numpy.ndarray=None,
+> y_insample:Optional[numpy.ndarray]=None,
+> y_hat_insample:Optional[numpy.ndarray]=None,
+> sigmah:Optional[numpy.ndarray]=None,
+> level:Optional[list[int]]=None,
+> intervals_method:Optional[str]=None,
+> num_samples:Optional[int]=None,
+> seed:Optional[int]=None)
+> ```
+
+\*Top Down Reconciliation Method.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `tags`: Each
+key is a level and each value its `S` indices. `idx_bottom`: Indices
+corresponding to the bottom level of `S`, size (`bottom`).
+`y_insample`: Insample values of size (`base`, `insample_size`).
+Optional for `forecast_proportions` method. `y_hat_insample`:
+Insample forecast values of size (`base`, `insample_size`). Optional for
+`forecast_proportions` method. `sigmah`: Estimated standard
+deviation of the conditional marginal distribution. `level`: float
+list 0-100, confidence levels for prediction intervals.
+`intervals_method`: Sampler for prediction intervals, one of
+`normality`, `bootstrap`, `permbu`. `num_samples`: Number of samples
+for probabilistic coherent distribution. `seed`: Seed for
+reproducibility.
+
+**Returns:** `y_tilde`: Reconciliated y_hat using the Top Down
+approach.\*
+
+------------------------------------------------------------------------
+
+source
+
+### TopDown.sample
+
+> ``` text
+> TopDown.sample (num_samples:int)
+> ```
+
+\*Sample probabilistic coherent distribution.
+
+Generates n samples from a probabilistic coherent distribution. The
+method uses fitted mean and probabilistic reconcilers, defined by the
+`intervals_method` selected during the reconciler’s instantiation.
+Currently available: `normality`, `bootstrap`, `permbu`.
+
+**Parameters:** `num_samples`: int, number of samples generated from
+coherent distribution.
+
+**Returns:** `samples`: Coherent samples of size (`num_series`,
+`horizon`, `num_samples`).\*
+
+------------------------------------------------------------------------
+
+source
+
+### TopDownSparse
+
+> ``` text
+> TopDownSparse (method:str)
+> ```
+
+\*TopDownSparse Reconciliation Class.
+
+This is an implementation of top-down reconciliation using the sparse
+matrix approach. It works much more efficiently on data sets with many
+time series.
+
+See the parent class for more details.\*
+
+------------------------------------------------------------------------
+
+source
+
+### TopDownSparse.fit
+
+> ``` text
+> TopDownSparse.fit (S, y_hat, y_insample:numpy.ndarray,
+> y_hat_insample:Optional[numpy.ndarray]=None,
+> sigmah:Optional[numpy.ndarray]=None,
+> intervals_method:Optional[str]=None,
+> num_samples:Optional[int]=None,
+> seed:Optional[int]=None,
+> tags:Optional[dict[str,numpy.ndarray]]=None,
+> idx_bottom:Optional[numpy.ndarray]=None)
+> ```
+
+\*TopDown Fit Method.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `y_insample`:
+Insample values of size (`base`, `insample_size`). Optional for
+`forecast_proportions` method. `y_hat_insample`: Insample forecast
+values of size (`base`, `insample_size`). Optional for
+`forecast_proportions` method. `sigmah`: Estimated standard
+deviation of the conditional marginal distribution.
+`interval_method`: Sampler for prediction intervals, one of `normality`,
+`bootstrap`, `permbu`. `num_samples`: Number of samples for
+probabilistic coherent distribution. `seed`: Seed for
+reproducibility. `tags`: Each key is a level and each value its `S`
+indices. `idx_bottom`: Indices corresponding to the bottom level of
+`S`, size (`bottom`).
+
+**Returns:** `self`: object, fitted reconciler.\*
+
+------------------------------------------------------------------------
+
+source
+
+### TopDownSparse.predict
+
+> ``` text
+> TopDownSparse.predict (S:numpy.ndarray, y_hat:numpy.ndarray,
+> level:Optional[list[int]]=None)
+> ```
+
+\*Predict using reconciler.
+
+Predict using fitted mean and probabilistic reconcilers.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `level`: float
+list 0-100, confidence levels for prediction intervals.
+
+**Returns:** `y_tilde`: Reconciliated predictions.\*
+
+------------------------------------------------------------------------
+
+source
+
+### TopDownSparse.fit_predict
+
+> ``` text
+> TopDownSparse.fit_predict (S:scipy.sparse._csr.csr_matrix,
+> y_hat:numpy.ndarray,
+> tags:dict[str,numpy.ndarray],
+> idx_bottom:numpy.ndarray=None,
+> y_insample:Optional[numpy.ndarray]=None,
+> y_hat_insample:Optional[numpy.ndarray]=None,
+> sigmah:Optional[numpy.ndarray]=None,
+> level:Optional[list[int]]=None,
+> intervals_method:Optional[str]=None,
+> num_samples:Optional[int]=None,
+> seed:Optional[int]=None)
+> ```
+
+\*Top Down Reconciliation Method.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `tags`: Each
+key is a level and each value its `S` indices. `idx_bottom`: Indices
+corresponding to the bottom level of `S`, size (`bottom`).
+`y_insample`: Insample values of size (`base`, `insample_size`).
+Optional for `forecast_proportions` method. `y_hat_insample`:
+Insample forecast values of size (`base`, `insample_size`). Optional for
+`forecast_proportions` method. `sigmah`: Estimated standard
+deviation of the conditional marginal distribution. `level`: float
+list 0-100, confidence levels for prediction intervals.
+`intervals_method`: Sampler for prediction intervals, one of
+`normality`, `bootstrap`, `permbu`. `num_samples`: Number of samples
+for probabilistic coherent distribution. `seed`: Seed for
+reproducibility.
+
+**Returns:** `y_tilde`: Reconciliated y_hat using the Top Down
+approach.\*
+
+------------------------------------------------------------------------
+
+source
+
+### TopDownSparse.sample
+
+> ``` text
+> TopDownSparse.sample (num_samples:int)
+> ```
+
+\*Sample probabilistic coherent distribution.
+
+Generates n samples from a probabilistic coherent distribution. The
+method uses fitted mean and probabilistic reconcilers, defined by the
+`intervals_method` selected during the reconciler’s instantiation.
+Currently available: `normality`, `bootstrap`, `permbu`.
+
+**Parameters:** `num_samples`: int, number of samples generated from
+coherent distribution.
+
+**Returns:** `samples`: Coherent samples of size (`num_series`,
+`horizon`, `num_samples`).\*
+
+
+```python
+cls_top_down(
+ S=S, y_hat=S @ y_hat_bottom, y_insample=S @ y_bottom, tags=tags
+ )["mean"]
+```
+
+
+```python
+#\ hide
+cls_top_down = TopDownSparse(method="average_proportions")
+test_fail(
+ cls_top_down,
+ contains="Top-down reconciliation requires strictly hierarchical structures.",
+ args=(sparse.csr_matrix(S_non_hier), None, tags_non_hier),
+)
+```
+
+# 3. Middle-Out
+
+------------------------------------------------------------------------
+
+source
+
+### MiddleOut
+
+> ``` text
+> MiddleOut (middle_level:str, top_down_method:str)
+> ```
+
+\*Middle Out Reconciliation Class.
+
+This method is only available for **strictly hierarchical structures**.
+It anchors the base predictions in a middle level. The levels above the
+base predictions use the Bottom-Up approach, while the levels below use
+a Top-Down.
+
+**Parameters:** `middle_level`: Middle level. `top_down_method`:
+One of `forecast_proportions`, `average_proportions` and
+`proportion_averages`.
+
+**References:** - [Hyndman, R.J., & Athanasopoulos, G. (2021).
+“Forecasting: principles and practice, 3rd edition: Chapter 11:
+Forecasting hierarchical and grouped series.”. OTexts: Melbourne,
+Australia. OTexts.com/fpp3 Accessed on July
+2022.](https://otexts.com/fpp3/hierarchical.html)\*
+
+------------------------------------------------------------------------
+
+source
+
+### MiddleOut.fit
+
+> ``` text
+> MiddleOut.fit (**kwargs)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### MiddleOut.predict
+
+> ``` text
+> MiddleOut.predict (**kwargs)
+> ```
+
+\*Predict using reconciler.
+
+Predict using fitted mean and probabilistic reconcilers.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `level`: float
+list 0-100, confidence levels for prediction intervals.
+
+**Returns:** `y_tilde`: Reconciliated predictions.\*
+
+------------------------------------------------------------------------
+
+source
+
+### MiddleOut.fit_predict
+
+> ``` text
+> MiddleOut.fit_predict (S:numpy.ndarray, y_hat:numpy.ndarray,
+> tags:dict[str,numpy.ndarray],
+> y_insample:Optional[numpy.ndarray]=None,
+> y_hat_insample:Optional[numpy.ndarray]=None,
+> sigmah:Optional[numpy.ndarray]=None,
+> level:Optional[list[int]]=None,
+> intervals_method:Optional[str]=None,
+> num_samples:Optional[int]=None,
+> seed:Optional[int]=None)
+> ```
+
+\*Middle Out Reconciliation Method.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `tags`: Each
+key is a level and each value its `S` indices. `y_insample`:
+Insample values of size (`base`, `insample_size`). Only used for
+`forecast_proportions` `y_hat_insample`: In-sample forecast values
+of size (`base`, `insample_size`). `sigmah`: Estimated standard
+deviation of the conditional marginal distribution.
+`level`: float list 0-100, confidence levels for prediction
+intervals. `intervals_method`: Sampler for prediction intervals, one
+of `normality`, `bootstrap`, `permbu`. `num_samples`: Number of
+samples for probabilistic coherent distribution. `seed`: Seed for
+reproducibility.
+
+**Returns:** `y_tilde`: Reconciliated y_hat using the Middle Out
+approach.\*
+
+------------------------------------------------------------------------
+
+source
+
+### MiddleOut.sample
+
+> ``` text
+> MiddleOut.sample (num_samples:int)
+> ```
+
+\*Sample probabilistic coherent distribution.
+
+Generates n samples from a probabilistic coherent distribution. The
+method uses fitted mean and probabilistic reconcilers, defined by the
+`intervals_method` selected during the reconciler’s instantiation.
+Currently available: `normality`, `bootstrap`, `permbu`.
+
+**Parameters:** `num_samples`: int, number of samples generated from
+coherent distribution.
+
+**Returns:** `samples`: Coherent samples of size (`num_series`,
+`horizon`, `num_samples`).\*
+
+------------------------------------------------------------------------
+
+source
+
+### MiddleOutSparse
+
+> ``` text
+> MiddleOutSparse (middle_level:str, top_down_method:str)
+> ```
+
+\*MiddleOutSparse Reconciliation Class.
+
+This is an implementation of middle-out reconciliation using the sparse
+matrix approach. It works much more efficiently on data sets with many
+time series.
+
+See the parent class for more details.\*
+
+------------------------------------------------------------------------
+
+source
+
+### MiddleOutSparse.fit
+
+> ``` text
+> MiddleOutSparse.fit (**kwargs)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### MiddleOutSparse.predict
+
+> ``` text
+> MiddleOutSparse.predict (**kwargs)
+> ```
+
+\*Predict using reconciler.
+
+Predict using fitted mean and probabilistic reconcilers.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `level`: float
+list 0-100, confidence levels for prediction intervals.
+
+**Returns:** `y_tilde`: Reconciliated predictions.\*
+
+------------------------------------------------------------------------
+
+source
+
+### MiddleOutSparse.fit_predict
+
+> ``` text
+> MiddleOutSparse.fit_predict (S:numpy.ndarray, y_hat:numpy.ndarray,
+> tags:dict[str,numpy.ndarray],
+> y_insample:Optional[numpy.ndarray]=None,
+> y_hat_insample:Optional[numpy.ndarray]=None,
+> sigmah:Optional[numpy.ndarray]=None,
+> level:Optional[list[int]]=None,
+> intervals_method:Optional[str]=None,
+> num_samples:Optional[int]=None,
+> seed:Optional[int]=None)
+> ```
+
+\*Middle Out Reconciliation Method.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `tags`: Each
+key is a level and each value its `S` indices. `y_insample`:
+Insample values of size (`base`, `insample_size`). Only used for
+`forecast_proportions` `y_hat_insample`: In-sample forecast values
+of size (`base`, `insample_size`). `sigmah`: Estimated standard
+deviation of the conditional marginal distribution.
+`level`: float list 0-100, confidence levels for prediction
+intervals. `intervals_method`: Sampler for prediction intervals, one
+of `normality`, `bootstrap`, `permbu`. `num_samples`: Number of
+samples for probabilistic coherent distribution. `seed`: Seed for
+reproducibility.
+
+**Returns:** `y_tilde`: Reconciliated y_hat using the Middle Out
+approach.\*
+
+------------------------------------------------------------------------
+
+source
+
+### MiddleOutSparse.sample
+
+> ``` text
+> MiddleOutSparse.sample (num_samples:int)
+> ```
+
+\*Sample probabilistic coherent distribution.
+
+Generates n samples from a probabilistic coherent distribution. The
+method uses fitted mean and probabilistic reconcilers, defined by the
+`intervals_method` selected during the reconciler’s instantiation.
+Currently available: `normality`, `bootstrap`, `permbu`.
+
+**Parameters:** `num_samples`: int, number of samples generated from
+coherent distribution.
+
+**Returns:** `samples`: Coherent samples of size (`num_series`,
+`horizon`, `num_samples`).\*
+
+# 4. Min-Trace
+
+------------------------------------------------------------------------
+
+source
+
+### MinTrace
+
+> ``` text
+> MinTrace (method:str, nonnegative:bool=False,
+> mint_shr_ridge:Optional[float]=2e-08, num_threads:int=1)
+> ```
+
+\*MinTrace Reconciliation Class.
+
+This reconciliation algorithm proposed by Wickramasuriya et al. depends
+on a generalized least squares estimator and an estimator of the
+covariance matrix of the coherency errors $\mathbf{W}_{h}$. The Min
+Trace algorithm minimizes the squared errors for the coherent forecasts
+under an unbiasedness assumption; the solution has a closed form.
+
+$$
+
+\mathbf{P}_{\text{MinT}}=\left(\mathbf{S}^{\intercal}\mathbf{W}_{h}\mathbf{S}\right)^{-1}
+\mathbf{S}^{\intercal}\mathbf{W}^{-1}_{h}
+
+$$
+
+**Parameters:** `method`: str, one of `ols`, `wls_struct`,
+`wls_var`, `mint_shrink`, `mint_cov`. `nonnegative`: bool,
+reconciled forecasts should be nonnegative? `mint_shr_ridge`:
+float=2e-8, ridge numeric protection to MinTrace-shr covariance
+estimator. `num_threads`: int=1, number of threads to use for
+solving the optimization problems (when nonnegative=True).
+
+**References:** - [Wickramasuriya, S. L., Athanasopoulos, G., &
+Hyndman, R. J. (2019). “Optimal forecast reconciliation for hierarchical
+and grouped time series through trace minimization”. Journal of the
+American Statistical Association, 114 , 804–819.
+doi:10.1080/01621459.2018.1448825.](https://robjhyndman.com/publications/mint/). -
+[Wickramasuriya, S.L., Turlach, B.A. & Hyndman, R.J. (2020). “Optimal
+non-negative forecast reconciliation”. Stat Comput 30, 1167–1182,
+https://doi.org/10.1007/s11222-020-09930-0](https://robjhyndman.com/publications/nnmint/).\*
+
+------------------------------------------------------------------------
+
+source
+
+### MinTrace.fit
+
+> ``` text
+> MinTrace.fit (S, y_hat, y_insample:Optional[numpy.ndarray]=None,
+> y_hat_insample:Optional[numpy.ndarray]=None,
+> sigmah:Optional[numpy.ndarray]=None,
+> intervals_method:Optional[str]=None,
+> num_samples:Optional[int]=None, seed:Optional[int]=None,
+> tags:Optional[dict[str,numpy.ndarray]]=None,
+> idx_bottom:Optional[numpy.ndarray]=None)
+> ```
+
+\*MinTrace Fit Method.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `y_insample`:
+Insample values of size (`base`, `insample_size`). Only used with
+“wls_var”, “mint_cov”, “mint_shrink”. `y_hat_insample`: Insample
+forecast values of size (`base`, `insample_size`). Only used with
+“wls_var”, “mint_cov”, “mint_shrink” `sigmah`: Estimated standard
+deviation of the conditional marginal distribution.
+`intervals_method`: Sampler for prediction intervals, one of
+`normality`, `bootstrap`, `permbu`. `num_samples`: Number of samples
+for probabilistic coherent distribution. `seed`: Seed for
+reproducibility. `tags`: Each key is a level and each value its `S`
+indices. `idx_bottom`: Indices corresponding to the bottom level of
+`S`, size (`bottom`).
+
+**Returns:** `self`: object, fitted reconciler.\*
+
+------------------------------------------------------------------------
+
+source
+
+### MinTrace.predict
+
+> ``` text
+> MinTrace.predict (S:numpy.ndarray, y_hat:numpy.ndarray,
+> level:Optional[list[int]]=None)
+> ```
+
+\*Predict using reconciler.
+
+Predict using fitted mean and probabilistic reconcilers.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `level`: float
+list 0-100, confidence levels for prediction intervals.
+
+**Returns:** `y_tilde`: Reconciliated predictions.\*
+
+------------------------------------------------------------------------
+
+source
+
+### MinTrace.fit_predict
+
+> ``` text
+> MinTrace.fit_predict (S:numpy.ndarray, y_hat:numpy.ndarray,
+> idx_bottom:numpy.ndarray=None,
+> y_insample:Optional[numpy.ndarray]=None,
+> y_hat_insample:Optional[numpy.ndarray]=None,
+> sigmah:Optional[numpy.ndarray]=None,
+> level:Optional[list[int]]=None,
+> intervals_method:Optional[str]=None,
+> num_samples:Optional[int]=None,
+> seed:Optional[int]=None,
+> tags:Optional[dict[str,numpy.ndarray]]=None)
+> ```
+
+\*MinTrace Reconciliation Method.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `idx_bottom`:
+Indices corresponding to the bottom level of `S`, size (`bottom`).
+`y_insample`: Insample values of size (`base`, `insample_size`). Only
+used by `wls_var`, `mint_cov`, `mint_shrink` `y_hat_insample`:
+Insample fitted values of size (`base`, `insample_size`). Only used by
+`wls_var`, `mint_cov`, `mint_shrink` `sigmah`: Estimated standard
+deviation of the conditional marginal distribution. `level`: float
+list 0-100, confidence levels for prediction intervals.
+`intervals_method`: Sampler for prediction intervals, one of
+`normality`, `bootstrap`, `permbu`. `num_samples`: Number of samples
+for probabilistic coherent distribution. `seed`: Seed for
+reproducibility. `tags`: Each key is a level and each value its `S`
+indices.
+
+**Returns:** `y_tilde`: Reconciliated y_hat using the MinTrace
+approach.\*
+
+------------------------------------------------------------------------
+
+source
+
+### MinTrace.sample
+
+> ``` text
+> MinTrace.sample (num_samples:int)
+> ```
+
+\*Sample probabilistic coherent distribution.
+
+Generates n samples from a probabilistic coherent distribution. The
+method uses fitted mean and probabilistic reconcilers, defined by the
+`intervals_method` selected during the reconciler’s instantiation.
+Currently available: `normality`, `bootstrap`, `permbu`.
+
+**Parameters:** `num_samples`: int, number of samples generated from
+coherent distribution.
+
+**Returns:** `samples`: Coherent samples of size (`num_series`,
+`horizon`, `num_samples`).\*
+
+------------------------------------------------------------------------
+
+source
+
+### MinTraceSparse
+
+> ``` text
+> MinTraceSparse (method:str, nonnegative:bool=False, num_threads:int=1,
+> qp:bool=True)
+> ```
+
+\*MinTraceSparse Reconciliation Class.
+
+This is the implementation of OLS and WLS estimators using sparse
+matrices. It is not guaranteed to give identical results to the
+non-sparse version, but works much more efficiently on data sets with
+many time series.
+
+See the parent class for more details.
+
+**Parameters:** `method`: str, one of `ols`, `wls_struct`, or
+`wls_var`. `nonnegative`: bool, return non-negative reconciled
+forecasts. `num_threads`: int, number of threads to execute
+non-negative quadratic programming calls. `qp`: bool, implement
+non-negativity constraint with a quadratic programming approach. Setting
+this to True generally gives better results, but at the expense of
+higher cost to compute. \*
+
+------------------------------------------------------------------------
+
+source
+
+### MinTraceSparse.fit
+
+> ``` text
+> MinTraceSparse.fit (S:scipy.sparse._csr.csr_matrix, y_hat:numpy.ndarray,
+> y_insample:Optional[numpy.ndarray]=None,
+> y_hat_insample:Optional[numpy.ndarray]=None,
+> sigmah:Optional[numpy.ndarray]=None,
+> intervals_method:Optional[str]=None,
+> num_samples:Optional[int]=None,
+> seed:Optional[int]=None,
+> tags:Optional[dict[str,numpy.ndarray]]=None,
+> idx_bottom:Optional[numpy.ndarray]=None)
+> ```
+
+\*MinTraceSparse Fit Method.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `y_insample`:
+Insample values of size (`base`, `insample_size`). Only used with
+“wls_var”. `y_hat_insample`: Insample forecast values of size
+(`base`, `insample_size`). Only used with “wls_var” `sigmah`:
+Estimated standard deviation of the conditional marginal
+distribution. `intervals_method`: Sampler for prediction intervals,
+one of `normality`, `bootstrap`, `permbu`. `num_samples`: Number of
+samples for probabilistic coherent distribution. `seed`: Seed for
+reproducibility. `tags`: Each key is a level and each value its `S`
+indices. `idx_bottom`: Indices corresponding to the bottom level of
+`S`, size (`bottom`).
+
+**Returns:** `self`: object, fitted reconciler.\*
+
+------------------------------------------------------------------------
+
+source
+
+### MinTraceSparse.predict
+
+> ``` text
+> MinTraceSparse.predict (S:numpy.ndarray, y_hat:numpy.ndarray,
+> level:Optional[list[int]]=None)
+> ```
+
+\*Predict using reconciler.
+
+Predict using fitted mean and probabilistic reconcilers.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `level`: float
+list 0-100, confidence levels for prediction intervals.
+
+**Returns:** `y_tilde`: Reconciliated predictions.\*
+
+------------------------------------------------------------------------
+
+source
+
+### MinTraceSparse.fit_predict
+
+> ``` text
+> MinTraceSparse.fit_predict (S:numpy.ndarray, y_hat:numpy.ndarray,
+> idx_bottom:numpy.ndarray=None,
+> y_insample:Optional[numpy.ndarray]=None,
+> y_hat_insample:Optional[numpy.ndarray]=None,
+> sigmah:Optional[numpy.ndarray]=None,
+> level:Optional[list[int]]=None,
+> intervals_method:Optional[str]=None,
+> num_samples:Optional[int]=None,
+> seed:Optional[int]=None,
+> tags:Optional[dict[str,numpy.ndarray]]=None)
+> ```
+
+\*MinTrace Reconciliation Method.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `idx_bottom`:
+Indices corresponding to the bottom level of `S`, size (`bottom`).
+`y_insample`: Insample values of size (`base`, `insample_size`). Only
+used by `wls_var`, `mint_cov`, `mint_shrink` `y_hat_insample`:
+Insample fitted values of size (`base`, `insample_size`). Only used by
+`wls_var`, `mint_cov`, `mint_shrink` `sigmah`: Estimated standard
+deviation of the conditional marginal distribution. `level`: float
+list 0-100, confidence levels for prediction intervals.
+`intervals_method`: Sampler for prediction intervals, one of
+`normality`, `bootstrap`, `permbu`. `num_samples`: Number of samples
+for probabilistic coherent distribution. `seed`: Seed for
+reproducibility. `tags`: Each key is a level and each value its `S`
+indices.
+
+**Returns:** `y_tilde`: Reconciliated y_hat using the MinTrace
+approach.\*
+
+------------------------------------------------------------------------
+
+source
+
+### MinTraceSparse.sample
+
+> ``` text
+> MinTraceSparse.sample (num_samples:int)
+> ```
+
+\*Sample probabilistic coherent distribution.
+
+Generates n samples from a probabilistic coherent distribution. The
+method uses fitted mean and probabilistic reconcilers, defined by the
+`intervals_method` selected during the reconciler’s instantiation.
+Currently available: `normality`, `bootstrap`, `permbu`.
+
+**Parameters:** `num_samples`: int, number of samples generated from
+coherent distribution.
+
+**Returns:** `samples`: Coherent samples of size (`num_series`,
+`horizon`, `num_samples`).\*
+
+# 5. Optimal Combination
+
+------------------------------------------------------------------------
+
+source
+
+### OptimalCombination
+
+> ``` text
+> OptimalCombination (method:str, nonnegative:bool=False,
+> num_threads:int=1)
+> ```
+
+\*Optimal Combination Reconciliation Class.
+
+This reconciliation algorithm was proposed by Hyndman et al. 2011, the
+method uses generalized least squares estimator using the coherency
+errors covariance matrix. Consider the covariance of the base forecast
+$\textrm{Var}(\epsilon_{h}) = \Sigma_{h}$, the $\mathbf{P}$ matrix of
+this method is defined by:
+$$ \mathbf{P} = \left(\mathbf{S}^{\intercal}\Sigma_{h}^{\dagger}\mathbf{S}\right)^{-1}\mathbf{S}^{\intercal}\Sigma^{\dagger}_{h}$$
+where $\Sigma_{h}^{\dagger}$ denotes the variance pseudo-inverse. The
+method was later proven equivalent to
+[`MinTrace`](https://Nixtla.github.io/hierarchicalforecast/src/methods.html#mintrace)
+variants.
+
+**Parameters:** `method`: str, allowed optimal combination methods:
+‘ols’, ‘wls_struct’. `nonnegative`: bool, reconciled forecasts
+should be nonnegative?
+
+**References:** - [Rob J. Hyndman, Roman A. Ahmed, George
+Athanasopoulos, Han Lin Shang (2010). “Optimal Combination Forecasts for
+Hierarchical Time
+Series”.](https://robjhyndman.com/papers/Hierarchical6.pdf). -
+[Shanika L. Wickramasuriya, George Athanasopoulos and Rob J. Hyndman
+(2010). “Optimal Combination Forecasts for Hierarchical Time
+Series”.](https://robjhyndman.com/papers/MinT.pdf). - [Wickramasuriya,
+S.L., Turlach, B.A. & Hyndman, R.J. (2020). “Optimal non-negative
+forecast reconciliation”. Stat Comput 30, 1167–1182,
+https://doi.org/10.1007/s11222-020-09930-0](https://robjhyndman.com/publications/nnmint/).\*
+
+------------------------------------------------------------------------
+
+source
+
+### OptimalCombination.fit
+
+> ``` text
+> OptimalCombination.fit (S, y_hat,
+> y_insample:Optional[numpy.ndarray]=None,
+> y_hat_insample:Optional[numpy.ndarray]=None,
+> sigmah:Optional[numpy.ndarray]=None,
+> intervals_method:Optional[str]=None,
+> num_samples:Optional[int]=None,
+> seed:Optional[int]=None,
+> tags:Optional[dict[str,numpy.ndarray]]=None,
+> idx_bottom:Optional[numpy.ndarray]=None)
+> ```
+
+\*MinTrace Fit Method.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `y_insample`:
+Insample values of size (`base`, `insample_size`). Only used with
+“wls_var”, “mint_cov”, “mint_shrink”. `y_hat_insample`: Insample
+forecast values of size (`base`, `insample_size`). Only used with
+“wls_var”, “mint_cov”, “mint_shrink” `sigmah`: Estimated standard
+deviation of the conditional marginal distribution.
+`intervals_method`: Sampler for prediction intervals, one of
+`normality`, `bootstrap`, `permbu`. `num_samples`: Number of samples
+for probabilistic coherent distribution. `seed`: Seed for
+reproducibility. `tags`: Each key is a level and each value its `S`
+indices. `idx_bottom`: Indices corresponding to the bottom level of
+`S`, size (`bottom`).
+
+**Returns:** `self`: object, fitted reconciler.\*
+
+------------------------------------------------------------------------
+
+source
+
+### OptimalCombination.predict
+
+> ``` text
+> OptimalCombination.predict (S:numpy.ndarray, y_hat:numpy.ndarray,
+> level:Optional[list[int]]=None)
+> ```
+
+\*Predict using reconciler.
+
+Predict using fitted mean and probabilistic reconcilers.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `level`: float
+list 0-100, confidence levels for prediction intervals.
+
+**Returns:** `y_tilde`: Reconciliated predictions.\*
+
+------------------------------------------------------------------------
+
+source
+
+### OptimalCombination.fit_predict
+
+> ``` text
+> OptimalCombination.fit_predict (S:numpy.ndarray, y_hat:numpy.ndarray,
+> idx_bottom:numpy.ndarray=None,
+> y_insample:Optional[numpy.ndarray]=None,
+> y_hat_insample:Optional[numpy.ndarray]=No
+> ne, sigmah:Optional[numpy.ndarray]=None,
+> level:Optional[list[int]]=None,
+> intervals_method:Optional[str]=None,
+> num_samples:Optional[int]=None,
+> seed:Optional[int]=None, tags:Optional[di
+> ct[str,numpy.ndarray]]=None)
+> ```
+
+\*MinTrace Reconciliation Method.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `idx_bottom`:
+Indices corresponding to the bottom level of `S`, size (`bottom`).
+`y_insample`: Insample values of size (`base`, `insample_size`). Only
+used by `wls_var`, `mint_cov`, `mint_shrink` `y_hat_insample`:
+Insample fitted values of size (`base`, `insample_size`). Only used by
+`wls_var`, `mint_cov`, `mint_shrink` `sigmah`: Estimated standard
+deviation of the conditional marginal distribution. `level`: float
+list 0-100, confidence levels for prediction intervals.
+`intervals_method`: Sampler for prediction intervals, one of
+`normality`, `bootstrap`, `permbu`. `num_samples`: Number of samples
+for probabilistic coherent distribution. `seed`: Seed for
+reproducibility. `tags`: Each key is a level and each value its `S`
+indices.
+
+**Returns:** `y_tilde`: Reconciliated y_hat using the MinTrace
+approach.\*
+
+------------------------------------------------------------------------
+
+source
+
+### OptimalCombination.sample
+
+> ``` text
+> OptimalCombination.sample (num_samples:int)
+> ```
+
+\*Sample probabilistic coherent distribution.
+
+Generates n samples from a probabilistic coherent distribution. The
+method uses fitted mean and probabilistic reconcilers, defined by the
+`intervals_method` selected during the reconciler’s instantiation.
+Currently available: `normality`, `bootstrap`, `permbu`.
+
+**Parameters:** `num_samples`: int, number of samples generated from
+coherent distribution.
+
+**Returns:** `samples`: Coherent samples of size (`num_series`,
+`horizon`, `num_samples`).\*
+
+# 6. Emp. Risk Minimization
+
+------------------------------------------------------------------------
+
+source
+
+### ERM
+
+> ``` text
+> ERM (method:str, lambda_reg:float=0.01)
+> ```
+
+\*Empirical Risk Minimization Reconciliation Class.
+
+The Empirical Risk Minimization reconciliation strategy relaxes the
+unbiasedness assumptions from previous reconciliation methods like MinT
+and optimizes square errors between the reconciled predictions and the
+validation data to obtain an optimal reconciliation matrix P.
+
+The exact solution for $\mathbf{P}$ (`method='closed'`) follows the
+expression:
+$$\mathbf{P}^{*} = \left(\mathbf{S}^{\intercal}\mathbf{S}\right)^{-1}\mathbf{Y}^{\intercal}\hat{\mathbf{Y}}\left(\hat{\mathbf{Y}}\hat{\mathbf{Y}}\right)^{-1}$$
+
+The alternative Lasso regularized $\mathbf{P}$ solution
+(`method='reg_bu'`) is useful when the observations of validation data
+is limited or the exact solution has low numerical stability.
+$$\mathbf{P}^{*} = \text{argmin}_{\mathbf{P}} ||\mathbf{Y}-\mathbf{S} \mathbf{P} \hat{Y} ||^{2}_{2} + \lambda ||\mathbf{P}-\mathbf{P}_{\text{BU}}||_{1}$$
+
+**Parameters:** `method`: str, one of `closed`, `reg` and
+`reg_bu`. `lambda_reg`: float, l1 regularizer for `reg` and
+`reg_bu`.
+
+**References:** - [Ben Taieb, S., & Koo, B. (2019). Regularized
+regression for hierarchical forecasting without unbiasedness conditions.
+In Proceedings of the 25th ACM SIGKDD International Conference on
+Knowledge Discovery & Data Mining KDD ’19 (p. 1337-1347). New York, NY,
+USA: Association for Computing
+Machinery.](https://doi.org/10.1145/3292500.3330976). \*
+
+------------------------------------------------------------------------
+
+source
+
+### ERM.fit
+
+> ``` text
+> ERM.fit (S, y_hat, y_insample, y_hat_insample,
+> sigmah:Optional[numpy.ndarray]=None,
+> intervals_method:Optional[str]=None,
+> num_samples:Optional[int]=None, seed:Optional[int]=None,
+> tags:Optional[dict[str,numpy.ndarray]]=None,
+> idx_bottom:Optional[numpy.ndarray]=None)
+> ```
+
+\*ERM Fit Method.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `y_insample`:
+Train values of size (`base`, `insample_size`). `y_hat_insample`:
+Insample train predictions of size (`base`, `insample_size`).
+`sigmah`: Estimated standard deviation of the conditional marginal
+distribution. `intervals_method`: Sampler for prediction intervals,
+one of `normality`, `bootstrap`, `permbu`. `num_samples`: Number of
+samples for probabilistic coherent distribution. `seed`: Seed for
+reproducibility. `tags`: Each key is a level and each value its `S`
+indices. `idx_bottom`: Indices corresponding to the bottom level of
+`S`, size (`bottom`).
+
+**Returns:** `self`: object, fitted reconciler.\*
+
+------------------------------------------------------------------------
+
+source
+
+### ERM.predict
+
+> ``` text
+> ERM.predict (S:numpy.ndarray, y_hat:numpy.ndarray,
+> level:Optional[list[int]]=None)
+> ```
+
+\*Predict using reconciler.
+
+Predict using fitted mean and probabilistic reconcilers.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `level`: float
+list 0-100, confidence levels for prediction intervals.
+
+**Returns:** `y_tilde`: Reconciliated predictions.\*
+
+------------------------------------------------------------------------
+
+source
+
+### ERM.fit_predict
+
+> ``` text
+> ERM.fit_predict (S:numpy.ndarray, y_hat:numpy.ndarray,
+> idx_bottom:numpy.ndarray=None,
+> y_insample:Optional[numpy.ndarray]=None,
+> y_hat_insample:Optional[numpy.ndarray]=None,
+> sigmah:Optional[numpy.ndarray]=None,
+> level:Optional[list[int]]=None,
+> intervals_method:Optional[str]=None,
+> num_samples:Optional[int]=None, seed:Optional[int]=None,
+> tags:Optional[dict[str,numpy.ndarray]]=None)
+> ```
+
+\*ERM Reconciliation Method.
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+`y_hat`: Forecast values of size (`base`, `horizon`). `idx_bottom`:
+Indices corresponding to the bottom level of `S`, size (`bottom`).
+`y_insample`: Train values of size (`base`, `insample_size`).
+`y_hat_insample`: Insample train predictions of size (`base`,
+`insample_size`). `sigmah`: Estimated standard deviation of the
+conditional marginal distribution. `level`: float list 0-100,
+confidence levels for prediction intervals. `intervals_method`:
+Sampler for prediction intervals, one of `normality`, `bootstrap`,
+`permbu`. `num_samples`: Number of samples for probabilistic
+coherent distribution. `seed`: Seed for reproducibility. `tags`:
+Each key is a level and each value its `S` indices.
+
+**Returns:** `y_tilde`: Reconciliated y_hat using the ERM
+approach.\*
+
+------------------------------------------------------------------------
+
+source
+
+### ERM.sample
+
+> ``` text
+> ERM.sample (num_samples:int)
+> ```
+
+\*Sample probabilistic coherent distribution.
+
+Generates n samples from a probabilistic coherent distribution. The
+method uses fitted mean and probabilistic reconcilers, defined by the
+`intervals_method` selected during the reconciler’s instantiation.
+Currently available: `normality`, `bootstrap`, `permbu`.
+
+**Parameters:** `num_samples`: int, number of samples generated from
+coherent distribution.
+
+**Returns:** `samples`: Coherent samples of size (`num_series`,
+`horizon`, `num_samples`).\*
+
+# References
+
+### General Reconciliation
+
+- [Orcutt, G.H., Watts, H.W., & Edwards, J.B.(1968). Data aggregation
+ and information loss. The American Economic Review, 58 ,
+ 773(787).](http://www.jstor.org/stable/1815532)
+- [Disaggregation methods to expedite product line forecasting.
+ Journal of Forecasting, 9 , 233–254.
+ doi:10.1002/for.3980090304](https://onlinelibrary.wiley.com/doi/abs/10.1002/for.3980090304).
+- [An investigation of aggregate variable time series forecast
+ strategies with specific subaggregate time series statistical
+ correlation. Computers and Operations Research, 26 , 1133–1149.
+ doi:10.1016/S0305-0548(99)00017-9.](https://doi.org/10.1016/S0305-0548(99)00017-9)
+- [Hyndman, R.J., & Athanasopoulos, G. (2021). “Forecasting:
+ principles and practice, 3rd edition: Chapter 11: Forecasting
+ hierarchical and grouped series.”. OTexts: Melbourne, Australia.
+ OTexts.com/fpp3 Accessed on July
+ 2022.](https://otexts.com/fpp3/hierarchical.html)
+
+### Optimal Reconciliation
+
+- [Rob J. Hyndman, Roman A. Ahmed, George Athanasopoulos, Han Lin
+ Shang. “Optimal Combination Forecasts for Hierarchical Time Series”
+ (2010).](https://robjhyndman.com/papers/Hierarchical6.pdf)
+- [Shanika L. Wickramasuriya, George Athanasopoulos and Rob J.
+ Hyndman. “Optimal Combination Forecasts for Hierarchical Time
+ Series” (2010).](https://robjhyndman.com/papers/MinT.pdf)
+- [Ben Taieb, S., & Koo, B. (2019). Regularized regression for
+ hierarchical forecasting without unbiasedness conditions. In
+ Proceedings of the 25th ACM SIGKDD International Conference on
+ Knowledge Discovery & Data Mining KDD ’19 (p. 1337-1347). New York,
+ NY, USA: Association for Computing
+ Machinery.](https://doi.org/10.1145/3292500.3330976)
+
+### Hierarchical Probabilistic Coherent Predictions
+
+- [Puwasala Gamakumara Ph. D. dissertation. Monash University,
+ Econometrics and Business Statistics. “Probabilistic Forecast
+ Reconciliation”.](https://bridges.monash.edu/articles/thesis/Probabilistic_Forecast_Reconciliation_Theory_and_Applications/11869533)
+- [Taieb, Souhaib Ben and Taylor, James W and Hyndman, Rob J. (2017).
+ Coherent probabilistic forecasts for hierarchical time series.
+ International conference on machine learning
+ ICML.](https://proceedings.mlr.press/v70/taieb17a.html)
+
diff --git a/hierarchicalforecast/src/probabilistic_methods.html.mdx b/hierarchicalforecast/src/probabilistic_methods.html.mdx
new file mode 100644
index 00000000..c2d6ea12
--- /dev/null
+++ b/hierarchicalforecast/src/probabilistic_methods.html.mdx
@@ -0,0 +1,261 @@
+---
+output-file: probabilistic_methods.html
+title: Probabilistic Methods
+---
+
+
+Here we provide a collection of methods designed to provide
+hierarchically coherent probabilistic distributions, which means that
+they generate samples of multivariate time series with hierarchical
+linear constraints.
+
+We designed these methods to extend the `core.HierarchicalForecast`
+capabilities class. Check their [usage example
+here](https://nixtlaverse.nixtla.io/hierarchicalforecast/examples/introduction.html).
+
+# 1. Normality
+
+------------------------------------------------------------------------
+
+source
+
+### Normality
+
+> ``` text
+> Normality (S:Union[numpy.ndarray,scipy.sparse._matrix.spmatrix],
+> P:Union[numpy.ndarray,scipy.sparse._matrix.spmatrix],
+> y_hat:numpy.ndarray, sigmah:numpy.ndarray,
+> W:Union[numpy.ndarray,scipy.sparse._matrix.spmatrix],
+> seed:int=0)
+> ```
+
+\*Normality Probabilistic Reconciliation Class.
+
+The Normality method leverages the Gaussian Distribution linearity, to
+generate hierarchically coherent prediction distributions. This class is
+meant to be used as the `sampler` input as other `HierarchicalForecast`
+[reconciliation
+classes](https://nixtla.github.io/hierarchicalforecast/src/methods.html).
+
+Given base forecasts under a normal distribution:
+$$\hat{y}_{h} \sim \mathrm{N}(\hat{\boldsymbol{\mu}}, \hat{\mathbf{W}}_{h})$$
+
+The reconciled forecasts are also normally distributed:
+
+$$
+
+\tilde{y}_{h} \sim \mathrm{N}(\mathbf{S}\mathbf{P}\hat{\boldsymbol{\mu}},
+\mathbf{S}\mathbf{P}\hat{\mathbf{W}}_{h} \mathbf{P}^{\intercal} \mathbf{S}^{\intercal})
+
+$$
+
+**Parameters:** `S`: np.array, summing matrix of size (`base`,
+`bottom`). `P`: np.array, reconciliation matrix of size (`bottom`,
+`base`). `y_hat`: Point forecasts values of size (`base`,
+`horizon`). `W`: np.array, hierarchical covariance matrix of size
+(`base`, `base`). `sigmah`: np.array, forecast standard dev. of size
+(`base`, `horizon`). `num_samples`: int, number of bootstraped
+samples generated. `seed`: int, random seed for numpy generator’s
+replicability.
+
+**References:** - [Panagiotelis A., Gamakumara P. Athanasopoulos G.,
+and Hyndman R. J. (2022). “Probabilistic forecast reconciliation:
+Properties, evaluation and score optimisation”. European Journal of
+Operational
+Research.](https://www.sciencedirect.com/science/article/pii/S0377221722006087)\*
+
+------------------------------------------------------------------------
+
+source
+
+### Normality.get_samples
+
+> ``` text
+> Normality.get_samples (num_samples:int)
+> ```
+
+\*Normality Coherent Samples.
+
+Obtains coherent samples under the Normality assumptions.
+
+**Parameters:** `num_samples`: int, number of samples generated from
+coherent distribution.
+
+**Returns:** `samples`: Coherent samples of size (`base`, `horizon`,
+`num_samples`).\*
+
+# 2. Bootstrap
+
+------------------------------------------------------------------------
+
+source
+
+### Bootstrap
+
+> ``` text
+> Bootstrap (S:Union[numpy.ndarray,scipy.sparse._matrix.spmatrix],
+> P:Union[numpy.ndarray,scipy.sparse._matrix.spmatrix],
+> y_hat:numpy.ndarray, y_insample:numpy.ndarray,
+> y_hat_insample:numpy.ndarray, num_samples:int=100, seed:int=0,
+> W:Union[numpy.ndarray,scipy.sparse._matrix.spmatrix]=None)
+> ```
+
+\*Bootstrap Probabilistic Reconciliation Class.
+
+This method goes beyond the normality assumption for the base forecasts,
+the technique simulates future sample paths and uses them to generate
+base sample paths that are latered reconciled. This clever idea and its
+simplicity allows to generate coherent bootstraped prediction intervals
+for any reconciliation strategy. This class is meant to be used as the
+`sampler` input as other `HierarchicalForecast` [reconciliation
+classes](https://nixtla.github.io/hierarchicalforecast/src/methods.html).
+
+Given a boostraped set of simulated sample paths:
+$$(\hat{\mathbf{y}}^{[1]}_{\tau}, \dots ,\hat{\mathbf{y}}^{[B]}_{\tau})$$
+
+The reconciled sample paths allow for reconciled distributional
+forecasts:
+$$(\mathbf{S}\mathbf{P}\hat{\mathbf{y}}^{[1]}_{\tau}, \dots ,\mathbf{S}\mathbf{P}\hat{\mathbf{y}}^{[B]}_{\tau})$$
+
+**Parameters:** `S`: np.array, summing matrix of size (`base`,
+`bottom`). `P`: np.array, reconciliation matrix of size (`bottom`,
+`base`). `y_hat`: Point forecasts values of size (`base`,
+`horizon`). `y_insample`: Insample values of size (`base`,
+`insample_size`). `y_hat_insample`: Insample point forecasts of size
+(`base`, `insample_size`). `num_samples`: int, number of bootstraped
+samples generated. `seed`: int, random seed for numpy generator’s
+replicability.
+
+**References:** - [Puwasala Gamakumara Ph. D. dissertation. Monash
+University, Econometrics and Business Statistics (2020). “Probabilistic
+Forecast
+Reconciliation”](https://bridges.monash.edu/articles/thesis/Probabilistic_Forecast_Reconciliation_Theory_and_Applications/11869533) -
+[Panagiotelis A., Gamakumara P. Athanasopoulos G., and Hyndman R. J.
+(2022). “Probabilistic forecast reconciliation: Properties, evaluation
+and score optimisation”. European Journal of Operational
+Research.](https://www.sciencedirect.com/science/article/pii/S0377221722006087)\*
+
+------------------------------------------------------------------------
+
+source
+
+### Bootstrap.get_samples
+
+> ``` text
+> Bootstrap.get_samples (num_samples:int)
+> ```
+
+\*Bootstrap Sample Reconciliation Method.
+
+Applies Bootstrap sample reconciliation method as defined by Gamakumara
+2020. Generating independent sample paths and reconciling them with
+Bootstrap.
+
+**Parameters:** `num_samples`: int, number of samples generated from
+coherent distribution.
+
+**Returns:** `samples`: Coherent samples of size (`base`, `horizon`,
+`num_samples`).\*
+
+# 3. PERMBU
+
+------------------------------------------------------------------------
+
+source
+
+### PERMBU
+
+> ``` text
+> PERMBU (S:Union[numpy.ndarray,scipy.sparse._matrix.spmatrix],
+> tags:dict[str,numpy.ndarray], y_hat:numpy.ndarray,
+> y_insample:numpy.ndarray, y_hat_insample:numpy.ndarray,
+> sigmah:numpy.ndarray, num_samples:Optional[int]=None, seed:int=0,
+> P:Union[numpy.ndarray,scipy.sparse._matrix.spmatrix]=None)
+> ```
+
+\*PERMBU Probabilistic Reconciliation Class.
+
+The PERMBU method leverages empirical bottom-level marginal
+distributions with empirical copula functions (describing bottom-level
+dependencies) to generate the distribution of aggregate-level
+distributions using BottomUp reconciliation. The sample reordering
+technique in the PERMBU method reinjects multivariate dependencies into
+independent bottom-level samples.
+
+``` text
+Algorithm:
+1. For all series compute conditional marginals distributions.
+2. Compute residuals $\hat{\epsilon}_{i,t}$ and obtain rank permutations.
+2. Obtain K-sample from the bottom-level series predictions.
+3. Apply recursively through the hierarchical structure:
+ 3.1. For a given aggregate series $i$ and its children series:
+ 3.2. Obtain children's empirical joint using sample reordering copula.
+ 3.2. From the children's joint obtain the aggregate series's samples.
+```
+
+**Parameters:** `S`: np.array, summing matrix of size (`base`,
+`bottom`). `tags`: Each key is a level and each value its `S`
+indices. `y_insample`: Insample values of size (`base`,
+`insample_size`). `y_hat_insample`: Insample point forecasts of size
+(`base`, `insample_size`). `sigmah`: np.array, forecast standard
+dev. of size (`base`, `horizon`). `num_samples`: int, number of
+normal prediction samples generated. `seed`: int, random seed for
+numpy generator’s replicability.
+
+**References:** - [Taieb, Souhaib Ben and Taylor, James W and
+Hyndman, Rob J. (2017). Coherent probabilistic forecasts for
+hierarchical time series. International conference on machine learning
+ICML.](https://proceedings.mlr.press/v70/taieb17a.html)\*
+
+------------------------------------------------------------------------
+
+source
+
+### PERMBU.get_samples
+
+> ``` text
+> PERMBU.get_samples (num_samples:Optional[int]=None)
+> ```
+
+\*PERMBU Sample Reconciliation Method.
+
+Applies PERMBU reconciliation method as defined by Taieb et. al 2017.
+Generating independent base prediction samples, restoring its
+multivariate dependence using estimated copula with reordering and
+applying the BottomUp aggregation to the new samples.
+
+**Parameters:** `num_samples`: int, number of samples generated from
+coherent distribution.
+
+**Returns:** `samples`: Coherent samples of size (`base`, `horizon`,
+`num_samples`).\*
+
+# References
+
+- [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ principles and practice, Reconciled distributional
+ forecasts”.](https://otexts.com/fpp3/rec-prob.html)
+- [Puwasala Gamakumara Ph. D. dissertation. Monash University,
+ Econometrics and Business Statistics (2020). “Probabilistic Forecast
+ Reconciliation”](https://bridges.monash.edu/articles/thesis/Probabilistic_Forecast_Reconciliation_Theory_and_Applications/11869533)
+- [Panagiotelis A., Gamakumara P. Athanasopoulos G., and Hyndman R. J.
+ (2022). “Probabilistic forecast reconciliation: Properties,
+ evaluation and score optimisation”. European Journal of Operational
+ Research.](https://www.sciencedirect.com/science/article/pii/S0377221722006087)
+- [Taieb, Souhaib Ben and Taylor, James W and Hyndman, Rob J. (2017).
+ Coherent probabilistic forecasts for hierarchical time series.
+ International conference on machine learning
+ ICML.](https://proceedings.mlr.press/v70/taieb17a.html)
+
diff --git a/hierarchicalforecast/src/utils.html.mdx b/hierarchicalforecast/src/utils.html.mdx
new file mode 100644
index 00000000..fcf23fc1
--- /dev/null
+++ b/hierarchicalforecast/src/utils.html.mdx
@@ -0,0 +1,403 @@
+---
+output-file: utils.html
+title: Aggregation/Visualization Utils
+---
+
+
+The `HierarchicalForecast` package contains utility functions to wrangle
+and visualize hierarchical series datasets. The
+[`aggregate`](https://Nixtla.github.io/hierarchicalforecast/src/utils.html#aggregate)
+function of the module allows you to create a hierarchy from categorical
+variables representing the structure levels, returning also the
+aggregation contraints matrix $\mathbf{S}$.
+
+In addition, `HierarchicalForecast` ensures compatibility of its
+reconciliation methods with other popular machine-learning libraries via
+its external forecast adapters that transform output base forecasts from
+external libraries into a compatible data frame format.
+
+# Aggregate Function
+
+------------------------------------------------------------------------
+
+source
+
+### aggregate
+
+> ``` text
+> aggregate
+> (df:Union[ForwardRef('DataFrame[Any]'),ForwardRef('LazyFrame[A
+> ny]')], spec:list[list[str]],
+> exog_vars:Optional[dict[str,Union[str,list[str]]]]=None,
+> sparse_s:bool=False, id_col:str='unique_id',
+> time_col:str='ds', id_time_col:Optional[str]=None,
+> target_cols:collections.abc.Sequence[str]=('y',))
+> ```
+
+*Utils Aggregation Function. Aggregates bottom level series contained in
+the DataFrame `df` according to levels defined in the `spec` list.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | Union | | Dataframe with columns `[time_col, *target_cols]`, columns to aggregate and optionally exog_vars. |
+| spec | list | | list of levels. Each element of the list should contain a list of columns of `df` to aggregate. |
+| exog_vars | Optional | None | |
+| sparse_s | bool | False | Return `S_df` as a sparse Pandas dataframe. |
+| id_col | str | unique_id | Column that will identify each serie after aggregation. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| id_time_col | Optional | None | Column that will identify each timestep after temporal aggregation. If provided, aggregate will operate temporally. |
+| target_cols | Sequence | (‘y’,) | list of columns that contains the targets to aggregate. |
+| **Returns** | **tuple** | | **Hierarchically structured series.** |
+
+------------------------------------------------------------------------
+
+source
+
+### aggregate_temporal
+
+> ``` text
+> aggregate_temporal
+> (df:Union[ForwardRef('DataFrame[Any]'),ForwardRef('La
+> zyFrame[Any]')], spec:dict[str,int], exog_vars:Option
+> al[dict[str,Union[str,list[str]]]]=None,
+> sparse_s:bool=False, id_col:str='unique_id',
+> time_col:str='ds', id_time_col:str='temporal_id',
+> target_cols:collections.abc.Sequence[str]=('y',),
+> aggregation_type:str='local')
+> ```
+
+*Utils Aggregation Function for Temporal aggregations. Aggregates bottom
+level timesteps contained in the DataFrame `df` according to temporal
+levels defined in the `spec` list.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | Union | | Dataframe with columns `[time_col, target_cols]` and columns to aggregate. |
+| spec | dict | | Dictionary of temporal levels. Each key should be a string with the value representing the number of bottom-level timesteps contained in the aggregation. |
+| exog_vars | Optional | None | |
+| sparse_s | bool | False | Return `S_df` as a sparse Pandas dataframe. |
+| id_col | str | unique_id | Column that will identify each serie after aggregation. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| id_time_col | str | temporal_id | Column that will identify each timestep after aggregation. |
+| target_cols | Sequence | (‘y’,) | List of columns that contain the targets to aggregate. |
+| aggregation_type | str | local | If ‘local’ the aggregation will be performed on the timestamps of each timeseries independently. If ‘global’ the aggregation will be performed on the unique timestamps of all timeseries. |
+| **Returns** | **tuple** | | **Temporally hierarchically structured series.** |
+
+------------------------------------------------------------------------
+
+source
+
+### make_future_dataframe
+
+> ``` text
+> make_future_dataframe
+> (df:Union[ForwardRef('DataFrame[Any]'),ForwardRef(
+> 'LazyFrame[Any]')], freq:Union[str,int], h:int,
+> id_col:str='unique_id', time_col:str='ds')
+> ```
+
+*Create future dataframe for forecasting.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | Union | | Dataframe with ids, times and values for the exogenous regressors. |
+| freq | Union | | Frequency of the data. Must be a valid pandas or polars offset alias, or an integer. |
+| h | int | | Forecast horizon. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| **Returns** | **FrameT** | | **DataFrame with future values** |
+
+------------------------------------------------------------------------
+
+source
+
+### get_cross_temporal_tags
+
+> ``` text
+> get_cross_temporal_tags
+> (df:Union[ForwardRef('DataFrame[Any]'),ForwardRe
+> f('LazyFrame[Any]')],
+> tags_cs:dict[str,numpy.ndarray],
+> tags_te:dict[str,numpy.ndarray], sep:str='//',
+> id_col:str='unique_id',
+> id_time_col:str='temporal_id',
+> cross_temporal_id_col:str='cross_temporal_id')
+> ```
+
+*Get cross-temporal tags.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | Union | | DataFrame with temporal ids. |
+| tags_cs | dict | | Tags for the cross-sectional hierarchies |
+| tags_te | dict | | Tags for the temporal hierarchies |
+| sep | str | // | Separator for the cross-temporal tags. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| id_time_col | str | temporal_id | Column that identifies each (aggregated) timestep. |
+| cross_temporal_id_col | str | cross_temporal_id | Column that will identify each cross-temporal aggregation. |
+| **Returns** | **tuple** | | **DataFrame with cross-temporal ids.** |
+
+# Hierarchical Visualization
+
+------------------------------------------------------------------------
+
+source
+
+### HierarchicalPlot
+
+> ``` text
+> HierarchicalPlot
+> (S:Union[ForwardRef('DataFrame[Any]'),ForwardRef('LazyF
+> rame[Any]')], tags:dict[str,numpy.ndarray],
+> S_id_col:str='unique_id')
+> ```
+
+\*Hierarchical Plot
+
+This class contains a collection of matplotlib visualization methods,
+suited for small to medium sized hierarchical series.
+
+**Parameters:** `S`: DataFrame with summing matrix of size
+`(base, bottom)`, see [aggregate
+function](https://nixtlaverse.nixtla.io/hierarchicalforecast/src/utils.html#aggregate).
+`tags`: np.ndarray, with hierarchical aggregation indexes, where each
+key is a level and its value contains tags associated to that level.
+`S_id_col` : str=‘unique_id’, column that identifies each
+aggregation. \*
+
+------------------------------------------------------------------------
+
+source
+
+### plot_summing_matrix
+
+> ``` text
+> plot_summing_matrix ()
+> ```
+
+\*Summation Constraints plot
+
+This method simply plots the hierarchical aggregation constraints matrix
+$\mathbf{S}$.
+
+**Returns:** `fig`: matplotlib.figure.Figure, figure object
+containing the plot of the summing matrix.\*
+
+------------------------------------------------------------------------
+
+source
+
+### plot_series
+
+> ``` text
+> plot_series (series:str,
+> Y_df:Union[ForwardRef('DataFrame[Any]'),ForwardRef('LazyFram
+> e[Any]')], models:Optional[list[str]]=None,
+> level:Optional[list[int]]=None, id_col:str='unique_id',
+> time_col:str='ds', target_col:str='y')
+> ```
+
+\*Single Series plot
+
+**Parameters:** `series`: str, string identifying the `'unique_id'`
+any-level series to plot. `Y_df`: DataFrame, hierarchically
+structured series ($\mathbf{y}_{[a,b]}$). It contains columns
+`['unique_id', 'ds', 'y']`, it may have `'models'`. `models`:
+list\[str\], string identifying filtering model columns. `level`:
+float list 0-100, confidence levels for prediction intervals available
+in `Y_df`. `id_col` : str=‘unique_id’, column that identifies each
+serie. `time_col` : str=‘ds’, column that identifies each timestep,
+its values can be timestamps or integers. `target_col` : str=‘y’,
+column that contains the target.
+
+**Returns:** `fig`: matplotlib.figure.Figure, figure object
+containing the plot of the single series.\*
+
+------------------------------------------------------------------------
+
+source
+
+### plot_hierarchically_linked_series
+
+> ``` text
+> plot_hierarchically_linked_series (bottom_series:str,
+> Y_df:Union[ForwardRef('DataFrame[Any]'
+> ),ForwardRef('LazyFrame[Any]')],
+> models:Optional[list[str]]=None,
+> level:Optional[list[int]]=None,
+> id_col:str='unique_id',
+> time_col:str='ds', target_col:str='y')
+> ```
+
+\*Hierarchically Linked Series plot
+
+**Parameters:** `bottom_series`: str, string identifying the
+`'unique_id'` bottom-level series to plot. `Y_df`: DataFrame,
+hierarchically structured series ($\mathbf{y}_{[a,b]}$). It contains
+columns \[‘unique_id’, ‘ds’, ‘y’\] and models. `models`:
+list\[str\], string identifying filtering model columns. `level`:
+float list 0-100, confidence levels for prediction intervals available
+in `Y_df`. `id_col` : str=‘unique_id’, column that identifies each
+serie. `time_col` : str=‘ds’, column that identifies each timestep,
+its values can be timestamps or integers. `target_col` : str=‘y’,
+column that contains the target.
+
+**Returns:** `fig`: matplotlib.figure.Figure, figure object
+containing the plots of the hierarchilly linked series.\*
+
+------------------------------------------------------------------------
+
+source
+
+### plot_hierarchical_predictions_gap
+
+> ``` text
+> plot_hierarchical_predictions_gap
+> (Y_df:Union[ForwardRef('DataFrame[Any]
+> '),ForwardRef('LazyFrame[Any]')],
+> models:Optional[list[str]]=None,
+> xlabel:Optional[str]=None,
+> ylabel:Optional[str]=None,
+> id_col:str='unique_id',
+> time_col:str='ds', target_col:str='y')
+> ```
+
+\*Hierarchically Predictions Gap plot
+
+**Parameters:** `Y_df`: DataFrame, hierarchically structured series
+($\mathbf{y}_{[a,b]}$). It contains columns \[‘unique_id’, ‘ds’, ‘y’\]
+and models. `models`: list\[str\], string identifying filtering
+model columns. `xlabel`: str, string for the plot’s x axis
+label. `ylabel`: str, string for the plot’s y axis label.
+`id_col` : str=‘unique_id’, column that identifies each serie.
+`time_col` : str=‘ds’, column that identifies each timestep, its values
+can be timestamps or integers. `target_col` : str=‘y’, column that
+contains the target.
+
+**Returns:** `fig`: matplotlib.figure.Figure, figure object
+containing the plot of the aggregated predictions at different levels of
+the hierarchical structure.\*
+
+
+```python
+from statsforecast.core import StatsForecast
+from statsforecast.models import AutoETS
+from datasetsforecast.hierarchical import HierarchicalData
+
+Y_df, S, tags = HierarchicalData.load('./data', 'Labour')
+Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+S = S.reset_index(names="unique_id")
+
+Y_test_df = Y_df.groupby('unique_id').tail(24)
+Y_train_df = Y_df.drop(Y_test_df.index)
+
+fcst = StatsForecast(
+ models=[AutoETS(season_length=12, model='AAZ')],
+ freq='MS',
+ n_jobs=-1
+)
+Y_hat_df = fcst.forecast(df=Y_train_df, h=24).reset_index()
+
+# Plot prediction difference of different aggregation
+# Levels Country, Country/Region, Country/Gender/Region ...
+hplots = HierarchicalPlot(S=S, tags=tags)
+
+hplots.plot_hierarchical_predictions_gap(
+ Y_df=Y_hat_df, models='AutoETS',
+ xlabel='Month', ylabel='Predictions',
+)
+```
+
+
+```python
+# polars
+from statsforecast.core import StatsForecast
+from statsforecast.models import AutoETS
+from datasetsforecast.hierarchical import HierarchicalData
+
+Y_df, S, tags = HierarchicalData.load('./data', 'Labour')
+Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+S = S.reset_index(names="unique_id")
+
+Y_test_df = Y_df.groupby('unique_id').tail(24)
+Y_train_df = Y_df.drop(Y_test_df.index)
+Y_test_df_pl = pl.from_pandas(Y_test_df)
+Y_train_df_pl = pl.from_pandas(Y_train_df)
+
+fcst = StatsForecast(
+ models=[AutoETS(season_length=12, model='AAZ')],
+ freq='1m',
+ n_jobs=-1
+)
+Y_hat_df = fcst.forecast(df=Y_train_df_pl, h=24)
+
+# Plot prediction difference of different aggregation
+# Levels Country, Country/Region, Country/Gender/Region ...
+hplots = HierarchicalPlot(S=S, tags=tags)
+
+hplots.plot_hierarchical_predictions_gap(
+ Y_df=Y_hat_df, models='AutoETS',
+ xlabel='Month', ylabel='Predictions',
+)
+```
+
+# External Forecast Adapters
+
+------------------------------------------------------------------------
+
+source
+
+### samples_to_quantiles_df
+
+> ``` text
+> samples_to_quantiles_df (samples:numpy.ndarray,
+> unique_ids:collections.abc.Sequence[str],
+> dates:list[str],
+> quantiles:Optional[list[float]]=None,
+> level:Optional[list[int]]=None,
+> model_name:str='model', id_col:str='unique_id',
+> time_col:str='ds', backend:str='pandas')
+> ```
+
+\*Transform Random Samples into HierarchicalForecast input. Auxiliary
+function to create compatible HierarchicalForecast input `Y_hat_df`
+dataframe.
+
+**Parameters:** `samples`: numpy array. Samples from forecast
+distribution of shape \[n_series, n_samples, horizon\].
+`unique_ids`: string list. Unique identifiers for each time series.
+`dates`: datetime list. list of forecast dates. `quantiles`: float
+list in \[0., 1.\]. Alternative to level, quantiles to estimate from y
+distribution. `level`: int list in \[0,100\]. Probability levels for
+prediction intervals. `model_name`: string. Name of forecasting
+model. `id_col` : str=‘unique_id’, column that identifies each
+serie. `time_col` : str=‘ds’, column that identifies each timestep,
+its values can be timestamps or integers. `backend` : str=‘pandas’,
+backend to use for the output dataframe, either ‘pandas’ or
+‘polars’.
+
+**Returns:** `quantiles`: float list in \[0., 1.\]. quantiles to
+estimate from y distribution . `Y_hat_df`: DataFrame. With base
+quantile forecasts with columns ds and models to reconcile indexed by
+unique_id.\*
+
diff --git a/index.mdx b/index.mdx
index dde2a2be..d41698c5 100644
--- a/index.mdx
+++ b/index.mdx
@@ -4,19 +4,70 @@ title: "Nixtlaverse"
The Nixtlaverse is composed by our open-source libraries, designed to provide a comprehensive, cutting-edge toolkit for time series forecasting. The Nixtla ecosystem is primarily built around five main libraries, each specializing in different aspects of time series forecasting:
-
+
+ Checkout our Enterprise offering for time series forecasting and anomaly detection!
+
-## StatsForecast
-Designed for high-speed forecasting, StatsForecast leverages statistical and econometric models. This library provides a set of robust algorithms and techniques that allow for quick, yet accurate predictions, making it an excellent choice for applications requiring rapid forecasting.
+
+
+ Lightning fast forecasting with statistical and econometric models.
+
+
+ Scalable machine learning for time series forecasting.
+
+
-## MLForecast
-MLForecast makes machine learning scalable for time series forecasting. This library is tailored for applications that require the processing of vast amounts of time-series data, implementing machine learning techniques to optimize accuracy, efficiency, and scalability.
+
+
+ Scalable and user friendly neural forecasting algorithms for time series data.
+
+
+ Probabilistic Hierarchical forecasting with statistical and econometric methods.
+
+
-## NeuralForecast
-NeuralForecast offers scalable and user-friendly neural forecasting algorithms for time series data. By combining the power of deep learning with the complexity of time series forecasting, this library makes advanced neural forecasting techniques accessible and easy to implement, even for non-experts.
+
-## HierarchicalForecast
-Hierarchical Forecast focuses on probabilistic hierarchical forecasting using statistical and econometric methods. It provides tools to model and forecast hierarchical or grouped time series data, capturing the underlying correlations and shared information between different levels of hierarchy.
+
+ Datasets for time series forecasting.
+
+
+ Forecasting utilities for plotting and robust evaluation.
+
+
+ Fast implementations of common forecasting routines.
+
+
-## TS Features
-TS Features is a Python implementation of the R package *tsfeatures*. It calculates various features from time series data, helping users to understand and extract meaningful information from their datasets. By identifying key characteristics and patterns, this library supports more informed and effective forecasting.
diff --git a/mint.json b/mint.json
deleted file mode 100644
index c413dae0..00000000
--- a/mint.json
+++ /dev/null
@@ -1,70 +0,0 @@
-{
- "$schema": "https://mintlify.com/schema.json",
- "name": "Nixtla",
- "logo": {
- "light": "/light.png",
- "dark": "/dark.png"
- },
- "favicon": "/favicon.svg",
- "colors": {
- "primary": "#0E0E0E",
- "light": "#FAFAFA",
- "dark": "#0E0E0E",
- "anchors": {
- "from": "#2AD0CA",
- "to": "#0E00F8"
- }
- },
- "topbarCtaButton": {
- "type": "github",
- "url": "https://github.com/Nixtla"
- },
- "anchors": [
- {
- "name": "TimeGPT",
- "url": "nixtla",
- "icon": "clock-nine"
- },
- {
- "name": "StatsForecast",
- "url": "statsforecast",
- "icon": "bolt"
- },
- {
- "name": "MLForecast",
- "url": "mlforecast",
- "icon": "robot"
- },
- {
- "name": "NeuralForecast",
- "url": "neuralforecast",
- "icon": "brain-circuit"
- },
- {
- "name": "HierarchicalForecast",
- "url": "hierarchicalforecast",
- "icon": "crown"
- },
- {
- "name": "UtilsForecast",
- "url": "utilsforecast",
- "icon": "wrench"
- },
- {
- "name": "DatasetsForecast",
- "url": "datasetsforecast",
- "icon": "chart-simple"
- },
- {
- "name": "CoreForecast",
- "url": "coreforecast",
- "icon": "truck-fast"
- }
- ],
- "navigation": [],
- "footerSocials": {
- "github": "https://github.com/Nixtla",
- "slack": "https://join.slack.com/t/nixtlaworkspace/shared_invite/zt-135dssye9-fWTzMpv2WBthq8NK0Yvu6A",
- "twitter": "https://twitter.com/nixtlainc"
- }
-}
diff --git a/mlforecast/.nojekyll b/mlforecast/.nojekyll
new file mode 100644
index 00000000..e69de29b
diff --git a/mlforecast/auto.html.mdx b/mlforecast/auto.html.mdx
new file mode 100644
index 00000000..5d77bc15
--- /dev/null
+++ b/mlforecast/auto.html.mdx
@@ -0,0 +1,571 @@
+---
+output-file: auto.html
+title: Auto
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### AutoRandomForest
+
+> ``` text
+> AutoRandomForest (config:Optional[Callable[[optuna.trial._trial.Trial],Di
+> ct[str,Any]]]=None)
+> ```
+
+*Structure to hold a model and its search space*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| config | Optional | None | function that takes an optuna trial and produces a configuration |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoElasticNet
+
+> ``` text
+> AutoElasticNet (config:Optional[Callable[[optuna.trial._trial.Trial],Dict
+> [str,Any]]]=None)
+> ```
+
+*Structure to hold a model and its search space*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| config | Optional | None | function that takes an optuna trial and produces a configuration |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoLasso
+
+> ``` text
+> AutoLasso (config:Optional[Callable[[optuna.trial._trial.Trial],Dict[str,
+> Any]]]=None)
+> ```
+
+*Structure to hold a model and its search space*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| config | Optional | None | function that takes an optuna trial and produces a configuration |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoRidge
+
+> ``` text
+> AutoRidge (config:Optional[Callable[[optuna.trial._trial.Trial],Dict[str,
+> Any]]]=None)
+> ```
+
+*Structure to hold a model and its search space*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| config | Optional | None | function that takes an optuna trial and produces a configuration |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoLinearRegression
+
+> ``` text
+> AutoLinearRegression (config:Optional[Callable[[optuna.trial._trial.Trial
+> ],Dict[str,Any]]]=None)
+> ```
+
+*Structure to hold a model and its search space*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| config | Optional | None | function that takes an optuna trial and produces a configuration |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoCatboost
+
+> ``` text
+> AutoCatboost (config:Optional[Callable[[optuna.trial._trial.Trial],Dict[s
+> tr,Any]]]=None)
+> ```
+
+*Structure to hold a model and its search space*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| config | Optional | None | function that takes an optuna trial and produces a configuration |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoXGBoost
+
+> ``` text
+> AutoXGBoost (config:Optional[Callable[[optuna.trial._trial.Trial],Dict[st
+> r,Any]]]=None)
+> ```
+
+*Structure to hold a model and its search space*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| config | Optional | None | function that takes an optuna trial and produces a configuration |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoLightGBM
+
+> ``` text
+> AutoLightGBM (config:Optional[Callable[[optuna.trial._trial.Trial],Dict[s
+> tr,Any]]]=None)
+> ```
+
+*Structure to hold a model and its search space*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| config | Optional | None | function that takes an optuna trial and produces a configuration |
+
+------------------------------------------------------------------------
+
+source
+
+### random_forest_space
+
+> ``` text
+> random_forest_space (trial:optuna.trial._trial.Trial)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### elastic_net_space
+
+> ``` text
+> elastic_net_space (trial:optuna.trial._trial.Trial)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### lasso_space
+
+> ``` text
+> lasso_space (trial:optuna.trial._trial.Trial)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### ridge_space
+
+> ``` text
+> ridge_space (trial:optuna.trial._trial.Trial)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### linear_regression_space
+
+> ``` text
+> linear_regression_space (trial:optuna.trial._trial.Trial)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### catboost_space
+
+> ``` text
+> catboost_space (trial:optuna.trial._trial.Trial)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### xgboost_space
+
+> ``` text
+> xgboost_space (trial:optuna.trial._trial.Trial)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### lightgbm_space
+
+> ``` text
+> lightgbm_space (trial:optuna.trial._trial.Trial)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoModel
+
+> ``` text
+> AutoModel (model:sklearn.base.BaseEstimator,
+> config:Callable[[optuna.trial._trial.Trial],Dict[str,Any]])
+> ```
+
+*Structure to hold a model and its search space*
+
+| | **Type** | **Details** |
+|--------|---------------------------|-------------------------------------|
+| model | BaseEstimator | scikit-learn compatible regressor |
+| config | Callable | function that takes an optuna trial and produces a configuration |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoMLForecast
+
+> ``` text
+> AutoMLForecast
+> (models:Union[List[__main__.AutoModel],Dict[str,__main__.
+> AutoModel]], freq:Union[int,str],
+> season_length:Optional[int]=None, init_config:Optional[Ca
+> llable[[optuna.trial._trial.Trial],Dict[str,Any]]]=None,
+> fit_config:Optional[Callable[[optuna.trial._trial.Trial],
+> Dict[str,Any]]]=None, num_threads:int=1)
+> ```
+
+*Hyperparameter optimization helper*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| models | Union | | Auto models to be optimized. |
+| freq | Union | | pandas’ or polars’ offset alias or integer denoting the frequency of the series. |
+| season_length | Optional | None | Length of the seasonal period. This is used for producing the feature space. Only required if `init_config` is None. |
+| init_config | Optional | None | Function that takes an optuna trial and produces a configuration passed to the MLForecast constructor. |
+| fit_config | Optional | None | Function that takes an optuna trial and produces a configuration passed to the MLForecast fit method. |
+| num_threads | int | 1 | Number of threads to use when computing the features. |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoMLForecast.fit
+
+> ``` text
+> AutoMLForecast.fit
+> (df:Union[pandas.core.frame.DataFrame,polars.datafram
+> e.frame.DataFrame], n_windows:int, h:int,
+> num_samples:int, step_size:Optional[int]=None,
+> input_size:Optional[int]=None,
+> refit:Union[bool,int]=False, loss:Optional[Callable[[
+> Union[pandas.core.frame.DataFrame,polars.dataframe.fr
+> ame.DataFrame],Union[pandas.core.frame.DataFrame,pola
+> rs.dataframe.frame.DataFrame]],float]]=None,
+> id_col:str='unique_id', time_col:str='ds',
+> target_col:str='y',
+> study_kwargs:Optional[Dict[str,Any]]=None,
+> optimize_kwargs:Optional[Dict[str,Any]]=None,
+> fitted:bool=False, prediction_intervals:Optional[mlfo
+> recast.utils.PredictionIntervals]=None)
+> ```
+
+*Carry out the optimization process. Each model is optimized
+independently and the best one is trained on all data*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | Union | | Series data in long format. |
+| n_windows | int | | Number of windows to evaluate. |
+| h | int | | Forecast horizon. |
+| num_samples | int | | Number of trials to run |
+| step_size | Optional | None | Step size between each cross validation window. If None it will be equal to `h`. |
+| input_size | Optional | None | Maximum training samples per serie in each window. If None, will use an expanding window. |
+| refit | Union | False | Retrain model for each cross validation window. If False, the models are trained at the beginning and then used to predict each window. If positive int, the models are retrained every `refit` windows. |
+| loss | Optional | None | Function that takes the validation and train dataframes and produces a float. If `None` will use the average SMAPE across series. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| study_kwargs | Optional | None | Keyword arguments to be passed to the optuna.Study constructor. |
+| optimize_kwargs | Optional | None | Keyword arguments to be passed to the optuna.Study.optimize method. |
+| fitted | bool | False | Whether to compute the fitted values when retraining the best model. |
+| prediction_intervals | Optional | None | Configuration to calibrate prediction intervals when retraining the best model. |
+| **Returns** | **AutoMLForecast** | | **object with best models and optimization results** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoMLForecast.predict
+
+> ``` text
+> AutoMLForecast.predict (h:int, X_df:Union[pandas.core.frame.DataFrame,pol
+> ars.dataframe.frame.DataFrame,NoneType]=None,
+> level:Optional[List[Union[int,float]]]=None)
+> ```
+
+*“Compute forecasts*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Number of periods to predict. |
+| X_df | Union | None | Dataframe with the future exogenous features. Should have the id column and the time column. |
+| level | Optional | None | Confidence levels between 0 and 100 for prediction intervals. |
+| **Returns** | **Union** | | **Predictions for each serie and timestep, with one column per model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoMLForecast.save
+
+> ``` text
+> AutoMLForecast.save (path:Union[str,pathlib.Path])
+> ```
+
+*Save AutoMLForecast objects*
+
+| | **Type** | **Details** |
+|-------------|----------|-------------------------------------------|
+| path | Union | Directory where artifacts will be stored. |
+| **Returns** | **None** | |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoMLForecast.forecast_fitted_values
+
+> ``` text
+> AutoMLForecast.forecast_fitted_values
+> (level:Optional[List[Union[int,flo
+> at]]]=None)
+> ```
+
+*Access in-sample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels between 0 and 100 for prediction intervals. |
+| **Returns** | **Union** | | **Dataframe with predictions for the training set** |
+
+```python
+import time
+
+import pandas as pd
+from datasetsforecast.m4 import M4, M4Evaluation, M4Info
+from sklearn.linear_model import Ridge
+from sklearn.compose import ColumnTransformer
+from sklearn.pipeline import make_pipeline
+from sklearn.preprocessing import OneHotEncoder
+```
+
+
+```python
+def train_valid_split(group):
+ df, *_ = M4.load(directory='data', group=group)
+ df['ds'] = df['ds'].astype('int')
+ horizon = M4Info[group].horizon
+ valid = df.groupby('unique_id').tail(horizon).copy()
+ train = df.drop(valid.index).reset_index(drop=True)
+ return train, valid
+```
+
+
+```python
+ridge_pipeline = make_pipeline(
+ ColumnTransformer(
+ [('encoder', OneHotEncoder(), ['unique_id'])],
+ remainder='passthrough',
+ ),
+ Ridge()
+)
+auto_ridge = AutoModel(ridge_pipeline, lambda trial: {f'ridge__{k}': v for k, v in ridge_space(trial).items()})
+```
+
+
+```python
+optuna.logging.set_verbosity(optuna.logging.ERROR)
+group = 'Weekly'
+train, valid = train_valid_split(group)
+train['unique_id'] = train['unique_id'].astype('category')
+valid['unique_id'] = valid['unique_id'].astype(train['unique_id'].dtype)
+info = M4Info[group]
+h = info.horizon
+season_length = info.seasonality
+auto_mlf = AutoMLForecast(
+ freq=1,
+ season_length=season_length,
+ models={
+ 'lgb': AutoLightGBM(),
+ 'ridge': auto_ridge,
+ },
+ fit_config=lambda trial: {'static_features': ['unique_id']},
+ num_threads=2,
+)
+auto_mlf.fit(
+ df=train,
+ n_windows=2,
+ h=h,
+ num_samples=2,
+ optimize_kwargs={'timeout': 60},
+ fitted=True,
+ prediction_intervals=PredictionIntervals(n_windows=2, h=h),
+)
+auto_mlf.predict(h, level=[80])
+```
+
+| | unique_id | ds | lgb | lgb-lo-80 | lgb-hi-80 | ridge | ridge-lo-80 | ridge-hi-80 |
+|----|----|----|----|----|----|----|----|----|
+| 0 | W1 | 2180 | 35529.435224 | 35061.835362 | 35997.035086 | 36110.921202 | 35880.445097 | 36341.397307 |
+| 1 | W1 | 2181 | 35521.764894 | 34973.035617 | 36070.494171 | 36195.175757 | 36051.013811 | 36339.337702 |
+| 2 | W1 | 2182 | 35537.417268 | 34960.050939 | 36114.783596 | 36107.528852 | 35784.062169 | 36430.995536 |
+| 3 | W1 | 2183 | 35538.058206 | 34823.640706 | 36252.475705 | 36027.139248 | 35612.635725 | 36441.642771 |
+| 4 | W1 | 2184 | 35614.611211 | 34627.023739 | 36602.198683 | 36092.858489 | 35389.690977 | 36796.026000 |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| 4662 | W99 | 2292 | 15071.536978 | 14484.617399 | 15658.456557 | 15319.146221 | 14869.410567 | 15768.881875 |
+| 4663 | W99 | 2293 | 15058.145278 | 14229.686322 | 15886.604234 | 15299.549555 | 14584.269352 | 16014.829758 |
+| 4664 | W99 | 2294 | 15042.493434 | 14096.380636 | 15988.606232 | 15271.744712 | 14365.349338 | 16178.140086 |
+| 4665 | W99 | 2295 | 15042.144846 | 14037.053904 | 16047.235787 | 15250.070504 | 14403.428791 | 16096.712216 |
+| 4666 | W99 | 2296 | 15038.729044 | 13944.821480 | 16132.636609 | 15232.127800 | 14325.059776 | 16139.195824 |
+
+```python
+auto_mlf.forecast_fitted_values(level=[95])
+```
+
+| | unique_id | ds | y | lgb | lgb-lo-95 | lgb-hi-95 | ridge | ridge-lo-95 | ridge-hi-95 |
+|----|----|----|----|----|----|----|----|----|----|
+| 0 | W1 | 15 | 1071.06 | 1060.584344 | 599.618355 | 1521.550334 | 1076.990151 | 556.535492 | 1597.444810 |
+| 1 | W1 | 16 | 1073.73 | 1072.669242 | 611.703252 | 1533.635232 | 1083.633276 | 563.178617 | 1604.087936 |
+| 2 | W1 | 17 | 1066.97 | 1072.452128 | 611.486139 | 1533.418118 | 1084.724311 | 564.269652 | 1605.178970 |
+| 3 | W1 | 18 | 1066.17 | 1065.837828 | 604.871838 | 1526.803818 | 1080.127197 | 559.672538 | 1600.581856 |
+| 4 | W1 | 19 | 1064.43 | 1065.214681 | 604.248691 | 1526.180671 | 1080.636826 | 560.182167 | 1601.091485 |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| 361881 | W99 | 2279 | 15738.54 | 15887.661228 | 15721.237195 | 16054.085261 | 15927.918181 | 15723.222760 | 16132.613603 |
+| 361882 | W99 | 2280 | 15388.13 | 15755.943789 | 15589.519756 | 15922.367823 | 15841.599064 | 15636.903642 | 16046.294485 |
+| 361883 | W99 | 2281 | 15187.62 | 15432.224701 | 15265.800668 | 15598.648735 | 15584.462232 | 15379.766811 | 15789.157654 |
+| 361884 | W99 | 2282 | 15172.27 | 15177.040831 | 15010.616797 | 15343.464864 | 15396.243223 | 15191.547801 | 15600.938644 |
+| 361885 | W99 | 2283 | 15101.03 | 15162.090803 | 14995.666770 | 15328.514836 | 15335.982465 | 15131.287044 | 15540.677887 |
+
+```python
+import polars as pl
+```
+
+
+```python
+train_pl = pl.from_pandas(train.astype({'unique_id': 'str'}))
+auto_mlf = AutoMLForecast(
+ freq=1,
+ season_length=season_length,
+ models={'ridge': AutoRidge()},
+ num_threads=2,
+)
+auto_mlf.fit(
+ df=train_pl,
+ n_windows=2,
+ h=h,
+ num_samples=2,
+ optimize_kwargs={'timeout': 60},
+ fitted=True,
+ prediction_intervals=PredictionIntervals(n_windows=2, h=h),
+)
+auto_mlf.predict(h, level=[80])
+```
+
+| unique_id | ds | ridge | ridge-lo-80 | ridge-hi-80 |
+|-----------|------|--------------|--------------|--------------|
+| str | i64 | f64 | f64 | f64 |
+| "W1" | 2180 | 35046.096663 | 34046.69521 | 36045.498116 |
+| "W1" | 2181 | 34743.269216 | 33325.847975 | 36160.690457 |
+| "W1" | 2182 | 34489.591086 | 32591.254559 | 36387.927614 |
+| "W1" | 2183 | 34270.768179 | 32076.507727 | 36465.02863 |
+| "W1" | 2184 | 34124.021857 | 31352.454121 | 36895.589593 |
+| … | … | … | … | … |
+| "W99" | 2292 | 14719.457096 | 13983.308582 | 15455.605609 |
+| "W99" | 2293 | 14631.552077 | 13928.874336 | 15334.229818 |
+| "W99" | 2294 | 14532.905239 | 13642.840118 | 15422.97036 |
+| "W99" | 2295 | 14446.065443 | 13665.088667 | 15227.04222 |
+| "W99" | 2296 | 14363.049604 | 13654.220051 | 15071.879157 |
+
+```python
+auto_mlf.forecast_fitted_values(level=[95])
+```
+
+| unique_id | ds | y | ridge | ridge-lo-95 | ridge-hi-95 |
+|-----------|------|----------|--------------|--------------|--------------|
+| str | i64 | f64 | f64 | f64 | f64 |
+| "W1" | 14 | 1061.96 | 1249.326428 | 488.765249 | 2009.887607 |
+| "W1" | 15 | 1071.06 | 1246.067836 | 485.506657 | 2006.629015 |
+| "W1" | 16 | 1073.73 | 1254.027897 | 493.466718 | 2014.589076 |
+| "W1" | 17 | 1066.97 | 1254.475948 | 493.914769 | 2015.037126 |
+| "W1" | 18 | 1066.17 | 1248.306754 | 487.745575 | 2008.867933 |
+| … | … | … | … | … | … |
+| "W99" | 2279 | 15738.54 | 15754.558812 | 15411.968645 | 16097.148979 |
+| "W99" | 2280 | 15388.13 | 15655.780865 | 15313.190698 | 15998.371032 |
+| "W99" | 2281 | 15187.62 | 15367.498468 | 15024.908301 | 15710.088635 |
+| "W99" | 2282 | 15172.27 | 15172.591423 | 14830.001256 | 15515.18159 |
+| "W99" | 2283 | 15101.03 | 15141.032886 | 14798.44272 | 15483.623053 |
+
diff --git a/mlforecast/callbacks.html.mdx b/mlforecast/callbacks.html.mdx
new file mode 100644
index 00000000..d538533b
--- /dev/null
+++ b/mlforecast/callbacks.html.mdx
@@ -0,0 +1,39 @@
+---
+output-file: callbacks.html
+title: Callbacks
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### SaveFeatures
+
+> ``` text
+> SaveFeatures ()
+> ```
+
+*Saves the features in every timestamp.*
+
+------------------------------------------------------------------------
+
+source
+
+### SaveFeatures.get_features
+
+> ``` text
+> SaveFeatures.get_features (with_step:bool=False)
+> ```
+
+*Retrieves the input features for every timestep*
+
+| | **Type** | **Default** | **Details** |
+|-------------|-----------|-------------|-----------------------------------|
+| with_step | bool | False | Add a column indicating the step |
+| **Returns** | **Union** | | **DataFrame with input features** |
+
diff --git a/mlforecast/compat.mdx b/mlforecast/compat.mdx
new file mode 100644
index 00000000..8b137891
--- /dev/null
+++ b/mlforecast/compat.mdx
@@ -0,0 +1 @@
+
diff --git a/mlforecast/core.html.mdx b/mlforecast/core.html.mdx
new file mode 100644
index 00000000..f87fe018
--- /dev/null
+++ b/mlforecast/core.html.mdx
@@ -0,0 +1,378 @@
+---
+output-file: core.html
+title: Core
+---
+
+
+```python
+import datetime
+
+import tempfile
+from nbdev import show_doc
+from fastcore.test import test_eq, test_fail, test_warns
+
+from mlforecast.callbacks import SaveFeatures
+from mlforecast.lag_transforms import ExpandingMean, RollingMean
+from mlforecast.target_transforms import Differences, LocalStandardScaler
+from mlforecast.utils import generate_daily_series, generate_prices_for_series
+```
+
+## Data format
+
+The required input format is a dataframe with at least the following
+columns: \* `unique_id` with a unique identifier for each time serie \*
+`ds` with the datestamp and a column \* `y` with the values of the
+serie.
+
+Every other column is considered a static feature unless stated
+otherwise in `TimeSeries.fit`
+
+```python
+series = generate_daily_series(20, n_static_features=2)
+series
+```
+
+| | unique_id | ds | y | static_0 | static_1 |
+|------|-----------|------------|------------|----------|----------|
+| 0 | id_00 | 2000-01-01 | 7.404529 | 27 | 53 |
+| 1 | id_00 | 2000-01-02 | 35.952624 | 27 | 53 |
+| 2 | id_00 | 2000-01-03 | 68.958353 | 27 | 53 |
+| 3 | id_00 | 2000-01-04 | 84.994505 | 27 | 53 |
+| 4 | id_00 | 2000-01-05 | 113.219810 | 27 | 53 |
+| ... | ... | ... | ... | ... | ... |
+| 4869 | id_19 | 2000-03-25 | 400.606807 | 97 | 45 |
+| 4870 | id_19 | 2000-03-26 | 538.794824 | 97 | 45 |
+| 4871 | id_19 | 2000-03-27 | 620.202104 | 97 | 45 |
+| 4872 | id_19 | 2000-03-28 | 20.625426 | 97 | 45 |
+| 4873 | id_19 | 2000-03-29 | 141.513169 | 97 | 45 |
+
+For simplicity we’ll just take one time serie here.
+
+```python
+uids = series['unique_id'].unique()
+serie = series[series['unique_id'].eq(uids[0])]
+serie
+```
+
+| | unique_id | ds | y | static_0 | static_1 |
+|-----|-----------|------------|------------|----------|----------|
+| 0 | id_00 | 2000-01-01 | 7.404529 | 27 | 53 |
+| 1 | id_00 | 2000-01-02 | 35.952624 | 27 | 53 |
+| 2 | id_00 | 2000-01-03 | 68.958353 | 27 | 53 |
+| 3 | id_00 | 2000-01-04 | 84.994505 | 27 | 53 |
+| 4 | id_00 | 2000-01-05 | 113.219810 | 27 | 53 |
+| ... | ... | ... | ... | ... | ... |
+| 217 | id_00 | 2000-08-05 | 13.263188 | 27 | 53 |
+| 218 | id_00 | 2000-08-06 | 38.231981 | 27 | 53 |
+| 219 | id_00 | 2000-08-07 | 59.555183 | 27 | 53 |
+| 220 | id_00 | 2000-08-08 | 86.986368 | 27 | 53 |
+| 221 | id_00 | 2000-08-09 | 119.254810 | 27 | 53 |
+
+------------------------------------------------------------------------
+
+source
+
+### TimeSeries
+
+> ``` text
+> TimeSeries (freq:Union[int,str], lags:Optional[Iterable[int]]=None, lag_t
+> ransforms:Optional[Dict[int,List[Union[Callable,Tuple[Callabl
+> e,Any]]]]]=None,
+> date_features:Optional[Iterable[Union[str,Callable]]]=None,
+> num_threads:int=1, target_transforms:Optional[List[Union[mlfo
+> recast.target_transforms.BaseTargetTransform,mlforecast.targe
+> t_transforms._BaseGroupedArrayTargetTransform]]]=None,
+> lag_transforms_namer:Optional[Callable]=None)
+> ```
+
+*Utility class for storing and transforming time series data.*
+
+The
+[`TimeSeries`](https://Nixtla.github.io/mlforecast/core.html#timeseries)
+class takes care of defining the transformations to be performed
+(`lags`, `lag_transforms` and `date_features`). The transformations can
+be computed using multithreading if `num_threads > 1`.
+
+```python
+def month_start_or_end(dates):
+ return dates.is_month_start | dates.is_month_end
+
+flow_config = dict(
+ freq='W-THU',
+ lags=[7],
+ lag_transforms={
+ 1: [ExpandingMean(), RollingMean(7)]
+ },
+ date_features=['dayofweek', 'week', month_start_or_end]
+)
+
+ts = TimeSeries(**flow_config)
+ts
+```
+
+``` text
+TimeSeries(freq=W-THU, transforms=['lag7', 'expanding_mean_lag1', 'rolling_mean_lag1_window_size7'], date_features=['dayofweek', 'week', 'month_start_or_end'], num_threads=1)
+```
+
+The frequency is converted to an offset.
+
+```python
+test_eq(ts.freq, pd.tseries.frequencies.to_offset(flow_config['freq']))
+```
+
+The date features are stored as they were passed to the constructor.
+
+```python
+test_eq(ts.date_features, flow_config['date_features'])
+```
+
+The transformations are stored as a dictionary where the key is the name
+of the transformation (name of the column in the dataframe with the
+computed features), which is built using `build_transform_name` and the
+value is a tuple where the first element is the lag it is applied to,
+then the function and then the function arguments.
+
+```python
+test_eq(
+ ts.transforms.keys(),
+ ['lag7', 'expanding_mean_lag1', 'rolling_mean_lag1_window_size7'],
+)
+```
+
+Note that for `lags` we define the transformation as the identity
+function applied to its corresponding lag. This is because
+[`_transform_series`](https://Nixtla.github.io/mlforecast/grouped_array.html#_transform_series)
+takes the lag as an argument and shifts the array before computing the
+transformation.
+
+------------------------------------------------------------------------
+
+source
+
+## TimeSeries.fit_transform
+
+> ``` text
+> TimeSeries.fit_transform (data:~DFType, id_col:str, time_col:str,
+> target_col:str,
+> static_features:Optional[List[str]]=None,
+> dropna:bool=True,
+> keep_last_n:Optional[int]=None,
+> max_horizon:Optional[int]=None,
+> return_X_y:bool=False, as_numpy:bool=False,
+> weight_col:Optional[str]=None)
+> ```
+
+\*Add the features to `data` and save the required information for the
+predictions step.
+
+If not all features are static, specify which ones are in
+`static_features`. If you don’t want to drop rows with null values after
+the transformations set `dropna=False` If `keep_last_n` is not None then
+that number of observations is kept across all series for updates.\*
+
+```python
+flow_config = dict(
+ freq='D',
+ lags=[7, 14],
+ lag_transforms={
+ 2: [
+ RollingMean(7),
+ RollingMean(14),
+ ]
+ },
+ date_features=['dayofweek', 'month', 'year'],
+ num_threads=2
+)
+
+ts = TimeSeries(**flow_config)
+_ = ts.fit_transform(series, id_col='unique_id', time_col='ds', target_col='y')
+```
+
+The series values are stored as a GroupedArray in an attribute `ga`. If
+the data type of the series values is an int then it is converted to
+`np.float32`, this is because lags generate `np.nan`s so we need a float
+data type for them.
+
+```python
+np.testing.assert_equal(
+ ts.ga.data,
+ series.groupby('unique_id', observed=True).tail(ts.keep_last_n)['y'],
+)
+```
+
+The series ids are stored in an `uids` attribute.
+
+```python
+test_eq(ts.uids, series['unique_id'].unique())
+```
+
+For each time serie, the last observed date is stored so that
+predictions start from the last date + the frequency.
+
+```python
+test_eq(ts.last_dates, series.groupby('unique_id', observed=True)['ds'].max().values)
+```
+
+The last row of every serie without the `y` and `ds` columns are taken
+as static features.
+
+```python
+pd.testing.assert_frame_equal(
+ ts.static_features_,
+ series.groupby('unique_id', observed=True).tail(1).drop(columns=['ds', 'y']).reset_index(drop=True),
+)
+```
+
+If you pass `static_features` to
+[`TimeSeries.fit_transform`](https://Nixtla.github.io/mlforecast/core.html#timeseries.fit_transform)
+then only these are kept.
+
+```python
+ts.fit_transform(series, id_col='unique_id', time_col='ds', target_col='y', static_features=['static_0'])
+
+pd.testing.assert_frame_equal(
+ ts.static_features_,
+ series.groupby('unique_id', observed=True).tail(1)[['unique_id', 'static_0']].reset_index(drop=True),
+)
+```
+
+You can also specify keep_last_n in TimeSeries.fit_transform, which
+means that after computing the features for training we want to keep
+only the last n samples of each time serie for computing the updates.
+This saves both memory and time, since the updates are performed by
+running the transformation functions on all time series again and
+keeping only the last value (the update).
+
+If you have very long time series and your updates only require a small
+sample it’s recommended that you set keep_last_n to the minimum number
+of samples required to compute the updates, which in this case is 15
+since we have a rolling mean of size 14 over the lag 2 and in the first
+update the lag 2 becomes the lag 1. This is because in the first update
+the lag 1 is the last value of the series (or the lag 0), the lag 2 is
+the lag 1 and so on.
+
+```python
+keep_last_n = 15
+
+ts = TimeSeries(**flow_config)
+df = ts.fit_transform(series, id_col='unique_id', time_col='ds', target_col='y', keep_last_n=keep_last_n)
+ts._predict_setup()
+
+expected_lags = ['lag7', 'lag14']
+expected_transforms = ['rolling_mean_lag2_window_size7',
+ 'rolling_mean_lag2_window_size14']
+expected_date_features = ['dayofweek', 'month', 'year']
+
+test_eq(ts.features, expected_lags + expected_transforms + expected_date_features)
+test_eq(ts.static_features_.columns.tolist() + ts.features, df.columns.drop(['ds', 'y']).tolist())
+# we dropped 2 rows because of the lag 2 and 13 more to have the window of size 14
+test_eq(df.shape[0], series.shape[0] - (2 + 13) * ts.ga.n_groups)
+test_eq(ts.ga.data.size, ts.ga.n_groups * keep_last_n)
+```
+
+[`TimeSeries.fit_transform`](https://Nixtla.github.io/mlforecast/core.html#timeseries.fit_transform)
+requires that the *y* column doesn’t have any null values. This is
+because the transformations could propagate them forward, so if you have
+null values in the *y* column you’ll get an error.
+
+```python
+series_with_nulls = series.copy()
+series_with_nulls.loc[1, 'y'] = np.nan
+test_fail(
+ lambda: ts.fit_transform(series_with_nulls, id_col='unique_id', time_col='ds', target_col='y'),
+ contains='y column contains null values'
+)
+```
+
+------------------------------------------------------------------------
+
+source
+
+## TimeSeries.predict
+
+> ``` text
+> TimeSeries.predict (models:Dict[str,Union[sklearn.base.BaseEstimator,List
+> [sklearn.base.BaseEstimator]]], horizon:int,
+> before_predict_callback:Optional[Callable]=None,
+> after_predict_callback:Optional[Callable]=None,
+> X_df:Optional[~DFType]=None,
+> ids:Optional[List[str]]=None)
+> ```
+
+Once we have a trained model we can use
+[`TimeSeries.predict`](https://Nixtla.github.io/mlforecast/core.html#timeseries.predict)
+passing the model and the horizon to get the predictions back.
+
+```python
+class DummyModel:
+ def predict(self, X: pd.DataFrame) -> np.ndarray:
+ return X['lag7'].values
+
+horizon = 7
+model = DummyModel()
+ts = TimeSeries(**flow_config)
+ts.fit_transform(series, id_col='unique_id', time_col='ds', target_col='y')
+predictions = ts.predict({'DummyModel': model}, horizon)
+
+grouped_series = series.groupby('unique_id', observed=True)
+expected_preds = grouped_series['y'].tail(7) # the model predicts the lag-7
+last_dates = grouped_series['ds'].max()
+expected_dsmin = last_dates + pd.offsets.Day()
+expected_dsmax = last_dates + horizon * pd.offsets.Day()
+grouped_preds = predictions.groupby('unique_id', observed=True)
+
+np.testing.assert_allclose(predictions['DummyModel'], expected_preds)
+pd.testing.assert_series_equal(grouped_preds['ds'].min(), expected_dsmin)
+pd.testing.assert_series_equal(grouped_preds['ds'].max(), expected_dsmax)
+```
+
+If we have dynamic features we can pass them to `X_df`.
+
+```python
+class PredictPrice:
+ def predict(self, X):
+ return X['price']
+
+series = generate_daily_series(20, n_static_features=2, equal_ends=True)
+dynamic_series = series.rename(columns={'static_1': 'product_id'})
+prices_catalog = generate_prices_for_series(dynamic_series)
+series_with_prices = dynamic_series.merge(prices_catalog, how='left')
+
+model = PredictPrice()
+ts = TimeSeries(**flow_config)
+ts.fit_transform(
+ series_with_prices,
+ id_col='unique_id',
+ time_col='ds',
+ target_col='y',
+ static_features=['static_0', 'product_id'],
+)
+predictions = ts.predict({'PredictPrice': model}, horizon=1, X_df=prices_catalog)
+pd.testing.assert_frame_equal(
+ predictions.rename(columns={'PredictPrice': 'price'}),
+ prices_catalog.merge(predictions[['unique_id', 'ds']])[['unique_id', 'ds', 'price']]
+)
+```
+
+------------------------------------------------------------------------
+
+source
+
+## TimeSeries.update
+
+> ``` text
+> TimeSeries.update
+> (df:Union[pandas.core.frame.DataFrame,polars.dataframe
+> .frame.DataFrame])
+> ```
+
+*Update the values of the stored series.*
+
diff --git a/mlforecast/dark.png b/mlforecast/dark.png
new file mode 100644
index 00000000..4142a0bb
Binary files /dev/null and b/mlforecast/dark.png differ
diff --git a/mlforecast/distributed.forecast.html.mdx b/mlforecast/distributed.forecast.html.mdx
new file mode 100644
index 00000000..6ce05f34
--- /dev/null
+++ b/mlforecast/distributed.forecast.html.mdx
@@ -0,0 +1,242 @@
+---
+description: Distributed pipeline encapsulation
+output-file: distributed.forecast.html
+title: Distributed Forecast
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### DistributedMLForecast
+
+> ``` text
+> DistributedMLForecast (models, freq:Union[int,str],
+> lags:Optional[Iterable[int]]=None, lag_transforms:
+> Optional[Dict[int,List[Union[Callable,Tuple[Callab
+> le,Any]]]]]=None, date_features:Optional[Iterable[
+> Union[str,Callable]]]=None, num_threads:int=1, tar
+> get_transforms:Optional[List[Union[mlforecast.targ
+> et_transforms.BaseTargetTransform,mlforecast.targe
+> t_transforms._BaseGroupedArrayTargetTransform]]]=N
+> one, engine=None,
+> num_partitions:Optional[int]=None,
+> lag_transforms_namer:Optional[Callable]=None)
+> ```
+
+*Multi backend distributed pipeline*
+
+------------------------------------------------------------------------
+
+source
+
+### DistributedMLForecast.fit
+
+> ``` text
+> DistributedMLForecast.fit (df:~AnyDataFrame, id_col:str='unique_id',
+> time_col:str='ds', target_col:str='y',
+> static_features:Optional[List[str]]=None,
+> dropna:bool=True,
+> keep_last_n:Optional[int]=None)
+> ```
+
+*Apply the feature engineering and train the models.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | AnyDataFrame | | Series data in long format. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| static_features | Optional | None | Names of the features that are static and will be repeated when forecasting. |
+| dropna | bool | True | Drop rows with missing values produced by the transformations. |
+| keep_last_n | Optional | None | Keep only these many records from each serie for the forecasting step. Can save time and memory if your features allow it. |
+| **Returns** | **DistributedMLForecast** | | **Forecast object with series values and trained models.** |
+
+------------------------------------------------------------------------
+
+source
+
+### DistributedMLForecast.predict
+
+> ``` text
+> DistributedMLForecast.predict (h:int,
+> before_predict_callback:Optional[Callable]
+> =None, after_predict_callback:Optional[Cal
+> lable]=None, X_df:Optional[pandas.core.fra
+> me.DataFrame]=None,
+> new_df:Optional[~AnyDataFrame]=None,
+> ids:Optional[List[str]]=None)
+> ```
+
+*Compute the predictions for the next `horizon` steps.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| before_predict_callback | Optional | None | Function to call on the features before computing the predictions. This function will take the input dataframe that will be passed to the model for predicting and should return a dataframe with the same structure. The series identifier is on the index. |
+| after_predict_callback | Optional | None | Function to call on the predictions before updating the targets. This function will take a pandas Series with the predictions and should return another one with the same structure. The series identifier is on the index. |
+| X_df | Optional | None | Dataframe with the future exogenous features. Should have the id column and the time column. |
+| new_df | Optional | None | Series data of new observations for which forecasts are to be generated. This dataframe should have the same structure as the one used to fit the model, including any features and time series data. If `new_df` is not None, the method will generate forecasts for the new observations. |
+| ids | Optional | None | List with subset of ids seen during training for which the forecasts should be computed. |
+| **Returns** | **AnyDataFrame** | | **Predictions for each serie and timestep, with one column per model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### DistributedMLForecast.save
+
+> ``` text
+> DistributedMLForecast.save (path:str)
+> ```
+
+*Save forecast object*
+
+| | **Type** | **Details** |
+|-------------|----------|-------------------------------------------|
+| path | str | Directory where artifacts will be stored. |
+| **Returns** | **None** | |
+
+------------------------------------------------------------------------
+
+source
+
+### DistributedMLForecast.load
+
+> ``` text
+> DistributedMLForecast.load (path:str, engine)
+> ```
+
+*Load forecast object*
+
+| | **Type** | **Details** |
+|--------|---------------------------|-------------------------------------|
+| path | str | Directory with saved artifacts. |
+| engine | fugue execution engine | Dask Client, Spark Session, etc to use for the distributed computation. |
+| **Returns** | **DistributedMLForecast** | |
+
+------------------------------------------------------------------------
+
+source
+
+### DistributedMLForecast.update
+
+> ``` text
+> DistributedMLForecast.update (df:pandas.core.frame.DataFrame)
+> ```
+
+*Update the values of the stored series.*
+
+| | **Type** | **Details** |
+|-------------|-----------|----------------------------------|
+| df | DataFrame | Dataframe with new observations. |
+| **Returns** | **None** | |
+
+------------------------------------------------------------------------
+
+source
+
+### DistributedMLForecast.to_local
+
+> ``` text
+> DistributedMLForecast.to_local ()
+> ```
+
+\*Convert this distributed forecast object into a local one
+
+This pulls all the data from the remote machines, so you have to be sure
+that it fits in the scheduler/driver. If you’re not sure use the save
+method instead.\*
+
+------------------------------------------------------------------------
+
+source
+
+### DistributedMLForecast.preprocess
+
+> ``` text
+> DistributedMLForecast.preprocess (df:~AnyDataFrame,
+> id_col:str='unique_id',
+> time_col:str='ds', target_col:str='y',
+> static_features:Optional[List[str]]=Non
+> e, dropna:bool=True,
+> keep_last_n:Optional[int]=None)
+> ```
+
+*Add the features to `data`.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | AnyDataFrame | | Series data in long format. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| static_features | Optional | None | Names of the features that are static and will be repeated when forecasting. |
+| dropna | bool | True | Drop rows with missing values produced by the transformations. |
+| keep_last_n | Optional | None | Keep only these many records from each serie for the forecasting step. Can save time and memory if your features allow it. |
+| **Returns** | **AnyDataFrame** | | **`df` with added features.** |
+
+------------------------------------------------------------------------
+
+source
+
+### DistributedMLForecast.cross_validation
+
+> ``` text
+> DistributedMLForecast.cross_validation (df:~AnyDataFrame, n_windows:int,
+> h:int, id_col:str='unique_id',
+> time_col:str='ds',
+> target_col:str='y',
+> step_size:Optional[int]=None, sta
+> tic_features:Optional[List[str]]=
+> None, dropna:bool=True,
+> keep_last_n:Optional[int]=None,
+> refit:bool=True, before_predict_c
+> allback:Optional[Callable]=None,
+> after_predict_callback:Optional[C
+> allable]=None,
+> input_size:Optional[int]=None)
+> ```
+
+*Perform time series cross validation. Creates `n_windows` splits where
+each window has `h` test periods, trains the models, computes the
+predictions and merges the actuals.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | AnyDataFrame | | Series data in long format. |
+| n_windows | int | | Number of windows to evaluate. |
+| h | int | | Number of test periods in each window. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| step_size | Optional | None | Step size between each cross validation window. If None it will be equal to `h`. |
+| static_features | Optional | None | Names of the features that are static and will be repeated when forecasting. |
+| dropna | bool | True | Drop rows with missing values produced by the transformations. |
+| keep_last_n | Optional | None | Keep only these many records from each serie for the forecasting step. Can save time and memory if your features allow it. |
+| refit | bool | True | Retrain model for each cross validation window. If False, the models are trained at the beginning and then used to predict each window. |
+| before_predict_callback | Optional | None | Function to call on the features before computing the predictions. This function will take the input dataframe that will be passed to the model for predicting and should return a dataframe with the same structure. The series identifier is on the index. |
+| after_predict_callback | Optional | None | Function to call on the predictions before updating the targets. This function will take a pandas Series with the predictions and should return another one with the same structure. The series identifier is on the index. |
+| input_size | Optional | None | Maximum training samples per serie in each window. If None, will use an expanding window. |
+| **Returns** | **AnyDataFrame** | | **Predictions for each window with the series id, timestamp, target value and predictions from each model.** |
+
diff --git a/mlforecast/distributed.models.dask.lgb.html.mdx b/mlforecast/distributed.models.dask.lgb.html.mdx
new file mode 100644
index 00000000..a0f83d25
--- /dev/null
+++ b/mlforecast/distributed.models.dask.lgb.html.mdx
@@ -0,0 +1,44 @@
+---
+description: dask LightGBM forecaster
+output-file: distributed.models.dask.lgb.html
+title: DaskLGBMForecast
+---
+
+
+Wrapper of `lightgbm.dask.DaskLGBMRegressor` that adds a `model_`
+property that contains the fitted booster and is sent to the workers to
+in the forecasting step.
+
+------------------------------------------------------------------------
+
+source
+
+### DaskLGBMForecast
+
+> ``` text
+> DaskLGBMForecast (boosting_type:str='gbdt', num_leaves:int=31,
+> max_depth:int=-1, learning_rate:float=0.1,
+> n_estimators:int=100, subsample_for_bin:int=200000, obj
+> ective:Union[str,Callable[[Optional[numpy.ndarray],nump
+> y.ndarray],Tuple[numpy.ndarray,numpy.ndarray]],Callable
+> [[Optional[numpy.ndarray],numpy.ndarray,Optional[numpy.
+> ndarray]],Tuple[numpy.ndarray,numpy.ndarray]],Callable[
+> [Optional[numpy.ndarray],numpy.ndarray,Optional[numpy.n
+> darray],Optional[numpy.ndarray]],Tuple[numpy.ndarray,nu
+> mpy.ndarray]],NoneType]=None,
+> class_weight:Union[dict,str,NoneType]=None,
+> min_split_gain:float=0.0, min_child_weight:float=0.001,
+> min_child_samples:int=20, subsample:float=1.0,
+> subsample_freq:int=0, colsample_bytree:float=1.0,
+> reg_alpha:float=0.0, reg_lambda:float=0.0, random_state
+> :Union[int,numpy.random.mtrand.RandomState,ForwardRef('
+> np.random.Generator'),NoneType]=None,
+> n_jobs:Optional[int]=None, importance_type:str='split',
+> client:Optional[distributed.client.Client]=None,
+> **kwargs:Any)
+> ```
+
+*Distributed version of lightgbm.LGBMRegressor.*
+
diff --git a/mlforecast/distributed.models.dask.xgb.html.mdx b/mlforecast/distributed.models.dask.xgb.html.mdx
new file mode 100644
index 00000000..db54efa7
--- /dev/null
+++ b/mlforecast/distributed.models.dask.xgb.html.mdx
@@ -0,0 +1,109 @@
+---
+description: dask XGBoost forecaster
+output-file: distributed.models.dask.xgb.html
+title: DaskXGBForecast
+---
+
+
+Wrapper of `xgboost.dask.DaskXGBRegressor` that adds a `model_` property
+that contains the fitted model and is sent to the workers in the
+forecasting step.
+
+------------------------------------------------------------------------
+
+source
+
+### DaskXGBForecast
+
+> ``` text
+> DaskXGBForecast (max_depth:Optional[int]=None,
+> max_leaves:Optional[int]=None,
+> max_bin:Optional[int]=None,
+> grow_policy:Optional[str]=None,
+> learning_rate:Optional[float]=None,
+> n_estimators:Optional[int]=None,
+> verbosity:Optional[int]=None, objective:Union[str,xgboos
+> t.sklearn._SklObjWProto,Callable[[Any,Any],Tuple[numpy.n
+> darray,numpy.ndarray]],NoneType]=None,
+> booster:Optional[str]=None,
+> tree_method:Optional[str]=None,
+> n_jobs:Optional[int]=None, gamma:Optional[float]=None,
+> min_child_weight:Optional[float]=None,
+> max_delta_step:Optional[float]=None,
+> subsample:Optional[float]=None,
+> sampling_method:Optional[str]=None,
+> colsample_bytree:Optional[float]=None,
+> colsample_bylevel:Optional[float]=None,
+> colsample_bynode:Optional[float]=None,
+> reg_alpha:Optional[float]=None,
+> reg_lambda:Optional[float]=None,
+> scale_pos_weight:Optional[float]=None,
+> base_score:Optional[float]=None, random_state:Union[nump
+> y.random.mtrand.RandomState,numpy.random._generator.Gene
+> rator,int,NoneType]=None, missing:float=nan,
+> num_parallel_tree:Optional[int]=None, monotone_constrain
+> ts:Union[Dict[str,int],str,NoneType]=None, interaction_c
+> onstraints:Union[str,Sequence[Sequence[str]],NoneType]=N
+> one, importance_type:Optional[str]=None,
+> device:Optional[str]=None,
+> validate_parameters:Optional[bool]=None,
+> enable_categorical:bool=False,
+> feature_types:Optional[Sequence[str]]=None,
+> max_cat_to_onehot:Optional[int]=None,
+> max_cat_threshold:Optional[int]=None,
+> multi_strategy:Optional[str]=None,
+> eval_metric:Union[str,List[str],Callable,NoneType]=None,
+> early_stopping_rounds:Optional[int]=None, callbacks:Opti
+> onal[List[xgboost.callback.TrainingCallback]]=None,
+> **kwargs:Any)
+> ```
+
+*Implementation of the Scikit-Learn API for XGBoost. See
+:doc:`/python/sklearn_estimator` for more information.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| max_depth | Optional | None | Maximum tree depth for base learners. |
+| max_leaves | Optional | None | Maximum number of leaves; 0 indicates no limit. |
+| max_bin | Optional | None | If using histogram-based algorithm, maximum number of bins per feature |
+| grow_policy | Optional | None | Tree growing policy.
- depthwise: Favors splitting at nodes closest to the node, - lossguide: Favors splitting at nodes with highest loss change. |
+| learning_rate | Optional | None | Boosting learning rate (xgb’s “eta”) |
+| n_estimators | Optional | None | Number of gradient boosted trees. Equivalent to number of boosting rounds. |
+| verbosity | Optional | None | The degree of verbosity. Valid values are 0 (silent) - 3 (debug). |
+| objective | Union | None | Specify the learning task and the corresponding learning objective or a custom objective function to be used.
For custom objective, see :doc:`/tutorials/custom_metric_obj` and :ref:`custom-obj-metric` for more information, along with the end note for function signatures. |
+| booster | Optional | None | |
+| tree_method | Optional | None | Specify which tree method to use. Default to auto. If this parameter is set to default, XGBoost will choose the most conservative option available. It’s recommended to study this option from the parameters document :doc:`tree method ` |
+| n_jobs | Optional | None | Number of parallel threads used to run xgboost. When used with other Scikit-Learn algorithms like grid search, you may choose which algorithm to parallelize and balance the threads. Creating thread contention will significantly slow down both algorithms. |
+| gamma | Optional | None | (min_split_loss) Minimum loss reduction required to make a further partition on a leaf node of the tree. |
+| min_child_weight | Optional | None | Minimum sum of instance weight(hessian) needed in a child. |
+| max_delta_step | Optional | None | Maximum delta step we allow each tree’s weight estimation to be. |
+| subsample | Optional | None | Subsample ratio of the training instance. |
+| sampling_method | Optional | None | Sampling method. Used only by the GPU version of `hist` tree method.
- `uniform`: Select random training instances uniformly. - `gradient_based`: Select random training instances with higher probability when the gradient and hessian are larger. (cf. CatBoost) |
+| colsample_bytree | Optional | None | Subsample ratio of columns when constructing each tree. |
+| colsample_bylevel | Optional | None | Subsample ratio of columns for each level. |
+| colsample_bynode | Optional | None | Subsample ratio of columns for each split. |
+| reg_alpha | Optional | None | L1 regularization term on weights (xgb’s alpha). |
+| reg_lambda | Optional | None | L2 regularization term on weights (xgb’s lambda). |
+| scale_pos_weight | Optional | None | Balancing of positive and negative weights. |
+| base_score | Optional | None | The initial prediction score of all instances, global bias. |
+| random_state | Union | None | Random number seed.
.. note::
Using gblinear booster with shotgun updater is nondeterministic as it uses Hogwild algorithm. |
+| missing | float | nan | Value in the data which needs to be present as a missing value. Default to :py:data:`numpy.nan`. |
+| num_parallel_tree | Optional | None | |
+| monotone_constraints | Union | None | Constraint of variable monotonicity. See :doc:`tutorial ` for more information. |
+| interaction_constraints | Union | None | Constraints for interaction representing permitted interactions. The constraints must be specified in the form of a nested list, e.g. `[[0, 1], [2, 3, 4]]`, where each inner list is a group of indices of features that are allowed to interact with each other. See :doc:`tutorial ` for more information |
+| importance_type | Optional | None | |
+| device | Optional | None | .. versionadded:: 2.0.0
Device ordinal, available options are `cpu`, `cuda`, and `gpu`. |
+| validate_parameters | Optional | None | Give warnings for unknown parameter. |
+| enable_categorical | bool | False | See the same parameter of :py:class:`DMatrix` for details. |
+| feature_types | Optional | None | .. versionadded:: 1.7.0
Used for specifying feature types without constructing a dataframe. See :py:class:`DMatrix` for details. |
+| max_cat_to_onehot | Optional | None | .. versionadded:: 1.6.0
.. note:: This parameter is experimental
A threshold for deciding whether XGBoost should use one-hot encoding based split for categorical data. When number of categories is lesser than the threshold then one-hot encoding is chosen, otherwise the categories will be partitioned into children nodes. Also, `enable_categorical` needs to be set to have categorical feature support. See :doc:`Categorical Data ` and :ref:`cat-param` for details. |
+| max_cat_threshold | Optional | None | .. versionadded:: 1.7.0
.. note:: This parameter is experimental
Maximum number of categories considered for each split. Used only by partition-based splits for preventing over-fitting. Also, `enable_categorical` needs to be set to have categorical feature support. See :doc:`Categorical Data ` and :ref:`cat-param` for details. |
+| multi_strategy | Optional | None | .. versionadded:: 2.0.0
.. note:: This parameter is working-in-progress.
The strategy used for training multi-target models, including multi-target regression and multi-class classification. See :doc:`/tutorials/multioutput` for more information.
- `one_output_per_tree`: One model for each target. - `multi_output_tree`: Use multi-target trees. |
+| eval_metric | Union | None | .. versionadded:: 1.6.0
Metric used for monitoring the training result and early stopping. It can be a string or list of strings as names of predefined metric in XGBoost (See doc/parameter.rst), one of the metrics in :py:mod:`sklearn.metrics`, or any other user defined metric that looks like `sklearn.metrics`.
If custom objective is also provided, then custom metric should implement the corresponding reverse link function.
Unlike the `scoring` parameter commonly used in scikit-learn, when a callable object is provided, it’s assumed to be a cost function and by default XGBoost will minimize the result during early stopping.
For advanced usage on Early stopping like directly choosing to maximize instead of minimize, see :py:obj:`xgboost.callback.EarlyStopping`.
See :doc:`/tutorials/custom_metric_obj` and :ref:`custom-obj-metric` for more information.
.. code-block:: python
from sklearn.datasets import load_diabetes from sklearn.metrics import mean_absolute_error X, y = load_diabetes(return_X_y=True) reg = xgb.XGBRegressor( tree_method=“hist”, eval_metric=mean_absolute_error, ) reg.fit(X, y, eval_set=\[(X, y)\]) |
+| early_stopping_rounds | Optional | None | .. versionadded:: 1.6.0
- Activates early stopping. Validation metric needs to improve at least once in every **early_stopping_rounds** round(s) to continue training. Requires at least one item in **eval_set** in :py:meth:`fit`.
- If early stopping occurs, the model will have two additional attributes: :py:attr:`best_score` and :py:attr:`best_iteration`. These are used by the :py:meth:`predict` and :py:meth:`apply` methods to determine the optimal number of trees during inference. If users want to access the full model (including trees built after early stopping), they can specify the `iteration_range` in these inference methods. In addition, other utilities like model plotting can also use the entire model.
- If you prefer to discard the trees after `best_iteration`, consider using the callback function :py:class:`xgboost.callback.EarlyStopping`.
- If there’s more than one item in **eval_set**, the last entry will be used for early stopping. If there’s more than one metric in **eval_metric**, the last metric will be used for early stopping. |
+| callbacks | Optional | None | List of callback functions that are applied at end of each iteration. It is possible to use predefined callbacks by using :ref:`Callback API `.
.. note::
States in callback are not preserved during training, which means callback objects can not be reused for multiple training sessions without reinitialization or deepcopy.
.. code-block:: python
for params in parameters_grid: \# be sure to (re)initialize the callbacks before each run callbacks = \[xgb.callback.LearningRateScheduler(custom_rates)\] reg = xgboost.XGBRegressor(\*\*params, callbacks=callbacks) reg.fit(X, y) |
+| kwargs | Any | | Keyword arguments for XGBoost Booster object. Full documentation of parameters can be found :doc:`here `. Attempting to set a parameter via the constructor args and \*\*kwargs dict simultaneously will result in a TypeError.
.. note:: \*\*kwargs unsupported by scikit-learn
\*\*kwargs is unsupported by scikit-learn. We do not guarantee that parameters passed via this argument will interact properly with scikit-learn. |
+| **Returns** | **None** | | |
+
diff --git a/mlforecast/distributed.models.ray.lgb.html.mdx b/mlforecast/distributed.models.ray.lgb.html.mdx
new file mode 100644
index 00000000..af06756e
--- /dev/null
+++ b/mlforecast/distributed.models.ray.lgb.html.mdx
@@ -0,0 +1,44 @@
+---
+description: ray LightGBM forecaster
+output-file: distributed.models.ray.lgb.html
+title: RayLGBMForecast
+---
+
+
+Wrapper of `lightgbm.ray.RayLGBMRegressor` that adds a `model_` property
+that contains the fitted booster and is sent to the workers to in the
+forecasting step.
+
+------------------------------------------------------------------------
+
+source
+
+### RayLGBMForecast
+
+> ``` text
+> RayLGBMForecast (boosting_type:str='gbdt', num_leaves:int=31,
+> max_depth:int=-1, learning_rate:float=0.1,
+> n_estimators:int=100, subsample_for_bin:int=200000, obje
+> ctive:Union[str,Callable[[Optional[numpy.ndarray],numpy.
+> ndarray],Tuple[numpy.ndarray,numpy.ndarray]],Callable[[O
+> ptional[numpy.ndarray],numpy.ndarray,Optional[numpy.ndar
+> ray]],Tuple[numpy.ndarray,numpy.ndarray]],Callable[[Opti
+> onal[numpy.ndarray],numpy.ndarray,Optional[numpy.ndarray
+> ],Optional[numpy.ndarray]],Tuple[numpy.ndarray,numpy.nda
+> rray]],NoneType]=None,
+> class_weight:Union[Dict,str,NoneType]=None,
+> min_split_gain:float=0.0, min_child_weight:float=0.001,
+> min_child_samples:int=20, subsample:float=1.0,
+> subsample_freq:int=0, colsample_bytree:float=1.0,
+> reg_alpha:float=0.0, reg_lambda:float=0.0, random_state:
+> Union[int,numpy.random.mtrand.RandomState,numpy.random._
+> generator.Generator,NoneType]=None,
+> n_jobs:Optional[int]=None, importance_type:str='split',
+> **kwargs:Any)
+> ```
+
+***PublicAPI (beta):** This API is in beta and may change before
+becoming stable.*
+
diff --git a/mlforecast/distributed.models.ray.xgb.html.mdx b/mlforecast/distributed.models.ray.xgb.html.mdx
new file mode 100644
index 00000000..b24b2d28
--- /dev/null
+++ b/mlforecast/distributed.models.ray.xgb.html.mdx
@@ -0,0 +1,34 @@
+---
+description: ray XGBoost forecaster
+output-file: distributed.models.ray.xgb.html
+title: RayXGBForecast
+---
+
+
+Wrapper of `xgboost.ray.RayXGBRegressor` that adds a `model_` property
+that contains the fitted model and is sent to the workers in the
+forecasting step.
+
+------------------------------------------------------------------------
+
+source
+
+### RayXGBForecast
+
+> ``` text
+> RayXGBForecast (objective:Union[str,xgboost.sklearn._SklObjWProto,Callabl
+> e[[Any,Any],Tuple[numpy.ndarray,numpy.ndarray]],NoneType]
+> ='reg:squarederror', **kwargs:Any)
+> ```
+
+*Implementation of the scikit-learn API for Ray-distributed XGBoost
+regression. See :doc:`/python/sklearn_estimator` for more information.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| objective | Union | reg:squarederror | Specify the learning task and the corresponding learning objective or a custom objective function to be used.
For custom objective, see :doc:`/tutorials/custom_metric_obj` and :ref:`custom-obj-metric` for more information, along with the end note for function signatures. |
+| kwargs | Any | | Keyword arguments for XGBoost Booster object. Full documentation of parameters can be found :doc:`here `. Attempting to set a parameter via the constructor args and \*\*kwargs dict simultaneously will result in a TypeError.
.. note:: \*\*kwargs unsupported by scikit-learn
\*\*kwargs is unsupported by scikit-learn. We do not guarantee that parameters passed via this argument will interact properly with scikit-learn.
.. note:: Custom objective function
A custom objective function can be provided for the `objective` parameter. In this case, it should have the signature `objective(y_true, y_pred) -> [grad, hess]` or `objective(y_true, y_pred, *, sample_weight) -> [grad, hess]`:
y_true: array_like of shape \[n_samples\] The target values y_pred: array_like of shape \[n_samples\] The predicted values sample_weight : Optional sample weights.
grad: array_like of shape \[n_samples\] The value of the gradient for each sample point. hess: array_like of shape \[n_samples\] The value of the second derivative for each sample point |
+| **Returns** | **None** | | |
+
diff --git a/mlforecast/distributed.models.spark.lgb.html.mdx b/mlforecast/distributed.models.spark.lgb.html.mdx
new file mode 100644
index 00000000..2afff0b0
--- /dev/null
+++ b/mlforecast/distributed.models.spark.lgb.html.mdx
@@ -0,0 +1,25 @@
+---
+description: spark LightGBM forecaster
+output-file: distributed.models.spark.lgb.html
+title: SparkLGBMForecast
+---
+
+
+Wrapper of `synapse.ml.lightgbm.LightGBMRegressor` that adds an
+`extract_local_model` method to get a local version of the trained model
+and broadcast it to the workers.
+
+------------------------------------------------------------------------
+
+source
+
+### SparkLGBMForecast
+
+> ``` text
+> SparkLGBMForecast ()
+> ```
+
+*Initialize self. See help(type(self)) for accurate signature.*
+
diff --git a/mlforecast/distributed.models.spark.xgb.html.mdx b/mlforecast/distributed.models.spark.xgb.html.mdx
new file mode 100644
index 00000000..eb98aa16
--- /dev/null
+++ b/mlforecast/distributed.models.spark.xgb.html.mdx
@@ -0,0 +1,62 @@
+---
+description: spark XGBoost forecaster
+output-file: distributed.models.spark.xgb.html
+title: SparkXGBForecast
+---
+
+
+Wrapper of `xgboost.spark.SparkXGBRegressor` that adds an
+`extract_local_model` method to get a local version of the trained model
+and broadcast it to the workers.
+
+``` text
+/opt/hostedtoolcache/Python/3.10.16/x64/lib/python3.10/site-packages/fastcore/docscrape.py:230: UserWarning: Unknown section Note
+ else: warn(msg)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### SparkXGBForecast
+
+> ``` text
+> SparkXGBForecast (features_col:Union[str,List[str]]='features',
+> label_col:str='label', prediction_col:str='prediction',
+> pred_contrib_col:Optional[str]=None,
+> validation_indicator_col:Optional[str]=None,
+> weight_col:Optional[str]=None,
+> base_margin_col:Optional[str]=None, num_workers:int=1,
+> use_gpu:Optional[bool]=None, device:Optional[str]=None,
+> force_repartition:bool=False,
+> repartition_random_shuffle:bool=False,
+> enable_sparse_data_optim:bool=False, **kwargs:Any)
+> ```
+
+\*SparkXGBRegressor is a PySpark ML estimator. It implements the XGBoost
+regression algorithm based on XGBoost python library, and it can be used
+in PySpark Pipeline and PySpark ML meta algorithms like -
+:py:class:`~pyspark.ml.tuning.CrossValidator`/ -
+:py:class:`~pyspark.ml.tuning.TrainValidationSplit`/ -
+:py:class:`~pyspark.ml.classification.OneVsRest`
+
+SparkXGBRegressor automatically supports most of the parameters in
+:py:class:`xgboost.XGBRegressor` constructor and most of the parameters
+used in :py:meth:`xgboost.XGBRegressor.fit` and
+:py:meth:`xgboost.XGBRegressor.predict` method.
+
+To enable GPU support, set `device` to `cuda` or `gpu`.
+
+SparkXGBRegressor doesn’t support setting `base_margin` explicitly as
+well, but support another param called `base_margin_col`. see doc below
+for more details.
+
+SparkXGBRegressor doesn’t support `validate_features` and
+`output_margin` param.
+
+SparkXGBRegressor doesn’t support setting `nthread` xgboost param,
+instead, the `nthread` param for each xgboost worker will be set equal
+to `spark.task.cpus` config value.\*
+
diff --git a/mlforecast/docs/getting-started/end_to_end_walkthrough.html.mdx b/mlforecast/docs/getting-started/end_to_end_walkthrough.html.mdx
new file mode 100644
index 00000000..237470e8
--- /dev/null
+++ b/mlforecast/docs/getting-started/end_to_end_walkthrough.html.mdx
@@ -0,0 +1,704 @@
+---
+description: Detailed description of all the functionalities that MLForecast provides.
+output-file: end_to_end_walkthrough.html
+title: End to end walkthrough
+---
+
+
+## Data setup
+
+For this example we’ll use a subset of the M4 hourly dataset. You can
+find the a notebook with the full dataset
+[here](https://www.kaggle.com/code/lemuz90/m4-competition).
+
+```python
+import random
+import tempfile
+from pathlib import Path
+
+import pandas as pd
+from datasetsforecast.m4 import M4
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+await M4.async_download('data', group='Hourly')
+df, *_ = M4.load('data', 'Hourly')
+uids = df['unique_id'].unique()
+random.seed(0)
+sample_uids = random.choices(uids, k=4)
+df = df[df['unique_id'].isin(sample_uids)].reset_index(drop=True)
+df['ds'] = df['ds'].astype('int64')
+df
+```
+
+| | unique_id | ds | y |
+|------|-----------|------|------|
+| 0 | H196 | 1 | 11.8 |
+| 1 | H196 | 2 | 11.4 |
+| 2 | H196 | 3 | 11.1 |
+| 3 | H196 | 4 | 10.8 |
+| 4 | H196 | 5 | 10.6 |
+| ... | ... | ... | ... |
+| 4027 | H413 | 1004 | 99.0 |
+| 4028 | H413 | 1005 | 88.0 |
+| 4029 | H413 | 1006 | 47.0 |
+| 4030 | H413 | 1007 | 41.0 |
+| 4031 | H413 | 1008 | 34.0 |
+
+## EDA
+
+We’ll take a look at our series to get ideas for transformations and
+features.
+
+```python
+fig = plot_series(df, max_insample_length=24 * 14)
+```
+
+
+
+We can use the
+[`MLForecast.preprocess`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.preprocess)
+method to explore different transformations. It looks like these series
+have a strong seasonality on the hour of the day, so we can subtract the
+value from the same hour in the previous day to remove it. This can be
+done with the
+[`mlforecast.target_transforms.Differences`](https://Nixtla.github.io/mlforecast/target_transforms.html#differences)
+transformer, which we pass through `target_transforms`.
+
+```python
+from mlforecast import MLForecast
+from mlforecast.target_transforms import Differences
+```
+
+
+```python
+fcst = MLForecast(
+ models=[], # we're not interested in modeling yet
+ freq=1, # our series have integer timestamps, so we'll just add 1 in every timestep
+ target_transforms=[Differences([24])],
+)
+prep = fcst.preprocess(df)
+prep
+```
+
+| | unique_id | ds | y |
+|------|-----------|------|------|
+| 24 | H196 | 25 | 0.3 |
+| 25 | H196 | 26 | 0.3 |
+| 26 | H196 | 27 | 0.1 |
+| 27 | H196 | 28 | 0.2 |
+| 28 | H196 | 29 | 0.2 |
+| ... | ... | ... | ... |
+| 4027 | H413 | 1004 | 39.0 |
+| 4028 | H413 | 1005 | 55.0 |
+| 4029 | H413 | 1006 | 14.0 |
+| 4030 | H413 | 1007 | 3.0 |
+| 4031 | H413 | 1008 | 4.0 |
+
+This has subtacted the lag 24 from each value, we can see what our
+series look like now.
+
+```python
+fig = plot_series(prep)
+```
+
+
+
+## Adding features
+
+### Lags
+
+Looks like the seasonality is gone, we can now try adding some lag
+features.
+
+```python
+fcst = MLForecast(
+ models=[],
+ freq=1,
+ lags=[1, 24],
+ target_transforms=[Differences([24])],
+)
+prep = fcst.preprocess(df)
+prep
+```
+
+| | unique_id | ds | y | lag1 | lag24 |
+|------|-----------|------|------|------|-------|
+| 48 | H196 | 49 | 0.1 | 0.1 | 0.3 |
+| 49 | H196 | 50 | 0.1 | 0.1 | 0.3 |
+| 50 | H196 | 51 | 0.2 | 0.1 | 0.1 |
+| 51 | H196 | 52 | 0.1 | 0.2 | 0.2 |
+| 52 | H196 | 53 | 0.1 | 0.1 | 0.2 |
+| ... | ... | ... | ... | ... | ... |
+| 4027 | H413 | 1004 | 39.0 | 29.0 | 1.0 |
+| 4028 | H413 | 1005 | 55.0 | 39.0 | -25.0 |
+| 4029 | H413 | 1006 | 14.0 | 55.0 | -20.0 |
+| 4030 | H413 | 1007 | 3.0 | 14.0 | 0.0 |
+| 4031 | H413 | 1008 | 4.0 | 3.0 | -16.0 |
+
+```python
+prep.drop(columns=['unique_id', 'ds']).corr()['y']
+```
+
+``` text
+y 1.000000
+lag1 0.622531
+lag24 -0.234268
+Name: y, dtype: float64
+```
+
+### Lag transforms
+
+Lag transforms are defined as a dictionary where the keys are the lags
+and the values are the transformations that we want to apply to that
+lag. The lag transformations can be either objects from the
+`mlforecast.lag_transforms` module or [numba](http://numba.pydata.org/)
+jitted functions (so that computing the features doesn’t become a
+bottleneck and we can bypass the GIL when using multithreading), we have
+some implemented in the [window-ops
+package](https://github.com/jmoralez/window_ops) but you can also
+implement your own.
+
+```python
+from mlforecast.lag_transforms import ExpandingMean, RollingMean
+```
+
+
+```python
+from numba import njit
+from window_ops.rolling import rolling_mean
+```
+
+
+```python
+@njit
+def rolling_mean_48(x):
+ return rolling_mean(x, window_size=48)
+
+
+fcst = MLForecast(
+ models=[],
+ freq=1,
+ target_transforms=[Differences([24])],
+ lag_transforms={
+ 1: [ExpandingMean()],
+ 24: [RollingMean(window_size=48), rolling_mean_48],
+ },
+)
+prep = fcst.preprocess(df)
+prep
+```
+
+| | unique_id | ds | y | expanding_mean_lag1 | rolling_mean_lag24_window_size48 | rolling_mean_48_lag24 |
+|----|----|----|----|----|----|----|
+| 95 | H196 | 96 | 0.1 | 0.174648 | 0.150000 | 0.150000 |
+| 96 | H196 | 97 | 0.3 | 0.173611 | 0.145833 | 0.145833 |
+| 97 | H196 | 98 | 0.3 | 0.175342 | 0.141667 | 0.141667 |
+| 98 | H196 | 99 | 0.3 | 0.177027 | 0.141667 | 0.141667 |
+| 99 | H196 | 100 | 0.3 | 0.178667 | 0.141667 | 0.141667 |
+| ... | ... | ... | ... | ... | ... | ... |
+| 4027 | H413 | 1004 | 39.0 | 0.242084 | 3.437500 | 3.437500 |
+| 4028 | H413 | 1005 | 55.0 | 0.281633 | 2.708333 | 2.708333 |
+| 4029 | H413 | 1006 | 14.0 | 0.337411 | 2.125000 | 2.125000 |
+| 4030 | H413 | 1007 | 3.0 | 0.351324 | 1.770833 | 1.770833 |
+| 4031 | H413 | 1008 | 4.0 | 0.354018 | 1.208333 | 1.208333 |
+
+You can see that both approaches get to the same result, you can use
+whichever one you feel most comfortable with.
+
+### Date features
+
+If your time column is made of timestamps then it might make sense to
+extract features like week, dayofweek, quarter, etc. You can do that by
+passing a list of strings with [pandas time/date
+components](https://pandas.pydata.org/docs/user_guide/timeseries.html#time-date-components).
+You can also pass functions that will take the time column as input, as
+we’ll show here.
+
+```python
+def hour_index(times):
+ return times % 24
+
+fcst = MLForecast(
+ models=[],
+ freq=1,
+ target_transforms=[Differences([24])],
+ date_features=[hour_index],
+)
+fcst.preprocess(df)
+```
+
+| | unique_id | ds | y | hour_index |
+|------|-----------|------|------|------------|
+| 24 | H196 | 25 | 0.3 | 1 |
+| 25 | H196 | 26 | 0.3 | 2 |
+| 26 | H196 | 27 | 0.1 | 3 |
+| 27 | H196 | 28 | 0.2 | 4 |
+| 28 | H196 | 29 | 0.2 | 5 |
+| ... | ... | ... | ... | ... |
+| 4027 | H413 | 1004 | 39.0 | 20 |
+| 4028 | H413 | 1005 | 55.0 | 21 |
+| 4029 | H413 | 1006 | 14.0 | 22 |
+| 4030 | H413 | 1007 | 3.0 | 23 |
+| 4031 | H413 | 1008 | 4.0 | 0 |
+
+### Target transformations
+
+If you want to do some transformation to your target before computing
+the features and then re-apply it after predicting you can use the
+`target_transforms` argument, which takes a list of transformations. You
+can find the implemented ones in `mlforecast.target_transforms` or you
+can implement your own as described in the [target transformations
+guide](../how-to-guides/target_transforms_guide.html#custom-transformations).
+
+```python
+from mlforecast.target_transforms import LocalStandardScaler
+```
+
+
+```python
+fcst = MLForecast(
+ models=[],
+ freq=1,
+ lags=[1],
+ target_transforms=[LocalStandardScaler()]
+)
+fcst.preprocess(df)
+```
+
+| | unique_id | ds | y | lag1 |
+|------|-----------|------|-----------|-----------|
+| 1 | H196 | 2 | -1.493026 | -1.383286 |
+| 2 | H196 | 3 | -1.575331 | -1.493026 |
+| 3 | H196 | 4 | -1.657635 | -1.575331 |
+| 4 | H196 | 5 | -1.712505 | -1.657635 |
+| 5 | H196 | 6 | -1.794810 | -1.712505 |
+| ... | ... | ... | ... | ... |
+| 4027 | H413 | 1004 | 3.062766 | 2.425012 |
+| 4028 | H413 | 1005 | 2.523128 | 3.062766 |
+| 4029 | H413 | 1006 | 0.511751 | 2.523128 |
+| 4030 | H413 | 1007 | 0.217403 | 0.511751 |
+| 4031 | H413 | 1008 | -0.126003 | 0.217403 |
+
+We can define a naive model to test this
+
+```python
+from sklearn.base import BaseEstimator
+
+class Naive(BaseEstimator):
+ def fit(self, X, y):
+ return self
+
+ def predict(self, X):
+ return X['lag1']
+```
+
+
+```python
+fcst = MLForecast(
+ models=[Naive()],
+ freq=1,
+ lags=[1],
+ target_transforms=[LocalStandardScaler()]
+)
+fcst.fit(df)
+preds = fcst.predict(1)
+preds
+```
+
+| | unique_id | ds | Naive |
+|-----|-----------|------|-------|
+| 0 | H196 | 1009 | 16.8 |
+| 1 | H256 | 1009 | 13.4 |
+| 2 | H381 | 1009 | 207.0 |
+| 3 | H413 | 1009 | 34.0 |
+
+We compare this with the last values of our serie
+
+```python
+last_vals = df.groupby('unique_id').tail(1)
+last_vals
+```
+
+| | unique_id | ds | y |
+|------|-----------|------|-------|
+| 1007 | H196 | 1008 | 16.8 |
+| 2015 | H256 | 1008 | 13.4 |
+| 3023 | H381 | 1008 | 207.0 |
+| 4031 | H413 | 1008 | 34.0 |
+
+```python
+import numpy as np
+```
+
+
+```python
+np.testing.assert_allclose(preds['Naive'], last_vals['y'])
+```
+
+## Training
+
+Once you’ve decided the features, transformations and models that you
+want to use you can use the
+[`MLForecast.fit`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.fit)
+method instead, which will do the preprocessing and then train the
+models. The models can be specified as a list (which will name them by
+using their class name and an index if there are repeated classes) or as
+a dictionary where the keys are the names you want to give to the
+models, i.e. the name of the column that will hold their predictions,
+and the values are the models themselves.
+
+```python
+import lightgbm as lgb
+```
+
+
+```python
+lgb_params = {
+ 'verbosity': -1,
+ 'num_leaves': 512,
+}
+
+fcst = MLForecast(
+ models={
+ 'avg': lgb.LGBMRegressor(**lgb_params),
+ 'q75': lgb.LGBMRegressor(**lgb_params, objective='quantile', alpha=0.75),
+ 'q25': lgb.LGBMRegressor(**lgb_params, objective='quantile', alpha=0.25),
+ },
+ freq=1,
+ target_transforms=[Differences([24])],
+ lags=[1, 24],
+ lag_transforms={
+ 1: [ExpandingMean()],
+ 24: [RollingMean(window_size=48)],
+ },
+ date_features=[hour_index],
+)
+fcst.fit(df)
+```
+
+``` text
+MLForecast(models=[avg, q75, q25], freq=1, lag_features=['lag1', 'lag24', 'expanding_mean_lag1', 'rolling_mean_lag24_window_size48'], date_features=[], num_threads=1)
+```
+
+This computed the features and trained three different models using
+them. We can now compute our forecasts.
+
+## Forecasting
+
+```python
+preds = fcst.predict(48)
+preds
+```
+
+| | unique_id | ds | avg | q75 | q25 |
+|-----|-----------|------|------------|------------|-----------|
+| 0 | H196 | 1009 | 16.295257 | 16.357148 | 16.315731 |
+| 1 | H196 | 1010 | 15.910282 | 16.007322 | 15.862261 |
+| 2 | H196 | 1011 | 15.728367 | 15.780183 | 15.658180 |
+| 3 | H196 | 1012 | 15.468414 | 15.513598 | 15.399717 |
+| 4 | H196 | 1013 | 15.081279 | 15.133848 | 15.007694 |
+| ... | ... | ... | ... | ... | ... |
+| 187 | H413 | 1052 | 100.450617 | 124.211150 | 47.025017 |
+| 188 | H413 | 1053 | 88.426800 | 108.303409 | 44.715380 |
+| 189 | H413 | 1054 | 59.675737 | 81.859964 | 19.239462 |
+| 190 | H413 | 1055 | 57.580356 | 72.703301 | 21.486674 |
+| 191 | H413 | 1056 | 42.669879 | 46.018271 | 24.392357 |
+
+```python
+fig = plot_series(df, preds, max_insample_length=24 * 7)
+```
+
+
+
+## Saving and loading
+
+The MLForecast class has the
+[`MLForecast.save`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.save)
+and
+[`MLForecast.load`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.load)
+to store and then load the forecast object.
+
+```python
+with tempfile.TemporaryDirectory() as tmpdir:
+ save_dir = Path(tmpdir) / 'mlforecast'
+ fcst.save(save_dir)
+ fcst2 = MLForecast.load(save_dir)
+ preds2 = fcst2.predict(48)
+ pd.testing.assert_frame_equal(preds, preds2)
+```
+
+## Updating series’ values
+
+After you’ve trained a forecast object you can save and load it with the
+previous methods. If by the time you want to use it you already know the
+following values of the target you can use the
+[`MLForecast.update`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.update)
+method to incorporate these, which will allow you to use these new
+values when computing predictions.
+
+- If no new values are provided for a serie that’s currently stored,
+ only the previous ones are kept.
+- If new series are included they are added to the existing ones.
+
+```python
+fcst = MLForecast(
+ models=[Naive()],
+ freq=1,
+ lags=[1, 2, 3],
+)
+fcst.fit(df)
+fcst.predict(1)
+```
+
+| | unique_id | ds | Naive |
+|-----|-----------|------|-------|
+| 0 | H196 | 1009 | 16.8 |
+| 1 | H256 | 1009 | 13.4 |
+| 2 | H381 | 1009 | 207.0 |
+| 3 | H413 | 1009 | 34.0 |
+
+```python
+new_values = pd.DataFrame({
+ 'unique_id': ['H196', 'H256'],
+ 'ds': [1009, 1009],
+ 'y': [17.0, 14.0],
+})
+fcst.update(new_values)
+preds = fcst.predict(1)
+preds
+```
+
+| | unique_id | ds | Naive |
+|-----|-----------|------|-------|
+| 0 | H196 | 1010 | 17.0 |
+| 1 | H256 | 1010 | 14.0 |
+| 2 | H381 | 1009 | 207.0 |
+| 3 | H413 | 1009 | 34.0 |
+
+## Estimating model performance
+
+### Cross validation
+
+In order to get an estimate of how well our model will be when
+predicting future data we can perform cross validation, which consist on
+training a few models independently on different subsets of the data,
+using them to predict a validation set and measuring their performance.
+
+Since our data depends on time, we make our splits by removing the last
+portions of the series and using them as validation sets. This process
+is implemented in
+[`MLForecast.cross_validation`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.cross_validation).
+
+```python
+fcst = MLForecast(
+ models=lgb.LGBMRegressor(**lgb_params),
+ freq=1,
+ target_transforms=[Differences([24])],
+ lags=[1, 24],
+ lag_transforms={
+ 1: [ExpandingMean()],
+ 24: [RollingMean(window_size=48)],
+ },
+ date_features=[hour_index],
+)
+cv_result = fcst.cross_validation(
+ df,
+ n_windows=4, # number of models to train/splits to perform
+ h=48, # length of the validation set in each window
+)
+cv_result
+```
+
+| | unique_id | ds | cutoff | y | LGBMRegressor |
+|-----|-----------|------|--------|------|---------------|
+| 0 | H196 | 817 | 816 | 15.3 | 15.383165 |
+| 1 | H196 | 818 | 816 | 14.9 | 14.923219 |
+| 2 | H196 | 819 | 816 | 14.6 | 14.667834 |
+| 3 | H196 | 820 | 816 | 14.2 | 14.275964 |
+| 4 | H196 | 821 | 816 | 13.9 | 13.973491 |
+| ... | ... | ... | ... | ... | ... |
+| 763 | H413 | 1004 | 960 | 99.0 | 65.644823 |
+| 764 | H413 | 1005 | 960 | 88.0 | 71.717097 |
+| 765 | H413 | 1006 | 960 | 47.0 | 76.704377 |
+| 766 | H413 | 1007 | 960 | 41.0 | 53.446638 |
+| 767 | H413 | 1008 | 960 | 34.0 | 54.902634 |
+
+```python
+fig = plot_series(forecasts_df=cv_result.drop(columns='cutoff'))
+```
+
+
+
+We can compute the RMSE on each split.
+
+```python
+from utilsforecast.losses import rmse
+```
+
+
+```python
+def evaluate_cv(df):
+ return rmse(df, models=['LGBMRegressor'], id_col='cutoff').set_index('cutoff')
+
+split_rmse = evaluate_cv(cv_result)
+split_rmse
+```
+
+| | LGBMRegressor |
+|--------|---------------|
+| cutoff | |
+| 816 | 29.418172 |
+| 864 | 34.257598 |
+| 912 | 13.145763 |
+| 960 | 35.066261 |
+
+And the average RMSE across splits.
+
+```python
+split_rmse.mean()
+```
+
+``` text
+LGBMRegressor 27.971949
+dtype: float64
+```
+
+You can quickly try different features and evaluate them this way. We
+can try removing the differencing and using an exponentially weighted
+average of the lag 1 instead of the expanding mean.
+
+```python
+from mlforecast.lag_transforms import ExponentiallyWeightedMean
+```
+
+
+```python
+fcst = MLForecast(
+ models=lgb.LGBMRegressor(**lgb_params),
+ freq=1,
+ lags=[1, 24],
+ lag_transforms={
+ 1: [ExponentiallyWeightedMean(alpha=0.5)],
+ 24: [RollingMean(window_size=48)],
+ },
+ date_features=[hour_index],
+)
+cv_result2 = fcst.cross_validation(
+ df,
+ n_windows=4,
+ h=48,
+)
+evaluate_cv(cv_result2).mean()
+```
+
+``` text
+LGBMRegressor 25.874439
+dtype: float64
+```
+
+### LightGBMCV
+
+In the same spirit of estimating our model’s performance,
+[`LightGBMCV`](https://Nixtla.github.io/mlforecast/lgb_cv.html#lightgbmcv)
+allows us to train a few
+[LightGBM](https://github.com/microsoft/LightGBM) models on different
+partitions of the data. The main differences with
+[`MLForecast.cross_validation`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.cross_validation)
+are:
+
+- It can only train LightGBM models.
+- It trains all models **simultaneously** and gives us per-iteration
+ averages of the errors across the complete forecasting window, which
+ allows us to find the best iteration.
+
+```python
+from mlforecast.lgb_cv import LightGBMCV
+```
+
+
+```python
+cv = LightGBMCV(
+ freq=1,
+ target_transforms=[Differences([24])],
+ lags=[1, 24],
+ lag_transforms={
+ 1: [ExpandingMean()],
+ 24: [RollingMean(window_size=48)],
+ },
+ date_features=[hour_index],
+ num_threads=2,
+)
+cv_hist = cv.fit(
+ df,
+ n_windows=4,
+ h=48,
+ params=lgb_params,
+ eval_every=5,
+ early_stopping_evals=5,
+ compute_cv_preds=True,
+)
+```
+
+``` text
+[5] mape: 0.158639
+[10] mape: 0.163739
+[15] mape: 0.161535
+[20] mape: 0.169491
+[25] mape: 0.163690
+[30] mape: 0.164198
+Early stopping at round 30
+Using best iteration: 5
+```
+
+As you can see this gives us the error by iteration (controlled by the
+`eval_every` argument) and performs early stopping (which can be
+configured with `early_stopping_evals` and `early_stopping_pct`). If you
+set `compute_cv_preds=True` the out-of-fold predictions are computed
+using the best iteration found and are saved in the `cv_preds_`
+attribute.
+
+```python
+cv.cv_preds_
+```
+
+| | unique_id | ds | y | Booster | window |
+|-----|-----------|------|------|-----------|--------|
+| 0 | H196 | 817 | 15.3 | 15.473182 | 0 |
+| 1 | H196 | 818 | 14.9 | 15.038571 | 0 |
+| 2 | H196 | 819 | 14.6 | 14.849409 | 0 |
+| 3 | H196 | 820 | 14.2 | 14.448379 | 0 |
+| 4 | H196 | 821 | 13.9 | 14.148379 | 0 |
+| ... | ... | ... | ... | ... | ... |
+| 187 | H413 | 1004 | 99.0 | 61.425396 | 3 |
+| 188 | H413 | 1005 | 88.0 | 62.886890 | 3 |
+| 189 | H413 | 1006 | 47.0 | 57.886890 | 3 |
+| 190 | H413 | 1007 | 41.0 | 38.849009 | 3 |
+| 191 | H413 | 1008 | 34.0 | 44.720562 | 3 |
+
+```python
+fig = plot_series(forecasts_df=cv.cv_preds_.drop(columns='window'))
+```
+
+
+
+You can use this class to quickly try different configurations of
+features and hyperparameters. Once you’ve found a combination that works
+you can train a model with those features and hyperparameters on all the
+data by creating an
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+object from the
+[`LightGBMCV`](https://Nixtla.github.io/mlforecast/lgb_cv.html#lightgbmcv)
+one as follows:
+
+```python
+final_fcst = MLForecast.from_cv(cv)
+final_fcst.fit(df)
+preds = final_fcst.predict(48)
+fig = plot_series(df, preds, max_insample_length=24 * 14)
+```
+
+
+
diff --git a/mlforecast/docs/getting-started/install.html.mdx b/mlforecast/docs/getting-started/install.html.mdx
new file mode 100644
index 00000000..192ff550
--- /dev/null
+++ b/mlforecast/docs/getting-started/install.html.mdx
@@ -0,0 +1,85 @@
+---
+description: Instructions to install the package from different sources.
+output-file: install.html
+title: Install
+---
+
+
+## Released versions
+
+### PyPI
+
+#### Latest release
+
+To install the latest release of mlforecast from
+[PyPI](https://pypi.org/project/mlforecast/) you just have to run the
+following in a terminal:
+
+`pip install mlforecast`
+
+#### Specific version
+
+If you want a specific version you can include a filter, for example:
+
+- `pip install "mlforecast==0.3.0"` to install the 0.3.0 version
+- `pip install "mlforecast<0.4.0"` to install any version prior to
+ 0.4.0
+
+#### Extras
+
+##### polars
+
+Using polars dataframes: `pip install "mlforecast[polars]"`
+
+##### Saving to remote storages
+
+If you want to save your forecast artifacts to a remote storage like S3
+or GCS you can use the following extras:
+
+- Saving to S3: `pip install "mlforecast[aws]"`
+- Saving to Google Cloud Storage: `pip install "mlforecast[gcp]"`
+- Saving to Azure Data Lake: `pip install "mlforecast[azure]"`
+
+##### Distributed training
+
+If you want to perform distributed training you can use either dask, ray
+or spark. Once you know which framework you want to use you can include
+its extra:
+
+- dask: `pip install "mlforecast[dask]"`
+- ray: `pip install "mlforecast[ray]"`
+- spark: `pip install "mlforecast[spark]"`
+
+### Conda
+
+#### Latest release
+
+The mlforecast package is also published to
+[conda-forge](https://anaconda.org/conda-forge/mlforecast), which you
+can install by running the following in a terminal:
+
+`conda install -c conda-forge mlforecast`
+
+Note that this happens about a day later after it is published to PyPI,
+so you may have to wait to get the latest release.
+
+#### Specific version
+
+If you want a specific version you can include a filter, for example:
+
+- `conda install -c conda-forge "mlforecast==0.3.0"` to install the
+ 0.3.0 version
+- `conda install -c conda-forge "mlforecast<0.4.0"` to install any
+ version prior to 0.4.0
+
+## Development version
+
+If you want to try out a new feature that hasn’t made it into a release
+yet you have the following options:
+
+- Install from github:
+ `pip install git+https://github.com/Nixtla/mlforecast`
+- Clone and install:
+ `git clone https://github.com/Nixtla/mlforecast mlforecast-dev && pip install mlforecast-dev/`,
+ which will install the version from the current main branch.
+
diff --git a/mlforecast/docs/getting-started/quick_start_distributed.html.mdx b/mlforecast/docs/getting-started/quick_start_distributed.html.mdx
new file mode 100644
index 00000000..a0ea3dbf
--- /dev/null
+++ b/mlforecast/docs/getting-started/quick_start_distributed.html.mdx
@@ -0,0 +1,667 @@
+---
+description: Minimal example of distributed training with MLForecast
+output-file: quick_start_distributed.html
+title: Quick start (distributed)
+---
+
+
+The
+[`DistributedMLForecast`](https://Nixtla.github.io/mlforecast/distributed.forecast.html#distributedmlforecast)
+class is a high level abstraction that encapsulates all the steps in the
+pipeline (preprocessing, fitting the model and computing predictions)
+and applies them in a distributed way.
+
+The different things that you need to use
+[`DistributedMLForecast`](https://Nixtla.github.io/mlforecast/distributed.forecast.html#distributedmlforecast)
+(as opposed to
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast))
+are:
+
+1. You need to set up a cluster. We currently support dask, ray and
+ spark.
+2. Your data needs to be a distributed collection (dask, ray or spark
+ dataframe).
+3. You need to use a model that implements distributed training in your
+ framework of choice, e.g. SynapseML for LightGBM in spark.
+
+```python
+import platform
+import sys
+import tempfile
+
+import matplotlib.pyplot as plt
+import git
+import numpy as np
+import pandas as pd
+import s3fs
+from sklearn.base import BaseEstimator
+from utilsforecast.feature_engineering import fourier
+
+from mlforecast.distributed import DistributedMLForecast
+from mlforecast.lag_transforms import ExpandingMean, ExponentiallyWeightedMean, RollingMean
+from mlforecast.target_transforms import Differences
+from mlforecast.utils import generate_daily_series, generate_prices_for_series
+```
+
+## Dask
+
+```python
+import dask.dataframe as dd
+from dask.distributed import Client
+```
+
+### Client setup
+
+```python
+client = Client(n_workers=2, threads_per_worker=1)
+```
+
+Here we define a client that connects to a
+`dask.distributed.LocalCluster`, however it could be any other kind of
+cluster.
+
+### Data setup
+
+For dask, the data must be a `dask.dataframe.DataFrame`. You need to
+make sure that each time serie is only in one partition and it is
+recommended that you have as many partitions as you have workers. If you
+have more partitions than workers make sure to set `num_threads=1` to
+avoid having nested parallelism.
+
+The required input format is the same as for
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast),
+except that it’s a `dask.dataframe.DataFrame` instead of a
+`pandas.Dataframe`.
+
+```python
+series = generate_daily_series(100, n_static_features=2, equal_ends=True, static_as_categorical=False, min_length=500, max_length=1_000)
+train, future = fourier(series, freq='d', season_length=7, k=2, h=7)
+npartitions = 10
+partitioned_series = dd.from_pandas(train.set_index('unique_id'), npartitions=npartitions) # make sure we split by the id_col
+partitioned_series = partitioned_series.map_partitions(lambda df: df.reset_index())
+partitioned_series['unique_id'] = partitioned_series['unique_id'].astype(str) # can't handle categoricals atm
+partitioned_series
+```
+
+| | unique_id | ds | y | static_0 | static_1 | sin1_7 | sin2_7 | cos1_7 | cos2_7 |
+|----|----|----|----|----|----|----|----|----|----|
+| npartitions=10 | | | | | | | | | |
+| id_00 | object | datetime64\[ns\] | float64 | int64 | int64 | float32 | float32 | float32 | float32 |
+| id_10 | ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| id_90 | ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| id_99 | ... | ... | ... | ... | ... | ... | ... | ... | ... |
+
+### Models
+
+In order to perform distributed forecasting, we need to use a model that
+is able to train in a distributed way using `dask`. The current
+implementations are in
+[`DaskLGBMForecast`](https://Nixtla.github.io/mlforecast/distributed.models.dask.lgb.html#dasklgbmforecast)
+and
+[`DaskXGBForecast`](https://Nixtla.github.io/mlforecast/distributed.models.dask.xgb.html#daskxgbforecast)
+which are just wrappers around the native implementations.
+
+```python
+from mlforecast.distributed.models.dask.lgb import DaskLGBMForecast
+from mlforecast.distributed.models.dask.xgb import DaskXGBForecast
+```
+
+
+```python
+models = [
+ DaskXGBForecast(random_state=0),
+ DaskLGBMForecast(random_state=0, verbosity=-1),
+]
+```
+
+### Training
+
+Once we have our models we instantiate a
+[`DistributedMLForecast`](https://Nixtla.github.io/mlforecast/distributed.forecast.html#distributedmlforecast)
+object defining our features. We can then call `fit` on this object
+passing our dask dataframe.
+
+```python
+fcst = DistributedMLForecast(
+ models=models,
+ freq='D',
+ target_transforms=[Differences([7])],
+ lags=[7],
+ lag_transforms={
+ 1: [ExpandingMean(), ExponentiallyWeightedMean(alpha=0.9)],
+ 7: [RollingMean(window_size=14)],
+ },
+ date_features=['dayofweek', 'month'],
+ num_threads=1,
+ engine=client,
+)
+fcst.fit(partitioned_series, static_features=['static_0', 'static_1'])
+```
+
+Once we have our fitted models we can compute the predictions for the
+next 7 timesteps.
+
+### Forecasting
+
+```python
+preds = fcst.predict(7, X_df=future).compute()
+preds.head()
+```
+
+| | unique_id | ds | DaskXGBForecast | DaskLGBMForecast |
+|-----|-----------|---------------------|-----------------|------------------|
+| 0 | id_00 | 2002-09-27 00:00:00 | 21.722841 | 21.725511 |
+| 1 | id_00 | 2002-09-28 00:00:00 | 84.918194 | 84.606362 |
+| 2 | id_00 | 2002-09-29 00:00:00 | 162.067624 | 163.36802 |
+| 3 | id_00 | 2002-09-30 00:00:00 | 249.001477 | 246.422894 |
+| 4 | id_00 | 2002-10-01 00:00:00 | 317.149512 | 315.538403 |
+
+### Saving and loading
+
+Once you’ve trained your model you can use the
+[`DistributedMLForecast.save`](https://Nixtla.github.io/mlforecast/distributed.forecast.html#distributedmlforecast.save)
+method to save the artifacts for inference. Keep in mind that if you’re
+on a remote cluster you should set a remote storage like S3 as the
+destination.
+
+mlforecast uses
+[fsspec](https://filesystem-spec.readthedocs.io/en/latest/) to handle
+the different filesystems, so if you’re using s3 for example you also
+need to install [s3fs](https://s3fs.readthedocs.io/en/latest/). If
+you’re using pip you can just include the aws extra,
+e.g. `pip install 'mlforecast[aws,dask]'`, which will install the
+required dependencies to perform distributed training with dask and
+saving to S3. If you’re using conda you’ll have to manually install them
+(`conda install dask fsspec fugue s3fs`).
+
+```python
+# define unique name for CI
+def build_unique_name(engine):
+ pyver = f'{sys.version_info.major}_{sys.version_info.minor}'
+ repo = git.Repo(search_parent_directories=True)
+ sha = repo.head.object.hexsha
+ return f'{sys.platform}-{pyver}-{engine}-{sha}'
+```
+
+
+```python
+save_dir = build_unique_name('dask')
+save_path = f's3://nixtla-tmp/mlf/{save_dir}'
+tmpdir = tempfile.TemporaryDirectory()
+try:
+ s3fs.S3FileSystem().ls('s3://nixtla-tmp/')
+ fcst.save(save_path)
+except Exception as e:
+ print(e)
+ save_path = f'{tmpdir.name}/{save_dir}'
+ fcst.save(save_path)
+```
+
+Once you’ve saved your forecast object you can then load it back by
+specifying the path where it was saved along with an engine, which will
+be used to perform the distributed computations (in this case the dask
+client).
+
+```python
+fcst2 = DistributedMLForecast.load(save_path, engine=client)
+```
+
+We can verify that this object produces the same results.
+
+```python
+preds = fa.as_pandas(fcst.predict(7, X_df=future)).sort_values(['unique_id', 'ds']).reset_index(drop=True)
+preds2 = fa.as_pandas(fcst2.predict(7, X_df=future)).sort_values(['unique_id', 'ds']).reset_index(drop=True)
+pd.testing.assert_frame_equal(preds, preds2)
+```
+
+### Converting to local
+
+Another option to store your distributed forecast object is to first
+turn it into a local one and then save it. Keep in mind that in order to
+do that all the remote data that is stored from the series will have to
+be pulled into a single machine (the scheduler in dask, driver in spark,
+etc.), so you have to be sure that it’ll fit in memory, it should
+consume about 2x the size of your target column (you can reduce this
+further by using the `keep_last_n` argument in the `fit` method).
+
+```python
+local_fcst = fcst.to_local()
+local_preds = local_fcst.predict(7, X_df=future)
+# we don't check the dtype because sometimes these are arrow dtypes
+# or different precisions of float
+pd.testing.assert_frame_equal(preds, local_preds, check_dtype=False)
+```
+
+### Cross validation
+
+```python
+cv_res = fcst.cross_validation(
+ partitioned_series,
+ n_windows=3,
+ h=14,
+ static_features=['static_0', 'static_1'],
+)
+```
+
+
+```python
+cv_res.compute().head()
+```
+
+| | unique_id | ds | DaskXGBForecast | DaskLGBMForecast | cutoff | y |
+|----|----|----|----|----|----|----|
+| 61 | id_04 | 2002-08-21 00:00:00 | 68.3418 | 68.944539 | 2002-08-15 00:00:00 | 69.699857 |
+| 83 | id_15 | 2002-08-29 00:00:00 | 199.315403 | 199.663555 | 2002-08-15 00:00:00 | 206.082864 |
+| 103 | id_17 | 2002-08-21 00:00:00 | 156.822598 | 158.018246 | 2002-08-15 00:00:00 | 152.227984 |
+| 61 | id_24 | 2002-08-21 00:00:00 | 136.598356 | 136.576865 | 2002-08-15 00:00:00 | 138.559945 |
+| 36 | id_33 | 2002-08-24 00:00:00 | 95.6072 | 96.249354 | 2002-08-15 00:00:00 | 102.068997 |
+
+```python
+client.close()
+```
+
+## Spark
+
+### Session setup
+
+```python
+from pyspark.sql import SparkSession
+```
+
+
+```python
+spark = (
+ SparkSession
+ .builder
+ .config("spark.jars.packages", "com.microsoft.azure:synapseml_2.12:0.10.2")
+ .config("spark.jars.repositories", "https://mmlspark.azureedge.net/maven")
+ .getOrCreate()
+)
+```
+
+### Data setup
+
+For spark, the data must be a `pyspark DataFrame`. You need to make sure
+that each time serie is only in one partition (which you can do using
+`repartitionByRange`, for example) and it is recommended that you have
+as many partitions as you have workers. If you have more partitions than
+workers make sure to set `num_threads=1` to avoid having nested
+parallelism.
+
+The required input format is the same as for
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast),
+i.e. it should have at least an id column, a time column and a target
+column.
+
+```python
+series = generate_daily_series(100, n_static_features=2, equal_ends=True, static_as_categorical=False, min_length=500, max_length=1_000)
+series['unique_id'] = series['unique_id'].astype(str) # can't handle categoricals atm
+train, future = fourier(series, freq='d', season_length=7, k=2, h=7)
+numPartitions = 4
+spark_series = spark.createDataFrame(train).repartitionByRange(numPartitions, 'unique_id')
+```
+
+### Models
+
+In order to perform distributed forecasting, we need to use a model that
+is able to train in a distributed way using `spark`. The current
+implementations are in
+[`SparkLGBMForecast`](https://Nixtla.github.io/mlforecast/distributed.models.spark.lgb.html#sparklgbmforecast)
+and
+[`SparkXGBForecast`](https://Nixtla.github.io/mlforecast/distributed.models.spark.xgb.html#sparkxgbforecast)
+which are just wrappers around the native implementations.
+
+```python
+from mlforecast.distributed.models.spark.lgb import SparkLGBMForecast
+from mlforecast.distributed.models.spark.xgb import SparkXGBForecast
+```
+
+
+```python
+models = [
+ SparkLGBMForecast(seed=0, verbosity=-1),
+ SparkXGBForecast(random_state=0),
+]
+```
+
+### Training
+
+```python
+fcst = DistributedMLForecast(
+ models,
+ freq='D',
+ target_transforms=[Differences([7])],
+ lags=[1],
+ lag_transforms={
+ 1: [ExpandingMean(), ExponentiallyWeightedMean(alpha=0.9)],
+ },
+ date_features=['dayofweek'],
+)
+fcst.fit(
+ spark_series,
+ static_features=['static_0', 'static_1'],
+)
+```
+
+### Forecasting
+
+```python
+preds = fcst.predict(7, X_df=future).toPandas()
+```
+
+``` text
+
+```
+
+```python
+preds.head()
+```
+
+| | unique_id | ds | SparkLGBMForecast | SparkXGBForecast |
+|-----|-----------|------------|-------------------|------------------|
+| 0 | id_00 | 2002-09-27 | 15.053577 | 18.631477 |
+| 1 | id_00 | 2002-09-28 | 93.010037 | 93.796269 |
+| 2 | id_00 | 2002-09-29 | 160.120148 | 159.582315 |
+| 3 | id_00 | 2002-09-30 | 250.445885 | 250.861651 |
+| 4 | id_00 | 2002-10-01 | 323.335956 | 321.564089 |
+
+### Saving and loading
+
+Once you’ve trained your model you can use the
+[`DistributedMLForecast.save`](https://Nixtla.github.io/mlforecast/distributed.forecast.html#distributedmlforecast.save)
+method to save the artifacts for inference. Keep in mind that if you’re
+on a remote cluster you should set a remote storage like S3 as the
+destination.
+
+mlforecast uses
+[fsspec](https://filesystem-spec.readthedocs.io/en/latest/) to handle
+the different filesystems, so if you’re using s3 for example you also
+need to install [s3fs](https://s3fs.readthedocs.io/en/latest/). If
+you’re using pip you can just include the aws extra,
+e.g. `pip install 'mlforecast[aws,spark]'`, which will install the
+required dependencies to perform distributed training with spark and
+saving to S3. If you’re using conda you’ll have to manually install them
+(`conda install fsspec fugue pyspark s3fs`).
+
+```python
+save_dir = build_unique_name('spark')
+save_path = f's3://nixtla-tmp/mlf/{save_dir}'
+try:
+ s3fs.S3FileSystem().ls('s3://nixtla-tmp/')
+ fcst.save(save_path)
+except Exception as e:
+ print(e)
+ save_path = f'{tmpdir.name}/{save_dir}'
+ fcst.save(save_path)
+```
+
+``` text
+
+```
+
+Once you’ve saved your forecast object you can then load it back by
+specifying the path where it was saved along with an engine, which will
+be used to perform the distributed computations (in this case the spark
+session).
+
+```python
+fcst2 = DistributedMLForecast.load(save_path, engine=spark)
+```
+
+``` text
+
+```
+
+We can verify that this object produces the same results.
+
+```python
+preds = fa.as_pandas(fcst.predict(7, X_df=future)).sort_values(['unique_id', 'ds']).reset_index(drop=True)
+preds2 = fa.as_pandas(fcst2.predict(7, X_df=future)).sort_values(['unique_id', 'ds']).reset_index(drop=True)
+pd.testing.assert_frame_equal(preds, preds2)
+```
+
+``` text
+
+```
+
+### Converting to local
+
+Another option to store your distributed forecast object is to first
+turn it into a local one and then save it. Keep in mind that in order to
+do that all the remote data that is stored from the series will have to
+be pulled into a single machine (the scheduler in dask, driver in spark,
+etc.), so you have to be sure that it’ll fit in memory, it should
+consume about 2x the size of your target column (you can reduce this
+further by using the `keep_last_n` argument in the `fit` method).
+
+```python
+local_fcst = fcst.to_local()
+local_preds = local_fcst.predict(7, X_df=future)
+# we don't check the dtype because sometimes these are arrow dtypes
+# or different precisions of float
+pd.testing.assert_frame_equal(preds, local_preds, check_dtype=False)
+```
+
+### Cross validation
+
+```python
+cv_res = fcst.cross_validation(
+ spark_series,
+ n_windows=3,
+ h=14,
+ static_features=['static_0', 'static_1'],
+).toPandas()
+```
+
+
+```python
+cv_res.head()
+```
+
+| | unique_id | ds | SparkLGBMForecast | SparkXGBForecast | cutoff | y |
+|----|----|----|----|----|----|----|
+| 0 | id_03 | 2002-08-18 | 3.272922 | 3.348874 | 2002-08-15 | 3.060194 |
+| 1 | id_09 | 2002-08-20 | 402.718091 | 402.622501 | 2002-08-15 | 398.784459 |
+| 2 | id_25 | 2002-08-22 | 87.189811 | 86.891632 | 2002-08-15 | 82.731377 |
+| 3 | id_06 | 2002-08-21 | 20.416790 | 20.478502 | 2002-08-15 | 19.196394 |
+| 4 | id_22 | 2002-08-23 | 357.718513 | 360.502024 | 2002-08-15 | 394.770699 |
+
+```python
+spark.stop()
+```
+
+## Ray
+
+### Session setup
+
+```python
+import ray
+from ray.cluster_utils import Cluster
+```
+
+
+```python
+ray_cluster = Cluster(
+ initialize_head=True,
+ head_node_args={"num_cpus": 2}
+)
+ray.init(address=ray_cluster.address, ignore_reinit_error=True)
+# add mock node to simulate a cluster
+mock_node = ray_cluster.add_node(num_cpus=2)
+```
+
+### Data setup
+
+For ray, the data must be a `ray DataFrame`. It is recommended that you
+have as many partitions as you have workers. If you have more partitions
+than workers make sure to set `num_threads=1` to avoid having nested
+parallelism.
+
+The required input format is the same as for
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast),
+i.e. it should have at least an id column, a time column and a target
+column.
+
+```python
+series = generate_daily_series(100, n_static_features=2, equal_ends=True, static_as_categorical=False, min_length=500, max_length=1_000)
+series['unique_id'] = series['unique_id'].astype(str) # can't handle categoricals atm
+train, future = fourier(series, freq='d', season_length=7, k=2, h=7)
+ray_series = ray.data.from_pandas(train)
+```
+
+### Models
+
+The ray integration allows to include `lightgbm` (`RayLGBMRegressor`),
+and `xgboost` (`RayXGBRegressor`).
+
+```python
+from mlforecast.distributed.models.ray.lgb import RayLGBMForecast
+from mlforecast.distributed.models.ray.xgb import RayXGBForecast
+```
+
+
+```python
+models = [
+ RayLGBMForecast(random_state=0, verbosity=-1),
+ RayXGBForecast(random_state=0),
+]
+```
+
+### Training
+
+To control the number of partitions to use using Ray, we have to include
+`num_partitions` to
+[`DistributedMLForecast`](https://Nixtla.github.io/mlforecast/distributed.forecast.html#distributedmlforecast).
+
+```python
+num_partitions = 4
+fcst = DistributedMLForecast(
+ models,
+ freq='D',
+ target_transforms=[Differences([7])],
+ lags=[1],
+ lag_transforms={
+ 1: [ExpandingMean(), ExponentiallyWeightedMean(alpha=0.9)],
+ },
+ date_features=['dayofweek'],
+ num_partitions=num_partitions, # Use num_partitions to reduce overhead
+)
+fcst.fit(
+ ray_series,
+ static_features=['static_0', 'static_1'],
+)
+```
+
+### Forecasting
+
+```python
+preds = fcst.predict(7, X_df=future).to_pandas()
+```
+
+
+```python
+preds.head()
+```
+
+| | unique_id | ds | RayLGBMForecast | RayXGBForecast |
+|-----|-----------|------------|-----------------|----------------|
+| 0 | id_00 | 2002-09-27 | 15.232455 | 10.38301 |
+| 1 | id_00 | 2002-09-28 | 92.288994 | 92.531502 |
+| 2 | id_00 | 2002-09-29 | 160.043472 | 160.722885 |
+| 3 | id_00 | 2002-09-30 | 250.03212 | 252.821899 |
+| 4 | id_00 | 2002-10-01 | 322.905182 | 324.387695 |
+
+### Saving and loading
+
+Once you’ve trained your model you can use the
+[`DistributedMLForecast.save`](https://Nixtla.github.io/mlforecast/distributed.forecast.html#distributedmlforecast.save)
+method to save the artifacts for inference. Keep in mind that if you’re
+on a remote cluster you should set a remote storage like S3 as the
+destination.
+
+mlforecast uses
+[fsspec](https://filesystem-spec.readthedocs.io/en/latest/) to handle
+the different filesystems, so if you’re using s3 for example you also
+need to install [s3fs](https://s3fs.readthedocs.io/en/latest/). If
+you’re using pip you can just include the aws extra,
+e.g. `pip install 'mlforecast[aws,ray]'`, which will install the
+required dependencies to perform distributed training with ray and
+saving to S3. If you’re using conda you’ll have to manually install them
+(`conda install fsspec fugue ray s3fs`).
+
+```python
+save_dir = build_unique_name('ray')
+save_path = f's3://nixtla-tmp/mlf/{save_dir}'
+try:
+ s3fs.S3FileSystem().ls('s3://nixtla-tmp/')
+ fcst.save(save_path)
+except Exception as e:
+ print(e)
+ save_path = f'{tmpdir.name}/{save_dir}'
+ fcst.save(save_path)
+```
+
+Once you’ve saved your forecast object you can then load it back by
+specifying the path where it was saved along with an engine, which will
+be used to perform the distributed computations (in this case the ‘ray’
+string).
+
+```python
+fcst2 = DistributedMLForecast.load(save_path, engine='ray')
+```
+
+We can verify that this object produces the same results.
+
+```python
+preds = fa.as_pandas(fcst.predict(7, X_df=future)).sort_values(['unique_id', 'ds']).reset_index(drop=True)
+preds2 = fa.as_pandas(fcst2.predict(7, X_df=future)).sort_values(['unique_id', 'ds']).reset_index(drop=True)
+pd.testing.assert_frame_equal(preds, preds2)
+```
+
+### Converting to local
+
+Another option to store your distributed forecast object is to first
+turn it into a local one and then save it. Keep in mind that in order to
+do that all the remote data that is stored from the series will have to
+be pulled into a single machine (the scheduler in dask, driver in spark,
+etc.), so you have to be sure that it’ll fit in memory, it should
+consume about 2x the size of your target column (you can reduce this
+further by using the `keep_last_n` argument in the `fit` method).
+
+```python
+local_fcst = fcst.to_local()
+local_preds = local_fcst.predict(7, X_df=future)
+# we don't check the dtype because sometimes these are arrow dtypes
+# or different precisions of float
+pd.testing.assert_frame_equal(preds, local_preds, check_dtype=False)
+```
+
+### Cross validation
+
+```python
+cv_res = fcst.cross_validation(
+ ray_series,
+ n_windows=3,
+ h=14,
+ static_features=['static_0', 'static_1'],
+).to_pandas()
+```
+
+
+```python
+cv_res.head()
+```
+
+| | unique_id | ds | RayLGBMForecast | RayXGBForecast | cutoff | y |
+|-----|-----------|------------|-----------------|----------------|------------|------------|
+| 0 | id_05 | 2002-09-21 | 108.285187 | 108.619698 | 2002-09-12 | 108.726387 |
+| 1 | id_08 | 2002-09-16 | 26.287956 | 26.589603 | 2002-09-12 | 27.980670 |
+| 2 | id_08 | 2002-09-25 | 83.210945 | 84.194962 | 2002-09-12 | 86.344885 |
+| 3 | id_11 | 2002-09-22 | 416.994843 | 417.106506 | 2002-09-12 | 425.434661 |
+| 4 | id_16 | 2002-09-14 | 377.916382 | 375.421600 | 2002-09-12 | 400.361977 |
+
+```python
+ray.shutdown()
+```
+
diff --git a/mlforecast/docs/getting-started/quick_start_local.html.mdx b/mlforecast/docs/getting-started/quick_start_local.html.mdx
new file mode 100644
index 00000000..440759ec
--- /dev/null
+++ b/mlforecast/docs/getting-started/quick_start_local.html.mdx
@@ -0,0 +1,151 @@
+---
+description: Minimal example of MLForecast
+output-file: quick_start_local.html
+title: Quick start (local)
+---
+
+
+## Main concepts
+
+The main component of mlforecast is the
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+class, which abstracts away:
+
+- Feature engineering and model training through
+ [`MLForecast.fit`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.fit)
+- Feature updates and multi step ahead predictions through
+ [`MLForecast.predict`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.predict)
+
+## Data format
+
+The data is expected to be a pandas dataframe in long format, that is,
+each row represents an observation of a single serie at a given time,
+with at least three columns:
+
+- `id_col`: column that identifies each serie.
+- `target_col`: column that has the series values at each timestamp.
+- `time_col`: column that contains the time the series value was
+ observed. These are usually timestamps, but can also be consecutive
+ integers.
+
+Here we present an example using the classic Box & Jenkins airline data,
+which measures monthly totals of international airline passengers from
+1949 to 1960. Source: Box, G. E. P., Jenkins, G. M. and Reinsel, G. C.
+(1976) Time Series Analysis, Forecasting and Control. Third Edition.
+Holden-Day. Series G.
+
+```python
+import pandas as pd
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+df = pd.read_csv('https://datasets-nixtla.s3.amazonaws.com/air-passengers.csv', parse_dates=['ds'])
+df.head()
+```
+
+| | unique_id | ds | y |
+|-----|---------------|------------|-----|
+| 0 | AirPassengers | 1949-01-01 | 112 |
+| 1 | AirPassengers | 1949-02-01 | 118 |
+| 2 | AirPassengers | 1949-03-01 | 132 |
+| 3 | AirPassengers | 1949-04-01 | 129 |
+| 4 | AirPassengers | 1949-05-01 | 121 |
+
+```python
+df['unique_id'].value_counts()
+```
+
+``` text
+AirPassengers 144
+Name: unique_id, dtype: int64
+```
+
+Here the `unique_id` column has the same value for all rows because this
+is a single time series, you can have multiple time series by stacking
+them together and having a column that differentiates them.
+
+We also have the `ds` column that contains the timestamps, in this case
+with a monthly frequency, and the `y` column that contains the series
+values in each timestamp.
+
+## Modeling
+
+```python
+fig = plot_series(df)
+```
+
+
+
+We can see that the serie has a clear trend, so we can take the first
+difference, i.e. take each value and subtract the value at the previous
+month. This can be achieved by passing an
+`mlforecast.target_transforms.Differences([1])` instance to
+`target_transforms`.
+
+We can then train a linear regression using the value from the same
+month at the previous year (lag 12) as a feature, this is done by
+passing `lags=[12]`.
+
+```python
+from mlforecast import MLForecast
+from mlforecast.target_transforms import Differences
+from sklearn.linear_model import LinearRegression
+```
+
+
+```python
+fcst = MLForecast(
+ models=LinearRegression(),
+ freq='MS', # our serie has a monthly frequency
+ lags=[12],
+ target_transforms=[Differences([1])],
+)
+fcst.fit(df)
+```
+
+``` text
+MLForecast(models=[LinearRegression], freq=MS, lag_features=['lag12'], date_features=[], num_threads=1)
+```
+
+The previous line computed the features and trained the model, so now
+we’re ready to compute our forecasts.
+
+## Forecasting
+
+Compute the forecast for the next 12 months
+
+```python
+preds = fcst.predict(12)
+preds
+```
+
+| | unique_id | ds | LinearRegression |
+|-----|---------------|------------|------------------|
+| 0 | AirPassengers | 1961-01-01 | 444.656555 |
+| 1 | AirPassengers | 1961-02-01 | 417.470734 |
+| 2 | AirPassengers | 1961-03-01 | 446.903046 |
+| 3 | AirPassengers | 1961-04-01 | 491.014130 |
+| 4 | AirPassengers | 1961-05-01 | 502.622223 |
+| 5 | AirPassengers | 1961-06-01 | 568.751465 |
+| 6 | AirPassengers | 1961-07-01 | 660.044312 |
+| 7 | AirPassengers | 1961-08-01 | 643.343323 |
+| 8 | AirPassengers | 1961-09-01 | 540.666687 |
+| 9 | AirPassengers | 1961-10-01 | 491.462708 |
+| 10 | AirPassengers | 1961-11-01 | 417.095154 |
+| 11 | AirPassengers | 1961-12-01 | 461.206238 |
+
+## Visualize results
+
+We can visualize what our prediction looks like.
+
+```python
+fig = plot_series(df, preds)
+```
+
+
+
+And that’s it! You’ve trained a linear regression to predict the air
+passengers for 1961.
+
diff --git a/mlforecast/docs/how-to-guides/analyzing_models.html.mdx b/mlforecast/docs/how-to-guides/analyzing_models.html.mdx
new file mode 100644
index 00000000..136bbec1
--- /dev/null
+++ b/mlforecast/docs/how-to-guides/analyzing_models.html.mdx
@@ -0,0 +1,204 @@
+---
+description: Access and interpret the models after fitting
+output-file: analyzing_models.html
+title: Analyzing the trained models
+---
+
+
+## Data setup
+
+```python
+from mlforecast.utils import generate_daily_series
+```
+
+
+```python
+series = generate_daily_series(10)
+series.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|----------|
+| 0 | id_0 | 2000-01-01 | 0.322947 |
+| 1 | id_0 | 2000-01-02 | 1.218794 |
+| 2 | id_0 | 2000-01-03 | 2.445887 |
+| 3 | id_0 | 2000-01-04 | 3.481831 |
+| 4 | id_0 | 2000-01-05 | 4.191721 |
+
+## Training
+
+Suppose that you want to train a linear regression model using the day
+of the week and lag1 as features.
+
+```python
+from sklearn.linear_model import LinearRegression
+
+from mlforecast import MLForecast
+```
+
+
+```python
+fcst = MLForecast(
+ freq='D',
+ models={'lr': LinearRegression()},
+ lags=[1],
+ date_features=['dayofweek'],
+)
+```
+
+
+```python
+fcst.fit(series)
+```
+
+``` text
+MLForecast(models=[lr], freq=, lag_features=['lag1'], date_features=['dayofweek'], num_threads=1)
+```
+
+What
+[`MLForecast.fit`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.fit)
+does is save the required data for the predict step and also train the
+models (in this case the linear regression). The trained models are
+available in the `MLForecast.models_` attribute, which is a dictionary
+where the keys are the model names and the values are the model
+themselves.
+
+```python
+fcst.models_
+```
+
+``` text
+{'lr': LinearRegression()}
+```
+
+## Inspect parameters
+
+We can access the linear regression coefficients in the following way:
+
+```python
+fcst.models_['lr'].intercept_, fcst.models_['lr'].coef_
+```
+
+``` text
+(3.2476337167384415, array([ 0.19896416, -0.21441331]))
+```
+
+## SHAP
+
+```python
+import shap
+```
+
+### Training set
+
+If you need to generate the training data you can use
+[`MLForecast.preprocess`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.preprocess).
+
+```python
+prep = fcst.preprocess(series)
+prep.head()
+```
+
+| | unique_id | ds | y | lag1 | dayofweek |
+|-----|-----------|------------|----------|----------|-----------|
+| 1 | id_0 | 2000-01-02 | 1.218794 | 0.322947 | 6 |
+| 2 | id_0 | 2000-01-03 | 2.445887 | 1.218794 | 0 |
+| 3 | id_0 | 2000-01-04 | 3.481831 | 2.445887 | 1 |
+| 4 | id_0 | 2000-01-05 | 4.191721 | 3.481831 | 2 |
+| 5 | id_0 | 2000-01-06 | 5.395863 | 4.191721 | 3 |
+
+We extract the X, which involves dropping the info columns (id + times)
+and the target
+
+```python
+X = prep.drop(columns=['unique_id', 'ds', 'y'])
+X.head()
+```
+
+| | lag1 | dayofweek |
+|-----|----------|-----------|
+| 1 | 0.322947 | 6 |
+| 2 | 1.218794 | 0 |
+| 3 | 2.445887 | 1 |
+| 4 | 3.481831 | 2 |
+| 5 | 4.191721 | 3 |
+
+We can now compute the shap values
+
+```python
+X100 = shap.utils.sample(X, 100)
+explainer = shap.Explainer(fcst.models_['lr'].predict, X100)
+shap_values = explainer(X)
+```
+
+And visualize them
+
+```python
+shap.plots.beeswarm(shap_values)
+```
+
+
+
+### Predictions
+
+Sometimes you want to determine why the model gave a specific
+prediction. In order to do this you need the input features, which
+aren’t returned by default, but you can retrieve them using a callback.
+
+```python
+from mlforecast.callbacks import SaveFeatures
+```
+
+
+```python
+save_feats = SaveFeatures()
+preds = fcst.predict(1, before_predict_callback=save_feats)
+preds.head()
+```
+
+| | unique_id | ds | lr |
+|-----|-----------|------------|----------|
+| 0 | id_0 | 2000-08-10 | 3.468643 |
+| 1 | id_1 | 2000-04-07 | 3.016877 |
+| 2 | id_2 | 2000-06-16 | 2.815249 |
+| 3 | id_3 | 2000-08-30 | 4.048894 |
+| 4 | id_4 | 2001-01-08 | 3.524532 |
+
+You can now retrieve the features by using
+[`SaveFeatures.get_features`](https://Nixtla.github.io/mlforecast/callbacks.html#savefeatures.get_features)
+
+```python
+features = save_feats.get_features()
+features.head()
+```
+
+| | lag1 | dayofweek |
+|-----|----------|-----------|
+| 0 | 4.343744 | 3 |
+| 1 | 3.150799 | 4 |
+| 2 | 2.137412 | 4 |
+| 3 | 6.182456 | 2 |
+| 4 | 1.391698 | 0 |
+
+And use those features to compute the shap values.
+
+```python
+shap_values_predictions = explainer(features)
+```
+
+We can now analyze what influenced the prediction for `'id_4'`.
+
+```python
+round(preds.loc[4, 'lr'], 3)
+```
+
+``` text
+3.525
+```
+
+```python
+shap.plots.waterfall(shap_values_predictions[4])
+```
+
+
+
diff --git a/mlforecast/docs/how-to-guides/analyzing_models_files/figure-markdown_strict/cell-13-output-1.png b/mlforecast/docs/how-to-guides/analyzing_models_files/figure-markdown_strict/cell-13-output-1.png
new file mode 100644
index 00000000..6ea6930a
Binary files /dev/null and b/mlforecast/docs/how-to-guides/analyzing_models_files/figure-markdown_strict/cell-13-output-1.png differ
diff --git a/mlforecast/docs/how-to-guides/analyzing_models_files/figure-markdown_strict/cell-19-output-1.png b/mlforecast/docs/how-to-guides/analyzing_models_files/figure-markdown_strict/cell-19-output-1.png
new file mode 100644
index 00000000..f0d925cc
Binary files /dev/null and b/mlforecast/docs/how-to-guides/analyzing_models_files/figure-markdown_strict/cell-19-output-1.png differ
diff --git a/mlforecast/docs/how-to-guides/cross_validation.html.mdx b/mlforecast/docs/how-to-guides/cross_validation.html.mdx
new file mode 100644
index 00000000..e6afac8b
--- /dev/null
+++ b/mlforecast/docs/how-to-guides/cross_validation.html.mdx
@@ -0,0 +1,237 @@
+---
+description: >-
+ In this example, we'll implement time series cross-validation to evaluate
+ model's performance.
+output-file: cross_validation.html
+title: Cross validation
+---
+
+
+> **Prerequesites**
+>
+> This tutorial assumes basic familiarity with
+> [`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast).
+> For a minimal example visit the [Quick Start](quick_start_local.html)
+
+## Introduction
+
+Time series cross-validation is a method for evaluating how a model
+would have performed in the past. It works by defining a sliding window
+across the historical data and predicting the period following it.
+
+
+
+[MLForecast](https://nixtla.github.io/mlforecast/) has an implementation
+of time series cross-validation that is fast and easy to use. This
+implementation makes cross-validation a efficient operation, which makes
+it less time-consuming. In this notebook, we’ll use it on a subset of
+the [M4
+Competition](https://www.sciencedirect.com/science/article/pii/S0169207019301128)
+hourly dataset.
+
+**Outline:**
+
+1. Install libraries
+2. Load and explore data
+3. Train model
+4. Perform time series cross-validation
+5. Evaluate results
+
+> **Tip**
+>
+> You can use Colab to run this Notebook interactively
+>
+
+## Install libraries
+
+We assume that you have
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+already installed. If not, check this guide for instructions on [how to
+install MLForecast](../getting-started/install.html).
+
+Install the necessary packages with `pip install mlforecast`.
+
+```python
+import pandas as pd
+
+from utilsforecast.plotting import plot_series
+
+from mlforecast import MLForecast # required to instantiate MLForecast object and use cross-validation method
+```
+
+## Load and explore the data
+
+As stated in the introduction, we’ll use the M4 Competition hourly
+dataset. We’ll first import the data from an URL using `pandas`.
+
+```python
+Y_df = pd.read_csv('https://datasets-nixtla.s3.amazonaws.com/m4-hourly.csv') # load the data
+Y_df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|-----|-------|
+| 0 | H1 | 1 | 605.0 |
+| 1 | H1 | 2 | 586.0 |
+| 2 | H1 | 3 | 586.0 |
+| 3 | H1 | 4 | 559.0 |
+| 4 | H1 | 5 | 511.0 |
+
+The input to
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+is a data frame in [long
+format](https://www.theanalysisfactor.com/wide-and-long-data/) with
+three columns: `unique_id`, `ds` and `y`:
+
+- The `unique_id` (string, int, or category) represents an identifier
+ for the series.
+- The `ds` (datestamp or int) column should be either an integer
+ indexing time or a datestamp in format YYYY-MM-DD or YYYY-MM-DD
+ HH:MM:SS.
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+The data in this example already has this format, so no changes are
+needed.
+
+We can plot the time series we’ll work with using the following
+function.
+
+```python
+fig = plot_series(Y_df, max_ids=4, plot_random=False, max_insample_length=24 * 14)
+```
+
+
+
+## Define forecast object
+
+For this example, we’ll use LightGBM. We first need to import it and
+then we need to instantiate a new
+[MLForecast](../../forecast.html#mlforecast) object.
+
+In this example, we are only using `differences` and `lags` to produce
+features. See [the full
+documentation](https://nixtla.github.io/mlforecast) to see all available
+features.
+
+Any settings are passed into the constructor. Then you call its `fit`
+method and pass in the historical data frame `df`.
+
+```python
+import lightgbm as lgb
+from mlforecast.target_transforms import Differences
+```
+
+
+```python
+models = [lgb.LGBMRegressor(verbosity=-1)]
+
+mlf = MLForecast(
+ models=models,
+ freq=1,# our series have integer timestamps, so we'll just add 1 in every timeste,
+ target_transforms=[Differences([24])],
+ lags=range(1, 25)
+)
+```
+
+## Perform time series cross-validation
+
+Once the
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+object has been instantiated, we can use the [cross_validation
+method](../../forecast.html#mlforecast.cross_validation).
+
+For this particular example, we’ll use 3 windows of 24 hours.
+
+```python
+cv_df = mlf.cross_validation(
+ df=Y_df,
+ h=24,
+ n_windows=3,
+)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id`: identifies each time series.
+- `ds`: datestamp or temporal index.
+- `cutoff`: the last datestamp or temporal index for the `n_windows`.
+- `y`: true value
+- `"model"`: columns with the model’s name and fitted value.
+
+```python
+cv_df.head()
+```
+
+| | unique_id | ds | cutoff | y | LGBMRegressor |
+|-----|-----------|-----|--------|-------|---------------|
+| 0 | H1 | 677 | 676 | 691.0 | 673.703191 |
+| 1 | H1 | 678 | 676 | 618.0 | 552.306270 |
+| 2 | H1 | 679 | 676 | 563.0 | 541.778027 |
+| 3 | H1 | 680 | 676 | 529.0 | 502.778027 |
+| 4 | H1 | 681 | 676 | 504.0 | 480.778027 |
+
+We’ll now plot the forecast for each cutoff period.
+
+```python
+import matplotlib.pyplot as plt
+```
+
+
+```python
+def plot_cv(df, df_cv, uid, fname, last_n=24 * 14):
+ cutoffs = df_cv.query('unique_id == @uid')['cutoff'].unique()
+ fig, ax = plt.subplots(nrows=len(cutoffs), ncols=1, figsize=(14, 6), gridspec_kw=dict(hspace=0.8))
+ for cutoff, axi in zip(cutoffs, ax.flat):
+ df.query('unique_id == @uid').tail(last_n).set_index('ds').plot(ax=axi, title=uid, y='y')
+ df_cv.query('unique_id == @uid & cutoff == @cutoff').set_index('ds').plot(ax=axi, title=uid, y='LGBMRegressor')
+ fig.savefig(fname, bbox_inches='tight')
+ plt.close()
+```
+
+
+```python
+plot_cv(Y_df, cv_df, 'H1', '../../figs/cross_validation__predictions.png')
+```
+
+
+
+Notice that in each cutoff period, we generated a forecast for the next
+24 hours using only the data `y` before said period.
+
+## Evaluate results
+
+We can now compute the accuracy of the forecast using an appropiate
+accuracy metric. Here we’ll use the [Root Mean Squared Error
+(RMSE).](https://en.wikipedia.org/wiki/Root-mean-square_deviation) To do
+this, we can use `utilsforecast`, a Python library developed by Nixtla
+that includes a function to compute the RMSE.
+
+```python
+from utilsforecast.evaluation import evaluate
+from utilsforecast.losses import rmse
+```
+
+
+```python
+cv_rmse = evaluate(
+ cv_df.drop(columns='cutoff'),
+ metrics=[rmse],
+ agg_fn='mean',
+)
+print(f"RMSE using cross-validation: {cv_rmse['LGBMRegressor'].item():.1f}")
+```
+
+``` text
+RMSE using cross-validation: 269.0
+```
+
+This measure should better reflect the predictive abilities of our
+model, since it used different time periods to test its accuracy.
+
+## References
+
+[Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+principles and practice, Time series
+cross-validation”](https://otexts.com/fpp3/tscv.html).
+
diff --git a/mlforecast/docs/how-to-guides/custom_date_features.html.mdx b/mlforecast/docs/how-to-guides/custom_date_features.html.mdx
new file mode 100644
index 00000000..995eeafa
--- /dev/null
+++ b/mlforecast/docs/how-to-guides/custom_date_features.html.mdx
@@ -0,0 +1,56 @@
+---
+description: Define your own functions to be used as date features
+output-file: custom_date_features.html
+title: Custom date features
+---
+
+
+```python
+from mlforecast import MLForecast
+from mlforecast.utils import generate_daily_series
+```
+
+The `date_features` argument of MLForecast can take pandas date
+attributes as well as functions that take a [pandas
+DatetimeIndex](https://pandas.pydata.org/docs/reference/api/pandas.DatetimeIndex.html)
+and return a numeric value. The name of the function is used as the name
+of the feature, so please use unique and descriptive names.
+
+```python
+series = generate_daily_series(1, min_length=6, max_length=6)
+```
+
+
+```python
+def even_day(dates):
+ """Day of month is even"""
+ return dates.day % 2 == 0
+
+def month_start_or_end(dates):
+ """Date is month start or month end"""
+ return dates.is_month_start | dates.is_month_end
+
+def is_monday(dates):
+ """Date is monday"""
+ return dates.dayofweek == 0
+```
+
+
+```python
+fcst = MLForecast(
+ [],
+ freq='D',
+ date_features=['dayofweek', 'dayofyear', even_day, month_start_or_end, is_monday]
+)
+fcst.preprocess(series)
+```
+
+| | unique_id | ds | y | dayofweek | dayofyear | even_day | month_start_or_end | is_monday |
+|----|----|----|----|----|----|----|----|----|
+| 0 | id_0 | 2000-01-01 | 0.274407 | 5 | 1 | False | True | False |
+| 1 | id_0 | 2000-01-02 | 1.357595 | 6 | 2 | True | False | False |
+| 2 | id_0 | 2000-01-03 | 2.301382 | 0 | 3 | False | False | True |
+| 3 | id_0 | 2000-01-04 | 3.272442 | 1 | 4 | True | False | False |
+| 4 | id_0 | 2000-01-05 | 4.211827 | 2 | 5 | False | False | False |
+| 5 | id_0 | 2000-01-06 | 5.322947 | 3 | 6 | True | False | False |
+
diff --git a/mlforecast/docs/how-to-guides/custom_training.html.mdx b/mlforecast/docs/how-to-guides/custom_training.html.mdx
new file mode 100644
index 00000000..bd92fe8c
--- /dev/null
+++ b/mlforecast/docs/how-to-guides/custom_training.html.mdx
@@ -0,0 +1,149 @@
+---
+description: Customize the training procedure for your models
+output-file: custom_training.html
+title: Custom training
+---
+
+
+mlforecast abstracts away most of the training details, which is useful
+for iterating quickly. However, sometimes you want more control over the
+fit parameters, the data that goes into the model, etc. This guide shows
+how you can train a model in a specific way and then giving it back to
+mlforecast to produce forecasts with it.
+
+## Data setup
+
+```python
+from mlforecast.utils import generate_daily_series
+```
+
+
+```python
+series = generate_daily_series(5)
+```
+
+## Creating forecast object
+
+```python
+import lightgbm as lgb
+import numpy as np
+from sklearn.linear_model import LinearRegression
+
+from mlforecast import MLForecast
+```
+
+Suppose we want to train a linear regression with the default settings.
+
+```python
+fcst = MLForecast(
+ models={'lr': LinearRegression()},
+ freq='D',
+ lags=[1],
+ date_features=['dayofweek'],
+)
+```
+
+## Generate training set
+
+Use
+[`MLForecast.preprocess`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.preprocess)
+to generate the training data.
+
+```python
+prep = fcst.preprocess(series)
+prep.head()
+```
+
+| | unique_id | ds | y | lag1 | dayofweek |
+|-----|-----------|------------|----------|----------|-----------|
+| 1 | id_0 | 2000-01-02 | 1.423626 | 0.428973 | 6 |
+| 2 | id_0 | 2000-01-03 | 2.311782 | 1.423626 | 0 |
+| 3 | id_0 | 2000-01-04 | 3.192191 | 2.311782 | 1 |
+| 4 | id_0 | 2000-01-05 | 4.148767 | 3.192191 | 2 |
+| 5 | id_0 | 2000-01-06 | 5.028356 | 4.148767 | 3 |
+
+```python
+X = prep.drop(columns=['unique_id', 'ds', 'y'])
+y = prep['y']
+```
+
+## Regular training
+
+Since we don’t want to do anything special in our training process for
+the linear regression, we can just call
+[`MLForecast.fit_models`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.fit_models)
+
+```python
+fcst.fit_models(X, y)
+```
+
+``` text
+MLForecast(models=[lr], freq=D, lag_features=['lag1'], date_features=['dayofweek'], num_threads=1)
+```
+
+This has trained the linear regression model and is now available in the
+`MLForecast.models_` attribute.
+
+```python
+fcst.models_
+```
+
+``` text
+{'lr': LinearRegression()}
+```
+
+## Custom training
+
+Now suppose you also want to train a LightGBM model on the same data,
+but treating the day of the week as a categorical feature and logging
+the train loss.
+
+```python
+model = lgb.LGBMRegressor(n_estimators=100, verbosity=-1)
+model.fit(
+ X,
+ y,
+ eval_set=[(X, y)],
+ categorical_feature=['dayofweek'],
+ callbacks=[lgb.log_evaluation(20)],
+);
+```
+
+``` text
+[20] training's l2: 0.0823528
+[40] training's l2: 0.0230292
+[60] training's l2: 0.0207829
+[80] training's l2: 0.019675
+[100] training's l2: 0.018778
+```
+
+## Computing forecasts
+
+Now we just assign this model to the `MLForecast.models_` dictionary.
+Note that you can assign as many models as you want.
+
+```python
+fcst.models_['lgbm'] = model
+fcst.models_
+```
+
+``` text
+{'lr': LinearRegression(), 'lgbm': LGBMRegressor(verbosity=-1)}
+```
+
+And now when calling
+[`MLForecast.predict`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.predict),
+mlforecast will use those models to compute the forecasts.
+
+```python
+fcst.predict(1)
+```
+
+| | unique_id | ds | lr | lgbm |
+|-----|-----------|------------|----------|----------|
+| 0 | id_0 | 2000-08-10 | 3.549124 | 5.166797 |
+| 1 | id_1 | 2000-04-07 | 3.154285 | 4.252490 |
+| 2 | id_2 | 2000-06-16 | 2.880933 | 3.224506 |
+| 3 | id_3 | 2000-08-30 | 4.061801 | 0.245443 |
+| 4 | id_4 | 2001-01-08 | 2.904872 | 2.225106 |
+
diff --git a/mlforecast/docs/how-to-guides/exogenous_features.html.mdx b/mlforecast/docs/how-to-guides/exogenous_features.html.mdx
new file mode 100644
index 00000000..3d0dfbe5
--- /dev/null
+++ b/mlforecast/docs/how-to-guides/exogenous_features.html.mdx
@@ -0,0 +1,233 @@
+---
+description: Use exogenous regressors for training and predicting
+output-file: exogenous_features.html
+title: Exogenous features
+---
+
+
+```python
+import lightgbm as lgb
+import pandas as pd
+from mlforecast import MLForecast
+from mlforecast.lag_transforms import ExpandingMean, RollingMean
+from mlforecast.utils import generate_daily_series, generate_prices_for_series
+```
+
+## Data setup
+
+```python
+series = generate_daily_series(
+ 100, equal_ends=True, n_static_features=2
+).rename(columns={'static_1': 'product_id'})
+series.head()
+```
+
+| | unique_id | ds | y | static_0 | product_id |
+|-----|-----------|------------|------------|----------|------------|
+| 0 | id_00 | 2000-10-05 | 39.811983 | 79 | 45 |
+| 1 | id_00 | 2000-10-06 | 103.274013 | 79 | 45 |
+| 2 | id_00 | 2000-10-07 | 176.574744 | 79 | 45 |
+| 3 | id_00 | 2000-10-08 | 258.987900 | 79 | 45 |
+| 4 | id_00 | 2000-10-09 | 344.940404 | 79 | 45 |
+
+## Use existing exogenous features
+
+In mlforecast the required columns are the series identifier, time and
+target. Any extra columns you have, like `static_0` and `product_id`
+here are considered to be static and are replicated when constructing
+the features for the next timestamp. You can disable this by passing
+`static_features` to
+[`MLForecast.preprocess`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.preprocess)
+or
+[`MLForecast.fit`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.fit),
+which will only keep the columns you define there as static. Keep in
+mind that all features in your input dataframe will be used for
+training, so you’ll have to provide the future values of exogenous
+features to
+[`MLForecast.predict`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.predict)
+through the `X_df` argument.
+
+Consider the following example. Suppose that we have a prices catalog
+for each id and date.
+
+```python
+prices_catalog = generate_prices_for_series(series)
+prices_catalog.head()
+```
+
+| | ds | unique_id | price |
+|-----|------------|-----------|----------|
+| 0 | 2000-10-05 | id_00 | 0.548814 |
+| 1 | 2000-10-06 | id_00 | 0.715189 |
+| 2 | 2000-10-07 | id_00 | 0.602763 |
+| 3 | 2000-10-08 | id_00 | 0.544883 |
+| 4 | 2000-10-09 | id_00 | 0.423655 |
+
+And that you have already merged these prices into your series
+dataframe.
+
+```python
+series_with_prices = series.merge(prices_catalog, how='left')
+series_with_prices.head()
+```
+
+| | unique_id | ds | y | static_0 | product_id | price |
+|-----|-----------|------------|------------|----------|------------|----------|
+| 0 | id_00 | 2000-10-05 | 39.811983 | 79 | 45 | 0.548814 |
+| 1 | id_00 | 2000-10-06 | 103.274013 | 79 | 45 | 0.715189 |
+| 2 | id_00 | 2000-10-07 | 176.574744 | 79 | 45 | 0.602763 |
+| 3 | id_00 | 2000-10-08 | 258.987900 | 79 | 45 | 0.544883 |
+| 4 | id_00 | 2000-10-09 | 344.940404 | 79 | 45 | 0.423655 |
+
+This dataframe will be passed to
+[`MLForecast.fit`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.fit)
+(or
+[`MLForecast.preprocess`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.preprocess)).
+However, since the price is dynamic we have to tell that method that
+only `static_0` and `product_id` are static.
+
+```python
+fcst = MLForecast(
+ models=lgb.LGBMRegressor(n_jobs=1, random_state=0, verbosity=-1),
+ freq='D',
+ lags=[7],
+ lag_transforms={
+ 1: [ExpandingMean()],
+ 7: [RollingMean(window_size=14)],
+ },
+ date_features=['dayofweek', 'month'],
+ num_threads=2,
+)
+fcst.fit(series_with_prices, static_features=['static_0', 'product_id'])
+```
+
+``` text
+MLForecast(models=[LGBMRegressor], freq=D, lag_features=['lag7', 'expanding_mean_lag1', 'rolling_mean_lag7_window_size14'], date_features=['dayofweek', 'month'], num_threads=2)
+```
+
+The features used for training are stored in
+`MLForecast.ts.features_order_`. As you can see `price` was used for
+training.
+
+```python
+fcst.ts.features_order_
+```
+
+``` text
+['static_0',
+ 'product_id',
+ 'price',
+ 'lag7',
+ 'expanding_mean_lag1',
+ 'rolling_mean_lag7_window_size14',
+ 'dayofweek',
+ 'month']
+```
+
+So in order to update the price in each timestep we just call
+[`MLForecast.predict`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.predict)
+with our forecast horizon and pass the prices catalog through `X_df`.
+
+```python
+preds = fcst.predict(h=7, X_df=prices_catalog)
+preds.head()
+```
+
+| | unique_id | ds | LGBMRegressor |
+|-----|-----------|------------|---------------|
+| 0 | id_00 | 2001-05-15 | 418.930093 |
+| 1 | id_00 | 2001-05-16 | 499.487368 |
+| 2 | id_00 | 2001-05-17 | 20.321885 |
+| 3 | id_00 | 2001-05-18 | 102.310778 |
+| 4 | id_00 | 2001-05-19 | 185.340281 |
+
+## Generating exogenous features
+
+Nixtla provides some utilities to generate exogenous features for both
+training and forecasting such as [statsforecast’s
+mstl_decomposition](https://nixtlaverse.nixtla.io/statsforecast/docs/how-to-guides/generating_features.html)
+or the [transform_exog function](transforming_exog.html). We also have
+[utilsforecast’s fourier
+function](https://nixtlaverse.nixtla.io/utilsforecast/feature_engineering.html#fourier),
+which we’ll demonstrate here.
+
+```python
+from sklearn.linear_model import LinearRegression
+from utilsforecast.feature_engineering import fourier
+```
+
+Suppose you start with some data like the one above where we have a
+couple of static features.
+
+```python
+series.head()
+```
+
+| | unique_id | ds | y | static_0 | product_id |
+|-----|-----------|------------|------------|----------|------------|
+| 0 | id_00 | 2000-10-05 | 39.811983 | 79 | 45 |
+| 1 | id_00 | 2000-10-06 | 103.274013 | 79 | 45 |
+| 2 | id_00 | 2000-10-07 | 176.574744 | 79 | 45 |
+| 3 | id_00 | 2000-10-08 | 258.987900 | 79 | 45 |
+| 4 | id_00 | 2000-10-09 | 344.940404 | 79 | 45 |
+
+Now we’d like to add some fourier terms to model the seasonality. We can
+do that with the following:
+
+```python
+transformed_df, future_df = fourier(series, freq='D', season_length=7, k=2, h=7)
+```
+
+This provides an extended training dataset.
+
+```python
+transformed_df.head()
+```
+
+| | unique_id | ds | y | static_0 | product_id | sin1_7 | sin2_7 | cos1_7 | cos2_7 |
+|----|----|----|----|----|----|----|----|----|----|
+| 0 | id_00 | 2000-10-05 | 39.811983 | 79 | 45 | 0.781832 | 0.974928 | 0.623490 | -0.222521 |
+| 1 | id_00 | 2000-10-06 | 103.274013 | 79 | 45 | 0.974928 | -0.433884 | -0.222521 | -0.900969 |
+| 2 | id_00 | 2000-10-07 | 176.574744 | 79 | 45 | 0.433884 | -0.781831 | -0.900969 | 0.623490 |
+| 3 | id_00 | 2000-10-08 | 258.987900 | 79 | 45 | -0.433884 | 0.781832 | -0.900969 | 0.623490 |
+| 4 | id_00 | 2000-10-09 | 344.940404 | 79 | 45 | -0.974928 | 0.433884 | -0.222521 | -0.900969 |
+
+Along with the future values of the features.
+
+```python
+future_df.head()
+```
+
+| | unique_id | ds | sin1_7 | sin2_7 | cos1_7 | cos2_7 |
+|-----|-----------|------------|-----------|-----------|-----------|-----------|
+| 0 | id_00 | 2001-05-15 | -0.781828 | -0.974930 | 0.623494 | -0.222511 |
+| 1 | id_00 | 2001-05-16 | 0.000006 | 0.000011 | 1.000000 | 1.000000 |
+| 2 | id_00 | 2001-05-17 | 0.781835 | 0.974925 | 0.623485 | -0.222533 |
+| 3 | id_00 | 2001-05-18 | 0.974927 | -0.433895 | -0.222527 | -0.900963 |
+| 4 | id_00 | 2001-05-19 | 0.433878 | -0.781823 | -0.900972 | 0.623500 |
+
+We can now train using only these features (and the static ones).
+
+```python
+fcst2 = MLForecast(models=LinearRegression(), freq='D')
+fcst2.fit(transformed_df, static_features=['static_0', 'product_id'])
+```
+
+``` text
+MLForecast(models=[LinearRegression], freq=D, lag_features=[], date_features=[], num_threads=1)
+```
+
+And provide the future values to the predict method.
+
+```python
+fcst2.predict(h=7, X_df=future_df).head()
+```
+
+| | unique_id | ds | LinearRegression |
+|-----|-----------|------------|------------------|
+| 0 | id_00 | 2001-05-15 | 275.822342 |
+| 1 | id_00 | 2001-05-16 | 262.258117 |
+| 2 | id_00 | 2001-05-17 | 238.195850 |
+| 3 | id_00 | 2001-05-18 | 240.997814 |
+| 4 | id_00 | 2001-05-19 | 262.247123 |
+
diff --git a/mlforecast/docs/how-to-guides/hyperparameter_optimization.html.mdx b/mlforecast/docs/how-to-guides/hyperparameter_optimization.html.mdx
new file mode 100644
index 00000000..a7fa8684
--- /dev/null
+++ b/mlforecast/docs/how-to-guides/hyperparameter_optimization.html.mdx
@@ -0,0 +1,367 @@
+---
+description: Tune your forecasting models
+output-file: hyperparameter_optimization.html
+title: Hyperparameter optimization
+---
+
+
+## Imports
+
+```python
+import os
+import tempfile
+
+import lightgbm as lgb
+import optuna
+import pandas as pd
+from datasetsforecast.m4 import M4, M4Evaluation, M4Info
+from sklearn.linear_model import Ridge
+from sklearn.compose import ColumnTransformer
+from sklearn.pipeline import make_pipeline
+from sklearn.preprocessing import OneHotEncoder
+from utilsforecast.plotting import plot_series
+
+from mlforecast import MLForecast
+from mlforecast.auto import (
+ AutoLightGBM,
+ AutoMLForecast,
+ AutoModel,
+ AutoRidge,
+ ridge_space,
+)
+from mlforecast.lag_transforms import ExponentiallyWeightedMean, RollingMean
+```
+
+## Data setup
+
+```python
+def get_data(group, horizon):
+ df, *_ = M4.load(directory='data', group=group)
+ df['ds'] = df['ds'].astype('int')
+ df['unique_id'] = df['unique_id'].astype('category')
+ return df.groupby('unique_id').head(-horizon).copy()
+
+group = 'Hourly'
+horizon = M4Info[group].horizon
+train = get_data(group, horizon)
+```
+
+## Optimization
+
+### Default optimization
+
+We have default search spaces for some models and we can define default
+features to look for based on the length of the seasonal period of your
+data. For this example we’ll use hourly data, for which we’ll set 24
+(one day) as the season length.
+
+```python
+optuna.logging.set_verbosity(optuna.logging.ERROR)
+auto_mlf = AutoMLForecast(
+ models={'lgb': AutoLightGBM(), 'ridge': AutoRidge()},
+ freq=1,
+ season_length=24,
+)
+auto_mlf.fit(
+ train,
+ n_windows=2,
+ h=horizon,
+ num_samples=2, # number of trials to run
+)
+```
+
+``` text
+AutoMLForecast(models={'lgb': AutoModel(model=LGBMRegressor), 'ridge': AutoModel(model=Ridge)})
+```
+
+We can now use these models to predict
+
+```python
+preds = auto_mlf.predict(horizon)
+preds.head()
+```
+
+| | unique_id | ds | lgb | ridge |
+|-----|-----------|-----|------------|------------|
+| 0 | H1 | 701 | 680.534943 | 604.140123 |
+| 1 | H1 | 702 | 599.038307 | 523.364874 |
+| 2 | H1 | 703 | 572.808421 | 479.174481 |
+| 3 | H1 | 704 | 564.573783 | 444.540062 |
+| 4 | H1 | 705 | 543.046026 | 419.987657 |
+
+And evaluate them
+
+```python
+def evaluate(df, group):
+ results = []
+ for model in df.columns.drop(['unique_id', 'ds']):
+ model_res = M4Evaluation.evaluate(
+ 'data', group, df[model].to_numpy().reshape(-1, horizon)
+ )
+ model_res.index = [model]
+ results.append(model_res)
+ return pd.concat(results).T.round(2)
+
+evaluate(preds, group)
+```
+
+| | lgb | ridge |
+|-------|-------|-------|
+| SMAPE | 18.78 | 20.00 |
+| MASE | 5.07 | 1.29 |
+| OWA | 1.57 | 0.81 |
+
+### Tuning model parameters
+
+You can provide your own model with its search space to perform the
+optimization. The search space should be a function that takes an optuna
+trial and returns the model parameters.
+
+```python
+def my_lgb_config(trial: optuna.Trial):
+ return {
+ 'learning_rate': 0.05,
+ 'verbosity': -1,
+ 'num_leaves': trial.suggest_int('num_leaves', 2, 128, log=True),
+ 'objective': trial.suggest_categorical('objective', ['l1', 'l2', 'mape']),
+ }
+
+my_lgb = AutoModel(
+ model=lgb.LGBMRegressor(),
+ config=my_lgb_config,
+)
+auto_mlf = AutoMLForecast(
+ models={'my_lgb': my_lgb},
+ freq=1,
+ season_length=24,
+).fit(
+ train,
+ n_windows=2,
+ h=horizon,
+ num_samples=2,
+)
+preds = auto_mlf.predict(horizon)
+evaluate(preds, group)
+```
+
+| | my_lgb |
+|-------|--------|
+| SMAPE | 18.67 |
+| MASE | 4.79 |
+| OWA | 1.51 |
+
+#### Tuning scikit-learn pipelines
+
+We internally use
+[BaseEstimator.set_params](https://scikit-learn.org/stable/modules/generated/sklearn.base.BaseEstimator.html#sklearn.base.BaseEstimator.set_params)
+for each configuration, so if you’re using a scikit-learn pipeline you
+can tune its parameters as you normally would with scikit-learn’s
+searches.
+
+```python
+ridge_pipeline = make_pipeline(
+ ColumnTransformer(
+ [('encoder', OneHotEncoder(), ['unique_id'])],
+ remainder='passthrough',
+ ),
+ Ridge()
+)
+my_auto_ridge = AutoModel(
+ ridge_pipeline,
+ # the space must have the name of the estimator followed by the parameter
+ # you could also tune the encoder here
+ lambda trial: {f'ridge__{k}': v for k, v in ridge_space(trial).items()},
+)
+auto_mlf = AutoMLForecast(
+ models={'ridge': my_auto_ridge},
+ freq=1,
+ season_length=24,
+ fit_config=lambda trial: {'static_features': ['unique_id']}
+).fit(
+ train,
+ n_windows=2,
+ h=horizon,
+ num_samples=2,
+)
+preds = auto_mlf.predict(horizon)
+evaluate(preds, group)
+```
+
+| | ridge |
+|-------|-------|
+| SMAPE | 18.50 |
+| MASE | 1.24 |
+| OWA | 0.76 |
+
+### Tuning features
+
+The
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+class defines the features to build in its constructor. You can tune the
+features by providing a function through the `init_config` argument,
+which will take an optuna trial and produce a configuration to pass to
+the
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+constructor.
+
+```python
+def my_init_config(trial: optuna.Trial):
+ lag_transforms = [
+ ExponentiallyWeightedMean(alpha=0.3),
+ RollingMean(window_size=24 * 7, min_samples=1),
+ ]
+ lag_to_transform = trial.suggest_categorical('lag_to_transform', [24, 48])
+ return {
+ 'lags': [24 * i for i in range(1, 7)], # this won't be tuned
+ 'lag_transforms': {lag_to_transform: lag_transforms},
+ }
+
+auto_mlf = AutoMLForecast(
+ models=[AutoRidge()],
+ freq=1,
+ season_length=24,
+ init_config=my_init_config,
+).fit(
+ train,
+ n_windows=2,
+ h=horizon,
+ num_samples=2,
+)
+preds = auto_mlf.predict(horizon)
+evaluate(preds, group)
+```
+
+| | AutoRidge |
+|-------|-----------|
+| SMAPE | 13.31 |
+| MASE | 1.67 |
+| OWA | 0.71 |
+
+### Tuning fit parameters
+
+The
+[`MLForecast.fit`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.fit)
+method takes some arguments that could improve the forecasting
+performance of your models, such as `dropna` and `static_features`. If
+you want to tune those you can provide a function to the `fit_config`
+argument.
+
+```python
+def my_fit_config(trial: optuna.Trial):
+ if trial.suggest_int('use_id', 0, 1):
+ static_features = ['unique_id']
+ else:
+ static_features = None
+ return {
+ 'static_features': static_features
+ }
+
+auto_mlf = AutoMLForecast(
+ models=[AutoLightGBM()],
+ freq=1,
+ season_length=24,
+ fit_config=my_fit_config,
+).fit(
+ train,
+ n_windows=2,
+ h=horizon,
+ num_samples=2,
+)
+preds = auto_mlf.predict(horizon)
+evaluate(preds, group)
+```
+
+| | AutoLightGBM |
+|-------|--------------|
+| SMAPE | 18.78 |
+| MASE | 5.07 |
+| OWA | 1.57 |
+
+## Accessing the optimization results
+
+After the process has finished the results are available under the
+`results_` attribute of the
+[`AutoMLForecast`](https://Nixtla.github.io/mlforecast/auto.html#automlforecast)
+object. There will be one result per model and the best configuration
+can be found under the `config` user attribute.
+
+```python
+len(auto_mlf.results_)
+```
+
+``` text
+1
+```
+
+```python
+auto_mlf.results_['AutoLightGBM'].best_trial.user_attrs['config']
+```
+
+``` text
+{'model_params': {'bagging_freq': 1,
+ 'learning_rate': 0.05,
+ 'verbosity': -1,
+ 'n_estimators': 169,
+ 'lambda_l1': 0.027334069690310565,
+ 'lambda_l2': 0.0026599310838681858,
+ 'num_leaves': 112,
+ 'feature_fraction': 0.7118273996694524,
+ 'bagging_fraction': 0.8229470565333281,
+ 'objective': 'l2'},
+ 'mlf_init_params': {'lags': [48],
+ 'target_transforms': None,
+ 'lag_transforms': {1: [ExponentiallyWeightedMean(alpha=0.9)]},
+ 'date_features': None,
+ 'num_threads': 1},
+ 'mlf_fit_params': {'static_features': None}}
+```
+
+### Individual models
+
+There is one optimization process per model. This is because different
+models can make use of different features. So after the optimization
+process is done for each model the best configuration is used to retrain
+the model using all of the data. These final models are
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+objects and are saved in the `models_` attribute.
+
+```python
+auto_mlf.models_
+```
+
+``` text
+{'AutoLightGBM': MLForecast(models=[AutoLightGBM], freq=1, lag_features=['lag48', 'exponentially_weighted_mean_lag1_alpha0.9'], date_features=[], num_threads=1)}
+```
+
+## Saving
+
+You can use the
+[`AutoMLForecast.save`](https://Nixtla.github.io/mlforecast/auto.html#automlforecast.save)
+method to save the best models found. This produces one directory per
+model.
+
+```python
+with tempfile.TemporaryDirectory() as tmpdir:
+ auto_mlf.save(tmpdir)
+ print(os.listdir(tmpdir))
+```
+
+``` text
+['AutoLightGBM']
+```
+
+Since each model is an
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+object you can load it by itself.
+
+```python
+with tempfile.TemporaryDirectory() as tmpdir:
+ auto_mlf.save(tmpdir)
+ loaded = MLForecast.load(f'{tmpdir}/AutoLightGBM')
+ print(loaded)
+```
+
+``` text
+MLForecast(models=[AutoLightGBM], freq=1, lag_features=['lag48', 'exponentially_weighted_mean_lag1_alpha0.9'], date_features=[], num_threads=1)
+```
+
diff --git a/mlforecast/docs/how-to-guides/lag_transforms_guide.html.mdx b/mlforecast/docs/how-to-guides/lag_transforms_guide.html.mdx
new file mode 100644
index 00000000..08e1b42d
--- /dev/null
+++ b/mlforecast/docs/how-to-guides/lag_transforms_guide.html.mdx
@@ -0,0 +1,221 @@
+---
+description: Compute features based on lags
+output-file: lag_transforms_guide.html
+title: Lag transformations
+---
+
+
+mlforecast allows you to define transformations on the lags to use as
+features. These are provided through the `lag_transforms` argument,
+which is a dict where the keys are the lags and the values are a list of
+transformations to apply to that lag.
+
+## Data setup
+
+```python
+import numpy as np
+
+from mlforecast import MLForecast
+from mlforecast.utils import generate_daily_series
+```
+
+
+```python
+data = generate_daily_series(10)
+```
+
+## Built-in transformations
+
+The built-in lag transformations are in the `mlforecast.lag_transforms`
+module.
+
+```python
+from mlforecast.lag_transforms import RollingMean, ExpandingStd
+```
+
+
+```python
+fcst = MLForecast(
+ models=[],
+ freq='D',
+ lag_transforms={
+ 1: [ExpandingStd()],
+ 7: [RollingMean(window_size=7, min_samples=1), RollingMean(window_size=14)]
+ },
+)
+```
+
+Once you define your transformations you can see what they look like
+with
+[`MLForecast.preprocess`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.preprocess).
+
+```python
+fcst.preprocess(data).head(2)
+```
+
+| | unique_id | ds | y | expanding_std_lag1 | rolling_mean_lag7_window_size7_min_samples1 | rolling_mean_lag7_window_size14 |
+|----|----|----|----|----|----|----|
+| 20 | id_0 | 2000-01-21 | 6.319961 | 1.956363 | 3.234486 | 3.283064 |
+| 21 | id_0 | 2000-01-22 | 0.071677 | 2.028545 | 3.256055 | 3.291068 |
+
+### Extending the built-in transformations
+
+You can compose the built-in transformations by using the
+[`Combine`](https://Nixtla.github.io/mlforecast/lag_transforms.html#combine)
+class, which takes two transformations and an operator.
+
+```python
+import operator
+
+from mlforecast.lag_transforms import Combine
+```
+
+
+```python
+fcst = MLForecast(
+ models=[],
+ freq='D',
+ lag_transforms={
+ 1: [
+ RollingMean(window_size=7),
+ RollingMean(window_size=14),
+ Combine(
+ RollingMean(window_size=7),
+ RollingMean(window_size=14),
+ operator.truediv,
+ )
+ ],
+ },
+)
+prep = fcst.preprocess(data)
+prep.head(2)
+```
+
+| | unique_id | ds | y | rolling_mean_lag1_window_size7 | rolling_mean_lag1_window_size14 | rolling_mean_lag1_window_size7_truediv_rolling_mean_lag1_window_size14 |
+|----|----|----|----|----|----|----|
+| 14 | id_0 | 2000-01-15 | 0.435006 | 3.234486 | 3.283064 | 0.985204 |
+| 15 | id_0 | 2000-01-16 | 1.489309 | 3.256055 | 3.291068 | 0.989361 |
+
+```python
+np.testing.assert_allclose(
+ prep['rolling_mean_lag1_window_size7'] / prep['rolling_mean_lag1_window_size14'],
+ prep['rolling_mean_lag1_window_size7_truediv_rolling_mean_lag1_window_size14']
+)
+```
+
+If you want one of the transformations in
+[`Combine`](https://Nixtla.github.io/mlforecast/lag_transforms.html#combine)
+to be applied to a different lag you can use the
+[`Offset`](https://Nixtla.github.io/mlforecast/lag_transforms.html#offset)
+class, which will apply the offset first and then the transformation.
+
+```python
+from mlforecast.lag_transforms import Offset
+```
+
+
+```python
+fcst = MLForecast(
+ models=[],
+ freq='D',
+ lag_transforms={
+ 1: [
+ RollingMean(window_size=7),
+ Combine(
+ RollingMean(window_size=7),
+ Offset(RollingMean(window_size=7), n=1),
+ operator.truediv,
+ )
+ ],
+ 2: [RollingMean(window_size=7)]
+ },
+)
+prep = fcst.preprocess(data)
+prep.head(2)
+```
+
+| | unique_id | ds | y | rolling_mean_lag1_window_size7 | rolling_mean_lag1_window_size7_truediv_rolling_mean_lag2_window_size7 | rolling_mean_lag2_window_size7 |
+|----|----|----|----|----|----|----|
+| 8 | id_0 | 2000-01-09 | 1.462798 | 3.326081 | 0.998331 | 3.331641 |
+| 9 | id_0 | 2000-01-10 | 2.035518 | 3.360938 | 1.010480 | 3.326081 |
+
+```python
+np.testing.assert_allclose(
+ prep['rolling_mean_lag1_window_size7'] / prep['rolling_mean_lag2_window_size7'],
+ prep['rolling_mean_lag1_window_size7_truediv_rolling_mean_lag2_window_size7']
+)
+```
+
+## numba-based transformations
+
+The [window-ops package](https://github.com/jmoralez/window_ops)
+provides transformations defined as [numba](https://numba.pydata.org/)
+[JIT compiled](https://en.wikipedia.org/wiki/Just-in-time_compilation)
+functions. We use numba because it makes them really fast and can also
+bypass [python’s
+GIL](https://wiki.python.org/moin/GlobalInterpreterLock), which allows
+running them concurrently with multithreading.
+
+The main benefit of using these transformations is that they’re very
+easy to implement. However, when we need to update their values on the
+predict step they can very slow, because we have to call the function
+again on the complete history and just keep the last value, so if
+performance is a concern you should try to use the built-in ones or set
+`keep_last_n` in
+[`MLForecast.preprocess`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.preprocess)
+or
+[`MLForecast.fit`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.fit)
+to the minimum number of samples that your transformations require.
+
+```python
+from numba import njit
+from window_ops.expanding import expanding_mean
+from window_ops.shift import shift_array
+```
+
+
+```python
+@njit
+def ratio_over_previous(x, offset=1):
+ """Computes the ratio between the current value and its `offset` lag"""
+ return x / shift_array(x, offset=offset)
+
+@njit
+def diff_over_previous(x, offset=1):
+ """Computes the difference between the current value and its `offset` lag"""
+ return x - shift_array(x, offset=offset)
+```
+
+If your function takes more arguments than the input array you can
+provide a tuple like: `(func, arg1, arg2, ...)`
+
+```python
+fcst = MLForecast(
+ models=[],
+ freq='D',
+ lags=[1, 2, 3],
+ lag_transforms={
+ 1: [expanding_mean, ratio_over_previous, (ratio_over_previous, 2)], # the second ratio sets offset=2
+ 2: [diff_over_previous],
+ },
+)
+prep = fcst.preprocess(data)
+prep.head(2)
+```
+
+| | unique_id | ds | y | lag1 | lag2 | lag3 | expanding_mean_lag1 | ratio_over_previous_lag1 | ratio_over_previous_lag1_offset2 | diff_over_previous_lag2 |
+|----|----|----|----|----|----|----|----|----|----|----|
+| 3 | id_0 | 2000-01-04 | 3.481831 | 2.445887 | 1.218794 | 0.322947 | 1.329209 | 2.006809 | 7.573645 | 0.895847 |
+| 4 | id_0 | 2000-01-05 | 4.191721 | 3.481831 | 2.445887 | 1.218794 | 1.867365 | 1.423546 | 2.856785 | 1.227093 |
+
+As you can see the name of the function is used as the transformation
+name plus the `_lag` suffix. If the function has other arguments and
+they’re not set to their default values they’re included as well, as is
+done with `offset=2` here.
+
+```python
+np.testing.assert_allclose(prep['lag1'] / prep['lag2'], prep['ratio_over_previous_lag1'])
+np.testing.assert_allclose(prep['lag1'] / prep['lag3'], prep['ratio_over_previous_lag1_offset2'])
+np.testing.assert_allclose(prep['lag2'] - prep['lag3'], prep['diff_over_previous_lag2'])
+```
+
diff --git a/mlforecast/docs/how-to-guides/mlflow.html.mdx b/mlforecast/docs/how-to-guides/mlflow.html.mdx
new file mode 100644
index 00000000..acc73083
--- /dev/null
+++ b/mlforecast/docs/how-to-guides/mlflow.html.mdx
@@ -0,0 +1,200 @@
+---
+description: Log your metrics and models
+output-file: mlflow.html
+title: MLflow
+---
+
+
+## Libraries
+
+```python
+import copy
+import subprocess
+import time
+
+import lightgbm as lgb
+import mlflow
+import pandas as pd
+import requests
+from sklearn.linear_model import LinearRegression
+from utilsforecast.data import generate_series
+from utilsforecast.losses import rmse, smape
+from utilsforecast.evaluation import evaluate
+from utilsforecast.feature_engineering import fourier
+
+import mlforecast.flavor
+from mlforecast import MLForecast
+from mlforecast.lag_transforms import ExponentiallyWeightedMean
+from mlforecast.utils import PredictionIntervals
+```
+
+## Data setup
+
+```python
+freq = 'h'
+h = 10
+series = generate_series(5, freq=freq)
+valid = series.groupby('unique_id', observed=True).tail(h)
+train = series.drop(valid.index)
+train, X_df = fourier(train, freq=freq, season_length=24, k=2, h=h)
+```
+
+## Parameters
+
+```python
+params = {
+ 'init': {
+ 'models': {
+ 'lgb': lgb.LGBMRegressor(
+ n_estimators=50, num_leaves=16, verbosity=-1
+ ),
+ 'lr': LinearRegression(),
+ },
+ 'freq': freq,
+ 'lags': [24],
+ 'lag_transforms': {
+ 1: [ExponentiallyWeightedMean(0.9)],
+ },
+ 'num_threads': 2,
+ },
+ 'fit': {
+ 'static_features': ['unique_id'],
+ 'prediction_intervals': PredictionIntervals(n_windows=2, h=h),
+ }
+}
+```
+
+## Logging
+
+If you have a tracking server, you can run
+`mlflow.set_tracking_uri(your_server_uri)` to connect to it.
+
+```python
+mlflow.set_experiment("mlforecast")
+with mlflow.start_run() as run:
+ train_ds = mlflow.data.from_pandas(train)
+ valid_ds = mlflow.data.from_pandas(valid)
+ mlflow.log_input(train_ds, context="training")
+ mlflow.log_input(valid_ds, context="validation")
+ logged_params = copy.deepcopy(params)
+ logged_params['init']['models'] = {
+ k: (v.__class__.__name__, v.get_params())
+ for k, v in params['init']['models'].items()
+ }
+ mlflow.log_params(logged_params)
+ mlf = MLForecast(**params['init'])
+ mlf.fit(train, **params['fit'])
+ preds = mlf.predict(h, X_df=X_df)
+ eval_result = evaluate(
+ valid.merge(preds, on=['unique_id', 'ds']),
+ metrics=[rmse, smape],
+ agg_fn='mean',
+ )
+ models = mlf.models_.keys()
+ logged_metrics = {}
+ for _, row in eval_result.iterrows():
+ metric = row['metric']
+ for model in models:
+ logged_metrics[f'{metric}_{model}'] = row[model]
+ mlflow.log_metrics(logged_metrics)
+ mlforecast.flavor.log_model(model=mlf, artifact_path="model")
+ model_uri = mlflow.get_artifact_uri("model")
+ run_id = run.info.run_id
+```
+
+``` text
+/home/ubuntu/repos/mlforecast/.venv/lib/python3.10/site-packages/mlflow/types/utils.py:406: UserWarning: Hint: Inferred schema contains integer column(s). Integer columns in Python cannot represent missing values. If your input data contains missing values at inference time, it will be encoded as floats and will cause a schema enforcement error. The best way to avoid this problem is to infer the model schema based on a realistic data sample (training dataset) that includes missing values. Alternatively, you can declare integer columns as doubles (float64) whenever these columns may have missing values. See `Handling Integers With Missing Values `_ for more details.
+ warnings.warn(
+2024/08/23 02:57:14 WARNING mlflow.models.model: Input example should be provided to infer model signature if the model signature is not provided when logging the model.
+```
+
+## Load model
+
+```python
+loaded_model = mlforecast.flavor.load_model(model_uri=model_uri)
+results = loaded_model.predict(h=h, X_df=X_df, ids=[3])
+results.head(2)
+```
+
+| | unique_id | ds | lgb | lr |
+|-----|-----------|---------------------|----------|----------|
+| 0 | 3 | 2000-01-10 16:00:00 | 0.333308 | 0.243017 |
+| 1 | 3 | 2000-01-10 17:00:00 | 0.127424 | 0.249742 |
+
+## PyFunc
+
+```python
+loaded_pyfunc = mlforecast.flavor.pyfunc.load_model(model_uri=model_uri)
+# single row dataframe
+predict_conf = pd.DataFrame(
+ [
+ {
+ "h": h,
+ "ids": [0, 2],
+ "X_df": X_df,
+ "level": [80]
+ }
+ ]
+)
+pyfunc_result = loaded_pyfunc.predict(predict_conf)
+pyfunc_result.head(2)
+```
+
+| | unique_id | ds | lgb | lr | lgb-lo-80 | lgb-hi-80 | lr-lo-80 | lr-hi-80 |
+|----|----|----|----|----|----|----|----|----|
+| 0 | 0 | 2000-01-09 20:00:00 | 0.260544 | 0.244128 | 0.140168 | 0.380921 | 0.114001 | 0.374254 |
+| 1 | 0 | 2000-01-09 21:00:00 | 0.250096 | 0.247742 | 0.072820 | 0.427372 | 0.047584 | 0.447900 |
+
+## Model serving
+
+```python
+host = 'localhost'
+port = '5000'
+cmd = f'mlflow models serve -m runs:/{run_id}/model -h {host} -p {port} --env-manager local'
+# initialize server
+process = subprocess.Popen(cmd.split())
+time.sleep(5)
+# single row dataframe. must be JSON serializable
+predict_conf = pd.DataFrame(
+ [
+ {
+ "h": h,
+ "ids": [3, 4],
+ "X_df": X_df.astype({'ds': 'str'}).to_dict(orient='list'),
+ "level": [95]
+ }
+ ]
+)
+payload = {'dataframe_split': predict_conf.to_dict(orient='split', index=False)}
+resp = requests.post(f'http://{host}:{port}/invocations', json=payload)
+print(pd.DataFrame(resp.json()['predictions']).head(2))
+process.terminate()
+process.wait(timeout=10)
+```
+
+``` text
+Downloading artifacts: 100%|██████████| 7/7 [00:00<00:00, 18430.71it/s]
+2024/08/23 02:57:16 INFO mlflow.models.flavor_backend_registry: Selected backend for flavor 'python_function'
+2024/08/23 02:57:16 INFO mlflow.pyfunc.backend: === Running command 'exec gunicorn --timeout=60 -b localhost:5000 -w 1 ${GUNICORN_CMD_ARGS} -- mlflow.pyfunc.scoring_server.wsgi:app'
+[2024-08-23 02:57:16 +0000] [23054] [INFO] Starting gunicorn 22.0.0
+[2024-08-23 02:57:16 +0000] [23054] [INFO] Listening at: http://127.0.0.1:5000 (23054)
+[2024-08-23 02:57:16 +0000] [23054] [INFO] Using worker: sync
+[2024-08-23 02:57:16 +0000] [23055] [INFO] Booting worker with pid: 23055
+```
+
+``` text
+ unique_id ds lgb lr lgb-lo-95 lgb-hi-95 \
+0 3 2000-01-10T16:00:00 0.333308 0.243017 0.174073 0.492544
+1 3 2000-01-10T17:00:00 0.127424 0.249742 -0.009993 0.264842
+
+ lr-lo-95 lr-hi-95
+0 0.032451 0.453583
+1 0.045525 0.453959
+```
+
+``` text
+[2024-08-23 02:57:20 +0000] [23054] [INFO] Handling signal: term
+[2024-08-23 02:57:20 +0000] [23055] [INFO] Worker exiting (pid: 23055)
+[2024-08-23 02:57:21 +0000] [23054] [INFO] Shutting down: Master
+```
+
diff --git a/mlforecast/docs/how-to-guides/one_model_per_horizon.html.mdx b/mlforecast/docs/how-to-guides/one_model_per_horizon.html.mdx
new file mode 100644
index 00000000..fd2674dd
--- /dev/null
+++ b/mlforecast/docs/how-to-guides/one_model_per_horizon.html.mdx
@@ -0,0 +1,115 @@
+---
+description: Train one model to predict each step of the forecasting horizon
+output-file: one_model_per_horizon.html
+title: One model per step
+---
+
+
+By default mlforecast uses the recursive strategy, i.e. a model is
+trained to predict the next value and if we’re predicting several values
+we do it one at a time and then use the model’s predictions as the new
+target, recompute the features and predict the next step.
+
+There’s another approach where if we want to predict 10 steps ahead we
+train 10 different models, where each model is trained to predict the
+value at each specific step, i.e. one model predicts the next value,
+another one predicts the value two steps ahead and so on. This can be
+very time consuming but can also provide better results. If you want to
+use this approach you can specify `max_horizon` in
+[`MLForecast.fit`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.fit),
+which will train that many models and each model will predict its
+corresponding horizon when you call
+[`MLForecast.predict`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.predict).
+
+## Setup
+
+```python
+import random
+import lightgbm as lgb
+import pandas as pd
+from datasetsforecast.m4 import M4, M4Info
+from utilsforecast.evaluation import evaluate
+from utilsforecast.losses import smape
+
+from mlforecast import MLForecast
+from mlforecast.lag_transforms import ExponentiallyWeightedMean, RollingMean
+from mlforecast.target_transforms import Differences
+```
+
+### Data
+
+We will use four random series from the M4 dataset
+
+```python
+group = 'Hourly'
+await M4.async_download('data', group=group)
+df, *_ = M4.load(directory='data', group=group)
+df['ds'] = df['ds'].astype('int')
+ids = df['unique_id'].unique()
+random.seed(0)
+sample_ids = random.choices(ids, k=4)
+sample_df = df[df['unique_id'].isin(sample_ids)]
+info = M4Info[group]
+horizon = info.horizon
+valid = sample_df.groupby('unique_id').tail(horizon)
+train = sample_df.drop(valid.index)
+```
+
+
+```python
+def avg_smape(df):
+ """Computes the SMAPE by serie and then averages it across all series."""
+ full = df.merge(valid)
+ return (
+ evaluate(full, metrics=[smape])
+ .drop(columns='metric')
+ .set_index('unique_id')
+ .squeeze()
+ )
+```
+
+## Model
+
+```python
+fcst = MLForecast(
+ models=lgb.LGBMRegressor(random_state=0, verbosity=-1),
+ freq=1,
+ lags=[24 * (i+1) for i in range(7)],
+ lag_transforms={
+ 1: [RollingMean(window_size=24)],
+ 24: [RollingMean(window_size=24)],
+ 48: [ExponentiallyWeightedMean(alpha=0.3)],
+ },
+ num_threads=1,
+ target_transforms=[Differences([24])],
+)
+```
+
+
+```python
+horizon = 24
+# the following will train 24 models, one for each horizon
+individual_fcst = fcst.fit(train, max_horizon=horizon)
+individual_preds = individual_fcst.predict(horizon)
+avg_smape_individual = avg_smape(individual_preds).rename('individual')
+# the following will train a single model and use the recursive strategy
+recursive_fcst = fcst.fit(train)
+recursive_preds = recursive_fcst.predict(horizon)
+avg_smape_recursive = avg_smape(recursive_preds).rename('recursive')
+# results
+print('Average SMAPE per method and serie')
+avg_smape_individual.to_frame().join(avg_smape_recursive).applymap('{:.1%}'.format)
+```
+
+``` text
+Average SMAPE per method and serie
+```
+
+| | individual | recursive |
+|-----------|------------|-----------|
+| unique_id | | |
+| H196 | 0.3% | 0.3% |
+| H256 | 0.4% | 0.3% |
+| H381 | 20.9% | 9.5% |
+| H413 | 11.9% | 13.6% |
+
diff --git a/mlforecast/docs/how-to-guides/predict_callbacks.html.mdx b/mlforecast/docs/how-to-guides/predict_callbacks.html.mdx
new file mode 100644
index 00000000..430b98f2
--- /dev/null
+++ b/mlforecast/docs/how-to-guides/predict_callbacks.html.mdx
@@ -0,0 +1,135 @@
+---
+description: Get access to the input features and predictions in each forecasting horizon
+output-file: predict_callbacks.html
+title: Predict callbacks
+---
+
+
+If you want to do something to the input before predicting or something
+to the output before it gets used to update the target (and thus the
+next features that rely on lags), you can pass a function to run at any
+of these times.
+
+Here are a couple of examples:
+
+```python
+import copy
+
+import lightgbm as lgb
+import numpy as np
+from IPython.display import display
+
+from mlforecast import MLForecast
+from mlforecast.utils import generate_daily_series
+```
+
+
+```python
+series = generate_daily_series(1)
+```
+
+## Before predicting
+
+### Inspecting the input
+
+We can define a function that displays our input dataframe before
+predicting.
+
+```python
+def inspect_input(new_x):
+ """Displays the model inputs to inspect them"""
+ display(new_x)
+ return new_x
+```
+
+And now we can pass this function to the `before_predict_callback`
+argument of
+[`MLForecast.predict`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.predict).
+
+```python
+fcst = MLForecast(lgb.LGBMRegressor(verbosity=-1), freq='D', lags=[1, 2])
+fcst.fit(series, static_features=['unique_id'])
+preds = fcst.predict(2, before_predict_callback=inspect_input)
+preds
+```
+
+| | unique_id | lag1 | lag2 |
+|-----|-----------|---------|----------|
+| 0 | id_0 | 4.15593 | 3.000028 |
+
+| | unique_id | lag1 | lag2 |
+|-----|-----------|----------|---------|
+| 0 | id_0 | 5.250205 | 4.15593 |
+
+| | unique_id | ds | LGBMRegressor |
+|-----|-----------|------------|---------------|
+| 0 | id_0 | 2000-08-10 | 5.250205 |
+| 1 | id_0 | 2000-08-11 | 6.241739 |
+
+### Saving the input features
+
+Saving the features that are sent as input to the model in each
+timestamp can be helpful, for example to estimate SHAP values. This can
+be easily achieved with the
+[`SaveFeatures`](https://Nixtla.github.io/mlforecast/callbacks.html#savefeatures)
+callback.
+
+```python
+from mlforecast.callbacks import SaveFeatures
+```
+
+
+```python
+fcst = MLForecast(lgb.LGBMRegressor(verbosity=-1), freq='D', lags=[1])
+fcst.fit(series, static_features=['unique_id'])
+save_features_cbk = SaveFeatures()
+fcst.predict(2, before_predict_callback=save_features_cbk);
+```
+
+Once we’ve called predict we can just retrieve the features.
+
+```python
+save_features_cbk.get_features()
+```
+
+| | unique_id | lag1 |
+|-----|-----------|----------|
+| 0 | id_0 | 4.155930 |
+| 1 | id_0 | 5.281643 |
+
+## After predicting
+
+When predicting with the recursive strategy (the default) the
+predictions for each timestamp are used to update the target and
+recompute the features. If you want to do something to these predictions
+before that happens you can use the `after_predict_callback` argument of
+[`MLForecast.predict`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.predict).
+
+### Increasing predictions values
+
+Suppose we know that our model always underestimates and we want to
+prevent that from happening by making our predictions 10% higher. We can
+achieve that with the following:
+
+```python
+def increase_predictions(predictions):
+ """Increases all predictions by 10%"""
+ return 1.1 * predictions
+```
+
+
+```python
+fcst = MLForecast(
+ {'model': lgb.LGBMRegressor(verbosity=-1)},
+ freq='D',
+ date_features=['dayofweek'],
+)
+fcst.fit(series)
+original_preds = fcst.predict(2)
+scaled_preds = fcst.predict(2, after_predict_callback=increase_predictions)
+np.testing.assert_array_less(
+ original_preds['model'].values,
+ scaled_preds['model'].values,
+)
+```
+
diff --git a/mlforecast/docs/how-to-guides/predict_subset.html.mdx b/mlforecast/docs/how-to-guides/predict_subset.html.mdx
new file mode 100644
index 00000000..614c83f1
--- /dev/null
+++ b/mlforecast/docs/how-to-guides/predict_subset.html.mdx
@@ -0,0 +1,52 @@
+---
+description: Compute predictions for only a subset of the training ids
+output-file: predict_subset.html
+title: Predicting a subset of ids
+---
+
+
+```python
+from lightgbm import LGBMRegressor
+from fastcore.test import test_fail
+
+from mlforecast import MLForecast
+from mlforecast.utils import generate_daily_series
+```
+
+
+```python
+series = generate_daily_series(5)
+fcst = MLForecast({'lgb': LGBMRegressor(verbosity=-1)}, freq='D', date_features=['dayofweek'])
+fcst.fit(series)
+all_preds = fcst.predict(1)
+all_preds
+```
+
+| | unique_id | ds | lgb |
+|-----|-----------|------------|----------|
+| 0 | id_0 | 2000-08-10 | 3.728396 |
+| 1 | id_1 | 2000-04-07 | 4.749133 |
+| 2 | id_2 | 2000-06-16 | 4.749133 |
+| 3 | id_3 | 2000-08-30 | 2.758949 |
+| 4 | id_4 | 2001-01-08 | 3.331394 |
+
+By default all series seen during training will be forecasted with the
+predict method. If you’re only interested in predicting a couple of them
+you can use the `ids` argument.
+
+```python
+fcst.predict(1, ids=['id_0', 'id_4'])
+```
+
+| | unique_id | ds | lgb |
+|-----|-----------|------------|----------|
+| 0 | id_0 | 2000-08-10 | 3.728396 |
+| 1 | id_4 | 2001-01-08 | 3.331394 |
+
+Note that the ids must’ve been seen during training, if you try to
+predict an id that wasn’t there you’ll get an error.
+
+```python
+test_fail(lambda: fcst.predict(1, ids=['fake_id']), contains='fake_id')
+```
+
diff --git a/mlforecast/docs/how-to-guides/prediction_intervals.html.mdx b/mlforecast/docs/how-to-guides/prediction_intervals.html.mdx
new file mode 100644
index 00000000..b37a8e78
--- /dev/null
+++ b/mlforecast/docs/how-to-guides/prediction_intervals.html.mdx
@@ -0,0 +1,349 @@
+---
+description: In this example, we'll implement prediction intervals
+output-file: prediction_intervals.html
+title: Probabilistic forecasting
+---
+
+
+> **Prerequesites**
+>
+> This tutorial assumes basic familiarity with MLForecast. For a minimal
+> example visit the [Quick
+> Start](https://nixtla.github.io/mlforecast/docs/getting-started/quick_start_local.html)
+
+## Introduction
+
+When we generate a forecast, we usually produce a single value known as
+the point forecast. This value, however, doesn’t tell us anything about
+the uncertainty associated with the forecast. To have a measure of this
+uncertainty, we need **prediction intervals**.
+
+A prediction interval is a range of values that the forecast can take
+with a given probability. Hence, a 95% prediction interval should
+contain a range of values that include the actual future value with
+probability 95%. Probabilistic forecasting aims to generate the full
+forecast distribution. Point forecasting, on the other hand, usually
+returns the mean or the median or said distribution. However, in
+real-world scenarios, it is better to forecast not only the most
+probable future outcome, but many alternative outcomes as well.
+
+With [MLForecast](https://nixtla.github.io/mlforecast/) you can train
+`sklearn` models to generate point forecasts. It also takes the
+advantages of `ConformalPrediction` to generate the same point forecasts
+and adds them prediction intervals. By the end of this tutorial, you’ll
+have a good understanding of how to add probabilistic intervals to
+`sklearn` models for time series forecasting. Furthermore, you’ll also
+learn how to generate plots with the historical data, the point
+forecasts, and the prediction intervals.
+
+> **Important**
+>
+> Although the terms are often confused, prediction intervals are not
+> the same as [confidence
+> intervals](https://robjhyndman.com/hyndsight/intervals/).
+
+> **Warning**
+>
+> In practice, most prediction intervals are too narrow since models do
+> not account for all sources of uncertainty. A discussion about this
+> can be found [here](https://robjhyndman.com/hyndsight/narrow-pi/).
+
+**Outline:**
+
+1. Install libraries
+2. Load and explore the data
+3. Train models
+4. Plot prediction intervals
+
+> **Tip**
+>
+> You can use Colab to run this Notebook interactively
+>
+
+## Install libraries
+
+Install the necessary packages using
+`pip install mlforecast utilsforecast`
+
+## Load and explore the data
+
+For this example, we’ll use the hourly dataset from the [M4
+Competition](https://www.sciencedirect.com/science/article/pii/S0169207019301128).
+We first need to download the data from a URL and then load it as a
+`pandas` dataframe. Notice that we’ll load the train and the test data
+separately. We’ll also rename the `y` column of the test data as
+`y_test`.
+
+```python
+import pandas as pd
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+train = pd.read_csv('https://auto-arima-results.s3.amazonaws.com/M4-Hourly.csv')
+test = pd.read_csv('https://auto-arima-results.s3.amazonaws.com/M4-Hourly-test.csv')
+```
+
+
+```python
+train.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|-----|-------|
+| 0 | H1 | 1 | 605.0 |
+| 1 | H1 | 2 | 586.0 |
+| 2 | H1 | 3 | 586.0 |
+| 3 | H1 | 4 | 559.0 |
+| 4 | H1 | 5 | 511.0 |
+
+```python
+test.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|-----|-------|
+| 0 | H1 | 701 | 619.0 |
+| 1 | H1 | 702 | 565.0 |
+| 2 | H1 | 703 | 532.0 |
+| 3 | H1 | 704 | 495.0 |
+| 4 | H1 | 705 | 481.0 |
+
+Since the goal of this notebook is to generate prediction intervals,
+we’ll only use the first 8 series of the dataset to reduce the total
+computational time.
+
+```python
+n_series = 8
+uids = train['unique_id'].unique()[:n_series] # select first n_series of the dataset
+train = train.query('unique_id in @uids')
+test = test.query('unique_id in @uids')
+```
+
+We can plot these series using the `plot_series` function from the
+[utilsforecast](https://nixtla.github.io/utilsforecast/plotting.html)
+library. This function has multiple parameters, and the required ones to
+generate the plots in this notebook are explained below.
+
+- `df`: A `pandas` dataframe with columns \[`unique_id`, `ds`, `y`\].
+- `forecasts_df`: A `pandas` dataframe with columns \[`unique_id`,
+ `ds`\] and models.
+- `plot_random`: bool = `True`. Plots the time series randomly.
+- `models`: List\[str\]. A list with the models we want to plot.
+- `level`: List\[float\]. A list with the prediction intervals we want
+ to plot.
+- `engine`: str = `matplotlib`. It can also be `plotly`. `plotly`
+ generates interactive plots, while `matplotlib` generates static
+ plots.
+
+```python
+fig = plot_series(train, test.rename(columns={'y': 'y_test'}), models=['y_test'], plot_random=False)
+```
+
+
+
+## Train models
+
+MLForecast can train multiple models that follow the `sklearn` syntax
+(`fit` and `predict`) on different time series efficiently.
+
+For this example, we’ll use the following `sklearn` baseline models:
+
+- [Lasso](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.Lasso.html)
+- [LinearRegression](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html)
+- [Ridge](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.Ridge.html)
+- [K-Nearest
+ Neighbors](https://scikit-learn.org/stable/modules/generated/sklearn.neighbors.KNeighborsRegressor.html)
+- [Multilayer Perceptron
+ (NeuralNetwork)](https://scikit-learn.org/stable/modules/generated/sklearn.neural_network.MLPRegressor.html)
+
+To use these models, we first need to import them from `sklearn` and
+then we need to instantiate them.
+
+```python
+from mlforecast import MLForecast
+from mlforecast.target_transforms import Differences
+from mlforecast.utils import PredictionIntervals
+from sklearn.linear_model import Lasso, LinearRegression, Ridge
+from sklearn.neighbors import KNeighborsRegressor
+from sklearn.neural_network import MLPRegressor
+```
+
+
+```python
+# Create a list of models and instantiation parameters
+models = [
+ KNeighborsRegressor(),
+ Lasso(),
+ LinearRegression(),
+ MLPRegressor(),
+ Ridge(),
+]
+```
+
+To instantiate a new MLForecast object, we need the following
+parameters:
+
+- `models`: The list of models defined in the previous step.
+- `target_transforms`: Transformations to apply to the target before
+ computing the features. These are restored at the forecasting step.
+- `lags`: Lags of the target to use as features.
+
+```python
+mlf = MLForecast(
+ models=[Ridge(), Lasso(), LinearRegression(), KNeighborsRegressor(), MLPRegressor(random_state=0)],
+ freq=1,
+ target_transforms=[Differences([1])],
+ lags=[24 * (i+1) for i in range(7)],
+)
+```
+
+Now we’re ready to generate the point forecasts and the prediction
+intervals. To do this, we’ll use the `fit` method, which takes the
+following arguments:
+
+- `data`: Series data in long format.
+- `id_col`: Column that identifies each series. In our case,
+ `unique_id`.
+- `time_col`: Column that identifies each timestep, its values can be
+ timestamps or integers. In our case, `ds`.
+- `target_col`: Column that contains the target. In our case, `y`.
+- `prediction_intervals`: A `PredicitonIntervals` class. The class
+ takes two parameters: `n_windows` and `h`. `n_windows` represents
+ the number of cross-validation windows used to calibrate the
+ intervals and `h` is the forecast horizon. The strategy will adjust
+ the intervals for each horizon step, resulting in different widths
+ for each step.
+
+```python
+mlf.fit(
+ train,
+ prediction_intervals=PredictionIntervals(n_windows=10, h=48),
+);
+```
+
+After fitting the models, we will call the `predict` method to generate
+forecasts with prediction intervals. The method takes the following
+arguments:
+
+- `horizon`: An integer that represent the forecasting horizon. In
+ this case, we’ll forecast the next 48 hours.
+- `level`: A list of floats with the confidence levels of the
+ prediction intervals. For example, `level=[95]` means that the range
+ of values should include the actual future value with probability
+ 95%.
+
+```python
+levels = [50, 80, 95]
+forecasts = mlf.predict(48, level=levels)
+forecasts.head()
+```
+
+| | unique_id | ds | Ridge | Lasso | LinearRegression | KNeighborsRegressor | MLPRegressor | Ridge-lo-95 | Ridge-lo-80 | Ridge-lo-50 | ... | KNeighborsRegressor-lo-50 | KNeighborsRegressor-hi-50 | KNeighborsRegressor-hi-80 | KNeighborsRegressor-hi-95 | MLPRegressor-lo-95 | MLPRegressor-lo-80 | MLPRegressor-lo-50 | MLPRegressor-hi-50 | MLPRegressor-hi-80 | MLPRegressor-hi-95 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | H1 | 701 | 612.418170 | 612.418079 | 612.418170 | 615.2 | 612.651532 | 590.473256 | 594.326570 | 603.409944 | ... | 609.45 | 620.95 | 627.20 | 631.310 | 584.736193 | 591.084898 | 597.462107 | 627.840957 | 634.218166 | 640.566870 |
+| 1 | H1 | 702 | 552.309298 | 552.308073 | 552.309298 | 551.6 | 548.791801 | 498.721501 | 518.433843 | 532.710850 | ... | 535.85 | 567.35 | 569.16 | 597.525 | 497.308756 | 500.417799 | 515.452396 | 582.131207 | 597.165804 | 600.274847 |
+| 2 | H1 | 703 | 494.943384 | 494.943367 | 494.943384 | 509.6 | 490.226796 | 448.253304 | 463.266064 | 475.006125 | ... | 492.70 | 526.50 | 530.92 | 544.180 | 424.587658 | 436.042788 | 448.682502 | 531.771091 | 544.410804 | 555.865935 |
+| 3 | H1 | 704 | 462.815779 | 462.815363 | 462.815779 | 474.6 | 459.619069 | 409.975219 | 422.243593 | 436.128272 | ... | 451.80 | 497.40 | 510.26 | 525.500 | 379.291083 | 392.580306 | 413.353178 | 505.884959 | 526.657832 | 539.947054 |
+| 4 | H1 | 705 | 440.141034 | 440.140586 | 440.141034 | 451.6 | 438.091712 | 377.999588 | 392.523016 | 413.474795 | ... | 427.40 | 475.80 | 488.96 | 503.945 | 348.618034 | 362.503767 | 386.303325 | 489.880099 | 513.679657 | 527.565389 |
+
+```python
+test = test.merge(forecasts, how='left', on=['unique_id', 'ds'])
+```
+
+## Plot prediction intervals
+
+To plot the point and the prediction intervals, we’ll use the
+`plot_series` function again. Notice that now we also need to specify
+the model and the levels that we want to plot.
+
+### KNeighborsRegressor
+
+```python
+fig = plot_series(
+ train,
+ test,
+ plot_random=False,
+ models=['KNeighborsRegressor'],
+ level=levels,
+ max_insample_length=48
+)
+```
+
+
+
+### Lasso
+
+```python
+fig = plot_series(
+ train,
+ test,
+ plot_random=False,
+ models=['Lasso'],
+ level=levels,
+ max_insample_length=48
+)
+```
+
+
+
+### LineaRegression
+
+```python
+fig = plot_series(
+ train,
+ test,
+ plot_random=False,
+ models=['LinearRegression'],
+ level=levels,
+ max_insample_length=48
+)
+```
+
+
+
+### MLPRegressor
+
+```python
+fig = plot_series(
+ train,
+ test,
+ plot_random=False,
+ models=['MLPRegressor'],
+ level=levels,
+ max_insample_length=48
+)
+```
+
+
+
+### Ridge
+
+```python
+fig = plot_series(
+ train,
+ test,
+ plot_random=False,
+ models=['Ridge'],
+ level=levels,
+ max_insample_length=48
+)
+```
+
+
+
+From these plots, we can conclude that the uncertainty around each
+forecast varies according to the model that is being used. For the same
+time series, one model can predict a wider range of possible future
+values than others.
+
+## References
+
+- [Kamile Stankeviciute, Ahmed M. Alaa and Mihaela van der Schaar
+ (2021). “Conformal Time-Series
+ Forecasting”](https://proceedings.neurips.cc/paper/2021/file/312f1ba2a72318edaaa995a67835fad5-Paper.pdf)
+- [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ principles and practice, The Statistical Forecasting
+ Perspective”](https://otexts.com/fpp3/perspective.html).
+
diff --git a/mlforecast/docs/how-to-guides/sample_weights.html.mdx b/mlforecast/docs/how-to-guides/sample_weights.html.mdx
new file mode 100644
index 00000000..16490b42
--- /dev/null
+++ b/mlforecast/docs/how-to-guides/sample_weights.html.mdx
@@ -0,0 +1,80 @@
+---
+description: Provide a column to pass through to the underlying models as sample weights
+output-file: sample_weights.html
+title: Sample weights
+---
+
+
+## Data setup
+
+```python
+import numpy as np
+from mlforecast.utils import generate_daily_series
+```
+
+
+```python
+series = generate_daily_series(2)
+series['weight'] = np.random.default_rng(seed=0).random(series.shape[0])
+series.head(2)
+```
+
+| | unique_id | ds | y | weight |
+|-----|-----------|------------|----------|----------|
+| 0 | id_0 | 2000-01-01 | 0.357595 | 0.636962 |
+| 1 | id_0 | 2000-01-02 | 1.301382 | 0.269787 |
+
+## Creating forecast object
+
+```python
+import lightgbm as lgb
+from sklearn.linear_model import LinearRegression
+
+from mlforecast import MLForecast
+```
+
+
+```python
+fcst = MLForecast(
+ models={
+ 'lr': LinearRegression(),
+ 'lgbm': lgb.LGBMRegressor(verbosity=-1),
+ },
+ freq='D',
+ lags=[1],
+ date_features=['dayofweek'],
+)
+```
+
+## Forecasting
+
+You can provide the `weight_col` argument to
+[`MLForecast.fit`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.fit)
+to indicate which column should be used as the sample weights.
+
+```python
+fcst.fit(series, weight_col='weight').predict(1)
+```
+
+| | unique_id | ds | lr | lgbm |
+|-----|-----------|------------|----------|----------|
+| 0 | id_0 | 2000-08-10 | 3.336019 | 5.283677 |
+| 1 | id_1 | 2000-04-07 | 3.300786 | 4.230655 |
+
+## Cross validation
+
+You can provide the `weight_col` argument to
+[`MLForecast.cross_validation`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.cross_validation)
+to indicate which column should be used as the sample weights.
+
+```python
+fcst.cross_validation(series, n_windows=2, h=1, weight_col='weight')
+```
+
+| | unique_id | ds | cutoff | y | lr | lgbm |
+|-----|-----------|------------|------------|----------|----------|----------|
+| 0 | id_0 | 2000-08-08 | 2000-08-07 | 3.436325 | 2.770717 | 3.242790 |
+| 1 | id_1 | 2000-04-05 | 2000-04-04 | 2.430276 | 2.687932 | 2.075247 |
+| 2 | id_0 | 2000-08-09 | 2000-08-08 | 4.136771 | 3.095140 | 4.239010 |
+| 3 | id_1 | 2000-04-06 | 2000-04-05 | 3.363522 | 3.016661 | 3.436962 |
+
diff --git a/mlforecast/docs/how-to-guides/sklearn_pipelines.html.mdx b/mlforecast/docs/how-to-guides/sklearn_pipelines.html.mdx
new file mode 100644
index 00000000..88d934f1
--- /dev/null
+++ b/mlforecast/docs/how-to-guides/sklearn_pipelines.html.mdx
@@ -0,0 +1,153 @@
+---
+description: Leverage scikit-learn's composability to define pipelines as models
+output-file: sklearn_pipelines.html
+title: Using scikit-learn pipelines
+---
+
+
+mlforecast takes scikit-learn estimators as models, which means you can
+provide [scikit-learn’s
+pipelines](https://scikit-learn.org/stable/modules/generated/sklearn.pipeline.Pipeline.html)
+as models in order to further apply transformations to the data before
+passing it to the model.
+
+## Data setup
+
+```python
+from mlforecast.utils import generate_daily_series
+```
+
+
+```python
+series = generate_daily_series(5)
+series.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|----------|
+| 0 | id_0 | 2000-01-01 | 0.428973 |
+| 1 | id_0 | 2000-01-02 | 1.423626 |
+| 2 | id_0 | 2000-01-03 | 2.311782 |
+| 3 | id_0 | 2000-01-04 | 3.192191 |
+| 4 | id_0 | 2000-01-05 | 4.148767 |
+
+## Pipelines definition
+
+Suppose that you want to use a linear regression model with the lag1 and
+the day of the week as features. mlforecast returns the day of the week
+as a single column, however, that’s not the optimal format for a linear
+regression model, which benefits more from having indicator columns for
+each day of the week (removing one to avoid colinearity). We can achieve
+this by using [scikit-learn’s
+OneHotEncoder](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.OneHotEncoder.html)
+and then fitting our linear regression model, which we can do in the
+following way:
+
+```python
+from mlforecast import MLForecast
+from sklearn.compose import ColumnTransformer
+from sklearn.linear_model import LinearRegression
+from sklearn.pipeline import make_pipeline
+from sklearn.preprocessing import OneHotEncoder
+```
+
+
+```python
+fcst = MLForecast(
+ models=[],
+ freq='D',
+ lags=[1],
+ date_features=['dayofweek']
+)
+X, y = fcst.preprocess(series, return_X_y=True)
+X.head()
+```
+
+| | lag1 | dayofweek |
+|-----|----------|-----------|
+| 1 | 0.428973 | 6 |
+| 2 | 1.423626 | 0 |
+| 3 | 2.311782 | 1 |
+| 4 | 3.192191 | 2 |
+| 5 | 4.148767 | 3 |
+
+This is what will be passed to our model, so we’d like to get the
+`dayofweek` column and perform one hot encoding, leaving the `lag1`
+column untouched. We can achieve that with the following:
+
+```python
+ohe = ColumnTransformer(
+ transformers=[
+ ('encoder', OneHotEncoder(drop='first'), ['dayofweek'])
+ ],
+ remainder='passthrough',
+)
+X_transformed = ohe.fit_transform(X)
+X_transformed.shape
+```
+
+``` text
+(1096, 7)
+```
+
+We can see that our data now has 7 columns, 1 for the lag plus 6 for the
+days of the week (we dropped the first one).
+
+```python
+ohe.get_feature_names_out()
+```
+
+``` text
+array(['encoder__dayofweek_1', 'encoder__dayofweek_2',
+ 'encoder__dayofweek_3', 'encoder__dayofweek_4',
+ 'encoder__dayofweek_5', 'encoder__dayofweek_6', 'remainder__lag1'],
+ dtype=object)
+```
+
+## Training
+
+We can now build a pipeline that does this and then passes it to our
+linear regression model.
+
+```python
+model = make_pipeline(ohe, LinearRegression())
+```
+
+And provide this as a model to mlforecast
+
+```python
+fcst = MLForecast(
+ models={'ohe_lr': model},
+ freq='D',
+ lags=[1],
+ date_features=['dayofweek']
+)
+fcst.fit(series)
+```
+
+``` text
+MLForecast(models=[ohe_lr], freq=, lag_features=['lag1'], date_features=['dayofweek'], num_threads=1)
+```
+
+## Forecasting
+
+Finally, we compute the forecasts.
+
+```python
+fcst.predict(1)
+```
+
+| | unique_id | ds | ohe_lr |
+|-----|-----------|------------|----------|
+| 0 | id_0 | 2000-08-10 | 4.312748 |
+| 1 | id_1 | 2000-04-07 | 4.537019 |
+| 2 | id_2 | 2000-06-16 | 4.160505 |
+| 3 | id_3 | 2000-08-30 | 3.777040 |
+| 4 | id_4 | 2001-01-08 | 2.676933 |
+
+## Summary
+
+You can provide complex scikit-learn pipelines as models to mlforecast,
+which allows you to perform different transformations depending on the
+model and use any of scikit-learn’s compatible estimators.
+
diff --git a/mlforecast/docs/how-to-guides/target_transforms_guide.html.mdx b/mlforecast/docs/how-to-guides/target_transforms_guide.html.mdx
new file mode 100644
index 00000000..9fb0c08c
--- /dev/null
+++ b/mlforecast/docs/how-to-guides/target_transforms_guide.html.mdx
@@ -0,0 +1,240 @@
+---
+description: Seamlessly transform target values
+output-file: target_transforms_guide.html
+title: Target transformations
+---
+
+
+Since mlforecast uses a single global model it can be helpful to apply
+some transformations to the target to ensure that all series have
+similar distributions. They can also help remove trend for models that
+can’t deal with it out of the box.
+
+## Data setup
+
+For this example we’ll use a single serie from the M4 dataset.
+
+```python
+import matplotlib.pyplot as plt
+import numpy as np
+import pandas as pd
+from datasetsforecast.m4 import M4
+from sklearn.base import BaseEstimator
+
+from mlforecast import MLForecast
+from mlforecast.target_transforms import Differences, LocalStandardScaler
+```
+
+
+```python
+data_path = 'data'
+await M4.async_download(data_path, group='Hourly')
+df, *_ = M4.load(data_path, 'Hourly')
+df['ds'] = df['ds'].astype('int32')
+serie = df[df['unique_id'].eq('H196')]
+```
+
+## Local transformations
+
+> Transformations applied per serie
+
+### Differences
+
+We’ll take a look at our serie to see possible differences that would
+help our models.
+
+```python
+def plot(series, fname):
+ n_series = len(series)
+ fig, ax = plt.subplots(ncols=n_series, figsize=(7 * n_series, 6), squeeze=False)
+ for (title, serie), axi in zip(series.items(), ax.flat):
+ serie.set_index('ds')['y'].plot(title=title, ax=axi)
+ fig.savefig(f'../../figs/{fname}', bbox_inches='tight')
+ plt.close()
+```
+
+
+```python
+plot({'original': serie}, 'target_transforms__eda.png')
+```
+
+
+
+We can see that our data has a trend as well as a clear seasonality. We
+can try removing the trend first.
+
+```python
+fcst = MLForecast(
+ models=[],
+ freq=1,
+ target_transforms=[Differences([1])],
+)
+without_trend = fcst.preprocess(serie)
+plot({'original': serie, 'without trend': without_trend}, 'target_transforms__diff1.png')
+```
+
+
+
+The trend is gone, we can now try taking the 24 difference (subtract the
+value at the same hour in the previous day).
+
+```python
+fcst = MLForecast(
+ models=[],
+ freq=1,
+ target_transforms=[Differences([1, 24])],
+)
+without_trend_and_seasonality = fcst.preprocess(serie)
+plot({'original': serie, 'without trend and seasonality': without_trend_and_seasonality}, 'target_transforms__diff2.png')
+```
+
+
+
+### LocalStandardScaler
+
+We see that our serie is random noise now. Suppose we also want to
+standardize it, i.e. make it have a mean of 0 and variance of 1. We can
+add the LocalStandardScaler transformation after these differences.
+
+```python
+fcst = MLForecast(
+ models=[],
+ freq=1,
+ target_transforms=[Differences([1, 24]), LocalStandardScaler()],
+)
+standardized = fcst.preprocess(serie)
+plot({'original': serie, 'standardized': standardized}, 'target_transforms__standardized.png')
+standardized['y'].agg(['mean', 'var']).round(2)
+```
+
+``` text
+mean -0.0
+var 1.0
+Name: y, dtype: float64
+```
+
+
+
+Now that we’ve captured the components of the serie (trend +
+seasonality), we could try forecasting it with a model that always
+predicts 0, which will basically project the trend and seasonality.
+
+```python
+class Zeros(BaseEstimator):
+ def fit(self, X, y=None):
+ return self
+
+ def predict(self, X, y=None):
+ return np.zeros(X.shape[0])
+
+fcst = MLForecast(
+ models={'zeros_model': Zeros()},
+ freq=1,
+ target_transforms=[Differences([1, 24]), LocalStandardScaler()],
+)
+preds = fcst.fit(serie).predict(48)
+fig, ax = plt.subplots()
+pd.concat([serie.tail(24 * 10), preds]).set_index('ds').plot(ax=ax)
+plt.close()
+```
+
+
+
+## Global transformations
+
+> Transformations applied to all series
+
+### GlobalSklearnTransformer
+
+There are some transformations that don’t require to learn any
+parameters, such as applying logarithm for example. These can be easily
+defined using the
+[`GlobalSklearnTransformer`](https://Nixtla.github.io/mlforecast/target_transforms.html#globalsklearntransformer),
+which takes a scikit-learn compatible transformer and applies it to all
+series. Here’s an example on how to define a transformation that applies
+logarithm to each value of the series + 1, which can help avoid
+computing the log of 0.
+
+```python
+import numpy as np
+from sklearn.preprocessing import FunctionTransformer
+
+from mlforecast.target_transforms import GlobalSklearnTransformer
+
+sk_log1p = FunctionTransformer(func=np.log1p, inverse_func=np.expm1)
+fcst = MLForecast(
+ models={'zeros_model': Zeros()},
+ freq=1,
+ target_transforms=[GlobalSklearnTransformer(sk_log1p)],
+)
+log1p_transformed = fcst.preprocess(serie)
+plot({'original': serie, 'Log transformed': log1p_transformed}, 'target_transforms__log.png')
+```
+
+
+
+We can also combine this with local transformations. For example we can
+apply log first and then differencing.
+
+```python
+fcst = MLForecast(
+ models=[],
+ freq=1,
+ target_transforms=[GlobalSklearnTransformer(sk_log1p), Differences([1, 24])],
+)
+log_diffs = fcst.preprocess(serie)
+plot({'original': serie, 'Log + Differences': log_diffs}, 'target_transforms__log_diffs.png')
+```
+
+
+
+## Custom transformations
+
+> Implementing your own target transformations
+
+In order to implement your own target transformation you have to define
+a class that inherits from
+[`mlforecast.target_transforms.BaseTargetTransform`](https://Nixtla.github.io/mlforecast/target_transforms.html#basetargettransform)
+(this takes care of setting the column names as the `id_col`, `time_col`
+and `target_col` attributes) and implement the `fit_transform` and
+`inverse_transform` methods. Here’s an example on how to define a
+min-max scaler.
+
+```python
+from mlforecast.target_transforms import BaseTargetTransform
+```
+
+
+```python
+class LocalMinMaxScaler(BaseTargetTransform):
+ """Scales each serie to be in the [0, 1] interval."""
+ def fit_transform(self, df: pd.DataFrame) -> pd.DataFrame:
+ self.stats_ = df.groupby(self.id_col)[self.target_col].agg(['min', 'max'])
+ df = df.merge(self.stats_, on=self.id_col)
+ df[self.target_col] = (df[self.target_col] - df['min']) / (df['max'] - df['min'])
+ df = df.drop(columns=['min', 'max'])
+ return df
+
+ def inverse_transform(self, df: pd.DataFrame) -> pd.DataFrame:
+ df = df.merge(self.stats_, on=self.id_col)
+ for col in df.columns.drop([self.id_col, self.time_col, 'min', 'max']):
+ df[col] = df[col] * (df['max'] - df['min']) + df['min']
+ df = df.drop(columns=['min', 'max'])
+ return df
+```
+
+And now you can pass an instance of this class to the
+`target_transforms` argument.
+
+```python
+fcst = MLForecast(
+ models=[],
+ freq=1,
+ target_transforms=[LocalMinMaxScaler()],
+)
+minmax_scaled = fcst.preprocess(serie)
+plot({'original': serie, 'min-max scaled': minmax_scaled}, 'target_transforms__minmax.png')
+```
+
+
+
diff --git a/mlforecast/docs/how-to-guides/training_with_numpy.html.mdx b/mlforecast/docs/how-to-guides/training_with_numpy.html.mdx
new file mode 100644
index 00000000..809f4344
--- /dev/null
+++ b/mlforecast/docs/how-to-guides/training_with_numpy.html.mdx
@@ -0,0 +1,165 @@
+---
+description: Convert your dataframes to arrays to use less memory and train faster
+output-file: training_with_numpy.html
+title: Training with numpy arrays
+---
+
+
+Most of the machine learning libraries use numpy arrays, even when you
+provide a dataframe it ends up being converted into a numpy array. By
+providing an array to those models we can make the process faster, since
+the conversion will only happen once.
+
+## Data setup
+
+```python
+from mlforecast.utils import generate_daily_series
+```
+
+
+```python
+series = generate_daily_series(5)
+```
+
+## fit and cross_validation methods
+
+```python
+import numpy as np
+from lightgbm import LGBMRegressor
+from sklearn.linear_model import LinearRegression
+
+from mlforecast import MLForecast
+```
+
+
+```python
+fcst = MLForecast(
+ models={'lr': LinearRegression(), 'lgbm': LGBMRegressor(verbosity=-1)},
+ freq='D',
+ lags=[7, 14],
+ date_features=['dayofweek'],
+)
+```
+
+If you’re using the fit/cross_validation methods from
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+all you have to do to train with numpy arrays is provide the `as_numpy`
+argument, which will cast the features to an array before passing them
+to the models.
+
+```python
+fcst.fit(series, as_numpy=True)
+```
+
+``` text
+MLForecast(models=[lr, lgbm], freq=, lag_features=['lag7', 'lag14'], date_features=['dayofweek'], num_threads=1)
+```
+
+When predicting, the new features will also be cast to arrays, so it can
+also be faster.
+
+```python
+fcst.predict(1)
+```
+
+| | unique_id | ds | lr | lgbm |
+|-----|-----------|------------|----------|----------|
+| 0 | id_0 | 2000-08-10 | 5.268787 | 6.322262 |
+| 1 | id_1 | 2000-04-07 | 4.437316 | 5.213255 |
+| 2 | id_2 | 2000-06-16 | 3.246518 | 4.373904 |
+| 3 | id_3 | 2000-08-30 | 0.144860 | 1.285219 |
+| 4 | id_4 | 2001-01-08 | 2.211318 | 3.236700 |
+
+For cross_validation we also just need to specify `as_numpy=True`.
+
+```python
+cv_res = fcst.cross_validation(series, n_windows=2, h=2, as_numpy=True)
+```
+
+## preprocess method
+
+Having the features as a numpy array can also be helpful in cases where
+you have categorical columns and the library doesn’t support them, for
+example LightGBM with polars. In order to use categorical features with
+LightGBM and polars we have to convert them to their integer
+representation and tell LightGBM to treat those features as categorical,
+which we can achieve in the following way:
+
+```python
+series_pl = generate_daily_series(5, n_static_features=1, engine='polars')
+series_pl.head(2)
+```
+
+| unique_id | ds | y | static_0 |
+|-----------|---------------------|------------|----------|
+| cat | datetime\[ns\] | f64 | cat |
+| "id_0" | 2000-01-01 00:00:00 | 36.462689 | "84" |
+| "id_0" | 2000-01-02 00:00:00 | 121.008199 | "84" |
+
+```python
+fcst = MLForecast(
+ models=[],
+ freq='1d',
+ lags=[7, 14],
+ date_features=['weekday'],
+)
+```
+
+In order to get the features as an array with the preprocess method we
+also have to ask for the X, y tuple.
+
+```python
+X, y = fcst.preprocess(series_pl, return_X_y=True, as_numpy=True)
+X[:2]
+```
+
+``` text
+array([[ 0. , 20.30076749, 36.46268875, 6. ],
+ [ 0. , 119.51717097, 121.0081989 , 7. ]])
+```
+
+The feature names are available in `fcst.ts.features_order_`
+
+```python
+fcst.ts.features_order_
+```
+
+``` text
+['static_0', 'lag7', 'lag14', 'weekday']
+```
+
+Now we can just train a LightGBM model specifying the feature names and
+which features should be treated as categorical.
+
+```python
+model = LGBMRegressor(verbosity=-1)
+model.fit(
+ X=X,
+ y=y,
+ feature_name=fcst.ts.features_order_,
+ categorical_feature=['static_0', 'weekday'],
+);
+```
+
+We can now add this model to our models dict, as described in the
+[custom training guide](./custom_training.html).
+
+```python
+fcst.models_ = {'lgbm': model}
+```
+
+And use it to predict.
+
+```python
+fcst.predict(1)
+```
+
+| unique_id | ds | lgbm |
+|-----------|---------------------|------------|
+| cat | datetime\[ns\] | f64 |
+| "id_0" | 2000-08-10 00:00:00 | 448.796188 |
+| "id_1" | 2000-04-07 00:00:00 | 81.058211 |
+| "id_2" | 2000-06-16 00:00:00 | 4.450549 |
+| "id_3" | 2000-08-30 00:00:00 | 14.219603 |
+| "id_4" | 2001-01-08 00:00:00 | 87.361881 |
+
diff --git a/mlforecast/docs/how-to-guides/transfer_learning.html.mdx b/mlforecast/docs/how-to-guides/transfer_learning.html.mdx
new file mode 100644
index 00000000..f9555d3f
--- /dev/null
+++ b/mlforecast/docs/how-to-guides/transfer_learning.html.mdx
@@ -0,0 +1,189 @@
+---
+output-file: transfer_learning.html
+title: Transfer Learning
+---
+
+
+Transfer learning refers to the process of pre-training a flexible model
+on a large dataset and using it later on other data with little to no
+training. It is one of the most outstanding 🚀 achievements in Machine
+Learning and has many practical applications.
+
+For time series forecasting, the technique allows you to get
+lightning-fast predictions ⚡ bypassing the tradeoff between accuracy
+and speed (more than 30 times faster than our already fast
+[AutoARIMA](https://github.com/Nixtla/statsforecast) for a similar
+accuracy).
+
+This notebook shows how to generate a pre-trained model to forecast new
+time series never seen by the model.
+
+Table of Contents
+
+- Installing MLForecast
+- Load M3 Monthly Data
+- Instantiate NeuralForecast core, Fit, and save
+- Use the pre-trained model to predict on AirPassengers
+- Evaluate Results
+
+You can run these experiments with Google Colab.
+
+
+
+## Installing Libraries
+
+```python
+# !pip install mlforecast datasetsforecast utilsforecast s3fs
+```
+
+
+```python
+import lightgbm as lgb
+import numpy as np
+import pandas as pd
+from datasetsforecast.m3 import M3
+from sklearn.metrics import mean_absolute_error
+from utilsforecast.plotting import plot_series
+
+from mlforecast import MLForecast
+from mlforecast.target_transforms import Differences
+```
+
+## Load M3 Data
+
+The `M3` class will automatically download the complete M3 dataset and
+process it.
+
+It return three Dataframes: `Y_df` contains the values for the target
+variables, `X_df` contains exogenous calendar features and `S_df`
+contains static features for each time-series. For this example we will
+only use `Y_df`.
+
+If you want to use your own data just replace `Y_df`. Be sure to use a
+long format and have a simmilar structure than our data set.
+
+```python
+Y_df_M3, _, _ = M3.load(directory='./', group='Monthly')
+```
+
+In this tutorial we are only using `1_000` series to speed up
+computations. Remove the filter to use the whole dataset.
+
+```python
+fig = plot_series(Y_df_M3)
+```
+
+
+
+## Model Training
+
+Using the
+[`MLForecast.fit`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.fit)
+method you can train a set of models to your dataset. You can modify the
+hyperparameters of the model to get a better accuracy, in this case we
+will use the default hyperparameters of `lgb.LGBMRegressor`.
+
+```python
+models = [lgb.LGBMRegressor(verbosity=-1)]
+```
+
+The
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+object has the following parameters:
+
+- `models`: a list of sklearn-like (`fit` and `predict`) models.
+- `freq`: a string indicating the frequency of the data. See [panda’s
+ available
+ frequencies.](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases)
+- `differences`: Differences to take of the target before computing
+ the features. These are restored at the forecasting step.
+- `lags`: Lags of the target to use as features.
+
+In this example, we are only using `differences` and `lags` to produce
+features. See [the full
+documentation](https://nixtla.github.io/mlforecast/forecast.html) to see
+all available features.
+
+Any settings are passed into the constructor. Then you call its `fit`
+method and pass in the historical data frame `Y_df_M3`.
+
+```python
+fcst = MLForecast(
+ models=models,
+ lags=range(1, 13),
+ freq='MS',
+ target_transforms=[Differences([1, 12])],
+)
+fcst.fit(Y_df_M3);
+```
+
+## Transfer M3 to AirPassengers
+
+Now we can transfer the trained model to forecast `AirPassengers` with
+the
+[`MLForecast.predict`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.predict)
+method, we just have to pass the new dataframe to the `new_data`
+argument.
+
+```python
+Y_df = pd.read_csv('https://datasets-nixtla.s3.amazonaws.com/air-passengers.csv', parse_dates=['ds'])
+
+# We define the train df.
+Y_train_df = Y_df[Y_df.ds<='1959-12-31'] # 132 train
+Y_test_df = Y_df[Y_df.ds>'1959-12-31'] # 12 test
+```
+
+
+```python
+Y_hat_df = fcst.predict(h=12, new_df=Y_train_df)
+Y_hat_df.head()
+```
+
+| | unique_id | ds | LGBMRegressor |
+|-----|---------------|------------|---------------|
+| 0 | AirPassengers | 1960-01-01 | 422.740096 |
+| 1 | AirPassengers | 1960-02-01 | 399.480193 |
+| 2 | AirPassengers | 1960-03-01 | 458.220289 |
+| 3 | AirPassengers | 1960-04-01 | 442.960385 |
+| 4 | AirPassengers | 1960-05-01 | 461.700482 |
+
+```python
+Y_hat_df = Y_test_df.merge(Y_hat_df, how='left', on=['unique_id', 'ds'])
+```
+
+
+```python
+fig = plot_series(Y_train_df, Y_hat_df)
+```
+
+
+
+## Evaluate Results
+
+We evaluate the forecasts of the pre-trained model with the Mean
+Absolute Error (`mae`).
+
+$$
+
+\qquad MAE = \frac{1}{Horizon} \sum_{\tau} |y_{\tau} - \hat{y}_{\tau}|\qquad
+
+$$
+
+```python
+y_true = Y_test_df.y.values
+y_hat = Y_hat_df['LGBMRegressor'].values
+```
+
+
+```python
+print(f'LGBMRegressor MAE: {mean_absolute_error(y_hat, y_true):.3f}')
+print('ETS MAE: 16.222')
+print('AutoARIMA MAE: 18.551')
+```
+
+``` text
+LGBMRegressor MAE: 13.560
+ETS MAE: 16.222
+AutoARIMA MAE: 18.551
+```
+
diff --git a/mlforecast/docs/how-to-guides/transforming_exog.html.mdx b/mlforecast/docs/how-to-guides/transforming_exog.html.mdx
new file mode 100644
index 00000000..5552b5b8
--- /dev/null
+++ b/mlforecast/docs/how-to-guides/transforming_exog.html.mdx
@@ -0,0 +1,172 @@
+---
+description: Compute transformations on your exogenous features for MLForecast
+output-file: transforming_exog.html
+title: Transforming exogenous features
+---
+
+
+The MLForecast class allows you to compute lag transformations on your
+target, however, sometimes you want to also compute transformations on
+your dynamic exogenous features. This guide shows you how to accomplish
+that.
+
+## Data setup
+
+```python
+from mlforecast.utils import generate_series, generate_prices_for_series
+```
+
+
+```python
+series = generate_series(10, equal_ends=True)
+prices = generate_prices_for_series(series)
+prices.head(2)
+```
+
+| | ds | unique_id | price |
+|-----|------------|-----------|----------|
+| 0 | 2000-10-05 | 0 | 0.548814 |
+| 1 | 2000-10-06 | 0 | 0.715189 |
+
+Suppose that you have some series along with their prices for each id
+and date and you want to compute forecasts for the next 7 days. Since
+the price is a dynamic feature you have to provide the future values
+through `X_df` in
+[`MLForecast.predict`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.predict).
+
+If you want to use not only the price but the lag7 of the price and the
+expanding mean of the lag1 for example, you can compute them before
+training, merge them with your series and then provide the future values
+through `X_df`. Consider the following example.
+
+## Computing the transformations
+
+```python
+from mlforecast.lag_transforms import ExpandingMean
+
+from mlforecast.feature_engineering import transform_exog
+```
+
+
+```python
+transformed_prices = transform_exog(prices, lags=[7], lag_transforms={1: [ExpandingMean()]})
+transformed_prices.head(10)
+```
+
+| | ds | unique_id | price | price_lag7 | price_expanding_mean_lag1 |
+|-----|------------|-----------|----------|------------|---------------------------|
+| 0 | 2000-10-05 | 0 | 0.548814 | NaN | NaN |
+| 1 | 2000-10-06 | 0 | 0.715189 | NaN | 0.548814 |
+| 2 | 2000-10-07 | 0 | 0.602763 | NaN | 0.632001 |
+| 3 | 2000-10-08 | 0 | 0.544883 | NaN | 0.622255 |
+| 4 | 2000-10-09 | 0 | 0.423655 | NaN | 0.602912 |
+| 5 | 2000-10-10 | 0 | 0.645894 | NaN | 0.567061 |
+| 6 | 2000-10-11 | 0 | 0.437587 | NaN | 0.580200 |
+| 7 | 2000-10-12 | 0 | 0.891773 | 0.548814 | 0.559827 |
+| 8 | 2000-10-13 | 0 | 0.963663 | 0.715189 | 0.601320 |
+| 9 | 2000-10-14 | 0 | 0.383442 | 0.602763 | 0.641580 |
+
+You can now merge this with your original series
+
+```python
+series_with_prices = series.merge(transformed_prices, on=['unique_id', 'ds'])
+series_with_prices.head(10)
+```
+
+| | unique_id | ds | y | price | price_lag7 | price_expanding_mean_lag1 |
+|----|----|----|----|----|----|----|
+| 0 | 0 | 2000-10-05 | 0.322947 | 0.548814 | NaN | NaN |
+| 1 | 0 | 2000-10-06 | 1.218794 | 0.715189 | NaN | 0.548814 |
+| 2 | 0 | 2000-10-07 | 2.445887 | 0.602763 | NaN | 0.632001 |
+| 3 | 0 | 2000-10-08 | 3.481831 | 0.544883 | NaN | 0.622255 |
+| 4 | 0 | 2000-10-09 | 4.191721 | 0.423655 | NaN | 0.602912 |
+| 5 | 0 | 2000-10-10 | 5.395863 | 0.645894 | NaN | 0.567061 |
+| 6 | 0 | 2000-10-11 | 6.264447 | 0.437587 | NaN | 0.580200 |
+| 7 | 0 | 2000-10-12 | 0.284022 | 0.891773 | 0.548814 | 0.559827 |
+| 8 | 0 | 2000-10-13 | 1.462798 | 0.963663 | 0.715189 | 0.601320 |
+| 9 | 0 | 2000-10-14 | 2.035518 | 0.383442 | 0.602763 | 0.641580 |
+
+You can then define your forecast object. Note that you can still
+compute lag features based on the target as you normally would.
+
+```python
+from sklearn.linear_model import LinearRegression
+
+from mlforecast import MLForecast
+```
+
+
+```python
+fcst = MLForecast(
+ models=[LinearRegression()],
+ freq='D',
+ lags=[1],
+ date_features=['dayofweek'],
+)
+fcst.preprocess(series_with_prices, static_features=[], dropna=True).head()
+```
+
+| | unique_id | ds | y | price | price_lag7 | price_expanding_mean_lag1 | lag1 | dayofweek |
+|----|----|----|----|----|----|----|----|----|
+| 1 | 0 | 2000-10-06 | 1.218794 | 0.715189 | NaN | 0.548814 | 0.322947 | 4 |
+| 2 | 0 | 2000-10-07 | 2.445887 | 0.602763 | NaN | 0.632001 | 1.218794 | 5 |
+| 3 | 0 | 2000-10-08 | 3.481831 | 0.544883 | NaN | 0.622255 | 2.445887 | 6 |
+| 4 | 0 | 2000-10-09 | 4.191721 | 0.423655 | NaN | 0.602912 | 3.481831 | 0 |
+| 5 | 0 | 2000-10-10 | 5.395863 | 0.645894 | NaN | 0.567061 | 4.191721 | 1 |
+
+It’s important to note that the `dropna` argument only considers the
+null values generated by the lag features based on the target. If you
+want to drop all rows containing null values you have to do that in your
+original series.
+
+```python
+series_with_prices2 = series_with_prices.dropna()
+fcst.preprocess(series_with_prices2, dropna=True, static_features=[]).head()
+```
+
+| | unique_id | ds | y | price | price_lag7 | price_expanding_mean_lag1 | lag1 | dayofweek |
+|----|----|----|----|----|----|----|----|----|
+| 8 | 0 | 2000-10-13 | 1.462798 | 0.963663 | 0.715189 | 0.601320 | 0.284022 | 4 |
+| 9 | 0 | 2000-10-14 | 2.035518 | 0.383442 | 0.602763 | 0.641580 | 1.462798 | 5 |
+| 10 | 0 | 2000-10-15 | 3.043565 | 0.791725 | 0.544883 | 0.615766 | 2.035518 | 6 |
+| 11 | 0 | 2000-10-16 | 4.010109 | 0.528895 | 0.423655 | 0.631763 | 3.043565 | 0 |
+| 12 | 0 | 2000-10-17 | 5.416310 | 0.568045 | 0.645894 | 0.623190 | 4.010109 | 1 |
+
+You can now train the model.
+
+```python
+fcst.fit(series_with_prices2, static_features=[])
+```
+
+``` text
+MLForecast(models=[LinearRegression], freq=D, lag_features=['lag1'], date_features=['dayofweek'], num_threads=1)
+```
+
+And predict using the prices. Note that you can provide the dataframe
+with the full history and mlforecast will filter the required dates for
+the forecasting horizon.
+
+```python
+fcst.predict(1, X_df=transformed_prices).head()
+```
+
+| | unique_id | ds | LinearRegression |
+|-----|-----------|------------|------------------|
+| 0 | 0 | 2001-05-15 | 3.803967 |
+| 1 | 1 | 2001-05-15 | 3.512489 |
+| 2 | 2 | 2001-05-15 | 3.170019 |
+| 3 | 3 | 2001-05-15 | 4.307121 |
+| 4 | 4 | 2001-05-15 | 3.018758 |
+
+In this example we have prices for the next 7 days, if you try to
+forecast a longer horizon you’ll get an error.
+
+```python
+from fastcore.test import test_fail
+```
+
+
+```python
+test_fail(lambda: fcst.predict(8, X_df=transformed_prices), contains='Found missing inputs in X_df')
+```
+
diff --git a/mlforecast/docs/tutorials/electricity_load_forecasting.html.mdx b/mlforecast/docs/tutorials/electricity_load_forecasting.html.mdx
new file mode 100644
index 00000000..88936bc3
--- /dev/null
+++ b/mlforecast/docs/tutorials/electricity_load_forecasting.html.mdx
@@ -0,0 +1,735 @@
+---
+description: >-
+ In this example we will show how to perform electricity load forecasting using
+ MLForecast alongside many models. We also compare them against the prophet
+ library.
+output-file: electricity_load_forecasting.html
+title: Electricity Load Forecast
+---
+
+
+## Introduction
+
+Some time series are generated from very low frequency data. These data
+generally exhibit multiple seasonalities. For example, hourly data may
+exhibit repeated patterns every hour (every 24 observations) or every
+day (every 24 \* 7, hours per day, observations). This is the case for
+electricity load. Electricity load may vary hourly, e.g., during the
+evenings electricity consumption may be expected to increase. But also,
+the electricity load varies by week. Perhaps on weekends there is an
+increase in electrical activity.
+
+In this example we will show how to model the two seasonalities of the
+time series to generate accurate forecasts in a short time. We will use
+hourly PJM electricity load data. The original data can be found
+[here](https://www.kaggle.com/datasets/robikscube/hourly-energy-consumption).
+
+## Libraries
+
+In this example we will use the following libraries:
+
+- [`mlforecast`](https://nixtla.github.io/mlforecast/). Accurate and
+ ⚡️ fast forecasting withc lassical machine learning models.
+- [`prophet`](https://github.com/facebook/prophet). Benchmark model
+ developed by Facebook.
+- [`utilsforecast`](https://nixtla.github.io/utilsforecast/). Library
+ with different functions for forecasting evaluation.
+
+If you have already installed the libraries you can skip the next cell,
+if not be sure to run it.
+
+```python
+# %%capture
+# !pip install prophet
+# !pip install -U mlforecast
+# !pip install -U utilsforecast
+```
+
+## Forecast using Multiple Seasonalities
+
+### Electricity Load Data
+
+According to the [dataset’s
+page](https://www.kaggle.com/datasets/robikscube/hourly-energy-consumption),
+
+> PJM Interconnection LLC (PJM) is a regional transmission organization
+> (RTO) in the United States. It is part of the Eastern Interconnection
+> grid operating an electric transmission system serving all or parts of
+> Delaware, Illinois, Indiana, Kentucky, Maryland, Michigan, New Jersey,
+> North Carolina, Ohio, Pennsylvania, Tennessee, Virginia, West
+> Virginia, and the District of Columbia. The hourly power consumption
+> data comes from PJM’s website and are in megawatts (MW).
+
+Let’s take a look to the data.
+
+```python
+import matplotlib.pyplot as plt
+import numpy as np
+import pandas as pd
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+pd.plotting.register_matplotlib_converters()
+plt.rc("figure", figsize=(10, 8))
+plt.rc("font", size=10)
+```
+
+
+```python
+data_url = 'https://raw.githubusercontent.com/panambY/Hourly_Energy_Consumption/master/data/PJM_Load_hourly.csv'
+df = pd.read_csv(data_url, parse_dates=['Datetime'])
+df.columns = ['ds', 'y']
+df.insert(0, 'unique_id', 'PJM_Load_hourly')
+df['ds'] = pd.to_datetime(df['ds'])
+df = df.sort_values(['unique_id', 'ds']).reset_index(drop=True)
+print(f'Shape of the data {df.shape}')
+df.tail()
+```
+
+``` text
+Shape of the data (32896, 3)
+```
+
+| | unique_id | ds | y |
+|-------|-----------------|---------------------|---------|
+| 32891 | PJM_Load_hourly | 2001-12-31 20:00:00 | 36392.0 |
+| 32892 | PJM_Load_hourly | 2001-12-31 21:00:00 | 35082.0 |
+| 32893 | PJM_Load_hourly | 2001-12-31 22:00:00 | 33890.0 |
+| 32894 | PJM_Load_hourly | 2001-12-31 23:00:00 | 32590.0 |
+| 32895 | PJM_Load_hourly | 2002-01-01 00:00:00 | 31569.0 |
+
+```python
+fig = plot_series(df)
+```
+
+
+
+We clearly observe that the time series exhibits seasonal patterns.
+Moreover, the time series contains `32,896` observations, so it is
+necessary to use very computationally efficient methods to display them
+in production.
+
+We are going to split our series in order to create a train and test
+set. The model will be tested using the last 24 hours of the timeseries.
+
+```python
+threshold_time = df['ds'].max() - pd.Timedelta(hours=24)
+
+# Split the dataframe
+df_train = df[df['ds'] <= threshold_time]
+df_last_24_hours = df[df['ds'] > threshold_time]
+```
+
+### Analizing Seasonalities
+
+First we must visualize the seasonalities of the model. As mentioned
+before, the electricity load presents seasonalities every 24 hours
+(Hourly) and every 24 \* 7 (Daily) hours. Therefore, we will use
+`[24, 24 * 7]` as the seasonalities for the model. In order to analize
+how they affect our series we are going to use the `Difference` method.
+
+```python
+from mlforecast import MLForecast
+from mlforecast.target_transforms import Differences
+```
+
+We can use the
+[`MLForecast.preprocess`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.preprocess)
+method to explore different transformations. It looks like these series
+have a strong seasonality on the hour of the day, so we can subtract the
+value from the same hour in the previous day to remove it. This can be
+done with the
+[`mlforecast.target_transforms.Differences`](https://Nixtla.github.io/mlforecast/target_transforms.html#differences)
+transformer, which we pass through `target_transforms`.
+
+In order to analize the trends individually and combined we are going to
+plot them individually and combined. Therefore, we can compare them
+against the original series. We can use the next function for that.
+
+```python
+def plot_differences(df, differences,fname):
+ prep = [df]
+ # Plot individual Differences
+ for d in differences:
+ fcst = MLForecast(
+ models=[], # we're not interested in modeling yet
+ freq='H', # our series have hourly frequency
+ target_transforms=[Differences([d])],
+ )
+ df_ = fcst.preprocess(df)
+ df_['unique_id'] = df_['unique_id'] + f'_{d}'
+ prep.append(df_)
+
+ # Plot combined Differences
+ fcst = MLForecast(
+ models=[], # we're not interested in modeling yet
+ freq='H', # our series have hourly frequency
+ target_transforms=[Differences([24, 24*7])],
+ )
+ df_ = fcst.preprocess(df)
+ df_['unique_id'] = df_['unique_id'] + f'_all_diff'
+ prep.append(df_)
+ prep = pd.concat(prep, ignore_index=True)
+ #return prep
+ n_series = len(prep['unique_id'].unique())
+ fig, ax = plt.subplots(nrows=n_series, figsize=(7 * n_series, 10*n_series), squeeze=False)
+ for title, axi in zip(prep['unique_id'].unique(), ax.flat):
+ df_ = prep[prep['unique_id'] == title]
+ df_.set_index('ds')['y'].plot(title=title, ax=axi)
+ fig.savefig(f'../../figs/{fname}', bbox_inches='tight')
+ plt.close()
+```
+
+Since the seasonalities are present at `24` hours (daily) and `24*7`
+(weekly) we are going to substract them from the serie using
+`Differences([24, 24*7])` and plot them.
+
+```python
+plot_differences(df=df_train, differences=[24, 24*7], fname='load_forecasting__differences.png')
+```
+
+
+
+As we can see when we extract the 24 difference (daily) in
+`PJM_Load_hourly_24` the series seem to stabilize sisnce the peaks seem
+more uniform in comparison with the original series `PJM_Load_hourly`.
+
+When we extrac the 24\*7 (weekly) `PJM_Load_hourly_168` difference we
+can see there is more periodicity in the peaks in comparison with the
+original series.
+
+Finally we can see the result from the combined result from substracting
+all the differences `PJM_Load_hourly_all_diff`.
+
+For modeling we are going to use both difference for the forecasting,
+therefore we are setting the argument `target_transforms` from the
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+object equal to `[Differences([24, 24*7])]`, if we wanted to include a
+yearly difference we would need to add the term `24*365`.
+
+```python
+fcst = MLForecast(
+ models=[], # we're not interested in modeling yet
+ freq='H', # our series have hourly frequency
+ target_transforms=[Differences([24, 24*7])],
+)
+prep = fcst.preprocess(df_train)
+prep
+```
+
+| | unique_id | ds | y |
+|-------|-----------------|---------------------|--------|
+| 192 | PJM_Load_hourly | 1998-04-09 02:00:00 | 831.0 |
+| 193 | PJM_Load_hourly | 1998-04-09 03:00:00 | 918.0 |
+| 194 | PJM_Load_hourly | 1998-04-09 04:00:00 | 760.0 |
+| 195 | PJM_Load_hourly | 1998-04-09 05:00:00 | 849.0 |
+| 196 | PJM_Load_hourly | 1998-04-09 06:00:00 | 710.0 |
+| ... | ... | ... | ... |
+| 32867 | PJM_Load_hourly | 2001-12-30 20:00:00 | 3417.0 |
+| 32868 | PJM_Load_hourly | 2001-12-30 21:00:00 | 3596.0 |
+| 32869 | PJM_Load_hourly | 2001-12-30 22:00:00 | 3501.0 |
+| 32870 | PJM_Load_hourly | 2001-12-30 23:00:00 | 3939.0 |
+| 32871 | PJM_Load_hourly | 2001-12-31 00:00:00 | 4235.0 |
+
+```python
+fig = plot_series(prep)
+```
+
+
+
+### Model Selection with Cross-Validation
+
+We can test many models simoultaneously using MLForecast
+`cross_validation`. We can import `lightgbm` and `scikit-learn` models
+and try different combinations of them, alongside different target
+transformations (as the ones we created previously) and historical
+variables.
+You can see an in-depth tutorial on how to use
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+[Cross Validation methods
+here](https://nixtla.github.io/mlforecast/docs/how-to-guides/cross_validation.html)
+
+```python
+import lightgbm as lgb
+from sklearn.base import BaseEstimator
+from sklearn.linear_model import Lasso, LinearRegression, Ridge
+from sklearn.neighbors import KNeighborsRegressor
+from sklearn.neural_network import MLPRegressor
+from sklearn.ensemble import RandomForestRegressor
+
+from mlforecast.lag_transforms import ExpandingMean, RollingMean
+from mlforecast.target_transforms import Differences
+```
+
+We can create a benchmark `Naive` model that uses the electricity load
+of the last hour as prediction `lag1` as showed in the next cell. You
+can create your own models and try them with
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+using the same structure.
+
+```python
+class Naive(BaseEstimator):
+ def fit(self, X, y):
+ return self
+
+ def predict(self, X):
+ return X['lag1']
+```
+
+Now let’s try differen models from the `scikit-learn` library: `Lasso`,
+`LinearRegression`, `Ridge`, `KNN`, `MLP` and `Random Forest` alongside
+the `LightGBM`. You can add any model to the dictionary to train and
+compare them by adding them to the dictionary (`models`) as shown.
+
+```python
+# Model dictionary
+models ={
+ 'naive': Naive(),
+ 'lgbm': lgb.LGBMRegressor(verbosity=-1),
+ 'lasso': Lasso(),
+ 'lin_reg': LinearRegression(),
+ 'ridge': Ridge(),
+ 'knn': KNeighborsRegressor(),
+ 'mlp': MLPRegressor(),
+ 'rf': RandomForestRegressor()
+ }
+```
+
+The we can instanciate the
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+class with the models we want to try along side `target_transforms`,
+`lags`, `lag_transforms`, and `date_features`. All this features are
+applied to the models we selected.
+
+In this case we use the 1st, 12th and 24th lag, which are passed as a
+list. Potentially you could pass a `range`.
+
+``` text
+lags=[1,12,24]
+```
+
+Lag transforms are defined as a dictionary where the keys are the lags
+and the values are lists of the transformations that we want to apply to
+that lag. You can refer to the [lag transformations
+guide](../how-to-guides/lag_transforms_guide.html) for more details.
+
+For using the date features you need to be sure that your time column is
+made of timestamps. Then it might make sense to extract features like
+week, dayofweek, quarter, etc. You can do that by passing a list of
+strings with [pandas time/date
+components](https://pandas.pydata.org/docs/user_guide/timeseries.html#time-date-components).
+You can also pass functions that will take the time column as input, as
+we’ll show here.
+Here we add month, hour and dayofweek features:
+
+``` text
+ date_features=['month', 'hour', 'dayofweek']
+```
+
+```python
+mlf = MLForecast(
+ models = models,
+ freq='H', # our series have hourly frequency
+ target_transforms=[Differences([24, 24*7])],
+ lags=[1,12,24], # Lags to be used as features
+ lag_transforms={
+ 1: [ExpandingMean()],
+ 24: [RollingMean(window_size=48)],
+ },
+ date_features=['month', 'hour', 'dayofweek']
+)
+```
+
+Now we use the `cross_validation` method to train and evalaute the
+models. + `df`: Receives the training data + `h`: Forecast horizon +
+`n_windows`: The number of folds we want to predict
+
+You can specify the names of the time series id, time and target
+columns. + `id_col`:Column that identifies each serie ( Default
+*unique_id* ) + `time_col`: Column that identifies each timestep, its
+values can be timestamps or integer( Default *ds* ) +
+`target_col`:Column that contains the target ( Default *y* )
+
+```python
+crossvalidation_df = mlf.cross_validation(
+ df=df_train,
+ h=24,
+ n_windows=4,
+ refit=False,
+)
+crossvalidation_df.head()
+```
+
+| | unique_id | ds | cutoff | y | naive | lgbm | lasso | lin_reg | ridge | knn | mlp | rf |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | PJM_Load_hourly | 2001-12-27 01:00:00 | 2001-12-27 | 28332.0 | 28837.0 | 28526.505572 | 28703.185712 | 28702.625949 | 28702.625956 | 28479.0 | 28660.021947 | 27995.17 |
+| 1 | PJM_Load_hourly | 2001-12-27 02:00:00 | 2001-12-27 | 27329.0 | 27969.0 | 27467.860847 | 27693.502318 | 27692.395954 | 27692.395969 | 27521.6 | 27584.635434 | 27112.50 |
+| 2 | PJM_Load_hourly | 2001-12-27 03:00:00 | 2001-12-27 | 26986.0 | 27435.0 | 26605.710615 | 26991.795124 | 26990.157567 | 26990.157589 | 26451.6 | 26809.412477 | 26529.72 |
+| 3 | PJM_Load_hourly | 2001-12-27 04:00:00 | 2001-12-27 | 27009.0 | 27401.0 | 26284.065138 | 26789.418399 | 26787.262262 | 26787.262291 | 26388.4 | 26523.416348 | 26490.83 |
+| 4 | PJM_Load_hourly | 2001-12-27 05:00:00 | 2001-12-27 | 27555.0 | 28169.0 | 26823.617078 | 27369.643789 | 27366.983075 | 27366.983111 | 26779.6 | 26986.355992 | 27180.69 |
+
+Now we can plot each model and window (fold) to see how it behaves
+
+```python
+def plot_cv(df, df_cv, uid, fname, last_n=24 * 14, models={}):
+ cutoffs = df_cv.query('unique_id == @uid')['cutoff'].unique()
+ fig, ax = plt.subplots(nrows=len(cutoffs), ncols=1, figsize=(14, 14), gridspec_kw=dict(hspace=0.8))
+ for cutoff, axi in zip(cutoffs, ax.flat):
+ max_date = df_cv.query('unique_id == @uid & cutoff == @cutoff')['ds'].max()
+ df[df['ds'] < max_date].query('unique_id == @uid').tail(last_n).set_index('ds').plot(ax=axi, title=uid, y='y')
+ for m in models.keys():
+ df_cv.query('unique_id == @uid & cutoff == @cutoff').set_index('ds').plot(ax=axi, title=uid, y=m)
+ fig.savefig(f'../../figs/{fname}', bbox_inches='tight')
+ plt.close()
+```
+
+
+```python
+plot_cv(df_train, crossvalidation_df, 'PJM_Load_hourly', 'load_forecasting__predictions.png', models=models)
+```
+
+
+
+Visually examining the forecasts can give us some idea of how the model
+is behaving, yet in order to asses the performace we need to evaluate
+them trough metrics. For that we use the
+[utilsforecast](https://nixtla.github.io/utilsforecast/) library that
+contains many useful metrics and an evaluate function.
+
+```python
+from utilsforecast.losses import mae, mape, rmse, smape
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+# Metrics to be used for evaluation
+metrics = [
+ mae,
+ rmse,
+ mape,
+ smape
+]
+```
+
+
+```python
+# Function to evaluate the crossvalidation
+def evaluate_crossvalidation(crossvalidation_df, metrics, models):
+ evaluations = []
+ for c in crossvalidation_df['cutoff'].unique():
+ df_cv = crossvalidation_df.query('cutoff == @c')
+ evaluation = evaluate(
+ df = df_cv,
+ metrics=metrics,
+ models=list(models.keys())
+ )
+ evaluations.append(evaluation)
+ evaluations = pd.concat(evaluations, ignore_index=True).drop(columns='unique_id')
+ evaluations = evaluations.groupby('metric').mean()
+ return evaluations.style.background_gradient(cmap='RdYlGn_r', axis=1)
+```
+
+
+```python
+evaluate_crossvalidation(crossvalidation_df, metrics, models)
+```
+
+| | naive | lgbm | lasso | lin_reg | ridge | knn | mlp | rf |
+|----|----|----|----|----|----|----|----|----|
+| metric | | | | | | | | |
+| mae | 1631.395833 | 971.536200 | 1003.796433 | 1007.998597 | 1007.998547 | 1248.145833 | 1870.547722 | 1017.957813 |
+| mape | 0.049759 | 0.030966 | 0.031760 | 0.031888 | 0.031888 | 0.038721 | 0.057504 | 0.032341 |
+| rmse | 1871.398919 | 1129.713256 | 1148.616156 | 1153.262719 | 1153.262664 | 1451.964390 | 2102.098238 | 1154.647164 |
+| smape | 0.024786 | 0.015886 | 0.016269 | 0.016338 | 0.016338 | 0.019549 | 0.029917 | 0.016563 |
+
+We can se that the model `lgbm` has top performance in most metrics
+folowed by the `lasso regression`. Both models perform way better than
+the `naive`.
+
+### Test Evaluation
+
+Now we are going to evaluate their perfonce in the test set. We can use
+both of them for forecasting the test alongside some prediction
+intervals. For that we can use the
+[\[`PredictionIntervals`\](https://Nixtla.github.io/mlforecast/utils.html#predictionintervals)](https://nixtla.github.io/mlforecast/utils.html#predictionintervals)
+function in `mlforecast.utils`.
+You can see an in-depth tutotorial of [Probabilistic Forecasting
+here](https://nixtlaverse.nixtla.io/mlforecast/docs/tutorials/prediction_intervals_in_forecasting_models.html)
+
+```python
+from mlforecast.utils import PredictionIntervals
+```
+
+
+```python
+models_evaluation ={
+ 'lgbm': lgb.LGBMRegressor(verbosity=-1),
+ 'lasso': Lasso(),
+ }
+
+mlf_evaluation = MLForecast(
+ models = models_evaluation,
+ freq='H', # our series have hourly frequency
+ target_transforms=[Differences([24, 24*7])],
+ lags=[1,12,24],
+ lag_transforms={
+ 1: [ExpandingMean()],
+ 24: [RollingMean(window_size=48)],
+ },
+ date_features=['month', 'hour', 'dayofweek']
+)
+```
+
+Now we’re ready to generate the point forecasts and the prediction
+intervals. To do this, we’ll use the `fit` method, which takes the
+following arguments:
+
+- `df`: Series data in long format.
+- `id_col`: Column that identifies each series. In our case,
+ unique_id.
+- `time_col`: Column that identifies each timestep, its values can be
+ timestamps or integers. In our case, ds.
+- `target_col`: Column that contains the target. In our case, y.
+
+The
+[`PredictionIntervals`](https://Nixtla.github.io/mlforecast/utils.html#predictionintervals)
+function is used to compute prediction intervals for the models using
+[Conformal
+Prediction](https://valeman.medium.com/how-to-predict-full-probability-distribution-using-machine-learning-conformal-predictive-f8f4d805e420).
+The function takes the following arguments: + `n_windows`: represents
+the number of cross-validation windows used to calibrate the intervals +
+`h`: the forecast horizon
+
+```python
+mlf_evaluation.fit(
+ df = df_train,
+ prediction_intervals=PredictionIntervals(n_windows=4, h=24)
+)
+```
+
+``` text
+MLForecast(models=[lgbm, lasso], freq=H, lag_features=['lag1', 'lag12', 'lag24', 'expanding_mean_lag1', 'rolling_mean_lag24_window_size48'], date_features=['month', 'hour', 'dayofweek'], num_threads=1)
+```
+
+Now that the model has been trained we are going to forecast the next 24
+hours using the `predict` method so we can compare them to our `test`
+data. Additionally, we are going to create prediction intervals at
+`levels` `[90,95]`.
+
+```python
+levels = [90, 95] # Levels for prediction intervals
+forecasts = mlf_evaluation.predict(24, level=levels)
+forecasts.head()
+```
+
+| | unique_id | ds | lgbm | lasso | lgbm-lo-95 | lgbm-lo-90 | lgbm-hi-90 | lgbm-hi-95 | lasso-lo-95 | lasso-lo-90 | lasso-hi-90 | lasso-hi-95 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | PJM_Load_hourly | 2001-12-31 01:00:00 | 28847.573176 | 29124.085976 | 28544.593464 | 28567.603130 | 29127.543222 | 29150.552888 | 28762.752269 | 28772.604275 | 29475.567677 | 29485.419682 |
+| 1 | PJM_Load_hourly | 2001-12-31 02:00:00 | 27862.589195 | 28365.330749 | 27042.311414 | 27128.839888 | 28596.338503 | 28682.866977 | 27528.548959 | 27619.065224 | 29111.596275 | 29202.112539 |
+| 2 | PJM_Load_hourly | 2001-12-31 03:00:00 | 27044.418960 | 27712.161676 | 25596.659896 | 25688.230426 | 28400.607493 | 28492.178023 | 26236.955369 | 26338.087102 | 29086.236251 | 29187.367984 |
+| 3 | PJM_Load_hourly | 2001-12-31 04:00:00 | 26976.104125 | 27661.572733 | 25249.961527 | 25286.024722 | 28666.183529 | 28702.246724 | 25911.133521 | 25959.815715 | 29363.329750 | 29412.011944 |
+| 4 | PJM_Load_hourly | 2001-12-31 05:00:00 | 26694.246238 | 27393.922370 | 25044.220845 | 25051.548832 | 28336.943644 | 28344.271631 | 25751.547897 | 25762.524815 | 29025.319924 | 29036.296843 |
+
+The `predict` method returns a DataFrame witht the predictions for each
+model (`lasso` and `lgbm`) along side the prediction tresholds. The
+high-threshold is indicated by the keyword `hi`, the low-threshold by
+the keyword `lo`, and the level by the number in the column names.
+
+```python
+test = df_last_24_hours.merge(forecasts, how='left', on=['unique_id', 'ds'])
+test.head()
+```
+
+| | unique_id | ds | y | lgbm | lasso | lgbm-lo-95 | lgbm-lo-90 | lgbm-hi-90 | lgbm-hi-95 | lasso-lo-95 | lasso-lo-90 | lasso-hi-90 | lasso-hi-95 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | PJM_Load_hourly | 2001-12-31 01:00:00 | 29001.0 | 28847.573176 | 29124.085976 | 28544.593464 | 28567.603130 | 29127.543222 | 29150.552888 | 28762.752269 | 28772.604275 | 29475.567677 | 29485.419682 |
+| 1 | PJM_Load_hourly | 2001-12-31 02:00:00 | 28138.0 | 27862.589195 | 28365.330749 | 27042.311414 | 27128.839888 | 28596.338503 | 28682.866977 | 27528.548959 | 27619.065224 | 29111.596275 | 29202.112539 |
+| 2 | PJM_Load_hourly | 2001-12-31 03:00:00 | 27830.0 | 27044.418960 | 27712.161676 | 25596.659896 | 25688.230426 | 28400.607493 | 28492.178023 | 26236.955369 | 26338.087102 | 29086.236251 | 29187.367984 |
+| 3 | PJM_Load_hourly | 2001-12-31 04:00:00 | 27874.0 | 26976.104125 | 27661.572733 | 25249.961527 | 25286.024722 | 28666.183529 | 28702.246724 | 25911.133521 | 25959.815715 | 29363.329750 | 29412.011944 |
+| 4 | PJM_Load_hourly | 2001-12-31 05:00:00 | 28427.0 | 26694.246238 | 27393.922370 | 25044.220845 | 25051.548832 | 28336.943644 | 28344.271631 | 25751.547897 | 25762.524815 | 29025.319924 | 29036.296843 |
+
+Now we can evaluate the metrics and performance in the `test` set.
+
+```python
+evaluate(
+ df = test,
+ metrics=metrics,
+ models=list(models_evaluation.keys())
+)
+```
+
+| | unique_id | metric | lgbm | lasso |
+|-----|-----------------|--------|-------------|-------------|
+| 0 | PJM_Load_hourly | mae | 1092.050817 | 899.979743 |
+| 1 | PJM_Load_hourly | rmse | 1340.422762 | 1163.695525 |
+| 2 | PJM_Load_hourly | mape | 0.033600 | 0.027688 |
+| 3 | PJM_Load_hourly | smape | 0.017137 | 0.013812 |
+
+We can see that the `lasso` regression performed slighty better than the
+`LightGBM` for the test set. Additonally, we can also plot the forecasts
+alongside their prediction intervals. For that we can use the
+`plot_series` method available in `utilsforecast.plotting`.
+
+We can plot one or many models at once alongside their coinfidence
+intervals.
+
+```python
+fig = plot_series(
+ df_train,
+ test,
+ models=['lasso', 'lgbm'],
+ plot_random=False,
+ level=levels,
+ max_insample_length=24
+)
+```
+
+
+
+### Comparison with Prophet
+
+One of the most widely used models for time series forecasting is
+`Prophet`. This model is known for its ability to model different
+seasonalities (weekly, daily yearly). We will use this model as a
+benchmark to see if the `lgbm` alongside
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+adds value for this time series.
+
+```python
+from prophet import Prophet
+from time import time
+```
+
+``` text
+Importing plotly failed. Interactive plots will not work.
+```
+
+```python
+# create prophet model
+prophet = Prophet(interval_width=0.9)
+init = time()
+prophet.fit(df_train)
+# produce forecasts
+future = prophet.make_future_dataframe(periods=len(df_last_24_hours), freq='H', include_history=False)
+forecast_prophet = prophet.predict(future)
+end = time()
+# data wrangling
+forecast_prophet = forecast_prophet[['ds', 'yhat', 'yhat_lower', 'yhat_upper']]
+forecast_prophet.columns = ['ds', 'Prophet', 'Prophet-lo-90', 'Prophet-hi-90']
+forecast_prophet.insert(0, 'unique_id', 'PJM_Load_hourly')
+forecast_prophet.head()
+```
+
+| | unique_id | ds | Prophet | Prophet-lo-90 | Prophet-hi-90 |
+|----|----|----|----|----|----|
+| 0 | PJM_Load_hourly | 2001-12-31 01:00:00 | 25333.448442 | 20589.873559 | 30370.174820 |
+| 1 | PJM_Load_hourly | 2001-12-31 02:00:00 | 24039.925936 | 18927.503487 | 29234.930903 |
+| 2 | PJM_Load_hourly | 2001-12-31 03:00:00 | 23363.998793 | 18428.462513 | 28292.424622 |
+| 3 | PJM_Load_hourly | 2001-12-31 04:00:00 | 23371.799609 | 18206.273446 | 28181.023448 |
+| 4 | PJM_Load_hourly | 2001-12-31 05:00:00 | 24146.468610 | 19356.171497 | 29006.546759 |
+
+```python
+time_prophet = (end - init)
+print(f'Prophet Time: {time_prophet:.2f} seconds')
+```
+
+``` text
+Prophet Time: 18.00 seconds
+```
+
+```python
+models_comparison ={
+ 'lgbm': lgb.LGBMRegressor(verbosity=-1)
+}
+
+mlf_comparison = MLForecast(
+ models = models_comparison,
+ freq='H', # our series have hourly frequency
+ target_transforms=[Differences([24, 24*7])],
+ lags=[1,12,24],
+ lag_transforms={
+ 1: [ExpandingMean()],
+ 24: [RollingMean(window_size=48)],
+ },
+ date_features=['month', 'hour', 'dayofweek']
+)
+
+init = time()
+mlf_comparison.fit(
+ df = df_train,
+ prediction_intervals=PredictionIntervals(n_windows=4, h=24)
+)
+
+levels = [90]
+forecasts_comparison = mlf_comparison.predict(24, level=levels)
+end = time()
+forecasts_comparison.head()
+```
+
+| | unique_id | ds | lgbm | lgbm-lo-90 | lgbm-hi-90 |
+|----|----|----|----|----|----|
+| 0 | PJM_Load_hourly | 2001-12-31 01:00:00 | 28847.573176 | 28567.603130 | 29127.543222 |
+| 1 | PJM_Load_hourly | 2001-12-31 02:00:00 | 27862.589195 | 27128.839888 | 28596.338503 |
+| 2 | PJM_Load_hourly | 2001-12-31 03:00:00 | 27044.418960 | 25688.230426 | 28400.607493 |
+| 3 | PJM_Load_hourly | 2001-12-31 04:00:00 | 26976.104125 | 25286.024722 | 28666.183529 |
+| 4 | PJM_Load_hourly | 2001-12-31 05:00:00 | 26694.246238 | 25051.548832 | 28336.943644 |
+
+```python
+time_lgbm = (end - init)
+print(f'LGBM Time: {time_lgbm:.2f} seconds')
+```
+
+``` text
+LGBM Time: 0.86 seconds
+```
+
+```python
+metrics_comparison = df_last_24_hours.merge(forecasts_comparison, how='left', on=['unique_id', 'ds']).merge(
+ forecast_prophet, how='left', on=['unique_id', 'ds'])
+metrics_comparison = evaluate(
+ df = metrics_comparison,
+ metrics=metrics,
+ models=['Prophet', 'lgbm']
+ )
+metrics_comparison.reset_index(drop=True).style.background_gradient(cmap='RdYlGn_r', axis=1)
+```
+
+| | unique_id | metric | Prophet | lgbm |
+|-----|-----------------|--------|-------------|-------------|
+| 0 | PJM_Load_hourly | mae | 2266.561642 | 1092.050817 |
+| 1 | PJM_Load_hourly | rmse | 2701.302779 | 1340.422762 |
+| 2 | PJM_Load_hourly | mape | 0.073226 | 0.033600 |
+| 3 | PJM_Load_hourly | smape | 0.038320 | 0.017137 |
+
+As we can see `lgbm` had consistently better metrics than `prophet`.
+
+```python
+metrics_comparison['improvement'] = metrics_comparison['Prophet'] / metrics_comparison['lgbm']
+metrics_comparison['improvement'] = metrics_comparison['improvement'].apply(lambda x: f'{x:.2f}')
+metrics_comparison.set_index('metric')[['improvement']]
+```
+
+| | improvement |
+|--------|-------------|
+| metric | |
+| mae | 2.08 |
+| rmse | 2.02 |
+| mape | 2.18 |
+| smape | 2.24 |
+
+```python
+print(f'lgbm with MLForecast has a speedup of {time_prophet/time_lgbm:.2f} compared with prophet')
+```
+
+``` text
+lgbm with MLForecast has a speedup of 20.95 compared with prophet
+```
+
+We can see that `lgbm` with
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+was able to provide metrics at least twice as good as `Prophet` as seen
+in the column `improvement` above, and way faster.
+
diff --git a/mlforecast/docs/tutorials/electricity_peak_forecasting.html.mdx b/mlforecast/docs/tutorials/electricity_peak_forecasting.html.mdx
new file mode 100644
index 00000000..86213949
--- /dev/null
+++ b/mlforecast/docs/tutorials/electricity_peak_forecasting.html.mdx
@@ -0,0 +1,274 @@
+---
+description: >-
+ In this example we will show how to perform electricity load forecasting on
+ the ERCOT (Texas) market for detecting daily peaks.
+output-file: electricity_peak_forecasting.html
+title: Detect Demand Peaks
+---
+
+
+## Introduction
+
+Predicting peaks in different markets is useful. In the electricity
+market, consuming electricity at peak demand is penalized with higher
+tarifs. When an individual or company consumes electricity when its most
+demanded, regulators calls that a coincident peak (CP).
+
+In the Texas electricity market (ERCOT), the peak is the monthly
+15-minute interval when the ERCOT Grid is at a point of highest
+capacity. The peak is caused by all consumers’ combined demand on the
+electrical grid. The coincident peak demand is an important factor used
+by ERCOT to determine final electricity consumption bills. ERCOT
+registers the CP demand of each client for 4 months, between June and
+September, and uses this to adjust electricity prices. Clients can
+therefore save on electricity bills by reducing the coincident peak
+demand.
+
+In this example we will train a `LightGBM` model on historic load data
+to forecast day-ahead peaks on September 2022. Multiple seasonality is
+traditionally present in low sampled electricity data. Demand exhibits
+daily and weekly seasonality, with clear patterns for specific hours of
+the day such as 6:00pm vs 3:00am or for specific days such as Sunday vs
+Friday.
+
+First, we will load ERCOT historic demand, then we will use the
+[`MLForecast.cross_validation`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.cross_validation)
+method to fit the `LightGBM` model and forecast daily load during
+September. Finally, we show how to use the forecasts to detect the
+coincident peak.
+
+**Outline**
+
+1. Install libraries
+2. Load and explore the data
+3. Fit LightGBM model and forecast
+4. Peak detection
+
+> **Tip**
+>
+> You can use Colab to run this Notebook interactively
+>
+
+## Libraries
+
+We assume you have MLForecast already installed. Check this guide for
+instructions on [how to install
+MLForecast](../getting-started/install.html).
+
+Install the necessary packages using `pip install mlforecast`.
+
+Also we have to install `LightGBM` using `pip install lightgbm`.
+
+## Load Data
+
+The input to MLForecast is always a data frame in [long
+format](https://www.theanalysisfactor.com/wide-and-long-data/) with
+three columns: `unique_id`, `ds` and `y`:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp or int) column should be either an integer
+ indexing time or a datestamp ideally like YYYY-MM-DD for a date or
+ YYYY-MM-DD HH:MM:SS for a timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast. We
+ will rename the
+
+First, read the 2022 historic total demand of the ERCOT market. We
+processed the original data (available
+[here](https://www.ercot.com/gridinfo/load/load_hist)), by adding the
+missing hour due to daylight saving time, parsing the date to datetime
+format, and filtering columns of interest.
+
+```python
+import numpy as np
+import pandas as pd
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+# Load data
+Y_df = pd.read_csv('https://datasets-nixtla.s3.amazonaws.com/ERCOT-clean.csv', parse_dates=['ds'])
+Y_df = Y_df.query("ds >= '2022-01-01' & ds <= '2022-10-01'")
+```
+
+
+```python
+fig = plot_series(Y_df)
+```
+
+
+
+We observe that the time series exhibits seasonal patterns. Moreover,
+the time series contains `6,552` observations, so it is necessary to use
+computationally efficient methods to deploy them in production.
+
+## Fit and Forecast LightGBM model
+
+Import the
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+class and the models you need.
+
+```python
+import lightgbm as lgb
+
+from mlforecast import MLForecast
+from mlforecast.target_transforms import Differences
+```
+
+First, instantiate the model and define the parameters.
+
+> **Tip**
+>
+> In this example we are using the default parameters of the
+> `lgb.LGBMRegressor` model, but you can change them to improve the
+> forecasting performance.
+
+```python
+models = [
+ lgb.LGBMRegressor(verbosity=-1) # you can include more models here
+]
+```
+
+We fit the model by instantiating a
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+object with the following required parameters:
+
+- `models`: a list of sklearn-like (fit and predict) models.
+
+- `freq`: a string indicating the frequency of the data. (See [panda’s
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `target_transforms`: Transformations to apply to the target before
+ computing the features. These are restored at the forecasting step.
+
+- `lags`: Lags of the target to use as features.
+
+```python
+# Instantiate MLForecast class as mlf
+mlf = MLForecast(
+ models=models,
+ freq='H',
+ target_transforms=[Differences([24])],
+ lags=range(1, 25)
+)
+```
+
+> **Tip**
+>
+> In this example, we are only using differences and lags to produce
+> features. See the [full
+> documentation](https://nixtla.github.io/mlforecast/forecast.html#mlforecast)
+> to see all available features.
+
+The `cross_validation` method allows the user to simulate multiple
+historic forecasts, greatly simplifying pipelines by replacing for loops
+with `fit` and `predict` methods. This method re-trains the model and
+forecast each window. See [this
+tutorial]https://nixtlaverse.nixtla.io/statsforecast/docs/getting-started/getting_started_complete.html#evaluate-the-model’s-performance)
+for an animation of how the windows are defined.
+
+Use the `cross_validation` method to produce all the daily forecasts for
+September. To produce daily forecasts set the forecasting horizon
+`window_size` as 24. In this example we are simulating deploying the
+pipeline during September, so set the number of windows as 30 (one for
+each day). Finally, the step size between windows is 24 (equal to the
+`window_size`). This ensure to only produce one forecast per day.
+
+Additionally,
+
+- `id_col`: identifies each time series.
+- `time_col`: indetifies the temporal column of the time series.
+- `target_col`: identifies the column to model.
+
+```python
+crossvalidation_df = mlf.cross_validation(
+ df=Y_df,
+ h=24,
+ n_windows=30,
+)
+```
+
+
+```python
+crossvalidation_df.head()
+```
+
+| | unique_id | ds | cutoff | y | LGBMRegressor |
+|----|----|----|----|----|----|
+| 0 | ERCOT | 2022-09-01 00:00:00 | 2022-08-31 23:00:00 | 45482.471757 | 45685.265537 |
+| 1 | ERCOT | 2022-09-01 01:00:00 | 2022-08-31 23:00:00 | 43602.658043 | 43779.819515 |
+| 2 | ERCOT | 2022-09-01 02:00:00 | 2022-08-31 23:00:00 | 42284.817342 | 42672.470923 |
+| 3 | ERCOT | 2022-09-01 03:00:00 | 2022-08-31 23:00:00 | 41663.156771 | 42091.768192 |
+| 4 | ERCOT | 2022-09-01 04:00:00 | 2022-08-31 23:00:00 | 41710.621904 | 42481.403168 |
+
+> **Important**
+>
+> When using `cross_validation` make sure the forecasts are produced at
+> the desired timestamps. Check the `cutoff` column which specifices the
+> last timestamp before the forecasting window.
+
+## Peak Detection
+
+Finally, we use the forecasts in `crossvaldation_df` to detect the daily
+hourly demand peaks. For each day, we set the detected peaks as the
+highest forecasts. In this case, we want to predict one peak (`npeaks`);
+depending on your setting and goals, this parameter might change. For
+example, the number of peaks can correspond to how many hours a battery
+can be discharged to reduce demand.
+
+```python
+npeaks = 1 # Number of peaks
+```
+
+For the ERCOT 4CP detection task we are interested in correctly
+predicting the highest monthly load. Next, we filter the day in
+September with the highest hourly demand and predict the peak.
+
+```python
+crossvalidation_df = crossvalidation_df.reset_index()[['ds','y','LGBMRegressor']]
+max_day = crossvalidation_df.iloc[crossvalidation_df['y'].argmax()].ds.day # Day with maximum load
+cv_df_day = crossvalidation_df.query('ds.dt.day == @max_day')
+max_hour = cv_df_day['y'].argmax()
+peaks = cv_df_day['LGBMRegressor'].argsort().iloc[-npeaks:].values # Predicted peaks
+```
+
+In the following plot we see how the LightGBM model is able to correctly
+detect the coincident peak for September 2022.
+
+```python
+import matplotlib.pyplot as plt
+```
+
+
+```python
+fig, ax = plt.subplots(figsize=(10, 5))
+ax.axvline(cv_df_day.iloc[max_hour]['ds'], color='black', label='True Peak')
+ax.scatter(cv_df_day.iloc[peaks]['ds'], cv_df_day.iloc[peaks]['LGBMRegressor'], color='green', label=f'Predicted Top-{npeaks}')
+ax.plot(cv_df_day['ds'], cv_df_day['y'], label='y', color='blue')
+ax.plot(cv_df_day['ds'], cv_df_day['LGBMRegressor'], label='Forecast', color='red')
+ax.set(xlabel='Time', ylabel='Load (MW)')
+ax.grid()
+ax.legend()
+fig.savefig('../../figs/electricity_peak_forecasting__predicted_peak.png', bbox_inches='tight')
+plt.close()
+```
+
+
+
+> **Important**
+>
+> In this example we only include September. However, MLForecast and
+> LightGBM can correctly predict the peaks for the 4 months of 2022. You
+> can try this by increasing the `n_windows` parameter of
+> `cross_validation` or filtering the `Y_df` dataset.
+
+## Next steps
+
+MLForecast and LightGBM in particular are good benchmarking models for
+peak detection. However, it might be useful to explore further and newer
+forecasting algorithms or perform hyperparameter optimization.
+
diff --git a/mlforecast/docs/tutorials/prediction_intervals_in_forecasting_models.html.mdx b/mlforecast/docs/tutorials/prediction_intervals_in_forecasting_models.html.mdx
new file mode 100644
index 00000000..50b6b1ad
--- /dev/null
+++ b/mlforecast/docs/tutorials/prediction_intervals_in_forecasting_models.html.mdx
@@ -0,0 +1,1015 @@
+---
+output-file: prediction_intervals_in_forecasting_models.html
+title: Prediction intervals
+---
+
+
+The objective of the following article is to obtain a step-by-step guide
+on building `Prediction intervals in forecasting models` using
+`mlforecast`.
+
+During this walkthrough, we will become familiar with the main
+`MlForecast` class and some relevant methods such as
+[`MLForecast.fit`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.fit),
+[`MLForecast.predict`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.predict)
+and
+[`MLForecast.cross_validation`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.cross_validation)
+in other.
+
+Let’s start!!!
+
+# Table of contents
+
+1. [Introduction](#introduction)
+2. [Forecasts and prediction
+ intervals](#forecasts-and-prediction-intervals)
+3. [Installing mlforecast](#installing-mlforecast)
+4. [Loading libraries and data](#loading-libraries-and-data)
+5. [Explore Data with the plot
+ method](#explore-data-with-the-plot-method)
+6. [Split the data into training and
+ testing](#split-the-data-into-training-and-testing)
+7. [Modeling with mlforecast](#modeling-with-mlforecast)
+8. [References](#references)
+
+# Introduction
+
+The target of our prediction is something unknown (otherwise we wouldn’t
+be making a prediction), so we can think of it as a random variable. For
+example, the total sales for the next month could have different
+possible values, and we won’t know what the exact value will be until we
+get the actual sales at the end of the month. Until next month’s sales
+are known, this is a random amount.
+
+By the time the next month draws near, we usually have a pretty good
+idea of possible sales values. However, if we are forecasting sales for
+the same month next year, the possible values can vary much more. In
+most forecasting cases, the variability associated with what we are
+forecasting reduces as we get closer to the event. In other words, the
+further back in time we make the prediction, the more uncertainty there
+is.
+
+We can imagine many possible future scenarios, each yielding a different
+value for what we are trying to forecast.
+
+When we obtain a forecast, we are estimating the middle of the range of
+possible values the random variable could take. Often, a forecast is
+accompanied by a prediction interval giving a range of values the random
+variable could take with relatively high probability. For example, a 95%
+prediction interval contains a range of values which should include the
+actual future value with probability 95%.
+
+Rather than plotting individual possible futures , we usually show these
+prediction intervals instead.
+
+When we generate a forecast, we usually produce a single value known as
+the point forecast. This value, however, doesn’t tell us anything about
+the uncertainty associated with the forecast. To have a measure of this
+uncertainty, we need prediction intervals.
+
+A prediction interval is a range of values that the forecast can take
+with a given probability. Hence, a 95% prediction interval should
+contain a range of values that include the actual future value with
+probability 95%. Probabilistic forecasting aims to generate the full
+forecast distribution. Point forecasting, on the other hand, usually
+returns the mean or the median or said distribution. However, in
+real-world scenarios, it is better to forecast not only the most
+probable future outcome, but many alternative outcomes as well.
+
+The problem is that some timeseries models provide forecast
+distributions, but some other ones only provide point forecasts. How can
+we then estimate the uncertainty of predictions?
+
+# Forecasts and prediction intervals
+
+There are at least four sources of uncertainty in forecasting using time
+series models:
+
+1. The random error term;
+2. The parameter estimates;
+3. The choice of model for the historical data;
+4. The continuation of the historical data generating process into the
+ future.
+
+When we produce prediction intervals for time series models, we
+generally only take into account the first of these sources of
+uncertainty. It would be possible to account for 2 and 3 using
+simulations, but that is almost never done because it would take too
+much time to compute. As computing speeds increase, it might become a
+viable approach in the future.
+
+Even if we ignore the model uncertainty and the DGP uncertainty (sources
+3 and 4), and just try to allow for parameter uncertainty as well as the
+random error term (sources 1 and 2), there are no closed form solutions
+apart from some simple special cases. see full article [Rob J
+Hyndman](https://robjhyndman.com/hyndsight/narrow-pi/)
+
+## Forecast distributions
+
+We use forecast distributions to express the uncertainty in our
+predictions. These probability distributions describe the probability of
+observing different future values using the fitted model. The point
+forecast corresponds to the mean of this distribution. Most time series
+models generate forecasts that follow a normal distribution, which
+implies that we assume that possible future values follow a normal
+distribution. However, later in this section we will look at some
+alternatives to normal distributions.
+
+### Importance of Confidence Interval Prediction in Time Series:
+
+1. Uncertainty Estimation: The confidence interval provides a measure
+ of the uncertainty associated with time series predictions. It
+ enables variability and the range of possible future values to be
+ quantified, which is essential for making informed decisions.
+
+2. Precision evaluation: By having a confidence interval, the precision
+ of the predictions can be evaluated. If the interval is narrow, it
+ indicates that the forecast is more accurate and reliable. On the
+ other hand, if the interval is wide, it indicates greater
+ uncertainty and less precision in the predictions.
+
+3. Risk management: The confidence interval helps in risk management by
+ providing information about possible future scenarios. It allows
+ identifying the ranges in which the real values could be located and
+ making decisions based on those possible scenarios.
+
+4. Effective communication: The confidence interval is a useful tool
+ for communicating predictions clearly and accurately. It allows the
+ variability and uncertainty associated with the predictions to be
+ conveyed to the stakeholders, avoiding a wrong or overly optimistic
+ interpretation of the results.
+
+Therefore, confidence interval prediction in time series is essential to
+understand and manage uncertainty, assess the accuracy of predictions,
+and make informed decisions based on possible future scenarios.
+
+## Prediction intervals
+
+A prediction interval gives us a range in which we expect $y_t$ to lie
+with a specified probability. For example, if we assume that the
+distribution of future observations follows a normal distribution, a 95%
+prediction interval for the forecast of step h would be represented by
+the range
+
+$$\hat{y}_{T+h|T} \pm 1.96 \hat\sigma_h,$$
+
+Where $\hat\sigma_h$ is an estimate of the standard deviation of the h
+-step forecast distribution.
+
+More generally, a prediction interval can be written as
+
+$$\hat{y}_{T+h|T} \pm c \hat\sigma_h$$
+
+In this context, the term “multiplier c” is associated with the
+probability of coverage. In this article, intervals of 80% and 95% are
+typically calculated, but any other percentage can be used. The table
+below shows the values of c corresponding to different coverage
+probabilities, assuming a normal forecast distribution.
+
+| Percentage | Multiplier |
+|------------|------------|
+| 50 | 0.67 |
+| 55 | 0.76 |
+| 60 | 0.84 |
+| 65 | 0.93 |
+| 70 | 1.04 |
+| 75 | 1.15 |
+| 80 | 1.28 |
+| 85 | 1.44 |
+| 90 | 1.64 |
+| 95 | 1.96 |
+| 96 | 2.05 |
+| 97 | 2.17 |
+| 98 | 2.33 |
+| 99 | 2.58 |
+
+Prediction intervals are valuable because they reflect the uncertainty
+in the predictions. If we only generate point forecasts, we cannot
+assess how accurate those forecasts are. However, by providing
+prediction intervals, the amount of uncertainty associated with each
+forecast becomes apparent. For this reason, point forecasts may lack
+significant value without the inclusion of corresponding forecast
+intervals.
+
+## One-step prediction intervals
+
+When making a prediction for a future step, it is possible to estimate
+the standard deviation of the forecast distribution using the standard
+deviation of the residuals, which is calculated by
+
+where $K$ is the number of parameters estimated in the forecasting
+method, and $M$ is the number of missing values in the residuals. (For
+example, $M=1$ for a naive forecast, because we can’t forecast the first
+observation.)
+
+## Multi-step prediction intervals
+
+A typical feature of forecast intervals is that they tend to increase in
+length as the forecast horizon lengthens. As we move further out in
+time, there is greater uncertainty associated with the prediction,
+resulting in wider prediction intervals. In general, σh tends to
+increase as h increases (although there are some nonlinear forecasting
+methods that do not follow this property).
+
+To generate a prediction interval, it is necessary to have an estimate
+of σh. As mentioned above, for one-step forecasts (h=1), equation (1)
+provides a good estimate of the standard deviation of the forecast, σ1.
+However, for multi-step forecasts, a more complex calculation method is
+required. These calculations assume that the residuals are uncorrelated
+with each other.
+
+## Benchmark methods
+
+For the four benchmark methods, it is possible to mathematically derive
+the forecast standard deviation under the assumption of uncorrelated
+residuals. If $\hat{\sigma}_h$ denotes the standard deviation of the $h$
+-step forecast distribution, and $\hat{\sigma}$ is the residual standard
+deviation given by (1), then we can use the expressions shown in next
+Table. Note that when $h=1$ and $T$ is large, these all give the same
+approximate value $\hat{\sigma}$.
+
+| Method | h-step forecast standard deviation |
+|--------------------------|--------------------------------------------|
+| Mean forecasts | $\hat\sigma_h = \hat\sigma\sqrt{1 + 1/T}$ |
+| Naïve forecasts | $\hat\sigma_h = \hat\sigma\sqrt{h}$ |
+| Seasonal naïve forecasts | $\hat\sigma_h = \hat\sigma\sqrt{k+1}$ |
+| Drift forecasts | $\hat\sigma_h = \hat\sigma\sqrt{h(1+h/T)}$ |
+
+Note that when $h=1$ and $T$ is large, these all give the same
+approximate value $\hat{\sigma}$.
+
+## Prediction intervals from bootstrapped residuals
+
+When a normal distribution for the residuals is an unreasonable
+assumption, one alternative is to use bootstrapping, which only assumes
+that the residuals are uncorrelated with constant variance. We will
+illustrate the procedure using a naïve forecasting method.
+
+A one-step forecast error is defined as $e_t = y_t - \hat{y}_{t|t-1}$.
+For a naïve forecasting method, $\hat{y}_{t|t-1} = y_{t-1}$, so we can
+rewrite this as $$y_t = y_{t-1} + e_t.$$
+
+Assuming future errors will be similar to past errors, when $t>T$ we can
+replace $e_{t}$ by sampling from the collection of errors we have seen
+in the past (i.e., the residuals). So we can simulate the next
+observation of a time series using
+
+$$y^*_{T+1} = y_{T} + e^*_{T+1}$$
+
+where $e^*_{T+1}$ is a randomly sampled error from the past, and
+$y^*_{T+1}$ is the possible future value that would arise if that
+particular error value occurred. We use We use a \* to indicate that
+this is not the observed $y_{T+1}$ value, but one possible future that
+could occur. Adding the new simulated observation to our data set, we
+can repeat the process to obtain
+
+$$y^*_{T+2} = y_{T+1}^* + e^*_{T+2},$$
+
+where $e^*_{T+2}$ is another draw from the collection of residuals.
+Continuing in this way, we can simulate an entire set of future values
+for our time series.
+
+## Conformal Prediction
+
+Multi-quantile losses and statistical models can provide provide
+prediction intervals, but the problem is that these are uncalibrated,
+meaning that the actual frequency of observations falling within the
+interval does not align with the confidence level associated with it.
+For example, a calibrated 95% prediction interval should contain the
+true value 95% of the time in repeated sampling. An uncalibrated 95%
+prediction interval, on the other hand, might contain the true value
+only 80% of the time, or perhaps 99% of the time. In the first case, the
+interval is too narrow and underestimates the uncertainty, while in the
+second case, it is too wide and overestimates the uncertainty.
+
+Statistical methods also assume normality. Here, we talk about another
+method called conformal prediction that doesn’t require any
+distributional assumptions.
+
+Conformal prediction intervals use cross-validation on a point
+forecaster model to generate the intervals. This means that no prior
+probabilities are needed, and the output is well-calibrated. No
+additional training is needed, and the model is treated as a black box.
+The approach is compatible with any model
+
+[mlforecast](https://github.com/nixtla/mlforecast) now supports
+Conformal Prediction on all available models.
+
+# Installing mlforecast
+
+- using pip:
+
+ - `pip install mlforecast`
+
+- using with conda:
+
+ - `conda install -c conda-forge mlforecast`
+
+# Loading libraries and data
+
+```python
+# Handling and processing of Data
+# ==============================================================================
+import numpy as np
+import pandas as pd
+
+import scipy.stats as stats
+
+# Handling and processing of Data for Date (time)
+# ==============================================================================
+import datetime
+import time
+from datetime import datetime, timedelta
+
+#
+# ==============================================================================
+from statsmodels.tsa.stattools import adfuller
+import statsmodels.api as sm
+import statsmodels.tsa.api as smt
+from statsmodels.tsa.seasonal import seasonal_decompose
+#
+# ==============================================================================
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+import xgboost as xgb
+
+from mlforecast import MLForecast
+from mlforecast.lag_transforms import ExpandingMean, ExponentiallyWeightedMean, RollingMean
+from mlforecast.target_transforms import Differences
+from mlforecast.utils import PredictionIntervals
+```
+
+
+```python
+# Plot
+# ==============================================================================
+import matplotlib.pyplot as plt
+import matplotlib.ticker as ticker
+from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
+```
+
+## Read Data
+
+```python
+data_url = "https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/nyc_taxi.csv"
+df = pd.read_csv(data_url, parse_dates=["timestamp"])
+df.head()
+```
+
+| | timestamp | value |
+|-----|---------------------|-------|
+| 0 | 2014-07-01 00:00:00 | 10844 |
+| 1 | 2014-07-01 00:30:00 | 8127 |
+| 2 | 2014-07-01 01:00:00 | 6210 |
+| 3 | 2014-07-01 01:30:00 | 4656 |
+| 4 | 2014-07-01 02:00:00 | 3820 |
+
+The input to MlForecast is always a data frame in long format with three
+columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"] = "1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|---------------------|-------|-----------|
+| 0 | 2014-07-01 00:00:00 | 10844 | 1 |
+| 1 | 2014-07-01 00:30:00 | 8127 | 1 |
+| 2 | 2014-07-01 01:00:00 | 6210 | 1 |
+| 3 | 2014-07-01 01:30:00 | 4656 | 1 |
+| 4 | 2014-07-01 02:00:00 | 3820 | 1 |
+
+```python
+df.info()
+```
+
+``` text
+
+RangeIndex: 10320 entries, 0 to 10319
+Data columns (total 3 columns):
+ # Column Non-Null Count Dtype
+--- ------ -------------- -----
+ 0 ds 10320 non-null datetime64[ns]
+ 1 y 10320 non-null int64
+ 2 unique_id 10320 non-null object
+dtypes: datetime64[ns](1), int64(1), object(1)
+memory usage: 242.0+ KB
+```
+
+# Explore Data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints 8 random series from the dataset and is useful for
+basic EDA.
+
+```python
+fig = plot_series(df)
+```
+
+
+
+## The Augmented Dickey-Fuller Test
+
+An Augmented Dickey-Fuller (ADF) test is a type of statistical test that
+determines whether a unit root is present in time series data. Unit
+roots can cause unpredictable results in time series analysis. A null
+hypothesis is formed in the unit root test to determine how strongly
+time series data is affected by a trend. By accepting the null
+hypothesis, we accept the evidence that the time series data is not
+stationary. By rejecting the null hypothesis or accepting the
+alternative hypothesis, we accept the evidence that the time series data
+is generated by a stationary process. This process is also known as
+stationary trend. The values of the ADF test statistic are negative.
+Lower ADF values indicate a stronger rejection of the null hypothesis.
+
+Augmented Dickey-Fuller Test is a common statistical test used to test
+whether a given time series is stationary or not. We can achieve this by
+defining the null and alternate hypothesis.
+
+- Null Hypothesis: Time Series is non-stationary. It gives a
+ time-dependent trend.
+
+- Alternate Hypothesis: Time Series is stationary. In another term,
+ the series doesn’t depend on time.
+
+- ADF or t Statistic \< critical values: Reject the null hypothesis,
+ time series is stationary.
+
+- ADF or t Statistic \> critical values: Failed to reject the null
+ hypothesis, time series is non-stationary.
+
+```python
+def augmented_dickey_fuller_test(series , column_name):
+ print (f'Dickey-Fuller test results for columns: {column_name}')
+ dftest = adfuller(series, autolag='AIC')
+ dfoutput = pd.Series(dftest[0:4], index=['Test Statistic','p-value','No Lags Used','Number of observations used'])
+ for key,value in dftest[4].items():
+ dfoutput['Critical Value (%s)'%key] = value
+ print (dfoutput)
+ if dftest[1] <= 0.05:
+ print("Conclusion:====>")
+ print("Reject the null hypothesis")
+ print("The data is stationary")
+ else:
+ print("Conclusion:====>")
+ print("The null hypothesis cannot be rejected")
+ print("The data is not stationary")
+```
+
+
+```python
+augmented_dickey_fuller_test(df["y"],'Ads')
+```
+
+``` text
+Dickey-Fuller test results for columns: Ads
+Test Statistic -1.076452e+01
+p-value 2.472132e-19
+No Lags Used 3.900000e+01
+Number of observations used 1.028000e+04
+Critical Value (1%) -3.430986e+00
+Critical Value (5%) -2.861821e+00
+Critical Value (10%) -2.566920e+00
+dtype: float64
+Conclusion:====>
+Reject the null hypothesis
+The data is stationary
+```
+
+## Autocorrelation plots
+
+### Autocorrelation Function
+
+**Definition 1.** Let $\{x_t;1 ≤ t ≤ n\}$ be a time series sample of
+size n from $\{X_t\}$. 1. $\bar x = \sum_{t=1}^n \frac{x_t}{n}$ is
+called the sample mean of $\{X_t\}$. 2.
+$c_k =\sum_{t=1}^{n−k} (x_{t+k}- \bar x)(x_t−\bar x)/n$ is known as the
+sample autocovariance function of $\{X_t\}$. 3. $r_k = c_k /c_0$ is said
+to be the sample autocorrelation function of $\{X_t\}$.
+
+Note the following remarks about this definition:
+
+- Like most literature, this guide uses ACF to denote the sample
+ autocorrelation function as well as the autocorrelation function.
+ What is denoted by ACF can easily be identified in context.
+
+- Clearly c0 is the sample variance of $\{X_t\}$. Besides,
+ $r_0 = c_0/c_0 = 1$ and for any integer $k, |r_k| ≤ 1$.
+
+- When we compute the ACF of any sample series with a fixed length
+ $n$, we cannot put too much confidence in the values of $r_k$ for
+ large k’s, since fewer pairs of $(x_{t +k }, x_t )$ are available
+ for calculating $r_k$ as $k$ is large. One rule of thumb is not to
+ estimate $r_k$ for $k > n/3$, and another is $n ≥ 50, k ≤ n/4$. In
+ any case, it is always a good idea to be careful.
+
+- We also compute the ACF of a nonstationary time series sample by
+ Definition 1. In this case, however, the ACF or $r_k$ very slowly or
+ hardly tapers off as $k$ increases.
+
+- Plotting the ACF $(r_k)$ against lag $k$ is easy but very helpful in
+ analyzing time series sample. Such an ACF plot is known as a
+ correlogram.
+
+- If $\{X_t\}$ is stationary with $E(X_t)=0$ and $\rho_k =0$ for all
+ $k \neq 0$,thatis,itisa white noise series, then the sampling
+ distribution of $r_k$ is asymptotically normal with the mean 0 and
+ the variance of $1/n$. Hence, there is about 95% chance that $r_k$
+ falls in the interval $[−1.96/√n, 1.96/√n]$.
+
+Now we can give a summary that (1) if the time series plot of a time
+series clearly shows a trend or/and seasonality, it is surely
+nonstationary; (2) if the ACF $r_k$ very slowly or hardly tapers off as
+lag $k$ increases, the time series should also be nonstationary.
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=30, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+# Grafico
+plot_pacf(df["y"], lags=30, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+plt.savefig("../../figs/prediction_intervals_in_forecasting_models__autocorrelation.png", bbox_inches='tight')
+plt.close();
+```
+
+
+
+## Decomposition of the time series
+
+How to decompose a time series and why?
+
+In time series analysis to forecast new values, it is very important to
+know past data. More formally, we can say that it is very important to
+know the patterns that values follow over time. There can be many
+reasons that cause our forecast values to fall in the wrong direction.
+Basically, a time series consists of four components. The variation of
+those components causes the change in the pattern of the time series.
+These components are:
+
+- **Level:** This is the primary value that averages over time.
+- **Trend:** The trend is the value that causes increasing or
+ decreasing patterns in a time series.
+- **Seasonality:** This is a cyclical event that occurs in a time
+ series for a short time and causes short-term increasing or
+ decreasing patterns in a time series.
+- **Residual/Noise:** These are the random variations in the time
+ series.
+
+Combining these components over time leads to the formation of a time
+series. Most time series consist of level and noise/residual and trend
+or seasonality are optional values.
+
+If seasonality and trend are part of the time series, then there will be
+effects on the forecast value. As the pattern of the forecasted time
+series may be different from the previous time series.
+
+The combination of the components in time series can be of two types: \*
+Additive \* multiplicative
+
+Additive time series
+
+If the components of the time series are added to make the time series.
+Then the time series is called the additive time series. By
+visualization, we can say that the time series is additive if the
+increasing or decreasing pattern of the time series is similar
+throughout the series. The mathematical function of any additive time
+series can be represented by:
+$$y(t) = level + Trend + seasonality + noise$$
+
+## Multiplicative time series
+
+If the components of the time series are multiplicative together, then
+the time series is called a multiplicative time series. For
+visualization, if the time series is having exponential growth or
+decline with time, then the time series can be considered as the
+multiplicative time series. The mathematical function of the
+multiplicative time series can be represented as.
+
+$$y(t) = Level * Trend * seasonality * Noise$$
+
+### Additive
+
+```python
+a = seasonal_decompose(df["y"], model = "additive", period=24).plot()
+a.savefig('../../figs/prediction_intervals_in_forecasting_models__seasonal_decompose_aditive.png', bbox_inches='tight')
+plt.close()
+```
+
+
+
+### Multiplicative
+
+```python
+b = seasonal_decompose(df["y"], model = "Multiplicative", period=24).plot()
+b.savefig('../../figs/prediction_intervals_in_forecasting_models__seasonal_decompose_multiplicative.png', bbox_inches='tight')
+plt.close();
+```
+
+
+
+# Split the data into training and testing
+
+Let’s divide our data into sets 1. Data to train our model. 2. Data to
+test our model
+
+For the test data we will use the last 500 hours to test and evaluate
+the performance of our model.
+
+```python
+train = df[df.ds<='2015-01-21 13:30:00']
+test = df[df.ds>'2015-01-21 13:30:00']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((9820, 3), (500, 3))
+```
+
+Now let’s plot the training data and the test data.
+
+```python
+fig = plot_series(train,test)
+```
+
+
+
+# Modeling with mlforecast
+
+## Building Model
+
+We define the model that we want to use, for our example we are going to
+use the `XGBoost model`.
+
+```python
+model1 = [xgb.XGBRegressor()]
+```
+
+We can use the
+[`MLForecast.preprocess`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.preprocess)
+method to explore different transformations.
+
+If it is true that the series we are working with is a stationary series
+see (Dickey fuller test), however for the sake of practice and
+instruction in this guide, we will apply the difference to our series,
+we will do this using the `target_transforms parameter` and calling the
+diff function like:
+[`mlforecast.target_transforms.Differences`](https://Nixtla.github.io/mlforecast/target_transforms.html#differences)
+
+```python
+mlf = MLForecast(models=model1,
+ freq='30min',
+ target_transforms=[Differences([1])],
+ )
+```
+
+It is important to take into account when we use the parameter
+`target_transforms=[Differences([1])]` in case the series is stationary
+we can use a difference, or in the case that the series is not
+stationary, we can use more than one difference so that the series is
+constant over time, that is, that it is constant in mean and in
+variance.
+
+```python
+prep = mlf.preprocess(df)
+prep
+```
+
+| | ds | y | unique_id |
+|-------|---------------------|---------|-----------|
+| 1 | 2014-07-01 00:30:00 | -2717.0 | 1 |
+| 2 | 2014-07-01 01:00:00 | -1917.0 | 1 |
+| 3 | 2014-07-01 01:30:00 | -1554.0 | 1 |
+| 4 | 2014-07-01 02:00:00 | -836.0 | 1 |
+| 5 | 2014-07-01 02:30:00 | -947.0 | 1 |
+| ... | ... | ... | ... |
+| 10315 | 2015-01-31 21:30:00 | 951.0 | 1 |
+| 10316 | 2015-01-31 22:00:00 | 1051.0 | 1 |
+| 10317 | 2015-01-31 22:30:00 | 1588.0 | 1 |
+| 10318 | 2015-01-31 23:00:00 | -718.0 | 1 |
+| 10319 | 2015-01-31 23:30:00 | -303.0 | 1 |
+
+This has subtacted the lag 1 from each value, we can see what our series
+look like now.
+
+```python
+fig = plot_series(prep)
+```
+
+
+
+## Adding features
+
+### Lags
+
+Looks like the seasonality is gone, we can now try adding some lag
+features.
+
+```python
+mlf = MLForecast(models=model1,
+ freq='30min',
+ lags=[1,24],
+ target_transforms=[Differences([1])],
+ )
+```
+
+
+```python
+prep = mlf.preprocess(df)
+prep
+```
+
+| | ds | y | unique_id | lag1 | lag24 |
+|-------|---------------------|--------|-----------|--------|---------|
+| 25 | 2014-07-01 12:30:00 | -22.0 | 1 | 445.0 | -2717.0 |
+| 26 | 2014-07-01 13:00:00 | -708.0 | 1 | -22.0 | -1917.0 |
+| 27 | 2014-07-01 13:30:00 | 1281.0 | 1 | -708.0 | -1554.0 |
+| 28 | 2014-07-01 14:00:00 | 87.0 | 1 | 1281.0 | -836.0 |
+| 29 | 2014-07-01 14:30:00 | 1045.0 | 1 | 87.0 | -947.0 |
+| ... | ... | ... | ... | ... | ... |
+| 10315 | 2015-01-31 21:30:00 | 951.0 | 1 | 428.0 | 4642.0 |
+| 10316 | 2015-01-31 22:00:00 | 1051.0 | 1 | 951.0 | -519.0 |
+| 10317 | 2015-01-31 22:30:00 | 1588.0 | 1 | 1051.0 | 2411.0 |
+| 10318 | 2015-01-31 23:00:00 | -718.0 | 1 | 1588.0 | 214.0 |
+| 10319 | 2015-01-31 23:30:00 | -303.0 | 1 | -718.0 | 2595.0 |
+
+```python
+prep.drop(columns=['unique_id', 'ds']).corr()['y']
+```
+
+``` text
+y 1.000000
+lag1 0.663082
+lag24 0.155366
+Name: y, dtype: float64
+```
+
+### Lag transforms
+
+Lag transforms are defined as a dictionary where the keys are the lags
+and the values are lists of the transformations that we want to apply to
+that lag. You can refer to the [lag transformations
+guide](../how-to-guides/lag_transforms_guide.html) for more details.
+
+```python
+mlf = MLForecast(models=model1,
+ freq='30min',
+ lags=[1,24],
+ lag_transforms={1: [ExpandingMean()], 24: [RollingMean(window_size=7)]},
+ target_transforms=[Differences([1])],
+ )
+```
+
+
+```python
+prep = mlf.preprocess(df)
+prep
+```
+
+| | ds | y | unique_id | lag1 | lag24 | expanding_mean_lag1 | rolling_mean_lag24_window_size7 |
+|----|----|----|----|----|----|----|----|
+| 31 | 2014-07-01 15:30:00 | -836.0 | 1 | -1211.0 | -305.0 | 284.533325 | -1254.285767 |
+| 32 | 2014-07-01 16:00:00 | -2316.0 | 1 | -836.0 | 157.0 | 248.387100 | -843.714294 |
+| 33 | 2014-07-01 16:30:00 | -1215.0 | 1 | -2316.0 | -63.0 | 168.250000 | -578.857117 |
+| 34 | 2014-07-01 17:00:00 | 2190.0 | 1 | -1215.0 | 357.0 | 126.333336 | -305.857147 |
+| 35 | 2014-07-01 17:30:00 | 2322.0 | 1 | 2190.0 | 1849.0 | 187.029419 | 77.714287 |
+| ... | ... | ... | ... | ... | ... | ... | ... |
+| 10315 | 2015-01-31 21:30:00 | 951.0 | 1 | 428.0 | 4642.0 | 1.248303 | 2064.285645 |
+| 10316 | 2015-01-31 22:00:00 | 1051.0 | 1 | 951.0 | -519.0 | 1.340378 | 1873.428589 |
+| 10317 | 2015-01-31 22:30:00 | 1588.0 | 1 | 1051.0 | 2411.0 | 1.442129 | 2179.000000 |
+| 10318 | 2015-01-31 23:00:00 | -718.0 | 1 | 1588.0 | 214.0 | 1.595910 | 1888.714233 |
+| 10319 | 2015-01-31 23:30:00 | -303.0 | 1 | -718.0 | 2595.0 | 1.526168 | 2071.714355 |
+
+You can see that both approaches get to the same result, you can use
+whichever one you feel most comfortable with.
+
+## Date features
+
+If your time column is made of timestamps then it might make sense to
+extract features like week, dayofweek, quarter, etc. You can do that by
+passing a list of strings with pandas time/date components. You can also
+pass functions that will take the time column as input, as we’ll show
+here.
+
+```python
+mlf = MLForecast(models=model1,
+ freq='30min',
+ lags=[1,24],
+ lag_transforms={1: [ExpandingMean()], 24: [RollingMean(window_size=7)]},
+ target_transforms=[Differences([1])],
+ date_features=["year", "month", "day", "hour"]) # Seasonal data
+```
+
+
+```python
+prep = mlf.preprocess(df)
+prep
+```
+
+| | ds | y | unique_id | lag1 | lag24 | expanding_mean_lag1 | rolling_mean_lag24_window_size7 | year | month | day | hour |
+|----|----|----|----|----|----|----|----|----|----|----|----|
+| 31 | 2014-07-01 15:30:00 | -836.0 | 1 | -1211.0 | -305.0 | 284.533325 | -1254.285767 | 2014 | 7 | 1 | 15 |
+| 32 | 2014-07-01 16:00:00 | -2316.0 | 1 | -836.0 | 157.0 | 248.387100 | -843.714294 | 2014 | 7 | 1 | 16 |
+| 33 | 2014-07-01 16:30:00 | -1215.0 | 1 | -2316.0 | -63.0 | 168.250000 | -578.857117 | 2014 | 7 | 1 | 16 |
+| 34 | 2014-07-01 17:00:00 | 2190.0 | 1 | -1215.0 | 357.0 | 126.333336 | -305.857147 | 2014 | 7 | 1 | 17 |
+| 35 | 2014-07-01 17:30:00 | 2322.0 | 1 | 2190.0 | 1849.0 | 187.029419 | 77.714287 | 2014 | 7 | 1 | 17 |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| 10315 | 2015-01-31 21:30:00 | 951.0 | 1 | 428.0 | 4642.0 | 1.248303 | 2064.285645 | 2015 | 1 | 31 | 21 |
+| 10316 | 2015-01-31 22:00:00 | 1051.0 | 1 | 951.0 | -519.0 | 1.340378 | 1873.428589 | 2015 | 1 | 31 | 22 |
+| 10317 | 2015-01-31 22:30:00 | 1588.0 | 1 | 1051.0 | 2411.0 | 1.442129 | 2179.000000 | 2015 | 1 | 31 | 22 |
+| 10318 | 2015-01-31 23:00:00 | -718.0 | 1 | 1588.0 | 214.0 | 1.595910 | 1888.714233 | 2015 | 1 | 31 | 23 |
+| 10319 | 2015-01-31 23:30:00 | -303.0 | 1 | -718.0 | 2595.0 | 1.526168 | 2071.714355 | 2015 | 1 | 31 | 23 |
+
+## Fit the Model
+
+```python
+# fit the models
+mlf.fit(df,
+ fitted=True,
+prediction_intervals=PredictionIntervals(n_windows=5, h=30, method="conformal_distribution" ) )
+```
+
+``` text
+MLForecast(models=[XGBRegressor], freq=30min, lag_features=['lag1', 'lag24', 'expanding_mean_lag1', 'rolling_mean_lag24_window_size7'], date_features=['year', 'month', 'day', 'hour'], num_threads=1)
+```
+
+Let’s see the results of our model in this case the `XGBoost model`. We
+can observe it with the following instruction:
+
+Let us now visualize the fitted values of our models.
+
+```python
+result = mlf.forecast_fitted_values()
+result = result.set_index("unique_id")
+result
+```
+
+| | ds | y | XGBRegressor |
+|-----------|---------------------|---------|--------------|
+| unique_id | | | |
+| 1 | 2014-07-01 15:30:00 | 18544.0 | 18441.443359 |
+| 1 | 2014-07-01 16:00:00 | 16228.0 | 16391.152344 |
+| 1 | 2014-07-01 16:30:00 | 15013.0 | 15260.714844 |
+| 1 | 2014-07-01 17:00:00 | 17203.0 | 17066.148438 |
+| 1 | 2014-07-01 17:30:00 | 19525.0 | 19714.404297 |
+| ... | ... | ... | ... |
+| 1 | 2015-01-31 21:30:00 | 24670.0 | 24488.646484 |
+| 1 | 2015-01-31 22:00:00 | 25721.0 | 25868.865234 |
+| 1 | 2015-01-31 22:30:00 | 27309.0 | 27290.125000 |
+| 1 | 2015-01-31 23:00:00 | 26591.0 | 27123.226562 |
+| 1 | 2015-01-31 23:30:00 | 26288.0 | 26241.205078 |
+
+```python
+from statsmodels.stats.diagnostic import normal_ad
+from scipy import stats
+```
+
+
+```python
+sw_result = stats.shapiro(result["XGBRegressor"])
+ad_result = normal_ad(np.array(result["XGBRegressor"]), axis=0)
+dag_result = stats.normaltest(result["XGBRegressor"], axis=0, nan_policy='propagate')
+```
+
+It’s important to note that we can only use this method if we assume
+that the residuals of our validation predictions are normally
+distributed. To see if this is the case, we will use a PP-plot and test
+its normality with the Anderson-Darling, Kolmogorov-Smirnov, and
+D’Agostino K^2 tests.
+
+The PP-plot(Probability-to-Probability) plots the data sample against
+the normal distribution plot in such a way that if normally distributed,
+the data points will form a straight line.
+
+The three normality tests determine how likely a data sample is from a
+normally distributed population using p-values. The null hypothesis for
+each test is that “the sample came from a normally distributed
+population”. This means that if the resulting p-values are below a
+chosen alpha value, then the null hypothesis is rejected. Thus there is
+evidence to suggest that the data comes from a non-normal distribution.
+For this article, we will use an Alpha value of 0.01.
+
+```python
+result=mlf.forecast_fitted_values()
+fig, axs = plt.subplots(nrows=2, ncols=2)
+
+# plot[1,1]
+result["XGBRegressor"].plot(ax=axs[0,0])
+axs[0,0].set_title("Residuals model");
+
+# plot
+axs[0,1].hist(result["XGBRegressor"], density=True,bins=50, alpha=0.5 )
+axs[0,1].set_title("Density plot - Residual");
+
+# plot
+stats.probplot(result["XGBRegressor"], dist="norm", plot=axs[1,0])
+axs[1,0].set_title('Plot Q-Q')
+axs[1,0].annotate("SW p-val: {:.4f}".format(sw_result[1]), xy=(0.05,0.9), xycoords='axes fraction', fontsize=15,
+ bbox=dict(boxstyle="round", fc="none", ec="gray", pad=0.6))
+
+axs[1,0].annotate("AD p-val: {:.4f}".format(ad_result[1]), xy=(0.05,0.8), xycoords='axes fraction', fontsize=15,
+ bbox=dict(boxstyle="round", fc="none", ec="gray", pad=0.6))
+
+axs[1,0].annotate("DAG p-val: {:.4f}".format(dag_result[1]), xy=(0.05,0.7), xycoords='axes fraction', fontsize=15,
+ bbox=dict(boxstyle="round", fc="none", ec="gray", pad=0.6))
+# plot
+plot_acf(result["XGBRegressor"], lags=35, ax=axs[1,1],color="fuchsia")
+axs[1,1].set_title("Autocorrelation");
+
+plt.savefig("../../figs/prediction_intervals_in_forecasting_models__plot_residual_model.png", bbox_inches='tight')
+plt.close();
+```
+
+
+
+## Predict method with prediction intervals
+
+To generate forecasts use the predict method.
+
+```python
+forecast_df = mlf.predict(h=30, level=[80,95])
+forecast_df.head()
+```
+
+| | unique_id | ds | XGBRegressor | XGBRegressor-lo-95 | XGBRegressor-lo-80 | XGBRegressor-hi-80 | XGBRegressor-hi-95 |
+|----|----|----|----|----|----|----|----|
+| 0 | 1 | 2015-02-01 00:00:00 | 26320.298828 | 25559.884241 | 25680.228369 | 26960.369287 | 27080.713416 |
+| 1 | 1 | 2015-02-01 00:30:00 | 26446.472656 | 24130.429614 | 25195.461621 | 27697.483691 | 28762.515698 |
+| 2 | 1 | 2015-02-01 01:00:00 | 24909.970703 | 23094.950537 | 23579.583398 | 26240.358008 | 26724.990869 |
+| 3 | 1 | 2015-02-01 01:30:00 | 24405.402344 | 21548.628296 | 22006.662598 | 26804.142090 | 27262.176392 |
+| 4 | 1 | 2015-02-01 02:00:00 | 22292.390625 | 20666.736963 | 21130.215430 | 23454.565820 | 23918.044287 |
+
+## Plot prediction intervals
+
+Now let’s visualize the result of our forecast and the historical data
+of our time series, also let’s draw the confidence interval that we have
+obtained when making the prediction with 95% confidence.
+
+```python
+fig = plot_series(df, forecast_df, level=[80,95], max_insample_length=200,engine="matplotlib")
+fig.get_axes()[0].set_title("Prediction intervals")
+fig.savefig('../../figs/prediction_intervals_in_forecasting_models__plot_forecasting_intervals.png', bbox_inches='tight')
+```
+
+
+
+The confidence interval is a range of values that has a high probability
+of containing the true value of a variable. In machine learning time
+series models, the confidence interval is used to estimate the
+uncertainty in the predictions.
+
+One of the main benefits of using the confidence interval is that it
+allows users to understand the accuracy of the predictions. For example,
+if the confidence interval is very wide, it means that the prediction is
+less accurate. Conversely, if the confidence interval is very narrow, it
+means that the prediction is more accurate.
+
+Another benefit of the confidence interval is that it helps users make
+informed decisions. For example, if a prediction is within the
+confidence interval, it means that it is likely to come true.
+Conversely, if a prediction is outside the confidence interval, it means
+that it is less likely to come true.
+
+In general, the confidence interval is an important tool for machine
+learning time series models. It helps users understand the accuracy of
+the forecasts and make informed decisions.
+
+# References
+
+1. Changquan Huang • Alla Petukhina. Springer series (2022). Applied
+ Time Series Analysis and Forecasting with Python.
+2. Ivan Svetunkov. [Forecasting and Analytics with the Augmented
+ Dynamic Adaptive Model (ADAM)](https://openforecast.org/adam/)
+3. [James D. Hamilton. Time Series Analysis Princeton University Press,
+ Princeton, New Jersey, 1st Edition,
+ 1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Nixtla Parameters for
+ Mlforecast](https://nixtla.github.io/mlforecast/forecast.html).
+5. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+6. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ principles and practice, Time series
+ cross-validation”.](https://otexts.com/fpp3/tscv.html).
+7. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/mlforecast/favicon.svg b/mlforecast/favicon.svg
new file mode 100644
index 00000000..e5f33342
--- /dev/null
+++ b/mlforecast/favicon.svg
@@ -0,0 +1,5 @@
+
diff --git a/mlforecast/feature_engineering.html.mdx b/mlforecast/feature_engineering.html.mdx
new file mode 100644
index 00000000..233c9dde
--- /dev/null
+++ b/mlforecast/feature_engineering.html.mdx
@@ -0,0 +1,112 @@
+---
+description: Compute transformations on exogenous regressors
+output-file: feature_engineering.html
+title: Feature engineering
+---
+
+
+```python
+import numpy as np
+import pandas as pd
+from nbdev import show_doc
+
+from mlforecast.lag_transforms import ExpandingMean
+from mlforecast.utils import generate_daily_series
+```
+
+## Setup
+
+```python
+rng = np.random.RandomState(0)
+series = generate_daily_series(100, equal_ends=True)
+starts_ends = series.groupby(
+ 'unique_id', observed=True, as_index=False
+)['ds'].agg(['min', 'max'])
+prices = []
+for r in starts_ends.itertuples():
+ dates = pd.date_range(r.min, r.max + 14 * pd.offsets.Day())
+ df = pd.DataFrame({'ds': dates, 'price': rng.rand(dates.size)})
+ df['unique_id'] = r.Index
+ prices.append(df)
+prices = pd.concat(prices)
+prices['price2'] = prices['price'] * rng.rand(prices.shape[0])
+prices.head()
+```
+
+| | ds | price | unique_id | price2 |
+|-----|------------|----------|-----------|----------|
+| 0 | 2000-10-05 | 0.548814 | 0 | 0.345011 |
+| 1 | 2000-10-06 | 0.715189 | 0 | 0.445598 |
+| 2 | 2000-10-07 | 0.602763 | 0 | 0.165147 |
+| 3 | 2000-10-08 | 0.544883 | 0 | 0.041373 |
+| 4 | 2000-10-09 | 0.423655 | 0 | 0.391577 |
+
+------------------------------------------------------------------------
+
+source
+
+## transform_exog
+
+> ``` text
+> transform_exog (df:~DFType, lags:Optional[Iterable[int]]=None, lag_transf
+> orms:Optional[Dict[int,List[Union[Callable,Tuple[Callable
+> ,Any]]]]]=None, id_col:str='unique_id',
+> time_col:str='ds', num_threads:int=1)
+> ```
+
+*Compute lag features for dynamic exogenous regressors.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Dataframe with ids, times and values for the exogenous regressors. |
+| lags | Optional | None | Lags of the target to use as features. |
+| lag_transforms | Optional | None | Mapping of target lags to their transformations. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| num_threads | int | 1 | Number of threads to use when computing the features. |
+| **Returns** | **DFType** | | **Original DataFrame with the computed features** |
+
+```python
+transformed = transform_exog(
+ prices,
+ lags=[1, 2],
+ lag_transforms={1: [ExpandingMean()]}
+)
+transformed.head()
+```
+
+| | ds | price | unique_id | price2 | price_lag1 | price_lag2 | price_expanding_mean_lag1 | price2_lag1 | price2_lag2 | price2_expanding_mean_lag1 |
+|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | 2000-10-05 | 0.548814 | 0 | 0.345011 | NaN | NaN | NaN | NaN | NaN | NaN |
+| 1 | 2000-10-06 | 0.715189 | 0 | 0.445598 | 0.548814 | NaN | 0.548814 | 0.345011 | NaN | 0.345011 |
+| 2 | 2000-10-07 | 0.602763 | 0 | 0.165147 | 0.715189 | 0.548814 | 0.632001 | 0.445598 | 0.345011 | 0.395304 |
+| 3 | 2000-10-08 | 0.544883 | 0 | 0.041373 | 0.602763 | 0.715189 | 0.622255 | 0.165147 | 0.445598 | 0.318585 |
+| 4 | 2000-10-09 | 0.423655 | 0 | 0.391577 | 0.544883 | 0.602763 | 0.602912 | 0.041373 | 0.165147 | 0.249282 |
+
+```python
+import polars as pl
+```
+
+
+```python
+prices_pl = pl.from_pandas(prices)
+transformed_pl = transform_exog(
+ prices_pl,
+ lags=[1, 2],
+ lag_transforms={1: [ExpandingMean()]},
+ num_threads=2,
+)
+transformed_pl.head()
+```
+
+| ds | price | unique_id | price2 | price_lag1 | price_lag2 | price_expanding_mean_lag1 | price2_lag1 | price2_lag2 | price2_expanding_mean_lag1 |
+|----|----|----|----|----|----|----|----|----|----|
+| datetime\[ns\] | f64 | i64 | f64 | f64 | f64 | f64 | f64 | f64 | f64 |
+| 2000-10-05 00:00:00 | 0.548814 | 0 | 0.345011 | NaN | NaN | NaN | NaN | NaN | NaN |
+| 2000-10-06 00:00:00 | 0.715189 | 0 | 0.445598 | 0.548814 | NaN | 0.548814 | 0.345011 | NaN | 0.345011 |
+| 2000-10-07 00:00:00 | 0.602763 | 0 | 0.165147 | 0.715189 | 0.548814 | 0.632001 | 0.445598 | 0.345011 | 0.395304 |
+| 2000-10-08 00:00:00 | 0.544883 | 0 | 0.041373 | 0.602763 | 0.715189 | 0.622255 | 0.165147 | 0.445598 | 0.318585 |
+| 2000-10-09 00:00:00 | 0.423655 | 0 | 0.391577 | 0.544883 | 0.602763 | 0.602912 | 0.041373 | 0.165147 | 0.249282 |
+
diff --git a/mlforecast/figs/cross_validation__predictions.png b/mlforecast/figs/cross_validation__predictions.png
new file mode 100644
index 00000000..2969e3a0
Binary files /dev/null and b/mlforecast/figs/cross_validation__predictions.png differ
diff --git a/mlforecast/figs/cross_validation__series.png b/mlforecast/figs/cross_validation__series.png
new file mode 100644
index 00000000..dc35d5b4
Binary files /dev/null and b/mlforecast/figs/cross_validation__series.png differ
diff --git a/mlforecast/figs/electricity_peak_forecasting__eda.png b/mlforecast/figs/electricity_peak_forecasting__eda.png
new file mode 100644
index 00000000..bce02f59
Binary files /dev/null and b/mlforecast/figs/electricity_peak_forecasting__eda.png differ
diff --git a/mlforecast/figs/electricity_peak_forecasting__predicted_peak.png b/mlforecast/figs/electricity_peak_forecasting__predicted_peak.png
new file mode 100644
index 00000000..c5ae03b8
Binary files /dev/null and b/mlforecast/figs/electricity_peak_forecasting__predicted_peak.png differ
diff --git a/mlforecast/figs/end_to_end_walkthrough__cv.png b/mlforecast/figs/end_to_end_walkthrough__cv.png
new file mode 100644
index 00000000..956e6dd0
Binary files /dev/null and b/mlforecast/figs/end_to_end_walkthrough__cv.png differ
diff --git a/mlforecast/figs/end_to_end_walkthrough__differences.png b/mlforecast/figs/end_to_end_walkthrough__differences.png
new file mode 100644
index 00000000..1132945e
Binary files /dev/null and b/mlforecast/figs/end_to_end_walkthrough__differences.png differ
diff --git a/mlforecast/figs/end_to_end_walkthrough__eda.png b/mlforecast/figs/end_to_end_walkthrough__eda.png
new file mode 100644
index 00000000..d5b6271e
Binary files /dev/null and b/mlforecast/figs/end_to_end_walkthrough__eda.png differ
diff --git a/mlforecast/figs/end_to_end_walkthrough__final_forecast.png b/mlforecast/figs/end_to_end_walkthrough__final_forecast.png
new file mode 100644
index 00000000..d03f35ce
Binary files /dev/null and b/mlforecast/figs/end_to_end_walkthrough__final_forecast.png differ
diff --git a/mlforecast/figs/end_to_end_walkthrough__lgbcv.png b/mlforecast/figs/end_to_end_walkthrough__lgbcv.png
new file mode 100644
index 00000000..832987c9
Binary files /dev/null and b/mlforecast/figs/end_to_end_walkthrough__lgbcv.png differ
diff --git a/mlforecast/figs/end_to_end_walkthrough__predictions.png b/mlforecast/figs/end_to_end_walkthrough__predictions.png
new file mode 100644
index 00000000..918e9fca
Binary files /dev/null and b/mlforecast/figs/end_to_end_walkthrough__predictions.png differ
diff --git a/mlforecast/figs/forecast__cross_validation.png b/mlforecast/figs/forecast__cross_validation.png
new file mode 100644
index 00000000..bfb3c422
Binary files /dev/null and b/mlforecast/figs/forecast__cross_validation.png differ
diff --git a/mlforecast/figs/forecast__cross_validation_intervals.png b/mlforecast/figs/forecast__cross_validation_intervals.png
new file mode 100644
index 00000000..80a18864
Binary files /dev/null and b/mlforecast/figs/forecast__cross_validation_intervals.png differ
diff --git a/mlforecast/figs/forecast__ercot.png b/mlforecast/figs/forecast__ercot.png
new file mode 100644
index 00000000..f7d99503
Binary files /dev/null and b/mlforecast/figs/forecast__ercot.png differ
diff --git a/mlforecast/figs/forecast__predict.png b/mlforecast/figs/forecast__predict.png
new file mode 100644
index 00000000..ffddf7b1
Binary files /dev/null and b/mlforecast/figs/forecast__predict.png differ
diff --git a/mlforecast/figs/forecast__predict_intervals.png b/mlforecast/figs/forecast__predict_intervals.png
new file mode 100644
index 00000000..3a82f5dc
Binary files /dev/null and b/mlforecast/figs/forecast__predict_intervals.png differ
diff --git a/mlforecast/figs/forecast__predict_intervals_window_size_1.png b/mlforecast/figs/forecast__predict_intervals_window_size_1.png
new file mode 100644
index 00000000..dd7c6007
Binary files /dev/null and b/mlforecast/figs/forecast__predict_intervals_window_size_1.png differ
diff --git a/mlforecast/figs/load_forecasting__differences.png b/mlforecast/figs/load_forecasting__differences.png
new file mode 100644
index 00000000..f52ab593
Binary files /dev/null and b/mlforecast/figs/load_forecasting__differences.png differ
diff --git a/mlforecast/figs/load_forecasting__prediction_intervals.png b/mlforecast/figs/load_forecasting__prediction_intervals.png
new file mode 100644
index 00000000..32903322
Binary files /dev/null and b/mlforecast/figs/load_forecasting__prediction_intervals.png differ
diff --git a/mlforecast/figs/load_forecasting__predictions.png b/mlforecast/figs/load_forecasting__predictions.png
new file mode 100644
index 00000000..d96bd858
Binary files /dev/null and b/mlforecast/figs/load_forecasting__predictions.png differ
diff --git a/mlforecast/figs/load_forecasting__raw.png b/mlforecast/figs/load_forecasting__raw.png
new file mode 100644
index 00000000..d35d9ded
Binary files /dev/null and b/mlforecast/figs/load_forecasting__raw.png differ
diff --git a/mlforecast/figs/load_forecasting__transformed.png b/mlforecast/figs/load_forecasting__transformed.png
new file mode 100644
index 00000000..1544c457
Binary files /dev/null and b/mlforecast/figs/load_forecasting__transformed.png differ
diff --git a/mlforecast/figs/prediction_intervals__eda.png b/mlforecast/figs/prediction_intervals__eda.png
new file mode 100644
index 00000000..2f4cde6d
Binary files /dev/null and b/mlforecast/figs/prediction_intervals__eda.png differ
diff --git a/mlforecast/figs/prediction_intervals__knn.png b/mlforecast/figs/prediction_intervals__knn.png
new file mode 100644
index 00000000..616da6ea
Binary files /dev/null and b/mlforecast/figs/prediction_intervals__knn.png differ
diff --git a/mlforecast/figs/prediction_intervals__lasso.png b/mlforecast/figs/prediction_intervals__lasso.png
new file mode 100644
index 00000000..150e08f1
Binary files /dev/null and b/mlforecast/figs/prediction_intervals__lasso.png differ
diff --git a/mlforecast/figs/prediction_intervals__lr.png b/mlforecast/figs/prediction_intervals__lr.png
new file mode 100644
index 00000000..de72ad92
Binary files /dev/null and b/mlforecast/figs/prediction_intervals__lr.png differ
diff --git a/mlforecast/figs/prediction_intervals__mlp.png b/mlforecast/figs/prediction_intervals__mlp.png
new file mode 100644
index 00000000..ed73510c
Binary files /dev/null and b/mlforecast/figs/prediction_intervals__mlp.png differ
diff --git a/mlforecast/figs/prediction_intervals__ridge.png b/mlforecast/figs/prediction_intervals__ridge.png
new file mode 100644
index 00000000..439f20af
Binary files /dev/null and b/mlforecast/figs/prediction_intervals__ridge.png differ
diff --git a/mlforecast/figs/prediction_intervals_in_forecasting_models__autocorrelation.png b/mlforecast/figs/prediction_intervals_in_forecasting_models__autocorrelation.png
new file mode 100644
index 00000000..773ad90c
Binary files /dev/null and b/mlforecast/figs/prediction_intervals_in_forecasting_models__autocorrelation.png differ
diff --git a/mlforecast/figs/prediction_intervals_in_forecasting_models__eda.png b/mlforecast/figs/prediction_intervals_in_forecasting_models__eda.png
new file mode 100644
index 00000000..88cc40d7
Binary files /dev/null and b/mlforecast/figs/prediction_intervals_in_forecasting_models__eda.png differ
diff --git a/mlforecast/figs/prediction_intervals_in_forecasting_models__plot_forecasting_intervals.png b/mlforecast/figs/prediction_intervals_in_forecasting_models__plot_forecasting_intervals.png
new file mode 100644
index 00000000..58f777f7
Binary files /dev/null and b/mlforecast/figs/prediction_intervals_in_forecasting_models__plot_forecasting_intervals.png differ
diff --git a/mlforecast/figs/prediction_intervals_in_forecasting_models__plot_residual_model.png b/mlforecast/figs/prediction_intervals_in_forecasting_models__plot_residual_model.png
new file mode 100644
index 00000000..fc7518a6
Binary files /dev/null and b/mlforecast/figs/prediction_intervals_in_forecasting_models__plot_residual_model.png differ
diff --git a/mlforecast/figs/prediction_intervals_in_forecasting_models__plot_values.png b/mlforecast/figs/prediction_intervals_in_forecasting_models__plot_values.png
new file mode 100644
index 00000000..40edf8bf
Binary files /dev/null and b/mlforecast/figs/prediction_intervals_in_forecasting_models__plot_values.png differ
diff --git a/mlforecast/figs/prediction_intervals_in_forecasting_models__seasonal_decompose_aditive.png b/mlforecast/figs/prediction_intervals_in_forecasting_models__seasonal_decompose_aditive.png
new file mode 100644
index 00000000..7afdcb5a
Binary files /dev/null and b/mlforecast/figs/prediction_intervals_in_forecasting_models__seasonal_decompose_aditive.png differ
diff --git a/mlforecast/figs/prediction_intervals_in_forecasting_models__seasonal_decompose_multiplicative.png b/mlforecast/figs/prediction_intervals_in_forecasting_models__seasonal_decompose_multiplicative.png
new file mode 100644
index 00000000..7afdcb5a
Binary files /dev/null and b/mlforecast/figs/prediction_intervals_in_forecasting_models__seasonal_decompose_multiplicative.png differ
diff --git a/mlforecast/figs/prediction_intervals_in_forecasting_models__train_test.png b/mlforecast/figs/prediction_intervals_in_forecasting_models__train_test.png
new file mode 100644
index 00000000..09464e3d
Binary files /dev/null and b/mlforecast/figs/prediction_intervals_in_forecasting_models__train_test.png differ
diff --git a/mlforecast/figs/quick_start_local__eda.png b/mlforecast/figs/quick_start_local__eda.png
new file mode 100644
index 00000000..c29ff25c
Binary files /dev/null and b/mlforecast/figs/quick_start_local__eda.png differ
diff --git a/mlforecast/figs/quick_start_local__predictions.png b/mlforecast/figs/quick_start_local__predictions.png
new file mode 100644
index 00000000..56149acf
Binary files /dev/null and b/mlforecast/figs/quick_start_local__predictions.png differ
diff --git a/mlforecast/figs/target_transforms__diff1.png b/mlforecast/figs/target_transforms__diff1.png
new file mode 100644
index 00000000..120e9b0c
Binary files /dev/null and b/mlforecast/figs/target_transforms__diff1.png differ
diff --git a/mlforecast/figs/target_transforms__diff2.png b/mlforecast/figs/target_transforms__diff2.png
new file mode 100644
index 00000000..2fbf954d
Binary files /dev/null and b/mlforecast/figs/target_transforms__diff2.png differ
diff --git a/mlforecast/figs/target_transforms__eda.png b/mlforecast/figs/target_transforms__eda.png
new file mode 100644
index 00000000..c5b1a470
Binary files /dev/null and b/mlforecast/figs/target_transforms__eda.png differ
diff --git a/mlforecast/figs/target_transforms__log.png b/mlforecast/figs/target_transforms__log.png
new file mode 100644
index 00000000..0a138b26
Binary files /dev/null and b/mlforecast/figs/target_transforms__log.png differ
diff --git a/mlforecast/figs/target_transforms__log_diffs.png b/mlforecast/figs/target_transforms__log_diffs.png
new file mode 100644
index 00000000..9973580b
Binary files /dev/null and b/mlforecast/figs/target_transforms__log_diffs.png differ
diff --git a/mlforecast/figs/target_transforms__minmax.png b/mlforecast/figs/target_transforms__minmax.png
new file mode 100644
index 00000000..9c6c09ab
Binary files /dev/null and b/mlforecast/figs/target_transforms__minmax.png differ
diff --git a/mlforecast/figs/target_transforms__standardized.png b/mlforecast/figs/target_transforms__standardized.png
new file mode 100644
index 00000000..a76be888
Binary files /dev/null and b/mlforecast/figs/target_transforms__standardized.png differ
diff --git a/mlforecast/figs/target_transforms__zeros.png b/mlforecast/figs/target_transforms__zeros.png
new file mode 100644
index 00000000..c02c201b
Binary files /dev/null and b/mlforecast/figs/target_transforms__zeros.png differ
diff --git a/mlforecast/figs/transfer_learning__eda.png b/mlforecast/figs/transfer_learning__eda.png
new file mode 100644
index 00000000..c77b05d0
Binary files /dev/null and b/mlforecast/figs/transfer_learning__eda.png differ
diff --git a/mlforecast/figs/transfer_learning__forecast.png b/mlforecast/figs/transfer_learning__forecast.png
new file mode 100644
index 00000000..5274dd03
Binary files /dev/null and b/mlforecast/figs/transfer_learning__forecast.png differ
diff --git a/mlforecast/forecast.html.mdx b/mlforecast/forecast.html.mdx
new file mode 100644
index 00000000..9fd48631
--- /dev/null
+++ b/mlforecast/forecast.html.mdx
@@ -0,0 +1,950 @@
+---
+description: Full pipeline encapsulation
+output-file: forecast.html
+title: MLForecast
+---
+
+
+### Data
+
+This shows an example with just 4 series of the M4 dataset. If you want
+to run it yourself on all of them, you can refer to [this
+notebook](https://www.kaggle.com/code/lemuz90/m4-competition).
+
+```python
+import random
+import tempfile
+
+import lightgbm as lgb
+import matplotlib.pyplot as plt
+import numpy as np
+import xgboost as xgb
+from sklearn.linear_model import LinearRegression
+from utilsforecast.feature_engineering import time_features
+from utilsforecast.plotting import plot_series
+
+from mlforecast.lag_transforms import ExpandingMean, ExponentiallyWeightedMean, RollingMean
+from mlforecast.lgb_cv import LightGBMCV
+from mlforecast.target_transforms import Differences, LocalStandardScaler
+from mlforecast.utils import generate_daily_series
+```
+
+
+```python
+df = pd.read_parquet('https://datasets-nixtla.s3.amazonaws.com/m4-hourly.parquet')
+ids = df['unique_id'].unique()
+random.seed(0)
+sample_ids = random.choices(ids, k=4)
+sample_df = df[df['unique_id'].isin(sample_ids)]
+sample_df
+```
+
+| | unique_id | ds | y |
+|--------|-----------|------|------|
+| 86796 | H196 | 1 | 11.8 |
+| 86797 | H196 | 2 | 11.4 |
+| 86798 | H196 | 3 | 11.1 |
+| 86799 | H196 | 4 | 10.8 |
+| 86800 | H196 | 5 | 10.6 |
+| ... | ... | ... | ... |
+| 325235 | H413 | 1004 | 99.0 |
+| 325236 | H413 | 1005 | 88.0 |
+| 325237 | H413 | 1006 | 47.0 |
+| 325238 | H413 | 1007 | 41.0 |
+| 325239 | H413 | 1008 | 34.0 |
+
+We now split this data into train and validation.
+
+```python
+horizon = 48
+valid = sample_df.groupby('unique_id').tail(horizon)
+train = sample_df.drop(valid.index)
+train.shape, valid.shape
+```
+
+``` text
+((3840, 3), (192, 3))
+```
+
+------------------------------------------------------------------------
+
+source
+
+### MLForecast
+
+> ``` text
+> MLForecast (models:Union[sklearn.base.BaseEstimator,List[sklearn.base.Bas
+> eEstimator],Dict[str,sklearn.base.BaseEstimator]],
+> freq:Union[int,str], lags:Optional[Iterable[int]]=None, lag_t
+> ransforms:Optional[Dict[int,List[Union[Callable,Tuple[Callabl
+> e,Any]]]]]=None,
+> date_features:Optional[Iterable[Union[str,Callable]]]=None,
+> num_threads:int=1, target_transforms:Optional[List[Union[mlfo
+> recast.target_transforms.BaseTargetTransform,mlforecast.targe
+> t_transforms._BaseGroupedArrayTargetTransform]]]=None,
+> lag_transforms_namer:Optional[Callable]=None)
+> ```
+
+*Forecasting pipeline*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| models | Union | | Models that will be trained and used to compute the forecasts. |
+| freq | Union | | Pandas offset, pandas offset alias, e.g. ‘D’, ‘W-THU’ or integer denoting the frequency of the series. |
+| lags | Optional | None | Lags of the target to use as features. |
+| lag_transforms | Optional | None | Mapping of target lags to their transformations. |
+| date_features | Optional | None | Features computed from the dates. Can be pandas date attributes or functions that will take the dates as input. |
+| num_threads | int | 1 | Number of threads to use when computing the features. |
+| target_transforms | Optional | None | Transformations that will be applied to the target before computing the features and restored after the forecasting step. |
+| lag_transforms_namer | Optional | None | Function that takes a transformation (either function or class), a lag and extra arguments and produces a name. |
+
+The MLForecast object encapsulates the feature engineering + training
+the models + forecasting
+
+```python
+fcst = MLForecast(
+ models=lgb.LGBMRegressor(random_state=0, verbosity=-1),
+ freq=1,
+ lags=[24 * (i+1) for i in range(7)],
+ lag_transforms={
+ 48: [ExponentiallyWeightedMean(alpha=0.3)],
+ },
+ num_threads=1,
+ target_transforms=[Differences([24])],
+)
+fcst
+```
+
+``` text
+MLForecast(models=[LGBMRegressor], freq=1, lag_features=['lag24', 'lag48', 'lag72', 'lag96', 'lag120', 'lag144', 'lag168', 'exponentially_weighted_mean_lag48_alpha0.3'], date_features=[], num_threads=1)
+```
+
+Once we have this setup we can compute the features and fit the model.
+
+------------------------------------------------------------------------
+
+source
+
+### MLForecast.fit
+
+> ``` text
+> MLForecast.fit
+> (df:Union[pandas.core.frame.DataFrame,polars.dataframe.fr
+> ame.DataFrame], id_col:str='unique_id',
+> time_col:str='ds', target_col:str='y',
+> static_features:Optional[List[str]]=None,
+> dropna:bool=True, keep_last_n:Optional[int]=None,
+> max_horizon:Optional[int]=None, prediction_intervals:Opti
+> onal[mlforecast.utils.PredictionIntervals]=None,
+> fitted:bool=False, as_numpy:bool=False,
+> weight_col:Optional[str]=None)
+> ```
+
+*Apply the feature engineering and train the models.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | Union | | Series data in long format. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| static_features | Optional | None | Names of the features that are static and will be repeated when forecasting. If `None`, will consider all columns (except id_col and time_col) as static. |
+| dropna | bool | True | Drop rows with missing values produced by the transformations. |
+| keep_last_n | Optional | None | Keep only these many records from each serie for the forecasting step. Can save time and memory if your features allow it. |
+| max_horizon | Optional | None | Train this many models, where each model will predict a specific horizon. |
+| prediction_intervals | Optional | None | Configuration to calibrate prediction intervals (Conformal Prediction). |
+| fitted | bool | False | Save in-sample predictions. |
+| as_numpy | bool | False | Cast features to numpy array. |
+| weight_col | Optional | None | Column that contains the sample weights. |
+| **Returns** | **MLForecast** | | **Forecast object with series values and trained models.** |
+
+```python
+fcst = MLForecast(
+ models=lgb.LGBMRegressor(random_state=0, verbosity=-1),
+ freq=1,
+ lags=[24 * (i+1) for i in range(7)],
+ lag_transforms={
+ 48: [ExponentiallyWeightedMean(alpha=0.3)],
+ },
+ num_threads=1,
+ target_transforms=[Differences([24])],
+)
+```
+
+
+```python
+train2 = train.copy()
+train2['weight'] = np.random.default_rng(seed=0).random(train2.shape[0])
+fcst.fit(train2, weight_col='weight', as_numpy=True).predict(5)
+```
+
+| | unique_id | ds | LGBMRegressor |
+|-----|-----------|-----|---------------|
+| 0 | H196 | 961 | 16.079737 |
+| 1 | H196 | 962 | 15.679737 |
+| 2 | H196 | 963 | 15.279737 |
+| 3 | H196 | 964 | 14.979737 |
+| 4 | H196 | 965 | 14.679737 |
+| 5 | H256 | 961 | 13.279737 |
+| 6 | H256 | 962 | 12.679737 |
+| 7 | H256 | 963 | 12.379737 |
+| 8 | H256 | 964 | 12.079737 |
+| 9 | H256 | 965 | 11.879737 |
+| 10 | H381 | 961 | 56.939977 |
+| 11 | H381 | 962 | 40.314608 |
+| 12 | H381 | 963 | 33.859013 |
+| 13 | H381 | 964 | 15.498139 |
+| 14 | H381 | 965 | 25.722674 |
+| 15 | H413 | 961 | 25.131194 |
+| 16 | H413 | 962 | 19.177421 |
+| 17 | H413 | 963 | 21.250829 |
+| 18 | H413 | 964 | 18.743132 |
+| 19 | H413 | 965 | 16.027263 |
+
+```python
+fcst.cross_validation(train2, n_windows=2, h=5, weight_col='weight', as_numpy=True)
+```
+
+| | unique_id | ds | cutoff | y | LGBMRegressor |
+|-----|-----------|-----|--------|-------|---------------|
+| 0 | H196 | 951 | 950 | 24.4 | 24.288850 |
+| 1 | H196 | 952 | 950 | 24.3 | 24.188850 |
+| 2 | H196 | 953 | 950 | 23.8 | 23.688850 |
+| 3 | H196 | 954 | 950 | 22.8 | 22.688850 |
+| 4 | H196 | 955 | 950 | 21.2 | 21.088850 |
+| 5 | H256 | 951 | 950 | 19.5 | 19.688850 |
+| 6 | H256 | 952 | 950 | 19.4 | 19.488850 |
+| 7 | H256 | 953 | 950 | 18.9 | 19.088850 |
+| 8 | H256 | 954 | 950 | 18.3 | 18.388850 |
+| 9 | H256 | 955 | 950 | 17.0 | 17.088850 |
+| 10 | H381 | 951 | 950 | 182.0 | 208.327270 |
+| 11 | H381 | 952 | 950 | 222.0 | 247.768326 |
+| 12 | H381 | 953 | 950 | 288.0 | 277.965997 |
+| 13 | H381 | 954 | 950 | 264.0 | 321.532857 |
+| 14 | H381 | 955 | 950 | 191.0 | 206.316903 |
+| 15 | H413 | 951 | 950 | 77.0 | 60.972692 |
+| 16 | H413 | 952 | 950 | 91.0 | 54.936494 |
+| 17 | H413 | 953 | 950 | 76.0 | 73.949203 |
+| 18 | H413 | 954 | 950 | 68.0 | 67.087417 |
+| 19 | H413 | 955 | 950 | 68.0 | 75.896022 |
+| 20 | H196 | 956 | 955 | 19.3 | 19.287891 |
+| 21 | H196 | 957 | 955 | 18.2 | 18.187891 |
+| 22 | H196 | 958 | 955 | 17.5 | 17.487891 |
+| 23 | H196 | 959 | 955 | 16.9 | 16.887891 |
+| 24 | H196 | 960 | 955 | 16.5 | 16.487891 |
+| 25 | H256 | 956 | 955 | 15.5 | 15.687891 |
+| 26 | H256 | 957 | 955 | 14.7 | 14.787891 |
+| 27 | H256 | 958 | 955 | 14.1 | 14.287891 |
+| 28 | H256 | 959 | 955 | 13.6 | 13.787891 |
+| 29 | H256 | 960 | 955 | 13.2 | 13.387891 |
+| 30 | H381 | 956 | 955 | 130.0 | 124.117828 |
+| 31 | H381 | 957 | 955 | 113.0 | 119.180350 |
+| 32 | H381 | 958 | 955 | 94.0 | 105.356552 |
+| 33 | H381 | 959 | 955 | 192.0 | 127.095338 |
+| 34 | H381 | 960 | 955 | 87.0 | 119.875754 |
+| 35 | H413 | 956 | 955 | 59.0 | 67.993133 |
+| 36 | H413 | 957 | 955 | 58.0 | 69.869815 |
+| 37 | H413 | 958 | 955 | 53.0 | 34.717960 |
+| 38 | H413 | 959 | 955 | 38.0 | 47.665581 |
+| 39 | H413 | 960 | 955 | 46.0 | 45.940137 |
+
+```python
+fcst.fit(train, fitted=True);
+```
+
+------------------------------------------------------------------------
+
+source
+
+### MLForecast.save
+
+> ``` text
+> MLForecast.save (path:Union[str,pathlib.Path])
+> ```
+
+*Save forecast object*
+
+| | **Type** | **Details** |
+|-------------|----------|-------------------------------------------|
+| path | Union | Directory where artifacts will be stored. |
+| **Returns** | **None** | |
+
+------------------------------------------------------------------------
+
+source
+
+### MLForecast.load
+
+> ``` text
+> MLForecast.load (path:Union[str,pathlib.Path])
+> ```
+
+*Load forecast object*
+
+| | **Type** | **Details** |
+|-------------|----------------|---------------------------------|
+| path | Union | Directory with saved artifacts. |
+| **Returns** | **MLForecast** | |
+
+------------------------------------------------------------------------
+
+source
+
+### MLForecast.update
+
+> ``` text
+> MLForecast.update
+> (df:Union[pandas.core.frame.DataFrame,polars.dataframe
+> .frame.DataFrame])
+> ```
+
+*Update the values of the stored series.*
+
+| | **Type** | **Details** |
+|-------------|----------|----------------------------------|
+| df | Union | Dataframe with new observations. |
+| **Returns** | **None** | |
+
+------------------------------------------------------------------------
+
+source
+
+### MLForecast.make_future_dataframe
+
+> ``` text
+> MLForecast.make_future_dataframe (h:int)
+> ```
+
+*Create a dataframe with all ids and future times in the forecasting
+horizon.*
+
+| | **Type** | **Details** |
+|-------------|-----------|--------------------------------------------------|
+| h | int | Number of periods to predict. |
+| **Returns** | **Union** | **DataFrame with expected ids and future times** |
+
+```python
+expected_future = fcst.make_future_dataframe(h=1)
+expected_future
+```
+
+| | unique_id | ds |
+|-----|-----------|-----|
+| 0 | H196 | 961 |
+| 1 | H256 | 961 |
+| 2 | H381 | 961 |
+| 3 | H413 | 961 |
+
+------------------------------------------------------------------------
+
+source
+
+### MLForecast.get_missing_future
+
+> ``` text
+> MLForecast.get_missing_future (h:int, X_df:~DFType)
+> ```
+
+*Get the missing id and time combinations in `X_df`.*
+
+| | **Type** | **Details** |
+|--------|---------------------------|-------------------------------------|
+| h | int | Number of periods to predict. |
+| X_df | DFType | Dataframe with the future exogenous features. Should have the id column and the time column. |
+| **Returns** | **DFType** | **DataFrame with expected ids and future times missing in `X_df`** |
+
+```python
+missing_future = fcst.get_missing_future(h=1, X_df=expected_future.head(2))
+pd.testing.assert_frame_equal(
+ missing_future,
+ expected_future.tail(2).reset_index(drop=True)
+)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### MLForecast.forecast_fitted_values
+
+> ``` text
+> MLForecast.forecast_fitted_values
+> (level:Optional[List[Union[int,float]]
+> ]=None)
+> ```
+
+*Access in-sample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels between 0 and 100 for prediction intervals. |
+| **Returns** | **Union** | | **Dataframe with predictions for the training set** |
+
+```python
+fcst.forecast_fitted_values()
+```
+
+| | unique_id | ds | y | LGBMRegressor |
+|------|-----------|-----|------|---------------|
+| 0 | H196 | 193 | 12.7 | 12.671271 |
+| 1 | H196 | 194 | 12.3 | 12.271271 |
+| 2 | H196 | 195 | 11.9 | 11.871271 |
+| 3 | H196 | 196 | 11.7 | 11.671271 |
+| 4 | H196 | 197 | 11.4 | 11.471271 |
+| ... | ... | ... | ... | ... |
+| 3067 | H413 | 956 | 59.0 | 68.280574 |
+| 3068 | H413 | 957 | 58.0 | 70.427570 |
+| 3069 | H413 | 958 | 53.0 | 44.767965 |
+| 3070 | H413 | 959 | 38.0 | 48.691257 |
+| 3071 | H413 | 960 | 46.0 | 46.652238 |
+
+```python
+fcst.forecast_fitted_values(level=[90])
+```
+
+| | unique_id | ds | y | LGBMRegressor | LGBMRegressor-lo-90 | LGBMRegressor-hi-90 |
+|------|-----------|-----|------|---------------|---------------------|---------------------|
+| 0 | H196 | 193 | 12.7 | 12.671271 | 12.540634 | 12.801909 |
+| 1 | H196 | 194 | 12.3 | 12.271271 | 12.140634 | 12.401909 |
+| 2 | H196 | 195 | 11.9 | 11.871271 | 11.740634 | 12.001909 |
+| 3 | H196 | 196 | 11.7 | 11.671271 | 11.540634 | 11.801909 |
+| 4 | H196 | 197 | 11.4 | 11.471271 | 11.340634 | 11.601909 |
+| ... | ... | ... | ... | ... | ... | ... |
+| 3067 | H413 | 956 | 59.0 | 68.280574 | 58.846640 | 77.714509 |
+| 3068 | H413 | 957 | 58.0 | 70.427570 | 60.993636 | 79.861504 |
+| 3069 | H413 | 958 | 53.0 | 44.767965 | 35.334031 | 54.201899 |
+| 3070 | H413 | 959 | 38.0 | 48.691257 | 39.257323 | 58.125191 |
+| 3071 | H413 | 960 | 46.0 | 46.652238 | 37.218304 | 56.086172 |
+
+Once we’ve run this we’re ready to compute our predictions.
+
+------------------------------------------------------------------------
+
+source
+
+### MLForecast.predict
+
+> ``` text
+> MLForecast.predict (h:int,
+> before_predict_callback:Optional[Callable]=None,
+> after_predict_callback:Optional[Callable]=None,
+> new_df:Optional[~DFType]=None,
+> level:Optional[List[Union[int,float]]]=None,
+> X_df:Optional[~DFType]=None,
+> ids:Optional[List[str]]=None)
+> ```
+
+*Compute the predictions for the next `h` steps.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Number of periods to predict. |
+| before_predict_callback | Optional | None | Function to call on the features before computing the predictions. This function will take the input dataframe that will be passed to the model for predicting and should return a dataframe with the same structure. The series identifier is on the index. |
+| after_predict_callback | Optional | None | Function to call on the predictions before updating the targets. This function will take a pandas Series with the predictions and should return another one with the same structure. The series identifier is on the index. |
+| new_df | Optional | None | Series data of new observations for which forecasts are to be generated. This dataframe should have the same structure as the one used to fit the model, including any features and time series data. If `new_df` is not None, the method will generate forecasts for the new observations. |
+| level | Optional | None | Confidence levels between 0 and 100 for prediction intervals. |
+| X_df | Optional | None | Dataframe with the future exogenous features. Should have the id column and the time column. |
+| ids | Optional | None | List with subset of ids seen during training for which the forecasts should be computed. |
+| **Returns** | **DFType** | | **Predictions for each serie and timestep, with one column per model.** |
+
+```python
+predictions = fcst.predict(horizon)
+```
+
+We can see at a couple of results.
+
+```python
+results = valid.merge(predictions, on=['unique_id', 'ds'])
+fig = plot_series(forecasts_df=results)
+```
+
+
+
+#### Prediction intervals
+
+With
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast),
+you can generate prediction intervals using Conformal Prediction. To
+configure Conformal Prediction, you need to pass an instance of the
+[`PredictionIntervals`](https://Nixtla.github.io/mlforecast/utils.html#predictionintervals)
+class to the `prediction_intervals` argument of the `fit` method. The
+class takes three parameters: `n_windows`, `h` and `method`.
+
+- `n_windows` represents the number of cross-validation windows used
+ to calibrate the intervals
+- `h` is the forecast horizon
+- `method` can be `conformal_distribution` or `conformal_error`;
+ `conformal_distribution` (default) creates forecasts paths based on
+ the cross-validation errors and calculate quantiles using those
+ paths, on the other hand `conformal_error` calculates the error
+ quantiles to produce prediction intervals. The strategy will adjust
+ the intervals for each horizon step, resulting in different widths
+ for each step. Please note that a minimum of 2 cross-validation
+ windows must be used.
+
+```python
+fcst.fit(
+ train,
+ prediction_intervals=PredictionIntervals(n_windows=3, h=48)
+);
+```
+
+After that, you just have to include your desired confidence levels to
+the `predict` method using the `level` argument. Levels must lie between
+0 and 100.
+
+```python
+predictions_w_intervals = fcst.predict(48, level=[50, 80, 95])
+predictions_w_intervals.head()
+```
+
+| | unique_id | ds | LGBMRegressor | LGBMRegressor-lo-95 | LGBMRegressor-lo-80 | LGBMRegressor-lo-50 | LGBMRegressor-hi-50 | LGBMRegressor-hi-80 | LGBMRegressor-hi-95 |
+|----|----|----|----|----|----|----|----|----|----|
+| 0 | H196 | 961 | 16.071271 | 15.958042 | 15.971271 | 16.005091 | 16.137452 | 16.171271 | 16.184501 |
+| 1 | H196 | 962 | 15.671271 | 15.553632 | 15.553632 | 15.578632 | 15.763911 | 15.788911 | 15.788911 |
+| 2 | H196 | 963 | 15.271271 | 15.153632 | 15.153632 | 15.162452 | 15.380091 | 15.388911 | 15.388911 |
+| 3 | H196 | 964 | 14.971271 | 14.858042 | 14.871271 | 14.905091 | 15.037452 | 15.071271 | 15.084501 |
+| 4 | H196 | 965 | 14.671271 | 14.553632 | 14.553632 | 14.562452 | 14.780091 | 14.788911 | 14.788911 |
+
+Let’s explore the generated intervals.
+
+```python
+results = valid.merge(predictions_w_intervals, on=['unique_id', 'ds'])
+fig = plot_series(forecasts_df=results, level=[50, 80, 95])
+```
+
+
+
+If you want to reduce the computational time and produce intervals with
+the same width for the whole forecast horizon, simple pass `h=1` to the
+[`PredictionIntervals`](https://Nixtla.github.io/mlforecast/utils.html#predictionintervals)
+class. The caveat of this strategy is that in some cases, variance of
+the absolute residuals maybe be small (even zero), so the intervals may
+be too narrow.
+
+```python
+fcst.fit(
+ train,
+ prediction_intervals=PredictionIntervals(n_windows=3, h=1)
+);
+```
+
+
+```python
+predictions_w_intervals_ws_1 = fcst.predict(48, level=[80, 90, 95])
+```
+
+Let’s explore the generated intervals.
+
+```python
+results = valid.merge(predictions_w_intervals_ws_1, on=['unique_id', 'ds'])
+fig = plot_series(forecasts_df=results, level=[90])
+```
+
+
+
+#### Forecast using a pretrained model
+
+MLForecast allows you to use a pretrained model to generate forecasts
+for a new dataset. Simply provide a pandas dataframe containing the new
+observations as the value for the `new_df` argument when calling the
+`predict` method. The dataframe should have the same structure as the
+one used to fit the model, including any features and time series data.
+The function will then use the pretrained model to generate forecasts
+for the new observations. This allows you to easily apply a pretrained
+model to a new dataset and generate forecasts without the need to
+retrain the model.
+
+```python
+ercot_df = pd.read_csv('https://datasets-nixtla.s3.amazonaws.com/ERCOT-clean.csv')
+# we have to convert the ds column to integers
+# since MLForecast was trained with that structure
+ercot_df['ds'] = np.arange(1, len(ercot_df) + 1)
+# use the `new_df` argument to pass the ercot dataset
+ercot_fcsts = fcst.predict(horizon, new_df=ercot_df)
+fig = plot_series(ercot_df, ercot_fcsts, max_insample_length=48 * 2)
+```
+
+
+
+If you want to take a look at the data that will be used to train the
+models you can call `Forecast.preprocess`.
+
+------------------------------------------------------------------------
+
+source
+
+### MLForecast.preprocess
+
+> ``` text
+> MLForecast.preprocess (df:~DFType, id_col:str='unique_id',
+> time_col:str='ds', target_col:str='y',
+> static_features:Optional[List[str]]=None,
+> dropna:bool=True, keep_last_n:Optional[int]=None,
+> max_horizon:Optional[int]=None,
+> return_X_y:bool=False, as_numpy:bool=False,
+> weight_col:Optional[str]=None)
+> ```
+
+*Add the features to `data`.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Series data in long format. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| static_features | Optional | None | Names of the features that are static and will be repeated when forecasting. |
+| dropna | bool | True | Drop rows with missing values produced by the transformations. |
+| keep_last_n | Optional | None | Keep only these many records from each serie for the forecasting step. Can save time and memory if your features allow it. |
+| max_horizon | Optional | None | Train this many models, where each model will predict a specific horizon. |
+| return_X_y | bool | False | Return a tuple with the features and the target. If False will return a single dataframe. |
+| as_numpy | bool | False | Cast features to numpy array. Only works for `return_X_y=True`. |
+| weight_col | Optional | None | Column that contains the sample weights. |
+| **Returns** | **Union** | | **`df` plus added features and target(s).** |
+
+```python
+prep_df = fcst.preprocess(train)
+prep_df
+```
+
+| | unique_id | ds | y | lag24 | lag48 | lag72 | lag96 | lag120 | lag144 | lag168 | exponentially_weighted_mean_lag48_alpha0.3 |
+|----|----|----|----|----|----|----|----|----|----|----|----|
+| 86988 | H196 | 193 | 0.1 | 0.0 | 0.0 | 0.0 | 0.3 | 0.1 | 0.1 | 0.3 | 0.002810 |
+| 86989 | H196 | 194 | 0.1 | -0.1 | 0.1 | 0.0 | 0.3 | 0.1 | 0.1 | 0.3 | 0.031967 |
+| 86990 | H196 | 195 | 0.1 | -0.1 | 0.1 | 0.0 | 0.3 | 0.1 | 0.2 | 0.1 | 0.052377 |
+| 86991 | H196 | 196 | 0.1 | 0.0 | 0.0 | 0.0 | 0.3 | 0.2 | 0.1 | 0.2 | 0.036664 |
+| 86992 | H196 | 197 | 0.0 | 0.0 | 0.0 | 0.1 | 0.2 | 0.2 | 0.1 | 0.2 | 0.025665 |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| 325187 | H413 | 956 | 0.0 | 10.0 | 1.0 | 6.0 | -53.0 | 44.0 | -21.0 | 21.0 | 7.963225 |
+| 325188 | H413 | 957 | 9.0 | 10.0 | 10.0 | -7.0 | -46.0 | 27.0 | -19.0 | 24.0 | 8.574257 |
+| 325189 | H413 | 958 | 16.0 | 8.0 | 5.0 | -9.0 | -36.0 | 32.0 | -13.0 | 8.0 | 7.501980 |
+| 325190 | H413 | 959 | -3.0 | 17.0 | -7.0 | 2.0 | -31.0 | 22.0 | 5.0 | -2.0 | 3.151386 |
+| 325191 | H413 | 960 | 15.0 | 11.0 | -6.0 | -5.0 | -17.0 | 22.0 | -18.0 | 10.0 | 0.405970 |
+
+If we do this we then have to call `Forecast.fit_models`, since this
+only stores the series information.
+
+------------------------------------------------------------------------
+
+source
+
+### MLForecast.fit_models
+
+> ``` text
+> MLForecast.fit_models (X:Union[pandas.core.frame.DataFrame,polars.datafra
+> me.frame.DataFrame,numpy.ndarray],
+> y:numpy.ndarray)
+> ```
+
+*Manually train models. Use this if you called
+[`MLForecast.preprocess`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.preprocess)
+beforehand.*
+
+| | **Type** | **Details** |
+|-------------|----------------|------------------------------------------|
+| X | Union | Features. |
+| y | ndarray | Target. |
+| **Returns** | **MLForecast** | **Forecast object with trained models.** |
+
+```python
+X, y = prep_df.drop(columns=['unique_id', 'ds', 'y']), prep_df['y']
+fcst.fit_models(X, y)
+```
+
+``` text
+MLForecast(models=[LGBMRegressor], freq=1, lag_features=['lag24', 'lag48', 'lag72', 'lag96', 'lag120', 'lag144', 'lag168', 'exponentially_weighted_mean_lag48_alpha0.3'], date_features=[], num_threads=1)
+```
+
+```python
+predictions2 = fcst.predict(horizon)
+pd.testing.assert_frame_equal(predictions, predictions2)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### MLForecast.cross_validation
+
+> ``` text
+> MLForecast.cross_validation (df:~DFType, n_windows:int, h:int,
+> id_col:str='unique_id', time_col:str='ds',
+> target_col:str='y',
+> step_size:Optional[int]=None,
+> static_features:Optional[List[str]]=None,
+> dropna:bool=True,
+> keep_last_n:Optional[int]=None,
+> refit:Union[bool,int]=True,
+> max_horizon:Optional[int]=None, before_predi
+> ct_callback:Optional[Callable]=None, after_p
+> redict_callback:Optional[Callable]=None, pre
+> diction_intervals:Optional[mlforecast.utils.
+> PredictionIntervals]=None,
+> level:Optional[List[Union[int,float]]]=None,
+> input_size:Optional[int]=None,
+> fitted:bool=False, as_numpy:bool=False,
+> weight_col:Optional[str]=None)
+> ```
+
+*Perform time series cross validation. Creates `n_windows` splits where
+each window has `h` test periods, trains the models, computes the
+predictions and merges the actuals.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Series data in long format. |
+| n_windows | int | | Number of windows to evaluate. |
+| h | int | | Forecast horizon. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| step_size | Optional | None | Step size between each cross validation window. If None it will be equal to `h`. |
+| static_features | Optional | None | Names of the features that are static and will be repeated when forecasting. |
+| dropna | bool | True | Drop rows with missing values produced by the transformations. |
+| keep_last_n | Optional | None | Keep only these many records from each serie for the forecasting step. Can save time and memory if your features allow it. |
+| refit | Union | True | Retrain model for each cross validation window. If False, the models are trained at the beginning and then used to predict each window. If positive int, the models are retrained every `refit` windows. |
+| max_horizon | Optional | None | |
+| before_predict_callback | Optional | None | Function to call on the features before computing the predictions. This function will take the input dataframe that will be passed to the model for predicting and should return a dataframe with the same structure. The series identifier is on the index. |
+| after_predict_callback | Optional | None | Function to call on the predictions before updating the targets. This function will take a pandas Series with the predictions and should return another one with the same structure. The series identifier is on the index. |
+| prediction_intervals | Optional | None | Configuration to calibrate prediction intervals (Conformal Prediction). |
+| level | Optional | None | Confidence levels between 0 and 100 for prediction intervals. |
+| input_size | Optional | None | Maximum training samples per serie in each window. If None, will use an expanding window. |
+| fitted | bool | False | Store the in-sample predictions. |
+| as_numpy | bool | False | Cast features to numpy array. |
+| weight_col | Optional | None | Column that contains the sample weights. |
+| **Returns** | **DFType** | | **Predictions for each window with the series id, timestamp, last train date, target value and predictions from each model.** |
+
+If we would like to know how good our forecast will be for a specific
+model and set of features then we can perform cross validation. What
+cross validation does is take our data and split it in two parts, where
+the first part is used for training and the second one for validation.
+Since the data is time dependant we usually take the last *x*
+observations from our data as the validation set.
+
+This process is implemented in
+[`MLForecast.cross_validation`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.cross_validation),
+which takes our data and performs the process described above for
+`n_windows` times where each window has `h` validation samples in it.
+For example, if we have 100 samples and we want to perform 2 backtests
+each of size 14, the splits will be as follows:
+
+1. Train: 1 to 72. Validation: 73 to 86.
+2. Train: 1 to 86. Validation: 87 to 100.
+
+You can control the size between each cross validation window using the
+`step_size` argument. For example, if we have 100 samples and we want to
+perform 2 backtests each of size 14 and move one step ahead in each fold
+(`step_size=1`), the splits will be as follows:
+
+1. Train: 1 to 85. Validation: 86 to 99.
+2. Train: 1 to 86. Validation: 87 to 100.
+
+You can also perform cross validation without refitting your models for
+each window by setting `refit=False`. This allows you to evaluate the
+performance of your models using multiple window sizes without having to
+retrain them each time.
+
+```python
+fcst = MLForecast(
+ models=lgb.LGBMRegressor(random_state=0, verbosity=-1),
+ freq=1,
+ lags=[24 * (i+1) for i in range(7)],
+ lag_transforms={
+ 1: [RollingMean(window_size=24)],
+ 24: [RollingMean(window_size=24)],
+ 48: [ExponentiallyWeightedMean(alpha=0.3)],
+ },
+ num_threads=1,
+ target_transforms=[Differences([24])],
+)
+cv_results = fcst.cross_validation(
+ train,
+ n_windows=2,
+ h=horizon,
+ step_size=horizon,
+ fitted=True,
+)
+cv_results
+```
+
+| | unique_id | ds | cutoff | y | LGBMRegressor |
+|-----|-----------|-----|--------|------|---------------|
+| 0 | H196 | 865 | 864 | 15.5 | 15.373393 |
+| 1 | H196 | 866 | 864 | 15.1 | 14.973393 |
+| 2 | H196 | 867 | 864 | 14.8 | 14.673393 |
+| 3 | H196 | 868 | 864 | 14.4 | 14.373393 |
+| 4 | H196 | 869 | 864 | 14.2 | 14.073393 |
+| ... | ... | ... | ... | ... | ... |
+| 379 | H413 | 956 | 912 | 59.0 | 64.284167 |
+| 380 | H413 | 957 | 912 | 58.0 | 64.830429 |
+| 381 | H413 | 958 | 912 | 53.0 | 40.726851 |
+| 382 | H413 | 959 | 912 | 38.0 | 42.739657 |
+| 383 | H413 | 960 | 912 | 46.0 | 52.802769 |
+
+Since we set `fitted=True` we can access the predictions for the
+training sets as well with the `cross_validation_fitted_values` method.
+
+```python
+fcst.cross_validation_fitted_values()
+```
+
+| | unique_id | ds | fold | y | LGBMRegressor |
+|------|-----------|-----|------|------|---------------|
+| 0 | H196 | 193 | 0 | 12.7 | 12.673393 |
+| 1 | H196 | 194 | 0 | 12.3 | 12.273393 |
+| 2 | H196 | 195 | 0 | 11.9 | 11.873393 |
+| 3 | H196 | 196 | 0 | 11.7 | 11.673393 |
+| 4 | H196 | 197 | 0 | 11.4 | 11.473393 |
+| ... | ... | ... | ... | ... | ... |
+| 5563 | H413 | 908 | 1 | 49.0 | 50.620196 |
+| 5564 | H413 | 909 | 1 | 39.0 | 35.972331 |
+| 5565 | H413 | 910 | 1 | 29.0 | 29.359678 |
+| 5566 | H413 | 911 | 1 | 24.0 | 25.784563 |
+| 5567 | H413 | 912 | 1 | 20.0 | 23.168413 |
+
+We can also compute prediction intervals by passing a configuration to
+`prediction_intervals` as well as values for the width through `levels`.
+
+```python
+cv_results_intervals = fcst.cross_validation(
+ train,
+ n_windows=2,
+ h=horizon,
+ step_size=horizon,
+ prediction_intervals=PredictionIntervals(h=horizon),
+ level=[80, 90]
+)
+cv_results_intervals
+```
+
+| | unique_id | ds | cutoff | y | LGBMRegressor | LGBMRegressor-lo-90 | LGBMRegressor-lo-80 | LGBMRegressor-hi-80 | LGBMRegressor-hi-90 |
+|----|----|----|----|----|----|----|----|----|----|
+| 0 | H196 | 865 | 864 | 15.5 | 15.373393 | 15.311379 | 15.316528 | 15.430258 | 15.435407 |
+| 1 | H196 | 866 | 864 | 15.1 | 14.973393 | 14.940556 | 14.940556 | 15.006230 | 15.006230 |
+| 2 | H196 | 867 | 864 | 14.8 | 14.673393 | 14.606230 | 14.606230 | 14.740556 | 14.740556 |
+| 3 | H196 | 868 | 864 | 14.4 | 14.373393 | 14.306230 | 14.306230 | 14.440556 | 14.440556 |
+| 4 | H196 | 869 | 864 | 14.2 | 14.073393 | 14.006230 | 14.006230 | 14.140556 | 14.140556 |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| 379 | H413 | 956 | 912 | 59.0 | 64.284167 | 29.890099 | 34.371545 | 94.196788 | 98.678234 |
+| 380 | H413 | 957 | 912 | 58.0 | 64.830429 | 56.874572 | 57.827689 | 71.833169 | 72.786285 |
+| 381 | H413 | 958 | 912 | 53.0 | 40.726851 | 35.296195 | 35.846206 | 45.607495 | 46.157506 |
+| 382 | H413 | 959 | 912 | 38.0 | 42.739657 | 35.292153 | 35.807640 | 49.671674 | 50.187161 |
+| 383 | H413 | 960 | 912 | 46.0 | 52.802769 | 42.465597 | 43.895670 | 61.709869 | 63.139941 |
+
+The `refit` argument allows us to control if we want to retrain the
+models in every window. It can either be:
+
+- A boolean: True will retrain on every window and False only on the
+ first one.
+- A positive integer: The models will be trained on the first window
+ and then every `refit` windows.
+
+```python
+fcst = MLForecast(
+ models=LinearRegression(),
+ freq=1,
+ lags=[1, 24],
+)
+for refit, expected_models in zip([True, False, 2], [4, 1, 2]):
+ fcst.cross_validation(
+ train,
+ n_windows=4,
+ h=horizon,
+ refit=refit,
+ )
+ test_eq(len(fcst.cv_models_), expected_models)
+```
+
+
+```python
+fig = plot_series(forecasts_df=cv_results.drop(columns='cutoff'))
+```
+
+
+
+```python
+fig = plot_series(forecasts_df=cv_results_intervals.drop(columns='cutoff'), level=[90])
+```
+
+
+
+------------------------------------------------------------------------
+
+source
+
+### MLForecast.from_cv
+
+> ``` text
+> MLForecast.from_cv (cv:mlforecast.lgb_cv.LightGBMCV)
+> ```
+
+Once you’ve found a set of features and parameters that work for your
+problem you can build a forecast object from it using
+[`MLForecast.from_cv`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast.from_cv),
+which takes the trained
+[`LightGBMCV`](https://Nixtla.github.io/mlforecast/lgb_cv.html#lightgbmcv)
+object and builds an
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+object that will use the same features and parameters. Then you can call
+fit and predict as you normally would.
+
+```python
+cv = LightGBMCV(
+ freq=1,
+ lags=[24 * (i+1) for i in range(7)],
+ lag_transforms={
+ 48: [ExponentiallyWeightedMean(alpha=0.3)],
+ },
+ num_threads=1,
+ target_transforms=[Differences([24])]
+)
+hist = cv.fit(
+ train,
+ n_windows=2,
+ h=horizon,
+ params={'verbosity': -1},
+)
+```
+
+``` text
+[10] mape: 0.118569
+[20] mape: 0.111506
+[30] mape: 0.107314
+[40] mape: 0.106089
+[50] mape: 0.106630
+Early stopping at round 50
+Using best iteration: 40
+```
+
+```python
+fcst = MLForecast.from_cv(cv)
+assert cv.best_iteration_ == fcst.models['LGBMRegressor'].n_estimators
+```
+
diff --git a/mlforecast/grouped_array.mdx b/mlforecast/grouped_array.mdx
new file mode 100644
index 00000000..457ee1da
--- /dev/null
+++ b/mlforecast/grouped_array.mdx
@@ -0,0 +1,116 @@
+------------------------------------------------------------------------
+
+source
+
+### GroupedArray
+
+> ``` text
+> GroupedArray (data:numpy.ndarray, indptr:numpy.ndarray)
+> ```
+
+\*Array made up of different groups. Can be thought of (and iterated) as
+a list of arrays.
+
+All the data is stored in a single 1d array `data`. The indices for the
+group boundaries are stored in another 1d array `indptr`.\*
+
+```python
+import copy
+
+from fastcore.test import test_eq, test_fail
+```
+
+
+```python
+# The `GroupedArray` is used internally for storing the series values and performing transformations.
+data = np.arange(10, dtype=np.float32)
+indptr = np.array([0, 2, 10]) # group 1: [0, 1], group 2: [2..9]
+ga = GroupedArray(data, indptr)
+test_eq(len(ga), 2)
+test_eq(str(ga), 'GroupedArray(ndata=10, n_groups=2)')
+```
+
+
+```python
+# Iterate through the groups
+ga_iter = iter(ga)
+np.testing.assert_equal(next(ga_iter), np.array([0, 1]))
+np.testing.assert_equal(next(ga_iter), np.arange(2, 10))
+```
+
+
+```python
+# Take the last two observations from every group
+last_2 = ga.take_from_groups(slice(-2, None))
+np.testing.assert_equal(last_2.data, np.array([0, 1, 8, 9]))
+np.testing.assert_equal(last_2.indptr, np.array([0, 2, 4]))
+```
+
+
+```python
+# Take the last four observations from every group. Note that since group 1 only has two elements, only these are returned.
+last_4 = ga.take_from_groups(slice(-4, None))
+np.testing.assert_equal(last_4.data, np.array([0, 1, 6, 7, 8, 9]))
+np.testing.assert_equal(last_4.indptr, np.array([0, 2, 6]))
+```
+
+
+```python
+# Select a specific subset of groups
+indptr = np.array([0, 2, 4, 7, 10])
+ga2 = GroupedArray(data, indptr)
+subset = ga2.take([0, 2])
+np.testing.assert_allclose(subset[0].data, ga2[0].data)
+np.testing.assert_allclose(subset[1].data, ga2[2].data)
+```
+
+
+```python
+# The groups are [0, 1], [2, ..., 9]. expand_target(2) should take rolling pairs of them and fill with nans when there aren't enough
+np.testing.assert_equal(
+ ga.expand_target(2),
+ np.array([
+ [0, 1],
+ [1, np.nan],
+ [2, 3],
+ [3, 4],
+ [4, 5],
+ [5, 6],
+ [6, 7],
+ [7, 8],
+ [8, 9],
+ [9, np.nan]
+ ])
+)
+```
+
+
+```python
+# append
+combined = ga.append(np.array([-1, -2]))
+np.testing.assert_equal(
+ combined.data,
+ np.hstack([ga.data[:2], np.array([-1]), ga.data[2:], np.array([-2])]),
+)
+# try to append new values that don't match the number of groups
+test_fail(lambda: ga.append(np.array([1., 2., 3.])), contains='`new_data` must be of size 2')
+```
+
+
+```python
+# __setitem__
+new_vals = np.array([10, 11])
+ga[0] = new_vals
+np.testing.assert_equal(ga.data, np.append(new_vals, np.arange(2, 10)))
+```
+
+
+```python
+ga_copy = copy.copy(ga)
+ga_copy.data[0] = 900
+assert ga.data[0] == 10
+assert ga.indptr is ga_copy.indptr
+```
+
diff --git a/mlforecast/index.html.mdx b/mlforecast/index.html.mdx
new file mode 100644
index 00000000..d86fbe96
--- /dev/null
+++ b/mlforecast/index.html.mdx
@@ -0,0 +1,236 @@
+---
+output-file: index.html
+title: MLForecast 🤖
+description: >-
+ **mlforecast** is a framework to perform time series forecasting using
+ machine learning models, with the option to scale to massive amounts of
+ data using remote clusters.
+---
+
+## Install
+
+### PyPI
+
+`pip install mlforecast`
+
+### conda-forge
+
+`conda install -c conda-forge mlforecast`
+
+For more detailed instructions you can refer to the [installation
+page](https://nixtla.github.io/mlforecast/docs/getting-started/install.html).
+
+## Quick Start
+
+**Get Started with this [quick
+guide](https://nixtla.github.io/mlforecast/docs/getting-started/quick_start_local.html).**
+
+**Follow this [end-to-end
+walkthrough](https://nixtla.github.io/mlforecast/docs/getting-started/end_to_end_walkthrough.html)
+for best practices.**
+
+### Videos
+
+- [Overview](https://www.youtube.com/live/EnhyJx8l2LE)
+
+### Sample notebooks
+
+- [m5](https://www.kaggle.com/code/lemuz90/m5-mlforecast-eval)
+- [m5-polars](https://www.kaggle.com/code/lemuz90/m5-mlforecast-eval-polars)
+- [m4](https://www.kaggle.com/code/lemuz90/m4-competition)
+- [m4-cv](https://www.kaggle.com/code/lemuz90/m4-competition-cv)
+- [favorita](https://www.kaggle.com/code/lemuz90/mlforecast-favorita)
+- [VN1](https://colab.research.google.com/drive/1UdhCAk49k6HgMezG-U_1ETnAB5pYvZk9)
+
+## Why?
+
+Current Python alternatives for machine learning models are slow,
+inaccurate and don’t scale well. So we created a library that can be
+used to forecast in production environments.
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+includes efficient feature engineering to train any machine learning
+model (with `fit` and `predict` methods such as
+[`sklearn`](https://scikit-learn.org/stable/)) to fit millions of time
+series.
+
+## Features
+
+- Fastest implementations of feature engineering for time series
+ forecasting in Python.
+- Out-of-the-box compatibility with pandas, polars, spark, dask, and
+ ray.
+- Probabilistic Forecasting with Conformal Prediction.
+- Support for exogenous variables and static covariates.
+- Familiar `sklearn` syntax: `.fit` and `.predict`.
+
+Missing something? Please open an issue or write us in
+[](https://join.slack.com/t/nixtlaworkspace/shared_invite/zt-135dssye9-fWTzMpv2WBthq8NK0Yvu6A)
+
+## Examples and Guides
+
+📚 [End to End
+Walkthrough](https://nixtla.github.io/mlforecast/docs/getting-started/end_to_end_walkthrough.html):
+model training, evaluation and selection for multiple time series.
+
+🔎 [Probabilistic
+Forecasting](https://nixtla.github.io/mlforecast/docs/how-to-guides/prediction_intervals.html):
+use Conformal Prediction to produce prediciton intervals.
+
+👩🔬 [Cross
+Validation](https://nixtla.github.io/mlforecast/docs/how-to-guides/cross_validation.html):
+robust model’s performance evaluation.
+
+🔌 [Predict Demand
+Peaks](https://nixtla.github.io/mlforecast/docs/tutorials/electricity_peak_forecasting.html):
+electricity load forecasting for detecting daily peaks and reducing
+electric bills.
+
+📈 [Transfer
+Learning](https://nixtla.github.io/mlforecast/docs/how-to-guides/transfer_learning.html):
+pretrain a model using a set of time series and then predict another one
+using that pretrained model.
+
+🌡️ [Distributed
+Training](https://nixtla.github.io/mlforecast/docs/getting-started/quick_start_distributed.html):
+use a Dask, Ray or Spark cluster to train models at scale.
+
+## How to use
+
+The following provides a very basic overview, for a more detailed
+description see the
+[documentation](https://nixtla.github.io/mlforecast/).
+
+### Data setup
+
+Store your time series in a pandas dataframe in long format, that is,
+each row represents an observation for a specific serie and timestamp.
+
+```python
+from mlforecast.utils import generate_daily_series
+
+series = generate_daily_series(
+ n_series=20,
+ max_length=100,
+ n_static_features=1,
+ static_as_categorical=False,
+ with_trend=True
+)
+series.head()
+```
+
+| | unique_id | ds | y | static_0 |
+|-----|-----------|------------|------------|----------|
+| 0 | id_00 | 2000-01-01 | 17.519167 | 72 |
+| 1 | id_00 | 2000-01-02 | 87.799695 | 72 |
+| 2 | id_00 | 2000-01-03 | 177.442975 | 72 |
+| 3 | id_00 | 2000-01-04 | 232.704110 | 72 |
+| 4 | id_00 | 2000-01-05 | 317.510474 | 72 |
+
+> Note: The unique_id serves as an identifier for each distinct time
+> series in your dataset. If you are using only single time series from
+> your dataset, set this column to a constant value.
+
+### Models
+
+Next define your models, each one will be trained on all series. These
+can be any regressor that follows the scikit-learn API.
+
+```python
+import lightgbm as lgb
+from sklearn.linear_model import LinearRegression
+```
+
+
+```python
+models = [
+ lgb.LGBMRegressor(random_state=0, verbosity=-1),
+ LinearRegression(),
+]
+```
+
+### Forecast object
+
+Now instantiate an
+[`MLForecast`](https://Nixtla.github.io/mlforecast/forecast.html#mlforecast)
+object with the models and the features that you want to use. The
+features can be lags, transformations on the lags and date features. You
+can also define transformations to apply to the target before fitting,
+which will be restored when predicting.
+
+```python
+from mlforecast import MLForecast
+from mlforecast.lag_transforms import ExpandingMean, RollingMean
+from mlforecast.target_transforms import Differences
+```
+
+
+```python
+fcst = MLForecast(
+ models=models,
+ freq='D',
+ lags=[7, 14],
+ lag_transforms={
+ 1: [ExpandingMean()],
+ 7: [RollingMean(window_size=28)]
+ },
+ date_features=['dayofweek'],
+ target_transforms=[Differences([1])],
+)
+```
+
+### Training
+
+To compute the features and train the models call `fit` on your
+`Forecast` object.
+
+```python
+fcst.fit(series)
+```
+
+``` text
+MLForecast(models=[LGBMRegressor, LinearRegression], freq=D, lag_features=['lag7', 'lag14', 'expanding_mean_lag1', 'rolling_mean_lag7_window_size28'], date_features=['dayofweek'], num_threads=1)
+```
+
+### Predicting
+
+To get the forecasts for the next `n` days call `predict(n)` on the
+forecast object. This will automatically handle the updates required by
+the features using a recursive strategy.
+
+```python
+predictions = fcst.predict(14)
+predictions
+```
+
+| | unique_id | ds | LGBMRegressor | LinearRegression |
+|-----|-----------|------------|---------------|------------------|
+| 0 | id_00 | 2000-04-04 | 299.923771 | 311.432371 |
+| 1 | id_00 | 2000-04-05 | 365.424147 | 379.466214 |
+| 2 | id_00 | 2000-04-06 | 432.562441 | 460.234028 |
+| 3 | id_00 | 2000-04-07 | 495.628000 | 524.278924 |
+| 4 | id_00 | 2000-04-08 | 60.786223 | 79.828767 |
+| ... | ... | ... | ... | ... |
+| 275 | id_19 | 2000-03-23 | 36.266780 | 28.333215 |
+| 276 | id_19 | 2000-03-24 | 44.370984 | 33.368228 |
+| 277 | id_19 | 2000-03-25 | 50.746222 | 38.613001 |
+| 278 | id_19 | 2000-03-26 | 58.906524 | 43.447398 |
+| 279 | id_19 | 2000-03-27 | 63.073949 | 48.666783 |
+
+### Visualize results
+
+```python
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+fig = plot_series(series, predictions, max_ids=4, plot_random=False)
+```
+
+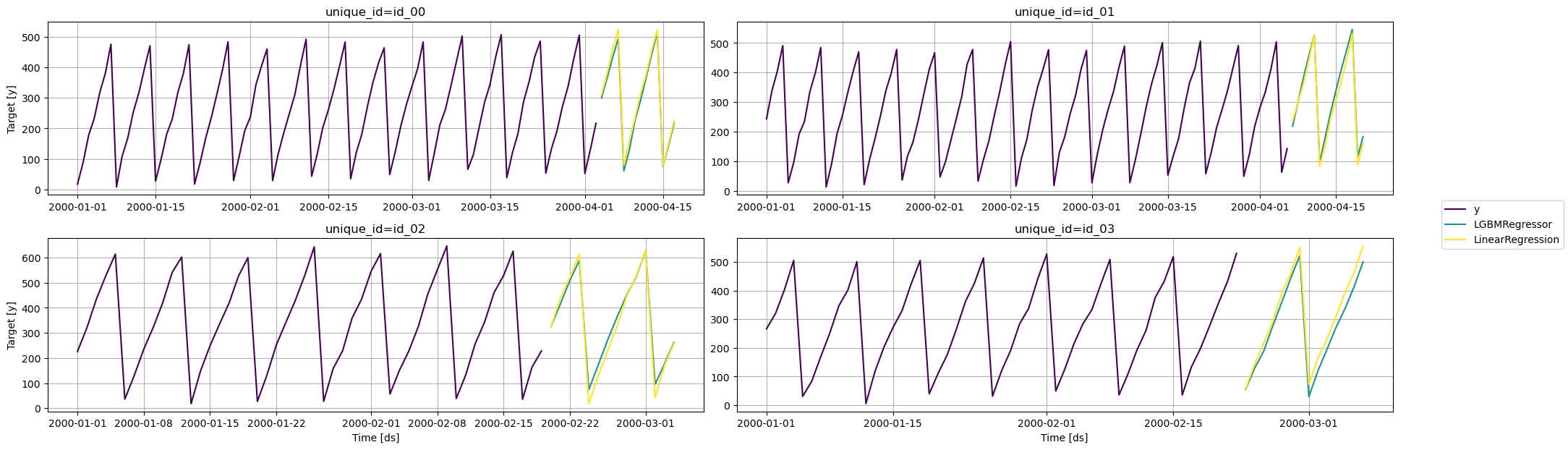
+
+## How to contribute
+
+See
+[CONTRIBUTING.md](https://github.com/Nixtla/mlforecast/blob/main/CONTRIBUTING.md).
+
diff --git a/mlforecast/lag_transforms.html.mdx b/mlforecast/lag_transforms.html.mdx
new file mode 100644
index 00000000..725c97df
--- /dev/null
+++ b/mlforecast/lag_transforms.html.mdx
@@ -0,0 +1,281 @@
+---
+description: Built-in lag transformations
+output-file: lag_transforms.html
+title: Lag transforms
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### RollingQuantile
+
+> ``` text
+> RollingQuantile (p:float, window_size:int,
+> min_samples:Optional[int]=None)
+> ```
+
+*Rolling statistic*
+
+------------------------------------------------------------------------
+
+source
+
+### RollingMax
+
+> ``` text
+> RollingMax (window_size:int, min_samples:Optional[int]=None)
+> ```
+
+*Rolling statistic*
+
+------------------------------------------------------------------------
+
+source
+
+### RollingMin
+
+> ``` text
+> RollingMin (window_size:int, min_samples:Optional[int]=None)
+> ```
+
+*Rolling statistic*
+
+------------------------------------------------------------------------
+
+source
+
+### RollingStd
+
+> ``` text
+> RollingStd (window_size:int, min_samples:Optional[int]=None)
+> ```
+
+*Rolling statistic*
+
+------------------------------------------------------------------------
+
+source
+
+### RollingMean
+
+> ``` text
+> RollingMean (window_size:int, min_samples:Optional[int]=None)
+> ```
+
+*Rolling statistic*
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalRollingQuantile
+
+> ``` text
+> SeasonalRollingQuantile (p:float, season_length:int, window_size:int,
+> min_samples:Optional[int]=None)
+> ```
+
+*Rolling statistic over seasonal periods*
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalRollingMax
+
+> ``` text
+> SeasonalRollingMax (season_length:int, window_size:int,
+> min_samples:Optional[int]=None)
+> ```
+
+*Rolling statistic over seasonal periods*
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalRollingMin
+
+> ``` text
+> SeasonalRollingMin (season_length:int, window_size:int,
+> min_samples:Optional[int]=None)
+> ```
+
+*Rolling statistic over seasonal periods*
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalRollingStd
+
+> ``` text
+> SeasonalRollingStd (season_length:int, window_size:int,
+> min_samples:Optional[int]=None)
+> ```
+
+*Rolling statistic over seasonal periods*
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalRollingMean
+
+> ``` text
+> SeasonalRollingMean (season_length:int, window_size:int,
+> min_samples:Optional[int]=None)
+> ```
+
+*Rolling statistic over seasonal periods*
+
+------------------------------------------------------------------------
+
+source
+
+### ExpandingQuantile
+
+> ``` text
+> ExpandingQuantile (p:float)
+> ```
+
+*Expanding statistic*
+
+------------------------------------------------------------------------
+
+source
+
+### ExpandingMax
+
+> ``` text
+> ExpandingMax ()
+> ```
+
+*Expanding statistic*
+
+------------------------------------------------------------------------
+
+source
+
+### ExpandingMin
+
+> ``` text
+> ExpandingMin ()
+> ```
+
+*Expanding statistic*
+
+------------------------------------------------------------------------
+
+source
+
+### ExpandingStd
+
+> ``` text
+> ExpandingStd ()
+> ```
+
+*Expanding statistic*
+
+------------------------------------------------------------------------
+
+source
+
+### ExpandingMean
+
+> ``` text
+> ExpandingMean ()
+> ```
+
+*Expanding statistic*
+
+------------------------------------------------------------------------
+
+source
+
+### ExponentiallyWeightedMean
+
+> ``` text
+> ExponentiallyWeightedMean (alpha:float)
+> ```
+
+*Exponentially weighted average*
+
+| | **Type** | **Details** |
+|-------|----------|-------------------|
+| alpha | float | Smoothing factor. |
+
+------------------------------------------------------------------------
+
+source
+
+### Offset
+
+> ``` text
+> Offset (tfm:__main__._BaseLagTransform, n:int)
+> ```
+
+*Shift series before computing transformation*
+
+| | **Type** | **Details** |
+|--------|---------------------------|-------------------------------------|
+| tfm | \_BaseLagTransform | Transformation to be applied |
+| n | int | Number of positions to shift (lag) series before applying the transformation |
+
+------------------------------------------------------------------------
+
+source
+
+### Combine
+
+> ``` text
+> Combine (tfm1:__main__._BaseLagTransform,
+> tfm2:__main__._BaseLagTransform, operator:Callable)
+> ```
+
+*Combine two lag transformations using an operator*
+
+| | **Type** | **Details** |
+|--------|---------------------------|-------------------------------------|
+| tfm1 | \_BaseLagTransform | First transformation. |
+| tfm2 | \_BaseLagTransform | Second transformation. |
+| operator | Callable | Binary operator that defines how to combine the two transformations. |
+
diff --git a/mlforecast/lgb_cv.html.mdx b/mlforecast/lgb_cv.html.mdx
new file mode 100644
index 00000000..277ea798
--- /dev/null
+++ b/mlforecast/lgb_cv.html.mdx
@@ -0,0 +1,567 @@
+---
+description: Time series cross validation with LightGBM.
+output-file: lgb_cv.html
+title: LightGBMCV
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### LightGBMCV
+
+> ``` text
+> LightGBMCV (freq:Union[int,str], lags:Optional[Iterable[int]]=None, lag_t
+> ransforms:Optional[Dict[int,List[Union[Callable,Tuple[Callabl
+> e,Any]]]]]=None,
+> date_features:Optional[Iterable[Union[str,Callable]]]=None,
+> num_threads:int=1, target_transforms:Optional[List[Union[mlfo
+> recast.target_transforms.BaseTargetTransform,mlforecast.targe
+> t_transforms._BaseGroupedArrayTargetTransform]]]=None)
+> ```
+
+*Create LightGBM CV object.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| freq | Union | | Pandas offset alias, e.g. ‘D’, ‘W-THU’ or integer denoting the frequency of the series. |
+| lags | Optional | None | Lags of the target to use as features. |
+| lag_transforms | Optional | None | Mapping of target lags to their transformations. |
+| date_features | Optional | None | Features computed from the dates. Can be pandas date attributes or functions that will take the dates as input. |
+| num_threads | int | 1 | Number of threads to use when computing the features. |
+| target_transforms | Optional | None | Transformations that will be applied to the target before computing the features and restored after the forecasting step. |
+
+## Example
+
+This shows an example with just 4 series of the M4 dataset. If you want
+to run it yourself on all of them, you can refer to [this
+notebook](https://www.kaggle.com/code/lemuz90/m4-competition-cv).
+
+```python
+import random
+
+from datasetsforecast.m4 import M4, M4Info
+from fastcore.test import test_eq, test_fail
+from mlforecast.target_transforms import Differences
+from nbdev import show_doc
+
+from mlforecast.lag_transforms import SeasonalRollingMean
+```
+
+
+```python
+group = 'Hourly'
+await M4.async_download('data', group=group)
+df, *_ = M4.load(directory='data', group=group)
+df['ds'] = df['ds'].astype('int')
+ids = df['unique_id'].unique()
+random.seed(0)
+sample_ids = random.choices(ids, k=4)
+sample_df = df[df['unique_id'].isin(sample_ids)]
+sample_df
+```
+
+| | unique_id | ds | y |
+|--------|-----------|------|------|
+| 86796 | H196 | 1 | 11.8 |
+| 86797 | H196 | 2 | 11.4 |
+| 86798 | H196 | 3 | 11.1 |
+| 86799 | H196 | 4 | 10.8 |
+| 86800 | H196 | 5 | 10.6 |
+| ... | ... | ... | ... |
+| 325235 | H413 | 1004 | 99.0 |
+| 325236 | H413 | 1005 | 88.0 |
+| 325237 | H413 | 1006 | 47.0 |
+| 325238 | H413 | 1007 | 41.0 |
+| 325239 | H413 | 1008 | 34.0 |
+
+```python
+info = M4Info[group]
+horizon = info.horizon
+valid = sample_df.groupby('unique_id').tail(horizon)
+train = sample_df.drop(valid.index)
+train.shape, valid.shape
+```
+
+``` text
+((3840, 3), (192, 3))
+```
+
+What LightGBMCV does is emulate [LightGBM’s cv
+function](https://lightgbm.readthedocs.io/en/v3.3.2/pythonapi/lightgbm.cv.html#lightgbm.cv)
+where several Boosters are trained simultaneously on different
+partitions of the data, that is, one boosting iteration is performed on
+all of them at a time. This allows to have an estimate of the error by
+iteration, so if we combine this with early stopping we can find the
+best iteration to train a final model using all the data or even use
+these individual models’ predictions to compute an ensemble.
+
+In order to have a good estimate of the forecasting performance of our
+model we compute predictions for the whole test period and compute a
+metric on that. Since this step can slow down training, there’s an
+`eval_every` parameter that can be used to control this, that is, if
+`eval_every=10` (the default) every 10 boosting iterations we’re going
+to compute forecasts for the complete window and report the error.
+
+We also have early stopping parameters:
+
+- `early_stopping_evals`: how many evaluations of the full window
+ should we go without improving to stop training?
+- `early_stopping_pct`: what’s the minimum percentage improvement we
+ want in these `early_stopping_evals` in order to keep training?
+
+This makes the LightGBMCV class a good tool to quickly test different
+configurations of the model. Consider the following example, where we’re
+going to try to find out which features can improve the performance of
+our model. We start just using lags.
+
+```python
+static_fit_config = dict(
+ n_windows=2,
+ h=horizon,
+ params={'verbose': -1},
+ compute_cv_preds=True,
+)
+cv = LightGBMCV(
+ freq=1,
+ lags=[24 * (i+1) for i in range(7)], # one week of lags
+)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### LightGBMCV.fit
+
+> ``` text
+> LightGBMCV.fit (df:pandas.core.frame.DataFrame, n_windows:int, h:int,
+> id_col:str='unique_id', time_col:str='ds',
+> target_col:str='y', step_size:Optional[int]=None,
+> num_iterations:int=100,
+> params:Optional[Dict[str,Any]]=None,
+> static_features:Optional[List[str]]=None,
+> dropna:bool=True, keep_last_n:Optional[int]=None,
+> eval_every:int=10,
+> weights:Optional[Sequence[float]]=None,
+> metric:Union[str,Callable]='mape',
+> verbose_eval:bool=True, early_stopping_evals:int=2,
+> early_stopping_pct:float=0.01,
+> compute_cv_preds:bool=False,
+> before_predict_callback:Optional[Callable]=None,
+> after_predict_callback:Optional[Callable]=None,
+> input_size:Optional[int]=None)
+> ```
+
+*Train boosters simultaneously and assess their performance on the
+complete forecasting window.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DataFrame | | Series data in long format. |
+| n_windows | int | | Number of windows to evaluate. |
+| h | int | | Forecast horizon. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| step_size | Optional | None | Step size between each cross validation window. If None it will be equal to `h`. |
+| num_iterations | int | 100 | Maximum number of boosting iterations to run. |
+| params | Optional | None | Parameters to be passed to the LightGBM Boosters. |
+| static_features | Optional | None | Names of the features that are static and will be repeated when forecasting. |
+| dropna | bool | True | Drop rows with missing values produced by the transformations. |
+| keep_last_n | Optional | None | Keep only these many records from each serie for the forecasting step. Can save time and memory if your features allow it. |
+| eval_every | int | 10 | Number of boosting iterations to train before evaluating on the whole forecast window. |
+| weights | Optional | None | Weights to multiply the metric of each window. If None, all windows have the same weight. |
+| metric | Union | mape | Metric used to assess the performance of the models and perform early stopping. |
+| verbose_eval | bool | True | Print the metrics of each evaluation. |
+| early_stopping_evals | int | 2 | Maximum number of evaluations to run without improvement. |
+| early_stopping_pct | float | 0.01 | Minimum percentage improvement in metric value in `early_stopping_evals` evaluations. |
+| compute_cv_preds | bool | False | Compute predictions for each window after finding the best iteration. |
+| before_predict_callback | Optional | None | Function to call on the features before computing the predictions. This function will take the input dataframe that will be passed to the model for predicting and should return a dataframe with the same structure. The series identifier is on the index. |
+| after_predict_callback | Optional | None | Function to call on the predictions before updating the targets. This function will take a pandas Series with the predictions and should return another one with the same structure. The series identifier is on the index. |
+| input_size | Optional | None | Maximum training samples per serie in each window. If None, will use an expanding window. |
+| **Returns** | **List** | | **List of (boosting rounds, metric value) tuples.** |
+
+```python
+hist = cv.fit(train, **static_fit_config)
+```
+
+``` text
+[LightGBM] [Info] Start training from score 51.745632
+[10] mape: 0.590690
+[20] mape: 0.251093
+[30] mape: 0.143643
+[40] mape: 0.109723
+[50] mape: 0.102099
+[60] mape: 0.099448
+[70] mape: 0.098349
+[80] mape: 0.098006
+[90] mape: 0.098718
+Early stopping at round 90
+Using best iteration: 80
+```
+
+By setting `compute_cv_preds` we get the predictions from each model on
+their corresponding validation fold.
+
+```python
+cv.cv_preds_
+```
+
+| | unique_id | ds | y | Booster | window |
+|-----|-----------|-----|------|-----------|--------|
+| 0 | H196 | 865 | 15.5 | 15.522924 | 0 |
+| 1 | H196 | 866 | 15.1 | 14.985832 | 0 |
+| 2 | H196 | 867 | 14.8 | 14.667901 | 0 |
+| 3 | H196 | 868 | 14.4 | 14.514592 | 0 |
+| 4 | H196 | 869 | 14.2 | 14.035793 | 0 |
+| ... | ... | ... | ... | ... | ... |
+| 187 | H413 | 956 | 59.0 | 77.227905 | 1 |
+| 188 | H413 | 957 | 58.0 | 80.589641 | 1 |
+| 189 | H413 | 958 | 53.0 | 53.986834 | 1 |
+| 190 | H413 | 959 | 38.0 | 36.749786 | 1 |
+| 191 | H413 | 960 | 46.0 | 36.281225 | 1 |
+
+The individual models we trained are saved, so calling `predict` returns
+the predictions from every model trained.
+
+------------------------------------------------------------------------
+
+source
+
+### LightGBMCV.predict
+
+> ``` text
+> LightGBMCV.predict (h:int,
+> before_predict_callback:Optional[Callable]=None,
+> after_predict_callback:Optional[Callable]=None,
+> X_df:Optional[pandas.core.frame.DataFrame]=None)
+> ```
+
+*Compute predictions with each of the trained boosters.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| before_predict_callback | Optional | None | Function to call on the features before computing the predictions. This function will take the input dataframe that will be passed to the model for predicting and should return a dataframe with the same structure. The series identifier is on the index. |
+| after_predict_callback | Optional | None | Function to call on the predictions before updating the targets. This function will take a pandas Series with the predictions and should return another one with the same structure. The series identifier is on the index. |
+| X_df | Optional | None | Dataframe with the future exogenous features. Should have the id column and the time column. |
+| **Returns** | **DataFrame** | | **Predictions for each serie and timestep, with one column per window.** |
+
+```python
+preds = cv.predict(horizon)
+preds
+```
+
+| | unique_id | ds | Booster0 | Booster1 |
+|-----|-----------|------|-----------|-----------|
+| 0 | H196 | 961 | 15.670252 | 15.848888 |
+| 1 | H196 | 962 | 15.522924 | 15.697399 |
+| 2 | H196 | 963 | 14.985832 | 15.166213 |
+| 3 | H196 | 964 | 14.985832 | 14.723238 |
+| 4 | H196 | 965 | 14.562152 | 14.451092 |
+| ... | ... | ... | ... | ... |
+| 187 | H413 | 1004 | 70.695242 | 65.917620 |
+| 188 | H413 | 1005 | 66.216580 | 62.615788 |
+| 189 | H413 | 1006 | 63.896573 | 67.848598 |
+| 190 | H413 | 1007 | 46.922797 | 50.981950 |
+| 191 | H413 | 1008 | 45.006541 | 42.752819 |
+
+We can average these predictions and evaluate them.
+
+```python
+def evaluate_on_valid(preds):
+ preds = preds.copy()
+ preds['final_prediction'] = preds.drop(columns=['unique_id', 'ds']).mean(1)
+ merged = preds.merge(valid, on=['unique_id', 'ds'])
+ merged['abs_err'] = abs(merged['final_prediction'] - merged['y']) / merged['y']
+ return merged.groupby('unique_id')['abs_err'].mean().mean()
+```
+
+
+```python
+eval1 = evaluate_on_valid(preds)
+eval1
+```
+
+``` text
+0.11036194712311806
+```
+
+Now, since these series are hourly, maybe we can try to remove the daily
+seasonality by taking the 168th (24 \* 7) difference, that is, substract
+the value at the same hour from one week ago, thus our target will be
+$z_t = y_{t} - y_{t-168}$. The features will be computed from this
+target and when we predict they will be automatically re-applied.
+
+```python
+cv2 = LightGBMCV(
+ freq=1,
+ target_transforms=[Differences([24 * 7])],
+ lags=[24 * (i+1) for i in range(7)],
+)
+hist2 = cv2.fit(train, **static_fit_config)
+```
+
+``` text
+[LightGBM] [Info] Start training from score 0.519010
+[10] mape: 0.089024
+[20] mape: 0.090683
+[30] mape: 0.092316
+Early stopping at round 30
+Using best iteration: 10
+```
+
+```python
+assert hist2[-1][1] < hist[-1][1]
+```
+
+Nice! We achieve a better score in less iterations. Let’s see if this
+improvement translates to the validation set as well.
+
+```python
+preds2 = cv2.predict(horizon)
+eval2 = evaluate_on_valid(preds2)
+eval2
+```
+
+``` text
+0.08956665504570135
+```
+
+```python
+assert eval2 < eval1
+```
+
+Great! Maybe we can try some lag transforms now. We’ll try the seasonal
+rolling mean that averages the values “every season”, that is, if we set
+`season_length=24` and `window_size=7` then we’ll average the value at
+the same hour for every day of the week.
+
+```python
+cv3 = LightGBMCV(
+ freq=1,
+ target_transforms=[Differences([24 * 7])],
+ lags=[24 * (i+1) for i in range(7)],
+ lag_transforms={
+ 48: [SeasonalRollingMean(season_length=24, window_size=7)],
+ },
+)
+hist3 = cv3.fit(train, **static_fit_config)
+```
+
+``` text
+[LightGBM] [Info] Start training from score 0.273641
+[10] mape: 0.086724
+[20] mape: 0.088466
+[30] mape: 0.090536
+Early stopping at round 30
+Using best iteration: 10
+```
+
+Seems like this is helping as well!
+
+```python
+assert hist3[-1][1] < hist2[-1][1]
+```
+
+Does this reflect on the validation set?
+
+```python
+preds3 = cv3.predict(horizon)
+eval3 = evaluate_on_valid(preds3)
+eval3
+```
+
+``` text
+0.08961279023129345
+```
+
+Nice! mlforecast also supports date features, but in this case our time
+column is made from integers so there aren’t many possibilites here. As
+you can see this allows you to iterate faster and get better estimates
+of the forecasting performance you can expect from your model.
+
+If you’re doing hyperparameter tuning it’s useful to be able to run a
+couple of iterations, assess the performance, and determine if this
+particular configuration isn’t promising and should be discarded. For
+example, [optuna](https://optuna.org/) has
+[pruners](https://optuna.readthedocs.io/en/stable/reference/pruners.html)
+that you can call with your current score and it decides if the trial
+should be discarded. We’ll now show how to do that.
+
+Since the CV requires a bit of setup, like the LightGBM datasets and the
+internal features, we have this `setup` method.
+
+------------------------------------------------------------------------
+
+source
+
+### LightGBMCV.setup
+
+> ``` text
+> LightGBMCV.setup (df:pandas.core.frame.DataFrame, n_windows:int, h:int,
+> id_col:str='unique_id', time_col:str='ds',
+> target_col:str='y', step_size:Optional[int]=None,
+> params:Optional[Dict[str,Any]]=None,
+> static_features:Optional[List[str]]=None,
+> dropna:bool=True, keep_last_n:Optional[int]=None,
+> weights:Optional[Sequence[float]]=None,
+> metric:Union[str,Callable]='mape',
+> input_size:Optional[int]=None)
+> ```
+
+*Initialize internal data structures to iteratively train the boosters.
+Use this before calling partial_fit.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DataFrame | | Series data in long format. |
+| n_windows | int | | Number of windows to evaluate. |
+| h | int | | Forecast horizon. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| step_size | Optional | None | Step size between each cross validation window. If None it will be equal to `h`. |
+| params | Optional | None | Parameters to be passed to the LightGBM Boosters. |
+| static_features | Optional | None | Names of the features that are static and will be repeated when forecasting. |
+| dropna | bool | True | Drop rows with missing values produced by the transformations. |
+| keep_last_n | Optional | None | Keep only these many records from each serie for the forecasting step. Can save time and memory if your features allow it. |
+| weights | Optional | None | Weights to multiply the metric of each window. If None, all windows have the same weight. |
+| metric | Union | mape | Metric used to assess the performance of the models and perform early stopping. |
+| input_size | Optional | None | Maximum training samples per serie in each window. If None, will use an expanding window. |
+| **Returns** | **LightGBMCV** | | **CV object with internal data structures for partial_fit.** |
+
+```python
+cv4 = LightGBMCV(
+ freq=1,
+ lags=[24 * (i+1) for i in range(7)],
+)
+cv4.setup(
+ train,
+ n_windows=2,
+ h=horizon,
+ params={'verbose': -1},
+)
+```
+
+``` text
+LightGBMCV(freq=1, lag_features=['lag24', 'lag48', 'lag72', 'lag96', 'lag120', 'lag144', 'lag168'], date_features=[], num_threads=1, bst_threads=8)
+```
+
+Once we have this we can call `partial_fit` to only train for some
+iterations and return the score of the forecast window.
+
+------------------------------------------------------------------------
+
+source
+
+### LightGBMCV.partial_fit
+
+> ``` text
+> LightGBMCV.partial_fit (num_iterations:int,
+> before_predict_callback:Optional[Callable]=None,
+> after_predict_callback:Optional[Callable]=None)
+> ```
+
+*Train the boosters for some iterations.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| num_iterations | int | | Number of boosting iterations to run |
+| before_predict_callback | Optional | None | Function to call on the features before computing the predictions. This function will take the input dataframe that will be passed to the model for predicting and should return a dataframe with the same structure. The series identifier is on the index. |
+| after_predict_callback | Optional | None | Function to call on the predictions before updating the targets. This function will take a pandas Series with the predictions and should return another one with the same structure. The series identifier is on the index. |
+| **Returns** | **float** | | **Weighted metric after training for num_iterations.** |
+
+```python
+score = cv4.partial_fit(10)
+score
+```
+
+``` text
+[LightGBM] [Info] Start training from score 51.745632
+```
+
+``` text
+0.5906900462828166
+```
+
+This is equal to the first evaluation from our first example.
+
+```python
+assert hist[0][1] == score
+```
+
+We can now use this score to decide if this configuration is promising.
+If we want to we can train some more iterations.
+
+```python
+score2 = cv4.partial_fit(20)
+```
+
+This is now equal to our third metric from the first example, since this
+time we trained for 20 iterations.
+
+```python
+assert hist[2][1] == score2
+```
+
+### Using a custom metric
+
+The built-in metrics are MAPE and RMSE, which are computed by serie and
+then averaged across all series. If you want to do something different
+or use a different metric entirely, you can define your own metric like
+the following:
+
+```python
+def weighted_mape(
+ y_true: pd.Series,
+ y_pred: pd.Series,
+ ids: pd.Series,
+ dates: pd.Series,
+):
+ """Weighs the MAPE by the magnitude of the series values"""
+ abs_pct_err = abs(y_true - y_pred) / abs(y_true)
+ mape_by_serie = abs_pct_err.groupby(ids).mean()
+ totals_per_serie = y_pred.groupby(ids).sum()
+ series_weights = totals_per_serie / totals_per_serie.sum()
+ return (mape_by_serie * series_weights).sum()
+```
+
+
+```python
+_ = LightGBMCV(
+ freq=1,
+ lags=[24 * (i+1) for i in range(7)],
+).fit(
+ train,
+ n_windows=2,
+ h=horizon,
+ params={'verbose': -1},
+ metric=weighted_mape,
+)
+```
+
+``` text
+[LightGBM] [Info] Start training from score 51.745632
+[10] weighted_mape: 0.480353
+[20] weighted_mape: 0.218670
+[30] weighted_mape: 0.161706
+[40] weighted_mape: 0.149992
+[50] weighted_mape: 0.149024
+[60] weighted_mape: 0.148496
+Early stopping at round 60
+Using best iteration: 60
+```
+
diff --git a/mlforecast/light.png b/mlforecast/light.png
new file mode 100644
index 00000000..bbb99b54
Binary files /dev/null and b/mlforecast/light.png differ
diff --git a/mlforecast/mint.json b/mlforecast/mint.json
new file mode 100644
index 00000000..1434c9a7
--- /dev/null
+++ b/mlforecast/mint.json
@@ -0,0 +1,109 @@
+{
+ "$schema": "https://mintlify.com/schema.json",
+ "name": "Nixtla",
+ "logo": {
+ "light": "/light.png",
+ "dark": "/dark.png"
+ },
+ "favicon": "/favicon.svg",
+ "colors": {
+ "primary": "#0E0E0E",
+ "light": "#FAFAFA",
+ "dark": "#0E0E0E",
+ "anchors": {
+ "from": "#2AD0CA",
+ "to": "#0E00F8"
+ }
+ },
+ "topbarCtaButton": {
+ "type": "github",
+ "url": "https://github.com/Nixtla/mlforecast"
+ },
+ "topAnchor": {
+ "name": "MLForecast",
+ "icon": "robot"
+ },
+ "navigation": [
+ {
+ "group": "",
+ "pages": ["index.html"]
+ },
+ {
+ "group": "Getting Started",
+ "pages": [
+ "docs/getting-started/install.html",
+ "docs/getting-started/quick_start_local.html",
+ "docs/getting-started/quick_start_distributed.html",
+ "docs/getting-started/end_to_end_walkthrough.html"
+ ]
+ },
+ {
+ "group": "How-to guides",
+ "pages": [
+ "docs/how-to-guides/exogenous_features.html",
+ "docs/how-to-guides/lag_transforms_guide.html",
+ "docs/how-to-guides/hyperparameter_optimization.html",
+ "docs/how-to-guides/sklearn_pipelines.html",
+ "docs/how-to-guides/sample_weights.html",
+ "docs/how-to-guides/cross_validation.html",
+ "docs/how-to-guides/prediction_intervals.html",
+ "docs/how-to-guides/target_transforms_guide.html",
+ "docs/how-to-guides/analyzing_models.html",
+ "docs/how-to-guides/mlflow.html",
+ "docs/how-to-guides/transforming_exog.html",
+ "docs/how-to-guides/custom_training.html",
+ "docs/how-to-guides/training_with_numpy.html",
+ "docs/how-to-guides/one_model_per_horizon.html",
+ "docs/how-to-guides/custom_date_features.html",
+ "docs/how-to-guides/predict_callbacks.html",
+ "docs/how-to-guides/predict_subset.html",
+ "docs/how-to-guides/transfer_learning.html"
+ ]
+ },
+ {
+ "group": "Tutorials",
+ "pages": [
+ "docs/tutorials/electricity_load_forecasting.html",
+ "docs/tutorials/electricity_peak_forecasting.html",
+ "docs/tutorials/prediction_intervals_in_forecasting_models.html"
+ ]
+ },
+ {
+ "group": "API Reference",
+ "pages": [
+ {
+ "group": "Local",
+ "pages": [
+ "forecast.html",
+ "auto.html",
+ "lgb_cv.html",
+ "optimization.html",
+ "utils.html",
+ "core.html",
+ "target_transforms.html",
+ "lag_transforms.html",
+ "feature_engineering.html",
+ "callbacks.html"
+ ]
+ },
+ {
+ "group": "Distributed",
+ "pages": [
+ "distributed.forecast.html",
+ {
+ "group": "Models",
+ "pages": [
+ "distributed.models.dask.lgb.html",
+ "distributed.models.dask.xgb.html",
+ "distributed.models.ray.lgb.html",
+ "distributed.models.ray.xgb.html",
+ "distributed.models.spark.lgb.html",
+ "distributed.models.spark.xgb.html"
+ ]
+ }
+ ]
+ }
+ ]
+ }
+ ]
+}
diff --git a/mlforecast/optimization.html.mdx b/mlforecast/optimization.html.mdx
new file mode 100644
index 00000000..8d9c7a19
--- /dev/null
+++ b/mlforecast/optimization.html.mdx
@@ -0,0 +1,171 @@
+---
+output-file: optimization.html
+title: Optimization
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### mlforecast_objective
+
+> ``` text
+> mlforecast_objective
+> (df:Union[pandas.core.frame.DataFrame,polars.datafr
+> ame.frame.DataFrame], config_fn:Callable[[optuna.tr
+> ial._trial.Trial],Dict[str,Any]], loss:Callable,
+> model:sklearn.base.BaseEstimator,
+> freq:Union[int,str], n_windows:int, h:int,
+> step_size:Optional[int]=None,
+> input_size:Optional[int]=None,
+> refit:Union[bool,int]=False,
+> id_col:str='unique_id', time_col:str='ds',
+> target_col:str='y')
+> ```
+
+*optuna objective function for the MLForecast class*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | Union | | |
+| config_fn | Callable | | Function that takes an optuna trial and produces a configuration with the following keys: - model_params - mlf_init_params - mlf_fit_params |
+| loss | Callable | | Function that takes the validation and train dataframes and produces a float. |
+| model | BaseEstimator | | scikit-learn compatible model to be trained |
+| freq | Union | | pandas’ or polars’ offset alias or integer denoting the frequency of the series. |
+| n_windows | int | | Number of windows to evaluate. |
+| h | int | | Forecast horizon. |
+| step_size | Optional | None | Step size between each cross validation window. If None it will be equal to `h`. |
+| input_size | Optional | None | Maximum training samples per serie in each window. If None, will use an expanding window. |
+| refit | Union | False | Retrain model for each cross validation window. If False, the models are trained at the beginning and then used to predict each window. If positive int, the models are retrained every `refit` windows. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **Callable** | | |
+
+```python
+import lightgbm as lgb
+from datasetsforecast.m4 import M4, M4Evaluation, M4Info
+from utilsforecast.losses import smape
+
+from mlforecast.lag_transforms import ExpandingMean, RollingMean
+from mlforecast.target_transforms import Differences, LocalBoxCox, LocalStandardScaler
+```
+
+
+```python
+def train_valid_split(group):
+ df, *_ = M4.load(directory='data', group=group)
+ df['ds'] = df['ds'].astype('int')
+ horizon = M4Info[group].horizon
+ valid = df.groupby('unique_id').tail(horizon)
+ train = df.drop(valid.index)
+ return train, valid
+```
+
+
+```python
+h = M4Info['Weekly'].horizon
+weekly_train, weekly_valid = train_valid_split('Weekly')
+weekly_train['unique_id'] = weekly_train['unique_id'].astype('category')
+weekly_valid['unique_id'] = weekly_valid['unique_id'].astype(weekly_train['unique_id'].dtype)
+```
+
+
+```python
+def config_fn(trial):
+ candidate_lags = [
+ [1],
+ [13],
+ [1, 13],
+ range(1, 33),
+ ]
+ lag_idx = trial.suggest_categorical('lag_idx', range(len(candidate_lags)))
+ candidate_lag_tfms = [
+ {
+ 1: [RollingMean(window_size=13)]
+ },
+ {
+ 1: [RollingMean(window_size=13)],
+ 13: [RollingMean(window_size=13)],
+ },
+ {
+ 13: [RollingMean(window_size=13)],
+ },
+ {
+ 4: [ExpandingMean(), RollingMean(window_size=4)],
+ 8: [ExpandingMean(), RollingMean(window_size=4)],
+ }
+ ]
+ lag_tfms_idx = trial.suggest_categorical('lag_tfms_idx', range(len(candidate_lag_tfms)))
+ candidate_targ_tfms = [
+ [Differences([1])],
+ [LocalBoxCox()],
+ [LocalStandardScaler()],
+ [LocalBoxCox(), Differences([1])],
+ [LocalBoxCox(), LocalStandardScaler()],
+ [LocalBoxCox(), Differences([1]), LocalStandardScaler()],
+ ]
+ targ_tfms_idx = trial.suggest_categorical('targ_tfms_idx', range(len(candidate_targ_tfms)))
+ return {
+ 'model_params': {
+ 'learning_rate': 0.05,
+ 'objective': 'l1',
+ 'bagging_freq': 1,
+ 'num_threads': 2,
+ 'verbose': -1,
+ 'force_col_wise': True,
+ 'n_estimators': trial.suggest_int('n_estimators', 10, 1000, log=True),
+ 'num_leaves': trial.suggest_int('num_leaves', 31, 1024, log=True),
+ 'lambda_l1': trial.suggest_float('lambda_l1', 0.01, 10, log=True),
+ 'lambda_l2': trial.suggest_float('lambda_l2', 0.01, 10, log=True),
+ 'bagging_fraction': trial.suggest_float('bagging_fraction', 0.75, 1.0),
+ 'feature_fraction': trial.suggest_float('feature_fraction', 0.75, 1.0),
+ },
+ 'mlf_init_params': {
+ 'lags': candidate_lags[lag_idx],
+ 'lag_transforms': candidate_lag_tfms[lag_tfms_idx],
+ 'target_transforms': candidate_targ_tfms[targ_tfms_idx],
+ },
+ 'mlf_fit_params': {
+ 'static_features': ['unique_id'],
+ }
+ }
+
+def loss(df, train_df):
+ return smape(df, models=['model'])['model'].mean()
+```
+
+
+```python
+optuna.logging.set_verbosity(optuna.logging.WARNING)
+objective = mlforecast_objective(
+ df=weekly_train,
+ config_fn=config_fn,
+ loss=loss,
+ model=lgb.LGBMRegressor(),
+ freq=1,
+ n_windows=2,
+ h=h,
+)
+study = optuna.create_study(
+ direction='minimize', sampler=optuna.samplers.TPESampler(seed=0)
+)
+study.optimize(objective, n_trials=2)
+best_cfg = study.best_trial.user_attrs['config']
+final_model = MLForecast(
+ models=[lgb.LGBMRegressor(**best_cfg['model_params'])],
+ freq=1,
+ **best_cfg['mlf_init_params'],
+)
+final_model.fit(weekly_train, **best_cfg['mlf_fit_params'])
+preds = final_model.predict(h)
+M4Evaluation.evaluate('data', 'Weekly', preds['LGBMRegressor'].values.reshape(-1, 13))
+```
+
+| | SMAPE | MASE | OWA |
+|--------|----------|----------|----------|
+| Weekly | 9.261538 | 2.614473 | 0.976158 |
+
diff --git a/mlforecast/target_transforms.html.mdx b/mlforecast/target_transforms.html.mdx
new file mode 100644
index 00000000..4f7c9031
--- /dev/null
+++ b/mlforecast/target_transforms.html.mdx
@@ -0,0 +1,322 @@
+---
+output-file: target_transforms.html
+title: Target transforms
+---
+
+
+```python
+import pandas as pd
+from fastcore.test import test_fail
+from sklearn.ensemble import HistGradientBoostingRegressor
+from sklearn.linear_model import LinearRegression
+from sklearn.preprocessing import PowerTransformer
+from utilsforecast.processing import counts_by_id
+
+from mlforecast import MLForecast
+from mlforecast.utils import generate_daily_series
+```
+
+------------------------------------------------------------------------
+
+source
+
+### BaseTargetTransform
+
+> ``` text
+> BaseTargetTransform ()
+> ```
+
+*Base class used for target transformations.*
+
+------------------------------------------------------------------------
+
+source
+
+### Differences
+
+> ``` text
+> Differences (differences:Iterable[int])
+> ```
+
+*Subtracts previous values of the serie. Can be used to remove trend or
+seasonalities.*
+
+```python
+series = generate_daily_series(10, min_length=50, max_length=100)
+```
+
+
+```python
+diffs = Differences([1, 2, 5])
+id_counts = counts_by_id(series, 'unique_id')
+indptr = np.append(0, id_counts['counts'].cumsum())
+ga = GroupedArray(series['y'].values, indptr)
+
+# differences are applied correctly
+transformed = diffs.fit_transform(ga)
+assert diffs.fitted_ == []
+expected = series.copy()
+for d in diffs.differences:
+ expected['y'] -= expected.groupby('unique_id', observed=True)['y'].shift(d)
+np.testing.assert_allclose(transformed.data, expected['y'].values)
+
+# fitted differences are restored correctly
+diffs.store_fitted = True
+transformed = diffs.fit_transform(ga)
+keep_mask = ~np.isnan(transformed.data)
+restored = diffs.inverse_transform_fitted(transformed)
+np.testing.assert_allclose(ga.data[keep_mask], restored.data[keep_mask])
+
+# test transform
+new_ga = GroupedArray(np.random.rand(10), np.arange(11))
+prev_orig = [diffs.scalers_[i].tails_[::d].copy() for i, d in enumerate(diffs.differences)]
+expected = new_ga.data - np.add.reduce(prev_orig)
+updates = diffs.update(new_ga)
+np.testing.assert_allclose(expected, updates.data)
+np.testing.assert_allclose(diffs.scalers_[0].tails_, new_ga.data)
+np.testing.assert_allclose(diffs.scalers_[1].tails_[1::2], new_ga.data - prev_orig[0])
+np.testing.assert_allclose(diffs.scalers_[2].tails_[4::5], new_ga.data - np.add.reduce(prev_orig[:2]))
+# variable sizes
+diff1 = Differences([1])
+ga = GroupedArray(np.arange(10), np.array([0, 3, 10]))
+diff1.fit_transform(ga)
+new_ga = GroupedArray(np.arange(4), np.array([0, 1, 4]))
+updates = diff1.update(new_ga)
+np.testing.assert_allclose(updates.data, np.array([0 - 2, 1 - 9, 2 - 1, 3 - 2]))
+np.testing.assert_allclose(diff1.scalers_[0].tails_, np.array([0, 3]))
+
+# short series
+ga = GroupedArray(np.arange(20), np.array([0, 2, 20]))
+test_fail(lambda: diffs.fit_transform(ga), contains="[0]")
+
+# stack
+diffs = Differences([1, 2, 5])
+ga = GroupedArray(series['y'].values, indptr)
+diffs.fit_transform(ga)
+stacked = Differences.stack([diffs, diffs])
+for i in range(len(diffs.differences)):
+ np.testing.assert_allclose(
+ stacked.scalers_[i].tails_,
+ np.tile(diffs.scalers_[i].tails_, 2)
+ )
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoDifferences
+
+> ``` text
+> AutoDifferences (max_diffs:int)
+> ```
+
+*Find and apply the optimal number of differences to each serie.*
+
+------------------------------------------------------------------------
+
+source
+
+### AutoSeasonalDifferences
+
+> ``` text
+> AutoSeasonalDifferences (season_length:int, max_diffs:int,
+> n_seasons:Optional[int]=10)
+> ```
+
+*Find and apply the optimal number of seasonal differences to each
+group.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| season_length | int | | Length of the seasonal period. |
+| max_diffs | int | | Maximum number of differences to apply. |
+| n_seasons | Optional | 10 | Number of seasons to use to determine the number of differences. Defaults to 10. If `None` will use all samples, otherwise `season_length` \* `n_seasons samples` will be used for the test. Smaller values will be faster but could be less accurate. |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoSeasonalityAndDifferences
+
+> ``` text
+> AutoSeasonalityAndDifferences (max_season_length:int, max_diffs:int,
+> n_seasons:Optional[int]=10)
+> ```
+
+*Find the length of the seasonal period and apply the optimal number of
+differences to each group.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| max_season_length | int | | Maximum length of the seasonal period. |
+| max_diffs | int | | Maximum number of differences to apply. |
+| n_seasons | Optional | 10 | Number of seasons to use to determine the number of differences. Defaults to 10. If `None` will use all samples, otherwise `max_season_length` \* `n_seasons samples` will be used for the test. Smaller values will be faster but could be less accurate. |
+
+```python
+def test_scaler(sc, series):
+ id_counts = counts_by_id(series, 'unique_id')
+ indptr = np.append(0, id_counts['counts'].cumsum())
+ ga = GroupedArray(series['y'].values, indptr)
+ transformed = sc.fit_transform(ga)
+ np.testing.assert_allclose(
+ sc.inverse_transform(transformed).data,
+ ga.data,
+ )
+ transformed2 = sc.update(ga)
+ np.testing.assert_allclose(transformed.data, transformed2.data)
+
+ idxs = [0, 7]
+ subset = ga.take(idxs)
+ transformed_subset = transformed.take(idxs)
+ subsc = sc.take(idxs)
+ np.testing.assert_allclose(
+ subsc.inverse_transform(transformed_subset).data,
+ subset.data,
+ )
+
+ stacked = sc.stack([sc, sc])
+ stacked_stats = stacked.scaler_.stats_
+ np.testing.assert_allclose(
+ stacked_stats,
+ np.tile(sc.scaler_.stats_, (2, 1)),
+ )
+```
+
+------------------------------------------------------------------------
+
+source
+
+### LocalStandardScaler
+
+> ``` text
+> LocalStandardScaler ()
+> ```
+
+*Standardizes each serie by subtracting its mean and dividing by its
+standard deviation.*
+
+```python
+test_scaler(LocalStandardScaler(), series)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### LocalMinMaxScaler
+
+> ``` text
+> LocalMinMaxScaler ()
+> ```
+
+*Scales each serie to be in the \[0, 1\] interval.*
+
+```python
+test_scaler(LocalMinMaxScaler(), series)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### LocalRobustScaler
+
+> ``` text
+> LocalRobustScaler (scale:str)
+> ```
+
+*Scaler robust to outliers.*
+
+| | **Type** | **Details** |
+|--------|---------------------------|-------------------------------------|
+| scale | str | Statistic to use for scaling. Can be either ‘iqr’ (Inter Quartile Range) or ‘mad’ (Median Asbolute Deviation) |
+
+```python
+test_scaler(LocalRobustScaler(scale='iqr'), series)
+```
+
+
+```python
+test_scaler(LocalRobustScaler(scale='mad'), series)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### LocalBoxCox
+
+> ``` text
+> LocalBoxCox ()
+> ```
+
+*Finds the optimum lambda for each serie and applies the Box-Cox
+transformation*
+
+```python
+test_scaler(LocalBoxCox(), series)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### GlobalSklearnTransformer
+
+> ``` text
+> GlobalSklearnTransformer (transformer:sklearn.base.TransformerMixin)
+> ```
+
+*Applies the same scikit-learn transformer to all series.*
+
+```python
+# need this import in order for isinstance to work
+from mlforecast.target_transforms import Differences as ExportedDifferences
+```
+
+
+```python
+sk_boxcox = PowerTransformer(method='box-cox', standardize=False)
+boxcox_global = GlobalSklearnTransformer(sk_boxcox)
+single_difference = ExportedDifferences([1])
+series = generate_daily_series(10)
+fcst = MLForecast(
+ models=[LinearRegression(), HistGradientBoostingRegressor()],
+ freq='D',
+ lags=[1, 2],
+ target_transforms=[boxcox_global, single_difference]
+)
+prep = fcst.preprocess(series, dropna=False)
+expected = (
+ pd.Series(
+ sk_boxcox.fit_transform(series[['y']])[:, 0], index=series['unique_id']
+ ).groupby('unique_id', observed=True)
+ .diff()
+ .dropna()
+ .values
+)
+np.testing.assert_allclose(prep['y'].values, expected)
+preds = fcst.fit(series).predict(5)
+```
+
diff --git a/mlforecast/utils.html.mdx b/mlforecast/utils.html.mdx
new file mode 100644
index 00000000..9406e546
--- /dev/null
+++ b/mlforecast/utils.html.mdx
@@ -0,0 +1,168 @@
+---
+output-file: utils.html
+title: Utils
+---
+
+
+```python
+from fastcore.test import test_eq, test_fail
+from nbdev import show_doc
+```
+
+------------------------------------------------------------------------
+
+source
+
+### generate_daily_series
+
+> ``` text
+> generate_daily_series (n_series:int, min_length:int=50,
+> max_length:int=500, n_static_features:int=0,
+> equal_ends:bool=False,
+> static_as_categorical:bool=True,
+> with_trend:bool=False, seed:int=0,
+> engine:str='pandas')
+> ```
+
+*Generate Synthetic Panel Series.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| n_series | int | | Number of series for synthetic panel. |
+| min_length | int | 50 | Minimum length of synthetic panel’s series. |
+| max_length | int | 500 | Maximum length of synthetic panel’s series. |
+| n_static_features | int | 0 | Number of static exogenous variables for synthetic panel’s series. |
+| equal_ends | bool | False | Series should end in the same date stamp `ds`. |
+| static_as_categorical | bool | True | Static features should have a categorical data type. |
+| with_trend | bool | False | Series should have a (positive) trend. |
+| seed | int | 0 | Random seed used for generating the data. |
+| engine | str | pandas | Output Dataframe type. |
+| **Returns** | **Union** | | **Synthetic panel with columns \[`unique_id`, `ds`, `y`\] and exogenous features.** |
+
+Generate 20 series with lengths between 100 and 1,000.
+
+```python
+n_series = 20
+min_length = 100
+max_length = 1000
+
+series = generate_daily_series(n_series, min_length, max_length)
+series
+```
+
+| | unique_id | ds | y |
+|-------|-----------|------------|----------|
+| 0 | id_00 | 2000-01-01 | 0.395863 |
+| 1 | id_00 | 2000-01-02 | 1.264447 |
+| 2 | id_00 | 2000-01-03 | 2.284022 |
+| 3 | id_00 | 2000-01-04 | 3.462798 |
+| 4 | id_00 | 2000-01-05 | 4.035518 |
+| ... | ... | ... | ... |
+| 12446 | id_19 | 2002-03-11 | 0.309275 |
+| 12447 | id_19 | 2002-03-12 | 1.189464 |
+| 12448 | id_19 | 2002-03-13 | 2.325032 |
+| 12449 | id_19 | 2002-03-14 | 3.333198 |
+| 12450 | id_19 | 2002-03-15 | 4.306117 |
+
+We can also add static features to each serie (these can be things like
+product_id or store_id). Only the first static feature (`static_0`) is
+relevant to the target.
+
+```python
+n_static_features = 2
+
+series_with_statics = generate_daily_series(n_series, min_length, max_length, n_static_features)
+series_with_statics
+```
+
+| | unique_id | ds | y | static_0 | static_1 |
+|-------|-----------|------------|------------|----------|----------|
+| 0 | id_00 | 2000-01-01 | 7.521388 | 18 | 10 |
+| 1 | id_00 | 2000-01-02 | 24.024502 | 18 | 10 |
+| 2 | id_00 | 2000-01-03 | 43.396423 | 18 | 10 |
+| 3 | id_00 | 2000-01-04 | 65.793168 | 18 | 10 |
+| 4 | id_00 | 2000-01-05 | 76.674843 | 18 | 10 |
+| ... | ... | ... | ... | ... | ... |
+| 12446 | id_19 | 2002-03-11 | 27.834771 | 89 | 42 |
+| 12447 | id_19 | 2002-03-12 | 107.051746 | 89 | 42 |
+| 12448 | id_19 | 2002-03-13 | 209.252845 | 89 | 42 |
+| 12449 | id_19 | 2002-03-14 | 299.987801 | 89 | 42 |
+| 12450 | id_19 | 2002-03-15 | 387.550536 | 89 | 42 |
+
+```python
+for i in range(n_static_features):
+ assert all(series_with_statics.groupby('unique_id')[f'static_{i}'].nunique() == 1)
+```
+
+If `equal_ends=False` (the default) then every serie has a different end
+date.
+
+```python
+assert series_with_statics.groupby('unique_id')['ds'].max().nunique() > 1
+```
+
+We can have all of them end at the same date by specifying
+`equal_ends=True`.
+
+```python
+series_equal_ends = generate_daily_series(n_series, min_length, max_length, equal_ends=True)
+
+assert series_equal_ends.groupby('unique_id')['ds'].max().nunique() == 1
+```
+
+------------------------------------------------------------------------
+
+source
+
+### generate_prices_for_series
+
+> ``` text
+> generate_prices_for_series (series:pandas.core.frame.DataFrame,
+> horizon:int=7, seed:int=0)
+> ```
+
+```python
+series_for_prices = generate_daily_series(20, n_static_features=2, equal_ends=True)
+series_for_prices.rename(columns={'static_1': 'product_id'}, inplace=True)
+prices_catalog = generate_prices_for_series(series_for_prices, horizon=7)
+prices_catalog
+```
+
+| | ds | unique_id | price |
+|------|------------|-----------|----------|
+| 0 | 2000-10-05 | id_00 | 0.548814 |
+| 1 | 2000-10-06 | id_00 | 0.715189 |
+| 2 | 2000-10-07 | id_00 | 0.602763 |
+| 3 | 2000-10-08 | id_00 | 0.544883 |
+| 4 | 2000-10-09 | id_00 | 0.423655 |
+| ... | ... | ... | ... |
+| 5009 | 2001-05-17 | id_19 | 0.288027 |
+| 5010 | 2001-05-18 | id_19 | 0.846305 |
+| 5011 | 2001-05-19 | id_19 | 0.791284 |
+| 5012 | 2001-05-20 | id_19 | 0.578636 |
+| 5013 | 2001-05-21 | id_19 | 0.288589 |
+
+```python
+test_eq(set(prices_catalog['unique_id']), set(series_for_prices['unique_id']))
+test_fail(lambda: generate_prices_for_series(series), contains='equal ends')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### PredictionIntervals
+
+> ``` text
+> PredictionIntervals (n_windows:int=2, h:int=1,
+> method:str='conformal_distribution')
+> ```
+
+*Class for storing prediction intervals metadata information.*
+
diff --git a/neuralforecast/.nojekyll b/neuralforecast/.nojekyll
new file mode 100644
index 00000000..e69de29b
diff --git a/neuralforecast/common.base_auto.html.mdx b/neuralforecast/common.base_auto.html.mdx
new file mode 100644
index 00000000..88b96b9e
--- /dev/null
+++ b/neuralforecast/common.base_auto.html.mdx
@@ -0,0 +1,199 @@
+---
+description: >-
+ Machine Learning forecasting methods are defined by many hyperparameters that
+ control their behavior, with effects ranging from their speed and memory
+ requirements to their predictive performance. For a long time, manual
+ hyperparameter tuning prevailed. This approach is time-consuming, **automated
+ hyperparameter optimization** methods have been introduced, proving more
+ efficient than manual tuning, grid search, and random search.
The
+ `BaseAuto` class offers shared API connections to hyperparameter optimization
+ algorithms like
+ [Optuna](https://docs.ray.io/en/latest/tune/examples/bayesopt_example.html),
+ [HyperOpt](https://docs.ray.io/en/latest/tune/examples/hyperopt_example.html),
+ [Dragonfly](https://docs.ray.io/en/releases-2.7.0/tune/examples/dragonfly_example.html)
+ among others through `ray`, which gives you access to grid search, bayesian
+ optimization and other state-of-the-art tools like
+ hyperband.
Comprehending the impacts of hyperparameters is still a
+ precious skill, as it can help guide the design of informed hyperparameter
+ spaces that are faster to explore automatically.
+output-file: common.base_auto.html
+title: Hyperparameter Optimization
+---
+
+
+
+
+Figure 1. Example of dataset split
+(left), validation (yellow) and test (orange). The hyperparameter
+optimization guiding signal is obtained from the validation
+set.
+
+
+------------------------------------------------------------------------
+
+### BaseAuto
+
+> ``` text
+> BaseAuto (cls_model, h, loss, valid_loss, config,
+> search_alg= object at 0x7f820028a2f0>, num_samples=10, cpus=4, gpus=0,
+> refit_with_val=False, verbose=False, alias=None, backend='ray',
+> callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| cls_model | PyTorch/PyTorchLightning model | | See `neuralforecast.models` [collection here](https://nixtla.github.io/neuralforecast/models.html). |
+| h | int | | Forecast horizon |
+| loss | PyTorch module | | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | PyTorch module | | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | dict or callable | | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+------------------------------------------------------------------------
+
+### BaseAuto.fit
+
+> ``` text
+> BaseAuto.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*BaseAuto.fit
+
+Perform the hyperparameter optimization as specified by the BaseAuto
+configuration dictionary `config`.
+
+The optimization is performed on the
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset)
+using temporal cross validation with the validation set that
+sequentially precedes the test set.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset)
+see details
+[here](https://nixtla.github.io/neuralforecast/tsdataset.html)
+`val_size`: int, size of temporal validation set (needs to be bigger
+than 0). `test_size`: int, size of temporal test set (default
+0). `random_seed`: int=None, random_seed for hyperparameter
+exploration algorithms, not yet implemented. **Returns:**
+`self`: fitted instance of `BaseAuto` with best hyperparameters and
+results .\*
+
+------------------------------------------------------------------------
+
+### BaseAuto.predict
+
+> ``` text
+> BaseAuto.predict (dataset, step_size=1, **data_kwargs)
+> ```
+
+\*BaseAuto.predict
+
+Predictions of the best performing model on validation.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset)
+see details
+[here](https://nixtla.github.io/neuralforecast/tsdataset.html)
+`step_size`: int, steps between sequential predictions, (default 1).
+`**data_kwarg`: additional parameters for the dataset module.
+`random_seed`: int=None, random_seed for hyperparameter exploration
+algorithms (not implemented). **Returns:** `y_hat`: numpy
+predictions of the
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+model. \*
+
+
+```python
+class RayLogLossesCallback(tune.Callback):
+ def on_trial_complete(self, iteration, trials, trial, **info):
+ result = trial.last_result
+ print(40 * '-' + 'Trial finished' + 40 * '-')
+ print(f'Train loss: {result["train_loss"]:.2f}. Valid loss: {result["loss"]:.2f}')
+ print(80 * '-')
+```
+
+
+```python
+config = {
+ "hidden_size": tune.choice([512]),
+ "num_layers": tune.choice([3, 4]),
+ "input_size": 12,
+ "max_steps": 10,
+ "val_check_steps": 5
+}
+auto = BaseAuto(h=12, loss=MAE(), valid_loss=MSE(), cls_model=MLP, config=config, num_samples=2, cpus=1, gpus=0, callbacks=[RayLogLossesCallback()])
+auto.fit(dataset=dataset)
+y_hat = auto.predict(dataset=dataset)
+assert mae(Y_test_df['y'].values, y_hat[:, 0]) < 200
+```
+
+
+```python
+def config_f(trial):
+ return {
+ "hidden_size": trial.suggest_categorical('hidden_size', [512]),
+ "num_layers": trial.suggest_categorical('num_layers', [3, 4]),
+ "input_size": 12,
+ "max_steps": 10,
+ "val_check_steps": 5
+ }
+
+class OptunaLogLossesCallback:
+ def __call__(self, study, trial):
+ metrics = trial.user_attrs['METRICS']
+ print(40 * '-' + 'Trial finished' + 40 * '-')
+ print(f'Train loss: {metrics["train_loss"]:.2f}. Valid loss: {metrics["loss"]:.2f}')
+ print(80 * '-')
+```
+
+
+```python
+auto2 = BaseAuto(h=12, loss=MAE(), valid_loss=MSE(), cls_model=MLP, config=config_f, search_alg=optuna.samplers.RandomSampler(), num_samples=2, backend='optuna', callbacks=[OptunaLogLossesCallback()])
+auto2.fit(dataset=dataset)
+assert isinstance(auto2.results, optuna.Study)
+y_hat2 = auto2.predict(dataset=dataset)
+assert mae(Y_test_df['y'].values, y_hat2[:, 0]) < 200
+```
+
+### References
+
+- [James Bergstra, Remi Bardenet, Yoshua Bengio, and Balazs Kegl
+ (2011). “Algorithms for Hyper-Parameter Optimization”. In: Advances
+ in Neural Information Processing Systems. url:
+ https://proceedings.neurips.cc/paper/2011/file/86e8f7ab32cfd12577bc2619bc635690-Paper.pdf](https://proceedings.neurips.cc/paper/2011/file/86e8f7ab32cfd12577bc2619bc635690-Paper.pdf)
+- [Kirthevasan Kandasamy, Karun Raju Vysyaraju, Willie Neiswanger,
+ Biswajit Paria, Christopher R. Collins, Jeff Schneider, Barnabas
+ Poczos, Eric P. Xing (2019). “Tuning Hyperparameters without Grad
+ Students: Scalable and Robust Bayesian Optimisation with Dragonfly”.
+ Journal of Machine Learning Research. url:
+ https://arxiv.org/abs/1903.06694](https://arxiv.org/abs/1903.06694)
+- [Lisha Li, Kevin Jamieson, Giulia DeSalvo, Afshin Rostamizadeh,
+ Ameet Talwalkar (2016). “Hyperband: A Novel Bandit-Based Approach to
+ Hyperparameter Optimization”. Journal of Machine Learning Research.
+ url:
+ https://arxiv.org/abs/1603.06560](https://arxiv.org/abs/1603.06560)
+
diff --git a/neuralforecast/common.base_model.mdx b/neuralforecast/common.base_model.mdx
new file mode 100644
index 00000000..7a99b673
--- /dev/null
+++ b/neuralforecast/common.base_model.mdx
@@ -0,0 +1,42 @@
+------------------------------------------------------------------------
+
+### DistributedConfig
+
+> ``` text
+> DistributedConfig (partitions_path:str, num_nodes:int, devices:int)
+> ```
+
+------------------------------------------------------------------------
+
+### BaseModel
+
+> ``` text
+> BaseModel (h:int, input_size:int, loss:Union[neuralforecast.losses.pytorc
+> h.BasePointLoss,neuralforecast.losses.pytorch.DistributionLoss
+> ,torch.nn.modules.module.Module], valid_loss:Union[neuralforec
+> ast.losses.pytorch.BasePointLoss,neuralforecast.losses.pytorch
+> .DistributionLoss,torch.nn.modules.module.Module],
+> learning_rate:float, max_steps:int, val_check_steps:int,
+> batch_size:int, valid_batch_size:Optional[int],
+> windows_batch_size:int,
+> inference_windows_batch_size:Optional[int],
+> start_padding_enabled:bool, n_series:Optional[int]=None,
+> n_samples:Optional[int]=100, h_train:int=1,
+> inference_input_size:Optional[int]=None, step_size:int=1,
+> num_lr_decays:int=0, early_stop_patience_steps:int=-1,
+> scaler_type:str='identity',
+> futr_exog_list:Optional[List]=None,
+> hist_exog_list:Optional[List]=None,
+> stat_exog_list:Optional[List]=None,
+> exclude_insample_y:Optional[bool]=False,
+> drop_last_loader:Optional[bool]=False,
+> random_seed:Optional[int]=1, alias:Optional[str]=None,
+> optimizer:Optional[torch.optim.optimizer.Optimizer]=None,
+> optimizer_kwargs:Optional[Dict]=None, lr_scheduler:Optional[to
+> rch.optim.lr_scheduler.LRScheduler]=None,
+> lr_scheduler_kwargs:Optional[Dict]=None,
+> dataloader_kwargs=None, **trainer_kwargs)
+> ```
+
+*Hooks to be used in LightningModule.*
+
diff --git a/neuralforecast/common.model_checks.html.mdx b/neuralforecast/common.model_checks.html.mdx
new file mode 100644
index 00000000..f8e5476e
--- /dev/null
+++ b/neuralforecast/common.model_checks.html.mdx
@@ -0,0 +1,37 @@
+---
+output-file: common.model_checks.html
+title: 1. Checks for models
+---
+
+
+This file provides a set of unit tests for all models
+
+------------------------------------------------------------------------
+
+### check_model
+
+> ``` text
+> check_model (model_class, checks=['losses', 'airpassengers'])
+> ```
+
+*Check model with various tests. Options for checks are: “losses”:
+test the model against all loss functions “airpassengers”: test the
+model against the airpassengers dataset for forecasting and
+cross-validation *
+
+------------------------------------------------------------------------
+
+### check_airpassengers
+
+> ``` text
+> check_airpassengers (model_class)
+> ```
+
+------------------------------------------------------------------------
+
+### check_loss_functions
+
+> ``` text
+> check_loss_functions (model_class)
+> ```
+
diff --git a/neuralforecast/common.modules.html.mdx b/neuralforecast/common.modules.html.mdx
new file mode 100644
index 00000000..f9ebc3eb
--- /dev/null
+++ b/neuralforecast/common.modules.html.mdx
@@ -0,0 +1,685 @@
+---
+output-file: common.modules.html
+title: NN Modules
+---
+
+
+## 1. MLP
+
+Multi-Layer Perceptron
+
+------------------------------------------------------------------------
+
+source
+
+### MLP
+
+> ``` text
+> MLP (in_features, out_features, activation, hidden_size, num_layers,
+> dropout)
+> ```
+
+\*Multi-Layer Perceptron Class
+
+**Parameters:** `in_features`: int, dimension of input.
+`out_features`: int, dimension of output. `activation`: str,
+activation function to use. `hidden_size`: int, dimension of hidden
+layers. `num_layers`: int, number of hidden layers. `dropout`:
+float, dropout rate. \*
+
+## 2. Temporal Convolutions
+
+For long time in deep learning, sequence modelling was synonymous with
+recurrent networks, yet several papers have shown that simple
+convolutional architectures can outperform canonical recurrent networks
+like LSTMs by demonstrating longer effective memory.
+
+**References** -[van den Oord, A., Dieleman, S., Zen, H., Simonyan,
+K., Vinyals, O., Graves, A., Kalchbrenner, N., Senior, A. W., &
+Kavukcuoglu, K. (2016). Wavenet: A generative model for raw audio.
+Computing Research Repository, abs/1609.03499. URL:
+http://arxiv.org/abs/1609.03499.
+arXiv:1609.03499.](https://arxiv.org/abs/1609.03499) -[Shaojie Bai,
+Zico Kolter, Vladlen Koltun. (2018). An Empirical Evaluation of Generic
+Convolutional and Recurrent Networks for Sequence Modeling. Computing
+Research Repository, abs/1803.01271. URL:
+https://arxiv.org/abs/1803.01271.](https://arxiv.org/abs/1803.01271)
+
+------------------------------------------------------------------------
+
+### Chomp1d
+
+> ``` text
+> Chomp1d (horizon)
+> ```
+
+\*Chomp1d
+
+Receives `x` input of dim \[N,C,T\], and trims it so that only ‘time
+available’ information is used. Used by one dimensional causal
+convolutions `CausalConv1d`.
+
+**Parameters:** `horizon`: int, length of outsample values to
+skip.\*
+
+------------------------------------------------------------------------
+
+### CausalConv1d
+
+> ``` text
+> CausalConv1d (in_channels, out_channels, kernel_size, padding, dilation,
+> activation, stride:int=1)
+> ```
+
+\*Causal Convolution 1d
+
+Receives `x` input of dim \[N,C_in,T\], and computes a causal
+convolution in the time dimension. Skipping the H steps of the forecast
+horizon, through its dilation. Consider a batch of one element, the
+dilated convolution operation on the $t$ time step is defined:
+
+$\mathrm{Conv1D}(\mathbf{x},\mathbf{w})(t) = (\mathbf{x}_{[*d]} \mathbf{w})(t) = \sum^{K}_{k=1} w_{k} \mathbf{x}_{t-dk}$
+
+where $d$ is the dilation factor, $K$ is the kernel size, $t-dk$ is the
+index of the considered past observation. The dilation effectively
+applies a filter with skip connections. If $d=1$ one recovers a normal
+convolution.
+
+**Parameters:** `in_channels`: int, dimension of `x` input’s initial
+channels. `out_channels`: int, dimension of `x` outputs’s
+channels. `activation`: str, identifying activations from PyTorch
+activations. select from ‘ReLU’,‘Softplus’,‘Tanh’,‘SELU’,
+‘LeakyReLU’,‘PReLU’,‘Sigmoid’. `padding`: int, number of zero
+padding used to the left. `kernel_size`: int, convolution’s kernel
+size. `dilation`: int, dilation skip connections.
+
+**Returns:** `x`: tensor, torch tensor of dim \[N,C_out,T\]
+activation(conv1d(inputs, kernel) + bias). \*
+
+------------------------------------------------------------------------
+
+### TemporalConvolutionEncoder
+
+> ``` text
+> TemporalConvolutionEncoder (in_channels, out_channels, kernel_size,
+> dilations, activation:str='ReLU')
+> ```
+
+\*Temporal Convolution Encoder
+
+Receives `x` input of dim \[N,T,C_in\], permutes it to \[N,C_in,T\]
+applies a deep stack of exponentially dilated causal convolutions. The
+exponentially increasing dilations of the convolutions allow for the
+creation of weighted averages of exponentially large long-term memory.
+
+**Parameters:** `in_channels`: int, dimension of `x` input’s initial
+channels. `out_channels`: int, dimension of `x` outputs’s
+channels. `kernel_size`: int, size of the convolving kernel.
+`dilations`: int list, controls the temporal spacing between the kernel
+points. `activation`: str, identifying activations from PyTorch
+activations. select from ‘ReLU’,‘Softplus’,‘Tanh’,‘SELU’,
+‘LeakyReLU’,‘PReLU’,‘Sigmoid’.
+
+**Returns:** `x`: tensor, torch tensor of dim \[N,T,C_out\]. \*
+
+## 3. Transformers
+
+**References** - [Haoyi Zhou, Shanghang Zhang, Jieqi Peng, Shuai
+Zhang, Jianxin Li, Hui Xiong, Wancai Zhang. “Informer: Beyond Efficient
+Transformer for Long Sequence Time-Series
+Forecasting”](https://arxiv.org/abs/2012.07436) - [Haixu Wu, Jiehui
+Xu, Jianmin Wang, Mingsheng Long.](https://arxiv.org/abs/2106.13008)
+
+------------------------------------------------------------------------
+
+### TransEncoder
+
+> ``` text
+> TransEncoder (attn_layers, conv_layers=None, norm_layer=None)
+> ```
+
+\*Base class for all neural network modules.
+
+Your models should also subclass this class.
+
+Modules can also contain other Modules, allowing them to be nested in a
+tree structure. You can assign the submodules as regular attributes::
+
+``` text
+import torch.nn as nn
+import torch.nn.functional as F
+
+class Model(nn.Module):
+ def __init__(self) -> None:
+ super().__init__()
+ self.conv1 = nn.Conv2d(1, 20, 5)
+ self.conv2 = nn.Conv2d(20, 20, 5)
+
+ def forward(self, x):
+ x = F.relu(self.conv1(x))
+ return F.relu(self.conv2(x))
+```
+
+Submodules assigned in this way will be registered, and will also have
+their parameters converted when you call :meth:`to`, etc.
+
+.. note:: As per the example above, an `__init__()` call to the parent
+class must be made before assignment on the child.
+
+:ivar training: Boolean represents whether this module is in training or
+evaluation mode. :vartype training: bool\*
+
+------------------------------------------------------------------------
+
+### TransEncoderLayer
+
+> ``` text
+> TransEncoderLayer (attention, hidden_size, conv_hidden_size=None,
+> dropout=0.1, activation='relu')
+> ```
+
+\*Base class for all neural network modules.
+
+Your models should also subclass this class.
+
+Modules can also contain other Modules, allowing them to be nested in a
+tree structure. You can assign the submodules as regular attributes::
+
+``` text
+import torch.nn as nn
+import torch.nn.functional as F
+
+class Model(nn.Module):
+ def __init__(self) -> None:
+ super().__init__()
+ self.conv1 = nn.Conv2d(1, 20, 5)
+ self.conv2 = nn.Conv2d(20, 20, 5)
+
+ def forward(self, x):
+ x = F.relu(self.conv1(x))
+ return F.relu(self.conv2(x))
+```
+
+Submodules assigned in this way will be registered, and will also have
+their parameters converted when you call :meth:`to`, etc.
+
+.. note:: As per the example above, an `__init__()` call to the parent
+class must be made before assignment on the child.
+
+:ivar training: Boolean represents whether this module is in training or
+evaluation mode. :vartype training: bool\*
+
+------------------------------------------------------------------------
+
+### TransDecoder
+
+> ``` text
+> TransDecoder (layers, norm_layer=None, projection=None)
+> ```
+
+\*Base class for all neural network modules.
+
+Your models should also subclass this class.
+
+Modules can also contain other Modules, allowing them to be nested in a
+tree structure. You can assign the submodules as regular attributes::
+
+``` text
+import torch.nn as nn
+import torch.nn.functional as F
+
+class Model(nn.Module):
+ def __init__(self) -> None:
+ super().__init__()
+ self.conv1 = nn.Conv2d(1, 20, 5)
+ self.conv2 = nn.Conv2d(20, 20, 5)
+
+ def forward(self, x):
+ x = F.relu(self.conv1(x))
+ return F.relu(self.conv2(x))
+```
+
+Submodules assigned in this way will be registered, and will also have
+their parameters converted when you call :meth:`to`, etc.
+
+.. note:: As per the example above, an `__init__()` call to the parent
+class must be made before assignment on the child.
+
+:ivar training: Boolean represents whether this module is in training or
+evaluation mode. :vartype training: bool\*
+
+------------------------------------------------------------------------
+
+### TransDecoderLayer
+
+> ``` text
+> TransDecoderLayer (self_attention, cross_attention, hidden_size,
+> conv_hidden_size=None, dropout=0.1, activation='relu')
+> ```
+
+\*Base class for all neural network modules.
+
+Your models should also subclass this class.
+
+Modules can also contain other Modules, allowing them to be nested in a
+tree structure. You can assign the submodules as regular attributes::
+
+``` text
+import torch.nn as nn
+import torch.nn.functional as F
+
+class Model(nn.Module):
+ def __init__(self) -> None:
+ super().__init__()
+ self.conv1 = nn.Conv2d(1, 20, 5)
+ self.conv2 = nn.Conv2d(20, 20, 5)
+
+ def forward(self, x):
+ x = F.relu(self.conv1(x))
+ return F.relu(self.conv2(x))
+```
+
+Submodules assigned in this way will be registered, and will also have
+their parameters converted when you call :meth:`to`, etc.
+
+.. note:: As per the example above, an `__init__()` call to the parent
+class must be made before assignment on the child.
+
+:ivar training: Boolean represents whether this module is in training or
+evaluation mode. :vartype training: bool\*
+
+------------------------------------------------------------------------
+
+### AttentionLayer
+
+> ``` text
+> AttentionLayer (attention, hidden_size, n_heads, d_keys=None,
+> d_values=None)
+> ```
+
+\*Base class for all neural network modules.
+
+Your models should also subclass this class.
+
+Modules can also contain other Modules, allowing them to be nested in a
+tree structure. You can assign the submodules as regular attributes::
+
+``` text
+import torch.nn as nn
+import torch.nn.functional as F
+
+class Model(nn.Module):
+ def __init__(self) -> None:
+ super().__init__()
+ self.conv1 = nn.Conv2d(1, 20, 5)
+ self.conv2 = nn.Conv2d(20, 20, 5)
+
+ def forward(self, x):
+ x = F.relu(self.conv1(x))
+ return F.relu(self.conv2(x))
+```
+
+Submodules assigned in this way will be registered, and will also have
+their parameters converted when you call :meth:`to`, etc.
+
+.. note:: As per the example above, an `__init__()` call to the parent
+class must be made before assignment on the child.
+
+:ivar training: Boolean represents whether this module is in training or
+evaluation mode. :vartype training: bool\*
+
+------------------------------------------------------------------------
+
+### FullAttention
+
+> ``` text
+> FullAttention (mask_flag=True, factor=5, scale=None,
+> attention_dropout=0.1, output_attention=False)
+> ```
+
+\*Base class for all neural network modules.
+
+Your models should also subclass this class.
+
+Modules can also contain other Modules, allowing them to be nested in a
+tree structure. You can assign the submodules as regular attributes::
+
+``` text
+import torch.nn as nn
+import torch.nn.functional as F
+
+class Model(nn.Module):
+ def __init__(self) -> None:
+ super().__init__()
+ self.conv1 = nn.Conv2d(1, 20, 5)
+ self.conv2 = nn.Conv2d(20, 20, 5)
+
+ def forward(self, x):
+ x = F.relu(self.conv1(x))
+ return F.relu(self.conv2(x))
+```
+
+Submodules assigned in this way will be registered, and will also have
+their parameters converted when you call :meth:`to`, etc.
+
+.. note:: As per the example above, an `__init__()` call to the parent
+class must be made before assignment on the child.
+
+:ivar training: Boolean represents whether this module is in training or
+evaluation mode. :vartype training: bool\*
+
+------------------------------------------------------------------------
+
+### TriangularCausalMask
+
+> ``` text
+> TriangularCausalMask (B, L, device='cpu')
+> ```
+
+*TriangularCausalMask*
+
+------------------------------------------------------------------------
+
+source
+
+### DataEmbedding_inverted
+
+> ``` text
+> DataEmbedding_inverted (c_in, hidden_size, dropout=0.1)
+> ```
+
+*DataEmbedding_inverted*
+
+------------------------------------------------------------------------
+
+### DataEmbedding
+
+> ``` text
+> DataEmbedding (c_in, exog_input_size, hidden_size, pos_embedding=True,
+> dropout=0.1)
+> ```
+
+\*Base class for all neural network modules.
+
+Your models should also subclass this class.
+
+Modules can also contain other Modules, allowing them to be nested in a
+tree structure. You can assign the submodules as regular attributes::
+
+``` text
+import torch.nn as nn
+import torch.nn.functional as F
+
+class Model(nn.Module):
+ def __init__(self) -> None:
+ super().__init__()
+ self.conv1 = nn.Conv2d(1, 20, 5)
+ self.conv2 = nn.Conv2d(20, 20, 5)
+
+ def forward(self, x):
+ x = F.relu(self.conv1(x))
+ return F.relu(self.conv2(x))
+```
+
+Submodules assigned in this way will be registered, and will also have
+their parameters converted when you call :meth:`to`, etc.
+
+.. note:: As per the example above, an `__init__()` call to the parent
+class must be made before assignment on the child.
+
+:ivar training: Boolean represents whether this module is in training or
+evaluation mode. :vartype training: bool\*
+
+------------------------------------------------------------------------
+
+### TemporalEmbedding
+
+> ``` text
+> TemporalEmbedding (d_model, embed_type='fixed', freq='h')
+> ```
+
+\*Base class for all neural network modules.
+
+Your models should also subclass this class.
+
+Modules can also contain other Modules, allowing them to be nested in a
+tree structure. You can assign the submodules as regular attributes::
+
+``` text
+import torch.nn as nn
+import torch.nn.functional as F
+
+class Model(nn.Module):
+ def __init__(self) -> None:
+ super().__init__()
+ self.conv1 = nn.Conv2d(1, 20, 5)
+ self.conv2 = nn.Conv2d(20, 20, 5)
+
+ def forward(self, x):
+ x = F.relu(self.conv1(x))
+ return F.relu(self.conv2(x))
+```
+
+Submodules assigned in this way will be registered, and will also have
+their parameters converted when you call :meth:`to`, etc.
+
+.. note:: As per the example above, an `__init__()` call to the parent
+class must be made before assignment on the child.
+
+:ivar training: Boolean represents whether this module is in training or
+evaluation mode. :vartype training: bool\*
+
+------------------------------------------------------------------------
+
+### FixedEmbedding
+
+> ``` text
+> FixedEmbedding (c_in, d_model)
+> ```
+
+\*Base class for all neural network modules.
+
+Your models should also subclass this class.
+
+Modules can also contain other Modules, allowing them to be nested in a
+tree structure. You can assign the submodules as regular attributes::
+
+``` text
+import torch.nn as nn
+import torch.nn.functional as F
+
+class Model(nn.Module):
+ def __init__(self) -> None:
+ super().__init__()
+ self.conv1 = nn.Conv2d(1, 20, 5)
+ self.conv2 = nn.Conv2d(20, 20, 5)
+
+ def forward(self, x):
+ x = F.relu(self.conv1(x))
+ return F.relu(self.conv2(x))
+```
+
+Submodules assigned in this way will be registered, and will also have
+their parameters converted when you call :meth:`to`, etc.
+
+.. note:: As per the example above, an `__init__()` call to the parent
+class must be made before assignment on the child.
+
+:ivar training: Boolean represents whether this module is in training or
+evaluation mode. :vartype training: bool\*
+
+------------------------------------------------------------------------
+
+### TimeFeatureEmbedding
+
+> ``` text
+> TimeFeatureEmbedding (input_size, hidden_size)
+> ```
+
+\*Base class for all neural network modules.
+
+Your models should also subclass this class.
+
+Modules can also contain other Modules, allowing them to be nested in a
+tree structure. You can assign the submodules as regular attributes::
+
+``` text
+import torch.nn as nn
+import torch.nn.functional as F
+
+class Model(nn.Module):
+ def __init__(self) -> None:
+ super().__init__()
+ self.conv1 = nn.Conv2d(1, 20, 5)
+ self.conv2 = nn.Conv2d(20, 20, 5)
+
+ def forward(self, x):
+ x = F.relu(self.conv1(x))
+ return F.relu(self.conv2(x))
+```
+
+Submodules assigned in this way will be registered, and will also have
+their parameters converted when you call :meth:`to`, etc.
+
+.. note:: As per the example above, an `__init__()` call to the parent
+class must be made before assignment on the child.
+
+:ivar training: Boolean represents whether this module is in training or
+evaluation mode. :vartype training: bool\*
+
+------------------------------------------------------------------------
+
+source
+
+### TokenEmbedding
+
+> ``` text
+> TokenEmbedding (c_in, hidden_size)
+> ```
+
+\*Base class for all neural network modules.
+
+Your models should also subclass this class.
+
+Modules can also contain other Modules, allowing them to be nested in a
+tree structure. You can assign the submodules as regular attributes::
+
+``` text
+import torch.nn as nn
+import torch.nn.functional as F
+
+class Model(nn.Module):
+ def __init__(self) -> None:
+ super().__init__()
+ self.conv1 = nn.Conv2d(1, 20, 5)
+ self.conv2 = nn.Conv2d(20, 20, 5)
+
+ def forward(self, x):
+ x = F.relu(self.conv1(x))
+ return F.relu(self.conv2(x))
+```
+
+Submodules assigned in this way will be registered, and will also have
+their parameters converted when you call :meth:`to`, etc.
+
+.. note:: As per the example above, an `__init__()` call to the parent
+class must be made before assignment on the child.
+
+:ivar training: Boolean represents whether this module is in training or
+evaluation mode. :vartype training: bool\*
+
+------------------------------------------------------------------------
+
+### PositionalEmbedding
+
+> ``` text
+> PositionalEmbedding (hidden_size, max_len=5000)
+> ```
+
+\*Base class for all neural network modules.
+
+Your models should also subclass this class.
+
+Modules can also contain other Modules, allowing them to be nested in a
+tree structure. You can assign the submodules as regular attributes::
+
+``` text
+import torch.nn as nn
+import torch.nn.functional as F
+
+class Model(nn.Module):
+ def __init__(self) -> None:
+ super().__init__()
+ self.conv1 = nn.Conv2d(1, 20, 5)
+ self.conv2 = nn.Conv2d(20, 20, 5)
+
+ def forward(self, x):
+ x = F.relu(self.conv1(x))
+ return F.relu(self.conv2(x))
+```
+
+Submodules assigned in this way will be registered, and will also have
+their parameters converted when you call :meth:`to`, etc.
+
+.. note:: As per the example above, an `__init__()` call to the parent
+class must be made before assignment on the child.
+
+:ivar training: Boolean represents whether this module is in training or
+evaluation mode. :vartype training: bool\*
+
+------------------------------------------------------------------------
+
+source
+
+### SeriesDecomp
+
+> ``` text
+> SeriesDecomp (kernel_size)
+> ```
+
+*Series decomposition block*
+
+------------------------------------------------------------------------
+
+source
+
+### MovingAvg
+
+> ``` text
+> MovingAvg (kernel_size, stride)
+> ```
+
+*Moving average block to highlight the trend of time series*
+
+------------------------------------------------------------------------
+
+### RevIN
+
+> ``` text
+> RevIN (num_features:int, eps=1e-05, affine=False, subtract_last=False,
+> non_norm=False)
+> ```
+
+*RevIN (Reversible-Instance-Normalization)*
+
+------------------------------------------------------------------------
+
+### RevINMultivariate
+
+> ``` text
+> RevINMultivariate (num_features:int, eps=1e-05, affine=False,
+> subtract_last=False, non_norm=False)
+> ```
+
+*ReversibleInstanceNorm1d for Multivariate models*
+
diff --git a/neuralforecast/common.scalers.html.mdx b/neuralforecast/common.scalers.html.mdx
new file mode 100644
index 00000000..787a5b1f
--- /dev/null
+++ b/neuralforecast/common.scalers.html.mdx
@@ -0,0 +1,390 @@
+---
+description: >-
+ Temporal normalization has proven to be essential in neural forecasting tasks,
+ as it enables network's non-linearities to express themselves. Forecasting
+ scaling methods take particular interest in the temporal dimension where most
+ of the variance dwells, contrary to other deep learning techniques like
+ `BatchNorm` that normalizes across batch and temporal dimensions, and
+ `LayerNorm` that normalizes across the feature dimension. Currently we support
+ the following techniques: `std`, `median`, `norm`, `norm1`, `invariant`,
+ `revin`.
+output-file: common.scalers.html
+title: TemporalNorm
+---
+
+
+## References
+
+- [Kin G. Olivares, David Luo, Cristian Challu, Stefania La Vattiata,
+ Max Mergenthaler, Artur Dubrawski (2023). “HINT: Hierarchical
+ Mixture Networks For Coherent Probabilistic Forecasting”. Neural
+ Information Processing Systems, submitted. Working Paper version
+ available at arxiv.](https://arxiv.org/abs/2305.07089)
+- [Taesung Kim and Jinhee Kim and Yunwon Tae and Cheonbok Park and
+ Jang-Ho Choi and Jaegul Choo. “Reversible Instance Normalization for
+ Accurate Time-Series Forecasting against Distribution Shift”. ICLR
+ 2022.](https://openreview.net/pdf?id=cGDAkQo1C0p)
+- [David Salinas, Valentin Flunkert, Jan Gasthaus, Tim Januschowski
+ (2020). “DeepAR: Probabilistic forecasting with autoregressive
+ recurrent networks”. International Journal of
+ Forecasting.](https://www.sciencedirect.com/science/article/pii/S0169207019301888)
+
+
+
+Figure 1. Illustration of temporal
+normalization (left), layer normalization (center) and batch
+normalization (right). The entries in green show the components used to
+compute the normalizing statistics.
+
+
+# 1. Auxiliary Functions
+
+------------------------------------------------------------------------
+
+### masked_median
+
+> ``` text
+> masked_median (x, mask, dim=-1, keepdim=True)
+> ```
+
+\*Masked Median
+
+Compute the median of tensor `x` along dim, ignoring values where `mask`
+is False. `x` and `mask` need to be broadcastable.
+
+**Parameters:** `x`: torch.Tensor to compute median of along `dim`
+dimension. `mask`: torch Tensor bool with same shape as `x`, where
+`x` is valid and False where `x` should be masked. Mask should not be
+all False in any column of dimension dim to avoid NaNs from zero
+division. `dim` (int, optional): Dimension to take median of.
+Defaults to -1. `keepdim` (bool, optional): Keep dimension of `x` or
+not. Defaults to True.
+
+**Returns:** `x_median`: torch.Tensor with normalized values.\*
+
+------------------------------------------------------------------------
+
+### masked_mean
+
+> ``` text
+> masked_mean (x, mask, dim=-1, keepdim=True)
+> ```
+
+\*Masked Mean
+
+Compute the mean of tensor `x` along dimension, ignoring values where
+`mask` is False. `x` and `mask` need to be broadcastable.
+
+**Parameters:** `x`: torch.Tensor to compute mean of along `dim`
+dimension. `mask`: torch Tensor bool with same shape as `x`, where
+`x` is valid and False where `x` should be masked. Mask should not be
+all False in any column of dimension dim to avoid NaNs from zero
+division. `dim` (int, optional): Dimension to take mean of. Defaults
+to -1. `keepdim` (bool, optional): Keep dimension of `x` or not.
+Defaults to True.
+
+**Returns:** `x_mean`: torch.Tensor with normalized values.\*
+
+# 2. Scalers
+
+------------------------------------------------------------------------
+
+### minmax_statistics
+
+> ``` text
+> minmax_statistics (x, mask, eps=1e-06, dim=-1)
+> ```
+
+\*MinMax Scaler
+
+Standardizes temporal features by ensuring its range dweels between
+\[0,1\] range. This transformation is often used as an alternative to
+the standard scaler. The scaled features are obtained as:
+
+$$
+
+\mathbf{z} = (\mathbf{x}_{[B,T,C]}-\mathrm{min}({\mathbf{x}})_{[B,1,C]})/
+ (\mathrm{max}({\mathbf{x}})_{[B,1,C]}- \mathrm{min}({\mathbf{x}})_{[B,1,C]})
+
+$$
+
+**Parameters:** `x`: torch.Tensor input tensor. `mask`: torch
+Tensor bool, same dimension as `x`, indicates where `x` is valid and
+False where `x` should be masked. Mask should not be all False in any
+column of dimension dim to avoid NaNs from zero division. `eps`
+(float, optional): Small value to avoid division by zero. Defaults to
+1e-6. `dim` (int, optional): Dimension over to compute min and max.
+Defaults to -1.
+
+**Returns:** `z`: torch.Tensor same shape as `x`, except scaled.\*
+
+------------------------------------------------------------------------
+
+### minmax1_statistics
+
+> ``` text
+> minmax1_statistics (x, mask, eps=1e-06, dim=-1)
+> ```
+
+\*MinMax1 Scaler
+
+Standardizes temporal features by ensuring its range dweels between
+\[-1,1\] range. This transformation is often used as an alternative to
+the standard scaler or classic Min Max Scaler. The scaled features are
+obtained as:
+
+$$\mathbf{z} = 2 (\mathbf{x}_{[B,T,C]}-\mathrm{min}({\mathbf{x}})_{[B,1,C]})/ (\mathrm{max}({\mathbf{x}})_{[B,1,C]}- \mathrm{min}({\mathbf{x}})_{[B,1,C]})-1$$
+
+**Parameters:** `x`: torch.Tensor input tensor. `mask`: torch
+Tensor bool, same dimension as `x`, indicates where `x` is valid and
+False where `x` should be masked. Mask should not be all False in any
+column of dimension dim to avoid NaNs from zero division. `eps`
+(float, optional): Small value to avoid division by zero. Defaults to
+1e-6. `dim` (int, optional): Dimension over to compute min and max.
+Defaults to -1.
+
+**Returns:** `z`: torch.Tensor same shape as `x`, except scaled.\*
+
+------------------------------------------------------------------------
+
+### std_statistics
+
+> ``` text
+> std_statistics (x, mask, dim=-1, eps=1e-06)
+> ```
+
+\*Standard Scaler
+
+Standardizes features by removing the mean and scaling to unit variance
+along the `dim` dimension.
+
+For example, for `base_windows` models, the scaled features are obtained
+as (with dim=1):
+
+$$\mathbf{z} = (\mathbf{x}_{[B,T,C]}-\bar{\mathbf{x}}_{[B,1,C]})/\hat{\sigma}_{[B,1,C]}$$
+
+**Parameters:** `x`: torch.Tensor. `mask`: torch Tensor bool,
+same dimension as `x`, indicates where `x` is valid and False where `x`
+should be masked. Mask should not be all False in any column of
+dimension dim to avoid NaNs from zero division. `eps` (float,
+optional): Small value to avoid division by zero. Defaults to 1e-6.
+`dim` (int, optional): Dimension over to compute mean and std. Defaults
+to -1.
+
+**Returns:** `z`: torch.Tensor same shape as `x`, except scaled.\*
+
+------------------------------------------------------------------------
+
+### robust_statistics
+
+> ``` text
+> robust_statistics (x, mask, dim=-1, eps=1e-06)
+> ```
+
+\*Robust Median Scaler
+
+Standardizes features by removing the median and scaling with the mean
+absolute deviation (mad) a robust estimator of variance. This scaler is
+particularly useful with noisy data where outliers can heavily influence
+the sample mean / variance in a negative way. In these scenarios the
+median and amd give better results.
+
+For example, for `base_windows` models, the scaled features are obtained
+as (with dim=1):
+
+$$\mathbf{z} = (\mathbf{x}_{[B,T,C]}-\textrm{median}(\mathbf{x})_{[B,1,C]})/\textrm{mad}(\mathbf{x})_{[B,1,C]}$$
+
+$$\textrm{mad}(\mathbf{x}) = \frac{1}{N} \sum_{}|\mathbf{x} - \mathrm{median}(x)|$$
+
+**Parameters:** `x`: torch.Tensor input tensor. `mask`: torch
+Tensor bool, same dimension as `x`, indicates where `x` is valid and
+False where `x` should be masked. Mask should not be all False in any
+column of dimension dim to avoid NaNs from zero division. `eps`
+(float, optional): Small value to avoid division by zero. Defaults to
+1e-6. `dim` (int, optional): Dimension over to compute median and
+mad. Defaults to -1.
+
+**Returns:** `z`: torch.Tensor same shape as `x`, except scaled.\*
+
+------------------------------------------------------------------------
+
+### invariant_statistics
+
+> ``` text
+> invariant_statistics (x, mask, dim=-1, eps=1e-06)
+> ```
+
+\*Invariant Median Scaler
+
+Standardizes features by removing the median and scaling with the mean
+absolute deviation (mad) a robust estimator of variance. Aditionally it
+complements the transformation with the arcsinh transformation.
+
+For example, for `base_windows` models, the scaled features are obtained
+as (with dim=1):
+
+$$\mathbf{z} = (\mathbf{x}_{[B,T,C]}-\textrm{median}(\mathbf{x})_{[B,1,C]})/\textrm{mad}(\mathbf{x})_{[B,1,C]}$$
+
+$$\mathbf{z} = \textrm{arcsinh}(\mathbf{z})$$
+
+**Parameters:** `x`: torch.Tensor input tensor. `mask`: torch
+Tensor bool, same dimension as `x`, indicates where `x` is valid and
+False where `x` should be masked. Mask should not be all False in any
+column of dimension dim to avoid NaNs from zero division. `eps`
+(float, optional): Small value to avoid division by zero. Defaults to
+1e-6. `dim` (int, optional): Dimension over to compute median and
+mad. Defaults to -1.
+
+**Returns:** `z`: torch.Tensor same shape as `x`, except scaled.\*
+
+------------------------------------------------------------------------
+
+### identity_statistics
+
+> ``` text
+> identity_statistics (x, mask, dim=-1, eps=1e-06)
+> ```
+
+\*Identity Scaler
+
+A placeholder identity scaler, that is argument insensitive.
+
+**Parameters:** `x`: torch.Tensor input tensor. `mask`: torch
+Tensor bool, same dimension as `x`, indicates where `x` is valid and
+False where `x` should be masked. Mask should not be all False in any
+column of dimension dim to avoid NaNs from zero division. `eps`
+(float, optional): Small value to avoid division by zero. Defaults to
+1e-6. `dim` (int, optional): Dimension over to compute median and
+mad. Defaults to -1.
+
+**Returns:** `x`: original torch.Tensor `x`.\*
+
+# 3. TemporalNorm Module
+
+------------------------------------------------------------------------
+
+### TemporalNorm
+
+> ``` text
+> TemporalNorm (scaler_type='robust', dim=-1, eps=1e-06, num_features=None)
+> ```
+
+\*Temporal Normalization
+
+Standardization of the features is a common requirement for many machine
+learning estimators, and it is commonly achieved by removing the level
+and scaling its variance. The `TemporalNorm` module applies temporal
+normalization over the batch of inputs as defined by the type of scaler.
+
+$$\mathbf{z}_{[B,T,C]} = \textrm{Scaler}(\mathbf{x}_{[B,T,C]})$$
+
+If `scaler_type` is `revin` learnable normalization parameters are added
+on top of the usual normalization technique, the parameters are learned
+through scale decouple global skip connections. The technique is
+available for point and probabilistic outputs.
+
+$$\mathbf{\hat{z}}_{[B,T,C]} = \boldsymbol{\hat{\gamma}}_{[1,1,C]} \mathbf{z}_{[B,T,C]} +\boldsymbol{\hat{\beta}}_{[1,1,C]}$$
+
+**Parameters:** `scaler_type`: str, defines the type of scaler used
+by TemporalNorm. Available \[`identity`, `standard`, `robust`, `minmax`,
+`minmax1`, `invariant`, `revin`\]. `dim` (int, optional): Dimension
+over to compute scale and shift. Defaults to -1. `eps` (float,
+optional): Small value to avoid division by zero. Defaults to 1e-6.
+`num_features`: int=None, for RevIN-like learnable affine parameters
+initialization.
+
+**References** - [Kin G. Olivares, David Luo, Cristian Challu,
+Stefania La Vattiata, Max Mergenthaler, Artur Dubrawski (2023). “HINT:
+Hierarchical Mixture Networks For Coherent Probabilistic Forecasting”.
+Neural Information Processing Systems, submitted. Working Paper version
+available at arxiv.](https://arxiv.org/abs/2305.07089) \*
+
+------------------------------------------------------------------------
+
+### TemporalNorm.transform
+
+> ``` text
+> TemporalNorm.transform (x, mask)
+> ```
+
+\*Center and scale the data.
+
+**Parameters:** `x`: torch.Tensor shape \[batch, time,
+channels\]. `mask`: torch Tensor bool, shape \[batch, time\] where
+`x` is valid and False where `x` should be masked. Mask should not be
+all False in any column of dimension dim to avoid NaNs from zero
+division.
+
+**Returns:** `z`: torch.Tensor same shape as `x`, except scaled.\*
+
+------------------------------------------------------------------------
+
+### TemporalNorm.inverse_transform
+
+> ``` text
+> TemporalNorm.inverse_transform (z, x_shift=None, x_scale=None)
+> ```
+
+\*Scale back the data to the original representation.
+
+**Parameters:** `z`: torch.Tensor shape \[batch, time, channels\],
+scaled.
+
+**Returns:** `x`: torch.Tensor original data.\*
+
+# Example
+
+
+```python
+import numpy as np
+```
+
+
+```python
+# Declare synthetic batch to normalize
+x1 = 10**0 * np.arange(36)[:, None]
+x2 = 10**1 * np.arange(36)[:, None]
+
+np_x = np.concatenate([x1, x2], axis=1)
+np_x = np.repeat(np_x[None, :,:], repeats=2, axis=0)
+np_x[0,:,:] = np_x[0,:,:] + 100
+
+np_mask = np.ones(np_x.shape)
+np_mask[:, -12:, :] = 0
+
+print(f'x.shape [batch, time, features]={np_x.shape}')
+print(f'mask.shape [batch, time, features]={np_mask.shape}')
+```
+
+
+```python
+# Validate scalers
+x = 1.0*torch.tensor(np_x)
+mask = torch.tensor(np_mask)
+scaler = TemporalNorm(scaler_type='standard', dim=1)
+x_scaled = scaler.transform(x=x, mask=mask)
+x_recovered = scaler.inverse_transform(x_scaled)
+
+plt.plot(x[0,:,0], label='x1', color='#78ACA8')
+plt.plot(x[0,:,1], label='x2', color='#E3A39A')
+plt.title('Before TemporalNorm')
+plt.xlabel('Time')
+plt.legend()
+plt.show()
+
+plt.plot(x_scaled[0,:,0], label='x1', color='#78ACA8')
+plt.plot(x_scaled[0,:,1]+0.1, label='x2+0.1', color='#E3A39A')
+plt.title(f'TemporalNorm \'{scaler.scaler_type}\' ')
+plt.xlabel('Time')
+plt.legend()
+plt.show()
+
+plt.plot(x_recovered[0,:,0], label='x1', color='#78ACA8')
+plt.plot(x_recovered[0,:,1], label='x2', color='#E3A39A')
+plt.title('Recovered')
+plt.xlabel('Time')
+plt.legend()
+plt.show()
+```
+
diff --git a/neuralforecast/compat.mdx b/neuralforecast/compat.mdx
new file mode 100644
index 00000000..8b137891
--- /dev/null
+++ b/neuralforecast/compat.mdx
@@ -0,0 +1 @@
+
diff --git a/neuralforecast/core.html.mdx b/neuralforecast/core.html.mdx
new file mode 100644
index 00000000..535b7266
--- /dev/null
+++ b/neuralforecast/core.html.mdx
@@ -0,0 +1,384 @@
+---
+description: >-
+ NeuralForecast contains two main components, PyTorch implementations deep
+ learning predictive models, as well as parallelization and distributed
+ computation utilities. The first component comprises low-level PyTorch model
+ estimator classes like `models.NBEATS` and `models.RNN`. The second component
+ is a high-level `core.NeuralForecast` wrapper class that operates with sets of
+ time series data stored in pandas DataFrames.
+output-file: core.html
+title: Core
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### NeuralForecast
+
+> ``` text
+> NeuralForecast (models:List[Any], freq:Union[str,int],
+> local_scaler_type:Optional[str]=None)
+> ```
+
+*The `core.StatsForecast` class allows you to efficiently fit multiple
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+models for large sets of time series. It operates with pandas DataFrame
+`df` that identifies series and datestamps with the `unique_id` and `ds`
+columns. The `y` column denotes the target time series variable.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| models | List | | Instantiated `neuralforecast.models` see [collection here](https://nixtla.github.io/neuralforecast/models.html). |
+| freq | Union | | Frequency of the data. Must be a valid pandas or polars offset alias, or an integer. |
+| local_scaler_type | Optional | None | Scaler to apply per-serie to all features before fitting, which is inverted after predicting. Can be ‘standard’, ‘robust’, ‘robust-iqr’, ‘minmax’ or ‘boxcox’ |
+| **Returns** | **NeuralForecast** | | **Returns instantiated [`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast) class.** |
+
+------------------------------------------------------------------------
+
+source
+
+### NeuralForecast.fit
+
+> ``` text
+> NeuralForecast.fit (df:Union[pandas.core.frame.DataFrame,polars.dataframe
+> .frame.DataFrame,neuralforecast.compat.SparkDataFrame
+> ,Sequence[str],NoneType]=None, static_df:Union[pandas
+> .core.frame.DataFrame,polars.dataframe.frame.DataFram
+> e,neuralforecast.compat.SparkDataFrame,NoneType]=None
+> , val_size:Optional[int]=0,
+> use_init_models:bool=False, verbose:bool=False,
+> id_col:str='unique_id', time_col:str='ds',
+> target_col:str='y', distributed_config:Optional[neura
+> lforecast.common._base_model.DistributedConfig]=None,
+> prediction_intervals:Optional[neuralforecast.utils.Pr
+> edictionIntervals]=None)
+> ```
+
+\*Fit the core.NeuralForecast.
+
+Fit `models` to a large set of time series from DataFrame `df`. and
+store fitted models for later inspection.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | Union | None | DataFrame with columns \[`unique_id`, `ds`, `y`\] and exogenous variables. If None, a previously stored dataset is required. |
+| static_df | Union | None | DataFrame with columns \[`unique_id`\] and static exogenous. |
+| val_size | Optional | 0 | Size of validation set. |
+| use_init_models | bool | False | Use initial model passed when NeuralForecast object was instantiated. |
+| verbose | bool | False | Print processing steps. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| distributed_config | Optional | None | Configuration to use for DDP training. Currently only spark is supported. |
+| prediction_intervals | Optional | None | Configuration to calibrate prediction intervals (Conformal Prediction). |
+| **Returns** | **NeuralForecast** | | **Returns [`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast) class with fitted `models`.** |
+
+------------------------------------------------------------------------
+
+source
+
+### NeuralForecast.predict
+
+> ``` text
+> NeuralForecast.predict (df:Union[pandas.core.frame.DataFrame,polars.dataf
+> rame.frame.DataFrame,neuralforecast.compat.SparkD
+> ataFrame,NoneType]=None, static_df:Union[pandas.c
+> ore.frame.DataFrame,polars.dataframe.frame.DataFr
+> ame,neuralforecast.compat.SparkDataFrame,NoneType
+> ]=None, futr_df:Union[pandas.core.frame.DataFrame
+> ,polars.dataframe.frame.DataFrame,neuralforecast.
+> compat.SparkDataFrame,NoneType]=None,
+> verbose:bool=False, engine=None,
+> level:Optional[List[Union[int,float]]]=None,
+> quantiles:Optional[List[float]]=None,
+> **data_kwargs)
+> ```
+
+\*Predict with core.NeuralForecast.
+
+Use stored fitted `models` to predict large set of time series from
+DataFrame `df`.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | Union | None | DataFrame with columns \[`unique_id`, `ds`, `y`\] and exogenous variables. If a DataFrame is passed, it is used to generate forecasts. |
+| static_df | Union | None | DataFrame with columns \[`unique_id`\] and static exogenous. |
+| futr_df | Union | None | DataFrame with \[`unique_id`, `ds`\] columns and `df`’s future exogenous. |
+| verbose | bool | False | Print processing steps. |
+| engine | NoneType | None | Distributed engine for inference. Only used if df is a spark dataframe or if fit was called on a spark dataframe. |
+| level | Optional | None | Confidence levels between 0 and 100. |
+| quantiles | Optional | None | Alternative to level, target quantiles to predict. |
+| data_kwargs | VAR_KEYWORD | | Extra arguments to be passed to the dataset within each model. |
+| **Returns** | **pandas or polars DataFrame** | | **DataFrame with insample `models` columns for point predictions and probabilistic predictions for all fitted `models`. ** |
+
+------------------------------------------------------------------------
+
+source
+
+### NeuralForecast.cross_validation
+
+> ``` text
+> NeuralForecast.cross_validation (df:Union[pandas.core.frame.DataFrame,pol
+> ars.dataframe.frame.DataFrame,NoneType]=
+> None, static_df:Union[pandas.core.frame.
+> DataFrame,polars.dataframe.frame.DataFra
+> me,NoneType]=None, n_windows:int=1,
+> step_size:int=1,
+> val_size:Optional[int]=0,
+> test_size:Optional[int]=None,
+> use_init_models:bool=False,
+> verbose:bool=False,
+> refit:Union[bool,int]=False,
+> id_col:str='unique_id',
+> time_col:str='ds', target_col:str='y', p
+> rediction_intervals:Optional[neuralforec
+> ast.utils.PredictionIntervals]=None, lev
+> el:Optional[List[Union[int,float]]]=None
+> , quantiles:Optional[List[float]]=None,
+> **data_kwargs)
+> ```
+
+\*Temporal Cross-Validation with core.NeuralForecast.
+
+`core.NeuralForecast`’s cross-validation efficiently fits a list of
+NeuralForecast models through multiple windows, in either chained or
+rolled manner.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | Union | None | DataFrame with columns \[`unique_id`, `ds`, `y`\] and exogenous variables. If None, a previously stored dataset is required. |
+| static_df | Union | None | DataFrame with columns \[`unique_id`\] and static exogenous. |
+| n_windows | int | 1 | Number of windows used for cross validation. |
+| step_size | int | 1 | Step size between each window. |
+| val_size | Optional | 0 | Length of validation size. If passed, set `n_windows=None`. |
+| test_size | Optional | None | Length of test size. If passed, set `n_windows=None`. |
+| use_init_models | bool | False | Use initial model passed when object was instantiated. |
+| verbose | bool | False | Print processing steps. |
+| refit | Union | False | Retrain model for each cross validation window. If False, the models are trained at the beginning and then used to predict each window. If positive int, the models are retrained every `refit` windows. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| prediction_intervals | Optional | None | Configuration to calibrate prediction intervals (Conformal Prediction). |
+| level | Optional | None | Confidence levels between 0 and 100. |
+| quantiles | Optional | None | Alternative to level, target quantiles to predict. |
+| data_kwargs | VAR_KEYWORD | | Extra arguments to be passed to the dataset within each model. |
+| **Returns** | **Union** | | **DataFrame with insample `models` columns for point predictions and probabilistic predictions for all fitted `models`. ** |
+
+------------------------------------------------------------------------
+
+source
+
+### NeuralForecast.predict_insample
+
+> ``` text
+> NeuralForecast.predict_insample (step_size:int=1,
+> level:Optional[List[Union[int,float]]]=N
+> one,
+> quantiles:Optional[List[float]]=None)
+> ```
+
+\*Predict insample with core.NeuralForecast.
+
+`core.NeuralForecast`’s `predict_insample` uses stored fitted `models`
+to predict historic values of a time series from the stored dataframe.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| step_size | int | 1 | Step size between each window. |
+| level | Optional | None | Confidence levels between 0 and 100. |
+| quantiles | Optional | None | Alternative to level, target quantiles to predict. |
+| **Returns** | **pandas.DataFrame** | | **DataFrame with insample predictions for all fitted `models`. ** |
+
+------------------------------------------------------------------------
+
+source
+
+### NeuralForecast.save
+
+> ``` text
+> NeuralForecast.save (path:str, model_index:Optional[List]=None,
+> save_dataset:bool=True, overwrite:bool=False)
+> ```
+
+\*Save NeuralForecast core class.
+
+`core.NeuralForecast`’s method to save current status of models,
+dataset, and configuration. Note that by default the `models` are not
+saving training checkpoints to save disk memory, to get them change the
+individual model `**trainer_kwargs` to include
+`enable_checkpointing=True`.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| path | str | | Directory to save current status. |
+| model_index | Optional | None | List to specify which models from list of self.models to save. |
+| save_dataset | bool | True | Whether to save dataset or not. |
+| overwrite | bool | False | Whether to overwrite files or not. |
+
+``` text
+/opt/hostedtoolcache/Python/3.10.17/x64/lib/python3.10/site-packages/fastcore/docscrape.py:230: UserWarning: potentially wrong underline length...
+Parameters
+----------- in
+Load NeuralForecast
+...
+ else: warn(msg)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### NeuralForecast.load
+
+> ``` text
+> NeuralForecast.load (path, verbose=False, **kwargs)
+> ```
+
+\*Load NeuralForecast
+
+`core.NeuralForecast`’s method to load checkpoint from path.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| path | str | | Directory with stored artifacts. |
+| verbose | bool | False | |
+| kwargs | VAR_KEYWORD | | Additional keyword arguments to be passed to the function `load_from_checkpoint`. |
+| **Returns** | **NeuralForecast** | | **Instantiated [`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast) class.** |
+
+
+```python
+# Test predict_insample step_size
+
+h = 12
+train_end = AirPassengersPanel_train['ds'].max()
+sizes = AirPassengersPanel_train['unique_id'].value_counts().to_numpy()
+for step_size, test_size in [(7, 0), (9, 0), (7, 5), (9, 5)]:
+ models = [NHITS(h=h, input_size=12, max_steps=1)]
+ nf = NeuralForecast(models=models, freq='M')
+ nf.fit(AirPassengersPanel_train)
+ # Note: only apply set_test_size() upon nf.fit(), otherwise it would have set the test_size = 0
+ nf.models[0].set_test_size(test_size)
+
+ forecasts = nf.predict_insample(step_size=step_size)
+ last_cutoff = train_end - test_size * pd.offsets.MonthEnd() - h * pd.offsets.MonthEnd()
+ n_expected_cutoffs = (sizes[0] - test_size - nf.h + step_size) // step_size
+
+ # compare cutoff values
+ expected_cutoffs = np.flip(np.array([last_cutoff - step_size * i * pd.offsets.MonthEnd() for i in range(n_expected_cutoffs)]))
+ actual_cutoffs = np.array([pd.Timestamp(x) for x in forecasts[forecasts['unique_id']==nf.uids[1]]['cutoff'].unique()])
+ np.testing.assert_array_equal(expected_cutoffs, actual_cutoffs, err_msg=f"{step_size=},{expected_cutoffs=},{actual_cutoffs=}")
+
+ # check forecast-points count per series
+ cutoffs_by_series = forecasts.groupby(['unique_id', 'cutoff']).size().unstack('unique_id')
+ pd.testing.assert_series_equal(cutoffs_by_series['Airline1'], cutoffs_by_series['Airline2'], check_names=False)
+```
+
+
+```python
+# Test predict_insample
+
+def get_expected_cols(model, level):
+ # index columns
+ n_cols = 4
+ for model in models:
+ if isinstance(loss, (DistributionLoss, PMM, GMM, NBMM)):
+ if level is None:
+ # Variations of DistributionLoss return the sample mean as well
+ n_cols += len(loss.quantiles) + 1
+ else:
+ # Variations of DistributionLoss return the sample mean as well
+ n_cols += 2 * len(level) + 1
+ else:
+ if level is None:
+ # Other probabilistic models return the sample mean as well
+ n_cols += 1
+ # Other probabilistic models return just the levels
+ else:
+ n_cols += len(level) + 1
+ return n_cols
+
+for loss in [
+ # IQLoss(),
+ DistributionLoss(distribution="Normal", level=[80]),
+ PMM(level=[80]),
+]:
+ for level in [None, [80, 90]]:
+ models = [
+ NHITS(h=12, input_size=12, loss=loss, max_steps=1),
+ LSTM(h=12, input_size=12, loss=loss, max_steps=1, recurrent=True),
+ ]
+ nf = NeuralForecast(models=models, freq='D')
+
+ nf.fit(df=AirPassengersPanel_train)
+ df = nf.predict_insample(step_size=1, level=level)
+ expected_cols = get_expected_cols(models, level)
+ assert df.shape[1] == expected_cols, f'Shape mismatch for {loss} and level={level} in predict_insample: cols={df.shape[1]}, expected_cols={expected_cols}'
+```
+
+
+```python
+def config_optuna(trial):
+ return {"input_size": trial.suggest_categorical('input_size', [12, 24]),
+ "hist_exog_list": trial.suggest_categorical('hist_exog_list', [['trend'], ['y_[lag12]'], ['trend', 'y_[lag12]']]),
+ "futr_exog_list": ['trend'],
+ "max_steps": 10,
+ "val_check_steps": 5}
+
+config_ray = {'input_size': tune.choice([12, 24]),
+ 'hist_exog_list': tune.choice([['trend'], ['y_[lag12]'], ['trend', 'y_[lag12]']]),
+ 'futr_exog_list': ['trend'],
+ 'max_steps': 10,
+ 'val_check_steps': 5}
+```
+
+
+```python
+# Test predict_insample step_size
+
+h = 12
+train_end = AirPassengers_pl['time'].max()
+sizes = AirPassengers_pl['uid'].value_counts().to_numpy()
+
+for step_size, test_size in [(7, 0), (9, 0), (7, 5), (9, 5)]:
+ models = [NHITS(h=h, input_size=12, max_steps=1)]
+ nf = NeuralForecast(models=models, freq='1mo')
+ nf.fit(
+ AirPassengers_pl,
+ id_col='uid',
+ time_col='time',
+ target_col='target',
+ )
+ # Note: only apply set_test_size() upon nf.fit(), otherwise it would have set the test_size = 0
+ nf.models[0].set_test_size(test_size)
+
+ forecasts = nf.predict_insample(step_size=step_size)
+ n_expected_cutoffs = (sizes[0][1] - test_size - nf.h + step_size) // step_size
+
+ # compare cutoff values
+ last_cutoff = train_end - test_size * pd.offsets.MonthEnd() - h * pd.offsets.MonthEnd()
+ expected_cutoffs = np.flip(np.array([last_cutoff - step_size * i * pd.offsets.MonthEnd() for i in range(n_expected_cutoffs)]))
+ pl_cutoffs = forecasts.filter(polars.col('uid') ==nf.uids[1]).select('cutoff').unique(maintain_order=True)
+ actual_cutoffs = np.sort(np.array([pd.Timestamp(x['cutoff']) for x in pl_cutoffs.rows(named=True)]))
+ np.testing.assert_array_equal(expected_cutoffs, actual_cutoffs, err_msg=f"{step_size=},{expected_cutoffs=},{actual_cutoffs=}")
+
+ # check forecast-points count per series
+ cutoffs_by_series = forecasts.group_by(['uid', 'cutoff']).count()
+ assert_frame_equal(cutoffs_by_series.filter(polars.col('uid') == "Airline1").select(['cutoff', 'count']), cutoffs_by_series.filter(polars.col('uid') == "Airline2").select(['cutoff', 'count'] ), check_row_order=False)
+```
+
diff --git a/neuralforecast/dark.png b/neuralforecast/dark.png
new file mode 100644
index 00000000..4142a0bb
Binary files /dev/null and b/neuralforecast/dark.png differ
diff --git a/neuralforecast/docs/api-reference/neuralforecast_map.html.mdx b/neuralforecast/docs/api-reference/neuralforecast_map.html.mdx
new file mode 100644
index 00000000..0716d85c
--- /dev/null
+++ b/neuralforecast/docs/api-reference/neuralforecast_map.html.mdx
@@ -0,0 +1,151 @@
+---
+description: Modules of the NeuralForecast library
+output-file: neuralforecast_map.html
+title: NeuralForecast Map
+---
+
+
+The `neuralforecast` library provides a comprehensive set of
+state-of-the-art deep learning models designed to power-up time series
+forecasting pipelines.
+
+The library is constructed using a modular approach, where different
+responsibilities are isolated within specific modules. These modules
+include the user interface functions (`core`), data processing and
+loading (`tsdataset`), scalers, losses, and base classes for models.
+
+This tutorial aims to explain the library’s structure and to describe
+how the different modules interact with each other.
+
+## I. Map
+
+The following diagram presents the modules of the `neuralforecast`
+library and their relations.
+
+
+
+Neuralforecast map
+
+
+## II. Modules
+
+### 1. Core (`core.py`)
+
+The `core` module acts as the primary interaction point for users of the
+`neuralforecast` library. It houses the
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+class, which incorporates a range of key user interface functions
+designed to simplify the process of training and forecasting models.
+Functions include `fit`, `predict`, `cross_validation`, and
+`predict_insample`, each one constructed to be intuitive and
+user-friendly. The design of the
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+class is centered around enabling users to streamline their forecasting
+pipelines and to comfortably train and evaluate models.
+
+### 2. Dataset and Loader (`tsdataset.py`)
+
+The
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset)
+class, located within the `tsdataset` module, is responsible for the
+storage and preprocessing of the input time series dataset. Once the
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset)
+class has prepared the data, it’s then consumed by the
+[`TimeSeriesLoader`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesloader)
+class, which samples batches (or subsets) of the time series during the
+training and inference stages.
+
+### 3. Base Model (`common`)
+
+The `common` module contains three `BaseModel` classes, which serve as
+the foundation for all the model structures provided in the library.
+These base classes allow for a level of abstraction and code-reusability
+in the design of the models. We currently support three type of models:
+
+- `BaseWindows`: designed for window-based models like
+ [`NBEATS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nbeats.html#nbeats)
+ and `Transformers`.
+- `BaseRecurrent`: designed for recurrent models like
+ [`RNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.rnn.html#rnn)
+ and
+ [`LSTM`](https://nixtlaverse.nixtla.io/neuralforecast/models.lstm.html#lstm).
+- `BaseMultivariate`: caters to multivariate models like
+ [`StemGNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.stemgnn.html#stemgnn).
+
+### 4. Model (`models`)
+
+The `models` module encompasses all the specific model classes available
+for use in the library. These include a variety of both simple and
+complex models such as
+[`RNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.rnn.html#rnn),
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits),
+[`LSTM`](https://nixtlaverse.nixtla.io/neuralforecast/models.lstm.html#lstm),
+[`StemGNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.stemgnn.html#stemgnn),
+and
+[`TFT`](https://nixtlaverse.nixtla.io/neuralforecast/models.tft.html#tft).
+Each model in this module extends from one of the `BaseModel` classes in
+the `common` module.
+
+### 5. Losses (`losses`)
+
+The `losses` module includes both `numpy` and `pytorch` losses, used for
+evalaution and training respectively. The module contains a wide range
+of losses, including
+[`MAE`](https://nixtlaverse.nixtla.io/neuralforecast/losses.pytorch.html#mae),
+[`MSE`](https://nixtlaverse.nixtla.io/neuralforecast/losses.pytorch.html#mse),
+[`MAPE`](https://nixtlaverse.nixtla.io/neuralforecast/losses.pytorch.html#mape),
+[`HuberLoss`](https://nixtlaverse.nixtla.io/neuralforecast/losses.pytorch.html#huberloss),
+among many others.
+
+### 6. Scalers (`_scalers.py`)
+
+The `_scalers.py` module houses the `TemporalNorm` class. This class is
+responsible for the scaling (normalization) and de-scaling (reversing
+the normalization) of time series data. This step is crucial because it
+ensures all data fed to the model have a similar range, leading to more
+stable and efficient training processes.
+
+## III. Flow
+
+The `user` first instantiates a model and the
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+core class. When they call the `fit` method, the following flow is
+executed:
+
+1. The `fit` method instantiates a
+ [`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset)
+ object to store and pre-process the input time series dataset, and
+ the
+ [`TimeSeriesLoader`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesloader)
+ object to sample batches.
+2. The `fit` method calls the model’s `fit` method (in the `BaseModel`
+ class).
+3. The model’s `fit` method instantiates a Pytorch-Lightning `Trainer`
+ object, in charge of training the model.
+4. The `Trainer` method samples a batch from the
+ [`TimeSeriesLoader`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesloader)
+ object, and calls the model’s `training_step` method (in the
+ `BaseModel` class).
+5. The model’s `training_step`:
+ - Samples windows from the original batch.
+ - Normalizes the windows with the `scaler` module.
+ - Calls the model’s `forward` method.
+ - Computes the loss using the `losses` module.
+ - Returns the loss.
+6. The `Trainer` object repeats step 4 and 5 until `max_steps`
+ iterations are completed.
+7. The model is fitted, and can be used for forecasting future values
+ (with the `predict` method) or recover insample predictions (using
+ the `predict_insample` method).
+
+## IV. Next Steps: add your own model
+
+Congratulations! You now know the internal details of the
+`neuralforecast` library.
+
+With this knowledge you can easily add new models to the library, by
+just creating a `model` class which only requires the `init` and
+`forward` methods.
+
+Check our detailed guide on how to add new models!
+
diff --git a/neuralforecast/docs/capabilities/03_exogenous_variables_files/figure-markdown_strict/cell-13-output-1.png b/neuralforecast/docs/capabilities/03_exogenous_variables_files/figure-markdown_strict/cell-13-output-1.png
new file mode 100644
index 00000000..15b76e44
Binary files /dev/null and b/neuralforecast/docs/capabilities/03_exogenous_variables_files/figure-markdown_strict/cell-13-output-1.png differ
diff --git a/neuralforecast/docs/capabilities/03_exogenous_variables_files/figure-markdown_strict/cell-5-output-1.png b/neuralforecast/docs/capabilities/03_exogenous_variables_files/figure-markdown_strict/cell-5-output-1.png
new file mode 100644
index 00000000..f84514bd
Binary files /dev/null and b/neuralforecast/docs/capabilities/03_exogenous_variables_files/figure-markdown_strict/cell-5-output-1.png differ
diff --git a/neuralforecast/docs/capabilities/04_hyperparameter_tuning_files/figure-markdown_strict/cell-23-output-1.png b/neuralforecast/docs/capabilities/04_hyperparameter_tuning_files/figure-markdown_strict/cell-23-output-1.png
new file mode 100644
index 00000000..3a77810d
Binary files /dev/null and b/neuralforecast/docs/capabilities/04_hyperparameter_tuning_files/figure-markdown_strict/cell-23-output-1.png differ
diff --git a/neuralforecast/docs/capabilities/05_predictInsample_files/figure-markdown_strict/cell-11-output-1.png b/neuralforecast/docs/capabilities/05_predictInsample_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..f9bbb1e8
Binary files /dev/null and b/neuralforecast/docs/capabilities/05_predictInsample_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/neuralforecast/docs/capabilities/05_predictInsample_files/figure-markdown_strict/cell-14-output-1.png b/neuralforecast/docs/capabilities/05_predictInsample_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..df772929
Binary files /dev/null and b/neuralforecast/docs/capabilities/05_predictInsample_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/neuralforecast/docs/capabilities/06_save_load_models_files/figure-markdown_strict/cell-11-output-1.png b/neuralforecast/docs/capabilities/06_save_load_models_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..0a9ef68a
Binary files /dev/null and b/neuralforecast/docs/capabilities/06_save_load_models_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/neuralforecast/docs/capabilities/06_save_load_models_files/figure-markdown_strict/cell-14-output-1.png b/neuralforecast/docs/capabilities/06_save_load_models_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..4bf09c3f
Binary files /dev/null and b/neuralforecast/docs/capabilities/06_save_load_models_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/neuralforecast/docs/capabilities/07_time_series_scaling_files/figure-markdown_strict/cell-11-output-1.png b/neuralforecast/docs/capabilities/07_time_series_scaling_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..72ee192a
Binary files /dev/null and b/neuralforecast/docs/capabilities/07_time_series_scaling_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/neuralforecast/docs/capabilities/07_time_series_scaling_files/figure-markdown_strict/cell-15-output-1.png b/neuralforecast/docs/capabilities/07_time_series_scaling_files/figure-markdown_strict/cell-15-output-1.png
new file mode 100644
index 00000000..37bba595
Binary files /dev/null and b/neuralforecast/docs/capabilities/07_time_series_scaling_files/figure-markdown_strict/cell-15-output-1.png differ
diff --git a/neuralforecast/docs/capabilities/08_cross_validation_files/figure-markdown_strict/cell-11-output-1.png b/neuralforecast/docs/capabilities/08_cross_validation_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..1afced23
Binary files /dev/null and b/neuralforecast/docs/capabilities/08_cross_validation_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/neuralforecast/docs/capabilities/08_cross_validation_files/figure-markdown_strict/cell-13-output-1.png b/neuralforecast/docs/capabilities/08_cross_validation_files/figure-markdown_strict/cell-13-output-1.png
new file mode 100644
index 00000000..bbdf0169
Binary files /dev/null and b/neuralforecast/docs/capabilities/08_cross_validation_files/figure-markdown_strict/cell-13-output-1.png differ
diff --git a/neuralforecast/docs/capabilities/08_cross_validation_files/figure-markdown_strict/cell-15-output-1.png b/neuralforecast/docs/capabilities/08_cross_validation_files/figure-markdown_strict/cell-15-output-1.png
new file mode 100644
index 00000000..fa1f4445
Binary files /dev/null and b/neuralforecast/docs/capabilities/08_cross_validation_files/figure-markdown_strict/cell-15-output-1.png differ
diff --git a/neuralforecast/docs/capabilities/08_cross_validation_files/figure-markdown_strict/cell-7-output-1.png b/neuralforecast/docs/capabilities/08_cross_validation_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..1e63f3e3
Binary files /dev/null and b/neuralforecast/docs/capabilities/08_cross_validation_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/neuralforecast/docs/capabilities/08_cross_validation_files/figure-markdown_strict/cell-9-output-1.png b/neuralforecast/docs/capabilities/08_cross_validation_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..4e4643dd
Binary files /dev/null and b/neuralforecast/docs/capabilities/08_cross_validation_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/neuralforecast/docs/capabilities/cross_validation.html.mdx b/neuralforecast/docs/capabilities/cross_validation.html.mdx
new file mode 100644
index 00000000..c6abe2ec
--- /dev/null
+++ b/neuralforecast/docs/capabilities/cross_validation.html.mdx
@@ -0,0 +1,315 @@
+---
+output-file: cross_validation.html
+title: Cross-validation
+---
+
+
+In this tutorial, we explore in detail the cross-validation function in
+`neuralforecast`.
+
+## 1. Libraries
+
+Make sure to install `neuralforecast` to follow along.
+
+
+```python
+!pip install neuralforecast
+```
+
+
+```python
+import logging
+import matplotlib.pyplot as plt
+import pandas as pd
+from utilsforecast.plotting import plot_series
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import NHITS
+```
+
+
+```python
+logging.getLogger("pytorch_lightning").setLevel(logging.ERROR)
+```
+
+## 2. Read the data
+
+For this tutorial, we use part of the hourly M4 dataset. It is stored in
+a parquet file for efficiency. You can use ordinary pandas operations to
+read your data in other formats likes `.csv`.
+
+The input to
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+is always a data frame in [long
+format](https://www.theanalysisfactor.com/wide-and-long-data/) with
+three columns: `unique_id`, `ds` and `y`:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp or int) column should be either an integer
+ indexing time or a datestampe ideally like YYYY-MM-DD for a date or
+ YYYY-MM-DD HH:MM:SS for a timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+Depending on your internet connection, this step should take around 10
+seconds.
+
+
+```python
+Y_df = pd.read_parquet('https://datasets-nixtla.s3.amazonaws.com/m4-hourly.parquet')
+Y_df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|-----|-------|
+| 0 | H1 | 1 | 605.0 |
+| 1 | H1 | 2 | 586.0 |
+| 2 | H1 | 3 | 586.0 |
+| 3 | H1 | 4 | 559.0 |
+| 4 | H1 | 5 | 511.0 |
+
+For simplicity, we use only a single series to explore in detail the
+cross-validation functionality. Also, let’s use the first 700 time
+steps, such that we work with round numbers, making it easier to
+visualize and understand cross-validation.
+
+
+```python
+Y_df = Y_df.query("unique_id == 'H1'")[:700]
+Y_df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|-----|-------|
+| 0 | H1 | 1 | 605.0 |
+| 1 | H1 | 2 | 586.0 |
+| 2 | H1 | 3 | 586.0 |
+| 3 | H1 | 4 | 559.0 |
+| 4 | H1 | 5 | 511.0 |
+
+
+```python
+plot_series(Y_df)
+```
+
+
+
+## 3. Using cross-validation
+
+### 3.1 Using `n_windows`
+
+To use the `cross_validation` method, we can either: - Set the sizes of
+a validation and test set - Set a number of cross-validation windows
+
+Let’s see how it works in a minimal example. Here, we use the NHITS
+model and set the horizon to 100, and give an input size of 200.
+
+First, let’s use `n_windows = 4`.
+
+We also set `step_size` equal to the horizon. This parameter controls
+the distance between each cross-validation window. By setting it equal
+to the horizon, we perform *chained cross-validation* where the windows
+do not overlap.
+
+
+```python
+h = 100
+nf = NeuralForecast(models=[NHITS(h=h, input_size=2*h, max_steps=500, enable_progress_bar=False, logger=False)], freq=1);
+cv_df = nf.cross_validation(Y_df, n_windows=4, step_size=h, verbose=0)
+cv_df.head()
+```
+
+``` text
+Seed set to 1
+```
+
+| | unique_id | ds | cutoff | NHITS | y |
+|-----|-----------|-----|--------|------------|-------|
+| 0 | H1 | 301 | 300 | 490.048950 | 485.0 |
+| 1 | H1 | 302 | 300 | 537.713867 | 525.0 |
+| 2 | H1 | 303 | 300 | 612.900635 | 585.0 |
+| 3 | H1 | 304 | 300 | 689.346313 | 670.0 |
+| 4 | H1 | 305 | 300 | 760.153992 | 747.0 |
+
+
+```python
+cutoffs = cv_df['cutoff'].unique()
+
+plt.figure(figsize=(15,5))
+plt.plot(Y_df['ds'], Y_df['y'])
+plt.plot(cv_df['ds'], cv_df['NHITS'], label='NHITS', ls='--')
+
+for cutoff in cutoffs:
+ plt.axvline(x=cutoff, color='black', ls=':')
+
+plt.xlabel('Time steps')
+plt.ylabel('Target [H1]')
+plt.legend()
+plt.tight_layout()
+```
+
+
+
+In the figure above, we see that we have 4 cutoff points, which
+correspond to our four cross-validation windows. Of course, notice that
+the windows are set from the end of the dataset. That way, the model
+trains on past data to predict future data.
+
+> **Important note**
+>
+> We start counting at 0, so counting from 0 to 99 results in a sequence
+> of 100 data points.
+
+Thus, the model is initially trained using time steps 0 to 299. Then, to
+make predictions, it takes time steps 100 to 299 (input size of 200) and
+it makes predictions for time steps 300 to 399 (horizon of 100).
+
+Then, the actual values from 200 to 399 (because our model has an
+`input_size` of 200) are used to generate predictions over the next
+window, from 400 to 499.
+
+This process is repeated until we run out of windows.
+
+### 3.2 Using a validation and test set
+
+Instead of setting a number of windows, we can define a validation and
+test set. In that case, we must set `n_windows=None`
+
+
+```python
+cv_df_val_test = nf.cross_validation(Y_df, val_size=200, test_size=200, step_size=h, n_windows=None)
+```
+
+
+```python
+cutoffs = cv_df_val_test['cutoff'].unique()
+plt.figure(figsize=(15,5))
+
+# Plot the original data and NHITS predictions
+plt.plot(Y_df['ds'], Y_df['y'])
+plt.plot(cv_df_val_test['ds'], cv_df_val_test['NHITS'], label='NHITS', ls='--')
+
+# Add highlighted areas for validation and test sets
+plt.axvspan(Y_df['ds'].iloc[300], Y_df['ds'].iloc[499], alpha=0.2, color='yellow', label='Validation Set')
+plt.axvspan(Y_df['ds'].iloc[500], Y_df['ds'].iloc[699], alpha=0.2, color='red', label='Test Set')
+
+# Add vertical lines for cutoffs
+for cutoff in cutoffs:
+ plt.axvline(x=cutoff, color='black', ls=':')
+
+# Set labels and legend
+plt.xlabel('Time steps')
+plt.ylabel('Target [H1]')
+plt.legend()
+
+plt.tight_layout()
+plt.show()
+```
+
+
+
+Here, we predict only the test set, which corresponds to the last 200
+time steps. Since the model has a forecast horizon of 100, and
+`step_size` is also set to 100, there are only two cross-validation
+windows in the test set (200/100 = 2). Thus, we only see two cutoff
+points.
+
+### 3.3 Cross-validation with refit
+
+In the previous sections, we trained the model only once and predicted
+over many cross-validation windows. However, in real life, we often
+retrain our model with new observed data before making the next set of
+predictions.
+
+We can simulate that process using `refit=True`. That way, the model is
+retrained at every step in the cross-validation process. In other words,
+the training set is gradually expanded with new observed values and the
+model is retrained before making the next set of predictions.
+
+
+```python
+cv_df_refit = nf.cross_validation(Y_df, n_windows=4, step_size=h, refit=True)
+```
+
+
+```python
+cutoffs = cv_df_refit['cutoff'].unique()
+
+plt.figure(figsize=(15,5))
+plt.plot(Y_df['ds'], Y_df['y'])
+plt.plot(cv_df_refit['ds'], cv_df_refit['NHITS'], label='NHITS', ls='--')
+
+for cutoff in cutoffs:
+ plt.axvline(x=cutoff, color='black', ls=':')
+
+plt.xlabel('Time steps')
+plt.ylabel('Target [H1]')
+plt.legend()
+plt.tight_layout()
+```
+
+
+
+Notice that when we run cross-validation with `refit=True`, there were 4
+training loops that were completed. This is expected because the model
+is now retrained with new data for each fold in the cross-validation: -
+fold 1: train on the first 300 steps, predict the next 100 - fold 2:
+train on the first 400 steps, predict the next 100 - fold 3: train on
+the first 500 steps, predict the next 100 - fold 4: train on the first
+600 steps, predict the next 100
+
+### 3.4 Overlapping windows in cross-validation
+
+In the case where `step_size` is smaller than the horizon, we get
+overlapping windows. This means that we make predictions more than once
+for some time steps.
+
+This is useful to test the model over more forecast windows, and it
+provides a more robust evaluation, as the model is tested across
+different segments of the series.
+
+However, it comes with a higher computation cost, as we are making
+predictions more than once for some of the time steps.
+
+
+```python
+cv_df_refit_overlap = nf.cross_validation(Y_df, n_windows=2, step_size=50, refit=True)
+```
+
+
+```python
+cutoffs = cv_df_refit_overlap['cutoff'].unique()
+
+fold1 = cv_df_refit_overlap.query("cutoff==550")
+fold2 = cv_df_refit_overlap.query("cutoff==600")
+
+plt.figure(figsize=(15,5))
+plt.plot(Y_df['ds'], Y_df['y'])
+plt.plot(fold1['ds'], fold1['NHITS'], label='NHITS (fold 1)', ls='--', color='blue')
+plt.plot(fold2['ds'], fold2['NHITS'], label='NHITS (fold 2)', ls='-.', color='red')
+
+for cutoff in cutoffs:
+ plt.axvline(x=cutoff, color='black', ls=':')
+
+plt.xlabel('Time steps')
+plt.ylabel('Target [H1]')
+plt.xlim(500, 700)
+plt.legend()
+plt.tight_layout()
+```
+
+
+
+In the figure above, we see that our two folds overlap between time
+steps 601 and 650, since the step size is 50. This happens because: -
+fold 1: model is trained using time steps 0 to 550 and predicts 551 to
+650 (h=100) - fold 2: model is trained using time steps 0 to 600
+(`step_size=50`) and predicts 601 to 700
+
+Be aware that when evaluating a model trained with overlapping
+cross-validation windows, some time steps have more than one prediction.
+This may bias your evaluation metric, as the repeated time steps are
+taken into account in the metric multiple times.
+
diff --git a/neuralforecast/docs/capabilities/exogenous_variables.html.mdx b/neuralforecast/docs/capabilities/exogenous_variables.html.mdx
new file mode 100644
index 00000000..ddc5d4cb
--- /dev/null
+++ b/neuralforecast/docs/capabilities/exogenous_variables.html.mdx
@@ -0,0 +1,292 @@
+---
+output-file: exogenous_variables.html
+title: Exogenous Variables
+---
+
+
+Exogenous variables can provide additional information to greatly
+improve forecasting accuracy. Some examples include price or future
+promotions variables for demand forecasting, and weather data for
+electricity load forecast. In this notebook we show an example on how to
+add different types of exogenous variables to NeuralForecast models for
+making day-ahead hourly electricity price forecasts (EPF) for France and
+Belgium markets.
+
+All NeuralForecast models are capable of incorporating exogenous
+variables to model the following conditional predictive distribution:
+$$\mathbb{P}(\mathbf{y}_{t+1:t+H} \;|\; \mathbf{y}_{[:t]},\; \mathbf{x}^{(h)}_{[:t]},\; \mathbf{x}^{(f)}_{[:t+H]},\; \mathbf{x}^{(s)} )$$
+
+where the regressors are static exogenous $\mathbf{x}^{(s)}$, historic
+exogenous $\mathbf{x}^{(h)}_{[:t]}$, exogenous available at the time of
+the prediction $\mathbf{x}^{(f)}_{[:t+H]}$ and autorregresive features
+$\mathbf{y}_{[:t]}$. Depending on the [train
+loss](https://nixtla.github.io/neuralforecast/losses.pytorch.html), the
+model outputs can be point forecasts (location estimators) or
+uncertainty intervals (quantiles).
+
+We will show you how to include exogenous variables in the data, specify
+variables to a model, and produce forecasts using future exogenous
+variables.
+
+> **Important**
+>
+> This Guide assumes basic knowledge on the NeuralForecast library. For
+> a minimal example visit the [Getting
+> Started](../getting-started/quickstart.html) guide.
+
+You can run these experiments using GPU with Google Colab.
+
+
+
+## 1. Libraries
+
+
+```python
+!pip install neuralforecast
+```
+
+## 2. Load data
+
+The `df` dataframe contains the target and exogenous variables past
+information to train the model. The `unique_id` column identifies the
+markets, `ds` contains the datestamps, and `y` the electricity price.
+
+Include both historic and future temporal variables as columns. In this
+example, we are adding the system load (`system_load`) as historic data.
+For future variables, we include a forecast of how much electricity will
+be produced (`gen_forecast`) and day of week (`week_day`). Both the
+electricity system demand and offer impact the price significantly,
+including these variables to the model greatly improve performance, as
+we demonstrate in Olivares et al. (2022).
+
+The distinction between historic and future variables will be made later
+as parameters of the model.
+
+
+```python
+import pandas as pd
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+df = pd.read_csv(
+ 'https://datasets-nixtla.s3.amazonaws.com/EPF_FR_BE.csv',
+ parse_dates=['ds'],
+)
+df.head()
+```
+
+| | unique_id | ds | y | gen_forecast | system_load | week_day |
+|-----|-----------|---------------------|-------|--------------|-------------|----------|
+| 0 | FR | 2015-01-01 00:00:00 | 53.48 | 76905.0 | 74812.0 | 3 |
+| 1 | FR | 2015-01-01 01:00:00 | 51.93 | 75492.0 | 71469.0 | 3 |
+| 2 | FR | 2015-01-01 02:00:00 | 48.76 | 74394.0 | 69642.0 | 3 |
+| 3 | FR | 2015-01-01 03:00:00 | 42.27 | 72639.0 | 66704.0 | 3 |
+| 4 | FR | 2015-01-01 04:00:00 | 38.41 | 69347.0 | 65051.0 | 3 |
+
+> **Tip**
+>
+> Calendar variables such as day of week, month, and year are very
+> useful to capture long seasonalities.
+
+
+```python
+plot_series(df)
+```
+
+
+
+Add the static variables in a separate `static_df` dataframe. In this
+example, we are using one-hot encoding of the electricity market. The
+`static_df` must include one observation (row) for each `unique_id` of
+the `df` dataframe, with the different statics variables as columns.
+
+
+```python
+static_df = pd.read_csv('https://datasets-nixtla.s3.amazonaws.com/EPF_FR_BE_static.csv')
+static_df.head()
+```
+
+| | unique_id | market_0 | market_1 |
+|-----|-----------|----------|----------|
+| 0 | FR | 1 | 0 |
+| 1 | BR | 0 | 1 |
+
+## 3. Training with exogenous variables
+
+We distinguish the exogenous variables by whether they reflect static or
+time-dependent aspects of the modeled data.
+
+- **Static exogenous variables**: The static exogenous variables carry
+ time-invariant information for each time series. When the model is
+ built with global parameters to forecast multiple time series, these
+ variables allow sharing information within groups of time series
+ with similar static variable levels. Examples of static variables
+ include designators such as identifiers of regions, groups of
+ products, etc.
+
+- **Historic exogenous variables**: This time-dependent exogenous
+ variable is restricted to past observed values. Its predictive power
+ depends on Granger-causality, as its past values can provide
+ significant information about future values of the target variable
+ $\mathbf{y}$.
+
+- **Future exogenous variables**: In contrast with historic exogenous
+ variables, future values are available at the time of the
+ prediction. Examples include calendar variables, weather forecasts,
+ and known events that can cause large spikes and dips such as
+ scheduled promotions.
+
+To add exogenous variables to the model, first specify the name of each
+variable from the previous dataframes to the corresponding model
+hyperparameter during initialization: `futr_exog_list`,
+`hist_exog_list`, and `stat_exog_list`. We also set `horizon` as 24 to
+produce the next day hourly forecasts, and set `input_size` to use the
+last 5 days of data as input.
+
+
+```python
+import logging
+
+from neuralforecast.auto import NHITS, BiTCN
+from neuralforecast.core import NeuralForecast
+```
+
+
+```python
+logging.getLogger("pytorch_lightning").setLevel(logging.WARNING)
+```
+
+
+```python
+horizon = 24 # day-ahead daily forecast
+models = [NHITS(h = horizon,
+ max_steps=100,
+ input_size = 5*horizon,
+ futr_exog_list = ['gen_forecast', 'week_day'], # <- Future exogenous variables
+ hist_exog_list = ['system_load'], # <- Historical exogenous variables
+ stat_exog_list = ['market_0', 'market_1'], # <- Static exogenous variables
+ scaler_type = 'robust'),
+ BiTCN(h = horizon,
+ input_size = 5*horizon,
+ max_steps=100,
+ futr_exog_list = ['gen_forecast', 'week_day'], # <- Future exogenous variables
+ hist_exog_list = ['system_load'], # <- Historical exogenous variables
+ stat_exog_list = ['market_0', 'market_1'], # <- Static exogenous variables
+ scaler_type = 'robust',
+ ),
+ ]
+```
+
+``` text
+Seed set to 1
+Seed set to 1
+```
+
+> **Tip**
+>
+> When including exogenous variables always use a scaler by setting the
+> `scaler_type` hyperparameter. The scaler will scale all the temporal
+> features: the target variable `y`, historic and future variables.
+
+> **Important**
+>
+> Make sure future and historic variables are correctly placed. Defining
+> historic variables as future variables will lead to data leakage.
+
+Next, pass the datasets to the `df` and `static_df` inputs of the `fit`
+method.
+
+
+```python
+nf = NeuralForecast(models=models, freq='h')
+nf.fit(df=df, static_df=static_df)
+```
+
+## 4. Forecasting with exogenous variables
+
+Before predicting the prices, we need to gather the future exogenous
+variables for the day we want to forecast. Define a new dataframe
+(`futr_df`) with the `unique_id`, `ds`, and future exogenous variables.
+There is no need to add the target variable `y` and historic variables
+as they won’t be used by the model.
+
+
+```python
+futr_df = pd.read_csv(
+ 'https://datasets-nixtla.s3.amazonaws.com/EPF_FR_BE_futr.csv',
+ parse_dates=['ds'],
+)
+futr_df.head()
+```
+
+| | unique_id | ds | gen_forecast | week_day |
+|-----|-----------|---------------------|--------------|----------|
+| 0 | FR | 2016-11-01 00:00:00 | 49118.0 | 1 |
+| 1 | FR | 2016-11-01 01:00:00 | 47890.0 | 1 |
+| 2 | FR | 2016-11-01 02:00:00 | 47158.0 | 1 |
+| 3 | FR | 2016-11-01 03:00:00 | 45991.0 | 1 |
+| 4 | FR | 2016-11-01 04:00:00 | 45378.0 | 1 |
+
+> **Important**
+>
+> Make sure `futr_df` has informations for the entire forecast horizon.
+> In this example, we are forecasting 24 hours ahead, so `futr_df` must
+> have 24 rows for each time series.
+
+Finally, use the `predict` method to forecast the day-ahead prices.
+
+
+```python
+Y_hat_df = nf.predict(futr_df=futr_df)
+Y_hat_df.head()
+```
+
+``` text
+Predicting: | | 0/? [00:00
+
+## 1. Install `Neuralforecast`
+
+
+```python
+!pip install neuralforecast hyperopt
+```
+
+## 2. Load Data
+
+In this example we will use the `AirPasengers`, a popular dataset with
+monthly airline passengers in the US from 1949 to 1960. Load the data,
+available at our `utils` methods in the required format. See
+https://nixtlaverse.nixtla.io/neuralforecast/examples/data_format.html for
+more details on the data input format.
+
+
+```python
+import logging
+
+from neuralforecast.utils import AirPassengersDF
+```
+
+
+```python
+logging.getLogger('pytorch_lightning').setLevel(logging.ERROR)
+```
+
+
+```python
+Y_df = AirPassengersDF
+Y_df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|-------|
+| 0 | 1.0 | 1949-01-31 | 112.0 |
+| 1 | 1.0 | 1949-02-28 | 118.0 |
+| 2 | 1.0 | 1949-03-31 | 132.0 |
+| 3 | 1.0 | 1949-04-30 | 129.0 |
+| 4 | 1.0 | 1949-05-31 | 121.0 |
+
+## 3. Ray’s `Tune` backend
+
+First, we show how to use the `Tune` backend. This backend is based on
+Ray’s `Tune` library, which is a scalable framework for hyperparameter
+tuning. It is a popular library in the machine learning community, and
+it is used by many companies and research labs. If you plan to use the
+`Optuna` backend, you can skip this section.
+
+### 3.a Define hyperparameter grid
+
+Each `Auto` model contains a default search space that was extensively
+tested on multiple large-scale datasets. Search spaces are specified
+with dictionaries, where keys corresponds to the model’s hyperparameter
+and the value is a `Tune` function to specify how the hyperparameter
+will be sampled. For example, use `randint` to sample integers
+uniformly, and `choice` to sample values of a list.
+
+### 3.a.1 Default hyperparameter grid
+
+The default search space dictionary can be accessed through the
+`get_default_config` function of the `Auto` model. This is useful if you
+wish to use the default parameter configuration but want to change one
+or more hyperparameter spaces without changing the other default values.
+
+To extract the default config, you need to define: \* `h`: forecasting
+horizon. \* `backend`: backend to use. \* `n_series`: Optional, the
+number of unique time series, required only for Multivariate models.
+
+In this example, we will use `h=12` and we use `ray` as backend. We will
+use the default hyperparameter space but only change `random_seed` range
+and `n_pool_kernel_size`.
+
+
+```python
+from ray import tune
+from neuralforecast.auto import AutoNHITS
+```
+
+
+```python
+nhits_config = AutoNHITS.get_default_config(h = 12, backend="ray") # Extract the default hyperparameter settings
+nhits_config["random_seed"] = tune.randint(1, 10) # Random seed
+nhits_config["n_pool_kernel_size"] = tune.choice([[2, 2, 2], [16, 8, 1]]) # MaxPool's Kernelsize
+```
+
+### 3.a.2 Custom hyperparameter grid
+
+More generally, users can define fully customized search spaces tailored
+for particular datasets and tasks, by fully specifying a hyperparameter
+search space dictionary.
+
+In the following example we are optimizing the `learning_rate` and two
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+specific hyperparameters: `n_pool_kernel_size` and `n_freq_downsample`.
+Additionaly, we use the search space to modify default hyperparameters,
+such as `max_steps` and `val_check_steps`.
+
+
+```python
+nhits_config = {
+ "max_steps": 100, # Number of SGD steps
+ "input_size": 24, # Size of input window
+ "learning_rate": tune.loguniform(1e-5, 1e-1), # Initial Learning rate
+ "n_pool_kernel_size": tune.choice([[2, 2, 2], [16, 8, 1]]), # MaxPool's Kernelsize
+ "n_freq_downsample": tune.choice([[168, 24, 1], [24, 12, 1], [1, 1, 1]]), # Interpolation expressivity ratios
+ "val_check_steps": 50, # Compute validation every 50 steps
+ "random_seed": tune.randint(1, 10), # Random seed
+}
+```
+
+> **Important**
+>
+> Configuration dictionaries are not interchangeable between models
+> since they have different hyperparameters. Refer to
+> https://nixtla.github.io/neuralforecast/models.html for a complete
+> list of each model’s hyperparameters.
+
+### 3.b Instantiate `Auto` model
+
+To instantiate an `Auto` model you need to define:
+
+- `h`: forecasting horizon.
+- `loss`: training and validation loss from
+ `neuralforecast.losses.pytorch`.
+- `config`: hyperparameter search space. If `None`, the `Auto` class
+ will use a pre-defined suggested hyperparameter space.
+- `search_alg`: search algorithm (from `tune.search`), default is
+ random search. Refer to
+ https://docs.ray.io/en/latest/tune/api_docs/suggestion.html for more
+ information on the different search algorithm options.
+- `backend`: backend to use, default is `ray`. If `optuna`, the `Auto`
+ class will use the `Optuna` backend.
+- `num_samples`: number of configurations explored.
+
+In this example we set horizon `h` as 12, use the
+[`MAE`](https://nixtlaverse.nixtla.io/neuralforecast/losses.pytorch.html#mae)
+loss for training and validation, and use the `HYPEROPT` search
+algorithm.
+
+
+```python
+from ray.tune.search.hyperopt import HyperOptSearch
+from neuralforecast.losses.pytorch import MAE
+from neuralforecast.auto import AutoNHITS
+```
+
+
+```python
+model = AutoNHITS(
+ h=12,
+ loss=MAE(),
+ config=nhits_config,
+ search_alg=HyperOptSearch(),
+ backend='ray',
+ num_samples=10,
+)
+```
+
+> **Tip**
+>
+> The number of samples, `num_samples`, is a crucial parameter! Larger
+> values will usually produce better results as we explore more
+> configurations in the search space, but it will increase training
+> times. Larger search spaces will usually require more samples. As a
+> general rule, we recommend setting `num_samples` higher than 20. We
+> set 10 in this example for demonstration purposes.
+
+### 3.c Train model and predict with `Core` class
+
+Next, we use the `Neuralforecast` class to train the `Auto` model. In
+this step, `Auto` models will automatically perform hyperparamter tuning
+training multiple models with different hyperparameters, producing the
+forecasts on the validation set, and evaluating them. The best
+configuration is selected based on the error on a validation set. Only
+the best model is stored and used during inference.
+
+
+```python
+from neuralforecast import NeuralForecast
+```
+
+Use the `val_size` parameter of the `fit` method to control the length
+of the validation set. In this case we set the validation set as twice
+the forecasting horizon.
+
+
+```python
+nf = NeuralForecast(models=[model], freq='ME')
+nf.fit(df=Y_df, val_size=24)
+```
+
+The results of the hyperparameter tuning are available in the `results`
+attribute of the `Auto` model. Use the `get_dataframe` method to get the
+results in a pandas dataframe.
+
+
+```python
+results = nf.models[0].results.get_dataframe()
+results.head()
+```
+
+| | loss | train_loss | timestamp | checkpoint_dir_name | done | training_iteration | trial_id | date | time_this_iter_s | time_total_s | ... | config/input_size | config/learning_rate | config/n_pool_kernel_size | config/n_freq_downsample | config/val_check_steps | config/random_seed | config/h | config/loss | config/valid_loss | logdir |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | 21.948565 | 11.748630 | 1732660404 | None | False | 2 | e684ab59 | 2024-11-26_22-33-24 | 0.473169 | 1.742914 | ... | 24 | 0.000583 | (16, 8, 1) | (1, 1, 1) | 50 | 9 | 12 | MAE() | MAE() | e684ab59 |
+| 1 | 23.497557 | 13.491600 | 1732660411 | None | False | 2 | 28016d96 | 2024-11-26_22-33-31 | 0.467711 | 1.767644 | ... | 24 | 0.000222 | (16, 8, 1) | (168, 24, 1) | 50 | 5 | 12 | MAE() | MAE() | 28016d96 |
+| 2 | 29.214516 | 16.968582 | 1732660419 | None | False | 2 | ded66a42 | 2024-11-26_22-33-39 | 0.969751 | 2.623766 | ... | 24 | 0.009816 | (16, 8, 1) | (24, 12, 1) | 50 | 5 | 12 | MAE() | MAE() | ded66a42 |
+| 3 | 45.178616 | 28.338690 | 1732660427 | None | False | 2 | 2964d41f | 2024-11-26_22-33-47 | 0.985556 | 2.656381 | ... | 24 | 0.012083 | (16, 8, 1) | (24, 12, 1) | 50 | 7 | 12 | MAE() | MAE() | 2964d41f |
+| 4 | 32.580570 | 21.667740 | 1732660434 | None | False | 2 | 766cc549 | 2024-11-26_22-33-54 | 0.418154 | 1.465539 | ... | 24 | 0.000040 | (2, 2, 2) | (1, 1, 1) | 50 | 4 | 12 | MAE() | MAE() | 766cc549 |
+
+Next, we use the `predict` method to forecast the next 12 months using
+the optimal hyperparameters.
+
+
+```python
+Y_hat_df = nf.predict()
+Y_hat_df.head()
+```
+
+``` text
+Predicting: | …
+```
+
+| | unique_id | ds | AutoNHITS |
+|-----|-----------|------------|------------|
+| 0 | 1.0 | 1961-01-31 | 438.724091 |
+| 1 | 1.0 | 1961-02-28 | 415.593628 |
+| 2 | 1.0 | 1961-03-31 | 493.484894 |
+| 3 | 1.0 | 1961-04-30 | 493.120728 |
+| 4 | 1.0 | 1961-05-31 | 499.806702 |
+
+## 4. `Optuna` backend
+
+In this section we show how to use the `Optuna` backend. `Optuna` is a
+lightweight and versatile platform for hyperparameter optimization. If
+you plan to use the `Tune` backend, you can skip this section.
+
+### 4.a Define hyperparameter grid
+
+Each `Auto` model contains a default search space that was extensively
+tested on multiple large-scale datasets. Search spaces are specified
+with a function that returns a dictionary, where keys corresponds to the
+model’s hyperparameter and the value is a `suggest` function to specify
+how the hyperparameter will be sampled. For example, use `suggest_int`
+to sample integers uniformly, and `suggest_categorical` to sample values
+of a list. See
+https://optuna.readthedocs.io/en/stable/reference/generated/optuna.trial.Trial.html
+for more details.
+
+### 4.a.1 Default hyperparameter grid
+
+The default search space dictionary can be accessed through the
+`get_default_config` function of the `Auto` model. This is useful if you
+wish to use the default parameter configuration but want to change one
+or more hyperparameter spaces without changing the other default values.
+
+To extract the default config, you need to define: \* `h`: forecasting
+horizon. \* `backend`: backend to use. \* `n_series`: Optional, the
+number of unique time series, required only for Multivariate models.
+
+In this example, we will use `h=12` and we use `optuna` as backend. We
+will use the default hyperparameter space but only change `random_seed`
+range and `n_pool_kernel_size`.
+
+
+```python
+import optuna
+```
+
+
+```python
+optuna.logging.set_verbosity(optuna.logging.WARNING) # Use this to disable training prints from optuna
+nhits_default_config = AutoNHITS.get_default_config(h = 12, backend="optuna") # Extract the default hyperparameter settings
+
+def config_nhits(trial):
+ config = {**nhits_default_config(trial)}
+ config.update({
+ "random_seed": trial.suggest_int("random_seed", 1, 10),
+ "n_pool_kernel_size": trial.suggest_categorical("n_pool_kernel_size", [[2, 2, 2], [16, 8, 1]])
+ })
+ return config
+```
+
+### 3.a.2 Custom hyperparameter grid
+
+More generally, users can define fully customized search spaces tailored
+for particular datasets and tasks, by fully specifying a hyperparameter
+search space function.
+
+In the following example we are optimizing the `learning_rate` and two
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+specific hyperparameters: `n_pool_kernel_size` and `n_freq_downsample`.
+Additionaly, we use the search space to modify default hyperparameters,
+such as `max_steps` and `val_check_steps`.
+
+
+```python
+def config_nhits(trial):
+ return {
+ "max_steps": 100, # Number of SGD steps
+ "input_size": 24, # Size of input window
+ "learning_rate": trial.suggest_loguniform("learning_rate", 1e-5, 1e-1), # Initial Learning rate
+ "n_pool_kernel_size": trial.suggest_categorical("n_pool_kernel_size", [[2, 2, 2], [16, 8, 1]]), # MaxPool's Kernelsize
+ "n_freq_downsample": trial.suggest_categorical("n_freq_downsample", [[168, 24, 1], [24, 12, 1], [1, 1, 1]]), # Interpolation expressivity ratios
+ "val_check_steps": 50, # Compute validation every 50 steps
+ "random_seed": trial.suggest_int("random_seed", 1, 10), # Random seed
+ }
+```
+
+### 4.b Instantiate `Auto` model
+
+To instantiate an `Auto` model you need to define:
+
+- `h`: forecasting horizon.
+- `loss`: training and validation loss from
+ `neuralforecast.losses.pytorch`.
+- `config`: hyperparameter search space. If `None`, the `Auto` class
+ will use a pre-defined suggested hyperparameter space.
+- `search_alg`: search algorithm (from `optuna.samplers`), default is
+ TPESampler (Tree-structured Parzen Estimator). Refer to
+ https://optuna.readthedocs.io/en/stable/reference/samplers/index.html
+ for more information on the different search algorithm options.
+- `backend`: backend to use, default is `ray`. If `optuna`, the `Auto`
+ class will use the `Optuna` backend.
+- `num_samples`: number of configurations explored.
+
+
+```python
+model = AutoNHITS(
+ h=12,
+ loss=MAE(),
+ config=config_nhits,
+ search_alg=optuna.samplers.TPESampler(seed=0),
+ backend='optuna',
+ num_samples=10,
+)
+```
+
+> **Important**
+>
+> Configuration dictionaries and search algorithms for `Tune` and
+> `Optuna` are not interchangeable! Use the appropriate type of search
+> algorithm and custom configuration dictionary for each backend.
+
+### 4.c Train model and predict with `Core` class
+
+Use the `val_size` parameter of the `fit` method to control the length
+of the validation set. In this case we set the validation set as twice
+the forecasting horizon.
+
+
+```python
+nf = NeuralForecast(models=[model], freq='ME')
+nf.fit(df=Y_df, val_size=24)
+```
+
+The results of the hyperparameter tuning are available in the `results`
+attribute of the `Auto` model. Use the `trials_dataframe` method to get
+the results in a pandas dataframe.
+
+
+```python
+results = nf.models[0].results.trials_dataframe()
+results.drop(columns='user_attrs_ALL_PARAMS')
+```
+
+| | number | value | datetime_start | datetime_complete | duration | params_learning_rate | params_n_freq_downsample | params_n_pool_kernel_size | params_random_seed | user_attrs_METRICS | state |
+|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | 0 | 1.827570e+01 | 2024-11-26 22:34:29.382448 | 2024-11-26 22:34:30.773811 | 0 days 00:00:01.391363 | 0.001568 | \[1, 1, 1\] | \[2, 2, 2\] | 5 | \{'loss': tensor(18.2757), 'train_loss': tensor... | COMPLETE |
+| 1 | 1 | 9.055198e+06 | 2024-11-26 22:34:30.774153 | 2024-11-26 22:34:32.090132 | 0 days 00:00:01.315979 | 0.036906 | \[168, 24, 1\] | \[2, 2, 2\] | 10 | \{'loss': tensor(9055198.), 'train_loss': tenso... | COMPLETE |
+| 2 | 2 | 5.554298e+01 | 2024-11-26 22:34:32.090466 | 2024-11-26 22:34:33.425103 | 0 days 00:00:01.334637 | 0.000019 | \[1, 1, 1\] | \[2, 2, 2\] | 10 | \{'loss': tensor(55.5430), 'train_loss': tensor... | COMPLETE |
+| 3 | 3 | 9.857751e+01 | 2024-11-26 22:34:33.425460 | 2024-11-26 22:34:34.962057 | 0 days 00:00:01.536597 | 0.015727 | \[24, 12, 1\] | \[16, 8, 1\] | 10 | \{'loss': tensor(98.5775), 'train_loss': tensor... | COMPLETE |
+| 4 | 4 | 1.966841e+01 | 2024-11-26 22:34:34.962357 | 2024-11-26 22:34:36.951450 | 0 days 00:00:01.989093 | 0.001223 | \[168, 24, 1\] | \[2, 2, 2\] | 1 | \{'loss': tensor(19.6684), 'train_loss': tensor... | COMPLETE |
+| 5 | 5 | 1.524971e+01 | 2024-11-26 22:34:36.951775 | 2024-11-26 22:34:38.280982 | 0 days 00:00:01.329207 | 0.002955 | \[168, 24, 1\] | \[16, 8, 1\] | 5 | \{'loss': tensor(15.2497), 'train_loss': tensor... | COMPLETE |
+| 6 | 6 | 1.678810e+01 | 2024-11-26 22:34:38.281381 | 2024-11-26 22:34:39.648595 | 0 days 00:00:01.367214 | 0.006173 | \[168, 24, 1\] | \[16, 8, 1\] | 4 | \{'loss': tensor(16.7881), 'train_loss': tensor... | COMPLETE |
+| 7 | 7 | 2.014485e+01 | 2024-11-26 22:34:39.649025 | 2024-11-26 22:34:41.075568 | 0 days 00:00:01.426543 | 0.000285 | \[168, 24, 1\] | \[2, 2, 2\] | 2 | \{'loss': tensor(20.1448), 'train_loss': tensor... | COMPLETE |
+| 8 | 8 | 2.109382e+01 | 2024-11-26 22:34:41.075891 | 2024-11-26 22:34:42.449451 | 0 days 00:00:01.373560 | 0.004097 | \[168, 24, 1\] | \[16, 8, 1\] | 7 | \{'loss': tensor(21.0938), 'train_loss': tensor... | COMPLETE |
+| 9 | 9 | 5.091650e+01 | 2024-11-26 22:34:42.449762 | 2024-11-26 22:34:43.804981 | 0 days 00:00:01.355219 | 0.000036 | \[1, 1, 1\] | \[16, 8, 1\] | 1 | \{'loss': tensor(50.9165), 'train_loss': tensor... | COMPLETE |
+
+Next, we use the `predict` method to forecast the next 12 months using
+the optimal hyperparameters.
+
+
+```python
+Y_hat_df_optuna = nf.predict()
+Y_hat_df_optuna.head()
+```
+
+``` text
+Predicting: | …
+```
+
+| | unique_id | ds | AutoNHITS |
+|-----|-----------|------------|------------|
+| 0 | 1.0 | 1961-01-31 | 446.410736 |
+| 1 | 1.0 | 1961-02-28 | 422.048523 |
+| 2 | 1.0 | 1961-03-31 | 508.271515 |
+| 3 | 1.0 | 1961-04-30 | 496.549133 |
+| 4 | 1.0 | 1961-05-31 | 506.865723 |
+
+## 5. Plots
+
+Finally, we compare the forecasts produced by the
+[`AutoNHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autonhits)
+model with both backends.
+
+
+```python
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+plot_series(
+ Y_df,
+ Y_hat_df.merge(
+ Y_hat_df_optuna,
+ on=['unique_id', 'ds'],
+ suffixes=['_ray', '_optuna'],
+ ),
+)
+```
+
+
+
+### References
+
+- [Cristian Challu, Kin G. Olivares, Boris N. Oreshkin, Federico
+ Garza, Max Mergenthaler-Canseco, Artur Dubrawski (2021). NHITS:
+ Neural Hierarchical Interpolation for Time Series Forecasting.
+ Accepted at AAAI 2023.](https://arxiv.org/abs/2201.12886)
+- [James Bergstra, Remi Bardenet, Yoshua Bengio, and Balazs Kegl
+ (2011). “Algorithms for Hyper-Parameter Optimization”. In: Advances
+ in Neural Information Processing Systems. url:
+ https://proceedings.neurips.cc/paper/2011/file/86e8f7ab32cfd12577bc2619bc635690-Paper.pdf](https://proceedings.neurips.cc/paper/2011/file/86e8f7ab32cfd12577bc2619bc635690-Paper.pdf)
+- [Kirthevasan Kandasamy, Karun Raju Vysyaraju, Willie Neiswanger,
+ Biswajit Paria, Christopher R. Collins, Jeff Schneider, Barnabas
+ Poczos, Eric P. Xing (2019). “Tuning Hyperparameters without Grad
+ Students: Scalable and Robust Bayesian Optimisation with Dragonfly”.
+ Journal of Machine Learning Research. url:
+ https://arxiv.org/abs/1903.06694](https://arxiv.org/abs/1903.06694)
+- [Lisha Li, Kevin Jamieson, Giulia DeSalvo, Afshin Rostamizadeh,
+ Ameet Talwalkar (2016). “Hyperband: A Novel Bandit-Based Approach to
+ Hyperparameter Optimization”. Journal of Machine Learning Research.
+ url:
+ https://arxiv.org/abs/1603.06560](https://arxiv.org/abs/1603.06560)
+
diff --git a/neuralforecast/docs/capabilities/objectives.html.mdx b/neuralforecast/docs/capabilities/objectives.html.mdx
new file mode 100644
index 00000000..c5e1b44e
--- /dev/null
+++ b/neuralforecast/docs/capabilities/objectives.html.mdx
@@ -0,0 +1,31 @@
+---
+output-file: objectives.html
+title: Optimization Objectives
+---
+
+
+NeuralForecast is a highly modular framework capable of augmenting a
+wide variety of robust neural network architectures with different point
+or probability outputs as defined by their optimization objectives.
+
+## Point losses
+
+| Scale-Dependent | Percentage-Errors | Scale-Independent | Robust |
+|:-----------------|:-------------------|:-----------------|:---------------|
+| [**MAE**](../../losses.pytorch.html#mean-absolute-error-mae) | [**MAPE**](../../losses.pytorch.html#mean-absolute-percentage-error-mape) | [**MASE**](../../losses.pytorch.html#mean-absolute-scaled-error-mase) | [**Huber**](../losses.pytorch.html#huber-loss) |
+| [**MSE**](../../losses.pytorch.html#mean-squared-error-mse) | [**sMAPE**](../../losses.pytorch.html#symmetric-mape-smape) | | [**Tukey**](../../losses.pytorch.html#tukey-loss) |
+| [**RMSE**](../../losses.pytorch.html#root-mean-squared-error-rmse) | | | [**HuberMQLoss**](../../losses.pytorch.html#huberized-mqloss) |
+
+## Probabilistic losses
+
+| Parametric Probabilities | Non-Parametric Probabilities |
+|:-----------------------------------|:-----------------------------------|
+| [**Normal**](../../losses.pytorch.html#distributionloss) | [**QuantileLoss**](../../losses.pytorch.html#quantile-loss) |
+| [**StudenT**](../../losses.pytorch.html#distributionloss) | [**MQLoss**](../../losses.pytorch.html#multi-quantile-loss-mqloss) |
+| [**Poisson**](../../losses.pytorch.html#distributionloss) | [**HuberQLoss**](../../losses.pytorch.html#huberized-quantile-loss) |
+| [**Negative Binomial**](../../losses.pytorch.html#distributionloss) | [**HuberMQLoss**](../../losses.pytorch.html#huberized-mqloss) |
+| [**Tweedie**](../../losses.pytorch.html#distributionloss) | [**IQLoss**](../../losses.pytorch.html#iqloss) |
+| [**PMM**](../../losses.pytorch.html#poisson-mixture-mesh-pmm) | [**HuberIQLoss**](../../losses.pytorch.html#huberized-iqloss) |
+| [**GMM**](../../losses.pytorch.html#gaussian-mixture-mesh-gmm) | [**ISQF**](../../losses.pytorch.html#isqf) |
+| [**NBMM**](../../losses.pytorch.html#negative-binomial-mixture-mesh-nbmm) | |
+
diff --git a/neuralforecast/docs/capabilities/overview.html.mdx b/neuralforecast/docs/capabilities/overview.html.mdx
new file mode 100644
index 00000000..fa3dc2f2
--- /dev/null
+++ b/neuralforecast/docs/capabilities/overview.html.mdx
@@ -0,0 +1,82 @@
+---
+output-file: overview.html
+title: Forecasting Models
+---
+
+
+NeuralForecast currently offers the following models.
+
+| Model1 | AutoModel2 | Family3 | Univariate / Multivariate4 | Forecast Type5 | Exogenous6 |
+|:------|:--------|:-----------|:--------------------|:-----------|:-------------|
+| [`Autoformer`](https://nixtlaverse.nixtla.io/neuralforecast/models.autoformer.html#autoformer) | [`AutoAutoformer`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autoautoformer) | Transformer | Univariate | Direct | F |
+| [`BiTCN`](https://nixtlaverse.nixtla.io/neuralforecast/models.bitcn.html#bitcn) | [`AutoBiTCN`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autobitcn) | CNN | Univariate | Direct | F/H/S |
+| [`DeepAR`](https://nixtlaverse.nixtla.io/neuralforecast/models.deepar.html#deepar) | [`AutoDeepAR`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autodeepar) | RNN | Univariate | Direct | F/S |
+| [`DeepNPTS`](https://nixtlaverse.nixtla.io/neuralforecast/models.deepnpts.html#deepnpts) | [`AutoDeepNPTS`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autodeepnpts) | MLP | Univariate | Direct | F/H/S |
+| [`DilatedRNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.dilated_rnn.html#dilatedrnn) | [`AutoDilatedRNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autodilatedrnn) | RNN | Univariate | Direct | F/H/S |
+| [`FEDformer`](https://nixtlaverse.nixtla.io/neuralforecast/models.fedformer.html#fedformer) | [`AutoFEDformer`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autofedformer) | Transformer | Univariate | Direct | F |
+| [`GRU`](https://nixtlaverse.nixtla.io/neuralforecast/models.gru.html#gru) | [`AutoGRU`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autogru) | RNN | Univariate | Both8 | F/H/S |
+| [`HINT`](https://nixtlaverse.nixtla.io/neuralforecast/models.hint.html#hint) | [`AutoHINT`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autohint) | Any7 | Both7 | Both7 | F/H/S |
+| [`Informer`](https://nixtlaverse.nixtla.io/neuralforecast/models.informer.html#informer) | [`AutoInformer`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autoinformer) | Transformer | Univariate | Direct | F |
+| [`iTransformer`](https://nixtlaverse.nixtla.io/neuralforecast/models.itransformer.html#itransformer) | [`AutoiTransformer`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autoitransformer) | Transformer | Multivariate | Direct | \- |
+| [`KAN`](https://nixtlaverse.nixtla.io/neuralforecast/models.kan.html#kan) | [`AutoKAN`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autokan) | KAN | Univariate | Direct | F/H/S |
+| [`LSTM`](https://nixtlaverse.nixtla.io/neuralforecast/models.lstm.html#lstm) | [`AutoLSTM`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autolstm) | RNN | Univariate | Both8 | F/H/S |
+| [`MLP`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlp.html#mlp) | [`AutoMLP`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#automlp) | MLP | Univariate | Direct | F/H/S |
+| [`MLPMultivariate`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlpmultivariate.html#mlpmultivariate) | [`AutoMLPMultivariate`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#automlpmultivariate) | MLP | Multivariate | Direct | F/H/S |
+| [`NBEATS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nbeats.html#nbeats) | [`AutoNBEATS`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autonbeats) | MLP | Univariate | Direct | \- |
+| [`NBEATSx`](https://nixtlaverse.nixtla.io/neuralforecast/models.nbeatsx.html#nbeatsx) | [`AutoNBEATSx`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autonbeatsx) | MLP | Univariate | Direct | F/H/S |
+| [`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits) | [`AutoNHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autonhits) | MLP | Univariate | Direct | F/H/S |
+| [`NLinear`](https://nixtlaverse.nixtla.io/neuralforecast/models.nlinear.html#nlinear) | [`AutoNLinear`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autonlinear) | MLP | Univariate | Direct | \- |
+| [`PatchTST`](https://nixtlaverse.nixtla.io/neuralforecast/models.patchtst.html#patchtst) | [`AutoPatchTST`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autopatchtst) | Transformer | Univariate | Direct | \- |
+| [`RMoK`](https://nixtlaverse.nixtla.io/neuralforecast/models.rmok.html#rmok) | [`AutoRMoK`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autormok) | KAN | Multivariate | Direct | \- |
+| [`RNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.rnn.html#rnn) | [`AutoRNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autornn) | RNN | Univariate | Both8 | F/H/S |
+| [`SOFTS`](https://nixtlaverse.nixtla.io/neuralforecast/models.softs.html#softs) | [`AutoSOFTS`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autosofts) | MLP | Multivariate | Direct | \- |
+| [`StemGNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.stemgnn.html#stemgnn) | [`AutoStemGNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autostemgnn) | GNN | Multivariate | Direct | \- |
+| [`TCN`](https://nixtlaverse.nixtla.io/neuralforecast/models.tcn.html#tcn) | [`AutoTCN`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotcn) | CNN | Univariate | Direct | F/H/S |
+| [`TFT`](https://nixtlaverse.nixtla.io/neuralforecast/models.tft.html#tft) | [`AutoTFT`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotft) | Transformer | Univariate | Direct | F/H/S |
+| [`TiDE`](https://nixtlaverse.nixtla.io/neuralforecast/models.tide.html#tide) | [`AutoTiDE`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotide) | MLP | Univariate | Direct | F/H/S |
+| [`TimeMixer`](https://nixtlaverse.nixtla.io/neuralforecast/models.timemixer.html#timemixer) | [`AutoTimeMixer`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotimemixer) | MLP | Multivariate | Direct | \- |
+| [`TimeLLM`](https://nixtlaverse.nixtla.io/neuralforecast/models.timellm.html#timellm) | \- | LLM | Univariate | Direct | \- |
+| [`TimesNet`](https://nixtlaverse.nixtla.io/neuralforecast/models.timesnet.html#timesnet) | [`AutoTimesNet`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotimesnet) | CNN | Univariate | Direct | F |
+| [`TimeXer`](https://nixtlaverse.nixtla.io/neuralforecast/models.timexer.html#timexer) | [`AutoTimeXer`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotimexer) | Transformer | Multivariate | Direct | F |
+| [`TSMixer`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixer.html#tsmixer) | [`AutoTSMixer`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotsmixer) | MLP | Multivariate | Direct | \- |
+| [`TSMixerx`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixerx.html#tsmixerx) | [`AutoTSMixerx`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotsmixerx) | MLP | Multivariate | Direct | F/H/S |
+| [`VanillaTransformer`](https://nixtlaverse.nixtla.io/neuralforecast/models.vanillatransformer.html#vanillatransformer) | [`AutoVanillaTransformer`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autovanillatransformer) | Transformer | Univariate | Direct | F |
+
+1. **Model**: The model name.
+2. **AutoModel**: NeuralForecast offers most models also in an Auto\*
+ version, in which the hyperparameters of the underlying model are
+ automatically optimized and the best-performing model for a
+ validation set is selected. The optimization methods include grid
+ search, random search, and Bayesian optimization.
+3. **Family**: The main neural network architecture underpinning the
+ model.
+4. **Univariate / Multivariate**: A multivariate model explicitly
+ models the interactions between multiple time series in a dataset
+ and will provide predictions for multiple time series concurrently.
+ In contrast, a univariate model trained on multiple time series
+ implicitly models interactions between multiple time series and
+ provides predictions for single time series concurrently.
+ Multivariate models are typically computationally expensive and
+ empirically do not necessarily offer better forecasting performance
+ compared to using a univariate model.
+5. **Forecast Type**: Direct forecast models are models that produce
+ all steps in the forecast horizon at once. In contrast, recursive
+ forecast models predict one-step ahead, and subsequently use the
+ prediction to compute the next step in the forecast horizon, and so
+ forth. Direct forecast models typically suffer less from bias and
+ variance propagation as compared to recursive forecast models,
+ whereas recursive models can be computationally less expensive.
+6. **Exogenous**: Whether the model accepts exogenous variables. This
+ can be exogenous variables that contain information about the past
+ and future (F), about the past only (*historical*, H), or that
+ contain static information (*static*, S).
+7. **HINT** is a modular framework that can combine any type of neural
+ architecture with task-specialized mixture probability and advanced
+ hierarchical reconciliation strategies.
+8. Models that can produce forecasts recursively and direct. For
+ example, the RNN model uses an RNN to encode the past sequence, and
+ subsequently the user can choose between producing forecasts
+ recursively using the RNN or direct using an MLP that uses the
+ encoded sequence as input. The models feature an `recursive=False`
+ feature that sets how they produce forecasts.
+
diff --git a/neuralforecast/docs/capabilities/predictinsample.html.mdx b/neuralforecast/docs/capabilities/predictinsample.html.mdx
new file mode 100644
index 00000000..d06401f5
--- /dev/null
+++ b/neuralforecast/docs/capabilities/predictinsample.html.mdx
@@ -0,0 +1,229 @@
+---
+description: Tutorial on how to produce insample predictions.
+output-file: predictinsample.html
+title: Predict Insample
+---
+
+
+This tutorial provides and example on how to use the `predict_insample`
+function of the `core` class to produce forecasts of the train and
+validation sets. In this example we will train the
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+model on the AirPassengers data, and show how to recover the insample
+predictions after model is fitted.
+
+*Predict Insample*: The process of producing forecasts of the train and
+validation sets.
+
+*Use Cases*: \* Debugging: producing insample predictions is useful for
+debugging purposes. For example, to check if the model is able to fit
+the train set. \* Training convergence: check if the the model has
+converged. \* Anomaly detection: insample predictions can be used to
+detect anomalous behavior in the train set (e.g. outliers). (Note: if a
+model is too flexible it might be able to perfectly forecast outliers)
+
+You can run these experiments using GPU with Google Colab.
+
+
+
+## 1. Installing NeuralForecast
+
+
+```python
+!pip install neuralforecast
+```
+
+## 2. Loading AirPassengers Data
+
+The `core.NeuralForecast` class contains shared, `fit`, `predict` and
+other methods that take as inputs pandas DataFrames with columns
+`['unique_id', 'ds', 'y']`, where `unique_id` identifies individual time
+series from the dataset, `ds` is the date, and `y` is the target
+variable.
+
+In this example dataset consists of a set of a single series, but you
+can easily fit your model to larger datasets in long format.
+
+
+```python
+from neuralforecast.utils import AirPassengersPanel
+```
+
+
+```python
+Y_df = AirPassengersPanel
+Y_df.head()
+```
+
+| | unique_id | ds | y | trend | y\_\[lag12\] |
+|-----|-----------|------------|-------|-------|--------------|
+| 0 | Airline1 | 1949-01-31 | 112.0 | 0 | 112.0 |
+| 1 | Airline1 | 1949-02-28 | 118.0 | 1 | 118.0 |
+| 2 | Airline1 | 1949-03-31 | 132.0 | 2 | 132.0 |
+| 3 | Airline1 | 1949-04-30 | 129.0 | 3 | 129.0 |
+| 4 | Airline1 | 1949-05-31 | 121.0 | 4 | 121.0 |
+
+## 3. Model Training
+
+First, we train the
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+models on the AirPassengers data. We will use the `fit` method of the
+`core` class to train the models.
+
+
+```python
+import logging
+import pandas as pd
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import NHITS, LSTM
+```
+
+
+```python
+logging.getLogger('pytorch_lightning').setLevel(logging.ERROR)
+```
+
+
+```python
+horizon = 12
+
+# Try different hyperparameters to improve accuracy.
+models = [NHITS(h=horizon, # Forecast horizon
+ input_size=2 * horizon, # Length of input sequence
+ max_steps=100, # Number of steps to train
+ n_freq_downsample=[2, 1, 1], # Downsampling factors for each stack output
+ mlp_units = 3 * [[1024, 1024]],
+ ) # Number of units in each block.
+ ]
+nf = NeuralForecast(models=models, freq='ME')
+nf.fit(df=Y_df, val_size=horizon)
+```
+
+## 4. Predict Insample
+
+Using the
+[`NeuralForecast.predict_insample`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast.predict_insample)
+method you can obtain the forecasts for the train and validation sets
+after the models are fitted. The function will always take the last
+dataset used for training in either the `fit` or `cross_validation`
+methods.
+
+With the `step_size` parameter you can specify the step size between
+consecutive windows to produce the forecasts. In this example we will
+set `step_size=horizon` to produce non-overlapping forecasts.
+
+The following diagram shows how the forecasts are produced based on the
+`step_size` parameter and `h` (horizon) of the model. In the diagram we
+set `step_size=2` and `h=4`.
+
+
+
+
+```python
+Y_hat_insample = nf.predict_insample(step_size=horizon)
+```
+
+The `predict_insample` function returns a pandas DataFrame with the
+following columns: \* `unique_id`: the unique identifier of the time
+series. \* `ds`: the datestamp of the forecast for each row. \*
+`cutoff`: the datestamp at which the forecast was made. \* `y`: the
+actual value of the target variable. \* `model_name`: the forecasted
+values for the models. In this case,
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits).
+
+
+```python
+Y_hat_insample.head()
+```
+
+| | unique_id | ds | cutoff | NHITS | y |
+|-----|-----------|------------|------------|----------|-------|
+| 0 | Airline1 | 1949-01-31 | 1948-12-31 | 0.064625 | 112.0 |
+| 1 | Airline1 | 1949-02-28 | 1948-12-31 | 0.074300 | 118.0 |
+| 2 | Airline1 | 1949-03-31 | 1948-12-31 | 0.133020 | 132.0 |
+| 3 | Airline1 | 1949-04-30 | 1948-12-31 | 0.221040 | 129.0 |
+| 4 | Airline1 | 1949-05-31 | 1948-12-31 | 0.176580 | 121.0 |
+
+> **Important**
+>
+> The function will produce forecasts from the first timestamp of the
+> time series. For these initial timestamps, the forecasts might not be
+> accurate given that models have very limited input information to
+> produce forecasts.
+
+## 5. Plot Predictions
+
+Finally, we plot the forecasts for the train and validation sets.
+
+
+```python
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+plot_series(forecasts_df=Y_hat_insample.drop(columns='cutoff'))
+```
+
+
+
+## 6. Insample predictions with prediction intervals
+
+We can also show insample prediction intervals for models trained with a
+distribution loss function. This can be achieved by simply specifying
+the required level in the `predict_insample` function.
+
+Note that the following settings are not yet supported: - Prediction
+intervals on insample predictions on models trained with conformal
+prediction intervals (e.g. a model trained with MAE and conformal
+prediction intervals); - Prediction intervals on insample predictions on
+multivariate models (e.g. a TSMixer model).
+
+
+```python
+from neuralforecast.losses.pytorch import DistributionLoss, GMM
+```
+
+
+```python
+horizon = 12
+
+# Try different hyperparameters to improve accuracy.
+models = [
+ NHITS(h=horizon,
+ input_size=2 * horizon,
+ loss=DistributionLoss(distribution="Poisson", num_samples=50),
+ max_steps=100,
+ scaler_type="robust",
+ ),
+ LSTM(h=horizon,
+ input_size=2 * horizon,
+ loss=GMM(),
+ max_steps=500,
+ scaler_type="robust",
+ ),
+ ]
+nf = NeuralForecast(models=models, freq='ME')
+nf.fit(df=Y_df, val_size=horizon)
+
+Y_hat_insample = nf.predict_insample(
+ step_size=horizon,
+ level=[80],
+)
+```
+
+
+```python
+plot_series(forecasts_df=Y_hat_insample.drop(columns=['cutoff']), level=[80])
+```
+
+
+
+## References
+
+- [Cristian Challu, Kin G. Olivares, Boris N. Oreshkin, Federico
+ Garza, Max Mergenthaler-Canseco, Artur Dubrawski (2021). NHITS:
+ Neural Hierarchical Interpolation for Time Series Forecasting.
+ Accepted at AAAI 2023.](https://arxiv.org/abs/2201.12886)
+
diff --git a/neuralforecast/docs/capabilities/save_load_models.html.mdx b/neuralforecast/docs/capabilities/save_load_models.html.mdx
new file mode 100644
index 00000000..07e7f9bb
--- /dev/null
+++ b/neuralforecast/docs/capabilities/save_load_models.html.mdx
@@ -0,0 +1,263 @@
+---
+output-file: save_load_models.html
+title: Save and Load Models
+---
+
+
+Saving and loading trained Deep Learning models has multiple valuable
+uses. These models are often costly to train; storing a pre-trained
+model can help reduce costs as it can be loaded and reused to forecast
+multiple times. Moreover, it enables Transfer learning capabilities,
+consisting of pre-training a flexible model on a large dataset and using
+it later on other data with little to no training. It is one of the most
+outstanding 🚀 achievements in Machine Learning 🧠 and has many
+practical applications.
+
+In this notebook we show an example on how to save and load
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+models.
+
+The two methods to consider are: 1.
+[`NeuralForecast.save`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast.save):
+Saves models into disk, allows save dataset and config. 2.
+[`NeuralForecast.load`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast.load):
+Loads models from a given path.
+
+> **Important**
+>
+> This Guide assumes basic knowledge on the NeuralForecast library. For
+> a minimal example visit the [Getting
+> Started](../getting-started/quickstart.html) guide.
+
+You can run these experiments using GPU with Google Colab.
+
+
+
+## 1. Installing NeuralForecast
+
+
+```python
+!pip install neuralforecast
+```
+
+## 2. Loading AirPassengers Data
+
+For this example we will use the classical [AirPassenger Data
+set](https://www.kaggle.com/datasets/rakannimer/air-passengers). Import
+the pre-processed AirPassenger from `utils`.
+
+
+```python
+from neuralforecast.utils import AirPassengersDF
+```
+
+
+```python
+Y_df = AirPassengersDF
+Y_df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|-------|
+| 0 | 1.0 | 1949-01-31 | 112.0 |
+| 1 | 1.0 | 1949-02-28 | 118.0 |
+| 2 | 1.0 | 1949-03-31 | 132.0 |
+| 3 | 1.0 | 1949-04-30 | 129.0 |
+| 4 | 1.0 | 1949-05-31 | 121.0 |
+
+## 3. Model Training
+
+Next, we instantiate and train three models:
+[`NBEATS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nbeats.html#nbeats),
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits),
+and
+[`AutoMLP`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#automlp).
+The models with their hyperparameters are defined in the `models` list.
+
+
+```python
+import logging
+
+from ray import tune
+
+from neuralforecast.core import NeuralForecast
+from neuralforecast.auto import AutoMLP
+from neuralforecast.models import NBEATS, NHITS
+```
+
+
+```python
+logging.getLogger('pytorch_lightning').setLevel(logging.ERROR)
+```
+
+
+```python
+horizon = 12
+models = [NBEATS(input_size=2 * horizon, h=horizon, max_steps=50),
+ NHITS(input_size=2 * horizon, h=horizon, max_steps=50),
+ AutoMLP(# Ray tune explore config
+ config=dict(max_steps=100, # Operates with steps not epochs
+ input_size=tune.choice([3*horizon]),
+ learning_rate=tune.choice([1e-3])),
+ h=horizon,
+ num_samples=1, cpus=1)]
+```
+
+``` text
+Seed set to 1
+Seed set to 1
+```
+
+
+```python
+nf = NeuralForecast(models=models, freq='ME')
+nf.fit(df=Y_df)
+```
+
+Produce the forecasts with the `predict` method.
+
+
+```python
+Y_hat_df = nf.predict()
+Y_hat_df.head()
+```
+
+``` text
+Predicting: | …
+```
+
+``` text
+Predicting: | …
+```
+
+``` text
+Predicting: | …
+```
+
+| | unique_id | ds | NBEATS | NHITS | AutoMLP |
+|-----|-----------|------------|------------|------------|------------|
+| 0 | 1.0 | 1961-01-31 | 446.882172 | 447.219238 | 454.914154 |
+| 1 | 1.0 | 1961-02-28 | 465.145813 | 464.558014 | 430.188446 |
+| 2 | 1.0 | 1961-03-31 | 469.978424 | 474.637238 | 458.478577 |
+| 3 | 1.0 | 1961-04-30 | 493.650665 | 502.670349 | 477.244507 |
+| 4 | 1.0 | 1961-05-31 | 537.569275 | 559.405212 | 522.252991 |
+
+We plot the forecasts for each model.
+
+
+```python
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+plot_series(Y_df, Y_hat_df)
+```
+
+
+
+## 4. Save models
+
+To save all the trained models use the `save` method. This method will
+save both the hyperparameters and the learnable weights (parameters).
+
+The `save` method has the following inputs:
+
+- `path`: directory where models will be saved.
+- `model_index`: optional list to specify which models to save. For
+ example, to only save the
+ [`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+ model use `model_index=[2]`.
+- `overwrite`: boolean to overwrite existing files in `path`. When
+ True, the method will only overwrite models with conflicting names.
+- `save_dataset`: boolean to save `Dataset` object with the dataset.
+
+
+```python
+nf.save(path='./checkpoints/test_run/',
+ model_index=None,
+ overwrite=True,
+ save_dataset=True)
+```
+
+For each model, two files are created and stored:
+
+- `[model_name]_[suffix].ckpt`: Pytorch Lightning checkpoint file with
+ the model parameters and hyperparameters.
+- `[model_name]_[suffix].pkl`: Dictionary with configuration
+ attributes.
+
+Where `model_name` corresponds to the name of the model in lowercase
+(eg. `nhits`). We use a numerical suffix to distinguish multiple models
+of each class. In this example the names will be `automlp_0`,
+`nbeats_0`, and `nhits_0`.
+
+> **Important**
+>
+> The `Auto` models will be stored as their base model. For example, the
+> [`AutoMLP`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#automlp)
+> trained above is stored as an
+> [`MLP`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlp.html#mlp)
+> model, with the best hyparparameters found during tuning.
+
+## 5. Load models
+
+Load the saved models with the `load` method, specifying the `path`, and
+use the new `nf2` object to produce forecasts.
+
+
+```python
+nf2 = NeuralForecast.load(path='./checkpoints/test_run/')
+Y_hat_df2 = nf2.predict()
+Y_hat_df2.head()
+```
+
+``` text
+Seed set to 1
+Seed set to 1
+Seed set to 1
+```
+
+``` text
+Predicting: | …
+```
+
+``` text
+Predicting: | …
+```
+
+``` text
+Predicting: | …
+```
+
+| | unique_id | ds | NHITS | NBEATS | AutoMLP |
+|-----|-----------|------------|------------|------------|------------|
+| 0 | 1.0 | 1961-01-31 | 447.219238 | 446.882172 | 454.914154 |
+| 1 | 1.0 | 1961-02-28 | 464.558014 | 465.145813 | 430.188446 |
+| 2 | 1.0 | 1961-03-31 | 474.637238 | 469.978424 | 458.478577 |
+| 3 | 1.0 | 1961-04-30 | 502.670349 | 493.650665 | 477.244507 |
+| 4 | 1.0 | 1961-05-31 | 559.405212 | 537.569275 | 522.252991 |
+
+Finally, plot the forecasts to confirm they are identical to the
+original forecasts.
+
+
+```python
+plot_series(Y_df, Y_hat_df2)
+```
+
+
+
+## References
+
+https://pytorch-lightning.readthedocs.io/en/stable/common/checkpointing_basic.html
+
+[Oreshkin, B. N., Carpov, D., Chapados, N., & Bengio, Y. (2019).
+N-BEATS: Neural basis expansion analysis for interpretable time series
+forecasting. ICLR 2020](https://arxiv.org/abs/1905.10437)
+
+[Cristian Challu, Kin G. Olivares, Boris N. Oreshkin, Federico Garza,
+Max Mergenthaler-Canseco, Artur Dubrawski (2021). N-HiTS: Neural
+Hierarchical Interpolation for Time Series Forecasting. Accepted at AAAI
+2023.](https://arxiv.org/abs/2201.12886)
+
diff --git a/neuralforecast/docs/capabilities/time_series_scaling.html.mdx b/neuralforecast/docs/capabilities/time_series_scaling.html.mdx
new file mode 100644
index 00000000..c1ab1bf4
--- /dev/null
+++ b/neuralforecast/docs/capabilities/time_series_scaling.html.mdx
@@ -0,0 +1,344 @@
+---
+output-file: time_series_scaling.html
+title: Time Series Scaling
+---
+
+
+Scaling time series data is an important preprocessing step when using
+neural forecasting methods for several reasons:
+
+1. **Convergence speed**: Neural forecasting models tend to converge
+ faster when the features are on a similar scale.
+2. **Avoiding vanishing or exploding gradients**: some architectures,
+ such as recurrent neural networks (RNNs), are sensitive to the scale
+ of input data. If the input values are too large, it could lead to
+ exploding gradients, where the gradients become too large and the
+ model becomes unstable. Conversely, very small input values could
+ lead to vanishing gradients, where weight updates during training
+ are negligible and the training fails to converge.
+3. **Ensuring consistent scale**: Neural forecasting models have shared
+ global parameters for the all time series of the task. In cases
+ where time series have different scale, scaling ensures that no
+ particular time series dominates the learning process.
+4. **Improving generalization**: time series with consistent scale can
+ lead to smoother loss surfaces. Moreover, scaling helps to
+ homogenize the distribution of the input data, which can also
+ improve generalization by avoiding out-of-range values.
+
+The `Neuralforecast` library integrates two types of temporal scaling:
+
+- **Time Series Scaling**: scaling each time series using all its data
+ on the train set before start training the model. This is done by
+ using the `local_scaler_type` parameter of the `Neuralforecast` core
+ class.
+- **Window scaling (TemporalNorm)**: scaling each input window
+ separetly for each element of the batch at every training iteration.
+ This is done by using the `scaler_type` parameter of each model
+ class.
+
+In this notebook, we will demonstrate how to scale the time series data
+with both methods on an Eletricity Price Forecasting (EPF) task.
+
+You can run these experiments using GPU with Google Colab.
+
+
+
+## 1. Install `Neuralforecast`
+
+
+```python
+!pip install neuralforecast
+!pip install hyperopt
+```
+
+## 2. Load Data
+
+The `df` dataframe contains the target and exogenous variables past
+information to train the model. The `unique_id` column identifies the
+markets, `ds` contains the datestamps, and `y` the electricity price.
+For future variables, we include a forecast of how much electricity will
+be produced (`gen_forecast`), and day of week (`week_day`). Both the
+electricity system demand and offer impact the price significantly,
+including these variables to the model greatly improve performance, as
+we demonstrate in Olivares et al. (2022).
+
+The `futr_df` dataframe includes the information of the future exogenous
+variables for the period we want to forecast (in this case, 24 hours
+after the end of the train dataset `df`).
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+```
+
+
+```python
+df = pd.read_csv(
+ 'https://datasets-nixtla.s3.amazonaws.com/EPF_FR_BE.csv',
+ parse_dates=['ds'],
+)
+futr_df = pd.read_csv(
+ 'https://datasets-nixtla.s3.amazonaws.com/EPF_FR_BE_futr.csv',
+ parse_dates=['ds'],
+)
+df.head()
+```
+
+| | unique_id | ds | y | gen_forecast | system_load | week_day |
+|-----|-----------|---------------------|-------|--------------|-------------|----------|
+| 0 | FR | 2015-01-01 00:00:00 | 53.48 | 76905.0 | 74812.0 | 3 |
+| 1 | FR | 2015-01-01 01:00:00 | 51.93 | 75492.0 | 71469.0 | 3 |
+| 2 | FR | 2015-01-01 02:00:00 | 48.76 | 74394.0 | 69642.0 | 3 |
+| 3 | FR | 2015-01-01 03:00:00 | 42.27 | 72639.0 | 66704.0 | 3 |
+| 4 | FR | 2015-01-01 04:00:00 | 38.41 | 69347.0 | 65051.0 | 3 |
+
+We can see that `y` and the exogenous variables are on largely different
+scales. Next, we show two methods to scale the data.
+
+## 3. Time Series Scaling with `Neuralforecast` class
+
+One of the most widely used approches for scaling time series is to
+treat it as a pre-processing step, where each time series and temporal
+exogenous variables are scaled based on their entire information in the
+train set. Models are then trained on the scaled data.
+
+To simplify pipelines, we added a scaling functionality to the
+`Neuralforecast` class. Each time series will be scaled before training
+the model with either `fit` or `cross_validation`, and scaling
+statistics are stored. The class then uses the stored statistics to
+scale the forecasts back to the original scale before returning the
+forecasts.
+
+### 3.a. Instantiate model and `Neuralforecast` class
+
+In this example we will use the
+[`TimesNet`](https://nixtlaverse.nixtla.io/neuralforecast/models.timesnet.html#timesnet)
+model, recently proposed in [Wu, Haixu, et
+al. (2022)](https://arxiv.org/abs/2210.02186). First instantiate the
+model with the desired parameters.
+
+
+```python
+import logging
+
+from neuralforecast.models import TimesNet
+from neuralforecast.core import NeuralForecast
+```
+
+
+```python
+logging.getLogger("pytorch_lightning").setLevel(logging.WARNING)
+```
+
+
+```python
+horizon = 24 # day-ahead daily forecast
+model = TimesNet(h = horizon, # Horizon
+ input_size = 5*horizon, # Length of input window
+ max_steps = 100, # Training iterations
+ top_k = 3, # Number of periods (for FFT).
+ num_kernels = 3, # Number of kernels for Inception module
+ batch_size = 2, # Number of time series per batch
+ windows_batch_size = 32, # Number of windows per batch
+ learning_rate = 0.001, # Learning rate
+ futr_exog_list = ['gen_forecast', 'week_day'], # Future exogenous variables
+ scaler_type = None) # We use the Core scaling method
+```
+
+``` text
+Seed set to 1
+```
+
+Fit the model by instantiating a
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+object and using the `fit` method. The `local_scaler_type` parameter is
+used to specify the type of scaling to be used. In this case, we will
+use `standard`, which scales the data to have zero mean and unit
+variance.Other supported scalers are `minmax`, `robust`, `robust-iqr`,
+`minmax`, and `boxcox`.
+
+
+```python
+nf = NeuralForecast(models=[model], freq='h', local_scaler_type='standard')
+nf.fit(df=df)
+```
+
+``` text
+Sanity Checking: | …
+```
+
+``` text
+Training: | …
+```
+
+``` text
+Validation: | …
+```
+
+### 3.b Forecast and plots
+
+Finally, use the `predict` method to forecast the day-ahead prices. The
+`Neuralforecast` class handles the inverse normalization, forecasts are
+returned in the original scale.
+
+
+```python
+Y_hat_df = nf.predict(futr_df=futr_df)
+Y_hat_df.head()
+```
+
+``` text
+Predicting: | …
+```
+
+| | unique_id | ds | TimesNet |
+|-----|-----------|---------------------|-----------|
+| 0 | BE | 2016-11-01 00:00:00 | 39.523182 |
+| 1 | BE | 2016-11-01 01:00:00 | 33.386608 |
+| 2 | BE | 2016-11-01 02:00:00 | 27.978468 |
+| 3 | BE | 2016-11-01 03:00:00 | 28.143955 |
+| 4 | BE | 2016-11-01 04:00:00 | 32.332230 |
+
+
+```python
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+plot_series(df, Y_hat_df, max_insample_length=24*5)
+```
+
+
+
+> **Important**
+>
+> The inverse scaling is performed by the `Neuralforecast` class before
+> returning the final forecasts. Therefore, the hyperparmater selection
+> with `Auto` models and validation loss for early stopping or model
+> selection are performed on the scaled data. Different types of scaling
+> with the `Neuralforecast` class can’t be automatically compared with
+> `Auto` models.
+
+## 4. Temporal Window normalization during training
+
+Temporal normalization scales each instance of the batch separately at
+the window level. It is performed at each training iteration for each
+window of the batch, for both target variable and temporal exogenous
+covariates. For more details, see [Olivares et
+al. (2023)](https://arxiv.org/abs/2305.07089) and
+https://nixtla.github.io/neuralforecast/common.scalers.html.
+
+### 4.a. Instantiate model and `Neuralforecast` class
+
+Temporal normalization is specified by the `scaler_type` argument.
+Currently, it is only supported for Windows-based models
+([`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits),
+[`NBEATS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nbeats.html#nbeats),
+[`MLP`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlp.html#mlp),
+[`TimesNet`](https://nixtlaverse.nixtla.io/neuralforecast/models.timesnet.html#timesnet),
+and all Transformers). In this example, we use the
+[`TimesNet`](https://nixtlaverse.nixtla.io/neuralforecast/models.timesnet.html#timesnet)
+model and `robust` scaler, recently proposed by Wu, Haixu, et
+al. (2022). First instantiate the model with the desired parameters.
+
+Visit https://nixtla.github.io/neuralforecast/common.scalers.html for a
+complete list of supported scalers.
+
+
+```python
+horizon = 24 # day-ahead daily forecast
+model = TimesNet(h = horizon, # Horizon
+ input_size = 5*horizon, # Length of input window
+ max_steps = 100, # Training iterations
+ top_k = 3, # Number of periods (for FFT).
+ num_kernels = 3, # Number of kernels for Inception module
+ batch_size = 2, # Number of time series per batch
+ windows_batch_size = 32, # Number of windows per batch
+ learning_rate = 0.001, # Learning rate
+ futr_exog_list = ['gen_forecast','week_day'], # Future exogenous variables
+ scaler_type = 'robust') # Robust scaling
+```
+
+``` text
+Seed set to 1
+```
+
+Fit the model by instantiating a
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+object and using the `fit` method. Note that `local_scaler_type` has
+`None` as default to avoid scaling the data before training.
+
+
+```python
+nf = NeuralForecast(models=[model], freq='h')
+nf.fit(df=df)
+```
+
+``` text
+Sanity Checking: | …
+```
+
+``` text
+Training: | …
+```
+
+``` text
+Validation: | …
+```
+
+### 4.b Forecast and plots
+
+Finally, use the `predict` method to forecast the day-ahead prices. The
+forecasts are returned in the original scale.
+
+
+```python
+Y_hat_df = nf.predict(futr_df=futr_df)
+Y_hat_df.head()
+```
+
+``` text
+Predicting: | …
+```
+
+| | unique_id | ds | TimesNet |
+|-----|-----------|---------------------|-----------|
+| 0 | BE | 2016-11-01 00:00:00 | 37.624653 |
+| 1 | BE | 2016-11-01 01:00:00 | 33.069824 |
+| 2 | BE | 2016-11-01 02:00:00 | 30.623751 |
+| 3 | BE | 2016-11-01 03:00:00 | 28.773439 |
+| 4 | BE | 2016-11-01 04:00:00 | 30.689444 |
+
+
+```python
+plot_series(df, Y_hat_df, max_insample_length=24*5)
+```
+
+
+
+> **Important**
+>
+> For most applications, models with temporal normalization (section 4)
+> produced more accurate forecasts than time series scaling (section 3).
+> However, with temporal normalization models lose the information of
+> the relative level between different windows. In some cases this
+> global information within time series is crucial, for instance when an
+> exogenous variables contains the dosage of a medication. In these
+> cases, time series scaling (section 3) is preferred.
+
+## References
+
+- [Kin G. Olivares, David Luo, Cristian Challu, Stefania La Vattiata,
+ Max Mergenthaler, Artur Dubrawski (2023). “HINT: Hierarchical
+ Mixture Networks For Coherent Probabilistic Forecasting”.
+ International Conference on Machine Learning (ICML). Workshop on
+ Structured Probabilistic Inference & Generative Modeling. Available
+ at
+ https://arxiv.org/abs/2305.07089.](https://arxiv.org/abs/2305.07089)
+- [Wu, Haixu, Tengge Hu, Yong Liu, Hang Zhou, Jianmin Wang, and
+ Mingsheng Long. “Timesnet: Temporal 2d-variation modeling for
+ general time series analysis.”, ICLR
+ 2023](https://openreview.net/forum?id=ju_Uqw384Oq)
+
diff --git a/neuralforecast/docs/getting-started/01_introduction_files/figure-markdown_strict/cell-4-output-2.png b/neuralforecast/docs/getting-started/01_introduction_files/figure-markdown_strict/cell-4-output-2.png
new file mode 100644
index 00000000..82961c7a
Binary files /dev/null and b/neuralforecast/docs/getting-started/01_introduction_files/figure-markdown_strict/cell-4-output-2.png differ
diff --git a/neuralforecast/docs/getting-started/02_quickstart_files/figure-markdown_strict/cell-11-output-1.png b/neuralforecast/docs/getting-started/02_quickstart_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..e4bb8a71
Binary files /dev/null and b/neuralforecast/docs/getting-started/02_quickstart_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/neuralforecast/docs/getting-started/datarequirements.html.mdx b/neuralforecast/docs/getting-started/datarequirements.html.mdx
new file mode 100644
index 00000000..306ef497
--- /dev/null
+++ b/neuralforecast/docs/getting-started/datarequirements.html.mdx
@@ -0,0 +1,135 @@
+---
+description: Dataset input requirments
+output-file: datarequirements.html
+title: Data Requirements
+---
+
+
+In this example we will go through the dataset input requirements of the
+`core.NeuralForecast` class.
+
+The `core.NeuralForecast` methods operate as global models that receive
+a set of time series rather than single series. The class uses
+cross-learning technique to fit flexible-shared models such as neural
+networks improving its generalization capabilities as shown by the M4
+international forecasting competition (Smyl 2019, Semenoglou 2021).
+
+You can run these experiments using GPU with Google Colab.
+
+
+
+## Long format
+
+### Multiple time series
+
+Store your time series in a pandas dataframe in long format, that is,
+each row represents an observation for a specific series and timestamp.
+Let’s see an example using the `datasetsforecast` library.
+
+`Y_df = pd.concat( [series1, series2, ...])`
+
+
+```python
+!pip install datasetsforecast
+```
+
+
+```python
+import pandas as pd
+from datasetsforecast.m3 import M3
+```
+
+
+```python
+Y_df, *_ = M3.load('./data', group='Yearly')
+```
+
+
+```python
+Y_df.groupby('unique_id').head(2)
+```
+
+| | unique_id | ds | y |
+|-------|-----------|------------|---------|
+| 0 | Y1 | 1975-12-31 | 940.66 |
+| 1 | Y1 | 1976-12-31 | 1084.86 |
+| 20 | Y10 | 1975-12-31 | 2160.04 |
+| 21 | Y10 | 1976-12-31 | 2553.48 |
+| 40 | Y100 | 1975-12-31 | 1424.70 |
+| ... | ... | ... | ... |
+| 18260 | Y97 | 1976-12-31 | 1618.91 |
+| 18279 | Y98 | 1975-12-31 | 1164.97 |
+| 18280 | Y98 | 1976-12-31 | 1277.87 |
+| 18299 | Y99 | 1975-12-31 | 1870.00 |
+| 18300 | Y99 | 1976-12-31 | 1307.20 |
+
+`Y_df` is a dataframe with three columns: `unique_id` with a unique
+identifier for each time series, a column `ds` with the datestamp and a
+column `y` with the values of the series.
+
+### Single time series
+
+If you have only one time series, you have to include the `unique_id`
+column. Consider, for example, the
+[AirPassengers](https://github.com/Nixtla/transfer-learning-time-series/blob/main/datasets/air_passengers.csv)
+dataset.
+
+
+```python
+Y_df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/air_passengers.csv')
+Y_df
+```
+
+| | timestamp | value |
+|-----|------------|-------|
+| 0 | 1949-01-01 | 112 |
+| 1 | 1949-02-01 | 118 |
+| 2 | 1949-03-01 | 132 |
+| 3 | 1949-04-01 | 129 |
+| 4 | 1949-05-01 | 121 |
+| ... | ... | ... |
+| 139 | 1960-08-01 | 606 |
+| 140 | 1960-09-01 | 508 |
+| 141 | 1960-10-01 | 461 |
+| 142 | 1960-11-01 | 390 |
+| 143 | 1960-12-01 | 432 |
+
+In this example `Y_df` only contains two columns: `timestamp`, and
+`value`. To use
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+we have to include the `unique_id` column and rename the previuos ones.
+
+
+```python
+Y_df['unique_id'] = 1. # We can add an integer as identifier
+Y_df = Y_df.rename(columns={'timestamp': 'ds', 'value': 'y'})
+Y_df = Y_df[['unique_id', 'ds', 'y']]
+Y_df
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|-----|
+| 0 | 1.0 | 1949-01-01 | 112 |
+| 1 | 1.0 | 1949-02-01 | 118 |
+| 2 | 1.0 | 1949-03-01 | 132 |
+| 3 | 1.0 | 1949-04-01 | 129 |
+| 4 | 1.0 | 1949-05-01 | 121 |
+| ... | ... | ... | ... |
+| 139 | 1.0 | 1960-08-01 | 606 |
+| 140 | 1.0 | 1960-09-01 | 508 |
+| 141 | 1.0 | 1960-10-01 | 461 |
+| 142 | 1.0 | 1960-11-01 | 390 |
+| 143 | 1.0 | 1960-12-01 | 432 |
+
+## References
+
+- [Slawek Smyl. (2019). “A hybrid method of exponential smoothing and
+ recurrent networks for time series forecasting”. International
+ Journal of
+ Forecasting.](https://www.sciencedirect.com/science/article/pii/S0169207019301153)
+- [Artemios-Anargyros Semenoglou, Evangelos Spiliotis, Spyros
+ Makridakis, and Vassilios Assimakopoulos. (2021). Investigating the
+ accuracy of cross-learning time series forecasting methods”.
+ International Journal of
+ Forecasting.](https://www.sciencedirect.com/science/article/pii/S0169207020301850)
+
diff --git a/neuralforecast/docs/getting-started/installation.html.mdx b/neuralforecast/docs/getting-started/installation.html.mdx
new file mode 100644
index 00000000..760cf36f
--- /dev/null
+++ b/neuralforecast/docs/getting-started/installation.html.mdx
@@ -0,0 +1,80 @@
+---
+description: Install NeuralForecast with pip or conda
+output-file: installation.html
+title: Installation
+---
+
+
+You can install the *released version* of
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+from the [Python package index](https://pypi.org) with:
+
+
+```shell
+pip install neuralforecast
+```
+
+or
+
+
+```shell
+conda install -c conda-forge neuralforecast
+```
+
+> **Tip**
+>
+> Neural Forecasting methods profit from using GPU computation. Be sure
+> to have Cuda installed.
+
+> **Warning**
+>
+> We are constantly updating neuralforecast, so we suggest fixing the
+> version to avoid issues. `pip install neuralforecast=="1.0.0"`
+
+> **Tip**
+>
+> We recommend installing your libraries inside a python virtual or
+> [conda
+> environment](https://docs.conda.io/projects/conda/en/latest/user-guide/install/macos.html).
+
+## Extras
+
+You can use the following extras to add optional functionality:
+
+- distributed training with spark: `pip install neuralforecast[spark]`
+- saving and loading from S3: `pip install neuralforecast[aws]`
+
+#### User our env (optional)
+
+If you don’t have a Conda environment and need tools like Numba, Pandas,
+NumPy, Jupyter, Tune, and Nbdev you can use ours by following these
+steps:
+
+1. Clone the NeuralForecast repo:
+
+
+```bash
+$ git clone https://github.com/Nixtla/neuralforecast.git && cd neuralforecast
+```
+
+1. Create the environment using the `environment.yml` file:
+
+
+```bash
+$ conda env create -f environment.yml
+```
+
+1. Activate the environment:
+
+
+```bash
+$ conda activate neuralforecast
+```
+
+1. Install NeuralForecast Dev
+
+
+```bash
+$ pip install -e ".[dev]"
+```
+
diff --git a/neuralforecast/docs/getting-started/introduction.html.mdx b/neuralforecast/docs/getting-started/introduction.html.mdx
new file mode 100644
index 00000000..cced67e8
--- /dev/null
+++ b/neuralforecast/docs/getting-started/introduction.html.mdx
@@ -0,0 +1,168 @@
+---
+description: >-
+ **NeuralForecast** offers a large collection of neural forecasting models
+ focused on their usability, and robustness. The models range from classic
+ networks like `MLP`, `RNN`s to novel proven contributions like `NBEATS`,
+ `NHITS`, `TFT` and other architectures.
+output-file: introduction.html
+title: About NeuralForecast
+---
+
+
+## 🎊 Features
+
+- **Exogenous Variables**: Static, historic and future exogenous
+ support.
+- **Forecast Interpretability**: Plot trend, seasonality and exogenous
+ [`NBEATS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nbeats.html#nbeats),
+ [`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits),
+ [`TFT`](https://nixtlaverse.nixtla.io/neuralforecast/models.tft.html#tft),
+ `ESRNN` prediction components.
+- **Probabilistic Forecasting**: Simple model adapters for quantile
+ losses and parametric distributions.
+- **Train and Evaluation Losses** Scale-dependent, percentage and
+ scale independent errors, and parametric likelihoods.
+- **Automatic Model Selection** Parallelized automatic hyperparameter
+ tuning, that efficiently searches best validation configuration.
+- **Simple Interface** Unified SKLearn Interface for `StatsForecast`
+ and `MLForecast` compatibility.
+- **Model Collection**: Out of the box implementation of
+ [`MLP`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlp.html#mlp),
+ [`LSTM`](https://nixtlaverse.nixtla.io/neuralforecast/models.lstm.html#lstm),
+ [`RNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.rnn.html#rnn),
+ [`TCN`](https://nixtlaverse.nixtla.io/neuralforecast/models.tcn.html#tcn),
+ [`DilatedRNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.dilated_rnn.html#dilatedrnn),
+ [`NBEATS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nbeats.html#nbeats),
+ [`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits),
+ `ESRNN`,
+ [`Informer`](https://nixtlaverse.nixtla.io/neuralforecast/models.informer.html#informer),
+ [`TFT`](https://nixtlaverse.nixtla.io/neuralforecast/models.tft.html#tft),
+ [`PatchTST`](https://nixtlaverse.nixtla.io/neuralforecast/models.patchtst.html#patchtst),
+ [`VanillaTransformer`](https://nixtlaverse.nixtla.io/neuralforecast/models.vanillatransformer.html#vanillatransformer),
+ [`StemGNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.stemgnn.html#stemgnn)
+ and
+ [`HINT`](https://nixtlaverse.nixtla.io/neuralforecast/models.hint.html#hint).
+ See the entire [collection
+ here](https://nixtlaverse.nixtla.io/neuralforecast/docs/capabilities/overview.html).
+
+## Why?
+
+There is a shared belief in Neural forecasting methods’ capacity to
+improve our pipeline’s accuracy and efficiency.
+
+Unfortunately, available implementations and published research are yet
+to realize neural networks’ potential. They are hard to use and
+continuously fail to improve over statistical methods while being
+computationally prohibitive. For this reason, we created
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast),
+a library favoring proven accurate and efficient models focusing on
+their usability.
+
+## 💻 Installation
+
+### PyPI
+
+You can install
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)’s
+*released version* from the Python package index
+[pip](https://pypi.org/project/neuralforecast/) with:
+
+
+```python
+pip install neuralforecast
+```
+
+(Installing inside a python virtualenvironment or a conda environment is
+recommended.)
+
+### Conda
+
+Also you can install
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)’s
+*released version* from
+[conda](https://anaconda.org/conda-forge/neuralforecast) with:
+
+
+```python
+conda install -c conda-forge neuralforecast
+```
+
+(Installing inside a python virtualenvironment or a conda environment is
+recommended.)
+
+### Dev Mode
+
+If you want to make some modifications to the code and see the effects
+in real time (without reinstalling), follow the steps below:
+
+
+```bash
+git clone https://github.com/Nixtla/neuralforecast.git
+cd neuralforecast
+pip install -e .
+```
+
+## How to Use
+
+
+```python
+import logging
+
+import pandas as pd
+from utilsforecast.plotting import plot_series
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import NBEATS, NHITS
+from neuralforecast.utils import AirPassengersDF
+```
+
+
+```python
+logging.getLogger('pytorch_lightning').setLevel(logging.ERROR)
+```
+
+
+```python
+# Split data and declare panel dataset
+Y_df = AirPassengersDF
+Y_train_df = Y_df[Y_df.ds<='1959-12-31'] # 132 train
+Y_test_df = Y_df[Y_df.ds>'1959-12-31'] # 12 test
+
+# Fit and predict with NBEATS and NHITS models
+horizon = len(Y_test_df)
+models = [NBEATS(input_size=2 * horizon, h=horizon, max_steps=100, enable_progress_bar=False),
+ NHITS(input_size=2 * horizon, h=horizon, max_steps=100, enable_progress_bar=False)]
+nf = NeuralForecast(models=models, freq='ME')
+nf.fit(df=Y_train_df)
+Y_hat_df = nf.predict()
+
+# Plot predictions
+plot_series(Y_train_df, Y_hat_df)
+```
+
+``` text
+Seed set to 1
+Seed set to 1
+```
+
+
+
+## 🙏 How to Cite
+
+If you enjoy or benefit from using these Python implementations, a
+citation to the repository will be greatly appreciated.
+
+``` text
+@misc{olivares2022library_neuralforecast,
+ author={Kin G. Olivares and
+ Cristian Challú and
+ Federico Garza and
+ Max Mergenthaler Canseco and
+ Artur Dubrawski},
+ title = {{NeuralForecast}: User friendly state-of-the-art neural forecasting models.},
+ year={2022},
+ howpublished={{PyCon} Salt Lake City, Utah, US 2022},
+ url={https://github.com/Nixtla/neuralforecast}
+}
+```
+
diff --git a/neuralforecast/docs/getting-started/quickstart.html.mdx b/neuralforecast/docs/getting-started/quickstart.html.mdx
new file mode 100644
index 00000000..65f007b2
--- /dev/null
+++ b/neuralforecast/docs/getting-started/quickstart.html.mdx
@@ -0,0 +1,192 @@
+---
+description: Fit an LSTM and NHITS model
+output-file: quickstart.html
+title: Quickstart
+---
+
+
+This notebook provides an example on how to start using the main
+functionalities of the NeuralForecast library. The
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+class allows users to easily interact with `NeuralForecast.models`
+PyTorch models. In this example we will forecast AirPassengers data with
+a classic
+[`LSTM`](https://nixtlaverse.nixtla.io/neuralforecast/models.lstm.html#lstm)
+and the recent
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+models. The full list of available models is available
+[here](https://nixtlaverse.nixtla.io/neuralforecast/docs/capabilities/overview.html).
+
+You can run these experiments using GPU with Google Colab.
+
+
+
+## 1. Installing NeuralForecast
+
+
+```python
+!pip install neuralforecast
+```
+
+## 2. Loading AirPassengers Data
+
+The `core.NeuralForecast` class contains shared, `fit`, `predict` and
+other methods that take as inputs pandas DataFrames with columns
+`['unique_id', 'ds', 'y']`, where `unique_id` identifies individual time
+series from the dataset, `ds` is the date, and `y` is the target
+variable.
+
+In this example dataset consists of a set of a single series, but you
+can easily fit your model to larger datasets in long format.
+
+
+```python
+from neuralforecast.utils import AirPassengersDF
+```
+
+
+```python
+Y_df = AirPassengersDF
+Y_df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|-------|
+| 0 | 1.0 | 1949-01-31 | 112.0 |
+| 1 | 1.0 | 1949-02-28 | 118.0 |
+| 2 | 1.0 | 1949-03-31 | 132.0 |
+| 3 | 1.0 | 1949-04-30 | 129.0 |
+| 4 | 1.0 | 1949-05-31 | 121.0 |
+
+> **Important**
+>
+> DataFrames must include all `['unique_id', 'ds', 'y']` columns. Make
+> sure `y` column does not have missing or non-numeric values.
+
+## 3. Model Training
+
+### Fit the models
+
+Using the
+[`NeuralForecast.fit`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast.fit)
+method you can train a set of models to your dataset. You can define the
+forecasting `horizon` (12 in this example), and modify the
+hyperparameters of the model. For example, for the
+[`LSTM`](https://nixtlaverse.nixtla.io/neuralforecast/models.lstm.html#lstm)
+we changed the default hidden size for both encoder and decoders.
+
+
+```python
+import logging
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import LSTM, NHITS, RNN
+```
+
+
+```python
+logging.getLogger('pytorch_lightning').setLevel(logging.ERROR)
+```
+
+
+```python
+horizon = 12
+
+# Try different hyperparmeters to improve accuracy.
+models = [LSTM(input_size=2 * horizon,
+ h=horizon, # Forecast horizon
+ max_steps=500, # Number of steps to train
+ scaler_type='standard', # Type of scaler to normalize data
+ encoder_hidden_size=64, # Defines the size of the hidden state of the LSTM
+ decoder_hidden_size=64,), # Defines the number of hidden units of each layer of the MLP decoder
+ NHITS(h=horizon, # Forecast horizon
+ input_size=2 * horizon, # Length of input sequence
+ max_steps=100, # Number of steps to train
+ n_freq_downsample=[2, 1, 1]) # Downsampling factors for each stack output
+ ]
+nf = NeuralForecast(models=models, freq='ME')
+nf.fit(df=Y_df)
+```
+
+> **Tip**
+>
+> The performance of Deep Learning models can be very sensitive to the
+> choice of hyperparameters. Tuning the correct hyperparameters is an
+> important step to obtain the best forecasts. The `Auto` version of
+> these models,
+> [`AutoLSTM`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autolstm)
+> and
+> [`AutoNHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autonhits),
+> already perform hyperparameter selection automatically.
+
+### Predict using the fitted models
+
+Using the
+[`NeuralForecast.predict`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast.predict)
+method you can obtain the `h` forecasts after the training data `Y_df`.
+
+
+```python
+Y_hat_df = nf.predict()
+```
+
+The
+[`NeuralForecast.predict`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast.predict)
+method returns a DataFrame with the forecasts for each `unique_id`,
+`ds`, and model.
+
+
+```python
+Y_hat_df = Y_hat_df
+Y_hat_df.head()
+```
+
+| | unique_id | ds | LSTM | NHITS |
+|-----|-----------|------------|------------|------------|
+| 0 | 1.0 | 1961-01-31 | 445.602112 | 447.531281 |
+| 1 | 1.0 | 1961-02-28 | 431.253510 | 439.081024 |
+| 2 | 1.0 | 1961-03-31 | 456.301270 | 481.924194 |
+| 3 | 1.0 | 1961-04-30 | 508.149750 | 501.501343 |
+| 4 | 1.0 | 1961-05-31 | 524.903870 | 514.664551 |
+
+## 4. Plot Predictions
+
+Finally, we plot the forecasts of both models againts the real values.
+
+
+```python
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+plot_series(Y_df, Y_hat_df)
+```
+
+
+
+> **Tip**
+>
+> For this guide we are using a simple
+> [`LSTM`](https://nixtlaverse.nixtla.io/neuralforecast/models.lstm.html#lstm)
+> model. More recent models, such as
+> [`TSMixer`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixer.html#tsmixer),
+> [`TFT`](https://nixtlaverse.nixtla.io/neuralforecast/models.tft.html#tft)
+> and
+> [`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+> achieve better accuracy than
+> [`LSTM`](https://nixtlaverse.nixtla.io/neuralforecast/models.lstm.html#lstm)
+> in most settings. The full list of available models is available
+> [here](https://nixtlaverse.nixtla.io/neuralforecast/docs/capabilities/overview.html).
+
+## References
+
+- [Boris N. Oreshkin, Dmitri Carpov, Nicolas Chapados, Yoshua Bengio
+ (2020). “N-BEATS: Neural basis expansion analysis for interpretable
+ time series forecasting”. International Conference on Learning
+ Representations.](https://arxiv.org/abs/1905.10437)
+- [Cristian Challu, Kin G. Olivares, Boris N. Oreshkin, Federico
+ Garza, Max Mergenthaler-Canseco, Artur Dubrawski (2021). NHITS:
+ Neural Hierarchical Interpolation for Time Series Forecasting.
+ Accepted at AAAI 2023.](https://arxiv.org/abs/2201.12886)
+
diff --git a/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-14-output-1.png b/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..1e895119
Binary files /dev/null and b/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-15-output-1.png b/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-15-output-1.png
new file mode 100644
index 00000000..0e59ebb4
Binary files /dev/null and b/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-15-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-16-output-1.png b/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-16-output-1.png
new file mode 100644
index 00000000..a770bf37
Binary files /dev/null and b/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-16-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-23-output-1.png b/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-23-output-1.png
new file mode 100644
index 00000000..3fba3e5a
Binary files /dev/null and b/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-23-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-23-output-2.png b/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-23-output-2.png
new file mode 100644
index 00000000..452b31b8
Binary files /dev/null and b/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-23-output-2.png differ
diff --git a/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-28-output-1.png b/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-28-output-1.png
new file mode 100644
index 00000000..081ac623
Binary files /dev/null and b/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-28-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-31-output-1.png b/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-31-output-1.png
new file mode 100644
index 00000000..5e523797
Binary files /dev/null and b/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-31-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-7-output-1.png b/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..cab000be
Binary files /dev/null and b/neuralforecast/docs/tutorials/01_getting_started_complete_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/02_cross_validation_files/figure-markdown_strict/cell-17-output-1.png b/neuralforecast/docs/tutorials/02_cross_validation_files/figure-markdown_strict/cell-17-output-1.png
new file mode 100644
index 00000000..441cd93a
Binary files /dev/null and b/neuralforecast/docs/tutorials/02_cross_validation_files/figure-markdown_strict/cell-17-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/02_cross_validation_files/figure-markdown_strict/cell-18-output-1.png b/neuralforecast/docs/tutorials/02_cross_validation_files/figure-markdown_strict/cell-18-output-1.png
new file mode 100644
index 00000000..19cf6d96
Binary files /dev/null and b/neuralforecast/docs/tutorials/02_cross_validation_files/figure-markdown_strict/cell-18-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/02_cross_validation_files/figure-markdown_strict/cell-18-output-2.png b/neuralforecast/docs/tutorials/02_cross_validation_files/figure-markdown_strict/cell-18-output-2.png
new file mode 100644
index 00000000..44c154d3
Binary files /dev/null and b/neuralforecast/docs/tutorials/02_cross_validation_files/figure-markdown_strict/cell-18-output-2.png differ
diff --git a/neuralforecast/docs/tutorials/02_cross_validation_files/figure-markdown_strict/cell-18-output-3.png b/neuralforecast/docs/tutorials/02_cross_validation_files/figure-markdown_strict/cell-18-output-3.png
new file mode 100644
index 00000000..973c76d4
Binary files /dev/null and b/neuralforecast/docs/tutorials/02_cross_validation_files/figure-markdown_strict/cell-18-output-3.png differ
diff --git a/neuralforecast/docs/tutorials/02_cross_validation_files/figure-markdown_strict/cell-8-output-1.png b/neuralforecast/docs/tutorials/02_cross_validation_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..cab000be
Binary files /dev/null and b/neuralforecast/docs/tutorials/02_cross_validation_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/03_uncertainty_quantification_files/figure-markdown_strict/cell-15-output-1.png b/neuralforecast/docs/tutorials/03_uncertainty_quantification_files/figure-markdown_strict/cell-15-output-1.png
new file mode 100644
index 00000000..b5095e0d
Binary files /dev/null and b/neuralforecast/docs/tutorials/03_uncertainty_quantification_files/figure-markdown_strict/cell-15-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/03_uncertainty_quantification_files/figure-markdown_strict/cell-16-output-1.png b/neuralforecast/docs/tutorials/03_uncertainty_quantification_files/figure-markdown_strict/cell-16-output-1.png
new file mode 100644
index 00000000..a17a862d
Binary files /dev/null and b/neuralforecast/docs/tutorials/03_uncertainty_quantification_files/figure-markdown_strict/cell-16-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/03_uncertainty_quantification_files/figure-markdown_strict/cell-8-output-1.png b/neuralforecast/docs/tutorials/03_uncertainty_quantification_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..0b837267
Binary files /dev/null and b/neuralforecast/docs/tutorials/03_uncertainty_quantification_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/04_longhorizon_nhits_files/figure-markdown_strict/cell-11-output-1.png b/neuralforecast/docs/tutorials/04_longhorizon_nhits_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..b9a6efc9
Binary files /dev/null and b/neuralforecast/docs/tutorials/04_longhorizon_nhits_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/04_longhorizon_nhits_files/figure-markdown_strict/cell-4-output-1.png b/neuralforecast/docs/tutorials/04_longhorizon_nhits_files/figure-markdown_strict/cell-4-output-1.png
new file mode 100644
index 00000000..04b53fcb
Binary files /dev/null and b/neuralforecast/docs/tutorials/04_longhorizon_nhits_files/figure-markdown_strict/cell-4-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/05_longhorizon_transformers_files/figure-markdown_strict/cell-10-output-1.png b/neuralforecast/docs/tutorials/05_longhorizon_transformers_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..cc3491a8
Binary files /dev/null and b/neuralforecast/docs/tutorials/05_longhorizon_transformers_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/06_longhorizon_probabilistic_files/figure-markdown_strict/cell-15-output-1.png b/neuralforecast/docs/tutorials/06_longhorizon_probabilistic_files/figure-markdown_strict/cell-15-output-1.png
new file mode 100644
index 00000000..a75682cd
Binary files /dev/null and b/neuralforecast/docs/tutorials/06_longhorizon_probabilistic_files/figure-markdown_strict/cell-15-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/06_longhorizon_probabilistic_files/figure-markdown_strict/cell-6-output-1.png b/neuralforecast/docs/tutorials/06_longhorizon_probabilistic_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..f45fea8c
Binary files /dev/null and b/neuralforecast/docs/tutorials/06_longhorizon_probabilistic_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/07_forecasting_tft_files/figure-markdown_strict/cell-12-output-1.png b/neuralforecast/docs/tutorials/07_forecasting_tft_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..edd6222e
Binary files /dev/null and b/neuralforecast/docs/tutorials/07_forecasting_tft_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/07_forecasting_tft_files/figure-markdown_strict/cell-9-output-1.png b/neuralforecast/docs/tutorials/07_forecasting_tft_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..499a918e
Binary files /dev/null and b/neuralforecast/docs/tutorials/07_forecasting_tft_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/08_multivariate_tsmixer_files/figure-markdown_strict/cell-11-output-1.png b/neuralforecast/docs/tutorials/08_multivariate_tsmixer_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..7390256f
Binary files /dev/null and b/neuralforecast/docs/tutorials/08_multivariate_tsmixer_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/09_hierarchical_forecasting_files/figure-markdown_strict/cell-12-output-1.png b/neuralforecast/docs/tutorials/09_hierarchical_forecasting_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..17fe9222
Binary files /dev/null and b/neuralforecast/docs/tutorials/09_hierarchical_forecasting_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/09_hierarchical_forecasting_files/figure-markdown_strict/cell-5-output-1.png b/neuralforecast/docs/tutorials/09_hierarchical_forecasting_files/figure-markdown_strict/cell-5-output-1.png
new file mode 100644
index 00000000..2f990298
Binary files /dev/null and b/neuralforecast/docs/tutorials/09_hierarchical_forecasting_files/figure-markdown_strict/cell-5-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/09_hierarchical_forecasting_files/figure-markdown_strict/cell-6-output-1.png b/neuralforecast/docs/tutorials/09_hierarchical_forecasting_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..1fb89d6e
Binary files /dev/null and b/neuralforecast/docs/tutorials/09_hierarchical_forecasting_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/10_distributed_neuralforecast_files/figure-markdown_strict/cell-14-output-1.png b/neuralforecast/docs/tutorials/10_distributed_neuralforecast_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..362dc71f
Binary files /dev/null and b/neuralforecast/docs/tutorials/10_distributed_neuralforecast_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/11_intermittent_data_files/figure-markdown_strict/cell-12-output-1.png b/neuralforecast/docs/tutorials/11_intermittent_data_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..525659cd
Binary files /dev/null and b/neuralforecast/docs/tutorials/11_intermittent_data_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/11_intermittent_data_files/figure-markdown_strict/cell-16-output-1.png b/neuralforecast/docs/tutorials/11_intermittent_data_files/figure-markdown_strict/cell-16-output-1.png
new file mode 100644
index 00000000..92b9364b
Binary files /dev/null and b/neuralforecast/docs/tutorials/11_intermittent_data_files/figure-markdown_strict/cell-16-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/11_intermittent_data_files/figure-markdown_strict/cell-16-output-2.png b/neuralforecast/docs/tutorials/11_intermittent_data_files/figure-markdown_strict/cell-16-output-2.png
new file mode 100644
index 00000000..b7a653d0
Binary files /dev/null and b/neuralforecast/docs/tutorials/11_intermittent_data_files/figure-markdown_strict/cell-16-output-2.png differ
diff --git a/neuralforecast/docs/tutorials/11_intermittent_data_files/figure-markdown_strict/cell-16-output-3.png b/neuralforecast/docs/tutorials/11_intermittent_data_files/figure-markdown_strict/cell-16-output-3.png
new file mode 100644
index 00000000..b1208aa8
Binary files /dev/null and b/neuralforecast/docs/tutorials/11_intermittent_data_files/figure-markdown_strict/cell-16-output-3.png differ
diff --git a/neuralforecast/docs/tutorials/11_intermittent_data_files/figure-markdown_strict/cell-6-output-1.png b/neuralforecast/docs/tutorials/11_intermittent_data_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..f3829802
Binary files /dev/null and b/neuralforecast/docs/tutorials/11_intermittent_data_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/12_using_mlflow_files/figure-markdown_strict/cell-6-output-1.png b/neuralforecast/docs/tutorials/12_using_mlflow_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..e88fe88d
Binary files /dev/null and b/neuralforecast/docs/tutorials/12_using_mlflow_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/13_robust_forecasting_files/figure-markdown_strict/cell-10-output-1.png b/neuralforecast/docs/tutorials/13_robust_forecasting_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..aa601633
Binary files /dev/null and b/neuralforecast/docs/tutorials/13_robust_forecasting_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/13_robust_forecasting_files/figure-markdown_strict/cell-5-output-1.png b/neuralforecast/docs/tutorials/13_robust_forecasting_files/figure-markdown_strict/cell-5-output-1.png
new file mode 100644
index 00000000..0806b638
Binary files /dev/null and b/neuralforecast/docs/tutorials/13_robust_forecasting_files/figure-markdown_strict/cell-5-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/13_robust_forecasting_files/figure-markdown_strict/cell-6-output-1.png b/neuralforecast/docs/tutorials/13_robust_forecasting_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..15aff1f5
Binary files /dev/null and b/neuralforecast/docs/tutorials/13_robust_forecasting_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/14_interpretable_decompositions_files/figure-markdown_strict/cell-11-output-1.png b/neuralforecast/docs/tutorials/14_interpretable_decompositions_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..be62b3b7
Binary files /dev/null and b/neuralforecast/docs/tutorials/14_interpretable_decompositions_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/14_interpretable_decompositions_files/figure-markdown_strict/cell-14-output-1.png b/neuralforecast/docs/tutorials/14_interpretable_decompositions_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..0e7edec2
Binary files /dev/null and b/neuralforecast/docs/tutorials/14_interpretable_decompositions_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/14_interpretable_decompositions_files/figure-markdown_strict/cell-6-output-1.png b/neuralforecast/docs/tutorials/14_interpretable_decompositions_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..c7393657
Binary files /dev/null and b/neuralforecast/docs/tutorials/14_interpretable_decompositions_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-25-output-1.png b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-25-output-1.png
new file mode 100644
index 00000000..6869a91a
Binary files /dev/null and b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-25-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-26-output-1.png b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-26-output-1.png
new file mode 100644
index 00000000..1b10d56b
Binary files /dev/null and b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-26-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-35-output-1.png b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-35-output-1.png
new file mode 100644
index 00000000..5fa8e643
Binary files /dev/null and b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-35-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-35-output-2.png b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-35-output-2.png
new file mode 100644
index 00000000..06395a31
Binary files /dev/null and b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-35-output-2.png differ
diff --git a/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-35-output-3.png b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-35-output-3.png
new file mode 100644
index 00000000..3607aeea
Binary files /dev/null and b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-35-output-3.png differ
diff --git a/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-38-output-1.png b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-38-output-1.png
new file mode 100644
index 00000000..aca29a5b
Binary files /dev/null and b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-38-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-38-output-2.png b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-38-output-2.png
new file mode 100644
index 00000000..21d613a2
Binary files /dev/null and b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-38-output-2.png differ
diff --git a/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-38-output-3.png b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-38-output-3.png
new file mode 100644
index 00000000..879355b4
Binary files /dev/null and b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-38-output-3.png differ
diff --git a/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-46-output-1.png b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-46-output-1.png
new file mode 100644
index 00000000..c091fde5
Binary files /dev/null and b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-46-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-47-output-1.png b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-47-output-1.png
new file mode 100644
index 00000000..de42d588
Binary files /dev/null and b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-47-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-48-output-1.png b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-48-output-1.png
new file mode 100644
index 00000000..701f4a76
Binary files /dev/null and b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-48-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-49-output-1.png b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-49-output-1.png
new file mode 100644
index 00000000..be658b1a
Binary files /dev/null and b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-49-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-51-output-1.png b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-51-output-1.png
new file mode 100644
index 00000000..d2e653d9
Binary files /dev/null and b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-51-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-7-output-1.png b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..f3829802
Binary files /dev/null and b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-8-output-1.png b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..7bb131f9
Binary files /dev/null and b/neuralforecast/docs/tutorials/15_comparing_methods_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/16_temporal_classification_files/figure-markdown_strict/cell-4-output-1.png b/neuralforecast/docs/tutorials/16_temporal_classification_files/figure-markdown_strict/cell-4-output-1.png
new file mode 100644
index 00000000..d0a7e708
Binary files /dev/null and b/neuralforecast/docs/tutorials/16_temporal_classification_files/figure-markdown_strict/cell-4-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/16_temporal_classification_files/figure-markdown_strict/cell-4-output-2.png b/neuralforecast/docs/tutorials/16_temporal_classification_files/figure-markdown_strict/cell-4-output-2.png
new file mode 100644
index 00000000..a8f01668
Binary files /dev/null and b/neuralforecast/docs/tutorials/16_temporal_classification_files/figure-markdown_strict/cell-4-output-2.png differ
diff --git a/neuralforecast/docs/tutorials/16_temporal_classification_files/figure-markdown_strict/cell-9-output-1.png b/neuralforecast/docs/tutorials/16_temporal_classification_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..f5e94e13
Binary files /dev/null and b/neuralforecast/docs/tutorials/16_temporal_classification_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/17_transfer_learning_files/figure-markdown_strict/cell-12-output-1.png b/neuralforecast/docs/tutorials/17_transfer_learning_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..8fae1aa6
Binary files /dev/null and b/neuralforecast/docs/tutorials/17_transfer_learning_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/20_conformal_prediction_files/figure-markdown_strict/cell-7-output-1.png b/neuralforecast/docs/tutorials/20_conformal_prediction_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..5c3382d7
Binary files /dev/null and b/neuralforecast/docs/tutorials/20_conformal_prediction_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/21_configure_optimizers_files/figure-markdown_strict/cell-5-output-1.png b/neuralforecast/docs/tutorials/21_configure_optimizers_files/figure-markdown_strict/cell-5-output-1.png
new file mode 100644
index 00000000..6186f0d2
Binary files /dev/null and b/neuralforecast/docs/tutorials/21_configure_optimizers_files/figure-markdown_strict/cell-5-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/22_explainability_files/figure-markdown_strict/cell-10-output-1.png b/neuralforecast/docs/tutorials/22_explainability_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..eae8f2f2
Binary files /dev/null and b/neuralforecast/docs/tutorials/22_explainability_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/22_explainability_files/figure-markdown_strict/cell-11-output-1.png b/neuralforecast/docs/tutorials/22_explainability_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..8044164e
Binary files /dev/null and b/neuralforecast/docs/tutorials/22_explainability_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/22_explainability_files/figure-markdown_strict/cell-12-output-1.png b/neuralforecast/docs/tutorials/22_explainability_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..4d6542fd
Binary files /dev/null and b/neuralforecast/docs/tutorials/22_explainability_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/neuralforecast/docs/tutorials/adding_models.html.mdx b/neuralforecast/docs/tutorials/adding_models.html.mdx
new file mode 100644
index 00000000..68c5ea2e
--- /dev/null
+++ b/neuralforecast/docs/tutorials/adding_models.html.mdx
@@ -0,0 +1,422 @@
+---
+description: Tutorial on how to add new models to NeuralForecast
+output-file: adding_models.html
+title: Adding Models to NeuralForecast
+---
+
+
+> **Prerequisites**
+>
+> This Guide assumes advanced familiarity with NeuralForecast.
+>
+> We highly recommend reading first the Getting Started and the
+> NeuralForecast Map tutorials!
+>
+> Additionally, refer to the [CONTRIBUTING
+> guide](https://github.com/Nixtla/neuralforecast/blob/main/CONTRIBUTING.md)
+> for the basics how to contribute to NeuralForecast.
+
+## Introduction
+
+This tutorial is aimed at contributors who want to add a new model to
+the NeuralForecast library. The library’s existing modules handle
+optimization, training, selection, and evaluation of deep learning
+models. The `core` class simplifies building entire pipelines, both for
+industry and academia, on any dataset, with user-friendly methods such
+as `fit` and `predict`.
+
+Adding a new model to NeuralForecast is simpler than building a new
+PyTorch model from scratch. You only need to write the forward method.
+
+**It has the following additional advantages:**
+
+- Existing modules in NeuralForecast already implement the essential
+ training and evaluating aspects for deep learning models.
+- Integrated with PyTorch-Lightning and Tune libraries for efficient
+ optimization and distributed computation.
+- The `BaseModel` classes provide common optimization components, such
+ as early stopping and learning rate schedulers.
+- Automatic performance tests are scheduled on Github to ensure
+ quality standards.
+- Users can easily compare the performance and computation of the new
+ model with existing models.
+- Opportunity for exposure to a large community of users and
+ contributors.
+
+### Example: simplified MLP model
+
+We will present the tutorial following an example on how to add a
+simplified version of the current
+[`MLP`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlp.html#mlp)
+model, which does not include exogenous covariates.
+
+At a given timestamp $t$, the
+[`MLP`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlp.html#mlp)
+model will forecast the next $h$ values of the univariate target time,
+$Y_{t+1:t+h}$, using as inputs the last $L$ historical values, given by
+$Y_{t-L:t}$. The following figure presents a diagram of the model.
+
+
+
+Figure 1. Three layer MLP with
+autoregresive inputs.
+
+
+## 0. Preliminaries
+
+Follow our tutorial on contributing
+[here](https://github.com/Nixtla/neuralforecast/blob/main/CONTRIBUTING.md)
+to set up your development environment.
+
+Here is a short list of the most important steps:
+
+1. Create a fork of the `neuralforecast` library.
+2. Clone the fork to your computer.
+3. Set an environment with the `neuralforecast` library, core
+ dependencies, and `nbdev` package to code your model in an
+ interactive notebook.
+
+## 1. Inherit the Base Class (`BaseModel`)
+
+The library contains a base model class: `BaseModel`. Using class
+attributes we can make this model recurrent or not, or multivariate or
+univariate, or allow the use of exogenous inputs.
+
+### a. Sampling process
+
+During training, the base class receives a sample of time series of the
+dataset from the
+[`TimeSeriesLoader`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesloader)
+module. The `BaseModel` models will sample individual windows of size
+`input_size+h`, starting from random timestamps.
+
+### b. `BaseModel`’ hyperparameters
+
+Get familiar with the hyperparameters specified in the base class,
+including `h` (horizon), `input_size`, and optimization hyperparameters
+such as `learning_rate`, `max_steps`, among others. The following list
+presents the hyperparameters related to the sampling of windows:
+
+- `h` (h): number of future values to predict.
+- `input_size` (L): number of historic values to use as input for the
+ model.
+- `batch_size` (bs): number of time series sampled by the loader
+ during training.
+- `valid_batch_size` (v_bs): number of time series sampled by the
+ loader during inference (validation and test).
+- `windows_batch_size` (w_bs): number of individual windows sampled
+ during training (from the previous time series) to form the batch.
+- `inference_windows_batch_size` (i_bs): number of individual windows
+ sampled during inference to form each batch. Used to control the GPU
+ memory.
+
+### c. Input and Output batch shapes
+
+The `forward` method receives a batch of data in a dictionary with the
+following keys:
+
+- `insample_y`: historic values of the time series.
+- `insample_mask`: mask indicating the available values of the time
+ series (1 if available, 0 if missing).
+- `futr_exog`: future exogenous covariates (if any).
+- `hist_exog`: historic exogenous covariates (if any).
+- `stat_exog`: static exogenous covariates (if any).
+
+The following table presents the shape for each tensor if the attribute
+`MULTIVARIATE = False` is set:
+
+| `tensor` | `BaseModel` |
+|-----------------|--------------------------|
+| `insample_y` | (`w_bs`, `L`, `1`) |
+| `insample_mask` | (`w_bs`, `L`) |
+| `futr_exog` | (`w_bs`, `L`+`h`, `n_f`) |
+| `hist_exog` | (`w_bs`, `L`, `n_h`) |
+| `stat_exog` | (`w_bs`,`n_s`) |
+
+The `forward` function should return a single tensor with the forecasts
+of the next `h` timestamps for each window. Use the attributes of the
+`loss` class to automatically parse the output to the correct shape (see
+the example below).
+
+> **Tip**
+>
+> Since we are using `nbdev`, you can easily add prints to the code and
+> see the shapes of the tensors during training.
+
+### d. `BaseModel`’ methods
+
+The `BaseModel` class contains several common methods for all
+windows-based models, simplifying the development of new models by
+preventing code duplication. The most important methods of the class
+are:
+
+- `_create_windows`: parses the time series from the
+ [`TimeSeriesLoader`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesloader)
+ into individual windows of size `input_size+h`.
+- `_normalization`: normalizes each window based on the `scaler` type.
+- `_inv_normalization`: inverse normalization of the forecasts.
+- `training_step`: training step of the model, called by
+ PyTorch-Lightning’s `Trainer` class during training (`fit` method).
+- `validation_step`: validation step of the model, called by
+ PyTorch-Lightning’s `Trainer` class during validation.
+- `predict_step`: prediction step of the model, called by
+ PyTorch-Lightning’s `Trainer` class during inference (`predict`
+ method).
+
+## 2. Create the model file and class
+
+Once familiar with the basics of the `BaseModel` class, the next step is
+creating your particular model.
+
+The main steps are:
+
+1. Create the file in the `nbs` folder
+ (https://github.com/Nixtla/neuralforecast/tree/main/nbs). It should
+ be named `models.YOUR_MODEL_NAME.ipynb`.
+2. Add the header of the `nbdev` file.
+3. Import libraries in the file.
+4. Define the `__init__` method with the model’s inherited and
+ particular hyperparameters and instantiate the architecture.
+5. Set the following model attributes:
+ - `EXOGENOUS_FUTR`: if the model can handle future exogenous
+ variables (True) or not (False)
+ - `EXOGENOUS_HIST`: if the model can handle historical exogenous
+ variables (True) or not (False)
+ - `EXOGENOUS_STAT`: if the model can handle static exogenous
+ variables (True) or not (False)
+ - `MULTIVARIATE`: If the model produces multivariate forecasts
+ (True) or univariate (False)
+ - `RECURRENT`: If the model produces forecasts recursively (True)
+ or direct (False)
+6. Define the `forward` method, which recieves the input batch
+ dictionary and returns the forecast.
+
+### a. Model class
+
+First, add the following **two cells** on top of the `nbdev` file.
+
+
+```python
+#| default_exp models.mlp
+```
+
+> **Important**
+>
+> Change `mlp` to your model’s name, using lowercase and underscores.
+> When you later run `nbdev_export`, it will create a `YOUR_MODEL.py`
+> script in the `neuralforecast/models/` directory.
+
+
+```python
+#| hide
+%load_ext autoreload
+%autoreload 2
+```
+
+Next, add the dependencies of the model.
+
+
+```python
+#| export
+from typing import Optional
+
+import torch
+import torch.nn as nn
+
+from neuralforecast.losses.pytorch import MAE
+from neuralforecast.common._base_model import BaseModel
+```
+
+> **Tip**
+>
+> Don’t forget to add the `#| export` tag on this cell.
+
+Next, create the class with the `init` and `forward` methods. The
+following example shows the example for the simplified
+[`MLP`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlp.html#mlp)
+model. We explain important details after the code.
+
+
+```python
+#| export
+class MLP(BaseModel): # <<---- Inherits from BaseModel
+ # Set class attributes to determine this model's characteristics
+ EXOGENOUS_FUTR = False # If the model can handle future exogenous variables
+ EXOGENOUS_HIST = False # If the model can handle historical exogenous variables
+ EXOGENOUS_STAT = False # If the model can handle static exogenous variables
+ MULTIVARIATE = False # If the model produces multivariate forecasts (True) or univariate (False)
+ RECURRENT = False # If the model produces forecasts recursively (True) or direct (False)
+
+ def __init__(self,
+ # Inhereted hyperparameters with no defaults
+ h,
+ input_size,
+ # Model specific hyperparameters
+ num_layers = 2,
+ hidden_size = 1024,
+ # Inhereted hyperparameters with defaults
+ futr_exog_list = None,
+ hist_exog_list = None,
+ stat_exog_list = None,
+ exclude_insample_y = False,
+ loss = MAE(),
+ valid_loss = None,
+ max_steps: int = 1000,
+ learning_rate: float = 1e-3,
+ num_lr_decays: int = -1,
+ early_stop_patience_steps: int =-1,
+ val_check_steps: int = 100,
+ batch_size: int = 32,
+ valid_batch_size: Optional[int] = None,
+ windows_batch_size = 1024,
+ inference_windows_batch_size = -1,
+ start_padding_enabled = False,
+ step_size: int = 1,
+ scaler_type: str = 'identity',
+ random_seed: int = 1,
+ drop_last_loader: bool = False,
+ optimizer = None,
+ optimizer_kwargs = None,
+ lr_scheduler = None,
+ lr_scheduler_kwargs = None,
+ dataloader_kwargs = None,
+ **trainer_kwargs):
+ # Inherit BaseWindows class
+ super(MLP, self).__init__(h=h,
+ input_size=input_size,
+ ..., # <<--- Add all inhereted hyperparameters
+ random_seed=random_seed,
+ **trainer_kwargs)
+
+ # Architecture
+ self.num_layers = num_layers
+ self.hidden_size = hidden_size
+
+ # MultiLayer Perceptron
+ layers = [nn.Linear(in_features=input_size, out_features=hidden_size)]
+ layers += [nn.ReLU()]
+ for i in range(num_layers - 1):
+ layers += [nn.Linear(in_features=hidden_size, out_features=hidden_size)]
+ layers += [nn.ReLU()]
+ self.mlp = nn.ModuleList(layers)
+
+ # Adapter with Loss dependent dimensions
+ self.out = nn.Linear(in_features=hidden_size,
+ out_features=h * self.loss.outputsize_multiplier) ## <<--- Use outputsize_multiplier to adjust output size
+
+ def forward(self, windows_batch): # <<--- Receives windows_batch dictionary
+ # Parse windows_batch
+ insample_y = windows_batch['insample_y'].squeeze(-1) # [batch_size, input_size]
+ # MLP
+ hidden = self.mlp(insample_y) # [batch_size, hidden_size]
+ y_pred = self.out(hidden) # [batch_size, h * n_outputs]
+
+ # Reshape
+ y_pred = y_pred.reshape(batch_size, self.h, self.loss.outputsize_multiplier) # [batch_size, h, n_outputs]
+
+ return y_pred
+
+```
+
+> **Tip**
+>
+> - Don’t forget to add the `#| export` tag on each cell.
+> - Larger architectures, such as Transformers, might require
+> splitting the `forward` by using intermediate functions.
+
+#### Important notes
+
+The base class has many hyperparameters, and models must have default
+values for all of them (except `h` and `input_size`). If you are unsure
+of what default value to use, we recommend copying the default values
+from existing models for most optimization and sampling hyperparameters.
+You can change the default values later at any time.
+
+The `reshape` method at the end of the `forward` step is used to adjust
+the output shape. The `loss` class contains an `outputsize_multiplier`
+attribute to automatically adjust the output size of the forecast
+depending on the `loss`. For example, for the Multi-quantile loss
+([`MQLoss`](https://nixtlaverse.nixtla.io/neuralforecast/losses.pytorch.html#mqloss)),
+the model needs to output each quantile for each horizon.
+
+### b. Tests and documentation
+
+`nbdev` allows for testing and documenting the model during the
+development process. It allows users to iterate the development within
+the notebook, testing the code in the same environment. Refer to
+existing models, such as the complete MLP model
+[here](https://github.com/Nixtla/neuralforecast/blob/main/nbs/models.mlp.html).
+These files already contain the tests, documentation, and usage examples
+that were used during the development process.
+
+### c. Export the new model to the library with `nbdev`
+
+Following the CONTRIBUTING guide, the next step is to export the new
+model from the development notebook to the `neuralforecast` folder with
+the actual scripts.
+
+To export the model, run `nbdev_export` in your terminal. You should see
+a new file with your model in the `neuralforecast/models/` folder.
+
+## 3. Core class and additional files
+
+Finally, add the model to the `core` class and additional files:
+
+1. Manually add the model in the following [init
+ file](https://github.com/Nixtla/neuralforecast/blob/main/neuralforecast/models/__init__.py).
+
+2. Add the model to the `core` class, using the `nbdev` file
+ [here](https://nixtlaverse.nixtla.io/neuralforecast/core.html):
+
+ 1. Add the model to the initial model list:
+
+
+ ```python
+ from neuralforecast.models import (
+ GRU, LSTM, RNN, TCN, DilatedRNN,
+ MLP, NHITS, NBEATS, NBEATSx,
+ TFT, VanillaTransformer,
+ Informer, Autoformer, FEDformer,
+ StemGNN, PatchTST
+ )
+ ```
+
+ 1. Add the model to the `MODEL_FILENAME_DICT` dictionary (used for
+ the `save` and `load` functions).
+
+## 4. Add the model to the documentation
+
+It’s important to add the model to the necessary documentation pages so
+that everyone can find the documentation:
+
+1. Add the model to the [model overview table](https://nixtlaverse.nixtla.io/neuralforecast/docs/capabilities/overview.html).
+2. Add the model to [mint.json](https://github.com/Nixtla/neuralforecast/blob/main/nbs/mint.json).
+
+## 5. Upload to GitHub
+
+Congratulations! The model is ready to be used in the library following
+the steps above.
+
+Follow our contributing guide’s final steps to upload the model to
+GitHub:
+[here](https://github.com/Nixtla/neuralforecast/blob/main/CONTRIBUTING.md).
+
+One of the maintainers will review the PR, request changes if necessary,
+and merge it into the library.
+
+## Quick Checklist
+
+- Get familiar with the `BaseModel` class hyperparameters and
+ input/output shapes of the `forward` method.
+- Create the notebook with your model class in the `nbs` folder:
+ `models.YOUR_MODEL_NAME.ipynb`
+- Add the header and import libraries.
+- Implement `init` and `forward` methods and set the class attributes.
+- Export model with `nbdev_export`.
+- Add model to this [init
+ file](https://github.com/Nixtla/neuralforecast/blob/main/neuralforecast/models/__init__.py).
+- Add the model to the `core` class
+ [here](https://nixtlaverse.nixtla.io/neuralforecast/core.html).
+- Follow the CONTRIBUTING guide to create the PR to upload the model.
+
diff --git a/neuralforecast/docs/tutorials/comparing_methods.html.mdx b/neuralforecast/docs/tutorials/comparing_methods.html.mdx
new file mode 100644
index 00000000..815a1d9e
--- /dev/null
+++ b/neuralforecast/docs/tutorials/comparing_methods.html.mdx
@@ -0,0 +1,946 @@
+---
+description: >-
+ In this notebook, you will make forecasts for the M5 dataset choosing the best
+ model for each time series using cross validation.
+output-file: comparing_methods.html
+title: Statistical, Machine Learning and Neural Forecasting methods
+---
+
+
+Statistical, Machine Learning, and Neural Forecasting Methods In this
+tutorial, we will explore the process of forecasting on the M5 dataset
+by utilizing the most suitable model for each time series. We’ll
+accomplish this through an essential technique known as
+cross-validation. This approach helps us in estimating the predictive
+performance of our models, and in selecting the model that yields the
+best performance for each time series.
+
+The M5 dataset comprises of hierarchical sales data, spanning five
+years, from Walmart. The aim is to forecast daily sales for the next 28
+days. The dataset is broken down into the 50 states of America, with 10
+stores in each state.
+
+In the realm of time series forecasting and analysis, one of the more
+complex tasks is identifying the model that is optimally suited for a
+specific group of series. Quite often, this selection process leans
+heavily on intuition, which may not necessarily align with the empirical
+reality of our dataset.
+
+In this tutorial, we aim to provide a more structured, data-driven
+approach to model selection for different groups of series within the M5
+benchmark dataset. This dataset, well-known in the field of forecasting,
+allows us to showcase the versatility and power of our methodology.
+
+We will train an assortment of models from various forecasting
+paradigms:
+
+*[StatsForecast]((https://github.com/Nixtla/statsforecast))*
+
+- Baseline models: These models are simple yet often highly effective
+ for providing an initial perspective on the forecasting problem. We
+ will use `SeasonalNaive` and `HistoricAverage` models for this
+ category.
+- Intermittent models: For series with sporadic, non-continuous
+ demand, we will utilize models like `CrostonOptimized`, `IMAPA`, and
+ `ADIDA`. These models are particularly suited for handling
+ zero-inflated series.
+- State Space Models: These are statistical models that use
+ mathematical descriptions of a system to make predictions. The
+ `AutoETS` model from the statsforecast library falls under this
+ category.
+
+*[MLForecast](https://github.com/Nixtla/mlforecast)*
+
+Machine Learning: Leveraging ML models like `LightGBM`, `XGBoost`, and
+`LinearRegression` can be advantageous due to their capacity to uncover
+intricate patterns in data. We’ll use the MLForecast library for this
+purpose.
+
+*[NeuralForecast](https://github.com/Nixtla/neuralforecast)*
+
+Deep Learning: DL models, such as Transformers
+([`AutoTFT`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotft))
+and Neural Networks
+([`AutoNHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autonhits)),
+allow us to handle complex non-linear dependencies in time series data.
+We’ll utilize the NeuralForecast library for these models.
+
+Using the Nixtla suite of libraries, we’ll be able to drive our model
+selection process with data, ensuring we utilize the most suitable
+models for specific groups of series in our dataset.
+
+Outline:
+
+- Reading Data: In this initial step, we load our dataset into memory,
+ making it available for our subsequent analysis and forecasting. It
+ is important to understand the structure and nuances of the dataset
+ at this stage.
+
+- Forecasting Using Statistical and Deep Learning Methods: We apply a
+ wide range of forecasting methods from basic statistical techniques
+ to advanced deep learning models. The aim is to generate predictions
+ for the next 28 days based on our dataset.
+
+- Model Performance Evaluation on Different Windows: We assess the
+ performance of our models on distinct windows.
+
+- Selecting the Best Model for a Group of Series: Using the
+ performance evaluation, we identify the optimal model for each group
+ of series. This step ensures that the chosen model is tailored to
+ the unique characteristics of each group.
+
+- Filtering the Best Possible Forecast: Finally, we filter the
+ forecasts generated by our chosen models to obtain the most
+ promising predictions. This is our final output and represents the
+ best possible forecast for each series according to our models.
+
+> **Warning**
+>
+> This tutorial was originally executed using a `c5d.24xlarge` EC2
+> instance.
+
+## Installing Libraries
+
+
+```python
+!pip install statsforecast mlforecast neuralforecast pyarrow
+```
+
+## Download and prepare data
+
+The example uses the [M5
+dataset](https://github.com/Mcompetitions/M5-methods/blob/master/M5-Competitors-Guide.pdf).
+It consists of `30,490` bottom time series.
+
+
+```python
+import pandas as pd
+```
+
+
+```python
+# Load the training target dataset from the provided URL
+Y_df = pd.read_parquet('https://m5-benchmarks.s3.amazonaws.com/data/train/target.parquet')
+
+# Rename columns to match the Nixtlaverse's expectations
+# The 'item_id' becomes 'unique_id' representing the unique identifier of the time series
+# The 'timestamp' becomes 'ds' representing the time stamp of the data points
+# The 'demand' becomes 'y' representing the target variable we want to forecast
+Y_df = Y_df.rename(
+ columns={
+ 'item_id': 'unique_id',
+ 'timestamp': 'ds',
+ 'demand': 'y'
+ }
+)
+
+# Convert the 'ds' column to datetime format to ensure proper handling of date-related operations in subsequent steps
+Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+```
+
+For simplicity sake we will keep just one category
+
+
+```python
+Y_df = Y_df.query('unique_id.str.startswith("FOODS_3")').reset_index(drop=True)
+Y_df['unique_id'] = Y_df['unique_id'].astype(str)
+```
+
+# Basic Plotting
+
+Plot some series using the `plot_series` function from the
+`utilsforecast` library. This method prints 8 random series from the
+dataset and is useful for basic EDA.
+
+
+```python
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+# Feature: plot random series for EDA
+plot_series(Y_df)
+```
+
+
+
+
+```python
+# Feature: plot groups of series for EDA
+plot_series(Y_df, ids=["FOODS_3_432_TX_2"])
+```
+
+
+
+# Create forecasts with Stats, Ml and Neural methods.
+
+## StatsForecast
+
+`StatsForecast` is a comprehensive library providing a suite of popular
+univariate time series forecasting models, all designed with a focus on
+high performance and scalability.
+
+Here’s what makes StatsForecast a powerful tool for time series
+forecasting:
+
+- **Collection of Local Models**: StatsForecast provides a diverse
+ collection of local models that can be applied to each time series
+ individually, allowing us to capture unique patterns within each
+ series.
+
+- **Simplicity**: With StatsForecast, training, forecasting, and
+ backtesting multiple models become a straightforward process,
+ requiring only a few lines of code. This simplicity makes it a
+ convenient tool for both beginners and experienced practitioners.
+
+- **Optimized for Speed**: The implementation of the models in
+ StatsForecast is optimized for speed, ensuring that large-scale
+ computations are performed efficiently, thereby reducing the overall
+ time for model training and prediction.
+
+- **Horizontal Scalability**: One of the distinguishing features of
+ StatsForecast is its ability to scale horizontally. It is compatible
+ with distributed computing frameworks such as Spark, Dask, and Ray.
+ This feature allows it to handle large datasets by distributing the
+ computations across multiple nodes in a cluster, making it a go-to
+ solution for large-scale time series forecasting tasks.
+
+`StatsForecast` receives a list of models to fit each time series. Since
+we are dealing with Daily data, it would be benefitial to use 7 as
+seasonality.
+
+
+```python
+from statsforecast import StatsForecast
+# Import necessary models from the statsforecast library
+from statsforecast.models import (
+ # SeasonalNaive: A model that uses the previous season's data as the forecast
+ SeasonalNaive,
+ # Naive: A simple model that uses the last observed value as the forecast
+ Naive,
+ # HistoricAverage: This model uses the average of all historical data as the forecast
+ HistoricAverage,
+ # CrostonOptimized: A model specifically designed for intermittent demand forecasting
+ CrostonOptimized,
+ # ADIDA: Adaptive combination of Intermittent Demand Approaches, a model designed for intermittent demand
+ ADIDA,
+ # IMAPA: Intermittent Multiplicative AutoRegressive Average, a model for intermittent series that incorporates autocorrelation
+ IMAPA,
+ # AutoETS: Automated Exponential Smoothing model that automatically selects the best Exponential Smoothing model based on AIC
+ AutoETS
+)
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+- `models`: a list of models. Select the models you want from models
+ and import them.
+- `freq`: a string indicating the frequency of the data. (See panda’s
+ available frequencies.)
+- `n_jobs`: int, number of jobs used in the parallel processing, use
+ -1 for all cores.
+- `fallback_model`: a model to be used if a model fails. Any settings
+ are passed into the constructor. Then you call its fit method and
+ pass in the historical data frame.
+
+
+```python
+horizon = 28
+models = [
+ SeasonalNaive(season_length=7),
+ Naive(),
+ HistoricAverage(),
+ CrostonOptimized(),
+ ADIDA(),
+ IMAPA(),
+ AutoETS(season_length=7)
+]
+```
+
+
+```python
+# Instantiate the StatsForecast class
+sf = StatsForecast(
+ models=models, # A list of models to be used for forecasting
+ freq='D', # The frequency of the time series data (in this case, 'D' stands for daily frequency)
+ n_jobs=-1, # The number of CPU cores to use for parallel execution (-1 means use all available cores)
+ verbose=True, # Show progress
+)
+```
+
+The forecast method produces predictions for the next `h` periods.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values.
+
+This block of code times how long it takes to run the forecasting
+function of the StatsForecast class, which predicts the next 28 days
+(h=28). The time is calculated in minutes and printed out at the end.
+
+
+```python
+from time import time
+
+# Get the current time before forecasting starts, this will be used to measure the execution time
+init = time()
+
+# Call the forecast method of the StatsForecast instance to predict the next 28 days (h=28)
+fcst_df = sf.forecast(df=Y_df, h=28)
+
+# Get the current time after the forecasting ends
+end = time()
+
+# Calculate and print the total time taken for the forecasting in minutes
+print(f'Forecast Minutes: {(end - init) / 60}')
+```
+
+``` text
+Forecast: 0%| | 0/2000 [Elapsed: 00:00]
+```
+
+``` text
+Forecast Minutes: 4.009805858135223
+```
+
+
+```python
+fcst_df.head()
+```
+
+| | unique_id | ds | SeasonalNaive | Naive | HistoricAverage | CrostonOptimized | ADIDA | IMAPA | AutoETS |
+|----|----|----|----|----|----|----|----|----|----|
+| 0 | FOODS_3_001_CA_1 | 2016-05-23 | 1.0 | 2.0 | 0.448738 | 0.345192 | 0.345477 | 0.347249 | 0.381414 |
+| 1 | FOODS_3_001_CA_1 | 2016-05-24 | 0.0 | 2.0 | 0.448738 | 0.345192 | 0.345477 | 0.347249 | 0.286933 |
+| 2 | FOODS_3_001_CA_1 | 2016-05-25 | 0.0 | 2.0 | 0.448738 | 0.345192 | 0.345477 | 0.347249 | 0.334987 |
+| 3 | FOODS_3_001_CA_1 | 2016-05-26 | 1.0 | 2.0 | 0.448738 | 0.345192 | 0.345477 | 0.347249 | 0.186851 |
+| 4 | FOODS_3_001_CA_1 | 2016-05-27 | 0.0 | 2.0 | 0.448738 | 0.345192 | 0.345477 | 0.347249 | 0.308112 |
+
+## MLForecast
+
+`MLForecast` is a powerful library that provides automated feature
+creation for time series forecasting, facilitating the use of global
+machine learning models. It is designed for high performance and
+scalability.
+
+Key features of MLForecast include:
+
+- **Support for sklearn models**: MLForecast is compatible with models
+ that follow the scikit-learn API. This makes it highly flexible and
+ allows it to seamlessly integrate with a wide variety of machine
+ learning algorithms.
+
+- **Simplicity**: With MLForecast, the tasks of training, forecasting,
+ and backtesting models can be accomplished in just a few lines of
+ code. This streamlined simplicity makes it user-friendly for
+ practitioners at all levels of expertise.
+
+- **Optimized for speed:** MLForecast is engineered to execute tasks
+ rapidly, which is crucial when handling large datasets and complex
+ models.
+
+- **Horizontal Scalability:** MLForecast is capable of horizontal
+ scaling using distributed computing frameworks such as Spark, Dask,
+ and Ray. This feature enables it to efficiently process massive
+ datasets by distributing the computations across multiple nodes in a
+ cluster, making it ideal for large-scale time series forecasting
+ tasks.
+
+
+```python
+from mlforecast import MLForecast
+from mlforecast.lag_transforms import ExpandingMean
+from mlforecast.target_transforms import Differences
+from mlforecast.utils import PredictionIntervals
+```
+
+
+```python
+!pip install lightgbm xgboost
+```
+
+
+```python
+# Import the necessary models from various libraries
+
+# LGBMRegressor: A gradient boosting framework that uses tree-based learning algorithms from the LightGBM library
+from lightgbm import LGBMRegressor
+
+# XGBRegressor: A gradient boosting regressor model from the XGBoost library
+from xgboost import XGBRegressor
+
+# LinearRegression: A simple linear regression model from the scikit-learn library
+from sklearn.linear_model import LinearRegression
+```
+
+To use `MLForecast` for time series forecasting, we instantiate a new
+`MLForecast` object and provide it with various parameters to tailor the
+modeling process to our specific needs:
+
+- `models`: This parameter accepts a list of machine learning models
+ you wish to use for forecasting. You can import your preferred
+ models from scikit-learn, lightgbm and xgboost.
+
+- `freq`: This is a string indicating the frequency of your data
+ (hourly, daily, weekly, etc.). The specific format of this string
+ should align with pandas’ recognized frequency strings.
+
+- `target_transforms`: These are transformations applied to the target
+ variable before model training and after model prediction. This can
+ be useful when working with data that may benefit from
+ transformations, such as log-transforms for highly skewed data.
+
+- `lags`: This parameter accepts specific lag values to be used as
+ regressors. Lags represent how many steps back in time you want to
+ look when creating features for your model. For example, if you want
+ to use the previous day’s data as a feature for predicting today’s
+ value, you would specify a lag of 1.
+
+- `lags_transforms`: These are specific transformations for each lag.
+ This allows you to apply transformations to your lagged features.
+
+- `date_features`: This parameter specifies date-related features to
+ be used as regressors. For instance, you might want to include the
+ day of the week or the month as a feature in your model.
+
+- `num_threads`: This parameter controls the number of threads to use
+ for parallelizing feature creation, helping to speed up this process
+ when working with large datasets.
+
+All these settings are passed to the `MLForecast` constructor. Once the
+`MLForecast` object is initialized with these settings, we call its
+`fit` method and pass the historical data frame as the argument. The
+`fit` method trains the models on the provided historical data, readying
+them for future forecasting tasks.
+
+
+```python
+# Instantiate the MLForecast object
+mlf = MLForecast(
+ models=[LGBMRegressor(verbosity=-1), XGBRegressor(), LinearRegression()], # List of models for forecasting: LightGBM, XGBoost and Linear Regression
+ freq='D', # Frequency of the data - 'D' for daily frequency
+ lags=list(range(1, 7)), # Specific lags to use as regressors: 1 to 6 days
+ lag_transforms = {
+ 1: [ExpandingMean()], # Apply expanding mean transformation to the lag of 1 day
+ },
+ date_features=['year', 'month', 'day', 'dayofweek', 'quarter', 'week'], # Date features to use as regressors
+)
+```
+
+Just call the `fit` models to train the select models. In this case we
+are generating conformal prediction intervals.
+
+
+```python
+# Start the timer to calculate the time taken for fitting the models
+init = time()
+
+# Fit the MLForecast models to the data
+mlf.fit(Y_df)
+
+# Calculate the end time after fitting the models
+end = time()
+
+# Print the time taken to fit the MLForecast models, in minutes
+print(f'MLForecast Minutes: {(end - init) / 60}')
+```
+
+``` text
+MLForecast Minutes: 0.5360581119855244
+```
+
+After that, just call `predict` to generate forecasts.
+
+
+```python
+fcst_mlf_df = mlf.predict(28)
+```
+
+
+```python
+fcst_mlf_df.head()
+```
+
+| | unique_id | ds | LGBMRegressor | XGBRegressor | LinearRegression |
+|-----|------------------|------------|---------------|--------------|------------------|
+| 0 | FOODS_3_001_CA_1 | 2016-05-23 | 0.549520 | 0.560123 | 0.332693 |
+| 1 | FOODS_3_001_CA_1 | 2016-05-24 | 0.553196 | 0.369337 | 0.055071 |
+| 2 | FOODS_3_001_CA_1 | 2016-05-25 | 0.599668 | 0.374338 | 0.127144 |
+| 3 | FOODS_3_001_CA_1 | 2016-05-26 | 0.638097 | 0.327176 | 0.101624 |
+| 4 | FOODS_3_001_CA_1 | 2016-05-27 | 0.763305 | 0.331631 | 0.269863 |
+
+## NeuralForecast
+
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+is a robust collection of neural forecasting models that focuses on
+usability and performance. It includes a variety of model architectures,
+from classic networks such as Multilayer Perceptrons (MLP) and Recurrent
+Neural Networks (RNN) to novel contributions like N-BEATS, N-HITS,
+Temporal Fusion Transformers (TFT), and more.
+
+Key features of
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+include:
+
+- A broad collection of global models. Out of the box implementation
+ of MLP, LSTM, RNN, TCN, DilatedRNN, NBEATS, NHITS, ESRNN, TFT,
+ Informer, PatchTST and HINT.
+- A simple and intuitive interface that allows training, forecasting,
+ and backtesting of various models in a few lines of code.
+- Support for GPU acceleration to improve computational speed.
+
+This machine doesn’t have GPU, but Google Colabs offers some for free.
+
+Using [Colab’s GPU to train
+NeuralForecast](https://nixtla.github.io/neuralforecast/docs/tutorials/intermittent_data.html).
+
+
+```python
+# Read the results from Colab
+fcst_nf_df = pd.read_parquet('https://m5-benchmarks.s3.amazonaws.com/data/forecast-nf.parquet')
+```
+
+
+```python
+fcst_nf_df.head()
+```
+
+| | unique_id | ds | AutoNHITS | AutoNHITS-lo-90 | AutoNHITS-hi-90 | AutoTFT | AutoTFT-lo-90 | AutoTFT-hi-90 |
+|----|----|----|----|----|----|----|----|----|
+| 0 | FOODS_3_001_CA_1 | 2016-05-23 | 0.0 | 0.0 | 2.0 | 0.0 | 0.0 | 2.0 |
+| 1 | FOODS_3_001_CA_1 | 2016-05-24 | 0.0 | 0.0 | 2.0 | 0.0 | 0.0 | 2.0 |
+| 2 | FOODS_3_001_CA_1 | 2016-05-25 | 0.0 | 0.0 | 2.0 | 0.0 | 0.0 | 1.0 |
+| 3 | FOODS_3_001_CA_1 | 2016-05-26 | 0.0 | 0.0 | 2.0 | 0.0 | 0.0 | 2.0 |
+| 4 | FOODS_3_001_CA_1 | 2016-05-27 | 0.0 | 0.0 | 2.0 | 0.0 | 0.0 | 2.0 |
+
+
+```python
+# Merge the forecasts from StatsForecast and NeuralForecast
+fcst_df = fcst_df.merge(fcst_nf_df, how='left', on=['unique_id', 'ds'])
+
+# Merge the forecasts from MLForecast into the combined forecast dataframe
+fcst_df = fcst_df.merge(fcst_mlf_df, how='left', on=['unique_id', 'ds'])
+```
+
+
+```python
+fcst_df.head()
+```
+
+| | unique_id | ds | SeasonalNaive | Naive | HistoricAverage | CrostonOptimized | ADIDA | IMAPA | AutoETS | AutoNHITS | AutoNHITS-lo-90 | AutoNHITS-hi-90 | AutoTFT | AutoTFT-lo-90 | AutoTFT-hi-90 | LGBMRegressor | XGBRegressor | LinearRegression |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | FOODS_3_001_CA_1 | 2016-05-23 | 1.0 | 2.0 | 0.448738 | 0.345192 | 0.345477 | 0.347249 | 0.381414 | 0.0 | 0.0 | 2.0 | 0.0 | 0.0 | 2.0 | 0.549520 | 0.560123 | 0.332693 |
+| 1 | FOODS_3_001_CA_1 | 2016-05-24 | 0.0 | 2.0 | 0.448738 | 0.345192 | 0.345477 | 0.347249 | 0.286933 | 0.0 | 0.0 | 2.0 | 0.0 | 0.0 | 2.0 | 0.553196 | 0.369337 | 0.055071 |
+| 2 | FOODS_3_001_CA_1 | 2016-05-25 | 0.0 | 2.0 | 0.448738 | 0.345192 | 0.345477 | 0.347249 | 0.334987 | 0.0 | 0.0 | 2.0 | 0.0 | 0.0 | 1.0 | 0.599668 | 0.374338 | 0.127144 |
+| 3 | FOODS_3_001_CA_1 | 2016-05-26 | 1.0 | 2.0 | 0.448738 | 0.345192 | 0.345477 | 0.347249 | 0.186851 | 0.0 | 0.0 | 2.0 | 0.0 | 0.0 | 2.0 | 0.638097 | 0.327176 | 0.101624 |
+| 4 | FOODS_3_001_CA_1 | 2016-05-27 | 0.0 | 2.0 | 0.448738 | 0.345192 | 0.345477 | 0.347249 | 0.308112 | 0.0 | 0.0 | 2.0 | 0.0 | 0.0 | 2.0 | 0.763305 | 0.331631 | 0.269863 |
+
+## Forecast plots
+
+
+```python
+plot_series(Y_df, fcst_df, max_insample_length=28 * 3)
+```
+
+
+
+Use the plot function to explore models and ID’s
+
+
+```python
+plot_series(
+ Y_df,
+ fcst_df,
+ max_insample_length=28 * 3,
+ models=['CrostonOptimized', 'AutoNHITS', 'SeasonalNaive', 'LGBMRegressor'],
+)
+```
+
+
+
+# Validate Model’s Performance
+
+The three libraries - `StatsForecast`, `MLForecast`, and
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast) -
+offer out-of-the-box cross-validation capabilities specifically designed
+for time series. This allows us to evaluate the model’s performance
+using historical data to obtain an unbiased assessment of how well each
+model is likely to perform on unseen data.
+
+## Cross Validation in StatsForecast
+
+The `cross_validation` method from the `StatsForecast` class accepts the
+following arguments:
+
+- `df`: A DataFrame representing the training data.
+- `h` (int): The forecast horizon, represented as the number of steps
+ into the future that we wish to predict. For example, if we’re
+ forecasting hourly data, `h=24` would represent a 24-hour forecast.
+- `step_size` (int): The step size between each cross-validation
+ window. This parameter determines how often we want to run the
+ forecasting process.
+- `n_windows` (int): The number of windows used for cross validation.
+ This parameter defines how many past forecasting processes we want
+ to evaluate.
+
+These parameters allow us to control the extent and granularity of our
+cross-validation process. By tuning these settings, we can balance
+between computational cost and the thoroughness of the cross-validation.
+
+
+```python
+sf.verbose = False
+init = time()
+cv_df = sf.cross_validation(df=Y_df, h=horizon, n_windows=3, step_size=horizon)
+end = time()
+print(f'CV Minutes: {(end - init) / 60}')
+```
+
+``` text
+CV Minutes: 10.829525109132131
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id` series identifier
+- `ds`: datestamp or temporal index
+- `cutoff`: the last datestamp or temporal index for the n_windows. If
+ n_windows=1, then one unique cuttoff value, if n_windows=2 then two
+ unique cutoff values.
+- `y`: true value
+- `"model"`: columns with the model’s name and fitted value.
+
+
+```python
+cv_df.head()
+```
+
+| | unique_id | ds | cutoff | y | SeasonalNaive | Naive | HistoricAverage | CrostonOptimized | ADIDA | IMAPA | AutoETS |
+|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | FOODS_3_001_CA_1 | 2016-02-29 | 2016-02-28 | 0.0 | 2.0 | 0.0 | 0.449111 | 0.618472 | 0.618375 | 0.617998 | 0.655286 |
+| 1 | FOODS_3_001_CA_1 | 2016-03-01 | 2016-02-28 | 1.0 | 0.0 | 0.0 | 0.449111 | 0.618472 | 0.618375 | 0.617998 | 0.568595 |
+| 2 | FOODS_3_001_CA_1 | 2016-03-02 | 2016-02-28 | 1.0 | 0.0 | 0.0 | 0.449111 | 0.618472 | 0.618375 | 0.617998 | 0.618805 |
+| 3 | FOODS_3_001_CA_1 | 2016-03-03 | 2016-02-28 | 0.0 | 1.0 | 0.0 | 0.449111 | 0.618472 | 0.618375 | 0.617998 | 0.455891 |
+| 4 | FOODS_3_001_CA_1 | 2016-03-04 | 2016-02-28 | 0.0 | 1.0 | 0.0 | 0.449111 | 0.618472 | 0.618375 | 0.617998 | 0.591197 |
+
+## MLForecast
+
+The `cross_validation` method from the `MLForecast` class takes the
+following arguments.
+
+- `df`: training data frame
+- `h` (int): represents the steps into the future that are being
+ forecasted. In this case, 24 hours ahead.
+- `step_size` (int): step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+- `n_windows` (int): number of windows used for cross-validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+
+```python
+init = time()
+cv_mlf_df = mlf.cross_validation(
+ df=Y_df,
+ h=horizon,
+ n_windows=3,
+)
+end = time()
+print(f'CV Minutes: {(end - init) / 60}')
+```
+
+``` text
+CV Minutes: 1.6215598344802857
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id` series identifier
+- `ds`: datestamp or temporal index
+- `cutoff`: the last datestamp or temporal index for the n_windows. If
+ n_windows=1, then one unique cuttoff value, if n_windows=2 then two
+ unique cutoff values.
+- `y`: true value
+- `"model"`: columns with the model’s name and fitted value.
+
+
+```python
+cv_mlf_df.head()
+```
+
+| | unique_id | ds | cutoff | y | LGBMRegressor | XGBRegressor | LinearRegression |
+|----|----|----|----|----|----|----|----|
+| 0 | FOODS_3_001_CA_1 | 2016-02-29 | 2016-02-28 | 0.0 | 0.435674 | 0.556261 | -0.353077 |
+| 1 | FOODS_3_001_CA_1 | 2016-03-01 | 2016-02-28 | 1.0 | 0.639676 | 0.625807 | -0.088985 |
+| 2 | FOODS_3_001_CA_1 | 2016-03-02 | 2016-02-28 | 1.0 | 0.792989 | 0.659651 | 0.217697 |
+| 3 | FOODS_3_001_CA_1 | 2016-03-03 | 2016-02-28 | 0.0 | 0.806868 | 0.535121 | 0.438713 |
+| 4 | FOODS_3_001_CA_1 | 2016-03-04 | 2016-02-28 | 0.0 | 0.829106 | 0.313354 | 0.637066 |
+
+## NeuralForecast
+
+This machine doesn’t have GPU, but Google Colabs offers some for free.
+
+Using [Colab’s GPU to train
+NeuralForecast](https://nixtla.github.io/neuralforecast/docs/tutorials/intermittent_data.html).
+
+
+```python
+cv_nf_df = pd.read_parquet('https://m5-benchmarks.s3.amazonaws.com/data/cross-validation-nf.parquet')
+```
+
+
+```python
+cv_nf_df.head()
+```
+
+| | unique_id | ds | cutoff | AutoNHITS | AutoNHITS-lo-90 | AutoNHITS-hi-90 | AutoTFT | AutoTFT-lo-90 | AutoTFT-hi-90 | y |
+|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | FOODS_3_001_CA_1 | 2016-02-29 | 2016-02-28 | 0.0 | 0.0 | 2.0 | 1.0 | 0.0 | 2.0 | 0.0 |
+| 1 | FOODS_3_001_CA_1 | 2016-03-01 | 2016-02-28 | 0.0 | 0.0 | 2.0 | 1.0 | 0.0 | 2.0 | 1.0 |
+| 2 | FOODS_3_001_CA_1 | 2016-03-02 | 2016-02-28 | 0.0 | 0.0 | 2.0 | 1.0 | 0.0 | 2.0 | 1.0 |
+| 3 | FOODS_3_001_CA_1 | 2016-03-03 | 2016-02-28 | 0.0 | 0.0 | 2.0 | 1.0 | 0.0 | 2.0 | 0.0 |
+| 4 | FOODS_3_001_CA_1 | 2016-03-04 | 2016-02-28 | 0.0 | 0.0 | 2.0 | 1.0 | 0.0 | 2.0 | 0.0 |
+
+## Merge cross validation forecasts
+
+
+```python
+cv_df = cv_df.merge(cv_nf_df.drop(columns=['y']), how='left', on=['unique_id', 'ds', 'cutoff'])
+cv_df = cv_df.merge(cv_mlf_df.drop(columns=['y']), how='left', on=['unique_id', 'ds', 'cutoff'])
+```
+
+## Plots CV
+
+
+```python
+cutoffs = cv_df['cutoff'].unique()
+```
+
+
+```python
+for cutoff in cutoffs:
+ display(
+ plot_series(
+ Y_df,
+ cv_df.query('cutoff == @cutoff').drop(columns=['y', 'cutoff']),
+ max_insample_length=28 * 5,
+ ids=['FOODS_3_001_CA_1'],
+ )
+ )
+```
+
+
+
+
+
+
+
+### Aggregate Demand
+
+
+```python
+agg_cv_df = cv_df.loc[:,~cv_df.columns.str.contains('hi|lo')].groupby(['ds', 'cutoff']).sum(numeric_only=True).reset_index()
+agg_cv_df.insert(0, 'unique_id', 'agg_demand')
+```
+
+
+```python
+agg_Y_df = Y_df.groupby(['ds']).sum(numeric_only=True).reset_index()
+agg_Y_df.insert(0, 'unique_id', 'agg_demand')
+```
+
+
+```python
+for cutoff in cutoffs:
+ display(
+ plot_series(
+ agg_Y_df,
+ agg_cv_df.query('cutoff == @cutoff').drop(columns=['y', 'cutoff']),
+ max_insample_length=28 * 5,
+ )
+ )
+```
+
+
+
+
+
+
+
+## Evaluation per series and CV window
+
+In this section, we will evaluate the performance of each model for each
+time series.
+
+
+```python
+from utilsforecast.evaluation import evaluate
+from utilsforecast.losses import mse, mae, smape
+```
+
+
+```python
+evaluation_df = evaluate(cv_df.drop(columns='cutoff'), metrics=[mse, mae, smape])
+evaluation_df
+```
+
+| | unique_id | metric | SeasonalNaive | Naive | HistoricAverage | CrostonOptimized | ADIDA | IMAPA | AutoETS | AutoNHITS | AutoTFT | LGBMRegressor | XGBRegressor | LinearRegression |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | FOODS_3_001_CA_1 | mse | 1.250000 | 0.892857 | 0.485182 | 0.507957 | 0.509299 | 0.516988 | 0.494235 | 0.630952 | 0.571429 | 0.648962 | 0.584722 | 0.529400 |
+| 1 | FOODS_3_001_CA_2 | mse | 6.273809 | 3.773809 | 3.477309 | 3.412580 | 3.432295 | 3.474050 | 3.426468 | 4.550595 | 3.607143 | 3.423646 | 3.856465 | 3.773264 |
+| 2 | FOODS_3_001_CA_3 | mse | 5.880952 | 4.357143 | 5.016396 | 4.173154 | 4.160645 | 4.176733 | 4.145148 | 4.005952 | 4.372024 | 4.928764 | 6.937792 | 5.317195 |
+| 3 | FOODS_3_001_CA_4 | mse | 1.071429 | 0.476190 | 0.402938 | 0.382559 | 0.380783 | 0.380877 | 0.380872 | 0.476190 | 0.476190 | 0.664270 | 0.424068 | 0.637221 |
+| 4 | FOODS_3_001_TX_1 | mse | 0.047619 | 0.047619 | 0.238824 | 0.261356 | 0.047619 | 0.047619 | 0.077575 | 0.047619 | 0.047619 | 0.718796 | 0.063564 | 0.187810 |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| 24685 | FOODS_3_827_TX_2 | smape | 0.083333 | 0.035714 | 0.989540 | 0.996362 | 0.987395 | 0.982847 | 0.981537 | 0.323810 | 0.335714 | 0.976356 | 0.994702 | 0.985058 |
+| 24686 | FOODS_3_827_TX_3 | smape | 0.708532 | 0.681495 | 0.662490 | 0.653057 | 0.655810 | 0.660161 | 0.649180 | 0.683947 | 0.712121 | 0.639518 | 0.856866 | 0.686547 |
+| 24687 | FOODS_3_827_WI_1 | smape | 0.608722 | 0.694328 | 0.470570 | 0.470846 | 0.480032 | 0.480032 | 0.466956 | 0.486852 | 0.475980 | 0.472336 | 0.484906 | 0.492277 |
+| 24688 | FOODS_3_827_WI_2 | smape | 0.531777 | 0.398156 | 0.433577 | 0.387718 | 0.388827 | 0.389371 | 0.389888 | 0.393774 | 0.374640 | 0.413559 | 0.430893 | 0.399131 |
+| 24689 | FOODS_3_827_WI_3 | smape | 0.643689 | 0.680178 | 0.588031 | 0.589143 | 0.599820 | 0.628673 | 0.591437 | 0.558201 | 0.567460 | 0.589870 | 0.698798 | 0.627255 |
+
+
+```python
+by_metric = evaluation_df.groupby('metric').mean(numeric_only=True)
+by_metric
+```
+
+| | SeasonalNaive | Naive | HistoricAverage | CrostonOptimized | ADIDA | IMAPA | AutoETS | AutoNHITS | AutoTFT | LGBMRegressor | XGBRegressor | LinearRegression |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| metric | | | | | | | | | | | | |
+| mae | 1.775415 | 2.045906 | 1.749080 | 1.634791 | 1.542097 | 1.543745 | 1.511545 | 1.438250 | 1.497647 | 1.697947 | 1.552061 | 1.592978 |
+| mse | 14.265773 | 20.453325 | 12.938136 | 11.484233 | 11.090195 | 11.094446 | 10.351927 | 9.606913 | 10.721251 | 10.502289 | 11.565916 | 10.830894 |
+| smape | 0.436414 | 0.446430 | 0.616884 | 0.613219 | 0.618910 | 0.619313 | 0.620084 | 0.400770 | 0.411018 | 0.579856 | 0.693615 | 0.641515 |
+
+Best models by metric
+
+
+```python
+by_metric.idxmin(axis=1)
+```
+
+``` text
+metric
+mae AutoNHITS
+mse AutoNHITS
+smape AutoNHITS
+dtype: object
+```
+
+### Distribution of errors
+
+
+```python
+!pip install seaborn
+```
+
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+```
+
+
+```python
+evaluation_df_long = pd.melt(evaluation_df, id_vars=['unique_id', 'metric'], var_name='model', value_name='error')
+```
+
+#### SMAPE
+
+
+```python
+sns.violinplot(evaluation_df_long.query('metric=="smape"'), x='error', y='model');
+```
+
+
+
+### Choose models for groups of series
+
+Feature:
+
+- A unified dataframe with forecasts for all different models
+- Easy Ensamble
+- E.g. Average predictions
+- Or MinMax (Choosing is ensembling)
+
+
+```python
+# Choose the best model for each time series, metric, and cross validation window
+evaluation_df['best_model'] = evaluation_df.idxmin(axis=1, numeric_only=True)
+# count how many times a model wins per metric and cross validation window
+count_best_model = evaluation_df.groupby(['metric', 'best_model']).size().rename('n').to_frame().reset_index()
+# plot results
+sns.barplot(count_best_model, x='n', y='best_model', hue='metric')
+```
+
+
+
+### Et pluribus unum: an inclusive forecasting Pie.
+
+
+```python
+# For the mse, calculate how many times a model wins
+eval_series_df = evaluation_df.query('metric == "mse"').groupby(['unique_id']).mean(numeric_only=True)
+eval_series_df['best_model'] = eval_series_df.idxmin(axis=1)
+counts_series = eval_series_df.value_counts('best_model')
+plt.pie(counts_series, labels=counts_series.index, autopct='%.0f%%')
+plt.show()
+```
+
+
+
+
+```python
+plot_series(
+ Y_df,
+ cv_df.drop(columns=['cutoff', 'y']),
+ max_insample_length=28 * 6,
+ models=['AutoNHITS'],
+)
+```
+
+
+
+# Choose Forecasting method for different groups of series
+
+
+```python
+# Merge the best model per time series dataframe
+# and filter the forecasts based on that dataframe
+# for each time series
+fcst_df = pd.melt(fcst_df.set_index('unique_id'), id_vars=['ds'], var_name='model', value_name='forecast', ignore_index=False)
+fcst_df = fcst_df.join(eval_series_df[['best_model']])
+fcst_df[['model', 'pred-interval']] = fcst_df['model'].str.split('-', expand=True, n=1)
+fcst_df = fcst_df.query('model == best_model')
+fcst_df['name'] = [f'forecast-{x}' if x is not None else 'forecast' for x in fcst_df['pred-interval']]
+fcst_df = pd.pivot_table(fcst_df, index=['unique_id', 'ds'], values=['forecast'], columns=['name']).droplevel(0, axis=1).reset_index()
+```
+
+
+```python
+plot_series(Y_df, fcst_df, max_insample_length=28 * 3)
+```
+
+
+
+# Further materials
+
+- [Available Models
+ StatsForecast](https://nixtlaverse.nixtla.io/statsforecast/index.html#models)
+- [Available Models
+ NeuralForecast](https://nixtlaverse.nixtla.io/neuralforecast/docs/capabilities/overview.html)
+- [Loss Functions in
+ NeuralForecast](https://nixtlaverse.nixtla.io/neuralforecast/docs/capabilities/objectives.html)
+- [Getting Started
+ NeuralForecast](https://nixtlaverse.nixtla.io/neuralforecast/docs/getting-started/quickstart.html)
+- [Hierarchical
+ Reconciliation](https://nixtlaverse.nixtla.io/hierarchicalforecast/examples/tourismsmall.html)
+- [Distributed ML Forecast
+ (trees)](https://nixtlaverse.nixtla.io/mlforecast/docs/getting-started/quick_start_distributed.html)
+- [Using StatsForecast to train millions of time
+ series](https://www.anyscale.com/blog/how-nixtla-uses-ray-to-accurately-predict-more-than-a-million-time-series)
+- [Intermittent Demand Forecasting With Nixtla on
+ Databricks](https://www.databricks.com/blog/2022/12/06/intermittent-demand-forecasting-nixtla-databricks.html)
+
diff --git a/neuralforecast/docs/tutorials/configure_optimizers.html.mdx b/neuralforecast/docs/tutorials/configure_optimizers.html.mdx
new file mode 100644
index 00000000..b88dcbee
--- /dev/null
+++ b/neuralforecast/docs/tutorials/configure_optimizers.html.mdx
@@ -0,0 +1,190 @@
+---
+description: >-
+ Tutorial on how to achieve a full control of the `configure_optimizers()`
+ behavior of NeuralForecast models
+output-file: configure_optimizers.html
+title: Modify the configure_optimizers() behavior of NeuralForecast models
+---
+
+
+NeuralForecast models allow us to customize the default optimizer and
+learning rate scheduler behaviors via `optimizer`, `optimizer_kwargs`,
+`lr_scheduler`, `lr_scheduler_kwargs`. However this is not sufficient to
+support the use of
+[ReduceLROnPlateau](https://pytorch.org/docs/stable/generated/torch.optim.lr_scheduler.ReduceLROnPlateau.html),
+for instance, as it requires the specification of `monitor` parameter.
+
+This tutorial provides an example of how to support the use of
+`ReduceLROnPlateau`.
+
+## Load libraries
+
+
+```python
+import numpy as np
+import pandas as pd
+import torch
+import matplotlib.pyplot as plt
+from neuralforecast import NeuralForecast
+from neuralforecast.models import NHITS
+from neuralforecast.utils import AirPassengersPanel
+
+from utilsforecast.plotting import plot_series
+```
+
+``` text
+/root/miniconda3/envs/neuralforecast/lib/python3.10/site-packages/tqdm/auto.py:21: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
+ from .autonotebook import tqdm as notebook_tqdm
+2025-02-25 15:57:21,708 INFO util.py:154 -- Missing packages: ['ipywidgets']. Run `pip install -U ipywidgets`, then restart the notebook server for rich notebook output.
+2025-02-25 15:57:21,760 INFO util.py:154 -- Missing packages: ['ipywidgets']. Run `pip install -U ipywidgets`, then restart the notebook server for rich notebook output.
+```
+
+## Data
+
+We use the AirPassengers dataset for the demonstration of conformal
+prediction.
+
+
+```python
+AirPassengersPanel_train = AirPassengersPanel[AirPassengersPanel['ds'] < AirPassengersPanel['ds'].values[-12]].reset_index(drop=True)
+AirPassengersPanel_test = AirPassengersPanel[AirPassengersPanel['ds'] >= AirPassengersPanel['ds'].values[-12]].reset_index(drop=True)
+AirPassengersPanel_test['y'] = np.nan
+AirPassengersPanel_test['y_[lag12]'] = np.nan
+```
+
+## Model training
+
+We now train a NHITS model on the above dataset. We consider two
+different predictions: 1. Training using the default
+`configure_optimizers()`. 2. Training by overwriting the
+`configure_optimizers()` of the subclass of NHITS model.
+
+
+```python
+horizon = 12
+input_size = 24
+
+class CustomNHITS(NHITS):
+ def configure_optimizers(self):
+ optimizer = torch.optim.Adadelta(params=self.parameters(), rho=0.75)
+ scheduler=torch.optim.lr_scheduler.ReduceLROnPlateau(
+ optimizer=optimizer, mode='min',factor=0.5, patience=2,
+ )
+ scheduler_config = {
+ 'scheduler': scheduler,
+ 'interval': 'step',
+ 'frequency': 1,
+ 'monitor': 'train_loss',
+ 'strict': True,
+ 'name': None,
+ }
+ return {'optimizer': optimizer, 'lr_scheduler': scheduler_config}
+
+models = [
+ NHITS(h=horizon, input_size=input_size, max_steps=100, alias='NHITS-default-scheduler'),
+ CustomNHITS(h=horizon, input_size=input_size, max_steps=100, alias='NHITS-ReduceLROnPlateau-scheduler'),
+]
+nf = NeuralForecast(models=models, freq='ME')
+nf.fit(AirPassengersPanel_train)
+preds = nf.predict(futr_df=AirPassengersPanel_test)
+```
+
+``` text
+Seed set to 1
+Seed set to 1
+GPU available: False, used: False
+TPU available: False, using: 0 TPU cores
+HPU available: False, using: 0 HPUs
+
+ | Name | Type | Params | Mode
+-------------------------------------------------------
+0 | loss | MAE | 0 | train
+1 | padder_train | ConstantPad1d | 0 | train
+2 | scaler | TemporalNorm | 0 | train
+3 | blocks | ModuleList | 2.4 M | train
+-------------------------------------------------------
+2.4 M Trainable params
+0 Non-trainable params
+2.4 M Total params
+9.751 Total estimated model params size (MB)
+34 Modules in train mode
+0 Modules in eval mode
+```
+
+``` text
+Epoch 99: 100%|██████████| 1/1 [00:00<00:00, 2.50it/s, v_num=85, train_loss_step=14.20, train_loss_epoch=14.20]
+```
+
+``` text
+`Trainer.fit` stopped: `max_steps=100` reached.
+```
+
+``` text
+Epoch 99: 100%|██████████| 1/1 [00:00<00:00, 2.49it/s, v_num=85, train_loss_step=14.20, train_loss_epoch=14.20]
+```
+
+``` text
+GPU available: False, used: False
+TPU available: False, using: 0 TPU cores
+HPU available: False, using: 0 HPUs
+
+ | Name | Type | Params | Mode
+-------------------------------------------------------
+0 | loss | MAE | 0 | train
+1 | padder_train | ConstantPad1d | 0 | train
+2 | scaler | TemporalNorm | 0 | train
+3 | blocks | ModuleList | 2.4 M | train
+-------------------------------------------------------
+2.4 M Trainable params
+0 Non-trainable params
+2.4 M Total params
+9.751 Total estimated model params size (MB)
+34 Modules in train mode
+0 Modules in eval mode
+```
+
+``` text
+
+Epoch 99: 100%|██████████| 1/1 [00:00<00:00, 2.78it/s, v_num=86, train_loss_step=24.10, train_loss_epoch=24.10]
+```
+
+``` text
+`Trainer.fit` stopped: `max_steps=100` reached.
+```
+
+``` text
+Epoch 99: 100%|██████████| 1/1 [00:00<00:00, 2.77it/s, v_num=86, train_loss_step=24.10, train_loss_epoch=24.10]
+```
+
+``` text
+GPU available: False, used: False
+TPU available: False, using: 0 TPU cores
+HPU available: False, using: 0 HPUs
+```
+
+``` text
+
+Predicting DataLoader 0: 100%|██████████| 1/1 [00:00<00:00, 33.39it/s]
+```
+
+``` text
+GPU available: False, used: False
+TPU available: False, using: 0 TPU cores
+HPU available: False, using: 0 HPUs
+```
+
+``` text
+
+Predicting DataLoader 0: 100%|██████████| 1/1 [00:00<00:00, 246.29it/s]
+```
+
+
+```python
+plot_series(AirPassengersPanel_train, preds)
+```
+
+
+
+We can clearly notice the prediction outputs are different due to the
+change in `configure_optimizers()`.
+
diff --git a/neuralforecast/docs/tutorials/conformal_prediction.html.mdx b/neuralforecast/docs/tutorials/conformal_prediction.html.mdx
new file mode 100644
index 00000000..be03ccdb
--- /dev/null
+++ b/neuralforecast/docs/tutorials/conformal_prediction.html.mdx
@@ -0,0 +1,139 @@
+---
+description: >-
+ Tutorial on how to train neuralforecast models and obtain prediction intervals
+ using the conformal prediction methods
+output-file: conformal_prediction.html
+title: Uncertainty quantification with Conformal Prediction
+---
+
+
+Conformal prediction uses cross-validation on a model trained with a
+point loss function to generate prediction intervals. No additional
+training is needed, and the model is treated as a black box. The
+approach is compatible with any model.
+
+In this notebook, we demonstrate how to obtain prediction intervals
+using conformal prediction.
+
+## Load libraries
+
+
+```python
+import logging
+
+import numpy as np
+import pandas as pd
+import matplotlib.pyplot as plt
+from neuralforecast import NeuralForecast
+from neuralforecast.models import NHITS
+from neuralforecast.utils import AirPassengersPanel
+from neuralforecast.utils import PredictionIntervals
+from neuralforecast.losses.pytorch import DistributionLoss, MAE
+```
+
+
+```python
+logging.getLogger('pytorch_lightning').setLevel(logging.ERROR)
+```
+
+## Data
+
+We use the AirPassengers dataset for the demonstration of conformal
+prediction.
+
+
+```python
+AirPassengersPanel_train = AirPassengersPanel[AirPassengersPanel['ds'] < AirPassengersPanel['ds'].values[-12]].reset_index(drop=True)
+AirPassengersPanel_test = AirPassengersPanel[AirPassengersPanel['ds'] >= AirPassengersPanel['ds'].values[-12]].reset_index(drop=True)
+AirPassengersPanel_test['y'] = np.nan
+AirPassengersPanel_test['y_[lag12]'] = np.nan
+```
+
+## Model training
+
+We now train a NHITS model on the above dataset. To support conformal
+predictions, we must first instantiate the
+[`PredictionIntervals`](https://nixtlaverse.nixtla.io/neuralforecast/utils.html#predictionintervals)
+class and pass this to the `fit` method. By default,
+[`PredictionIntervals`](https://nixtlaverse.nixtla.io/neuralforecast/utils.html#predictionintervals)
+class employs `n_windows=2` for the corss-validation during the
+computation of conformity scores. We also train a MLP model using
+DistributionLoss to demonstate the difference between conformal
+prediction and quantiled outputs.
+
+By default,
+[`PredictionIntervals`](https://nixtlaverse.nixtla.io/neuralforecast/utils.html#predictionintervals)
+class employs `method=conformal_distribution` for the conformal
+predictions, but it also supports `method=conformal_error`. The
+`conformal_distribution` method calculates forecast paths using the
+absolute errors and based on them calculates quantiles. The
+`conformal_error` method calculates quantiles directly from errors.
+
+We consider two models below:
+
+1. A model trained using a point loss function
+ ([`MAE`](https://nixtlaverse.nixtla.io/neuralforecast/losses.pytorch.html#mae)),
+ where we quantify the uncertainty using conformal prediction. This
+ case is labeled with
+ [`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits).
+2. A model trained using a `DistributionLoss('Normal')`, where we
+ quantify the uncertainty by training the model to fit the parameters
+ of a Normal distribution. This case is labeled with `NHITS1`.
+
+
+```python
+horizon = 12
+input_size = 24
+
+prediction_intervals = PredictionIntervals()
+
+models = [NHITS(h=horizon, input_size=input_size, max_steps=100, loss=MAE(), scaler_type="robust"),
+ NHITS(h=horizon, input_size=input_size, max_steps=100, loss=DistributionLoss("Normal", level=[90]), scaler_type="robust")]
+nf = NeuralForecast(models=models, freq='ME')
+nf.fit(AirPassengersPanel_train, prediction_intervals=prediction_intervals)
+```
+
+## Forecasting
+
+To generate conformal intervals, we specify the desired levels in the
+`predict` method.
+
+
+```python
+preds = nf.predict(futr_df=AirPassengersPanel_test, level=[90])
+```
+
+
+```python
+fig, (ax1, ax2) = plt.subplots(2, 1, figsize = (20, 7))
+plot_df = pd.concat([AirPassengersPanel_train, preds])
+
+plot_df = plot_df[plot_df['unique_id']=='Airline1'].drop(['unique_id','trend','y_[lag12]'], axis=1).iloc[-50:]
+
+ax1.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+ax1.plot(plot_df['ds'], plot_df['NHITS'], c='blue', label='median')
+ax1.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['NHITS-lo-90'][-12:].values,
+ y2=plot_df['NHITS-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+ax1.set_title('AirPassengers Forecast - Uncertainty quantification using Conformal Prediction', fontsize=18)
+ax1.set_ylabel('Monthly Passengers', fontsize=15)
+ax1.set_xticklabels([])
+ax1.legend(prop={'size': 10})
+ax1.grid()
+
+ax2.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+ax2.plot(plot_df['ds'], plot_df['NHITS1'], c='blue', label='median')
+ax2.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['NHITS1-lo-90'][-12:].values,
+ y2=plot_df['NHITS1-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+ax2.set_title('AirPassengers Forecast - Uncertainty quantification using Normal distribution', fontsize=18)
+ax2.set_ylabel('Monthly Passengers', fontsize=15)
+ax2.set_xlabel('Timestamp [t]', fontsize=15)
+ax2.legend(prop={'size': 10})
+ax2.grid()
+```
+
+
+
diff --git a/neuralforecast/docs/tutorials/cross_validation.html.mdx b/neuralforecast/docs/tutorials/cross_validation.html.mdx
new file mode 100644
index 00000000..04b9fc40
--- /dev/null
+++ b/neuralforecast/docs/tutorials/cross_validation.html.mdx
@@ -0,0 +1,372 @@
+---
+description: Implement cross-validation to evaluate models on historical data
+output-file: cross_validation.html
+title: Cross-validation
+---
+
+
+Time series cross-validation is a method for evaluating how a model
+would have performed on historical data. It works by defining a sliding
+window across past observations and predicting the period following it.
+It differs from standard cross-validation by maintaining the
+chronological order of the data instead of randomly splitting it.
+
+This method allows for a better estimation of our model’s predictive
+capabilities by considering multiple periods. When only one window is
+used, it resembles a standard train-test split, where the test data is
+the last set of observations, and the training set consists of the
+earlier data.
+
+The following graph showcases how time series cross-validation works.
+
+
+
+In this tutorial we’ll explain how to perform cross-validation in
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast).
+
+**Outline:** 1. Install NeuralForecast
+
+1. Load and plot the data
+
+2. Train multiple models using cross-validation
+
+3. Evaluate models and select the best for each series
+
+4. Plot cross-validation results
+
+> **Prerequesites**
+>
+> This guide assumes basic familiarity with `neuralforecast`. For a
+> minimal example visit the [Quick
+> Start](../getting-started/quickstart.html)
+
+## 1. Install NeuralForecast
+
+
+```python
+!pip install neuralforecast
+```
+
+## 2. Load and plot the data
+
+We’ll use pandas to load the hourly dataset from the [M4 Forecasting
+Competition](https://www.sciencedirect.com/science/article/pii/S0169207019301128),
+which has been stored in a parquet file for efficiency.
+
+
+```python
+import os
+import logging
+
+import pandas as pd
+from IPython.display import display
+```
+
+
+```python
+os.environ['PL_TRAINER_ENABLE_PROGRESS_BAR'] = '0'
+logging.getLogger('pytorch_lightning').setLevel(logging.ERROR)
+```
+
+
+```python
+Y_df = pd.read_parquet('https://datasets-nixtla.s3.amazonaws.com/m4-hourly.parquet')
+Y_df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|-----|-------|
+| 0 | H1 | 1 | 605.0 |
+| 1 | H1 | 2 | 586.0 |
+| 2 | H1 | 3 | 586.0 |
+| 3 | H1 | 4 | 559.0 |
+| 4 | H1 | 5 | 511.0 |
+
+The input to `neuralforecast` should be a data frame in long format with
+three columns: `unique_id`, `ds`, and `y`.
+
+- `unique_id` (string, int, or category): A unique identifier for each
+ time series.
+
+- `ds` (int or timestamp): An integer indexing time or a timestamp in
+ format YYYY-MM-DD or YYYY-MM-DD HH:MM:SS.
+
+- `y` (numeric): The target variable to forecast.
+
+This dataset contains 414 unique time series. To reduce the total
+execution time, we’ll use only the first 10.
+
+
+```python
+uids = Y_df['unique_id'].unique()[:10] # Select 10 ids to make the example run faster
+Y_df = Y_df.query('unique_id in @uids').reset_index(drop=True)
+```
+
+To plot the series, we’ll use the `plot_series` method from
+`utilsforecast.plotting`. `utilsforecast` is a dependency of
+`neuralforecast` so it should be already installed.
+
+
+```python
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+plot_series(Y_df)
+```
+
+
+
+## 3. Train multiple models using cross-validation
+
+We’ll train different models from `neuralforecast` using the
+`cross-validation` method to decide which one perfoms best on the
+historical data. To do this, we need to import the
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+class and the models that we want to compare.
+
+
+```python
+from neuralforecast import NeuralForecast
+from neuralforecast.auto import MLP, NBEATS, NHITS
+from neuralforecast.losses.pytorch import MQLoss
+```
+
+In this tutorial, we will use `neuralforecast's`
+[MPL](https://nixtlaverse.nixtla.io/neuralforecast/models.mlp.html),
+[NBEATS](https://nixtlaverse.nixtla.io/neuralforecast/models.nbeats.html),
+and
+[NHITS](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html)
+models.
+
+First, we need to create a list of models and then instantiate the
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+class. For each model, we’ll define the following hyperparameters:
+
+- `h`: The forecast horizon. Here, we will use the same horizon as in
+ the M4 competition, which was 48 steps ahead.
+
+- `input_size`: The number of historical observations (lags) that the
+ model uses to make predictions. In this case, it will be twice the
+ forecast horizon.
+
+- `loss`: The loss function to optimize. Here, we’ll use the Multi
+ Quantile Loss (MQLoss) from `neuralforecast.losses.pytorch`.
+
+> **Warning**
+>
+> The Multi Quantile Loss (MQLoss) is the sum of the quantile losses for
+> each target quantile. The quantile loss for a single quantile measures
+> how well a model has predicted a specific quantile of the actual
+> distribution, penalizing overestimations and underestimations
+> asymmetrically based on the quantile’s value. For more details see
+> [here](https://nixtlaverse.nixtla.io/neuralforecast/losses.pytorch.html#multi-quantile-loss-mqloss).
+
+While there are other hyperparameters that can be defined for each
+model, we’ll use the default values for the purposes of this tutorial.
+To learn more about the hyperparameters of each model, please check out
+the corresponding documentation.
+
+
+```python
+horizon = 48
+models = [MLP(h=horizon, input_size=2*horizon, loss=MQLoss()),
+ NBEATS(h=horizon, input_size=2*horizon, loss=MQLoss()),
+ NHITS(h=horizon, input_size=2*horizon, loss=MQLoss()),]
+nf = NeuralForecast(models=models, freq=1)
+```
+
+The `cross_validation` method takes the following arguments:
+
+- `df`: The data frame in the format described in section 2.
+
+- `n_windows` (int): The number of windows to evaluate. Default is 1
+ and here we’ll use 3.
+
+- `step_size` (int): The number of steps between consecutive windows
+ to produce the forecasts. In this example, we’ll set
+ `step_size=horizon` to produce non-overlapping forecasts. The
+ following diagram shows how the forecasts are produced based on the
+ `step_size` parameter and forecast horizon `h` of a model. In this
+ diagram `step_size=2` and `h=4`.
+
+
+
+- `refit` (bool or int): Whether to retrain models for each
+ cross-validation window. If `False`, the models are trained at the
+ beginning and then used to predict each window. If a positive
+ integer, the models are retrained every `refit` windows. Default is
+ `False`, but here we’ll use `refit=1` so that the models are
+ retrained after each window using the data with timestamps up to and
+ including the cutoff.
+
+
+```python
+cv_df = nf.cross_validation(Y_df, n_windows=3, step_size=horizon, refit=1)
+```
+
+It’s worth mentioning that the default version of the `cross_validation`
+method in `neuralforecast` diverges from other libraries, where models
+are typically retrained at the start of each window. By default, it
+trains the models once and then uses them to generate predictions over
+all the windows, thus reducing the total execution time. For scenarios
+where the models need to be retrained, you can use the `refit` parameter
+to specify the number of windows after which the models should be
+retrained.
+
+
+```python
+cv_df.head()
+```
+
+| | unique_id | ds | cutoff | MLP-median | MLP-lo-90 | MLP-lo-80 | MLP-hi-80 | MLP-hi-90 | NBEATS-median | NBEATS-lo-90 | NBEATS-lo-80 | NBEATS-hi-80 | NBEATS-hi-90 | NHITS-median | NHITS-lo-90 | NHITS-lo-80 | NHITS-hi-80 | NHITS-hi-90 | y |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | H1 | 605 | 604 | 638.964111 | 528.127747 | 546.731812 | 714.415466 | 750.265259 | 623.230896 | 580.549744 | 587.317688 | 647.942505 | 654.148682 | 625.377930 | 556.786926 | 577.746765 | 657.901611 | 670.458069 | 622.0 |
+| 1 | H1 | 606 | 604 | 588.216370 | 445.395081 | 483.736542 | 684.394592 | 670.042358 | 552.829407 | 501.618988 | 529.007507 | 593.528564 | 603.152527 | 555.956177 | 511.696350 | 526.399597 | 604.318970 | 622.839722 | 558.0 |
+| 2 | H1 | 607 | 604 | 542.242737 | 419.206757 | 439.244476 | 617.775269 | 638.583923 | 495.155548 | 451.871613 | 467.183533 | 550.048950 | 574.697021 | 502.860077 | 462.284668 | 460.950287 | 555.336731 | 571.852722 | 513.0 |
+| 3 | H1 | 608 | 604 | 494.055573 | 414.775085 | 427.531647 | 583.965759 | 602.303772 | 465.182556 | 403.593140 | 410.033203 | 500.744019 | 518.277954 | 460.588684 | 406.762390 | 418.040710 | 501.833740 | 515.022095 | 476.0 |
+| 4 | H1 | 609 | 604 | 469.330688 | 361.437927 | 378.501373 | 557.875244 | 569.767273 | 441.072388 | 371.541504 | 401.923584 | 483.667877 | 485.047729 | 441.463043 | 393.917725 | 394.483337 | 475.985229 | 499.001373 | 449.0 |
+
+The output of the `cross-validation` method is a data frame that
+includes the following columns:
+
+- `unique_id`: The unique identifier for each time series.
+
+- `ds`: The timestamp or temporal index.
+
+- `cutoff`: The last timestamp or temporal index used in that
+ cross-validation window.
+
+- `"model"`: Columns with the model’s point forecasts (median) and
+ prediction intervals. By default, the 80 and 90% prediction
+ intervals are included when using the MQLoss.
+
+- `y`: The actual value.
+
+## 4. Evaluate models and select the best for each series
+
+To evaluate the point forecasts of the models, we’ll use the Root Mean
+Squared Error (RMSE), defined as the square root of the mean of the
+squared differences between the actual and the predicted values.
+
+For convenience, we’ll use the `evaluate` and the
+[`rmse`](https://nixtlaverse.nixtla.io/neuralforecast/losses.numpy.html#rmse)
+functions from `utilsforecast`.
+
+
+```python
+from utilsforecast.evaluation import evaluate
+from utilsforecast.losses import rmse
+```
+
+The `evaluate` function takes the following arguments:
+
+- `df`: The data frame with the forecasts to evaluate.
+
+- `metrics` (list): The metrics to compute.
+
+- `models` (list): Names of the models to evaluate. Default is `None`,
+ which uses all columns after removing `id_col`, `time_col`, and
+ `target_col`.
+
+- `id_col` (str): Column that identifies unique ids of the series.
+ Default is `unique_id`.
+
+- `time_col` (str): Column with the timestamps or the temporal index.
+ Default is `ds`.
+
+- `target_col` (str): Column with the target variable. Default is `y`.
+
+Notice that if we use the default value of `models`, then we need to
+exclude the `cutoff` column from the cross-validation data frame.
+
+
+```python
+evaluation_df = evaluate(cv_df.drop(columns='cutoff'), metrics=[rmse])
+```
+
+For each unique id, we’ll select the model with the lowest RMSE.
+
+
+```python
+evaluation_df['best_model'] = evaluation_df.drop(columns=['metric', 'unique_id']).idxmin(axis=1)
+evaluation_df
+```
+
+| | unique_id | metric | MLP-median | NBEATS-median | NHITS-median | best_model |
+|-----|-----------|--------|-------------|---------------|--------------|---------------|
+| 0 | H1 | rmse | 46.654390 | 49.595304 | 47.651201 | MLP-median |
+| 1 | H10 | rmse | 24.192081 | 21.580142 | 16.887989 | NHITS-median |
+| 2 | H100 | rmse | 171.958998 | 178.820952 | 170.452623 | NHITS-median |
+| 3 | H101 | rmse | 331.270162 | 260.021871 | 169.453119 | NHITS-median |
+| 4 | H102 | rmse | 440.470939 | 362.602167 | 326.571391 | NHITS-median |
+| 5 | H103 | rmse | 9069.937603 | 9267.925257 | 8578.535681 | NHITS-median |
+| 6 | H104 | rmse | 189.534415 | 169.017976 | 226.442403 | NBEATS-median |
+| 7 | H105 | rmse | 341.029706 | 284.038751 | 262.140145 | NHITS-median |
+| 8 | H106 | rmse | 203.723728 | 328.128422 | 298.377068 | MLP-median |
+| 9 | H107 | rmse | 212.384943 | 161.445838 | 231.303421 | NBEATS-median |
+
+We can summarize the results to see how many times each model won.
+
+
+```python
+summary_df = evaluation_df.groupby(['metric', 'best_model']).size().sort_values().to_frame()
+summary_df = summary_df.reset_index()
+summary_df.columns = ['metric', 'model', 'num. of unique_ids']
+summary_df
+```
+
+| | metric | model | num. of unique_ids |
+|-----|--------|---------------|--------------------|
+| 0 | rmse | MLP-median | 2 |
+| 1 | rmse | NBEATS-median | 2 |
+| 2 | rmse | NHITS-median | 6 |
+
+With this information, we now know which model performs best for each
+series in the historical data.
+
+## 5. Plot cross-validation results
+
+To visualize the cross-validation results, we will use the `plot_series`
+method again. We’ll need to rename the `y` column in the
+cross-validation output to avoid duplicates with the original data
+frame. We’ll also exclude the `cutoff` column and use the
+`max_insample_length argument` to plot only the last 300 observations
+for better visualization.
+
+
+```python
+cv_df.rename(columns = {'y': 'actual'}, inplace=True) # rename actual values
+plot_series(Y_df, cv_df.drop(columns='cutoff'), max_insample_length=300)
+```
+
+
+
+To clarify the concept of cross-validation further, we’ll plot the
+forecasts generated at each cutoff for the series with `unique_id='H1'`.
+There are three cutoffs because we set `n_windows=3`. In this example,
+we used `refit=1`, so each model is retrained for each window using data
+with timestamps up to and including the respective cutoff. Additionally,
+since `step_size` is equal to the forecast horizon, the resulting
+forecasts are non-overlapping
+
+
+```python
+cutoff1, cutoff2, cutoff3 = cv_df['cutoff'].unique()
+for cutoff in cv_df['cutoff'].unique():
+ display(
+ plot_series(
+ Y_df,
+ cv_df[cv_df['cutoff'] == cutoff].drop(columns='cutoff'),
+ ids=['H1'], # use ids parameter to select specific series
+ )
+ )
+```
+
+
+
+
+
+
+
diff --git a/neuralforecast/docs/tutorials/distributed_neuralforecast.html.mdx b/neuralforecast/docs/tutorials/distributed_neuralforecast.html.mdx
new file mode 100644
index 00000000..b519d054
--- /dev/null
+++ b/neuralforecast/docs/tutorials/distributed_neuralforecast.html.mdx
@@ -0,0 +1,395 @@
+---
+output-file: distributed_neuralforecast.html
+title: Distributed Training
+---
+
+
+## Prerequisites
+
+This notebook was ran in databricks using the following configuration:
+
+- Databricks Runtime Version: 14.3 LTS ML (Spark 3.5, GPU, Scala 2.12)
+- Worker and executors instance type: g4dn.xlarge
+- Cluster libraries:
+ - neuralforecast==1.7.0
+ - fugue
+ - protobuf\<=3.20.1
+ - s3fs
+
+## Load libraries
+
+
+```python
+import logging
+
+import numpy as np
+import pandas as pd
+
+from neuralforecast import NeuralForecast, DistributedConfig
+from neuralforecast.auto import AutoNHITS
+from neuralforecast.models import NHITS, LSTM
+from utilsforecast.evaluation import evaluate
+from utilsforecast.losses import mae, rmse, smape
+from utilsforecast.plotting import plot_series
+```
+
+``` text
+2024-06-12 21:29:32.857491: I tensorflow/core/util/port.cc:111] oneDNN custom operations are on. You may see slightly different numerical results due to floating-point round-off errors from different computation orders. To turn them off, set the environment variable `TF_ENABLE_ONEDNN_OPTS=0`.
+2024-06-12 21:29:32.901906: E tensorflow/compiler/xla/stream_executor/cuda/cuda_dnn.cc:9342] Unable to register cuDNN factory: Attempting to register factory for plugin cuDNN when one has already been registered
+2024-06-12 21:29:32.901946: E tensorflow/compiler/xla/stream_executor/cuda/cuda_fft.cc:609] Unable to register cuFFT factory: Attempting to register factory for plugin cuFFT when one has already been registered
+2024-06-12 21:29:32.901973: E tensorflow/compiler/xla/stream_executor/cuda/cuda_blas.cc:1518] Unable to register cuBLAS factory: Attempting to register factory for plugin cuBLAS when one has already been registered
+2024-06-12 21:29:32.909956: I tensorflow/core/platform/cpu_feature_guard.cc:182] This TensorFlow binary is optimized to use available CPU instructions in performance-critical operations.
+To enable the following instructions: AVX2 AVX512F AVX512_VNNI FMA, in other operations, rebuild TensorFlow with the appropriate compiler flags.
+```
+
+
+```python
+logging.getLogger('pytorch_lightning').setLevel(logging.ERROR)
+```
+
+## Data
+
+
+```python
+df = pd.read_parquet('https://datasets-nixtla.s3.amazonaws.com/m4-hourly.parquet')
+df['exog_0'] = np.random.rand(df.shape[0])
+static = df.groupby('unique_id').head(1).copy()
+static['stat_0'] = static['unique_id'].astype('category').cat.codes
+static = static[['unique_id', 'stat_0']]
+valid = df.groupby('unique_id').tail(24)
+train = df.drop(valid.index)
+# save for loading in spark
+s3_prefix = 's3://nixtla-tmp/distributed'
+train.to_parquet(f'{s3_prefix}/train.parquet', index=False)
+valid.to_parquet(f'{s3_prefix}/valid.parquet', index=False)
+static.to_parquet(f'{s3_prefix}/static.parquet', index=False)
+# load in spark
+spark_train = spark.read.parquet(f'{s3_prefix}/train.parquet')
+spark_valid = spark.read.parquet(f'{s3_prefix}/valid.parquet')
+spark_static = spark.read.parquet(f'{s3_prefix}/static.parquet')
+```
+
+## Configuration
+
+
+```python
+# Configuration required for distributed training
+dist_cfg = DistributedConfig(
+ partitions_path=f'{s3_prefix}/partitions', # path where the partitions will be saved
+ num_nodes=2, # number of nodes to use during training (machines)
+ devices=1, # number of GPUs in each machine
+)
+
+# pytorch lightning configuration
+# the executors don't have permission to write on the filesystem, so we disable saving artifacts
+distributed_kwargs = dict(
+ accelerator='gpu',
+ enable_progress_bar=False,
+ logger=False,
+ enable_checkpointing=False,
+)
+
+# exogenous features
+exogs = {
+ 'futr_exog_list': ['exog_0'],
+ 'stat_exog_list': ['stat_0'],
+}
+
+# for the AutoNHITS
+def config(trial):
+ return dict(
+ input_size=48,
+ max_steps=2_000,
+ learning_rate=trial.suggest_float('learning_rate', 1e-4, 1e-1, log=True),
+ **exogs,
+ **distributed_kwargs
+ )
+```
+
+## Model training
+
+
+```python
+nf = NeuralForecast(
+ models=[
+ NHITS(h=24, input_size=48, max_steps=2_000, **exogs, **distributed_kwargs),
+ AutoNHITS(h=24, config=config, backend='optuna', num_samples=2, alias='tuned_nhits'),
+ LSTM(h=24, input_size=48, max_steps=2_000, **exogs, **distributed_kwargs),
+ ],
+ freq=1,
+)
+nf.fit(spark_train, static_df=spark_static, distributed_config=dist_cfg, val_size=24)
+```
+
+``` text
+[rank: 0] Seed set to 1
+/local_disk0/.ephemeral_nfs/cluster_libraries/python/lib/python3.10/site-packages/pytorch_lightning/utilities/parsing.py:199: Attribute 'loss' is an instance of `nn.Module` and is already saved during checkpointing. It is recommended to ignore them using `self.save_hyperparameters(ignore=['loss'])`.
+[rank: 0] Seed set to 1
+INFO:TorchDistributor:Started distributed training with 2 executor processes
+[rank: 1] Seed set to 1
+[rank: 0] Seed set to 1
+[rank: 1] Seed set to 1
+Initializing distributed: GLOBAL_RANK: 1, MEMBER: 2/2
+GPU available: True (cuda), used: True
+TPU available: False, using: 0 TPU cores
+IPU available: False, using: 0 IPUs
+HPU available: False, using: 0 HPUs
+[rank: 0] Seed set to 1
+Initializing distributed: GLOBAL_RANK: 0, MEMBER: 1/2
+----------------------------------------------------------------------------------------------------
+distributed_backend=nccl
+All distributed processes registered. Starting with 2 processes
+----------------------------------------------------------------------------------------------------
+
+LOCAL_RANK: 0 - CUDA_VISIBLE_DEVICES: [0]
+
+ | Name | Type | Params
+-----------------------------------------------
+0 | loss | MAE | 0
+1 | padder_train | ConstantPad1d | 0
+2 | scaler | TemporalNorm | 0
+3 | blocks | ModuleList | 2.6 M
+-----------------------------------------------
+2.6 M Trainable params
+0 Non-trainable params
+2.6 M Total params
+10.341 Total estimated model params size (MB)
+LOCAL_RANK: 0 - CUDA_VISIBLE_DEVICES: [0]
+`Trainer.fit` stopped: `max_steps=2000` reached.
+INFO:TorchDistributor:Finished distributed training with 2 executor processes
+[I 2024-06-12 21:31:09,627] A new study created in memory with name: no-name-849c3a84-28d7-417b-a48d-f0feac64cbc3
+[rank: 0] Seed set to 1
+INFO:TorchDistributor:Started distributed training with 2 executor processes
+[rank: 1] Seed set to 1
+[rank: 0] Seed set to 1
+[rank: 1] Seed set to 1
+Initializing distributed: GLOBAL_RANK: 1, MEMBER: 2/2
+GPU available: True (cuda), used: True
+TPU available: False, using: 0 TPU cores
+IPU available: False, using: 0 IPUs
+HPU available: False, using: 0 HPUs
+[rank: 0] Seed set to 1
+Initializing distributed: GLOBAL_RANK: 0, MEMBER: 1/2
+----------------------------------------------------------------------------------------------------
+distributed_backend=nccl
+All distributed processes registered. Starting with 2 processes
+----------------------------------------------------------------------------------------------------
+
+LOCAL_RANK: 0 - CUDA_VISIBLE_DEVICES: [0]
+LOCAL_RANK: 0 - CUDA_VISIBLE_DEVICES: [0]
+
+ | Name | Type | Params
+-----------------------------------------------
+0 | loss | MAE | 0
+1 | padder_train | ConstantPad1d | 0
+2 | scaler | TemporalNorm | 0
+3 | blocks | ModuleList | 2.6 M
+-----------------------------------------------
+2.6 M Trainable params
+0 Non-trainable params
+2.6 M Total params
+10.341 Total estimated model params size (MB)
+`Trainer.fit` stopped: `max_steps=2000` reached.
+INFO:TorchDistributor:Finished distributed training with 2 executor processes
+[I 2024-06-12 21:32:26,716] Trial 0 finished with value: 240.63693237304688 and parameters: {'learning_rate': 0.0008137359313625077}. Best is trial 0 with value: 240.63693237304688.
+[rank: 0] Seed set to 1
+INFO:TorchDistributor:Started distributed training with 2 executor processes
+[rank: 1] Seed set to 1
+[rank: 0] Seed set to 1
+[rank: 1] Seed set to 1
+Initializing distributed: GLOBAL_RANK: 1, MEMBER: 2/2
+GPU available: True (cuda), used: True
+TPU available: False, using: 0 TPU cores
+IPU available: False, using: 0 IPUs
+HPU available: False, using: 0 HPUs
+[rank: 0] Seed set to 1
+Initializing distributed: GLOBAL_RANK: 0, MEMBER: 1/2
+----------------------------------------------------------------------------------------------------
+distributed_backend=nccl
+All distributed processes registered. Starting with 2 processes
+----------------------------------------------------------------------------------------------------
+
+LOCAL_RANK: 0 - CUDA_VISIBLE_DEVICES: [0]
+
+ | Name | Type | Params
+-----------------------------------------------
+0 | loss | MAE | 0
+1 | padder_train | ConstantPad1d | 0
+2 | scaler | TemporalNorm | 0
+3 | blocks | ModuleList | 2.6 M
+-----------------------------------------------
+2.6 M Trainable params
+0 Non-trainable params
+2.6 M Total params
+10.341 Total estimated model params size (MB)
+LOCAL_RANK: 0 - CUDA_VISIBLE_DEVICES: [0]
+`Trainer.fit` stopped: `max_steps=2000` reached.
+INFO:TorchDistributor:Finished distributed training with 2 executor processes
+[I 2024-06-12 21:33:43,744] Trial 1 finished with value: 269.3470153808594 and parameters: {'learning_rate': 0.0007824692588634985}. Best is trial 0 with value: 240.63693237304688.
+[rank: 0] Seed set to 1
+INFO:TorchDistributor:Started distributed training with 2 executor processes
+[rank: 1] Seed set to 1
+[rank: 0] Seed set to 1
+[rank: 1] Seed set to 1
+Initializing distributed: GLOBAL_RANK: 1, MEMBER: 2/2
+GPU available: True (cuda), used: True
+TPU available: False, using: 0 TPU cores
+IPU available: False, using: 0 IPUs
+HPU available: False, using: 0 HPUs
+[rank: 0] Seed set to 1
+Initializing distributed: GLOBAL_RANK: 0, MEMBER: 1/2
+----------------------------------------------------------------------------------------------------
+distributed_backend=nccl
+All distributed processes registered. Starting with 2 processes
+----------------------------------------------------------------------------------------------------
+
+LOCAL_RANK: 0 - CUDA_VISIBLE_DEVICES: [0]
+LOCAL_RANK: 0 - CUDA_VISIBLE_DEVICES: [0]
+
+ | Name | Type | Params
+-----------------------------------------------
+0 | loss | MAE | 0
+1 | padder_train | ConstantPad1d | 0
+2 | scaler | TemporalNorm | 0
+3 | blocks | ModuleList | 2.6 M
+-----------------------------------------------
+2.6 M Trainable params
+0 Non-trainable params
+2.6 M Total params
+10.341 Total estimated model params size (MB)
+`Trainer.fit` stopped: `max_steps=2000` reached.
+INFO:TorchDistributor:Finished distributed training with 2 executor processes
+INFO:TorchDistributor:Started distributed training with 2 executor processes
+[rank: 0] Seed set to 1
+[rank: 1] Seed set to 1
+GPU available: True (cuda), used: True
+TPU available: False, using: 0 TPU cores
+IPU available: False, using: 0 IPUs
+HPU available: False, using: 0 HPUs
+[rank: 0] Seed set to 1
+Initializing distributed: GLOBAL_RANK: 0, MEMBER: 1/2
+----------------------------------------------------------------------------------------------------
+distributed_backend=nccl
+All distributed processes registered. Starting with 2 processes
+----------------------------------------------------------------------------------------------------
+
+[rank: 1] Seed set to 1
+Initializing distributed: GLOBAL_RANK: 1, MEMBER: 2/2
+LOCAL_RANK: 0 - CUDA_VISIBLE_DEVICES: [0]
+LOCAL_RANK: 0 - CUDA_VISIBLE_DEVICES: [0]
+
+ | Name | Type | Params
+--------------------------------------------------
+0 | loss | MAE | 0
+1 | padder | ConstantPad1d | 0
+2 | scaler | TemporalNorm | 0
+3 | hist_encoder | LSTM | 484 K
+4 | context_adapter | Linear | 54.0 K
+5 | mlp_decoder | MLP | 2.6 K
+--------------------------------------------------
+541 K Trainable params
+0 Non-trainable params
+541 K Total params
+2.166 Total estimated model params size (MB)
+`Trainer.fit` stopped: `max_steps=2000` reached.
+INFO:TorchDistributor:Finished distributed training with 2 executor processes
+```
+
+## Forecasting
+
+When we’re done training the model in a distributed way we can predict
+using the stored dataset. If we have future exogenous features we can
+provide a spark dataframe as `futr_df`. Note that if you want to load
+the stored dataset you need to provide the spark session through the
+`engine` argument.
+
+
+```python
+saved_ds_preds = nf.predict(futr_df=spark_valid.drop("y"), engine=spark).toPandas()
+```
+
+We can also provide a spark dataframe as `df` as well as `static_df` and
+`futr_df` (if applicable) to compute predictions on different data or
+after loading a saved model.
+
+
+```python
+new_df_preds = nf.predict(df=spark_train, static_df=spark_static, futr_df=spark_valid.drop("y")).toPandas()
+```
+
+Either of the above methods will yield the same results.
+
+
+```python
+pd.testing.assert_frame_equal(
+ saved_ds_preds.sort_values(['unique_id', 'ds']).reset_index(drop=True),
+ new_df_preds.sort_values(['unique_id', 'ds']).reset_index(drop=True),
+ atol=1e-3,
+)
+```
+
+## Saving for inference
+
+We can now persist the trained models
+
+
+```python
+save_path = f'{s3_prefix}/model-artifacts'
+nf.save(save_path, save_dataset=False, overwrite=True)
+```
+
+And load them back
+
+
+```python
+nf2 = NeuralForecast.load(save_path)
+```
+
+``` text
+[rank: 0] Seed set to 1
+[rank: 0] Seed set to 1
+[rank: 0] Seed set to 1
+```
+
+We can now use this object to compute forecasts. We can provide either
+local dataframes (pandas, polars) as well as spark dataframes
+
+
+```python
+preds = nf.predict(df=train, static_df=static, futr_df=valid.drop(columns='y'))
+preds2 = nf2.predict(df=train, static_df=static, futr_df=valid.drop(columns='y'))[preds.columns]
+pd.testing.assert_frame_equal(saved_ds_preds, preds)
+pd.testing.assert_frame_equal(preds, preds2)
+```
+
+## Evaluation
+
+
+```python
+(
+ evaluate(
+ preds.merge(valid.drop(columns='exog_0'), on=['unique_id', 'ds']),
+ metrics=[mae, rmse, smape],
+ )
+ .drop(columns='unique_id')
+ .groupby('metric')
+ .mean()
+)
+```
+
+| | NHITS | tuned_nhits | LSTM |
+|--------|------------|-------------|------------|
+| metric | | | |
+| mae | 417.075336 | 322.751522 | 270.423775 |
+| rmse | 485.304941 | 410.998659 | 330.579283 |
+| smape | 0.063995 | 0.066046 | 0.063975 |
+
+## Plotting a sample
+
+
+```python
+plot_series(train, preds)
+```
+
+
+
diff --git a/neuralforecast/docs/tutorials/explainability.mdx b/neuralforecast/docs/tutorials/explainability.mdx
new file mode 100644
index 00000000..5a543344
--- /dev/null
+++ b/neuralforecast/docs/tutorials/explainability.mdx
@@ -0,0 +1,635 @@
+---
+title: Explainability for Deep Learning Forecasting Models
+---
+
+In this detailed tutorial, we discover how to explain forecasts made
+with models from *neuralforecast*.
+
+Note that the functionality is currently in beta. It can only be applied
+on univariate models, but support for multivariate models is coming
+soon.
+
+## Prerequisites
+
+- We assume you have *neuralforecast* already installed.
+- Explanations are obtained with [Captum](https://captum.ai/): an
+ open-source library for model interpretability in PyTorch. Make sure
+ to install the package with `pip install captum` to use the features
+ demonstrated below.
+- You can optionally install
+ [SHAP](https://shap.readthedocs.io/en/latest/) to access their
+ visualizations capabilities. This can be done with
+ `pip install shap`.
+
+## Load libraries
+
+```python
+import numpy as np
+import pandas as pd
+import torch
+
+from neuralforecast.core import NeuralForecast
+from neuralforecast.models import NHITS
+from neuralforecast.losses.pytorch import MQLoss
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+```
+
+
+```python
+# Set random seeds for reproducibility
+np.random.seed(42);
+torch.manual_seed(42);
+```
+
+## Load the data
+
+We demonstrate the explainability capabilities with the AirPassengers
+dataset. This dataset has:
+- 2 unique series
+- a future exogenous variable (`trend`)
+- a historical exogenous variable (`y_lag[12]`)
+- static exogenous variable (`Airline1`)
+
+That way, we see that we can handle attributions for all types of
+exogenous features. For more information on the types of exogenous
+features, read [this
+tutorial](https://nixtlaverse.nixtla.io/neuralforecast/docs/capabilities/exogenous_variables.html).
+
+```python
+Y_train_df = AirPassengersPanel[AirPassengersPanel['ds'] < AirPassengersPanel['ds'].values[-12]].reset_index(drop=True)
+Y_test_df = AirPassengersPanel[AirPassengersPanel['ds'] >= AirPassengersPanel['ds'].values[-12]].reset_index(drop=True)
+futr_df = Y_test_df.drop(columns=["y", "y_[lag12]"])
+```
+
+## Basic usage
+
+### Train a model
+
+Before explaining forecasts, we need to train a forecasting model. Here,
+we use the NHITS model, but you can use any univariate model. For now,
+we don’t support multivariate models just yet, this feature will be
+implemented soon.
+
+```python
+models = [
+ NHITS(
+ h=12,
+ input_size=24,
+ hist_exog_list=["y_[lag12]"],
+ futr_exog_list=["trend"],
+ stat_exog_list=['airline1'],
+ max_steps=20,
+ scaler_type="robust",
+ ),
+]
+
+nf = NeuralForecast(
+ models=models,
+ freq="ME",
+)
+
+nf.fit(
+ df=Y_train_df,
+ static_df=AirPassengersStatic
+)
+```
+
+### Get features attributions
+
+Once the model is trained, we can get feature attributions using the `nf.explain` method.
+
+This method takes the following parameters:
+- `horizons`: List of horizons to explain. If None, all horizons are explained. Defaults to None.
+- `outputs`: List of outputs to explain for models with multiple outputs. Defaults to \[0\] (first output). This is useful when we have models trained with a probabilistic loss. We will explore that later in the tutorial.
+- `explainer`: Name of the explainer to use. Options are ‘IntegratedGradients’, ‘ShapleyValueSampling’, ‘InputXGradient’. Defaults to ‘IntegratedGradients’.
+- `df` (pandas, polars or spark DataFrame): DataFrame with columns \[`unique_id`, `ds`, `y`\] and exogenous variables. If a DataFrame is passed, it is used to generate forecasts. Defaults to None.
+- `static_df` (pandas, polars or spark DataFrame): DataFrame with columns \[`unique_id`\] and static exogenous. Defaults to None. Only use it if you trained your model with static exogenous features.
+- `futr_df` (pandas, polars or spark DataFrame): DataFrame with \[`unique_id`, `ds`\] columns and `df`’s future exogenous. Defaults to None. Only use it if you trained your model with future exogenous features.
+- `verbose`: Print warnings. Defaults to True.
+- `engine`: Distributed engine for inference. Only used if df is a spark dataframe or if fit was called on a spark dataframe.
+- `level`: Confidence levels between 0 and 100. Defaults to None.
+- `quantiles`: Alternative to level, target quantiles to predict. Defaults to None.
+- `data_kwargs`: Extra arguments to be passed to the dataset within each model.
+
+Note that parameters from `df` and onwards act exactly the same way as
+in the `nf.predict()` method.
+
+In this case, let’s explain each horizon step, so we keep
+`horizons=None`. Since our model used a point loss, there is only one
+output, so we also keep the default value `outputs=[0]`. Finally, we
+choose the “IntegratedGradients” explainer, as it is one of the fastest
+method for interpretability in deep learning.
+
+```python
+preds_df, explanations = nf.explain(
+ static_df=AirPassengersStatic,
+ futr_df=futr_df,
+ explainer="IntegratedGradients"
+)
+```
+
+We can see that `nf.explain()` returns two values:
+
+1. A dataframe with the forecasts from the fitted models
+2. A dictionary with the feature attributions for each model
+
+Thus, you can access the attribution score of each features used for
+training the NHITS model by accessing `explanations["NHITS"]`. Note that
+if you used an alias when initializing the model, then the key is the
+value of the alias.
+
+```python
+explanations["NHITS"].keys()
+```
+
+``` text
+dict_keys(['insample', 'futr_exog', 'hist_exog', 'stat_exog', 'baseline_predictions'])
+```
+
+From above, we can see that we have stored the attributions for each
+feature type as well as the baseline predictions.
+- `insample` contains
+the attributions for past lags and availability mask
+- `futr_exog`
+contains the attributions for future exogenous features
+- `hist_exog`
+contains the attributions for historical exogenous features
+- `stat_exog` contains the attributions for static exogenous features
+- `baseline_predictions` contains the baseline prediction of the model if
+none of the features above were available. Note that if the selected
+explainer does not have the additivity property, the value will be set
+to None.
+
+We will touch upon the additivity property in a later section. For now,
+just know that `IntegratedGradients` has the additive property, meaning
+that taking the sum of baseline predictions and feature attributions
+results in the final forecast made by the model.
+
+Now, because we are using Captum, we work directly with tensors, keeping
+the entire process fast, efficient, and allowing us to leverage GPUs
+when available. As such, the attributions are also stored as tensors as
+shown below.
+
+```python
+for key in list(explanations["NHITS"].keys()):
+ print(f"Shape of {key}: {explanations['NHITS'][key].shape}")
+```
+
+``` text
+Shape of insample: torch.Size([2, 12, 1, 1, 24, 2])
+Shape of futr_exog: torch.Size([2, 12, 1, 1, 36, 1])
+Shape of hist_exog: torch.Size([2, 12, 1, 1, 24, 1])
+Shape of stat_exog: torch.Size([2, 12, 1, 1, 1])
+Shape of baseline_predictions: torch.Size([2, 12, 1, 1])
+```
+
+For each element above, the shape is defined as:
+
+- `insample`: \[batch_size, horizon, n_series, n_output, input_size, 2
+ (y attribution, mask attribution)\]
+- `futr_exog`: \[batch_size, horizon, n_series, n_output,
+ input_size+horizon, n_futr_features\]
+- `hist_exog`: \[batch_size, horizon, n_series, n_output, input_size,
+ n_hist_features\]
+- `stat_exog`: \[batch_size, horizon, n_series, n_output,
+ n_static_features\]
+- `baseline_predictions`: \[batch_size, horizon, n_series, n_output\]
+
+Here, `batch_size` is 2 for all, because we are explaining two different
+series. `n_series` however is 1 because NHITS is a univariate model.
+Also note that for `insample`, the last shape is always 2, because we
+score the attribution of the values of the past lags and their
+availability.
+
+At this point, we have all the information needed to analyze the
+attribution scores and make visualizations.
+
+### Plotting feature attributions
+
+You can now use any method you want to plot feature attributions. You
+can make plots manually using any visualization library like
+`matplotlib` or `seaborn`, but `shap` has dedicated plots for
+explainability, so let’s see how we can use them.
+
+Basically, with the information we have, we can easily create a
+`shap.Explanation` object that can then be used to create different
+plots from the `shap` package.
+
+Specifically, a `shap.Explanation` object needs:
+- `values`: the attribution scores
+- `base_values`: the baseline predictions of the model
+- `feature_names`: a list to display nice feature names
+
+Here, let’s create a waterfall plot to visualize the attributions of
+each features, for the first series (Airline1), and for the first step
+in the horizon.
+
+```python
+import shap
+```
+
+
+```python
+batch_idx = 0 # Attributions for the first series (Airline1)
+horizon_idx = 0 # Attributions for the first horizon step
+output_idx = 0
+
+attributions = []
+feature_names = []
+
+# Insample attributions
+y_attr = explanations["NHITS"]["insample"][batch_idx, horizon_idx, 0, output_idx, :, 0]
+mask_attr = explanations["NHITS"]["insample"][batch_idx, horizon_idx, 0, output_idx, :, 1]
+combined_insample = (y_attr + mask_attr).cpu().numpy()
+for i, attr in enumerate(combined_insample):
+ attributions.append(attr)
+ feature_names.append(f"y_lag{i+1}")
+
+# hist_exog attributions
+hist_attr = explanations["NHITS"]["hist_exog"][batch_idx, horizon_idx, 0, output_idx]
+hist_attr = hist_attr.cpu().numpy()
+for t in range(hist_attr.shape[0]):
+ attributions.append(hist_attr[t, 0])
+ feature_names.append(f"y_lag12_t{t+1}")
+
+# futr_exog attributions
+futr_attr = explanations["NHITS"]["futr_exog"][batch_idx, horizon_idx, 0, output_idx]
+futr_attr = futr_attr.cpu().numpy()
+for t in range(futr_attr.shape[0]):
+ attributions.append(futr_attr[t, 0])
+ if t < 24:
+ feature_names.append(f"trend_hist_t{t+1}") # Known values in the past
+ else:
+ feature_names.append(f"trend_futr_h{t-23}") # Knwon values in the future
+
+# stat_exog attributions
+stat_attr = explanations["NHITS"]["stat_exog"][batch_idx, horizon_idx, 0, output_idx]
+attributions.append(float(stat_attr.cpu()))
+feature_names.append("airline1")
+
+shap_values = np.array(attributions)
+
+# baseline_predictions
+baseline = float(explanations["NHITS"]["baseline_predictions"][batch_idx, horizon_idx, 0, output_idx].cpu())
+
+# Create SHAP Explanation
+shap_explanation = shap.Explanation(
+ values=shap_values,
+ base_values=baseline,
+ feature_names=feature_names
+)
+
+shap.plots.waterfall(shap_explanation)
+```
+
+
+
+As you can see, we now have a nice waterfall plot showing the baseline
+prediction, E\[f(X)\] = 396.163, and how each features contributes to
+the final forecast f(x) = 418.631.
+
+Of course, we can do a wide variery of different plots from `shap`. For
+example, we can do a simple bar plot as shown below.
+
+```python
+shap.plots.bar(shap_explanation)
+```
+
+
+
+In both figures above, we have the breakdown of each feature at each
+timestep. This can make the plots crowded or it can be a level of
+granularity that is not necessary for analysis. So, you can also decide
+to combine all time steps together for a cleaner plot.
+
+```python
+batch_idx = 0
+horizon_idx = 0
+output_idx = 0
+
+baseline = float(explanations["NHITS"]["baseline_predictions"][batch_idx, horizon_idx, output_idx, output_idx].cpu())
+insample_sum = float(explanations["NHITS"]["insample"][batch_idx, horizon_idx, output_idx, output_idx, :, :].sum().cpu())
+
+futr_exog_sum = 0
+futr_exog_sum = float(explanations["NHITS"]["futr_exog"][batch_idx, horizon_idx, output_idx, output_idx, :, :].sum().cpu())
+
+hist_exog_sum = 0
+hist_exog_sum = float(explanations["NHITS"]["hist_exog"][batch_idx, horizon_idx, output_idx, output_idx, :, :].sum().cpu())
+
+stat_exog_sum = 0
+stat_exog_sum = float(explanations["NHITS"]["stat_exog"][batch_idx, horizon_idx, output_idx, output_idx, :].sum().cpu())
+
+feature_names = []
+shap_values = []
+
+if insample_sum != 0:
+ feature_names.append("Historical Y (all lags)")
+ shap_values.append(insample_sum)
+
+if hist_exog_sum != 0:
+ feature_names.append("Historical Exog (y_lag12)")
+ shap_values.append(hist_exog_sum)
+
+if futr_exog_sum != 0:
+ feature_names.append("Future Exog (trend)")
+ shap_values.append(futr_exog_sum)
+
+if stat_exog_sum != 0:
+ feature_names.append("Static (airline1)")
+ shap_values.append(stat_exog_sum)
+
+shap_values = np.array(shap_values)
+
+# Create SHAP Explanation
+shap_explanation = shap.Explanation(
+ values=shap_values,
+ base_values=baseline,
+ feature_names=feature_names
+)
+
+shap.plots.waterfall(shap_explanation)
+```
+
+
+
+As you can see from the plot above, we have combined all inputs of each
+type of feature into a single category, so we can see how each overall
+feature contributes to the final forecast.
+
+### Verifying additivity
+
+As mentioned above, “IntegratedGradients” has the additive property,
+meaning that when we sum the baseline predictions with the total
+attribution scores of each features, we get the final forecasts made by
+the model.
+
+```python
+attribution_predictions = []
+
+# Process each series
+for batch_idx in range(2): # 2 series
+ # Process each horizon for this series
+ for horizon_idx in range(12): # horizon = 12
+ # Get baseline
+ baseline = float(explanations["NHITS"]["baseline_predictions"][batch_idx, horizon_idx, 0, 0].cpu())
+
+ # Sum all attribution components
+ total_attr = 0
+
+ # Insample (y + mask)
+ insample_attr = explanations["NHITS"]["insample"][batch_idx, horizon_idx, 0, 0, :, :].sum()
+ total_attr += float(insample_attr.cpu())
+
+ # Historical exogenous
+ if explanations["NHITS"]["hist_exog"] is not None:
+ hist_attr = explanations["NHITS"]["hist_exog"][batch_idx, horizon_idx, 0, 0, :, :].sum()
+ total_attr += float(hist_attr.cpu())
+
+ # Future exogenous
+ if explanations["NHITS"]["futr_exog"] is not None:
+ futr_attr = explanations["NHITS"]["futr_exog"][batch_idx, horizon_idx, 0, 0, :, :].sum()
+ total_attr += float(futr_attr.cpu())
+
+ # Static exogenous
+ if explanations["NHITS"]["stat_exog"] is not None:
+ stat_attr = explanations["NHITS"]["stat_exog"][batch_idx, horizon_idx, 0, 0, :].sum()
+ total_attr += float(stat_attr.cpu())
+
+ # Compute final prediction from attributions
+ pred_from_attr = baseline + total_attr
+ attribution_predictions.append(pred_from_attr)
+
+# Add as new column to preds_df
+preds_df['NHITS_attribution'] = attribution_predictions
+```
+
+
+```python
+np.testing.assert_allclose(
+ preds_df['NHITS'].values,
+ preds_df['NHITS_attribution'].values,
+ rtol=1e-3,
+ err_msg="Attribution predictions do not match model predictions"
+)
+```
+
+From the code above, we can see that reconstructed forecasts from the
+baseline predictions and attributions are within 0.1% of the original
+forecasts, so additivity is verified.
+
+## Advanced concepts
+
+### Choosing an explainer
+
+In this section, we outline the different explainers supported in
+*neuralforecast*. Different algorithms will produce different
+attribution scores, and so we must choose which applies best to our
+scenario.
+
+| Explainer | Local/Global | Additivity Property | Speed |
+|------------------------|--------------|---------------------|-----------|
+| Integrated Gradients | Local | Yes | Fast |
+| Shapley Value Sampling | Local | Yes | Very slow |
+| Input X Gradient | Local | No | Very fast |
+
+**Notes:**
+- **Local/Global**: All explainers are local, because they only explain how a specific input affects a specific forecast.
+- **Additivity Property**: Whether the sum of the feature attributions and baseline predictions result in the final forecast.
+- **Speed**:
+ - Very fast: Single gradient computation
+ - Fast: Multiple gradient computations (Integrated Gradients)
+ - Medium: Multiple model evaluations
+ - Slow: Many model evaluations for sampling-based methods
+ - Very Slow: Exponential complexity in worst case (exact Shapley values)
+
+#### Integrated Gradients
+
+Integrated Gradients computes attributions by integrating gradients
+along the straight-line path from a chosen baseline input (e.g., black
+image, zero embedding) to the actual input. The method calculates the
+path integral, which is approximated using a Riemann sum with typically
+20-300 gradient computations. Learn more in the [original
+paper](https://arxiv.org/pdf/1703.01365).
+
+**Advantages**
+- Theoretically grounded: Satisfies the axioms of
+sensitivity (features that affect the output get non-zero attribution)
+and implementation invariance (functionally equivalent networks produce
+identical attributions)
+- Has the additivity property
+
+**Disadvantages**
+- Relies on choosing an appropriate baseline that
+represents “absence of signal”. By default, we use as input only 0
+values.
+
+#### Shapley Value Sampling
+
+Shapley Value Sampling approximates Shapley values using Monte Carlo
+sampling of feature permutations. The method randomly samples different
+orderings of features and computes how much each feature contributes by
+comparing model predictions when that feature is included versus
+excluded from the subset. The approach simulates “missing” features by
+drawing random values from the training data distribution. Learn more in
+the [original
+paper](https://www.sciencedirect.com/science/article/abs/pii/S0305054808000804).
+
+**Advantages**
+- All subsets of input features are perturbed, so interactions and redundancies between features are taken into account
+- Uses simple permutation sampling that is easy to understand
+
+**Disadvantages**
+- High computational cost: requires many model evaluations (typically hundreds to thousands) to achieve reasonable approximation accuracy
+- Very slow due to the high number of model evaluations
+- Simulates missing features by sampling from marginal distributions, which may create unrealistic data instances when features are correlated
+
+#### Input X Gradient
+
+Input X Gradient computes feature attribution by simply multiplying each
+input value by the gradient of the model output with respect to that
+input. This corresponds to a first-order Taylor approximation of how the
+output would change if the input were set to zero. This means each time
+step’s input values are multiplied by the gradients to show which
+historical observations most influence the prediction. Learn more in the
+[original paper](https://arxiv.org/pdf/1605.01713).
+
+**Advantages**
+- Computational efficiency: it requires only a single pass through the model
+- No approximations as it uses the gradient of the model
+
+**Disadvantages**
+- No additivity
+- A bit problematic with the ReLu function, because their gradient can be 0, but it can still carry some information
+- Functions like tanh or sigmoid can have very low gradients, even though the input is significant, so it’s problematic for LSTM and GRU models.
+
+### Explaining models with different loss functions
+
+Currently, explanations are supported for models trained with:
+- Point loss functions (MAE, MSE, RMSE, etc.)
+- Non-parametric probabilistic losses (IQLoss, MQLoss, etc.)
+
+We do not support yet explaining models trained with parametric loss functions, like Normal, Student’s T, etc.
+
+For more information on the different loss functions supported in
+*neuralforecast*, read
+[here](https://nixtlaverse.nixtla.io/neuralforecast/docs/capabilities/objectives.html).
+
+#### Explaning a model with a probablistic loss function
+
+If you are explaining a model with a non-parametric loss function, then
+by default, we only explain the median forecast. This is controlled by
+the `ouputs` parameter. Let’s see an example.
+
+```python
+# Initialize model
+models = [
+ NHITS(
+ h=12,
+ input_size=24,
+ hist_exog_list=["y_[lag12]"],
+ futr_exog_list=["trend"],
+ stat_exog_list=['airline1'],
+ loss=MQLoss(level=[80]),
+ max_steps=20,
+ scaler_type="robust",
+ ),
+]
+
+nf = NeuralForecast(
+ models=models,
+ freq="ME",
+)
+
+# Fit model
+nf.fit(
+ df=Y_train_df,
+ static_df=AirPassengersStatic
+)
+
+# Get predictions and attributions
+preds_df, explanations = nf.explain(
+ outputs=[0], # Explain only the median forecast
+ static_df=AirPassengersStatic,
+ futr_df=futr_df,
+ explainer="IntegratedGradients"
+)
+```
+
+Above, by specifying `outputs=[0]`, which is the default value, we only
+explain the median forecast. However, we can explain up to three
+ouputs:
+1. Median forecast
+2. Lower bound
+3. Upper bound
+
+As such, to explain all outputs, we must set `ouputs=[0,1,2]`.
+
+```python
+preds_df, explanations = nf.explain(
+ outputs=[0, 1, 2], # Explain all outputs
+ static_df=AirPassengersStatic,
+ futr_df=futr_df,
+ explainer="IntegratedGradients"
+)
+```
+
+
+```python
+for key in list(explanations["NHITS"].keys()):
+ print(f"Shape of {key}: {explanations['NHITS'][key].shape}")
+```
+
+``` text
+Shape of insample: torch.Size([2, 12, 1, 3, 24, 2])
+Shape of futr_exog: torch.Size([2, 12, 1, 3, 36, 1])
+Shape of hist_exog: torch.Size([2, 12, 1, 3, 24, 1])
+Shape of stat_exog: torch.Size([2, 12, 1, 3, 1])
+Shape of baseline_predictions: torch.Size([2, 12, 1, 3])
+```
+
+As you can see, the fourth dimension, which represents the number of
+ouputs, is now equal to 3, because we are explaining the median, the
+lower bound and upper bound.
+
+### Explaining models with a scaler (`local_scaler_type`)
+
+If you specify a `local_scaler_type` in your `NeuralForecast` object,
+note that the attribution scores will be scaled. This is because the
+data is scaled before the training process. The relative importance is
+still relevant, but note that additivity will not hold.
+
+If additivtiy is important, then you must use `scaler_type` when
+initializing the model, as we do in this tutorial. This scales each
+window of data during training, so we can easily inverse transform the
+attribution scores.
+
+Again, no matter which approach you choose, the relative attribution
+scores are still valid and comparable. It’s only additivity that is
+impacted. If you specify a `local_scaler_type`, then a warning is issued
+about additivity.
+
+### Explaining recurrent models
+
+You can explain recurrent models (LSTM, GRU). Just note that if you set
+`recurrent=True`, then the Integrated Gradients explainer is not
+supported. If `recurrent=False`, you can use any explainer.
+
+## References
+
+- M. Sundararajan, A. Taly, and Q. Yan, “Axiomatic Attribution for Deep
+Networks.” Available: https://arxiv.org/pdf/1703.01365
+
+- S. Lundberg, P. Allen, and S.-I. Lee, “A Unified Approach to
+Interpreting Model Predictions,” Nov. 2017. Available:
+https://arxiv.org/pdf/1705.07874
+
+- J. Castro, D. Gómez, and J. Tejada, “Polynomial calculation of the
+Shapley value based on sampling,” Computers & Operations Research,
+vol. 36, no. 5, pp. 1726–1730, May 2009, doi:
+https://doi.org/10.1016/j.cor.2008.04.004.
+
+- A. Shrikumar, P. Greenside, A. Shcherbina, and A. Kundaje, “Not Just a
+Black Box: Learning Important Features Through Propagating Activation
+Differences,” arXiv:1605.01713 \[cs\], Apr. 2017, Available:
+https://arxiv.org/abs/1605.01713
+
diff --git a/neuralforecast/docs/tutorials/forecasting_tft.html.mdx b/neuralforecast/docs/tutorials/forecasting_tft.html.mdx
new file mode 100644
index 00000000..b15c7ce4
--- /dev/null
+++ b/neuralforecast/docs/tutorials/forecasting_tft.html.mdx
@@ -0,0 +1,310 @@
+---
+output-file: forecasting_tft.html
+title: 'Forecasting with TFT: Temporal Fusion Transformer'
+---
+
+
+Temporal Fusion Transformer (TFT) proposed by Lim et al. \[1\] is one of
+the most popular transformer-based model for time-series forecasting. In
+summary, TFT combines gating layers, an LSTM recurrent encoder, with
+multi-head attention layers for a multi-step forecasting strategy
+decoder. For more details on the Nixtla’s TFT implementation visit [this
+link](https://nixtla.github.io/neuralforecast/models.tft.html).
+
+In this notebook we show how to train the TFT model on the Texas
+electricity market load data (ERCOT). Accurately forecasting electricity
+markets is of great interest, as it is useful for planning distribution
+and consumption.
+
+We will show you how to load the data, train the TFT performing
+automatic hyperparameter tuning, and produce forecasts. Then, we will
+show you how to perform multiple historical forecasts for cross
+validation.
+
+You can run these experiments using GPU with Google Colab.
+
+
+
+## 1. Libraries
+
+
+```python
+!pip install neuralforecast
+```
+
+
+```python
+import pandas as pd
+```
+
+## 2. Load ERCOT Data
+
+The input to NeuralForecast is always a data frame in [long
+format](https://www.theanalysisfactor.com/wide-and-long-data/) with
+three columns: `unique_id`, `ds` and `y`:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp or int) column should be either an integer
+ indexing time or a datestamp ideally like YYYY-MM-DD for a date or
+ YYYY-MM-DD HH:MM:SS for a timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast. We
+ will rename the
+
+First, read the 2022 historic total demand of the ERCOT market. We
+processed the original data (available
+[here](https://www.ercot.com/gridinfo/load/load_hist)), by adding the
+missing hour due to daylight saving time, parsing the date to datetime
+format, and filtering columns of interest.
+
+
+```python
+Y_df = pd.read_csv('https://datasets-nixtla.s3.amazonaws.com/ERCOT-clean.csv')
+Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+Y_df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|---------------------|--------------|
+| 0 | ERCOT | 2021-01-01 00:00:00 | 43719.849616 |
+| 1 | ERCOT | 2021-01-01 01:00:00 | 43321.050347 |
+| 2 | ERCOT | 2021-01-01 02:00:00 | 43063.067063 |
+| 3 | ERCOT | 2021-01-01 03:00:00 | 43090.059203 |
+| 4 | ERCOT | 2021-01-01 04:00:00 | 43486.590073 |
+
+## 3. Model training and forecast
+
+First, instantiate the
+[`AutoTFT`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotft)
+model. The
+[`AutoTFT`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotft)
+class will automatically perform hyperparamter tunning using [Tune
+library](https://docs.ray.io/en/latest/tune/index.html), exploring a
+user-defined or default search space. Models are selected based on the
+error on a validation set and the best model is then stored and used
+during inference.
+
+To instantiate
+[`AutoTFT`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotft)
+you need to define:
+
+- `h`: forecasting horizon
+- `loss`: training loss
+- `config`: hyperparameter search space. If `None`, the
+ [`AutoTFT`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotft)
+ class will use a pre-defined suggested hyperparameter space.
+- `num_samples`: number of configurations explored.
+
+
+```python
+from ray import tune
+
+from neuralforecast.auto import AutoTFT
+from neuralforecast.core import NeuralForecast
+from neuralforecast.losses.pytorch import MAE
+
+import logging
+logging.getLogger("pytorch_lightning").setLevel(logging.WARNING)
+```
+
+> **Tip**
+>
+> Increase the `num_samples` parameter to explore a wider set of
+> configurations for the selected models. As a rule of thumb choose it
+> to be bigger than `15`.
+>
+> With `num_samples=3` this example should run in around 20 minutes.
+
+
+```python
+horizon = 24
+models = [AutoTFT(h=horizon,
+ loss=MAE(),
+ config=None,
+ num_samples=3)]
+```
+
+> **Tip**
+>
+> All our models can be used for both point and probabilistic
+> forecasting. For producing probabilistic outputs, simply modify the
+> loss to one of our
+> [`DistributionLoss`](https://nixtlaverse.nixtla.io/neuralforecast/losses.pytorch.html#distributionloss).
+> The complete list of losses is available in [this
+> link](https://nixtla.github.io/neuralforecast/losses.pytorch.html)
+
+> **Important**
+>
+> TFT is a very large model and can require a lot of memory! If you are
+> running out of GPU memory, try declaring your config search space and
+> decrease the `hidden_size`, `n_heads`, and `windows_batch_size`
+> parameters.
+>
+> This are all the parameters of the config:
+>
+>
+> ```python
+> config = {
+> "input_size": tune.choice([horizon]),
+> "hidden_size": tune.choice([32]),
+> "n_head": tune.choice([2]),
+> "learning_rate": tune.loguniform(1e-4, 1e-1),
+> "scaler_type": tune.choice(['robust', 'standard']),
+> "max_steps": tune.choice([500, 1000]),
+> "windows_batch_size": tune.choice([32]),
+> "check_val_every_n_epoch": tune.choice([100]),
+> "random_seed": tune.randint(1, 20),
+> }
+> ```
+
+The
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+class has built-in methods to simplify the forecasting pipelines, such
+as `fit`, `predit`, and `cross_validation`. Instantiate a
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+object with the following required parameters:
+
+- `models`: a list of models.
+
+- `freq`: a string indicating the frequency of the data. (See [panda’s
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+Then, use the `fit` method to train the
+[`AutoTFT`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotft)
+model on the ERCOT data. The total training time will depend on the
+hardware and the explored configurations, it should take between 10 and
+30 minutes.
+
+
+```python
+nf = NeuralForecast(
+ models=models,
+ freq='h')
+
+nf.fit(df=Y_df)
+```
+
+Finally, use the `predict` method to forecast the next 24 hours after
+the training data and plot the forecasts.
+
+
+```python
+Y_hat_df = nf.predict()
+Y_hat_df.head()
+```
+
+``` text
+c:\Users\ospra\miniconda3\envs\neuralforecast\lib\site-packages\utilsforecast\processing.py:384: FutureWarning: 'H' is deprecated and will be removed in a future version, please use 'h' instead.
+ freq = pd.tseries.frequencies.to_offset(freq)
+c:\Users\ospra\miniconda3\envs\neuralforecast\lib\site-packages\utilsforecast\processing.py:440: FutureWarning: 'H' is deprecated and will be removed in a future version, please use 'h' instead.
+ freq = pd.tseries.frequencies.to_offset(freq)
+```
+
+``` text
+Predicting: | | 0/? [00:00 **Prerequesites**
+>
+> This Guide assumes basic familiarity with NeuralForecast. For a
+> minimal example visit the [Quick
+> Start](../getting-started/quickstart.html)
+
+Follow this article for a step to step guide on building a
+production-ready forecasting pipeline for multiple time series.
+
+During this guide you will gain familiarity with the core
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)class
+and some relevant methods like
+[`NeuralForecast.fit`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast.fit),
+[`NeuralForecast.predict`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast.predict),
+and `StatsForecast.cross_validation.`
+
+We will use a classical benchmarking dataset from the M4 competition.
+The dataset includes time series from different domains like finance,
+economy and sales. In this example, we will use a subset of the Hourly
+dataset.
+
+We will model each time series globally Therefore, you will train a set
+of models for the whole dataset, and then select the best model for each
+individual time series. NeuralForecast focuses on speed, simplicity, and
+scalability, which makes it ideal for this task.
+
+**Outline:**
+
+1. Install packages.
+2. Read the data.
+3. Explore the data.
+4. Train many models globally for the entire dataset.
+5. Evaluate the model’s performance using cross-validation.
+6. Select the best model for every unique time series.
+
+> **Not Covered in this guide**
+>
+> - Using external regressors or exogenous variables
+> - Follow this tutorial to [include exogenous
+> variables](../capabilities/exogenous_variables.html) like
+> weather or holidays or static variables like category or
+> family.
+> - Probabilistic forecasting
+> - Follow this tutorial to [generate probabilistic
+> forecasts](../tutorials/uncertainty_quantification.html)
+> - Transfer Learning
+> - Train a model and use it to forecast on different data using
+> [this tutorial](../tutorials/transfer_learning.html)
+
+> **Tip**
+>
+> You can use Colab to run this Notebook interactively
+>
+
+> **Warning**
+>
+> To reduce the computation time, it is recommended to use GPU. Using
+> Colab, do not forget to activate it. Just go to
+> `Runtime>Change runtime type` and select GPU as hardware accelerator.
+
+## 1. Install libraries
+
+We assume you have
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+already installed. Check this guide for instructions on [how to install
+NeuralForecast](../getting-started/installation.html).
+
+
+```python
+! pip install neuralforecast
+```
+
+## 2. Read the data
+
+We will use pandas to read the M4 Hourly data set stored in a parquet
+file for efficiency. You can use ordinary pandas operations to read your
+data in other formats likes `.csv`.
+
+The input to
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+is always a data frame in [long
+format](https://www.theanalysisfactor.com/wide-and-long-data/) with
+three columns: `unique_id`, `ds` and `y`:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp or int) column should be either an integer
+ indexing time or a datestampe ideally like YYYY-MM-DD for a date or
+ YYYY-MM-DD HH:MM:SS for a timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast. We
+ will rename the
+
+This data set already satisfies the requirement.
+
+Depending on your internet connection, this step should take around 10
+seconds.
+
+
+```python
+import pandas as pd
+```
+
+
+```python
+Y_df = pd.read_parquet('https://datasets-nixtla.s3.amazonaws.com/m4-hourly.parquet')
+Y_df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|-----|-------|
+| 0 | H1 | 1 | 605.0 |
+| 1 | H1 | 2 | 586.0 |
+| 2 | H1 | 3 | 586.0 |
+| 3 | H1 | 4 | 559.0 |
+| 4 | H1 | 5 | 511.0 |
+
+This dataset contains 414 unique series with 900 observations on
+average. For this example and reproducibility’s sake, we will select
+only 10 unique IDs. Depending on your processing infrastructure feel
+free to select more or less series.
+
+> **Note**
+>
+> Processing time is dependent on the available computing resources.
+> Running this example with the complete dataset takes around 10 minutes
+> in a c5d.24xlarge (96 cores) instance from AWS.
+
+
+```python
+uids = Y_df['unique_id'].unique()[:10] # Select 10 ids to make the example faster
+Y_df = Y_df.query('unique_id in @uids').reset_index(drop=True)
+```
+
+## 3. Explore Data with the plot_series function
+
+Plot some series using the `plot_series` function from the
+`utilsforecast` library. This method prints 8 random series from the
+dataset and is useful for basic EDA.
+
+> **Note**
+>
+> The `plot_series` function uses matplotlib as a default engine. You
+> can change to plotly by setting `engine="plotly"`.
+
+
+```python
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+plot_series(Y_df)
+```
+
+
+
+## 4. Train multiple models for many series
+
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+can train many models on many time series globally and efficiently.
+
+
+```python
+import logging
+
+import optuna
+import ray.tune as tune
+import torch
+
+from neuralforecast import NeuralForecast
+from neuralforecast.auto import AutoNHITS, AutoLSTM
+from neuralforecast.losses.pytorch import MQLoss
+```
+
+
+```python
+optuna.logging.set_verbosity(optuna.logging.WARNING)
+logging.getLogger('pytorch_lightning').setLevel(logging.ERROR)
+torch.set_float32_matmul_precision('high')
+```
+
+Each `Auto` model contains a default search space that was extensively
+tested on multiple large-scale datasets. Additionally, users can define
+specific search spaces tailored for particular datasets and tasks.
+
+First, we create a custom search space for the
+[`AutoNHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autonhits)
+and
+[`AutoLSTM`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autolstm)
+models. Search spaces are specified with dictionaries, where keys
+corresponds to the model’s hyperparameter and the value is a `Tune`
+function to specify how the hyperparameter will be sampled. For example,
+use `randint` to sample integers uniformly, and `choice` to sample
+values of a list.
+
+
+```python
+def config_nhits(trial):
+ return {
+ "input_size": trial.suggest_categorical( # Length of input window
+ "input_size", (48, 48*2, 48*3)
+ ),
+ "start_padding_enabled": True,
+ "n_blocks": 5 * [1], # Length of input window
+ "mlp_units": 5 * [[64, 64]], # Length of input window
+ "n_pool_kernel_size": trial.suggest_categorical( # MaxPooling Kernel size
+ "n_pool_kernel_size",
+ (5*[1], 5*[2], 5*[4], [8, 4, 2, 1, 1])
+ ),
+ "n_freq_downsample": trial.suggest_categorical( # Interpolation expressivity ratios
+ "n_freq_downsample",
+ ([8, 4, 2, 1, 1], [1, 1, 1, 1, 1])
+ ),
+ "learning_rate": trial.suggest_float( # Initial Learning rate
+ "learning_rate",
+ low=1e-4,
+ high=1e-2,
+ log=True,
+ ),
+ "scaler_type": None, # Scaler type
+ "max_steps": 1000, # Max number of training iterations
+ "batch_size": trial.suggest_categorical( # Number of series in batch
+ "batch_size",
+ (1, 4, 10),
+ ),
+ "windows_batch_size": trial.suggest_categorical( # Number of windows in batch
+ "windows_batch_size",
+ (128, 256, 512),
+ ),
+ "random_seed": trial.suggest_int( # Random seed
+ "random_seed",
+ low=1,
+ high=20,
+ ),
+ }
+
+def config_lstm(trial):
+ return {
+ "input_size": trial.suggest_categorical( # Length of input window
+ "input_size",
+ (48, 48*2, 48*3)
+ ),
+ "encoder_hidden_size": trial.suggest_categorical( # Hidden size of LSTM cells
+ "encoder_hidden_size",
+ (64, 128),
+ ),
+ "encoder_n_layers": trial.suggest_categorical( # Number of layers in LSTM
+ "encoder_n_layers",
+ (2,4),
+ ),
+ "learning_rate": trial.suggest_float( # Initial Learning rate
+ "learning_rate",
+ low=1e-4,
+ high=1e-2,
+ log=True,
+ ),
+ "scaler_type": 'robust', # Scaler type
+ "max_steps": trial.suggest_categorical( # Max number of training iterations
+ "max_steps",
+ (500, 1000)
+ ),
+ "batch_size": trial.suggest_categorical( # Number of series in batch
+ "batch_size",
+ (1, 4)
+ ),
+ "random_seed": trial.suggest_int( # Random seed
+ "random_seed",
+ low=1,
+ high=20
+ ),
+ }
+```
+
+To instantiate an `Auto` model you need to define:
+
+- `h`: forecasting horizon.
+- `loss`: training and validation loss from
+ `neuralforecast.losses.pytorch`.
+- `config`: hyperparameter search space. If `None`, the `Auto` class
+ will use a pre-defined suggested hyperparameter space.
+- `search_alg`: search algorithm
+- `num_samples`: number of configurations explored.
+
+In this example we set horizon `h` as 48, use the
+[`MQLoss`](https://nixtlaverse.nixtla.io/neuralforecast/losses.pytorch.html#mqloss)
+distribution loss for training and validation, and use the default
+search algorithm.
+
+
+```python
+nf = NeuralForecast(
+ models=[
+ AutoNHITS(h=48, config=config_nhits, loss=MQLoss(), backend='optuna', num_samples=5),
+ AutoLSTM(h=48, config=config_lstm, loss=MQLoss(), backend='optuna', num_samples=2),
+ ],
+ freq=1,
+)
+```
+
+> **Tip**
+>
+> The number of samples, `num_samples`, is a crucial parameter! Larger
+> values will usually produce better results as we explore more
+> configurations in the search space, but it will increase training
+> times. Larger search spaces will usually require more samples. As a
+> general rule, we recommend setting `num_samples` higher than 20.
+
+Next, we use the `Neuralforecast` class to train the `Auto` model. In
+this step, `Auto` models will automatically perform hyperparameter
+tuning training multiple models with different hyperparameters,
+producing the forecasts on the validation set, and evaluating them. The
+best configuration is selected based on the error on a validation set.
+Only the best model is stored and used during inference.
+
+
+```python
+nf.fit(df=Y_df)
+```
+
+Next, we use the `predict` method to forecast the next 48 days using the
+optimal hyperparameters.
+
+
+```python
+fcst_df = nf.predict()
+fcst_df.columns = fcst_df.columns.str.replace('-median', '')
+fcst_df.head()
+```
+
+
+```python
+plot_series(Y_df, fcst_df, plot_random=False, max_insample_length=48 * 3, level=[80, 90])
+```
+
+
+
+The `plot_series` function allows for further customization. For
+example, plot the results of the different models and unique ids.
+
+
+```python
+# Plot to unique_ids and some selected models
+plot_series(Y_df, fcst_df, models=["AutoLSTM"], ids=["H107", "H104"], level=[80, 90])
+```
+
+
+
+
+```python
+# Explore other models
+plot_series(Y_df, fcst_df, models=["AutoNHITS"], ids=["H10", "H105"], level=[80, 90])
+```
+
+
+
+## 5. Evaluate the model’s performance
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, **Cross Validation** is done by defining a
+sliding window across the historical data and predicting the period
+following it. This form of cross-validation allows us to arrive at a
+better estimation of our model’s predictive abilities across a wider
+range of temporal instances while also keeping the data in the training
+set contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+> **Tip**
+>
+> Setting `n_windows=1` mirrors a traditional train-test split with our
+> historical data serving as the training set and the last 48 hours
+> serving as the testing set.
+
+The `cross_validation` method from the
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+class takes the following arguments.
+
+- `df`: training data frame
+
+- `step_size` (int): step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows` (int): number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+
+```python
+from neuralforecast.auto import AutoNHITS, AutoLSTM
+```
+
+
+```python
+nf = NeuralForecast(
+ models=[
+ AutoNHITS(h=48, config=config_nhits, loss=MQLoss(), num_samples=5, backend="optuna"),
+ AutoLSTM(h=48, config=config_lstm, loss=MQLoss(), num_samples=2, backend="optuna"),
+ ],
+ freq=1,
+)
+```
+
+
+```python
+cv_df = nf.cross_validation(Y_df, n_windows=2)
+```
+
+The `cv_df` object is a new data frame that includes the following
+columns:
+
+- `unique_id`: identifies each time series
+- `ds`: datestamp or temporal index
+- `cutoff`: the last datestamp or temporal index for the n_windows. If
+ n_windows=1, then one unique cuttoff value, if n_windows=2 then two
+ unique cutoff values.
+- `y`: true value
+- `"model"`: columns with the model’s name and fitted value.
+
+
+```python
+cv_df.columns = cv_df.columns.str.replace('-median', '')
+```
+
+
+```python
+cv_df.head()
+```
+
+| | unique_id | ds | cutoff | AutoNHITS | AutoNHITS-lo-90 | AutoNHITS-lo-80 | AutoNHITS-hi-80 | AutoNHITS-hi-90 | AutoLSTM | AutoLSTM-lo-90 | AutoLSTM-lo-80 | AutoLSTM-hi-80 | AutoLSTM-hi-90 | y |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | H1 | 700 | 699 | 654.506348 | 615.993774 | 616.021851 | 693.879272 | 712.376587 | 777.396362 | 511.052124 | 585.006470 | 992.880249 | 1084.980957 | 684.0 |
+| 1 | H1 | 701 | 699 | 619.320068 | 573.836060 | 577.762695 | 663.133301 | 683.214478 | 691.002991 | 417.614349 | 488.192810 | 905.101135 | 1002.091919 | 619.0 |
+| 2 | H1 | 702 | 699 | 546.807922 | 486.383362 | 498.541748 | 599.284302 | 623.889038 | 569.914795 | 314.173462 | 389.398865 | 763.250244 | 852.974121 | 565.0 |
+| 3 | H1 | 703 | 699 | 483.149811 | 420.416351 | 435.613708 | 536.380005 | 561.349487 | 548.401917 | 305.305054 | 379.597839 | 732.263123 | 817.543152 | 532.0 |
+| 4 | H1 | 704 | 699 | 434.347931 | 381.605713 | 394.665619 | 481.329041 | 501.715546 | 511.798950 | 269.810272 | 346.146484 | 692.443542 | 776.531921 | 495.0 |
+
+
+```python
+from IPython.display import display
+```
+
+
+```python
+for cutoff in cv_df['cutoff'].unique():
+ display(
+ plot_series(
+ Y_df,
+ cv_df.query('cutoff == @cutoff').drop(columns=['y', 'cutoff']),
+ max_insample_length=48 * 4,
+ ids=['H102'],
+ )
+ )
+```
+
+
+
+
+
+Now, let’s evaluate the models’ performance.
+
+
+```python
+from utilsforecast.evaluation import evaluate
+from utilsforecast.losses import mse, mae, rmse
+```
+
+> **Warning**
+>
+> You can also use Mean Average Percentage Error (MAPE), however for
+> granular forecasts, MAPE values are extremely [hard to
+> judge](%22https://blog.blueyonder.com/mean-absolute-percentage-error-mape-has-served-its-duty-and-should-now-retire/%22)
+> and not useful to assess forecasting quality.
+
+Create the data frame with the results of the evaluation of your
+cross-validation data frame using a Mean Squared Error metric.
+
+
+```python
+evaluation_df = evaluate(cv_df.drop(columns='cutoff'), metrics=[mse, mae, rmse])
+evaluation_df['best_model'] = evaluation_df.drop(columns=['metric', 'unique_id']).idxmin(axis=1)
+evaluation_df.head()
+```
+
+| | unique_id | metric | AutoNHITS | AutoLSTM | best_model |
+|-----|-----------|--------|--------------|--------------|------------|
+| 0 | H1 | mse | 2295.630068 | 1889.340182 | AutoLSTM |
+| 1 | H10 | mse | 724.468906 | 362.463659 | AutoLSTM |
+| 2 | H100 | mse | 62943.031250 | 17063.347107 | AutoLSTM |
+| 3 | H101 | mse | 48771.973540 | 12213.554997 | AutoLSTM |
+| 4 | H102 | mse | 30671.342050 | 84569.434859 | AutoNHITS |
+
+Create a summary table with a model column and the number of series
+where that model performs best.
+
+
+```python
+summary_df = evaluation_df.groupby(['metric', 'best_model']).size().sort_values().to_frame()
+summary_df = summary_df.reset_index()
+summary_df.columns = ['metric', 'model', 'nr. of unique_ids']
+summary_df
+```
+
+| | metric | model | nr. of unique_ids |
+|-----|--------|-----------|-------------------|
+| 0 | mae | AutoNHITS | 3 |
+| 1 | mse | AutoNHITS | 4 |
+| 2 | rmse | AutoNHITS | 4 |
+| 3 | mse | AutoLSTM | 6 |
+| 4 | rmse | AutoLSTM | 6 |
+| 5 | mae | AutoLSTM | 7 |
+
+
+```python
+summary_df.query('metric == "mse"')
+```
+
+| | metric | model | nr. of unique_ids |
+|-----|--------|-----------|-------------------|
+| 1 | mse | AutoNHITS | 4 |
+| 3 | mse | AutoLSTM | 6 |
+
+You can further explore your results by plotting the unique_ids where a
+specific model wins.
+
+
+```python
+nhits_ids = evaluation_df.query('best_model == "AutoNHITS" and metric == "mse"')['unique_id'].unique()
+
+plot_series(Y_df, fcst_df, ids=nhits_ids)
+```
+
+
+
+## 6. Select the best model for every unique series
+
+Define a utility function that takes your forecast’s data frame with the
+predictions and the evaluation data frame and returns a data frame with
+the best possible forecast for every unique_id.
+
+
+```python
+def get_best_model_forecast(forecasts_df, evaluation_df, metric):
+ metric_eval = evaluation_df.loc[evaluation_df['metric'] == metric, ['unique_id', 'best_model']]
+ with_best = forecasts_df.merge(metric_eval)
+ res = with_best[['unique_id', 'ds']].copy()
+ for suffix in ('', '-lo-90', '-hi-90'):
+ res[f'best_model{suffix}'] = with_best.apply(lambda row: row[row['best_model'] + suffix], axis=1)
+ return res
+```
+
+Create your production-ready data frame with the best forecast for every
+unique_id.
+
+
+```python
+prod_forecasts_df = get_best_model_forecast(fcst_df, evaluation_df, metric='mse')
+prod_forecasts_df
+```
+
+| | unique_id | ds | best_model | best_model-lo-90 | best_model-hi-90 |
+|-----|-----------|-----|-------------|------------------|------------------|
+| 0 | H1 | 749 | 603.923767 | 437.270447 | 786.502686 |
+| 1 | H1 | 750 | 533.691284 | 383.289154 | 702.944397 |
+| 2 | H1 | 751 | 490.400085 | 349.417816 | 648.831299 |
+| 3 | H1 | 752 | 463.768066 | 327.452026 | 616.572144 |
+| 4 | H1 | 753 | 454.710266 | 320.023468 | 605.468018 |
+| ... | ... | ... | ... | ... | ... |
+| 475 | H107 | 792 | 4720.256348 | 4142.459961 | 5235.727051 |
+| 476 | H107 | 793 | 4394.605469 | 3952.059082 | 4992.124023 |
+| 477 | H107 | 794 | 4161.221191 | 3664.091553 | 4632.160645 |
+| 478 | H107 | 795 | 3945.432617 | 3453.011963 | 4437.968750 |
+| 479 | H107 | 796 | 3666.446045 | 3177.937744 | 4059.684570 |
+
+Plot the results.
+
+
+```python
+plot_series(Y_df, prod_forecasts_df, level=[90])
+```
+
+
+
diff --git a/neuralforecast/docs/tutorials/hierarchical_forecasting.html.mdx b/neuralforecast/docs/tutorials/hierarchical_forecasting.html.mdx
new file mode 100644
index 00000000..a348043d
--- /dev/null
+++ b/neuralforecast/docs/tutorials/hierarchical_forecasting.html.mdx
@@ -0,0 +1,318 @@
+---
+output-file: hierarchical_forecasting.html
+title: Hierarchical Forecast
+---
+
+
+This notebook offers a step by step guide to create a hierarchical
+forecasting pipeline.
+
+In the pipeline we will use
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+and
+[`HINT`](https://nixtlaverse.nixtla.io/neuralforecast/models.hint.html#hint)
+class, to create fit, predict and reconcile forecasts.
+
+We will use the TourismL dataset that summarizes large Australian
+national visitor survey.
+
+Outline 1. Installing packages 2. Load hierarchical dataset
+3. Fit and Predict HINT 4. Benchmark methods 5. Forecast
+Evaluation
+
+You can run these experiments using GPU with Google Colab.
+
+
+
+## 1. Installing packages
+
+
+```python
+!pip install datasetsforecast hierarchicalforecast neuralforecast statsforecast
+```
+
+## 2. Load hierarchical dataset
+
+This detailed Australian Tourism Dataset comes from the National Visitor
+Survey, managed by the Tourism Research Australia, it is composed of 555
+monthly series from 1998 to 2016, it is organized geographically, and
+purpose of travel. The natural geographical hierarchy comprises seven
+states, divided further in 27 zones and 76 regions. The purpose of
+travel categories are holiday, visiting friends and relatives (VFR),
+business and other. The MinT (Wickramasuriya et al., 2019), among other
+hierarchical forecasting studies has used the dataset it in the past.
+The dataset can be accessed in the [MinT reconciliation
+webpage](https://robjhyndman.com/publications/mint/), although other
+sources are available.
+
+| Geographical Division | Number of series per division | Number of series per purpose | Total |
+|------------------|------------------|------------------|------------------|
+| Australia | 1 | 4 | 5 |
+| States | 7 | 28 | 35 |
+| Zones | 27 | 108 | 135 |
+| Regions | 76 | 304 | 380 |
+| Total | 111 | 444 | 555 |
+
+
+```python
+import pandas as pd
+
+from datasetsforecast.hierarchical import HierarchicalData
+from hierarchicalforecast.utils import aggregate, HierarchicalPlot
+from neuralforecast.utils import augment_calendar_df
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+# Load hierarchical dataset
+Y_df, S_df, tags = HierarchicalData.load('./data', 'TourismLarge')
+Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+Y_df, _ = augment_calendar_df(df=Y_df, freq='M')
+S_df = S_df.reset_index(names="unique_id")
+```
+
+Mathematically a hierarchical multivariate time series can be denoted by
+the vector $\mathbf{y}_{[a,b],t}$ defined by the following aggregation
+constraint:
+
+$$
+
+\mathbf{y}_{[a,b],t} = \mathbf{S}_{[a,b][b]} \mathbf{y}_{[b],t} \quad \Leftrightarrow \quad
+\begin{bmatrix}\mathbf{y}_{[a],t}
+\\ %\hline
+\mathbf{y}_{[b],t}\end{bmatrix}
+= \begin{bmatrix}
+\mathbf{A}_{[a][b]}\\ %\hline
+\mathbf{I}_{[b][b]}
+\end{bmatrix}
+\mathbf{y}_{[b],t}
+
+$$
+
+where $\mathbf{y}_{[a],t}$ are the aggregate series,
+$\mathbf{y}_{[b],t}$ are the bottom level series and
+$\mathbf{S}_{[a,b][b]}$ are the hierarchical aggregation constraints.
+
+
+```python
+# Here we plot the hierarchical constraints matrix
+hplot = HierarchicalPlot(S=S_df, tags=tags)
+hplot.plot_summing_matrix()
+```
+
+
+
+
+```python
+plot_series(forecasts_df=Y_df[["unique_id", "ds", "y"]], ids=['TotalAll'])
+```
+
+
+
+## 3. Fit and Predict HINT
+
+The Hierarchical Forecast Network (HINT) combines into an easy to use
+model three components: 1. SoTA neural forecast model. 2. An
+efficient and flexible multivariate probability distribution. 3.
+Builtin reconciliation capabilities.
+
+
+```python
+import logging
+
+import numpy as np
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import NHITS, HINT
+from neuralforecast.losses.pytorch import GMM, sCRPS
+```
+
+
+```python
+# Train test splits
+horizon = 12
+Y_test_df = Y_df.groupby('unique_id', observed=True).tail(horizon)
+Y_train_df = Y_df.drop(Y_test_df.index)
+```
+
+
+```python
+# Horizon and quantiles
+level = np.arange(0, 100, 2)
+qs = [[50-lv/2, 50+lv/2] if lv!=0 else [50] for lv in level]
+quantiles = np.sort(np.concatenate(qs)/100)
+
+# HINT := BaseNetwork + Distribution + Reconciliation
+nhits = NHITS(h=horizon,
+ input_size=24,
+ loss=GMM(n_components=10, quantiles=quantiles),
+ hist_exog_list=['month'],
+ max_steps=2000,
+ early_stop_patience_steps=10,
+ val_check_steps=50,
+ scaler_type='robust',
+ learning_rate=1e-3,
+ valid_loss=sCRPS(quantiles=quantiles))
+
+model = HINT(h=horizon, S=S_df.drop(columns='unique_id').values,
+ model=nhits, reconciliation='BottomUp')
+```
+
+``` text
+INFO:lightning_fabric.utilities.seed:Seed set to 1
+```
+
+
+```python
+logging.getLogger('pytorch_lightning').setLevel(logging.ERROR)
+```
+
+
+```python
+Y_df['y'] = Y_df['y'] * (Y_df['y'] > 0)
+nf = NeuralForecast(models=[model], freq='MS')
+nf.fit(df=Y_train_df, val_size=12)
+Y_hat_df = nf.predict()
+
+Y_hat_df = Y_hat_df.rename(columns=lambda x: x.replace('.0', ''))
+```
+
+
+```python
+plot_series(
+ Y_df,
+ Y_hat_df.drop(columns='NHITS-median'),
+ ids=['TotalAll'],
+ level=[90],
+ max_insample_length=12*5,
+)
+```
+
+
+
+## 4. Benchmark methods
+
+We compare against AutoARIMA, a well-established traditional forecasting
+method from the
+[StatsForecast](https://nixtlaverse.nixtla.io/statsforecast/index.html)
+package, for which we reconcile the forecasts using
+[HierarchicalForecast](https://nixtlaverse.nixtla.io/hierarchicalforecast/index.html).
+
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import AutoARIMA
+from hierarchicalforecast.methods import BottomUp, MinTrace
+from hierarchicalforecast.core import HierarchicalReconciliation
+```
+
+We define the model, and create the forecasts.
+
+
+```python
+sf = StatsForecast(models=[AutoARIMA()],
+ freq='MS', n_jobs=-1)
+Y_hat_df_arima = sf.forecast(df=Y_train_df,
+ h=12,
+ fitted=True,
+ X_df=Y_test_df.drop(columns="y"),
+ level = np.arange(2, 100, 2))
+Y_fitted_df_arima = sf.forecast_fitted_values()
+```
+
+Next, we reconcile the forecasts using `BottomUp` and `MinTrace`
+reconciliation techniques:
+
+
+```python
+reconcilers = [
+ BottomUp(),
+ MinTrace(method='mint_shrink'),
+]
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+Y_rec_df = hrec.reconcile(Y_hat_df=Y_hat_df_arima,
+ Y_df=Y_fitted_df_arima,
+ S=S_df,
+ tags=tags,
+ level = np.arange(2, 100, 2),
+ intervals_method="bootstrap")
+```
+
+## 5. Forecast Evaluation
+
+To evaluate the coherent probabilistic predictions we use the scaled
+Continuous Ranked Probability Score (sCRPS), defined as follows:
+
+$$
+
+\mathrm{CRPS}(\hat{F}_{[a,b],\tau},\mathbf{y}_{[a,b],\tau}) =
+ \frac{2}{N_{a}+N_{b}} \sum_{i} \int^{1}_{0} \mathrm{QL}(\hat{F}_{i,\tau}, y_{i,\tau})_{q} dq
+
+$$
+
+$$
+
+\mathrm{sCRPS}(\hat{F}_{[a,b\,],\tau},\mathbf{y}_{[a,b\,],\tau}) =
+ \frac{\mathrm{CRPS}(\hat{F}_{[a,b\,],\tau},\mathbf{y}_{[a,b\,],\tau})}{\sum_{i} | y_{i,\tau} |}
+
+$$
+
+As you can see the HINT model (using NHITS as base model) efficiently
+achieves state of the art accuracy under minimal tuning.
+
+
+```python
+from utilsforecast.losses import scaled_crps
+from hierarchicalforecast.evaluation import evaluate
+```
+
+
+```python
+df_metrics = Y_hat_df.merge(Y_test_df.drop(columns="month"), on=['unique_id', 'ds'])
+df_metrics = df_metrics.merge(Y_rec_df, on=['unique_id', 'ds'])
+
+metrics = evaluate(df = df_metrics,
+ tags = tags,
+ metrics = [scaled_crps],
+ models= ["NHITS", "AutoARIMA"],
+ level = np.arange(2, 100, 2),
+ train_df = Y_train_df.drop(columns="month"),
+ )
+
+metrics
+```
+
+| | level | metric | NHITS | AutoARIMA |
+|-----|-----------------------------------|-------------|----------|-----------|
+| 0 | Country | scaled_crps | 0.044431 | 0.131136 |
+| 1 | Country/State | scaled_crps | 0.063411 | 0.147516 |
+| 2 | Country/State/Zone | scaled_crps | 0.106060 | 0.174071 |
+| 3 | Country/State/Zone/Region | scaled_crps | 0.151988 | 0.205654 |
+| 4 | Country/Purpose | scaled_crps | 0.075821 | 0.133664 |
+| 5 | Country/State/Purpose | scaled_crps | 0.114674 | 0.181850 |
+| 6 | Country/State/Zone/Purpose | scaled_crps | 0.180491 | 0.244324 |
+| 7 | Country/State/Zone/Region/Purpose | scaled_crps | 0.245466 | 0.310656 |
+| 8 | Overall | scaled_crps | 0.122793 | 0.191109 |
+
+## References
+
+- [Kin G. Olivares, David Luo, Cristian Challu, Stefania La Vattiata,
+ Max Mergenthaler, Artur Dubrawski (2023). “HINT: Hierarchical
+ Mixture Networks For Coherent Probabilistic Forecasting”.
+ International Conference on Machine Learning (ICML). Workshop on
+ Structured Probabilistic Inference & Generative Modeling. Available
+ at
+ https://arxiv.org/abs/2305.07089.](https://arxiv.org/abs/2305.07089)
+- [Kin G. Olivares, O. Nganba Meetei, Ruijun Ma, Rohan Reddy, Mengfei
+ Cao, Lee Dicker (2023).”Probabilistic Hierarchical Forecasting with
+ Deep Poisson Mixtures”. International Journal Forecasting, accepted
+ paper. URL
+ https://arxiv.org/pdf/2110.13179.pdf.](https://arxiv.org/pdf/2110.13179.pdf)
+- [Kin G. Olivares, Federico Garza, David Luo, Cristian Challu, Max
+ Mergenthaler, Souhaib Ben Taieb, Shanika Wickramasuriya, and Artur
+ Dubrawski (2023). “HierarchicalForecast: A reference framework for
+ hierarchical forecasting”. Journal of Machine Learning Research,
+ submitted. URL
+ https://arxiv.org/abs/2207.03517](https://arxiv.org/abs/2207.03517)
+
diff --git a/neuralforecast/docs/tutorials/intermittent_data.html.mdx b/neuralforecast/docs/tutorials/intermittent_data.html.mdx
new file mode 100644
index 00000000..2b6ae945
--- /dev/null
+++ b/neuralforecast/docs/tutorials/intermittent_data.html.mdx
@@ -0,0 +1,369 @@
+---
+description: >-
+ In this notebook, we'll implement models for intermittent or sparse data using
+ the M5 dataset.
+output-file: intermittent_data.html
+title: Intermittent Data
+---
+
+
+Intermittent or sparse data has very few non-zero observations. This
+type of data is hard to forecast because the zero values increase the
+uncertainty about the underlying patterns in the data. Furthermore, once
+a non-zero observation occurs, there can be considerable variation in
+its size. Intermittent time series are common in many industries,
+including finance, retail, transportation, and energy. Given the
+ubiquity of this type of series, special methods have been developed to
+forecast them. The first was from [Croston (1972)](#ref), followed by
+several variants and by different aggregation frameworks.
+
+The models of [NeuralForecast](https://nixtla.github.io/statsforecast/)
+can be trained to model sparse or intermittent time series using a
+`Poisson` distribution loss. By the end of this tutorial, you’ll have a
+good understanding of these models and how to use them.
+
+**Outline:**
+
+1. Install libraries
+2. Load and explore the data
+3. Train models for intermittent data
+4. Perform Cross Validation
+
+> **Tip**
+>
+> You can use Colab to run this Notebook interactively
+>
+
+> **Warning**
+>
+> To reduce the computation time, it is recommended to use GPU. Using
+> Colab, do not forget to activate it. Just go to
+> `Runtime>Change runtime type` and select GPU as hardware accelerator.
+
+## 1. Install libraries
+
+We assume that you have NeuralForecast already installed. If not, check
+this guide for instructions on [how to install
+NeuralForecast](https://nixtla.github.io/neuralforecast/docs/getting-started/installation.html)
+
+Install the necessary packages using `pip install neuralforecast`
+
+
+```python
+!pip install statsforecast s3fs fastparquet neuralforecast
+```
+
+## 2. Load and explore the data
+
+For this example, we’ll use a subset of the [M5
+Competition](https://www.sciencedirect.com/science/article/pii/S0169207021001187#:~:text=The%20objective%20of%20the%20M5,the%20uncertainty%20around%20these%20forecasts)
+dataset. Each time series represents the unit sales of a particular
+product in a given Walmart store. At this level (product-store), most of
+the data is intermittent. We first need to import the data.
+
+
+```python
+import pandas as pd
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+Y_df = pd.read_parquet('https://m5-benchmarks.s3.amazonaws.com/data/train/target.parquet')
+Y_df = Y_df.rename(columns={
+ 'item_id': 'unique_id',
+ 'timestamp': 'ds',
+ 'demand': 'y'
+})
+Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+```
+
+For simplicity sake we will keep just one category
+
+
+```python
+Y_df = Y_df.query('unique_id.str.startswith("FOODS_3")')
+Y_df['unique_id'] = Y_df['unique_id'].astype(str)
+Y_df = Y_df.reset_index(drop=True)
+```
+
+Plot some series using the plot method from the `StatsForecast` class.
+This method prints 8 random series from the dataset and is useful for
+basic
+[EDA](https://nixtla.github.io/statsforecast/src/core/core.html#statsforecast.plot).
+
+
+```python
+plot_series(Y_df)
+```
+
+
+
+## 3. Train models for intermittent data
+
+
+```python
+from ray import tune
+
+from neuralforecast import NeuralForecast
+from neuralforecast.auto import AutoNHITS, AutoTFT
+from neuralforecast.losses.pytorch import DistributionLoss
+```
+
+Each `Auto` model contains a default search space that was extensively
+tested on multiple large-scale datasets. Additionally, users can define
+specific search spaces tailored for particular datasets and tasks.
+
+First, we create a custom search space for the
+[`AutoNHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autonhits)
+and
+[`AutoTFT`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotft)
+models. Search spaces are specified with dictionaries, where keys
+corresponds to the model’s hyperparameter and the value is a `Tune`
+function to specify how the hyperparameter will be sampled. For example,
+use `randint` to sample integers uniformly, and `choice` to sample
+values of a list.
+
+
+```python
+config_nhits = {
+ "input_size": tune.choice([28, 28*2, 28*3, 28*5]), # Length of input window
+ "n_blocks": 5*[1], # Length of input window
+ "mlp_units": 5 * [[512, 512]], # Length of input window
+ "n_pool_kernel_size": tune.choice([5*[1], 5*[2], 5*[4],
+ [8, 4, 2, 1, 1]]), # MaxPooling Kernel size
+ "n_freq_downsample": tune.choice([[8, 4, 2, 1, 1],
+ [1, 1, 1, 1, 1]]), # Interpolation expressivity ratios
+ "learning_rate": tune.loguniform(1e-4, 1e-2), # Initial Learning rate
+ "scaler_type": tune.choice([None]), # Scaler type
+ "max_steps": tune.choice([1000]), # Max number of training iterations
+ "batch_size": tune.choice([32, 64, 128, 256]), # Number of series in batch
+ "windows_batch_size": tune.choice([128, 256, 512, 1024]), # Number of windows in batch
+ "random_seed": tune.randint(1, 20), # Random seed
+}
+
+config_tft = {
+ "input_size": tune.choice([28, 28*2, 28*3]), # Length of input window
+ "hidden_size": tune.choice([64, 128, 256]), # Size of embeddings and encoders
+ "learning_rate": tune.loguniform(1e-4, 1e-2), # Initial learning rate
+ "scaler_type": tune.choice([None]), # Scaler type
+ "max_steps": tune.choice([500, 1000]), # Max number of training iterations
+ "batch_size": tune.choice([32, 64, 128, 256]), # Number of series in batch
+ "windows_batch_size": tune.choice([128, 256, 512, 1024]), # Number of windows in batch
+ "random_seed": tune.randint(1, 20), # Random seed
+ }
+```
+
+To instantiate an `Auto` model you need to define:
+
+- `h`: forecasting horizon.
+- `loss`: training and validation loss from
+ `neuralforecast.losses.pytorch`.
+- `config`: hyperparameter search space. If `None`, the `Auto` class
+ will use a pre-defined suggested hyperparameter space.
+- `search_alg`: search algorithm (from `tune.search`), default is
+ random search. Refer to
+ https://docs.ray.io/en/latest/tune/api_docs/suggestion.html for more
+ information on the different search algorithm options.
+- `num_samples`: number of configurations explored.
+
+In this example we set horizon `h` as 28, use the `Poisson` distribution
+loss (ideal for count data) for training and validation, and use the
+default search algorithm.
+
+
+```python
+nf = NeuralForecast(
+ models=[
+ AutoNHITS(h=28, config=config_nhits, loss=DistributionLoss(distribution='Poisson', level=[80, 90]), num_samples=5),
+ AutoTFT(h=28, config=config_tft, loss=DistributionLoss(distribution='Poisson', level=[80, 90]), num_samples=2),
+ ],
+ freq='D'
+)
+```
+
+> **Tip**
+>
+> The number of samples, `num_samples`, is a crucial parameter! Larger
+> values will usually produce better results as we explore more
+> configurations in the search space, but it will increase training
+> times. Larger search spaces will usually require more samples. As a
+> general rule, we recommend setting `num_samples` higher than 20.
+
+Next, we use the `Neuralforecast` class to train the `Auto` model. In
+this step, `Auto` models will automatically perform hyperparamter tuning
+training multiple models with different hyperparameters, producing the
+forecasts on the validation set, and evaluating them. The best
+configuration is selected based on the error on a validation set. Only
+the best model is stored and used during inference.
+
+
+```python
+nf.fit(df=Y_df)
+```
+
+Next, we use the `predict` method to forecast the next 28 days using the
+optimal hyperparameters.
+
+
+```python
+fcst_df = nf.predict()
+```
+
+``` text
+GPU available: True (cuda), used: True
+TPU available: False, using: 0 TPU cores
+HPU available: False, using: 0 HPUs
+LOCAL_RANK: 0 - CUDA_VISIBLE_DEVICES: [0]
+```
+
+``` text
+Predicting: | | 0/? [00:00 - Installing
+NeuralForecast. - Simulate a Harmonic Signal. - NHITS’ forecast
+decomposition. - NBEATSx’ forecast decomposition.
+
+You can run these experiments using GPU with Google Colab.
+
+
+
+## 1. Installing NeuralForecast
+
+
+```python
+!pip install neuralforecast
+```
+
+## 2. Simulate a Harmonic Signal
+
+In this example, we will consider a Harmonic signal comprising two
+frequencies: one low-frequency and one high-frequency.
+
+
+```python
+import numpy as np
+import pandas as pd
+```
+
+
+```python
+N = 10_000
+T = 1.0 / 800.0 # sample spacing
+x = np.linspace(0.0, N*T, N, endpoint=False)
+
+y1 = np.sin(10.0 * 2.0*np.pi*x)
+y2 = 0.5 * np.sin(100 * 2.0*np.pi*x)
+y = y1 + y2
+```
+
+
+```python
+import matplotlib.pyplot as plt
+plt.rcParams["axes.grid"]=True
+```
+
+
+```python
+fig, ax = plt.subplots(figsize=(6, 2.5))
+plt.plot(y[-80:], label='True')
+plt.plot(y1[-80:], label='Low Frequency', alpha=0.4)
+plt.plot(y2[-80:], label='High Frequency', alpha=0.4)
+plt.ylabel('Harmonic Signal')
+plt.xlabel('Time')
+plt.legend()
+plt.show()
+plt.close()
+```
+
+
+
+
+```python
+# Split dataset into train/test
+# Last horizon observations for test
+horizon = 96
+Y_df = pd.DataFrame(dict(unique_id=1, ds=np.arange(len(x)), y=y))
+Y_train_df = Y_df.groupby('unique_id').head(len(Y_df)-horizon)
+Y_test_df = Y_df.groupby('unique_id').tail(horizon)
+Y_test_df
+```
+
+| | unique_id | ds | y |
+|------|-----------|------|-----------|
+| 9904 | 1 | 9904 | -0.951057 |
+| 9905 | 1 | 9905 | -0.570326 |
+| 9906 | 1 | 9906 | -0.391007 |
+| 9907 | 1 | 9907 | -0.499087 |
+| 9908 | 1 | 9908 | -0.809017 |
+| ... | ... | ... | ... |
+| 9995 | 1 | 9995 | -0.029130 |
+| 9996 | 1 | 9996 | -0.309017 |
+| 9997 | 1 | 9997 | -0.586999 |
+| 9998 | 1 | 9998 | -0.656434 |
+| 9999 | 1 | 9999 | -0.432012 |
+
+## 3. NHITS decomposition
+
+We will employ
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+stack-specialization to recover the latent harmonic functions.
+
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits),
+a Wavelet-inspired algorithm, allows for breaking down a time series
+into various scales or resolutions, aiding in the identification of
+localized patterns or features. The expressivity ratios for each layer
+enable control over the model’s stack specialization.
+
+
+```python
+from neuralforecast.models import NHITS, NBEATSx
+from neuralforecast import NeuralForecast
+from neuralforecast.losses.pytorch import HuberLoss, MQLoss
+```
+
+
+```python
+models = [NHITS(h=horizon, # Forecast horizon
+ input_size=2 * horizon, # Length of input sequence
+ loss=HuberLoss(), # Robust Huber Loss
+ max_steps=1000, # Number of steps to train
+ dropout_prob_theta=0.5,
+ interpolation_mode='linear',
+ stack_types=['identity']*2,
+ n_blocks=[1, 1],
+ mlp_units=[[64, 64],[64, 64]],
+ n_freq_downsample=[10, 1], # Inverse expressivity ratios for NHITS' stacks specialization
+ val_check_steps=10, # Frequency of validation signal (affects early stopping)
+ )
+ ]
+nf = NeuralForecast(models=models, freq=1)
+nf.fit(df=Y_train_df)
+```
+
+
+```python
+from neuralforecast.tsdataset import TimeSeriesDataset
+
+# NHITS decomposition plot
+model = nf.models[0]
+dataset, *_ = TimeSeriesDataset.from_df(df = Y_train_df)
+y_hat = model.decompose(dataset=dataset)
+```
+
+``` text
+GPU available: True (cuda), used: True
+TPU available: False, using: 0 TPU cores
+HPU available: False, using: 0 HPUs
+LOCAL_RANK: 0 - CUDA_VISIBLE_DEVICES: [0]
+```
+
+``` text
+Predicting: | | 0/? [00:00
+- [Boris N. Oreshkin, Dmitri Carpov, Nicolas Chapados, Yoshua Bengio
+ (2019). “N-BEATS: Neural basis expansion analysis for interpretable
+ time series forecasting”.](https://arxiv.org/abs/1905.10437)
+- [Kin G. Olivares, Cristian Challu, Grzegorz Marcjasz, Rafał Weron,
+ Artur Dubrawski (2021). “Neural basis expansion analysis with
+ exogenous variables: Forecasting electricity prices with
+ NBEATSx”.](https://arxiv.org/abs/2104.05522)
+
diff --git a/neuralforecast/docs/tutorials/large_datasets.html.mdx b/neuralforecast/docs/tutorials/large_datasets.html.mdx
new file mode 100644
index 00000000..e164f0e8
--- /dev/null
+++ b/neuralforecast/docs/tutorials/large_datasets.html.mdx
@@ -0,0 +1,244 @@
+---
+description: >-
+ Tutorial on how to train neuralforecast models on datasets that cannot fit
+ into memory
+output-file: large_datasets.html
+title: Using Large Datasets
+---
+
+
+The standard DataLoader class used by NeuralForecast expects the dataset
+to be represented by a single DataFrame, which is entirely loaded into
+memory when fitting the model. However, when the dataset is too large
+for this, we can instead use the custom large-scale DataLoader. This
+custom loader assumes that each timeseries is split across a collection
+of Parquet files, and ensure that only one batch is ever loaded into
+memory at a given time.
+
+In this notebook, we will demonstrate the expected format of these
+files, how to train the model and and how to perform inference using
+this large-scale DataLoader.
+
+## Load libraries
+
+
+```python
+import logging
+import os
+import tempfile
+
+import pandas as pd
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import NHITS
+from utilsforecast.evaluation import evaluate
+from utilsforecast.losses import mae, rmse, smape
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+```
+
+
+```python
+logging.getLogger('pytorch_lightning').setLevel(logging.ERROR)
+```
+
+## Data
+
+Each timeseries should be stored in a directory named
+**unique_id=timeseries_id**. Within this directory, the timeseries can
+be entirely contained in a single Parquet file or split across multiple
+Parquet files. Regardless of the format, the timeseries must be ordered
+by time.
+
+For example, the following code splits the AirPassengers DataFrame (of
+which each timeseries is already sorted by time) into the below format:
+
+
+**\>** data
+ **\>** unique_id=Airline1
+ - a59945617fdb40d1bc6caa4aadad881c-0.parquet
+ **\>** unique_id=Airline2
+ - a59945617fdb40d1bc6caa4aadad881c-0.parquet
+
+ We then simply input a list of the paths to these directories.
+
+
+```python
+Y_df = AirPassengersPanel.copy()
+Y_df
+```
+
+| | unique_id | ds | y | trend | y\_\[lag12\] |
+|-----|-----------|------------|-------|-------|--------------|
+| 0 | Airline1 | 1949-01-31 | 112.0 | 0 | 112.0 |
+| 1 | Airline1 | 1949-02-28 | 118.0 | 1 | 118.0 |
+| 2 | Airline1 | 1949-03-31 | 132.0 | 2 | 132.0 |
+| 3 | Airline1 | 1949-04-30 | 129.0 | 3 | 129.0 |
+| 4 | Airline1 | 1949-05-31 | 121.0 | 4 | 121.0 |
+| ... | ... | ... | ... | ... | ... |
+| 283 | Airline2 | 1960-08-31 | 906.0 | 283 | 859.0 |
+| 284 | Airline2 | 1960-09-30 | 808.0 | 284 | 763.0 |
+| 285 | Airline2 | 1960-10-31 | 761.0 | 285 | 707.0 |
+| 286 | Airline2 | 1960-11-30 | 690.0 | 286 | 662.0 |
+| 287 | Airline2 | 1960-12-31 | 732.0 | 287 | 705.0 |
+
+
+```python
+valid = Y_df.groupby('unique_id').tail(72)
+# from now on we will use the id_col as the unique identifier for the timeseries (this is because we are using the unique_id column to partition the data into parquet files)
+valid = valid.rename(columns={'unique_id': 'id_col'})
+
+train = Y_df.drop(valid.index)
+train['id_col'] = train['unique_id'].copy()
+
+# we generate the files using a temporary directory here to demonstrate the expected file structure
+tmpdir = tempfile.TemporaryDirectory()
+train.to_parquet(tmpdir.name, partition_cols=['unique_id'], index=False)
+files_list = [f"{tmpdir.name}/{dir}" for dir in os.listdir(tmpdir.name)]
+files_list
+```
+
+``` text
+['C:\\Users\\ospra\\AppData\\Local\\Temp\\tmpxe__gjoo/unique_id=Airline1',
+ 'C:\\Users\\ospra\\AppData\\Local\\Temp\\tmpxe__gjoo/unique_id=Airline2']
+```
+
+You can also create this directory structure with a spark dataframe
+using the following:
+
+
+```python
+spark.conf.set("spark.sql.parquet.outputTimestampType", "TIMESTAMP_MICROS")
+(
+ spark_df
+ .repartition(id_col)
+ .sortWithinPartitions(id_col, time_col)
+ .write
+ .partitionBy(id_col)
+ .parquet(out_dir)
+)
+```
+
+The DataLoader class still expects the static data to be passed in as a
+single DataFrame with one row per timeseries.
+
+
+```python
+static = AirPassengersStatic.rename(columns={'unique_id': 'id_col'})
+static
+```
+
+| | id_col | airline1 | airline2 |
+|-----|----------|----------|----------|
+| 0 | Airline1 | 0 | 1 |
+| 1 | Airline2 | 1 | 0 |
+
+## Model training
+
+We now train a NHITS model on the above dataset. It is worth noting that
+NeuralForecast currently does not support scaling when using this
+DataLoader. If you want to scale the timeseries this should be done
+before passing it in to the `fit` method.
+
+
+```python
+horizon = 12
+stacks = 3
+models = [NHITS(input_size=5 * horizon,
+ h=horizon,
+ futr_exog_list=['trend', 'y_[lag12]'],
+ stat_exog_list=['airline1', 'airline2'],
+ max_steps=100,
+ stack_types = stacks*['identity'],
+ n_blocks = stacks*[1],
+ mlp_units = [[256,256] for _ in range(stacks)],
+ n_pool_kernel_size = stacks*[1],
+ interpolation_mode="nearest")]
+nf = NeuralForecast(models=models, freq='ME')
+nf.fit(df=files_list, static_df=static, id_col='id_col')
+```
+
+``` text
+Seed set to 1
+```
+
+``` text
+Sanity Checking: | | 0/? [00:00
+
+## 1. Installing NeuralForecast
+
+
+```python
+!pip install neuralforecast datasetsforecast
+```
+
+## 2. Load ETTm2 Data
+
+The `LongHorizon` class will automatically download the complete ETTm2
+dataset and process it.
+
+It return three Dataframes: `Y_df` contains the values for the target
+variables, `X_df` contains exogenous calendar features and `S_df`
+contains static features for each time-series (none for ETTm2). For this
+example we will only use `Y_df`.
+
+If you want to use your own data just replace `Y_df`. Be sure to use a
+long format and have a simmilar structure than our data set.
+
+
+```python
+import pandas as pd
+from datasetsforecast.long_horizon import LongHorizon
+
+# Change this to your own data to try the model
+Y_df, _, _ = LongHorizon.load(directory='./', group='ETTm2')
+Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+
+# For this excercise we are going to take 20% of the DataSet
+n_time = len(Y_df.ds.unique())
+val_size = int(.2 * n_time)
+test_size = int(.2 * n_time)
+
+Y_df.groupby('unique_id').head(2)
+```
+
+| | unique_id | ds | y |
+|--------|-----------|---------------------|-----------|
+| 0 | HUFL | 2016-07-01 00:00:00 | -0.041413 |
+| 1 | HUFL | 2016-07-01 00:15:00 | -0.185467 |
+| 57600 | HULL | 2016-07-01 00:00:00 | 0.040104 |
+| 57601 | HULL | 2016-07-01 00:15:00 | -0.214450 |
+| 115200 | LUFL | 2016-07-01 00:00:00 | 0.695804 |
+| 115201 | LUFL | 2016-07-01 00:15:00 | 0.434685 |
+| 172800 | LULL | 2016-07-01 00:00:00 | 0.434430 |
+| 172801 | LULL | 2016-07-01 00:15:00 | 0.428168 |
+| 230400 | MUFL | 2016-07-01 00:00:00 | -0.599211 |
+| 230401 | MUFL | 2016-07-01 00:15:00 | -0.658068 |
+| 288000 | MULL | 2016-07-01 00:00:00 | -0.393536 |
+| 288001 | MULL | 2016-07-01 00:15:00 | -0.659338 |
+| 345600 | OT | 2016-07-01 00:00:00 | 1.018032 |
+| 345601 | OT | 2016-07-01 00:15:00 | 0.980124 |
+
+
+```python
+import matplotlib.pyplot as plt
+
+# We are going to plot the temperature of the transformer
+# and marking the validation and train splits
+u_id = 'HUFL'
+x_plot = pd.to_datetime(Y_df[Y_df.unique_id==u_id].ds)
+y_plot = Y_df[Y_df.unique_id==u_id].y.values
+
+x_val = x_plot[n_time - val_size - test_size]
+x_test = x_plot[n_time - test_size]
+
+fig = plt.figure(figsize=(10, 5))
+fig.tight_layout()
+
+plt.plot(x_plot, y_plot)
+plt.xlabel('Date', fontsize=17)
+plt.ylabel('HUFL [15 min temperature]', fontsize=17)
+
+plt.axvline(x_val, color='black', linestyle='-.')
+plt.axvline(x_test, color='black', linestyle='-.')
+plt.text(x_val, 5, ' Validation', fontsize=12)
+plt.text(x_test, 5, ' Test', fontsize=12)
+
+plt.grid()
+```
+
+
+
+## 3. Hyperparameter selection and forecasting
+
+The
+[`AutoNHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autonhits)
+class will automatically perform hyperparamter tunning using [Tune
+library](https://docs.ray.io/en/latest/tune/index.html), exploring a
+user-defined or default search space. Models are selected based on the
+error on a validation set and the best model is then stored and used
+during inference.
+
+The `AutoNHITS.default_config` attribute contains a suggested
+hyperparameter space. Here, we specify a different search space
+following the paper’s hyperparameters. Notice that *1000 Stochastic
+Gradient Steps* are enough to achieve SoTA performance. Feel free to
+play around with this space.
+
+
+```python
+from ray import tune
+from neuralforecast.auto import AutoNHITS
+from neuralforecast.core import NeuralForecast
+```
+
+
+```python
+horizon = 96 # 24hrs = 4 * 15 min.
+
+# Use your own config or AutoNHITS.default_config
+nhits_config = {
+ "learning_rate": tune.choice([1e-3]), # Initial Learning rate
+ "max_steps": tune.choice([1000]), # Number of SGD steps
+ "input_size": tune.choice([5 * horizon]), # input_size = multiplier * horizon
+ "batch_size": tune.choice([7]), # Number of series in windows
+ "windows_batch_size": tune.choice([256]), # Number of windows in batch
+ "n_pool_kernel_size": tune.choice([[2, 2, 2], [16, 8, 1]]), # MaxPool's Kernelsize
+ "n_freq_downsample": tune.choice([[168, 24, 1], [24, 12, 1], [1, 1, 1]]), # Interpolation expressivity ratios
+ "activation": tune.choice(['ReLU']), # Type of non-linear activation
+ "n_blocks": tune.choice([[1, 1, 1]]), # Blocks per each 3 stacks
+ "mlp_units": tune.choice([[[512, 512], [512, 512], [512, 512]]]), # 2 512-Layers per block for each stack
+ "interpolation_mode": tune.choice(['linear']), # Type of multi-step interpolation
+ "val_check_steps": tune.choice([100]), # Compute validation every 100 epochs
+ "random_seed": tune.randint(1, 10),
+ }
+```
+
+> **Tip**
+>
+> Refer to https://docs.ray.io/en/latest/tune/index.html for more
+> information on the different space options, such as lists and
+> continous intervals.m
+
+To instantiate
+[`AutoNHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autonhits)
+you need to define:
+
+- `h`: forecasting horizon
+- `loss`: training loss. Use the
+ [`DistributionLoss`](https://nixtlaverse.nixtla.io/neuralforecast/losses.pytorch.html#distributionloss)
+ to produce probabilistic forecasts.
+- `config`: hyperparameter search space. If `None`, the
+ [`AutoNHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autonhits)
+ class will use a pre-defined suggested hyperparameter space.
+- `num_samples`: number of configurations explored.
+
+
+```python
+models = [AutoNHITS(h=horizon,
+ config=nhits_config,
+ num_samples=5)]
+```
+
+Fit the model by instantiating a
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+object with the following required parameters:
+
+- `models`: a list of models.
+
+- `freq`: a string indicating the frequency of the data. (See [panda’s
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+The `cross_validation` method allows you to simulate multiple historic
+forecasts, greatly simplifying pipelines by replacing for loops with
+`fit` and `predict` methods.
+
+With time series data, cross validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The `cross_validation` method will use the validation set for
+hyperparameter selection, and will then produce the forecasts for the
+test set.
+
+
+```python
+nf = NeuralForecast(
+ models=models,
+ freq='15min')
+
+Y_hat_df = nf.cross_validation(df=Y_df, val_size=val_size,
+ test_size=test_size, n_windows=None)
+```
+
+## 4. Evaluate Results
+
+The
+[`AutoNHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autonhits)
+class contains a `results` tune attribute that stores information of
+each configuration explored. It contains the validation loss and best
+validation hyperparameter.
+
+
+```python
+nf.models[0].results.get_best_result().config
+```
+
+``` text
+{'learning_rate': 0.001,
+ 'max_steps': 1000,
+ 'input_size': 480,
+ 'batch_size': 7,
+ 'windows_batch_size': 256,
+ 'n_pool_kernel_size': [2, 2, 2],
+ 'n_freq_downsample': [24, 12, 1],
+ 'activation': 'ReLU',
+ 'n_blocks': [1, 1, 1],
+ 'mlp_units': [[512, 512], [512, 512], [512, 512]],
+ 'interpolation_mode': 'linear',
+ 'val_check_steps': 100,
+ 'random_seed': 8,
+ 'h': 96,
+ 'loss': MAE(),
+ 'valid_loss': MAE()}
+```
+
+
+```python
+y_true = Y_hat_df.y.values
+y_hat = Y_hat_df['AutoNHITS'].values
+
+n_series = len(Y_df.unique_id.unique())
+
+y_true = y_true.reshape(n_series, -1, horizon)
+y_hat = y_hat.reshape(n_series, -1, horizon)
+
+print('Parsed results')
+print('2. y_true.shape (n_series, n_windows, n_time_out):\t', y_true.shape)
+print('2. y_hat.shape (n_series, n_windows, n_time_out):\t', y_hat.shape)
+```
+
+``` text
+Parsed results
+2. y_true.shape (n_series, n_windows, n_time_out): (7, 11425, 96)
+2. y_hat.shape (n_series, n_windows, n_time_out): (7, 11425, 96)
+```
+
+
+```python
+fig, axs = plt.subplots(nrows=3, ncols=1, figsize=(10, 11))
+fig.tight_layout()
+
+series = ['HUFL','HULL','LUFL','LULL','MUFL','MULL','OT']
+series_idx = 3
+
+for idx, w_idx in enumerate([200, 300, 400]):
+ axs[idx].plot(y_true[series_idx, w_idx,:],label='True')
+ axs[idx].plot(y_hat[series_idx, w_idx,:],label='Forecast')
+ axs[idx].grid()
+ axs[idx].set_ylabel(series[series_idx]+f' window {w_idx}',
+ fontsize=17)
+ if idx==2:
+ axs[idx].set_xlabel('Forecast Horizon', fontsize=17)
+plt.legend()
+plt.show()
+plt.close()
+```
+
+
+
+Finally, we compute the test errors for the two metrics of interest:
+
+$\qquad MAE = \frac{1}{Windows * Horizon} \sum_{\tau} |y_{\tau} - \hat{y}_{\tau}| \qquad$
+and
+$\qquad MSE = \frac{1}{Windows * Horizon} \sum_{\tau} (y_{\tau} - \hat{y}_{\tau})^{2} \qquad$
+
+
+```python
+from neuralforecast.losses.numpy import mae, mse
+
+print('MAE: ', mae(y_hat, y_true))
+print('MSE: ', mse(y_hat, y_true))
+```
+
+``` text
+MAE: 0.24862242128243706
+MSE: 0.17257850996828134
+```
+
+For reference we can check the performance when compared to previous
+‘state-of-the-art’ long-horizon Transformer-based forecasting methods
+from the [NHITS paper](https://arxiv.org/abs/2201.12886). To recover or
+improve the paper results try setting `hyperopt_max_evals=30` in
+[Hyperparameter Tuning](#cell-4).
+
+Mean Absolute Error (MAE):
+
+| Horizon | NHITS | AutoFormer | InFormer | ARIMA |
+|---------|-----------|------------|----------|-------|
+| 96 | **0.249** | 0.339 | 0.453 | 0.301 |
+| 192 | 0.305 | 0.340 | 0.563 | 0.345 |
+| 336 | 0.346 | 0.372 | 0.887 | 0.386 |
+| 720 | 0.426 | 0.419 | 1.388 | 0.445 |
+
+Mean Squared Error (MSE):
+
+| Horizon | NHITS | AutoFormer | InFormer | ARIMA |
+|---------|-----------|------------|----------|-------|
+| 96 | **0.173** | 0.255 | 0.365 | 0.225 |
+| 192 | 0.245 | 0.281 | 0.533 | 0.298 |
+| 336 | 0.295 | 0.339 | 1.363 | 0.370 |
+| 720 | 0.401 | 0.422 | 3.379 | 0.478 |
+
+## References
+
+[Cristian Challu, Kin G. Olivares, Boris N. Oreshkin, Federico Garza,
+Max Mergenthaler-Canseco, Artur Dubrawski (2021). NHITS: Neural
+Hierarchical Interpolation for Time Series Forecasting. Accepted at AAAI
+2023.](https://arxiv.org/abs/2201.12886)
+
diff --git a/neuralforecast/docs/tutorials/longhorizon_probabilistic.html.mdx b/neuralforecast/docs/tutorials/longhorizon_probabilistic.html.mdx
new file mode 100644
index 00000000..d3df1019
--- /dev/null
+++ b/neuralforecast/docs/tutorials/longhorizon_probabilistic.html.mdx
@@ -0,0 +1,290 @@
+---
+output-file: longhorizon_probabilistic.html
+title: Long-Horizon Probabilistic Forecasting
+---
+
+
+Long-horizon forecasting is challenging because of the *volatility* of
+the predictions and the *computational complexity*. To solve this
+problem we created the [NHITS](https://arxiv.org/abs/2201.12886) model
+and made the code available [NeuralForecast
+library](https://nixtla.github.io/neuralforecast/models.nhits.html).
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+specializes its partial outputs in the different frequencies of the time
+series through hierarchical interpolation and multi-rate input
+processing. We model the target time-series with Student’s
+t-distribution. The
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+will output the distribution parameters for each timestamp.
+
+In this notebook we show how to use
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+on the [ETTm2](https://github.com/zhouhaoyi/ETDataset) benchmark dataset
+for probabilistic forecasting. This data set includes data points for 2
+Electricity Transformers at 2 stations, including load, oil temperature.
+
+We will show you how to load data, train, and perform automatic
+hyperparameter tuning, **to achieve SoTA performance**, outperforming
+even the latest Transformer architectures for a fraction of their
+computational cost (50x faster).
+
+You can run these experiments using GPU with Google Colab.
+
+
+
+## 1. Libraries
+
+
+```python
+!pip install neuralforecast datasetsforecast
+```
+
+## 2. Load ETTm2 Data
+
+The `LongHorizon` class will automatically download the complete ETTm2
+dataset and process it.
+
+It return three Dataframes: `Y_df` contains the values for the target
+variables, `X_df` contains exogenous calendar features and `S_df`
+contains static features for each time-series (none for ETTm2). For this
+example we will only use `Y_df`.
+
+If you want to use your own data just replace `Y_df`. Be sure to use a
+long format and have a simmilar structure than our data set.
+
+
+```python
+import pandas as pd
+from datasetsforecast.long_horizon import LongHorizon
+```
+
+
+```python
+# Change this to your own data to try the model
+Y_df, _, _ = LongHorizon.load(directory='./', group='ETTm2')
+Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+
+# For this excercise we are going to take 960 timestamps as validation and test
+n_time = len(Y_df.ds.unique())
+val_size = 96*10
+test_size = 96*10
+
+Y_df.groupby('unique_id').head(2)
+```
+
+| | unique_id | ds | y |
+|--------|-----------|---------------------|-----------|
+| 0 | HUFL | 2016-07-01 00:00:00 | -0.041413 |
+| 1 | HUFL | 2016-07-01 00:15:00 | -0.185467 |
+| 57600 | HULL | 2016-07-01 00:00:00 | 0.040104 |
+| 57601 | HULL | 2016-07-01 00:15:00 | -0.214450 |
+| 115200 | LUFL | 2016-07-01 00:00:00 | 0.695804 |
+| 115201 | LUFL | 2016-07-01 00:15:00 | 0.434685 |
+| 172800 | LULL | 2016-07-01 00:00:00 | 0.434430 |
+| 172801 | LULL | 2016-07-01 00:15:00 | 0.428168 |
+| 230400 | MUFL | 2016-07-01 00:00:00 | -0.599211 |
+| 230401 | MUFL | 2016-07-01 00:15:00 | -0.658068 |
+| 288000 | MULL | 2016-07-01 00:00:00 | -0.393536 |
+| 288001 | MULL | 2016-07-01 00:15:00 | -0.659338 |
+| 345600 | OT | 2016-07-01 00:00:00 | 1.018032 |
+| 345601 | OT | 2016-07-01 00:15:00 | 0.980124 |
+
+> **Important**
+>
+> DataFrames must include all `['unique_id', 'ds', 'y']` columns. Make
+> sure `y` column does not have missing or non-numeric values.
+
+Next, plot the `HUFL` variable marking the validation and train splits.
+
+
+```python
+import matplotlib.pyplot as plt
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+u_id = 'HUFL'
+fig = plot_series(Y_df, ids=[u_id])
+ax = fig.axes[0]
+
+x_plot = pd.to_datetime(Y_df[Y_df.unique_id==u_id].ds)
+y_plot = Y_df[Y_df.unique_id==u_id].y.values
+x_val = x_plot[n_time - val_size - test_size]
+x_test = x_plot[n_time - test_size]
+
+ax.axvline(x_val, color='black', linestyle='-.')
+ax.axvline(x_test, color='black', linestyle='-.')
+ax.text(x_val, 5, ' Validation', fontsize=12)
+ax.text(x_test, 3, ' Test', fontsize=12)
+fig
+```
+
+
+
+## 3. Hyperparameter selection and forecasting
+
+The
+[`AutoNHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autonhits)
+class will automatically perform hyperparamter tunning using [Tune
+library](https://docs.ray.io/en/latest/tune/index.html), exploring a
+user-defined or default search space. Models are selected based on the
+error on a validation set and the best model is then stored and used
+during inference.
+
+The `AutoNHITS.default_config` attribute contains a suggested
+hyperparameter space. Here, we specify a different search space
+following the paper’s hyperparameters. Notice that *1000 Stochastic
+Gradient Steps* are enough to achieve SoTA performance. Feel free to
+play around with this space.
+
+
+```python
+import logging
+
+import torch
+from neuralforecast.auto import AutoNHITS
+from neuralforecast.core import NeuralForecast
+from neuralforecast.losses.pytorch import DistributionLoss
+from ray import tune
+```
+
+
+```python
+logging.getLogger("pytorch_lightning").setLevel(logging.WARNING)
+torch.set_float32_matmul_precision('high')
+```
+
+
+```python
+horizon = 96 # 24hrs = 4 * 15 min.
+
+# Use your own config or AutoNHITS.default_config
+nhits_config = {
+ "learning_rate": tune.choice([1e-3]), # Initial Learning rate
+ "max_steps": tune.choice([1000]), # Number of SGD steps
+ "input_size": tune.choice([5 * horizon]), # input_size = multiplier * horizon
+ "batch_size": tune.choice([7]), # Number of series in windows
+ "windows_batch_size": tune.choice([256]), # Number of windows in batch
+ "n_pool_kernel_size": tune.choice([[2, 2, 2], [16, 8, 1]]), # MaxPool's Kernelsize
+ "n_freq_downsample": tune.choice([[168, 24, 1], [24, 12, 1], [1, 1, 1]]), # Interpolation expressivity ratios
+ "activation": tune.choice(['ReLU']), # Type of non-linear activation
+ "n_blocks": tune.choice([[1, 1, 1]]), # Blocks per each 3 stacks
+ "mlp_units": tune.choice([[[512, 512], [512, 512], [512, 512]]]), # 2 512-Layers per block for each stack
+ "interpolation_mode": tune.choice(['linear']), # Type of multi-step interpolation
+ "random_seed": tune.randint(1, 10),
+ "scaler_type": tune.choice(['robust']),
+ "val_check_steps": tune.choice([100])
+ }
+```
+
+> **Tip**
+>
+> Refer to https://docs.ray.io/en/latest/tune/index.html for more
+> information on the different space options, such as lists and
+> continous intervals.m
+
+To instantiate
+[`AutoNHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autonhits)
+you need to define:
+
+- `h`: forecasting horizon
+- `loss`: training loss. Use the
+ [`DistributionLoss`](https://nixtlaverse.nixtla.io/neuralforecast/losses.pytorch.html#distributionloss)
+ to produce probabilistic forecasts.
+- `config`: hyperparameter search space. If `None`, the
+ [`AutoNHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autonhits)
+ class will use a pre-defined suggested hyperparameter space.
+- `num_samples`: number of configurations explored.
+
+
+```python
+models = [AutoNHITS(h=horizon,
+ loss=DistributionLoss(distribution='StudentT', level=[80, 90]),
+ config=nhits_config,
+ num_samples=5)]
+```
+
+Fit the model by instantiating a
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+object with the following required parameters:
+
+- `models`: a list of models.
+
+- `freq`: a string indicating the frequency of the data. (See [panda’s
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+
+```python
+# Fit and predict
+nf = NeuralForecast(models=models, freq='15min')
+```
+
+The `cross_validation` method allows you to simulate multiple historic
+forecasts, greatly simplifying pipelines by replacing for loops with
+`fit` and `predict` methods.
+
+With time series data, cross validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The `cross_validation` method will use the validation set for
+hyperparameter selection, and will then produce the forecasts for the
+test set.
+
+
+```python
+Y_hat_df = nf.cross_validation(df=Y_df, val_size=val_size,
+ test_size=test_size, n_windows=None)
+```
+
+## 4. Visualization
+
+Finally, we merge the forecasts with the `Y_df` dataset and plot the
+forecasts.
+
+
+```python
+Y_hat_df
+```
+
+| | unique_id | ds | cutoff | AutoNHITS | AutoNHITS-median | AutoNHITS-lo-90 | AutoNHITS-lo-80 | AutoNHITS-hi-80 | AutoNHITS-hi-90 | y |
+|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | HUFL | 2018-02-11 00:00:00 | 2018-02-10 23:45:00 | -0.922304 | -0.914175 | -1.217987 | -1.138274 | -0.708157 | -0.617799 | -0.849571 |
+| 1 | HUFL | 2018-02-11 00:15:00 | 2018-02-10 23:45:00 | -0.954299 | -0.957198 | -1.403932 | -1.263984 | -0.618467 | -0.442688 | -1.049700 |
+| 2 | HUFL | 2018-02-11 00:30:00 | 2018-02-10 23:45:00 | -0.987538 | -0.972558 | -1.512509 | -1.310191 | -0.621673 | -0.444359 | -1.185730 |
+| 3 | HUFL | 2018-02-11 00:45:00 | 2018-02-10 23:45:00 | -1.067760 | -1.063188 | -1.614276 | -1.475302 | -0.665729 | -0.521775 | -1.329785 |
+| 4 | HUFL | 2018-02-11 01:00:00 | 2018-02-10 23:45:00 | -1.001276 | -1.001494 | -1.508795 | -1.390156 | -0.629212 | -0.470608 | -1.369715 |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| 581275 | OT | 2018-02-20 22:45:00 | 2018-02-19 23:45:00 | -1.200041 | -1.200862 | -1.591271 | -1.490571 | -0.907190 | -0.779424 | -1.581325 |
+| 581276 | OT | 2018-02-20 23:00:00 | 2018-02-19 23:45:00 | -1.237206 | -1.225333 | -1.618691 | -1.518204 | -0.960075 | -0.838512 | -1.581325 |
+| 581277 | OT | 2018-02-20 23:15:00 | 2018-02-19 23:45:00 | -1.232434 | -1.229675 | -1.591164 | -1.481251 | -0.989993 | -0.870404 | -1.581325 |
+| 581278 | OT | 2018-02-20 23:30:00 | 2018-02-19 23:45:00 | -1.259237 | -1.258848 | -1.659239 | -1.536979 | -0.985581 | -0.822370 | -1.562328 |
+| 581279 | OT | 2018-02-20 23:45:00 | 2018-02-19 23:45:00 | -1.247161 | -1.251899 | -1.631909 | -1.520350 | -0.949529 | -0.832602 | -1.562328 |
+
+
+```python
+Y_hat_df = Y_hat_df.reset_index(drop=True)
+Y_hat_df = Y_hat_df[(Y_hat_df['unique_id']=='OT') & (Y_hat_df['cutoff']=='2018-02-11 12:00:00')]
+Y_hat_df = Y_hat_df.drop(columns=['y','cutoff'])
+```
+
+
+```python
+plot_df = Y_df.merge(Y_hat_df, on=['unique_id','ds'], how='outer').tail(96*10+50+96*4).head(96*2+96*4)
+plot_series(forecasts_df=plot_df.drop(columns='AutoNHITS').rename(columns={'AutoNHITS-median': 'AutoNHITS'}), level=[90])
+```
+
+
+
+## References
+
+[Cristian Challu, Kin G. Olivares, Boris N. Oreshkin, Federico Garza,
+Max Mergenthaler-Canseco, Artur Dubrawski (2021). NHITS: Neural
+Hierarchical Interpolation for Time Series Forecasting. Accepted at AAAI
+2023.](https://arxiv.org/abs/2201.12886)
+
diff --git a/neuralforecast/docs/tutorials/longhorizon_transformers.html.mdx b/neuralforecast/docs/tutorials/longhorizon_transformers.html.mdx
new file mode 100644
index 00000000..4d9a894c
--- /dev/null
+++ b/neuralforecast/docs/tutorials/longhorizon_transformers.html.mdx
@@ -0,0 +1,316 @@
+---
+description: Tutorial on how to train and forecast Transformer models.
+output-file: longhorizon_transformers.html
+title: Long-Horizon Forecasting with Transformer models
+---
+
+
+Transformer models, originally proposed for applications in natural
+language processing, have seen increasing adoption in the field of time
+series forecasting. The transformative power of these models lies in
+their novel architecture that relies heavily on the self-attention
+mechanism, which helps the model to focus on different parts of the
+input sequence to make predictions, while capturing long-range
+dependencies within the data. In the context of time series forecasting,
+Transformer models leverage this self-attention mechanism to identify
+relevant information across different periods in the time series, making
+them exceptionally effective in predicting future values for complex and
+noisy sequences.
+
+Long horizon forecasting consists of predicting a large number of
+timestamps. It is a challenging task because of the *volatility* of the
+predictions and the *computational complexity*. To solve this problem,
+recent studies proposed a variety of Transformer-based models.
+
+The Neuralforecast library includes implementations of the following
+popular recent models:
+[`Informer`](https://nixtlaverse.nixtla.io/neuralforecast/models.informer.html#informer)
+(Zhou, H. et al. 2021),
+[`Autoformer`](https://nixtlaverse.nixtla.io/neuralforecast/models.autoformer.html#autoformer)
+(Wu et al. 2021),
+[`FEDformer`](https://nixtlaverse.nixtla.io/neuralforecast/models.fedformer.html#fedformer)
+(Zhou, T. et al. 2022), and
+[`PatchTST`](https://nixtlaverse.nixtla.io/neuralforecast/models.patchtst.html#patchtst)
+(Nie et al. 2023).
+
+Our implementation of all these models are univariate, meaning that only
+autoregressive values of each feature are used for forecasting. **We
+observed that these unvivariate models are more accurate and faster than
+their multivariate couterpart**.
+
+In this notebook we will show how to: \* Load the
+[ETTm2](https://github.com/zhouhaoyi/ETDataset) benchmark dataset, used
+in the academic literature. \* Train models \* Forecast the test set
+
+**The results achieved in this notebook outperform the original
+self-reported results in the respective original paper, with a fraction
+of the computational cost. Additionally, all models are trained with the
+default recommended parameters, results can be further improved using
+our `auto` models with automatic hyperparameter selection.**
+
+You can run these experiments using GPU with Google Colab.
+
+
+
+## 1. Installing libraries
+
+
+```python
+!pip install neuralforecast datasetsforecast
+```
+
+## 2. Load ETTm2 Data
+
+The `LongHorizon` class will automatically download the complete ETTm2
+dataset and process it.
+
+It return three Dataframes: `Y_df` contains the values for the target
+variables, `X_df` contains exogenous calendar features and `S_df`
+contains static features for each time-series (none for ETTm2). For this
+example we will only use `Y_df`.
+
+If you want to use your own data just replace `Y_df`. Be sure to use a
+long format and have a simmilar structure than our data set.
+
+
+```python
+import pandas as pd
+
+from datasetsforecast.long_horizon import LongHorizon
+```
+
+
+```python
+# Change this to your own data to try the model
+Y_df, _, _ = LongHorizon.load(directory='./', group='ETTm2')
+Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+
+n_time = len(Y_df.ds.unique())
+val_size = int(.2 * n_time)
+test_size = int(.2 * n_time)
+
+Y_df.groupby('unique_id').head(2)
+```
+
+| | unique_id | ds | y |
+|--------|-----------|---------------------|-----------|
+| 0 | HUFL | 2016-07-01 00:00:00 | -0.041413 |
+| 1 | HUFL | 2016-07-01 00:15:00 | -0.185467 |
+| 57600 | HULL | 2016-07-01 00:00:00 | 0.040104 |
+| 57601 | HULL | 2016-07-01 00:15:00 | -0.214450 |
+| 115200 | LUFL | 2016-07-01 00:00:00 | 0.695804 |
+| 115201 | LUFL | 2016-07-01 00:15:00 | 0.434685 |
+| 172800 | LULL | 2016-07-01 00:00:00 | 0.434430 |
+| 172801 | LULL | 2016-07-01 00:15:00 | 0.428168 |
+| 230400 | MUFL | 2016-07-01 00:00:00 | -0.599211 |
+| 230401 | MUFL | 2016-07-01 00:15:00 | -0.658068 |
+| 288000 | MULL | 2016-07-01 00:00:00 | -0.393536 |
+| 288001 | MULL | 2016-07-01 00:15:00 | -0.659338 |
+| 345600 | OT | 2016-07-01 00:00:00 | 1.018032 |
+| 345601 | OT | 2016-07-01 00:15:00 | 0.980124 |
+
+## 3. Train models
+
+We will train models using the `cross_validation` method, which allows
+users to automatically simulate multiple historic forecasts (in the test
+set).
+
+The `cross_validation` method will use the validation set for
+hyperparameter selection and early stopping, and will then produce the
+forecasts for the test set.
+
+First, instantiate each model in the `models` list, specifying the
+`horizon`, `input_size`, and training iterations.
+
+(NOTE: The
+[`FEDformer`](https://nixtlaverse.nixtla.io/neuralforecast/models.fedformer.html#fedformer)
+model was excluded due to extremely long training times.)
+
+
+```python
+from neuralforecast.core import NeuralForecast
+from neuralforecast.models import Informer, Autoformer, FEDformer, PatchTST
+```
+
+``` text
+INFO:torch.distributed.nn.jit.instantiator:Created a temporary directory at /tmp/tmpopb2vyyt
+INFO:torch.distributed.nn.jit.instantiator:Writing /tmp/tmpopb2vyyt/_remote_module_non_scriptable.py
+```
+
+
+```python
+horizon = 96 # 24hrs = 4 * 15 min.
+models = [Informer(h=horizon, # Forecasting horizon
+ input_size=horizon, # Input size
+ max_steps=1000, # Number of training iterations
+ val_check_steps=100, # Compute validation loss every 100 steps
+ early_stop_patience_steps=3), # Stop training if validation loss does not improve
+ Autoformer(h=horizon,
+ input_size=horizon,
+ max_steps=1000,
+ val_check_steps=100,
+ early_stop_patience_steps=3),
+ PatchTST(h=horizon,
+ input_size=horizon,
+ max_steps=1000,
+ val_check_steps=100,
+ early_stop_patience_steps=3),
+ ]
+```
+
+``` text
+INFO:lightning_fabric.utilities.seed:Global seed set to 1
+INFO:lightning_fabric.utilities.seed:Global seed set to 1
+INFO:lightning_fabric.utilities.seed:Global seed set to 1
+```
+
+> **Tip**
+>
+> Check our `auto` models for automatic hyperparameter optimization.
+
+Instantiate a
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+object with the following required parameters:
+
+- `models`: a list of models.
+
+- `freq`: a string indicating the frequency of the data. (See [panda’s
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+Second, use the `cross_validation` method, specifying the dataset
+(`Y_df`), validation size and test size.
+
+
+```python
+nf = NeuralForecast(
+ models=models,
+ freq='15min')
+
+Y_hat_df = nf.cross_validation(df=Y_df,
+ val_size=val_size,
+ test_size=test_size,
+ n_windows=None)
+```
+
+The `cross_validation` method will return the forecasts for each model
+on the test set.
+
+
+```python
+Y_hat_df.head()
+```
+
+| | unique_id | ds | cutoff | Informer | Autoformer | PatchTST | y |
+|----|----|----|----|----|----|----|----|
+| 0 | HUFL | 2017-10-24 00:00:00 | 2017-10-23 23:45:00 | -1.055062 | -0.861487 | -0.860189 | -0.977673 |
+| 1 | HUFL | 2017-10-24 00:15:00 | 2017-10-23 23:45:00 | -1.021247 | -0.873399 | -0.865730 | -0.865620 |
+| 2 | HUFL | 2017-10-24 00:30:00 | 2017-10-23 23:45:00 | -1.057297 | -0.900345 | -0.944296 | -0.961624 |
+| 3 | HUFL | 2017-10-24 00:45:00 | 2017-10-23 23:45:00 | -0.886652 | -0.867466 | -0.974849 | -1.049700 |
+| 4 | HUFL | 2017-10-24 01:00:00 | 2017-10-23 23:45:00 | -1.000431 | -0.887454 | -1.008530 | -0.953600 |
+
+## 4. Evaluate Results
+
+Next, we plot the forecasts on the test set for the `OT` variable for
+all models.
+
+
+```python
+import matplotlib.pyplot as plt
+```
+
+
+```python
+Y_plot = Y_hat_df[Y_hat_df['unique_id']=='OT'] # OT dataset
+cutoffs = Y_hat_df['cutoff'].unique()[::horizon]
+Y_plot = Y_plot[Y_hat_df['cutoff'].isin(cutoffs)]
+
+plt.figure(figsize=(20,5))
+plt.plot(Y_plot['ds'], Y_plot['y'], label='True')
+plt.plot(Y_plot['ds'], Y_plot['Informer'], label='Informer')
+plt.plot(Y_plot['ds'], Y_plot['Autoformer'], label='Autoformer')
+plt.plot(Y_plot['ds'], Y_plot['PatchTST'], label='PatchTST')
+plt.xlabel('Datestamp')
+plt.ylabel('OT')
+plt.grid()
+plt.legend()
+```
+
+
+
+Finally, we compute the test errors using the Mean Absolute Error (MAE):
+
+$\qquad MAE = \frac{1}{Windows * Horizon} \sum_{\tau} |y_{\tau} - \hat{y}_{\tau}| \qquad$
+
+
+```python
+from neuralforecast.losses.numpy import mae
+```
+
+
+```python
+mae_informer = mae(Y_hat_df['y'], Y_hat_df['Informer'])
+mae_autoformer = mae(Y_hat_df['y'], Y_hat_df['Autoformer'])
+mae_patchtst = mae(Y_hat_df['y'], Y_hat_df['PatchTST'])
+
+print(f'Informer: {mae_informer:.3f}')
+print(f'Autoformer: {mae_autoformer:.3f}')
+print(f'PatchTST: {mae_patchtst:.3f}')
+```
+
+``` text
+Informer: 0.339
+Autoformer: 0.316
+PatchTST: 0.251
+```
+
+For reference, we can check the performance when compared to
+self-reported performance in their respective papers.
+
+| Horizon | PatchTST | AutoFormer | Informer | ARIMA |
+|---------|-----------|------------|----------|-------|
+| 96 | **0.256** | 0.339 | 0.453 | 0.301 |
+| 192 | 0.296 | 0.340 | 0.563 | 0.345 |
+| 336 | 0.329 | 0.372 | 0.887 | 0.386 |
+| 720 | 0.385 | 0.419 | 1.388 | 0.445 |
+
+## Next steps
+
+We proposed an alternative model for long-horizon forecasting, the
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits),
+based on feed-forward networks in (Challu et al. 2023). It achieves on
+par performance with
+[`PatchTST`](https://nixtlaverse.nixtla.io/neuralforecast/models.patchtst.html#patchtst),
+with a fraction of the computational cost. The
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+tutorial is available
+[here](https://nixtlaverse.nixtla.io/neuralforecast/docs/tutorials/longhorizon_nhits.html).
+
+## References
+
+[Zhou, H., Zhang, S., Peng, J., Zhang, S., Li, J., Xiong, H., & Zhang,
+W. (2021, May). Informer: Beyond efficient transformer for long sequence
+time-series forecasting. In Proceedings of the AAAI conference on
+artificial intelligence (Vol. 35, No. 12,
+pp. 11106-11115)](https://ojs.aaai.org/index.php/AAAI/article/view/17325)
+
+[Wu, H., Xu, J., Wang, J., & Long, M. (2021). Autoformer: Decomposition
+transformers with auto-correlation for long-term series forecasting.
+Advances in Neural Information Processing Systems, 34,
+22419-22430.](https://proceedings.neurips.cc/paper/2021/hash/bcc0d400288793e8bdcd7c19a8ac0c2b-Abstract.html)
+
+[Zhou, T., Ma, Z., Wen, Q., Wang, X., Sun, L., & Jin, R. (2022, June).
+Fedformer: Frequency enhanced decomposed transformer for long-term
+series forecasting. In International Conference on Machine Learning
+(pp. 27268-27286).
+PMLR.](https://proceedings.mlr.press/v162/zhou22g.html)
+
+[Nie, Y., Nguyen, N. H., Sinthong, P., & Kalagnanam, J. (2022). A Time
+Series is Worth 64 Words: Long-term Forecasting with
+Transformers.](https://arxiv.org/pdf/2211.14730.pdf)
+
+[Cristian Challu, Kin G. Olivares, Boris N. Oreshkin, Federico Garza,
+Max Mergenthaler-Canseco, Artur Dubrawski (2021). NHITS: Neural
+Hierarchical Interpolation for Time Series Forecasting. Accepted at AAAI
+2023.](https://arxiv.org/abs/2201.12886)
+
diff --git a/neuralforecast/docs/tutorials/multivariate_tsmixer.html.mdx b/neuralforecast/docs/tutorials/multivariate_tsmixer.html.mdx
new file mode 100644
index 00000000..7f7c0973
--- /dev/null
+++ b/neuralforecast/docs/tutorials/multivariate_tsmixer.html.mdx
@@ -0,0 +1,549 @@
+---
+description: Tutorial on how to do multivariate forecasting using TSMixer models.
+output-file: multivariate_tsmixer.html
+title: Multivariate Forecasting with TSMixer
+---
+
+
+In *multivariate* forecasting, we use the information from every time
+series to produce all forecasts for all time series jointly. In
+contrast, in *univariate* forecasting we only consider the information
+from every individual time series and produce forecasts for every time
+series separately. Multivariate forecasting methods thus use more
+information to produce every forecast, and thus should be able to
+provide better forecasting results. However, multivariate forecasting
+methods also scale with the number of time series, which means these
+methods are commonly less well suited for large-scale problems
+(i.e. forecasting many, many time series).
+
+In this notebook, we will demonstrate the performance of a
+state-of-the-art multivariate forecasting architecture
+[`TSMixer`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixer.html#tsmixer)
+/
+[`TSMixerx`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixerx.html#tsmixerx)
+when compared to a univariate forecasting method
+([`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits))
+and a simple MLP-based multivariate method
+([`MLPMultivariate`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlpmultivariate.html#mlpmultivariate)).
+
+We will show how to: \* Load the
+[ETTm2](https://github.com/zhouhaoyi/ETDataset) benchmark dataset, used
+in the academic literature. \* Train a
+[`TSMixer`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixer.html#tsmixer),
+[`TSMixerx`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixerx.html#tsmixerx)
+and
+[`MLPMultivariate`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlpmultivariate.html#mlpmultivariate)
+model \* Forecast the test set \* Optimize the hyperparameters
+
+You can run these experiments using GPU with Google Colab.
+
+
+
+## 1. Installing libraries
+
+
+```python
+!pip install neuralforecast datasetsforecast
+```
+
+## 2. Load ETTm2 Data
+
+The `LongHorizon` class will automatically download the complete ETTm2
+dataset and process it.
+
+It return three Dataframes: `Y_df` contains the values for the target
+variables, `X_df` contains exogenous calendar features and `S_df`
+contains static features for each time-series (none for ETTm2). For this
+example we will use `Y_df` and `X_df`.
+
+In
+[`TSMixerx`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixerx.html#tsmixerx),
+we can make use of the additional exogenous features contained in
+`X_df`. In
+[`TSMixer`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixer.html#tsmixer),
+there is *no* support for exogenous features. Hence, if you want to use
+exogenous features, you should use
+[`TSMixerx`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixerx.html#tsmixerx).
+
+If you want to use your own data just replace `Y_df` and `X_df`. Be sure
+to use a long format and make sure to have a similar structure as our
+data set.
+
+
+```python
+import pandas as pd
+
+from datasetsforecast.long_horizon import LongHorizon
+```
+
+
+```python
+# Change this to your own data to try the model
+Y_df, X_df, _ = LongHorizon.load(directory='./', group='ETTm2')
+Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+
+# X_df contains the exogenous features, which we add to Y_df
+X_df['ds'] = pd.to_datetime(X_df['ds'])
+Y_df = Y_df.merge(X_df, on=['unique_id', 'ds'], how='left')
+
+# We make validation and test splits
+n_time = len(Y_df.ds.unique())
+val_size = int(.2 * n_time)
+test_size = int(.2 * n_time)
+```
+
+
+```python
+Y_df
+```
+
+| | unique_id | ds | y | ex_1 | ex_2 | ex_3 | ex_4 |
+|----|----|----|----|----|----|----|----|
+| 0 | HUFL | 2016-07-01 00:00:00 | -0.041413 | -0.500000 | 0.166667 | -0.500000 | -0.001370 |
+| 1 | HUFL | 2016-07-01 00:15:00 | -0.185467 | -0.500000 | 0.166667 | -0.500000 | -0.001370 |
+| 2 | HUFL | 2016-07-01 00:30:00 | -0.257495 | -0.500000 | 0.166667 | -0.500000 | -0.001370 |
+| 3 | HUFL | 2016-07-01 00:45:00 | -0.577510 | -0.500000 | 0.166667 | -0.500000 | -0.001370 |
+| 4 | HUFL | 2016-07-01 01:00:00 | -0.385501 | -0.456522 | 0.166667 | -0.500000 | -0.001370 |
+| ... | ... | ... | ... | ... | ... | ... | ... |
+| 403195 | OT | 2018-02-20 22:45:00 | -1.581325 | 0.456522 | -0.333333 | 0.133333 | -0.363014 |
+| 403196 | OT | 2018-02-20 23:00:00 | -1.581325 | 0.500000 | -0.333333 | 0.133333 | -0.363014 |
+| 403197 | OT | 2018-02-20 23:15:00 | -1.581325 | 0.500000 | -0.333333 | 0.133333 | -0.363014 |
+| 403198 | OT | 2018-02-20 23:30:00 | -1.562328 | 0.500000 | -0.333333 | 0.133333 | -0.363014 |
+| 403199 | OT | 2018-02-20 23:45:00 | -1.562328 | 0.500000 | -0.333333 | 0.133333 | -0.363014 |
+
+## 3. Train models
+
+We will train models using the `cross_validation` method, which allows
+users to automatically simulate multiple historic forecasts (in the test
+set).
+
+The `cross_validation` method will use the validation set for
+hyperparameter selection and early stopping, and will then produce the
+forecasts for the test set.
+
+First, instantiate each model in the `models` list, specifying the
+`horizon`, `input_size`, and training iterations. In this notebook, we
+compare against the univariate
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+and multivariate
+[`MLPMultivariate`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlpmultivariate.html#mlpmultivariate)
+models.
+
+
+```python
+import logging
+
+import torch
+from neuralforecast.core import NeuralForecast
+from neuralforecast.models import TSMixer, TSMixerx, NHITS, MLPMultivariate
+from neuralforecast.losses.pytorch import MAE
+```
+
+
+```python
+logging.getLogger('pytorch_lightning').setLevel(logging.ERROR)
+torch.set_float32_matmul_precision('high')
+```
+
+
+```python
+horizon = 96
+input_size = 512
+models = [
+ TSMixer(h=horizon,
+ input_size=input_size,
+ n_series=7,
+ max_steps=1000,
+ val_check_steps=100,
+ early_stop_patience_steps=5,
+ scaler_type='identity',
+ valid_loss=MAE(),
+ random_seed=12345678,
+ ),
+ TSMixerx(h=horizon,
+ input_size=input_size,
+ n_series=7,
+ max_steps=1000,
+ val_check_steps=100,
+ early_stop_patience_steps=5,
+ scaler_type='identity',
+ dropout=0.7,
+ valid_loss=MAE(),
+ random_seed=12345678,
+ futr_exog_list=['ex_1', 'ex_2', 'ex_3', 'ex_4'],
+ ),
+ MLPMultivariate(h=horizon,
+ input_size=input_size,
+ n_series=7,
+ max_steps=1000,
+ val_check_steps=100,
+ early_stop_patience_steps=5,
+ scaler_type='standard',
+ hidden_size=256,
+ valid_loss=MAE(),
+ random_seed=12345678,
+ ),
+ NHITS(h=horizon,
+ input_size=horizon,
+ max_steps=1000,
+ val_check_steps=100,
+ early_stop_patience_steps=5,
+ scaler_type='robust',
+ valid_loss=MAE(),
+ random_seed=12345678,
+ ),
+ ]
+```
+
+> **Tip**
+>
+> Check our `auto` models for automatic hyperparameter optimization, and
+> see the end of this tutorial for an example of hyperparameter tuning.
+
+Instantiate a
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+object with the following required parameters:
+
+- `models`: a list of models.
+
+- `freq`: a string indicating the frequency of the data. (See [panda’s
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+Second, use the `cross_validation` method, specifying the dataset
+(`Y_df`), validation size and test size.
+
+
+```python
+nf = NeuralForecast(
+ models=models,
+ freq='15min',
+)
+
+Y_hat_df = nf.cross_validation(
+ df=Y_df,
+ val_size=val_size,
+ test_size=test_size,
+ n_windows=None,
+)
+```
+
+The `cross_validation` method will return the forecasts for each model
+on the test set.
+
+## 4. Evaluate Results
+
+Next, we plot the forecasts on the test set for the `OT` variable for
+all models.
+
+
+```python
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+cutoffs = Y_hat_df['cutoff'].unique()[::horizon]
+Y_plot = Y_hat_df[Y_hat_df['cutoff'].isin(cutoffs)].drop(columns='cutoff')
+plot_series(forecasts_df=Y_plot, ids=['OT'])
+```
+
+
+
+Finally, we compute the test errors using the Mean Absolute Error (MAE)
+and Mean Squared Error (MSE):
+
+$\qquad MAE = \frac{1}{Windows * Horizon} \sum_{\tau} |y_{\tau} - \hat{y}_{\tau}| \qquad$
+and
+$\qquad MSE = \frac{1}{Windows * Horizon} \sum_{\tau} (y_{\tau} - \hat{y}_{\tau})^{2} \qquad$
+
+
+```python
+from utilsforecast.evaluation import evaluate
+from utilsforecast.losses import mae, mse
+```
+
+
+```python
+evaluate(Y_hat_df.drop(columns='cutoff'), metrics=[mae, mse], agg_fn='mean')
+```
+
+| | metric | TSMixer | TSMixerx | MLPMultivariate | NHITS |
+|-----|--------|----------|----------|-----------------|----------|
+| 0 | mae | 0.245435 | 0.249727 | 0.263579 | 0.251008 |
+| 1 | mse | 0.162566 | 0.163098 | 0.176594 | 0.178864 |
+
+For reference, we can check the performance when compared to
+self-reported performance in the paper. We find that
+[`TSMixer`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixer.html#tsmixer)
+provides better results than the *univariate* method
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits).
+Also, our implementation of
+[`TSMixer`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixer.html#tsmixer)
+very closely tracks the results of the original paper. Finally, it seems
+that there is little benefit of using the additional exogenous variables
+contained in the dataframe `X_df` as
+[`TSMixerx`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixerx.html#tsmixerx)
+performs worse than
+[`TSMixer`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixer.html#tsmixer),
+especially on longer horizons. Note also that
+[`MLPMultivariate`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlpmultivariate.html#mlpmultivariate)
+clearly underperforms as compared to the other methods, which can be
+somewhat expected given its relative simplicity.
+
+Mean Absolute Error (MAE)
+
+| Horizon | TSMixer (this notebook) | TSMixer (paper) | TSMixerx (this notebook) | NHITS (this notebook) | NHITS (paper) | MLPMultivariate (this notebook) |
+|-----------|-----------|-----------|-----------|-----------|-----------|-----------|
+| 96 | **0.245** | 0.252 | 0.250 | 0.251 | 0.251 | 0.263 |
+| 192 | **0.288** | 0.290 | 0.300 | 0.291 | 0.305 | 0.361 |
+| 336 | **0.323** | 0.324 | 0.380 | 0.344 | 0.346 | 0.390 |
+| 720 | **0.377** | 0.422 | 0.464 | 0.417 | 0.413 | 0.608 |
+
+Mean Squared Error (MSE)
+
+| Horizon | TSMixer (this notebook) | TSMixer (paper) | TSMixerx (this notebook) | NHITS (this notebook) | NHITS (paper) | MLPMultivariate (this notebook) |
+|-----------|-----------|-----------|-----------|-----------|-----------|-----------|
+| 96 | **0.163** | **0.163** | 0.163 | 0.179 | 0.179 | 0.177 |
+| 192 | 0.220 | **0.216** | 0.231 | 0.239 | 0.245 | 0.330 |
+| 336 | 0.272 | **0.268** | 0.361 | 0.311 | 0.295 | 0.376 |
+| 720 | **0.356** | 0.420 | 0.493 | 0.451 | 0.401 | 3.421 |
+
+Note that for the table above, we use the same hyperparameters for all
+methods for all horizons, whereas the original papers tune the
+hyperparameters for each horizon.
+
+## 5. Tuning the hyperparameters
+
+The
+[`AutoTSMixer`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotsmixer)
+/
+[`AutoTSMixerx`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotsmixerx)
+class will automatically perform hyperparamter tunning using the [Tune
+library](https://docs.ray.io/en/latest/tune/index.html), exploring a
+user-defined or default search space. Models are selected based on the
+error on a validation set and the best model is then stored and used
+during inference.
+
+The `AutoTSMixer.default_config` / `AutoTSMixerx.default_config`
+attribute contains a suggested hyperparameter space. Here, we specify a
+different search space following the paper’s hyperparameters. Feel free
+to play around with this space.
+
+For this example, we will optimize the hyperparameters for
+`horizon = 96`.
+
+
+```python
+from ray import tune
+from ray.tune.search.hyperopt import HyperOptSearch
+from neuralforecast.auto import AutoTSMixer, AutoTSMixerx
+```
+
+
+```python
+horizon = 96 # 24hrs = 4 * 15 min.
+
+tsmixer_config = {
+ "input_size": input_size, # Size of input window
+ "max_steps": tune.choice([500, 1000, 2000]), # Number of training iterations
+ "val_check_steps": 100, # Compute validation every x steps
+ "early_stop_patience_steps": 5, # Early stopping steps
+ "learning_rate": tune.loguniform(1e-4, 1e-2), # Initial Learning rate
+ "n_block": tune.choice([1, 2, 4, 6, 8]), # Number of mixing layers
+ "dropout": tune.uniform(0.0, 0.99), # Dropout
+ "ff_dim": tune.choice([32, 64, 128]), # Dimension of the feature linear layer
+ "scaler_type": 'identity',
+ }
+
+tsmixerx_config = tsmixer_config.copy()
+tsmixerx_config['futr_exog_list'] = ['ex_1', 'ex_2', 'ex_3', 'ex_4']
+```
+
+To instantiate
+[`AutoTSMixer`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotsmixer)
+and
+[`AutoTSMixerx`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotsmixerx)
+you need to define:
+
+- `h`: forecasting horizon
+- `n_series`: number of time series in the multivariate time series
+ problem.
+
+In addition, we define the following parameters (if these are not given,
+the
+[`AutoTSMixer`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotsmixer)/[`AutoTSMixerx`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotsmixerx)
+class will use a pre-defined value): \* `loss`: training loss. Use the
+[`DistributionLoss`](https://nixtlaverse.nixtla.io/neuralforecast/losses.pytorch.html#distributionloss)
+to produce probabilistic forecasts. \* `config`: hyperparameter search
+space. If `None`, the
+[`AutoTSMixer`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotsmixer)
+class will use a pre-defined suggested hyperparameter space. \*
+`num_samples`: number of configurations explored. For this example, we
+only use a limited amount of `10`. \* `search_alg`: type of search
+algorithm used for selecting parameter values within the hyperparameter
+space. \* `backend`: the backend used for the hyperparameter
+optimization search, either `ray` or `optuna`. \* `valid_loss`: the loss
+used for the validation sets in the optimization procedure.
+
+
+```python
+model = AutoTSMixer(h=horizon,
+ n_series=7,
+ loss=MAE(),
+ config=tsmixer_config,
+ num_samples=10,
+ search_alg=HyperOptSearch(),
+ backend='ray',
+ valid_loss=MAE())
+
+modelx = AutoTSMixerx(h=horizon,
+ n_series=7,
+ loss=MAE(),
+ config=tsmixerx_config,
+ num_samples=10,
+ search_alg=HyperOptSearch(),
+ backend='ray',
+ valid_loss=MAE())
+```
+
+Now, we fit the model by instantiating a
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+object with the following required parameters:
+
+- `models`: a list of models.
+
+- `freq`: a string indicating the frequency of the data. (See [panda’s
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+The `cross_validation` method allows you to simulate multiple historic
+forecasts, greatly simplifying pipelines by replacing for loops with
+`fit` and `predict` methods.
+
+With time series data, cross validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The `cross_validation` method will use the validation set for
+hyperparameter selection, and will then produce the forecasts for the
+test set.
+
+
+```python
+nf = NeuralForecast(models=[model, modelx], freq='15min')
+Y_hat_df = nf.cross_validation(df=Y_df, val_size=val_size,
+ test_size=test_size, n_windows=None)
+```
+
+## 6. Evaluate Results
+
+The
+[`AutoTSMixer`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotsmixer)/[`AutoTSMixerx`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotsmixerx)
+class contains a `results` attribute that stores information of each
+configuration explored. It contains the validation loss and best
+validation hyperparameter. The result dataframe `Y_hat_df` that we
+obtained in the previous step is based on the best config of the
+hyperparameter search. For
+[`AutoTSMixer`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotsmixer),
+the best config is:
+
+
+```python
+nf.models[0].results.get_best_result().config
+```
+
+``` text
+{'input_size': 512,
+ 'max_steps': 2000,
+ 'val_check_steps': 100,
+ 'early_stop_patience_steps': 5,
+ 'learning_rate': 0.00034884229033995355,
+ 'n_block': 4,
+ 'dropout': 0.7592667651473878,
+ 'ff_dim': 128,
+ 'scaler_type': 'identity',
+ 'n_series': 7,
+ 'h': 96,
+ 'loss': MAE(),
+ 'valid_loss': MAE()}
+```
+
+and for
+[`AutoTSMixerx`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autotsmixerx):
+
+
+```python
+nf.models[1].results.get_best_result().config
+```
+
+``` text
+{'input_size': 512,
+ 'max_steps': 2000,
+ 'val_check_steps': 100,
+ 'early_stop_patience_steps': 5,
+ 'learning_rate': 0.00019039338576148522,
+ 'n_block': 6,
+ 'dropout': 0.5902743834953548,
+ 'ff_dim': 128,
+ 'scaler_type': 'identity',
+ 'futr_exog_list': ('ex_1', 'ex_2', 'ex_3', 'ex_4'),
+ 'n_series': 7,
+ 'h': 96,
+ 'loss': MAE(),
+ 'valid_loss': MAE()}
+```
+
+We compute the test errors of the best config for the two metrics of
+interest:
+
+$\qquad MAE = \frac{1}{Windows * Horizon} \sum_{\tau} |y_{\tau} - \hat{y}_{\tau}| \qquad$
+and
+$\qquad MSE = \frac{1}{Windows * Horizon} \sum_{\tau} (y_{\tau} - \hat{y}_{\tau})^{2} \qquad$
+
+
+```python
+evaluate(Y_hat_df.drop(columns='cutoff'), metrics=[mae, mse], agg_fn='mean')
+```
+
+| | metric | AutoTSMixer | AutoTSMixerx |
+|-----|--------|-------------|--------------|
+| 0 | mae | 0.243749 | 0.251972 |
+| 1 | mse | 0.162212 | 0.164347 |
+
+We can compare the error metrics for our optimized setting to the
+earlier setting in which we used the default hyperparameters. In this
+case, for a horizon of 96, we got slightly improved results for
+[`TSMixer`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixer.html#tsmixer)
+on
+[`MAE`](https://nixtlaverse.nixtla.io/neuralforecast/losses.pytorch.html#mae).
+Interestingly, we did not improve for
+[`TSMixerx`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixerx.html#tsmixerx)
+as compared to the default settings. For this dataset, it seems there is
+limited value in using exogenous features with the
+[`TSMixerx`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixerx.html#tsmixerx)
+architecture for a horizon of 96.
+
+| Metric | TSMixer (optimized) | TSMixer (default) | TSMixer (paper) | TSMixerx (optimized) | TSMixerx (default) |
+|------------|------------|------------|------------|------------|------------|
+| MAE | **0.244** | 0.245 | 0.252 | 0.252 | 0.250 |
+| MSE | **0.162** | 0.163 | 0.163 | 0.164 | 0.163 |
+
+Note that we only evaluated 10 hyperparameter configurations
+(`num_samples=10`), which may suggest that it is possible to further
+improve forecasting performance by exploring more hyperparameter
+configurations.
+
+## References
+
+[Chen, Si-An, Chun-Liang Li, Nate Yoder, Sercan O. Arik, and Tomas
+Pfister (2023). “TSMixer: An All-MLP Architecture for Time Series
+Forecasting.”](http://arxiv.org/abs/2303.06053) [Cristian Challu,
+Kin G. Olivares, Boris N. Oreshkin, Federico Garza, Max
+Mergenthaler-Canseco, Artur Dubrawski (2021). NHITS: Neural Hierarchical
+Interpolation for Time Series Forecasting. Accepted at AAAI
+2023.](https://arxiv.org/abs/2201.12886)
+
diff --git a/neuralforecast/docs/tutorials/robust_forecasting.html.mdx b/neuralforecast/docs/tutorials/robust_forecasting.html.mdx
new file mode 100644
index 00000000..4851dfd1
--- /dev/null
+++ b/neuralforecast/docs/tutorials/robust_forecasting.html.mdx
@@ -0,0 +1,290 @@
+---
+output-file: robust_forecasting.html
+title: Robust Forecasting
+---
+
+
+When outliers are present in a dataset, they can disrupt the calculated
+summary statistics, such as the mean and standard deviation, leading the
+model to favor the outlier values and deviate from most observations.
+Consequently, models need help in achieving a balance between accurately
+accommodating outliers and performing well on normal data, resulting in
+improved overall performance on both types of data. [Robust regression
+algorithms](https://en.wikipedia.org/wiki/Robust_regression) tackle this
+issue, explicitly accounting for outliers in the dataset.
+
+In this notebook we will show how to fit robust NeuralForecast methods.
+We will: - Installing NeuralForecast. - Loading Noisy
+AirPassengers. - Fit and predict robustified NeuralForecast. -
+Plot and evaluate predictions.
+
+You can run these experiments using GPU with Google Colab.
+
+
+
+## 1. Installing NeuralForecast
+
+
+```python
+!pip install neuralforecast
+```
+
+
+```python
+import logging
+import numpy as np
+import pandas as pd
+
+import matplotlib.pyplot as plt
+from random import random
+from random import randint
+from random import seed
+
+from neuralforecast import NeuralForecast
+from neuralforecast.utils import AirPassengersDF
+
+from neuralforecast.models import NHITS
+from neuralforecast.losses.pytorch import MQLoss, DistributionLoss, HuberMQLoss
+
+from utilsforecast.losses import mape, mqloss
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+logging.getLogger("pytorch_lightning").setLevel(logging.ERROR)
+```
+
+## 2. Loading Noisy AirPassengers
+
+For this example we will use the classic Box-Cox AirPassengers dataset
+that we will augment it by introducing outliers.
+
+In particular, we will focus on introducing outliers to the target
+variable altering it to deviate from its original observation by a
+specified factor, such as 2-to-4 times the standard deviation.
+
+
+```python
+# Original Box-Cox AirPassengers
+# as defined in neuralforecast.utils
+Y_df = AirPassengersDF.copy()
+plt.plot(Y_df.y)
+plt.ylabel('Monthly Passengers')
+plt.xlabel('Timestamp [t]')
+plt.grid()
+```
+
+
+
+
+```python
+# Here we add some artificial outliers to AirPassengers
+seed(1)
+for i in range(len(Y_df)):
+ factor = randint(2, 4)
+ if random() > 0.97:
+ Y_df.loc[i, "y"] += factor * Y_df["y"].std()
+
+plt.plot(Y_df.y)
+plt.ylabel('Monthly Passengers + Noise')
+plt.xlabel('Timestamp [t]')
+plt.grid()
+```
+
+
+
+
+```python
+# Split datasets into train/test
+# Last 12 months for test
+Y_train_df = Y_df.groupby('unique_id').head(-12)
+Y_test_df = Y_df.groupby('unique_id').tail(12)
+Y_test_df
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|-------|
+| 132 | 1.0 | 1960-01-31 | 417.0 |
+| 133 | 1.0 | 1960-02-29 | 391.0 |
+| 134 | 1.0 | 1960-03-31 | 419.0 |
+| 135 | 1.0 | 1960-04-30 | 461.0 |
+| 136 | 1.0 | 1960-05-31 | 472.0 |
+| 137 | 1.0 | 1960-06-30 | 535.0 |
+| 138 | 1.0 | 1960-07-31 | 622.0 |
+| 139 | 1.0 | 1960-08-31 | 606.0 |
+| 140 | 1.0 | 1960-09-30 | 508.0 |
+| 141 | 1.0 | 1960-10-31 | 461.0 |
+| 142 | 1.0 | 1960-11-30 | 390.0 |
+| 143 | 1.0 | 1960-12-31 | 432.0 |
+
+## 3. Fit and predict robustified NeuralForecast
+
+### Huber MQ Loss
+
+The Huber loss, employed in robust regression, is a loss function that
+exhibits reduced sensitivity to outliers in data when compared to the
+squared error loss. The Huber loss function is quadratic for small
+errors and linear for large errors. Here we will use a slight
+modification for probabilistic predictions. Feel free to play with the
+$\delta$ parameter.
+
+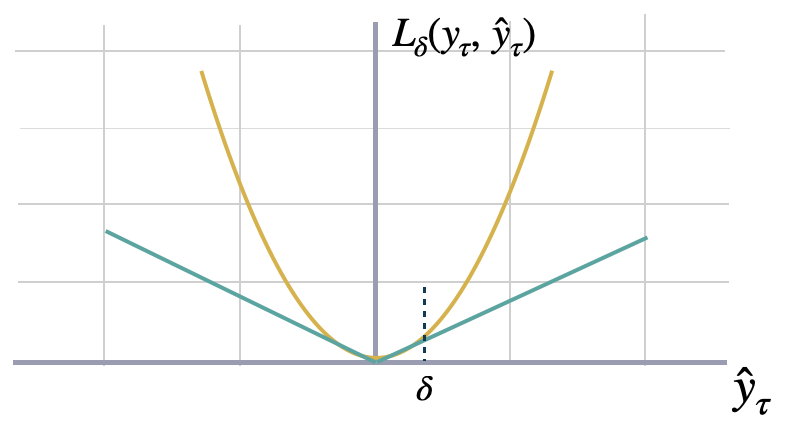
+
+### Dropout Regularization
+
+The dropout technique is a regularization method used in neural networks
+to prevent overfitting. During training, dropout randomly sets a
+fraction of the input units or neurons in a layer to zero at each
+update, effectively “dropping out” those units. This means that the
+network cannot rely on any individual unit because it may be dropped out
+at any time. By doing so, dropout forces the network to learn more
+robust and generalizable representations by preventing units from
+co-adapting too much.
+
+The dropout method, can help us to robustify the network to outliers in
+the auto-regressive features. You can explore it through the
+`dropout_prob_theta` parameter.
+
+### Fit NeuralForecast models
+
+Using the
+[`NeuralForecast.fit`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast.fit)
+method you can train a set of models to your dataset. You can define the
+forecasting `horizon` (12 in this example), and modify the
+hyperparameters of the model. For example, for the
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+we changed the default hidden size for both encoder and decoders.
+
+See the
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+and
+[`MLP`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlp.html#mlp)
+[model
+documentation](https://nixtla.github.io/neuralforecast/models.mlp.html).
+
+
+```python
+horizon = 12
+level = [50, 80]
+
+# Try different hyperparmeters to improve accuracy.
+models = [NHITS(h=horizon, # Forecast horizon
+ input_size=2 * horizon, # Length of input sequence
+ loss=HuberMQLoss(level=level), # Robust Huber Loss
+ valid_loss=MQLoss(level=level), # Validation signal
+ max_steps=500, # Number of steps to train
+ dropout_prob_theta=0.6, # Dropout to robustify vs outlier lag inputs
+ #early_stop_patience_steps=2, # Early stopping regularization patience
+ val_check_steps=10, # Frequency of validation signal (affects early stopping)
+ alias='Huber',
+ ),
+ NHITS(h=horizon,
+ input_size=2 * horizon,
+ loss=DistributionLoss(distribution='Normal',
+ level=level), # Classic Normal distribution
+ valid_loss=MQLoss(level=level),
+ max_steps=500,
+ #early_stop_patience_steps=2,
+ dropout_prob_theta=0.6,
+ val_check_steps=10,
+ alias='Normal',
+ )
+ ]
+nf = NeuralForecast(models=models, freq='M')
+nf.fit(df=Y_train_df)
+Y_hat_df = nf.predict()
+```
+
+
+```python
+# By default NeuralForecast produces forecast intervals
+# In this case the lo-x and high-x levels represent the
+# low and high bounds of the prediction accumulating x% probability
+Y_hat_df
+```
+
+| | unique_id | ds | Huber-median | Huber-lo-80 | Huber-lo-50 | Huber-hi-50 | Huber-hi-80 | Normal | Normal-median | Normal-lo-80 | Normal-lo-50 | Normal-hi-50 | Normal-hi-80 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | 1.0 | 1960-01-31 | 412.738525 | 401.058044 | 406.131958 | 420.779266 | 432.124268 | 406.459717 | 416.787842 | -124.278656 | 135.413223 | 680.997070 | 904.871765 |
+| 1 | 1.0 | 1960-02-29 | 403.913544 | 384.403534 | 391.904419 | 420.288208 | 469.040375 | 399.827148 | 418.305725 | -137.291870 | 103.988327 | 661.940430 | 946.699219 |
+| 2 | 1.0 | 1960-03-31 | 472.311523 | 446.644531 | 460.767334 | 486.710999 | 512.552979 | 380.263947 | 378.253998 | -105.411003 | 117.415565 | 647.887695 | 883.611633 |
+| 3 | 1.0 | 1960-04-30 | 460.996674 | 444.471039 | 452.971802 | 467.544189 | 480.843903 | 432.131378 | 442.395844 | -104.205200 | 135.457123 | 729.306885 | 974.661743 |
+| 4 | 1.0 | 1960-05-31 | 465.534790 | 452.048889 | 457.472626 | 476.141022 | 490.311005 | 417.186279 | 417.956543 | -117.399597 | 150.915833 | 692.936523 | 930.934814 |
+| 5 | 1.0 | 1960-06-30 | 538.116028 | 518.049866 | 527.238159 | 551.501709 | 563.818848 | 444.510834 | 440.168396 | -54.501572 | 189.301392 | 703.502014 | 946.068909 |
+| 6 | 1.0 | 1960-07-31 | 613.937866 | 581.048035 | 597.368408 | 629.111450 | 645.550659 | 423.707275 | 431.251526 | -97.069489 | 164.821259 | 687.764526 | 942.432251 |
+| 7 | 1.0 | 1960-08-31 | 616.188660 | 581.982300 | 599.544128 | 632.137512 | 643.219543 | 386.655823 | 383.755157 | -134.702011 | 139.954285 | 658.973022 | 897.393494 |
+| 8 | 1.0 | 1960-09-30 | 537.559143 | 513.477478 | 526.664856 | 551.563293 | 573.146667 | 388.874817 | 379.827057 | -139.859344 | 110.772484 | 673.086182 | 926.355774 |
+| 9 | 1.0 | 1960-10-31 | 471.107605 | 449.207916 | 459.288025 | 486.402985 | 515.082458 | 401.483643 | 412.114990 | -185.928085 | 95.805717 | 703.490784 | 970.837830 |
+| 10 | 1.0 | 1960-11-30 | 412.758423 | 389.203308 | 398.727295 | 431.723602 | 451.208588 | 425.829895 | 425.018799 | -172.022018 | 108.840889 | 723.424011 | 1035.656128 |
+| 11 | 1.0 | 1960-12-31 | 457.254761 | 438.565582 | 446.097168 | 468.809296 | 483.967865 | 406.916595 | 399.852051 | -199.963684 | 110.715050 | 729.735107 | 951.728577 |
+
+## 4. Plot and Evaluate Predictions
+
+Finally, we plot the forecasts of both models againts the real values.
+
+And evaluate the accuracy of the `NHITS-Huber` and `NHITS-Normal`
+forecasters.
+
+
+```python
+fig, ax = plt.subplots(1, 1, figsize = (20, 7))
+plot_df = pd.concat([Y_train_df, Y_hat_df]).set_index('ds') # Concatenate the train and forecast dataframes
+plot_df[['y', 'Huber-median', 'Normal-median']].plot(ax=ax, linewidth=2)
+
+ax.set_title('Noisy AirPassengers Forecast', fontsize=22)
+ax.set_ylabel('Monthly Passengers', fontsize=20)
+ax.set_xlabel('Timestamp [t]', fontsize=20)
+ax.legend(prop={'size': 15})
+ax.grid()
+```
+
+
+
+To evaluate the median predictions we use the mean average percentage
+error (MAPE), defined as follows:
+
+$$\mathrm{MAPE}(\mathbf{y}_{\tau}, \hat{\mathbf{y}}_{\tau}) = \mathrm{mean}\left(\frac{|\mathbf{y}_{\tau}-\hat{\mathbf{y}}_{\tau}|}{|\mathbf{y}_{\tau}|}\right)$$
+
+To evaluate the coherent probabilistic predictions we use the Continuous
+Ranked Probability Score (CRPS), defined as follows:
+
+$$\mathrm{CRPS}(\hat{F}_{\tau},\mathbf{y}_{\tau}) = \int^{1}_{0} \mathrm{QL}(\hat{F}_{\tau}, y_{\tau})_{q} dq$$
+
+As you can see, robust regression improvements reflect in both the
+normal and probabilistic forecast setting.
+
+
+```python
+df_metrics = Y_hat_df.merge(Y_test_df, on=['ds', 'unique_id'])
+df_metrics.rename(columns={'Huber-median': 'Huber'}, inplace=True)
+
+metrics = evaluate(df_metrics,
+ metrics=[mape, mqloss],
+ models=['Huber', 'Normal'],
+ level = [50, 80],
+ agg_fn="mean")
+
+metrics
+```
+
+| | metric | Huber | Normal |
+|-----|--------|----------|-----------|
+| 0 | mape | 0.034726 | 0.140207 |
+| 1 | mqloss | 5.511535 | 61.891651 |
+
+## References
+
+- [Huber Peter, J (1964). “Robust Estimation of a Location Parameter”.
+ Annals of
+ Statistics.](https://projecteuclid.org/journals/annals-of-mathematical-statistics/volume-35/issue-1/Robust-Estimation-of-a-Location-Parameter/10.1214/aoms/1177703732.full)
+- [Nitish Srivastava, Geoffrey Hinton, Alex Krizhevsky, Ilya
+ Sutskever, Ruslan Salakhutdinov (2014).”Dropout: A Simple Way to
+ Prevent Neural Networks from Overfitting”. Journal of Machine
+ Learning
+ Research.](https://jmlr.org/papers/v15/srivastava14a.html)
+- [Cristian Challu, Kin G. Olivares, Boris N. Oreshkin, Federico
+ Garza, Max Mergenthaler-Canseco, Artur Dubrawski (2023). NHITS:
+ Neural Hierarchical Interpolation for Time Series Forecasting.
+ Accepted at AAAI 2023.](https://arxiv.org/abs/2201.12886)
+
diff --git a/neuralforecast/docs/tutorials/temporal_classification.html.mdx b/neuralforecast/docs/tutorials/temporal_classification.html.mdx
new file mode 100644
index 00000000..fc92a354
--- /dev/null
+++ b/neuralforecast/docs/tutorials/temporal_classification.html.mdx
@@ -0,0 +1,274 @@
+---
+output-file: temporal_classification.html
+title: Temporal Classification
+---
+
+
+A logistic regression analyzes the relationship between a binary target
+variable and its predictor variables to estimate the probability of the
+dependent variable taking the value 1. In the presence of temporal data
+where observations along time aren’t independent, the errors of the
+model will be correlated through time and incorporating autoregressive
+features or lags can capture temporal dependencies and enhance the
+predictive power of logistic regression.
+
+ NHITS’s inputs are static exogenous $\mathbf{x}^{(s)}$, historic
+exogenous $\mathbf{x}^{(h)}_{[:t]}$, exogenous available at the time of
+the prediction $\mathbf{x}^{(f)}_{[:t+H]}$ and autorregresive features
+$\mathbf{y}_{[:t]}$, each of these inputs is further decomposed into
+categorical and continuous. The network uses a multi-quantile regression
+to model the following conditional
+probability:$$\mathbb{P}(\mathbf{y}_{[t+1:t+H]}|\;\mathbf{y}_{[:t]},\; \mathbf{x}^{(h)}_{[:t]},\; \mathbf{x}^{(f)}_{[:t+H]},\; \mathbf{x}^{(s)})$$
+
+In this notebook we show how to fit NeuralForecast methods for binary
+sequences regression. We will: - Installing NeuralForecast. - Loading
+binary sequence data. - Fit and predict temporal classifiers. - Plot and
+evaluate predictions.
+
+You can run these experiments using GPU with Google Colab.
+
+
+
+## 1. Installing NeuralForecast
+
+
+```python
+!pip install neuralforecast
+```
+
+
+```python
+import numpy as np
+import pandas as pd
+from sklearn import datasets
+
+import matplotlib.pyplot as plt
+from neuralforecast import NeuralForecast
+from neuralforecast.models import MLP, NHITS, LSTM
+from neuralforecast.losses.pytorch import DistributionLoss, Accuracy
+```
+
+## 2. Loading Binary Sequence Data
+
+The `core.NeuralForecast` class contains shared, `fit`, `predict` and
+other methods that take as inputs pandas DataFrames with columns
+`['unique_id', 'ds', 'y']`, where `unique_id` identifies individual time
+series from the dataset, `ds` is the date, and `y` is the target binary
+variable.
+
+In this motivation example we convert 8x8 digits images into 64-length
+sequences and define a classification problem, to identify when the
+pixels surpass certain threshold. We declare a pandas dataframe in long
+format, to match NeuralForecast’s inputs.
+
+
+```python
+digits = datasets.load_digits()
+images = digits.images[:100]
+
+plt.imshow(images[0,:,:], cmap=plt.cm.gray,
+ vmax=16, interpolation="nearest")
+
+pixels = np.reshape(images, (len(images), 64))
+ytarget = (pixels > 10) * 1
+
+fig, ax1 = plt.subplots()
+ax2 = ax1.twinx()
+ax1.plot(pixels[10])
+ax2.plot(ytarget[10], color='purple')
+ax1.set_xlabel('Pixel index')
+ax1.set_ylabel('Pixel value')
+ax2.set_ylabel('Pixel threshold', color='purple')
+plt.grid()
+plt.show()
+```
+
+
+
+
+
+
+```python
+# We flat the images and create an input dataframe
+# with 'unique_id' series identifier and 'ds' time stamp identifier.
+Y_df = pd.DataFrame.from_dict({
+ 'unique_id': np.repeat(np.arange(100), 64),
+ 'ds': np.tile(np.arange(64)+1910, 100),
+ 'y': ytarget.flatten(), 'pixels': pixels.flatten()})
+Y_df
+```
+
+| | unique_id | ds | y | pixels |
+|------|-----------|------|-----|--------|
+| 0 | 0 | 1910 | 0 | 0.0 |
+| 1 | 0 | 1911 | 0 | 0.0 |
+| 2 | 0 | 1912 | 0 | 5.0 |
+| 3 | 0 | 1913 | 1 | 13.0 |
+| 4 | 0 | 1914 | 0 | 9.0 |
+| ... | ... | ... | ... | ... |
+| 6395 | 99 | 1969 | 1 | 14.0 |
+| 6396 | 99 | 1970 | 1 | 16.0 |
+| 6397 | 99 | 1971 | 0 | 3.0 |
+| 6398 | 99 | 1972 | 0 | 0.0 |
+| 6399 | 99 | 1973 | 0 | 0.0 |
+
+## 3. Fit and predict temporal classifiers
+
+### Fit the models
+
+Using the
+[`NeuralForecast.fit`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast.fit)
+method you can train a set of models to your dataset. You can define the
+forecasting `horizon` (12 in this example), and modify the
+hyperparameters of the model. For example, for the
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+we changed the default hidden size for both encoder and decoders.
+
+See the
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+and
+[`MLP`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlp.html#mlp)
+[model
+documentation](https://nixtla.github.io/neuralforecast/models.mlp.html).
+
+> **Warning**
+>
+> For the moment Recurrent-based model family is not available to
+> operate with Bernoulli distribution output. This affects the following
+> methods
+> [`LSTM`](https://nixtlaverse.nixtla.io/neuralforecast/models.lstm.html#lstm),
+> [`GRU`](https://nixtlaverse.nixtla.io/neuralforecast/models.gru.html#gru),
+> [`DilatedRNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.dilated_rnn.html#dilatedrnn),
+> and
+> [`TCN`](https://nixtlaverse.nixtla.io/neuralforecast/models.tcn.html#tcn).
+> This feature is work in progress.
+
+
+```python
+# %%capture
+horizon = 12
+
+# Try different hyperparmeters to improve accuracy.
+models = [MLP(h=horizon, # Forecast horizon
+ input_size=2 * horizon, # Length of input sequence
+ loss=DistributionLoss('Bernoulli'), # Binary classification loss
+ valid_loss=Accuracy(), # Accuracy validation signal
+ max_steps=500, # Number of steps to train
+ scaler_type='standard', # Type of scaler to normalize data
+ hidden_size=64, # Defines the size of the hidden state of the LSTM
+ #early_stop_patience_steps=2, # Early stopping regularization patience
+ val_check_steps=10, # Frequency of validation signal (affects early stopping)
+ ),
+ NHITS(h=horizon, # Forecast horizon
+ input_size=2 * horizon, # Length of input sequence
+ loss=DistributionLoss('Bernoulli'), # Binary classification loss
+ valid_loss=Accuracy(), # Accuracy validation signal
+ max_steps=500, # Number of steps to train
+ n_freq_downsample=[2, 1, 1], # Downsampling factors for each stack output
+ #early_stop_patience_steps=2, # Early stopping regularization patience
+ val_check_steps=10, # Frequency of validation signal (affects early stopping)
+ interpolation_mode="nearest",
+ )
+ ]
+nf = NeuralForecast(models=models, freq=1)
+Y_hat_df = nf.cross_validation(df=Y_df, n_windows=1)
+```
+
+
+```python
+# By default NeuralForecast produces forecast intervals
+# In this case the lo-x and high-x levels represent the
+# low and high bounds of the prediction accumulating x% probability
+Y_hat_df
+```
+
+| | unique_id | ds | cutoff | MLP | MLP-median | MLP-lo-90 | MLP-lo-80 | MLP-hi-80 | MLP-hi-90 | NHITS | NHITS-median | NHITS-lo-90 | NHITS-lo-80 | NHITS-hi-80 | NHITS-hi-90 | y |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | 0 | 1962 | 1961 | 0.173 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0.761 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0 |
+| 1 | 0 | 1963 | 1961 | 0.784 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0.571 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 1 |
+| 2 | 0 | 1964 | 1961 | 0.042 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.009 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 |
+| 3 | 0 | 1965 | 1961 | 0.072 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.054 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0 |
+| 4 | 0 | 1966 | 1961 | 0.059 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.000 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| 1195 | 99 | 1969 | 1961 | 0.551 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0.697 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 1 |
+| 1196 | 99 | 1970 | 1961 | 0.662 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0.465 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 | 1 |
+| 1197 | 99 | 1971 | 1961 | 0.369 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0.382 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0 |
+| 1198 | 99 | 1972 | 1961 | 0.056 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.000 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 |
+| 1199 | 99 | 1973 | 1961 | 0.000 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.000 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 |
+
+
+```python
+# Define classification threshold for final predictions
+# If (prob > threshold) -> 1
+Y_hat_df['NHITS'] = (Y_hat_df['NHITS'] > 0.5) * 1
+Y_hat_df['MLP'] = (Y_hat_df['MLP'] > 0.5) * 1
+Y_hat_df
+```
+
+| | unique_id | ds | cutoff | MLP | MLP-median | MLP-lo-90 | MLP-lo-80 | MLP-hi-80 | MLP-hi-90 | NHITS | NHITS-median | NHITS-lo-90 | NHITS-lo-80 | NHITS-hi-80 | NHITS-hi-90 | y |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | 0 | 1962 | 1961 | 0 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 | 1 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0 |
+| 1 | 0 | 1963 | 1961 | 1 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 1 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 1 |
+| 2 | 0 | 1964 | 1961 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 |
+| 3 | 0 | 1965 | 1961 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0 |
+| 4 | 0 | 1966 | 1961 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| 1195 | 99 | 1969 | 1961 | 1 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 1 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 1 |
+| 1196 | 99 | 1970 | 1961 | 1 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 | 1 |
+| 1197 | 99 | 1971 | 1961 | 0 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0 |
+| 1198 | 99 | 1972 | 1961 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 |
+| 1199 | 99 | 1973 | 1961 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 |
+
+## 4. Plot and Evaluate Predictions
+
+Finally, we plot the forecasts of both models againts the real values.
+And evaluate the accuracy of the
+[`MLP`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlp.html#mlp)
+and
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+temporal classifiers.
+
+
+```python
+plot_df = Y_hat_df[Y_hat_df.unique_id==10]
+
+fig, ax = plt.subplots(1, 1, figsize = (20, 7))
+plt.plot(plot_df.ds, plot_df.y, label='target signal')
+plt.plot(plot_df.ds, plot_df['MLP'] * 1.1, label='MLP prediction')
+plt.plot(plot_df.ds, plot_df['NHITS'] * .9, label='NHITS prediction')
+ax.set_title('Binary Sequence Forecast', fontsize=22)
+ax.set_ylabel('Pixel Threshold and Prediction', fontsize=20)
+ax.set_xlabel('Timestamp [t]', fontsize=20)
+ax.legend(prop={'size': 15})
+ax.grid()
+```
+
+
+
+
+```python
+def accuracy(y, y_hat):
+ return np.mean(y==y_hat)
+
+mlp_acc = accuracy(y=Y_hat_df['y'], y_hat=Y_hat_df['MLP'])
+nhits_acc = accuracy(y=Y_hat_df['y'], y_hat=Y_hat_df['NHITS'])
+
+print(f'MLP Accuracy: {mlp_acc:.1%}')
+print(f'NHITS Accuracy: {nhits_acc:.1%}')
+```
+
+``` text
+MLP Accuracy: 77.8%
+NHITS Accuracy: 74.5%
+```
+
+## References
+
+- [Cox D. R. (1958). “The Regression Analysis of Binary Sequences.”
+ Journal of the Royal Statistical Society B, 20(2),
+ 215–242.](https://arxiv.org/abs/2201.12886)
+- [Cristian Challu, Kin G. Olivares, Boris N. Oreshkin, Federico
+ Garza, Max Mergenthaler-Canseco, Artur Dubrawski (2023). NHITS:
+ Neural Hierarchical Interpolation for Time Series Forecasting.
+ Accepted at AAAI 2023.](https://arxiv.org/abs/2201.12886)
+
diff --git a/neuralforecast/docs/tutorials/transfer_learning.html.mdx b/neuralforecast/docs/tutorials/transfer_learning.html.mdx
new file mode 100644
index 00000000..2ffc6870
--- /dev/null
+++ b/neuralforecast/docs/tutorials/transfer_learning.html.mdx
@@ -0,0 +1,222 @@
+---
+output-file: transfer_learning.html
+title: Transfer Learning
+---
+
+
+Transfer learning refers to the process of pre-training a flexible model
+on a large dataset and using it later on other data with little to no
+training. It is one of the most outstanding 🚀 achievements in Machine
+Learning 🧠 and has many practical applications.
+
+For time series forecasting, the technique allows you to get
+lightning-fast predictions ⚡ bypassing the tradeoff between accuracy
+and speed (more than 30 times faster than our alreadsy fast
+[autoARIMA](https://github.com/Nixtla/statsforecast) for a similar
+accuracy).
+
+This notebook shows how to generate a pre-trained model and store it in
+a checkpoint to make it available to forecast new time series never seen
+by the model.
+
+Table of Contents 1. Installing NeuralForecast/DatasetsForecast
+2. Load M4 Data 3. Instantiate NeuralForecast core, Fit, and
+save 4. Load pre-trained model and predict on AirPassengers 5.
+Evaluate Results
+
+You can run these experiments using GPU with Google Colab.
+
+
+
+## 1. Installing Libraries
+
+
+```python
+!pip install datasetsforecast neuralforecast
+```
+
+
+```python
+import logging
+
+import numpy as np
+import pandas as pd
+import torch
+from datasetsforecast.m4 import M4
+from neuralforecast.core import NeuralForecast
+from neuralforecast.models import NHITS
+from neuralforecast.utils import AirPassengersDF
+from utilsforecast.losses import mae
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+logging.getLogger("pytorch_lightning").setLevel(logging.WARNING)
+```
+
+This example will automatically run on GPUs if available. **Make sure**
+cuda is available. (If you need help to put this into production send us
+an email or join or community, we also offer a fully hosted solution)
+
+
+```python
+torch.cuda.is_available()
+```
+
+``` text
+True
+```
+
+## 2. Load M4 Data
+
+The `M4` class will automatically download the complete M4 dataset and
+process it.
+
+It return three Dataframes: `Y_df` contains the values for the target
+variables, `X_df` contains exogenous calendar features and `S_df`
+contains static features for each time-series (none for M4). For this
+example we will only use `Y_df`.
+
+If you want to use your own data just replace `Y_df`. Be sure to use a
+long format and have a simmilar structure than our data set.
+
+
+```python
+Y_df, _, _ = M4.load(directory='./', group='Monthly', cache=True)
+Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+Y_df
+```
+
+| | unique_id | ds | y |
+|----------|-----------|-------------------------------|--------|
+| 0 | M1 | 1970-01-01 00:00:00.000000001 | 8000.0 |
+| 1 | M1 | 1970-01-01 00:00:00.000000002 | 8350.0 |
+| 2 | M1 | 1970-01-01 00:00:00.000000003 | 8570.0 |
+| 3 | M1 | 1970-01-01 00:00:00.000000004 | 7700.0 |
+| 4 | M1 | 1970-01-01 00:00:00.000000005 | 7080.0 |
+| ... | ... | ... | ... |
+| 11246406 | M9999 | 1970-01-01 00:00:00.000000083 | 4200.0 |
+| 11246407 | M9999 | 1970-01-01 00:00:00.000000084 | 4300.0 |
+| 11246408 | M9999 | 1970-01-01 00:00:00.000000085 | 3800.0 |
+| 11246409 | M9999 | 1970-01-01 00:00:00.000000086 | 4400.0 |
+| 11246410 | M9999 | 1970-01-01 00:00:00.000000087 | 4300.0 |
+
+## 3. Model Train and Save
+
+Using the
+[`NeuralForecast.fit`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast.fit)
+method you can train a set of models to your dataset. You just have to
+define the `input_size` and `horizon` of your model. The `input_size` is
+the number of historic observations (lags) that the model will use to
+learn to predict `h` steps in the future. Also, you can modify the
+hyperparameters of the model to get a better accuracy.
+
+
+```python
+horizon = 12
+stacks = 3
+models = [NHITS(input_size=5 * horizon,
+ h=horizon,
+ max_steps=100,
+ stack_types = stacks*['identity'],
+ n_blocks = stacks*[1],
+ mlp_units = [[256,256] for _ in range(stacks)],
+ n_pool_kernel_size = stacks*[1],
+ batch_size = 32,
+ scaler_type='standard',
+ n_freq_downsample=[12,4,1],
+ enable_progress_bar=False,
+ interpolation_mode="nearest",
+ )]
+nf = NeuralForecast(models=models, freq='ME')
+nf.fit(df=Y_df)
+```
+
+``` text
+INFO:lightning_fabric.utilities.seed:Seed set to 1
+```
+
+Save model with `core.NeuralForecast.save` method. This method uses
+PytorchLightning `save_checkpoint` function. We set `save_dataset=False`
+to only save the model.
+
+
+```python
+nf.save(path='./results/transfer/', model_index=None, overwrite=True, save_dataset=False)
+```
+
+## 4. Transfer M4 to AirPassengers
+
+We load the stored model with the `core.NeuralForecast.load` method, and
+forecast `AirPassenger` with the `core.NeuralForecast.predict` function.
+
+
+```python
+fcst2 = NeuralForecast.load(path='./results/transfer/')
+```
+
+``` text
+c:\Nixtla\Repositories\neuralforecast\neuralforecast\common\_base_model.py:133: UserWarning: NHITS is a univariate model. Parameter n_series is ignored.
+ warnings.warn(
+INFO:lightning_fabric.utilities.seed:Seed set to 1
+```
+
+
+```python
+# We define the train df.
+Y_df = AirPassengersDF.copy()
+mean = Y_df[Y_df.ds<='1959-12-31']['y'].mean()
+std = Y_df[Y_df.ds<='1959-12-31']['y'].std()
+
+Y_train_df = Y_df[Y_df.ds<='1959-12-31'] # 132 train
+Y_test_df = Y_df[Y_df.ds>'1959-12-31'] # 12 test
+```
+
+
+```python
+Y_hat_df = fcst2.predict(df=Y_train_df)
+Y_hat_df.head()
+```
+
+| | unique_id | ds | NHITS |
+|-----|-----------|------------|------------|
+| 0 | 1.0 | 1960-01-31 | 422.038757 |
+| 1 | 1.0 | 1960-02-29 | 424.678040 |
+| 2 | 1.0 | 1960-03-31 | 439.538879 |
+| 3 | 1.0 | 1960-04-30 | 447.967072 |
+| 4 | 1.0 | 1960-05-31 | 470.603333 |
+
+
+```python
+plot_series(Y_train_df, Y_hat_df)
+```
+
+
+
+## 5. Evaluate Results
+
+We evaluate the forecasts of the pre-trained model with the Mean
+Absolute Error
+([`mae`](https://nixtlaverse.nixtla.io/neuralforecast/losses.numpy.html#mae)).
+
+$$
+
+\qquad MAE = \frac{1}{Horizon} \sum_{\tau} |y_{\tau} - \hat{y}_{\tau}|\qquad
+
+$$
+
+
+```python
+fcst_mae = mae(Y_test_df.merge(Y_hat_df), models=['NHITS'])['NHITS'].item()
+print(f'NHITS MAE: {fcst_mae:.3f}')
+print('ETS MAE: 16.222')
+print('AutoARIMA MAE: 18.551')
+```
+
+``` text
+NHITS MAE: 17.245
+ETS MAE: 16.222
+AutoARIMA MAE: 18.551
+```
+
diff --git a/neuralforecast/docs/tutorials/uncertainty_quantification.html.mdx b/neuralforecast/docs/tutorials/uncertainty_quantification.html.mdx
new file mode 100644
index 00000000..945d1b0a
--- /dev/null
+++ b/neuralforecast/docs/tutorials/uncertainty_quantification.html.mdx
@@ -0,0 +1,226 @@
+---
+description: Quantify uncertainty
+output-file: uncertainty_quantification.html
+title: Probabilistic Forecasting
+---
+
+
+Probabilistic forecasting is a natural answer to quantify the
+uncertainty of target variable’s future. The task requires to model the
+following conditional predictive distribution:
+
+$$\mathbb{P}(\mathbf{y}_{t+1:t+H} \;|\; \mathbf{y}_{:t})$$
+
+We will show you how to tackle the task with
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+by combining a classic Long Short Term Memory Network
+[(LSTM)](https://arxiv.org/abs/2201.12886) and the Neural Hierarchical
+Interpolation [(NHITS)](https://arxiv.org/abs/2201.12886) with the multi
+quantile loss function (MQLoss).
+
+$$ \mathrm{MQLoss}(y_{\tau}, [\hat{y}^{(q1)}_{\tau},\hat{y}^{(q2)}_{\tau},\dots,\hat{y}^{(Q)}_{\tau}]) = \frac{1}{H} \sum_{q} \mathrm{QL}(y_{\tau}, \hat{y}^{(q)}_{\tau}) $$
+
+In this notebook we will: 1. Install NeuralForecast Library 2.
+Explore the M4-Hourly data. 3. Train the LSTM and NHITS 4.
+Visualize the LSTM/NHITS prediction intervals.
+
+You can run these experiments using GPU with Google Colab.
+
+
+
+## 1. Installing NeuralForecast
+
+
+```python
+!pip install neuralforecast
+```
+
+#### Useful functions
+
+The `plot_grid` auxiliary function defined below will be useful to plot
+different time series, and different models’ forecasts.
+
+
+```python
+import logging
+import warnings
+
+import torch
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+warnings.filterwarnings("ignore")
+```
+
+## 2. Loading M4 Data
+
+For testing purposes, we will use the Hourly dataset from the [M4
+competition](https://www.researchgate.net/publication/325901666_The_M4_Competition_Results_findings_conclusion_and_way_forward).
+
+
+```python
+import pandas as pd
+```
+
+
+```python
+Y_train_df = pd.read_csv('https://auto-arima-results.s3.amazonaws.com/M4-Hourly.csv')
+Y_test_df = pd.read_csv(
+ 'https://auto-arima-results.s3.amazonaws.com/M4-Hourly-test.csv'
+).rename(columns={'y': 'y_test'})
+```
+
+In this example we will use a subset of the data to avoid waiting too
+long. You can modify the number of series if you want.
+
+
+```python
+n_series = 8
+uids = Y_train_df['unique_id'].unique()[:n_series]
+Y_train_df = Y_train_df.query('unique_id in @uids')
+Y_test_df = Y_test_df.query('unique_id in @uids')
+```
+
+
+```python
+plot_series(Y_train_df, Y_test_df)
+```
+
+
+
+## 3. Model Training
+
+The `core.NeuralForecast` provides a high-level interface with our
+collection of PyTorch models.
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+is instantiated with a list of `models=[LSTM(...), NHITS(...)]`,
+configured for the forecasting task.
+
+- The `horizon` parameter controls the number of steps ahead of the
+ predictions, in this example 48 hours ahead (2 days).
+- The
+ [`MQLoss`](https://nixtlaverse.nixtla.io/neuralforecast/losses.pytorch.html#mqloss)
+ with `levels=[80,90]` specializes the network’s output into the 80%
+ and 90% prediction intervals.
+- The `max_steps=2000`, controls the duration of the network’s
+ training.
+
+For more network’s instantiation details check their
+[documentation](https://nixtla.github.io/neuralforecast/models.dilated_rnn.html).
+
+
+```python
+from neuralforecast import NeuralForecast
+from neuralforecast.losses.pytorch import MQLoss
+from neuralforecast.models import LSTM, NHITS
+```
+
+
+```python
+logging.getLogger('pytorch_lightning').setLevel(logging.ERROR)
+torch.set_float32_matmul_precision('high')
+```
+
+
+```python
+horizon = 48
+levels = [80, 90]
+models = [LSTM(input_size=3*horizon, h=horizon,
+ loss=MQLoss(level=levels), max_steps=1000),
+ NHITS(input_size=7*horizon, h=horizon,
+ n_freq_downsample=[24, 12, 1],
+ loss=MQLoss(level=levels), max_steps=2000),]
+nf = NeuralForecast(models=models, freq=1)
+```
+
+``` text
+Seed set to 1
+Seed set to 1
+```
+
+All the models of the library are global, meaning that all time series
+in `Y_train_df` is used during a shared optimization to train a single
+model with shared parameters. This is the most common practice in the
+forecasting literature for deep learning models, and it is known as
+“cross-learning”.
+
+
+```python
+nf.fit(df=Y_train_df)
+```
+
+
+```python
+Y_hat_df = nf.predict()
+Y_hat_df.head()
+```
+
+``` text
+Predicting: | | 0/? [00:00
+- [Jeffrey L. Elman (1990). “Finding Structure in
+ Time”.](https://onlinelibrary.wiley.com/doi/abs/10.1207/s15516709cog1402_1)
+- [Cristian Challu, Kin G. Olivares, Boris N. Oreshkin, Federico
+ Garza, Max Mergenthaler-Canseco, Artur Dubrawski (2021). NHITS:
+ Neural Hierarchical Interpolation for Time Series Forecasting.
+ Accepted at AAAI 2023.](https://arxiv.org/abs/2201.12886)
+
diff --git a/neuralforecast/docs/tutorials/using_mlflow.html.mdx b/neuralforecast/docs/tutorials/using_mlflow.html.mdx
new file mode 100644
index 00000000..12f74db5
--- /dev/null
+++ b/neuralforecast/docs/tutorials/using_mlflow.html.mdx
@@ -0,0 +1,129 @@
+---
+description: Log your neuralforecast experiments to MLflow
+output-file: using_mlflow.html
+title: Using MLflow
+---
+
+
+## Installing dependencies
+
+To install Neuralforecast refer to
+[Installation](https://nixtlaverse.nixtla.io/neuralforecast/docs/getting-started/installation.html).
+
+To install mlflow: `pip install mlflow`
+
+## Imports
+
+
+```python
+import logging
+import warnings
+
+import matplotlib.pyplot as plt
+import mlflow
+import mlflow.data
+import numpy as np
+import pandas as pd
+from mlflow.client import MlflowClient
+from mlflow.data.pandas_dataset import PandasDataset
+from utilsforecast.plotting import plot_series
+
+from neuralforecast.core import NeuralForecast
+from neuralforecast.models import NBEATSx
+from neuralforecast.utils import AirPassengersDF
+from neuralforecast.losses.pytorch import MAE
+```
+
+
+```python
+logging.getLogger("mlflow").setLevel(logging.ERROR)
+logging.getLogger("pytorch_lightning").setLevel(logging.ERROR)
+warnings.filterwarnings("ignore")
+```
+
+## Splitting the data
+
+
+```python
+# Split data and declare panel dataset
+Y_df = AirPassengersDF
+Y_train_df = Y_df[Y_df.ds<='1959-12-31'] # 132 train
+Y_test_df = Y_df[Y_df.ds>'1959-12-31'] # 12 test
+Y_df.tail()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|-------|
+| 139 | 1.0 | 1960-08-31 | 606.0 |
+| 140 | 1.0 | 1960-09-30 | 508.0 |
+| 141 | 1.0 | 1960-10-31 | 461.0 |
+| 142 | 1.0 | 1960-11-30 | 390.0 |
+| 143 | 1.0 | 1960-12-31 | 432.0 |
+
+## MLflow UI
+
+Run the following command from the terminal to start the UI:
+`mlflow ui`. You can then go to the printed URL to visualize the
+experiments.
+
+## Model training
+
+
+```python
+mlflow.pytorch.autolog(checkpoint=False)
+
+with mlflow.start_run() as run:
+ # Log the dataset to the MLflow Run. Specify the "training" context to indicate that the
+ # dataset is used for model training
+ dataset: PandasDataset = mlflow.data.from_pandas(Y_df, source="AirPassengersDF")
+ mlflow.log_input(dataset, context="training")
+
+ # Define and log parameters
+ horizon = len(Y_test_df)
+ model_params = dict(
+ input_size=1 * horizon,
+ h=horizon,
+ max_steps=300,
+ loss=MAE(),
+ valid_loss=MAE(),
+ activation='ReLU',
+ scaler_type='robust',
+ random_seed=42,
+ enable_progress_bar=False,
+ )
+ mlflow.log_params(model_params)
+
+ # Fit NBEATSx model
+ models = [NBEATSx(**model_params)]
+ nf = NeuralForecast(models=models, freq='M')
+ train = nf.fit(df=Y_train_df, val_size=horizon)
+
+ # Save conda environment used to run the model
+ mlflow.pytorch.get_default_conda_env()
+
+ # Save pip requirements
+ mlflow.pytorch.get_default_pip_requirements()
+
+mlflow.pytorch.autolog(disable=True)
+
+# Save the neural forecast model
+nf.save(path='./checkpoints/test_run_1/',
+ model_index=None,
+ overwrite=True,
+ save_dataset=True)
+```
+
+``` text
+Seed set to 42
+```
+
+## Forecasting the future
+
+
+```python
+Y_hat_df = nf.predict(futr_df=Y_test_df)
+plot_series(Y_train_df, Y_hat_df, palette='tab20b')
+```
+
+
+
diff --git a/neuralforecast/docs/use-cases/electricity_peak_forecasting.html.mdx b/neuralforecast/docs/use-cases/electricity_peak_forecasting.html.mdx
new file mode 100644
index 00000000..2c110142
--- /dev/null
+++ b/neuralforecast/docs/use-cases/electricity_peak_forecasting.html.mdx
@@ -0,0 +1,264 @@
+---
+description: >-
+ In this example we will show how to perform electricity load forecasting on
+ the ERCOT (Texas) market for detecting daily peaks.
+output-file: electricity_peak_forecasting.html
+title: Detect Demand Peaks
+---
+
+
+## Introduction
+
+Predicting peaks in different markets is useful. In the electricity
+market, consuming electricity at peak demand is penalized with higher
+tarifs. When an individual or company consumes electricity when its most
+demanded, regulators calls that a coincident peak (CP).
+
+In the Texas electricity market (ERCOT), the peak is the monthly
+15-minute interval when the ERCOT Grid is at a point of highest
+capacity. The peak is caused by all consumers’ combined demand on the
+electrical grid. The coincident peak demand is an important factor used
+by ERCOT to determine final electricity consumption bills. ERCOT
+registers the CP demand of each client for 4 months, between June and
+September, and uses this to adjust electricity prices. Clients can
+therefore save on electricity bills by reducing the coincident peak
+demand.
+
+In this example we will train an
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+model on historic load data to forecast day-ahead peaks on September
+2022. Multiple seasonality is traditionally present in low sampled
+electricity data. Demand exhibits daily and weekly seasonality, with
+clear patterns for specific hours of the day such as 6:00pm vs 3:00am or
+for specific days such as Sunday vs Friday.
+
+First, we will load ERCOT historic demand, then we will use the
+`Neuralforecast.cross_validation` method to fit the model and forecast
+daily load during September. Finally, we show how to use the forecasts
+to detect the coincident peak.
+
+**Outline**
+
+1. Install libraries
+2. Load and explore the data
+3. Fit NHITS model and forecast
+4. Peak detection
+
+> **Tip**
+>
+> You can use Colab to run this Notebook interactively
+>
+
+## Libraries
+
+We assume you have NeuralForecast already installed. Check this guide
+for instructions on [how to install
+NeuralForecast](../getting-started/installation.html).
+
+Install the necessary packages using `pip install neuralforecast`
+
+## Load Data
+
+The input to NeuralForecast models is always a data frame in [long
+format](https://www.theanalysisfactor.com/wide-and-long-data/) with
+three columns: `unique_id`, `ds` and `y`:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp or int) column should be either an integer
+ indexing time or a datestamp ideally like YYYY-MM-DD for a date or
+ YYYY-MM-DD HH:MM:SS for a timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast. We
+ will rename the
+
+First, download and read the 2022 historic total demand of the ERCOT
+market, available [here](https://www.ercot.com/gridinfo/load/load_hist).
+The data processing includes adding the missing hour due to daylight
+saving time, parsing the date to datetime format, and filtering columns
+of interest.
+
+
+```python
+import numpy as np
+import pandas as pd
+```
+
+
+```python
+# Load data
+Y_df = pd.read_csv('https://datasets-nixtla.s3.amazonaws.com/ERCOT-clean.csv', parse_dates=['ds'])
+Y_df = Y_df.query("ds >= '2022-01-01' & ds <= '2022-10-01'")
+```
+
+
+```python
+Y_df.plot(x='ds', y='y', figsize=(20, 7))
+```
+
+
+
+## Fit and Forecast with NHITS
+
+Import the
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+class and the models you need.
+
+
+```python
+from neuralforecast.core import NeuralForecast
+from neuralforecast.auto import AutoNHITS
+```
+
+First, instantiate the model and define the parameters. To instantiate
+[`AutoNHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autonhits)
+you need to define:
+
+- `h`: forecasting horizon
+- `loss`: training loss. Use the
+ [`DistributionLoss`](https://nixtlaverse.nixtla.io/neuralforecast/losses.pytorch.html#distributionloss)
+ to produce probabilistic forecasts. Default:
+ [`MAE`](https://nixtlaverse.nixtla.io/neuralforecast/losses.pytorch.html#mae).
+- `config`: hyperparameter search space. If `None`, the
+ [`AutoNHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.html#autonhits)
+ class will use a pre-defined suggested hyperparameter space.
+- `num_samples`: number of configurations explored.
+
+
+```python
+models = [AutoNHITS(h=24,
+ config=None, # Uses default config
+ num_samples=10
+ )
+ ]
+```
+
+We fit the model by instantiating a
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+object with the following required parameters:
+
+- `models`: a list of models. Select the models you want from
+ [models](../capabilities/overview.html) and import them.
+
+- `freq`: a string indicating the frequency of the data. (See [panda’s
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+
+```python
+# Instantiate StatsForecast class as sf
+nf = NeuralForecast(
+ models=models,
+ freq='h',
+)
+```
+
+The `cross_validation` method allows the user to simulate multiple
+historic forecasts, greatly simplifying pipelines by replacing for loops
+with `fit` and `predict` methods. This method re-trains the model and
+forecast each window. See [this
+tutorial](https://nixtla.github.io/statsforecast/docs/getting-started/getting_started_complete.html)
+for an animation of how the windows are defined.
+
+Use the `cross_validation` method to produce all the daily forecasts for
+September. To produce daily forecasts set the forecasting horizon `h` as
+24. In this example we are simulating deploying the pipeline during
+September, so set the number of windows as 30 (one for each day).
+Finally, set the step size between windows as 24, to only produce one
+forecast per day.
+
+
+```python
+crossvalidation_df = nf.cross_validation(
+ df=Y_df,
+ step_size=24,
+ n_windows=30
+ )
+```
+
+
+```python
+crossvalidation_df.head()
+```
+
+| | unique_id | ds | cutoff | AutoNHITS | y |
+|----|----|----|----|----|----|
+| 0 | ERCOT | 2022-09-01 00:00:00 | 2022-08-31 23:00:00 | 45841.601562 | 45482.471757 |
+| 1 | ERCOT | 2022-09-01 01:00:00 | 2022-08-31 23:00:00 | 43613.394531 | 43602.658043 |
+| 2 | ERCOT | 2022-09-01 02:00:00 | 2022-08-31 23:00:00 | 41968.945312 | 42284.817342 |
+| 3 | ERCOT | 2022-09-01 03:00:00 | 2022-08-31 23:00:00 | 41038.539062 | 41663.156771 |
+| 4 | ERCOT | 2022-09-01 04:00:00 | 2022-08-31 23:00:00 | 41237.203125 | 41710.621904 |
+
+> **Important**
+>
+> When using `cross_validation` make sure the forecasts are produced at
+> the desired timestamps. Check the `cutoff` column which specifices the
+> last timestamp before the forecasting window.
+
+## Peak Detection
+
+Finally, we use the forecasts in `crossvaldation_df` to detect the daily
+hourly demand peaks. For each day, we set the detected peaks as the
+highest forecasts. In this case, we want to predict one peak (`npeaks`);
+depending on your setting and goals, this parameter might change. For
+example, the number of peaks can correspond to how many hours a battery
+can be discharged to reduce demand.
+
+
+```python
+npeaks = 1 # Number of peaks
+```
+
+For the ERCOT 4CP detection task we are interested in correctly
+predicting the highest monthly load. Next, we filter the day in
+September with the highest hourly demand and predict the peak.
+
+
+```python
+crossvalidation_df = crossvalidation_df[['ds','y','AutoNHITS']]
+max_day = crossvalidation_df.iloc[crossvalidation_df['y'].argmax()].ds.day # Day with maximum load
+cv_df_day = crossvalidation_df.query('ds.dt.day == @max_day')
+max_hour = cv_df_day['y'].argmax()
+peaks = cv_df_day['AutoNHITS'].argsort().iloc[-npeaks:].values # Predicted peaks
+```
+
+In the following plot we see how the model is able to correctly detect
+the coincident peak for September 2022.
+
+
+```python
+import matplotlib.pyplot as plt
+```
+
+
+```python
+plt.figure(figsize=(10, 5))
+plt.axvline(cv_df_day.iloc[max_hour]['ds'], color='black', label='True Peak')
+plt.scatter(cv_df_day.iloc[peaks]['ds'], cv_df_day.iloc[peaks]['AutoNHITS'], color='green', label=f'Predicted Top-{npeaks}')
+plt.plot(cv_df_day['ds'], cv_df_day['y'], label='y', color='blue')
+plt.plot(cv_df_day['ds'], cv_df_day['AutoNHITS'], label='Forecast', color='red')
+plt.xlabel('Time')
+plt.ylabel('Load (MW)')
+plt.grid()
+plt.legend()
+```
+
+
+
+> **Important**
+>
+> In this example we only include September. However,
+> [`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+> can correctly predict the peaks for the 4 months of 2022. You can try
+> this by increasing the `nwindows` parameter of `cross_validation` or
+> filtering the `Y_df` dataset. The complete run for all months take
+> only 10 minutes.
+
+## References
+
+- [Cristian Challu, Kin G. Olivares, Boris N. Oreshkin, Federico
+ Garza, Max Mergenthaler-Canseco, Artur Dubrawski (2021). “NHITS:
+ Neural Hierarchical Interpolation for Time Series Forecasting”.
+ Accepted at AAAI 2023.](https://arxiv.org/abs/2201.12886)
+
diff --git a/neuralforecast/docs/use-cases/electricity_peak_forecasting_files/figure-markdown_strict/cell-13-output-1.png b/neuralforecast/docs/use-cases/electricity_peak_forecasting_files/figure-markdown_strict/cell-13-output-1.png
new file mode 100644
index 00000000..d733662a
Binary files /dev/null and b/neuralforecast/docs/use-cases/electricity_peak_forecasting_files/figure-markdown_strict/cell-13-output-1.png differ
diff --git a/neuralforecast/docs/use-cases/electricity_peak_forecasting_files/figure-markdown_strict/cell-4-output-1.png b/neuralforecast/docs/use-cases/electricity_peak_forecasting_files/figure-markdown_strict/cell-4-output-1.png
new file mode 100644
index 00000000..88b9b105
Binary files /dev/null and b/neuralforecast/docs/use-cases/electricity_peak_forecasting_files/figure-markdown_strict/cell-4-output-1.png differ
diff --git a/neuralforecast/docs/use-cases/predictive_maintenance.html.mdx b/neuralforecast/docs/use-cases/predictive_maintenance.html.mdx
new file mode 100644
index 00000000..9e756f5e
--- /dev/null
+++ b/neuralforecast/docs/use-cases/predictive_maintenance.html.mdx
@@ -0,0 +1,255 @@
+---
+output-file: predictive_maintenance.html
+title: Predictive Maintenance
+---
+
+
+Predictive maintenance (PdM) is a data-driven preventive maintanance
+program. It is a proactive maintenance strategy that uses sensors to
+monitor the performance and equipment conditions during operation. The
+PdM methods constantly analyze the data to predict when optimal
+maintenance schedules. It can reduce maintenance costs and prevent
+catastrophic equipment failure when used correctly.
+
+In this notebook, we will apply NeuralForecast to perform a supervised
+Remaining Useful Life (RUL) estimation on the classic PHM2008 aircraft
+degradation dataset.
+
+Outline 1. Installing Packages 2. Load PHM2008 aircraft
+degradation dataset 3. Fit and Predict NeuralForecast 4.
+Evaluate Predictions
+
+You can run these experiments using GPU with Google Colab.
+
+
+
+## 1. Installing Packages
+
+
+```python
+!pip install neuralforecast datasetsforecast
+```
+
+
+```python
+import logging
+import numpy as np
+import pandas as pd
+
+import matplotlib.pyplot as plt
+plt.rcParams['font.family'] = 'serif'
+
+from neuralforecast.models import NBEATSx
+from neuralforecast import NeuralForecast
+from neuralforecast.losses.pytorch import HuberLoss
+
+from datasetsforecast.phm2008 import PHM2008
+```
+
+
+```python
+logging.getLogger("pytorch_lightning").setLevel(logging.ERROR)
+```
+
+## 2. Load PHM2008 aircraft degradation dataset
+
+Here we will load the Prognosis and Health Management 2008 challenge
+dataset. This dataset used the Commercial Modular Aero-Propulsion System
+Simulation to recreate the degradation process of turbofan engines for
+different aircraft with varying wear and manufacturing starting under
+normal conditions. The training dataset consists of complete
+run-to-failure simulations, while the test dataset comprises sequences
+before failure.
+
+
+
+
+```python
+Y_train_df, Y_test_df = PHM2008.load(directory='./data', group='FD001', clip_rul=False)
+Y_train_df
+```
+
+``` text
+100%|██████████| 12.4M/12.4M [00:00<00:00, 21.6MiB/s]
+INFO:datasetsforecast.utils:Successfully downloaded CMAPSSData.zip, 12437405, bytes.
+INFO:datasetsforecast.utils:Decompressing zip file...
+INFO:datasetsforecast.utils:Successfully decompressed data\phm2008\CMAPSSData.zip
+```
+
+| | unique_id | ds | s_2 | s_3 | s_4 | s_7 | s_8 | s_9 | s_11 | s_12 | s_13 | s_14 | s_15 | s_17 | s_20 | s_21 | y |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | 1 | 1 | 641.82 | 1589.70 | 1400.60 | 554.36 | 2388.06 | 9046.19 | 47.47 | 521.66 | 2388.02 | 8138.62 | 8.4195 | 392 | 39.06 | 23.4190 | 191 |
+| 1 | 1 | 2 | 642.15 | 1591.82 | 1403.14 | 553.75 | 2388.04 | 9044.07 | 47.49 | 522.28 | 2388.07 | 8131.49 | 8.4318 | 392 | 39.00 | 23.4236 | 190 |
+| 2 | 1 | 3 | 642.35 | 1587.99 | 1404.20 | 554.26 | 2388.08 | 9052.94 | 47.27 | 522.42 | 2388.03 | 8133.23 | 8.4178 | 390 | 38.95 | 23.3442 | 189 |
+| 3 | 1 | 4 | 642.35 | 1582.79 | 1401.87 | 554.45 | 2388.11 | 9049.48 | 47.13 | 522.86 | 2388.08 | 8133.83 | 8.3682 | 392 | 38.88 | 23.3739 | 188 |
+| 4 | 1 | 5 | 642.37 | 1582.85 | 1406.22 | 554.00 | 2388.06 | 9055.15 | 47.28 | 522.19 | 2388.04 | 8133.80 | 8.4294 | 393 | 38.90 | 23.4044 | 187 |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| 20626 | 100 | 196 | 643.49 | 1597.98 | 1428.63 | 551.43 | 2388.19 | 9065.52 | 48.07 | 519.49 | 2388.26 | 8137.60 | 8.4956 | 397 | 38.49 | 22.9735 | 4 |
+| 20627 | 100 | 197 | 643.54 | 1604.50 | 1433.58 | 550.86 | 2388.23 | 9065.11 | 48.04 | 519.68 | 2388.22 | 8136.50 | 8.5139 | 395 | 38.30 | 23.1594 | 3 |
+| 20628 | 100 | 198 | 643.42 | 1602.46 | 1428.18 | 550.94 | 2388.24 | 9065.90 | 48.09 | 520.01 | 2388.24 | 8141.05 | 8.5646 | 398 | 38.44 | 22.9333 | 2 |
+| 20629 | 100 | 199 | 643.23 | 1605.26 | 1426.53 | 550.68 | 2388.25 | 9073.72 | 48.39 | 519.67 | 2388.23 | 8139.29 | 8.5389 | 395 | 38.29 | 23.0640 | 1 |
+| 20630 | 100 | 200 | 643.85 | 1600.38 | 1432.14 | 550.79 | 2388.26 | 9061.48 | 48.20 | 519.30 | 2388.26 | 8137.33 | 8.5036 | 396 | 38.37 | 23.0522 | 0 |
+
+
+```python
+plot_df1 = Y_train_df[Y_train_df['unique_id']==1]
+plot_df2 = Y_train_df[Y_train_df['unique_id']==2]
+plot_df3 = Y_train_df[Y_train_df['unique_id']==3]
+
+plt.plot(plot_df1.ds, np.minimum(plot_df1.y, 125), color='#2D6B8F', linestyle='--')
+plt.plot(plot_df1.ds, plot_df1.y, color='#2D6B8F', label='Engine 1')
+
+plt.plot(plot_df2.ds, np.minimum(plot_df2.y, 125)+1.5, color='#CA6F6A', linestyle='--')
+plt.plot(plot_df2.ds, plot_df2.y+1.5, color='#CA6F6A', label='Engine 2')
+
+plt.plot(plot_df3.ds, np.minimum(plot_df3.y, 125)-1.5, color='#D5BC67', linestyle='--')
+plt.plot(plot_df3.ds, plot_df3.y-1.5, color='#D5BC67', label='Engine 3')
+
+plt.ylabel('Remaining Useful Life (RUL)', fontsize=15)
+plt.xlabel('Time Cycle', fontsize=15)
+plt.legend()
+plt.grid()
+```
+
+
+
+
+```python
+def smooth(s, b = 0.98):
+ v = np.zeros(len(s)+1) #v_0 is already 0.
+ bc = np.zeros(len(s)+1)
+ for i in range(1, len(v)): #v_t = 0.95
+ v[i] = (b * v[i-1] + (1-b) * s[i-1])
+ bc[i] = 1 - b**i
+ sm = v[1:] / bc[1:]
+ return sm
+
+unique_id = 1
+plot_df = Y_train_df[Y_train_df.unique_id == unique_id].copy()
+
+fig, axes = plt.subplots(2,3, figsize = (8,5))
+fig.tight_layout()
+
+j = -1
+#, 's_11', 's_12', 's_13', 's_14', 's_15', 's_17', 's_20', 's_21'
+for feature in ['s_2', 's_3', 's_4', 's_7', 's_8', 's_9']:
+ if ('s' in feature) and ('smoothed' not in feature):
+ j += 1
+ axes[j // 3, j % 3].plot(plot_df.ds, plot_df[feature],
+ c = '#2D6B8F', label = 'original')
+ axes[j // 3, j % 3].plot(plot_df.ds, smooth(plot_df[feature].values),
+ c = '#CA6F6A', label = 'smoothed')
+ #axes[j // 3, j % 3].plot([10,10],[0,1], c = 'black')
+ axes[j // 3, j % 3].set_title(feature)
+ axes[j // 3, j % 3].grid()
+ axes[j // 3, j % 3].legend()
+
+plt.suptitle(f'Engine {unique_id} sensor records')
+plt.tight_layout()
+```
+
+
+
+## 3. Fit and Predict NeuralForecast
+
+NeuralForecast methods are capable of addressing regression problems
+involving various variables. The regression problem involves predicting
+the target variable $y_{t+h}$ based on its lags $y_{:t}$, temporal
+exogenous features $x^{(h)}_{:t}$, exogenous features available at the
+time of prediction $x^{(f)}_{:t+h}$, and static features $x^{(s)}$.
+
+The task of estimating the remaining useful life (RUL) simplifies the
+problem to a single horizon prediction $h=1$, where the objective is to
+predict $y_{t+1}$ based on the exogenous features $x^{(f)}_{:t+1}$ and
+static features $x^{(s)}$. In the RUL estimation task, the exogenous
+features typically correspond to sensor monitoring information, while
+the target variable represents the RUL itself.
+
+$$P(y_{t+1}\;|\;x^{(f)}_{:t+1},x^{(s)})$$
+
+
+```python
+Y_train_df, Y_test_df = PHM2008.load(directory='./data', group='FD001', clip_rul=True)
+max_ds = Y_train_df.groupby('unique_id')["ds"].max()
+Y_test_df = Y_test_df.merge(max_ds, on='unique_id', how='left', suffixes=('', '_train_max_date'))
+Y_test_df["ds"] = Y_test_df["ds"] + Y_test_df["ds_train_max_date"]
+Y_test_df = Y_test_df.drop(columns=["ds_train_max_date"])
+```
+
+
+```python
+futr_exog_list =['s_2', 's_3', 's_4', 's_7', 's_8', 's_9', 's_11',
+ 's_12', 's_13', 's_14', 's_15', 's_17', 's_20', 's_21']
+
+model = NBEATSx(h=1,
+ input_size=24,
+ loss=HuberLoss(),
+ scaler_type='robust',
+ stack_types=['identity', 'identity', 'identity'],
+ dropout_prob_theta=0.5,
+ futr_exog_list=futr_exog_list,
+ exclude_insample_y = True,
+ max_steps=1000)
+nf = NeuralForecast(models=[model], freq=1)
+
+nf.fit(df=Y_train_df)
+Y_hat_df = nf.predict(futr_df=Y_test_df)
+```
+
+## 4. Evaluate Predictions
+
+In the original PHM2008 dataset the true RUL values for the test set are
+only provided for the last time cycle of each enginge. We will filter
+the predictions to only evaluate the last time cycle.
+
+$$RMSE(\mathbf{y}_{T},\hat{\mathbf{y}}_{T}) = \sqrt{\frac{1}{|\mathcal{D}_{test}|} \sum_{i} (y_{i,T}-\hat{y}_{i,T})^{2}}$$
+
+
+```python
+from utilsforecast.evaluation import evaluate
+from utilsforecast.losses import rmse
+```
+
+
+```python
+metrics = evaluate(Y_hat_df.merge(Y_test_df[["unique_id", "ds", "y"]], on=['unique_id', 'ds']),
+ metrics=[rmse],
+ agg_fn='mean')
+
+metrics
+```
+
+| | metric | NBEATSx |
+|-----|--------|------------|
+| 0 | rmse | 118.179373 |
+
+
+```python
+plot_df1 = Y_hat_df2[Y_hat_df2['unique_id']==1]
+plot_df2 = Y_hat_df2[Y_hat_df2['unique_id']==2]
+plot_df3 = Y_hat_df2[Y_hat_df2['unique_id']==3]
+
+plt.plot(plot_df1.ds, plot_df1['y'], c='#2D6B8F', label='E1 true RUL')
+plt.plot(plot_df1.ds, plot_df1[model_name]+1, c='#2D6B8F', linestyle='--', label='E1 predicted RUL')
+
+plt.plot(plot_df1.ds, plot_df2['y'], c='#CA6F6A', label='E2 true RUL')
+plt.plot(plot_df1.ds, plot_df2[model_name]+1, c='#CA6F6A', linestyle='--', label='E2 predicted RUL')
+
+plt.plot(plot_df1.ds, plot_df3['y'], c='#D5BC67', label='E3 true RUL')
+plt.plot(plot_df1.ds, plot_df3[model_name]+1, c='#D5BC67', linestyle='--', label='E3 predicted RUL')
+
+plt.legend()
+plt.grid()
+```
+
+
+
+## References
+
+- [R. Keith Mobley (2002). “An Introduction to Predictive
+ Maintenance”](https://www.irantpm.ir/wp-content/uploads/2008/02/an-introduction-to-predictive-maintenance.pdf)
+- [Saxena, A., Goebel, K., Simon, D.,&Eklund, N. (2008). “Damage
+ propagation modeling for aircraft engine run-to-failure simulation”.
+ International conference on prognostics and health
+ management.](https://ntrs.nasa.gov/api/citations/20090029214/downloads/20090029214.pdf)
+
diff --git a/neuralforecast/docs/use-cases/predictive_maintenance_files/figure-markdown_strict/cell-12-output-1.png b/neuralforecast/docs/use-cases/predictive_maintenance_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..4d3f648d
Binary files /dev/null and b/neuralforecast/docs/use-cases/predictive_maintenance_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/neuralforecast/docs/use-cases/predictive_maintenance_files/figure-markdown_strict/cell-6-output-1.png b/neuralforecast/docs/use-cases/predictive_maintenance_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..fe3bb4eb
Binary files /dev/null and b/neuralforecast/docs/use-cases/predictive_maintenance_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/neuralforecast/docs/use-cases/predictive_maintenance_files/figure-markdown_strict/cell-7-output-1.png b/neuralforecast/docs/use-cases/predictive_maintenance_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..784a730f
Binary files /dev/null and b/neuralforecast/docs/use-cases/predictive_maintenance_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/neuralforecast/favicon.svg b/neuralforecast/favicon.svg
new file mode 100644
index 00000000..e5f33342
--- /dev/null
+++ b/neuralforecast/favicon.svg
@@ -0,0 +1,5 @@
+
diff --git a/neuralforecast/imgs_indx/nf_map.png b/neuralforecast/imgs_indx/nf_map.png
new file mode 100644
index 00000000..07d01677
Binary files /dev/null and b/neuralforecast/imgs_indx/nf_map.png differ
diff --git a/neuralforecast/imgs_indx/predict_insample.png b/neuralforecast/imgs_indx/predict_insample.png
new file mode 100644
index 00000000..46112179
Binary files /dev/null and b/neuralforecast/imgs_indx/predict_insample.png differ
diff --git a/neuralforecast/imgs_losses/gmm.png b/neuralforecast/imgs_losses/gmm.png
new file mode 100644
index 00000000..eb2ec2c6
Binary files /dev/null and b/neuralforecast/imgs_losses/gmm.png differ
diff --git a/neuralforecast/imgs_losses/hmq_loss.png b/neuralforecast/imgs_losses/hmq_loss.png
new file mode 100644
index 00000000..393f9179
Binary files /dev/null and b/neuralforecast/imgs_losses/hmq_loss.png differ
diff --git a/neuralforecast/imgs_losses/huber_loss.png b/neuralforecast/imgs_losses/huber_loss.png
new file mode 100644
index 00000000..3eaf667e
Binary files /dev/null and b/neuralforecast/imgs_losses/huber_loss.png differ
diff --git a/neuralforecast/imgs_losses/huber_qloss.png b/neuralforecast/imgs_losses/huber_qloss.png
new file mode 100644
index 00000000..5be63e16
Binary files /dev/null and b/neuralforecast/imgs_losses/huber_qloss.png differ
diff --git a/neuralforecast/imgs_losses/mae_loss.png b/neuralforecast/imgs_losses/mae_loss.png
new file mode 100644
index 00000000..c9d3b7fa
Binary files /dev/null and b/neuralforecast/imgs_losses/mae_loss.png differ
diff --git a/neuralforecast/imgs_losses/mape_loss.png b/neuralforecast/imgs_losses/mape_loss.png
new file mode 100644
index 00000000..d0f9a66a
Binary files /dev/null and b/neuralforecast/imgs_losses/mape_loss.png differ
diff --git a/neuralforecast/imgs_losses/mase_loss.png b/neuralforecast/imgs_losses/mase_loss.png
new file mode 100644
index 00000000..90db8c90
Binary files /dev/null and b/neuralforecast/imgs_losses/mase_loss.png differ
diff --git a/neuralforecast/imgs_losses/mq_loss.png b/neuralforecast/imgs_losses/mq_loss.png
new file mode 100644
index 00000000..7e3f6da3
Binary files /dev/null and b/neuralforecast/imgs_losses/mq_loss.png differ
diff --git a/neuralforecast/imgs_losses/mse_loss.png b/neuralforecast/imgs_losses/mse_loss.png
new file mode 100644
index 00000000..d175d5e0
Binary files /dev/null and b/neuralforecast/imgs_losses/mse_loss.png differ
diff --git a/neuralforecast/imgs_losses/pmm.png b/neuralforecast/imgs_losses/pmm.png
new file mode 100644
index 00000000..151296a4
Binary files /dev/null and b/neuralforecast/imgs_losses/pmm.png differ
diff --git a/neuralforecast/imgs_losses/q_loss.png b/neuralforecast/imgs_losses/q_loss.png
new file mode 100644
index 00000000..942dbc30
Binary files /dev/null and b/neuralforecast/imgs_losses/q_loss.png differ
diff --git a/neuralforecast/imgs_losses/rmae_loss.png b/neuralforecast/imgs_losses/rmae_loss.png
new file mode 100644
index 00000000..39a05b2e
Binary files /dev/null and b/neuralforecast/imgs_losses/rmae_loss.png differ
diff --git a/neuralforecast/imgs_losses/rmse_loss.png b/neuralforecast/imgs_losses/rmse_loss.png
new file mode 100644
index 00000000..0ceadef0
Binary files /dev/null and b/neuralforecast/imgs_losses/rmse_loss.png differ
diff --git a/neuralforecast/imgs_losses/tukey_loss.png b/neuralforecast/imgs_losses/tukey_loss.png
new file mode 100644
index 00000000..6eec62e5
Binary files /dev/null and b/neuralforecast/imgs_losses/tukey_loss.png differ
diff --git a/neuralforecast/imgs_models/autoformer.png b/neuralforecast/imgs_models/autoformer.png
new file mode 100644
index 00000000..3721f560
Binary files /dev/null and b/neuralforecast/imgs_models/autoformer.png differ
diff --git a/neuralforecast/imgs_models/bitcn.png b/neuralforecast/imgs_models/bitcn.png
new file mode 100644
index 00000000..3241705b
Binary files /dev/null and b/neuralforecast/imgs_models/bitcn.png differ
diff --git a/neuralforecast/imgs_models/data_splits.png b/neuralforecast/imgs_models/data_splits.png
new file mode 100644
index 00000000..b7733514
Binary files /dev/null and b/neuralforecast/imgs_models/data_splits.png differ
diff --git a/neuralforecast/imgs_models/deepar.jpeg b/neuralforecast/imgs_models/deepar.jpeg
new file mode 100644
index 00000000..b9ae87b5
Binary files /dev/null and b/neuralforecast/imgs_models/deepar.jpeg differ
diff --git a/neuralforecast/imgs_models/dilated_rnn.png b/neuralforecast/imgs_models/dilated_rnn.png
new file mode 100644
index 00000000..f17ddde4
Binary files /dev/null and b/neuralforecast/imgs_models/dilated_rnn.png differ
diff --git a/neuralforecast/imgs_models/dlinear.png b/neuralforecast/imgs_models/dlinear.png
new file mode 100644
index 00000000..24df0454
Binary files /dev/null and b/neuralforecast/imgs_models/dlinear.png differ
diff --git a/neuralforecast/imgs_models/fedformer.png b/neuralforecast/imgs_models/fedformer.png
new file mode 100644
index 00000000..8aab614d
Binary files /dev/null and b/neuralforecast/imgs_models/fedformer.png differ
diff --git a/neuralforecast/imgs_models/gru.png b/neuralforecast/imgs_models/gru.png
new file mode 100644
index 00000000..a258e576
Binary files /dev/null and b/neuralforecast/imgs_models/gru.png differ
diff --git a/neuralforecast/imgs_models/hint.png b/neuralforecast/imgs_models/hint.png
new file mode 100644
index 00000000..5cd6f242
Binary files /dev/null and b/neuralforecast/imgs_models/hint.png differ
diff --git a/neuralforecast/imgs_models/hint_notation.png b/neuralforecast/imgs_models/hint_notation.png
new file mode 100644
index 00000000..411cb66d
Binary files /dev/null and b/neuralforecast/imgs_models/hint_notation.png differ
diff --git a/neuralforecast/imgs_models/informer_architecture.png b/neuralforecast/imgs_models/informer_architecture.png
new file mode 100644
index 00000000..2f39fdeb
Binary files /dev/null and b/neuralforecast/imgs_models/informer_architecture.png differ
diff --git a/neuralforecast/imgs_models/kan.png b/neuralforecast/imgs_models/kan.png
new file mode 100644
index 00000000..ce0b80cf
Binary files /dev/null and b/neuralforecast/imgs_models/kan.png differ
diff --git a/neuralforecast/imgs_models/lstm.png b/neuralforecast/imgs_models/lstm.png
new file mode 100644
index 00000000..47a131cd
Binary files /dev/null and b/neuralforecast/imgs_models/lstm.png differ
diff --git a/neuralforecast/imgs_models/mlp.png b/neuralforecast/imgs_models/mlp.png
new file mode 100644
index 00000000..0fd3e1ad
Binary files /dev/null and b/neuralforecast/imgs_models/mlp.png differ
diff --git a/neuralforecast/imgs_models/nbeats.png b/neuralforecast/imgs_models/nbeats.png
new file mode 100644
index 00000000..ba2d1df6
Binary files /dev/null and b/neuralforecast/imgs_models/nbeats.png differ
diff --git a/neuralforecast/imgs_models/nbeatsx.png b/neuralforecast/imgs_models/nbeatsx.png
new file mode 100644
index 00000000..f4f3e37f
Binary files /dev/null and b/neuralforecast/imgs_models/nbeatsx.png differ
diff --git a/neuralforecast/imgs_models/nhits.png b/neuralforecast/imgs_models/nhits.png
new file mode 100644
index 00000000..f3a83497
Binary files /dev/null and b/neuralforecast/imgs_models/nhits.png differ
diff --git a/neuralforecast/imgs_models/patchtst.png b/neuralforecast/imgs_models/patchtst.png
new file mode 100644
index 00000000..67689873
Binary files /dev/null and b/neuralforecast/imgs_models/patchtst.png differ
diff --git a/neuralforecast/imgs_models/rmok.png b/neuralforecast/imgs_models/rmok.png
new file mode 100644
index 00000000..11c7cf4d
Binary files /dev/null and b/neuralforecast/imgs_models/rmok.png differ
diff --git a/neuralforecast/imgs_models/rnn.png b/neuralforecast/imgs_models/rnn.png
new file mode 100644
index 00000000..51976d64
Binary files /dev/null and b/neuralforecast/imgs_models/rnn.png differ
diff --git a/neuralforecast/imgs_models/tcn.png b/neuralforecast/imgs_models/tcn.png
new file mode 100644
index 00000000..94d7cb6c
Binary files /dev/null and b/neuralforecast/imgs_models/tcn.png differ
diff --git a/neuralforecast/imgs_models/temporal_norm.png b/neuralforecast/imgs_models/temporal_norm.png
new file mode 100644
index 00000000..724895a0
Binary files /dev/null and b/neuralforecast/imgs_models/temporal_norm.png differ
diff --git a/neuralforecast/imgs_models/tft_architecture.png b/neuralforecast/imgs_models/tft_architecture.png
new file mode 100644
index 00000000..2d34d6b6
Binary files /dev/null and b/neuralforecast/imgs_models/tft_architecture.png differ
diff --git a/neuralforecast/imgs_models/tft_grn.png b/neuralforecast/imgs_models/tft_grn.png
new file mode 100644
index 00000000..2eb900b7
Binary files /dev/null and b/neuralforecast/imgs_models/tft_grn.png differ
diff --git a/neuralforecast/imgs_models/tft_vsn.png b/neuralforecast/imgs_models/tft_vsn.png
new file mode 100644
index 00000000..bb773f27
Binary files /dev/null and b/neuralforecast/imgs_models/tft_vsn.png differ
diff --git a/neuralforecast/imgs_models/tide.png b/neuralforecast/imgs_models/tide.png
new file mode 100644
index 00000000..d10b5b01
Binary files /dev/null and b/neuralforecast/imgs_models/tide.png differ
diff --git a/neuralforecast/imgs_models/timellm.png b/neuralforecast/imgs_models/timellm.png
new file mode 100644
index 00000000..17c27b43
Binary files /dev/null and b/neuralforecast/imgs_models/timellm.png differ
diff --git a/neuralforecast/imgs_models/timemixer.png b/neuralforecast/imgs_models/timemixer.png
new file mode 100644
index 00000000..b0812191
Binary files /dev/null and b/neuralforecast/imgs_models/timemixer.png differ
diff --git a/neuralforecast/imgs_models/timesnet.png b/neuralforecast/imgs_models/timesnet.png
new file mode 100644
index 00000000..c9027664
Binary files /dev/null and b/neuralforecast/imgs_models/timesnet.png differ
diff --git a/neuralforecast/imgs_models/timexer.png b/neuralforecast/imgs_models/timexer.png
new file mode 100644
index 00000000..99a6e22a
Binary files /dev/null and b/neuralforecast/imgs_models/timexer.png differ
diff --git a/neuralforecast/imgs_models/tsmixer.png b/neuralforecast/imgs_models/tsmixer.png
new file mode 100644
index 00000000..145a0415
Binary files /dev/null and b/neuralforecast/imgs_models/tsmixer.png differ
diff --git a/neuralforecast/imgs_models/tsmixerx.png b/neuralforecast/imgs_models/tsmixerx.png
new file mode 100644
index 00000000..467b4dff
Binary files /dev/null and b/neuralforecast/imgs_models/tsmixerx.png differ
diff --git a/neuralforecast/imgs_models/vanilla_transformer.png b/neuralforecast/imgs_models/vanilla_transformer.png
new file mode 100644
index 00000000..67804e99
Binary files /dev/null and b/neuralforecast/imgs_models/vanilla_transformer.png differ
diff --git a/neuralforecast/light.png b/neuralforecast/light.png
new file mode 100644
index 00000000..bbb99b54
Binary files /dev/null and b/neuralforecast/light.png differ
diff --git a/neuralforecast/losses.numpy.html.mdx b/neuralforecast/losses.numpy.html.mdx
new file mode 100644
index 00000000..1355398c
--- /dev/null
+++ b/neuralforecast/losses.numpy.html.mdx
@@ -0,0 +1,417 @@
+---
+description: >-
+ NeuralForecast contains a collection NumPy loss functions aimed to be used
+ during the models' evaluation.
+output-file: losses.numpy.html
+title: NumPy Evaluation
+---
+
+
+The most important train signal is the forecast error, which is the
+difference between the observed value $y_{\tau}$ and the prediction
+$\hat{y}_{\tau}$, at time $y_{\tau}$:
+
+$$e_{\tau} = y_{\tau}-\hat{y}_{\tau} \qquad \qquad \tau \in \{t+1,\dots,t+H \}$$
+
+The train loss summarizes the forecast errors in different evaluation
+metrics.
+
+# 1. Scale-dependent Errors
+
+These metrics are on the same scale as the data.
+
+## Mean Absolute Error
+
+------------------------------------------------------------------------
+
+source
+
+### mae
+
+> ``` text
+> mae (y:numpy.ndarray, y_hat:numpy.ndarray,
+> weights:Optional[numpy.ndarray]=None, axis:Optional[int]=None)
+> ```
+
+\*Mean Absolute Error
+
+Calculates Mean Absolute Error between `y` and `y_hat`. MAE measures the
+relative prediction accuracy of a forecasting method by calculating the
+deviation of the prediction and the true value at a given time and
+averages these devations over the length of the series.
+
+$$ \mathrm{MAE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}) = \frac{1}{H} \sum^{t+H}_{\tau=t+1} |y_{\tau} - \hat{y}_{\tau}| $$
+
+**Parameters:** `y`: numpy array, Actual values. `y_hat`: numpy
+array, Predicted values. `mask`: numpy array, Specifies date stamps
+per serie to consider in loss.
+
+**Returns:**
+[`mae`](https://nixtlaverse.nixtla.io/neuralforecast/losses.numpy.html#mae):
+numpy array, (single value).\*
+
+
+
+## Mean Squared Error
+
+------------------------------------------------------------------------
+
+source
+
+### mse
+
+> ``` text
+> mse (y:numpy.ndarray, y_hat:numpy.ndarray,
+> weights:Optional[numpy.ndarray]=None, axis:Optional[int]=None)
+> ```
+
+\*Mean Squared Error
+
+Calculates Mean Squared Error between `y` and `y_hat`. MSE measures the
+relative prediction accuracy of a forecasting method by calculating the
+squared deviation of the prediction and the true value at a given time,
+and averages these devations over the length of the series.
+
+$$ \mathrm{MSE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}) = \frac{1}{H} \sum^{t+H}_{\tau=t+1} (y_{\tau} - \hat{y}_{\tau})^{2} $$
+
+**Parameters:** `y`: numpy array, Actual values. `y_hat`: numpy
+array, Predicted values. `mask`: numpy array, Specifies date stamps
+per serie to consider in loss.
+
+**Returns:**
+[`mse`](https://nixtlaverse.nixtla.io/neuralforecast/losses.numpy.html#mse):
+numpy array, (single value).\*
+
+
+
+## Root Mean Squared Error
+
+------------------------------------------------------------------------
+
+source
+
+### rmse
+
+> ``` text
+> rmse (y:numpy.ndarray, y_hat:numpy.ndarray,
+> weights:Optional[numpy.ndarray]=None, axis:Optional[int]=None)
+> ```
+
+\*Root Mean Squared Error
+
+Calculates Root Mean Squared Error between `y` and `y_hat`. RMSE
+measures the relative prediction accuracy of a forecasting method by
+calculating the squared deviation of the prediction and the observed
+value at a given time and averages these devations over the length of
+the series. Finally the RMSE will be in the same scale as the original
+time series so its comparison with other series is possible only if they
+share a common scale. RMSE has a direct connection to the L2 norm.
+
+$$ \mathrm{RMSE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}) = \sqrt{\frac{1}{H} \sum^{t+H}_{\tau=t+1} (y_{\tau} - \hat{y}_{\tau})^{2}} $$
+
+**Parameters:** `y`: numpy array, Actual values. `y_hat`: numpy
+array, Predicted values. `mask`: numpy array, Specifies date stamps
+per serie to consider in loss.
+
+**Returns:**
+[`rmse`](https://nixtlaverse.nixtla.io/neuralforecast/losses.numpy.html#rmse):
+numpy array, (single value).\*
+
+
+
+# 2. Percentage errors
+
+These metrics are unit-free, suitable for comparisons across series.
+
+## Mean Absolute Percentage Error
+
+------------------------------------------------------------------------
+
+source
+
+### mape
+
+> ``` text
+> mape (y:numpy.ndarray, y_hat:numpy.ndarray,
+> weights:Optional[numpy.ndarray]=None, axis:Optional[int]=None)
+> ```
+
+\*Mean Absolute Percentage Error
+
+Calculates Mean Absolute Percentage Error between `y` and `y_hat`. MAPE
+measures the relative prediction accuracy of a forecasting method by
+calculating the percentual deviation of the prediction and the observed
+value at a given time and averages these devations over the length of
+the series. The closer to zero an observed value is, the higher penalty
+MAPE loss assigns to the corresponding error.
+
+$$ \mathrm{MAPE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}) = \frac{1}{H} \sum^{t+H}_{\tau=t+1} \frac{|y_{\tau}-\hat{y}_{\tau}|}{|y_{\tau}|} $$
+
+**Parameters:** `y`: numpy array, Actual values. `y_hat`: numpy
+array, Predicted values. `mask`: numpy array, Specifies date stamps
+per serie to consider in loss.
+
+**Returns:**
+[`mape`](https://nixtlaverse.nixtla.io/neuralforecast/losses.numpy.html#mape):
+numpy array, (single value).\*
+
+
+
+## SMAPE
+
+------------------------------------------------------------------------
+
+source
+
+### smape
+
+> ``` text
+> smape (y:numpy.ndarray, y_hat:numpy.ndarray,
+> weights:Optional[numpy.ndarray]=None, axis:Optional[int]=None)
+> ```
+
+\*Symmetric Mean Absolute Percentage Error
+
+Calculates Symmetric Mean Absolute Percentage Error between `y` and
+`y_hat`. SMAPE measures the relative prediction accuracy of a
+forecasting method by calculating the relative deviation of the
+prediction and the observed value scaled by the sum of the absolute
+values for the prediction and observed value at a given time, then
+averages these devations over the length of the series. This allows the
+SMAPE to have bounds between 0% and 200% which is desirable compared to
+normal MAPE that may be undetermined when the target is zero.
+
+$$ \mathrm{sMAPE}_{2}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}) = \frac{1}{H} \sum^{t+H}_{\tau=t+1} \frac{|y_{\tau}-\hat{y}_{\tau}|}{|y_{\tau}|+|\hat{y}_{\tau}|} $$
+
+**Parameters:** `y`: numpy array, Actual values. `y_hat`: numpy
+array, Predicted values. `mask`: numpy array, Specifies date stamps
+per serie to consider in loss.
+
+**Returns:**
+[`smape`](https://nixtlaverse.nixtla.io/neuralforecast/losses.numpy.html#smape):
+numpy array, (single value).
+
+**References:** [Makridakis S., “Accuracy measures: theoretical and
+practical
+concerns”.](https://www.sciencedirect.com/science/article/pii/0169207093900793)\*
+
+# 3. Scale-independent Errors
+
+These metrics measure the relative improvements versus baselines.
+
+## Mean Absolute Scaled Error
+
+------------------------------------------------------------------------
+
+source
+
+### mase
+
+> ``` text
+> mase (y:numpy.ndarray, y_hat:numpy.ndarray, y_train:numpy.ndarray,
+> seasonality:int, weights:Optional[numpy.ndarray]=None,
+> axis:Optional[int]=None)
+> ```
+
+\*Mean Absolute Scaled Error Calculates the Mean Absolute Scaled Error
+between `y` and `y_hat`. MASE measures the relative prediction accuracy
+of a forecasting method by comparinng the mean absolute errors of the
+prediction and the observed value against the mean absolute errors of
+the seasonal naive model. The MASE partially composed the Overall
+Weighted Average (OWA), used in the M4 Competition.
+
+$$ \mathrm{MASE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}, \mathbf{\hat{y}}^{season}_{\tau}) = \frac{1}{H} \sum^{t+H}_{\tau=t+1} \frac{|y_{\tau}-\hat{y}_{\tau}|}{\mathrm{MAE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{season}_{\tau})} $$
+
+**Parameters:** `y`: numpy array, (batch_size, output_size), Actual
+values. `y_hat`: numpy array, (batch_size, output_size)), Predicted
+values. `y_insample`: numpy array, (batch_size, input_size), Actual
+insample Seasonal Naive predictions. `seasonality`: int. Main
+frequency of the time series; Hourly 24, Daily 7, Weekly 52, Monthly 12,
+Quarterly 4, Yearly 1.
+`mask`: numpy array, Specifies date stamps per serie to consider in
+loss.
+
+**Returns:**
+[`mase`](https://nixtlaverse.nixtla.io/neuralforecast/losses.numpy.html#mase):
+numpy array, (single value).
+
+**References:** [Rob J. Hyndman, & Koehler, A. B. “Another look at
+measures of forecast
+accuracy”.](https://www.sciencedirect.com/science/article/pii/S0169207006000239)
+[Spyros Makridakis, Evangelos Spiliotis, Vassilios Assimakopoulos, “The
+M4 Competition: 100,000 time series and 61 forecasting
+methods”.](https://www.sciencedirect.com/science/article/pii/S0169207019301128)\*
+
+
+
+## Relative Mean Absolute Error
+
+------------------------------------------------------------------------
+
+source
+
+### rmae
+
+> ``` text
+> rmae (y:numpy.ndarray, y_hat1:numpy.ndarray, y_hat2:numpy.ndarray,
+> weights:Optional[numpy.ndarray]=None, axis:Optional[int]=None)
+> ```
+
+\*RMAE
+
+Calculates Relative Mean Absolute Error (RMAE) between two sets of
+forecasts (from two different forecasting methods). A number smaller
+than one implies that the forecast in the numerator is better than the
+forecast in the denominator.
+
+$$ \mathrm{rMAE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}, \mathbf{\hat{y}}^{base}_{\tau}) = \frac{1}{H} \sum^{t+H}_{\tau=t+1} \frac{|y_{\tau}-\hat{y}_{\tau}|}{\mathrm{MAE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{base}_{\tau})} $$
+
+**Parameters:** `y`: numpy array, observed values. `y_hat1`:
+numpy array. Predicted values of first model. `y_hat2`: numpy array.
+Predicted values of baseline model. `weights`: numpy array,
+optional. Weights for weighted average. `axis`: None or int,
+optional.Axis or axes along which to average a. The default,
+axis=None, will average over all of the elements of the input array.
+
+**Returns:**
+[`rmae`](https://nixtlaverse.nixtla.io/neuralforecast/losses.numpy.html#rmae):
+numpy array or double.
+
+**References:** [Rob J. Hyndman, & Koehler, A. B. “Another look at
+measures of forecast
+accuracy”.](https://www.sciencedirect.com/science/article/pii/S0169207006000239)\*
+
+
+
+# 4. Probabilistic Errors
+
+These measure absolute deviation non-symmetrically, that produce
+under/over estimation.
+
+## Quantile Loss
+
+------------------------------------------------------------------------
+
+source
+
+### quantile_loss
+
+> ``` text
+> quantile_loss (y:numpy.ndarray, y_hat:numpy.ndarray, q:float=0.5,
+> weights:Optional[numpy.ndarray]=None,
+> axis:Optional[int]=None)
+> ```
+
+\*Quantile Loss
+
+Computes the quantile loss between `y` and `y_hat`. QL measures the
+deviation of a quantile forecast. By weighting the absolute deviation in
+a non symmetric way, the loss pays more attention to under or over
+estimation. A common value for q is 0.5 for the deviation from the
+median (Pinball loss).
+
+$$ \mathrm{QL}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{(q)}_{\tau}) = \frac{1}{H} \sum^{t+H}_{\tau=t+1} \Big( (1-q)\,( \hat{y}^{(q)}_{\tau} - y_{\tau} )_{+} + q\,( y_{\tau} - \hat{y}^{(q)}_{\tau} )_{+} \Big) $$
+
+**Parameters:** `y`: numpy array, Actual values. `y_hat`: numpy
+array, Predicted values. `q`: float, between 0 and 1. The slope of
+the quantile loss, in the context of quantile regression, the q
+determines the conditional quantile level. `mask`: numpy array,
+Specifies date stamps per serie to consider in loss.
+
+**Returns:**
+[`quantile_loss`](https://nixtlaverse.nixtla.io/neuralforecast/losses.numpy.html#quantile_loss):
+numpy array, (single value).
+
+**References:** [Roger Koenker and Gilbert Bassett, Jr., “Regression
+Quantiles”.](https://www.jstor.org/stable/1913643)\*
+
+
+
+## Multi-Quantile Loss
+
+------------------------------------------------------------------------
+
+source
+
+### mqloss
+
+> ``` text
+> mqloss (y:numpy.ndarray, y_hat:numpy.ndarray, quantiles:numpy.ndarray,
+> weights:Optional[numpy.ndarray]=None, axis:Optional[int]=None)
+> ```
+
+\*Multi-Quantile loss
+
+Calculates the Multi-Quantile loss (MQL) between `y` and `y_hat`. MQL
+calculates the average multi-quantile Loss for a given set of quantiles,
+based on the absolute difference between predicted quantiles and
+observed values.
+
+$$ \mathrm{MQL}(\mathbf{y}_{\tau},[\mathbf{\hat{y}}^{(q_{1})}_{\tau}, ... ,\hat{y}^{(q_{n})}_{\tau}]) = \frac{1}{n} \sum_{q_{i}} \mathrm{QL}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{(q_{i})}_{\tau}) $$
+
+The limit behavior of MQL allows to measure the accuracy of a full
+predictive distribution $\mathbf{\hat{F}}_{\tau}$ with the continuous
+ranked probability score (CRPS). This can be achieved through a
+numerical integration technique, that discretizes the quantiles and
+treats the CRPS integral with a left Riemann approximation, averaging
+over uniformly distanced quantiles.
+
+$$ \mathrm{CRPS}(y_{\tau}, \mathbf{\hat{F}}_{\tau}) = \int^{1}_{0} \mathrm{QL}(y_{\tau}, \hat{y}^{(q)}_{\tau}) dq $$
+
+**Parameters:** `y`: numpy array, Actual values. `y_hat`: numpy
+array, Predicted values. `quantiles`: numpy array,(n_quantiles).
+Quantiles to estimate from the distribution of y. `mask`: numpy
+array, Specifies date stamps per serie to consider in loss.
+
+**Returns:**
+[`mqloss`](https://nixtlaverse.nixtla.io/neuralforecast/losses.numpy.html#mqloss):
+numpy array, (single value).
+
+**References:** [Roger Koenker and Gilbert Bassett, Jr., “Regression
+Quantiles”.](https://www.jstor.org/stable/1913643) [James E.
+Matheson and Robert L. Winkler, “Scoring Rules for Continuous
+Probability Distributions”.](https://www.jstor.org/stable/2629907)\*
+
+
+
+# Examples and Validation
+
+
+```python
+import unittest
+import torch as t
+import numpy as np
+
+from neuralforecast.losses.pytorch import (
+ MAE, MSE, RMSE, # unscaled errors
+ MAPE, SMAPE, # percentage errors
+ MASE, # scaled error
+ QuantileLoss, MQLoss # probabilistic errors
+)
+
+from neuralforecast.losses.numpy import (
+ mae, mse, rmse, # unscaled errors
+ mape, smape, # percentage errors
+ mase, # scaled error
+ quantile_loss, mqloss # probabilistic errors
+)
+```
+
diff --git a/neuralforecast/losses.pytorch.html.mdx b/neuralforecast/losses.pytorch.html.mdx
new file mode 100644
index 00000000..b9f6185b
--- /dev/null
+++ b/neuralforecast/losses.pytorch.html.mdx
@@ -0,0 +1,1499 @@
+---
+description: >-
+ NeuralForecast contains a collection PyTorch Loss classes aimed to be used
+ during the models' optimization.
+output-file: losses.pytorch.html
+title: PyTorch Losses
+---
+
+
+The most important train signal is the forecast error, which is the
+difference between the observed value $y_{\tau}$ and the prediction
+$\hat{y}_{\tau}$, at time $y_{\tau}$:
+
+$$e_{\tau} = y_{\tau}-\hat{y}_{\tau} \qquad \qquad \tau \in \{t+1,\dots,t+H \}$$
+
+The train loss summarizes the forecast errors in different train
+optimization objectives.
+
+All the losses are `torch.nn.modules` which helps to automatically moved
+them across CPU/GPU/TPU devices with Pytorch Lightning.
+
+------------------------------------------------------------------------
+
+source
+
+### BasePointLoss
+
+> ``` text
+> BasePointLoss (horizon_weight=None, outputsize_multiplier=None,
+> output_names=None)
+> ```
+
+\*Base class for point loss functions.
+
+**Parameters:** `horizon_weight`: Tensor of size h, weight for each
+timestamp of the forecasting window. `outputsize_multiplier`:
+Multiplier for the output size. `output_names`: Names of the
+outputs. \*
+
+# 1. Scale-dependent Errors
+
+These metrics are on the same scale as the data.
+
+## Mean Absolute Error (MAE)
+
+------------------------------------------------------------------------
+
+source
+
+### MAE.\_\_init\_\_
+
+> ``` text
+> MAE.__init__ (horizon_weight=None)
+> ```
+
+\*Mean Absolute Error
+
+Calculates Mean Absolute Error between `y` and `y_hat`. MAE measures the
+relative prediction accuracy of a forecasting method by calculating the
+deviation of the prediction and the true value at a given time and
+averages these devations over the length of the series.
+
+$$ \mathrm{MAE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}) = \frac{1}{H} \sum^{t+H}_{\tau=t+1} |y_{\tau} - \hat{y}_{\tau}| $$
+
+**Parameters:** `horizon_weight`: Tensor of size h, weight for each
+timestamp of the forecasting window. \*
+
+------------------------------------------------------------------------
+
+source
+
+### MAE.\_\_call\_\_
+
+> ``` text
+> MAE.__call__ (y:torch.Tensor, y_hat:torch.Tensor,
+> mask:Optional[torch.Tensor]=None,
+> y_insample:Optional[torch.Tensor]=None)
+> ```
+
+\***Parameters:** `y`: tensor, Actual values. `y_hat`: tensor,
+Predicted values. `mask`: tensor, Specifies datapoints to consider
+in loss.
+
+**Returns:**
+[`mae`](https://nixtlaverse.nixtla.io/neuralforecast/losses.numpy.html#mae):
+tensor (single value).\*
+
+
+
+## Mean Squared Error (MSE)
+
+------------------------------------------------------------------------
+
+source
+
+### MSE.\_\_init\_\_
+
+> ``` text
+> MSE.__init__ (horizon_weight=None)
+> ```
+
+\*Mean Squared Error
+
+Calculates Mean Squared Error between `y` and `y_hat`. MSE measures the
+relative prediction accuracy of a forecasting method by calculating the
+squared deviation of the prediction and the true value at a given time,
+and averages these devations over the length of the series.
+
+$$ \mathrm{MSE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}) = \frac{1}{H} \sum^{t+H}_{\tau=t+1} (y_{\tau} - \hat{y}_{\tau})^{2} $$
+
+**Parameters:** `horizon_weight`: Tensor of size h, weight for each
+timestamp of the forecasting window. \*
+
+------------------------------------------------------------------------
+
+source
+
+### MSE.\_\_call\_\_
+
+> ``` text
+> MSE.__call__ (y:torch.Tensor, y_hat:torch.Tensor,
+> y_insample:torch.Tensor, mask:Optional[torch.Tensor]=None)
+> ```
+
+\***Parameters:** `y`: tensor, Actual values. `y_hat`: tensor,
+Predicted values. `mask`: tensor, Specifies datapoints to consider
+in loss.
+
+**Returns:**
+[`mse`](https://nixtlaverse.nixtla.io/neuralforecast/losses.numpy.html#mse):
+tensor (single value).\*
+
+
+
+## Root Mean Squared Error (RMSE)
+
+------------------------------------------------------------------------
+
+source
+
+### RMSE.\_\_init\_\_
+
+> ``` text
+> RMSE.__init__ (horizon_weight=None)
+> ```
+
+\*Root Mean Squared Error
+
+Calculates Root Mean Squared Error between `y` and `y_hat`. RMSE
+measures the relative prediction accuracy of a forecasting method by
+calculating the squared deviation of the prediction and the observed
+value at a given time and averages these devations over the length of
+the series. Finally the RMSE will be in the same scale as the original
+time series so its comparison with other series is possible only if they
+share a common scale. RMSE has a direct connection to the L2 norm.
+
+$$ \mathrm{RMSE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}) = \sqrt{\frac{1}{H} \sum^{t+H}_{\tau=t+1} (y_{\tau} - \hat{y}_{\tau})^{2}} $$
+
+**Parameters:** `horizon_weight`: Tensor of size h, weight for each
+timestamp of the forecasting window. \*
+
+------------------------------------------------------------------------
+
+source
+
+### RMSE.\_\_call\_\_
+
+> ``` text
+> RMSE.__call__ (y:torch.Tensor, y_hat:torch.Tensor,
+> mask:Optional[torch.Tensor]=None,
+> y_insample:Optional[torch.Tensor]=None)
+> ```
+
+\***Parameters:** `y`: tensor, Actual values. `y_hat`: tensor,
+Predicted values. `mask`: tensor, Specifies datapoints to consider
+in loss.
+
+**Returns:**
+[`rmse`](https://nixtlaverse.nixtla.io/neuralforecast/losses.numpy.html#rmse):
+tensor (single value).\*
+
+
+
+# 2. Percentage errors
+
+These metrics are unit-free, suitable for comparisons across series.
+
+## Mean Absolute Percentage Error (MAPE)
+
+------------------------------------------------------------------------
+
+source
+
+### MAPE.\_\_init\_\_
+
+> ``` text
+> MAPE.__init__ (horizon_weight=None)
+> ```
+
+\*Mean Absolute Percentage Error
+
+Calculates Mean Absolute Percentage Error between `y` and `y_hat`. MAPE
+measures the relative prediction accuracy of a forecasting method by
+calculating the percentual deviation of the prediction and the observed
+value at a given time and averages these devations over the length of
+the series. The closer to zero an observed value is, the higher penalty
+MAPE loss assigns to the corresponding error.
+
+$$ \mathrm{MAPE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}) = \frac{1}{H} \sum^{t+H}_{\tau=t+1} \frac{|y_{\tau}-\hat{y}_{\tau}|}{|y_{\tau}|} $$
+
+**Parameters:** `horizon_weight`: Tensor of size h, weight for each
+timestamp of the forecasting window.
+
+**References:** [Makridakis S., “Accuracy measures: theoretical and
+practical
+concerns”.](https://www.sciencedirect.com/science/article/pii/0169207093900793)\*
+
+------------------------------------------------------------------------
+
+source
+
+### MAPE.\_\_call\_\_
+
+> ``` text
+> MAPE.__call__ (y:torch.Tensor, y_hat:torch.Tensor,
+> y_insample:torch.Tensor, mask:Optional[torch.Tensor]=None)
+> ```
+
+\***Parameters:** `y`: tensor, Actual values. `y_hat`: tensor,
+Predicted values. `mask`: tensor, Specifies date stamps per serie to
+consider in loss.
+
+**Returns:**
+[`mape`](https://nixtlaverse.nixtla.io/neuralforecast/losses.numpy.html#mape):
+tensor (single value).\*
+
+
+
+## Symmetric MAPE (sMAPE)
+
+------------------------------------------------------------------------
+
+source
+
+### SMAPE.\_\_init\_\_
+
+> ``` text
+> SMAPE.__init__ (horizon_weight=None)
+> ```
+
+\*Symmetric Mean Absolute Percentage Error
+
+Calculates Symmetric Mean Absolute Percentage Error between `y` and
+`y_hat`. SMAPE measures the relative prediction accuracy of a
+forecasting method by calculating the relative deviation of the
+prediction and the observed value scaled by the sum of the absolute
+values for the prediction and observed value at a given time, then
+averages these devations over the length of the series. This allows the
+SMAPE to have bounds between 0% and 200% which is desireble compared to
+normal MAPE that may be undetermined when the target is zero.
+
+$$ \mathrm{sMAPE}_{2}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}) = \frac{1}{H} \sum^{t+H}_{\tau=t+1} \frac{|y_{\tau}-\hat{y}_{\tau}|}{|y_{\tau}|+|\hat{y}_{\tau}|} $$
+
+**Parameters:** `horizon_weight`: Tensor of size h, weight for each
+timestamp of the forecasting window.
+
+**References:** [Makridakis S., “Accuracy measures: theoretical and
+practical
+concerns”.](https://www.sciencedirect.com/science/article/pii/0169207093900793)\*
+
+------------------------------------------------------------------------
+
+source
+
+### SMAPE.\_\_call\_\_
+
+> ``` text
+> SMAPE.__call__ (y:torch.Tensor, y_hat:torch.Tensor,
+> mask:Optional[torch.Tensor]=None,
+> y_insample:Optional[torch.Tensor]=None)
+> ```
+
+\***Parameters:** `y`: tensor, Actual values. `y_hat`: tensor,
+Predicted values. `mask`: tensor, Specifies date stamps per serie to
+consider in loss.
+
+**Returns:**
+[`smape`](https://nixtlaverse.nixtla.io/neuralforecast/losses.numpy.html#smape):
+tensor (single value).\*
+
+# 3. Scale-independent Errors
+
+These metrics measure the relative improvements versus baselines.
+
+## Mean Absolute Scaled Error (MASE)
+
+------------------------------------------------------------------------
+
+source
+
+### MASE.\_\_init\_\_
+
+> ``` text
+> MASE.__init__ (seasonality:int, horizon_weight=None)
+> ```
+
+\*Mean Absolute Scaled Error Calculates the Mean Absolute Scaled Error
+between `y` and `y_hat`. MASE measures the relative prediction accuracy
+of a forecasting method by comparinng the mean absolute errors of the
+prediction and the observed value against the mean absolute errors of
+the seasonal naive model. The MASE partially composed the Overall
+Weighted Average (OWA), used in the M4 Competition.
+
+$$ \mathrm{MASE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}, \mathbf{\hat{y}}^{season}_{\tau}) = \frac{1}{H} \sum^{t+H}_{\tau=t+1} \frac{|y_{\tau}-\hat{y}_{\tau}|}{\mathrm{MAE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{season}_{\tau})} $$
+
+**Parameters:** `seasonality`: int. Main frequency of the time
+series; Hourly 24, Daily 7, Weekly 52, Monthly 12, Quarterly 4,
+Yearly 1. `horizon_weight`: Tensor of size h, weight for each timestamp
+of the forecasting window.
+
+**References:** [Rob J. Hyndman, & Koehler, A. B. “Another look at
+measures of forecast
+accuracy”.](https://www.sciencedirect.com/science/article/pii/S0169207006000239)
+[Spyros Makridakis, Evangelos Spiliotis, Vassilios Assimakopoulos, “The
+M4 Competition: 100,000 time series and 61 forecasting
+methods”.](https://www.sciencedirect.com/science/article/pii/S0169207019301128)\*
+
+------------------------------------------------------------------------
+
+source
+
+### MASE.\_\_call\_\_
+
+> ``` text
+> MASE.__call__ (y:torch.Tensor, y_hat:torch.Tensor,
+> y_insample:torch.Tensor, mask:Optional[torch.Tensor]=None)
+> ```
+
+\***Parameters:** `y`: tensor (batch_size, output_size), Actual
+values. `y_hat`: tensor (batch_size, output_size)), Predicted
+values. `y_insample`: tensor (batch_size, input_size), Actual
+insample values. `mask`: tensor, Specifies date stamps per serie to
+consider in loss.
+
+**Returns:**
+[`mase`](https://nixtlaverse.nixtla.io/neuralforecast/losses.numpy.html#mase):
+tensor (single value).\*
+
+
+
+## Relative Mean Squared Error (relMSE)
+
+------------------------------------------------------------------------
+
+source
+
+### relMSE.\_\_init\_\_
+
+> ``` text
+> relMSE.__init__ (y_train=None, horizon_weight=None)
+> ```
+
+\*Relative Mean Squared Error Computes Relative Mean Squared Error
+(relMSE), as proposed by Hyndman & Koehler (2006) as an alternative to
+percentage errors, to avoid measure unstability.
+$$
+ \mathrm{relMSE}(\mathbf{y}, \mathbf{\hat{y}}, \mathbf{\hat{y}}^{benchmark}) =
+\frac{\mathrm{MSE}(\mathbf{y}, \mathbf{\hat{y}})}{\mathrm{MSE}(\mathbf{y}, \mathbf{\hat{y}}^{benchmark})}
+$$
+
+**Parameters:** `y_train`: numpy array, deprecated.
+`horizon_weight`: Tensor of size h, weight for each timestamp of the
+forecasting window.
+
+**References:** - [Hyndman, R. J and Koehler, A. B. (2006). “Another
+look at measures of forecast accuracy”, International Journal of
+Forecasting, Volume 22, Issue
+4.](https://www.sciencedirect.com/science/article/pii/S0169207006000239) -
+[Kin G. Olivares, O. Nganba Meetei, Ruijun Ma, Rohan Reddy, Mengfei Cao,
+Lee Dicker. “Probabilistic Hierarchical Forecasting with Deep Poisson
+Mixtures. Submitted to the International Journal Forecasting, Working
+paper available at arxiv.](https://arxiv.org/pdf/2110.13179.pdf)\*
+
+------------------------------------------------------------------------
+
+source
+
+### relMSE.\_\_call\_\_
+
+> ``` text
+> relMSE.__call__ (y:torch.Tensor, y_hat:torch.Tensor,
+> y_benchmark:torch.Tensor,
+> mask:Optional[torch.Tensor]=None)
+> ```
+
+\***Parameters:** `y`: tensor (batch_size, output_size), Actual
+values. `y_hat`: tensor (batch_size, output_size)), Predicted
+values. `y_benchmark`: tensor (batch_size, output_size), Benchmark
+predicted values. `mask`: tensor, Specifies date stamps per serie to
+consider in loss.
+
+**Returns:**
+[`relMSE`](https://nixtlaverse.nixtla.io/neuralforecast/losses.pytorch.html#relmse):
+tensor (single value).\*
+
+# 4. Probabilistic Errors
+
+These methods use statistical approaches for estimating unknown
+probability distributions using observed data.
+
+Maximum likelihood estimation involves finding the parameter values that
+maximize the likelihood function, which measures the probability of
+obtaining the observed data given the parameter values. MLE has good
+theoretical properties and efficiency under certain satisfied
+assumptions.
+
+On the non-parametric approach, quantile regression measures
+non-symmetrically deviation, producing under/over estimation.
+
+## Quantile Loss
+
+------------------------------------------------------------------------
+
+source
+
+### QuantileLoss.\_\_init\_\_
+
+> ``` text
+> QuantileLoss.__init__ (q, horizon_weight=None)
+> ```
+
+\*Quantile Loss
+
+Computes the quantile loss between `y` and `y_hat`. QL measures the
+deviation of a quantile forecast. By weighting the absolute deviation in
+a non symmetric way, the loss pays more attention to under or over
+estimation. A common value for q is 0.5 for the deviation from the
+median (Pinball loss).
+
+$$ \mathrm{QL}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{(q)}_{\tau}) = \frac{1}{H} \sum^{t+H}_{\tau=t+1} \Big( (1-q)\,( \hat{y}^{(q)}_{\tau} - y_{\tau} )_{+} + q\,( y_{\tau} - \hat{y}^{(q)}_{\tau} )_{+} \Big) $$
+
+**Parameters:** `q`: float, between 0 and 1. The slope of the
+quantile loss, in the context of quantile regression, the q determines
+the conditional quantile level. `horizon_weight`: Tensor of size h,
+weight for each timestamp of the forecasting window.
+
+**References:** [Roger Koenker and Gilbert Bassett, Jr., “Regression
+Quantiles”.](https://www.jstor.org/stable/1913643)\*
+
+------------------------------------------------------------------------
+
+source
+
+### QuantileLoss.\_\_call\_\_
+
+> ``` text
+> QuantileLoss.__call__ (y:torch.Tensor, y_hat:torch.Tensor,
+> y_insample:torch.Tensor,
+> mask:Optional[torch.Tensor]=None)
+> ```
+
+\***Parameters:** `y`: tensor, Actual values. `y_hat`: tensor,
+Predicted values. `mask`: tensor, Specifies datapoints to consider
+in loss.
+
+**Returns:**
+[`quantile_loss`](https://nixtlaverse.nixtla.io/neuralforecast/losses.numpy.html#quantile_loss):
+tensor (single value).\*
+
+
+
+## Multi Quantile Loss (MQLoss)
+
+------------------------------------------------------------------------
+
+source
+
+### MQLoss.\_\_init\_\_
+
+> ``` text
+> MQLoss.__init__ (level=[80, 90], quantiles=None, horizon_weight=None)
+> ```
+
+\*Multi-Quantile loss
+
+Calculates the Multi-Quantile loss (MQL) between `y` and `y_hat`. MQL
+calculates the average multi-quantile Loss for a given set of quantiles,
+based on the absolute difference between predicted quantiles and
+observed values.
+
+$$ \mathrm{MQL}(\mathbf{y}_{\tau},[\mathbf{\hat{y}}^{(q_{1})}_{\tau}, ... ,\hat{y}^{(q_{n})}_{\tau}]) = \frac{1}{n} \sum_{q_{i}} \mathrm{QL}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{(q_{i})}_{\tau}) $$
+
+The limit behavior of MQL allows to measure the accuracy of a full
+predictive distribution $\mathbf{\hat{F}}_{\tau}$ with the continuous
+ranked probability score (CRPS). This can be achieved through a
+numerical integration technique, that discretizes the quantiles and
+treats the CRPS integral with a left Riemann approximation, averaging
+over uniformly distanced quantiles.
+
+$$ \mathrm{CRPS}(y_{\tau}, \mathbf{\hat{F}}_{\tau}) = \int^{1}_{0} \mathrm{QL}(y_{\tau}, \hat{y}^{(q)}_{\tau}) dq $$
+
+**Parameters:** `level`: int list \[0,100\]. Probability levels for
+prediction intervals (Defaults median). `quantiles`: float list \[0.,
+1.\]. Alternative to level, quantiles to estimate from y distribution.
+`horizon_weight`: Tensor of size h, weight for each timestamp of the
+forecasting window.
+
+**References:** [Roger Koenker and Gilbert Bassett, Jr., “Regression
+Quantiles”.](https://www.jstor.org/stable/1913643) [James E.
+Matheson and Robert L. Winkler, “Scoring Rules for Continuous
+Probability Distributions”.](https://www.jstor.org/stable/2629907)\*
+
+------------------------------------------------------------------------
+
+source
+
+### MQLoss.\_\_call\_\_
+
+> ``` text
+> MQLoss.__call__ (y:torch.Tensor, y_hat:torch.Tensor,
+> y_insample:torch.Tensor,
+> mask:Optional[torch.Tensor]=None)
+> ```
+
+\***Parameters:** `y`: tensor, Actual values. `y_hat`: tensor,
+Predicted values. `mask`: tensor, Specifies date stamps per serie to
+consider in loss.
+
+**Returns:**
+[`mqloss`](https://nixtlaverse.nixtla.io/neuralforecast/losses.numpy.html#mqloss):
+tensor (single value).\*
+
+
+
+## Implicit Quantile Loss (IQLoss)
+
+------------------------------------------------------------------------
+
+source
+
+### QuantileLayer
+
+> ``` text
+> QuantileLayer (num_output:int, cos_embedding_dim:int=128)
+> ```
+
+\*Implicit Quantile Layer from the paper
+`IQN for Distributional Reinforcement Learning`
+(https://arxiv.org/abs/1806.06923) by Dabney et al. 2018.
+
+Code from GluonTS:
+https://github.com/awslabs/gluonts/blob/dev/src/gluonts/torch/distributions/implicit_quantile_network.py\*
+
+------------------------------------------------------------------------
+
+source
+
+### IQLoss.\_\_init\_\_
+
+> ``` text
+> IQLoss.__init__ (cos_embedding_dim=64, concentration0=1.0,
+> concentration1=1.0, horizon_weight=None)
+> ```
+
+\*Implicit Quantile Loss
+
+Computes the quantile loss between `y` and `y_hat`, with the quantile
+`q` provided as an input to the network. IQL measures the deviation of a
+quantile forecast. By weighting the absolute deviation in a non
+symmetric way, the loss pays more attention to under or over estimation.
+
+$$ \mathrm{QL}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{(q)}_{\tau}) = \frac{1}{H} \sum^{t+H}_{\tau=t+1} \Big( (1-q)\,( \hat{y}^{(q)}_{\tau} - y_{\tau} )_{+} + q\,( y_{\tau} - \hat{y}^{(q)}_{\tau} )_{+} \Big) $$
+
+**Parameters:** `quantile_sampling`: str, default=‘uniform’,
+sampling distribution used to sample the quantiles during training.
+Choose from \[‘uniform’, ‘beta’\]. `horizon_weight`: Tensor of size
+h, weight for each timestamp of the forecasting window.
+
+**References:** [Gouttes, Adèle, Kashif Rasul, Mateusz Koren,
+Johannes Stephan, and Tofigh Naghibi, “Probabilistic Time Series
+Forecasting with Implicit Quantile
+Networks”.](http://arxiv.org/abs/2107.03743)\*
+
+------------------------------------------------------------------------
+
+source
+
+### IQLoss.\_\_call\_\_
+
+> ``` text
+> IQLoss.__call__ (y:torch.Tensor, y_hat:torch.Tensor,
+> y_insample:torch.Tensor,
+> mask:Optional[torch.Tensor]=None)
+> ```
+
+\***Parameters:** `y`: tensor, Actual values. `y_hat`: tensor,
+Predicted values. `mask`: tensor, Specifies datapoints to consider
+in loss.
+
+**Returns:**
+[`quantile_loss`](https://nixtlaverse.nixtla.io/neuralforecast/losses.numpy.html#quantile_loss):
+tensor (single value).\*
+
+## DistributionLoss
+
+------------------------------------------------------------------------
+
+source
+
+### DistributionLoss.\_\_init\_\_
+
+> ``` text
+> DistributionLoss.__init__ (distribution, level=[80, 90], quantiles=None,
+> num_samples=1000, return_params=False,
+> horizon_weight=None, **distribution_kwargs)
+> ```
+
+\*DistributionLoss
+
+This PyTorch module wraps the `torch.distribution` classes allowing it
+to interact with NeuralForecast models modularly. It shares the negative
+log-likelihood as the optimization objective and a sample method to
+generate empirically the quantiles defined by the `level` list.
+
+Additionally, it implements a distribution transformation that
+factorizes the scale-dependent likelihood parameters into a base scale
+and a multiplier efficiently learnable within the network’s
+non-linearities operating ranges.
+
+Available distributions: - Poisson - Normal - StudentT -
+NegativeBinomial - Tweedie - Bernoulli (Temporal
+Classifiers) - ISQF (Incremental Spline Quantile Function)
+
+**Parameters:** `distribution`: str, identifier of a
+torch.distributions.Distribution class. `level`: float list
+\[0,100\], confidence levels for prediction intervals. `quantiles`:
+float list \[0,1\], alternative to level list, target quantiles.
+`num_samples`: int=500, number of samples for the empirical
+quantiles. `return_params`: bool=False, wether or not return the
+Distribution parameters. `horizon_weight`: Tensor of size h, weight
+for each timestamp of the forecasting window.
+
+**References:** - [PyTorch Probability Distributions Package:
+StudentT.](https://pytorch.org/docs/stable/distributions.html#studentt) -
+[David Salinas, Valentin Flunkert, Jan Gasthaus, Tim Januschowski
+(2020). “DeepAR: Probabilistic forecasting with autoregressive recurrent
+networks”. International Journal of
+Forecasting.](https://www.sciencedirect.com/science/article/pii/S0169207019301888) -
+[Park, Youngsuk, Danielle Maddix, François-Xavier Aubet, Kelvin Kan, Jan
+Gasthaus, and Yuyang Wang (2022). “Learning Quantile Functions without
+Quantile Crossing for Distribution-free Time Series
+Forecasting”.](https://proceedings.mlr.press/v151/park22a.html)\*
+
+------------------------------------------------------------------------
+
+source
+
+### DistributionLoss.sample
+
+> ``` text
+> DistributionLoss.sample (distr_args:torch.Tensor,
+> num_samples:Optional[int]=None)
+> ```
+
+\*Construct the empirical quantiles from the estimated Distribution,
+sampling from it `num_samples` independently.
+
+**Parameters** `distr_args`: Constructor arguments for the
+underlying Distribution type. `num_samples`: int, overwrite number
+of samples for the empirical quantiles.
+
+**Returns** `samples`: tensor, shape \[B,H,`num_samples`\].
+`quantiles`: tensor, empirical quantiles defined by `levels`. \*
+
+------------------------------------------------------------------------
+
+source
+
+### DistributionLoss.\_\_call\_\_
+
+> ``` text
+> DistributionLoss.__call__ (y:torch.Tensor, distr_args:torch.Tensor,
+> mask:Optional[torch.Tensor]=None)
+> ```
+
+\*Computes the negative log-likelihood objective function. To estimate
+the following predictive distribution:
+
+$$\mathrm{P}(\mathbf{y}_{\tau}\,|\,\theta) \quad \mathrm{and} \quad -\log(\mathrm{P}(\mathbf{y}_{\tau}\,|\,\theta))$$
+
+where $\theta$ represents the distributions parameters. It aditionally
+summarizes the objective signal using a weighted average using the
+`mask` tensor.
+
+**Parameters** `y`: tensor, Actual values. `distr_args`:
+Constructor arguments for the underlying Distribution type. `loc`:
+Optional tensor, of the same shape as the batch_shape + event_shape of
+the resulting distribution. `scale`: Optional tensor, of the same
+shape as the batch_shape+event_shape of the resulting distribution.
+`mask`: tensor, Specifies date stamps per serie to consider in loss.
+
+**Returns** `loss`: scalar, weighted loss function against which
+backpropagation will be performed. \*
+
+## Poisson Mixture Mesh (PMM)
+
+------------------------------------------------------------------------
+
+source
+
+### PMM.\_\_init\_\_
+
+> ``` text
+> PMM.__init__ (n_components=10, level=[80, 90], quantiles=None,
+> num_samples=1000, return_params=False,
+> batch_correlation=False, horizon_correlation=False,
+> weighted=False)
+> ```
+
+\*Poisson Mixture Mesh
+
+This Poisson Mixture statistical model assumes independence across
+groups of data $\mathcal{G}=\{[g_{i}]\}$, and estimates relationships
+within the group.
+
+$$
+ \mathrm{P}\left(\mathbf{y}_{[b][t+1:t+H]}\right) =
+\prod_{ [g_{i}] \in \mathcal{G}} \mathrm{P} \left(\mathbf{y}_{[g_{i}][\tau]} \right) =
+\prod_{\beta\in[g_{i}]}
+\left(\sum_{k=1}^{K} w_k \prod_{(\beta,\tau) \in [g_i][t+1:t+H]} \mathrm{Poisson}(y_{\beta,\tau}, \hat{\lambda}_{\beta,\tau,k}) \right)
+$$
+
+**Parameters:** `n_components`: int=10, the number of mixture
+components. `level`: float list \[0,100\], confidence levels for
+prediction intervals. `quantiles`: float list \[0,1\], alternative
+to level list, target quantiles. `return_params`: bool=False, wether
+or not return the Distribution parameters. `batch_correlation`:
+bool=False, wether or not model batch correlations.
+`horizon_correlation`: bool=False, wether or not model horizon
+correlations.
+
+**References:** [Kin G. Olivares, O. Nganba Meetei, Ruijun Ma, Rohan
+Reddy, Mengfei Cao, Lee Dicker. Probabilistic Hierarchical Forecasting
+with Deep Poisson Mixtures. Submitted to the International Journal
+Forecasting, Working paper available at
+arxiv.](https://arxiv.org/pdf/2110.13179.pdf)\*
+
+------------------------------------------------------------------------
+
+source
+
+### PMM.sample
+
+> ``` text
+> PMM.sample (distr_args:torch.Tensor, num_samples:Optional[int]=None)
+> ```
+
+\*Construct the empirical quantiles from the estimated Distribution,
+sampling from it `num_samples` independently.
+
+**Parameters** `distr_args`: Constructor arguments for the
+underlying Distribution type. `num_samples`: int, overwrite number
+of samples for the empirical quantiles.
+
+**Returns** `samples`: tensor, shape \[B,H,`num_samples`\].
+`quantiles`: tensor, empirical quantiles defined by `levels`. \*
+
+------------------------------------------------------------------------
+
+source
+
+### PMM.\_\_call\_\_
+
+> ``` text
+> PMM.__call__ (y:torch.Tensor, distr_args:torch.Tensor,
+> mask:Optional[torch.Tensor]=None)
+> ```
+
+\*Computes the negative log-likelihood objective function. To estimate
+the following predictive distribution:
+
+$$\mathrm{P}(\mathbf{y}_{\tau}\,|\,\theta) \quad \mathrm{and} \quad -\log(\mathrm{P}(\mathbf{y}_{\tau}\,|\,\theta))$$
+
+where $\theta$ represents the distributions parameters. It aditionally
+summarizes the objective signal using a weighted average using the
+`mask` tensor.
+
+**Parameters** `y`: tensor, Actual values. `distr_args`:
+Constructor arguments for the underlying Distribution type. `mask`:
+tensor, Specifies date stamps per serie to consider in loss.
+
+**Returns** `loss`: scalar, weighted loss function against which
+backpropagation will be performed. \*
+
+
+
+## Gaussian Mixture Mesh (GMM)
+
+------------------------------------------------------------------------
+
+source
+
+### GMM.\_\_init\_\_
+
+> ``` text
+> GMM.__init__ (n_components=1, level=[80, 90], quantiles=None,
+> num_samples=1000, return_params=False,
+> batch_correlation=False, horizon_correlation=False,
+> weighted=False)
+> ```
+
+\*Gaussian Mixture Mesh
+
+This Gaussian Mixture statistical model assumes independence across
+groups of data $\mathcal{G}=\{[g_{i}]\}$, and estimates relationships
+within the group.
+
+$$
+ \mathrm{P}\left(\mathbf{y}_{[b][t+1:t+H]}\right) =
+\prod_{ [g_{i}] \in \mathcal{G}} \mathrm{P}\left(\mathbf{y}_{[g_{i}][\tau]}\right)=
+\prod_{\beta\in[g_{i}]}
+\left(\sum_{k=1}^{K} w_k \prod_{(\beta,\tau) \in [g_i][t+1:t+H]}
+\mathrm{Gaussian}(y_{\beta,\tau}, \hat{\mu}_{\beta,\tau,k}, \sigma_{\beta,\tau,k})\right)
+$$
+
+**Parameters:** `n_components`: int=10, the number of mixture
+components. `level`: float list \[0,100\], confidence levels for
+prediction intervals. `quantiles`: float list \[0,1\], alternative
+to level list, target quantiles. `return_params`: bool=False, wether
+or not return the Distribution parameters. `batch_correlation`:
+bool=False, wether or not model batch correlations.
+`horizon_correlation`: bool=False, wether or not model horizon
+correlations.
+
+**References:** [Kin G. Olivares, O. Nganba Meetei, Ruijun Ma, Rohan
+Reddy, Mengfei Cao, Lee Dicker. Probabilistic Hierarchical Forecasting
+with Deep Poisson Mixtures. Submitted to the International Journal
+Forecasting, Working paper available at
+arxiv.](https://arxiv.org/pdf/2110.13179.pdf)\*
+
+------------------------------------------------------------------------
+
+source
+
+### GMM.sample
+
+> ``` text
+> GMM.sample (distr_args:torch.Tensor, num_samples:Optional[int]=None)
+> ```
+
+\*Construct the empirical quantiles from the estimated Distribution,
+sampling from it `num_samples` independently.
+
+**Parameters** `distr_args`: Constructor arguments for the
+underlying Distribution type. `num_samples`: int, overwrite number
+of samples for the empirical quantiles.
+
+**Returns** `samples`: tensor, shape \[B,H,`num_samples`\].
+`quantiles`: tensor, empirical quantiles defined by `levels`. \*
+
+------------------------------------------------------------------------
+
+source
+
+### GMM.\_\_call\_\_
+
+> ``` text
+> GMM.__call__ (y:torch.Tensor, distr_args:torch.Tensor,
+> mask:Optional[torch.Tensor]=None)
+> ```
+
+\*Computes the negative log-likelihood objective function. To estimate
+the following predictive distribution:
+
+$$\mathrm{P}(\mathbf{y}_{\tau}\,|\,\theta) \quad \mathrm{and} \quad -\log(\mathrm{P}(\mathbf{y}_{\tau}\,|\,\theta))$$
+
+where $\theta$ represents the distributions parameters. It aditionally
+summarizes the objective signal using a weighted average using the
+`mask` tensor.
+
+**Parameters** `y`: tensor, Actual values. `distr_args`:
+Constructor arguments for the underlying Distribution type. `mask`:
+tensor, Specifies date stamps per serie to consider in loss.
+
+**Returns** `loss`: scalar, weighted loss function against which
+backpropagation will be performed. \*
+
+
+
+## Negative Binomial Mixture Mesh (NBMM)
+
+------------------------------------------------------------------------
+
+source
+
+### NBMM.\_\_init\_\_
+
+> ``` text
+> NBMM.__init__ (n_components=1, level=[80, 90], quantiles=None,
+> num_samples=1000, return_params=False, weighted=False)
+> ```
+
+\*Negative Binomial Mixture Mesh
+
+This N. Binomial Mixture statistical model assumes independence across
+groups of data $\mathcal{G}=\{[g_{i}]\}$, and estimates relationships
+within the group.
+
+$$
+ \mathrm{P}\left(\mathbf{y}_{[b][t+1:t+H]}\right) =
+\prod_{ [g_{i}] \in \mathcal{G}} \mathrm{P}\left(\mathbf{y}_{[g_{i}][\tau]}\right)=
+\prod_{\beta\in[g_{i}]}
+\left(\sum_{k=1}^{K} w_k \prod_{(\beta,\tau) \in [g_i][t+1:t+H]}
+\mathrm{NBinomial}(y_{\beta,\tau}, \hat{r}_{\beta,\tau,k}, \hat{p}_{\beta,\tau,k})\right)
+$$
+
+**Parameters:** `n_components`: int=10, the number of mixture
+components. `level`: float list \[0,100\], confidence levels for
+prediction intervals. `quantiles`: float list \[0,1\], alternative
+to level list, target quantiles. `return_params`: bool=False, wether
+or not return the Distribution parameters.
+
+**References:** [Kin G. Olivares, O. Nganba Meetei, Ruijun Ma, Rohan
+Reddy, Mengfei Cao, Lee Dicker. Probabilistic Hierarchical Forecasting
+with Deep Poisson Mixtures. Submitted to the International Journal
+Forecasting, Working paper available at
+arxiv.](https://arxiv.org/pdf/2110.13179.pdf)\*
+
+------------------------------------------------------------------------
+
+source
+
+### NBMM.sample
+
+> ``` text
+> NBMM.sample (distr_args:torch.Tensor, num_samples:Optional[int]=None)
+> ```
+
+\*Construct the empirical quantiles from the estimated Distribution,
+sampling from it `num_samples` independently.
+
+**Parameters** `distr_args`: Constructor arguments for the
+underlying Distribution type. `num_samples`: int, overwrite number
+of samples for the empirical quantiles.
+
+**Returns** `samples`: tensor, shape \[B,H,`num_samples`\].
+`quantiles`: tensor, empirical quantiles defined by `levels`. \*
+
+------------------------------------------------------------------------
+
+source
+
+### NBMM.\_\_call\_\_
+
+> ``` text
+> NBMM.__call__ (y:torch.Tensor, distr_args:torch.Tensor,
+> mask:Optional[torch.Tensor]=None)
+> ```
+
+\*Computes the negative log-likelihood objective function. To estimate
+the following predictive distribution:
+
+$$\mathrm{P}(\mathbf{y}_{\tau}\,|\,\theta) \quad \mathrm{and} \quad -\log(\mathrm{P}(\mathbf{y}_{\tau}\,|\,\theta))$$
+
+where $\theta$ represents the distributions parameters. It aditionally
+summarizes the objective signal using a weighted average using the
+`mask` tensor.
+
+**Parameters** `y`: tensor, Actual values. `distr_args`:
+Constructor arguments for the underlying Distribution type. `mask`:
+tensor, Specifies date stamps per serie to consider in loss.
+
+**Returns** `loss`: scalar, weighted loss function against which
+backpropagation will be performed. \*
+
+# 5. Robustified Errors
+
+This type of errors from robust statistic focus on methods resistant to
+outliers and violations of assumptions, providing reliable estimates and
+inferences. Robust estimators are used to reduce the impact of outliers,
+offering more stable results.
+
+## Huber Loss
+
+------------------------------------------------------------------------
+
+source
+
+### HuberLoss.\_\_init\_\_
+
+> ``` text
+> HuberLoss.__init__ (delta:float=1.0, horizon_weight=None)
+> ```
+
+\*Huber Loss
+
+The Huber loss, employed in robust regression, is a loss function that
+exhibits reduced sensitivity to outliers in data when compared to the
+squared error loss. This function is also refered as SmoothL1.
+
+The Huber loss function is quadratic for small errors and linear for
+large errors, with equal values and slopes of the different sections at
+the two points where
+$(y_{\tau}-\hat{y}_{\tau})^{2}$=$|y_{\tau}-\hat{y}_{\tau}|$.
+
+$$
+ L_{\delta}(y_{\tau},\; \hat{y}_{\tau})
+=\begin{cases}{\frac{1}{2}}(y_{\tau}-\hat{y}_{\tau})^{2}\;{\text{for }}|y_{\tau}-\hat{y}_{\tau}|\leq \delta \\
+\delta \ \cdot \left(|y_{\tau}-\hat{y}_{\tau}|-{\frac {1}{2}}\delta \right),\;{\text{otherwise.}}\end{cases}
+$$
+
+where $\delta$ is a threshold parameter that determines the point at
+which the loss transitions from quadratic to linear, and can be tuned to
+control the trade-off between robustness and accuracy in the
+predictions.
+
+**Parameters:** `delta`: float=1.0, Specifies the threshold at which
+to change between delta-scaled L1 and L2 loss. `horizon_weight`: Tensor
+of size h, weight for each timestamp of the forecasting window.
+
+**References:** [Huber Peter, J (1964). “Robust Estimation of a
+Location Parameter”. Annals of
+Statistics](https://projecteuclid.org/journals/annals-of-mathematical-statistics/volume-35/issue-1/Robust-Estimation-of-a-Location-Parameter/10.1214/aoms/1177703732.full)\*
+
+------------------------------------------------------------------------
+
+source
+
+### HuberLoss.\_\_call\_\_
+
+> ``` text
+> HuberLoss.__call__ (y:torch.Tensor, y_hat:torch.Tensor,
+> y_insample:torch.Tensor,
+> mask:Optional[torch.Tensor]=None)
+> ```
+
+\***Parameters:** `y`: tensor, Actual values. `y_hat`: tensor,
+Predicted values. `mask`: tensor, Specifies date stamps per serie to
+consider in loss.
+
+**Returns:** `huber_loss`: tensor (single value).\*
+
+
+
+## Tukey Loss
+
+------------------------------------------------------------------------
+
+source
+
+### TukeyLoss.\_\_init\_\_
+
+> ``` text
+> TukeyLoss.__init__ (c:float=4.685, normalize:bool=True)
+> ```
+
+\*Tukey Loss
+
+The Tukey loss function, also known as Tukey’s biweight function, is a
+robust statistical loss function used in robust statistics. Tukey’s loss
+exhibits quadratic behavior near the origin, like the Huber loss;
+however, it is even more robust to outliers as the loss for large
+residuals remains constant instead of scaling linearly.
+
+The parameter $c$ in Tukey’s loss determines the ‘’saturation’’ point of
+the function: Higher values of $c$ enhance sensitivity, while lower
+values increase resistance to outliers.
+
+$$
+ L_{c}(y_{\tau},\; \hat{y}_{\tau})
+=\begin{cases}{
+\frac{c^{2}}{6}} \left[1-(\frac{y_{\tau}-\hat{y}_{\tau}}{c})^{2} \right]^{3} \;\text{for } |y_{\tau}-\hat{y}_{\tau}|\leq c \\
+\frac{c^{2}}{6} \qquad \text{otherwise.} \end{cases}
+$$
+
+Please note that the Tukey loss function assumes the data to be
+stationary or normalized beforehand. If the error values are excessively
+large, the algorithm may need help to converge during optimization. It
+is advisable to employ small learning rates.
+
+**Parameters:** `c`: float=4.685, Specifies the Tukey loss’
+threshold on which residuals are no longer considered. `normalize`:
+bool=True, Wether normalization is performed within Tukey loss’
+computation.
+
+**References:** [Beaton, A. E., and Tukey, J. W. (1974). “The
+Fitting of Power Series, Meaning Polynomials, Illustrated on
+Band-Spectroscopic Data.”](https://www.jstor.org/stable/1267936)\*
+
+------------------------------------------------------------------------
+
+source
+
+### TukeyLoss.\_\_call\_\_
+
+> ``` text
+> TukeyLoss.__call__ (y:torch.Tensor, y_hat:torch.Tensor,
+> y_insample:torch.Tensor,
+> mask:Optional[torch.Tensor]=None)
+> ```
+
+\***Parameters:** `y`: tensor, Actual values. `y_hat`: tensor,
+Predicted values. `mask`: tensor, Specifies date stamps per serie to
+consider in loss.
+
+**Returns:** `tukey_loss`: tensor (single value).\*
+
+
+
+## Huberized Quantile Loss
+
+------------------------------------------------------------------------
+
+source
+
+### HuberQLoss.\_\_init\_\_
+
+> ``` text
+> HuberQLoss.__init__ (q, delta:float=1.0, horizon_weight=None)
+> ```
+
+\*Huberized Quantile Loss
+
+The Huberized quantile loss is a modified version of the quantile loss
+function that combines the advantages of the quantile loss and the Huber
+loss. It is commonly used in regression tasks, especially when dealing
+with data that contains outliers or heavy tails.
+
+The Huberized quantile loss between `y` and `y_hat` measure the Huber
+Loss in a non-symmetric way. The loss pays more attention to
+under/over-estimation depending on the quantile parameter $q$; and
+controls the trade-off between robustness and accuracy in the
+predictions with the parameter $delta$.
+
+$$
+ \mathrm{HuberQL}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{(q)}_{\tau}) =
+(1-q)\, L_{\delta}(y_{\tau},\; \hat{y}^{(q)}_{\tau}) \mathbb{1}\{ \hat{y}^{(q)}_{\tau} \geq y_{\tau} \} +
+q\, L_{\delta}(y_{\tau},\; \hat{y}^{(q)}_{\tau}) \mathbb{1}\{ \hat{y}^{(q)}_{\tau} < y_{\tau} \}
+$$
+
+**Parameters:** `delta`: float=1.0, Specifies the threshold at which
+to change between delta-scaled L1 and L2 loss. `q`: float, between 0
+and 1. The slope of the quantile loss, in the context of quantile
+regression, the q determines the conditional quantile level.
+`horizon_weight`: Tensor of size h, weight for each timestamp of the
+forecasting window.
+
+**References:** [Huber Peter, J (1964). “Robust Estimation of a
+Location Parameter”. Annals of
+Statistics](https://projecteuclid.org/journals/annals-of-mathematical-statistics/volume-35/issue-1/Robust-Estimation-of-a-Location-Parameter/10.1214/aoms/1177703732.full)
+[Roger Koenker and Gilbert Bassett, Jr., “Regression
+Quantiles”.](https://www.jstor.org/stable/1913643)\*
+
+------------------------------------------------------------------------
+
+source
+
+### HuberQLoss.\_\_call\_\_
+
+> ``` text
+> HuberQLoss.__call__ (y:torch.Tensor, y_hat:torch.Tensor,
+> y_insample:torch.Tensor,
+> mask:Optional[torch.Tensor]=None)
+> ```
+
+\***Parameters:** `y`: tensor, Actual values. `y_hat`: tensor,
+Predicted values. `mask`: tensor, Specifies datapoints to consider
+in loss.
+
+**Returns:** `huber_qloss`: tensor (single value).\*
+
+
+
+## Huberized MQLoss
+
+------------------------------------------------------------------------
+
+source
+
+### HuberMQLoss.\_\_init\_\_
+
+> ``` text
+> HuberMQLoss.__init__ (level=[80, 90], quantiles=None, delta:float=1.0,
+> horizon_weight=None)
+> ```
+
+\*Huberized Multi-Quantile loss
+
+The Huberized Multi-Quantile loss (HuberMQL) is a modified version of
+the multi-quantile loss function that combines the advantages of the
+quantile loss and the Huber loss. HuberMQL is commonly used in
+regression tasks, especially when dealing with data that contains
+outliers or heavy tails. The loss function pays more attention to
+under/over-estimation depending on the quantile list
+$[q_{1},q_{2},\dots]$ parameter. It controls the trade-off between
+robustness and prediction accuracy with the parameter $\delta$.
+
+$$
+ \mathrm{HuberMQL}_{\delta}(\mathbf{y}_{\tau},[\mathbf{\hat{y}}^{(q_{1})}_{\tau}, ... ,\hat{y}^{(q_{n})}_{\tau}]) =
+\frac{1}{n} \sum_{q_{i}} \mathrm{HuberQL}_{\delta}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{(q_{i})}_{\tau})
+$$
+
+**Parameters:** `level`: int list \[0,100\]. Probability levels for
+prediction intervals (Defaults median). `quantiles`: float list \[0.,
+1.\]. Alternative to level, quantiles to estimate from y distribution.
+`delta`: float=1.0, Specifies the threshold at which to change between
+delta-scaled L1 and L2 loss.
+`horizon_weight`: Tensor of size h, weight for each timestamp of the
+forecasting window.
+
+**References:** [Huber Peter, J (1964). “Robust Estimation of a
+Location Parameter”. Annals of
+Statistics](https://projecteuclid.org/journals/annals-of-mathematical-statistics/volume-35/issue-1/Robust-Estimation-of-a-Location-Parameter/10.1214/aoms/1177703732.full)
+[Roger Koenker and Gilbert Bassett, Jr., “Regression
+Quantiles”.](https://www.jstor.org/stable/1913643)\*
+
+------------------------------------------------------------------------
+
+source
+
+### HuberMQLoss.\_\_call\_\_
+
+> ``` text
+> HuberMQLoss.__call__ (y:torch.Tensor, y_hat:torch.Tensor,
+> y_insample:torch.Tensor,
+> mask:Optional[torch.Tensor]=None)
+> ```
+
+\***Parameters:** `y`: tensor, Actual values. `y_hat`: tensor,
+Predicted values. `mask`: tensor, Specifies date stamps per serie to
+consider in loss.
+
+**Returns:** `hmqloss`: tensor (single value).\*
+
+
+
+## Huberized IQLoss
+
+------------------------------------------------------------------------
+
+source
+
+### HuberIQLoss.\_\_init\_\_
+
+> ``` text
+> HuberIQLoss.__init__ (cos_embedding_dim=64, concentration0=1.0,
+> concentration1=1.0, delta=1.0, horizon_weight=None)
+> ```
+
+\*Implicit Huber Quantile Loss
+
+Computes the huberized quantile loss between `y` and `y_hat`, with the
+quantile `q` provided as an input to the network. HuberIQLoss measures
+the deviation of a huberized quantile forecast. By weighting the
+absolute deviation in a non symmetric way, the loss pays more attention
+to under or over estimation.
+
+$$
+ \mathrm{HuberQL}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{(q)}_{\tau}) =
+(1-q)\, L_{\delta}(y_{\tau},\; \hat{y}^{(q)}_{\tau}) \mathbb{1}\{ \hat{y}^{(q)}_{\tau} \geq y_{\tau} \} +
+q\, L_{\delta}(y_{\tau},\; \hat{y}^{(q)}_{\tau}) \mathbb{1}\{ \hat{y}^{(q)}_{\tau} < y_{\tau} \}
+$$
+
+**Parameters:** `quantile_sampling`: str, default=‘uniform’,
+sampling distribution used to sample the quantiles during training.
+Choose from \[‘uniform’, ‘beta’\]. `horizon_weight`: Tensor of size
+h, weight for each timestamp of the forecasting window. `delta`:
+float=1.0, Specifies the threshold at which to change between
+delta-scaled L1 and L2 loss.
+
+**References:** [Gouttes, Adèle, Kashif Rasul, Mateusz Koren,
+Johannes Stephan, and Tofigh Naghibi, “Probabilistic Time Series
+Forecasting with Implicit Quantile
+Networks”.](http://arxiv.org/abs/2107.03743) [Huber Peter, J (1964).
+“Robust Estimation of a Location Parameter”. Annals of
+Statistics](https://projecteuclid.org/journals/annals-of-mathematical-statistics/volume-35/issue-1/Robust-Estimation-of-a-Location-Parameter/10.1214/aoms/1177703732.full)
+[Roger Koenker and Gilbert Bassett, Jr., “Regression
+Quantiles”.](https://www.jstor.org/stable/1913643)\*
+
+------------------------------------------------------------------------
+
+source
+
+### HuberIQLoss.\_\_call\_\_
+
+> ``` text
+> HuberIQLoss.__call__ (y:torch.Tensor, y_hat:torch.Tensor,
+> y_insample:torch.Tensor,
+> mask:Optional[torch.Tensor]=None)
+> ```
+
+\***Parameters:** `y`: tensor, Actual values. `y_hat`: tensor,
+Predicted values. `mask`: tensor, Specifies datapoints to consider
+in loss.
+
+**Returns:** `huber_qloss`: tensor (single value).\*
+
+# 6. Others
+
+## Accuracy
+
+------------------------------------------------------------------------
+
+source
+
+### Accuracy.\_\_init\_\_
+
+> ``` text
+> Accuracy.__init__ ()
+> ```
+
+\*Accuracy
+
+Computes the accuracy between categorical `y` and `y_hat`. This
+evaluation metric is only meant for evalution, as it is not
+differentiable.
+
+$$ \mathrm{Accuracy}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}) = \frac{1}{H} \sum^{t+H}_{\tau=t+1} \mathrm{1}\{\mathbf{y}_{\tau}==\mathbf{\hat{y}}_{\tau}\} $$\*
+
+------------------------------------------------------------------------
+
+source
+
+### Accuracy.\_\_call\_\_
+
+> ``` text
+> Accuracy.__call__ (y:torch.Tensor, y_hat:torch.Tensor,
+> y_insample:torch.Tensor,
+> mask:Optional[torch.Tensor]=None)
+> ```
+
+\***Parameters:** `y`: tensor, Actual values. `y_hat`: tensor,
+Predicted values. `mask`: tensor, Specifies date stamps per serie to
+consider in loss.
+
+**Returns:** `accuracy`: tensor (single value).\*
+
+## Scaled Continuous Ranked Probability Score (sCRPS)
+
+------------------------------------------------------------------------
+
+source
+
+### sCRPS.\_\_init\_\_
+
+> ``` text
+> sCRPS.__init__ (level=[80, 90], quantiles=None)
+> ```
+
+\*Scaled Continues Ranked Probability Score
+
+Calculates a scaled variation of the CRPS, as proposed by Rangapuram
+(2021), to measure the accuracy of predicted quantiles `y_hat` compared
+to the observation `y`.
+
+This metric averages percentual weighted absolute deviations as defined
+by the quantile losses.
+
+$$
+ \mathrm{sCRPS}(\mathbf{\hat{y}}^{(q)}_{\tau}, \mathbf{y}_{\tau}) = \frac{2}{N} \sum_{i}
+\int^{1}_{0}
+\frac{\mathrm{QL}(\mathbf{\hat{y}}^{(q}_{\tau} y_{i,\tau})_{q}}{\sum_{i} | y_{i,\tau} |} dq
+$$
+
+where $\mathbf{\hat{y}}^{(q}_{\tau}$ is the estimated quantile, and
+$y_{i,\tau}$ are the target variable realizations.
+
+**Parameters:** `level`: int list \[0,100\]. Probability levels for
+prediction intervals (Defaults median). `quantiles`: float list \[0.,
+1.\]. Alternative to level, quantiles to estimate from y distribution.
+
+**References:** - [Gneiting, Tilmann. (2011). “Quantiles as optimal
+point forecasts”. International Journal of
+Forecasting.](https://www.sciencedirect.com/science/article/pii/S0169207010000063) -
+[Spyros Makridakis, Evangelos Spiliotis, Vassilios Assimakopoulos, Zhi
+Chen, Anil Gaba, Ilia Tsetlin, Robert L. Winkler. (2022). “The M5
+uncertainty competition: Results, findings and conclusions”.
+International Journal of
+Forecasting.](https://www.sciencedirect.com/science/article/pii/S0169207021001722) -
+[Syama Sundar Rangapuram, Lucien D Werner, Konstantinos Benidis, Pedro
+Mercado, Jan Gasthaus, Tim Januschowski. (2021). “End-to-End Learning of
+Coherent Probabilistic Forecasts for Hierarchical Time Series”.
+Proceedings of the 38th International Conference on Machine Learning
+(ICML).](https://proceedings.mlr.press/v139/rangapuram21a.html)\*
+
+------------------------------------------------------------------------
+
+source
+
+### sCRPS.\_\_call\_\_
+
+> ``` text
+> sCRPS.__call__ (y:torch.Tensor, y_hat:torch.Tensor,
+> y_insample:torch.Tensor,
+> mask:Optional[torch.Tensor]=None)
+> ```
+
+\***Parameters:** `y`: tensor, Actual values. `y_hat`: tensor,
+Predicted values. `mask`: tensor, Specifies date stamps per series
+to consider in loss.
+
+**Returns:** `scrps`: tensor (single value).\*
+
diff --git a/neuralforecast/mint.json b/neuralforecast/mint.json
new file mode 100644
index 00000000..18f86829
--- /dev/null
+++ b/neuralforecast/mint.json
@@ -0,0 +1,168 @@
+{
+ "$schema": "https://mintlify.com/schema.json",
+ "name": "Nixtla",
+ "logo": {
+ "light": "/light.png",
+ "dark": "/dark.png"
+ },
+ "favicon": "/favicon.svg",
+ "colors": {
+ "primary": "#0E0E0E",
+ "light": "#FAFAFA",
+ "dark": "#0E0E0E",
+ "anchors": {
+ "from": "#2AD0CA",
+ "to": "#0E00F8"
+ }
+ },
+ "topbarCtaButton": {
+ "type": "github",
+ "url": "https://github.com/Nixtla/neuralforecast"
+ },
+ "topAnchor": {
+ "name": "NeuralForecast",
+ "icon": "brain-circuit"
+ },
+ "navigation": [
+ {
+ "group": "Getting Started",
+ "pages": [
+ "docs/getting-started/introduction.html",
+ "docs/getting-started/quickstart.html",
+ "docs/getting-started/installation.html",
+ "docs/getting-started/datarequirements.html"
+ ]
+ },
+ {
+ "group": "Capabilities",
+ "pages": [
+ "docs/capabilities/overview.html",
+ "docs/capabilities/objectives.html",
+ "docs/capabilities/exogenous_variables.html",
+ "docs/capabilities/cross_validation.html",
+ "docs/capabilities/hyperparameter_tuning.html",
+ "docs/capabilities/predictinsample.html",
+ "docs/capabilities/save_load_models.html",
+ "docs/capabilities/time_series_scaling.html"
+ ]
+ },
+ {
+ "group": "Tutorials",
+ "pages": [
+ {
+ "group":"Forecasting",
+ "pages":[
+ "docs/tutorials/getting_started_complete.html",
+ "docs/tutorials/cross_validation_tutorial.html",
+ "docs/tutorials/longhorizon_nhits.html",
+ "docs/tutorials/longhorizon_transformers.html",
+ "docs/tutorials/forecasting_tft.html",
+ "docs/tutorials/multivariate_tsmixer.html"
+ ]
+ },
+ {
+ "group":"Probabilistic Forecasting",
+ "pages":[
+ "docs/tutorials/uncertainty_quantification.html",
+ "docs/tutorials/longhorizon_probabilistic.html",
+ "docs/tutorials/conformal_prediction.html"
+ ]
+ },
+ {
+ "group":"Special Topics",
+ "pages":[
+ "docs/tutorials/hierarchical_forecasting.html",
+ "docs/tutorials/distributed_neuralforecast.html",
+ "docs/tutorials/intermittent_data.html",
+ "docs/tutorials/using_mlflow.html",
+ "docs/tutorials/robust_forecasting.html",
+ "docs/tutorials/interpretable_decompositions.html",
+ "docs/tutorials/comparing_methods.html",
+ "docs/tutorials/temporal_classification.html",
+ "docs/tutorials/transfer_learning.html",
+ "docs/tutorials/adding_models.html",
+ "docs/tutorials/large_datasets.html"
+ ]
+ }
+ ]
+ },
+ {
+ "group": "Use cases",
+ "pages": [
+ "docs/use-cases/electricity_peak_forecasting.html",
+ "docs/use-cases/predictive_maintenance.html"
+ ]
+ },
+ {
+ "group": "API Reference",
+ "pages": [
+ "docs/tutorials/neuralforecasting_map.html",
+ "core.html",
+ {
+ "group": "Models",
+ "pages": [
+ "models.autoformer.html",
+ "models.bitcn.html",
+ "models.deepar.html",
+ "models.deepnpts.html",
+ "models.dilated_rnn.html",
+ "models.dlinear.html",
+ "models.fedformer.html",
+ "models.gru.html",
+ "models.hint.html",
+ "models.informer.html",
+ "models.itransformer.html",
+ "models.kan.html",
+ "models.lstm.html",
+ "models.mlp.html",
+ "models.mlpmultivariate.html",
+ "models.nbeats.html",
+ "models.nbeatsx.html",
+ "models.nhits.html",
+ "models.nlinear.html",
+ "models.patchtst.html",
+ "models.rmok.html",
+ "models.rnn.html",
+ "models.softs.html",
+ "models.stemgnn.html",
+ "models.tcn.html",
+ "models.tft.html",
+ "models.tide.html",
+ "models.timellm.html",
+ "models.timemixer.html",
+ "models.timesnet.html",
+ "models.timexer.html",
+ "models.tsmixer.html",
+ "models.tsmixerx.html",
+ "models.vanillatransformer.html"
+ ]
+ },
+ "models.html",
+ {
+ "group": "Train/Evaluation",
+ "pages": [
+ "losses.pytorch.html",
+ "losses.numpy.html"
+ ]
+ },
+ {
+ "group": "Common Components",
+ "pages": [
+ "common.base_auto.html",
+ "common.base_recurrent.html",
+ "common.base_windows.html",
+ "common.scalers.html",
+ "common.modules.html"
+ ]
+ },
+ {
+ "group": "Utils",
+ "pages": [
+ "tsdataset.html",
+ "utils.html"
+ ]
+ }
+ ]
+ }
+ ]
+}
diff --git a/neuralforecast/models.autoformer.html.mdx b/neuralforecast/models.autoformer.html.mdx
new file mode 100644
index 00000000..6dbdd835
--- /dev/null
+++ b/neuralforecast/models.autoformer.html.mdx
@@ -0,0 +1,374 @@
+---
+output-file: models.autoformer.html
+title: Autoformer
+---
+
+
+The Autoformer model tackles the challenge of finding reliable
+dependencies on intricate temporal patterns of long-horizon forecasting.
+
+The architecture has the following distinctive features: - In-built
+progressive decomposition in trend and seasonal compontents based on a
+moving average filter. - Auto-Correlation mechanism that discovers the
+period-based dependencies by calculating the autocorrelation and
+aggregating similar sub-series based on the periodicity. - Classic
+encoder-decoder proposed by Vaswani et al. (2017) with a multi-head
+attention mechanism.
+
+The Autoformer model utilizes a three-component approach to define its
+embedding: - It employs encoded autoregressive features obtained from a
+convolution network. - Absolute positional embeddings obtained from
+calendar features are utilized.
+
+**References** - [Wu, Haixu, Jiehui Xu, Jianmin Wang, and Mingsheng
+Long. “Autoformer: Decomposition transformers with auto-correlation for
+long-term series
+forecasting”](https://proceedings.neurips.cc/paper/2021/hash/bcc0d400288793e8bdcd7c19a8ac0c2b-Abstract.html)
+
+
+
+Figure 1. Autoformer
+Architecture.
+
+
+## 1. Auxiliary Functions
+
+------------------------------------------------------------------------
+
+source
+
+### Decoder
+
+> ``` text
+> Decoder (layers, norm_layer=None, projection=None)
+> ```
+
+*Autoformer decoder*
+
+------------------------------------------------------------------------
+
+source
+
+### DecoderLayer
+
+> ``` text
+> DecoderLayer (self_attention, cross_attention, hidden_size, c_out,
+> conv_hidden_size=None, MovingAvg=25, dropout=0.1,
+> activation='relu')
+> ```
+
+*Autoformer decoder layer with the progressive decomposition
+architecture*
+
+------------------------------------------------------------------------
+
+source
+
+### Encoder
+
+> ``` text
+> Encoder (attn_layers, conv_layers=None, norm_layer=None)
+> ```
+
+*Autoformer encoder*
+
+------------------------------------------------------------------------
+
+source
+
+### EncoderLayer
+
+> ``` text
+> EncoderLayer (attention, hidden_size, conv_hidden_size=None,
+> MovingAvg=25, dropout=0.1, activation='relu')
+> ```
+
+*Autoformer encoder layer with the progressive decomposition
+architecture*
+
+------------------------------------------------------------------------
+
+source
+
+### LayerNorm
+
+> ``` text
+> LayerNorm (channels)
+> ```
+
+*Special designed layernorm for the seasonal part*
+
+------------------------------------------------------------------------
+
+source
+
+### AutoCorrelationLayer
+
+> ``` text
+> AutoCorrelationLayer (correlation, hidden_size, n_head, d_keys=None,
+> d_values=None)
+> ```
+
+*Auto Correlation Layer*
+
+------------------------------------------------------------------------
+
+source
+
+### AutoCorrelation
+
+> ``` text
+> AutoCorrelation (mask_flag=True, factor=1, scale=None,
+> attention_dropout=0.1, output_attention=False)
+> ```
+
+*AutoCorrelation Mechanism with the following two phases: (1)
+period-based dependencies discovery (2) time delay aggregation This
+block can replace the self-attention family mechanism seamlessly.*
+
+## 2. Autoformer
+
+------------------------------------------------------------------------
+
+source
+
+### Autoformer
+
+> ``` text
+> Autoformer (h:int, input_size:int, stat_exog_list=None,
+> hist_exog_list=None, futr_exog_list=None,
+> exclude_insample_y=False,
+> decoder_input_size_multiplier:float=0.5, hidden_size:int=128,
+> dropout:float=0.05, factor:int=3, n_head:int=4,
+> conv_hidden_size:int=32, activation:str='gelu',
+> encoder_layers:int=2, decoder_layers:int=1,
+> MovingAvg_window:int=25, loss=MAE(), valid_loss=None,
+> max_steps:int=5000, learning_rate:float=0.0001,
+> num_lr_decays:int=-1, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size:int=32,
+> valid_batch_size:Optional[int]=None, windows_batch_size=1024,
+> inference_windows_batch_size=1024,
+> start_padding_enabled=False, step_size:int=1,
+> scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*Autoformer
+
+The Autoformer model tackles the challenge of finding reliable
+dependencies on intricate temporal patterns of long-horizon forecasting.
+
+The architecture has the following distinctive features: - In-built
+progressive decomposition in trend and seasonal compontents based on a
+moving average filter. - Auto-Correlation mechanism that discovers the
+period-based dependencies by calculating the autocorrelation and
+aggregating similar sub-series based on the periodicity. - Classic
+encoder-decoder proposed by Vaswani et al. (2017) with a multi-head
+attention mechanism.
+
+The Autoformer model utilizes a three-component approach to define its
+embedding: - It employs encoded autoregressive features obtained from a
+convolution network. - Absolute positional embeddings obtained from
+calendar features are utilized.
+
+*Parameters:* `h`: int, forecast horizon. `input_size`: int,
+maximum sequence length for truncated train backpropagation. Default -1
+uses all history. `futr_exog_list`: str list, future exogenous
+columns. `hist_exog_list`: str list, historic exogenous columns.
+`stat_exog_list`: str list, static exogenous columns.
+`exclude_insample_y`: bool=False, the model skips the autoregressive
+features y\[t-input_size:t\] if True.
+`decoder_input_size_multiplier`: float = 0.5, . `hidden_size`:
+int=128, units of embeddings and encoders. `n_head`: int=4, controls
+number of multi-head’s attention. `dropout`: float (0, 1), dropout
+throughout Autoformer architecture. `factor`: int=3, Probsparse
+attention factor. `conv_hidden_size`: int=32, channels of the
+convolutional encoder. `activation`: str=`GELU`, activation from
+\[‘ReLU’, ‘Softplus’, ‘Tanh’, ‘SELU’, ‘LeakyReLU’, ‘PReLU’, ‘Sigmoid’,
+‘GELU’\]. `encoder_layers`: int=2, number of layers for the TCN
+encoder. `decoder_layers`: int=1, number of layers for the MLP
+decoder. `MovingAvg_window`: int=25, window size for the moving
+average filter. `loss`: PyTorch module, instantiated train loss
+class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module, instantiated validation loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=1024, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=1024, number of windows to sample in
+each inference batch. `start_padding_enabled`: bool=False, if True,
+the model will pad the time series with zeros at the beginning, by input
+size. `scaler_type`: str=‘robust’, type of scaler for temporal
+inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`. `dataloader_kwargs`:
+dict, optional, list of parameters passed into the PyTorch Lightning
+dataloader by the `TimeSeriesDataLoader`. `**trainer_kwargs`: int,
+keyword trainer arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+``` text
+*References*
+- [Wu, Haixu, Jiehui Xu, Jianmin Wang, and Mingsheng Long. "Autoformer: Decomposition transformers with auto-correlation for long-term series forecasting"](https://proceedings.neurips.cc/paper/2021/hash/bcc0d400288793e8bdcd7c19a8ac0c2b-Abstract.html) *
+```
+
+------------------------------------------------------------------------
+
+### Autoformer.fit
+
+> ``` text
+> Autoformer.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### Autoformer.predict
+
+> ``` text
+> Autoformer.predict (dataset, test_size=None, step_size=1,
+> random_seed=None, quantiles=None,
+> **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import Autoformer
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic, augment_calendar_df
+
+AirPassengersPanel, calendar_cols = augment_calendar_df(df=AirPassengersPanel, freq='M')
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = Autoformer(h=12,
+ input_size=24,
+ hidden_size = 16,
+ conv_hidden_size = 32,
+ n_head=2,
+ loss=MAE(),
+ futr_exog_list=calendar_cols,
+ scaler_type='robust',
+ learning_rate=1e-3,
+ max_steps=300,
+ val_check_steps=50,
+ early_stop_patience_steps=2)
+
+nf = NeuralForecast(
+ models=[model],
+ freq='ME'
+)
+nf.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+forecasts = nf.predict(futr_df=Y_test_df)
+
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+if model.loss.is_distribution_output:
+ plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+ plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+ plt.plot(plot_df['ds'], plot_df['Autoformer-median'], c='blue', label='median')
+ plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['Autoformer-lo-90'][-12:].values,
+ y2=plot_df['Autoformer-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+ plt.grid()
+ plt.legend()
+ plt.plot()
+else:
+ plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+ plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+ plt.plot(plot_df['ds'], plot_df['Autoformer'], c='blue', label='Forecast')
+ plt.legend()
+ plt.grid()
+```
+
diff --git a/neuralforecast/models.bitcn.html.mdx b/neuralforecast/models.bitcn.html.mdx
new file mode 100644
index 00000000..c55c7fb4
--- /dev/null
+++ b/neuralforecast/models.bitcn.html.mdx
@@ -0,0 +1,273 @@
+---
+output-file: models.bitcn.html
+title: BiTCN
+---
+
+
+Bidirectional Temporal Convolutional Network (BiTCN) is a forecasting
+architecture based on two temporal convolutional networks (TCNs). The
+first network (‘forward’) encodes future covariates of the time series,
+whereas the second network (‘backward’) encodes past observations and
+covariates. This method allows to preserve the temporal information of
+sequence data, and is computationally more efficient than common RNN
+methods (LSTM, GRU, …). As compared to Transformer-based methods, BiTCN
+has a lower space complexity, i.e. it requires orders of magnitude less
+parameters.
+
+This model may be a good choice if you seek a small model (small amount
+of trainable parameters) with few hyperparameters to tune (only 2).
+
+**References** -[Olivier Sprangers, Sebastian Schelter, Maarten de
+Rijke (2023). Parameter-Efficient Deep Probabilistic Forecasting.
+International Journal of Forecasting 39, no. 1 (1 January 2023): 332–45.
+URL:
+https://doi.org/10.1016/j.ijforecast.2021.11.011.](https://doi.org/10.1016/j.ijforecast.2021.11.011)
+-[Shaojie Bai, Zico Kolter, Vladlen Koltun. (2018). An Empirical
+Evaluation of Generic Convolutional and Recurrent Networks for Sequence
+Modeling. Computing Research Repository, abs/1803.01271. URL:
+https://arxiv.org/abs/1803.01271.](https://arxiv.org/abs/1803.01271)
+-[van den Oord, A., Dieleman, S., Zen, H., Simonyan, K., Vinyals, O.,
+Graves, A., Kalchbrenner, N., Senior, A. W., & Kavukcuoglu, K. (2016).
+Wavenet: A generative model for raw audio. Computing Research
+Repository, abs/1609.03499. URL: http://arxiv.org/abs/1609.03499.
+arXiv:1609.03499.](https://arxiv.org/abs/1609.03499)
+
+
+
+Figure 1. Visualization of a stack of
+dilated causal convolutional layers.
+
+
+## 1. Auxiliary Functions
+
+------------------------------------------------------------------------
+
+source
+
+### TCNCell
+
+> ``` text
+> TCNCell (in_channels, out_channels, kernel_size, padding, dilation, mode,
+> groups, dropout)
+> ```
+
+*Temporal Convolutional Network Cell, consisting of CustomConv1D
+modules.*
+
+------------------------------------------------------------------------
+
+source
+
+### CustomConv1d
+
+> ``` text
+> CustomConv1d (in_channels, out_channels, kernel_size, padding=0,
+> dilation=1, mode='backward', groups=1)
+> ```
+
+*Forward- and backward looking Conv1D*
+
+## 2. BiTCN
+
+------------------------------------------------------------------------
+
+source
+
+### BiTCN
+
+> ``` text
+> BiTCN (h:int, input_size:int, hidden_size:int=16, dropout:float=0.5,
+> futr_exog_list=None, hist_exog_list=None, stat_exog_list=None,
+> exclude_insample_y=False, loss=MAE(), valid_loss=None,
+> max_steps:int=1000, learning_rate:float=0.001,
+> num_lr_decays:int=-1, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size:int=32,
+> valid_batch_size:Optional[int]=None, windows_batch_size=1024,
+> inference_windows_batch_size=1024, start_padding_enabled=False,
+> step_size:int=1, scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*BiTCN
+
+Bidirectional Temporal Convolutional Network (BiTCN) is a forecasting
+architecture based on two temporal convolutional networks (TCNs). The
+first network (‘forward’) encodes future covariates of the time series,
+whereas the second network (‘backward’) encodes past observations and
+covariates. This is a univariate model.
+
+**Parameters:** `h`: int, forecast horizon. `input_size`: int,
+considered autorregresive inputs (lags), y=\[1,2,3,4\] input_size=2 -\>
+lags=\[1,2\]. `hidden_size`: int=16, units for the TCN’s hidden
+state size. `dropout`: float=0.1, dropout rate used for the dropout
+layers throughout the architecture. `futr_exog_list`: str list,
+future exogenous columns. `hist_exog_list`: str list, historic
+exogenous columns. `stat_exog_list`: str list, static exogenous
+columns. `exclude_insample_y`: bool=False, the model skips the
+autoregressive features y\[t-input_size:t\] if True. `loss`: PyTorch
+module, instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=1024, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=1024, number of windows to sample in
+each inference batch, -1 uses all. `start_padding_enabled`:
+bool=False, if True, the model will pad the time series with zeros at
+the beginning, by input size. `step_size`: int=1, step size between
+each window of temporal data. `scaler_type`: str=‘identity’, type of
+scaler for temporal inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`. `dataloader_kwargs`:
+dict, optional, list of parameters passed into the PyTorch Lightning
+dataloader by the `TimeSeriesDataLoader`. `**trainer_kwargs`: int,
+keyword trainer arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+**References**
+- [Olivier Sprangers, Sebastian Schelter, Maarten de Rijke (2023).
+Parameter-Efficient Deep Probabilistic Forecasting. International
+Journal of Forecasting 39, no. 1 (1 January 2023): 332–45. URL:
+https://doi.org/10.1016/j.ijforecast.2021.11.011.](https://doi.org/10.1016/j.ijforecast.2021.11.011) \*
+
+------------------------------------------------------------------------
+
+### BiTCN.fit
+
+> ``` text
+> BiTCN.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### BiTCN.predict
+
+> ``` text
+> BiTCN.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.losses.pytorch import GMM
+from neuralforecast.models import BiTCN
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+fcst = NeuralForecast(
+ models=[
+ BiTCN(h=12,
+ input_size=24,
+ loss=GMM(n_components=7, level=[80,90]),
+ max_steps=100,
+ scaler_type='standard',
+ futr_exog_list=['y_[lag12]'],
+ hist_exog_list=None,
+ stat_exog_list=['airline1'],
+ windows_batch_size=2048,
+ val_check_steps=10,
+ early_stop_patience_steps=-1,
+ ),
+ ],
+ freq='ME'
+)
+fcst.fit(df=Y_train_df, static_df=AirPassengersStatic)
+forecasts = fcst.predict(futr_df=Y_test_df)
+
+# Plot quantile predictions
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['BiTCN-median'], c='blue', label='median')
+plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['BiTCN-lo-90'][-12:].values,
+ y2=plot_df['BiTCN-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+plt.legend()
+plt.grid()
+```
+
diff --git a/neuralforecast/models.deepar.html.mdx b/neuralforecast/models.deepar.html.mdx
new file mode 100644
index 00000000..0a001e3b
--- /dev/null
+++ b/neuralforecast/models.deepar.html.mdx
@@ -0,0 +1,285 @@
+---
+output-file: models.deepar.html
+title: DeepAR
+---
+
+
+The DeepAR model produces probabilistic forecasts based on an
+autoregressive recurrent neural network optimized on panel data using
+cross-learning. DeepAR obtains its forecast distribution uses a Markov
+Chain Monte Carlo sampler with the following conditional probability:
+$$\mathbb{P}(\mathbf{y}_{[t+1:t+H]}|\;\mathbf{y}_{[:t]},\; \mathbf{x}^{(f)}_{[:t+H]},\; \mathbf{x}^{(s)})$$
+
+where $\mathbf{x}^{(s)}$ are static exogenous inputs,
+$\mathbf{x}^{(f)}_{[:t+H]}$ are future exogenous available at the time
+of the prediction. The predictions are obtained by transforming the
+hidden states $\mathbf{h}_{t}$ into predictive distribution parameters
+$\theta_{t}$, and then generating samples $\mathbf{\hat{y}}_{[t+1:t+H]}$
+through Monte Carlo sampling trajectories.
+
+$$
+
+\begin{align}
+\mathbf{h}_{t} &= \textrm{RNN}([\mathbf{y}_{t},\mathbf{x}^{(f)}_{t+1},\mathbf{x}^{(s)}], \mathbf{h}_{t-1})\\
+\mathbf{\theta}_{t}&=\textrm{Linear}(\mathbf{h}_{t}) \\
+\hat{y}_{t+1}&=\textrm{sample}(\;\mathrm{P}(y_{t+1}\;|\;\mathbf{\theta}_{t})\;)
+\end{align}
+
+$$
+
+**References** - [David Salinas, Valentin Flunkert, Jan Gasthaus,
+Tim Januschowski (2020). “DeepAR: Probabilistic forecasting with
+autoregressive recurrent networks”. International Journal of
+Forecasting.](https://www.sciencedirect.com/science/article/pii/S0169207019301888) -
+[Alexander Alexandrov et. al (2020). “GluonTS: Probabilistic and Neural
+Time Series Modeling in Python”. Journal of Machine Learning
+Research.](https://www.jmlr.org/papers/v21/19-820.html)
+
+> **Exogenous Variables, Losses, and Parameters Availability**
+>
+> Given the sampling procedure during inference, DeepAR only supports
+> [`DistributionLoss`](https://nixtlaverse.nixtla.io/neuralforecast/losses.pytorch.html#distributionloss)
+> as training loss.
+>
+> Note that DeepAR generates a non-parametric forecast distribution
+> using Monte Carlo. We use this sampling procedure also during
+> validation to make it closer to the inference procedure. Therefore,
+> only the
+> [`MQLoss`](https://nixtlaverse.nixtla.io/neuralforecast/losses.pytorch.html#mqloss)
+> is available for validation.
+>
+> Aditionally, Monte Carlo implies that historic exogenous variables are
+> not available for the model.
+
+
+
+Figure 1. DeepAR model, during training
+the optimization signal comes from likelihood of observations, during
+inference a recurrent multi-step strategy is used to generate predictive
+distributions.
+
+
+------------------------------------------------------------------------
+
+source
+
+### Decoder
+
+> ``` text
+> Decoder (in_features, out_features, hidden_size, hidden_layers)
+> ```
+
+\*Multi-Layer Perceptron Decoder
+
+**Parameters:** `in_features`: int, dimension of input.
+`out_features`: int, dimension of output. `hidden_size`: int,
+dimension of hidden layers. `num_layers`: int, number of hidden
+layers. \*
+
+------------------------------------------------------------------------
+
+source
+
+### DeepAR
+
+> ``` text
+> DeepAR (h, input_size:int=-1, h_train:int=1, lstm_n_layers:int=2,
+> lstm_hidden_size:int=128, lstm_dropout:float=0.1,
+> decoder_hidden_layers:int=0, decoder_hidden_size:int=0,
+> trajectory_samples:int=100, stat_exog_list=None,
+> hist_exog_list=None, futr_exog_list=None,
+> exclude_insample_y=False, loss=DistributionLoss(),
+> valid_loss=MAE(), max_steps:int=1000, learning_rate:float=0.001,
+> num_lr_decays:int=3, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size:int=32,
+> valid_batch_size:Optional[int]=None, windows_batch_size:int=1024,
+> inference_windows_batch_size:int=-1, start_padding_enabled=False,
+> step_size:int=1, scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader=False, alias:Optional[str]=None, optimizer=None,
+> optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*DeepAR
+
+**Parameters:** `h`: int, Forecast horizon. `input_size`: int,
+maximum sequence length for truncated train backpropagation. Default -1
+uses 3 \* horizon `h_train`: int, maximum sequence length for
+truncated train backpropagation. Default 1. `lstm_n_layers`: int=2,
+number of LSTM layers. `lstm_hidden_size`: int=128, LSTM hidden
+size. `lstm_dropout`: float=0.1, LSTM dropout.
+`decoder_hidden_layers`: int=0, number of decoder MLP hidden layers.
+Default: 0 for linear layer. `decoder_hidden_size`: int=0, decoder
+MLP hidden size. Default: 0 for linear layer. `trajectory_samples`:
+int=100, number of Monte Carlo trajectories during inference.
+`stat_exog_list`: str list, static exogenous columns.
+`hist_exog_list`: str list, historic exogenous columns.
+`futr_exog_list`: str list, future exogenous columns.
+`exclude_insample_y`: bool=False, the model skips the autoregressive
+features y\[t-input_size:t\] if True. `loss`: PyTorch module,
+instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=1024, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=-1, number of windows to sample in
+each inference batch, -1 uses all. `start_padding_enabled`:
+bool=False, if True, the model will pad the time series with zeros at
+the beginning, by input size. `step_size`: int=1, step size between
+each window of temporal data. `scaler_type`: str=‘identity’, type of
+scaler for temporal inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`. `dataloader_kwargs`:
+dict, optional, list of parameters passed into the PyTorch Lightning
+dataloader by the `TimeSeriesDataLoader`. `**trainer_kwargs`: int,
+keyword trainer arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+**References** - [David Salinas, Valentin Flunkert, Jan Gasthaus,
+Tim Januschowski (2020). “DeepAR: Probabilistic forecasting with
+autoregressive recurrent networks”. International Journal of
+Forecasting.](https://www.sciencedirect.com/science/article/pii/S0169207019301888) -
+[Alexander Alexandrov et. al (2020). “GluonTS: Probabilistic and Neural
+Time Series Modeling in Python”. Journal of Machine Learning
+Research.](https://www.jmlr.org/papers/v21/19-820.html) \*
+
+------------------------------------------------------------------------
+
+### DeepAR.fit
+
+> ``` text
+> DeepAR.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### DeepAR.predict
+
+> ``` text
+> DeepAR.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import DeepAR
+from neuralforecast.losses.pytorch import DistributionLoss, MQLoss
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+nf = NeuralForecast(
+ models=[DeepAR(h=12,
+ input_size=24,
+ lstm_n_layers=1,
+ trajectory_samples=100,
+ loss=DistributionLoss(distribution='StudentT', level=[80, 90], return_params=True),
+ valid_loss=MQLoss(level=[80, 90]),
+ learning_rate=0.005,
+ stat_exog_list=['airline1'],
+ futr_exog_list=['trend'],
+ max_steps=100,
+ val_check_steps=10,
+ early_stop_patience_steps=-1,
+ scaler_type='standard',
+ enable_progress_bar=True,
+ ),
+ ],
+ freq='ME'
+)
+nf.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+Y_hat_df = nf.predict(futr_df=Y_test_df)
+
+# Plot quantile predictions
+Y_hat_df = Y_hat_df.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['DeepAR-median'], c='blue', label='median')
+plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['DeepAR-lo-90'][-12:].values,
+ y2=plot_df['DeepAR-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+plt.legend()
+plt.grid()
+plt.plot()
+```
+
diff --git a/neuralforecast/models.deepnpts.html.mdx b/neuralforecast/models.deepnpts.html.mdx
new file mode 100644
index 00000000..a624b4fb
--- /dev/null
+++ b/neuralforecast/models.deepnpts.html.mdx
@@ -0,0 +1,223 @@
+---
+output-file: models.deepnpts.html
+title: DeepNPTS
+---
+
+
+Deep Non-Parametric Time Series Forecaster
+([`DeepNPTS`](https://nixtlaverse.nixtla.io/neuralforecast/models.deepnpts.html#deepnpts))
+is a non-parametric baseline model for time-series forecasting. This
+model generates predictions by sampling from the empirical distribution
+according to a tunable strategy. This strategy is learned by exploiting
+the information across multiple related time series. This model provides
+a strong, simple baseline for time series forecasting.
+
+**References** [Rangapuram, Syama Sundar, Jan Gasthaus, Lorenzo
+Stella, Valentin Flunkert, David Salinas, Yuyang Wang, and Tim
+Januschowski (2023). “Deep Non-Parametric Time Series Forecaster”.
+arXiv.](https://arxiv.org/abs/2312.14657)
+
+> **Losses**
+>
+> This implementation differs from the original work in that a weighted
+> sum of the empirical distribution is returned as forecast. Therefore,
+> it only supports point losses.
+
+------------------------------------------------------------------------
+
+source
+
+### DeepNPTS
+
+> ``` text
+> DeepNPTS (h, input_size:int, hidden_size:int=32, batch_norm:bool=True,
+> dropout:float=0.1, n_layers:int=2, stat_exog_list=None,
+> hist_exog_list=None, futr_exog_list=None,
+> exclude_insample_y=False, loss=MAE(), valid_loss=MAE(),
+> max_steps:int=1000, learning_rate:float=0.001,
+> num_lr_decays:int=3, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size:int=32,
+> valid_batch_size:Optional[int]=None,
+> windows_batch_size:int=1024,
+> inference_windows_batch_size:int=1024,
+> start_padding_enabled=False, step_size:int=1,
+> scaler_type:str='standard', random_seed:int=1,
+> drop_last_loader=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*DeepNPTS
+
+Deep Non-Parametric Time Series Forecaster
+([`DeepNPTS`](https://nixtlaverse.nixtla.io/neuralforecast/models.deepnpts.html#deepnpts))
+is a baseline model for time-series forecasting. This model generates
+predictions by (weighted) sampling from the empirical distribution
+according to a learnable strategy. The strategy is learned by exploiting
+the information across multiple related time series.
+
+**Parameters:** `h`: int, Forecast horizon. `input_size`: int,
+autorregresive inputs size, y=\[1,2,3,4\] input_size=2 -\>
+y\_\[t-2:t\]=\[1,2\]. `hidden_size`: int=32, hidden size of dense
+layers. `batch_norm`: bool=True, if True, applies Batch
+Normalization after each dense layer in the network. `dropout`:
+float=0.1, dropout. `n_layers`: int=2, number of dense layers.
+`stat_exog_list`: str list, static exogenous columns.
+`hist_exog_list`: str list, historic exogenous columns.
+`futr_exog_list`: str list, future exogenous columns.
+`exclude_insample_y`: bool=False, the model skips the autoregressive
+features y\[t-input_size:t\] if True. `loss`: PyTorch module,
+instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=1024, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=-1, number of windows to sample in
+each inference batch, -1 uses all. `start_padding_enabled`:
+bool=False, if True, the model will pad the time series with zeros at
+the beginning, by input size. `step_size`: int=1, step size between
+each window of temporal data. `scaler_type`: str=‘identity’, type of
+scaler for temporal inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`. `dataloader_kwargs`:
+dict, optional, list of parameters passed into the PyTorch Lightning
+dataloader by the `TimeSeriesDataLoader`. `**trainer_kwargs`: int,
+keyword trainer arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+**References** - [Rangapuram, Syama Sundar, Jan Gasthaus, Lorenzo
+Stella, Valentin Flunkert, David Salinas, Yuyang Wang, and Tim
+Januschowski (2023). “Deep Non-Parametric Time Series Forecaster”.
+arXiv.](https://arxiv.org/abs/2312.14657) \*
+
+------------------------------------------------------------------------
+
+### DeepNPTS.fit
+
+> ``` text
+> DeepNPTS.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### DeepNPTS.predict
+
+> ``` text
+> DeepNPTS.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+
+```python
+check_model(DeepNPTS, ["airpassengers"])
+```
+
+## Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import DeepNPTS
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+nf = NeuralForecast(
+ models=[DeepNPTS(h=12,
+ input_size=24,
+ stat_exog_list=['airline1'],
+ futr_exog_list=['trend'],
+ max_steps=1000,
+ val_check_steps=10,
+ early_stop_patience_steps=3,
+ scaler_type='robust',
+ enable_progress_bar=True),
+ ],
+ freq='ME'
+)
+nf.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+Y_hat_df = nf.predict(futr_df=Y_test_df)
+
+# Plot quantile predictions
+Y_hat_df = Y_hat_df.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['DeepNPTS'], c='red', label='mean')
+plt.grid()
+plt.plot()
+```
+
diff --git a/neuralforecast/models.dilated_rnn.html.mdx b/neuralforecast/models.dilated_rnn.html.mdx
new file mode 100644
index 00000000..dd013799
--- /dev/null
+++ b/neuralforecast/models.dilated_rnn.html.mdx
@@ -0,0 +1,234 @@
+---
+output-file: models.dilated_rnn.html
+title: Dilated RNN
+---
+
+
+The Dilated Recurrent Neural Network
+([`DilatedRNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.dilated_rnn.html#dilatedrnn))
+addresses common challenges of modeling long sequences like vanishing
+gradients, computational efficiency, and improved model flexibility to
+model complex relationships while maintaining its parsimony. The
+[`DilatedRNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.dilated_rnn.html#dilatedrnn)
+builds a deep stack of RNN layers using skip conditions on the temporal
+and the network’s depth dimensions. The temporal dilated recurrent skip
+connections offer the capability to focus on multi-resolution inputs.The
+predictions are obtained by transforming the hidden states into contexts
+$\mathbf{c}_{[t+1:t+H]}$, that are decoded and adapted into
+$\mathbf{\hat{y}}_{[t+1:t+H],[q]}$ through MLPs.
+
+where $\mathbf{h}_{t}$, is the hidden state for time $t$,
+$\mathbf{y}_{t}$ is the input at time $t$ and $\mathbf{h}_{t-1}$ is the
+hidden state of the previous layer at $t-1$, $\mathbf{x}^{(s)}$ are
+static exogenous inputs, $\mathbf{x}^{(h)}_{t}$ historic exogenous,
+$\mathbf{x}^{(f)}_{[:t+H]}$ are future exogenous available at the time
+of the prediction.
+
+**References** -[Shiyu Chang, et al. “Dilated Recurrent Neural
+Networks”.](https://arxiv.org/abs/1710.02224) -[Yao Qin, et al. “A
+Dual-Stage Attention-Based recurrent neural network for time series
+prediction”.](https://arxiv.org/abs/1704.02971) -[Kashif Rasul, et
+al. “Zalando Research: PyTorch Dilated Recurrent Neural
+Networks”.](https://arxiv.org/abs/1710.02224)
+
+
+
+Figure 1. Three layer DilatedRNN with
+dilation 1, 2, 4.
+
+
+------------------------------------------------------------------------
+
+source
+
+### DilatedRNN
+
+> ``` text
+> DilatedRNN (h:int, input_size:int=-1,
+> inference_input_size:Optional[int]=None,
+> cell_type:str='LSTM', dilations:List[List[int]]=[[1, 2], [4,
+> 8]], encoder_hidden_size:int=128, context_size:int=10,
+> decoder_hidden_size:int=128, decoder_layers:int=2,
+> futr_exog_list=None, hist_exog_list=None,
+> stat_exog_list=None, exclude_insample_y=False, loss=MAE(),
+> valid_loss=None, max_steps:int=1000,
+> learning_rate:float=0.001, num_lr_decays:int=3,
+> early_stop_patience_steps:int=-1, val_check_steps:int=100,
+> batch_size=32, valid_batch_size:Optional[int]=None,
+> windows_batch_size=128, inference_windows_batch_size=1024,
+> start_padding_enabled=False, step_size:int=1,
+> scaler_type:str='robust', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*DilatedRNN
+
+**Parameters:** `h`: int, forecast horizon. `input_size`: int,
+maximum sequence length for truncated train backpropagation. Default -1
+uses 3 \* horizon `inference_input_size`: int, maximum sequence
+length for truncated inference. Default None uses input_size
+history. `cell_type`: str, type of RNN cell to use. Options: ‘GRU’,
+‘RNN’, ‘LSTM’, ‘ResLSTM’, ‘AttentiveLSTM’. `dilations`: int list,
+dilations betweem layers. `encoder_hidden_size`: int=200, units for
+the RNN’s hidden state size. `context_size`: int=10, size of context
+vector for each timestamp on the forecasting window.
+`decoder_hidden_size`: int=200, size of hidden layer for the MLP
+decoder. `decoder_layers`: int=2, number of layers for the MLP
+decoder. `futr_exog_list`: str list, future exogenous columns.
+`hist_exog_list`: str list, historic exogenous columns.
+`stat_exog_list`: str list, static exogenous columns.
+`exclude_insample_y`: bool=False, the model skips the autoregressive
+features y\[t-input_size:t\] if True. `loss`: PyTorch module,
+instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int, maximum number of training steps. `learning_rate`:
+float, Learning rate between (0, 1). `num_lr_decays`: int, Number of
+learning rate decays, evenly distributed across max_steps.
+`early_stop_patience_steps`: int, Number of validation iterations before
+early stopping. `val_check_steps`: int, Number of training steps
+between every validation loss check. `batch_size`: int=32, number of
+different series in each batch. `valid_batch_size`: int=None, number
+of different series in each validation and test batch.
+`windows_batch_size`: int=128, number of windows to sample in each
+training batch, default uses all. `inference_windows_batch_size`:
+int=1024, number of windows to sample in each inference batch, -1 uses
+all. `start_padding_enabled`: bool=False, if True, the model will
+pad the time series with zeros at the beginning, by input size.
+`step_size`: int=1, step size between each window of temporal data.
+`scaler_type`: str=‘robust’, type of scaler for temporal inputs
+normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`. `dataloader_kwargs`:
+dict, optional, list of parameters passed into the PyTorch Lightning
+dataloader by the `TimeSeriesDataLoader`. `**trainer_kwargs`: int,
+keyword trainer arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer). \*
+
+------------------------------------------------------------------------
+
+### DilatedRNN.fit
+
+> ``` text
+> DilatedRNN.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### DilatedRNN.predict
+
+> ``` text
+> DilatedRNN.predict (dataset, test_size=None, step_size=1,
+> random_seed=None, quantiles=None,
+> **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import DilatedRNN
+from neuralforecast.losses.pytorch import DistributionLoss
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+fcst = NeuralForecast(
+ models=[DilatedRNN(h=12,
+ input_size=-1,
+ loss=DistributionLoss(distribution='Normal', level=[80, 90]),
+ scaler_type='robust',
+ encoder_hidden_size=100,
+ max_steps=200,
+ futr_exog_list=['y_[lag12]'],
+ hist_exog_list=None,
+ stat_exog_list=['airline1'],
+ )
+ ],
+ freq='ME'
+)
+fcst.fit(df=Y_train_df, static_df=AirPassengersStatic)
+forecasts = fcst.predict(futr_df=Y_test_df)
+
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['DilatedRNN-median'], c='blue', label='median')
+plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['DilatedRNN-lo-90'][-12:].values,
+ y2=plot_df['DilatedRNN-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+plt.legend()
+plt.grid()
+plt.plot()
+```
+
diff --git a/neuralforecast/models.dlinear.html.mdx b/neuralforecast/models.dlinear.html.mdx
new file mode 100644
index 00000000..caf82819
--- /dev/null
+++ b/neuralforecast/models.dlinear.html.mdx
@@ -0,0 +1,250 @@
+---
+output-file: models.dlinear.html
+title: DLinear
+---
+
+
+DLinear is a simple and fast yet accurate time series forecasting model
+for long-horizon forecasting.
+
+The architecture has the following distinctive features: - Uses
+Autoformmer’s trend and seasonality decomposition. - Simple linear
+layers for trend and seasonality component.
+
+**References** - [Zeng, Ailing, et al. “Are transformers effective
+for time series forecasting?.” Proceedings of the AAAI conference on
+artificial intelligence. Vol. 37. No. 9.
+2023.”](https://ojs.aaai.org/index.php/AAAI/article/view/26317)
+
+
+
+Figure 1. DLinear
+Architecture.
+
+
+## 1. Auxiliary Functions
+
+------------------------------------------------------------------------
+
+source
+
+### SeriesDecomp
+
+> ``` text
+> SeriesDecomp (kernel_size)
+> ```
+
+*Series decomposition block*
+
+------------------------------------------------------------------------
+
+source
+
+### MovingAvg
+
+> ``` text
+> MovingAvg (kernel_size, stride)
+> ```
+
+*Moving average block to highlight the trend of time series*
+
+## 2. DLinear
+
+------------------------------------------------------------------------
+
+source
+
+### DLinear
+
+> ``` text
+> DLinear (h:int, input_size:int, stat_exog_list=None, hist_exog_list=None,
+> futr_exog_list=None, exclude_insample_y=False,
+> moving_avg_window:int=25, loss=MAE(), valid_loss=None,
+> max_steps:int=5000, learning_rate:float=0.0001,
+> num_lr_decays:int=-1, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size:int=32,
+> valid_batch_size:Optional[int]=None, windows_batch_size=1024,
+> inference_windows_batch_size=1024, start_padding_enabled=False,
+> step_size:int=1, scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*DLinear
+
+*Parameters:* `h`: int, forecast horizon. `input_size`: int,
+maximum sequence length for truncated train backpropagation.
+`stat_exog_list`: str list, static exogenous columns.
+`hist_exog_list`: str list, historic exogenous columns.
+`futr_exog_list`: str list, future exogenous columns.
+`exclude_insample_y`: bool=False, the model skips the autoregressive
+features y\[t-input_size:t\] if True. `moving_avg_window`: int=25,
+window size for trend-seasonality decomposition. Should be uneven.
+`loss`: PyTorch module, instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=1024, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=1024, number of windows to sample in
+each inference batch. `start_padding_enabled`: bool=False, if True,
+the model will pad the time series with zeros at the beginning, by input
+size. `step_size`: int=1, step size between each window of temporal
+data. `scaler_type`: str=‘robust’, type of scaler for temporal
+inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`. `dataloader_kwargs`:
+dict, optional, list of parameters passed into the PyTorch Lightning
+dataloader by the `TimeSeriesDataLoader`. `**trainer_kwargs`: int,
+keyword trainer arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+``` text
+*References*
+- Zeng, Ailing, et al. "Are transformers effective for time series forecasting?." Proceedings of the AAAI conference on artificial intelligence. Vol. 37. No. 9. 2023."*
+```
+
+------------------------------------------------------------------------
+
+### DLinear.fit
+
+> ``` text
+> DLinear.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### DLinear.predict
+
+> ``` text
+> DLinear.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import DLinear
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic, augment_calendar_df
+
+AirPassengersPanel, calendar_cols = augment_calendar_df(df=AirPassengersPanel, freq='M')
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = DLinear(h=12,
+ input_size=24,
+ loss=MAE(),
+ scaler_type='robust',
+ learning_rate=1e-3,
+ max_steps=500,
+ val_check_steps=50,
+ early_stop_patience_steps=2)
+
+nf = NeuralForecast(
+ models=[model],
+ freq='ME'
+)
+nf.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+forecasts = nf.predict(futr_df=Y_test_df)
+
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+if model.loss.is_distribution_output:
+ plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+ plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+ plt.plot(plot_df['ds'], plot_df['DLinear-median'], c='blue', label='median')
+ plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['DLinear-lo-90'][-12:].values,
+ y2=plot_df['DLinear-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+ plt.grid()
+ plt.legend()
+ plt.plot()
+else:
+ plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+ plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+ plt.plot(plot_df['ds'], plot_df['DLinear'], c='blue', label='Forecast')
+ plt.legend()
+ plt.grid()
+```
+
diff --git a/neuralforecast/models.fedformer.html.mdx b/neuralforecast/models.fedformer.html.mdx
new file mode 100644
index 00000000..9210782d
--- /dev/null
+++ b/neuralforecast/models.fedformer.html.mdx
@@ -0,0 +1,401 @@
+---
+output-file: models.fedformer.html
+title: FEDformer
+---
+
+
+The FEDformer model tackles the challenge of finding reliable
+dependencies on intricate temporal patterns of long-horizon forecasting.
+
+The architecture has the following distinctive features: - In-built
+progressive decomposition in trend and seasonal components based on a
+moving average filter. - Frequency Enhanced Block and Frequency Enhanced
+Attention to perform attention in the sparse representation on basis
+such as Fourier transform. - Classic encoder-decoder proposed by Vaswani
+et al. (2017) with a multi-head attention mechanism.
+
+The FEDformer model utilizes a three-component approach to define its
+embedding: - It employs encoded autoregressive features obtained from a
+convolution network. - Absolute positional embeddings obtained from
+calendar features are utilized.
+
+**References** - [Zhou, Tian, Ziqing Ma, Qingsong Wen, Xue Wang,
+Liang Sun, and Rong Jin.. “FEDformer: Frequency enhanced decomposed
+transformer for long-term series
+forecasting”](https://proceedings.mlr.press/v162/zhou22g.html)
+
+
+
+Figure 1. FEDformer
+Architecture.
+
+
+## 1. Auxiliary functions
+
+------------------------------------------------------------------------
+
+source
+
+### AutoCorrelationLayer
+
+> ``` text
+> AutoCorrelationLayer (correlation, hidden_size, n_head, d_keys=None,
+> d_values=None)
+> ```
+
+*Auto Correlation Layer*
+
+------------------------------------------------------------------------
+
+source
+
+### LayerNorm
+
+> ``` text
+> LayerNorm (channels)
+> ```
+
+*Special designed layernorm for the seasonal part*
+
+------------------------------------------------------------------------
+
+source
+
+### Decoder
+
+> ``` text
+> Decoder (layers, norm_layer=None, projection=None)
+> ```
+
+*FEDformer decoder*
+
+------------------------------------------------------------------------
+
+source
+
+### DecoderLayer
+
+> ``` text
+> DecoderLayer (self_attention, cross_attention, hidden_size, c_out,
+> conv_hidden_size=None, MovingAvg=25, dropout=0.1,
+> activation='relu')
+> ```
+
+*FEDformer decoder layer with the progressive decomposition
+architecture*
+
+------------------------------------------------------------------------
+
+source
+
+### Encoder
+
+> ``` text
+> Encoder (attn_layers, conv_layers=None, norm_layer=None)
+> ```
+
+*FEDformer encoder*
+
+------------------------------------------------------------------------
+
+source
+
+### EncoderLayer
+
+> ``` text
+> EncoderLayer (attention, hidden_size, conv_hidden_size=None,
+> MovingAvg=25, dropout=0.1, activation='relu')
+> ```
+
+*FEDformer encoder layer with the progressive decomposition
+architecture*
+
+------------------------------------------------------------------------
+
+source
+
+### FourierCrossAttention
+
+> ``` text
+> FourierCrossAttention (in_channels, out_channels, seq_len_q, seq_len_kv,
+> modes=64, mode_select_method='random',
+> activation='tanh', policy=0)
+> ```
+
+*Fourier Cross Attention layer*
+
+------------------------------------------------------------------------
+
+source
+
+### FourierBlock
+
+> ``` text
+> FourierBlock (in_channels, out_channels, seq_len, modes=0,
+> mode_select_method='random')
+> ```
+
+*Fourier block*
+
+------------------------------------------------------------------------
+
+source
+
+### get_frequency_modes
+
+> ``` text
+> get_frequency_modes (seq_len, modes=64, mode_select_method='random')
+> ```
+
+*Get modes on frequency domain: ‘random’ for sampling randomly ‘else’
+for sampling the lowest modes;*
+
+## 2. Model
+
+------------------------------------------------------------------------
+
+source
+
+### FEDformer
+
+> ``` text
+> FEDformer (h:int, input_size:int, stat_exog_list=None,
+> hist_exog_list=None, futr_exog_list=None,
+> decoder_input_size_multiplier:float=0.5,
+> version:str='Fourier', modes:int=64, mode_select:str='random',
+> hidden_size:int=128, dropout:float=0.05, n_head:int=8,
+> conv_hidden_size:int=32, activation:str='gelu',
+> encoder_layers:int=2, decoder_layers:int=1,
+> MovingAvg_window:int=25, loss=MAE(), valid_loss=None,
+> max_steps:int=5000, learning_rate:float=0.0001,
+> num_lr_decays:int=-1, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size:int=32,
+> valid_batch_size:Optional[int]=None, windows_batch_size=1024,
+> inference_windows_batch_size=1024,
+> start_padding_enabled=False, step_size:int=1,
+> scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*FEDformer
+
+The FEDformer model tackles the challenge of finding reliable
+dependencies on intricate temporal patterns of long-horizon forecasting.
+
+The architecture has the following distinctive features: - In-built
+progressive decomposition in trend and seasonal components based on a
+moving average filter. - Frequency Enhanced Block and Frequency Enhanced
+Attention to perform attention in the sparse representation on basis
+such as Fourier transform. - Classic encoder-decoder proposed by Vaswani
+et al. (2017) with a multi-head attention mechanism.
+
+The FEDformer model utilizes a three-component approach to define its
+embedding: - It employs encoded autoregressive features obtained from a
+convolution network. - Absolute positional embeddings obtained from
+calendar features are utilized.
+
+*Parameters:* `h`: int, forecast horizon. `input_size`: int,
+maximum sequence length for truncated train backpropagation.
+`stat_exog_list`: str list, static exogenous columns.
+`hist_exog_list`: str list, historic exogenous columns.
+`futr_exog_list`: str list, future exogenous columns.
+`decoder_input_size_multiplier`: float = 0.5, . `version`: str =
+‘Fourier’, version of the model. `modes`: int = 64, number of modes
+for the Fourier block. `mode_select`: str = ‘random’, method to
+select the modes for the Fourier block. `hidden_size`: int=128,
+units of embeddings and encoders. `dropout`: float (0, 1), dropout
+throughout Autoformer architecture. `n_head`: int=8, controls number
+of multi-head’s attention. `conv_hidden_size`: int=32, channels of
+the convolutional encoder. `activation`: str=`GELU`, activation from
+\[‘ReLU’, ‘Softplus’, ‘Tanh’, ‘SELU’, ‘LeakyReLU’, ‘PReLU’, ‘Sigmoid’,
+‘GELU’\]. `encoder_layers`: int=2, number of layers for the TCN
+encoder. `decoder_layers`: int=1, number of layers for the MLP
+decoder. `MovingAvg_window`: int=25, window size for the moving
+average filter. `loss`: PyTorch module, instantiated train loss
+class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module, instantiated validation loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=1024, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=1024, number of windows to sample in
+each inference batch. `start_padding_enabled`: bool=False, if True,
+the model will pad the time series with zeros at the beginning, by input
+size. `step_size`: int=1, step size between each window of temporal
+data.
+`scaler_type`: str=‘robust’, type of scaler for temporal inputs
+normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`. `dataloader_kwargs`:
+dict, optional, list of parameters passed into the PyTorch Lightning
+dataloader by the `TimeSeriesDataLoader`. `**trainer_kwargs`: int,
+keyword trainer arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer). \*
+
+------------------------------------------------------------------------
+
+### FEDformer.fit
+
+> ``` text
+> FEDformer.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### FEDformer.predict
+
+> ``` text
+> FEDformer.predict (dataset, test_size=None, step_size=1,
+> random_seed=None, quantiles=None,
+> **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+# Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import FEDformer
+from neuralforecast.utils import AirPassengersPanel, augment_calendar_df
+
+AirPassengersPanel, calendar_cols = augment_calendar_df(df=AirPassengersPanel, freq='M')
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = FEDformer(h=12,
+ input_size=24,
+ modes=64,
+ hidden_size=64,
+ conv_hidden_size=128,
+ n_head=8,
+ loss=MAE(),
+ futr_exog_list=calendar_cols,
+ scaler_type='robust',
+ learning_rate=1e-3,
+ max_steps=500,
+ batch_size=2,
+ windows_batch_size=32,
+ val_check_steps=50,
+ early_stop_patience_steps=2)
+
+nf = NeuralForecast(
+ models=[model],
+ freq='ME',
+)
+nf.fit(df=Y_train_df, static_df=None, val_size=12)
+forecasts = nf.predict(futr_df=Y_test_df)
+
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+if model.loss.is_distribution_output:
+ plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+ plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+ plt.plot(plot_df['ds'], plot_df['FEDformer-median'], c='blue', label='median')
+ plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['FEDformer-lo-90'][-12:].values,
+ y2=plot_df['FEDformer-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+ plt.grid()
+ plt.legend()
+ plt.plot()
+else:
+ plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+ plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+ plt.plot(plot_df['ds'], plot_df['FEDformer'], c='blue', label='Forecast')
+ plt.legend()
+ plt.grid()
+```
+
diff --git a/neuralforecast/models.gru.html.mdx b/neuralforecast/models.gru.html.mdx
new file mode 100644
index 00000000..9d6b46f7
--- /dev/null
+++ b/neuralforecast/models.gru.html.mdx
@@ -0,0 +1,240 @@
+---
+output-file: models.gru.html
+title: GRU
+---
+
+
+Cho et. al proposed the Gated Recurrent Unit
+([`GRU`](https://nixtlaverse.nixtla.io/neuralforecast/models.gru.html#gru))
+to improve on LSTM and Elman cells. The predictions at each time are
+given by a MLP decoder. This architecture follows closely the original
+Multi Layer Elman
+[`RNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.rnn.html#rnn)
+with the main difference being its use of the GRU cells. The predictions
+are obtained by transforming the hidden states into contexts
+$\mathbf{c}_{[t+1:t+H]}$, that are decoded and adapted into
+$\mathbf{\hat{y}}_{[t+1:t+H],[q]}$ through MLPs.
+
+where $\mathbf{h}_{t}$, is the hidden state for time $t$,
+$\mathbf{y}_{t}$ is the input at time $t$ and $\mathbf{h}_{t-1}$ is the
+hidden state of the previous layer at $t-1$, $\mathbf{x}^{(s)}$ are
+static exogenous inputs, $\mathbf{x}^{(h)}_{t}$ historic exogenous,
+$\mathbf{x}^{(f)}_{[:t+H]}$ are future exogenous available at the time
+of the prediction.
+
+**References** -[Junyoung Chung, Caglar Gulcehre, KyungHyun Cho,
+Yoshua Bengio (2014). “Empirical Evaluation of Gated Recurrent Neural
+Networks on Sequence Modeling”.](https:arxivorg/abs/1412.3555)
+-[Kyunghyun Cho, Bart van Merrienboer, Dzmitry Bahdanau, Yoshua Bengio
+(2014). “On the Properties of Neural Machine Translation:
+Encoder-Decoder Approaches”.](https://arxiv.org/abs/1409.1259)
+
+
+
+Figure 1. Gated Recurrent Unit
+Cell.
+
+
+------------------------------------------------------------------------
+
+source
+
+### GRU
+
+> ``` text
+> GRU (h:int, input_size:int=-1, inference_input_size:Optional[int]=None,
+> h_train:int=1, encoder_n_layers:int=2, encoder_hidden_size:int=200,
+> encoder_activation:Optional[str]=None, encoder_bias:bool=True,
+> encoder_dropout:float=0.0, context_size:Optional[int]=None,
+> decoder_hidden_size:int=128, decoder_layers:int=2,
+> futr_exog_list=None, hist_exog_list=None, stat_exog_list=None,
+> exclude_insample_y=False, recurrent=False, loss=MAE(),
+> valid_loss=None, max_steps:int=1000, learning_rate:float=0.001,
+> num_lr_decays:int=-1, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size=32,
+> valid_batch_size:Optional[int]=None, windows_batch_size=128,
+> inference_windows_batch_size=1024, start_padding_enabled=False,
+> step_size:int=1, scaler_type:str='robust', random_seed=1,
+> drop_last_loader=False, alias:Optional[str]=None, optimizer=None,
+> optimizer_kwargs=None, lr_scheduler=None, lr_scheduler_kwargs=None,
+> dataloader_kwargs=None, **trainer_kwargs)
+> ```
+
+\*GRU
+
+Multi Layer Recurrent Network with Gated Units (GRU), and MLP decoder.
+The network has non-linear activation functions, it is trained using
+ADAM stochastic gradient descent. The network accepts static, historic
+and future exogenous data, flattens the inputs.
+
+**Parameters:** `h`: int, forecast horizon. `input_size`: int,
+maximum sequence length for truncated train backpropagation. Default -1
+uses 3 \* horizon `inference_input_size`: int, maximum sequence
+length for truncated inference. Default None uses input_size
+history. `h_train`: int, maximum sequence length for truncated train
+backpropagation. Default 1. `encoder_n_layers`: int=2, number of
+layers for the GRU. `encoder_hidden_size`: int=200, units for the
+GRU’s hidden state size. `encoder_activation`: Optional\[str\]=None,
+Deprecated. Activation function in GRU is frozen in PyTorch.
+`encoder_bias`: bool=True, whether or not to use biases b_ih, b_hh
+within GRU units. `encoder_dropout`: float=0., dropout
+regularization applied to GRU outputs. `context_size`:
+deprecated. `decoder_hidden_size`: int=200, size of hidden layer for
+the MLP decoder. `decoder_layers`: int=2, number of layers for the
+MLP decoder. `futr_exog_list`: str list, future exogenous
+columns. `hist_exog_list`: str list, historic exogenous columns.
+`stat_exog_list`: str list, static exogenous columns.
+`exclude_insample_y`: bool=False, whether to exclude the target variable
+from the input. `recurrent`: bool=False, whether to produce
+forecasts recursively (True) or direct (False). `loss`: PyTorch
+module, instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of
+differentseries in each batch. `valid_batch_size`: int=None, number
+of different series in each validation and test batch.
+`windows_batch_size`: int=128, number of windows to sample in each
+training batch, default uses all. `inference_windows_batch_size`:
+int=1024, number of windows to sample in each inference batch, -1 uses
+all. `start_padding_enabled`: bool=False, if True, the model will
+pad the time series with zeros at the beginning, by input size.
+`step_size`: int=1, step size between each window of temporal data.
+`scaler_type`: str=‘robust’, type of scaler for temporal inputs
+normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`. `dataloader_kwargs`:
+dict, optional, list of parameters passed into the PyTorch Lightning
+dataloader by the `TimeSeriesDataLoader`. `**trainer_kwargs`: int,
+keyword trainer arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer). \*
+
+------------------------------------------------------------------------
+
+### GRU.fit
+
+> ``` text
+> GRU.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### GRU.predict
+
+> ``` text
+> GRU.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+# from neuralforecast.models import GRU
+from neuralforecast.losses.pytorch import DistributionLoss
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+fcst = NeuralForecast(
+ models=[GRU(h=12, input_size=24,
+ loss=DistributionLoss(distribution='Normal', level=[80, 90]),
+ scaler_type='robust',
+ encoder_n_layers=2,
+ encoder_hidden_size=128,
+ decoder_hidden_size=128,
+ decoder_layers=2,
+ max_steps=200,
+ futr_exog_list=None,
+ hist_exog_list=['y_[lag12]'],
+ stat_exog_list=['airline1'],
+ )
+ ],
+ freq='ME'
+)
+fcst.fit(df=Y_train_df, static_df=AirPassengersStatic)
+forecasts = fcst.predict(futr_df=Y_test_df)
+
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['GRU-median'], c='blue', label='median')
+plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['GRU-lo-90'][-12:].values,
+ y2=plot_df['GRU-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+plt.legend()
+plt.grid()
+plt.plot()
+```
+
diff --git a/neuralforecast/models.hint.html.mdx b/neuralforecast/models.hint.html.mdx
new file mode 100644
index 00000000..fb7c2685
--- /dev/null
+++ b/neuralforecast/models.hint.html.mdx
@@ -0,0 +1,334 @@
+---
+output-file: models.hint.html
+title: HINT
+---
+
+
+The Hierarchical Mixture Networks (HINT) are a highly modular framework
+that combines SoTA neural forecast architectures with task-specialized
+mixture probability and advanced hierarchical reconciliation strategies.
+This powerful combination allows HINT to produce accurate and coherent
+probabilistic forecasts.
+
+HINT’s incorporates a `TemporalNorm` module into any neural forecast
+architecture, the module normalizes inputs into the network’s
+non-linearities operating range and recomposes its output’s scales
+through a global skip connection, improving accuracy and training
+robustness. HINT ensures the forecast coherence via bootstrap sample
+reconciliation that restores the aggregation constraints into its base
+samples.
+
+**References** - [Kin G. Olivares, David Luo, Cristian Challu,
+Stefania La Vattiata, Max Mergenthaler, Artur Dubrawski (2023). “HINT:
+Hierarchical Mixture Networks For Coherent Probabilistic Forecasting”.
+Neural Information Processing Systems, submitted. Working Paper version
+available at arxiv.](https://arxiv.org/abs/2305.07089) - [Kin G.
+Olivares, O. Nganba Meetei, Ruijun Ma, Rohan Reddy, Mengfei Cao, Lee
+Dicker (2022).”Probabilistic Hierarchical Forecasting with Deep Poisson
+Mixtures”. International Journal Forecasting, accepted paper available
+at arxiv.](https://arxiv.org/pdf/2110.13179.pdf) - [Kin G. Olivares,
+Federico Garza, David Luo, Cristian Challu, Max Mergenthaler, Souhaib
+Ben Taieb, Shanika Wickramasuriya, and Artur Dubrawski (2022).
+“HierarchicalForecast: A reference framework for hierarchical
+forecasting in python”. Journal of Machine Learning Research, submitted,
+abs/2207.03517, 2022b.](https://arxiv.org/abs/2207.03517)
+
+
+
+Figure 1. Hierarchical Mixture Networks
+(HINT).
+
+
+## Reconciliation Methods
+
+------------------------------------------------------------------------
+
+source
+
+### get_identity_P
+
+> ``` text
+> get_identity_P (S:numpy.ndarray)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### get_bottomup_P
+
+> ``` text
+> get_bottomup_P (S:numpy.ndarray)
+> ```
+
+\*BottomUp Reconciliation Matrix.
+
+Creates BottomUp hierarchical “projection” matrix is defined as:
+$$\mathbf{P}_{\text{BU}} = [\mathbf{0}_{\mathrm{[b],[a]}}\;|\;\mathbf{I}_{\mathrm{[b][b]}}]$$
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+
+**Returns:** `P`: Reconciliation matrix of size (`bottom`,
+`base`).
+
+**References:** - [Orcutt, G.H., Watts, H.W., & Edwards, J.B.(1968).
+“Data aggregation and information loss”. The American Economic Review,
+58 , 773(787)](http://www.jstor.org/stable/1815532).\*
+
+------------------------------------------------------------------------
+
+source
+
+### get_mintrace_ols_P
+
+> ``` text
+> get_mintrace_ols_P (S:numpy.ndarray)
+> ```
+
+\*MinTraceOLS Reconciliation Matrix.
+
+Creates MinTraceOLS reconciliation matrix as proposed by Wickramasuriya
+et al.
+
+$$\mathbf{P}_{\text{MinTraceOLS}}=\left(\mathbf{S}^{\intercal}\mathbf{S}\right)^{-1}\mathbf{S}^{\intercal}$$
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+
+**Returns:** `P`: Reconciliation matrix of size (`bottom`,
+`base`).
+
+**References:** - [Wickramasuriya, S.L., Turlach, B.A. & Hyndman,
+R.J. (2020). “Optimal non-negative forecast reconciliation”. Stat Comput
+30, 1167–1182,
+https://doi.org/10.1007/s11222-020-09930-0](https://robjhyndman.com/publications/nnmint/).\*
+
+------------------------------------------------------------------------
+
+source
+
+### get_mintrace_wls_P
+
+> ``` text
+> get_mintrace_wls_P (S:numpy.ndarray)
+> ```
+
+\*MinTraceOLS Reconciliation Matrix.
+
+Creates MinTraceOLS reconciliation matrix as proposed by Wickramasuriya
+et al. Depending on a weighted GLS estimator and an estimator of the
+covariance matrix of the coherency errors $\mathbf{W}_{h}$.
+
+$$ \mathbf{W}_{h} = \mathrm{Diag}(\mathbf{S} \mathbb{1}_{[b]})$$
+
+$$
+\mathbf{P}_{\text{MinTraceWLS}}=\left(\mathbf{S}^{\intercal}\mathbf{W}_{h}\mathbf{S}\right)^{-1}
+\mathbf{S}^{\intercal}\mathbf{W}^{-1}_{h}
+$$
+
+**Parameters:** `S`: Summing matrix of size (`base`, `bottom`).
+
+**Returns:** `P`: Reconciliation matrix of size (`bottom`,
+`base`).
+
+**References:** - [Wickramasuriya, S.L., Turlach, B.A. & Hyndman,
+R.J. (2020). “Optimal non-negative forecast reconciliation”. Stat Comput
+30, 1167–1182,
+https://doi.org/10.1007/s11222-020-09930-0](https://robjhyndman.com/publications/nnmint/).\*
+
+## HINT
+
+------------------------------------------------------------------------
+
+source
+
+### HINT
+
+> ``` text
+> HINT (h:int, S:numpy.ndarray, model, reconciliation:str,
+> alias:Optional[str]=None)
+> ```
+
+\*HINT
+
+The Hierarchical Mixture Networks (HINT) are a highly modular framework
+that combines SoTA neural forecast architectures with a task-specialized
+mixture probability and advanced hierarchical reconciliation strategies.
+This powerful combination allows HINT to produce accurate and coherent
+probabilistic forecasts.
+
+HINT’s incorporates a `TemporalNorm` module into any neural forecast
+architecture, the module normalizes inputs into the network’s
+non-linearities operating range and recomposes its output’s scales
+through a global skip connection, improving accuracy and training
+robustness. HINT ensures the forecast coherence via bootstrap sample
+reconciliation that restores the aggregation constraints into its base
+samples.
+
+Available reconciliations: - BottomUp - MinTraceOLS -
+MinTraceWLS - Identity
+
+**Parameters:** `h`: int, Forecast horizon. `model`:
+NeuralForecast model, instantiated model class from [architecture
+collection](https://nixtlaverse.nixtla.io/neuralforecast/models.html).
+`S`: np.ndarray, dumming matrix of size (`base`, `bottom`) see
+HierarchicalForecast’s [aggregate
+method](https://nixtlaverse.nixtla.io/hierarchicalforecast/src/utils.html#aggregate).
+`reconciliation`: str, HINT’s reconciliation method from \[‘BottomUp’,
+‘MinTraceOLS’, ‘MinTraceWLS’\]. `alias`: str, optional, Custom name
+of the model. \*
+
+------------------------------------------------------------------------
+
+source
+
+### HINT.fit
+
+> ``` text
+> HINT.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*HINT.fit
+
+HINT trains on the entire hierarchical dataset, by minimizing a
+composite log likelihood objective. HINT framework integrates
+`TemporalNorm` into the neural forecast architecture for a
+scale-decoupled optimization that robustifies cross-learning the
+hierachy’s series scales.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset)
+see details
+[here](https://nixtla.github.io/neuralforecast/tsdataset.html)
+`val_size`: int, size of the validation set, (default 0).
+`test_size`: int, size of the test set, (default 0). `random_seed`:
+int, random seed for the prediction.
+
+**Returns:** `self`: A fitted base
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+model. \*
+
+------------------------------------------------------------------------
+
+source
+
+### HINT.predict
+
+> ``` text
+> HINT.predict (dataset, step_size=1, random_seed=None,
+> **data_module_kwargs)
+> ```
+
+\*HINT.predict
+
+After fitting a base model on the entire hierarchical dataset. HINT
+restores the hierarchical aggregation constraints using bootstrapped
+sample reconciliation.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset)
+see details
+[here](https://nixtla.github.io/neuralforecast/tsdataset.html)
+`step_size`: int, steps between sequential predictions, (default 1).
+`random_seed`: int, random seed for the prediction. `**data_kwarg`:
+additional parameters for the dataset module.
+
+**Returns:** `y_hat`: numpy predictions of the
+[`NeuralForecast`](https://nixtlaverse.nixtla.io/neuralforecast/core.html#neuralforecast)
+model. \*
+
+## Usage Example
+
+In this example we will use HINT for the hierarchical forecast task, a
+multivariate regression problem with aggregation constraints. The
+aggregation constraints can be compactcly represented by the summing
+matrix $\mathbf{S}_{[i][b]}$, the Figure belows shows an example.
+
+In this example we will make coherent predictions for the TourismL
+dataset.
+
+Outline 1. Import packages 2. Load hierarchical dataset 3.
+Fit and Predict HINT 4. Forecast Plot
+
+
+
+
+```python
+import matplotlib.pyplot as plt
+
+from neuralforecast.losses.pytorch import GMM, sCRPS
+from datasetsforecast.hierarchical import HierarchicalData
+
+# Auxiliary sorting
+def sort_df_hier(Y_df, S_df):
+ # NeuralForecast core, sorts unique_id lexicographically
+ # by default, this class matches S_df and Y_hat_df order.
+ Y_df.unique_id = Y_df.unique_id.astype('category')
+ Y_df.unique_id = Y_df.unique_id.cat.set_categories(S_df.index)
+ Y_df = Y_df.sort_values(by=['unique_id', 'ds'])
+ return Y_df
+
+# Load TourismSmall dataset
+horizon = 12
+Y_df, S_df, tags = HierarchicalData.load('./data', 'TourismLarge')
+Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+Y_df = sort_df_hier(Y_df, S_df)
+level = [80,90]
+
+# Instantiate HINT
+# BaseNetwork + Distribution + Reconciliation
+nhits = NHITS(h=horizon,
+ input_size=24,
+ loss=GMM(n_components=10, level=level),
+ max_steps=2000,
+ early_stop_patience_steps=10,
+ val_check_steps=50,
+ scaler_type='robust',
+ learning_rate=1e-3,
+ valid_loss=sCRPS(level=level))
+
+model = HINT(h=horizon, S=S_df.values,
+ model=nhits, reconciliation='BottomUp')
+
+# Fit and Predict
+nf = NeuralForecast(models=[model], freq='MS')
+Y_hat_df = nf.cross_validation(df=Y_df, val_size=12, n_windows=1)
+Y_hat_df = Y_hat_df.reset_index()
+```
+
+
+```python
+# Plot coherent probabilistic forecast
+unique_id = 'TotalAll'
+Y_plot_df = Y_df[Y_df.unique_id==unique_id]
+plot_df = Y_hat_df[Y_hat_df.unique_id==unique_id]
+plot_df = Y_plot_df.merge(plot_df, on=['ds', 'unique_id'], how='left')
+n_years = 5
+
+plt.plot(plot_df['ds'][-12*n_years:], plot_df['y_x'][-12*n_years:], c='black', label='True')
+plt.plot(plot_df['ds'][-12*n_years:], plot_df['HINT'][-12*n_years:], c='purple', label='mean')
+plt.plot(plot_df['ds'][-12*n_years:], plot_df['HINT-median'][-12*n_years:], c='blue', label='median')
+plt.fill_between(x=plot_df['ds'][-12*n_years:],
+ y1=plot_df['HINT-lo-90'][-12*n_years:].values,
+ y2=plot_df['HINT-hi-90'][-12*n_years:].values,
+ alpha=0.4, label='level 90')
+plt.legend()
+plt.grid()
+plt.plot()
+```
+
diff --git a/neuralforecast/models.html.mdx b/neuralforecast/models.html.mdx
new file mode 100644
index 00000000..33e713a3
--- /dev/null
+++ b/neuralforecast/models.html.mdx
@@ -0,0 +1,2046 @@
+---
+description: >-
+ NeuralForecast contains user-friendly implementations of neural forecasting
+ models that allow for easy transition of computing capabilities (GPU/CPU),
+ computation parallelization, and hyperparameter tuning.
+output-file: models.html
+title: AutoModels
+---
+
+
+All the NeuralForecast models are “global” because we train them with
+all the series from the input pd.DataFrame data `Y_df`, yet the
+optimization objective is, momentarily, “univariate” as it does not
+consider the interaction between the output predictions across time
+series. Like the StatsForecast library, `core.NeuralForecast` allows you
+to explore collections of models efficiently and contains functions for
+convenient wrangling of input and output pd.DataFrames predictions.
+
+First we load the AirPassengers dataset such that you can run all the
+examples.
+
+
+```python
+import numpy as np
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast.tsdataset import TimeSeriesDataset
+from neuralforecast.utils import AirPassengersDF as Y_df
+```
+
+
+```python
+# Split train/test and declare time series dataset
+Y_train_df = Y_df[Y_df.ds<='1959-12-31'] # 132 train
+Y_test_df = Y_df[Y_df.ds>'1959-12-31'] # 12 test
+dataset, *_ = TimeSeriesDataset.from_df(Y_train_df)
+```
+
+# 1. Automatic Forecasting
+
+## A. RNN-Based
+
+------------------------------------------------------------------------
+
+source
+
+### AutoRNN
+
+> ``` text
+> AutoRNN (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= object at 0x7f1320942da0>, num_samples=10, refit_with_val=False,
+> cpus=4, gpus=0, verbose=False, alias=None, backend='ray',
+> callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoRNN.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=-1, encoder_hidden_size=8)
+model = AutoRNN(h=12, config=config, num_samples=1, cpus=1)
+
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoRNN(h=12, config=None, num_samples=1, cpus=1, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoLSTM
+
+> ``` text
+> AutoLSTM (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= object at 0x7f1320937310>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoLSTM.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=-1, encoder_hidden_size=8)
+model = AutoLSTM(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoLSTM(h=12, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoGRU
+
+> ``` text
+> AutoGRU (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= object at 0x7f1320e7c2b0>, num_samples=10, refit_with_val=False,
+> cpus=4, gpus=0, verbose=False, alias=None, backend='ray',
+> callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoGRU.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=-1, encoder_hidden_size=8)
+model = AutoGRU(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoGRU(h=12, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoTCN
+
+> ``` text
+> AutoTCN (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= object at 0x7f13208f1ae0>, num_samples=10, refit_with_val=False,
+> cpus=4, gpus=0, verbose=False, alias=None, backend='ray',
+> callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoTCN.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=-1, encoder_hidden_size=8)
+model = AutoTCN(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoTCN(h=12, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoDeepAR
+
+> ``` text
+> AutoDeepAR (h, loss=DistributionLoss(), valid_loss=MQLoss(), config=None,
+> search_alg= or object at 0x7f1320ecec80>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | DistributionLoss | DistributionLoss() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | MQLoss | MQLoss() | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoDeepAR.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12, lstm_hidden_size=8)
+model = AutoDeepAR(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoDeepAR(h=12, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoDilatedRNN
+
+> ``` text
+> AutoDilatedRNN (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= erator object at 0x7f132090feb0>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoDilatedRNN.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=-1, encoder_hidden_size=8)
+model = AutoDilatedRNN(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoDilatedRNN(h=12, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoBiTCN
+
+> ``` text
+> AutoBiTCN (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= r object at 0x7f1320a9c9d0>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoBiTCN.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12, hidden_size=8)
+model = AutoBiTCN(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoBiTCN(h=12, config=None, backend='optuna')
+```
+
+## B. MLP-Based
+
+------------------------------------------------------------------------
+
+source
+
+### AutoMLP
+
+> ``` text
+> AutoMLP (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= object at 0x7f1320ad7a60>, num_samples=10, refit_with_val=False,
+> cpus=4, gpus=0, verbose=False, alias=None, backend='ray',
+> callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoMLP.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12, hidden_size=8)
+model = AutoMLP(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoMLP(h=12, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoNBEATS
+
+> ``` text
+> AutoNBEATS (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= or object at 0x7f1320ad5390>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoNBEATS.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12,
+ mlp_units=3*[[8, 8]])
+model = AutoNBEATS(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoNBEATS(h=12, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoNBEATSx
+
+> ``` text
+> AutoNBEATSx (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= tor object at 0x7f1320ac6cb0>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoNBEATSx.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12,
+ mlp_units=3*[[8, 8]])
+model = AutoNBEATSx(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoNBEATSx(h=12, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoNHITS
+
+> ``` text
+> AutoNHITS (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= r object at 0x7f1320ab4100>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoNHITS.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12,
+ mlp_units=3 * [[8, 8]])
+model = AutoNHITS(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoNHITS(h=12, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoDLinear
+
+> ``` text
+> AutoDLinear (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= tor object at 0x7f1320a9cd90>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoDLinear.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12)
+model = AutoDLinear(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoDLinear(h=12, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoNLinear
+
+> ``` text
+> AutoNLinear (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= tor object at 0x7f1320ab6b00>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoNLinear.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12)
+model = AutoNLinear(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoNLinear(h=12, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoTiDE
+
+> ``` text
+> AutoTiDE (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= object at 0x7f1320ad4a60>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoTiDE.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12)
+model = AutoTiDE(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoTiDE(h=12, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoDeepNPTS
+
+> ``` text
+> AutoDeepNPTS (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= ator object at 0x7f1320dd1000>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoDeepNPTS.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12)
+model = AutoDeepNPTS(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoDeepNPTS(h=12, config=None, backend='optuna')
+```
+
+## C. KAN-Based
+
+------------------------------------------------------------------------
+
+source
+
+### AutoKAN
+
+> ``` text
+> AutoKAN (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= object at 0x7f1320a491e0>, num_samples=10, refit_with_val=False,
+> cpus=4, gpus=0, verbose=False, alias=None, backend='ray',
+> callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoKAN.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12)
+model = AutoKAN(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoKAN(h=12, config=None, backend='optuna')
+```
+
+## D. Transformer-Based
+
+------------------------------------------------------------------------
+
+source
+
+### AutoTFT
+
+> ``` text
+> AutoTFT (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= object at 0x7f1320de5780>, num_samples=10, refit_with_val=False,
+> cpus=4, gpus=0, verbose=False, alias=None, backend='ray',
+> callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoTFT.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12, hidden_size=8)
+model = AutoTFT(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoTFT(h=12, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoVanillaTransformer
+
+> ``` text
+> AutoVanillaTransformer (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= riantGenerator object at 0x7f1320a54a90>,
+> num_samples=10, refit_with_val=False, cpus=4,
+> gpus=0, verbose=False, alias=None, backend='ray',
+> callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoVanillaTransformer.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12, hidden_size=8)
+model = AutoVanillaTransformer(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoVanillaTransformer(h=12, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoInformer
+
+> ``` text
+> AutoInformer (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= ator object at 0x7f1320a31660>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoInformer.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12, hidden_size=8)
+model = AutoInformer(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoInformer(h=12, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoAutoformer
+
+> ``` text
+> AutoAutoformer (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= erator object at 0x7f1320a489d0>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoAutoformer.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12, hidden_size=8)
+model = AutoAutoformer(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoAutoformer(h=12, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoFEDformer
+
+> ``` text
+> AutoFEDformer (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= rator object at 0x7f1320c40220>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoFEDFormer.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12, hidden_size=64)
+model = AutoFEDformer(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoFEDformer(h=12, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoPatchTST
+
+> ``` text
+> AutoPatchTST (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= ator object at 0x7f13209ed420>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoPatchTST.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12, hidden_size=16)
+model = AutoPatchTST(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoPatchTST(h=12, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoiTransformer
+
+> ``` text
+> AutoiTransformer (h, n_series, loss=MAE(), valid_loss=None, config=None,
+> search_alg= enerator object at 0x7f1320da8490>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| n_series | | | |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoiTransformer.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12, hidden_size=16)
+model = AutoiTransformer(h=12, n_series=1, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoiTransformer(h=12, n_series=1, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoTimeXer
+
+> ``` text
+> AutoTimeXer (h, n_series, loss=MAE(), valid_loss=None, config=None,
+> search_alg= tor object at 0x7f13209ff2e0>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| n_series | | | |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoTimeXer.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12, patch_len=12)
+model = AutoTimeXer(h=12, n_series=1, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoTimeXer(h=12, n_series=1, config=None, backend='optuna')
+```
+
+## E. CNN Based
+
+------------------------------------------------------------------------
+
+source
+
+### AutoTimesNet
+
+> ``` text
+> AutoTimesNet (h, loss=MAE(), valid_loss=None, config=None,
+> search_alg= ator object at 0x7f1320a13760>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoTimesNet.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12, hidden_size=32)
+model = AutoTimesNet(h=12, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoTimesNet(h=12, config=None, backend='optuna')
+```
+
+## F. Multivariate
+
+------------------------------------------------------------------------
+
+source
+
+### AutoStemGNN
+
+> ``` text
+> AutoStemGNN (h, n_series, loss=MAE(), valid_loss=None, config=None,
+> search_alg= tor object at 0x7f1320bce500>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| n_series | | | |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoStemGNN.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12)
+model = AutoStemGNN(h=12, n_series=1, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoStemGNN(h=12, n_series=1, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoHINT
+
+> ``` text
+> AutoHINT (cls_model, h, loss, valid_loss, S, config,
+> search_alg= object at 0x7f13209ff790>, num_samples=10, cpus=4, gpus=0,
+> refit_with_val=False, verbose=False, alias=None, backend='ray',
+> callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| cls_model | PyTorch/PyTorchLightning model | | See `neuralforecast.models` [collection here](https://nixtla.github.io/neuralforecast/models.html). |
+| h | int | | Forecast horizon |
+| loss | PyTorch module | | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | PyTorch module | | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| S | | | |
+| config | dict or callable | | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Perform a simple hyperparameter optimization with
+# NHITS and then reconcile with HINT
+from neuralforecast.losses.pytorch import GMM, sCRPS
+
+base_config = dict(max_steps=1, val_check_steps=1, input_size=8)
+base_model = AutoNHITS(h=4, loss=GMM(n_components=2, quantiles=quantiles),
+ config=base_config, num_samples=1, cpus=1)
+model = HINT(h=4, S=S_df.values,
+ model=base_model, reconciliation='MinTraceOLS')
+
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=hint_dataset)
+
+# Perform a conjunct hyperparameter optimization with
+# NHITS + HINT reconciliation configurations
+nhits_config = {
+ "learning_rate": tune.choice([1e-3]), # Initial Learning rate
+ "max_steps": tune.choice([1]), # Number of SGD steps
+ "val_check_steps": tune.choice([1]), # Number of steps between validation
+ "input_size": tune.choice([5 * 12]), # input_size = multiplier * horizon
+ "batch_size": tune.choice([7]), # Number of series in windows
+ "windows_batch_size": tune.choice([256]), # Number of windows in batch
+ "n_pool_kernel_size": tune.choice([[2, 2, 2], [16, 8, 1]]), # MaxPool's Kernelsize
+ "n_freq_downsample": tune.choice([[168, 24, 1], [24, 12, 1], [1, 1, 1]]), # Interpolation expressivity ratios
+ "activation": tune.choice(['ReLU']), # Type of non-linear activation
+ "n_blocks": tune.choice([[1, 1, 1]]), # Blocks per each 3 stacks
+ "mlp_units": tune.choice([[[512, 512], [512, 512], [512, 512]]]), # 2 512-Layers per block for each stack
+ "interpolation_mode": tune.choice(['linear']), # Type of multi-step interpolation
+ "random_seed": tune.randint(1, 10),
+ "reconciliation": tune.choice(['BottomUp', 'MinTraceOLS', 'MinTraceWLS'])
+ }
+model = AutoHINT(h=4, S=S_df.values,
+ cls_model=NHITS,
+ config=nhits_config,
+ loss=GMM(n_components=2, level=[80, 90]),
+ valid_loss=sCRPS(level=[80, 90]),
+ num_samples=1, cpus=1)
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=hint_dataset)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoTSMixer
+
+> ``` text
+> AutoTSMixer (h, n_series, loss=MAE(), valid_loss=None, config=None,
+> search_alg= tor object at 0x7f1320a1f490>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| n_series | | | |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoTSMixer.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12)
+model = AutoTSMixer(h=12, n_series=1, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoTSMixer(h=12, n_series=1, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoTSMixerx
+
+> ``` text
+> AutoTSMixerx (h, n_series, loss=MAE(), valid_loss=None, config=None,
+> search_alg= ator object at 0x7f1320bcdea0>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| n_series | | | |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoTSMixerx.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12)
+model = AutoTSMixerx(h=12, n_series=1, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoTSMixerx(h=12, n_series=1, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoMLPMultivariate
+
+> ``` text
+> AutoMLPMultivariate (h, n_series, loss=MAE(), valid_loss=None,
+> config=None, search_alg= nt.BasicVariantGenerator object at 0x7f1320bbbc70>,
+> num_samples=10, refit_with_val=False, cpus=4,
+> gpus=0, verbose=False, alias=None, backend='ray',
+> callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| n_series | | | |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoMLPMultivariate.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12)
+model = AutoMLPMultivariate(h=12, n_series=1, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoMLPMultivariate(h=12, n_series=1, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoSOFTS
+
+> ``` text
+> AutoSOFTS (h, n_series, loss=MAE(), valid_loss=None, config=None,
+> search_alg= r object at 0x7f1320bae470>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| n_series | | | |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoSOFTS.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12, hidden_size=16)
+model = AutoSOFTS(h=12, n_series=1, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoSOFTS(h=12, n_series=1, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoTimeMixer
+
+> ``` text
+> AutoTimeMixer (h, n_series, loss=MAE(), valid_loss=None, config=None,
+> search_alg= rator object at 0x7f1320ba16c0>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| n_series | | | |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoTimeMixer.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12, d_model=16)
+model = AutoTimeMixer(h=12, n_series=1, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoTimeMixer(h=12, n_series=1, config=None, backend='optuna')
+```
+
+------------------------------------------------------------------------
+
+source
+
+### AutoRMoK
+
+> ``` text
+> AutoRMoK (h, n_series, loss=MAE(), valid_loss=None, config=None,
+> search_alg= object at 0x7f1320ba3340>, num_samples=10,
+> refit_with_val=False, cpus=4, gpus=0, verbose=False,
+> alias=None, backend='ray', callbacks=None)
+> ```
+
+\*Class for Automatic Hyperparameter Optimization, it builds on top of
+`ray` to give access to a wide variety of hyperparameter optimization
+tools ranging from classic grid search, to Bayesian optimization and
+HyperBand algorithm.
+
+The validation loss to be optimized is defined by the `config['loss']`
+dictionary value, the config also contains the rest of the
+hyperparameter search space.
+
+It is important to note that the success of this hyperparameter
+optimization heavily relies on a strong correlation between the
+validation and test periods.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon |
+| n_series | | | |
+| loss | MAE | MAE() | Instantiated train loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| valid_loss | NoneType | None | Instantiated valid loss class from [losses collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html). |
+| config | NoneType | None | Dictionary with ray.tune defined search space or function that takes an optuna trial and returns a configuration dict. |
+| search_alg | BasicVariantGenerator | \ | For ray see https://docs.ray.io/en/latest/tune/api_docs/suggestion.html For optuna see https://optuna.readthedocs.io/en/stable/reference/samplers/index.html. |
+| num_samples | int | 10 | Number of hyperparameter optimization steps/samples. |
+| refit_with_val | bool | False | Refit of best model should preserve val_size. |
+| cpus | int | 4 | Number of cpus to use during optimization. Only used with ray tune. |
+| gpus | int | 0 | Number of gpus to use during optimization, default all available. Only used with ray tune. |
+| verbose | bool | False | Track progress. |
+| alias | NoneType | None | Custom name of the model. |
+| backend | str | ray | Backend to use for searching the hyperparameter space, can be either ‘ray’ or ‘optuna’. |
+| callbacks | NoneType | None | List of functions to call during the optimization process. ray reference: https://docs.ray.io/en/latest/tune/tutorials/tune-metrics.html optuna reference: https://optuna.readthedocs.io/en/stable/tutorial/20_recipes/007_optuna_callback.html |
+
+
+```python
+# Use your own config or AutoRMoK.default_config
+config = dict(max_steps=1, val_check_steps=1, input_size=12, learning_rate=1e-2)
+model = AutoRMoK(h=12, n_series=1, config=config, num_samples=1, cpus=1)
+
+# Fit and predict
+model.fit(dataset=dataset)
+y_hat = model.predict(dataset=dataset)
+
+# Optuna
+model = AutoRMoK(h=12, n_series=1, config=None, backend='optuna')
+```
+
+# TESTS
+
diff --git a/neuralforecast/models.informer.html.mdx b/neuralforecast/models.informer.html.mdx
new file mode 100644
index 00000000..4e60fc7d
--- /dev/null
+++ b/neuralforecast/models.informer.html.mdx
@@ -0,0 +1,307 @@
+---
+output-file: models.informer.html
+title: Informer
+---
+
+
+The Informer model tackles the vanilla Transformer computational
+complexity challenges for long-horizon forecasting.
+
+The architecture has three distinctive features: - A ProbSparse
+self-attention mechanism with an O time and memory complexity Llog(L). -
+A self-attention distilling process that prioritizes attention and
+efficiently handles long input sequences. - An MLP multi-step decoder
+that predicts long time-series sequences in a single forward operation
+rather than step-by-step.
+
+The Informer model utilizes a three-component approach to define its
+embedding: - It employs encoded autoregressive features obtained from a
+convolution network. - It uses window-relative positional embeddings
+derived from harmonic functions. - Absolute positional embeddings
+obtained from calendar features are utilized.
+
+**References** - [Haoyi Zhou, Shanghang Zhang, Jieqi Peng, Shuai
+Zhang, Jianxin Li, Hui Xiong, Wancai Zhang. “Informer: Beyond Efficient
+Transformer for Long Sequence Time-Series
+Forecasting”](https://arxiv.org/abs/2012.07436)
+
+
+
+Figure 1. Temporal Fusion Transformer
+Architecture.
+
+
+## 1. Auxiliary Functions
+
+------------------------------------------------------------------------
+
+source
+
+### ConvLayer
+
+> ``` text
+> ConvLayer (c_in)
+> ```
+
+*ConvLayer*
+
+------------------------------------------------------------------------
+
+source
+
+### ProbAttention
+
+> ``` text
+> ProbAttention (mask_flag=True, factor=5, scale=None,
+> attention_dropout=0.1, output_attention=False)
+> ```
+
+*ProbAttention*
+
+------------------------------------------------------------------------
+
+source
+
+### ProbMask
+
+> ``` text
+> ProbMask (B, H, L, index, scores, device='cpu')
+> ```
+
+*ProbMask*
+
+## 2. Informer
+
+------------------------------------------------------------------------
+
+source
+
+### Informer
+
+> ``` text
+> Informer (h:int, input_size:int, futr_exog_list=None,
+> hist_exog_list=None, stat_exog_list=None,
+> exclude_insample_y=False,
+> decoder_input_size_multiplier:float=0.5, hidden_size:int=128,
+> dropout:float=0.05, factor:int=3, n_head:int=4,
+> conv_hidden_size:int=32, activation:str='gelu',
+> encoder_layers:int=2, decoder_layers:int=1, distil:bool=True,
+> loss=MAE(), valid_loss=None, max_steps:int=5000,
+> learning_rate:float=0.0001, num_lr_decays:int=-1,
+> early_stop_patience_steps:int=-1, val_check_steps:int=100,
+> batch_size:int=32, valid_batch_size:Optional[int]=None,
+> windows_batch_size=1024, inference_windows_batch_size=1024,
+> start_padding_enabled=False, step_size:int=1,
+> scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*Informer
+
+``` text
+The Informer model tackles the vanilla Transformer computational complexity challenges for long-horizon forecasting.
+The architecture has three distinctive features:
+1) A ProbSparse self-attention mechanism with an O time and memory complexity Llog(L).
+2) A self-attention distilling process that prioritizes attention and efficiently handles long input sequences.
+3) An MLP multi-step decoder that predicts long time-series sequences in a single forward operation rather than step-by-step.
+```
+
+The Informer model utilizes a three-component approach to define its
+embedding: 1) It employs encoded autoregressive features obtained from a
+convolution network. 2) It uses window-relative positional embeddings
+derived from harmonic functions. 3) Absolute positional embeddings
+obtained from calendar features are utilized.
+
+*Parameters:* `h`: int, forecast horizon. `input_size`: int,
+maximum sequence length for truncated train backpropagation.
+`futr_exog_list`: str list, future exogenous columns.
+`hist_exog_list`: str list, historic exogenous columns.
+`stat_exog_list`: str list, static exogenous columns.
+`exclude_insample_y`: bool=False, the model skips the autoregressive
+features y\[t-input_size:t\] if True.
+`decoder_input_size_multiplier`: float = 0.5, . `hidden_size`:
+int=128, units of embeddings and encoders. `dropout`: float (0, 1),
+dropout throughout Informer architecture. `factor`: int=3,
+Probsparse attention factor. `n_head`: int=4, controls number of
+multi-head’s attention. `conv_hidden_size`: int=32, channels of the
+convolutional encoder. `activation`: str=`GELU`, activation from
+\[‘ReLU’, ‘Softplus’, ‘Tanh’, ‘SELU’, ‘LeakyReLU’, ‘PReLU’, ‘Sigmoid’,
+‘GELU’\]. `encoder_layers`: int=2, number of layers for the TCN
+encoder. `decoder_layers`: int=1, number of layers for the MLP
+decoder. `distil`: bool = True, wether the Informer decoder uses
+bottlenecks. `loss`: PyTorch module, instantiated train loss class
+from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=1024, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=1024, number of windows to sample in
+each inference batch. `start_padding_enabled`: bool=False, if True,
+the model will pad the time series with zeros at the beginning, by input
+size. `step_size`: int=1, step size between each window of temporal
+data. `scaler_type`: str=‘robust’, type of scaler for temporal
+inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`. `dataloader_kwargs`:
+dict, optional, list of parameters passed into the PyTorch Lightning
+dataloader by the `TimeSeriesDataLoader`. `**trainer_kwargs`: int,
+keyword trainer arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+``` text
+*References*
+- [Haoyi Zhou, Shanghang Zhang, Jieqi Peng, Shuai Zhang, Jianxin Li, Hui Xiong, Wancai Zhang. "Informer: Beyond Efficient Transformer for Long Sequence Time-Series Forecasting"](https://arxiv.org/abs/2012.07436) *
+```
+
+------------------------------------------------------------------------
+
+### Informer.fit
+
+> ``` text
+> Informer.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### Informer.predict
+
+> ``` text
+> Informer.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import Informer
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic, augment_calendar_df
+
+AirPassengersPanel, calendar_cols = augment_calendar_df(df=AirPassengersPanel, freq='M')
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = Informer(h=12,
+ input_size=24,
+ hidden_size = 16,
+ conv_hidden_size = 32,
+ n_head = 2,
+ loss=MAE(),
+ futr_exog_list=calendar_cols,
+ scaler_type='robust',
+ learning_rate=1e-3,
+ max_steps=200,
+ val_check_steps=50,
+ early_stop_patience_steps=2)
+
+nf = NeuralForecast(
+ models=[model],
+ freq='ME'
+)
+nf.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+forecasts = nf.predict(futr_df=Y_test_df)
+
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+if model.loss.is_distribution_output:
+ plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+ plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+ plt.plot(plot_df['ds'], plot_df['Informer-median'], c='blue', label='median')
+ plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['Informer-lo-90'][-12:].values,
+ y2=plot_df['Informer-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+ plt.grid()
+ plt.legend()
+ plt.plot()
+else:
+ plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+ plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+ plt.plot(plot_df['ds'], plot_df['Informer'], c='blue', label='Forecast')
+ plt.legend()
+ plt.grid()
+```
+
diff --git a/neuralforecast/models.itransformer.html.mdx b/neuralforecast/models.itransformer.html.mdx
new file mode 100644
index 00000000..6b0c10d5
--- /dev/null
+++ b/neuralforecast/models.itransformer.html.mdx
@@ -0,0 +1,228 @@
+---
+output-file: models.itransformer.html
+title: iTransformer
+---
+
+
+The iTransformer model simply takes the Transformer architecture but it
+applies the attention and feed-forward network on the inverted
+dimensions. This means that time points of each individual series are
+embedded into tokens. That way, the attention mechanisms learn
+multivariate correlation and the feed-forward network learns non-linear
+relationships.
+
+**References** - [Yong Liu, Tengge Hu, Haoran Zhang, Haixu Wu, Shiyu
+Wang, Lintao Ma, Mingsheng Long. “iTransformer: Inverted Transformers
+Are Effective for Time Series
+Forecasting”](https://arxiv.org/abs/2310.06625)
+
+
+
+Figure 1. Architecture of
+iTransformer.
+
+
+# 1. Model
+
+------------------------------------------------------------------------
+
+source
+
+### iTransformer
+
+> ``` text
+> iTransformer (h, input_size, n_series, futr_exog_list=None,
+> hist_exog_list=None, stat_exog_list=None,
+> exclude_insample_y=False, hidden_size:int=512,
+> n_heads:int=8, e_layers:int=2, d_layers:int=1,
+> d_ff:int=2048, factor:int=1, dropout:float=0.1,
+> use_norm:bool=True, loss=MAE(), valid_loss=None,
+> max_steps:int=1000, learning_rate:float=0.001,
+> num_lr_decays:int=-1, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size:int=32,
+> valid_batch_size:Optional[int]=None, windows_batch_size=32,
+> inference_windows_batch_size=32,
+> start_padding_enabled=False, step_size:int=1,
+> scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*iTransformer
+
+**Parameters:** `h`: int, Forecast horizon. `input_size`: int,
+autorregresive inputs size, y=\[1,2,3,4\] input_size=2 -\>
+y\_\[t-2:t\]=\[1,2\]. `n_series`: int, number of time-series.
+`futr_exog_list`: str list, future exogenous columns.
+`hist_exog_list`: str list, historic exogenous columns.
+`stat_exog_list`: str list, static exogenous columns.
+`exclude_insample_y`: bool=False, the model skips the autoregressive
+features y\[t-input_size:t\] if True.
+`hidden_size`: int, dimension of the model. `n_heads`: int, number
+of heads. `e_layers`: int, number of encoder layers. `d_layers`:
+int, number of decoder layers. `d_ff`: int, dimension of
+fully-connected layer. `factor`: int, attention factor.
+`dropout`: float, dropout rate. `use_norm`: bool, whether to
+normalize or not. `loss`: PyTorch module, instantiated train loss
+class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=32, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=32, number of windows to sample in
+each inference batch, -1 uses all. `start_padding_enabled`:
+bool=False, if True, the model will pad the time series with zeros at
+the beginning, by input size. `step_size`: int=1, step size between
+each window of temporal data. `scaler_type`: str=‘identity’, type of
+scaler for temporal inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`. `dataloader_kwargs`:
+dict, optional, list of parameters passed into the PyTorch Lightning
+dataloader by the `TimeSeriesDataLoader`. `**trainer_kwargs`: int,
+keyword trainer arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+**References** - [Yong Liu, Tengge Hu, Haoran Zhang, Haixu Wu, Shiyu
+Wang, Lintao Ma, Mingsheng Long. “iTransformer: Inverted Transformers
+Are Effective for Time Series
+Forecasting”](https://arxiv.org/abs/2310.06625)\*
+
+------------------------------------------------------------------------
+
+### iTransformer.fit
+
+> ``` text
+> iTransformer.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### iTransformer.predict
+
+> ``` text
+> iTransformer.predict (dataset, test_size=None, step_size=1,
+> random_seed=None, quantiles=None,
+> **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+# 2. Usage example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import iTransformer
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+from neuralforecast.losses.pytorch import MSE
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = iTransformer(h=12,
+ input_size=24,
+ n_series=2,
+ hidden_size=128,
+ n_heads=2,
+ e_layers=2,
+ d_layers=1,
+ d_ff=4,
+ factor=1,
+ dropout=0.1,
+ use_norm=True,
+ loss=MSE(),
+ valid_loss=MAE(),
+ early_stop_patience_steps=3,
+ batch_size=32,
+ max_steps=100)
+
+fcst = NeuralForecast(models=[model], freq='ME')
+fcst.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+forecasts = fcst.predict(futr_df=Y_test_df)
+
+# Plot predictions
+fig, ax = plt.subplots(1, 1, figsize = (20, 7))
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['iTransformer'], c='blue', label='Forecast')
+ax.set_title('AirPassengers Forecast', fontsize=22)
+ax.set_ylabel('Monthly Passengers', fontsize=20)
+ax.set_xlabel('Year', fontsize=20)
+ax.legend(prop={'size': 15})
+ax.grid()
+```
+
diff --git a/neuralforecast/models.kan.html.mdx b/neuralforecast/models.kan.html.mdx
new file mode 100644
index 00000000..39038a4d
--- /dev/null
+++ b/neuralforecast/models.kan.html.mdx
@@ -0,0 +1,239 @@
+---
+output-file: models.kan.html
+title: KAN
+---
+
+
+Kolmogorov-Arnold Networks (KANs) are an alternative to Multi-Layer
+Perceptrons (MLPs). This model uses KANs similarly as our MLP model.
+
+**References** - [Ziming Liu, Yixuan Wang, Sachin Vaidya, Fabian Ruehle,
+James Halverson, Marin Soljačić, Thomas Y. Hou, Max Tegmark. “KAN:
+Kolmogorov–Arnold Networks”](https://arxiv.org/html/2404.19756v1)
+
+
+
+Figure 1. KAN compared to
+MLP.
+
+
+------------------------------------------------------------------------
+
+source
+
+### KANLinear
+
+> ``` text
+> KANLinear (in_features, out_features, grid_size=5, spline_order=3,
+> scale_noise=0.1, scale_base=1.0, scale_spline=1.0,
+> enable_standalone_scale_spline=True, base_activation= 'torch.nn.modules.activation.SiLU'>, grid_eps=0.02,
+> grid_range=[-1, 1])
+> ```
+
+*KANLinear*
+
+------------------------------------------------------------------------
+
+source
+
+### KAN
+
+> ``` text
+> KAN (h, input_size, grid_size:int=5, spline_order:int=3,
+> scale_noise:float=0.1, scale_base:float=1.0, scale_spline:float=1.0,
+> enable_standalone_scale_spline:bool=True, grid_eps:float=0.02,
+> grid_range:list=[-1, 1], n_hidden_layers:int=1,
+> hidden_size:Union[int,list]=512, stat_exog_list=None,
+> hist_exog_list=None, futr_exog_list=None, exclude_insample_y=False,
+> loss=MAE(), valid_loss=None, max_steps:int=1000,
+> learning_rate:float=0.001, num_lr_decays:int=-1,
+> early_stop_patience_steps:int=-1, val_check_steps:int=100,
+> batch_size:int=32, valid_batch_size:Optional[int]=None,
+> windows_batch_size=1024, inference_windows_batch_size=-1,
+> start_padding_enabled=False, step_size:int=1,
+> scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*KAN
+
+Simple Kolmogorov-Arnold Network (KAN). This network uses the
+Kolmogorov-Arnold approximation theorem, where splines are learned to
+approximate more complex functions. Unlike the MLP, the non-linear
+function are learned at the edges, and the nodes simply sum the
+different learned functions.
+
+**Parameters:** `h`: int, forecast horizon. `input_size`: int,
+considered autorregresive inputs (lags), y=\[1,2,3,4\] input_size=2 -\>
+lags=\[1,2\]. `grid_size`: int, number of intervals used by the
+splines to approximate the function. `spline_order`: int, order of
+the B-splines. `scale_noise`: float, regularization coefficient for
+the splines. `scale_base`: float, scaling coefficient for the base
+function. `scale_spline`: float, scaling coefficient for the
+splines. `enable_standalone_scale_spline`: bool, whether each spline
+is scaled individually. `grid_eps`: float, used for numerical
+stability. `grid_range`: list, range of the grid used for spline
+approximation. `n_hidden_layers`: int, number of hidden layers for
+the KAN. `hidden_size`: int or list, number of units for each hidden
+layer of the KAN. If an integer, all hidden layers will have the same
+size. Use a list to specify the size of each hidden layer.
+`stat_exog_list`: str list, static exogenous columns.
+`hist_exog_list`: str list, historic exogenous columns.
+`futr_exog_list`: str list, future exogenous columns.
+`exclude_insample_y`: bool=False, the model skips the autoregressive
+features y\[t-input_size:t\] if True. `loss`: PyTorch module,
+instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=1024, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=-1, number of windows to sample in
+each inference batch, -1 uses all. `start_padding_enabled`:
+bool=False, if True, the model will pad the time series with zeros at
+the beginning, by input size. `step_size`: int=1, step size between
+each window of temporal data. `scaler_type`: str=‘identity’, type of
+scaler for temporal inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`dataloader_kwargs`: dict, optional, list of parameters passed into the
+PyTorch Lightning dataloader by the `TimeSeriesDataLoader`.
+`**trainer_kwargs`: int, keyword trainer arguments inherited from
+[PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+**References** - [Ziming Liu, Yixuan Wang, Sachin Vaidya, Fabian
+Ruehle, James Halverson, Marin Soljačić, Thomas Y. Hou, Max Tegmark.
+“KAN: Kolmogorov-Arnold Networks”](https://arxiv.org/abs/2404.19756)\*
+
+------------------------------------------------------------------------
+
+### KAN.fit
+
+> ``` text
+> KAN.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### KAN.predict
+
+> ``` text
+> KAN.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import KAN
+from neuralforecast.losses.pytorch import DistributionLoss
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+fcst = NeuralForecast(
+ models=[
+ KAN(h=12,
+ input_size=24,
+ loss = DistributionLoss(distribution="Normal"),
+ max_steps=100,
+ scaler_type='standard',
+ futr_exog_list=['y_[lag12]'],
+ hist_exog_list=None,
+ stat_exog_list=['airline1'],
+ ),
+ ],
+ freq='ME'
+)
+fcst.fit(df=Y_train_df, static_df=AirPassengersStatic)
+forecasts = fcst.predict(futr_df=Y_test_df)
+
+# Plot quantile predictions
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['KAN-median'], c='blue', label='median')
+plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['KAN-lo-90'][-12:].values,
+ y2=plot_df['KAN-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+plt.legend()
+plt.grid()
+```
+
diff --git a/neuralforecast/models.lstm.html.mdx b/neuralforecast/models.lstm.html.mdx
new file mode 100644
index 00000000..a12ff124
--- /dev/null
+++ b/neuralforecast/models.lstm.html.mdx
@@ -0,0 +1,245 @@
+---
+output-file: models.lstm.html
+title: LSTM
+---
+
+
+The Long Short-Term Memory Recurrent Neural Network
+([`LSTM`](https://nixtlaverse.nixtla.io/neuralforecast/models.lstm.html#lstm)),
+uses a multilayer
+[`LSTM`](https://nixtlaverse.nixtla.io/neuralforecast/models.lstm.html#lstm)
+encoder and an
+[`MLP`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlp.html#mlp)
+decoder. It builds upon the LSTM-cell that improves the exploding and
+vanishing gradients of classic
+[`RNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.rnn.html#rnn)’s.
+This network has been extensively used in sequential prediction tasks
+like language modeling, phonetic labeling, and forecasting. The
+predictions are obtained by transforming the hidden states into contexts
+$\mathbf{c}_{[t+1:t+H]}$, that are decoded and adapted into
+$\mathbf{\hat{y}}_{[t+1:t+H],[q]}$ through MLPs.
+
+where $\mathbf{h}_{t}$, is the hidden state for time $t$,
+$\mathbf{y}_{t}$ is the input at time $t$ and $\mathbf{h}_{t-1}$ is the
+hidden state of the previous layer at $t-1$, $\mathbf{x}^{(s)}$ are
+static exogenous inputs, $\mathbf{x}^{(h)}_{t}$ historic exogenous,
+$\mathbf{x}^{(f)}_{[:t+H]}$ are future exogenous available at the time
+of the prediction.
+
+**References** -[Jeffrey L. Elman (1990). “Finding Structure in
+Time”.](https://onlinelibrary.wiley.com/doi/abs/10.1207/s15516709cog1402_1) -[Haşim
+Sak, Andrew Senior, Françoise Beaufays (2014). “Long Short-Term Memory
+Based Recurrent Neural Network Architectures for Large Vocabulary Speech
+Recognition.”](https://arxiv.org/abs/1402.1128)
+
+
+
+Figure 1. Long Short-Term Memory
+Cell.
+
+
+------------------------------------------------------------------------
+
+source
+
+### LSTM
+
+> ``` text
+> LSTM (h:int, input_size:int=-1, inference_input_size:Optional[int]=None,
+> h_train:int=1, encoder_n_layers:int=2, encoder_hidden_size:int=128,
+> encoder_bias:bool=True, encoder_dropout:float=0.0,
+> context_size:Optional[int]=None, decoder_hidden_size:int=128,
+> decoder_layers:int=2, futr_exog_list=None, hist_exog_list=None,
+> stat_exog_list=None, exclude_insample_y=False, recurrent=False,
+> loss=MAE(), valid_loss=None, max_steps:int=1000,
+> learning_rate:float=0.001, num_lr_decays:int=-1,
+> early_stop_patience_steps:int=-1, val_check_steps:int=100,
+> batch_size=32, valid_batch_size:Optional[int]=None,
+> windows_batch_size=128, inference_windows_batch_size=1024,
+> start_padding_enabled=False, step_size:int=1,
+> scaler_type:str='robust', random_seed=1, drop_last_loader=False,
+> alias:Optional[str]=None, optimizer=None, optimizer_kwargs=None,
+> lr_scheduler=None, lr_scheduler_kwargs=None,
+> dataloader_kwargs=None, **trainer_kwargs)
+> ```
+
+\*LSTM
+
+LSTM encoder, with MLP decoder. The network has `tanh` or `relu`
+non-linearities, it is trained using ADAM stochastic gradient descent.
+The network accepts static, historic and future exogenous data.
+
+**Parameters:** `h`: int, forecast horizon. `input_size`: int,
+maximum sequence length for truncated train backpropagation. Default -1
+uses 3 \* horizon `inference_input_size`: int, maximum sequence
+length for truncated inference. Default None uses input_size
+history. `h_train`: int, maximum sequence length for truncated train
+backpropagation. Default 1. `encoder_n_layers`: int=2, number of
+layers for the LSTM. `encoder_hidden_size`: int=200, units for the
+LSTM’s hidden state size. `encoder_bias`: bool=True, whether or not
+to use biases b_ih, b_hh within LSTM units. `encoder_dropout`:
+float=0., dropout regularization applied to LSTM outputs.
+`context_size`: deprecated. `decoder_hidden_size`: int=200, size of
+hidden layer for the MLP decoder. `decoder_layers`: int=2, number of
+layers for the MLP decoder. `futr_exog_list`: str list, future
+exogenous columns. `hist_exog_list`: str list, historic exogenous
+columns. `stat_exog_list`: str list, static exogenous columns.
+`exclude_insample_y`: bool=False, whether to exclude the target variable
+from the input. `recurrent`: bool=False, whether to produce
+forecasts recursively (True) or direct (False). `loss`: PyTorch
+module, instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of
+differentseries in each batch. `valid_batch_size`: int=None, number
+of different series in each validation and test batch.
+`windows_batch_size`: int=128, number of windows to sample in each
+training batch, default uses all. `inference_windows_batch_size`:
+int=1024, number of windows to sample in each inference batch, -1 uses
+all. `start_padding_enabled`: bool=False, if True, the model will
+pad the time series with zeros at the beginning, by input size.
+`step_size`: int=1, step size between each window of temporal
+data.
+`scaler_type`: str=‘robust’, type of scaler for temporal inputs
+normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`.
+`dataloader_kwargs`: dict, optional, list of parameters passed into the
+PyTorch Lightning dataloader by the `TimeSeriesDataLoader`.
+`**trainer_kwargs`: int, keyword trainer arguments inherited from
+[PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer). \*
+
+------------------------------------------------------------------------
+
+### LSTM.fit
+
+> ``` text
+> LSTM.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### LSTM.predict
+
+> ``` text
+> LSTM.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import LSTM
+from neuralforecast.losses.pytorch import DistributionLoss
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+nf = NeuralForecast(
+ models=[LSTM(h=12,
+ input_size=8,
+ loss=DistributionLoss(distribution="Normal", level=[80, 90]),
+ scaler_type='robust',
+ encoder_n_layers=2,
+ encoder_hidden_size=128,
+ decoder_hidden_size=128,
+ decoder_layers=2,
+ max_steps=200,
+ futr_exog_list=['y_[lag12]'],
+ stat_exog_list=['airline1'],
+ recurrent=True,
+ h_train=1,
+ )
+ ],
+ freq='ME'
+)
+nf.fit(df=Y_train_df, static_df=AirPassengersStatic)
+Y_hat_df = nf.predict(futr_df=Y_test_df)
+
+# Plots
+Y_hat_df = Y_hat_df.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['LSTM-median'], c='blue', label='median')
+plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['LSTM-lo-90'][-12:].values,
+ y2=plot_df['LSTM-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+plt.grid()
+plt.plot()
+```
+
diff --git a/neuralforecast/models.mlp.html.mdx b/neuralforecast/models.mlp.html.mdx
new file mode 100644
index 00000000..ee6ba633
--- /dev/null
+++ b/neuralforecast/models.mlp.html.mdx
@@ -0,0 +1,217 @@
+---
+description: >-
+ One of the simplest neural architectures are Multi Layer Perceptrons (`MLP`)
+ composed of stacked Fully Connected Neural Networks trained with
+ backpropagation. Each node in the architecture is capable of modeling
+ non-linear relationships granted by their activation functions. Novel
+ activations like Rectified Linear Units (`ReLU`) have greatly improved the
+ ability to fit deeper networks overcoming gradient vanishing problems that
+ were associated with `Sigmoid` and `TanH` activations. For the forecasting
+ task the last layer is changed to follow a auto-regression
+ problem.
**References** -[Rosenblatt, F. (1958). "The perceptron: A
+ probabilistic model for information storage and organization in the
+ brain."](https://psycnet.apa.org/record/1959-09865-001) -[Fukushima, K.
+ (1975). "Cognitron: A self-organizing multilayered neural
+ network."](https://pascal-francis.inist.fr/vibad/index.php?action=getRecordDetail&idt=PASCAL7750396723) -[Vinod
+ Nair, Geoffrey E. Hinton (2010). "Rectified Linear Units Improve Restricted
+ Boltzmann Machines"](https://www.cs.toronto.edu/~fritz/absps/reluICML.pdf)
+output-file: models.mlp.html
+title: MLP
+---
+
+
+
+
+Figure 1. Three layer MLP with
+autorregresive inputs.
+
+
+------------------------------------------------------------------------
+
+source
+
+### MLP
+
+> ``` text
+> MLP (h, input_size, stat_exog_list=None, hist_exog_list=None,
+> futr_exog_list=None, exclude_insample_y=False, num_layers=2,
+> hidden_size=1024, loss=MAE(), valid_loss=None, max_steps:int=1000,
+> learning_rate:float=0.001, num_lr_decays:int=-1,
+> early_stop_patience_steps:int=-1, val_check_steps:int=100,
+> batch_size:int=32, valid_batch_size:Optional[int]=None,
+> windows_batch_size=1024, inference_windows_batch_size=-1,
+> start_padding_enabled=False, step_size:int=1,
+> scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None, **trainer_kwargs)
+> ```
+
+\*MLP
+
+Simple Multi Layer Perceptron architecture (MLP). This deep neural
+network has constant units through its layers, each with ReLU
+non-linearities, it is trained using ADAM stochastic gradient descent.
+The network accepts static, historic and future exogenous data, flattens
+the inputs and learns fully connected relationships against the target
+variable.
+
+**Parameters:** `h`: int, forecast horizon. `input_size`: int,
+considered autorregresive inputs (lags), y=\[1,2,3,4\] input_size=2 -\>
+lags=\[1,2\]. `stat_exog_list`: str list, static exogenous
+columns. `hist_exog_list`: str list, historic exogenous columns.
+`futr_exog_list`: str list, future exogenous columns.
+`exclude_insample_y`: bool=False, the model skips the autoregressive
+features y\[t-input_size:t\] if True. `num_layers`: int, number of
+layers for the MLP. `hidden_size`: int, number of units for each
+layer of the MLP. `loss`: PyTorch module, instantiated train loss
+class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=1024, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=-1, number of windows to sample in
+each inference batch, -1 uses all. `start_padding_enabled`:
+bool=False, if True, the model will pad the time series with zeros at
+the beginning, by input size. `step_size`: int=1, step size between
+each window of temporal data. `scaler_type`: str=‘identity’, type of
+scaler for temporal inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`.
+`dataloader_kwargs`: dict, optional, list of parameters passed into the
+PyTorch Lightning dataloader by the `TimeSeriesDataLoader`.
+`**trainer_kwargs`: int, keyword trainer arguments inherited from
+[PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer). \*
+
+------------------------------------------------------------------------
+
+### MLP.fit
+
+> ``` text
+> MLP.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### MLP.predict
+
+> ``` text
+> MLP.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import MLP
+from neuralforecast.losses.pytorch import DistributionLoss
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = MLP(h=12, input_size=24,
+ loss=DistributionLoss(distribution='Normal', level=[80, 90]),
+ scaler_type='robust',
+ learning_rate=1e-3,
+ max_steps=200,
+ val_check_steps=10,
+ early_stop_patience_steps=2)
+
+fcst = NeuralForecast(
+ models=[model],
+ freq='ME'
+)
+fcst.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+forecasts = fcst.predict(futr_df=Y_test_df)
+
+# Plot predictions
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['MLP-median'], c='blue', label='median')
+plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['MLP-lo-90'][-12:].values,
+ y2=plot_df['MLP-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+plt.grid()
+plt.legend()
+plt.plot()
+```
+
diff --git a/neuralforecast/models.mlpmultivariate.html.mdx b/neuralforecast/models.mlpmultivariate.html.mdx
new file mode 100644
index 00000000..14163974
--- /dev/null
+++ b/neuralforecast/models.mlpmultivariate.html.mdx
@@ -0,0 +1,222 @@
+---
+description: >-
+ One of the simplest neural architectures are Multi Layer Perceptrons (`MLP`)
+ composed of stacked Fully Connected Neural Networks trained with
+ backpropagation. Each node in the architecture is capable of modeling
+ non-linear relationships granted by their activation functions. Novel
+ activations like Rectified Linear Units (`ReLU`) have greatly improved the
+ ability to fit deeper networks overcoming gradient vanishing problems that
+ were associated with `Sigmoid` and `TanH` activations. For the forecasting
+ task the last layer is changed to follow a auto-regression problem. This
+ version is multivariate, indicating that it will predict all time series of
+ the forecasting problem jointly.
**References** -[Rosenblatt, F.
+ (1958). "The perceptron: A probabilistic model for information storage and
+ organization in the
+ brain."](https://psycnet.apa.org/record/1959-09865-001) -[Fukushima, K.
+ (1975). "Cognitron: A self-organizing multilayered neural
+ network."](https://pascal-francis.inist.fr/vibad/index.php?action=getRecordDetail&idt=PASCAL7750396723) -[Vinod
+ Nair, Geoffrey E. Hinton (2010). "Rectified Linear Units Improve Restricted
+ Boltzmann Machines"](https://www.cs.toronto.edu/~fritz/absps/reluICML.pdf)
+output-file: models.mlpmultivariate.html
+title: MLPMultivariate
+---
+
+
+
+
+Figure 1. Three layer MLP with
+autorregresive inputs.
+
+
+------------------------------------------------------------------------
+
+source
+
+### MLPMultivariate
+
+> ``` text
+> MLPMultivariate (h, input_size, n_series, stat_exog_list=None,
+> hist_exog_list=None, futr_exog_list=None,
+> exclude_insample_y=False, num_layers=2,
+> hidden_size=1024, loss=MAE(), valid_loss=None,
+> max_steps:int=1000, learning_rate:float=0.001,
+> num_lr_decays:int=-1, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size:int=32,
+> valid_batch_size:Optional[int]=None,
+> windows_batch_size=32, inference_windows_batch_size=32,
+> start_padding_enabled=False, step_size:int=1,
+> scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None,
+> lr_scheduler=None, lr_scheduler_kwargs=None,
+> dataloader_kwargs=None, **trainer_kwargs)
+> ```
+
+\*MLPMultivariate
+
+Simple Multi Layer Perceptron architecture (MLP) for multivariate
+forecasting. This deep neural network has constant units through its
+layers, each with ReLU non-linearities, it is trained using ADAM
+stochastic gradient descent. The network accepts static, historic and
+future exogenous data, flattens the inputs and learns fully connected
+relationships against the target variables.
+
+**Parameters:** `h`: int, forecast horizon. `input_size`: int,
+considered autorregresive inputs (lags), y=\[1,2,3,4\] input_size=2 -\>
+lags=\[1,2\]. `n_series`: int, number of time-series.
+`stat_exog_list`: str list, static exogenous columns.
+`hist_exog_list`: str list, historic exogenous columns.
+`futr_exog_list`: str list, future exogenous columns. `num_layers`:
+int, number of layers for the MLP. `hidden_size`: int, number of
+units for each layer of the MLP. `loss`: PyTorch module,
+instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=32, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=32, number of windows to sample in
+each inference batch, -1 uses all. `start_padding_enabled`:
+bool=False, if True, the model will pad the time series with zeros at
+the beginning, by input size. `step_size`: int=1, step size between
+each window of temporal data. `scaler_type`: str=‘identity’, type of
+scaler for temporal inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`. `dataloader_kwargs`:
+dict, optional, list of parameters passed into the PyTorch Lightning
+dataloader by the `TimeSeriesDataLoader`. `**trainer_kwargs`: int,
+keyword trainer arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer). \*
+
+------------------------------------------------------------------------
+
+### MLPMultivariate.fit
+
+> ``` text
+> MLPMultivariate.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### MLPMultivariate.predict
+
+> ``` text
+> MLPMultivariate.predict (dataset, test_size=None, step_size=1,
+> random_seed=None, quantiles=None,
+> **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import MLPMultivariate
+from neuralforecast.losses.pytorch import MAE
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = MLPMultivariate(h=12,
+ input_size=24,
+ n_series=2,
+ stat_exog_list=['airline1'],
+ futr_exog_list=['trend'],
+ loss = MAE(),
+ scaler_type='robust',
+ learning_rate=1e-3,
+ stat_exog_list=['airline1'],
+ max_steps=200,
+ val_check_steps=10,
+ early_stop_patience_steps=2)
+
+fcst = NeuralForecast(
+ models=[model],
+ freq='ME'
+)
+fcst.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+forecasts = fcst.predict(futr_df=Y_test_df)
+
+# Plot predictions
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['MLPMultivariate'], c='blue', label='median')
+plt.grid()
+plt.legend()
+plt.plot()
+```
+
diff --git a/neuralforecast/models.nbeats.html.mdx b/neuralforecast/models.nbeats.html.mdx
new file mode 100644
index 00000000..b94361e4
--- /dev/null
+++ b/neuralforecast/models.nbeats.html.mdx
@@ -0,0 +1,250 @@
+---
+output-file: models.nbeats.html
+title: NBEATS
+---
+
+
+The Neural Basis Expansion Analysis
+([`NBEATS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nbeats.html#nbeats))
+is an
+[`MLP`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlp.html#mlp)-based
+deep neural architecture with backward and forward residual links. The
+network has two variants: (1) in its interpretable configuration,
+[`NBEATS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nbeats.html#nbeats)
+sequentially projects the signal into polynomials and harmonic basis to
+learn trend and seasonality components; (2) in its generic
+configuration, it substitutes the polynomial and harmonic basis for
+identity basis and larger network’s depth. The Neural Basis Expansion
+Analysis with Exogenous
+([`NBEATSx`](https://nixtlaverse.nixtla.io/neuralforecast/models.nbeatsx.html#nbeatsx)),
+incorporates projections to exogenous temporal variables available at
+the time of the prediction.
+
+This method proved state-of-the-art performance on the M3, M4, and
+Tourism Competition datasets, improving accuracy by 3% over the `ESRNN`
+M4 competition winner.
+
+**References** -[Boris N. Oreshkin, Dmitri Carpov, Nicolas Chapados,
+Yoshua Bengio (2019). “N-BEATS: Neural basis expansion analysis for
+interpretable time series
+forecasting”.](https://arxiv.org/abs/1905.10437)
+
+
+
+Figure 1. Neural Basis Expansion
+Analysis.
+
+
+------------------------------------------------------------------------
+
+source
+
+### NBEATS
+
+> ``` text
+> NBEATS (h, input_size, n_harmonics:int=2,
+> n_polynomials:Optional[int]=None, n_basis:int=2,
+> basis:str='polynomial', stack_types:list=['identity', 'trend',
+> 'seasonality'], n_blocks:list=[1, 1, 1], mlp_units:list=[[512,
+> 512], [512, 512], [512, 512]], dropout_prob_theta:float=0.0,
+> activation:str='ReLU', shared_weights:bool=False, loss=MAE(),
+> valid_loss=None, max_steps:int=1000, learning_rate:float=0.001,
+> num_lr_decays:int=3, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size:int=32,
+> valid_batch_size:Optional[int]=None, windows_batch_size:int=1024,
+> inference_windows_batch_size:int=-1, start_padding_enabled=False,
+> step_size:int=1, scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*NBEATS
+
+The Neural Basis Expansion Analysis for Time Series (NBEATS), is a
+simple and yet effective architecture, it is built with a deep stack of
+MLPs with the doubly residual connections. It has a generic and
+interpretable architecture depending on the blocks it uses. Its
+interpretable architecture is recommended for scarce data settings, as
+it regularizes its predictions through projections unto harmonic and
+trend basis well-suited for most forecasting tasks.
+
+**Parameters:** `h`: int, forecast horizon. `input_size`: int,
+considered autorregresive inputs (lags), y=\[1,2,3,4\] input_size=2 -\>
+lags=\[1,2\]. `n_harmonics`: int, Number of harmonic terms for
+seasonality stack type. Note that len(n_harmonics) = len(stack_types).
+Note that it will only be used if a seasonality stack is used.
+`n_polynomials`: int, DEPRECATED - polynomial degree for trend stack.
+Note that len(n_polynomials) = len(stack_types). Note that it will only
+be used if a trend stack is used. `basis`: str, Type of basis
+function to use in the trend stack. Choose one from \[‘legendre’,
+‘polynomial’, ‘changepoint’, ‘piecewise_linear’, ‘linear_hat’, ‘spline’,
+‘chebyshev’\] `n_basis`: int, the degree of the basis function for
+the trend stack. Note that it will only be used if a trend stack is
+used. `stack_types`: List\[str\], List of stack types. Subset from
+\[‘seasonality’, ‘trend’, ‘identity’\]. `n_blocks`: List\[int\],
+Number of blocks for each stack. Note that len(n_blocks) =
+len(stack_types). `mlp_units`: List\[List\[int\]\], Structure of
+hidden layers for each stack type. Each internal list should contain the
+number of units of each hidden layer. Note that len(n_hidden) =
+len(stack_types). `dropout_prob_theta`: float, Float between (0, 1).
+Dropout for N-BEATS basis. `activation`: str, activation from
+\[‘ReLU’, ‘Softplus’, ‘Tanh’, ‘SELU’, ‘LeakyReLU’, ‘PReLU’,
+‘Sigmoid’\]. `shared_weights`: bool, If True, all blocks within each
+stack will share parameters. `loss`: PyTorch module, instantiated
+train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=3, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=1024, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=-1, number of windows to sample in
+each inference batch, -1 uses all. `start_padding_enabled`:
+bool=False, if True, the model will pad the time series with zeros at
+the beginning, by input size. `step_size`: int=1, step size between
+each window of temporal data. `scaler_type`: str=‘identity’, type of
+scaler for temporal inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`. `dataloader_kwargs`:
+dict, optional, list of parameters passed into the PyTorch Lightning
+dataloader by the `TimeSeriesDataLoader`. `**trainer_kwargs`: int,
+keyword trainer arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+**References:** -[Boris N. Oreshkin, Dmitri Carpov, Nicolas
+Chapados, Yoshua Bengio (2019). “N-BEATS: Neural basis expansion
+analysis for interpretable time series
+forecasting”.](https://arxiv.org/abs/1905.10437)\*
+
+------------------------------------------------------------------------
+
+### NBEATS.fit
+
+> ``` text
+> NBEATS.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### NBEATS.predict
+
+> ``` text
+> NBEATS.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import NBEATS
+from neuralforecast.losses.pytorch import DistributionLoss
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = NBEATS(h=12, input_size=24,
+ basis='changepoint',
+ n_basis=2,
+ loss=DistributionLoss(distribution='Poisson', level=[80, 90]),
+ stack_types = ['identity', 'trend', 'seasonality'],
+ max_steps=100,
+ val_check_steps=10,
+ early_stop_patience_steps=2)
+
+fcst = NeuralForecast(
+ models=[model],
+ freq='ME'
+)
+fcst.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+forecasts = fcst.predict(futr_df=Y_test_df)
+
+# Plot quantile predictions
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['NBEATS-median'], c='blue', label='median')
+plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['NBEATS-lo-90'][-12:].values,
+ y2=plot_df['NBEATS-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+plt.grid()
+plt.legend()
+plt.plot()
+```
+
diff --git a/neuralforecast/models.nbeatsx.html.mdx b/neuralforecast/models.nbeatsx.html.mdx
new file mode 100644
index 00000000..bbf58a1c
--- /dev/null
+++ b/neuralforecast/models.nbeatsx.html.mdx
@@ -0,0 +1,258 @@
+---
+output-file: models.nbeatsx.html
+title: NBEATSx
+---
+
+
+The Neural Basis Expansion Analysis
+([`NBEATS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nbeats.html#nbeats))
+is an
+[`MLP`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlp.html#mlp)-based
+deep neural architecture with backward and forward residual links. The
+network has two variants: (1) in its interpretable configuration,
+[`NBEATS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nbeats.html#nbeats)
+sequentially projects the signal into polynomials and harmonic basis to
+learn trend and seasonality components; (2) in its generic
+configuration, it substitutes the polynomial and harmonic basis for
+identity basis and larger network’s depth. The Neural Basis Expansion
+Analysis with Exogenous
+([`NBEATSx`](https://nixtlaverse.nixtla.io/neuralforecast/models.nbeatsx.html#nbeatsx)),
+incorporates projections to exogenous temporal variables available at
+the time of the prediction.
This method proved state-of-the-art
+performance on the M3, M4, and Tourism Competition datasets, improving
+accuracy by 3% over the `ESRNN` M4 competition winner. For Electricity
+Price Forecasting tasks
+[`NBEATSx`](https://nixtlaverse.nixtla.io/neuralforecast/models.nbeatsx.html#nbeatsx)
+model improved accuracy by 20% and 5% over `ESRNN` and
+[`NBEATS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nbeats.html#nbeats),
+and 5% on task-specialized
+architectures.
**References** -[Boris N. Oreshkin, Dmitri
+Carpov, Nicolas Chapados, Yoshua Bengio (2019). “N-BEATS: Neural basis
+expansion analysis for interpretable time series
+forecasting”.](https://arxiv.org/abs/1905.10437) -[Kin G. Olivares,
+Cristian Challu, Grzegorz Marcjasz, Rafał Weron, Artur Dubrawski (2021).
+“Neural basis expansion analysis with exogenous variables: Forecasting
+electricity prices with NBEATSx”.](https://arxiv.org/abs/2104.05522)
+
+
+
+Figure 1. Neural Basis Expansion Analysis
+with Exogenous Variables.
+
+
+------------------------------------------------------------------------
+
+source
+
+### NBEATSx
+
+> ``` text
+> NBEATSx (h, input_size, futr_exog_list=None, hist_exog_list=None,
+> stat_exog_list=None, exclude_insample_y=False, n_harmonics=2,
+> n_polynomials=2, stack_types:list=['identity', 'trend',
+> 'seasonality'], n_blocks:list=[1, 1, 1], mlp_units:list=[[512,
+> 512], [512, 512], [512, 512]], dropout_prob_theta=0.0,
+> activation='ReLU', shared_weights=False, loss=MAE(),
+> valid_loss=None, max_steps:int=1000, learning_rate:float=0.001,
+> num_lr_decays:int=3, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size=32,
+> valid_batch_size:Optional[int]=None,
+> windows_batch_size:int=1024,
+> inference_windows_batch_size:int=-1,
+> start_padding_enabled:bool=False, step_size:int=1,
+> scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*NBEATSx
+
+The Neural Basis Expansion Analysis with Exogenous variables (NBEATSx)
+is a simple and effective deep learning architecture. It is built with a
+deep stack of MLPs with doubly residual connections. The NBEATSx
+architecture includes additional exogenous blocks, extending NBEATS
+capabilities and interpretability. With its interpretable version,
+NBEATSx decomposes its predictions on seasonality, trend, and exogenous
+effects.
+
+**Parameters:** `h`: int, Forecast horizon. `input_size`: int,
+autorregresive inputs size, y=\[1,2,3,4\] input_size=2 -\>
+y\_\[t-2:t\]=\[1,2\]. `futr_exog_list`: str list, future exogenous
+columns. `hist_exog_list`: str list, historic exogenous columns.
+`stat_exog_list`: str list, static exogenous columns.
+`exclude_insample_y`: bool=False, the model skips the autoregressive
+features y\[t-input_size:t\] if True. `n_harmonics`: int, Number of
+harmonic oscillations in the SeasonalityBasis \[cos(i \* t/n_harmonics),
+sin(i \* t/n_harmonics)\]. Note that it will only be used if
+‘seasonality’ is in `stack_types`. `n_polynomials`: int, Number of
+polynomial terms for TrendBasis \[1,t,…,t^n_poly\]. Note that it will
+only be used if ‘trend’ is in `stack_types`. `stack_types`:
+List\[str\], List of stack types. Subset from \[‘seasonality’, ‘trend’,
+‘identity’, ‘exogenous’\]. `n_blocks`: List\[int\], Number of blocks
+for each stack. Note that len(n_blocks) = len(stack_types).
+`mlp_units`: List\[List\[int\]\], Structure of hidden layers for each
+stack type. Each internal list should contain the number of units of
+each hidden layer. Note that len(n_hidden) = len(stack_types).
+`dropout_prob_theta`: float, Float between (0, 1). Dropout for N-BEATS
+basis. `activation`: str, activation from \[‘ReLU’, ‘Softplus’,
+‘Tanh’, ‘SELU’, ‘LeakyReLU’, ‘PReLU’, ‘Sigmoid’\]. `loss`: PyTorch
+module, instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=3, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=1024, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=-1, number of windows to sample in
+each inference batch, -1 uses all. `start_padding_enabled`:
+bool=False, if True, the model will pad the time series with zeros at
+the beginning, by input size. `step_size`: int=1, step size between
+each window of temporal data. `scaler_type`: str=‘identity’, type of
+scaler for temporal inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int, random seed initialization for replicability.
+`drop_last_loader`: bool=False, if True `TimeSeriesDataLoader` drops
+last non-full batch. `alias`: str, optional, Custom name of the
+model. `optimizer`: Subclass of ‘torch.optim.Optimizer’, optional,
+user specified optimizer instead of the default choice (Adam).
+`optimizer_kwargs`: dict, optional, list of parameters used by the user
+specified `optimizer`. `lr_scheduler`: Subclass of
+‘torch.optim.lr_scheduler.LRScheduler’, optional, user specified
+lr_scheduler instead of the default choice (StepLR).
+`lr_scheduler_kwargs`: dict, optional, list of parameters used by the
+user specified `lr_scheduler`. `dataloader_kwargs`: dict, optional,
+list of parameters passed into the PyTorch Lightning dataloader by the
+`TimeSeriesDataLoader`. `**trainer_kwargs`: int, keyword trainer
+arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+**References:** -[Kin G. Olivares, Cristian Challu, Grzegorz
+Marcjasz, Rafał Weron, Artur Dubrawski (2021). “Neural basis expansion
+analysis with exogenous variables: Forecasting electricity prices with
+NBEATSx”.](https://arxiv.org/abs/2104.05522)\*
+
+------------------------------------------------------------------------
+
+### NBEATSx.fit
+
+> ``` text
+> NBEATSx.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### NBEATSx.predict
+
+> ``` text
+> NBEATSx.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import NBEATSx
+from neuralforecast.losses.pytorch import MQLoss
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = NBEATSx(h=12, input_size=24,
+ loss=MQLoss(level=[80, 90]),
+ scaler_type='robust',
+ dropout_prob_theta=0.5,
+ stat_exog_list=['airline1'],
+ futr_exog_list=['trend'],
+ stack_types = ["identity", "trend", "seasonality", "exogenous"],
+ n_blocks = [1,1,1,1],
+ max_steps=200,
+ val_check_steps=10,
+ early_stop_patience_steps=2)
+
+nf = NeuralForecast(
+ models=[model],
+ freq='ME'
+)
+nf.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+Y_hat_df = nf.predict(futr_df=Y_test_df)
+
+# Plot quantile predictions
+Y_hat_df = Y_hat_df.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['NBEATSx-median'], c='blue', label='median')
+plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['NBEATSx-lo-90'][-12:].values,
+ y2=plot_df['NBEATSx-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+plt.legend()
+plt.grid()
+plt.plot()
+```
+
diff --git a/neuralforecast/models.nhits.html.mdx b/neuralforecast/models.nhits.html.mdx
new file mode 100644
index 00000000..7b58e3c8
--- /dev/null
+++ b/neuralforecast/models.nhits.html.mdx
@@ -0,0 +1,270 @@
+---
+output-file: models.nhits.html
+title: NHITS
+---
+
+
+Long-horizon forecasting is challenging because of the *volatility* of
+the predictions and the *computational complexity*. To solve this
+problem we created the Neural Hierarchical Interpolation for Time Series
+(NHITS).
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+builds upon
+[`NBEATS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nbeats.html#nbeats)
+and specializes its partial outputs in the different frequencies of the
+time series through hierarchical interpolation and multi-rate input
+processing. On the long-horizon forecasting task
+[`NHITS`](https://nixtlaverse.nixtla.io/neuralforecast/models.nhits.html#nhits)
+improved accuracy by 25% on AAAI’s best paper award the
+[`Informer`](https://nixtlaverse.nixtla.io/neuralforecast/models.informer.html#informer),
+while being 50x faster.
+
+The model is composed of several MLPs with ReLU non-linearities. Blocks
+are connected via doubly residual stacking principle with the backcast
+$\mathbf{\tilde{y}}_{t-L:t,l}$ and forecast
+$\mathbf{\hat{y}}_{t+1:t+H,l}$ outputs of the l-th block. Multi-rate
+input pooling, hierarchical interpolation and backcast residual
+connections together induce the specialization of the additive
+predictions in different signal bands, reducing memory footprint and
+computational time, thus improving the architecture parsimony and
+accuracy.
+
+**References** -[Boris N. Oreshkin, Dmitri Carpov, Nicolas Chapados,
+Yoshua Bengio (2019). “N-BEATS: Neural basis expansion analysis for
+interpretable time series
+forecasting”.](https://arxiv.org/abs/1905.10437) -[Cristian Challu,
+Kin G. Olivares, Boris N. Oreshkin, Federico Garza, Max
+Mergenthaler-Canseco, Artur Dubrawski (2023). “NHITS: Neural
+Hierarchical Interpolation for Time Series Forecasting”. Accepted at the
+Thirty-Seventh AAAI Conference on Artificial
+Intelligence.](https://arxiv.org/abs/2201.12886) -[Zhou, H.; Zhang,
+S.; Peng, J.; Zhang, S.; Li, J.; Xiong, H.; and Zhang, W. (2020).
+“Informer: Beyond Efficient Transformer for Long Sequence Time-Series
+Forecasting”. Association for the Advancement of Artificial Intelligence
+Conference 2021 (AAAI 2021).](https://arxiv.org/abs/2012.07436)
+
+
+
+Figure 1. Neural Hierarchical
+Interpolation for Time Series (NHITS).
+
+
+------------------------------------------------------------------------
+
+source
+
+### NHITS
+
+> ``` text
+> NHITS (h, input_size, futr_exog_list=None, hist_exog_list=None,
+> stat_exog_list=None, exclude_insample_y=False,
+> stack_types:list=['identity', 'identity', 'identity'],
+> n_blocks:list=[1, 1, 1], mlp_units:list=[[512, 512], [512, 512],
+> [512, 512]], n_pool_kernel_size:list=[2, 2, 1],
+> n_freq_downsample:list=[4, 2, 1], pooling_mode:str='MaxPool1d',
+> interpolation_mode:str='linear', dropout_prob_theta=0.0,
+> activation='ReLU', loss=MAE(), valid_loss=None,
+> max_steps:int=1000, learning_rate:float=0.001,
+> num_lr_decays:int=3, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size:int=32,
+> valid_batch_size:Optional[int]=None, windows_batch_size:int=1024,
+> inference_windows_batch_size:int=-1, start_padding_enabled=False,
+> step_size:int=1, scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader=False, alias:Optional[str]=None, optimizer=None,
+> optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*NHITS
+
+The Neural Hierarchical Interpolation for Time Series (NHITS), is an
+MLP-based deep neural architecture with backward and forward residual
+links. NHITS tackles volatility and memory complexity challenges, by
+locally specializing its sequential predictions into the signals
+frequencies with hierarchical interpolation and pooling.
+
+**Parameters:** `h`: int, Forecast horizon. `input_size`: int,
+autorregresive inputs size, y=\[1,2,3,4\] input_size=2 -\>
+y\_\[t-2:t\]=\[1,2\]. `futr_exog_list`: str list, future exogenous
+columns. `hist_exog_list`: str list, historic exogenous columns.
+`stat_exog_list`: str list, static exogenous columns.
+`exclude_insample_y`: bool=False, the model skips the autoregressive
+features y\[t-input_size:t\] if True. `stack_types`: List\[str\],
+stacks list in the form N \* \[‘identity’\], to be deprecated in favor
+of `n_stacks`. Note that
+len(stack_types)=len(n_freq_downsample)=len(n_pool_kernel_size).
+`n_blocks`: List\[int\], Number of blocks for each stack. Note that
+len(n_blocks) = len(stack_types). `mlp_units`: List\[List\[int\]\],
+Structure of hidden layers for each stack type. Each internal list
+should contain the number of units of each hidden layer. Note that
+len(n_hidden) = len(stack_types). `n_pool_kernel_size`: List\[int\],
+list with the size of the windows to take a max/avg over. Note that
+len(stack_types)=len(n_freq_downsample)=len(n_pool_kernel_size).
+`n_freq_downsample`: List\[int\], list with the stack’s coefficients
+(inverse expressivity ratios). Note that
+len(stack_types)=len(n_freq_downsample)=len(n_pool_kernel_size).
+`pooling_mode`: str, input pooling module from \[‘MaxPool1d’,
+‘AvgPool1d’\]. `interpolation_mode`: str=‘linear’, interpolation
+basis from \[‘linear’, ‘nearest’, ‘cubic’\]. `dropout_prob_theta`:
+float, Float between (0, 1). Dropout for NHITS basis. `activation`:
+str, activation from \[‘ReLU’, ‘Softplus’, ‘Tanh’, ‘SELU’, ‘LeakyReLU’,
+‘PReLU’, ‘Sigmoid’\]. `loss`: PyTorch module, instantiated train
+loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=1024, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=-1, number of windows to sample in
+each inference batch, -1 uses all. `start_padding_enabled`:
+bool=False, if True, the model will pad the time series with zeros at
+the beginning, by input size. `step_size`: int=1, step size between
+each window of temporal data. `scaler_type`: str=‘identity’, type of
+scaler for temporal inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`.
+`dataloader_kwargs`: dict, optional, list of parameters passed into the
+PyTorch Lightning dataloader by the `TimeSeriesDataLoader`.
+`**trainer_kwargs`: int, keyword trainer arguments inherited from
+[PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+**References:** -[Cristian Challu, Kin G. Olivares, Boris N.
+Oreshkin, Federico Garza, Max Mergenthaler-Canseco, Artur Dubrawski
+(2023). “NHITS: Neural Hierarchical Interpolation for Time Series
+Forecasting”. Accepted at the Thirty-Seventh AAAI Conference on
+Artificial Intelligence.](https://arxiv.org/abs/2201.12886)\*
+
+------------------------------------------------------------------------
+
+### NHITS.fit
+
+> ``` text
+> NHITS.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### NHITS.predict
+
+> ``` text
+> NHITS.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import NHITS
+from neuralforecast.losses.pytorch import DistributionLoss
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = NHITS(h=12,
+ input_size=24,
+ loss=DistributionLoss(distribution='StudentT', level=[80, 90], return_params=True),
+ stat_exog_list=['airline1'],
+ futr_exog_list=['trend'],
+ n_freq_downsample=[2, 1, 1],
+ scaler_type='robust',
+ max_steps=200,
+ early_stop_patience_steps=2,
+ inference_windows_batch_size=1,
+ val_check_steps=10,
+ learning_rate=1e-3)
+
+fcst = NeuralForecast(models=[model], freq='ME')
+fcst.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+forecasts = fcst.predict(futr_df=Y_test_df)
+
+# Plot quantile predictions
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['NHITS-median'], c='blue', label='median')
+plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['NHITS-lo-90'][-12:].values,
+ y2=plot_df['NHITS-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+plt.legend()
+plt.grid()
+plt.plot()
+```
+
diff --git a/neuralforecast/models.nlinear.html.mdx b/neuralforecast/models.nlinear.html.mdx
new file mode 100644
index 00000000..f222f229
--- /dev/null
+++ b/neuralforecast/models.nlinear.html.mdx
@@ -0,0 +1,221 @@
+---
+output-file: models.nlinear.html
+title: NLinear
+---
+
+
+NLinear is a simple and fast yet accurate time series forecasting model
+for long-horizon forecasting.
+
+The architecture aims to boost the performance when there is a
+distribution shift in the dataset: 1. NLinear first subtracts the input
+by the last value of the sequence; 2. Then, the input goes through a
+linear layer, and the subtracted part is added back before making the
+final prediction.
+
+**References** - [Zeng, Ailing, et al. “Are transformers effective
+for time series forecasting?.” Proceedings of the AAAI conference on
+artificial intelligence. Vol. 37. No. 9.
+2023.”](https://ojs.aaai.org/index.php/AAAI/article/view/26317)
+
+
+
+Figure 1. DLinear
+Architecture.
+
+
+------------------------------------------------------------------------
+
+source
+
+### NLinear
+
+> ``` text
+> NLinear (h:int, input_size:int, stat_exog_list=None, hist_exog_list=None,
+> futr_exog_list=None, exclude_insample_y=False, loss=MAE(),
+> valid_loss=None, max_steps:int=5000, learning_rate:float=0.0001,
+> num_lr_decays:int=-1, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size:int=32,
+> valid_batch_size:Optional[int]=None, windows_batch_size=1024,
+> inference_windows_batch_size=1024, start_padding_enabled=False,
+> step_size:int=1, scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*NLinear
+
+*Parameters:* `h`: int, forecast horizon. `input_size`: int,
+maximum sequence length for truncated train backpropagation.
+`stat_exog_list`: str list, static exogenous columns.
+`hist_exog_list`: str list, historic exogenous columns.
+`futr_exog_list`: str list, future exogenous columns.
+`exclude_insample_y`: bool=False, the model skips the autoregressive
+features y\[t-input_size:t\] if True. `loss`: PyTorch module,
+instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=1024, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=1024, number of windows to sample in
+each inference batch. `start_padding_enabled`: bool=False, if True,
+the model will pad the time series with zeros at the beginning, by input
+size. `step_size`: int=1, step size between each window of temporal
+data.
+`scaler_type`: str=‘robust’, type of scaler for temporal inputs
+normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`.
+`dataloader_kwargs`: dict, optional, list of parameters passed into the
+PyTorch Lightning dataloader by the `TimeSeriesDataLoader`.
+`**trainer_kwargs`: int, keyword trainer arguments inherited from
+[PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+``` text
+*References*
+- Zeng, Ailing, et al. "Are transformers effective for time series forecasting?." Proceedings of the AAAI conference on artificial intelligence. Vol. 37. No. 9. 2023."*
+```
+
+------------------------------------------------------------------------
+
+### NLinear.fit
+
+> ``` text
+> NLinear.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### NLinear.predict
+
+> ``` text
+> NLinear.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import NLinear
+from neuralforecast.losses.pytorch import DistributionLoss
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic, augment_calendar_df
+
+AirPassengersPanel, calendar_cols = augment_calendar_df(df=AirPassengersPanel, freq='M')
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = NLinear(h=12,
+ input_size=24,
+ loss=DistributionLoss(distribution='StudentT', level=[80, 90], return_params=True),
+ scaler_type='robust',
+ learning_rate=1e-3,
+ max_steps=500,
+ val_check_steps=50,
+ early_stop_patience_steps=2)
+
+nf = NeuralForecast(
+ models=[model],
+ freq='ME'
+)
+nf.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+forecasts = nf.predict(futr_df=Y_test_df)
+
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+if model.loss.is_distribution_output:
+ plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+ plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+ plt.plot(plot_df['ds'], plot_df['NLinear-median'], c='blue', label='median')
+ plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['NLinear-lo-90'][-12:].values,
+ y2=plot_df['NLinear-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+ plt.grid()
+ plt.legend()
+ plt.plot()
+else:
+ plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+ plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+ plt.plot(plot_df['ds'], plot_df['NLinear'], c='blue', label='Forecast')
+ plt.legend()
+ plt.grid()
+```
+
diff --git a/neuralforecast/models.patchtst.html.mdx b/neuralforecast/models.patchtst.html.mdx
new file mode 100644
index 00000000..67e0a4ea
--- /dev/null
+++ b/neuralforecast/models.patchtst.html.mdx
@@ -0,0 +1,438 @@
+---
+output-file: models.patchtst.html
+title: PatchTST
+---
+
+
+The PatchTST model is an efficient Transformer-based model for
+multivariate time series forecasting.
+
+It is based on two key components: - segmentation of time series into
+windows (patches) which are served as input tokens to Transformer -
+channel-independence. where each channel contains a single univariate
+time series.
+
+**References** - [Nie, Y., Nguyen, N. H., Sinthong, P., &
+Kalagnanam, J. (2022). “A Time Series is Worth 64 Words: Long-term
+Forecasting with
+Transformers”](https://arxiv.org/pdf/2211.14730.pdf)
+
+
+
+Figure 1. PatchTST.
+
+
+## 1. Backbone
+
+### Auxiliary Functions
+
+------------------------------------------------------------------------
+
+source
+
+### get_activation_fn
+
+> ``` text
+> get_activation_fn (activation)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### Transpose
+
+> ``` text
+> Transpose (*dims, contiguous=False)
+> ```
+
+*Transpose*
+
+### Positional Encoding
+
+------------------------------------------------------------------------
+
+source
+
+### positional_encoding
+
+> ``` text
+> positional_encoding (pe, learn_pe, q_len, hidden_size)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### Coord1dPosEncoding
+
+> ``` text
+> Coord1dPosEncoding (q_len, exponential=False, normalize=True)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### Coord2dPosEncoding
+
+> ``` text
+> Coord2dPosEncoding (q_len, hidden_size, exponential=False,
+> normalize=True, eps=0.001)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### PositionalEncoding
+
+> ``` text
+> PositionalEncoding (q_len, hidden_size, normalize=True)
+> ```
+
+### Encoder
+
+------------------------------------------------------------------------
+
+source
+
+### TSTEncoderLayer
+
+> ``` text
+> TSTEncoderLayer (q_len, hidden_size, n_heads, d_k=None, d_v=None,
+> linear_hidden_size=256, store_attn=False,
+> norm='BatchNorm', attn_dropout=0, dropout=0.0,
+> bias=True, activation='gelu', res_attention=False,
+> pre_norm=False)
+> ```
+
+*TSTEncoderLayer*
+
+------------------------------------------------------------------------
+
+source
+
+### TSTEncoder
+
+> ``` text
+> TSTEncoder (q_len, hidden_size, n_heads, d_k=None, d_v=None,
+> linear_hidden_size=None, norm='BatchNorm', attn_dropout=0.0,
+> dropout=0.0, activation='gelu', res_attention=False,
+> n_layers=1, pre_norm=False, store_attn=False)
+> ```
+
+*TSTEncoder*
+
+------------------------------------------------------------------------
+
+source
+
+### TSTiEncoder
+
+> ``` text
+> TSTiEncoder (c_in, patch_num, patch_len, max_seq_len=1024, n_layers=3,
+> hidden_size=128, n_heads=16, d_k=None, d_v=None,
+> linear_hidden_size=256, norm='BatchNorm', attn_dropout=0.0,
+> dropout=0.0, act='gelu', store_attn=False,
+> key_padding_mask='auto', padding_var=None, attn_mask=None,
+> res_attention=True, pre_norm=False, pe='zeros',
+> learn_pe=True)
+> ```
+
+*TSTiEncoder*
+
+------------------------------------------------------------------------
+
+source
+
+### Flatten_Head
+
+> ``` text
+> Flatten_Head (individual, n_vars, nf, h, c_out, head_dropout=0)
+> ```
+
+*Flatten_Head*
+
+------------------------------------------------------------------------
+
+source
+
+### PatchTST_backbone
+
+> ``` text
+> PatchTST_backbone (c_in:int, c_out:int, input_size:int, h:int,
+> patch_len:int, stride:int,
+> max_seq_len:Optional[int]=1024, n_layers:int=3,
+> hidden_size=128, n_heads=16, d_k:Optional[int]=None,
+> d_v:Optional[int]=None, linear_hidden_size:int=256,
+> norm:str='BatchNorm', attn_dropout:float=0.0,
+> dropout:float=0.0, act:str='gelu',
+> key_padding_mask:str='auto',
+> padding_var:Optional[int]=None,
+> attn_mask:Optional[torch.Tensor]=None,
+> res_attention:bool=True, pre_norm:bool=False,
+> store_attn:bool=False, pe:str='zeros',
+> learn_pe:bool=True, fc_dropout:float=0.0,
+> head_dropout=0, padding_patch=None,
+> pretrain_head:bool=False, head_type='flatten',
+> individual=False, revin=True, affine=True,
+> subtract_last=False)
+> ```
+
+*PatchTST_backbone*
+
+## 2. Model
+
+------------------------------------------------------------------------
+
+source
+
+### PatchTST
+
+> ``` text
+> PatchTST (h, input_size, stat_exog_list=None, hist_exog_list=None,
+> futr_exog_list=None, exclude_insample_y=False,
+> encoder_layers:int=3, n_heads:int=16, hidden_size:int=128,
+> linear_hidden_size:int=256, dropout:float=0.2,
+> fc_dropout:float=0.2, head_dropout:float=0.0,
+> attn_dropout:float=0.0, patch_len:int=16, stride:int=8,
+> revin:bool=True, revin_affine:bool=False,
+> revin_subtract_last:bool=True, activation:str='gelu',
+> res_attention:bool=True, batch_normalization:bool=False,
+> learn_pos_embed:bool=True, loss=MAE(), valid_loss=None,
+> max_steps:int=5000, learning_rate:float=0.0001,
+> num_lr_decays:int=-1, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size:int=32,
+> valid_batch_size:Optional[int]=None, windows_batch_size=1024,
+> inference_windows_batch_size:int=1024,
+> start_padding_enabled=False, step_size:int=1,
+> scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*PatchTST
+
+The PatchTST model is an efficient Transformer-based model for
+multivariate time series forecasting.
+
+It is based on two key components: - segmentation of time series into
+windows (patches) which are served as input tokens to Transformer -
+channel-independence, where each channel contains a single univariate
+time series.
+
+**Parameters:** `h`: int, Forecast horizon. `input_size`: int,
+autorregresive inputs size, y=\[1,2,3,4\] input_size=2 -\>
+y\_\[t-2:t\]=\[1,2\]. `stat_exog_list`: str list, static exogenous
+columns. `hist_exog_list`: str list, historic exogenous columns.
+`futr_exog_list`: str list, future exogenous columns.
+`exclude_insample_y`: bool=False, the model skips the autoregressive
+features y\[t-input_size:t\] if True. `encoder_layers`: int, number
+of layers for encoder. `n_heads`: int=16, number of multi-head’s
+attention. `hidden_size`: int=128, units of embeddings and
+encoders. `linear_hidden_size`: int=256, units of linear layer.
+`dropout`: float=0.1, dropout rate for residual connection.
+`fc_dropout`: float=0.1, dropout rate for linear layer.
+`head_dropout`: float=0.1, dropout rate for Flatten head layer.
+`attn_dropout`: float=0.1, dropout rate for attention layer.
+`patch_len`: int=32, length of patch. Note: patch_len = min(patch_len,
+input_size + stride). `stride`: int=16, stride of patch.
+`revin`: bool=True, bool to use RevIn. `revin_affine`: bool=False,
+bool to use affine in RevIn. `revin_subtract_last`: bool=False, bool
+to use substract last in RevIn. `activation`: str=‘ReLU’, activation
+from \[‘gelu’,‘relu’\]. `res_attention`: bool=False, bool to use
+residual attention. `batch_normalization`: bool=False, bool to use
+batch normalization. `learn_pos_embed`: bool=True, bool to learn
+positional embedding. `loss`: PyTorch module, instantiated train
+loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=1024, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=1024, number of windows to sample in
+each inference batch. `start_padding_enabled`: bool=False, if True,
+the model will pad the time series with zeros at the beginning, by input
+size. `step_size`: int=1, step size between each window of temporal
+data. `scaler_type`: str=‘identity’, type of scaler for temporal
+inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`.
+`dataloader_kwargs`: dict, optional, list of parameters passed into the
+PyTorch Lightning dataloader by the `TimeSeriesDataLoader`.
+`**trainer_kwargs`: int, keyword trainer arguments inherited from
+[PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+**References:** -[Nie, Y., Nguyen, N. H., Sinthong, P., &
+Kalagnanam, J. (2022). “A Time Series is Worth 64 Words: Long-term
+Forecasting with Transformers”](https://arxiv.org/pdf/2211.14730.pdf)\*
+
+------------------------------------------------------------------------
+
+### PatchTST.fit
+
+> ``` text
+> PatchTST.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### PatchTST.predict
+
+> ``` text
+> PatchTST.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import PatchTST
+from neuralforecast.losses.pytorch import DistributionLoss
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic, augment_calendar_df
+
+AirPassengersPanel, calendar_cols = augment_calendar_df(df=AirPassengersPanel, freq='M')
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = PatchTST(h=12,
+ input_size=104,
+ patch_len=24,
+ stride=24,
+ revin=False,
+ hidden_size=16,
+ n_heads=4,
+ scaler_type='robust',
+ loss=DistributionLoss(distribution='StudentT', level=[80, 90]),
+ learning_rate=1e-3,
+ max_steps=500,
+ val_check_steps=50,
+ early_stop_patience_steps=2)
+
+nf = NeuralForecast(
+ models=[model],
+ freq='ME'
+)
+nf.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+forecasts = nf.predict(futr_df=Y_test_df)
+
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+if model.loss.is_distribution_output:
+ plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+ plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+ plt.plot(plot_df['ds'], plot_df['PatchTST-median'], c='blue', label='median')
+ plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['PatchTST-lo-90'][-12:].values,
+ y2=plot_df['PatchTST-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+ plt.grid()
+ plt.legend()
+ plt.plot()
+else:
+ plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+ plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+ plt.plot(plot_df['ds'], plot_df['PatchTST'], c='blue', label='Forecast')
+ plt.legend()
+ plt.grid()
+```
+
diff --git a/neuralforecast/models.rmok.html.mdx b/neuralforecast/models.rmok.html.mdx
new file mode 100644
index 00000000..c0a93079
--- /dev/null
+++ b/neuralforecast/models.rmok.html.mdx
@@ -0,0 +1,260 @@
+---
+output-file: models.rmok.html
+title: Reversible Mixture of KAN - RMoK
+---
+
+
+
+
+Figure 1. Architecture of
+RMoK.
+
+
+## 1. Auxiliary functions
+
+### 1.1 WaveKAN
+
+------------------------------------------------------------------------
+
+source
+
+### WaveKANLayer
+
+> ``` text
+> WaveKANLayer (in_features, out_features, wavelet_type='mexican_hat',
+> with_bn=True, device='cpu')
+> ```
+
+\*This is a sample code for the simulations of the paper: Bozorgasl,
+Zavareh and Chen, Hao, Wav-KAN: Wavelet Kolmogorov-Arnold Networks (May,
+2024)
+
+https://arxiv.org/abs/2405.12832 and also available at:
+https://papers.ssrn.com/sol3/papers.cfm?abstract_id=4835325 We used
+efficient KAN notation and some part of the code:+\*
+
+### 1.2 TaylorKAN
+
+------------------------------------------------------------------------
+
+source
+
+### TaylorKANLayer
+
+> ``` text
+> TaylorKANLayer (input_dim, out_dim, order, addbias=True)
+> ```
+
+*https://github.com/Muyuzhierchengse/TaylorKAN/*
+
+### 1.3. JacobiKAN
+
+------------------------------------------------------------------------
+
+source
+
+### JacobiKANLayer
+
+> ``` text
+> JacobiKANLayer (input_dim, output_dim, degree, a=1.0, b=1.0)
+> ```
+
+*https://github.com/SpaceLearner/JacobiKAN/blob/main/JacobiKANLayer.py*
+
+## 2. Model
+
+------------------------------------------------------------------------
+
+source
+
+### RMoK
+
+> ``` text
+> RMoK (h, input_size, n_series:int, futr_exog_list=None,
+> hist_exog_list=None, stat_exog_list=None, taylor_order:int=3,
+> jacobi_degree:int=6, wavelet_function:str='mexican_hat',
+> dropout:float=0.1, revin_affine:bool=True, loss=MAE(),
+> valid_loss=None, max_steps:int=1000, learning_rate:float=0.001,
+> num_lr_decays:int=-1, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size:int=32,
+> valid_batch_size:Optional[int]=None, windows_batch_size=32,
+> inference_windows_batch_size=32, start_padding_enabled=False,
+> step_size:int=1, scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None, **trainer_kwargs)
+> ```
+
+\*Reversible Mixture of KAN
+
+**Parameters:** `h`: int, Forecast horizon. `input_size`: int,
+autorregresive inputs size, y=\[1,2,3,4\] input_size=2 -\>
+y\_\[t-2:t\]=\[1,2\]. `n_series`: int, number of time-series.
+`futr_exog_list`: str list, future exogenous columns.
+`hist_exog_list`: str list, historic exogenous columns.
+`stat_exog_list`: str list, static exogenous columns.
+`taylor_order`: int, order of the Taylor polynomial.
+`jacobi_degree`: int, degree of the Jacobi polynomial.
+`wavelet_function`: str, wavelet function to use in the WaveKAN. Choose
+from \[“mexican_hat”, “morlet”, “dog”, “meyer”, “shannon”\]
+`dropout`: float, dropout rate. `revin_affine`: bool=False, bool to
+use affine in RevIn. `loss`: PyTorch module, instantiated train loss
+class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=32, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=32, number of windows to sample in
+each inference batch, -1 uses all. `start_padding_enabled`:
+bool=False, if True, the model will pad the time series with zeros at
+the beginning, by input size. `step_size`: int=1, step size between
+each window of temporal data. `scaler_type`: str=‘identity’, type of
+scaler for temporal inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`. `dataloader_kwargs`:
+dict, optional, list of parameters passed into the PyTorch Lightning
+dataloader by the `TimeSeriesDataLoader`. `**trainer_kwargs`: int,
+keyword trainer arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+**References** - [Xiao Han, Xinfeng Zhang, Yiling Wu, Zhenduo Zhang,
+Zhe Wu.”KAN4TSF: Are KAN and KAN-based models Effective for Time Series
+Forecasting?“. arXiv.](https://arxiv.org/abs/2408.11306) \*
+
+------------------------------------------------------------------------
+
+### RMoK.fit
+
+> ``` text
+> RMoK.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### RMoK.predict
+
+> ``` text
+> RMoK.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## 3. Usage example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import RMoK
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+from neuralforecast.losses.pytorch import MSE
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = RMoK(h=12,
+ input_size=24,
+ n_series=2,
+ taylor_order=3,
+ jacobi_degree=6,
+ wavelet_function='mexican_hat',
+ dropout=0.1,
+ revin_affine=True,
+ loss=MSE(),
+ valid_loss=MAE(),
+ early_stop_patience_steps=3,
+ batch_size=32)
+
+fcst = NeuralForecast(models=[model], freq='ME')
+fcst.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+forecasts = fcst.predict(futr_df=Y_test_df)
+
+# Plot predictions
+fig, ax = plt.subplots(1, 1, figsize = (20, 7))
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['RMoK'], c='blue', label='Forecast')
+ax.set_title('AirPassengers Forecast', fontsize=22)
+ax.set_ylabel('Monthly Passengers', fontsize=20)
+ax.set_xlabel('Year', fontsize=20)
+ax.legend(prop={'size': 15})
+ax.grid()
+```
+
diff --git a/neuralforecast/models.rnn.html.mdx b/neuralforecast/models.rnn.html.mdx
new file mode 100644
index 00000000..24c7efde
--- /dev/null
+++ b/neuralforecast/models.rnn.html.mdx
@@ -0,0 +1,241 @@
+---
+output-file: models.rnn.html
+title: RNN
+---
+
+
+Elman proposed this classic recurrent neural network
+([`RNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.rnn.html#rnn))
+in 1990, where each layer uses the following recurrent transformation:
+$$\mathbf{h}^{l}_{t} = \mathrm{Activation}([\mathbf{y}_{t},\mathbf{x}^{(h)}_{t},\mathbf{x}^{(s)}] W^{\intercal}_{ih} + b_{ih} + \mathbf{h}^{l}_{t-1} W^{\intercal}_{hh} + b_{hh})$$
+
+where $\mathbf{h}^{l}_{t}$, is the hidden state of RNN layer $l$ for
+time $t$, $\mathbf{y}_{t}$ is the input at time $t$ and
+$\mathbf{h}_{t-1}$ is the hidden state of the previous layer at $t-1$,
+$\mathbf{x}^{(s)}$ are static exogenous inputs, $\mathbf{x}^{(h)}_{t}$
+historic exogenous, $\mathbf{x}^{(f)}_{[:t+H]}$ are future exogenous
+available at the time of the prediction. The available activations are
+`tanh`, and `relu`. The predictions are obtained by transforming the
+hidden states into contexts $\mathbf{c}_{[t+1:t+H]}$, that are decoded
+and adapted into $\mathbf{\hat{y}}_{[t+1:t+H],[q]}$ through MLPs.
+
+**References** -[Jeffrey L. Elman (1990). “Finding Structure in
+Time”.](https://onlinelibrary.wiley.com/doi/10.1207/s15516709cog1402_1)
+-[Cho, K., van Merrienboer, B., Gülcehre, C., Bougares, F., Schwenk, H.,
+& Bengio, Y. (2014). Learning phrase representations using RNN
+encoder-decoder for statistical machine
+translation.](http://arxiv.org/abs/1406.1078)
+
+
+
+Figure 1. Single Layer Elman RNN with MLP
+decoder.
+
+
+------------------------------------------------------------------------
+
+source
+
+### RNN
+
+> ``` text
+> RNN (h:int, input_size:int=-1, inference_input_size:Optional[int]=None,
+> h_train:int=1, encoder_n_layers:int=2, encoder_hidden_size:int=128,
+> encoder_activation:str='tanh', encoder_bias:bool=True,
+> encoder_dropout:float=0.0, context_size:Optional[int]=None,
+> decoder_hidden_size:int=128, decoder_layers:int=2,
+> futr_exog_list=None, hist_exog_list=None, stat_exog_list=None,
+> exclude_insample_y=False, recurrent=False, loss=MAE(),
+> valid_loss=None, max_steps:int=1000, learning_rate:float=0.001,
+> num_lr_decays:int=-1, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size=32,
+> valid_batch_size:Optional[int]=None, windows_batch_size=128,
+> inference_windows_batch_size=1024, start_padding_enabled=False,
+> step_size:int=1, scaler_type:str='robust', random_seed=1,
+> drop_last_loader=False, alias:Optional[str]=None, optimizer=None,
+> optimizer_kwargs=None, lr_scheduler=None, lr_scheduler_kwargs=None,
+> dataloader_kwargs=None, **trainer_kwargs)
+> ```
+
+\*RNN
+
+Multi Layer Elman RNN (RNN), with MLP decoder. The network has `tanh` or
+`relu` non-linearities, it is trained using ADAM stochastic gradient
+descent. The network accepts static, historic and future exogenous data.
+
+**Parameters:** `h`: int, forecast horizon. `input_size`: int,
+maximum sequence length for truncated train backpropagation. Default -1
+uses 3 \* horizon `inference_input_size`: int, maximum sequence
+length for truncated inference. Default None uses input_size
+history. `h_train`: int, maximum sequence length for truncated train
+backpropagation. Default 1. `encoder_n_layers`: int=2, number of
+layers for the RNN. `encoder_hidden_size`: int=200, units for the
+RNN’s hidden state size. `encoder_activation`: str=`tanh`, type of
+RNN activation from `tanh` or `relu`. `encoder_bias`: bool=True,
+whether or not to use biases b_ih, b_hh within RNN units.
+`encoder_dropout`: float=0., dropout regularization applied to RNN
+outputs. `context_size`: deprecated. `decoder_hidden_size`:
+int=200, size of hidden layer for the MLP decoder. `decoder_layers`:
+int=2, number of layers for the MLP decoder. `futr_exog_list`: str
+list, future exogenous columns. `hist_exog_list`: str list, historic
+exogenous columns. `stat_exog_list`: str list, static exogenous
+columns. `exclude_insample_y`: bool=False, whether to exclude the
+target variable from the historic exogenous data. `recurrent`:
+bool=False, whether to produce forecasts recursively (True) or direct
+(False). `loss`: PyTorch module, instantiated train loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of
+differentseries in each batch. `valid_batch_size`: int=None, number
+of different series in each validation and test batch.
+`windows_batch_size`: int=128, number of windows to sample in each
+training batch, default uses all. `inference_windows_batch_size`:
+int=1024, number of windows to sample in each inference batch, -1 uses
+all. `start_padding_enabled`: bool=False, if True, the model will
+pad the time series with zeros at the beginning, by input size.
+`step_size`: int=1, step size between each window of temporal
+data.
+`scaler_type`: str=‘robust’, type of scaler for temporal inputs
+normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`.
+`dataloader_kwargs`: dict, optional, list of parameters passed into the
+PyTorch Lightning dataloader by the `TimeSeriesDataLoader`.
+
+`**trainer_kwargs`: int, keyword trainer arguments inherited from
+[PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer). \*
+
+------------------------------------------------------------------------
+
+### RNN.fit
+
+> ``` text
+> RNN.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### RNN.predict
+
+> ``` text
+> RNN.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import RNN
+from neuralforecast.losses.pytorch import MQLoss
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+fcst = NeuralForecast(
+ models=[RNN(h=12,
+ input_size=24,
+ inference_input_size=24,
+ loss=MQLoss(level=[80, 90]),
+ valid_loss=MQLoss(level=[80, 90]),
+ scaler_type='standard',
+ encoder_n_layers=2,
+ encoder_hidden_size=128,
+ decoder_hidden_size=128,
+ decoder_layers=2,
+ max_steps=200,
+ futr_exog_list=['y_[lag12]'],
+ stat_exog_list=['airline1'],
+ )
+ ],
+ freq='ME'
+)
+fcst.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+forecasts = fcst.predict(futr_df=Y_test_df)
+
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['RNN-median'], c='blue', label='median')
+plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['RNN-lo-90'][-12:].values,
+ y2=plot_df['RNN-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+plt.legend()
+plt.grid()
+plt.plot()
+```
+
diff --git a/neuralforecast/models.softs.html.mdx b/neuralforecast/models.softs.html.mdx
new file mode 100644
index 00000000..99553283
--- /dev/null
+++ b/neuralforecast/models.softs.html.mdx
@@ -0,0 +1,231 @@
+---
+output-file: models.softs.html
+title: SOFTS
+---
+
+
+## 1. Auxiliary functions
+
+### 1.1 Embedding
+
+------------------------------------------------------------------------
+
+source
+
+### DataEmbedding_inverted
+
+> ``` text
+> DataEmbedding_inverted (c_in, d_model, dropout=0.1)
+> ```
+
+*Data Embedding*
+
+### 1.2 STAD (STar Aggregate Dispatch)
+
+------------------------------------------------------------------------
+
+source
+
+### STAD
+
+> ``` text
+> STAD (d_series, d_core)
+> ```
+
+*STar Aggregate Dispatch Module*
+
+## 2. Model
+
+------------------------------------------------------------------------
+
+source
+
+### SOFTS
+
+> ``` text
+> SOFTS (h, input_size, n_series, futr_exog_list=None, hist_exog_list=None,
+> stat_exog_list=None, exclude_insample_y=False,
+> hidden_size:int=512, d_core:int=512, e_layers:int=2,
+> d_ff:int=2048, dropout:float=0.1, use_norm:bool=True, loss=MAE(),
+> valid_loss=None, max_steps:int=1000, learning_rate:float=0.001,
+> num_lr_decays:int=-1, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size:int=32,
+> valid_batch_size:Optional[int]=None, windows_batch_size=32,
+> inference_windows_batch_size=32, start_padding_enabled=False,
+> step_size:int=1, scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*SOFTS
+
+**Parameters:** `h`: int, Forecast horizon. `input_size`: int,
+autorregresive inputs size, y=\[1,2,3,4\] input_size=2 -\>
+y\_\[t-2:t\]=\[1,2\]. `n_series`: int, number of time-series.
+`futr_exog_list`: str list, future exogenous columns.
+`hist_exog_list`: str list, historic exogenous columns.
+`stat_exog_list`: str list, static exogenous columns.
+`exclude_insample_y`: bool=False, whether to exclude the target variable
+from the input.
+`hidden_size`: int, dimension of the model. `d_core`: int, dimension
+of core in STAD. `e_layers`: int, number of encoder layers.
+`d_ff`: int, dimension of fully-connected layer. `dropout`: float,
+dropout rate. `use_norm`: bool, whether to normalize or not.
+`loss`: PyTorch module, instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=32, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=32, number of windows to sample in
+each inference batch, -1 uses all. `start_padding_enabled`:
+bool=False, if True, the model will pad the time series with zeros at
+the beginning, by input size. `step_size`: int=1, step size between
+each window of temporal data. `scaler_type`: str=‘identity’, type of
+scaler for temporal inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`. `dataloader_kwargs`:
+dict, optional, list of parameters passed into the PyTorch Lightning
+dataloader by the `TimeSeriesDataLoader`. `**trainer_kwargs`: int,
+keyword trainer arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+**References** [Lu Han, Xu-Yang Chen, Han-Jia Ye, De-Chuan Zhan.
+“SOFTS: Efficient Multivariate Time Series Forecasting with Series-Core
+Fusion”](https://arxiv.org/pdf/2404.14197)\*
+
+------------------------------------------------------------------------
+
+### SOFTS.fit
+
+> ``` text
+> SOFTS.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### SOFTS.predict
+
+> ``` text
+> SOFTS.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## 3. Usage example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import SOFTS
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+from neuralforecast.losses.pytorch import MASE
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = SOFTS(h=12,
+ input_size=24,
+ n_series=2,
+ hidden_size=256,
+ d_core=256,
+ e_layers=2,
+ d_ff=64,
+ dropout=0.1,
+ use_norm=True,
+ loss=MASE(seasonality=4),
+ early_stop_patience_steps=3,
+ batch_size=32)
+
+fcst = NeuralForecast(models=[model], freq='ME')
+fcst.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+forecasts = fcst.predict(futr_df=Y_test_df)
+
+# Plot predictions
+fig, ax = plt.subplots(1, 1, figsize = (20, 7))
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['SOFTS'], c='blue', label='Forecast')
+ax.set_title('AirPassengers Forecast', fontsize=22)
+ax.set_ylabel('Monthly Passengers', fontsize=20)
+ax.set_xlabel('Year', fontsize=20)
+ax.legend(prop={'size': 15})
+ax.grid()
+```
+
diff --git a/neuralforecast/models.stemgnn.html.mdx b/neuralforecast/models.stemgnn.html.mdx
new file mode 100644
index 00000000..ca7e6bd0
--- /dev/null
+++ b/neuralforecast/models.stemgnn.html.mdx
@@ -0,0 +1,271 @@
+---
+output-file: models.stemgnn.html
+title: StemGNN
+---
+
+
+The Spectral Temporal Graph Neural Network
+([`StemGNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.stemgnn.html#stemgnn))
+is a Graph-based multivariate time-series forecasting model.
+[`StemGNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.stemgnn.html#stemgnn)
+jointly learns temporal dependencies and inter-series correlations in
+the spectral domain, by combining Graph Fourier Transform (GFT) and
+Discrete Fourier Transform (DFT).
+
+This method proved state-of-the-art performance on geo-temporal datasets
+such as `Solar`, `METR-LA`, and `PEMS-BAY`, and
+
+**References** -[Defu Cao, Yujing Wang, Juanyong Duan, Ce Zhang, Xia
+Zhu, Congrui Huang, Yunhai Tong, Bixiong Xu, Jing Bai, Jie Tong, Qi
+Zhang (2020). “Spectral Temporal Graph Neural Network for Multivariate
+Time-series
+Forecasting”.](https://proceedings.neurips.cc/paper/2020/hash/cdf6581cb7aca4b7e19ef136c6e601a5-Abstract.html)
+
+
+
+Figure 1. StemGNN.
+
+
+------------------------------------------------------------------------
+
+source
+
+### GLU
+
+> ``` text
+> GLU (input_channel, output_channel)
+> ```
+
+*GLU*
+
+------------------------------------------------------------------------
+
+source
+
+### StockBlockLayer
+
+> ``` text
+> StockBlockLayer (time_step, unit, multi_layer, stack_cnt=0)
+> ```
+
+*StockBlockLayer*
+
+------------------------------------------------------------------------
+
+source
+
+### StemGNN
+
+> ``` text
+> StemGNN (h, input_size, n_series, futr_exog_list=None,
+> hist_exog_list=None, stat_exog_list=None,
+> exclude_insample_y=False, n_stacks=2, multi_layer:int=5,
+> dropout_rate:float=0.5, leaky_rate:float=0.2, loss=MAE(),
+> valid_loss=None, max_steps:int=1000, learning_rate:float=0.001,
+> num_lr_decays:int=3, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size:int=32,
+> valid_batch_size:Optional[int]=None, windows_batch_size=32,
+> inference_windows_batch_size=32, start_padding_enabled=False,
+> step_size:int=1, scaler_type:str='robust', random_seed:int=1,
+> drop_last_loader=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*StemGNN
+
+The Spectral Temporal Graph Neural Network
+([`StemGNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.stemgnn.html#stemgnn))
+is a Graph-based multivariate time-series forecasting model.
+[`StemGNN`](https://nixtlaverse.nixtla.io/neuralforecast/models.stemgnn.html#stemgnn)
+jointly learns temporal dependencies and inter-series correlations in
+the spectral domain, by combining Graph Fourier Transform (GFT) and
+Discrete Fourier Transform (DFT).
+
+**Parameters:** `h`: int, Forecast horizon. `input_size`: int,
+autorregresive inputs size, y=\[1,2,3,4\] input_size=2 -\>
+y\_\[t-2:t\]=\[1,2\]. `n_series`: int, number of time-series.
+`futr_exog_list`: str list, future exogenous columns.
+`hist_exog_list`: str list, historic exogenous columns.
+`stat_exog_list`: str list, static exogenous columns. `n_stacks`:
+int=2, number of stacks in the model. `multi_layer`: int=5,
+multiplier for FC hidden size on StemGNN blocks. `dropout_rate`:
+float=0.5, dropout rate. `leaky_rate`: float=0.2, alpha for
+LeakyReLU layer on Latent Correlation layer. `loss`: PyTorch module,
+instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int, number of windows in each
+batch. `valid_batch_size`: int=None, number of different series in
+each validation and test batch, if None uses batch_size.
+`windows_batch_size`: int=32, number of windows to sample in each
+training batch, default uses all. `inference_windows_batch_size`:
+int=32, number of windows to sample in each inference batch, -1 uses
+all. `start_padding_enabled`: bool=False, if True, the model will
+pad the time series with zeros at the beginning, by input size.
+`step_size`: int=1, step size between each window of temporal data.
+`scaler_type`: str=‘robust’, type of scaler for temporal inputs
+normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`. `dataloader_kwargs`:
+dict, optional, list of parameters passed into the PyTorch Lightning
+dataloader by the `TimeSeriesDataLoader`. `**trainer_kwargs`: int,
+keyword trainer arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer). \*
+
+------------------------------------------------------------------------
+
+### StemGNN.fit
+
+> ``` text
+> StemGNN.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### StemGNN.predict
+
+> ``` text
+> StemGNN.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage Examples
+
+Train model and forecast future values with `predict` method.
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import StemGNN
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+from neuralforecast.losses.pytorch import MAE
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = StemGNN(h=12,
+ input_size=24,
+ n_series=2,
+ scaler_type='standard',
+ max_steps=500,
+ early_stop_patience_steps=-1,
+ val_check_steps=10,
+ learning_rate=1e-3,
+ loss=MAE(),
+ valid_loss=MAE(),
+ batch_size=32
+ )
+
+fcst = NeuralForecast(models=[model], freq='ME')
+fcst.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+forecasts = fcst.predict(futr_df=Y_test_df)
+
+# Plot predictions
+fig, ax = plt.subplots(1, 1, figsize = (20, 7))
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['StemGNN'], c='blue', label='Forecast')
+ax.set_title('AirPassengers Forecast', fontsize=22)
+ax.set_ylabel('Monthly Passengers', fontsize=20)
+ax.set_xlabel('Year', fontsize=20)
+ax.legend(prop={'size': 15})
+ax.grid()
+```
+
+Using `cross_validation` to forecast multiple historic values.
+
+
+```python
+fcst = NeuralForecast(models=[model], freq='M')
+forecasts = fcst.cross_validation(df=AirPassengersPanel, static_df=AirPassengersStatic, n_windows=2, step_size=12)
+
+# Plot predictions
+fig, ax = plt.subplots(1, 1, figsize = (20, 7))
+Y_hat_df = forecasts.loc['Airline1']
+Y_df = AirPassengersPanel[AirPassengersPanel['unique_id']=='Airline1']
+
+plt.plot(Y_df['ds'], Y_df['y'], c='black', label='True')
+plt.plot(Y_hat_df['ds'], Y_hat_df['StemGNN'], c='blue', label='Forecast')
+ax.set_title('AirPassengers Forecast', fontsize=22)
+ax.set_ylabel('Monthly Passengers', fontsize=20)
+ax.set_xlabel('Year', fontsize=20)
+ax.legend(prop={'size': 15})
+ax.grid()
+```
+
diff --git a/neuralforecast/models.tcn.html.mdx b/neuralforecast/models.tcn.html.mdx
new file mode 100644
index 00000000..7021c3c7
--- /dev/null
+++ b/neuralforecast/models.tcn.html.mdx
@@ -0,0 +1,248 @@
+---
+output-file: models.tcn.html
+title: TCN
+---
+
+
+For long time in deep learning, sequence modelling was synonymous with
+recurrent networks, yet several papers have shown that simple
+convolutional architectures can outperform canonical recurrent networks
+like LSTMs by demonstrating longer effective memory. By skipping
+temporal connections the causal convolution filters can be applied to
+larger time spans while remaining computationally efficient.
+
+The predictions are obtained by transforming the hidden states into
+contexts $\mathbf{c}_{[t+1:t+H]}$, that are decoded and adapted into
+$\mathbf{\hat{y}}_{[t+1:t+H],[q]}$ through MLPs.
+
+where $\mathbf{h}_{t}$, is the hidden state for time $t$,
+$\mathbf{y}_{t}$ is the input at time $t$ and $\mathbf{h}_{t-1}$ is the
+hidden state of the previous layer at $t-1$, $\mathbf{x}^{(s)}$ are
+static exogenous inputs, $\mathbf{x}^{(h)}_{t}$ historic exogenous,
+$\mathbf{x}^{(f)}_{[:t+H]}$ are future exogenous available at the time
+of the prediction.
+
+**References** -[van den Oord, A., Dieleman, S., Zen, H., Simonyan,
+K., Vinyals, O., Graves, A., Kalchbrenner, N., Senior, A. W., &
+Kavukcuoglu, K. (2016). Wavenet: A generative model for raw audio.
+Computing Research Repository, abs/1609.03499. URL:
+http://arxiv.org/abs/1609.03499.
+arXiv:1609.03499.](https://arxiv.org/abs/1609.03499) -[Shaojie Bai,
+Zico Kolter, Vladlen Koltun. (2018). An Empirical Evaluation of Generic
+Convolutional and Recurrent Networks for Sequence Modeling. Computing
+Research Repository, abs/1803.01271. URL:
+https://arxiv.org/abs/1803.01271.](https://arxiv.org/abs/1803.01271)
+
+
+
+Figure 1. Visualization of a stack of
+dilated causal convolutional layers.
+
+
+------------------------------------------------------------------------
+
+source
+
+### TCN
+
+> ``` text
+> TCN (h:int, input_size:int=-1, inference_input_size:Optional[int]=None,
+> kernel_size:int=2, dilations:List[int]=[1, 2, 4, 8, 16],
+> encoder_hidden_size:int=128, encoder_activation:str='ReLU',
+> context_size:int=10, decoder_hidden_size:int=128,
+> decoder_layers:int=2, futr_exog_list=None, hist_exog_list=None,
+> stat_exog_list=None, loss=MAE(), valid_loss=None,
+> max_steps:int=1000, learning_rate:float=0.001, num_lr_decays:int=-1,
+> early_stop_patience_steps:int=-1, val_check_steps:int=100,
+> batch_size:int=32, valid_batch_size:Optional[int]=None,
+> windows_batch_size=128, inference_windows_batch_size=1024,
+> start_padding_enabled=False, step_size:int=1,
+> scaler_type:str='robust', random_seed:int=1, drop_last_loader=False,
+> alias:Optional[str]=None, optimizer=None, optimizer_kwargs=None,
+> lr_scheduler=None, lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*TCN
+
+Temporal Convolution Network (TCN), with MLP decoder. The historical
+encoder uses dilated skip connections to obtain efficient long memory,
+while the rest of the architecture allows for future exogenous
+alignment.
+
+**Parameters:** `h`: int, forecast horizon. `input_size`: int,
+maximum sequence length for truncated train backpropagation. Default -1
+uses 3 \* horizon `inference_input_size`: int, maximum sequence
+length for truncated inference. Default None uses input_size
+history. `kernel_size`: int, size of the convolving kernel.
+`dilations`: int list, ontrols the temporal spacing between the kernel
+points; also known as the à trous algorithm. `encoder_hidden_size`:
+int=200, units for the TCN’s hidden state size.
+`encoder_activation`: str=`tanh`, type of TCN activation from `tanh` or
+`relu`. `context_size`: int=10, size of context vector for each
+timestamp on the forecasting window. `decoder_hidden_size`: int=200,
+size of hidden layer for the MLP decoder. `decoder_layers`: int=2,
+number of layers for the MLP decoder. `futr_exog_list`: str list,
+future exogenous columns. `hist_exog_list`: str list, historic
+exogenous columns. `stat_exog_list`: str list, static exogenous
+columns. `loss`: PyTorch module, instantiated train loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of
+differentseries in each batch. `batch_size`: int=32, number of
+differentseries in each batch. `valid_batch_size`: int=None, number
+of different series in each validation and test batch.
+`windows_batch_size`: int=128, number of windows to sample in each
+training batch, default uses all. `inference_windows_batch_size`:
+int=1024, number of windows to sample in each inference batch, -1 uses
+all. `start_padding_enabled`: bool=False, if True, the model will
+pad the time series with zeros at the beginning, by input size.
+`step_size`: int=1, step size between each window of temporal
+data.
+`scaler_type`: str=‘robust’, type of scaler for temporal inputs
+normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`.
+`dataloader_kwargs`: dict, optional, list of parameters passed into the
+PyTorch Lightning dataloader by the `TimeSeriesDataLoader`.
+`**trainer_kwargs`: int, keyword trainer arguments inherited from
+[PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer). \*
+
+------------------------------------------------------------------------
+
+### TCN.fit
+
+> ``` text
+> TCN.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### TCN.predict
+
+> ``` text
+> TCN.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import TCN
+from neuralforecast.losses.pytorch import DistributionLoss
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+fcst = NeuralForecast(
+ models=[TCN(h=12,
+ input_size=-1,
+ loss=DistributionLoss(distribution='Normal', level=[80, 90]),
+ learning_rate=5e-4,
+ kernel_size=2,
+ dilations=[1,2,4,8,16],
+ encoder_hidden_size=128,
+ context_size=10,
+ decoder_hidden_size=128,
+ decoder_layers=2,
+ max_steps=500,
+ scaler_type='robust',
+ futr_exog_list=['y_[lag12]'],
+ hist_exog_list=None,
+ stat_exog_list=['airline1'],
+ )
+ ],
+ freq='ME'
+)
+fcst.fit(df=Y_train_df, static_df=AirPassengersStatic)
+forecasts = fcst.predict(futr_df=Y_test_df)
+
+# Plot quantile predictions
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['TCN-median'], c='blue', label='median')
+plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['TCN-lo-90'][-12:].values,
+ y2=plot_df['TCN-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+plt.legend()
+plt.grid()
+plt.plot()
+```
+
diff --git a/neuralforecast/models.tft.html.mdx b/neuralforecast/models.tft.html.mdx
new file mode 100644
index 00000000..33a2b9cd
--- /dev/null
+++ b/neuralforecast/models.tft.html.mdx
@@ -0,0 +1,696 @@
+---
+output-file: models.tft.html
+title: TFT
+---
+
+
+In summary Temporal Fusion Transformer (TFT) combines gating layers, an
+LSTM recurrent encoder, with multi-head attention layers for a
+multi-step forecasting strategy decoder. TFT’s inputs are static
+exogenous $\mathbf{x}^{(s)}$, historic exogenous
+$\mathbf{x}^{(h)}_{[:t]}$, exogenous available at the time of the
+prediction $\mathbf{x}^{(f)}_{[:t+H]}$ and autorregresive features
+$\mathbf{y}_{[:t]}$, each of these inputs is further decomposed into
+categorical and continuous. The network uses a multi-quantile regression
+to model the following conditional
+probability:$$\mathbb{P}(\mathbf{y}_{[t+1:t+H]}|\;\mathbf{y}_{[:t]},\; \mathbf{x}^{(h)}_{[:t]},\; \mathbf{x}^{(f)}_{[:t+H]},\; \mathbf{x}^{(s)})$$
+
+**References** - [Jan Golda, Krzysztof Kudrynski. “NVIDIA, Deep
+Learning Forecasting
+Examples”](https://github.com/NVIDIA/DeepLearningExamples/tree/master/PyTorch/Forecasting/TFT) -
+[Bryan Lim, Sercan O. Arik, Nicolas Loeff, Tomas Pfister, “Temporal
+Fusion Transformers for interpretable multi-horizon time series
+forecasting”](https://www.sciencedirect.com/science/article/pii/S0169207021000637)
+
+
+
+Figure 1. Temporal Fusion Transformer
+Architecture.
+
+
+## 1. Auxiliary Functions
+
+### 1.1 Gating Mechanisms
+
+The Gated Residual Network (GRN) provides adaptive depth and network
+complexity capable of accommodating different size datasets. As residual
+connections allow for the network to skip the non-linear transformation
+of input $\mathbf{a}$ and context $\mathbf{c}$.
+
+The Gated Linear Unit (GLU) provides the flexibility of supressing
+unnecesary parts of the GRN. Consider GRN’s output $\gamma$ then GLU
+transformation is defined by:
+
+$$\mathrm{GLU}(\gamma) = \sigma(\mathbf{W}_{4}\gamma +b_{4}) \odot (\mathbf{W}_{5}\gamma +b_{5})$$
+
+
+
+Figure 2. Gated Residual
+Network.
+
+
+### 1.2 Variable Selection Networks
+
+TFT includes automated variable selection capabilities, through its
+variable selection network (VSN) components. The VSN takes the original
+input
+$\{\mathbf{x}^{(s)}, \mathbf{x}^{(h)}_{[:t]}, \mathbf{x}^{(f)}_{[:t]}\}$
+and transforms it through embeddings or linear transformations into a
+high dimensional space
+$\{\mathbf{E}^{(s)}, \mathbf{E}^{(h)}_{[:t]}, \mathbf{E}^{(f)}_{[:t+H]}\}$.
+
+For the observed historic data, the embedding matrix
+$\mathbf{E}^{(h)}_{t}$ at time $t$ is a concatenation of $j$ variable
+$e^{(h)}_{t,j}$ embeddings:
+
+The variable selection weights are given by:
+$$s^{(h)}_{t}=\mathrm{SoftMax}(\mathrm{GRN}(\mathbf{E}^{(h)}_{t},\mathbf{E}^{(s)}))$$
+
+The VSN processed features are then:
+$$\tilde{\mathbf{E}}^{(h)}_{t}= \sum_{j} s^{(h)}_{j} \tilde{e}^{(h)}_{t,j}$$
+
+
+
+Figure 3. Variable Selection
+Network.
+
+
+### 1.3. Multi-Head Attention
+
+To avoid information bottlenecks from the classic Seq2Seq architecture,
+TFT incorporates a decoder-encoder attention mechanism inherited
+transformer architectures ([Li et. al
+2019](https://arxiv.org/abs/1907.00235), [Vaswani et. al
+2017](https://arxiv.org/abs/1706.03762)). It transform the the outputs
+of the LSTM encoded temporal features, and helps the decoder better
+capture long-term relationships.
+
+The original multihead attention for each component $H_{m}$ and its
+query, key, and value representations are denoted by
+$Q_{m}, K_{m}, V_{m}$, its transformation is given by:
+
+TFT modifies the original multihead attention to improve its
+interpretability. To do it it uses shared values $\tilde{V}$ across
+heads and employs additive aggregation,
+$\mathrm{InterpretableMultiHead}(Q,K,V) = \tilde{H} W_{M}$. The
+mechanism has a great resemblence to a single attention layer, but it
+allows for $M$ multiple attention weights, and can be therefore be
+interpreted as the average ensemble of $M$ single attention layers.
+
+## 2. TFT Architecture
+
+The first TFT’s step is embed the original input
+$\{\mathbf{x}^{(s)}, \mathbf{x}^{(h)}, \mathbf{x}^{(f)}\}$ into a high
+dimensional space
+$\{\mathbf{E}^{(s)}, \mathbf{E}^{(h)}, \mathbf{E}^{(f)}\}$, after which
+each embedding is gated by a variable selection network (VSN). The
+static embedding $\mathbf{E}^{(s)}$ is used as context for variable
+selection and as initial condition to the LSTM. Finally the encoded
+variables are fed into the multi-head attention decoder.
+
+### 2.1 Static Covariate Encoder
+
+The static embedding $\mathbf{E}^{(s)}$ is transformed by the
+StaticCovariateEncoder into contexts $c_{s}, c_{e}, c_{h}, c_{c}$. Where
+$c_{s}$ are temporal variable selection contexts, $c_{e}$ are
+TemporalFusionDecoder enriching contexts, and $c_{h}, c_{c}$ are LSTM’s
+hidden/contexts for the TemporalCovariateEncoder.
+
+### 2.2 Temporal Covariate Encoder
+
+TemporalCovariateEncoder encodes the embeddings
+$\mathbf{E}^{(h)}, \mathbf{E}^{(f)}$ and contexts $(c_{h}, c_{c})$ with
+an LSTM.
+
+An analogous process is repeated for the future data, with the main
+difference that $\mathbf{E}^{(f)}$ contains the future available
+information.
+
+### 2.3 Temporal Fusion Decoder
+
+The TemporalFusionDecoder enriches the LSTM’s outputs with $c_{e}$ and
+then uses an attention layer, and multi-step adapter.
+
+------------------------------------------------------------------------
+
+source
+
+### TFT
+
+> ``` text
+> TFT (h, input_size, tgt_size:int=1, stat_exog_list=None,
+> hist_exog_list=None, futr_exog_list=None, hidden_size:int=128,
+> n_head:int=4, attn_dropout:float=0.0, grn_activation:str='ELU',
+> n_rnn_layers:int=1, rnn_type:str='lstm',
+> one_rnn_initial_state:bool=False, dropout:float=0.1, loss=MAE(),
+> valid_loss=None, max_steps:int=1000, learning_rate:float=0.001,
+> num_lr_decays:int=-1, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size:int=32,
+> valid_batch_size:Optional[int]=None, windows_batch_size:int=1024,
+> inference_windows_batch_size:int=1024, start_padding_enabled=False,
+> step_size:int=1, scaler_type:str='robust', random_seed:int=1,
+> drop_last_loader=False, alias:Optional[str]=None, optimizer=None,
+> optimizer_kwargs=None, lr_scheduler=None, lr_scheduler_kwargs=None,
+> dataloader_kwargs=None, **trainer_kwargs)
+> ```
+
+\*TFT
+
+The Temporal Fusion Transformer architecture (TFT) is an
+Sequence-to-Sequence model that combines static, historic and future
+available data to predict an univariate target. The method combines
+gating layers, an LSTM recurrent encoder, with and interpretable
+multi-head attention layer and a multi-step forecasting strategy
+decoder.
+
+**Parameters:** `h`: int, Forecast horizon. `input_size`: int,
+autorregresive inputs size, y=\[1,2,3,4\] input_size=2 -\>
+y\_\[t-2:t\]=\[1,2\]. `tgt_size`: int=1, target size.
+`stat_exog_list`: str list, static continuous columns.
+`hist_exog_list`: str list, historic continuous columns.
+`futr_exog_list`: str list, future continuous columns.
+`hidden_size`: int, units of embeddings and encoders. `n_head`:
+int=4, number of attention heads in temporal fusion decoder.
+`attn_dropout`: float (0, 1), dropout of fusion decoder’s attention
+layer. `grn_activation`: str, activation for the GRN module from
+\[‘ReLU’, ‘Softplus’, ‘Tanh’, ‘SELU’, ‘LeakyReLU’, ‘Sigmoid’, ‘ELU’,
+‘GLU’\]. `n_rnn_layers`: int=1, number of RNN layers.
+`rnn_type`: str=“lstm”, recurrent neural network (RNN) layer type from
+\[“lstm”,“gru”\]. `one_rnn_initial_state`:str=False, Initialize all
+rnn layers with the same initial states computed from static
+covariates. `dropout`: float (0, 1), dropout of inputs VSNs.
+`loss`: PyTorch module, instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int, number of different series
+in each batch. `valid_batch_size`: int=None, number of different
+series in each validation and test batch. `windows_batch_size`:
+int=None, windows sampled from rolled data, default uses all.
+`inference_windows_batch_size`: int=-1, number of windows to sample in
+each inference batch, -1 uses all. `start_padding_enabled`:
+bool=False, if True, the model will pad the time series with zeros at
+the beginning, by input size. `step_size`: int=1, step size between
+each window of temporal data. `scaler_type`: str=‘robust’, type of
+scaler for temporal inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int, random seed initialization for replicability.
+`drop_last_loader`: bool=False, if True `TimeSeriesDataLoader` drops
+last non-full batch. `alias`: str, optional, Custom name of the
+model. `optimizer`: Subclass of ‘torch.optim.Optimizer’, optional,
+user specified optimizer instead of the default choice (Adam).
+`optimizer_kwargs`: dict, optional, list of parameters used by the user
+specified `optimizer`. `lr_scheduler`: Subclass of
+‘torch.optim.lr_scheduler.LRScheduler’, optional, user specified
+lr_scheduler instead of the default choice (StepLR).
+`lr_scheduler_kwargs`: dict, optional, list of parameters used by the
+user specified `lr_scheduler`. `dataloader_kwargs`: dict, optional,
+list of parameters passed into the PyTorch Lightning dataloader by the
+`TimeSeriesDataLoader`. `**trainer_kwargs`: int, keyword trainer
+arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+**References:** - [Bryan Lim, Sercan O. Arik, Nicolas Loeff, Tomas
+Pfister, “Temporal Fusion Transformers for interpretable multi-horizon
+time series
+forecasting”](https://www.sciencedirect.com/science/article/pii/S0169207021000637)\*
+
+## 3. TFT methods
+
+------------------------------------------------------------------------
+
+### TFT.fit
+
+> ``` text
+> TFT.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### TFT.predict
+
+> ``` text
+> TFT.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+------------------------------------------------------------------------
+
+source
+
+### TFT.feature_importances,
+
+> ``` text
+> TFT.feature_importances, ()
+> ```
+
+\*Compute the feature importances for historical, future, and static
+features.
+
+Returns: dict: A dictionary containing the feature importances for each
+feature type. The keys are ‘hist_vsn’, ‘future_vsn’, and ‘static_vsn’,
+and the values are pandas DataFrames with the corresponding feature
+importances.\*
+
+------------------------------------------------------------------------
+
+source
+
+### TFT.attention_weights
+
+> ``` text
+> TFT.attention_weights ()
+> ```
+
+\*Batch average attention weights
+
+Returns: np.ndarray: A 1D array containing the attention weights for
+each time step.\*
+
+------------------------------------------------------------------------
+
+source
+
+### TFT.attention_weights
+
+> ``` text
+> TFT.attention_weights ()
+> ```
+
+\*Batch average attention weights
+
+Returns: np.ndarray: A 1D array containing the attention weights for
+each time step.\*
+
+------------------------------------------------------------------------
+
+source
+
+### TFT.feature_importance_correlations
+
+> ``` text
+> TFT.feature_importance_correlations ()
+> ```
+
+\*Compute the correlation between the past and future feature
+importances and the mean attention weights.
+
+Returns: pd.DataFrame: A DataFrame containing the correlation
+coefficients between the past feature importances and the mean attention
+weights.\*
+
+## Usage Example
+
+
+```python
+import matplotlib.pyplot as plt
+import numpy as np
+import pandas as pd
+
+from neuralforecast import NeuralForecast
+
+# from neuralforecast.models import TFT
+from neuralforecast.losses.pytorch import DistributionLoss
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+
+AirPassengersPanel["month"] = AirPassengersPanel.ds.dt.month
+Y_train_df = AirPassengersPanel[
+ AirPassengersPanel.ds < AirPassengersPanel["ds"].values[-12]
+] # 132 train
+Y_test_df = AirPassengersPanel[
+ AirPassengersPanel.ds >= AirPassengersPanel["ds"].values[-12]
+].reset_index(drop=True) # 12 test
+
+nf = NeuralForecast(
+ models=[
+ TFT(
+ h=12,
+ input_size=48,
+ hidden_size=20,
+ grn_activation="ELU",
+ rnn_type="lstm",
+ n_rnn_layers=1,
+ one_rnn_initial_state=False,
+ loss=DistributionLoss(distribution="StudentT", level=[80, 90]),
+ learning_rate=0.005,
+ stat_exog_list=["airline1"],
+ futr_exog_list=["y_[lag12]", "month"],
+ hist_exog_list=["trend"],
+ max_steps=300,
+ val_check_steps=10,
+ early_stop_patience_steps=10,
+ scaler_type="robust",
+ windows_batch_size=None,
+ enable_progress_bar=True,
+ ),
+ ],
+ freq="ME",
+)
+nf.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+Y_hat_df = nf.predict(futr_df=Y_test_df)
+
+# Plot quantile predictions
+Y_hat_df = Y_hat_df.reset_index(drop=False).drop(columns=["unique_id", "ds"])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id == "Airline1"].drop("unique_id", axis=1)
+plt.plot(plot_df["ds"], plot_df["y"], c="black", label="True")
+plt.plot(plot_df["ds"], plot_df["TFT"], c="purple", label="mean")
+plt.plot(plot_df["ds"], plot_df["TFT-median"], c="blue", label="median")
+plt.fill_between(
+ x=plot_df["ds"][-12:],
+ y1=plot_df["TFT-lo-90"][-12:].values,
+ y2=plot_df["TFT-hi-90"][-12:].values,
+ alpha=0.4,
+ label="level 90",
+)
+plt.legend()
+plt.grid()
+plt.plot()
+```
+
+# Interpretability
+
+## 1. Attention Weights
+
+
+```python
+attention = nf.models[0].attention_weights()
+```
+
+
+```python
+def plot_attention(
+ self, plot: str = "time", output: str = "plot", width: int = 800, height: int = 400
+):
+ """
+ Plot the attention weights.
+
+ Args:
+ plot (str, optional): The type of plot to generate. Can be one of the following:
+ - 'time': Display the mean attention weights over time.
+ - 'all': Display the attention weights for each horizon.
+ - 'heatmap': Display the attention weights as a heatmap.
+ - An integer in the range [1, model.h) to display the attention weights for a specific horizon.
+ output (str, optional): The type of output to generate. Can be one of the following:
+ - 'plot': Display the plot directly.
+ - 'figure': Return the plot as a figure object.
+ width (int, optional): Width of the plot in pixels. Default is 800.
+ height (int, optional): Height of the plot in pixels. Default is 400.
+
+ Returns:
+ matplotlib.figure.Figure: If `output` is 'figure', the function returns the plot as a figure object.
+ """
+
+ attention = (
+ self.mean_on_batch(self.interpretability_params["attn_wts"])
+ .mean(dim=0)
+ .cpu()
+ .numpy()
+ )
+
+ fig, ax = plt.subplots(figsize=(width / 100, height / 100))
+
+ if plot == "time":
+ attention = attention[self.input_size :, :].mean(axis=0)
+ ax.plot(np.arange(-self.input_size, self.h), attention)
+ ax.axvline(
+ x=0, color="black", linewidth=3, linestyle="--", label="prediction start"
+ )
+ ax.set_title("Mean Attention")
+ ax.set_xlabel("time")
+ ax.set_ylabel("Attention")
+ ax.legend()
+
+ elif plot == "all":
+ for i in range(self.input_size, attention.shape[0]):
+ ax.plot(
+ np.arange(-self.input_size, self.h),
+ attention[i, :],
+ label=f"horizon {i-self.input_size+1}",
+ )
+ ax.axvline(
+ x=0, color="black", linewidth=3, linestyle="--", label="prediction start"
+ )
+ ax.set_title("Attention per horizon")
+ ax.set_xlabel("time")
+ ax.set_ylabel("Attention")
+ ax.legend()
+
+ elif plot == "heatmap":
+ cax = ax.imshow(
+ attention,
+ aspect="auto",
+ cmap="viridis",
+ extent=[-self.input_size, self.h, -self.input_size, self.h],
+ )
+ fig.colorbar(cax)
+ ax.set_title("Attention Heatmap")
+ ax.set_xlabel("Attention (current time step)")
+ ax.set_ylabel("Attention (previous time step)")
+
+ elif isinstance(plot, int) and (plot in np.arange(1, self.h + 1)):
+ i = self.input_size + plot - 1
+ ax.plot(
+ np.arange(-self.input_size, self.h),
+ attention[i, :],
+ label=f"horizon {plot}",
+ )
+ ax.axvline(
+ x=0, color="black", linewidth=3, linestyle="--", label="prediction start"
+ )
+ ax.set_title(f"Attention weight for horizon {plot}")
+ ax.set_xlabel("time")
+ ax.set_ylabel("Attention")
+ ax.legend()
+
+ else:
+ raise ValueError(
+ 'plot has to be in ["time","all","heatmap"] or integer in range(1,model.h)'
+ )
+
+ plt.tight_layout()
+
+ if output == "plot":
+ plt.show()
+ elif output == "figure":
+ return fig
+ else:
+ raise ValueError(f"Invalid output: {output}. Expected 'plot' or 'figure'.")
+```
+
+#### 1.1 Mean attention
+
+
+```python
+plot_attention(nf.models[0], plot="time")
+```
+
+#### 1.2 Attention of all future time steps
+
+
+```python
+plot_attention(nf.models[0], plot="all")
+```
+
+#### 1.3 Attention of a specific future time step
+
+
+```python
+plot_attention(nf.models[0], plot=8)
+```
+
+## 2. Feature Importance
+
+### 2.1 Global feature importance
+
+
+```python
+feature_importances = nf.models[0].feature_importances()
+feature_importances.keys()
+```
+
+#### Static variable importances
+
+
+```python
+feature_importances["Static covariates"].sort_values(by="importance").plot(kind="barh")
+```
+
+#### Past variable importances
+
+
+```python
+feature_importances["Past variable importance over time"].mean().sort_values().plot(
+ kind="barh"
+)
+```
+
+#### Future variable importances
+
+
+```python
+feature_importances["Future variable importance over time"].mean().sort_values().plot(
+ kind="barh"
+)
+```
+
+### 2.2 Variable importances over time
+
+#### Future variable importance over time
+
+Importance of each future covariate at each future time step
+
+
+```python
+df = feature_importances["Future variable importance over time"]
+
+
+fig, ax = plt.subplots(figsize=(20, 10))
+bottom = np.zeros(len(df.index))
+for col in df.columns:
+ p = ax.bar(np.arange(-len(df), 0), df[col].values, 0.6, label=col, bottom=bottom)
+ bottom += df[col]
+ax.set_title("Future variable importance over time ponderated by attention")
+ax.set_ylabel("Importance")
+ax.set_xlabel("Time")
+ax.grid(True)
+ax.legend()
+plt.show()
+```
+
+2.3
+
+#### Past variable importance over time
+
+
+```python
+df = feature_importances["Past variable importance over time"]
+
+fig, ax = plt.subplots(figsize=(20, 10))
+bottom = np.zeros(len(df.index))
+
+for col in df.columns:
+ p = ax.bar(np.arange(-len(df), 0), df[col].values, 0.6, label=col, bottom=bottom)
+ bottom += df[col]
+ax.set_title("Past variable importance over time")
+ax.set_ylabel("Importance")
+ax.set_xlabel("Time")
+ax.legend()
+ax.grid(True)
+
+plt.show()
+```
+
+#### Past variable importance over time ponderated by attention
+
+Decomposition of the importance of each time step based on importance of
+each variable at that time step
+
+
+```python
+df = feature_importances["Past variable importance over time"]
+mean_attention = (
+ nf.models[0]
+ .attention_weights()[nf.models[0].input_size :, :]
+ .mean(axis=0)[: nf.models[0].input_size]
+)
+df = df.multiply(mean_attention, axis=0)
+
+fig, ax = plt.subplots(figsize=(20, 10))
+bottom = np.zeros(len(df.index))
+
+for col in df.columns:
+ p = ax.bar(np.arange(-len(df), 0), df[col].values, 0.6, label=col, bottom=bottom)
+ bottom += df[col]
+ax.set_title("Past variable importance over time ponderated by attention")
+ax.set_ylabel("Importance")
+ax.set_xlabel("Time")
+ax.legend()
+ax.grid(True)
+plt.plot(
+ np.arange(-len(df), 0),
+ mean_attention,
+ color="black",
+ marker="o",
+ linestyle="-",
+ linewidth=2,
+ label="mean_attention",
+)
+plt.legend()
+plt.show()
+```
+
+### 3. Variable importance correlations over time
+
+Variables which gain and lose importance at same moments
+
+
+```python
+nf.models[0].feature_importance_correlations()
+```
+
diff --git a/neuralforecast/models.tide.html.mdx b/neuralforecast/models.tide.html.mdx
new file mode 100644
index 00000000..f45d92c6
--- /dev/null
+++ b/neuralforecast/models.tide.html.mdx
@@ -0,0 +1,239 @@
+---
+description: >-
+ Time-series Dense Encoder (`TiDE`) is a MLP-based univariate time-series
+ forecasting model. `TiDE` uses Multi-layer Perceptrons (MLPs) in an
+ encoder-decoder model for long-term time-series forecasting. In addition, this
+ model can handle exogenous inputs.
+output-file: models.tide.html
+title: TiDE
+---
+
+
+
+
+Figure 1. TiDE architecture.
+
+
+## 1. Auxiliary Functions
+
+## 1.1 MLP residual
+
+An MLP block with a residual connection.
+
+------------------------------------------------------------------------
+
+source
+
+### MLPResidual
+
+> ``` text
+> MLPResidual (input_dim, hidden_size, output_dim, dropout, layernorm)
+> ```
+
+*MLPResidual*
+
+## 2. Model
+
+------------------------------------------------------------------------
+
+source
+
+### TiDE
+
+> ``` text
+> TiDE (h, input_size, hidden_size=512, decoder_output_dim=32,
+> temporal_decoder_dim=128, dropout=0.3, layernorm=True,
+> num_encoder_layers=1, num_decoder_layers=1, temporal_width=4,
+> futr_exog_list=None, hist_exog_list=None, stat_exog_list=None,
+> exclude_insample_y=False, loss=MAE(), valid_loss=None,
+> max_steps:int=1000, learning_rate:float=0.001,
+> num_lr_decays:int=-1, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size:int=32,
+> valid_batch_size:Optional[int]=None, windows_batch_size=1024,
+> inference_windows_batch_size=1024, start_padding_enabled=False,
+> step_size:int=1, scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None, **trainer_kwargs)
+> ```
+
+\*TiDE
+
+Time-series Dense Encoder
+([`TiDE`](https://nixtlaverse.nixtla.io/neuralforecast/models.tide.html#tide))
+is a MLP-based univariate time-series forecasting model.
+[`TiDE`](https://nixtlaverse.nixtla.io/neuralforecast/models.tide.html#tide)
+uses Multi-layer Perceptrons (MLPs) in an encoder-decoder model for
+long-term time-series forecasting.
+
+**Parameters:** `h`: int, forecast horizon. `input_size`: int,
+considered autorregresive inputs (lags), y=\[1,2,3,4\] input_size=2 -\>
+lags=\[1,2\]. `hidden_size`: int=1024, number of units for the dense
+MLPs. `decoder_output_dim`: int=32, number of units for the output
+of the decoder. `temporal_decoder_dim`: int=128, number of units for
+the hidden sizeof the temporal decoder. `dropout`: float=0.0,
+dropout rate between (0, 1) . `layernorm`: bool=True, if True uses
+Layer Normalization on the MLP residual block outputs.
+`num_encoder_layers`: int=1, number of encoder layers.
+`num_decoder_layers`: int=1, number of decoder layers.
+`temporal_width`: int=4, lower temporal projected dimension.
+`futr_exog_list`: str list, future exogenous columns.
+`hist_exog_list`: str list, historic exogenous columns.
+`stat_exog_list`: str list, static exogenous columns.
+`exclude_insample_y`: bool=False, whether to exclude the target variable
+from the historic exogenous data. `loss`: PyTorch module,
+instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch.
+`windows_batch_size`: int=1024, number of windows to sample in each
+training batch, default uses all. `inference_windows_batch_size`:
+int=1024, number of windows to sample in each inference batch, -1 uses
+all. `start_padding_enabled`: bool=False, if True, the model will
+pad the time series with zeros at the beginning, by input size.
+`step_size`: int=1, step size between each window of temporal data.
+`scaler_type`: str=‘identity’, type of scaler for temporal inputs
+normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`. `dataloader_kwargs`:
+dict, optional, list of parameters passed into the PyTorch Lightning
+dataloader by the `TimeSeriesDataLoader`. `**trainer_kwargs`: int,
+keyword trainer arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+**References:** - [Das, Abhimanyu, Weihao Kong, Andrew Leach, Shaan
+Mathur, Rajat Sen, and Rose Yu (2024). “Long-term Forecasting with TiDE:
+Time-series Dense Encoder.”](http://arxiv.org/abs/2304.08424)\*
+
+------------------------------------------------------------------------
+
+### TiDE.fit
+
+> ``` text
+> TiDE.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### TiDE.predict
+
+> ``` text
+> TiDE.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## 3. Usage Examples
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import TiDE
+from neuralforecast.losses.pytorch import GMM
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+fcst = NeuralForecast(
+ models=[
+ TiDE(h=12,
+ input_size=24,
+ loss=GMM(n_components=7, return_params=True, level=[80,90], weighted=True),
+ max_steps=100,
+ scaler_type='standard',
+ futr_exog_list=['y_[lag12]'],
+ hist_exog_list=None,
+ stat_exog_list=['airline1'],
+ ),
+ ],
+ freq='ME'
+)
+fcst.fit(df=Y_train_df, static_df=AirPassengersStatic)
+forecasts = fcst.predict(futr_df=Y_test_df)
+
+# Plot quantile predictions
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['TiDE-median'], c='blue', label='median')
+plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['TiDE-lo-90'][-12:].values,
+ y2=plot_df['TiDE-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+plt.legend()
+plt.grid()
+```
+
diff --git a/neuralforecast/models.timellm.html.mdx b/neuralforecast/models.timellm.html.mdx
new file mode 100644
index 00000000..a87d22b6
--- /dev/null
+++ b/neuralforecast/models.timellm.html.mdx
@@ -0,0 +1,301 @@
+---
+output-file: models.timellm.html
+title: Time-LLM
+---
+
+
+Time-LLM is a reprogramming framework to repurpose LLMs for general time
+series forecasting with the backbone language models kept intact. In
+other words, it transforms a forecasting task into a “language task”
+that can be tackled by an off-the-shelf LLM.
+
+**References** - [Ming Jin, Shiyu Wang, Lintao Ma, Zhixuan Chu,
+James Y. Zhang, Xiaoming Shi, Pin-Yu Chen, Yuxuan Liang, Yuan-Fang Li,
+Shirui Pan, Qingsong Wen. “Time-LLM: Time Series Forecasting by
+Reprogramming Large Language
+Models”](https://arxiv.org/abs/2310.01728)
+
+
+
+Figure 1. Time-LLM
+Architecture.
+
+
+## 1. Auxiliary Functions
+
+------------------------------------------------------------------------
+
+source
+
+### ReprogrammingLayer
+
+> ``` text
+> ReprogrammingLayer (d_model, n_heads, d_keys=None, d_llm=None,
+> attention_dropout=0.1)
+> ```
+
+*ReprogrammingLayer*
+
+------------------------------------------------------------------------
+
+source
+
+### FlattenHead
+
+> ``` text
+> FlattenHead (n_vars, nf, target_window, head_dropout=0)
+> ```
+
+*FlattenHead*
+
+------------------------------------------------------------------------
+
+source
+
+### PatchEmbedding
+
+> ``` text
+> PatchEmbedding (d_model, patch_len, stride, dropout)
+> ```
+
+*PatchEmbedding*
+
+------------------------------------------------------------------------
+
+source
+
+### TokenEmbedding
+
+> ``` text
+> TokenEmbedding (c_in, d_model)
+> ```
+
+*TokenEmbedding*
+
+------------------------------------------------------------------------
+
+source
+
+### ReplicationPad1d
+
+> ``` text
+> ReplicationPad1d (padding)
+> ```
+
+*ReplicationPad1d*
+
+## 2. Model
+
+------------------------------------------------------------------------
+
+source
+
+### TimeLLM
+
+> ``` text
+> TimeLLM (h, input_size, patch_len:int=16, stride:int=8, d_ff:int=128,
+> top_k:int=5, d_llm:int=768, d_model:int=32, n_heads:int=8,
+> enc_in:int=7, dec_in:int=7, llm=None, llm_config=None,
+> llm_tokenizer=None, llm_num_hidden_layers=32,
+> llm_output_attention:bool=True,
+> llm_output_hidden_states:bool=True,
+> prompt_prefix:Optional[str]=None, dropout:float=0.1,
+> stat_exog_list=None, hist_exog_list=None, futr_exog_list=None,
+> loss=MAE(), valid_loss=None, learning_rate:float=0.0001,
+> max_steps:int=5, val_check_steps:int=100, batch_size:int=32,
+> valid_batch_size:Optional[int]=None,
+> windows_batch_size:int=1024,
+> inference_windows_batch_size:int=1024,
+> start_padding_enabled:bool=False, step_size:int=1,
+> num_lr_decays:int=0, early_stop_patience_steps:int=-1,
+> scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*TimeLLM
+
+Time-LLM is a reprogramming framework to repurpose an off-the-shelf LLM
+for time series forecasting.
+
+It trains a reprogramming layer that translates the observed series into
+a language task. This is fed to the LLM and an output projection layer
+translates the output back to numerical predictions.
+
+**Parameters:** `h`: int, Forecast horizon. `input_size`: int,
+autorregresive inputs size, y=\[1,2,3,4\] input_size=2 -\>
+y\_\[t-2:t\]=\[1,2\]. `patch_len`: int=16, length of patch.
+`stride`: int=8, stride of patch. `d_ff`: int=128, dimension of
+fcn. `top_k`: int=5, top tokens to consider. `d_llm`: int=768,
+hidden dimension of LLM. \# LLama7b:4096; GPT2-small:768;
+BERT-base:768 `d_model`: int=32, dimension of model. `n_heads`:
+int=8, number of heads in attention layer. `enc_in`: int=7, encoder
+input size. `dec_in`: int=7, decoder input size. `llm` = None,
+Path to pretrained LLM model to use. If not specified, it will use GPT-2
+from https://huggingface.co/openai-community/gpt2” `llm_config` =
+Deprecated, configuration of LLM. If not specified, it will use the
+configuration of GPT-2 from
+https://huggingface.co/openai-community/gpt2” `llm_tokenizer` =
+Deprecated, tokenizer of LLM. If not specified, it will use the GPT-2
+tokenizer from https://huggingface.co/openai-community/gpt2”
+`llm_num_hidden_layers` = 32, hidden layers in LLM
+`llm_output_attention`: bool = True, whether to output attention in
+encoder. `llm_output_hidden_states`: bool = True, whether to output
+hidden states. `prompt_prefix`: str=None, prompt to inform the LLM
+about the dataset. `dropout`: float=0.1, dropout rate.
+`stat_exog_list`: str list, static exogenous columns.
+`hist_exog_list`: str list, historic exogenous columns.
+`futr_exog_list`: str list, future exogenous columns. `loss`:
+PyTorch module, instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`max_steps`: int=1000, maximum number of training steps.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=1024, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=1024, number of windows to sample in
+each inference batch. `start_padding_enabled`: bool=False, if True,
+the model will pad the time series with zeros at the beginning, by input
+size. `step_size`: int=1, step size between each window of temporal
+data. `num_lr_decays`: int=-1, Number of learning rate decays,
+evenly distributed across max_steps. `early_stop_patience_steps`:
+int=-1, Number of validation iterations before early stopping.
+`scaler_type`: str=‘identity’, type of scaler for temporal inputs
+normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`. `dataloader_kwargs`:
+dict, optional, list of parameters passed into the PyTorch Lightning
+dataloader by the `TimeSeriesDataLoader`. `**trainer_kwargs`: int,
+keyword trainer arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+**References:** -[Ming Jin, Shiyu Wang, Lintao Ma, Zhixuan Chu,
+James Y. Zhang, Xiaoming Shi, Pin-Yu Chen, Yuxuan Liang, Yuan-Fang Li,
+Shirui Pan, Qingsong Wen. “Time-LLM: Time Series Forecasting by
+Reprogramming Large Language
+Models”](https://arxiv.org/abs/2310.01728)\*
+
+------------------------------------------------------------------------
+
+### TimeLLM.fit
+
+> ``` text
+> TimeLLM.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### TimeLLM.predict
+
+> ``` text
+> TimeLLM.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import TimeLLM
+from neuralforecast.utils import AirPassengersPanel
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+prompt_prefix = "The dataset contains data on monthly air passengers. There is a yearly seasonality"
+
+timellm = TimeLLM(h=12,
+ input_size=36,
+ llm='openai-community/gpt2',
+ prompt_prefix=prompt_prefix,
+ batch_size=16,
+ valid_batch_size=16,
+ windows_batch_size=16)
+
+nf = NeuralForecast(
+ models=[timellm],
+ freq='ME'
+)
+
+nf.fit(df=Y_train_df, val_size=12)
+forecasts = nf.predict(futr_df=Y_test_df)
+```
+
diff --git a/neuralforecast/models.timemixer.html.mdx b/neuralforecast/models.timemixer.html.mdx
new file mode 100644
index 00000000..cfffca2e
--- /dev/null
+++ b/neuralforecast/models.timemixer.html.mdx
@@ -0,0 +1,317 @@
+---
+output-file: models.timemixer.html
+title: TimeMixer
+---
+
+
+
+
+Figure 1. Architecture of
+SOFTS.
+
+
+### Embedding
+
+------------------------------------------------------------------------
+
+source
+
+### DataEmbedding_wo_pos
+
+> ``` text
+> DataEmbedding_wo_pos (c_in, d_model, dropout=0.1, embed_type='fixed',
+> freq='h')
+> ```
+
+*DataEmbedding_wo_pos*
+
+### DFT decomposition
+
+------------------------------------------------------------------------
+
+source
+
+### DFT_series_decomp
+
+> ``` text
+> DFT_series_decomp (top_k)
+> ```
+
+*Series decomposition block*
+
+### Mixing
+
+------------------------------------------------------------------------
+
+source
+
+### PastDecomposableMixing
+
+> ``` text
+> PastDecomposableMixing (seq_len, pred_len, down_sampling_window,
+> down_sampling_layers, d_model, dropout,
+> channel_independence, decomp_method, d_ff,
+> moving_avg, top_k)
+> ```
+
+*PastDecomposableMixing*
+
+------------------------------------------------------------------------
+
+source
+
+### MultiScaleTrendMixing
+
+> ``` text
+> MultiScaleTrendMixing (seq_len, down_sampling_window,
+> down_sampling_layers)
+> ```
+
+*Top-down mixing trend pattern*
+
+------------------------------------------------------------------------
+
+source
+
+### MultiScaleSeasonMixing
+
+> ``` text
+> MultiScaleSeasonMixing (seq_len, down_sampling_window,
+> down_sampling_layers)
+> ```
+
+*Bottom-up mixing season pattern*
+
+## 2. Model
+
+------------------------------------------------------------------------
+
+source
+
+### TimeMixer
+
+> ``` text
+> TimeMixer (h, input_size, n_series, stat_exog_list=None,
+> hist_exog_list=None, futr_exog_list=None, d_model:int=32,
+> d_ff:int=32, dropout:float=0.1, e_layers:int=4, top_k:int=5,
+> decomp_method:str='moving_avg', moving_avg:int=25,
+> channel_independence:int=0, down_sampling_layers:int=1,
+> down_sampling_window:int=2, down_sampling_method:str='avg',
+> use_norm:bool=True, decoder_input_size_multiplier:float=0.5,
+> loss=MAE(), valid_loss=None, max_steps:int=1000,
+> learning_rate:float=0.001, num_lr_decays:int=-1,
+> early_stop_patience_steps:int=-1, val_check_steps:int=100,
+> batch_size:int=32, valid_batch_size:Optional[int]=None,
+> windows_batch_size=32, inference_windows_batch_size=32,
+> start_padding_enabled=False, step_size:int=1,
+> scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*TimeMixer **Parameters** `h`: int, Forecast horizon.
+`input_size`: int, autorregresive inputs size, y=\[1,2,3,4\]
+input_size=2 -\> y\_\[t-2:t\]=\[1,2\]. `n_series`: int, number of
+time-series. `stat_exog_list`: str list, static exogenous
+columns. `hist_exog_list`: str list, historic exogenous columns.
+`futr_exog_list`: str list, future exogenous columns. `d_model`:
+int, dimension of the model. `d_ff`: int, dimension of the
+fully-connected network. `dropout`: float, dropout rate.
+`e_layers`: int, number of encoder layers. `top_k`: int, number of
+selected frequencies. `decomp_method`: str, method of series
+decomposition \[moving_avg, dft_decomp\]. `moving_avg`: int, window
+size of moving average. `channel_independence`: int, 0: channel
+dependence, 1: channel independence. `down_sampling_layers`: int,
+number of downsampling layers. `down_sampling_window`: int, size of
+downsampling window. `down_sampling_method`: str, down sampling
+method \[avg, max, conv\]. `use_norm`: bool, whether to normalize or
+not. `decoder_input_size_multiplier`: float = 0.5. `loss`:
+PyTorch module, instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=32, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=32, number of windows to sample in
+each inference batch, -1 uses all. `start_padding_enabled`:
+bool=False, if True, the model will pad the time series with zeros at
+the beginning, by input size. `step_size`: int=1, step size between
+each window of temporal data. `scaler_type`: str=‘identity’, type of
+scaler for temporal inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`. `dataloader_kwargs`:
+dict, optional, list of parameters passed into the PyTorch Lightning
+dataloader by the `TimeSeriesDataLoader`. `**trainer_kwargs`:
+keyword trainer arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+**References** [Shiyu Wang, Haixu Wu, Xiaoming Shi, Tengge Hu,
+Huakun Luo, Lintao Ma, James Y. Zhang, Jun Zhou.”TimeMixer: Decomposable
+Multiscale Mixing For Time Series
+Forecasting”](https://openreview.net/pdf?id=7oLshfEIC2) \*
+
+------------------------------------------------------------------------
+
+### TimeMixer.fit
+
+> ``` text
+> TimeMixer.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### TimeMixer.predict
+
+> ``` text
+> TimeMixer.predict (dataset, test_size=None, step_size=1,
+> random_seed=None, quantiles=None,
+> **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## 3. Usage example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import TimeMixer
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+from neuralforecast.losses.pytorch import MAE
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = TimeMixer(h=12,
+ input_size=24,
+ n_series=2,
+ scaler_type='standard',
+ max_steps=500,
+ early_stop_patience_steps=-1,
+ val_check_steps=5,
+ learning_rate=1e-3,
+ loss = MAE(),
+ valid_loss=MAE(),
+ batch_size=32
+ )
+
+fcst = NeuralForecast(models=[model], freq='ME')
+fcst.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+forecasts = fcst.predict(futr_df=Y_test_df)
+
+# Plot predictions
+fig, ax = plt.subplots(1, 1, figsize = (20, 7))
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['TimeMixer'], c='blue', label='median')
+ax.set_title('AirPassengers Forecast', fontsize=22)
+ax.set_ylabel('Monthly Passengers', fontsize=20)
+ax.set_xlabel('Year', fontsize=20)
+ax.legend(prop={'size': 15})
+ax.grid()
+```
+
+Using `cross_validation` to forecast multiple historic values.
+
+
+```python
+fcst = NeuralForecast(models=[model], freq='M')
+forecasts = fcst.cross_validation(df=AirPassengersPanel, static_df=AirPassengersStatic, n_windows=2, step_size=12)
+
+# Plot predictions
+fig, ax = plt.subplots(1, 1, figsize = (20, 7))
+Y_hat_df = forecasts.loc['Airline1']
+Y_df = AirPassengersPanel[AirPassengersPanel['unique_id']=='Airline1']
+
+plt.plot(Y_df['ds'], Y_df['y'], c='black', label='True')
+plt.plot(Y_hat_df['ds'], Y_hat_df['TimeMixer'], c='blue', label='Forecast')
+ax.set_title('AirPassengers Forecast', fontsize=22)
+ax.set_ylabel('Monthly Passengers', fontsize=20)
+ax.set_xlabel('Year', fontsize=20)
+ax.legend(prop={'size': 15})
+ax.grid()
+```
+
diff --git a/neuralforecast/models.timesnet.html.mdx b/neuralforecast/models.timesnet.html.mdx
new file mode 100644
index 00000000..13ce87b0
--- /dev/null
+++ b/neuralforecast/models.timesnet.html.mdx
@@ -0,0 +1,278 @@
+---
+output-file: models.timesnet.html
+title: TimesNet
+---
+
+
+The TimesNet univariate model tackles the challenge of modeling multiple
+intraperiod and interperiod temporal variations.
+
+The architecture has the following distinctive features: - An embedding
+layer that maps the input sequence into a latent space. - Transformation
+of 1D time seires into 2D tensors, based on periods found by FFT. - A
+convolutional Inception block that captures temporal variations at
+different scales and between periods.
+
+**References** - [Haixu Wu and Tengge Hu and Yong Liu and Hang Zhou
+and Jianmin Wang and Mingsheng Long. TimesNet: Temporal 2D-Variation
+Modeling for General Time Series
+Analysis](https://openreview.net/pdf?id=ju_Uqw384Oq) - Based on the
+implementation in https://github.com/thuml/Time-Series-Library (license:
+https://github.com/thuml/Time-Series-Library/blob/main/LICENSE)
+
+
+
+Figure 1. TimesNet
+Architecture.
+
+
+## 1. Auxiliary Functions
+
+------------------------------------------------------------------------
+
+source
+
+### Inception_Block_V1
+
+> ``` text
+> Inception_Block_V1 (in_channels, out_channels, num_kernels=6,
+> init_weight=True)
+> ```
+
+*Inception_Block_V1*
+
+------------------------------------------------------------------------
+
+source
+
+### TimesBlock
+
+> ``` text
+> TimesBlock (input_size, h, k, hidden_size, conv_hidden_size, num_kernels)
+> ```
+
+*TimesBlock*
+
+------------------------------------------------------------------------
+
+source
+
+### FFT_for_Period
+
+> ``` text
+> FFT_for_Period (x, k=2)
+> ```
+
+## 2. TimesNet
+
+------------------------------------------------------------------------
+
+source
+
+### TimesNet
+
+> ``` text
+> TimesNet (h:int, input_size:int, stat_exog_list=None,
+> hist_exog_list=None, futr_exog_list=None,
+> exclude_insample_y=False, hidden_size:int=64,
+> dropout:float=0.1, conv_hidden_size:int=64, top_k:int=5,
+> num_kernels:int=6, encoder_layers:int=2, loss=MAE(),
+> valid_loss=None, max_steps:int=1000,
+> learning_rate:float=0.0001, num_lr_decays:int=-1,
+> early_stop_patience_steps:int=-1, val_check_steps:int=100,
+> batch_size:int=32, valid_batch_size:Optional[int]=None,
+> windows_batch_size=64, inference_windows_batch_size=256,
+> start_padding_enabled=False, step_size:int=1,
+> scaler_type:str='standard', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*TimesNet
+
+The TimesNet univariate model tackles the challenge of modeling multiple
+intraperiod and interperiod temporal variations.
+
+**Parameters** `h` : int, Forecast horizon. `input_size` : int,
+Length of input window (lags). `stat_exog_list` : list of str,
+optional (default=None), Static exogenous columns. `hist_exog_list`
+: list of str, optional (default=None), Historic exogenous columns.
+`futr_exog_list` : list of str, optional (default=None), Future
+exogenous columns. `exclude_insample_y` : bool (default=False), The
+model skips the autoregressive features y\[t-input_size:t\] if True.
+`hidden_size` : int (default=64), Size of embedding for embedding and
+encoders. `dropout` : float between \[0, 1) (default=0.1), Dropout
+for embeddings. `conv_hidden_size`: int (default=64), Channels of
+the Inception block. `top_k`: int (default=5), Number of
+periods. `num_kernels`: int (default=6), Number of kernels for the
+Inception block. `encoder_layers` : int, (default=2), Number of
+encoder layers. `loss`: PyTorch module (default=MAE()), Instantiated
+train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module (default=None, uses loss), Instantiated
+validation loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int (default=1000), Maximum number of training steps.
+`learning_rate` : float (default=1e-4), Learning rate.
+`num_lr_decays`: int (default=-1), Number of learning rate decays,
+evenly distributed across max_steps. If -1, no learning rate decay is
+performed. `early_stop_patience_steps` : int (default=-1), Number of
+validation iterations before early stopping. If -1, no early stopping is
+performed. `val_check_steps` : int (default=100), Number of training
+steps between every validation loss check. `batch_size` : int
+(default=32), Number of different series in each batch.
+`valid_batch_size` : int (default=None), Number of different series in
+each validation and test batch, if None uses batch_size.
+`windows_batch_size` : int (default=64), Number of windows to sample in
+each training batch. `inference_windows_batch_size` : int
+(default=256), Number of windows to sample in each inference batch.
+`start_padding_enabled` : bool (default=False), If True, the model will
+pad the time series with zeros at the beginning by input size.
+`step_size` : int (default=1), Step size between each window of temporal
+data. `scaler_type` : str (default=‘standard’), Type of scaler for
+temporal inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed` : int (default=1), Random_seed for pytorch initializer and
+numpy generators. `drop_last_loader` : bool (default=False), If True
+`TimeSeriesDataLoader` drops last non-full batch. `alias` : str,
+optional (default=None), Custom name of the model. `optimizer`:
+Subclass of ‘torch.optim.Optimizer’, optional (default=None), User
+specified optimizer instead of the default choice (Adam).
+`optimizer_kwargs`: dict, optional (defualt=None), List of parameters
+used by the user specified `optimizer`. `lr_scheduler`: Subclass of
+‘torch.optim.lr_scheduler.LRScheduler’, optional, user specified
+lr_scheduler instead of the default choice (StepLR).
+`lr_scheduler_kwargs`: dict, optional, list of parameters used by the
+user specified `lr_scheduler`.
+`dataloader_kwargs`: dict, optional (default=None), List of parameters
+passed into the PyTorch Lightning dataloader by the
+`TimeSeriesDataLoader`. `**trainer_kwargs`: Keyword trainer
+arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer)\*
+
+------------------------------------------------------------------------
+
+### TimesNet.fit
+
+> ``` text
+> TimesNet.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### TimesNet.predict
+
+> ``` text
+> TimesNet.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.losses.pytorch import DistributionLoss
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = TimesNet(h=12,
+ input_size=24,
+ hidden_size = 16,
+ conv_hidden_size = 32,
+ loss=DistributionLoss(distribution='Normal', level=[80, 90]),
+ scaler_type='standard',
+ learning_rate=1e-3,
+ max_steps=100,
+ val_check_steps=50,
+ early_stop_patience_steps=2)
+
+nf = NeuralForecast(
+ models=[model],
+ freq='ME'
+)
+nf.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+forecasts = nf.predict(futr_df=Y_test_df)
+
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+if model.loss.is_distribution_output:
+ plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+ plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+ plt.plot(plot_df['ds'], plot_df['TimesNet-median'], c='blue', label='median')
+ plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['TimesNet-lo-90'][-12:].values,
+ y2=plot_df['TimesNet-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+ plt.grid()
+ plt.legend()
+ plt.plot()
+else:
+ plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+ plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+ plt.plot(plot_df['ds'], plot_df['TimesNet'], c='blue', label='Forecast')
+ plt.legend()
+ plt.grid()
+```
+
diff --git a/neuralforecast/models.timexer.html.mdx b/neuralforecast/models.timexer.html.mdx
new file mode 100644
index 00000000..96e896be
--- /dev/null
+++ b/neuralforecast/models.timexer.html.mdx
@@ -0,0 +1,400 @@
+---
+output-file: models.timexer.html
+title: TimeXer
+---
+
+
+
+
+Figure 1. Architecture of
+TimeXer.
+
+
+# 1. Auxiliary functions
+
+------------------------------------------------------------------------
+
+source
+
+### FlattenHead
+
+> ``` text
+> FlattenHead (n_vars, nf, target_window, head_dropout=0)
+> ```
+
+\*Base class for all neural network modules.
+
+Your models should also subclass this class.
+
+Modules can also contain other Modules, allowing them to be nested in a
+tree structure. You can assign the submodules as regular attributes::
+
+``` text
+import torch.nn as nn
+import torch.nn.functional as F
+
+class Model(nn.Module):
+ def __init__(self) -> None:
+ super().__init__()
+ self.conv1 = nn.Conv2d(1, 20, 5)
+ self.conv2 = nn.Conv2d(20, 20, 5)
+
+ def forward(self, x):
+ x = F.relu(self.conv1(x))
+ return F.relu(self.conv2(x))
+```
+
+Submodules assigned in this way will be registered, and will also have
+their parameters converted when you call :meth:`to`, etc.
+
+.. note:: As per the example above, an `__init__()` call to the parent
+class must be made before assignment on the child.
+
+:ivar training: Boolean represents whether this module is in training or
+evaluation mode. :vartype training: bool\*
+
+------------------------------------------------------------------------
+
+source
+
+### Encoder
+
+> ``` text
+> Encoder (layers, norm_layer=None, projection=None)
+> ```
+
+\*Base class for all neural network modules.
+
+Your models should also subclass this class.
+
+Modules can also contain other Modules, allowing them to be nested in a
+tree structure. You can assign the submodules as regular attributes::
+
+``` text
+import torch.nn as nn
+import torch.nn.functional as F
+
+class Model(nn.Module):
+ def __init__(self) -> None:
+ super().__init__()
+ self.conv1 = nn.Conv2d(1, 20, 5)
+ self.conv2 = nn.Conv2d(20, 20, 5)
+
+ def forward(self, x):
+ x = F.relu(self.conv1(x))
+ return F.relu(self.conv2(x))
+```
+
+Submodules assigned in this way will be registered, and will also have
+their parameters converted when you call :meth:`to`, etc.
+
+.. note:: As per the example above, an `__init__()` call to the parent
+class must be made before assignment on the child.
+
+:ivar training: Boolean represents whether this module is in training or
+evaluation mode. :vartype training: bool\*
+
+------------------------------------------------------------------------
+
+source
+
+### EncoderLayer
+
+> ``` text
+> EncoderLayer (self_attention, cross_attention, d_model, d_ff=None,
+> dropout=0.1, activation='relu')
+> ```
+
+\*Base class for all neural network modules.
+
+Your models should also subclass this class.
+
+Modules can also contain other Modules, allowing them to be nested in a
+tree structure. You can assign the submodules as regular attributes::
+
+``` text
+import torch.nn as nn
+import torch.nn.functional as F
+
+class Model(nn.Module):
+ def __init__(self) -> None:
+ super().__init__()
+ self.conv1 = nn.Conv2d(1, 20, 5)
+ self.conv2 = nn.Conv2d(20, 20, 5)
+
+ def forward(self, x):
+ x = F.relu(self.conv1(x))
+ return F.relu(self.conv2(x))
+```
+
+Submodules assigned in this way will be registered, and will also have
+their parameters converted when you call :meth:`to`, etc.
+
+.. note:: As per the example above, an `__init__()` call to the parent
+class must be made before assignment on the child.
+
+:ivar training: Boolean represents whether this module is in training or
+evaluation mode. :vartype training: bool\*
+
+------------------------------------------------------------------------
+
+source
+
+### EnEmbedding
+
+> ``` text
+> EnEmbedding (n_vars, d_model, patch_len, dropout)
+> ```
+
+\*Base class for all neural network modules.
+
+Your models should also subclass this class.
+
+Modules can also contain other Modules, allowing them to be nested in a
+tree structure. You can assign the submodules as regular attributes::
+
+``` text
+import torch.nn as nn
+import torch.nn.functional as F
+
+class Model(nn.Module):
+ def __init__(self) -> None:
+ super().__init__()
+ self.conv1 = nn.Conv2d(1, 20, 5)
+ self.conv2 = nn.Conv2d(20, 20, 5)
+
+ def forward(self, x):
+ x = F.relu(self.conv1(x))
+ return F.relu(self.conv2(x))
+```
+
+Submodules assigned in this way will be registered, and will also have
+their parameters converted when you call :meth:`to`, etc.
+
+.. note:: As per the example above, an `__init__()` call to the parent
+class must be made before assignment on the child.
+
+:ivar training: Boolean represents whether this module is in training or
+evaluation mode. :vartype training: bool\*
+
+# 2. Model
+
+------------------------------------------------------------------------
+
+source
+
+### TimeXer
+
+> ``` text
+> TimeXer (h, input_size, n_series, futr_exog_list=None,
+> hist_exog_list=None, stat_exog_list=None,
+> exclude_insample_y:bool=False, patch_len:int=16,
+> hidden_size:int=512, n_heads:int=8, e_layers:int=2,
+> d_ff:int=2048, factor:int=1, dropout:float=0.1,
+> use_norm:bool=True, loss=MAE(), valid_loss=None,
+> max_steps:int=1000, learning_rate:float=0.001,
+> num_lr_decays:int=-1, early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size:int=32,
+> valid_batch_size:Optional[int]=None, windows_batch_size=32,
+> inference_windows_batch_size=32, start_padding_enabled=False,
+> step_size:int=1, scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*TimeXer
+
+**Parameters:** `h`: int, Forecast horizon. `input_size`: int,
+autorregresive inputs size, y=\[1,2,3,4\] input_size=2 -\>
+y\_\[t-2:t\]=\[1,2\]. `n_series`: int, number of time-series.
+`futr_exog_list`: str list, future exogenous columns.
+`hist_exog_list`: str list, historic exogenous columns.
+`stat_exog_list`: str list, static exogenous columns. `patch_len`:
+int, length of patches. `hidden_size`: int, dimension of the
+model. `n_heads`: int, number of heads. `e_layers`: int, number
+of encoder layers. `d_ff`: int, dimension of fully-connected
+layer. `factor`: int, attention factor. `dropout`: float,
+dropout rate. `use_norm`: bool, whether to normalize or not.
+`loss`: PyTorch module, instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=32, number of windows in each
+batch.
+`inference_windows_batch_size`: int=32, number of windows to sample in
+each inference batch, -1 uses all. `start_padding_enabled`:
+bool=False, if True, the model will pad the time series with zeros at
+the beginning, by input size. `step_size`: int=1, step size between
+each window of temporal data. `scaler_type`: str=‘identity’, type of
+scaler for temporal inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`. `dataloader_kwargs`:
+dict, optional, list of parameters passed into the PyTorch Lightning
+dataloader by the `TimeSeriesDataLoader`. `**trainer_kwargs`: int,
+keyword trainer arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+**Parameters:**
+
+**References** - [Yuxuan Wang, Haixu Wu, Jiaxiang Dong, Guo Qin, Haoran
+Zhang, Yong Liu, Yunzhong Qiu, Jianmin Wang, Mingsheng Long. “TimeXer:
+Empowering Transformers for Time Series Forecasting with Exogenous
+Variables”](https://arxiv.org/abs/2402.19072)\*
+
+------------------------------------------------------------------------
+
+### TimeXer.fit
+
+> ``` text
+> TimeXer.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### TimeXer.predict
+
+> ``` text
+> TimeXer.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+
+```python
+# Unit tests for models
+logging.getLogger("pytorch_lightning").setLevel(logging.ERROR)
+logging.getLogger("lightning_fabric").setLevel(logging.ERROR)
+with warnings.catch_warnings():
+ warnings.simplefilter("ignore")
+ check_model(TimeXer, ["airpassengers"])
+```
+
+# 3. Usage example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import TimeXer
+from neuralforecast.losses.pytorch import MSE
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic, augment_calendar_df
+
+AirPassengersPanel, calendar_cols = augment_calendar_df(df=AirPassengersPanel, freq='M')
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = TimeXer(h=12,
+ input_size=24,
+ n_series=2,
+ futr_exog_list=["trend", "month"],
+ patch_len=12,
+ hidden_size=128,
+ n_heads=16,
+ e_layers=2,
+ d_ff=256,
+ factor=1,
+ dropout=0.1,
+ use_norm=True,
+ loss=MSE(),
+ valid_loss=MAE(),
+ early_stop_patience_steps=3,
+ batch_size=32)
+
+fcst = NeuralForecast(models=[model], freq='ME')
+fcst.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+forecasts = fcst.predict(futr_df=Y_test_df)
+
+# Plot predictions
+fig, ax = plt.subplots(1, 1, figsize = (20, 7))
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['TimeXer'], c='blue', label='Forecast')
+ax.set_title('AirPassengers Forecast', fontsize=22)
+ax.set_ylabel('Monthly Passengers', fontsize=20)
+ax.set_xlabel('Year', fontsize=20)
+ax.legend(prop={'size': 15})
+ax.grid()
+```
+
diff --git a/neuralforecast/models.tsmixer.html.mdx b/neuralforecast/models.tsmixer.html.mdx
new file mode 100644
index 00000000..b52302a5
--- /dev/null
+++ b/neuralforecast/models.tsmixer.html.mdx
@@ -0,0 +1,314 @@
+---
+description: >-
+ Time-Series Mixer (`TSMixer`) is a MLP-based multivariate time-series
+ forecasting model. `TSMixer` jointly learns temporal and cross-sectional
+ representations of the time-series by repeatedly combining time- and feature
+ information using stacked mixing layers. A mixing layer consists of a
+ sequential time- and feature Multi Layer Perceptron (`MLP`). Note: this model
+ cannot handle exogenous inputs. If you want to use additional exogenous
+ inputs, use `TSMixerx`.
+output-file: models.tsmixer.html
+title: TSMixer
+---
+
+
+
+
+Figure 1. TSMixer for multivariate time
+series forecasting.
+
+
+## 1. Auxiliary Functions
+
+## 1.1 Mixing layers
+
+A mixing layer consists of a sequential time- and feature Multi Layer
+Perceptron
+([`MLP`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlp.html#mlp)).
+
+------------------------------------------------------------------------
+
+source
+
+### MixingLayer
+
+> ``` text
+> MixingLayer (n_series, input_size, dropout, ff_dim)
+> ```
+
+*MixingLayer*
+
+------------------------------------------------------------------------
+
+source
+
+### FeatureMixing
+
+> ``` text
+> FeatureMixing (n_series, input_size, dropout, ff_dim)
+> ```
+
+*FeatureMixing*
+
+------------------------------------------------------------------------
+
+source
+
+### TemporalMixing
+
+> ``` text
+> TemporalMixing (n_series, input_size, dropout)
+> ```
+
+*TemporalMixing*
+
+## 2. Model
+
+------------------------------------------------------------------------
+
+source
+
+### TSMixer
+
+> ``` text
+> TSMixer (h, input_size, n_series, futr_exog_list=None,
+> hist_exog_list=None, stat_exog_list=None,
+> exclude_insample_y=False, n_block=2, ff_dim=64, dropout=0.9,
+> revin=True, loss=MAE(), valid_loss=None, max_steps:int=1000,
+> learning_rate:float=0.001, num_lr_decays:int=-1,
+> early_stop_patience_steps:int=-1, val_check_steps:int=100,
+> batch_size:int=32, valid_batch_size:Optional[int]=None,
+> windows_batch_size=32, inference_windows_batch_size=32,
+> start_padding_enabled=False, step_size:int=1,
+> scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*TSMixer
+
+Time-Series Mixer
+([`TSMixer`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixer.html#tsmixer))
+is a MLP-based multivariate time-series forecasting model.
+[`TSMixer`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixer.html#tsmixer)
+jointly learns temporal and cross-sectional representations of the
+time-series by repeatedly combining time- and feature information using
+stacked mixing layers. A mixing layer consists of a sequential time- and
+feature Multi Layer Perceptron
+([`MLP`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlp.html#mlp)).
+
+**Parameters:** `h`: int, forecast horizon. `input_size`: int,
+considered autorregresive inputs (lags), y=\[1,2,3,4\] input_size=2 -\>
+lags=\[1,2\]. `n_series`: int, number of time-series.
+`futr_exog_list`: str list, future exogenous columns.
+`hist_exog_list`: str list, historic exogenous columns.
+`stat_exog_list`: str list, static exogenous columns.
+`exclude_insample_y`: bool=False, if True excludes the target variable
+from the input features. `n_block`: int=2, number of mixing layers
+in the model. `ff_dim`: int=64, number of units for the second
+feed-forward layer in the feature MLP. `dropout`: float=0.9, dropout
+rate between (0, 1) . `revin`: bool=True, if True uses Reverse
+Instance Normalization to process inputs and outputs. `loss`:
+PyTorch module, instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=32, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=32, number of windows to sample in
+each inference batch, -1 uses all. `start_padding_enabled`:
+bool=False, if True, the model will pad the time series with zeros at
+the beginning, by input size. `step_size`: int=1, step size between
+each window of temporal data. `scaler_type`: str=‘identity’, type of
+scaler for temporal inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`.
+`dataloader_kwargs`: dict, optional, list of parameters passed into the
+PyTorch Lightning dataloader by the `TimeSeriesDataLoader`.
+`**trainer_kwargs`: int, keyword trainer arguments inherited from
+[PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+**References:** - [Chen, Si-An, Chun-Liang Li, Nate Yoder, Sercan O.
+Arik, and Tomas Pfister (2023). “TSMixer: An All-MLP Architecture for
+Time Series Forecasting.”](http://arxiv.org/abs/2303.06053)\*
+
+------------------------------------------------------------------------
+
+### TSMixer.fit
+
+> ``` text
+> TSMixer.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### TSMixer.predict
+
+> ``` text
+> TSMixer.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+
+```python
+# Unit tests for models
+logging.getLogger("pytorch_lightning").setLevel(logging.ERROR)
+logging.getLogger("lightning_fabric").setLevel(logging.ERROR)
+with warnings.catch_warnings():
+ warnings.simplefilter("ignore")
+ check_model(TSMixer, ["airpassengers"])
+```
+
+## 3. Usage Examples
+
+Train model and forecast future values with `predict` method.
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import TSMixer
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+from neuralforecast.losses.pytorch import MAE, MQLoss
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = TSMixer(h=12,
+ input_size=24,
+ n_series=2,
+ n_block=4,
+ ff_dim=4,
+ dropout=0,
+ revin=True,
+ scaler_type='standard',
+ max_steps=500,
+ early_stop_patience_steps=-1,
+ val_check_steps=5,
+ learning_rate=1e-3,
+ loss=MQLoss(),
+ batch_size=32
+ )
+
+fcst = NeuralForecast(models=[model], freq='ME')
+fcst.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+forecasts = fcst.predict(futr_df=Y_test_df)
+
+# Plot predictions
+fig, ax = plt.subplots(1, 1, figsize = (20, 7))
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline2'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['TSMixer-median'], c='blue', label='median')
+plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['TSMixer-lo-90'][-12:].values,
+ y2=plot_df['TSMixer-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+ax.set_title('AirPassengers Forecast', fontsize=22)
+ax.set_ylabel('Monthly Passengers', fontsize=20)
+ax.set_xlabel('Year', fontsize=20)
+ax.legend(prop={'size': 15})
+ax.grid()
+```
+
+Using `cross_validation` to forecast multiple historic values.
+
+
+```python
+fcst = NeuralForecast(models=[model], freq='M')
+forecasts = fcst.cross_validation(df=AirPassengersPanel, static_df=AirPassengersStatic, n_windows=2, step_size=12)
+
+# Plot predictions
+fig, ax = plt.subplots(1, 1, figsize = (20, 7))
+Y_hat_df = forecasts.loc['Airline1']
+Y_df = AirPassengersPanel[AirPassengersPanel['unique_id']=='Airline1']
+
+plt.plot(Y_df['ds'], Y_df['y'], c='black', label='True')
+plt.plot(Y_hat_df['ds'], Y_hat_df['TSMixer-median'], c='blue', label='Forecast')
+ax.set_title('AirPassengers Forecast', fontsize=22)
+ax.set_ylabel('Monthly Passengers', fontsize=20)
+ax.set_xlabel('Year', fontsize=20)
+ax.legend(prop={'size': 15})
+ax.grid()
+```
+
diff --git a/neuralforecast/models.tsmixerx.html.mdx b/neuralforecast/models.tsmixerx.html.mdx
new file mode 100644
index 00000000..0c4f3b8f
--- /dev/null
+++ b/neuralforecast/models.tsmixerx.html.mdx
@@ -0,0 +1,335 @@
+---
+description: >-
+ Time-Series Mixer exogenous (`TSMixerx`) is a MLP-based multivariate
+ time-series forecasting model, with capability for additional exogenous
+ inputs. `TSMixerx` jointly learns temporal and cross-sectional representations
+ of the time-series by repeatedly combining time- and feature information using
+ stacked mixing layers. A mixing layer consists of a sequential time- and
+ feature Multi Layer Perceptron (`MLP`).
+output-file: models.tsmixerx.html
+title: TSMixerx
+---
+
+
+
+
+Figure 2. TSMixerX for multivariate time
+series forecasting.
+
+
+## 1. Auxiliary Functions
+
+## 1.1 Mixing layers
+
+A mixing layer consists of a sequential time- and feature Multi Layer
+Perceptron
+([`MLP`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlp.html#mlp)).
+
+------------------------------------------------------------------------
+
+source
+
+### MixingLayerWithStaticExogenous
+
+> ``` text
+> MixingLayerWithStaticExogenous (h, dropout, ff_dim, stat_input_size)
+> ```
+
+*MixingLayerWithStaticExogenous*
+
+------------------------------------------------------------------------
+
+source
+
+### MixingLayer
+
+> ``` text
+> MixingLayer (in_features, out_features, h, dropout, ff_dim)
+> ```
+
+*MixingLayer*
+
+------------------------------------------------------------------------
+
+source
+
+### FeatureMixing
+
+> ``` text
+> FeatureMixing (in_features, out_features, h, dropout, ff_dim)
+> ```
+
+*FeatureMixing*
+
+------------------------------------------------------------------------
+
+source
+
+### TemporalMixing
+
+> ``` text
+> TemporalMixing (num_features, h, dropout)
+> ```
+
+*TemporalMixing*
+
+## 1.2 Reversible InstanceNormalization
+
+An Instance Normalization Layer that is reversible, based on [this
+reference
+implementation](https://github.com/google-research/google-research/blob/master/tsmixer/tsmixer_basic/models/rev_in.py).
+
+## 2. Model
+
+------------------------------------------------------------------------
+
+source
+
+### TSMixerx
+
+> ``` text
+> TSMixerx (h, input_size, n_series, futr_exog_list=None,
+> hist_exog_list=None, stat_exog_list=None,
+> exclude_insample_y=False, n_block=2, ff_dim=64, dropout=0.0,
+> revin=True, loss=MAE(), valid_loss=None, max_steps:int=1000,
+> learning_rate:float=0.001, num_lr_decays:int=-1,
+> early_stop_patience_steps:int=-1, val_check_steps:int=100,
+> batch_size:int=32, valid_batch_size:Optional[int]=None,
+> windows_batch_size=32, inference_windows_batch_size=32,
+> start_padding_enabled=False, step_size:int=1,
+> scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False, alias:Optional[str]=None,
+> optimizer=None, optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*TSMixerx
+
+Time-Series Mixer exogenous
+([`TSMixerx`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixerx.html#tsmixerx))
+is a MLP-based multivariate time-series forecasting model, with
+capability for additional exogenous inputs.
+[`TSMixerx`](https://nixtlaverse.nixtla.io/neuralforecast/models.tsmixerx.html#tsmixerx)
+jointly learns temporal and cross-sectional representations of the
+time-series by repeatedly combining time- and feature information using
+stacked mixing layers. A mixing layer consists of a sequential time- and
+feature Multi Layer Perceptron
+([`MLP`](https://nixtlaverse.nixtla.io/neuralforecast/models.mlp.html#mlp)).
+
+**Parameters:** `h`: int, forecast horizon. `input_size`: int,
+considered autorregresive inputs (lags), y=\[1,2,3,4\] input_size=2 -\>
+lags=\[1,2\]. `n_series`: int, number of time-series.
+`futr_exog_list`: str list, future exogenous columns.
+`hist_exog_list`: str list, historic exogenous columns.
+`stat_exog_list`: str list, static exogenous columns.
+`exclude_insample_y`: bool=False, if True excludes insample_y from the
+model. `n_block`: int=2, number of mixing layers in the model.
+`ff_dim`: int=64, number of units for the second feed-forward layer in
+the feature MLP. `dropout`: float=0.0, dropout rate between (0, 1)
+. `revin`: bool=True, if True uses Reverse Instance Normalization on
+`insample_y` and applies it to the outputs.
+`loss`: PyTorch module, instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=32, number of windows to
+sample in each training batch. `inference_windows_batch_size`:
+int=32, number of windows to sample in each inference batch, -1 uses
+all. `start_padding_enabled`: bool=False, if True, the model will
+pad the time series with zeros at the beginning, by input size.
+`step_size`: int=1, step size between each window of temporal data.
+`scaler_type`: str=‘identity’, type of scaler for temporal inputs
+normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`.
+`dataloader_kwargs`: dict, optional, list of parameters passed into the
+PyTorch Lightning dataloader by the `TimeSeriesDataLoader`.
+`**trainer_kwargs`: int, keyword trainer arguments inherited from
+[PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+**References:** - [Chen, Si-An, Chun-Liang Li, Nate Yoder, Sercan O.
+Arik, and Tomas Pfister (2023). “TSMixer: An All-MLP Architecture for
+Time Series Forecasting.”](http://arxiv.org/abs/2303.06053)\*
+
+------------------------------------------------------------------------
+
+### TSMixerx.fit
+
+> ``` text
+> TSMixerx.fit (dataset, val_size=0, test_size=0, random_seed=None,
+> distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### TSMixerx.predict
+
+> ``` text
+> TSMixerx.predict (dataset, test_size=None, step_size=1, random_seed=None,
+> quantiles=None, **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+
+```python
+# Unit tests for models
+logging.getLogger("pytorch_lightning").setLevel(logging.ERROR)
+logging.getLogger("lightning_fabric").setLevel(logging.ERROR)
+with warnings.catch_warnings():
+ warnings.simplefilter("ignore")
+ check_model(TSMixerx, ["airpassengers"])
+```
+
+## 3. Usage Examples
+
+Train model and forecast future values with `predict` method.
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import TSMixerx
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+from neuralforecast.losses.pytorch import GMM
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = TSMixerx(h=12,
+ input_size=24,
+ n_series=2,
+ stat_exog_list=['airline1'],
+ futr_exog_list=['trend'],
+ n_block=4,
+ ff_dim=4,
+ revin=True,
+ scaler_type='robust',
+ max_steps=500,
+ early_stop_patience_steps=-1,
+ val_check_steps=5,
+ learning_rate=1e-3,
+ loss = GMM(n_components=10, weighted=True),
+ batch_size=32
+ )
+
+fcst = NeuralForecast(models=[model], freq='ME')
+fcst.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+forecasts = fcst.predict(futr_df=Y_test_df)
+
+# Plot predictions
+fig, ax = plt.subplots(1, 1, figsize = (20, 7))
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+plt.plot(plot_df['ds'], plot_df['TSMixerx-median'], c='blue', label='median')
+plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['TSMixerx-lo-90'][-12:].values,
+ y2=plot_df['TSMixerx-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+ax.set_title('AirPassengers Forecast', fontsize=22)
+ax.set_ylabel('Monthly Passengers', fontsize=20)
+ax.set_xlabel('Year', fontsize=20)
+ax.legend(prop={'size': 15})
+ax.grid()
+```
+
+Using `cross_validation` to forecast multiple historic values.
+
+
+```python
+fcst = NeuralForecast(models=[model], freq='M')
+forecasts = fcst.cross_validation(df=AirPassengersPanel, static_df=AirPassengersStatic, n_windows=2, step_size=12)
+
+# Plot predictions
+fig, ax = plt.subplots(1, 1, figsize = (20, 7))
+Y_hat_df = forecasts.loc['Airline1']
+Y_df = AirPassengersPanel[AirPassengersPanel['unique_id']=='Airline1']
+
+plt.plot(Y_df['ds'], Y_df['y'], c='black', label='True')
+plt.plot(Y_hat_df['ds'], Y_hat_df['TSMixerx-median'], c='blue', label='Forecast')
+ax.set_title('AirPassengers Forecast', fontsize=22)
+ax.set_ylabel('Monthly Passengers', fontsize=20)
+ax.set_xlabel('Year', fontsize=20)
+ax.legend(prop={'size': 15})
+ax.grid()
+```
+
diff --git a/neuralforecast/models.vanillatransformer.html.mdx b/neuralforecast/models.vanillatransformer.html.mdx
new file mode 100644
index 00000000..ee2becb2
--- /dev/null
+++ b/neuralforecast/models.vanillatransformer.html.mdx
@@ -0,0 +1,261 @@
+---
+output-file: models.vanillatransformer.html
+title: Vanilla Transformer
+---
+
+
+Vanilla Transformer, following implementation of the Informer paper,
+used as baseline.
+
+The architecture has three distinctive features: - Full-attention
+mechanism with O(L^2) time and memory complexity. - Classic
+encoder-decoder proposed by Vaswani et al. (2017) with a multi-head
+attention mechanism. - An MLP multi-step decoder that predicts long
+time-series sequences in a single forward operation rather than
+step-by-step.
+
+The Vanilla Transformer model utilizes a three-component approach to
+define its embedding: - It employs encoded autoregressive features
+obtained from a convolution network. - It uses window-relative
+positional embeddings derived from harmonic functions. - Absolute
+positional embeddings obtained from calendar features are utilized.
+
+**References** - [Haoyi Zhou, Shanghang Zhang, Jieqi Peng, Shuai
+Zhang, Jianxin Li, Hui Xiong, Wancai Zhang. “Informer: Beyond Efficient
+Transformer for Long Sequence Time-Series
+Forecasting”](https://arxiv.org/abs/2012.07436)
+
+
+
+Figure 1. Transformer
+Architecture.
+
+
+## 1. VanillaTransformer
+
+------------------------------------------------------------------------
+
+source
+
+### VanillaTransformer
+
+> ``` text
+> VanillaTransformer (h:int, input_size:int, stat_exog_list=None,
+> hist_exog_list=None, futr_exog_list=None,
+> exclude_insample_y=False,
+> decoder_input_size_multiplier:float=0.5,
+> hidden_size:int=128, dropout:float=0.05,
+> n_head:int=4, conv_hidden_size:int=32,
+> activation:str='gelu', encoder_layers:int=2,
+> decoder_layers:int=1, loss=MAE(), valid_loss=None,
+> max_steps:int=5000, learning_rate:float=0.0001,
+> num_lr_decays:int=-1,
+> early_stop_patience_steps:int=-1,
+> val_check_steps:int=100, batch_size:int=32,
+> valid_batch_size:Optional[int]=None,
+> windows_batch_size=1024,
+> inference_windows_batch_size:int=1024,
+> start_padding_enabled=False, step_size:int=1,
+> scaler_type:str='identity', random_seed:int=1,
+> drop_last_loader:bool=False,
+> alias:Optional[str]=None, optimizer=None,
+> optimizer_kwargs=None, lr_scheduler=None,
+> lr_scheduler_kwargs=None, dataloader_kwargs=None,
+> **trainer_kwargs)
+> ```
+
+\*VanillaTransformer
+
+Vanilla Transformer, following implementation of the Informer paper,
+used as baseline.
+
+The architecture has three distinctive features: - Full-attention
+mechanism with O(L^2) time and memory complexity. - An MLP multi-step
+decoder that predicts long time-series sequences in a single forward
+operation rather than step-by-step.
+
+The Vanilla Transformer model utilizes a three-component approach to
+define its embedding: - It employs encoded autoregressive features
+obtained from a convolution network. - It uses window-relative
+positional embeddings derived from harmonic functions. - Absolute
+positional embeddings obtained from calendar features are utilized.
+
+*Parameters:* `h`: int, forecast horizon. `input_size`: int,
+maximum sequence length for truncated train backpropagation.
+`stat_exog_list`: str list, static exogenous columns.
+`hist_exog_list`: str list, historic exogenous columns.
+`futr_exog_list`: str list, future exogenous columns.
+`exclude_insample_y`: bool=False, whether to exclude the target variable
+from the input. `decoder_input_size_multiplier`: float = 0.5, .
+`hidden_size`: int=128, units of embeddings and encoders. `dropout`:
+float (0, 1), dropout throughout Informer architecture. `n_head`:
+int=4, controls number of multi-head’s attention.
+`conv_hidden_size`: int=32, channels of the convolutional encoder.
+`activation`: str=`GELU`, activation from \[‘ReLU’, ‘Softplus’, ‘Tanh’,
+‘SELU’, ‘LeakyReLU’, ‘PReLU’, ‘Sigmoid’, ‘GELU’\]. `encoder_layers`:
+int=2, number of layers for the TCN encoder. `decoder_layers`:
+int=1, number of layers for the MLP decoder. `loss`: PyTorch module,
+instantiated train loss class from [losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`valid_loss`: PyTorch module=`loss`, instantiated valid loss class from
+[losses
+collection](https://nixtla.github.io/neuralforecast/losses.pytorch.html).
+`max_steps`: int=1000, maximum number of training steps.
+`learning_rate`: float=1e-3, Learning rate between (0, 1).
+`num_lr_decays`: int=-1, Number of learning rate decays, evenly
+distributed across max_steps. `early_stop_patience_steps`: int=-1,
+Number of validation iterations before early stopping.
+`val_check_steps`: int=100, Number of training steps between every
+validation loss check. `batch_size`: int=32, number of different
+series in each batch. `valid_batch_size`: int=None, number of
+different series in each validation and test batch, if None uses
+batch_size. `windows_batch_size`: int=1024, number of windows to
+sample in each training batch, default uses all.
+`inference_windows_batch_size`: int=1024, number of windows to sample in
+each inference batch. `start_padding_enabled`: bool=False, if True,
+the model will pad the time series with zeros at the beginning, by input
+size. `step_size`: int=1, step size between each window of temporal
+data. `scaler_type`: str=‘robust’, type of scaler for temporal
+inputs normalization see [temporal
+scalers](https://nixtla.github.io/neuralforecast/common.scalers.html).
+`random_seed`: int=1, random_seed for pytorch initializer and numpy
+generators. `drop_last_loader`: bool=False, if True
+`TimeSeriesDataLoader` drops last non-full batch. `alias`: str,
+optional, Custom name of the model. `optimizer`: Subclass of
+‘torch.optim.Optimizer’, optional, user specified optimizer instead of
+the default choice (Adam). `optimizer_kwargs`: dict, optional, list
+of parameters used by the user specified `optimizer`.
+`lr_scheduler`: Subclass of ‘torch.optim.lr_scheduler.LRScheduler’,
+optional, user specified lr_scheduler instead of the default choice
+(StepLR). `lr_scheduler_kwargs`: dict, optional, list of parameters
+used by the user specified `lr_scheduler`. `dataloader_kwargs`:
+dict, optional, list of parameters passed into the PyTorch Lightning
+dataloader by the `TimeSeriesDataLoader`. `**trainer_kwargs`: int,
+keyword trainer arguments inherited from [PyTorch Lighning’s
+trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+``` text
+*References*
+- [Haoyi Zhou, Shanghang Zhang, Jieqi Peng, Shuai Zhang, Jianxin Li, Hui Xiong, Wancai Zhang. "Informer: Beyond Efficient Transformer for Long Sequence Time-Series Forecasting"](https://arxiv.org/abs/2012.07436) *
+```
+
+------------------------------------------------------------------------
+
+### VanillaTransformer.fit
+
+> ``` text
+> VanillaTransformer.fit (dataset, val_size=0, test_size=0,
+> random_seed=None, distributed_config=None)
+> ```
+
+\*Fit.
+
+The `fit` method, optimizes the neural network’s weights using the
+initialization parameters (`learning_rate`, `windows_batch_size`, …) and
+the `loss` function as defined during the initialization. Within `fit`
+we use a PyTorch Lightning `Trainer` that inherits the initialization’s
+`self.trainer_kwargs`, to customize its inputs, see [PL’s trainer
+arguments](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+The method is designed to be compatible with SKLearn-like classes and in
+particular to be compatible with the StatsForecast library.
+
+By default the `model` is not saving training checkpoints to protect
+disk memory, to get them change `enable_checkpointing=True` in
+`__init__`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`val_size`: int, validation size for temporal cross-validation.
+`random_seed`: int=None, random_seed for pytorch initializer and numpy
+generators, overwrites model.\_\_init\_\_’s. `test_size`: int, test
+size for temporal cross-validation. \*
+
+------------------------------------------------------------------------
+
+### VanillaTransformer.predict
+
+> ``` text
+> VanillaTransformer.predict (dataset, test_size=None, step_size=1,
+> random_seed=None, quantiles=None,
+> **data_module_kwargs)
+> ```
+
+\*Predict.
+
+Neural network prediction with PL’s `Trainer` execution of
+`predict_step`.
+
+**Parameters:** `dataset`: NeuralForecast’s
+[`TimeSeriesDataset`](https://nixtlaverse.nixtla.io/neuralforecast/tsdataset.html#timeseriesdataset),
+see
+[documentation](https://nixtla.github.io/neuralforecast/tsdataset.html).
+`test_size`: int=None, test size for temporal cross-validation.
+`step_size`: int=1, Step size between each window. `random_seed`:
+int=None, random_seed for pytorch initializer and numpy generators,
+overwrites model.\_\_init\_\_’s. `quantiles`: list of floats,
+optional (default=None), target quantiles to predict.
+`**data_module_kwargs`: PL’s TimeSeriesDataModule args, see
+[documentation](https://pytorch-lightning.readthedocs.io/en/1.6.1/extensions/datamodules.html#using-a-datamodule).\*
+
+## Usage Example
+
+
+```python
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from neuralforecast import NeuralForecast
+from neuralforecast.models import VanillaTransformer
+from neuralforecast.utils import AirPassengersPanel, AirPassengersStatic
+
+Y_train_df = AirPassengersPanel[AirPassengersPanel.ds=AirPassengersPanel['ds'].values[-12]].reset_index(drop=True) # 12 test
+
+model = VanillaTransformer(h=12,
+ input_size=24,
+ hidden_size=16,
+ conv_hidden_size=32,
+ n_head=2,
+ loss=MAE(),
+ scaler_type='robust',
+ learning_rate=1e-3,
+ max_steps=500,
+ val_check_steps=50,
+ early_stop_patience_steps=2)
+
+nf = NeuralForecast(
+ models=[model],
+ freq='ME'
+)
+nf.fit(df=Y_train_df, static_df=AirPassengersStatic, val_size=12)
+forecasts = nf.predict(futr_df=Y_test_df)
+
+Y_hat_df = forecasts.reset_index(drop=False).drop(columns=['unique_id','ds'])
+plot_df = pd.concat([Y_test_df, Y_hat_df], axis=1)
+plot_df = pd.concat([Y_train_df, plot_df])
+
+if model.loss.is_distribution_output:
+ plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+ plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+ plt.plot(plot_df['ds'], plot_df['VanillaTransformer-median'], c='blue', label='median')
+ plt.fill_between(x=plot_df['ds'][-12:],
+ y1=plot_df['VanillaTransformer-lo-90'][-12:].values,
+ y2=plot_df['VanillaTransformer-hi-90'][-12:].values,
+ alpha=0.4, label='level 90')
+ plt.grid()
+ plt.legend()
+ plt.plot()
+else:
+ plot_df = plot_df[plot_df.unique_id=='Airline1'].drop('unique_id', axis=1)
+ plt.plot(plot_df['ds'], plot_df['y'], c='black', label='True')
+ plt.plot(plot_df['ds'], plot_df['VanillaTransformer'], c='blue', label='Forecast')
+ plt.legend()
+ plt.grid()
+```
+
diff --git a/neuralforecast/models.xlstm.mdx b/neuralforecast/models.xlstm.mdx
new file mode 100644
index 00000000..41ea9e64
--- /dev/null
+++ b/neuralforecast/models.xlstm.mdx
@@ -0,0 +1,278 @@
+---
+output-file: models.xlstm
+title: xLSTM
+---
+
+
+
+
+# module `neuralforecast.models.xlstm`
+
+
+
+
+**Global Variables**
+---------------
+- **IS_XLSTM_INSTALLED**
+
+
+---
+
+
+
+## class `xLSTM`
+xLSTM
+
+xLSTM encoder, with MLP decoder.
+
+
+
+**Args:**
+
+ - `h` (int): forecast horizon.
+ - `input_size` (int): considered autorregresive inputs (lags), y=[1,2,3,4] input_size=2 -> lags=[1,2].
+ - `encoder_n_blocks` (int): number of blocks for the xLSTM.
+ - `encoder_hidden_size` (int): units for the xLSTM's hidden state size.
+ - `encoder_bias` (bool): whether or not to use biases within xLSTM blocks.
+ - `encoder_dropout` (float): dropout regularization applied within xLSTM blocks.
+ - `decoder_hidden_size` (int): size of hidden layer for the MLP decoder.
+ - `decoder_layers` (int): number of layers for the MLP decoder.
+ - `decoder_dropout` (float): dropout regularization applied within the MLP decoder.
+ - `decoder_activation` (str): activation function for the MLP decoder, see [activations collection](https://docs.pytorch.org/docs/stable/nn.html#non-linear-activations-weighted-sum-nonlinearity).
+ - `backbone` (str): backbone for the xLSTM, either 'sLSTM' or 'mLSTM'.
+ - `futr_exog_list` (List[str]): future exogenous columns.
+ - `hist_exog_list` (list): historic exogenous columns.
+ - `stat_exog_list` (list): static exogenous columns.
+ - `exclude_insample_y` (bool): whether to exclude the target variable from the input.
+ - `recurrent` (bool): whether to produce forecasts recursively (True) or direct (False).
+ - `loss` (nn.Module): instantiated train loss class from [losses collection](./losses.pytorch).
+ - `valid_loss` (nn.Module): instantiated valid loss class from [losses collection](./losses.pytorch).
+ - `max_steps` (int): maximum number of training steps.
+ - `learning_rate` (float): Learning rate between (0, 1).
+ - `num_lr_decays` (int): Number of learning rate decays, evenly distributed across max_steps.
+ - `early_stop_patience_steps` (int): Number of validation iterations before early stopping.
+ - `val_check_steps` (int): Number of training steps between every validation loss check.
+ - `batch_size` (int): number of differentseries in each batch.
+ - `valid_batch_size` (int): number of different series in each validation and test batch.
+ - `windows_batch_size` (int): number of windows to sample in each training batch, default uses all.
+ - `inference_windows_batch_size` (int): number of windows to sample in each inference batch, -1 uses all.
+ - `start_padding_enabled` (bool): if True, the model will pad the time series with zeros at the beginning, by input size.
+ - `training_data_availability_threshold` (Union[float, List[float]]): minimum fraction of valid data points required for training windows. Single float applies to both insample and outsample; list of two floats specifies [insample_fraction, outsample_fraction]. Default 0.0 allows windows with only 1 valid data point (current behavior).
+ - `step_size` (int): step size between each window of temporal data.
+ - `scaler_type` (str): type of scaler for temporal inputs normalization see [temporal scalers](https://github.com/Nixtla/neuralforecast/blob/main/neuralforecast/common/_scalers.py).
+ - `random_seed` (int): random_seed for pytorch initializer and numpy generators.
+ - `drop_last_loader` (bool): if True `TimeSeriesDataLoader` drops last non-full batch.
+ - `alias` (str): optional, Custom name of the model.
+ - `optimizer` (Subclass of 'torch.optim.Optimizer'): optional, user specified optimizer instead of the default choice (Adam).
+ - `optimizer_kwargs` (dict): optional, list of parameters used by the user specified `optimizer`.
+ - `lr_scheduler` (Subclass of 'torch.optim.lr_scheduler.LRScheduler'): optional, user specified lr_scheduler instead of the default choice (StepLR).
+ - `lr_scheduler_kwargs` (dict): optional, list of parameters used by the user specified `lr_scheduler`.
+ - `dataloader_kwargs` (dict): optional, list of parameters passed into the PyTorch Lightning dataloader by the `TimeSeriesDataLoader`.
+ - `**trainer_kwargs (int)`: keyword trainer arguments inherited from [PyTorch Lighning's trainer](https://pytorch-lightning.readthedocs.io/en/stable/api/pytorch_lightning.trainer.trainer.Trainer.html?highlight=trainer).
+
+References:
+ - [Maximilian Beck, Korbinian Pöppel, Markus Spanring, Andreas Auer, Oleksandra Prudnikova, Michael Kopp, Günter Klambauer, Johannes Brandstetter, Sepp Hochreiter (2024). "xLSTM: Extended Long Short-Term Memory"](https://arxiv.org/abs/2405.04517)
+
+
+
+### method `__init__`
+
+```python
+__init__(
+ h: int,
+ input_size: int = -1,
+ inference_input_size: Optional[int] = None,
+ h_train: int = 1,
+ encoder_n_blocks: int = 2,
+ encoder_hidden_size: int = 128,
+ encoder_bias: bool = True,
+ encoder_dropout: float = 0.1,
+ decoder_hidden_size: int = 128,
+ decoder_layers: int = 1,
+ decoder_dropout: float = 0.0,
+ decoder_activation: str = 'GELU',
+ backbone: str = 'mLSTM',
+ futr_exog_list=None,
+ hist_exog_list=None,
+ stat_exog_list=None,
+ exclude_insample_y=False,
+ recurrent=False,
+ loss=MAE(),
+ valid_loss=None,
+ max_steps: int = 1000,
+ learning_rate: float = 0.001,
+ num_lr_decays: int = -1,
+ early_stop_patience_steps: int = -1,
+ val_check_steps: int = 100,
+ batch_size=32,
+ valid_batch_size: Optional[int] = None,
+ windows_batch_size=128,
+ inference_windows_batch_size=1024,
+ start_padding_enabled=False,
+ training_data_availability_threshold=0.0,
+ step_size: int = 1,
+ scaler_type: str = 'robust',
+ random_seed=1,
+ drop_last_loader=False,
+ alias: Optional[str] = None,
+ optimizer=None,
+ optimizer_kwargs=None,
+ lr_scheduler=None,
+ lr_scheduler_kwargs=None,
+ dataloader_kwargs=None,
+ **trainer_kwargs
+)
+```
+
+
+
+
+
+
+---
+
+#### property automatic_optimization
+
+If set to ``False`` you are responsible for calling ``.backward()``, ``.step()``, ``.zero_grad()``.
+
+---
+
+#### property current_epoch
+
+The current epoch in the ``Trainer``, or 0 if not attached.
+
+---
+
+#### property device
+
+
+
+
+
+---
+
+#### property device_mesh
+
+Strategies like ``ModelParallelStrategy`` will create a device mesh that can be accessed in the :meth:`~pytorch_lightning.core.hooks.ModelHooks.configure_model` hook to parallelize the LightningModule.
+
+---
+
+#### property dtype
+
+
+
+
+
+---
+
+#### property example_input_array
+
+The example input array is a specification of what the module can consume in the :meth:`forward` method. The return type is interpreted as follows:
+
+
+- Single tensor: It is assumed the model takes a single argument, i.e., ``model.forward(model.example_input_array)``
+- Tuple: The input array should be interpreted as a sequence of positional arguments, i.e., ``model.forward(*model.example_input_array)``
+- Dict: The input array represents named keyword arguments, i.e., ``model.forward(**model.example_input_array)``
+
+---
+
+#### property fabric
+
+
+
+
+
+---
+
+#### property global_rank
+
+The index of the current process across all nodes and devices.
+
+---
+
+#### property global_step
+
+Total training batches seen across all epochs.
+
+If no Trainer is attached, this property is 0.
+
+---
+
+#### property hparams
+
+The collection of hyperparameters saved with :meth:`save_hyperparameters`. It is mutable by the user. For the frozen set of initial hyperparameters, use :attr:`hparams_initial`.
+
+
+
+**Returns:**
+ Mutable hyperparameters dictionary
+
+---
+
+#### property hparams_initial
+
+The collection of hyperparameters saved with :meth:`save_hyperparameters`. These contents are read-only. Manual updates to the saved hyperparameters can instead be performed through :attr:`hparams`.
+
+
+
+**Returns:**
+
+ - `AttributeDict`: immutable initial hyperparameters
+
+---
+
+#### property local_rank
+
+The index of the current process within a single node.
+
+---
+
+#### property logger
+
+Reference to the logger object in the Trainer.
+
+---
+
+#### property loggers
+
+Reference to the list of loggers in the Trainer.
+
+---
+
+#### property on_gpu
+
+Returns ``True`` if this model is currently located on a GPU.
+
+Useful to set flags around the LightningModule for different CPU vs GPU behavior.
+
+---
+
+#### property strict_loading
+
+Determines how Lightning loads this model using `.load_state_dict(..., strict=model.strict_loading)`.
+
+---
+
+#### property trainer
+
+
+
+
+
+
+
+---
+
+
+
+### method `forward`
+
+```python
+forward(windows_batch)
+```
+
+
+
+
+
+
diff --git a/neuralforecast/tsdataset.html.mdx b/neuralforecast/tsdataset.html.mdx
new file mode 100644
index 00000000..9f201e24
--- /dev/null
+++ b/neuralforecast/tsdataset.html.mdx
@@ -0,0 +1,200 @@
+---
+description: Torch Dataset for Time Series
+output-file: tsdataset.html
+title: PyTorch Dataset/Loader
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### TimeSeriesLoader
+
+> ``` text
+> TimeSeriesLoader (dataset, **kwargs)
+> ```
+
+\*TimeSeriesLoader DataLoader. [Source
+code](https://github.com/Nixtla/neuralforecast/blob/main/neuralforecast/tsdataset.py).
+
+Small change to PyTorch’s Data loader. Combines a dataset and a sampler,
+and provides an iterable over the given dataset.
+
+The class `~torch.utils.data.DataLoader` supports both map-style and
+iterable-style datasets with single- or multi-process loading,
+customizing loading order and optional automatic batching (collation)
+and memory pinning.
+
+**Parameters:** `batch_size`: (int, optional): how many samples per
+batch to load (default: 1). `shuffle`: (bool, optional): set to
+`True` to have the data reshuffled at every epoch (default:
+`False`). `sampler`: (Sampler or Iterable, optional): defines the
+strategy to draw samples from the dataset. Can be any `Iterable`
+with `__len__` implemented. If specified, `shuffle` must not be
+specified. \*
+
+------------------------------------------------------------------------
+
+source
+
+### BaseTimeSeriesDataset
+
+> ``` text
+> BaseTimeSeriesDataset (temporal_cols, max_size:int, min_size:int,
+> y_idx:int, static=None, static_cols=None)
+> ```
+
+\*An abstract class representing a :class:`Dataset`.
+
+All datasets that represent a map from keys to data samples should
+subclass it. All subclasses should overwrite :meth:`__getitem__`,
+supporting fetching a data sample for a given key. Subclasses could also
+optionally overwrite :meth:`__len__`, which is expected to return the
+size of the dataset by many :class:`~torch.utils.data.Sampler`
+implementations and the default options of
+:class:`~torch.utils.data.DataLoader`. Subclasses could also optionally
+implement :meth:`__getitems__`, for speedup batched samples loading.
+This method accepts list of indices of samples of batch and returns list
+of samples.
+
+.. note:: :class:`~torch.utils.data.DataLoader` by default constructs an
+index sampler that yields integral indices. To make it work with a
+map-style dataset with non-integral indices/keys, a custom sampler must
+be provided.\*
+
+------------------------------------------------------------------------
+
+source
+
+### LocalFilesTimeSeriesDataset
+
+> ``` text
+> LocalFilesTimeSeriesDataset (files_ds:List[str], temporal_cols,
+> id_col:str, time_col:str, target_col:str,
+> last_times, indices, max_size:int,
+> min_size:int, y_idx:int, static=None,
+> static_cols=None)
+> ```
+
+\*An abstract class representing a :class:`Dataset`.
+
+All datasets that represent a map from keys to data samples should
+subclass it. All subclasses should overwrite :meth:`__getitem__`,
+supporting fetching a data sample for a given key. Subclasses could also
+optionally overwrite :meth:`__len__`, which is expected to return the
+size of the dataset by many :class:`~torch.utils.data.Sampler`
+implementations and the default options of
+:class:`~torch.utils.data.DataLoader`. Subclasses could also optionally
+implement :meth:`__getitems__`, for speedup batched samples loading.
+This method accepts list of indices of samples of batch and returns list
+of samples.
+
+.. note:: :class:`~torch.utils.data.DataLoader` by default constructs an
+index sampler that yields integral indices. To make it work with a
+map-style dataset with non-integral indices/keys, a custom sampler must
+be provided.\*
+
+------------------------------------------------------------------------
+
+source
+
+### TimeSeriesDataset
+
+> ``` text
+> TimeSeriesDataset (temporal, temporal_cols, indptr, y_idx:int,
+> static=None, static_cols=None)
+> ```
+
+\*An abstract class representing a :class:`Dataset`.
+
+All datasets that represent a map from keys to data samples should
+subclass it. All subclasses should overwrite :meth:`__getitem__`,
+supporting fetching a data sample for a given key. Subclasses could also
+optionally overwrite :meth:`__len__`, which is expected to return the
+size of the dataset by many :class:`~torch.utils.data.Sampler`
+implementations and the default options of
+:class:`~torch.utils.data.DataLoader`. Subclasses could also optionally
+implement :meth:`__getitems__`, for speedup batched samples loading.
+This method accepts list of indices of samples of batch and returns list
+of samples.
+
+.. note:: :class:`~torch.utils.data.DataLoader` by default constructs an
+index sampler that yields integral indices. To make it work with a
+map-style dataset with non-integral indices/keys, a custom sampler must
+be provided.\*
+
+------------------------------------------------------------------------
+
+source
+
+### TimeSeriesDataModule
+
+> ``` text
+> TimeSeriesDataModule (dataset:__main__.BaseTimeSeriesDataset,
+> batch_size=32, valid_batch_size=1024,
+> drop_last=False, shuffle_train=True,
+> **dataloaders_kwargs)
+> ```
+
+\*A DataModule standardizes the training, val, test splits, data
+preparation and transforms. The main advantage is consistent data
+splits, data preparation and transforms across models.
+
+Example::
+
+``` text
+import lightning.pytorch as L
+import torch.utils.data as data
+from pytorch_lightning.demos.boring_classes import RandomDataset
+
+class MyDataModule(L.LightningDataModule):
+ def prepare_data(self):
+ # download, IO, etc. Useful with shared filesystems
+ # only called on 1 GPU/TPU in distributed
+ ...
+
+ def setup(self, stage):
+ # make assignments here (val/train/test split)
+ # called on every process in DDP
+ dataset = RandomDataset(1, 100)
+ self.train, self.val, self.test = data.random_split(
+ dataset, [80, 10, 10], generator=torch.Generator().manual_seed(42)
+ )
+
+ def train_dataloader(self):
+ return data.DataLoader(self.train)
+
+ def val_dataloader(self):
+ return data.DataLoader(self.val)
+
+ def test_dataloader(self):
+ return data.DataLoader(self.test)
+
+ def on_exception(self, exception):
+ # clean up state after the trainer faced an exception
+ ...
+
+ def teardown(self):
+ # clean up state after the trainer stops, delete files...
+ # called on every process in DDP
+ ...*
+```
+
+
+```python
+# To test correct future_df wrangling of the `update_df` method
+# We are checking that we are able to recover the AirPassengers dataset
+# using the dataframe or splitting it into parts and initializing.
+```
+
diff --git a/neuralforecast/utils.html.mdx b/neuralforecast/utils.html.mdx
new file mode 100644
index 00000000..c347a916
--- /dev/null
+++ b/neuralforecast/utils.html.mdx
@@ -0,0 +1,461 @@
+---
+description: >-
+ The `core.NeuralForecast` class allows you to efficiently fit multiple
+ `NeuralForecast` models for large sets of time series. It operates with pandas
+ DataFrame `df` that identifies individual series and datestamps with the
+ `unique_id` and `ds` columns, and the `y` column denotes the target time
+ series variable. To assist development, we declare useful datasets that we use
+ throughout all `NeuralForecast`'s unit tests.
+output-file: utils.html
+title: Example Data
+---
+
+
+# 1. Synthetic Panel Data
+
+------------------------------------------------------------------------
+
+source
+
+### generate_series
+
+> ``` text
+> generate_series (n_series:int, freq:str='D', min_length:int=50,
+> max_length:int=500, n_temporal_features:int=0,
+> n_static_features:int=0, equal_ends:bool=False,
+> seed:int=0)
+> ```
+
+\*Generate Synthetic Panel Series.
+
+Generates `n_series` of frequency `freq` of different lengths in the
+interval \[`min_length`, `max_length`\]. If `n_temporal_features > 0`,
+then each serie gets temporal features with random values. If
+`n_static_features > 0`, then a static dataframe is returned along the
+temporal dataframe. If `equal_ends == True` then all series end at the
+same date.
+
+**Parameters:** `n_series`: int, number of series for synthetic
+panel. `min_length`: int, minimal length of synthetic panel’s
+series. `max_length`: int, minimal length of synthetic panel’s
+series. `n_temporal_features`: int, default=0, number of temporal
+exogenous variables for synthetic panel’s series.
+`n_static_features`: int, default=0, number of static exogenous
+variables for synthetic panel’s series. `equal_ends`: bool, if True,
+series finish in the same date stamp `ds`. `freq`: str, frequency of
+the data, [panda’s available
+frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+
+**Returns:** `freq`: pandas.DataFrame, synthetic panel with columns
+\[`unique_id`, `ds`, `y`\] and exogenous.\*
+
+
+```python
+synthetic_panel = generate_series(n_series=2)
+synthetic_panel.groupby('unique_id').head(4)
+```
+
+
+```python
+temporal_df, static_df = generate_series(n_series=1000, n_static_features=2,
+ n_temporal_features=4, equal_ends=False)
+static_df.head(2)
+```
+
+# 2. AirPassengers Data
+
+The classic Box & Jenkins airline data. Monthly totals of international
+airline passengers, 1949 to 1960.
+
+It has been used as a reference on several forecasting libraries, since
+it is a series that shows clear trends and seasonalities it offers a
+nice opportunity to quickly showcase a model’s predictions performance.
+
+
+```python
+AirPassengersDF.head(12)
+```
+
+
+```python
+#We are going to plot the ARIMA predictions, and the prediction intervals.
+fig, ax = plt.subplots(1, 1, figsize = (20, 7))
+plot_df = AirPassengersDF.set_index('ds')
+
+plot_df[['y']].plot(ax=ax, linewidth=2)
+ax.set_title('AirPassengers Forecast', fontsize=22)
+ax.set_ylabel('Monthly Passengers', fontsize=20)
+ax.set_xlabel('Timestamp [t]', fontsize=20)
+ax.legend(prop={'size': 15})
+ax.grid()
+```
+
+
+```python
+import numpy as np
+import pandas as pd
+```
+
+
+```python
+n_static_features = 3
+n_series = 5
+
+static_features = np.random.uniform(low=0.0, high=1.0,
+ size=(n_series, n_static_features))
+static_df = pd.DataFrame.from_records(static_features,
+ columns = [f'static_{i}'for i in range(n_static_features)])
+static_df['unique_id'] = np.arange(n_series)
+```
+
+
+```python
+static_df
+```
+
+# 3. Panel AirPassengers Data
+
+Extension to classic Box & Jenkins airline data. Monthly totals of
+international airline passengers, 1949 to 1960.
+
+It includes two series with static, temporal and future exogenous
+variables, that can help to explore the performance of models like
+[`NBEATSx`](https://nixtlaverse.nixtla.io/neuralforecast/models.nbeatsx.html#nbeatsx)
+and
+[`TFT`](https://nixtlaverse.nixtla.io/neuralforecast/models.tft.html#tft).
+
+
+```python
+fig, ax = plt.subplots(1, 1, figsize = (20, 7))
+plot_df = AirPassengersPanel.set_index('ds')
+
+plot_df.groupby('unique_id')['y'].plot(legend=True)
+ax.set_title('AirPassengers Panel Data', fontsize=22)
+ax.set_ylabel('Monthly Passengers', fontsize=20)
+ax.set_xlabel('Timestamp [t]', fontsize=20)
+ax.legend(title='unique_id', prop={'size': 15})
+ax.grid()
+```
+
+
+```python
+fig, ax = plt.subplots(1, 1, figsize = (20, 7))
+plot_df = AirPassengersPanel[AirPassengersPanel.unique_id=='Airline1'].set_index('ds')
+
+plot_df[['y', 'trend', 'y_[lag12]']].plot(ax=ax, linewidth=2)
+ax.set_title('Box-Cox AirPassengers Data', fontsize=22)
+ax.set_ylabel('Monthly Passengers', fontsize=20)
+ax.set_xlabel('Timestamp [t]', fontsize=20)
+ax.legend(prop={'size': 15})
+ax.grid()
+```
+
+# 4. Time Features
+
+We have developed a utility that generates normalized calendar features
+for use as absolute positional embeddings in Transformer-based models.
+These embeddings capture seasonal patterns in time series data and can
+be easily incorporated into the model architecture. Additionally, the
+features can be used as exogenous variables in other models to inform
+them of calendar patterns in the data.
+
+**References** - [Haoyi Zhou, Shanghang Zhang, Jieqi Peng, Shuai
+Zhang, Jianxin Li, Hui Xiong, Wancai Zhang. “Informer: Beyond Efficient
+Transformer for Long Sequence Time-Series
+Forecasting”](https://arxiv.org/abs/2012.07436)
+
+------------------------------------------------------------------------
+
+source
+
+### augment_calendar_df
+
+> ``` text
+> augment_calendar_df (df, freq='H')
+> ```
+
+*\> * Q - \[month\] \> \* M - \[month\] \> \* W - \[Day of month, week
+of year\] \> \* D - \[Day of week, day of month, day of year\] \> \* B -
+\[Day of week, day of month, day of year\] \> \* H - \[Hour of day, day
+of week, day of month, day of year\] \> \* T - \[Minute of hour\*, hour
+of day, day of week, day of month, day of year\] \> \* S - \[Second of
+minute, minute of hour, hour of day, day of week, day of month, day of
+year\] *minute returns a number from 0-3 corresponding to the 15 minute
+period it falls into.*
+
+------------------------------------------------------------------------
+
+source
+
+### time_features_from_frequency_str
+
+> ``` text
+> time_features_from_frequency_str (freq_str:str)
+> ```
+
+*Returns a list of time features that will be appropriate for the given
+frequency string. Parameters ———- freq_str Frequency string of the form
+\[multiple\]\[granularity\] such as “12H”, “5min”, “1D” etc.*
+
+------------------------------------------------------------------------
+
+source
+
+### WeekOfYear
+
+> ``` text
+> WeekOfYear ()
+> ```
+
+*Week of year encoded as value between \[-0.5, 0.5\]*
+
+------------------------------------------------------------------------
+
+source
+
+### MonthOfYear
+
+> ``` text
+> MonthOfYear ()
+> ```
+
+*Month of year encoded as value between \[-0.5, 0.5\]*
+
+------------------------------------------------------------------------
+
+source
+
+### DayOfYear
+
+> ``` text
+> DayOfYear ()
+> ```
+
+*Day of year encoded as value between \[-0.5, 0.5\]*
+
+------------------------------------------------------------------------
+
+source
+
+### DayOfMonth
+
+> ``` text
+> DayOfMonth ()
+> ```
+
+*Day of month encoded as value between \[-0.5, 0.5\]*
+
+------------------------------------------------------------------------
+
+source
+
+### DayOfWeek
+
+> ``` text
+> DayOfWeek ()
+> ```
+
+*Hour of day encoded as value between \[-0.5, 0.5\]*
+
+------------------------------------------------------------------------
+
+source
+
+### HourOfDay
+
+> ``` text
+> HourOfDay ()
+> ```
+
+*Hour of day encoded as value between \[-0.5, 0.5\]*
+
+------------------------------------------------------------------------
+
+source
+
+### MinuteOfHour
+
+> ``` text
+> MinuteOfHour ()
+> ```
+
+*Minute of hour encoded as value between \[-0.5, 0.5\]*
+
+------------------------------------------------------------------------
+
+source
+
+### SecondOfMinute
+
+> ``` text
+> SecondOfMinute ()
+> ```
+
+*Minute of hour encoded as value between \[-0.5, 0.5\]*
+
+------------------------------------------------------------------------
+
+source
+
+### TimeFeature
+
+> ``` text
+> TimeFeature ()
+> ```
+
+*Initialize self. See help(type(self)) for accurate signature.*
+
+
+```python
+AirPassengerPanelCalendar, calendar_cols = augment_calendar_df(df=AirPassengersPanel, freq='M')
+AirPassengerPanelCalendar.head()
+```
+
+
+```python
+plot_df = AirPassengerPanelCalendar[AirPassengerPanelCalendar.unique_id=='Airline1'].set_index('ds')
+plt.plot(plot_df['month'])
+plt.grid()
+plt.xlabel('Datestamp')
+plt.ylabel('Normalized Month')
+plt.show()
+```
+
+------------------------------------------------------------------------
+
+source
+
+### get_indexer_raise_missing
+
+> ``` text
+> get_indexer_raise_missing (idx:pandas.core.indexes.base.Index,
+> vals:List[str])
+> ```
+
+# 5. Prediction Intervals
+
+------------------------------------------------------------------------
+
+source
+
+### PredictionIntervals
+
+> ``` text
+> PredictionIntervals (n_windows:int=2,
+> method:str='conformal_distribution')
+> ```
+
+*Class for storing prediction intervals metadata information.*
+
+------------------------------------------------------------------------
+
+source
+
+### add_conformal_distribution_intervals
+
+> ``` text
+> add_conformal_distribution_intervals (model_fcsts: infunctionarray>, cs_df:~DFType,
+> model:str, cs_n_windows:int,
+> n_series:int, horizon:int, level:Op
+> tional[List[Union[int,float]]]=None
+> , quantiles:Optional[List[float]]=N
+> one)
+> ```
+
+*Adds conformal intervals to a `fcst_df` based on conformal scores
+`cs_df`. `level` should be already sorted. This strategy creates
+forecasts paths based on errors and calculate quantiles using those
+paths.*
+
+------------------------------------------------------------------------
+
+source
+
+### add_conformal_error_intervals
+
+> ``` text
+> add_conformal_error_intervals (model_fcsts:,
+> cs_df:~DFType, model:str,
+> cs_n_windows:int, n_series:int,
+> horizon:int, level:Optional[List[Union[int
+> ,float]]]=None,
+> quantiles:Optional[List[float]]=None)
+> ```
+
+*Adds conformal intervals to a `fcst_df` based on conformal scores
+`cs_df`. `level` should be already sorted. This startegy creates
+prediction intervals based on the absolute errors.*
+
+------------------------------------------------------------------------
+
+source
+
+### get_prediction_interval_method
+
+> ``` text
+> get_prediction_interval_method (method:str)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### quantiles_to_level
+
+> ``` text
+> quantiles_to_level (quantiles:List[float])
+> ```
+
+*Converts a list of quantiles to a list of levels.*
+
+------------------------------------------------------------------------
+
+source
+
+### level_to_quantiles
+
+> ``` text
+> level_to_quantiles (level:List[Union[int,float]])
+> ```
+
+*Converts a list of levels to a list of quantiles.*
+
diff --git a/nixtla/.nojekyll b/nixtla/.nojekyll
new file mode 100644
index 00000000..e69de29b
diff --git a/nixtla/dark.png b/nixtla/dark.png
new file mode 100644
index 00000000..4142a0bb
Binary files /dev/null and b/nixtla/dark.png differ
diff --git a/nixtla/docs/capabilities/capabilities.html.mdx b/nixtla/docs/capabilities/capabilities.html.mdx
new file mode 100644
index 00000000..0eee3a72
--- /dev/null
+++ b/nixtla/docs/capabilities/capabilities.html.mdx
@@ -0,0 +1,8 @@
+---
+output-file: capabilities.html
+title: Capabilities
+---
+
+
+This section offers an overview of capabilities of TimeGPT
+
diff --git a/nixtla/docs/capabilities/forecast/01_quickstart_files/figure-markdown_strict/cell-5-output-2.png b/nixtla/docs/capabilities/forecast/01_quickstart_files/figure-markdown_strict/cell-5-output-2.png
new file mode 100644
index 00000000..350b8f8a
Binary files /dev/null and b/nixtla/docs/capabilities/forecast/01_quickstart_files/figure-markdown_strict/cell-5-output-2.png differ
diff --git a/nixtla/docs/capabilities/forecast/10_prediction_intervals_files/figure-markdown_strict/cell-5-output-2.png b/nixtla/docs/capabilities/forecast/10_prediction_intervals_files/figure-markdown_strict/cell-5-output-2.png
new file mode 100644
index 00000000..1a5a5ca7
Binary files /dev/null and b/nixtla/docs/capabilities/forecast/10_prediction_intervals_files/figure-markdown_strict/cell-5-output-2.png differ
diff --git a/nixtla/docs/capabilities/forecast/11_irregular_timestamps_files/figure-markdown_strict/cell-13-output-1.png b/nixtla/docs/capabilities/forecast/11_irregular_timestamps_files/figure-markdown_strict/cell-13-output-1.png
new file mode 100644
index 00000000..92966d4a
Binary files /dev/null and b/nixtla/docs/capabilities/forecast/11_irregular_timestamps_files/figure-markdown_strict/cell-13-output-1.png differ
diff --git a/nixtla/docs/capabilities/forecast/11_irregular_timestamps_files/figure-markdown_strict/cell-6-output-1.png b/nixtla/docs/capabilities/forecast/11_irregular_timestamps_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..375fff8a
Binary files /dev/null and b/nixtla/docs/capabilities/forecast/11_irregular_timestamps_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/nixtla/docs/capabilities/forecast/11_irregular_timestamps_files/figure-markdown_strict/cell-8-output-1.png b/nixtla/docs/capabilities/forecast/11_irregular_timestamps_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..ccd66b94
Binary files /dev/null and b/nixtla/docs/capabilities/forecast/11_irregular_timestamps_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/nixtla/docs/capabilities/forecast/categorical_variables.html.mdx b/nixtla/docs/capabilities/forecast/categorical_variables.html.mdx
new file mode 100644
index 00000000..625e36e1
--- /dev/null
+++ b/nixtla/docs/capabilities/forecast/categorical_variables.html.mdx
@@ -0,0 +1,89 @@
+---
+output-file: categorical_variables.html
+title: Add categorical variables
+---
+
+
+TimeGPT supports categorical variables and we can create them using
+[`SpecialDates`](https://Nixtla.github.io/nixtla/src/date_features.html#specialdates).
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/capabilities/forecast/04_categorical_variables.ipynb)
+
+```python
+import pandas as pd
+import datetime
+from nixtla import NixtlaClient
+from nixtla.date_features import SpecialDates
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+```python
+# Read the data
+df = pd.read_csv("https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/air_passengers.csv")
+
+# Create categorical variables to label Christmas and summer vacations
+categories_dates = SpecialDates(
+ special_dates={
+ 'christmas_vacations': [datetime.date(year, 12, 1) for year in range(1949, 1960 + 1)],
+ 'summer_vacations': [datetime.date(year, month, 1) for year in range(1949, 1960 + 1) for month in (6, 7)]
+ }
+)
+
+dates = pd.date_range('1949-01-01', '1960-12-01', freq='MS')
+
+categories_df = categories_dates(dates).reset_index(drop=True)
+
+# Merge with the dataset
+cat_df = pd.concat([df, categories_df], axis=1)
+
+# Forecast
+forecast_df = nixtla_client.forecast(
+ df=cat_df,
+ h=24,
+ target_col='value',
+ time_col='timestamp'
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+WARNING:nixtla.nixtla_client:You did not provide X_df. Exogenous variables in df are ignored. To surpress this warning, please add X_df with exogenous variables: christmas_vacations, summer_vacations
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon. This may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+For a detailed guide on using categorical variables for forecasting,
+read our in-depth tutorial on [Categorical
+variables](https://docs.nixtla.io/docs/tutorials-categorical_variables).
+
diff --git a/nixtla/docs/capabilities/forecast/cross_validation.html.mdx b/nixtla/docs/capabilities/forecast/cross_validation.html.mdx
new file mode 100644
index 00000000..319984fb
--- /dev/null
+++ b/nixtla/docs/capabilities/forecast/cross_validation.html.mdx
@@ -0,0 +1,69 @@
+---
+output-file: cross_validation.html
+title: Cross validation
+---
+
+
+We can perform cross-validation by simply using the `cross-validation`
+method. Specify the number of windows using the `n_windows` argument.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/capabilities/forecast/09_cross_validation.ipynb)
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+```python
+# Read the data
+df = pd.read_csv("https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/air_passengers.csv")
+
+# Cross-validation using two windows
+forecast_cv_df = nixtla_client.cross_validation(
+ df=df,
+ h=12,
+ n_windows=2,
+ time_col='timestamp',
+ target_col="value",
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Cross Validation Endpoint...
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+For more details, check out our [cross-validation
+tutorial](https://docs.nixtla.io/docs/tutorials-cross_validation).
+
diff --git a/nixtla/docs/capabilities/forecast/custom_loss_function.html.mdx b/nixtla/docs/capabilities/forecast/custom_loss_function.html.mdx
new file mode 100644
index 00000000..2d02c243
--- /dev/null
+++ b/nixtla/docs/capabilities/forecast/custom_loss_function.html.mdx
@@ -0,0 +1,84 @@
+---
+output-file: custom_loss_function.html
+title: Finetuning with a custom loss function
+---
+
+
+When fine-tuning, we can specify a loss function to be used usin the
+`finetune_loss` argument.
+
+The possible values are:
+
+- `"mae"`
+
+- `"mse"`
+
+- `"rmse"`
+
+- `"mape"`
+
+- `"smape"`
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/capabilities/forecast/08_custom_loss_function.ipynb)
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+```python
+# Read data
+df = pd.read_csv("https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/air_passengers.csv")
+
+# Fine-tune with a specified loss function and make predictions
+forecast_df = nixtla_client.forecast(
+ df=df,
+ h=12,
+ finetune_steps=5,
+ finetune_loss="mae",
+ time_col='timestamp',
+ target_col="value"
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+For more details on specifying a loss function and how it impacts the
+performance of the model, read our in-depth tutorial on [Fine-tuning
+with a specific loss
+function](https://docs.nixtla.io/docs/tutorials-fine_tuning_with_a_specific_loss_function).
+
diff --git a/nixtla/docs/capabilities/forecast/exogenous_variables.html.mdx b/nixtla/docs/capabilities/forecast/exogenous_variables.html.mdx
new file mode 100644
index 00000000..8c99d3d7
--- /dev/null
+++ b/nixtla/docs/capabilities/forecast/exogenous_variables.html.mdx
@@ -0,0 +1,117 @@
+---
+output-file: exogenous_variables.html
+title: Add exogenous variables
+---
+
+
+To model with exogenous features, you have two options: 1. Use
+historical exogenous variables: include these variables in the DataFrame
+you pass to the `forecast` method 2. Use future exogenous variables:
+include these variables in the DataFrame you pass to the `forecast`
+method and provide the future values of these exogenous features over
+the forecast horizon using the `X_df` parameter.
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, set the `base_url` argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 1. Historical exogenous variables
+
+```python
+# Read data
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short-with-ex-vars.csv')
+
+# Forecast
+forecast_df = nixtla_client.forecast(
+ df=df,
+ h=24,
+ id_col='unique_id',
+ target_col='y',
+ time_col='ds',
+ # Add the columns of `df` that will be considered as historical
+ hist_exog_list=['Exogenous1', 'Exogenous2', 'day_0', 'day_1', 'day_2', 'day_3', 'day_4', 'day_5', 'day_6']
+)
+```
+
+## 2. Future exogenous variables
+
+```python
+# Read data
+import numpy as np
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short-with-ex-vars.csv')
+
+# Load the future value of exogenous variables over the forecast horizon
+future_ex_vars_df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short-future-ex-vars.csv')
+
+# Forecast
+forecast_df = nixtla_client.forecast(
+ df=df,
+ X_df=future_ex_vars_df,
+ h=24,
+ id_col='unique_id',
+ target_col='y',
+ time_col='ds'
+)
+```
+
+## 3. Historical and future exogenous variables
+
+```python
+# Read data
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short-with-ex-vars.csv')
+
+# Load the future value of exogenous variables over the forecast horizon
+future_ex_vars_df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short-future-ex-vars.csv')
+
+# We will only use 2 exogenous of future_ex_vars_df
+future_ex_vars_df = future_ex_vars_df[["unique_id", "ds", "Exogenous1", "Exogenous2"]]
+# To pass historical exogenous variables, we need to add the list of columns
+# in the `hist_exog_list` as follows.
+
+# Forecast
+forecast_df = nixtla_client.forecast(
+ df=df,
+ X_df=future_ex_vars_df,
+ h=24,
+ id_col='unique_id',
+ target_col='y',
+ time_col='ds',
+ # Add the columns of `df` that will be considered as historical
+ hist_exog_list=['day_0', 'day_1', 'day_2', 'day_3', 'day_4', 'day_5', 'day_6']
+)
+```
+
+> 📘 Available models in Azure AI
+>
+> If you use an Azure AI endpoint, set `model="azureai"`
+>
+> `nixtla_client.detect_anomalies(..., model="azureai")`
+>
+> For the public API, two models are supported: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. See [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> for details on using `timegpt-1-long-horizon`.
+
+For more details on using exogenous features with TimeGPT, read our
+in-depth tutorials on [Exogenous
+variables](https://docs.nixtla.io/docs/tutorials-exogenous_variables)
+and on [Categorical
+variables](https://docs.nixtla.io/docs/tutorials-categorical_variables).
+
diff --git a/nixtla/docs/capabilities/forecast/finetuning.html.mdx b/nixtla/docs/capabilities/forecast/finetuning.html.mdx
new file mode 100644
index 00000000..5eec59ca
--- /dev/null
+++ b/nixtla/docs/capabilities/forecast/finetuning.html.mdx
@@ -0,0 +1,83 @@
+---
+output-file: finetuning.html
+title: Fine-tuning
+---
+
+
+We can fine-tune TimeGPT by specifying the `finetune_steps` parameter.
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+```python
+# Read data
+df = pd.read_csv("https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/air_passengers.csv")
+
+# Forecast with fine-tuning.
+# Here, we fine-tune for 5 steps
+forecast_df = nixtla_client.forecast(
+ df=df,
+ h=12,
+ finetune_steps=5,
+ time_col='timestamp',
+ target_col="value"
+)
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+By default, only a small amount of finetuning is applied
+(`finetune_depth=1`). We can increase the intensity of finetuning by
+increasing the `finetune_depth` parameter. Note that increasing
+`finetune_depth` and `finetune_steps` increases wall time for generating
+predictions.
+
+```python
+# Read data
+df = pd.read_csv("https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/air_passengers.csv")
+
+# Forecast with fine-tuning.
+# Here, we fine-tune for 5 steps
+# and we finetune more than just the last layer
+forecast_df = nixtla_client.forecast(
+ df=df,
+ h=12,
+ finetune_steps=5,
+ finetune_depth=2,
+ time_col='timestamp',
+ target_col="value"
+)
+```
+
+For more information on fine-tuning, read our [fine-tuning
+tutorial](https://docs.nixtla.io/docs/tutorials-fine_tuning).
+
diff --git a/nixtla/docs/capabilities/forecast/forecast.html.mdx b/nixtla/docs/capabilities/forecast/forecast.html.mdx
new file mode 100644
index 00000000..22d40a77
--- /dev/null
+++ b/nixtla/docs/capabilities/forecast/forecast.html.mdx
@@ -0,0 +1,46 @@
+---
+output-file: forecast.html
+title: Forecast
+---
+
+
+This section shows the capabilities TimeGPT offers for forecasting.
+
+TimeGPT is capable of zero-shot forecasting a wide variety of time
+series from different domains, thanks to its pretraining on a vast
+amount of time series data.
+
+Here, you will find recipes for the following tasks:
+
+- [Zero-shot
+ forecasting](https://docs.nixtla.io/docs/capabilities-forecast-quickstart)
+
+- [Forecasting with exogenous
+ variables](https://docs.nixtla.io/docs/capabilities-forecast-add_exogenous_variables)
+
+- [Forecasting with holidays and special
+ dates](https://docs.nixtla.io/docs/capabilities-forecast-add_holidays_and_special_dates)
+
+- [Forecasting with categorical
+ variables](https://docs.nixtla.io/docs/capabilities-forecast-add_categorical_variables)
+
+- [Long-horizon
+ forecasting](https://docs.nixtla.io/docs/capabilities-forecast-long_horizon_forecasting)
+
+- [Forecasting multiple
+ series](https://docs.nixtla.io/docs/capabilities-forecast-multiple_series_forecasting)
+
+- [Fine-tuning
+ TimeGPT](https://docs.nixtla.io/docs/capabilities-forecast-fine_tuning)
+
+- [Fine-tuning with a specific loss
+ function](https://docs.nixtla.io/docs/capabilities-forecast-finetuning_with_a_custom_loss_function)
+
+- [Cross-validation](https://docs.nixtla.io/docs/capabilities-forecast-cross_validation)
+
+- [Adding prediction
+ intervals](https://docs.nixtla.io/docs/capabilities-forecast-predictions_intervals)
+
+- [Dealing with irregular
+ timestamps](https://docs.nixtla.io/docs/capabilities-forecast-irregular_timestamps)
+
diff --git a/nixtla/docs/capabilities/forecast/holidays_special_dates.html.mdx b/nixtla/docs/capabilities/forecast/holidays_special_dates.html.mdx
new file mode 100644
index 00000000..0c3fad1a
--- /dev/null
+++ b/nixtla/docs/capabilities/forecast/holidays_special_dates.html.mdx
@@ -0,0 +1,57 @@
+---
+output-file: holidays_special_dates.html
+title: Add holidays and special dates
+---
+
+
+You can create DataFrames specifying holidays for particular countries
+and specify your own special dates to include them as features for
+forecasting.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/capabilities/forecast/03_holidays_special_dates.ipynb)
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+from nixtla.date_features import CountryHolidays
+from nixtla.date_features import SpecialDates
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+```python
+# Get country holidays for the US
+c_holidays = CountryHolidays(countries=['US'])
+periods = 365 * 1
+dates = pd.date_range(end='2023-09-01', periods=periods)
+holidays_df = c_holidays(dates)
+
+# Specify your own special dates
+special_dates = SpecialDates(
+ special_dates={
+ 'Important Dates': ['2021-02-26', '2020-02-26'],
+ 'Very Important Dates': ['2021-01-26', '2020-01-26', '2019-01-26']
+ }
+)
+periods = 365 * 1
+dates = pd.date_range(end='2023-09-01', periods=periods)
+special_dates_df = special_dates(dates)
+```
+
+For a detailed guide on using special dates and holidays, read our
+tutorial on [Holidays and special
+dates](https://docs.nixtla.io/docs/tutorials-holidays_and_special_dates).
+
diff --git a/nixtla/docs/capabilities/forecast/irregular_timestamps.html.mdx b/nixtla/docs/capabilities/forecast/irregular_timestamps.html.mdx
new file mode 100644
index 00000000..3553ff3a
--- /dev/null
+++ b/nixtla/docs/capabilities/forecast/irregular_timestamps.html.mdx
@@ -0,0 +1,264 @@
+---
+output-file: irregular_timestamps.html
+title: Irregular timestamps
+---
+
+
+When working with time series data, it is important to specify its
+frequency correctly, as this can significantly impact forecasting
+results. TimeGPT is designed to automatically infer the frequency of
+your timestamps. For commonly used frequencies, such as hourly, daily,
+or monthly, TimeGPT reliably infers the frequency automatically, so no
+additional input is required.
+
+However, for irregular frequencies, where observations are not recorded
+at consistent or regular intervals, such as the days the U.S. stock
+market is open, it is necessary to specify the frequency directly.
+
+TimeGPT requires that your data does not contain missing values, as this
+is not currently supported. In other words, the irregularity of the data
+should stem from the nature of the recorded phenomenon, not from missing
+observations. If your data contains missing values, please refer to our
+[tutorial on missing
+dates](https://docs.nixtla.io/docs/tutorials-missing_values).
+
+In this tutorial, we will show you how to handle irregular and custom
+frequencies in TimeGPT.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/capabilities/forecast/11_irregular_timestamps.ipynb)
+
+## 1. Import packages
+
+First, we import the required packages and initialize the Nixtla client.
+
+```python
+import pandas as pd
+import pandas_market_calendars as mcal
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 2. Handling regular frequencies
+
+As discussed in the introduction, for time series data with regular
+frequencies, where observations are recorded at consistent intervals,
+TimeGPT can automatically infer the frequency of your timestamps if the
+input data is a **pandas DataFrame**. If you prefer not to rely on
+TimeGPT’s automatic inference, you can set the `freq` parameter to a
+valid [pandas frequency
+string](https://pandas.pydata.org/docs/user_guide/timeseries.html#offset-aliases),
+such as `MS` for month-start frequency or `min` for minutely frequency.
+
+When working with **Polars DataFrames**, you must specify the frequency
+explicitly by using a valid [polars
+offset](https://docs.pola.rs/api/python/stable/reference/expressions/api/polars.Expr.dt.offset_by.html),
+such as `1d` for daily frequency or `1h` for hourly frequency.
+
+Below is an example of how to specify the frequency for a Polars
+DataFrame.
+
+```python
+import polars as pl
+
+url = 'https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/air_passengers.csv'
+
+polars_df = pl.read_csv(url, try_parse_dates=True)
+
+fcst_df = nixtla_client.forecast(
+ df=polars_df,
+ h=12,
+ freq='1mo',
+ time_col='timestamp',
+ target_col='value',
+ level=[80, 95]
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+```python
+nixtla_client.plot(polars_df, fcst_df, time_col='timestamp', target_col='value', level=[80,95])
+```
+
+
+
+## 3. Handling irregular frequencies
+
+In this section, we will discuss cases where observations are not
+recorded at consistent intervals.
+
+### 3.1 Load data
+
+We will use the daily stock prices of Palantir Technologies (PLTR) from
+2020 to 2023. The dataset includes data up to 2023-09-22, but for this
+tutorial, we will exclude any data before 2023-08-28. This allows us to
+show how a custom frequency can handle days when the stock market is
+closed, such as Labor Day in the U.S.
+
+```python
+url = 'https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/openbb/pltr.csv'
+pltr_df = pd.read_csv(url, parse_dates=['date'])
+pltr_df = pltr_df.query('date < "2023-08-28"')
+pltr_df.head()
+```
+
+| | date | Open | High | Low | Close | Adj Close | Volume | Dividends | Stock Splits |
+|-----|------------|-------|-------|------|-------|-----------|-----------|-----------|--------------|
+| 0 | 2020-09-30 | 10.00 | 11.41 | 9.11 | 9.50 | 9.50 | 338584400 | 0.0 | 0.0 |
+| 1 | 2020-10-01 | 9.69 | 10.10 | 9.23 | 9.46 | 9.46 | 124297600 | 0.0 | 0.0 |
+| 2 | 2020-10-02 | 9.06 | 9.28 | 8.94 | 9.20 | 9.20 | 55018300 | 0.0 | 0.0 |
+| 3 | 2020-10-05 | 9.43 | 9.49 | 8.92 | 9.03 | 9.03 | 36316900 | 0.0 | 0.0 |
+| 4 | 2020-10-06 | 9.04 | 10.18 | 8.90 | 9.90 | 9.90 | 90864000 | 0.0 | 0.0 |
+
+We will forecast the **adjusted closing price**, which represents the
+stock’s closing price adjusted for corporate actions such as stock
+splits, dividends, and rights offerings. Hence, we will exclude the
+other columns from the dataset.
+
+```python
+pltr_df = pltr_df[['date', 'Adj Close']]
+
+nixtla_client.plot(pltr_df, time_col = "date", target_col = "Adj Close")
+```
+
+
+
+### 3.2 Define the frequency
+
+To define a custom frequency, we will first extract and sort the dates
+from the input data, ensuring they are in the correct datetime format.
+Next, we will use the
+[`pandas_market_calendars package`](https://pypi.org/project/pandas-market-calendars/),
+specifically the `get_calendar` method, to obtain the New York Stock
+Exchange (NYSE) calendar. Using this calendar, we can create a custom
+frequency that includes only the days the stock market is open.
+
+```python
+dates = pd.DatetimeIndex(sorted(pltr_df['date'].unique())) # sort all dates in the dataset
+
+nyse = mcal.get_calendar('NYSE') # New Yor Stock Exchange calendar
+```
+
+Note that the days the stock market is open need to include all the
+dates in the input data plus the forecast horizon. In this example, we
+will forecast 7 days ahead, so we need to make sure our trading days
+include the last date in the input data as well as the next 7 valid
+trading days.
+
+To avoid dealing with holidays or weekends during the forecast horizon,
+we will specify an end date well beyond the forecast horizon. For this
+example, we will use January 1, 2024, as a safe cutoff.
+
+```python
+trading_days = nyse.valid_days(start_date=dates.min(), end_date="2024-01-01").tz_localize(None)
+```
+
+Now, with the list of trading days, we can identify the days the stock
+market is closed. These are all weekdays (Monday to Friday) within the
+range that are not trading days. Using this information, we can define a
+custom frequency that skips the stock market’s closed days.
+
+```python
+all_weekdays = pd.date_range(start=dates.min(), end="2024-01-01", freq='B')
+
+closed_days = all_weekdays.difference(trading_days)
+
+custom_bday = pd.offsets.CustomBusinessDay(holidays=closed_days)
+```
+
+### 3.3 Forecast with TimeGPT
+
+With the custom frequency defined, we can now use the `forecast` method,
+specifying the `custom_bday` frequency in the `freq` argument. This will
+make the forecast respect the trading schedule of the stock market.
+
+```python
+fcst_pltr_df = nixtla_client.forecast(
+ df=pltr_df,
+ h=7,
+ freq=custom_bday,
+ time_col='date',
+ target_col='Adj Close',
+ level=[80, 95]
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Querying model metadata...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+```python
+nixtla_client.plot(pltr_df, fcst_pltr_df, time_col = "date", target_col = "Adj Close", level=[80, 95], max_insample_length = 180)
+```
+
+
+
+```python
+fcst_pltr_df[['date']].head(7)
+```
+
+| | date |
+|-----|------------|
+| 0 | 2023-08-28 |
+| 1 | 2023-08-29 |
+| 2 | 2023-08-30 |
+| 3 | 2023-08-31 |
+| 4 | 2023-09-01 |
+| 5 | 2023-09-05 |
+| 6 | 2023-09-06 |
+
+Note that the forecast excludes 2023-09-04, which was a Monday when the
+stock market was closed for Labor Day in the United States.
+
+## 4. Summary
+
+Below are the key takeaways of this tutorial:
+
+- TimeGPT can reliably infer regular frequencies, but you can override
+ this by setting the `freq` parameter to the corresponding pandas
+ alias.
+
+- When working with polars data frames, you must always specify the
+ frequency using the correct polars offset.
+
+- TimeGPT supports irregular frequencies and allows you to define a
+ custom frequency, generating forecasts exclusively for the specified
+ dates.
+
diff --git a/nixtla/docs/capabilities/forecast/longhorizon.html.mdx b/nixtla/docs/capabilities/forecast/longhorizon.html.mdx
new file mode 100644
index 00000000..264a8f58
--- /dev/null
+++ b/nixtla/docs/capabilities/forecast/longhorizon.html.mdx
@@ -0,0 +1,73 @@
+---
+output-file: longhorizon.html
+title: Long-horizon forecasting
+---
+
+
+Long-horizon forecasting is when you wish to predict more than one
+seasonal cycle into the future. TimeGPT supports long-horizon
+forecasting simply by setting `model=timegpt-1-long-horizon`.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/capabilities/forecast/05_longhorizon.ipynb)
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+```python
+# Read the data
+df = pd.read_csv("https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/air_passengers.csv")
+
+# Forecast
+forecast_df = nixtla_client.forecast(
+ df=df,
+ h=36,
+ model='timegpt-1-long-horizon',
+ time_col='timestamp',
+ target_col="value"
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon. This may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+For a detailed guide on long-horizon forecasting, read our in-depth
+tutorial on [Long-horizon
+forecasting](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting).
+
diff --git a/nixtla/docs/capabilities/forecast/multiple_series.html.mdx b/nixtla/docs/capabilities/forecast/multiple_series.html.mdx
new file mode 100644
index 00000000..9cb617ee
--- /dev/null
+++ b/nixtla/docs/capabilities/forecast/multiple_series.html.mdx
@@ -0,0 +1,69 @@
+---
+output-file: multiple_series.html
+title: Multiple series forecasting
+---
+
+
+TimeGPT can concurrently forecast many series at the same time. Simply
+use a DataFrame with more than one unique value in the `unique_id`
+column.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/capabilities/forecast/06_multiple_series.ipynb)
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+```python
+# Read the data
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short.csv')
+
+# Forecast
+forecast_df = nixtla_client.forecast(
+ df=df,
+ h=24
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: H
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+For more details on forecasting multiple series, read our in-depth
+tutorial on [Multiple series
+forecasting](https://docs.nixtla.io/docs/tutorials-multiple_series_forecasting).
+
diff --git a/nixtla/docs/capabilities/forecast/prediction_intervals.html.mdx b/nixtla/docs/capabilities/forecast/prediction_intervals.html.mdx
new file mode 100644
index 00000000..4cc47e22
--- /dev/null
+++ b/nixtla/docs/capabilities/forecast/prediction_intervals.html.mdx
@@ -0,0 +1,85 @@
+---
+output-file: prediction_intervals.html
+title: Predictions intervals
+---
+
+
+We can generate prediction intervals using the `level` parameter in the
+`forecast` method. It takes any values between 0 and 100, including
+decimal numbers.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/capabilities/forecast/10_prediction_intervals.ipynb)
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+```python
+# Read the data
+df = pd.read_csv("https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/air_passengers.csv")
+
+# Forecast using a 80% confidence interval
+forecast_df = nixtla_client.forecast(
+ df=df,
+ h=12,
+ time_col='timestamp',
+ target_col="value",
+ level=[80]
+)
+
+# Plot predictions with intervals
+nixtla_client.plot(
+ df=df,
+ forecasts_df=forecast_df,
+ time_col='timestamp',
+ target_col='value',
+ level=[80]
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+For more details on uncertainty quantification, read our tutorials on
+using [quantile
+forecasts](https://docs.nixtla.io/docs/tutorials-quantile_forecasts) and
+[prediction
+intervals](https://docs.nixtla.io/docs/tutorials-prediction_intervals).
+
diff --git a/nixtla/docs/capabilities/forecast/quickstart.html.mdx b/nixtla/docs/capabilities/forecast/quickstart.html.mdx
new file mode 100644
index 00000000..d62f89ce
--- /dev/null
+++ b/nixtla/docs/capabilities/forecast/quickstart.html.mdx
@@ -0,0 +1,77 @@
+---
+output-file: quickstart.html
+title: Quickstart
+---
+
+
+To forecast with TimeGPT, call the `forecast` method. Pass your
+DataFrame and specify your target and time column names. Then plot the
+predictions using the `plot` method. You can read about data
+requierments
+[here](https://docs.nixtla.io/docs/getting-started-data_requirements).
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/capabilities/forecast/01_quickstart.ipynb)
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, set the `base_url` argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+```python
+# Read the data
+df = pd.read_csv("https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/air_passengers.csv")
+
+# Forecast
+forecast_df = nixtla_client.forecast(
+ df=df,
+ h=12,
+ time_col='timestamp',
+ target_col="value"
+)
+
+# Plot predictions
+nixtla_client.plot(
+ df=df,
+ forecasts_df=forecast_df,
+ time_col='timestamp',
+ target_col='value'
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+
+
+> 📘 Available models in Azure AI
+>
+> If you use an Azure AI endpoint, set `model="azureai"`
+>
+> `nixtla_client.detect_anomalies(..., model="azureai")`
+>
+> For the public API, two models are supported: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. See [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> for details on using `timegpt-1-long-horizon`.
+
diff --git a/nixtla/docs/capabilities/historical-anomaly-detection/01_quickstart_files/figure-markdown_strict/cell-5-output-2.png b/nixtla/docs/capabilities/historical-anomaly-detection/01_quickstart_files/figure-markdown_strict/cell-5-output-2.png
new file mode 100644
index 00000000..be325c02
Binary files /dev/null and b/nixtla/docs/capabilities/historical-anomaly-detection/01_quickstart_files/figure-markdown_strict/cell-5-output-2.png differ
diff --git a/nixtla/docs/capabilities/historical-anomaly-detection/02_anomaly_exogenous_files/figure-markdown_strict/cell-5-output-2.png b/nixtla/docs/capabilities/historical-anomaly-detection/02_anomaly_exogenous_files/figure-markdown_strict/cell-5-output-2.png
new file mode 100644
index 00000000..d9f53df1
Binary files /dev/null and b/nixtla/docs/capabilities/historical-anomaly-detection/02_anomaly_exogenous_files/figure-markdown_strict/cell-5-output-2.png differ
diff --git a/nixtla/docs/capabilities/historical-anomaly-detection/02_anomaly_exogenous_files/figure-markdown_strict/cell-6-output-1.png b/nixtla/docs/capabilities/historical-anomaly-detection/02_anomaly_exogenous_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..f3befb99
Binary files /dev/null and b/nixtla/docs/capabilities/historical-anomaly-detection/02_anomaly_exogenous_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/nixtla/docs/capabilities/historical-anomaly-detection/03_anomaly_detection_date_features_files/figure-markdown_strict/cell-5-output-2.png b/nixtla/docs/capabilities/historical-anomaly-detection/03_anomaly_detection_date_features_files/figure-markdown_strict/cell-5-output-2.png
new file mode 100644
index 00000000..5b09920c
Binary files /dev/null and b/nixtla/docs/capabilities/historical-anomaly-detection/03_anomaly_detection_date_features_files/figure-markdown_strict/cell-5-output-2.png differ
diff --git a/nixtla/docs/capabilities/historical-anomaly-detection/03_anomaly_detection_date_features_files/figure-markdown_strict/cell-6-output-1.png b/nixtla/docs/capabilities/historical-anomaly-detection/03_anomaly_detection_date_features_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..ad523f49
Binary files /dev/null and b/nixtla/docs/capabilities/historical-anomaly-detection/03_anomaly_detection_date_features_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/nixtla/docs/capabilities/historical-anomaly-detection/04_confidence_levels_files/figure-markdown_strict/cell-5-output-2.png b/nixtla/docs/capabilities/historical-anomaly-detection/04_confidence_levels_files/figure-markdown_strict/cell-5-output-2.png
new file mode 100644
index 00000000..c043dd31
Binary files /dev/null and b/nixtla/docs/capabilities/historical-anomaly-detection/04_confidence_levels_files/figure-markdown_strict/cell-5-output-2.png differ
diff --git a/nixtla/docs/capabilities/historical-anomaly-detection/anomaly_detection_date_features.html.mdx b/nixtla/docs/capabilities/historical-anomaly-detection/anomaly_detection_date_features.html.mdx
new file mode 100644
index 00000000..963c0335
--- /dev/null
+++ b/nixtla/docs/capabilities/historical-anomaly-detection/anomaly_detection_date_features.html.mdx
@@ -0,0 +1,83 @@
+---
+output-file: anomaly_detection_date_features.html
+title: Add date features
+---
+
+
+If your dataset lacks exogenous variables, add date features to inform
+the model for historical anomaly detection. Use the `date_features`
+argument. Set it to `True` to extract all possible features, or pass a
+list of specific features to include.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/capabilities/anomaly-detection/03_anomaly_detection_date_features.ipynb)
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, set the `base_url` argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+```python
+# Read the data
+df = pd.read_csv('https://datasets-nixtla.s3.amazonaws.com/peyton-manning.csv')
+
+# Add date features for anomaly detection
+# Here, we use date features at the month and year levels
+anomalies_df_x = nixtla_client.detect_anomalies(
+ df,
+ freq='D',
+ date_features=['month', 'year'],
+ date_features_to_one_hot=True,
+ level=99.99,
+)
+
+# Plot anomalies
+nixtla_client.plot(df, anomalies_df_x)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Querying model metadata...
+INFO:nixtla.nixtla_client:Using the following exogenous features: ['month_1.0', 'month_2.0', 'month_3.0', 'month_4.0', 'month_5.0', 'month_6.0', 'month_7.0', 'month_8.0', 'month_9.0', 'month_10.0', 'month_11.0', 'month_12.0', 'year_2007.0', 'year_2008.0', 'year_2009.0', 'year_2010.0', 'year_2011.0', 'year_2012.0', 'year_2013.0', 'year_2014.0', 'year_2015.0', 'year_2016.0']
+INFO:nixtla.nixtla_client:Calling Anomaly Detector Endpoint...
+```
+
+
+
+```python
+# Plot weights of date features
+nixtla_client.weights_x.plot.barh(x='features', y='weights')
+```
+
+
+
+> 📘 Available models in Azure AI
+>
+> If you use an Azure AI endpoint, set `model="azureai"`
+>
+> `nixtla_client.detect_anomalies(..., model="azureai")`
+>
+> For the public API, two models are supported: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. See [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> for details on using `timegpt-1-long-horizon`.
+
+For more details, check out our in-depth tutorial on [anomaly
+detection](https://docs.nixtla.io/docs/tutorials/anomaly_detection).
+
diff --git a/nixtla/docs/capabilities/historical-anomaly-detection/anomaly_exogenous.html.mdx b/nixtla/docs/capabilities/historical-anomaly-detection/anomaly_exogenous.html.mdx
new file mode 100644
index 00000000..2abf165a
--- /dev/null
+++ b/nixtla/docs/capabilities/historical-anomaly-detection/anomaly_exogenous.html.mdx
@@ -0,0 +1,82 @@
+---
+output-file: anomaly_exogenous.html
+title: Add exogenous variables
+---
+
+
+To detect anomalies with exogenous variables, load a dataset with the
+exogenous features as columns and use the same `detect_anomalies`
+method.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/capabilities/anomaly-detection/02_anomaly_exogenous.ipynb)
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, set the `base_url` argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+```python
+# Read the dataset
+# The dataset has exogenous features in its columns
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short-with-ex-vars.csv')
+
+# Detect anomalies
+anomalies_df = nixtla_client.detect_anomalies(
+ df=df,
+ time_col='ds',
+ target_col='y',
+)
+
+# Plot anomalies
+nixtla_client.plot(df, anomalies_df)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: H
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Querying model metadata...
+INFO:nixtla.nixtla_client:Using the following exogenous features: ['Exogenous1', 'Exogenous2', 'day_0', 'day_1', 'day_2', 'day_3', 'day_4', 'day_5', 'day_6']
+INFO:nixtla.nixtla_client:Calling Anomaly Detector Endpoint...
+```
+
+
+
+```python
+# Plot weights of date features
+nixtla_client.weights_x.plot.barh(x='features', y='weights')
+```
+
+
+
+> 📘 Available models in Azure AI
+>
+> If you use an Azure AI endpoint, set `model="azureai"`
+>
+> `nixtla_client.detect_anomalies(..., model="azureai")`
+>
+> For the public API, two models are supported: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. See [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> for details on using `timegpt-1-long-horizon`.
+
+Read our detailed guide on [anomaly
+detection](https://docs.nixtla.io/docs/tutorials/anomaly_detection) for
+more information.
+
diff --git a/nixtla/docs/capabilities/historical-anomaly-detection/confidence_levels.html.mdx b/nixtla/docs/capabilities/historical-anomaly-detection/confidence_levels.html.mdx
new file mode 100644
index 00000000..1f1bc752
--- /dev/null
+++ b/nixtla/docs/capabilities/historical-anomaly-detection/confidence_levels.html.mdx
@@ -0,0 +1,77 @@
+---
+output-file: confidence_levels.html
+title: Add confidence levels
+---
+
+
+Tweak the confidence level used for historical anomaly detection. By
+default, if a value falls outside the 99% confidence interval, it is
+labeled as an anomaly.
+
+Modify this with the `level` parameter, which accepts any value between
+0 and 100, including decimals.
+
+Increasing the `level` results in fewer anomalies detected, while
+decreasing the `level` increases the number of anomalies detected.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/capabilities/anomaly-detection/04_confidence_levels.ipynb)
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, set the `base_url` argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+```python
+# Read the data
+df = pd.read_csv('https://datasets-nixtla.s3.amazonaws.com/peyton-manning.csv')
+
+# Anomaly detection using a 70% confidence interval
+anomalies_df = nixtla_client.detect_anomalies(
+ df,
+ freq='D',
+ level=70
+)
+
+# Plot anomalies
+nixtla_client.plot(df, anomalies_df)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Querying model metadata...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Calling Anomaly Detector Endpoint...
+```
+
+
+
+> 📘 Available models in Azure AI
+>
+> If you use an Azure AI endpoint, set `model="azureai"`
+>
+> `nixtla_client.detect_anomalies(..., model="azureai")`
+>
+> For the public API, two models are supported: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. See [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> for details on using `timegpt-1-long-horizon`.
+
+For more information, read our detailed tutorial on [anomaly
+detection](https://docs.nixtla.io/docs/tutorials/anomaly_detection).
+
diff --git a/nixtla/docs/capabilities/historical-anomaly-detection/historical_anomaly_detection.html.mdx b/nixtla/docs/capabilities/historical-anomaly-detection/historical_anomaly_detection.html.mdx
new file mode 100644
index 00000000..28b65727
--- /dev/null
+++ b/nixtla/docs/capabilities/historical-anomaly-detection/historical_anomaly_detection.html.mdx
@@ -0,0 +1,31 @@
+---
+output-file: historical_anomaly_detection.html
+title: Historical anomaly detection
+---
+
+
+This section provides various recipes for performing historical anomaly
+detection using TimeGPT.
+
+Historical anomaly detection identifies data points that deviate from
+the expected behavior over a given historical time series, helping to
+spot fraudulent activity, security breaches, or significant outliers.
+
+The process involves generating predictions and constructing a 99%
+confidence interval. Data points falling outside this interval are
+considered anomalies.
+
+This section covers:
+
+- [Historical anomaly
+ detection](https://docs.nixtla.io/docs/capabilities-historical-anomaly-detection-quickstart)
+
+- [Historical anomaly detection with exogenous
+ features](https://docs.nixtla.io/docs/capabilities-historical-anomaly-detection-add_exogenous_variables)
+
+- [Historical anomaly detection with date
+ features](https://docs.nixtla.io/docs/capabilities-historical-anomaly-detection-add_date_features)
+
+- [Modifying the confidence
+ intervals](https://docs.nixtla.io/docs/capabilities-historical-anomaly-detection-add_confidence_levels)
+
diff --git a/nixtla/docs/capabilities/historical-anomaly-detection/quickstart.html.mdx b/nixtla/docs/capabilities/historical-anomaly-detection/quickstart.html.mdx
new file mode 100644
index 00000000..8d230d76
--- /dev/null
+++ b/nixtla/docs/capabilities/historical-anomaly-detection/quickstart.html.mdx
@@ -0,0 +1,66 @@
+---
+output-file: quickstart.html
+title: Quickstart
+---
+
+
+To perform historical anomaly detection, use the `detect_anomalies`
+method. Then, plot the anomalies using the `plot` method.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/capabilities/anomaly-detection/01_quickstart.ipynb)
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, set the `base_url` argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+```python
+# Read the dataset
+df = pd.read_csv('https://datasets-nixtla.s3.amazonaws.com/peyton-manning.csv')
+
+# Detect anomalies
+anomalies_df = nixtla_client.detect_anomalies(df, freq='D')
+
+# Plot anomalies
+nixtla_client.plot(df, anomalies_df)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Calling Anomaly Detector Endpoint...
+```
+
+
+
+> 📘 Available models in Azure AI
+>
+> If you use an Azure AI endpoint, set `model="azureai"`
+>
+> `nixtla_client.detect_anomalies(..., model="azureai")`
+>
+> For the public API, two models are supported: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. See [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> for details on using `timegpt-1-long-horizon`.
+
+For an in-depth guide on historical anomaly detection with TimeGPT,
+check out our
+[tutorial](https://docs.nixtla.io/docs/tutorials-anomaly_detection).
+
diff --git a/nixtla/docs/capabilities/online-anomaly-detection/01_quickstart_files/figure-markdown_strict/cell-6-output-1.png b/nixtla/docs/capabilities/online-anomaly-detection/01_quickstart_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..2b543d3b
Binary files /dev/null and b/nixtla/docs/capabilities/online-anomaly-detection/01_quickstart_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/nixtla/docs/capabilities/online-anomaly-detection/01_quickstart_files/figure-markdown_strict/cell-8-output-1.png b/nixtla/docs/capabilities/online-anomaly-detection/01_quickstart_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..b9d643d2
Binary files /dev/null and b/nixtla/docs/capabilities/online-anomaly-detection/01_quickstart_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/nixtla/docs/capabilities/online-anomaly-detection/02_adjusting_detection_process_files/figure-markdown_strict/cell-10-output-1.png b/nixtla/docs/capabilities/online-anomaly-detection/02_adjusting_detection_process_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..7cd92af5
Binary files /dev/null and b/nixtla/docs/capabilities/online-anomaly-detection/02_adjusting_detection_process_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/nixtla/docs/capabilities/online-anomaly-detection/02_adjusting_detection_process_files/figure-markdown_strict/cell-12-output-1.png b/nixtla/docs/capabilities/online-anomaly-detection/02_adjusting_detection_process_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..bc27ae6d
Binary files /dev/null and b/nixtla/docs/capabilities/online-anomaly-detection/02_adjusting_detection_process_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/nixtla/docs/capabilities/online-anomaly-detection/02_adjusting_detection_process_files/figure-markdown_strict/cell-8-output-1.png b/nixtla/docs/capabilities/online-anomaly-detection/02_adjusting_detection_process_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..f268f834
Binary files /dev/null and b/nixtla/docs/capabilities/online-anomaly-detection/02_adjusting_detection_process_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/nixtla/docs/capabilities/online-anomaly-detection/03_univariate_vs_multivariate_anomaly_detection_files/figure-markdown_strict/cell-10-output-1.png b/nixtla/docs/capabilities/online-anomaly-detection/03_univariate_vs_multivariate_anomaly_detection_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..4dcff566
Binary files /dev/null and b/nixtla/docs/capabilities/online-anomaly-detection/03_univariate_vs_multivariate_anomaly_detection_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/nixtla/docs/capabilities/online-anomaly-detection/03_univariate_vs_multivariate_anomaly_detection_files/figure-markdown_strict/cell-8-output-1.png b/nixtla/docs/capabilities/online-anomaly-detection/03_univariate_vs_multivariate_anomaly_detection_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..e4e20c6a
Binary files /dev/null and b/nixtla/docs/capabilities/online-anomaly-detection/03_univariate_vs_multivariate_anomaly_detection_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/nixtla/docs/capabilities/online-anomaly-detection/adjusting_detection_process.html.mdx b/nixtla/docs/capabilities/online-anomaly-detection/adjusting_detection_process.html.mdx
new file mode 100644
index 00000000..d0af28c7
--- /dev/null
+++ b/nixtla/docs/capabilities/online-anomaly-detection/adjusting_detection_process.html.mdx
@@ -0,0 +1,200 @@
+---
+output-file: adjusting_detection_process.html
+title: Adjusting the Anomaly Detection Process
+---
+
+
+This notebook explores methods to improve anomaly detection by refining
+the detection process. TimeGPT leverages its forecasting capabilities to
+identify anomalies based on forecast errors. By optimizing forecast
+parameters and accuracy, you can align anomaly detection with specific
+use cases and improve its accuracy.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/capabilities/online-anomaly-detection/02_adjusting_detection_process.ipynb)
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+import matplotlib.pyplot as plt
+```
+
+
+```python
+# Utility function to plot anomalies
+def plot_anomaly(df, anomaly_df, time_col = 'ts', target_col = 'y'):
+ merged_df = pd.merge(df.tail(300), anomaly_df[[time_col, 'anomaly', 'TimeGPT']], on=time_col, how='left')
+ plt.figure(figsize=(12, 2))
+ plt.plot(merged_df[time_col], merged_df[target_col], label='y', color='navy', alpha=0.8)
+ plt.plot(merged_df[time_col], merged_df['TimeGPT'], label='TimeGPT', color='orchid', alpha=0.7)
+ plt.scatter(merged_df.loc[merged_df['anomaly'] == True, time_col], merged_df.loc[merged_df['anomaly'] == True, target_col], color='orchid', label='Anomalies Detected')
+ plt.legend()
+ plt.tight_layout()
+ plt.show()
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, set the `base_url` argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 1. Conduct anomaly detection
+
+After initializing an instance of
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient),
+let’s explore an example using the Peyton Manning dataset.
+
+```python
+df = pd.read_csv('https://datasets-nixtla.s3.amazonaws.com/peyton-manning.csv',parse_dates = ['ds']).tail(200)
+df.head()
+```
+
+| | unique_id | ds | y |
+|------|-----------|------------|----------|
+| 2764 | 0 | 2015-07-05 | 6.499787 |
+| 2765 | 0 | 2015-07-06 | 6.859615 |
+| 2766 | 0 | 2015-07-07 | 6.881411 |
+| 2767 | 0 | 2015-07-08 | 6.997596 |
+| 2768 | 0 | 2015-07-09 | 7.152269 |
+
+First, let’s set a baseline by using only the default parameters of the
+method.
+
+```python
+# Base case for anomaly detection using detect_anomaly_online
+anomaly_df = nixtla_client.detect_anomalies_online(
+ df,
+ freq='D',
+ h=14,
+ level=80,
+ detection_size=150
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+WARNING:nixtla.nixtla_client:Detection size is large. Using the entire series to compute the anomaly threshold...
+INFO:nixtla.nixtla_client:Calling Online Anomaly Detector Endpoint...
+```
+
+```python
+plot_anomaly(df, anomaly_df, time_col = 'ds', target_col = 'y')
+```
+
+
+
+## 2. Adjusting the Anomaly Detection Process
+
+This section explores two key approaches to enhancing anomaly detection:
+
+1. fine-tuning the model to boost forecast accuracy
+2. adjusting forecast horizon and step sizes to optimize time series
+ segmentation and analysis.
+
+These strategies allow for a more tailored and effective anomaly
+detection process.
+
+### 2.1 Fine-tune TimeGPT
+
+TimeGPT uses forecast errors for anomaly detection, so improving
+forecast accuracy reduces noise in the errors, leading to better anomaly
+detection. You can fine-tune the model using the following parameters:
+
+- `finetune_steps`: Number of steps for finetuning TimeGPT on new
+ data.
+- `finetune_depth`: Level of fine-tuning controlling the quantity of
+ parameters being fine-tuned (see our [in-depth
+ tutorial](https://docs.nixtla.io/docs/tutorials-controlling_the_level_of_fine_tuning))
+- `finetune_loss`: Loss function to be used during the fine-tuning
+ process.
+
+```python
+anomaly_online_ft = nixtla_client.detect_anomalies_online(
+ df,
+ freq='D',
+ h=14,
+ level=80,
+ detection_size=150,
+ finetune_steps = 10, # Number of steps for fine-tuning TimeGPT on new data
+ finetune_depth = 2, # Intensity of finetuning
+ finetune_loss = 'mae' # Loss function used during the finetuning process
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+WARNING:nixtla.nixtla_client:Detection size is large. Using the entire series to compute the anomaly threshold...
+INFO:nixtla.nixtla_client:Calling Online Anomaly Detector Endpoint...
+```
+
+```python
+plot_anomaly(df, anomaly_online_ft, time_col = 'ds', target_col = 'y')
+```
+
+
+
+From the plot above, we can see that fewer anomalies were detected by
+the model, since the fine-tuning process helps TimeGPT better forecast
+the series.
+
+### 2.2 Change forecast horizon and step
+
+Similar to cross-validation, the anomaly detection method generates
+forecasts for historical data by splitting the time series into multiple
+windows. The way these windows are defined can impact the anomaly
+detection results. Two key parameters control this process:
+
+- `h`: Specifies how many steps into the future the forecast is made
+ for each window.
+- `step_size`: Determines the interval between the starting points of
+ consecutive windows.
+
+Note that when `step_size` is smaller than `h`, then we get overlapping
+windows. This can make the detection process more robust, as TimeGPT
+will see the same time step more than once. However, this comes with a
+computational cost, since the same time step will be predicted more than
+once.
+
+```python
+anomaly_df_horizon = nixtla_client.detect_anomalies_online(
+ df,
+ time_col='ds',
+ target_col='y',
+ freq='D',
+ h=2, # Forecast horizon
+ step_size = 1, # Step size for moving through the time series data
+ level=80,
+ detection_size=150
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+WARNING:nixtla.nixtla_client:Detection size is large. Using the entire series to compute the anomaly threshold...
+INFO:nixtla.nixtla_client:Calling Online Anomaly Detector Endpoint...
+```
+
+```python
+plot_anomaly(df, anomaly_df_horizon, time_col = 'ds', target_col = 'y')
+```
+
+
+
+> 📘 **Balancing h and step_size depends on your data:** For frequent,
+> short-lived anomalies, use a smaller `h` to focus on short-term
+> predictions and a smaller `step_size` to increase overlap and
+> sensitivity. For smooth trends or long-term patterns, use a larger `h`
+> to capture broader anomalies and a larger `step_size` to reduce noise
+> and computational cost.
+
diff --git a/nixtla/docs/capabilities/online-anomaly-detection/online_anomaly_detection.html.mdx b/nixtla/docs/capabilities/online-anomaly-detection/online_anomaly_detection.html.mdx
new file mode 100644
index 00000000..baadf129
--- /dev/null
+++ b/nixtla/docs/capabilities/online-anomaly-detection/online_anomaly_detection.html.mdx
@@ -0,0 +1,30 @@
+---
+output-file: online_anomaly_detection.html
+title: Online (Real-Time) Anomaly Detection
+---
+
+
+Online anomaly detection dynamically identifies anomalies as data
+streams in, allowing users to specify the number of timestamps to
+monitor. This method is well-suited for immediate applications, such as
+fraud detection, live sensor monitoring, or tracking real-time demand
+changes. By focusing on recent data and continuously generating
+forecasts, it enables timely responses to anomalies in critical
+scenarios.
+
+This section provides various recipes for performing real-time anomaly
+detection using TimeGPT, offering users the ability to detect outliers
+and unusual patterns as they emerge, ensuring prompt intervention in
+time-sensitive situations.
+
+This section covers:
+
+- [Online anomaly
+ detection](https://docs.nixtla.io/docs/capabilities-online-anomaly-detection-quickstart)
+
+- [How to adjust the detection
+ process](https://docs.nixtla.io/docs/capabilities-online-anomaly-detection-adjusting_detection_process.html)
+
+- [Univariate vs. multiseries anomaly
+ detection](https://docs.nixtla.io/docs/capabilities-online-anomaly-detection-univariate_vs_multivariate_anomaly_detection)
+
diff --git a/nixtla/docs/capabilities/online-anomaly-detection/quickstart.html.mdx b/nixtla/docs/capabilities/online-anomaly-detection/quickstart.html.mdx
new file mode 100644
index 00000000..f5debe49
--- /dev/null
+++ b/nixtla/docs/capabilities/online-anomaly-detection/quickstart.html.mdx
@@ -0,0 +1,116 @@
+---
+output-file: quickstart.html
+title: Introduction to Online (Real-Time) Anomaly Detection
+---
+
+
+In this notebook, we introduce the `detect_anomalies_online` method. You
+will learn how to quickly start using this new endpoint and understand
+its key differences from the historical anomaly detection endpoint. New
+features include: \* More flexibility and control over the anomaly
+detection process \* Perform univariate and multivariate anomaly
+detection \* Detect anomalies on stream data
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/capabilities/online-anomaly-detection/01_quickstart.ipynb)
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+import matplotlib.pyplot as plt
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, set the `base_url` argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 1. Dataset
+
+In this notebook, we use a minute-level time series dataset that
+monitors server usage. This is a good example of a streaming data
+scenario, as the task is to detect server failures or downtime.
+
+```python
+df = pd.read_csv('https://datasets-nixtla.s3.us-east-1.amazonaws.com/machine-1-1.csv', parse_dates=['ts'])
+```
+
+We observe that the time series remains stable during the initial
+period; however, a spike occurs in the last 20 steps, indicating an
+anomalous behavior. Our goal is to capture this abnormal jump as soon as
+it appears. Let’s see how the real-time anomaly detection capability of
+TimeGPT performs in this scenario!
+
+
+
+## 2. Detect anomalies in real time
+
+The `detect_anomalies_online` method detect anomalies in a time series
+leveraging TimeGPT’s forecast power. It uses the forecast error in
+deciding the anomalous step so you can specify and tune the parameters
+like that of the `forecast` method. This function will return a
+dataframe that contains anomaly flags and anomaly score (its absolute
+value quantifies the abnormality of the value).
+
+To perfom real-time anomaly detection, set the following parameters:
+
+- `df`: A pandas DataFrame containing the time series data.
+- `time_col`: The column that identifies the datestamp.
+- `target_col`: The variable to forecast.
+- `h`: Horizon is the number of steps ahead to make forecast.
+- `freq`: The frequency of the time series in Pandas format.
+- `level`: Percentile of scores distribution at which the threshold is
+ set, controlling how strictly anomalies are flagged. Default at 99%.
+- `detection_size`: The number of steps to analyze for anomaly at the
+ end of time series.
+
+```python
+anomaly_online = nixtla_client.detect_anomalies_online(
+ df,
+ time_col='ts',
+ target_col='y',
+ freq='min', # Specify the frequency of the data
+ h=10, # Specify the forecast horizon
+ level=99, # Set the confidence level for anomaly detection
+ detection_size=100 # How many steps you want for analyzing anomalies
+)
+anomaly_online.tail()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Calling Online Anomaly Detector Endpoint...
+```
+
+| | unique_id | ts | y | TimeGPT | anomaly | anomaly_score | TimeGPT-hi-99 | TimeGPT-lo-99 |
+|----|----|----|----|----|----|----|----|----|
+| 95 | machine-1-1_y_29 | 2020-02-01 22:11:00 | 0.606017 | 0.544625 | True | 18.463266 | 0.553161 | 0.536090 |
+| 96 | machine-1-1_y_29 | 2020-02-01 22:12:00 | 0.044413 | 0.570869 | True | -158.933850 | 0.579404 | 0.562333 |
+| 97 | machine-1-1_y_29 | 2020-02-01 22:13:00 | 0.038682 | 0.560303 | True | -157.474880 | 0.568839 | 0.551767 |
+| 98 | machine-1-1_y_29 | 2020-02-01 22:14:00 | 0.024355 | 0.521797 | True | -150.178240 | 0.530333 | 0.513261 |
+| 99 | machine-1-1_y_29 | 2020-02-01 22:15:00 | 0.044413 | 0.467860 | True | -127.848560 | 0.476396 | 0.459325 |
+
+> 📘 In this example, we use a detection size of 100 to illustrate the
+> anomaly detection process. In practice, using a smaller detection size
+> and running the detection more frequently improves granularity and
+> enables more timely identification of anomalies as they occur.
+
+From the plot, we observe that both anomalous periods were detected
+right as they arose. For further methods on improving detection accuracy
+and customizing anomaly detection, read our other tutorials on online
+anomaly detection.
+
+
+
+For an in-depth analysis of the `detect_anomalies_online` method, refer
+to the tutorial (coming soon).
+
diff --git a/nixtla/docs/capabilities/online-anomaly-detection/univariate_vs_multivariate_anomaly_detection.html.mdx b/nixtla/docs/capabilities/online-anomaly-detection/univariate_vs_multivariate_anomaly_detection.html.mdx
new file mode 100644
index 00000000..148ab82a
--- /dev/null
+++ b/nixtla/docs/capabilities/online-anomaly-detection/univariate_vs_multivariate_anomaly_detection.html.mdx
@@ -0,0 +1,157 @@
+---
+output-file: univariate_vs_multivariate_anomaly_detection.html
+title: Univariate vs. Multivariate Anomaly Detection
+---
+
+
+In this notebook, we show how to detect anomalies across multiple time
+series using the multivariate method. This method is great for
+situations where you have several sensors or related time series. We
+also explain how it works differently from the univariate method.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/capabilities/online-anomaly-detection/03_univariate_vs_multivariate_anomaly_detection.ipynb)
+
+```python
+import numpy as np
+import pandas as pd
+import matplotlib.pyplot as plt
+from nixtla import NixtlaClient
+```
+
+
+```python
+# Utility function to plot anomalies
+def plot_anomalies(df, unique_ids, rows, cols):
+ fig, axes = plt.subplots(rows, cols, figsize=(12, rows * 2))
+ for i, (ax, uid) in enumerate(zip(axes.flatten(), unique_ids)):
+ filtered_df = df[df['unique_id'] == uid]
+ ax.plot(filtered_df['ts'], filtered_df['y'], color='navy', alpha=0.8, label='y')
+ ax.plot(filtered_df['ts'], filtered_df['TimeGPT'], color='orchid', alpha=0.7, label='TimeGPT')
+ ax.scatter(filtered_df.loc[filtered_df['anomaly'] == 1, 'ts'], filtered_df.loc[filtered_df['anomaly'] == 1, 'y'], color='orchid', label='Anomalies Detected')
+ ax.set_title(f"Unique_id: {uid}", fontsize=8); ax.tick_params(axis='x', labelsize=6)
+ fig.legend(loc='upper center', ncol=3, fontsize=8, labels=['y', 'TimeGPT', 'Anomaly'])
+ plt.tight_layout(rect=[0, 0, 1, 0.95])
+ plt.show()
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, set the `base_url` argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 1. Dataset
+
+In this notebook example from SMD dataset. SMD (Server Machine Dataset)
+is a benchmark dataset for anomaly detection with multiple time series.
+It monitors abnormal patterns in server machine data.
+
+Here, we use a set of monitoring data from a single server
+machine(machine-1-1) that has 38 time series. Each time series
+represents a different metric being monitored, such as CPU usage, memory
+usage, disk I/O, and network I/O.
+
+```python
+df = pd.read_csv('https://datasets-nixtla.s3.us-east-1.amazonaws.com/SMD_test.csv', parse_dates=['ts'])
+df.unique_id.nunique()
+```
+
+``` text
+38
+```
+
+## 2. Univariate vs. Multivariate Method
+
+### 2.1 Univariate Method
+
+Univariate anomaly detection analyzes each time series independently,
+flagging anomalies based on deviations from its historical patterns.
+This method is effective for detecting issues within a single metric but
+ignores dependencies across multiple series. As a result, it may miss
+collective anomalies or flag irrelevant ones in scenarios where
+anomalies arise from patterns across multiple series, such as
+system-wide failures, correlated financial metrics, or interconnected
+processes. That’s when multivariate anomaly detection comes into play.
+
+```python
+anomaly_online = nixtla_client.detect_anomalies_online(
+ df[['ts', 'y', 'unique_id']],
+ time_col='ts',
+ target_col='y',
+ freq='h',
+ h=24,
+ level=95,
+ detection_size=475,
+ threshold_method = 'univariate' # Specify the threshold_method as 'univariate'
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+WARNING:nixtla.nixtla_client:Detection size is large. Using the entire series to compute the anomaly threshold...
+INFO:nixtla.nixtla_client:Calling Online Anomaly Detector Endpoint...
+```
+
+```python
+display_ids = ['machine-1-1_y_0', 'machine-1-1_y_1', 'machine-1-1_y_6', 'machine-1-1_y_29']
+plot_anomalies(anomaly_online, display_ids, rows=2, cols=2)
+```
+
+
+
+### 2.2 Multivariate Method
+
+The multivariate anomaly detection method considers multiple time series
+simultaneously. Instead of treating each series in isolation, it
+accumulates the anomaly scores for the same time step across all series
+and determines whether the step is anomalous based on the combined
+score. This method is particularly useful in scenarios where anomalies
+are only significant when multiple series collectively indicate an
+issue. To apply multivariate detection, simply set the
+`threshold_method` parameter to `multivariate`.
+
+We can see that the anomalies detected for each time series occur at the
+same time step since they rely on the accumulated error across all
+series.
+
+```python
+anomaly_online_multi = nixtla_client.detect_anomalies_online(
+ df[['ts', 'y', 'unique_id']],
+ time_col='ts',
+ target_col='y',
+ freq='h',
+ h=24,
+ level=95,
+ detection_size=475,
+ threshold_method = 'multivariate' # Specify the threshold_method as 'multivariate'
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+WARNING:nixtla.nixtla_client:Detection size is large. Using the entire series to compute the anomaly threshold...
+INFO:nixtla.nixtla_client:Calling Online Anomaly Detector Endpoint...
+```
+
+```python
+plot_anomalies(anomaly_online_multi, display_ids, rows=2, cols=2)
+```
+
+
+
+> 📘 In multiseries anomaly detection, error scores from all time series
+> are aggregated at each time step, and a threshold is applied to
+> identify significant deviations. If the aggregated error exceeds the
+> threshold, the time step is flagged as anomalous across all series,
+> capturing system-wide patterns.
+
diff --git a/nixtla/docs/deployment/azure_ai.html.mdx b/nixtla/docs/deployment/azure_ai.html.mdx
new file mode 100644
index 00000000..4528f9bb
--- /dev/null
+++ b/nixtla/docs/deployment/azure_ai.html.mdx
@@ -0,0 +1,52 @@
+---
+description: >-
+ The foundational models for time series by Nixtla can be deployed on your
+ Azure subscription. This page explains how to easily get started with
+ TimeGEN-1 deployed as an Azure AI endpoint. If you use the `nixtla` library,
+ it should be a drop-in replacement where you only need to change the client
+ parameters (endpoint URL, API key, model name).
+output-file: azure_ai.html
+title: AzureAI
+---
+
+
+## Deploying TimeGEN-1
+
+## Using the model
+
+Once your model is deployed and provided that you have the relevant
+permissions, consuming it will basically be the same process as for a
+Nixtla endpoint.
+
+To run the examples below, you will need to define the following
+environment variables:
+
+- `AZURE_AI_NIXTLA_BASE_URL` is your api URL, should be of the form
+ `https://your-endpoint.inference.ai.azure.com/`.
+- `AZURE_AI_NIXTLA_API_KEY` is your authentication key.
+
+## How to use
+
+Just import the library, set your credentials, and start forecasting in
+two lines of code!
+
+```bash
+pip install nixtla
+```
+
+
+```python
+import os
+from nixtla import NixtlaClient
+
+base_url = os.environ["AZURE_AI_NIXTLA_BASE_URL"]
+api_key = os.environ["AZURE_AI_NIXTLA_API_KEY"]
+model = "azureai"
+
+nixtla_client = NixtlaClient(api_key=api_key, base_url=base_url)
+nixtla_client.forecast(
+ ...,
+ model=model,
+)
+```
+
diff --git a/nixtla/docs/getting-started/21_polars_quickstart_files/figure-markdown_strict/cell-10-output-1.png b/nixtla/docs/getting-started/21_polars_quickstart_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..1b26f156
Binary files /dev/null and b/nixtla/docs/getting-started/21_polars_quickstart_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/nixtla/docs/getting-started/21_polars_quickstart_files/figure-markdown_strict/cell-12-output-1.png b/nixtla/docs/getting-started/21_polars_quickstart_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..d4ffdb57
Binary files /dev/null and b/nixtla/docs/getting-started/21_polars_quickstart_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/nixtla/docs/getting-started/21_polars_quickstart_files/figure-markdown_strict/cell-13-output-2.png b/nixtla/docs/getting-started/21_polars_quickstart_files/figure-markdown_strict/cell-13-output-2.png
new file mode 100644
index 00000000..0584d27a
Binary files /dev/null and b/nixtla/docs/getting-started/21_polars_quickstart_files/figure-markdown_strict/cell-13-output-2.png differ
diff --git a/nixtla/docs/getting-started/21_polars_quickstart_files/figure-markdown_strict/cell-8-output-1.png b/nixtla/docs/getting-started/21_polars_quickstart_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..fc49b593
Binary files /dev/null and b/nixtla/docs/getting-started/21_polars_quickstart_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/nixtla/docs/getting-started/22_azure_quickstart_files/figure-markdown_strict/cell-7-output-1.png b/nixtla/docs/getting-started/22_azure_quickstart_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..ac0fdc2f
Binary files /dev/null and b/nixtla/docs/getting-started/22_azure_quickstart_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/nixtla/docs/getting-started/22_azure_quickstart_files/figure-markdown_strict/cell-9-output-1.png b/nixtla/docs/getting-started/22_azure_quickstart_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..350b8f8a
Binary files /dev/null and b/nixtla/docs/getting-started/22_azure_quickstart_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/nixtla/docs/getting-started/2_quickstart_files/figure-markdown_strict/cell-10-output-1.png b/nixtla/docs/getting-started/2_quickstart_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..350b8f8a
Binary files /dev/null and b/nixtla/docs/getting-started/2_quickstart_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/nixtla/docs/getting-started/2_quickstart_files/figure-markdown_strict/cell-12-output-1.png b/nixtla/docs/getting-started/2_quickstart_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..c89e5ac3
Binary files /dev/null and b/nixtla/docs/getting-started/2_quickstart_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/nixtla/docs/getting-started/2_quickstart_files/figure-markdown_strict/cell-13-output-2.png b/nixtla/docs/getting-started/2_quickstart_files/figure-markdown_strict/cell-13-output-2.png
new file mode 100644
index 00000000..bfc31fd1
Binary files /dev/null and b/nixtla/docs/getting-started/2_quickstart_files/figure-markdown_strict/cell-13-output-2.png differ
diff --git a/nixtla/docs/getting-started/2_quickstart_files/figure-markdown_strict/cell-8-output-1.png b/nixtla/docs/getting-started/2_quickstart_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..ac0fdc2f
Binary files /dev/null and b/nixtla/docs/getting-started/2_quickstart_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/nixtla/docs/getting-started/7_why_timegpt_files/figure-markdown_strict/cell-20-output-1.png b/nixtla/docs/getting-started/7_why_timegpt_files/figure-markdown_strict/cell-20-output-1.png
new file mode 100644
index 00000000..7603cf79
Binary files /dev/null and b/nixtla/docs/getting-started/7_why_timegpt_files/figure-markdown_strict/cell-20-output-1.png differ
diff --git a/nixtla/docs/getting-started/azure_quickstart.html.mdx b/nixtla/docs/getting-started/azure_quickstart.html.mdx
new file mode 100644
index 00000000..d38e013f
--- /dev/null
+++ b/nixtla/docs/getting-started/azure_quickstart.html.mdx
@@ -0,0 +1,175 @@
+---
+description: >-
+ TimeGEN-1 is TimeGPT optimized for the Azure infrastructure. It is a
+ production ready, generative pretrained transformer for time series. It's
+ capable of accurately predicting various domains such as retail, electricity,
+ finance, and IoT with just a few lines of code 🚀.
+output-file: azure_quickstart.html
+title: TimeGEN-1 Quickstart (Azure)
+---
+
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/getting-started/22_azure_quickstart.ipynb)
+
+## Step 1: Set up a TimeGEN-1 endpoint account and generate your API key on Azure
+
+- Go to [ml.azure.com](https://ml.azure.com/)
+- Sign in or create an account at Microsoft
+- Click on ‘Models’ in the sidebar
+- Search for ‘TimeGEN’ in the model catalog
+- Select TimeGEN-1
+
+
+
+Azure Model Catalog landing page. Search
+for forecast and TimeGEN-1 comes up as the only option.
+
+
+- Click ‘Deploy’ and this will create an Endpoint
+
+
+
+TimeGEN-1 in the model catalog. Cursor is
+on the Deploy button indicating what to select to deploy
+TimeGEN-1.
+
+
+- Go to ‘Endpoint’ in the sidebar and you will see your TimeGEN-1
+ endpoint there
+- In that Endpoint are the base URL and API Key you will use
+
+
+
+Endpoint is highlighted in the side
+panel. The main panel shows the endpoint and API key for the TimeGEN-1
+endpoint, with buttons where you can copy the information.
+
+
+## Step 2: Install Nixtla
+
+In your favorite Python development environment:
+
+Install `nixtla` with `pip`:
+
+```shell
+pip install nixtla
+```
+
+## Step 3: Import the Nixtla TimeGPT client
+
+```python
+from nixtla import NixtlaClient
+```
+
+You can instantiate the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class providing your authentication API key.
+
+```python
+nixtla_client = NixtlaClient(
+ base_url = "YOUR_BASE_URL",
+ api_key = "YOUR_API_KEY"
+)
+```
+
+## Step 4: Start making forecasts!
+
+Now you can start making forecasts! Let’s import an example using the
+classic `AirPassengers` dataset. This dataset contains the monthly
+number of airline passengers in Australia between 1949 and 1960. First,
+load the dataset and plot it:
+
+```python
+import pandas as pd
+```
+
+
+```python
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/air_passengers.csv')
+df.head()
+```
+
+| | timestamp | value |
+|-----|------------|-------|
+| 0 | 1949-01-01 | 112 |
+| 1 | 1949-02-01 | 118 |
+| 2 | 1949-03-01 | 132 |
+| 3 | 1949-04-01 | 129 |
+| 4 | 1949-05-01 | 121 |
+
+```python
+nixtla_client.plot(df, time_col='timestamp', target_col='value')
+```
+
+
+
+> 📘 Data Requirements
+>
+> - Make sure the target variable column does not have missing or
+> non-numeric values.
+> - Do not include gaps/jumps in the datestamps (for the given
+> frequency) between the first and late datestamps. The forecast
+> function will not impute missing dates.
+> - The format of the datestamp column should be readable by Pandas
+> (see [this
+> link](https://pandas.pydata.org/docs/reference/api/pandas.to_datetime.html)
+> for more details).
+>
+> For further details go to [Data
+> Requirements](https://docs.nixtla.io/docs/getting-started-data_requirements).
+
+> 👍 Save figures made with TimeGEN
+>
+> The `plot` method automatically displays figures when in a notebook
+> environment. To save figures locally, you can do:
+>
+> `fig = nixtla_client.plot(df, time_col='timestamp', target_col='value')`
+>
+> `fig.savefig('plot.png', bbox_inches='tight')`
+
+### Make forecasts
+
+Next, forecast the next 12 months using the SDK `forecast` method. Set
+the following parameters:
+
+- `df`: A pandas DataFrame containing the time series data.
+- `h`: Horizons is the number of steps ahead to forecast.
+- `freq`: The frequency of the time series in Pandas format. See
+ [pandas’ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+ (If you don’t provide any frequency, the SDK will try to infer it)
+- `time_col`: The column that identifies the datestamp.
+- `target_col`: The variable to forecast.
+
+```python
+timegen_fcst_df = nixtla_client.forecast(df=df, h=12, freq='MS', time_col='timestamp', target_col='value')
+timegen_fcst_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| | timestamp | TimeGPT |
+|-----|------------|------------|
+| 0 | 1961-01-01 | 437.837921 |
+| 1 | 1961-02-01 | 426.062714 |
+| 2 | 1961-03-01 | 463.116547 |
+| 3 | 1961-04-01 | 478.244507 |
+| 4 | 1961-05-01 | 505.646484 |
+
+```python
+nixtla_client.plot(df, timegen_fcst_df, time_col='timestamp', target_col='value')
+```
+
+
+
diff --git a/nixtla/docs/getting-started/data_requirements.html.mdx b/nixtla/docs/getting-started/data_requirements.html.mdx
new file mode 100644
index 00000000..6f365203
--- /dev/null
+++ b/nixtla/docs/getting-started/data_requirements.html.mdx
@@ -0,0 +1,255 @@
+---
+description: This section explains the data requirements for `TimeGPT`.
+output-file: data_requirements.html
+title: Data Requirements
+---
+
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/getting-started/5_data_requirements.ipynb)
+
+`TimeGPT` accepts `pandas` and `polars` dataframes in [long
+format](https://www.theanalysisfactor.com/wide-and-long-data/#comments)
+with the following necessary columns:
+
+- `ds` (timestamp): timestamp in format `YYYY-MM-DD` or
+ `YYYY-MM-DD HH:MM:SS`.
+- `y` (numeric): The target variable to forecast.
+
+(Optionally, you can also pass a DataFrame without the `ds` column as
+long as it has DatetimeIndex)
+
+`TimeGPT` also works with distributed dataframes like `dask`, `spark`
+and `ray`.
+
+You can also include exogenous features in the DataFrame as additional
+columns. For more information, follow this
+[tutorial](https://docs.nixtla.io/docs/tutorials-exogenous_variables).
+
+Below is an example of a valid input dataframe for `TimeGPT`.
+
+```python
+import pandas as pd
+
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/air_passengers.csv')
+df.head()
+```
+
+| | timestamp | value |
+|-----|------------|-------|
+| 0 | 1949-01-01 | 112 |
+| 1 | 1949-02-01 | 118 |
+| 2 | 1949-03-01 | 132 |
+| 3 | 1949-04-01 | 129 |
+| 4 | 1949-05-01 | 121 |
+
+Note that in this example, the `ds` column is named `timestamp` and the
+`y` column is named `value`. You can either:
+
+1. Rename the columns to `ds` and `y`, respectively, or
+
+2. Keep the current column names and specify them when using any method
+ from the
+ [`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+ class with the `time_col` and `target_col` arguments.
+
+For example, when using the `forecast` method from the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class, you must instantiate the class and then specify the columns names
+as follows.
+
+```python
+from nixtla import NixtlaClient
+
+nixtla_client = NixtlaClient(
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+
+```python
+fcst = nixtla_client.forecast(df=df, h=12, time_col='timestamp', target_col='value')
+fcst.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Querying model metadata...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| | timestamp | TimeGPT |
+|-----|------------|-----------|
+| 0 | 1961-01-01 | 437.83792 |
+| 1 | 1961-02-01 | 426.06270 |
+| 2 | 1961-03-01 | 463.11655 |
+| 3 | 1961-04-01 | 478.24450 |
+| 4 | 1961-05-01 | 505.64648 |
+
+In this example, the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+is infereing the frequency, but you can explicitly specify it with the
+`freq` argument.
+
+To learn more about how to instantiate the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class, refer to the [TimeGPT
+Quickstart](https://docs.nixtla.io/docs/getting-started-timegpt_quickstart)
+
+## Multiple Series
+
+If you’re working with multiple time series, make sure that each series
+has a unique identifier. You can name this column `unique_id` or specify
+its name using the `id_col` argument when calling any method from the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class. This column should be a string, integer, or category.
+
+In this example, we have five series representing hourly electricity
+prices in five different markets. The columns already have the default
+names, so it’s unnecessary to specify the `id_col`, `time_col`, or
+`target_col` arguments. If your columns have different names, specify
+these arguments as required.
+
+```python
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short.csv')
+df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|---------------------|-------|
+| 0 | BE | 2016-10-22 00:00:00 | 70.00 |
+| 1 | BE | 2016-10-22 01:00:00 | 37.10 |
+| 2 | BE | 2016-10-22 02:00:00 | 37.10 |
+| 3 | BE | 2016-10-22 03:00:00 | 44.75 |
+| 4 | BE | 2016-10-22 04:00:00 | 37.10 |
+
+```python
+fcst = nixtla_client.forecast(df=df, h=24) # use id_col, time_col and target_col here if needed.
+fcst.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: h
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Querying model metadata...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| | unique_id | ds | TimeGPT |
+|-----|-----------|---------------------|-----------|
+| 0 | BE | 2016-12-31 00:00:00 | 45.190582 |
+| 1 | BE | 2016-12-31 01:00:00 | 43.244987 |
+| 2 | BE | 2016-12-31 02:00:00 | 41.958897 |
+| 3 | BE | 2016-12-31 03:00:00 | 39.796680 |
+| 4 | BE | 2016-12-31 04:00:00 | 39.204865 |
+
+When working with a large number of time series, consider using a
+[distributed computing
+framework](https://docs.nixtla.io/docs/tutorials-computing_at_scale) to
+handle the data efficiently. `TimeGPT` supports frameworks such as
+[Spark](https://docs.nixtla.io/docs/tutorials-spark),
+[Dask](https://docs.nixtla.io/docs/tutorials-dask), and
+[Ray](https://docs.nixtla.io/docs/tutorials-ray).
+
+## Exogenous Variables
+
+`TimeGPT` also accepts exogenous variables. You can add exogenous
+variables to your dataframe by including additional columns after the
+`y` column.
+
+```python
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short-with-ex-vars.csv')
+df.head()
+```
+
+| | unique_id | ds | y | Exogenous1 | Exogenous2 | day_0 | day_1 | day_2 | day_3 | day_4 | day_5 | day_6 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | BE | 2016-10-22 00:00:00 | 70.00 | 57253.0 | 49593.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 1 | BE | 2016-10-22 01:00:00 | 37.10 | 51887.0 | 46073.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 2 | BE | 2016-10-22 02:00:00 | 37.10 | 51896.0 | 44927.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 3 | BE | 2016-10-22 03:00:00 | 44.75 | 48428.0 | 44483.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 4 | BE | 2016-10-22 04:00:00 | 37.10 | 46721.0 | 44338.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+
+When using exogenous variables, you also need to provide its future
+values.
+
+```python
+future_ex_vars_df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short-future-ex-vars.csv')
+future_ex_vars_df.head()
+```
+
+| | unique_id | ds | Exogenous1 | Exogenous2 | day_0 | day_1 | day_2 | day_3 | day_4 | day_5 | day_6 |
+|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | BE | 2016-12-31 00:00:00 | 70318.0 | 64108.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 1 | BE | 2016-12-31 01:00:00 | 67898.0 | 62492.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 2 | BE | 2016-12-31 02:00:00 | 68379.0 | 61571.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 3 | BE | 2016-12-31 03:00:00 | 64972.0 | 60381.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 4 | BE | 2016-12-31 04:00:00 | 62900.0 | 60298.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+
+```python
+fcst = nixtla_client.forecast(df=df, X_df=future_ex_vars_df, h=24)
+fcst.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: h
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Using future exogenous features: ['Exogenous1', 'Exogenous2', 'day_0', 'day_1', 'day_2', 'day_3', 'day_4', 'day_5', 'day_6']
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| | unique_id | ds | TimeGPT |
+|-----|-----------|---------------------|-----------|
+| 0 | BE | 2016-12-31 00:00:00 | 51.632830 |
+| 1 | BE | 2016-12-31 01:00:00 | 45.750877 |
+| 2 | BE | 2016-12-31 02:00:00 | 39.650543 |
+| 3 | BE | 2016-12-31 03:00:00 | 34.000072 |
+| 4 | BE | 2016-12-31 04:00:00 | 33.785370 |
+
+To learn more about how to use exogenous variables with `TimeGPT`,
+consult the [Exogenous
+Variables](https://docs.nixtla.io/docs/tutorials-exogenous_variables)
+tutorial.
+
+## Important Considerations
+
+When using `TimeGPT`, the data cannot contain missing values. This means
+that for every series, there should be no gaps in the timestamps and no
+missing values in the target variable.
+
+For more, please refer to the tutorial on [Dealing with Missing Values
+in
+TimeGPT](https://docs.nixtla.io/docs/tutorials-dealing_with_missing_values_in_timegpt).
+
+### Minimum Data Requirements (for AzureAI)
+
+`TimeGPT` currently supports any amount of data for generating point
+forecasts. That is, the minimum size per series to expect results from
+this call `nixtla_client.forecast(df=df, h=h, freq=freq)` is one,
+regardless of the frequency.
+
+For Azure AI, when using the arguments `level`, `finetune_steps`, `X_df`
+(exogenous variables), or `add_history`, the API requires a minimum
+number of data points depending on the frequency. Here are the minimum
+sizes for each frequency:
+
+| Frequency | Minimum Size |
+|------------------------------------------------------|--------------|
+| Hourly and subhourly (e.g., “H”, “min”, “15T”) | 1008 |
+| Daily (“D”) | 300 |
+| Weekly (e.g., “W-MON”,…, “W-SUN”) | 64 |
+| Monthly and other frequencies (e.g., “M”, “MS”, “Y”) | 48 |
+
+For cross-validation, you need to consider these numbers as well as the
+forecast horizon (`h`), the number of windows (`n_windows`), and the gap
+between windows (`step_size`). Thus, the minimum number of observations
+per series in this case would be determined by the following
+relationship:
+
+Minimum number described previously + h + step_size + (n_windows - 1)
+
diff --git a/nixtla/docs/getting-started/faq.html.mdx b/nixtla/docs/getting-started/faq.html.mdx
new file mode 100644
index 00000000..fa28258c
--- /dev/null
+++ b/nixtla/docs/getting-started/faq.html.mdx
@@ -0,0 +1,487 @@
+---
+output-file: faq.html
+title: FAQ
+---
+
+
+Commonly asked questions about TimeGPT
+
+## Table of contents
+
+- [TimeGPT](#timegpt)
+- [TimeGPT API Key](#timegpt-api-key)
+- [Features and Capabilities](#features-and-capabilities)
+- [Fine-tuning](#finetuning)
+- [Pricing and Billing](#pricing-and-billing)
+- [Privacy and Security](#privacy-and-security)
+- [Troubleshooting](#troubleshooting)
+- [Additional Support](#additional-support)
+
+## TimeGPT
+
+What is TimeGPT?
+
+`TimeGPT` is the first foundation model for time series forecasting. It
+can produce accurate forecasts for new time series across a diverse
+array of domains using only historical values as inputs. The model
+“reads” time series data sequentially from left to right, similarly to
+how humans read a sentence. It looks at windows of past data, which we
+can think of as “tokens”, and then predicts what comes next. This
+prediction is based on patterns the model identifies and that it
+extrapolates into the future. Beyond forecasting, `TimeGPT` supports
+other time series related tasks, such as what-if-scenarios, anomaly
+detection, and more.
+
+Is TimeGPT based on a Large Language Model (LLM)?
+
+No, `TimeGPT` is not based on any large language model. While it follows
+the same principle of training a large transformer model on a vast
+dataset, its architecture is specifically designed to handle time series
+data and it has been trained to minimize forecasting errors.
+
+How do I get started with TimeGPT?
+
+To get started with `TimeGPT`, you need to register for an account
+[here](https://dashboard.nixtla.io/). You will receive an email asking
+you to confirm your signup. After confirming, you will be able to access
+your dashboard, which contains the details of your account.
+
+How accessible is TimeGPT and what are the usage costs?
+
+For a more in-depth understanding of `TimeGPT`, please refer to the
+[research paper](https://arxiv.org/pdf/2310.03589.pdf). While certain
+aspects of the model’s architecture remain confidential, registration
+for `TimeGPT` is open to all. New users receive \$1,000 USD in free
+credits and subsequent usage fees are based on token consumption. For
+more details, please refer to the [Pricing and
+Billing](#pricing-and-billing) section
+
+How can I use TimeGPT?
+
+- Through the [Python SDK](https://github.com/Nixtla/nixtla)
+
+- Via the `TimeGPT` API. For instructions on how to call the API using
+ different languages, please refer to the [API
+ documentation](https://docs.nixtla.io/reference/timegpt_timegpt_post)
+
+Both methods require you to have a [API key](#timegpt-api-key), which is
+obtained upon registration and can be found in your dashboard under
+`API Keys`.
+
+## TimeGPT API Key
+
+What is an API key?
+
+An API key is a unique string of characters that serves as a key to
+authenticate your requests when using the Nixtla SDK. It ensures that
+the person making the requests is authorized to do so.
+
+Where can I get an API key?
+
+Upon registration, you will receive an API key that can be found in your
+[dashboard](https://dashboard.nixtla.io/) under `API Keys`. Remember
+that your API key is personal and should not be shared with anyone.
+
+How do I use my API key?
+
+To integrate your API key into your development workflow, please refer
+to the tutorial on [Setting Up Your API
+Key](https://docs.nixtla.io/docs/getting-started-setting_up_your_api_key).
+
+How can I check the status of my API key?
+
+If you want to check the status of your API key, you can use the
+[`validate_api_key`
+method](https://nixtlaverse.nixtla.io/nixtla/nixtla_client.html#nixtlaclient-validate-api-key)
+of the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class.
+
+`nixtla_client = NixtlaClient( api_key = 'my_api_key_provided_by_nixtla' )`
+
+`nixtla_client.validate_api_key()`
+
+If your key is validating correctly, this will return
+
+``` text
+INFO:nixtla.nixtla_client:Happy Forecasting! :), If you have questions or need support, please email support@nixtla.io
+
+True
+```
+
+What if my API key isn’t validating?
+
+When you validate your API key and it returns `False`:
+
+- If you are targeting an Azure endpoint, getting `False` from the
+ [`NixtlaClient.validate_api_key`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient.validate_api_key)
+ method is expected. You can skip this step when taregting an Azure
+ endpoint and proceed diretly to forecasting instead.
+- If you are not taregting an Azure endpoint, then you should check
+ the following:
+ - Make sure you are using the latest version of the SDK (Python or
+ R).
+ - Check that your API key is active in your dashboard by visiting
+ https://dashboard.nixtla.io/
+ - Consider any firewalls your organization might have. There may
+ be restricted access. If so, you can whitelist our endpoint
+ https://api.nixtla.io/.
+ - To use Nixtla’s API, you need to let your system know that
+ our endpoint is ok, so it will let you access it.
+ Whitelisting the endpoint isn’t something that Nixtla can do
+ on our side. It’s something that needs to be done on the
+ user’s system. This is a bit of an [overview on
+ whitelisting](https://www.csoonline.com/article/569493/whitelisting-explained-how-it-works-and-where-it-fits-in-a-security-program.html).
+ - If you work in an organization, please work with an IT team.
+ They’re likely the ones setting the security and you can
+ talk with them to get it addressed. If you run your own
+ systems, then it’s something you should be able to update,
+ depending on the system you’re using.
+
+## Features and Capabilities
+
+What is the input to TimeGPT?
+
+`TimeGPT` accepts `pandas` dataframes in [long
+format](https://www.theanalysisfactor.com/wide-and-long-data/#comments)
+with the following necessary columns:
+
+- `ds` (timestamp): timestamp in format `YYYY-MM-DD` or
+ `YYYY-MM-DD HH:MM:SS`.
+- `y` (numeric): The target variable to forecast.
+
+(Optionally, you can also pass a DataFrame without the `ds` column as
+long as it has DatetimeIndex)
+
+`TimeGPT` also works with [distributed
+dataframes](https://docs.nixtla.io/docs/tutorials-computing_at_scale)
+like `dask`, `spark` and `ray`.
+
+Can TimeGPT handle multiple time series?
+
+Yes. For guidance on forecasting multiple time series at once, consult
+the [Multiple
+Series](https://docs.nixtla.io/docs/tutorials-multiple_series_forecasting)
+tutorial.
+
+Does TimeGPT support forecasting with exogenous variables?
+
+Yes. For instructions on how to incorporate exogenous variables to
+`TimeGPT`, see the [Exogenous
+Variables](https://docs.nixtla.io/docs/tutorials-exogenous_variables)
+tutorial. For incorporating calendar dates specifically, you may find
+the [Holidays and Special
+Dates](https://docs.nixtla.io/docs/tutorials-holidays_and_special_dates)
+tutorial useful. For categorical variables, refer to the [Categorical
+Variables](https://docs.nixtla.io/docs/tutorials-categorical_variables)
+tutorial.
+
+Can TimeGPT be used for anomaly detection?
+
+Yes. To learn how to use `TimeGPT` for anomaly detection, refer to the
+[Anomaly
+Detection](https://docs.nixtla.io/docs/capabilities-anomaly-detection-anomaly_detection)
+tutorial.
+
+Does TimeGPT support cross-validation?
+
+Yes. To learn how to use `TimeGPT` for cross-validation, refer to the
+[Cross-Validation](https://docs.nixtla.io/docs/tutorials-cross_validation)
+tutorial.
+
+Can TimeGPT be used to forecast historical data?
+
+Yes. To find out how to forecast historical data using `TimeGPT`, see
+the [Historical
+Forecast](https://docs.nixtla.io/docs/tutorials-historical_forecast)
+tutorial.
+
+Can TimeGPT be used for uncertainty quantification?
+
+Yes. For more information, explore the [Prediction
+Intervals](https://docs.nixtla.io/docs/tutorials-prediction_intervals)
+and [Quantile
+Forecasts](https://docs.nixtla.io/docs/tutorials-quantile_forecasts)
+tutorials.
+
+Can TimeGPT handle large datasets?
+
+Yes. When dealing with large datasets that contain hundreds of thousands
+or millions of time series, we recommend using a distributed backend.
+`TimeGPT` is compatible with several [distributed computing
+frameworks](https://docs.nixtla.io/docs/tutorials-computing_at_scale),
+including [Spark](https://docs.nixtla.io/docs/tutorials-spark),
+[Ray](https://docs.nixtla.io/docs/tutorials-ray), and
+[Dask](https://docs.nixtla.io/docs/tutorials-dask). Both the `TimeGPT`
+SDK and API don’t have a limit on the size of the dataset as long as a
+distributed backend is used.
+
+Can TimeGPT be used with limited/short data?
+
+`TimeGPT` supports any amount of data for generating point forecasts and
+is capable of producing results with just one observation per series.
+When using arguments such as `level`, `finetune_steps`, `X_df`
+(exogenous variables), or `add_history`, additional data points are
+necessary depending on the data frequency. For more details, please
+refer to the [Data
+Requirements](https://docs.nixtla.io/docs/getting-started-data_requirements)
+tutorial.
+
+What is the maximum forecast horizon allowed by TimeGPT?
+
+While `TimeGPT` does not have a maximum forecast horizon, its
+performance will decrease as the horizon increases. When the forecast
+horizon exceeds the season length of the data (for example, more than 12
+months for monthly data), you will get this message:
+`WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon. This may lead to less accurate forecasts. Please consider using a smaller horizon`.
+
+For details, refer to the tutorial on [Long Horizon in Time
+Series](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting).
+
+Can TimeGPT handle missing values?
+
+`TimeGPT` cannot handle missing values or series with irregular
+timestamps. For more information, see the [Forecasting Time Series with
+Irregular
+Timestamps](https://docs.nixtla.io/docs/capabilities-forecast-irregular_timestamps)
+and the [Dealing with Missing
+Values](https://docs.nixtla.io/docs/tutorials-dealing_with_missing_values_in_timegpt)
+tutorial.
+
+How can I plot the TimeGPT forecast?
+
+The
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class has a [`plot`
+method](https://nixtlaverse.nixtla.io/nixtla/nixtla_client.html#nixtlaclient-validate-token)
+that can be used to visualize the forecast. This method only works in
+interactive environments such as Jupyter notebooks and it doesn’t work
+on Python scripts.
+
+Does TimeGPT support polars?
+
+As of now, `TimeGPT` does not offer support for polars.
+
+Does TimeGPT produce stable predictions?
+
+`TimeGPT` is engineered for stability, ensuring consistent results for
+identical input data. This means that given the same dataset, the model
+will produce the same forecasts.
+
+Can TimeGPT forecast data with simple pattern such as a straight line or
+sine wave?
+
+While this is not the primary use case for `TimeGPT`, it is capable of
+generating solid results on simple data such as a straight line. While
+zero-shot predictions might not always meet expectations, a little help
+with fine-tuning allows TimeGPT to quickly grasp the trend and produce
+accurate forecasts. For more details, please refer to the [Improve
+Forecast Accuracy with
+TimeGPT](https://docs.nixtla.io/docs/tutorials-improve_forecast_accuracy_with_timegpt)
+tutorial.
+
+## Fine-tuning
+
+What is fine-tuning?
+
+`TimeGPT` was trained on the largest publicly available time series
+dataset, covering a wide range of domains such as finance, retail,
+healthcare, and more. This comprehensive training enables `TimeGPT` to
+produce accurate forecasts for new time series without additional
+training, a capability known as zero-shot learning.
+
+While the zero-shot model provides a solid baseline, the performance of
+`TimeGPT` can often be improved through fine-tuning. During this
+process, the `TimeGPT` model undergoes additional training using your
+specific dataset, starting from the pre-trained paramaters. The updated
+model then produces the forecasts. You can control the number of
+training iterations and the loss function for fine-tuning with the
+`finetune_steps` and the `finetune_loss` parameters in the `forecast`
+method from the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class, respectively.
+
+For a comprehensive guide on how to apply fine-tuning, please refer to
+the [fine-tuning](https://docs.nixtla.io/docs/tutorials-fine_tuning) and
+the [fine-tuning with a specific loss
+function](https://docs.nixtla.io/docs/tutorials-fine_tuning_with_a_specific_loss_function)
+tutorials.
+
+Do I have to fine-tune every series?
+
+No, you do not need to fine-tune every series individually. When using
+the `finetune_steps` parameter, the model undergoes fine-tuning across
+all series in your dataset simultaneously. This method uses a
+cross-learning approach, allowing the model to learn from multiple
+series at once, which can improve individual forecasts.
+
+Keep in mind that selecting the right number of fine-tuning steps may
+require some trial and error. As the number of fine-tuning steps
+increases, the model becomes more specialized to your dataset, but will
+take longer to train and may be more prone to overfitting.
+
+Can I save fine-tuned parameters?
+
+Yes! You can fine-tune the TimeGPT model, save it, and reuse it later.
+For detailed instructions, see our guide on [Re-using Fine-tuned
+Models](https://docs.nixtla.io/docs/tutorials-re_using_fine_tuned_models).
+
+## Pricing and Billing
+
+How does pricing work?
+
+See our [Pricing
+page](https://docs.nixtla.io/docs/getting-started-subscription_plans)
+for information about pricing.
+
+[Start for Free](https://dashboard.nixtla.io/) \*No credit card needed.
+
+For customized plan details and offerings, book a demo or contact us at
+`support@nixtla.io`.
+
+Are there free options or discounts?
+
+Yes! We provide some discounted options for academic research. If you
+would like to learn more, please email us at `support@nixtla.io`.
+
+What counts as an API call?
+
+An API call is a request made to TimeGPT to perform an action like
+forecasting or detecting anomalies. API Usage is as follows:
+
+### Forecasting:
+
+1. When not requesting historical forecasts (`add_history=False`)
+ - If you do not set `num_partitions`, all calls to perform
+ [forecasting](https://docs.nixtla.io/docs/getting-started-timegpt_quickstart),
+ [finetuning](https://docs.nixtla.io/docs/tutorials-fine_tuning),
+ or
+ [cross-validation](https://docs.nixtla.io/docs/tutorials-cross_validation)
+ increase the usage by 1. Note that addition of [exogenous
+ variables](https://docs.nixtla.io/docs/tutorials-exogenous_variables),
+ requesting [uncertainity
+ quantification](https://docs.nixtla.io/docs/tutorials-uncertainty_quantification)
+ or forecasting [multiple
+ series](https://docs.nixtla.io/docs/tutorials-multiple_series_forecasting)
+ does not increase the usage further.
+ - If the API call requires to send more than 200MB of data, the
+ API will return an error and will require you to use the
+ `num_partitions` parameter in order to partition your request.
+ Every partition will count as an API call, hence the usage will
+ increase by the value you set for `num_partitions` (e.g. for
+ num_partitions=2, the usage will increase by 2). If you set
+ `num_partitions`, all calls to perform forecasting, finetuning,
+ or cross-validation increase the usage by num_partitions.
+2. When requesting [in-sample
+ predictions](https://docs.nixtla.io/docs/tutorials-historical_forecast)
+ (`add_history=True`), the usage from #1 above is multipled by 2.
+
+**Examples**
+
+1. A user uses TimeGPT to forecast daily data, using the `timegpt-1`
+ model. How many API calls are made? (*Ans*: 1)
+2. A user calls the `cross_validation` method on a dataset. How many
+ API calls are made (*Ans*: 1)
+3. A user decides to forecast on a longer horizon, so they use the
+ `timegpt-1-long-horizon` model. How many API calls are made (*Ans*:
+ 1)
+4. A user needs to get the in-sample predicitons when forecasting using
+ `add_history=True`. How many API calls are made (*Ans*: 2)
+5. A user has a very large dataset, with a daily frequency, and they
+ must set `num_partitions=4` when forecasting. How many API calls are
+ made (*Ans*: 4)
+6. A user has to set `num_partitions=4` and is also interesed in
+ getting the in-sample predicitons (`add_history=True`) when
+ forecasting. How many API calls are made (*Ans*: 8)
+
+### Anomaly Detection:
+
+1. If you do not set `num_partitions`, all calls to perform [anomaly
+ detection](https://docs.nixtla.io/docs/capabilities-anomaly-detection-quickstart)
+ increase the usage by 1. Note that addition of [exogenous
+ variables](https://docs.nixtla.io/docs/capabilities-anomaly-detection-add_exogenous_variables)
+ does not increase the usage further.
+2. If the API call requires to send more than 200MB of data, the API
+ will return an error and will require you to use the
+ `num_partitions` parameter in order to partition your request. Every
+ partition will count as an API call, hence the usage will increase
+ by the value you set for `num_partitions` (e.g. for
+ num_partitions=2, the usage will increase by 2).
+
+How does billing work?
+
+Billing is done through Stripe. We’ve partnered with Stripe to handle
+all payment processing. You can view your invoices and payment history
+in your [dashboard](https://dashboard.nixtla.io/) under `Billing`.
+
+## Privacy and Security
+
+How do you ensure the privacy and security of my data?
+
+At Nixtla, we take your privacy and security very seriously. To ensure
+you are fully informed about our policies regarding your data, please
+refer to the following documents:
+
+- [Privacy Notice](https://docs.nixtla.io/docs/privacy-notice)
+
+- For the Python SDK, please review the [license
+ agreement](https://github.com/Nixtla/nixtla/blob/main/LICENSE).
+
+- For `TimeGPT`, please refer to our [terms and
+ conditions](https://docs.nixtla.io/docs/terms-and-conditions).
+
+In addtion, we are currently developing a self-hosted version of
+`TimeGPT`, tailored for the unique security requirements of enterprise
+data. This version is currently in beta. If you are interested in
+exploring this option, please contact us at `support@nixtla.io`.
+
+## Troubleshooting
+
+The following section contains some common errors and warnings
+
+Error message: Invalid API key
+
+```python
+ApiError: status_code: 401, body: {'data': None, 'message': 'Invalid API key', 'details': 'Key not found', 'code': 'A12', 'requestID': 'E7F2BBTB2P', 'support': 'If you have questions or need support, please email support@nixtla.io'}
+```
+
+**Solution:** This error occurs when your `TimeGPT` API key is either
+invalid or has not been set up correctly. Please use the
+`validate_api_key` method to verify it or make sure it was copied
+correctly from the `API Keys` section of your
+[dashboard](https://dashboard.nixtla.io/).
+
+Error message: Too many requests
+
+```python
+ApiError: status_code: 429, body: {'data': None, 'message': 'Too many requests', 'details': 'You need to add a payment method to continue using the API, do so from https://dashboard.nixtla.io', 'code': 'A21', 'requestID': 'NCJDK7KSJ6', 'support': 'If you have questions or need support, please email support@nixtla.io'}
+```
+
+**Solution:** This error occurs when you have exhausted your free
+credits and need to add a payment method to continue using `TimeGPT`.
+You can add a payment method in the `Billing` section of your
+[dashboard](https://dashboard.nixtla.io/).
+
+Error message: WriteTimeout
+
+**Solution:** If you encounter a `WriteTimeout` error, it means that the
+request has exceeded the allowable processing time. This is a common
+issue when working with large datasets. To fix this, consider increasing
+the `num_partitions` parameter in the [`forecast`
+method](https://nixtlaverse.nixtla.io/nixtla/nixtla_client.html#nixtlaclient-forecast)
+of the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class, or use a distributed backend if not already in use.
+
+## Additional Support
+
+If you have any more questions or need support, please reach out by:
+
+- Opening an [issue](https://github.com/Nixtla/nixtla/issues) on
+ GitHub for technical questions or bugs.
+- Sending an email to `support@nixtla.io` for general inquiries or
+ support.
+- Joining our
+ [Slack](https://join.slack.com/t/nixtlacommunity/shared_invite/zt-2ebtgjbip-QMSnvm6ED1NF5vi4xj_13Q)
+ community to connect with our team and the forecasting community.
+
diff --git a/nixtla/docs/getting-started/glossary.html.mdx b/nixtla/docs/getting-started/glossary.html.mdx
new file mode 100644
index 00000000..4687e6bb
--- /dev/null
+++ b/nixtla/docs/getting-started/glossary.html.mdx
@@ -0,0 +1,162 @@
+---
+output-file: glossary.html
+title: Glossary
+---
+
+
+> These are some key concepts related to time series forecasting,
+> designed to help you better understand and leverage the capabilities
+> of TimeGPT.
+
+- [Time Series](#time-series)
+- [Forecasting](#forecasting)
+- [Foundation Model](#foundation-model)
+- [TimeGPT](#timegpt)
+- [Tokens](#tokens)
+- [Fine-tuning](#fine-tuning)
+- [Historical Forecasts](#historical-forecasts)
+- [Anomaly Detection](#anomaly-detection)
+- [Time Series Cross-Validation](#time-series-cross-validation)
+- [Exogenous Variables](#exogenous-variables)
+
+## Time Series
+
+A time series is a sequence of data points indexed by time, used to
+model phenomena that changes over time, such as stock prices,
+temperature, or product sales. A time series can generally be thought of
+as comprising the following components:
+
+- **Trend**: The consistent, long-term direction of the data, whether
+ upward or downward. It reflects the persistent, overall movement in
+ the series over time.
+
+- **Seasonality**: A repeated cycle around a known and fixed period.
+
+- **Remainder**: The residuals or random noise left in the data after
+ the trend and seasonal effects have been accounted for.
+
+## Forecasting
+
+Forecasting is the process of predicting the future values of a time
+series based on historical data. It plays a crucial role in the
+decision-making process across various fields such as finance,
+healthcare, retail, and economics, among others.
+
+Forecasting can use a variety of approaches, from statistical approaches
+to novel techniques such as machine learning, deep learning, and
+foundation models. These models can be further classified into
+univariate and multivariate models, depending on the number of variables
+used to make the predictions, or local or global models, with local
+models estimating parameters independently for each series and global
+models estimating parameters jointly across multiple series.
+
+Forecasts themselves can be presented as point forecasts, which predict
+a single future value, or as probabilistic forecasts, which provide a
+full probability distribution of future values, and hence, providing a
+measure of uncertainty.
+
+## Foundation Model
+
+Foundation model refers to a type of large, pre-trained model that can
+be adapted to a wide range of tasks, including time series forecasting.
+Originally developed for domains such as natural language processing and
+computer vision, foundation models are now increasingly applied to
+sequential data like time series. These models are typically trained on
+extensive datasets, capturing complex patterns and dependencies that can
+be fine-tuned for specific tasks.
+
+## TimeGPT
+
+Developed by Nixtla, `TimeGPT` is the first foundation model for time
+series forecasting. `TimeGPT` was trained on billions of observations
+from publicly available datasets across multiple domains and can produce
+accurate forecasts for new time series without additional training,
+using only historical values as inputs. The model ‘reads’ time series
+data similarly to how humans read a sentence—sequentially from left to
+right. It looks at windows of past data, which we can think of as
+‘tokens’, and predicts what comes next. This prediction is based on
+patterns the model identifies in past data and extrapolates into the
+future.
+
+## Tokens
+
+`TimeGPT` processes time series data in chunks. Each data point in a
+series can be thought of as a ‘token’, akin to how individual words or
+characters are treated in natural language processing (NLP).
+
+## Fine-tuning
+
+Fine-tuning is a process used in machine learning where a pre-trained
+model like `TimeGPT` undergoes additional training to adapt it for a
+specific dataset. Initially, `TimeGPT` can operate in a zero-shot
+manner, meaning it can generate forecasts as-is. While this zero-shot
+approach provides a solid baseline, the performance of `TimeGPT` can
+often be improved through fine-tuning. During this process, the
+`TimeGPT` model undergoes additional training using the specific
+dataset, starting from the pre-trained parameters. The updated model
+then produces the forecasts.
+
+[Learn how to fine-tune
+TimeGPT](https://docs.nixtla.io/docs/tutorials-fine_tuning)
+
+## Historical Forecasts
+
+Historical forecasts, also known as in-sample forecasts, are the
+predictions made for the historical data. These forecasts are commonly
+used to evaluate the performance of forecasting models by comparing the
+predicted values against the actual values.
+
+[Learn how to make historical forecasts with
+TimeGPT](https://docs.nixtla.io/docs/tutorials-historical_forecast)
+
+## Anomaly Detection
+
+Anomaly detection refers to the process of identifying unusual
+observations that deviate significantly from the expected behavior of
+the data. Anomalies, also known as outliers, can be caused by a variety
+of factors, such as errors in the data collection process, sudden
+changes in the underlying patterns of the data, or unexpected events.
+These anomalies can pose challenges for many forecasting models, as they
+may distort trends, seasonal patterns, or estimates of autocorrelation.
+Consequently, anomalies can significantly impact the accuracy of
+forecasts. Therefore, it is crucial to be able to identify them
+accurately.
+
+Anomaly detection has many applications across different industries,
+including detecting fraud in financial transactions, monitoring the
+performance of online services, or identifying unusual patterns in
+energy usage.
+
+[Learn how to detect anomalies with
+TimeGPT](https://docs.nixtla.io/docs/capabilities-anomaly-detection-anomaly_detection)
+
+## Time Series Cross Validation
+
+Time series cross-validation is a method for evaluating how a model
+would have performed on historical data. It works by defining a sliding
+window across past observations and predicting the period following it.
+It differs from standard cross-validation by maintaining the
+chronological order of the data instead of randomly splitting it.
+
+This method allows for a more accurate estimation of a forecasting
+model’s predictive capabilities by considering multiple sequential
+periods. When only one window is used, this method resembles a standard
+train-test split, with the last set of observations serving as the test
+data and all preceding data as the training set.
+
+[Learn how to perform cross-validation with
+TimeGPT](https://docs.nixtla.io/docs/tutorials-cross_validation)
+
+## Exogenous Variables
+
+Exogenous variables are external factors that can influence the behavior
+of a time series but are not directly affected by it. For example, in
+retail sales forecasting, exogenous variables could include factors such
+as holidays, promotions, prices, or weather data for electricity load
+forecasts. By incorporating these variables into the forecasting model,
+it is possible to capture the relationships between the target series
+and external factors, leading to more accurate predictions.
+
+[Learn how to include exogenous variables in
+TimeGPT](https://docs.nixtla.io/docs/tutorials-exogenous_variables)
+
diff --git a/nixtla/docs/getting-started/introduction.html.mdx b/nixtla/docs/getting-started/introduction.html.mdx
new file mode 100644
index 00000000..2248b918
--- /dev/null
+++ b/nixtla/docs/getting-started/introduction.html.mdx
@@ -0,0 +1,147 @@
+---
+output-file: introduction.html
+title: About TimeGPT
+---
+
+
+TimeGPT is a production-ready generative pretrained transformer for time
+series. It’s capable of accurately predicting various domains such as
+retail, electricity, finance, and IoT with just a few lines of code.
+
+It is user-friendly and low-code. Users can simply upload their time
+series data and generate forecasts or detect anomalies with just a
+single line of code.
+
+TimeGPT is the only out of-the-box foundation model for time series that
+can be used through our public APIs, through [Azure Studio as
+TimeGEN-1](https://techcommunity.microsoft.com/t5/ai-machine-learning-blog/announcing-timegen-1-in-azure-ai-leap-forward-in-time-series/ba-p/4140446)
+or on your own infrastructure.
+
+Get started! [Activate your free
+trial](https://dashboard.nixtla.io/freetrial) and see our [Quickstart
+Guide](https://docs.nixtla.io/docs/getting-started-timegpt_quickstart).
+
+## Features and capabilities
+
+- **[Zero-shot
+ Inference](https://docs.nixtla.io/docs/capabilities-forecast-quickstart)**:
+ TimeGPT can generate forecasts and detect anomalies straight out of
+ the box, requiring no prior training data. This allows for immediate
+ deployment and quick insights from any time series data.
+
+- **[Fine-tuning](https://docs.nixtla.io/docs/tutorials-fine_tuning)**:
+ Enhance TimeGPT’s capabilities by fine-tuning the model on your
+ specific datasets, enabling the model to adapt to the nuances of
+ your unique time series data and improving performance on tailored
+ tasks.
+
+- **[API Access](https://dashboard.nixtla.io/sign_in)**: Integrate
+ TimeGPT seamlessly into your applications via our robust API (obtain
+ an API key through our
+ [Dashboard](https://dashboard.nixtla.io/sign_in)). TimeGPT is also
+ supported through [Azure
+ Studio](https://docs.nixtla.io/docs/deployment-azureai) for even
+ more flexible integration options. Alternatively, deploy TimeGPT on
+ your own infrastructure to maintain full control over your data and
+ workflows.
+
+- **[Add Exogenous
+ Variables](https://docs.nixtla.io/docs/tutorials-exogenous_variables)**:
+ Incorporate additional variables that might influence your
+ predictions to enhance forecast accuracy. (E.g. Special Dates,
+ events or prices)
+
+- **[Multiple Series
+ Forecasting](https://docs.nixtla.io/docs/tutorials-multiple_series_forecasting)**:
+ Simultaneously forecast multiple time series data, optimizing
+ workflows and resources.
+
+- **[Specific Loss
+ Function](https://docs.nixtla.io/docs/tutorials-fine_tuning_with_a_specific_loss_function)**:
+ Tailor the fine-tuning process by choosing from many loss functions
+ to meet specific performance metrics.
+
+- **[Cross-validation](https://docs.nixtla.io/docs/tutorials-cross_validation)**:
+ Implement out of the box cross-validation techniques to ensure model
+ robustness and generalizability.
+
+- **[Prediction
+ Intervals](https://docs.nixtla.io/docs/tutorials-prediction_intervals)**:
+ Provide intervals in your predictions to quantify uncertainty
+ effectively.
+
+- **[Irregular
+ Timestamps](https://docs.nixtla.io/docs/capabilities-forecast-irregular_timestamps)**:
+ Handle data with irregular timestamps, accommodating non-uniform
+ interval series without preprocessing.
+
+- **[Anomaly
+ Detection](https://docs.nixtla.io/docs/tutorials-anomaly_detection)**:
+ Automatically detect anomalies in time series, and use exogenous
+ features for enhanced performance.
+
+**Get started with our [QuickStart
+guide](https://docs.nixtla.io/docs/getting-started-timegpt_quickstart),
+walk through tutorials on the different capabilities, and learn from
+real-world use cases in our documentation.**
+
+## Architecture
+
+Self-attention, the revolutionary concept introduced by the paper
+[Attention is all you need](https://arxiv.org/abs/1706.03762), is the
+basis of this foundation model. TimeGPT model is not based on any
+existing large language model(LLM). Instead, it is independently trained
+on a vast amount of time series data, and the large transformer model is
+designed to minimize the forecasting error.
+
+
+
+The architecture consists of an encoder-decoder structure with multiple
+layers, each with residual connections and layer normalization. Finally,
+a linear layer maps the decoder’s output to the forecasting window
+dimension. The general intuition is that attention-based mechanisms are
+able to capture the diversity of past events and correctly extrapolate
+potential future distributions.
+
+To make prediction, TimeGPT “reads” the input series much like the way
+humans read a sentence – from left to right. It looks at windows of past
+data, which we can think of as “tokens”, and predicts what comes next.
+This prediction is based on patterns the model identifies in past data
+and extrapolates into the future.
+
+## Explore examples and use cases
+
+Visit our comprehensive documentation to explore a wide range of
+examples and practical use cases for TimeGPT. Whether you’re getting
+started with our [Quickstart
+Guide](https://docs.nixtla.io/docs/getting-started-timegpt_quickstart),
+[setting up your API
+key](https://docs.nixtla.io/docs/getting-started-setting_up_your_api_key),
+or looking for advanced forecasting techniques, our resources are
+designed to guide you through every step of the process.
+
+Learn how to handle [anomaly
+detection](https://docs.nixtla.io/docs/capabilities-anomaly-detection-quickstart),
+[fine-tune
+models](https://docs.nixtla.io/docs/tutorials-fine_tuning_with_a_specific_loss_function)
+with specific loss functions, and scale your computing using frameworks
+like [Spark](https://docs.nixtla.io/docs/tutorials-spark),
+[Dask](https://docs.nixtla.io/docs/tutorials-dask), and
+[Ray](https://docs.nixtla.io/docs/tutorials-ray).
+
+Additionally, our documentation covers specialized topics such as
+handling [exogenous
+variables](https://docs.nixtla.io/docs/tutorials-exogenous_variables),
+validating models through
+[cross-validation](https://docs.nixtla.io/docs/tutorials-cross_validation),
+and forecasting under uncertainty with [quantile
+forecasts](https://docs.nixtla.io/docs/tutorials-quantile_forecasts) and
+[prediction
+intervals](https://docs.nixtla.io/docs/tutorials-prediction_intervals).
+
+For those interested in real-world applications, discover how TimeGPT
+can be used for [forecasting web
+traffic](https://docs.nixtla.io/docs/use-cases-forecasting_web_traffic)
+or [predicting Bitcoin
+prices](https://docs.nixtla.io/docs/use-cases-bitcoin_price_prediction).
+
diff --git a/nixtla/docs/getting-started/polars_quickstart.html.mdx b/nixtla/docs/getting-started/polars_quickstart.html.mdx
new file mode 100644
index 00000000..70a127b7
--- /dev/null
+++ b/nixtla/docs/getting-started/polars_quickstart.html.mdx
@@ -0,0 +1,207 @@
+---
+description: >-
+ TimeGPT is a production ready, generative pretrained transformer for time
+ series. It's capable of accurately predicting various domains such as retail,
+ electricity, finance, and IoT with just a few lines of code 🚀.
+output-file: polars_quickstart.html
+title: TimeGPT Quickstart (Polars)
+---
+
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/getting-started/21_polars_quickstart.ipynb)
+
+## Step 1: Create a TimeGPT account and generate your API key
+
+- Go to [dashboard.nixtla.io](https://dashboard.nixtla.io/)
+- Sign in with Google, GitHub or your email
+- Create your API key by going to ‘API Keys’ in the menu and clicking
+ on ‘Create New API Key’
+- Your new key will appear. Copy the API key using the button on the
+ right.
+
+
+
+## Step 2: Install Nixtla
+
+In your favorite Python development environment:
+
+Install `nixtla` with `pip`:
+
+```shell
+pip install nixtla
+```
+
+## Step 3: Import the Nixtla TimeGPT client
+
+```python
+from nixtla import NixtlaClient
+```
+
+You can instantiate the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class providing your authentication API key.
+
+```python
+nixtla_client = NixtlaClient(
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+Check your API key status with the `validate_api_key` method.
+
+```python
+nixtla_client.validate_api_key()
+```
+
+``` text
+True
+```
+
+**This will get you started, but for more secure usage, see [Setting Up
+your API
+Key](https://docs.nixtla.io/docs/getting-started-setting_up_your_api_key).**
+
+## Step 4: Start making forecasts!
+
+Now you can start making forecasts! Let’s import an example using the
+classic `AirPassengers` dataset. This dataset contains the monthly
+number of airline passengers in Australia between 1949 and 1960. First,
+load the dataset and plot it:
+
+```python
+import polars as pl
+```
+
+
+```python
+df = pl.read_csv(
+ 'https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/air_passengers.csv',
+ try_parse_dates=True,
+)
+df.head()
+```
+
+| timestamp | value |
+|------------|-------|
+| date | i64 |
+| 1949-01-01 | 112 |
+| 1949-02-01 | 118 |
+| 1949-03-01 | 132 |
+| 1949-04-01 | 129 |
+| 1949-05-01 | 121 |
+
+```python
+nixtla_client.plot(df, time_col='timestamp', target_col='value')
+```
+
+
+
+> 📘 Data Requirements
+>
+> - Make sure the target variable column does not have missing or
+> non-numeric values.
+> - Do not include gaps/jumps in the timestamps (for the given
+> frequency) between the first and late timestamps. The forecast
+> function will not impute missing dates.
+> - The time column should be of type
+> [Date](https://docs.pola.rs/api/python/stable/reference/api/polars.datatypes.Date.html)
+> or
+> [Datetime](https://docs.pola.rs/api/python/stable/reference/api/polars.datatypes.Datetime.html).
+>
+> For further details go to [Data
+> Requeriments](https://docs.nixtla.io/docs/getting-started-data_requirements).
+
+### Forecast a longer horizon into the future
+
+Next, forecast the next 12 months using the SDK `forecast` method. Set
+the following parameters:
+
+- `df`: A pandas DataFrame containing the time series data.
+- `h`: Horizons is the number of steps ahead to forecast.
+- `freq`: The polars offset alias, see the possible values
+ [here](https://docs.pola.rs/api/python/stable/reference/expressions/api/polars.Expr.dt.offset_by.html).
+- `time_col`: The column that identifies the datestamp.
+- `target_col`: The variable to forecast.
+
+```python
+timegpt_fcst_df = nixtla_client.forecast(df=df, h=12, freq='1mo', time_col='timestamp', target_col='value')
+timegpt_fcst_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Querying model metadata...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| timestamp | TimeGPT |
+|------------|------------|
+| date | f64 |
+| 1961-01-01 | 437.837921 |
+| 1961-02-01 | 426.062714 |
+| 1961-03-01 | 463.116547 |
+| 1961-04-01 | 478.244507 |
+| 1961-05-01 | 505.646484 |
+
+```python
+nixtla_client.plot(df, timegpt_fcst_df, time_col='timestamp', target_col='value')
+```
+
+
+
+You can also produce longer forecasts by increasing the horizon
+parameter and selecting the `timegpt-1-long-horizon` model. Use this
+model if you want to predict more than one seasonal period of your data.
+
+For example, let’s forecast the next 36 months:
+
+```python
+timegpt_fcst_df = nixtla_client.forecast(df=df, h=36, time_col='timestamp', target_col='value', freq='1mo', model='timegpt-1-long-horizon')
+timegpt_fcst_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Querying model metadata...
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon. This may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| timestamp | TimeGPT |
+|------------|------------|
+| date | f64 |
+| 1961-01-01 | 436.843414 |
+| 1961-02-01 | 419.351532 |
+| 1961-03-01 | 458.943146 |
+| 1961-04-01 | 477.876068 |
+| 1961-05-01 | 505.656921 |
+
+```python
+nixtla_client.plot(df, timegpt_fcst_df, time_col='timestamp', target_col='value')
+```
+
+
+
+### Produce a shorter forecast
+
+You can also produce a shorter forecast. For this, we recommend using
+the default model, `timegpt-1`.
+
+```python
+timegpt_fcst_df = nixtla_client.forecast(df=df, h=6, time_col='timestamp', target_col='value', freq='1mo')
+nixtla_client.plot(df, timegpt_fcst_df, time_col='timestamp', target_col='value')
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+
+
diff --git a/nixtla/docs/getting-started/pricing.html.mdx b/nixtla/docs/getting-started/pricing.html.mdx
new file mode 100644
index 00000000..8a3b00de
--- /dev/null
+++ b/nixtla/docs/getting-started/pricing.html.mdx
@@ -0,0 +1,29 @@
+---
+output-file: pricing.html
+title: Subscription Plans
+---
+
+
+We offer various Enterprise plans tailored to your forecasting needs.
+The number of API calls, number of users, and support levels can be
+customized based on your needs. We also offer an option for a
+self-hosted version and a version hosted on Azure.
+
+Please get in touch with us at `support@nixtla.io` for more information
+regarding pricing options and to discuss your specific requirements. For
+organizations interested in exploring our solution further, you can
+schedule a demo
+[here](https://meetings.hubspot.com/cristian-challu/enterprise-contact-us?uuid=dc037f5a-d93b-4%5B…%5D90b-a611dd9460af&utm_source=github&utm_medium=pricing_page).
+
+**Free trial available**
+
+When you [create your account](https://dashboard.nixtla.io), you’ll
+receive a 30-day free trial, no credit card required. After 30 days,
+access will expire unless you upgrade to a paid plan. Contact us to
+continue leveraging TimeGPT for accurate and easy to use forecasting!
+
+**More information on pricing and billing**
+
+For additional information on pricing and billing please see [our
+FAQ](https://docs.nixtla.io/docs/getting-started-faq#pricing-and-billing).
+
diff --git a/nixtla/docs/getting-started/quickstart.html.mdx b/nixtla/docs/getting-started/quickstart.html.mdx
new file mode 100644
index 00000000..09c33c86
--- /dev/null
+++ b/nixtla/docs/getting-started/quickstart.html.mdx
@@ -0,0 +1,215 @@
+---
+description: >-
+ TimeGPT is a production ready, generative pretrained transformer for time
+ series. It's capable of accurately predicting various domains such as retail,
+ electricity, finance, and IoT with just a few lines of code 🚀.
+output-file: quickstart.html
+title: TimeGPT Quickstart
+---
+
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/getting-started/2_quickstart.ipynb)
+
+## Step 1: Create a TimeGPT account and generate your API key
+
+- Go to [dashboard.nixtla.io](https://dashboard.nixtla.io) to activate
+ your free trial and set up an account.
+- Sign in with Google, GitHub or your email
+- Create your API key by going to ‘API Keys’ in the menu and clicking
+ on ‘Create New API Key’
+- Your new key will appear. Copy the API key using the button on the
+ right.
+
+
+
+## Step 2: Install Nixtla
+
+In your favorite Python development environment:
+
+Install `nixtla` with `pip`:
+
+```shell
+pip install nixtla
+```
+
+## Step 3: Import the Nixtla TimeGPT client
+
+```python
+from nixtla import NixtlaClient
+```
+
+You can instantiate the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class providing your authentication API key.
+
+```python
+nixtla_client = NixtlaClient(
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+Check your API key status with the `validate_api_key` method.
+
+```python
+nixtla_client.validate_api_key()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Happy Forecasting! :), If you have questions or need support, please email support@nixtla.io
+```
+
+``` text
+True
+```
+
+**This will get you started, but for more secure usage, see [Setting Up
+your API
+Key](https://docs.nixtla.io/docs/getting-started-setting_up_your_api_key).**
+
+## Step 4: Start making forecasts!
+
+Now you can start making forecasts! Let’s import an example using the
+classic `AirPassengers` dataset. This dataset contains the monthly
+number of airline passengers in Australia between 1949 and 1960. First,
+load the dataset and plot it:
+
+```python
+import pandas as pd
+```
+
+
+```python
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/air_passengers.csv')
+df.head()
+```
+
+| | timestamp | value |
+|-----|------------|-------|
+| 0 | 1949-01-01 | 112 |
+| 1 | 1949-02-01 | 118 |
+| 2 | 1949-03-01 | 132 |
+| 3 | 1949-04-01 | 129 |
+| 4 | 1949-05-01 | 121 |
+
+```python
+nixtla_client.plot(df, time_col='timestamp', target_col='value')
+```
+
+
+
+> 📘 Data Requirements
+>
+> - Make sure the target variable column does not have missing or
+> non-numeric values.
+> - Do not include gaps/jumps in the datestamps (for the given
+> frequency) between the first and late datestamps. The forecast
+> function will not impute missing dates.
+> - The format of the datestamp column should be readable by Pandas
+> (see [this
+> link](https://pandas.pydata.org/docs/reference/api/pandas.to_datetime.html)
+> for more details).
+>
+> For further details go to [Data
+> Requirements](https://docs.nixtla.io/docs/getting-started-data_requirements).
+
+> 👍 Save figures made with TimeGPT
+>
+> The `plot` method automatically displays figures when in a notebook
+> environment. To save figures locally, you can do:
+>
+> `fig = nixtla_client.plot(df, time_col='timestamp', target_col='value')`
+>
+> `fig.savefig('plot.png', bbox_inches='tight')`
+
+### Forecast a longer horizon into the future
+
+Next, forecast the next 12 months using the SDK `forecast` method. Set
+the following parameters:
+
+- `df`: A pandas DataFrame containing the time series data.
+- `h`: Horizons is the number of steps ahead to forecast.
+- `freq`: The frequency of the time series in Pandas format. See
+ [pandas’ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+ (If you don’t provide any frequency, the SDK will try to infer it)
+- `time_col`: The column that identifies the datestamp.
+- `target_col`: The variable to forecast.
+
+```python
+timegpt_fcst_df = nixtla_client.forecast(df=df, h=12, freq='MS', time_col='timestamp', target_col='value')
+timegpt_fcst_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| | timestamp | TimeGPT |
+|-----|------------|------------|
+| 0 | 1961-01-01 | 437.837921 |
+| 1 | 1961-02-01 | 426.062714 |
+| 2 | 1961-03-01 | 463.116547 |
+| 3 | 1961-04-01 | 478.244507 |
+| 4 | 1961-05-01 | 505.646484 |
+
+```python
+nixtla_client.plot(df, timegpt_fcst_df, time_col='timestamp', target_col='value')
+```
+
+
+
+You can also produce longer forecasts by increasing the horizon
+parameter and selecting the `timegpt-1-long-horizon` model. Use this
+model if you want to predict more than one seasonal period of your data.
+
+For example, let’s forecast the next 36 months:
+
+```python
+timegpt_fcst_df = nixtla_client.forecast(df=df, h=36, time_col='timestamp', target_col='value', freq='MS', model='timegpt-1-long-horizon')
+timegpt_fcst_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon. This may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| | timestamp | TimeGPT |
+|-----|------------|------------|
+| 0 | 1961-01-01 | 436.843414 |
+| 1 | 1961-02-01 | 419.351532 |
+| 2 | 1961-03-01 | 458.943146 |
+| 3 | 1961-04-01 | 477.876068 |
+| 4 | 1961-05-01 | 505.656921 |
+
+```python
+nixtla_client.plot(df, timegpt_fcst_df, time_col='timestamp', target_col='value')
+```
+
+
+
+### Produce a shorter forecast
+
+You can also produce a shorter forecast. For this, we recommend using
+the default model, `timegpt-1`.
+
+```python
+timegpt_fcst_df = nixtla_client.forecast(df=df, h=6, time_col='timestamp', target_col='value', freq='MS')
+nixtla_client.plot(df, timegpt_fcst_df, time_col='timestamp', target_col='value')
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+
+
diff --git a/nixtla/docs/getting-started/setting_up_your_api_key.html.mdx b/nixtla/docs/getting-started/setting_up_your_api_key.html.mdx
new file mode 100644
index 00000000..44fbb73f
--- /dev/null
+++ b/nixtla/docs/getting-started/setting_up_your_api_key.html.mdx
@@ -0,0 +1,125 @@
+---
+output-file: setting_up_your_api_key.html
+title: Setting up your API key
+---
+
+
+This tutorial will explain how to set up your API key when using the
+Nixtla SDK. To create an `Api Key` go to your
+[Dashboard](https://dashboard.nixtla.io/).
+
+There are different ways to set up your API key. We provide some
+examples below. A scematic is given below.
+
+
+
+## 1. Copy and paste your key directly into your Python code
+
+This approach is straightforward and best for quick tests or scripts
+that won’t be shared.
+
+- **Step 1**: Copy the API key found in the `API Keys` of your [Nixtla
+ dashboard](https://dashboard.nixtla.io/).
+- **Step 2**: Paste the key directly into your Python code, by
+ instantiating the
+ [`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+ with your API key:
+
+```python
+from nixtla import NixtlaClient
+nixtla_client = NixtlaClient(api_key ='your API key here')
+```
+
+> **Important**
+>
+> This approach is considered unsecure, as your API key will be part of
+> your source code.
+
+## 2. Secure: using an environment variable
+
+- **Step 1**: Store your API key in an environment variable named
+ `NIXTLA_API_KEY`. This can be done (a) temporarily for a session
+ or (b) permanently, depending on your preference.
+- **Step 2**: When you instantiate the
+ [`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+ class, the SDK will automatically look for the `NIXTLA_API_KEY`
+ environment variable and use it to authenticate your requests.
+
+> **Important**
+>
+> The environment variable must be named exactly `NIXTLA_API_KEY`, with
+> all capital letters and no deviations in spelling, for the SDK to
+> recognize it.
+
+### a. Temporary: From the Terminal
+
+This approach is useful if you are working from a terminal, and need a
+temporary solution.
+
+#### Linux / Mac
+
+Open a terminal and use the `export` command to set `NIXTLA_API_KEY`.
+
+```bash
+export NIXTLA_API_KEY=your_api_key
+```
+
+#### Windows
+
+For Windows users, open a Powershell window and use the `Set` command to
+set `NIXTLA_API_KEY`.
+
+```powershell
+Set NIXTLA_API_KEY=your_api_key
+```
+
+### b. Permanent: Using a `.env` file
+
+For a more persistent solution place your API key in a `.env` file
+located in the folder of your Python script. In this file, include the
+following:
+
+```python
+NIXTLA_API_KEY=your_api_key
+```
+
+You can now load the environment variable within your Python script. Use
+the `dotenv` package to load the `.env` file and then instantiate the
+`NIXTLA_API_KEY` class. For example:
+
+```python
+from dotenv import load_dotenv
+load_dotenv()
+
+from nixtla import NixtlaClient
+nixtla_client = NixtlaClient()
+```
+
+This approach is more secure and suitable for applications that will be
+deployed or shared, as it keeps API keys out of the source code.
+
+> **Important**
+>
+> Remember, your API key is like a password - keep it secret, keep it
+> safe!
+
+## 3. Validate your API key
+
+You can always find your API key in the `API Keys` section of your
+dashboard. To check the status of your API key, use the
+`validate_api_key` method of the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class. This method will return `True` if the API key is valid and
+`False` otherwise.
+
+```python
+nixtla_client.validate_api_key()
+```
+
+You don’t need to validate your API key every time you use `TimeGPT`.
+This function is provided for your convenience to ensure its validity.
+For full access to `TimeGPT`’s functionalities, in addition to a valid
+API key, you also need sufficient credits in your account. You can check
+your credits in the `Usage` section of your
+[dashboard](https://dashboard.nixtla.io/).
+
diff --git a/nixtla/docs/getting-started/why_timegpt.html.mdx b/nixtla/docs/getting-started/why_timegpt.html.mdx
new file mode 100644
index 00000000..e2f4247e
--- /dev/null
+++ b/nixtla/docs/getting-started/why_timegpt.html.mdx
@@ -0,0 +1,374 @@
+---
+output-file: why_timegpt.html
+title: Why TimeGPT?
+---
+
+
+In this notebook, we compare the performance of TimeGPT against three
+forecasting models: the classical model (ARIMA), the machine learning
+model (LightGBM), and the deep learning model (N-HiTS), using a subset
+of data from the M5 Forecasting competition. We want to highlight three
+top-rated benefits our users love about TimeGPT:
+
+🎯 **Accuracy**: TimeGPT consistently outperforms traditional models by
+capturing complex patterns with precision.
+
+⚡ **Speed**: Generate forecasts faster without needing extensive
+training or tuning for each series.
+
+🚀 **Ease of Use**: Minimal setup and no complex preprocessing make
+TimeGPT accessible and ready to use right out of the box!
+
+Before diving into the notebook, please visit our
+[dashboard](https://dashboard.nixtla.io) to generate your TimeGPT
+`api_key` and give it a try yourself!
+
+# Table of Contents
+
+1. [Data Introduction](#1-data-introduction)
+2. [Model Fitting](#2-model-fitting-timegpt-arima-lightgbm-n-hits)
+ 1. [Fitting TimeGPT](#21-timegpt)
+ 2. [Fitting ARIMA](#22-classical-models-arima)
+ 3. [Fitting Light GBM](#23-machine-learning-models-lightgbm)
+ 4. [Fitting NHITS](#24-n-hits)
+3. [Results and Evaluation](#3-performance-comparison-and-results)
+4. [Conclusion](#4-conclusion)
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/getting-started/7_why_timegpt.ipynb)
+
+```python
+import os
+import numpy as np
+import pandas as pd
+import matplotlib.pyplot as plt
+
+from nixtla import NixtlaClient
+from utilsforecast.plotting import plot_series
+from utilsforecast.losses import mae, rmse, smape
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+## 1. Data introduction
+
+In this notebook, we’re working with an aggregated dataset from the M5
+Forecasting - Accuracy competition. This dataset includes **7 daily time
+series**, each with **1,941 data points**. The last **28 data points**
+of each series are set aside as the test set, allowing us to evaluate
+model performance on unseen data.
+
+```python
+df = pd.read_csv('https://datasets-nixtla.s3.amazonaws.com/demand_example.csv', parse_dates=['ds'])
+```
+
+
+```python
+df.groupby('unique_id').agg({"ds":["min","max","count"],\
+ "y":["min","mean","median","max"]})
+```
+
+| | ds | | | y | | | |
+|-------------|------------|------------|-------|------|--------------|---------|---------|
+| | min | max | count | min | mean | median | max |
+| unique_id | | | | | | | |
+| FOODS_1 | 2011-01-29 | 2016-05-22 | 1941 | 0.0 | 2674.085523 | 2665.0 | 5493.0 |
+| FOODS_2 | 2011-01-29 | 2016-05-22 | 1941 | 0.0 | 4015.984029 | 3894.0 | 9069.0 |
+| FOODS_3 | 2011-01-29 | 2016-05-22 | 1941 | 10.0 | 16969.089129 | 16548.0 | 28663.0 |
+| HOBBIES_1 | 2011-01-29 | 2016-05-22 | 1941 | 0.0 | 2936.122617 | 2908.0 | 5009.0 |
+| HOBBIES_2 | 2011-01-29 | 2016-05-22 | 1941 | 0.0 | 279.053065 | 248.0 | 871.0 |
+| HOUSEHOLD_1 | 2011-01-29 | 2016-05-22 | 1941 | 0.0 | 6039.594539 | 5984.0 | 11106.0 |
+| HOUSEHOLD_2 | 2011-01-29 | 2016-05-22 | 1941 | 0.0 | 1566.840289 | 1520.0 | 2926.0 |
+
+```python
+df_train = df.query('ds <= "2016-04-24"')
+df_test = df.query('ds > "2016-04-24"')
+
+print(df_train.shape, df_test.shape)
+```
+
+``` text
+(13391, 3) (196, 3)
+```
+
+## 2. Model Fitting (TimeGPT, ARIMA, LightGBM, N-HiTS)
+
+### 2.1 TimeGPT
+
+TimeGPT offers a powerful, streamlined solution for time series
+forecasting, delivering state-of-the-art results with minimal effort.
+With TimeGPT, there’s no need for data preprocessing or feature
+engineering – simply initiate the Nixtla client and call
+`nixtla_client.forecast` to produce accurate, high-performance forecasts
+tailored to your unique time series.
+
+```python
+# Forecast with TimeGPT
+fcst_timegpt = nixtla_client.forecast(df = df_train,
+ target_col = 'y',
+ h=28, # Forecast horizon, predicts the next 28 time steps
+ model='timegpt-1-long-horizon', # Use the model for long-horizon forecasting
+ finetune_steps=10, # Number of finetuning steps
+ level = [90]) # Generate a 90% confidence interval
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: D
+INFO:nixtla.nixtla_client:Querying model metadata...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+```python
+# Evaluate performance and plot forecast
+fcst_timegpt['ds'] = pd.to_datetime(fcst_timegpt['ds'])
+test_df = pd.merge(df_test, fcst_timegpt, 'left', ['unique_id', 'ds'])
+evaluation_timegpt = evaluate(test_df, metrics=[rmse, smape], models=["TimeGPT"])
+evaluation_timegpt.groupby(['metric'])['TimeGPT'].mean()
+```
+
+``` text
+metric
+rmse 592.607378
+smape 0.049403
+Name: TimeGPT, dtype: float64
+```
+
+### 2.2 Classical Models (ARIMA):
+
+Next, we applied ARIMA, a traditional statistical model, to the same
+forecasting task. Classical models use historical trends and seasonality
+to make predictions by relying on linear assumptions. However, they
+struggled to capture the complex, non-linear patterns within the data,
+leading to lower accuracy compared to other approaches. Additionally,
+ARIMA was slower due to its iterative parameter estimation process,
+which becomes computationally intensive for larger datasets.
+
+> 📘 Why Use TimeGPT over Classical Models?
+>
+> - **Complex Patterns**: TimeGPT captures non-linear trends classical
+> models miss.
+>
+> - **Minimal Preprocessing**: TimeGPT requires little to no data
+> preparation.
+>
+> - **Scalability**: TimeGPT can efficiently scales across multiple
+> series without retraining.
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import AutoARIMA
+```
+
+
+```python
+#Initiate ARIMA model
+sf = StatsForecast(
+ models=[AutoARIMA(season_length=7)],
+ freq='D'
+)
+# Fit and forecast
+fcst_arima = sf.forecast(h=28, df=df_train)
+```
+
+
+```python
+fcst_arima.reset_index(inplace=True)
+test_df = pd.merge(df_test, fcst_arima, 'left', ['unique_id', 'ds'])
+evaluation_arima = evaluate(test_df, metrics=[rmse, smape], models=["AutoARIMA"])
+evaluation_arima.groupby(['metric'])['AutoARIMA'].mean()
+```
+
+``` text
+metric
+rmse 724.957364
+smape 0.055018
+Name: AutoARIMA, dtype: float64
+```
+
+### 2.3 Machine Learning Models (LightGBM)
+
+Thirdly, we used a machine learning model, LightGBM, for the same
+forecasting task, implemented through the automated pipeline provided by
+our mlforecast library. While LightGBM can capture seasonality and
+patterns, achieving the best performance often requires detailed feature
+engineering, careful hyperparameter tuning, and domain knowledge. You
+can try our mlforecast library to simplify this process and get started
+quickly!
+
+> 📘 Why Use TimeGPT over Machine Learning Models?
+>
+> - **Automatic Pattern Recognition**: Captures complex patterns from
+> raw data, bypassing the need for feature engineering.
+>
+> - **Minimal Tuning**: Works well without extensive tuning.
+>
+> - **Scalability**: Forecasts across multiple series without
+> retraining.
+
+```python
+import optuna
+from mlforecast.auto import AutoMLForecast, AutoLightGBM
+
+# Suppress Optuna's logging output
+optuna.logging.set_verbosity(optuna.logging.ERROR)
+```
+
+
+```python
+# Initialize an automated forecasting pipeline using AutoMLForecast.
+mlf = AutoMLForecast(
+ models=[AutoLightGBM()],
+ freq='D',
+ season_length=7,
+ fit_config=lambda trial: {'static_features': ['unique_id']}
+)
+
+# Fit the model to the training dataset.
+mlf.fit(
+ df=df_train.astype({'unique_id': 'category'}),
+ n_windows=1,
+ h=28,
+ num_samples=10,
+)
+fcst_lgbm = mlf.predict(28)
+```
+
+
+```python
+test_df = pd.merge(df_test, fcst_lgbm, 'left', ['unique_id', 'ds'])
+evaluation_lgbm = evaluate(test_df, metrics=[rmse, smape], models=["AutoLightGBM"])
+evaluation_lgbm.groupby(['metric'])['AutoLightGBM'].mean()
+```
+
+``` text
+metric
+rmse 687.773744
+smape 0.051448
+Name: AutoLightGBM, dtype: float64
+```
+
+### 2.4 N-HiTS
+
+Lastly, we used N-HiTS, a state-of-the-art deep learning model designed
+for time series forecasting. The model produced accurate results,
+demonstrating its ability to capture complex, non-linear patterns within
+the data. However, setting up and tuning N-HiTS required significantly
+more time and computational resources compared to TimeGPT.
+
+> 📘 Why Use TimeGPT Over Deep Learning Models?
+>
+> - **Faster Setup**: Quick setup and forecasting, unlike the lengthy
+> configuration and training times of neural networks.
+>
+> - **Less Tuning**: Performs well with minimal tuning and
+> preprocessing, while neural networks often need extensive
+> adjustments.
+>
+> - **Ease of Use**: Simple deployment with high accuracy, making it
+> accessible without deep technical expertise.
+
+```python
+from neuralforecast.core import NeuralForecast
+from neuralforecast.models import NHITS
+```
+
+
+```python
+# Initialize the N-HiTS model.
+models = [NHITS(h=28,
+ input_size=28,
+ max_steps=100)]
+
+# Fit the model using training data
+nf = NeuralForecast(models=models, freq='D')
+nf.fit(df=df_train)
+fcst_nhits = nf.predict()
+```
+
+
+```python
+test_df = pd.merge(df_test,fcst_nhits, 'left', ['unique_id', 'ds'])
+evaluation_nhits = evaluate(test_df, metrics=[rmse, smape], models=["NHITS"])
+evaluation_nhits.groupby(['metric'])['NHITS'].mean()
+```
+
+``` text
+metric
+rmse 605.011948
+smape 0.053446
+Name: NHITS, dtype: float64
+```
+
+## 3. Performance Comparison and Results:
+
+The performance of each model is evaluated using RMSE (Root Mean Squared
+Error) and SMAPE (Symmetric Mean Absolute Percentage Error). While RMSE
+emphasizes the models’ ability to control significant errors, SMAPE
+provides a relative performance perspective by normalizing errors as
+percentages. Below, we present a snapshot of performance across all
+groups. The results demonstrate that TimeGPT outperforms other models on
+both metrics.
+
+🌟 For a deeper dive into benchmarking, check out our benchmark
+repository. The summarized results are displayed below:
+
+#### Overall Performance Metrics
+
+| **Model** | **RMSE** | **SMAPE** |
+|-------------|-----------|-----------|
+| ARIMA | 724.9 | 5.50% |
+| LightGBM | 687.8 | 5.14% |
+| N-HiTS | 605.0 | 5.34% |
+| **TimeGPT** | **592.6** | **4.94%** |
+
+#### Breakdown for Each Time-series
+
+Followed below are the metrics for each individual time series groups.
+TimeGPT consistently delivers accurate forecasts across all time series
+groups. In many cases, it performs as well as or better than
+data-specific models, showing its versatility and reliability across
+different datasets.
+
+
+
+#### Benchmark Results
+
+For a more comprehensive dive into model accuracy and performance,
+explore our [Time Series Model
+Arena](https://github.com/Nixtla/nixtla/tree/main/experiments/foundation-time-series-arena)!
+TimeGPT continues to lead the pack with exceptional performance across
+benchmarks! 🌟
+
+
+
+## 4. Conclusion
+
+At the end of this notebook, we’ve put together a handy table to show
+you exactly where TimeGPT shines brightest compared to other forecasting
+models. ☀️ Think of it as your quick guide to choosing the best model
+for your unique project needs. We’re confident that TimeGPT will be a
+valuable tool in your forecasting journey. Don’t forget to visit our
+[dashboard](https://dashboard.nixtla.io) to generate your TimeGPT
+`api_key` and get started today! Happy forecasting, and enjoy the
+insights ahead!
+
+| Scenario | TimeGPT | Classical Models (e.g., ARIMA) | Machine Learning Models (e.g., XGB, LGBM) | Deep Learning Models (e.g., N-HITS) |
+|-----------|-------------|----------------|-----------------|----------------|
+| **Seasonal Patterns** | ✅ Performs well with minimal setup | ✅ Handles seasonality with adjustments (e.g., SARIMA) | ✅ Performs well with feature engineering | ✅ Captures seasonal patterns effectively |
+| **Non-Linear Patterns** | ✅ Excels, especially with complex non-linear patterns | ❌ Limited performance | ❌ Struggles without extensive feature engineering | ✅ Performs well with non-linear relationships |
+| **Large Dataset** | ✅ Highly scalable across many series | ❌ Slow and resource-intensive | ✅ Scalable with optimized implementations | ❌ Requires significant resources for large datasets |
+| **Small Dataset** | ✅ Performs well; requires only one data point to start | ✅ Performs well; may struggle with very sparse data | ✅ Performs adequately if enough features are extracted | ❌ May need a minimum data size to learn effectively |
+| **Preprocessing Required** | ✅ Minimal preprocessing needed | ❌ Requires scaling, log-transform, etc., to meet model assumptions | ❌ Requires extensive feature engineering for complex patterns | ❌ Needs data normalization and preprocessing |
+| **Accuracy Requirement** | ✅ Achieves high accuracy with minimal tuning | ❌ May struggle with complex accuracy requirements | ✅ Can achieve good accuracy with tuning | ✅ High accuracy possible but with significant resource use |
+| **Scalability** | ✅ Highly scalable with minimal task-specific configuration | ❌ Not easily scalable | ✅ Moderate scalability, with feature engineering and tuning per task | ❌ Limited scalability due to resource demands |
+| **Computational Resources** | ✅ Highly efficient, operates seamlessly on CPU, no GPU needed | ✅ Light to moderate, scales poorly with large datasets | ❌ Moderate, depends on feature complexity | ❌ High resource consumption, often requires GPU |
+| **Memory Requirement** | ✅ Efficient memory usage for large datasets | ✅ Moderate memory requirements | ❌ High memory usage for larger datasets or many series cases | ❌ High memory consumption for larger datasets and multiple series |
+| **Technical Requirements & Domain Knowledge** | ✅ Low; minimal technical setup and no domain expertise needed | ✅ Low to moderate; needs understanding of stationarity | ❌ Moderate to high; requires feature engineering and tuning | ❌ High; complex architecture and tuning |
+
diff --git a/nixtla/docs/reference/date_features.html.mdx b/nixtla/docs/reference/date_features.html.mdx
new file mode 100644
index 00000000..12ee07ea
--- /dev/null
+++ b/nixtla/docs/reference/date_features.html.mdx
@@ -0,0 +1,77 @@
+---
+output-file: date_features.html
+title: Date Features
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+#### CountryHolidays
+
+> ``` text
+> CountryHolidays (countries:list[str])
+> ```
+
+*Given a list of countries, returns a dataframe with holidays for each
+country.*
+
+```python
+import pandas as pd
+```
+
+| | US_New Year's Day | US_Memorial Day | US_Independence Day | US_Labor Day | US_Veterans Day | US_Veterans Day (observed) | US_Thanksgiving | US_Christmas Day | US_Martin Luther King Jr. Day | US_Washington's Birthday | ... | US_Juneteenth National Independence Day (observed) | US_Christmas Day (observed) | MX_Año Nuevo | MX_Día de la Constitución | MX_Natalicio de Benito Juárez | MX_Día del Trabajo | MX_Día de la Independencia | MX_Día de la Revolución | MX_Transmisión del Poder Ejecutivo Federal | MX_Navidad |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 2018-09-03 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
+| 2018-09-04 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
+| 2018-09-05 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
+| 2018-09-06 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
+| 2018-09-07 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | ... | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
+
+```python
+c_holidays = CountryHolidays(countries=['US', 'MX'])
+periods = 365 * 5
+dates = pd.date_range(end='2023-09-01', periods=periods)
+holidays_df = c_holidays(dates)
+holidays_df.head()
+```
+
+------------------------------------------------------------------------
+
+source
+
+#### SpecialDates
+
+> ``` text
+> SpecialDates (special_dates:dict[str,list[str]])
+> ```
+
+*Given a dictionary of categories and dates, returns a dataframe with
+the special dates.*
+
+```python
+special_dates = SpecialDates(
+ special_dates={
+ 'Important Dates': ['2021-02-26', '2020-02-26'],
+ 'Very Important Dates': ['2021-01-26', '2020-01-26', '2019-01-26']
+ }
+)
+periods = 365 * 5
+dates = pd.date_range(end='2023-09-01', periods=periods)
+holidays_df = special_dates(dates)
+holidays_df.head()
+```
+
+| | Important Dates | Very Important Dates |
+|------------|-----------------|----------------------|
+| 2018-09-03 | 0 | 0 |
+| 2018-09-04 | 0 | 0 |
+| 2018-09-05 | 0 | 0 |
+| 2018-09-06 | 0 | 0 |
+| 2018-09-07 | 0 | 0 |
+
diff --git a/nixtla/docs/reference/excel_addin.html.mdx b/nixtla/docs/reference/excel_addin.html.mdx
new file mode 100644
index 00000000..20861145
--- /dev/null
+++ b/nixtla/docs/reference/excel_addin.html.mdx
@@ -0,0 +1,103 @@
+---
+output-file: excel_addin.html
+title: TimeGPT Excel Add-in (Beta)
+---
+
+
+## Installation
+
+Head to the [TimeGTP excel add-in page in Microsoft
+Appsource](https://appsource.microsoft.com/en-us/product/office/WA200006429?tab=Overview)
+and click on “Get it now”
+
+## Usage
+
+> 📘 Access token required
+>
+> The TimeGPT Excel Add-in requires an access token. Get your API Key on
+> the [Nixtla Dashboard](http://dashboard.nixtla.io).
+
+## Support
+
+If you have questions or need support, please email `support@nixtla.io`.
+
+## How-to
+
+### Settings
+
+If this is your first time using Excel add-ins, find information on how
+to add Excel add-ins with your version of Excel. In the Office Add-ins
+Store, you’ll search for “TimeGPT”.
+
+Once you have installed the TimeGPT add-in, the add-in comes up in a
+sidebar task pane. \* Read through the Welcome screen. \* Click on the
+**‘Get Started’** button. \* The API URL is already set to:
+https://api.nixtla.io. \* Copy your API key from [Nixtla
+Dashboard](http://dashboard.nixtla.io). Paste it into the box that say
+**API Key, Bearer**. \* Click the gray arrow next to that box on the
+right. \* You’ll get to a screen with options for ‘Forecast’ and
+‘Anomaly Detection’.
+
+To access the settings later, click the gear icon in the top left.
+
+### Data Requirements
+
+- Put your dates in one column and your values in another.
+- Ensure your date format is recognized as a valid date by excel.
+- Ensure your values are recognized as valid number by excel.
+- All data inputs must exist in the same worksheet. The add-in does
+ not support forecasting using multiple worksheets.
+- Do not include headers
+
+Example:
+
+| dates | values |
+|:--------------|:-------|
+| 12/1/16 0:00 | 72 |
+| 12/1/16 1:00 | 65.8 |
+| 12/1/16 2:00 | 59.99 |
+| 12/1/16 3:00 | 50.69 |
+| 12/1/16 4:00 | 52.58 |
+| 12/1/16 5:00 | 65.05 |
+| 12/1/16 6:00 | 80.4 |
+| 12/1/16 7:00 | 200 |
+| 12/1/16 8:00 | 200.63 |
+| 12/1/16 9:00 | 155.47 |
+| 12/1/16 10:00 | 150.91 |
+
+#### Forecasting
+
+Once you’ve configured your token and formatted your input data then
+you’re all ready to forecast!
+
+With the add-in open, configure the forecasting settings by selecting
+the column for each input.
+
+- **Frequency** - The frequency of the data (hourly / daily / weekly /
+ monthly)
+
+- **Horizon** - The forecasting horizon. This represents the number of
+ time steps into the future that the forecast should predict.
+
+- **Dates Range** - The column and range of the timeseries timestamps.
+ Must not include header data, and should be formatted as a range,
+ e.g. A2:A145.
+
+- **Values Range** - The column and range of the timeseries values for
+ each point in time. Must not include header data, and should be
+ formatted as a range, e.g. B2:B145.
+
+When you’re ready, click **Make Prediction** to generate the predicted
+values. The add-in will generate a plot and append the forecasted data
+to the end of the column of your existing data and highlight them in
+green. So, scroll to the end of your data to see the predicted values.
+
+#### Anomaly Detection
+
+The requirements are the same as for the forecasting functionality, so
+if you already tried it you are ready to run the anomaly detection one.
+Go to the main page in the add-in and select “Anomaly Detection”, then
+choose your dates and values cell ranges and click on submit. We’ll run
+the model and mark the anomalies cells in yellow while adding a third
+column for expected values with a green background.
+
diff --git a/nixtla/docs/reference/nixtla_client.html.mdx b/nixtla/docs/reference/nixtla_client.html.mdx
new file mode 100644
index 00000000..b504f18c
--- /dev/null
+++ b/nixtla/docs/reference/nixtla_client.html.mdx
@@ -0,0 +1,378 @@
+---
+output-file: nixtla_client.html
+title: SDK Reference
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+## NixtlaClient
+
+> ``` text
+> NixtlaClient (api_key:Optional[str]=None, base_url:Optional[str]=None,
+> timeout:Optional[int]=60, max_retries:int=6,
+> retry_interval:int=10, max_wait_time:int=360)
+> ```
+
+*Client to interact with the Nixtla API.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| api_key | Optional | None | The authorization api_key interacts with the Nixtla API. If not provided, will use the NIXTLA_API_KEY environment variable. |
+| base_url | Optional | None | Custom base_url. If not provided, will use the NIXTLA_BASE_URL environment variable. |
+| timeout | Optional | 60 | Request timeout in seconds. Set this to `None` to disable it. |
+| max_retries | int | 6 | The maximum number of attempts to make when calling the API before giving up. It defines how many times the client will retry the API call if it fails. Default value is 6, indicating the client will attempt the API call up to 6 times in total |
+| retry_interval | int | 10 | The interval in seconds between consecutive retry attempts. This is the waiting period before the client tries to call the API again after a failed attempt. Default value is 10 seconds, meaning the client waits for 10 seconds between retries. |
+| max_wait_time | int | 360 | The maximum total time in seconds that the client will spend on all retry attempts before giving up. This sets an upper limit on the cumulative waiting time for all retry attempts. If this time is exceeded, the client will stop retrying and raise an exception. Default value is 360 seconds, meaning the client will cease retrying if the total time spent on retries exceeds 360 seconds. The client throws a ReadTimeout error after 60 seconds of inactivity. If you want to catch these errors, use max_wait_time \>\> 60. |
+
+------------------------------------------------------------------------
+
+source
+
+## NixtlaClient.validate_api_key
+
+> ``` text
+> NixtlaClient.validate_api_key (log:bool=True)
+> ```
+
+*Check API key status.*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|-------------------------------|
+| log | bool | True | Show the endpoint’s response. |
+| **Returns** | **bool** | | **Whether API key is valid.** |
+
+------------------------------------------------------------------------
+
+source
+
+## NixtlaClient.forecast
+
+> ``` text
+> NixtlaClient.forecast (df:~AnyDFType, h:typing.Annotated[int,Gt(gt=0)],
+> freq:Union[str,int,pandas._libs.tslibs.offsets.Bas
+> eOffset,NoneType]=None, id_col:str='unique_id',
+> time_col:str='ds', target_col:str='y',
+> X_df:Optional[~AnyDFType]=None,
+> level:Optional[list[Union[int,float]]]=None,
+> quantiles:Optional[list[float]]=None,
+> finetune_steps:typing.Annotated[int,Ge(ge=0)]=0,
+> finetune_depth:Literal[1,2,3,4,5]=1, finetune_loss
+> :Literal['default','mae','mse','rmse','mape','smap
+> e']='default',
+> finetuned_model_id:Optional[str]=None,
+> clean_ex_first:bool=True,
+> hist_exog_list:Optional[list[str]]=None,
+> validate_api_key:bool=False,
+> add_history:bool=False, date_features:Union[bool,l
+> ist[Union[str,Callable]]]=False, date_features_to_
+> one_hot:Union[bool,list[str]]=False, model:Literal
+> ['azureai','timegpt-1','timegpt-1-long-
+> horizon']='timegpt-1', num_partitions:Optional[Ann
+> otated[int,Gt(gt=0)]]=None,
+> feature_contributions:bool=False)
+> ```
+
+*Forecast your time series using TimeGPT.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | AnyDFType | | The DataFrame on which the function will operate. Expected to contain at least the following columns: - time_col: Column name in `df` that contains the time indices of the time series. This is typically a datetime column with regular intervals, e.g., hourly, daily, monthly data points. - target_col: Column name in `df` that contains the target variable of the time series, i.e., the variable we wish to predict or analyze. Additionally, you can pass multiple time series (stacked in the dataframe) considering an additional column: - id_col: Column name in `df` that identifies unique time series. Each unique value in this column corresponds to a unique time series. |
+| h | Annotated | | Forecast horizon. |
+| freq | Union | None | Frequency of the timestamps. If `None`, it will be inferred automatically. See [pandas’ available frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases). |
+| id_col | str | unique_id | Column that identifies each series. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| X_df | Optional | None | DataFrame with \[`unique_id`, `ds`\] columns and `df`’s future exogenous. |
+| level | Optional | None | Confidence levels between 0 and 100 for prediction intervals. |
+| quantiles | Optional | None | Quantiles to forecast, list between (0, 1). `level` and `quantiles` should not be used simultaneously. The output dataframe will have the quantile columns formatted as TimeGPT-q-(100 \* q) for each q. 100 \* q represents percentiles but we choose this notation to avoid having dots in column names. |
+| finetune_steps | Annotated | 0 | Number of steps used to finetune learning TimeGPT in the new data. |
+| finetune_depth | Literal | 1 | The depth of the finetuning. Uses a scale from 1 to 5, where 1 means little finetuning, and 5 means that the entire model is finetuned. |
+| finetune_loss | Literal | default | Loss function to use for finetuning. Options are: `default`, `mae`, `mse`, `rmse`, `mape`, and `smape`. |
+| finetuned_model_id | Optional | None | ID of previously fine-tuned model to use. |
+| clean_ex_first | bool | True | Clean exogenous signal before making forecasts using TimeGPT. |
+| hist_exog_list | Optional | None | Column names of the historical exogenous features. |
+| validate_api_key | bool | False | If True, validates api_key before sending requests. |
+| add_history | bool | False | Return fitted values of the model. |
+| date_features | Union | False | Features computed from the dates. Can be pandas date attributes or functions that will take the dates as input. If True automatically adds most used date features for the frequency of `df`. |
+| date_features_to_one_hot | Union | False | Apply one-hot encoding to these date features. If `date_features=True`, then all date features are one-hot encoded by default. |
+| model | Literal | timegpt-1 | Model to use as a string. Options are: `timegpt-1`, and `timegpt-1-long-horizon`. We recommend using `timegpt-1-long-horizon` for forecasting if you want to predict more than one seasonal period given the frequency of your data. |
+| num_partitions | Optional | None | Number of partitions to use. If None, the number of partitions will be equal to the available parallel resources in distributed environments. |
+| feature_contributions | bool | False | |
+| **Returns** | **AnyDFType** | | **DataFrame with TimeGPT forecasts for point predictions and probabilistic predictions (if level is not None).** |
+
+------------------------------------------------------------------------
+
+source
+
+## NixtlaClient.cross_validation
+
+> ``` text
+> NixtlaClient.cross_validation (df:~AnyDFType,
+> h:typing.Annotated[int,Gt(gt=0)], freq:Uni
+> on[str,int,pandas._libs.tslibs.offsets.Bas
+> eOffset,NoneType]=None,
+> id_col:str='unique_id', time_col:str='ds',
+> target_col:str='y', level:Optional[list[Un
+> ion[int,float]]]=None,
+> quantiles:Optional[list[float]]=None,
+> validate_api_key:bool=False, n_windows:typ
+> ing.Annotated[int,Gt(gt=0)]=1, step_size:O
+> ptional[Annotated[int,Gt(gt=0)]]=None, fin
+> etune_steps:typing.Annotated[int,Ge(ge=0)]
+> =0, finetune_depth:Literal[1,2,3,4,5]=1, f
+> inetune_loss:Literal['default','mae','mse'
+> ,'rmse','mape','smape']='default',
+> finetuned_model_id:Optional[str]=None,
+> refit:bool=True, clean_ex_first:bool=True,
+> hist_exog_list:Optional[list[str]]=None,
+> date_features:Union[bool,list[str]]=False,
+> date_features_to_one_hot:Union[bool,list[s
+> tr]]=False, model:Literal['azureai','timeg
+> pt-1','timegpt-1-long-
+> horizon']='timegpt-1', num_partitions:Opti
+> onal[Annotated[int,Gt(gt=0)]]=None)
+> ```
+
+*Perform cross validation in your time series using TimeGPT.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | AnyDFType | | The DataFrame on which the function will operate. Expected to contain at least the following columns: - time_col: Column name in `df` that contains the time indices of the time series. This is typically a datetime column with regular intervals, e.g., hourly, daily, monthly data points. - target_col: Column name in `df` that contains the target variable of the time series, i.e., the variable we wish to predict or analyze. Additionally, you can pass multiple time series (stacked in the dataframe) considering an additional column: - id_col: Column name in `df` that identifies unique time series. Each unique value in this column corresponds to a unique time series. |
+| h | Annotated | | Forecast horizon. |
+| freq | Union | None | Frequency of the timestamps. If `None`, it will be inferred automatically. See [pandas’ available frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases). |
+| id_col | str | unique_id | Column that identifies each series. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| level | Optional | None | Confidence level between 0 and 100 for prediction intervals. |
+| quantiles | Optional | None | Quantiles to forecast, list between (0, 1). `level` and `quantiles` should not be used simultaneously. The output dataframe will have the quantile columns formatted as TimeGPT-q-(100 \* q) for each q. 100 \* q represents percentiles but we choose this notation to avoid having dots in column names. |
+| validate_api_key | bool | False | If True, validates api_key before sending requests. |
+| n_windows | Annotated | 1 | Number of windows to evaluate. |
+| step_size | Optional | None | Step size between each cross validation window. If None it will be equal to `h`. |
+| finetune_steps | Annotated | 0 | Number of steps used to finetune TimeGPT in the new data. |
+| finetune_depth | Literal | 1 | The depth of the finetuning. Uses a scale from 1 to 5, where 1 means little finetuning, and 5 means that the entire model is finetuned. |
+| finetune_loss | Literal | default | Loss function to use for finetuning. Options are: `default`, `mae`, `mse`, `rmse`, `mape`, and `smape`. |
+| finetuned_model_id | Optional | None | ID of previously fine-tuned model to use. |
+| refit | bool | True | Fine-tune the model in each window. If `False`, only fine-tunes on the first window. Only used if `finetune_steps` \> 0. |
+| clean_ex_first | bool | True | Clean exogenous signal before making forecasts using TimeGPT. |
+| hist_exog_list | Optional | None | Column names of the historical exogenous features. |
+| date_features | Union | False | Features computed from the dates. Can be pandas date attributes or functions that will take the dates as input. If True automatically adds most used date features for the frequency of `df`. |
+| date_features_to_one_hot | Union | False | Apply one-hot encoding to these date features. If `date_features=True`, then all date features are one-hot encoded by default. |
+| model | Literal | timegpt-1 | Model to use as a string. Options are: `timegpt-1`, and `timegpt-1-long-horizon`. We recommend using `timegpt-1-long-horizon` for forecasting if you want to predict more than one seasonal period given the frequency of your data. |
+| num_partitions | Optional | None | Number of partitions to use. If None, the number of partitions will be equal to the available parallel resources in distributed environments. |
+| **Returns** | **AnyDFType** | | **DataFrame with cross validation forecasts.** |
+
+------------------------------------------------------------------------
+
+source
+
+## NixtlaClient.detect_anomalies
+
+> ``` text
+> NixtlaClient.detect_anomalies (df:~AnyDFType,
+> freq:Union[str,int,pandas._libs.tslibs.off
+> sets.BaseOffset,NoneType]=None,
+> id_col:str='unique_id', time_col:str='ds',
+> target_col:str='y',
+> level:Union[int,float]=99,
+> finetuned_model_id:Optional[str]=None,
+> clean_ex_first:bool=True,
+> validate_api_key:bool=False,
+> date_features:Union[bool,list[str]]=False,
+> date_features_to_one_hot:Union[bool,list[s
+> tr]]=False, model:Literal['azureai','timeg
+> pt-1','timegpt-1-long-
+> horizon']='timegpt-1', num_partitions:Opti
+> onal[Annotated[int,Gt(gt=0)]]=None)
+> ```
+
+*Detect anomalies in your time series using TimeGPT.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | AnyDFType | | The DataFrame on which the function will operate. Expected to contain at least the following columns: - time_col: Column name in `df` that contains the time indices of the time series. This is typically a datetime column with regular intervals, e.g., hourly, daily, monthly data points. - target_col: Column name in `df` that contains the target variable of the time series, i.e., the variable we wish to predict or analyze. Additionally, you can pass multiple time series (stacked in the dataframe) considering an additional column: - id_col: Column name in `df` that identifies unique time series. Each unique value in this column corresponds to a unique time series. |
+| freq | Union | None | Frequency of the timestamps. If `None`, it will be inferred automatically. See [pandas’ available frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases). |
+| id_col | str | unique_id | Column that identifies each series. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| level | Union | 99 | Confidence level between 0 and 100 for detecting the anomalies. |
+| finetuned_model_id | Optional | None | ID of previously fine-tuned model to use. |
+| clean_ex_first | bool | True | Clean exogenous signal before making forecasts using TimeGPT. |
+| validate_api_key | bool | False | If True, validates api_key before sending requests. |
+| date_features | Union | False | Features computed from the dates. Can be pandas date attributes or functions that will take the dates as input. If True automatically adds most used date features for the frequency of `df`. |
+| date_features_to_one_hot | Union | False | Apply one-hot encoding to these date features. If `date_features=True`, then all date features are one-hot encoded by default. |
+| model | Literal | timegpt-1 | Model to use as a string. Options are: `timegpt-1`, and `timegpt-1-long-horizon`. We recommend using `timegpt-1-long-horizon` for forecasting if you want to predict more than one seasonal period given the frequency of your data. |
+| num_partitions | Optional | None | Number of partitions to use. If None, the number of partitions will be equal to the available parallel resources in distributed environments. |
+| **Returns** | **AnyDFType** | | **DataFrame with anomalies flagged by TimeGPT.** |
+
+------------------------------------------------------------------------
+
+source
+
+## NixtlaClient.usage
+
+> ``` text
+> NixtlaClient.usage ()
+> ```
+
+*Query consumed requests and limits*
+
+------------------------------------------------------------------------
+
+source
+
+## NixtlaClient.finetune
+
+> ``` text
+> NixtlaClient.finetune
+> (df:Union[pandas.core.frame.DataFrame,polars.dataf
+> rame.frame.DataFrame], freq:Union[str,int,pandas._
+> libs.tslibs.offsets.BaseOffset,NoneType]=None,
+> id_col:str='unique_id', time_col:str='ds',
+> target_col:str='y',
+> finetune_steps:typing.Annotated[int,Ge(ge=0)]=10,
+> finetune_depth:Literal[1,2,3,4,5]=1, finetune_loss
+> :Literal['default','mae','mse','rmse','mape','smap
+> e']='default', output_model_id:Optional[str]=None,
+> finetuned_model_id:Optional[str]=None, model:Liter
+> al['azureai','timegpt-1','timegpt-1-long-
+> horizon']='timegpt-1')
+> ```
+
+*Fine-tune TimeGPT to your series.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | Union | | The DataFrame on which the function will operate. Expected to contain at least the following columns: - time_col: Column name in `df` that contains the time indices of the time series. This is typically a datetime column with regular intervals, e.g., hourly, daily, monthly data points. - target_col: Column name in `df` that contains the target variable of the time series, i.e., the variable we wish to predict or analyze. Additionally, you can pass multiple time series (stacked in the dataframe) considering an additional column: - id_col: Column name in `df` that identifies unique time series. Each unique value in this column corresponds to a unique time series. |
+| freq | Union | None | Frequency of the timestamps. If `None`, it will be inferred automatically. See [pandas’ available frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases). |
+| id_col | str | unique_id | Column that identifies each series. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| finetune_steps | Annotated | 10 | Number of steps used to finetune learning TimeGPT in the new data. |
+| finetune_depth | Literal | 1 | The depth of the finetuning. Uses a scale from 1 to 5, where 1 means little finetuning, and 5 means that the entire model is finetuned. |
+| finetune_loss | Literal | default | Loss function to use for finetuning. Options are: `default`, `mae`, `mse`, `rmse`, `mape`, and `smape`. |
+| output_model_id | Optional | None | ID to assign to the fine-tuned model. If `None`, an UUID is used. |
+| finetuned_model_id | Optional | None | ID of previously fine-tuned model to use as base. |
+| model | Literal | timegpt-1 | Model to use as a string. Options are: `timegpt-1`, and `timegpt-1-long-horizon`. We recommend using `timegpt-1-long-horizon` for forecasting if you want to predict more than one seasonal period given the frequency of your data. |
+| **Returns** | **str** | | **ID of the fine-tuned model** |
+
+------------------------------------------------------------------------
+
+source
+
+## NixtlaClient.finetuned_models
+
+> ``` text
+> NixtlaClient.finetuned_models (as_df:bool=False)
+> ```
+
+*List fine-tuned models*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| as_df | bool | False | Return the fine-tuned models as a pandas dataframe |
+| **Returns** | **Union** | | **List of available fine-tuned models.** |
+
+------------------------------------------------------------------------
+
+source
+
+## NixtlaClient.finetuned_model
+
+> ``` text
+> NixtlaClient.finetuned_model (finetuned_model_id:str)
+> ```
+
+*Get fine-tuned model metadata*
+
+| | **Type** | **Details** |
+|--------|---------------------------|-------------------------------------|
+| finetuned_model_id | str | ID of the fine-tuned model to get metadata from. |
+| **Returns** | **FinetunedModel** | **Fine-tuned model metadata.** |
+
+------------------------------------------------------------------------
+
+source
+
+## NixtlaClient.delete_finetuned_model
+
+> ``` text
+> NixtlaClient.delete_finetuned_model (finetuned_model_id:str)
+> ```
+
+*Delete a previously fine-tuned model*
+
+| | **Type** | **Details** |
+|--------------------|----------|-------------------------------------------|
+| finetuned_model_id | str | ID of the fine-tuned model to be deleted. |
+| **Returns** | **bool** | **Whether delete was successful.** |
+
+------------------------------------------------------------------------
+
+source
+
+## NixtlaClient.plot
+
+> ``` text
+> NixtlaClient.plot (df:Union[pandas.core.frame.DataFrame,polars.dataframe.
+> frame.DataFrame,NoneType]=None, forecasts_df:Union[pan
+> das.core.frame.DataFrame,polars.dataframe.frame.DataFr
+> ame,NoneType]=None, id_col:str='unique_id',
+> time_col:str='ds', target_col:str='y', unique_ids:Unio
+> n[list[str],NoneType,numpy.ndarray]=None,
+> plot_random:bool=True, max_ids:int=8,
+> models:Optional[list[str]]=None,
+> level:Optional[list[Union[int,float]]]=None,
+> max_insample_length:Optional[int]=None,
+> plot_anomalies:bool=False,
+> engine:Literal['matplotlib','plotly','plotly-
+> resampler']='matplotlib',
+> resampler_kwargs:Optional[dict]=None, ax:Union[Forward
+> Ref('plt.Axes'),numpy.ndarray,ForwardRef('plotly.graph
+> _objects.Figure'),NoneType]=None)
+> ```
+
+*Plot forecasts and insample values.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | Union | None | The DataFrame on which the function will operate. Expected to contain at least the following columns: - time_col: Column name in `df` that contains the time indices of the time series. This is typically a datetime column with regular intervals, e.g., hourly, daily, monthly data points. - target_col: Column name in `df` that contains the target variable of the time series, i.e., the variable we wish to predict or analyze. Additionally, you can pass multiple time series (stacked in the dataframe) considering an additional column: - id_col: Column name in `df` that identifies unique time series. Each unique value in this column corresponds to a unique time series. |
+| forecasts_df | Union | None | DataFrame with columns \[`unique_id`, `ds`\] and models. |
+| id_col | str | unique_id | Column that identifies each series. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| unique_ids | Union | None | Time Series to plot. If None, time series are selected randomly. |
+| plot_random | bool | True | Select time series to plot randomly. |
+| max_ids | int | 8 | Maximum number of ids to plot. |
+| models | Optional | None | list of models to plot. |
+| level | Optional | None | list of prediction intervals to plot if paseed. |
+| max_insample_length | Optional | None | Max number of train/insample observations to be plotted. |
+| plot_anomalies | bool | False | Plot anomalies for each prediction interval. |
+| engine | Literal | matplotlib | Library used to plot. ‘matplotlib’, ‘plotly’ or ‘plotly-resampler’. |
+| resampler_kwargs | Optional | None | Kwargs to be passed to plotly-resampler constructor. For further custumization (“show_dash”) call the method, store the plotting object and add the extra arguments to its `show_dash` method. |
+| ax | Union | None | Object where plots will be added. |
+
diff --git a/nixtla/docs/reference/nixtlar.html.mdx b/nixtla/docs/reference/nixtlar.html.mdx
new file mode 100644
index 00000000..0750ac6c
--- /dev/null
+++ b/nixtla/docs/reference/nixtlar.html.mdx
@@ -0,0 +1,25 @@
+---
+output-file: nixtlar.html
+title: TimeGPT in R
+---
+
+
+
+
+## How to use
+
+To learn how to use `nixtlar`, please refer to the
+[documentation](https://nixtla.github.io/nixtlar/).
+
+To view directly on CRAN, please use this
+[link](https://cloud.r-project.org/web/packages/nixtlar/index.html).
+
+> 📘 API key required
+>
+> The `nixtlar` package requires an API key. Get yours on the [Nixtla
+> Dashboard](http://dashboard.nixtla.io).
+
+## Support
+
+If you have questions or need support, please email `support@nixtla.io`.
+
diff --git a/nixtla/docs/tutorials/01_exogenous_variables_files/figure-markdown_strict/cell-11-output-1.png b/nixtla/docs/tutorials/01_exogenous_variables_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..605136ce
Binary files /dev/null and b/nixtla/docs/tutorials/01_exogenous_variables_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/nixtla/docs/tutorials/01_exogenous_variables_files/figure-markdown_strict/cell-13-output-1.png b/nixtla/docs/tutorials/01_exogenous_variables_files/figure-markdown_strict/cell-13-output-1.png
new file mode 100644
index 00000000..f42b16f4
Binary files /dev/null and b/nixtla/docs/tutorials/01_exogenous_variables_files/figure-markdown_strict/cell-13-output-1.png differ
diff --git a/nixtla/docs/tutorials/01_exogenous_variables_files/figure-markdown_strict/cell-23-output-1.png b/nixtla/docs/tutorials/01_exogenous_variables_files/figure-markdown_strict/cell-23-output-1.png
new file mode 100644
index 00000000..1ff1ac09
Binary files /dev/null and b/nixtla/docs/tutorials/01_exogenous_variables_files/figure-markdown_strict/cell-23-output-1.png differ
diff --git a/nixtla/docs/tutorials/01_exogenous_variables_files/figure-markdown_strict/cell-8-output-1.png b/nixtla/docs/tutorials/01_exogenous_variables_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..b1112758
Binary files /dev/null and b/nixtla/docs/tutorials/01_exogenous_variables_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/nixtla/docs/tutorials/01_exogenous_variables_files/figure-markdown_strict/cell-9-output-1.png b/nixtla/docs/tutorials/01_exogenous_variables_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..6d601b5e
Binary files /dev/null and b/nixtla/docs/tutorials/01_exogenous_variables_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/nixtla/docs/tutorials/02_holidays_files/figure-markdown_strict/cell-11-output-1.png b/nixtla/docs/tutorials/02_holidays_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..364e211c
Binary files /dev/null and b/nixtla/docs/tutorials/02_holidays_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/nixtla/docs/tutorials/02_holidays_files/figure-markdown_strict/cell-12-output-1.png b/nixtla/docs/tutorials/02_holidays_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..3aa492ca
Binary files /dev/null and b/nixtla/docs/tutorials/02_holidays_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/nixtla/docs/tutorials/03_categorical_variables_files/figure-markdown_strict/cell-13-output-1.png b/nixtla/docs/tutorials/03_categorical_variables_files/figure-markdown_strict/cell-13-output-1.png
new file mode 100644
index 00000000..c954ff42
Binary files /dev/null and b/nixtla/docs/tutorials/03_categorical_variables_files/figure-markdown_strict/cell-13-output-1.png differ
diff --git a/nixtla/docs/tutorials/03_categorical_variables_files/figure-markdown_strict/cell-15-output-1.png b/nixtla/docs/tutorials/03_categorical_variables_files/figure-markdown_strict/cell-15-output-1.png
new file mode 100644
index 00000000..fb714504
Binary files /dev/null and b/nixtla/docs/tutorials/03_categorical_variables_files/figure-markdown_strict/cell-15-output-1.png differ
diff --git a/nixtla/docs/tutorials/04_longhorizon_files/figure-markdown_strict/cell-8-output-1.png b/nixtla/docs/tutorials/04_longhorizon_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..fbdff06c
Binary files /dev/null and b/nixtla/docs/tutorials/04_longhorizon_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/nixtla/docs/tutorials/05_multiple_series_files/figure-markdown_strict/cell-10-output-1.png b/nixtla/docs/tutorials/05_multiple_series_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..6bc23c47
Binary files /dev/null and b/nixtla/docs/tutorials/05_multiple_series_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/nixtla/docs/tutorials/05_multiple_series_files/figure-markdown_strict/cell-6-output-1.png b/nixtla/docs/tutorials/05_multiple_series_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..788ffdb2
Binary files /dev/null and b/nixtla/docs/tutorials/05_multiple_series_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/nixtla/docs/tutorials/05_multiple_series_files/figure-markdown_strict/cell-8-output-1.png b/nixtla/docs/tutorials/05_multiple_series_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..79ec0018
Binary files /dev/null and b/nixtla/docs/tutorials/05_multiple_series_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/nixtla/docs/tutorials/06_finetuning_files/figure-markdown_strict/cell-7-output-1.png b/nixtla/docs/tutorials/06_finetuning_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..b3cf360e
Binary files /dev/null and b/nixtla/docs/tutorials/06_finetuning_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/nixtla/docs/tutorials/07_loss_function_finetuning_files/figure-markdown_strict/cell-7-output-1.png b/nixtla/docs/tutorials/07_loss_function_finetuning_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..58514318
Binary files /dev/null and b/nixtla/docs/tutorials/07_loss_function_finetuning_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-11-output-1.png b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..7147f80b
Binary files /dev/null and b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-11-output-2.png b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-11-output-2.png
new file mode 100644
index 00000000..8d020bc5
Binary files /dev/null and b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-11-output-2.png differ
diff --git a/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-11-output-3.png b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-11-output-3.png
new file mode 100644
index 00000000..d6dbcf98
Binary files /dev/null and b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-11-output-3.png differ
diff --git a/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-11-output-4.png b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-11-output-4.png
new file mode 100644
index 00000000..d3e117d8
Binary files /dev/null and b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-11-output-4.png differ
diff --git a/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-11-output-5.png b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-11-output-5.png
new file mode 100644
index 00000000..c13e3d32
Binary files /dev/null and b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-11-output-5.png differ
diff --git a/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-13-output-2.png b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-13-output-2.png
new file mode 100644
index 00000000..d0f59853
Binary files /dev/null and b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-13-output-2.png differ
diff --git a/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-13-output-3.png b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-13-output-3.png
new file mode 100644
index 00000000..f207c4ca
Binary files /dev/null and b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-13-output-3.png differ
diff --git a/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-14-output-2.png b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-14-output-2.png
new file mode 100644
index 00000000..83fc885f
Binary files /dev/null and b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-14-output-2.png differ
diff --git a/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-14-output-3.png b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-14-output-3.png
new file mode 100644
index 00000000..4f44d572
Binary files /dev/null and b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-14-output-3.png differ
diff --git a/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-7-output-1.png b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..e68cc77e
Binary files /dev/null and b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-7-output-2.png b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-7-output-2.png
new file mode 100644
index 00000000..b9e0e5ad
Binary files /dev/null and b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-7-output-2.png differ
diff --git a/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-7-output-3.png b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-7-output-3.png
new file mode 100644
index 00000000..33f7b749
Binary files /dev/null and b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-7-output-3.png differ
diff --git a/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-7-output-4.png b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-7-output-4.png
new file mode 100644
index 00000000..bfee6116
Binary files /dev/null and b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-7-output-4.png differ
diff --git a/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-7-output-5.png b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-7-output-5.png
new file mode 100644
index 00000000..fc6236a7
Binary files /dev/null and b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-7-output-5.png differ
diff --git a/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-9-output-1.png b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..b2c72d41
Binary files /dev/null and b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-9-output-2.png b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-9-output-2.png
new file mode 100644
index 00000000..1dbe580e
Binary files /dev/null and b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-9-output-2.png differ
diff --git a/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-9-output-3.png b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-9-output-3.png
new file mode 100644
index 00000000..373ae3c4
Binary files /dev/null and b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-9-output-3.png differ
diff --git a/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-9-output-4.png b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-9-output-4.png
new file mode 100644
index 00000000..a171c3d3
Binary files /dev/null and b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-9-output-4.png differ
diff --git a/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-9-output-5.png b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-9-output-5.png
new file mode 100644
index 00000000..8e373834
Binary files /dev/null and b/nixtla/docs/tutorials/08_cross_validation_files/figure-markdown_strict/cell-9-output-5.png differ
diff --git a/nixtla/docs/tutorials/09_historical_forecast_files/figure-markdown_strict/cell-6-output-1.png b/nixtla/docs/tutorials/09_historical_forecast_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..b6fb5ba7
Binary files /dev/null and b/nixtla/docs/tutorials/09_historical_forecast_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/nixtla/docs/tutorials/09_historical_forecast_files/figure-markdown_strict/cell-9-output-1.png b/nixtla/docs/tutorials/09_historical_forecast_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..15ff6920
Binary files /dev/null and b/nixtla/docs/tutorials/09_historical_forecast_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-11-output-1.png b/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..0ff5930d
Binary files /dev/null and b/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-11-output-2.png b/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-11-output-2.png
new file mode 100644
index 00000000..c97034a9
Binary files /dev/null and b/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-11-output-2.png differ
diff --git a/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-11-output-3.png b/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-11-output-3.png
new file mode 100644
index 00000000..016708ed
Binary files /dev/null and b/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-11-output-3.png differ
diff --git a/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-11-output-4.png b/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-11-output-4.png
new file mode 100644
index 00000000..fcfac1b1
Binary files /dev/null and b/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-11-output-4.png differ
diff --git a/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-11-output-5.png b/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-11-output-5.png
new file mode 100644
index 00000000..4aae80bc
Binary files /dev/null and b/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-11-output-5.png differ
diff --git a/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-7-output-1.png b/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..9feacd2e
Binary files /dev/null and b/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-9-output-1.png b/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..23b2b046
Binary files /dev/null and b/nixtla/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/nixtla/docs/tutorials/11_uncertainty_quantification_with_prediction_intervals_files/figure-markdown_strict/cell-7-output-1.png b/nixtla/docs/tutorials/11_uncertainty_quantification_with_prediction_intervals_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..5ba7a01e
Binary files /dev/null and b/nixtla/docs/tutorials/11_uncertainty_quantification_with_prediction_intervals_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/nixtla/docs/tutorials/11_uncertainty_quantification_with_prediction_intervals_files/figure-markdown_strict/cell-9-output-1.png b/nixtla/docs/tutorials/11_uncertainty_quantification_with_prediction_intervals_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..0d008df0
Binary files /dev/null and b/nixtla/docs/tutorials/11_uncertainty_quantification_with_prediction_intervals_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/nixtla/docs/tutorials/13_bounded_forecasts_files/figure-markdown_strict/cell-11-output-1.png b/nixtla/docs/tutorials/13_bounded_forecasts_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..2b10015d
Binary files /dev/null and b/nixtla/docs/tutorials/13_bounded_forecasts_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/nixtla/docs/tutorials/13_bounded_forecasts_files/figure-markdown_strict/cell-13-output-1.png b/nixtla/docs/tutorials/13_bounded_forecasts_files/figure-markdown_strict/cell-13-output-1.png
new file mode 100644
index 00000000..1a5fb5ac
Binary files /dev/null and b/nixtla/docs/tutorials/13_bounded_forecasts_files/figure-markdown_strict/cell-13-output-1.png differ
diff --git a/nixtla/docs/tutorials/13_bounded_forecasts_files/figure-markdown_strict/cell-7-output-1.png b/nixtla/docs/tutorials/13_bounded_forecasts_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..d9ca609d
Binary files /dev/null and b/nixtla/docs/tutorials/13_bounded_forecasts_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/nixtla/docs/tutorials/14_hierarchical_forecasting_files/figure-markdown_strict/cell-12-output-1.png b/nixtla/docs/tutorials/14_hierarchical_forecasting_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..76b30f49
Binary files /dev/null and b/nixtla/docs/tutorials/14_hierarchical_forecasting_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/nixtla/docs/tutorials/14_hierarchical_forecasting_files/figure-markdown_strict/cell-16-output-1.png b/nixtla/docs/tutorials/14_hierarchical_forecasting_files/figure-markdown_strict/cell-16-output-1.png
new file mode 100644
index 00000000..118caa58
Binary files /dev/null and b/nixtla/docs/tutorials/14_hierarchical_forecasting_files/figure-markdown_strict/cell-16-output-1.png differ
diff --git a/nixtla/docs/tutorials/15_missing_values_files/figure-markdown_strict/cell-10-output-1.png b/nixtla/docs/tutorials/15_missing_values_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..ce34bc47
Binary files /dev/null and b/nixtla/docs/tutorials/15_missing_values_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/nixtla/docs/tutorials/15_missing_values_files/figure-markdown_strict/cell-11-output-1.png b/nixtla/docs/tutorials/15_missing_values_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..9c9fc61f
Binary files /dev/null and b/nixtla/docs/tutorials/15_missing_values_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/nixtla/docs/tutorials/15_missing_values_files/figure-markdown_strict/cell-17-output-1.png b/nixtla/docs/tutorials/15_missing_values_files/figure-markdown_strict/cell-17-output-1.png
new file mode 100644
index 00000000..6a65b384
Binary files /dev/null and b/nixtla/docs/tutorials/15_missing_values_files/figure-markdown_strict/cell-17-output-1.png differ
diff --git a/nixtla/docs/tutorials/20_anomaly_detection_files/figure-markdown_strict/cell-10-output-1.png b/nixtla/docs/tutorials/20_anomaly_detection_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..e6a9d0e7
Binary files /dev/null and b/nixtla/docs/tutorials/20_anomaly_detection_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/nixtla/docs/tutorials/20_anomaly_detection_files/figure-markdown_strict/cell-12-output-1.png b/nixtla/docs/tutorials/20_anomaly_detection_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..95cf55c7
Binary files /dev/null and b/nixtla/docs/tutorials/20_anomaly_detection_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/nixtla/docs/tutorials/20_anomaly_detection_files/figure-markdown_strict/cell-6-output-1.png b/nixtla/docs/tutorials/20_anomaly_detection_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..bdd27162
Binary files /dev/null and b/nixtla/docs/tutorials/20_anomaly_detection_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/nixtla/docs/tutorials/20_anomaly_detection_files/figure-markdown_strict/cell-8-output-1.png b/nixtla/docs/tutorials/20_anomaly_detection_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..fb148b15
Binary files /dev/null and b/nixtla/docs/tutorials/20_anomaly_detection_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/nixtla/docs/tutorials/21_shap_values_files/figure-markdown_strict/cell-10-output-1.png b/nixtla/docs/tutorials/21_shap_values_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..5aa97626
Binary files /dev/null and b/nixtla/docs/tutorials/21_shap_values_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/nixtla/docs/tutorials/21_shap_values_files/figure-markdown_strict/cell-11-output-1.png b/nixtla/docs/tutorials/21_shap_values_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..2d4164c4
Binary files /dev/null and b/nixtla/docs/tutorials/21_shap_values_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/nixtla/docs/tutorials/21_shap_values_files/figure-markdown_strict/cell-9-output-1.png b/nixtla/docs/tutorials/21_shap_values_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..791eda05
Binary files /dev/null and b/nixtla/docs/tutorials/21_shap_values_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-11-output-1.png b/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..39dba094
Binary files /dev/null and b/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-14-output-1.png b/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..dbfa4e3a
Binary files /dev/null and b/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-17-output-1.png b/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-17-output-1.png
new file mode 100644
index 00000000..d4728fd4
Binary files /dev/null and b/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-17-output-1.png differ
diff --git a/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-20-output-1.png b/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-20-output-1.png
new file mode 100644
index 00000000..f08c108d
Binary files /dev/null and b/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-20-output-1.png differ
diff --git a/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-25-output-1.png b/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-25-output-1.png
new file mode 100644
index 00000000..d305de40
Binary files /dev/null and b/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-25-output-1.png differ
diff --git a/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-28-output-1.png b/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-28-output-1.png
new file mode 100644
index 00000000..6c4e8dc1
Binary files /dev/null and b/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-28-output-1.png differ
diff --git a/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-7-output-1.png b/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..700a442c
Binary files /dev/null and b/nixtla/docs/tutorials/22_how_to_improve_forecast_accuracy_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/nixtla/docs/tutorials/23_temporalhierarchical_files/figure-markdown_strict/cell-20-output-1.png b/nixtla/docs/tutorials/23_temporalhierarchical_files/figure-markdown_strict/cell-20-output-1.png
new file mode 100644
index 00000000..853d865c
Binary files /dev/null and b/nixtla/docs/tutorials/23_temporalhierarchical_files/figure-markdown_strict/cell-20-output-1.png differ
diff --git a/nixtla/docs/tutorials/23_temporalhierarchical_files/figure-markdown_strict/cell-21-output-1.png b/nixtla/docs/tutorials/23_temporalhierarchical_files/figure-markdown_strict/cell-21-output-1.png
new file mode 100644
index 00000000..ab0fff11
Binary files /dev/null and b/nixtla/docs/tutorials/23_temporalhierarchical_files/figure-markdown_strict/cell-21-output-1.png differ
diff --git a/nixtla/docs/tutorials/23_temporalhierarchical_files/figure-markdown_strict/cell-7-output-1.png b/nixtla/docs/tutorials/23_temporalhierarchical_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..1ce33ae9
Binary files /dev/null and b/nixtla/docs/tutorials/23_temporalhierarchical_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/nixtla/docs/tutorials/anomaly_detection.html.mdx b/nixtla/docs/tutorials/anomaly_detection.html.mdx
new file mode 100644
index 00000000..ac003e99
--- /dev/null
+++ b/nixtla/docs/tutorials/anomaly_detection.html.mdx
@@ -0,0 +1,180 @@
+---
+output-file: anomaly_detection.html
+title: Anomaly detection
+---
+
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/20_anomaly_detection.ipynb)
+
+## Import packages
+
+First, we import the required packages for this tutorial and create an
+instance of
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient).
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, set the `base_url` argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## Load dataset
+
+Now, let’s load the dataset for this tutorial. We use the Peyton Manning
+dataset which tracks the visits to the Wikipedia page of Peyton Mannig.
+
+```python
+df = pd.read_csv('https://datasets-nixtla.s3.amazonaws.com/peyton-manning.csv')
+df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|----------|
+| 0 | 0 | 2007-12-10 | 9.590761 |
+| 1 | 0 | 2007-12-11 | 8.519590 |
+| 2 | 0 | 2007-12-12 | 8.183677 |
+| 3 | 0 | 2007-12-13 | 8.072467 |
+| 4 | 0 | 2007-12-14 | 7.893572 |
+
+```python
+nixtla_client.plot(df, max_insample_length=365)
+```
+
+
+
+## Anomaly detection
+
+We now perform anomaly detection. By default, TimeGPT uses a 99%
+confidence interval. If a point falls outisde of that interval, it is
+considered to be an anomaly.
+
+```python
+anomalies_df = nixtla_client.detect_anomalies(df, freq='D')
+anomalies_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Querying model metadata...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Calling Anomaly Detector Endpoint...
+```
+
+| | unique_id | ds | y | TimeGPT | TimeGPT-hi-99 | TimeGPT-lo-99 | anomaly |
+|-----|-----------|------------|-----------|----------|---------------|---------------|---------|
+| 0 | 0 | 2008-01-10 | 8.281724 | 8.224187 | 9.503586 | 6.944788 | False |
+| 1 | 0 | 2008-01-11 | 8.292799 | 8.151533 | 9.430932 | 6.872135 | False |
+| 2 | 0 | 2008-01-12 | 8.199189 | 8.127243 | 9.406642 | 6.847845 | False |
+| 3 | 0 | 2008-01-13 | 9.996522 | 8.917259 | 10.196658 | 7.637861 | False |
+| 4 | 0 | 2008-01-14 | 10.127071 | 9.002326 | 10.281725 | 7.722928 | False |
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.detect_anomalies(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+As you can see, `False` is assigned to “normal” values, as they fall
+inside the confidence interval. A label of `True` is then assigned to
+abnormal points.
+
+We can also plot the anomalies using
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient).
+
+```python
+nixtla_client.plot(df, anomalies_df)
+```
+
+
+
+## Anomaly detection with exogenous features
+
+Previously, we performed anomaly detection without using any exogenous
+features. Now, it is possible to create features specifically for this
+scenario to inform the model in its task of anomaly detection.
+
+Here, we create date features that can be used by the model.
+
+This is done using the `date_features` argument. We can set it to `True`
+and it will generate all possible features from the given dates and
+frequency of the data. Alternatively, we can specify a list of features
+that we want. In this case, we want only features at the *month* and
+*year* level.
+
+```python
+anomalies_df_x = nixtla_client.detect_anomalies(
+ df,
+ freq='D',
+ date_features=['month', 'year'],
+ date_features_to_one_hot=True,
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Using the following exogenous features: ['month_1.0', 'month_2.0', 'month_3.0', 'month_4.0', 'month_5.0', 'month_6.0', 'month_7.0', 'month_8.0', 'month_9.0', 'month_10.0', 'month_11.0', 'month_12.0', 'year_2007.0', 'year_2008.0', 'year_2009.0', 'year_2010.0', 'year_2011.0', 'year_2012.0', 'year_2013.0', 'year_2014.0', 'year_2015.0', 'year_2016.0']
+INFO:nixtla.nixtla_client:Calling Anomaly Detector Endpoint...
+```
+
+Then, we can plot the detected anomalies where the model now used
+additional information from exogenous features.
+
+```python
+nixtla_client.plot(df, anomalies_df_x)
+```
+
+
+
+## Modifying the confidence intervals
+
+We can tweak the confidence intervals using the `level` argument. This
+takes any values between 0 and 100, including decimal numbers.
+
+Reducing the confidence interval resutls in more anomalies being
+detected, while increasing it will reduce the number of anomalies.
+
+Here, for example, we reduce the interval to 70%, and we will notice
+more anomalies being plotted (red dots).
+
+```python
+anomalies_df = nixtla_client.detect_anomalies(
+ df,
+ freq='D',
+ level=70
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Calling Anomaly Detector Endpoint...
+```
+
+```python
+nixtla_client.plot(df, anomalies_df)
+```
+
+
+
diff --git a/nixtla/docs/tutorials/bounded_forecasts.html.mdx b/nixtla/docs/tutorials/bounded_forecasts.html.mdx
new file mode 100644
index 00000000..c4583db9
--- /dev/null
+++ b/nixtla/docs/tutorials/bounded_forecasts.html.mdx
@@ -0,0 +1,225 @@
+---
+output-file: bounded_forecasts.html
+title: Bounded forecasts
+---
+
+
+In forecasting, we often want to make sure the predictions stay within a
+certain range. For example, for predicting the sales of a product, we
+may require all forecasts to be positive. Thus, the forecasts may need
+to be bounded.
+
+With TimeGPT, you can create bounded forecasts by transforming your data
+prior to calling the forecast function.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/13_bounded_forecasts.ipynb)
+
+## 1. Import packages
+
+First, we install and import the required packages
+
+```python
+import pandas as pd
+import numpy as np
+
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, set the `base_url` argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 2. Load data
+
+We use the [annual egg
+prices](https://github.com/robjhyndman/fpp3package/tree/master/data)
+dataset from [Forecasting, Principles and
+Practices](https://otexts.com/fpp3/). We expect egg prices to be
+strictly positive, so we want to bound our forecasts to be positive.
+
+> **Note**
+>
+> You can install `pyreadr` with `pip`:
+>
+> ```shell
+> pip install pyreadr
+> ```
+
+```python
+import pyreadr
+from pathlib import Path
+
+# Download and store the dataset
+url = 'https://github.com/robjhyndman/fpp3package/raw/master/data/prices.rda'
+dst_path = str(Path.cwd().joinpath('prices.rda'))
+result = pyreadr.read_r(pyreadr.download_file(url, dst_path), dst_path)
+```
+
+
+```python
+# Perform some preprocessing
+df = result['prices'][['year', 'eggs']]
+df = df.dropna().reset_index(drop=True)
+df = df.rename(columns={'year':'ds', 'eggs':'y'})
+df['ds'] = pd.to_datetime(df['ds'], format='%Y')
+df['unique_id'] = 'eggs'
+
+df.tail(10)
+```
+
+| | ds | y | unique_id |
+|-----|------------|--------|-----------|
+| 84 | 1984-01-01 | 100.58 | eggs |
+| 85 | 1985-01-01 | 76.84 | eggs |
+| 86 | 1986-01-01 | 81.10 | eggs |
+| 87 | 1987-01-01 | 69.60 | eggs |
+| 88 | 1988-01-01 | 64.55 | eggs |
+| 89 | 1989-01-01 | 80.36 | eggs |
+| 90 | 1990-01-01 | 79.79 | eggs |
+| 91 | 1991-01-01 | 74.79 | eggs |
+| 92 | 1992-01-01 | 64.86 | eggs |
+| 93 | 1993-01-01 | 62.27 | eggs |
+
+We can have a look at how the prices have evolved in the 20th century,
+demonstrating that the price is trending down.
+
+```python
+nixtla_client.plot(df)
+```
+
+
+
+## 3. Bounded forecasts with TimeGPT
+
+First, we transform the target data. In this case, we will log-transform
+the data prior to forecasting, such that we can only forecast positive
+prices.
+
+```python
+df_transformed = df.copy()
+df_transformed['y'] = np.log(df_transformed['y'])
+```
+
+We will create forecasts for the next 10 years, and we include an 80, 90
+and 99.5 percentile of our forecast distribution.
+
+```python
+timegpt_fcst_with_transform = nixtla_client.forecast(df=df_transformed, h=10, freq='Y', level=[80, 90, 99.5])
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: AS-JAN
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+After having created the forecasts, we need to inverse the
+transformation that we applied earlier. With a log-transformation, this
+simply means we need to exponentiate the forecasts:
+
+```python
+cols_to_transform = [col for col in timegpt_fcst_with_transform if col not in ['unique_id', 'ds']]
+for col in cols_to_transform:
+ timegpt_fcst_with_transform[col] = np.exp(timegpt_fcst_with_transform[col])
+```
+
+Now, we can plot the forecasts. We include a number of prediction
+intervals, indicating the 80, 90 and 99.5 percentile of our forecast
+distribution.
+
+```python
+nixtla_client.plot(
+ df,
+ timegpt_fcst_with_transform,
+ level=[80, 90, 99.5],
+ max_insample_length=20
+)
+```
+
+
+
+The forecast and the prediction intervals look reasonable.
+
+Let’s compare these forecasts to the situation where we don’t apply a
+transformation. In this case, it may be possible to forecast a negative
+price.
+
+```python
+timegpt_fcst_without_transform = nixtla_client.forecast(df=df, h=10, freq='Y', level=[80, 90, 99.5])
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: AS-JAN
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+Indeed, we now observe prediction intervals that become negative:
+
+```python
+nixtla_client.plot(
+ df,
+ timegpt_fcst_without_transform,
+ level=[80, 90, 99.5],
+ max_insample_length=20
+)
+```
+
+
+
+For example, in 1995:
+
+```python
+timegpt_fcst_without_transform
+```
+
+| | unique_id | ds | TimeGPT | TimeGPT-lo-99.5 | TimeGPT-lo-90 | TimeGPT-lo-80 | TimeGPT-hi-80 | TimeGPT-hi-90 | TimeGPT-hi-99.5 |
+|----|----|----|----|----|----|----|----|----|----|
+| 0 | eggs | 1994-01-01 | 66.859756 | 43.103240 | 46.131448 | 49.319034 | 84.400479 | 87.588065 | 90.616273 |
+| 1 | eggs | 1995-01-01 | 64.993477 | -20.924112 | -4.750041 | 12.275298 | 117.711656 | 134.736995 | 150.911066 |
+| 2 | eggs | 1996-01-01 | 66.695808 | 6.499170 | 8.291150 | 10.177444 | 123.214173 | 125.100467 | 126.892446 |
+| 3 | eggs | 1997-01-01 | 66.103325 | 17.304282 | 24.966939 | 33.032894 | 99.173756 | 107.239711 | 114.902368 |
+| 4 | eggs | 1998-01-01 | 67.906517 | 4.995371 | 12.349648 | 20.090992 | 115.722042 | 123.463386 | 130.817663 |
+| 5 | eggs | 1999-01-01 | 66.147575 | 29.162207 | 31.804460 | 34.585779 | 97.709372 | 100.490691 | 103.132943 |
+| 6 | eggs | 2000-01-01 | 66.062637 | 14.671932 | 19.305822 | 24.183601 | 107.941673 | 112.819453 | 117.453343 |
+| 7 | eggs | 2001-01-01 | 68.045769 | 3.915282 | 13.188964 | 22.950736 | 113.140802 | 122.902573 | 132.176256 |
+| 8 | eggs | 2002-01-01 | 66.718903 | -42.212631 | -30.583703 | -18.342726 | 151.780531 | 164.021508 | 175.650436 |
+| 9 | eggs | 2003-01-01 | 67.344078 | -86.239911 | -44.959745 | -1.506939 | 136.195095 | 179.647901 | 220.928067 |
+
+This demonstrates the value of the log-transformation to obtain bounded
+forecasts with TimeGPT, which allows us to obtain better calibrated
+prediction intervals.
+
+**References**
+
+- [Hyndman, Rob J., and George Athanasopoulos (2021). “Forecasting:
+ Principles and Practice (3rd Ed)”](https://otexts.com/fpp3/)
+
diff --git a/nixtla/docs/tutorials/categorical_variables.html.mdx b/nixtla/docs/tutorials/categorical_variables.html.mdx
new file mode 100644
index 00000000..5c74f441
--- /dev/null
+++ b/nixtla/docs/tutorials/categorical_variables.html.mdx
@@ -0,0 +1,373 @@
+---
+output-file: categorical_variables.html
+title: Categorical variables
+---
+
+
+Categorical variables are external factors that can influence a
+forecast. These variables take on one of a limited, fixed number of
+possible values, and induce a grouping of your observations.
+
+For example, if you’re forecasting daily product demand for a retailer,
+you could benefit from an event variable that may tell you what kind of
+event takes place on a given day, for example ‘None’, ‘Sporting’, or
+‘Cultural’.
+
+To incorporate categorical variables in TimeGPT, you’ll need to pair
+each point in your time series data with the corresponding external
+data.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/03_categorical_variables.ipynb)
+
+## 1. Import packages
+
+First, we install and import the required packages and initialize the
+Nixtla client.
+
+```python
+import pandas as pd
+import os
+
+from nixtla import NixtlaClient
+from datasetsforecast.m5 import M5
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 2. Load M5 data
+
+Let’s see an example on predicting sales of products of the [M5
+dataset](https://nixtlaverse.nixtla.io/datasetsforecast/m5.html). The M5
+dataset contains daily product demand (sales) for 10 retail stores in
+the US.
+
+First, we load the data using `datasetsforecast`. This returns:
+
+- `Y_df`, containing the sales (`y` column), for each unique product
+ (`unique_id` column) at every timestamp (`ds` column).
+- `X_df`, containing additional relevant information for each unique
+ product (`unique_id` column) at every timestamp (`ds` column).
+
+```python
+Y_df, X_df, _ = M5.load(directory=os.getcwd())
+Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+X_df['ds'] = pd.to_datetime(X_df['ds'])
+Y_df.head(10)
+```
+
+| | unique_id | ds | y |
+|-----|------------------|------------|-----|
+| 0 | FOODS_1_001_CA_1 | 2011-01-29 | 3.0 |
+| 1 | FOODS_1_001_CA_1 | 2011-01-30 | 0.0 |
+| 2 | FOODS_1_001_CA_1 | 2011-01-31 | 0.0 |
+| 3 | FOODS_1_001_CA_1 | 2011-02-01 | 1.0 |
+| 4 | FOODS_1_001_CA_1 | 2011-02-02 | 4.0 |
+| 5 | FOODS_1_001_CA_1 | 2011-02-03 | 2.0 |
+| 6 | FOODS_1_001_CA_1 | 2011-02-04 | 0.0 |
+| 7 | FOODS_1_001_CA_1 | 2011-02-05 | 2.0 |
+| 8 | FOODS_1_001_CA_1 | 2011-02-06 | 0.0 |
+| 9 | FOODS_1_001_CA_1 | 2011-02-07 | 0.0 |
+
+For this example, we will only keep the additional relevant information
+from the column `event_type_1`. This column is a *categorical variable*
+that indicates whether an important event that might affect the sales of
+the product takes place at a certain date.
+
+```python
+X_df = X_df[['unique_id', 'ds', 'event_type_1']]
+
+X_df.head(10)
+```
+
+| | unique_id | ds | event_type_1 |
+|-----|------------------|------------|--------------|
+| 0 | FOODS_1_001_CA_1 | 2011-01-29 | nan |
+| 1 | FOODS_1_001_CA_1 | 2011-01-30 | nan |
+| 2 | FOODS_1_001_CA_1 | 2011-01-31 | nan |
+| 3 | FOODS_1_001_CA_1 | 2011-02-01 | nan |
+| 4 | FOODS_1_001_CA_1 | 2011-02-02 | nan |
+| 5 | FOODS_1_001_CA_1 | 2011-02-03 | nan |
+| 6 | FOODS_1_001_CA_1 | 2011-02-04 | nan |
+| 7 | FOODS_1_001_CA_1 | 2011-02-05 | nan |
+| 8 | FOODS_1_001_CA_1 | 2011-02-06 | Sporting |
+| 9 | FOODS_1_001_CA_1 | 2011-02-07 | nan |
+
+As you can see, on February 6th 2011, there is a Sporting event.
+
+## 3. Forecasting product demand using categorical variables
+
+We will forecast the demand for a single product only. We choose a high
+selling food product identified by `FOODS_3_090_CA_3`.
+
+```python
+product = 'FOODS_3_090_CA_3'
+Y_df_product = Y_df.query('unique_id == @product')
+X_df_product = X_df.query('unique_id == @product')
+```
+
+We merge our two dataframes to create the dataset to be used in TimeGPT.
+
+```python
+df = Y_df_product.merge(X_df_product)
+
+df.head(10)
+```
+
+| | unique_id | ds | y | event_type_1 |
+|-----|------------------|------------|-------|--------------|
+| 0 | FOODS_3_090_CA_3 | 2011-01-29 | 108.0 | nan |
+| 1 | FOODS_3_090_CA_3 | 2011-01-30 | 132.0 | nan |
+| 2 | FOODS_3_090_CA_3 | 2011-01-31 | 102.0 | nan |
+| 3 | FOODS_3_090_CA_3 | 2011-02-01 | 120.0 | nan |
+| 4 | FOODS_3_090_CA_3 | 2011-02-02 | 106.0 | nan |
+| 5 | FOODS_3_090_CA_3 | 2011-02-03 | 123.0 | nan |
+| 6 | FOODS_3_090_CA_3 | 2011-02-04 | 279.0 | nan |
+| 7 | FOODS_3_090_CA_3 | 2011-02-05 | 175.0 | nan |
+| 8 | FOODS_3_090_CA_3 | 2011-02-06 | 186.0 | Sporting |
+| 9 | FOODS_3_090_CA_3 | 2011-02-07 | 120.0 | nan |
+
+In order to use *categorical variables* with TimeGPT, it is necessary to
+numerically encode the variables. We will use *one-hot encoding* in this
+tutorial.
+
+We can one-hot encode the `event_type_1` column by using pandas built-in
+`get_dummies` functionality. After one-hot encoding the `event_type_1`
+variable, we can add it to the dataframe and remove the original column.
+
+```python
+event_type_1_ohe = pd.get_dummies(df['event_type_1'], dtype=int)
+df = pd.concat([df, event_type_1_ohe], axis=1)
+df = df.drop(columns = 'event_type_1')
+
+df.tail(10)
+```
+
+| | unique_id | ds | y | Cultural | National | Religious | Sporting | nan |
+|------|------------------|------------|-------|----------|----------|-----------|----------|-----|
+| 1959 | FOODS_3_090_CA_3 | 2016-06-10 | 140.0 | 0 | 0 | 0 | 0 | 1 |
+| 1960 | FOODS_3_090_CA_3 | 2016-06-11 | 151.0 | 0 | 0 | 0 | 0 | 1 |
+| 1961 | FOODS_3_090_CA_3 | 2016-06-12 | 87.0 | 0 | 0 | 0 | 0 | 1 |
+| 1962 | FOODS_3_090_CA_3 | 2016-06-13 | 67.0 | 0 | 0 | 0 | 0 | 1 |
+| 1963 | FOODS_3_090_CA_3 | 2016-06-14 | 50.0 | 0 | 0 | 0 | 0 | 1 |
+| 1964 | FOODS_3_090_CA_3 | 2016-06-15 | 58.0 | 0 | 0 | 0 | 0 | 1 |
+| 1965 | FOODS_3_090_CA_3 | 2016-06-16 | 116.0 | 0 | 0 | 0 | 0 | 1 |
+| 1966 | FOODS_3_090_CA_3 | 2016-06-17 | 124.0 | 0 | 0 | 0 | 0 | 1 |
+| 1967 | FOODS_3_090_CA_3 | 2016-06-18 | 167.0 | 0 | 0 | 0 | 0 | 1 |
+| 1968 | FOODS_3_090_CA_3 | 2016-06-19 | 118.0 | 0 | 0 | 0 | 1 | 0 |
+
+As you can see, we have now added 5 columns, each with a binary
+indicator (`1` or `0`) whether there is a `Cultural`, `National`,
+`Religious`, `Sporting` or no (`nan`) event on that particular day. For
+example, on June 19th 2016, there is a `Sporting` event.
+
+Let’s turn to our forecasting task. We will forecast the first 7 days of
+February 2016. This includes 7 February 2016 - the date on which [Super
+Bowl 50](https://en.wikipedia.org/wiki/Super_Bowl_50) was held. Such
+large, national events typically impact retail product sales.
+
+To use the encoded categorical variables in TimeGPT, we have to add them
+as future values. Therefore, we create a future values dataframe, that
+contains the `unique_id`, the timestamp `ds`, and the encoded
+categorical variables.
+
+Of course, we drop the target column as this is normally not available -
+this is the quantity that we seek to forecast!
+
+```python
+future_ex_vars_df = df.drop(columns = ['y'])
+future_ex_vars_df = future_ex_vars_df.query("ds >= '2016-02-01' & ds <= '2016-02-07'")
+
+future_ex_vars_df.head(10)
+```
+
+| | unique_id | ds | Cultural | National | Religious | Sporting | nan |
+|------|------------------|------------|----------|----------|-----------|----------|-----|
+| 1829 | FOODS_3_090_CA_3 | 2016-02-01 | 0 | 0 | 0 | 0 | 1 |
+| 1830 | FOODS_3_090_CA_3 | 2016-02-02 | 0 | 0 | 0 | 0 | 1 |
+| 1831 | FOODS_3_090_CA_3 | 2016-02-03 | 0 | 0 | 0 | 0 | 1 |
+| 1832 | FOODS_3_090_CA_3 | 2016-02-04 | 0 | 0 | 0 | 0 | 1 |
+| 1833 | FOODS_3_090_CA_3 | 2016-02-05 | 0 | 0 | 0 | 0 | 1 |
+| 1834 | FOODS_3_090_CA_3 | 2016-02-06 | 0 | 0 | 0 | 0 | 1 |
+| 1835 | FOODS_3_090_CA_3 | 2016-02-07 | 0 | 0 | 0 | 1 | 0 |
+
+Next, we limit our input dataframe to all but the 7 forecast days:
+
+```python
+df_train = df.query("ds < '2016-02-01'")
+
+df_train.tail(10)
+```
+
+| | unique_id | ds | y | Cultural | National | Religious | Sporting | nan |
+|------|------------------|------------|-------|----------|----------|-----------|----------|-----|
+| 1819 | FOODS_3_090_CA_3 | 2016-01-22 | 94.0 | 0 | 0 | 0 | 0 | 1 |
+| 1820 | FOODS_3_090_CA_3 | 2016-01-23 | 144.0 | 0 | 0 | 0 | 0 | 1 |
+| 1821 | FOODS_3_090_CA_3 | 2016-01-24 | 146.0 | 0 | 0 | 0 | 0 | 1 |
+| 1822 | FOODS_3_090_CA_3 | 2016-01-25 | 87.0 | 0 | 0 | 0 | 0 | 1 |
+| 1823 | FOODS_3_090_CA_3 | 2016-01-26 | 73.0 | 0 | 0 | 0 | 0 | 1 |
+| 1824 | FOODS_3_090_CA_3 | 2016-01-27 | 62.0 | 0 | 0 | 0 | 0 | 1 |
+| 1825 | FOODS_3_090_CA_3 | 2016-01-28 | 64.0 | 0 | 0 | 0 | 0 | 1 |
+| 1826 | FOODS_3_090_CA_3 | 2016-01-29 | 102.0 | 0 | 0 | 0 | 0 | 1 |
+| 1827 | FOODS_3_090_CA_3 | 2016-01-30 | 113.0 | 0 | 0 | 0 | 0 | 1 |
+| 1828 | FOODS_3_090_CA_3 | 2016-01-31 | 98.0 | 0 | 0 | 0 | 0 | 1 |
+
+Let’s call the `forecast` method, first *without* the categorical
+variables.
+
+```python
+timegpt_fcst_without_cat_vars_df = nixtla_client.forecast(df=df_train, h=7, level=[80, 90])
+timegpt_fcst_without_cat_vars_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: D
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| | unique_id | ds | TimeGPT | TimeGPT-lo-90 | TimeGPT-lo-80 | TimeGPT-hi-80 | TimeGPT-hi-90 |
+|----|----|----|----|----|----|----|----|
+| 0 | FOODS_3_090_CA_3 | 2016-02-01 | 73.304092 | 53.449049 | 54.795078 | 91.813107 | 93.159136 |
+| 1 | FOODS_3_090_CA_3 | 2016-02-02 | 66.335518 | 47.510669 | 50.274136 | 82.396899 | 85.160367 |
+| 2 | FOODS_3_090_CA_3 | 2016-02-03 | 65.881630 | 36.218617 | 41.388896 | 90.374364 | 95.544643 |
+| 3 | FOODS_3_090_CA_3 | 2016-02-04 | 72.371864 | -26.683115 | 25.097362 | 119.646367 | 171.426844 |
+| 4 | FOODS_3_090_CA_3 | 2016-02-05 | 95.141045 | -2.084882 | 34.027078 | 156.255011 | 192.366971 |
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+We plot the forecast and the last 28 days before the forecast period:
+
+```python
+nixtla_client.plot(
+ df[['unique_id', 'ds', 'y']].query("ds <= '2016-02-07'"),
+ timegpt_fcst_without_cat_vars_df,
+ max_insample_length=28,
+)
+```
+
+
+
+TimeGPT already provides a reasonable forecast, but it seems to somewhat
+underforecast the peak on the 6th of February 2016 - the day before the
+Super Bowl.
+
+Let’s call the `forecast` method again, now *with* the categorical
+variables.
+
+```python
+timegpt_fcst_with_cat_vars_df = nixtla_client.forecast(df=df_train, X_df=future_ex_vars_df, h=7, level=[80, 90])
+timegpt_fcst_with_cat_vars_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: D
+INFO:nixtla.nixtla_client:Using the following exogenous variables: Cultural, National, Religious, Sporting, nan
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| | unique_id | ds | TimeGPT | TimeGPT-lo-90 | TimeGPT-lo-80 | TimeGPT-hi-80 | TimeGPT-hi-90 |
+|----|----|----|----|----|----|----|----|
+| 0 | FOODS_3_090_CA_3 | 2016-02-01 | 70.661271 | -0.204378 | 14.593348 | 126.729194 | 141.526919 |
+| 1 | FOODS_3_090_CA_3 | 2016-02-02 | 65.566941 | -20.394326 | 11.654239 | 119.479643 | 151.528208 |
+| 2 | FOODS_3_090_CA_3 | 2016-02-03 | 68.510010 | -33.713710 | 6.732952 | 130.287069 | 170.733731 |
+| 3 | FOODS_3_090_CA_3 | 2016-02-04 | 75.417710 | -40.974649 | 4.751767 | 146.083653 | 191.810069 |
+| 4 | FOODS_3_090_CA_3 | 2016-02-05 | 97.340302 | -57.385361 | 18.253812 | 176.426792 | 252.065965 |
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+We plot the forecast and the last 28 days before the forecast period:
+
+```python
+nixtla_client.plot(
+ df[['unique_id', 'ds', 'y']].query("ds <= '2016-02-07'"),
+ timegpt_fcst_with_cat_vars_df,
+ max_insample_length=28,
+)
+```
+
+
+
+We can visually verify that the forecast is closer to the actual
+observed value, which is the result of including the categorical
+variable in our forecast.
+
+Let’s verify this conclusion by computing the [Mean Absolute
+Error](https://en.wikipedia.org/wiki/Mean_absolute_error) on the
+forecasts we created.
+
+```python
+from utilsforecast.losses import mae
+```
+
+
+```python
+# Create target dataframe
+df_target = df[['unique_id', 'ds', 'y']].query("ds >= '2016-02-01' & ds <= '2016-02-07'")
+
+# Rename forecast columns
+timegpt_fcst_without_cat_vars_df = timegpt_fcst_without_cat_vars_df.rename(columns={'TimeGPT': 'TimeGPT-without-cat-vars'})
+timegpt_fcst_with_cat_vars_df = timegpt_fcst_with_cat_vars_df.rename(columns={'TimeGPT': 'TimeGPT-with-cat-vars'})
+
+# Merge forecasts with target dataframe
+df_target = df_target.merge(timegpt_fcst_without_cat_vars_df[['unique_id', 'ds', 'TimeGPT-without-cat-vars']])
+df_target = df_target.merge(timegpt_fcst_with_cat_vars_df[['unique_id', 'ds', 'TimeGPT-with-cat-vars']])
+
+# Compute errors
+mean_absolute_errors = mae(df_target, ['TimeGPT-without-cat-vars', 'TimeGPT-with-cat-vars'])
+```
+
+
+```python
+mean_absolute_errors
+```
+
+| | unique_id | TimeGPT-without-cat-vars | TimeGPT-with-cat-vars |
+|-----|------------------|--------------------------|-----------------------|
+| 0 | FOODS_3_090_CA_3 | 24.285649 | 20.028514 |
+
+Indeed, we find that the error when using TimeGPT with the categorical
+variable is approx. 20% lower than when using TimeGPT without the
+categorical variables, indicating better performance when we include the
+categorical variable.
+
diff --git a/nixtla/docs/tutorials/computing_at_scale.html.mdx b/nixtla/docs/tutorials/computing_at_scale.html.mdx
new file mode 100644
index 00000000..8ca34e87
--- /dev/null
+++ b/nixtla/docs/tutorials/computing_at_scale.html.mdx
@@ -0,0 +1,91 @@
+---
+output-file: computing_at_scale.html
+title: Computing at scale
+---
+
+
+Handling large datasets is a common challenge in time series
+forecasting. For example, when working with retail data, you may have to
+forecast sales for thousands of products across hundreds of stores.
+Similarly, when dealing with electricity consumption data, you may need
+to predict consumption for thousands of households across various
+regions.
+
+Nixtla’s `TimeGPT` enables you to use several distributed computing
+frameworks to manage large datasets efficiently. `TimeGPT` currently
+supports `Spark`, `Dask`, and `Ray` through `Fugue`.
+
+In this notebook, we will explain how to leverage these frameworks using
+`TimeGPT`.
+
+**Outline:**
+
+1. [Getting Started](#1-getting-started)
+
+2. [Forecasting at Scale](#2-forecasting-at-scale)
+
+3. [Important Considerations](#3-important-considerations)
+
+## Getting started
+
+To use `TimeGPT` with any of the supported distributed computing
+frameworks, you first need an API Key, just as you would when not using
+any distributed computing.
+
+Upon [registration](https://dashboard.nixtla.io/), you will receive an
+email asking you to confirm your signup. After confirming, you will
+receive access to your dashboard. There, under`API Keys`, you will find
+your API Key. Next, you need to integrate your API Key into your
+development workflow with the Nixtla SDK. For guidance on how to do
+this, please refer to the [Setting Up Your Authentication Key
+tutorial](https://docs.nixtla.io/docs/getting-started-setting_up_your_api_key).
+
+## Forecasting at Scale
+
+Using `TimeGPT` with any of the supported distributed computing
+frameworks is straightforward and its usage is almost identical to the
+non-distributed case.
+
+1. Instantiate a
+ [`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+ class.
+2. Load your data as a `pandas` DataFrame.
+3. Initialize the distributed computing framework.
+ - [Spark](https://docs.nixtla.io/docs/tutorials-spark)
+ - [Dask](https://docs.nixtla.io/docs/tutorials-dask)
+ - [Ray](https://docs.nixtla.io/docs/tutorials-ray)
+4. Use any of the
+ [`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+ class methods.
+5. Stop the distributed computing framework, if necessary.
+
+These are the general steps that you will need to follow to use
+`TimeGPT` with any of the supported distributed computing frameworks.
+For a detailed explanation and a complete example, please refer to the
+guide for the specific framework linked above.
+
+> **Important**
+>
+> Parallelization in these frameworks is done along the various time
+> series within your dataset. Therefore, it is essential that your
+> dataset includes multiple time series, each with a unique id.
+
+## Important Considerations
+
+### When to Use a Distributed Computing Framework
+
+Consider using a distributed computing framework if your dataset:
+
+- Consists of millions of observations over multiple time series.
+- Is too large to fit into the memory of a single machine.
+- Would be too slow to process on a single machine.
+
+### Choosing the Right Framework
+
+When selecting a distributed computing framework, take into account your
+existing infrastructure and the skill set of your team. Although
+`TimeGPT` can be used with any of the supported frameworks with minimal
+code changes, choosing the right one should align with your specific
+needs and resources. This will ensure that you leverage the full
+potential of `TimeGPT` while handling large datasets efficiently.
+
diff --git a/nixtla/docs/tutorials/computing_at_scale_dask_distributed.html.mdx b/nixtla/docs/tutorials/computing_at_scale_dask_distributed.html.mdx
new file mode 100644
index 00000000..8cd610f3
--- /dev/null
+++ b/nixtla/docs/tutorials/computing_at_scale_dask_distributed.html.mdx
@@ -0,0 +1,170 @@
+---
+description: Run TimeGPT distributedly on top of Dask
+output-file: computing_at_scale_dask_distributed.html
+title: Dask
+---
+
+
+[Dask](https://www.dask.org/get-started) is an open source parallel
+computing library for Python. In this guide, we will explain how to use
+`TimeGPT` on top of Dask.
+
+**Outline:**
+
+1. [Installation](#installation)
+
+2. [Load Your Data](#load-your-data)
+
+3. [Import Dask](#import-dask)
+
+4. [Use TimeGPT on Dask](#use-timegpt-on-dask)
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/17_computing_at_scale_dask_distributed.ipynb)
+
+## 1. Installation
+
+Install Dask through [Fugue](https://fugue-tutorials.readthedocs.io/).
+Fugue provides an easy-to-use interface for distributed computing that
+lets users execute Python code on top of several distributed computing
+frameworks, including Dask.
+
+> **Note**
+>
+> You can install `fugue` with `pip`:
+>
+> ```shell
+> pip install fugue[dask]
+> ```
+
+If executing on a distributed `Dask` cluster, ensure that the `nixtla`
+library is installed across all the workers.
+
+## 2. Load Data
+
+You can load your data as a `pandas` DataFrame. In this tutorial, we
+will use a dataset that contains hourly electricity prices from
+different markets.
+
+```python
+import pandas as pd
+```
+
+
+```python
+df = pd.read_csv(
+ 'https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short.csv',
+ parse_dates=['ds'],
+)
+df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|---------------------|-------|
+| 0 | BE | 2016-10-22 00:00:00 | 70.00 |
+| 1 | BE | 2016-10-22 01:00:00 | 37.10 |
+| 2 | BE | 2016-10-22 02:00:00 | 37.10 |
+| 3 | BE | 2016-10-22 03:00:00 | 44.75 |
+| 4 | BE | 2016-10-22 04:00:00 | 37.10 |
+
+## 3. Import Dask
+
+Import Dask and convert the `pandas` DataFrame to a Dask DataFrame.
+
+```python
+import dask.dataframe as dd
+```
+
+
+```python
+dask_df = dd.from_pandas(df, npartitions=2)
+dask_df
+```
+
+| | unique_id | ds | y |
+|---------------|-----------|--------|---------|
+| npartitions=2 | | | |
+| 0 | string | string | float64 |
+| 4200 | ... | ... | ... |
+| 8399 | ... | ... | ... |
+
+## 4. Use TimeGPT on Dask
+
+Using `TimeGPT` on top of `Dask` is almost identical to the
+non-distributed case. The only difference is that you need to use a
+`Dask` DataFrame, which we already defined in the previous step.
+
+First, instantiate the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class.
+
+```python
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, set the `base_url` argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+Then use any method from the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class such as
+[`forecast`](https://docs.nixtla.io/docs/reference-sdk_reference#nixtlaclientforecast)
+or
+[`cross_validation`](https://docs.nixtla.io/docs/reference-sdk_reference#nixtlaclientcross_validation).
+
+```python
+fcst_df = nixtla_client.forecast(dask_df, h=12)
+fcst_df.compute().head()
+```
+
+| | unique_id | ds | TimeGPT |
+|-----|-----------|---------------------|-----------|
+| 0 | BE | 2016-12-31 00:00:00 | 45.190453 |
+| 1 | BE | 2016-12-31 01:00:00 | 43.244446 |
+| 2 | BE | 2016-12-31 02:00:00 | 41.958389 |
+| 3 | BE | 2016-12-31 03:00:00 | 39.796486 |
+| 4 | BE | 2016-12-31 04:00:00 | 39.204533 |
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+```python
+cv_df = nixtla_client.cross_validation(dask_df, h=12, n_windows=5, step_size=2)
+cv_df.compute().head()
+```
+
+| | unique_id | ds | cutoff | TimeGPT |
+|-----|-----------|---------------------|---------------------|-----------|
+| 0 | BE | 2016-12-30 04:00:00 | 2016-12-30 03:00:00 | 39.375439 |
+| 1 | BE | 2016-12-30 05:00:00 | 2016-12-30 03:00:00 | 40.039215 |
+| 2 | BE | 2016-12-30 06:00:00 | 2016-12-30 03:00:00 | 43.455849 |
+| 3 | BE | 2016-12-30 07:00:00 | 2016-12-30 03:00:00 | 47.716408 |
+| 4 | BE | 2016-12-30 08:00:00 | 2016-12-30 03:00:00 | 50.31665 |
+
+You can also use exogenous variables with `TimeGPT` on top of `Dask`. To
+do this, please refer to the [Exogenous
+Variables](https://docs.nixtla.io/docs/tutorials-exogenous_variables)
+tutorial. Just keep in mind that instead of using a pandas DataFrame,
+you need to use a `Dask` DataFrame instead.
+
diff --git a/nixtla/docs/tutorials/computing_at_scale_ray_distributed.html.mdx b/nixtla/docs/tutorials/computing_at_scale_ray_distributed.html.mdx
new file mode 100644
index 00000000..a8e84886
--- /dev/null
+++ b/nixtla/docs/tutorials/computing_at_scale_ray_distributed.html.mdx
@@ -0,0 +1,213 @@
+---
+description: Run TimeGPT distributedly on top of Ray
+output-file: computing_at_scale_ray_distributed.html
+title: Ray
+---
+
+
+[Ray](https://www.ray.io/) is an open source unified compute framework
+to scale Python workloads. In this guide, we will explain how to use
+`TimeGPT` on top of Ray.
+
+**Outline:**
+
+1. [Installation](#installation)
+
+2. [Load Your Data](#load-your-data)
+
+3. [Initialize Ray](#initialize-ray)
+
+4. [Use TimeGPT on Ray](#use-timegpt-on-ray)
+
+5. [Shutdown Ray](#shutdown-ray)
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/19_computing_at_scale_ray_distributed.ipynb)
+
+## 1. Installation
+
+Install Ray through [Fugue](https://fugue-tutorials.readthedocs.io/).
+Fugue provides an easy-to-use interface for distributed computing that
+lets users execute Python code on top of several distributed computing
+frameworks, including Ray.
+
+> **Note**
+>
+> You can install `fugue` with `pip`:
+>
+> ```shell
+> pip install fugue[ray]
+> ```
+
+If executing on a distributed `Ray` cluster, ensure that the `nixtla`
+library is installed across all the workers.
+
+## 2. Load Data
+
+You can load your data as a `pandas` DataFrame. In this tutorial, we
+will use a dataset that contains hourly electricity prices from
+different markets.
+
+```python
+import pandas as pd
+```
+
+
+```python
+df = pd.read_csv(
+ 'https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short.csv',
+ parse_dates=['ds'],
+)
+df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|---------------------|-------|
+| 0 | BE | 2016-10-22 00:00:00 | 70.00 |
+| 1 | BE | 2016-10-22 01:00:00 | 37.10 |
+| 2 | BE | 2016-10-22 02:00:00 | 37.10 |
+| 3 | BE | 2016-10-22 03:00:00 | 44.75 |
+| 4 | BE | 2016-10-22 04:00:00 | 37.10 |
+
+## 3. Initialize Ray
+
+Initialize `Ray` and convert the pandas DataFrame to a `Ray` DataFrame.
+
+```python
+import ray
+from ray.cluster_utils import Cluster
+```
+
+
+```python
+ray_cluster = Cluster(
+ initialize_head=True,
+ head_node_args={"num_cpus": 2}
+)
+ray.init(address=ray_cluster.address, ignore_reinit_error=True)
+```
+
+
+```python
+ray_df = ray.data.from_pandas(df)
+ray_df
+```
+
+``` text
+MaterializedDataset(
+ num_blocks=1,
+ num_rows=6720,
+ schema={unique_id: object, ds: datetime64[ns], y: float64}
+)
+```
+
+## 4. Use TimeGPT on Ray
+
+Using `TimeGPT` on top of `Ray` is almost identical to the
+non-distributed case. The only difference is that you need to use a
+`Ray` DataFrame.
+
+First, instantiate the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class.
+
+```python
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, set the `base_url` argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+Then use any method from the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class such as
+[`forecast`](https://docs.nixtla.io/docs/reference-sdk_reference#nixtlaclientforecast)
+or
+[`cross_validation`](https://docs.nixtla.io/docs/reference-sdk_reference#nixtlaclientcross_validation).
+
+```python
+ray_df
+```
+
+``` text
+MaterializedDataset(
+ num_blocks=1,
+ num_rows=6720,
+ schema={unique_id: object, ds: datetime64[ns], y: float64}
+)
+```
+
+```python
+fcst_df = nixtla_client.forecast(ray_df, h=12)
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+To visualize the result, use the `to_pandas` method to convert the
+output of `Ray` to a `pandas` DataFrame.
+
+```python
+fcst_df.to_pandas().tail()
+```
+
+| | unique_id | ds | TimeGPT |
+|-----|-----------|---------------------|-----------|
+| 55 | NP | 2018-12-24 07:00:00 | 55.387066 |
+| 56 | NP | 2018-12-24 08:00:00 | 56.115517 |
+| 57 | NP | 2018-12-24 09:00:00 | 56.090714 |
+| 58 | NP | 2018-12-24 10:00:00 | 55.813717 |
+| 59 | NP | 2018-12-24 11:00:00 | 55.528519 |
+
+```python
+cv_df = nixtla_client.cross_validation(ray_df, h=12, freq='H', n_windows=5, step_size=2)
+```
+
+
+```python
+cv_df.to_pandas().tail()
+```
+
+| | unique_id | ds | cutoff | TimeGPT |
+|-----|-----------|---------------------|---------------------|-----------|
+| 295 | NP | 2018-12-23 19:00:00 | 2018-12-23 11:00:00 | 53.632019 |
+| 296 | NP | 2018-12-23 20:00:00 | 2018-12-23 11:00:00 | 52.512775 |
+| 297 | NP | 2018-12-23 21:00:00 | 2018-12-23 11:00:00 | 51.894035 |
+| 298 | NP | 2018-12-23 22:00:00 | 2018-12-23 11:00:00 | 51.06572 |
+| 299 | NP | 2018-12-23 23:00:00 | 2018-12-23 11:00:00 | 50.32592 |
+
+You can also use exogenous variables with `TimeGPT` on top of `Ray`. To
+do this, please refer to the [Exogenous
+Variables](https://docs.nixtla.io/docs/tutorials-exogenous_variables)
+tutorial. Just keep in mind that instead of using a pandas DataFrame,
+you need to use a `Ray` DataFrame instead.
+
+## 5. Shutdown Ray
+
+When you are done, shutdown the `Ray` session.
+
+```python
+ray.shutdown()
+```
+
diff --git a/nixtla/docs/tutorials/computing_at_scale_spark_distributed.html.mdx b/nixtla/docs/tutorials/computing_at_scale_spark_distributed.html.mdx
new file mode 100644
index 00000000..d84dcfbe
--- /dev/null
+++ b/nixtla/docs/tutorials/computing_at_scale_spark_distributed.html.mdx
@@ -0,0 +1,163 @@
+---
+description: Run TimeGPT distributedly on top of Spark
+output-file: computing_at_scale_spark_distributed.html
+title: Spark
+---
+
+
+[Spark](https://spark.apache.org/) is an open-source distributed
+computing framework designed for large-scale data processing. In this
+guide, we will explain how to use `TimeGPT` on top of Spark.
+
+**Outline:**
+
+1. [Installation](#installation)
+
+2. [Load Your Data](#load-your-data)
+
+3. [Initialize Spark](#initialize-spark)
+
+4. [Use TimeGPT on Spark](#use-timegpt-on-spark)
+
+5. [Stop Spark](#stop-spark)
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/16_computing_at_scale_spark_distributed.ipynb)
+
+## 1. Installation
+
+Install Spark through [Fugue](https://fugue-tutorials.readthedocs.io/).
+Fugue provides an easy-to-use interface for distributed computing that
+lets users execute Python code on top of several distributed computing
+frameworks, including Spark.
+
+> **Note**
+>
+> You can install `fugue` with `pip`:
+>
+> ```shell
+> pip install fugue[spark]
+> ```
+
+If executing on a distributed `Spark` cluster, ensure that the `nixtla`
+library is installed across all the workers.
+
+## 2. Load Data
+
+You can load your data as a `pandas` DataFrame. In this tutorial, we
+will use a dataset that contains hourly electricity prices from
+different markets.
+
+```python
+import pandas as pd
+```
+
+
+```python
+df = pd.read_csv(
+ 'https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short.csv',
+ parse_dates=['ds'],
+)
+df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|---------------------|-------|
+| 0 | BE | 2016-10-22 00:00:00 | 70.00 |
+| 1 | BE | 2016-10-22 01:00:00 | 37.10 |
+| 2 | BE | 2016-10-22 02:00:00 | 37.10 |
+| 3 | BE | 2016-10-22 03:00:00 | 44.75 |
+| 4 | BE | 2016-10-22 04:00:00 | 37.10 |
+
+## 3. Initialize Spark
+
+Initialize `Spark` and convert the pandas DataFrame to a `Spark`
+DataFrame.
+
+```python
+from pyspark.sql import SparkSession
+```
+
+
+```python
+spark = SparkSession.builder.getOrCreate()
+```
+
+
+```python
+spark_df = spark.createDataFrame(df)
+spark_df.show(5)
+```
+
+## 4. Use TimeGPT on Spark
+
+Using `TimeGPT` on top of `Spark` is almost identical to the
+non-distributed case. The only difference is that you need to use a
+`Spark` DataFrame.
+
+First, instantiate the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class.
+
+```python
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, set the `base_url` argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+Then use any method from the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class such as
+[`forecast`](https://docs.nixtla.io/docs/reference-sdk_reference#nixtlaclientforecast)
+or
+[`cross_validation`](https://docs.nixtla.io/docs/reference-sdk_reference#nixtlaclientcross_validation).
+
+```python
+fcst_df = nixtla_client.forecast(spark_df, h=12)
+fcst_df.show(5)
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+```python
+cv_df = nixtla_client.cross_validation(spark_df, h=12, n_windows=5, step_size=2)
+cv_df.show(5)
+```
+
+You can also use exogenous variables with `TimeGPT` on top of `Spark`.
+To do this, please refer to the [Exogenous
+Variables](https://docs.nixtla.io/docs/tutorials-exogenous_variables)
+tutorial. Just keep in mind that instead of using a pandas DataFrame,
+you need to use a `Spark` DataFrame instead.
+
+## 5. Stop Spark
+
+When you are done, stop the `Spark` session.
+
+```python
+spark.stop()
+```
+
diff --git a/nixtla/docs/tutorials/cross_validation.html.mdx b/nixtla/docs/tutorials/cross_validation.html.mdx
new file mode 100644
index 00000000..8341ac3d
--- /dev/null
+++ b/nixtla/docs/tutorials/cross_validation.html.mdx
@@ -0,0 +1,402 @@
+---
+output-file: cross_validation.html
+title: Cross-validation
+---
+
+
+One of the primary challenges in time series forecasting is the inherent
+uncertainty and variability over time, making it crucial to validate the
+accuracy and reliability of the models employed. Cross-validation, a
+robust model validation technique, is particularly adapted for this
+task, as it provides insights into the expected performance of a model
+on unseen data, ensuring the forecasts are reliable and resilient before
+being deployed in real-world scenarios.
+
+`TimeGPT`, understanding the intricate needs of time series forecasting,
+incorporates the `cross_validation` method, designed to streamline the
+validation process for time series models. This functionality enables
+practitioners to rigorously test their forecasting models against
+historical data, assessing their effectiveness while tuning them for
+optimal performance. This tutorial will guide you through the nuanced
+process of conducting cross-validation within the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class, ensuring your time series forecasting models are not just
+well-constructed, but also validated for trustworthiness and precision.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/08_cross_validation.ipynb)
+
+## 1. Import packages
+
+First, we install and import the required packages and initialize the
+Nixtla client.
+
+We start off by initializing an instance of
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient).
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+
+from IPython.display import display
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 2. Load data
+
+Let’s see an example, using the Peyton Manning dataset.
+
+```python
+pm_df = pd.read_csv('https://datasets-nixtla.s3.amazonaws.com/peyton-manning.csv')
+```
+
+## 3. Cross-validation
+
+The `cross_validation` method within the `TimeGPT` class is an advanced
+functionality crafted to perform systematic validation on time series
+forecasting models. This method necessitates a dataframe comprising
+time-ordered data and employs a rolling-window scheme to meticulously
+evaluate the model’s performance across different time periods, thereby
+ensuring the model’s reliability and stability over time. The animation
+below shows how TimeGPT performs cross-validation.
+
+
+
+Key parameters include `freq`, which denotes the data’s frequency and is
+automatically inferred if not specified. The `id_col`, `time_col`, and
+`target_col` parameters designate the respective columns for each
+series’ identifier, time step, and target values. The method offers
+customization through parameters like `n_windows`, indicating the number
+of separate time windows on which the model is assessed, and
+`step_size`, determining the gap between these windows. If `step_size`
+is unspecified, it defaults to the forecast horizon `h`.
+
+The process also allows for model refinement via `finetune_steps`,
+specifying the number of iterations for model fine-tuning on new data.
+Data pre-processing is manageable through `clean_ex_first`, deciding
+whether to cleanse the exogenous signal prior to forecasting.
+Additionally, the method supports enhanced feature engineering from time
+data through the `date_features` parameter, which can automatically
+generate crucial date-related features or accept custom functions for
+bespoke feature creation. The `date_features_to_one_hot` parameter
+further enables the transformation of categorical date features into a
+format suitable for machine learning models.
+
+In execution, `cross_validation` assesses the model’s forecasting
+accuracy in each window, providing a robust view of the model’s
+performance variability over time and potential overfitting. This
+detailed evaluation ensures the forecasts generated are not only
+accurate but also consistent across diverse temporal contexts.
+
+```python
+timegpt_cv_df = nixtla_client.cross_validation(
+ pm_df,
+ h=7,
+ n_windows=5,
+ freq='D',
+)
+timegpt_cv_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Querying model metadata...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Cross Validation Endpoint...
+```
+
+| | unique_id | ds | cutoff | y | TimeGPT |
+|-----|-----------|------------|------------|----------|----------|
+| 0 | 0 | 2015-12-17 | 2015-12-16 | 7.591862 | 7.939553 |
+| 1 | 0 | 2015-12-18 | 2015-12-16 | 7.528869 | 7.887512 |
+| 2 | 0 | 2015-12-19 | 2015-12-16 | 7.171657 | 7.766617 |
+| 3 | 0 | 2015-12-20 | 2015-12-16 | 7.891331 | 7.931502 |
+| 4 | 0 | 2015-12-21 | 2015-12-16 | 8.360071 | 8.312632 |
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.cross_validation(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+```python
+cutoffs = timegpt_cv_df['cutoff'].unique()
+for cutoff in cutoffs:
+ fig = nixtla_client.plot(
+ pm_df.tail(100),
+ timegpt_cv_df.query('cutoff == @cutoff').drop(columns=['cutoff', 'y']),
+ )
+ display(fig)
+```
+
+
+
+
+
+
+
+
+
+
+
+## 4. Cross-validation with prediction intervals
+
+It is also possible to generate prediction intervals during
+cross-validation. To do so, we simply use the `level` argument.
+
+```python
+timegpt_cv_df = nixtla_client.cross_validation(
+ pm_df,
+ h=7,
+ n_windows=5,
+ freq='D',
+ level=[80, 90],
+)
+timegpt_cv_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Cross Validation Endpoint...
+```
+
+| | unique_id | ds | cutoff | y | TimeGPT | TimeGPT-hi-80 | TimeGPT-hi-90 | TimeGPT-lo-80 | TimeGPT-lo-90 |
+|----|----|----|----|----|----|----|----|----|----|
+| 0 | 0 | 2015-12-17 | 2015-12-16 | 7.591862 | 7.939553 | 8.201465 | 8.314956 | 7.677642 | 7.564151 |
+| 1 | 0 | 2015-12-18 | 2015-12-16 | 7.528869 | 7.887512 | 8.175414 | 8.207470 | 7.599609 | 7.567553 |
+| 2 | 0 | 2015-12-19 | 2015-12-16 | 7.171657 | 7.766617 | 8.267363 | 8.386674 | 7.265871 | 7.146560 |
+| 3 | 0 | 2015-12-20 | 2015-12-16 | 7.891331 | 7.931502 | 8.205929 | 8.369983 | 7.657075 | 7.493020 |
+| 4 | 0 | 2015-12-21 | 2015-12-16 | 8.360071 | 8.312632 | 9.184893 | 9.625794 | 7.440371 | 6.999469 |
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.cross_validation(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+```python
+cutoffs = timegpt_cv_df['cutoff'].unique()
+for cutoff in cutoffs:
+ fig = nixtla_client.plot(
+ pm_df.tail(100),
+ timegpt_cv_df.query('cutoff == @cutoff').drop(columns=['cutoff', 'y']),
+ level=[80, 90],
+ models=['TimeGPT']
+ )
+ display(fig)
+```
+
+
+
+
+
+
+
+
+
+
+
+## 5. Cross-validation with exogenous variables
+
+### Time features
+
+It is possible to include exogenous variables when performing
+cross-validation. Here we use the `date_features` parameter to create
+labels for each month. These features are then used by the model to make
+predictions during cross-validation.
+
+```python
+timegpt_cv_df = nixtla_client.cross_validation(
+ pm_df,
+ h=7,
+ n_windows=5,
+ freq='D',
+ level=[80, 90],
+ date_features=['month'],
+ date_features_to_one_hot=True,
+)
+timegpt_cv_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Using the following exogenous features: ['month_1.0', 'month_2.0', 'month_3.0', 'month_4.0', 'month_5.0', 'month_6.0', 'month_7.0', 'month_8.0', 'month_9.0', 'month_10.0', 'month_11.0', 'month_12.0']
+INFO:nixtla.nixtla_client:Calling Cross Validation Endpoint...
+```
+
+| | unique_id | ds | cutoff | y | TimeGPT | TimeGPT-hi-80 | TimeGPT-hi-90 | TimeGPT-lo-80 | TimeGPT-lo-90 |
+|----|----|----|----|----|----|----|----|----|----|
+| 0 | 0.0 | 2015-12-17 | 2015-12-16 | 7.591862 | 8.426320 | 8.721996 | 8.824101 | 8.130644 | 8.028540 |
+| 1 | 0.0 | 2015-12-18 | 2015-12-16 | 7.528869 | 8.049962 | 8.452083 | 8.658603 | 7.647842 | 7.441321 |
+| 2 | 0.0 | 2015-12-19 | 2015-12-16 | 7.171657 | 7.509098 | 7.984788 | 8.138017 | 7.033409 | 6.880180 |
+| 3 | 0.0 | 2015-12-20 | 2015-12-16 | 7.891331 | 7.739536 | 8.306914 | 8.641355 | 7.172158 | 6.837718 |
+| 4 | 0.0 | 2015-12-21 | 2015-12-16 | 8.360071 | 8.027471 | 8.722828 | 9.152306 | 7.332113 | 6.902636 |
+
+```python
+cutoffs = timegpt_cv_df['cutoff'].unique()
+for cutoff in cutoffs:
+ fig = nixtla_client.plot(
+ pm_df.tail(100),
+ timegpt_cv_df.query('cutoff == @cutoff').drop(columns=['cutoff', 'y']),
+ level=[80, 90],
+ models=['TimeGPT']
+ )
+ display(fig)
+```
+
+
+
+
+
+
+
+
+
+
+
+### Dynamic features
+
+Additionally you can pass dynamic exogenous variables to better inform
+`TimeGPT` about the data. You just simply have to add the exogenous
+regressors after the target column.
+
+```python
+Y_df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity.csv')
+X_df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/exogenous-vars-electricity.csv')
+df = Y_df.merge(X_df)
+```
+
+Now let’s cross validate `TimeGPT` considering this information
+
+```python
+timegpt_cv_df_x = nixtla_client.cross_validation(
+ df.groupby('unique_id').tail(100 * 48),
+ h=48,
+ n_windows=2,
+ level=[80, 90]
+)
+cutoffs = timegpt_cv_df_x.query('unique_id == "BE"')['cutoff'].unique()
+for cutoff in cutoffs:
+ fig = nixtla_client.plot(
+ df.query('unique_id == "BE"').tail(24 * 7),
+ timegpt_cv_df_x.query('cutoff == @cutoff & unique_id == "BE"').drop(columns=['cutoff', 'y']),
+ models=['TimeGPT'],
+ level=[80, 90],
+ )
+ display(fig)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: h
+INFO:nixtla.nixtla_client:Querying model metadata...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Using the following exogenous features: ['Exogenous1', 'Exogenous2', 'day_0', 'day_1', 'day_2', 'day_3', 'day_4', 'day_5', 'day_6']
+INFO:nixtla.nixtla_client:Calling Cross Validation Endpoint...
+```
+
+
+
+
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.cross_validation(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+## 6. Cross-validation with different TimeGPT instances
+
+Also, you can generate cross validation for different instances of
+`TimeGPT` using the `model` argument. Here we use the base model and the
+model for long-horizon forecasting.
+
+```python
+timegpt_cv_df_x_long_horizon = nixtla_client.cross_validation(
+ df.groupby('unique_id').tail(100 * 48),
+ h=48,
+ n_windows=2,
+ level=[80, 90],
+ model='timegpt-1-long-horizon',
+)
+timegpt_cv_df_x_long_horizon.columns = timegpt_cv_df_x_long_horizon.columns.str.replace('TimeGPT', 'TimeGPT-LongHorizon')
+timegpt_cv_df_x_models = timegpt_cv_df_x_long_horizon.merge(timegpt_cv_df_x)
+cutoffs = timegpt_cv_df_x_models.query('unique_id == "BE"')['cutoff'].unique()
+for cutoff in cutoffs:
+ fig = nixtla_client.plot(
+ df.query('unique_id == "BE"').tail(24 * 7),
+ timegpt_cv_df_x_models.query('cutoff == @cutoff & unique_id == "BE"').drop(columns=['cutoff', 'y']),
+ models=['TimeGPT', 'TimeGPT-LongHorizon'],
+ level=[80, 90],
+ )
+ display(fig)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: h
+INFO:nixtla.nixtla_client:Querying model metadata...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Using the following exogenous features: ['Exogenous1', 'Exogenous2', 'day_0', 'day_1', 'day_2', 'day_3', 'day_4', 'day_5', 'day_6']
+INFO:nixtla.nixtla_client:Calling Cross Validation Endpoint...
+```
+
+
+
+
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.cross_validation(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
diff --git a/nixtla/docs/tutorials/exogenous_variables.html.mdx b/nixtla/docs/tutorials/exogenous_variables.html.mdx
new file mode 100644
index 00000000..eba25556
--- /dev/null
+++ b/nixtla/docs/tutorials/exogenous_variables.html.mdx
@@ -0,0 +1,445 @@
+---
+output-file: exogenous_variables.html
+title: Exogenous variables
+---
+
+
+Exogenous variables or external factors are crucial in time series
+forecasting as they provide additional information that might influence
+the prediction. These variables could include holiday markers, marketing
+spending, weather data, or any other external data that correlate with
+the time series data you are forecasting.
+
+For example, if you’re forecasting ice cream sales, temperature data
+could serve as a useful exogenous variable. On hotter days, ice cream
+sales may increase.
+
+To incorporate exogenous variables in TimeGPT, you’ll need to pair each
+point in your time series data with the corresponding external data.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/01_exogenous_variables.ipynb)
+
+## 1. Import packages
+
+First, we import the required packages and initialize the Nixtla client.
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 2. Load data
+
+Let’s see an example on predicting day-ahead electricity prices. The
+following dataset contains the hourly electricity price (`y` column) for
+five markets in Europe and US, identified by the `unique_id` column. The
+columns from `Exogenous1` to `day_6` are exogenous variables that
+TimeGPT will use to predict the prices.
+
+```python
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short-with-ex-vars.csv')
+df.head()
+```
+
+| | unique_id | ds | y | Exogenous1 | Exogenous2 | day_0 | day_1 | day_2 | day_3 | day_4 | day_5 | day_6 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | BE | 2016-10-22 00:00:00 | 70.00 | 57253.0 | 49593.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 1 | BE | 2016-10-22 01:00:00 | 37.10 | 51887.0 | 46073.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 2 | BE | 2016-10-22 02:00:00 | 37.10 | 51896.0 | 44927.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 3 | BE | 2016-10-22 03:00:00 | 44.75 | 48428.0 | 44483.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 4 | BE | 2016-10-22 04:00:00 | 37.10 | 46721.0 | 44338.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+
+## 3a. Forecasting electricity prices using future exogenous variables
+
+To produce forecasts with future exogenous variables we have to add the
+future values of the exogenous variables. Let’s read this dataset. In
+this case, we want to predict 24 steps ahead, therefore each `unique_id`
+will have 24 observations.
+
+```python
+future_ex_vars_df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short-future-ex-vars.csv')
+future_ex_vars_df.head()
+```
+
+| | unique_id | ds | Exogenous1 | Exogenous2 | day_0 | day_1 | day_2 | day_3 | day_4 | day_5 | day_6 |
+|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | BE | 2016-12-31 00:00:00 | 70318.0 | 64108.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 1 | BE | 2016-12-31 01:00:00 | 67898.0 | 62492.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 2 | BE | 2016-12-31 02:00:00 | 68379.0 | 61571.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 3 | BE | 2016-12-31 03:00:00 | 64972.0 | 60381.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 4 | BE | 2016-12-31 04:00:00 | 62900.0 | 60298.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+
+Let’s call the `forecast` method, adding this information:
+
+```python
+timegpt_fcst_ex_vars_df = nixtla_client.forecast(df=df, X_df=future_ex_vars_df, h=24, level=[80, 90])
+timegpt_fcst_ex_vars_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: h
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Querying model metadata...
+INFO:nixtla.nixtla_client:Using future exogenous features: ['Exogenous1', 'Exogenous2', 'day_0', 'day_1', 'day_2', 'day_3', 'day_4', 'day_5', 'day_6']
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| | unique_id | ds | TimeGPT | TimeGPT-hi-80 | TimeGPT-hi-90 | TimeGPT-lo-80 | TimeGPT-lo-90 |
+|----|----|----|----|----|----|----|----|
+| 0 | BE | 2016-12-31 00:00:00 | 51.632830 | 61.598820 | 66.088295 | 41.666843 | 37.177372 |
+| 1 | BE | 2016-12-31 01:00:00 | 45.750877 | 54.611988 | 60.176445 | 36.889767 | 31.325312 |
+| 2 | BE | 2016-12-31 02:00:00 | 39.650543 | 46.256210 | 52.842808 | 33.044876 | 26.458277 |
+| 3 | BE | 2016-12-31 03:00:00 | 34.000072 | 44.015310 | 47.429000 | 23.984835 | 20.571144 |
+| 4 | BE | 2016-12-31 04:00:00 | 33.785370 | 43.140503 | 48.581240 | 24.430239 | 18.989498 |
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+```python
+nixtla_client.plot(
+ df[['unique_id', 'ds', 'y']],
+ timegpt_fcst_ex_vars_df,
+ max_insample_length=365,
+ level=[80, 90],
+)
+```
+
+
+
+We can also show the importance of the features.
+
+```python
+nixtla_client.weights_x.plot.barh(x='features', y='weights')
+```
+
+
+
+This plot shows that `Exogenous1` and `Exogenous2` are the most
+important for this forecasting task, as they have the largest weight.
+
+## 3b. Forecasting electricity prices using historic exogenous variables
+
+In the example above, we just loaded the future exogenous variables.
+Often, these are not available because these variables are unknown. We
+can also make forecasts using only historic exogenous variables. This
+can be done by adding the `hist_exog_list` argument with the list of
+columns of `df` to be considered as historical. In that case, we can
+pass all extra columns available in `df` as historic exogenous variables
+using
+`hist_exog_list=['Exogenous1', 'Exogenous2', 'day_0', 'day_1', 'day_2', 'day_3', 'day_4', 'day_5', 'day_6']`.
+
+> **Important**
+>
+> If you include historic exogenous variables in your model, you are
+> *implicitly* making assumptions about the future of these exogenous
+> variables in your forecast. It is recommended to make these
+> assumptions explicit by making use of future exogenous variables.
+
+Let’s call the `forecast` method, adding `hist_exog_list`:
+
+```python
+timegpt_fcst_hist_ex_vars_df = nixtla_client.forecast(
+ df=df,
+ h=24,
+ level=[80, 90],
+ hist_exog_list=['Exogenous1', 'Exogenous2', 'day_0', 'day_1', 'day_2', 'day_3', 'day_4', 'day_5', 'day_6'],
+)
+timegpt_fcst_hist_ex_vars_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: h
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Using historical exogenous features: ['Exogenous1', 'Exogenous2', 'day_0', 'day_1', 'day_2', 'day_3', 'day_4', 'day_5', 'day_6']
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| | unique_id | ds | TimeGPT | TimeGPT-hi-80 | TimeGPT-hi-90 | TimeGPT-lo-80 | TimeGPT-lo-90 |
+|----|----|----|----|----|----|----|----|
+| 0 | BE | 2016-12-31 00:00:00 | 47.311330 | 57.277317 | 61.766790 | 37.345340 | 32.855870 |
+| 1 | BE | 2016-12-31 01:00:00 | 47.142740 | 56.003850 | 61.568306 | 38.281628 | 32.717170 |
+| 2 | BE | 2016-12-31 02:00:00 | 47.311474 | 53.917137 | 60.503740 | 40.705810 | 34.119210 |
+| 3 | BE | 2016-12-31 03:00:00 | 47.224514 | 57.239750 | 60.653442 | 37.209280 | 33.795586 |
+| 4 | BE | 2016-12-31 04:00:00 | 47.266945 | 56.622078 | 62.062817 | 37.911810 | 32.471073 |
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+```python
+nixtla_client.plot(
+ df[['unique_id', 'ds', 'y']],
+ timegpt_fcst_hist_ex_vars_df,
+ max_insample_length=365,
+ level=[80, 90],
+)
+```
+
+
+
+## 3c. Forecasting electricity prices using future and historic exogenous variables
+
+A third option is to use both historic and future exogenous variables.
+For example, we might not have available the future information for
+`Exogenous1` and `Exogenous2`. In this example, we drop these variables
+from our future exogenous dataframe (because we assume we do not know
+the future value of these variables), and add them to `hist_exog_list`
+to be considered as historical exogenous variables.
+
+```python
+hist_cols = ["Exogenous1", "Exogenous2"]
+future_ex_vars_df_limited = future_ex_vars_df.drop(columns=hist_cols)
+timegpt_fcst_ex_vars_df_limited = nixtla_client.forecast(df=df, X_df=future_ex_vars_df_limited, h=24, level=[80, 90], hist_exog_list=hist_cols)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: h
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Using future exogenous features: ['day_0', 'day_1', 'day_2', 'day_3', 'day_4', 'day_5', 'day_6']
+INFO:nixtla.nixtla_client:Using historical exogenous features: ['Exogenous1', 'Exogenous2']
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+```python
+nixtla_client.plot(
+ df[['unique_id', 'ds', 'y']],
+ timegpt_fcst_ex_vars_df_limited,
+ max_insample_length=365,
+ level=[80, 90],
+)
+```
+
+
+
+Note that TimeGPT informs you which variables are used as historic
+exogenous and which are used as future exogenous.
+
+## 3d. Forecasting future exogenous variables
+
+A fourth option in case the future exogenous variables are not available
+is to forecast them. Below, we’ll show you how we can also forecast
+`Exogenous1` and `Exogenous2` separately, so that you can generate the
+future exogenous variables in case they are not available.
+
+```python
+# We read the data and create separate dataframes for the historic exogenous that we want to forecast separately.
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short-with-ex-vars.csv')
+df_exog1 = df[['unique_id', 'ds', 'Exogenous1']]
+df_exog2 = df[['unique_id', 'ds', 'Exogenous2']]
+```
+
+Next, we can use TimeGPT to forecast `Exogenous1` and `Exogenous2`. In
+this case, we assume these quantities can be separately forecast.
+
+```python
+timegpt_fcst_ex1 = nixtla_client.forecast(df=df_exog1, h=24, target_col='Exogenous1')
+timegpt_fcst_ex2 = nixtla_client.forecast(df=df_exog2, h=24, target_col='Exogenous2')
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: h
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: h
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+We can now start creating `X_df`, which contains the future exogenous
+variables.
+
+```python
+timegpt_fcst_ex1 = timegpt_fcst_ex1.rename(columns={'TimeGPT':'Exogenous1'})
+timegpt_fcst_ex2 = timegpt_fcst_ex2.rename(columns={'TimeGPT':'Exogenous2'})
+```
+
+
+```python
+X_df = timegpt_fcst_ex1.merge(timegpt_fcst_ex2)
+```
+
+Next, we also need to add the `day_0` to `day_6` future exogenous
+variables. These are easy: this is just the weekday, which we can
+extract from the `ds` column.
+
+```python
+# We have 7 days, for each day a separate column denoting 1/0
+for i in range(7):
+ X_df[f'day_{i}'] = 1 * (pd.to_datetime(X_df['ds']).dt.weekday == i)
+```
+
+We have now created `X_df`, let’s investigate it:
+
+```python
+X_df.head(10)
+```
+
+| | unique_id | ds | Exogenous1 | Exogenous2 | day_0 | day_1 | day_2 | day_3 | day_4 | day_5 | day_6 |
+|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | BE | 2016-12-31 00:00:00 | 70861.410 | 66282.560 | 0 | 0 | 0 | 0 | 0 | 1 | 0 |
+| 1 | BE | 2016-12-31 01:00:00 | 67851.830 | 64465.370 | 0 | 0 | 0 | 0 | 0 | 1 | 0 |
+| 2 | BE | 2016-12-31 02:00:00 | 67246.660 | 63257.117 | 0 | 0 | 0 | 0 | 0 | 1 | 0 |
+| 3 | BE | 2016-12-31 03:00:00 | 64027.203 | 62059.316 | 0 | 0 | 0 | 0 | 0 | 1 | 0 |
+| 4 | BE | 2016-12-31 04:00:00 | 61524.086 | 61247.062 | 0 | 0 | 0 | 0 | 0 | 1 | 0 |
+| 5 | BE | 2016-12-31 05:00:00 | 63054.086 | 62052.312 | 0 | 0 | 0 | 0 | 0 | 1 | 0 |
+| 6 | BE | 2016-12-31 06:00:00 | 65199.473 | 63457.720 | 0 | 0 | 0 | 0 | 0 | 1 | 0 |
+| 7 | BE | 2016-12-31 07:00:00 | 68285.770 | 65388.656 | 0 | 0 | 0 | 0 | 0 | 1 | 0 |
+| 8 | BE | 2016-12-31 08:00:00 | 72038.484 | 67406.836 | 0 | 0 | 0 | 0 | 0 | 1 | 0 |
+| 9 | BE | 2016-12-31 09:00:00 | 72821.190 | 68057.240 | 0 | 0 | 0 | 0 | 0 | 1 | 0 |
+
+Let’s compare it to our pre-loaded version:
+
+```python
+future_ex_vars_df.head(10)
+```
+
+| | unique_id | ds | Exogenous1 | Exogenous2 | day_0 | day_1 | day_2 | day_3 | day_4 | day_5 | day_6 |
+|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | BE | 2016-12-31 00:00:00 | 70318.0 | 64108.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 1 | BE | 2016-12-31 01:00:00 | 67898.0 | 62492.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 2 | BE | 2016-12-31 02:00:00 | 68379.0 | 61571.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 3 | BE | 2016-12-31 03:00:00 | 64972.0 | 60381.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 4 | BE | 2016-12-31 04:00:00 | 62900.0 | 60298.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 5 | BE | 2016-12-31 05:00:00 | 62364.0 | 60339.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 6 | BE | 2016-12-31 06:00:00 | 64242.0 | 62576.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 7 | BE | 2016-12-31 07:00:00 | 65884.0 | 63732.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 8 | BE | 2016-12-31 08:00:00 | 68217.0 | 66235.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 9 | BE | 2016-12-31 09:00:00 | 69921.0 | 66801.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+
+As you can see, the values for `Exogenous1` and `Exogenous2` are
+slightly different, which makes sense because we’ve made a forecast of
+these values with TimeGPT.
+
+Let’s create a new forecast of our electricity prices with TimeGPT using
+our new `X_df`:
+
+```python
+timegpt_fcst_ex_vars_df_new = nixtla_client.forecast(df=df, X_df=X_df, h=24, level=[80, 90])
+timegpt_fcst_ex_vars_df_new.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: h
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Using future exogenous features: ['Exogenous1', 'Exogenous2', 'day_0', 'day_1', 'day_2', 'day_3', 'day_4', 'day_5', 'day_6']
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| | unique_id | ds | TimeGPT | TimeGPT-hi-80 | TimeGPT-hi-90 | TimeGPT-lo-80 | TimeGPT-lo-90 |
+|----|----|----|----|----|----|----|----|
+| 0 | BE | 2016-12-31 00:00:00 | 46.987225 | 56.953213 | 61.442684 | 37.021236 | 32.531765 |
+| 1 | BE | 2016-12-31 01:00:00 | 25.719133 | 34.580242 | 40.144700 | 16.858023 | 11.293568 |
+| 2 | BE | 2016-12-31 02:00:00 | 38.553528 | 45.159195 | 51.745792 | 31.947860 | 25.361261 |
+| 3 | BE | 2016-12-31 03:00:00 | 35.771927 | 45.787163 | 49.200855 | 25.756690 | 22.342999 |
+| 4 | BE | 2016-12-31 04:00:00 | 34.555115 | 43.910248 | 49.350986 | 25.199984 | 19.759243 |
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+Let’s create a combined dataframe with the two forecasts and plot the
+values to compare the forecasts.
+
+```python
+timegpt_fcst_ex_vars_df = timegpt_fcst_ex_vars_df.rename(columns={'TimeGPT':'TimeGPT-provided_exogenous'})
+timegpt_fcst_ex_vars_df_new = timegpt_fcst_ex_vars_df_new.rename(columns={'TimeGPT':'TimeGPT-forecasted_exogenous'})
+
+forecasts = timegpt_fcst_ex_vars_df[['unique_id', 'ds', 'TimeGPT-provided_exogenous']].merge(timegpt_fcst_ex_vars_df_new[['unique_id', 'ds', 'TimeGPT-forecasted_exogenous']])
+```
+
+
+```python
+nixtla_client.plot(
+ df[['unique_id', 'ds', 'y']],
+ forecasts,
+ max_insample_length=365,
+)
+```
+
+
+
+As you can see, we obtain a slightly different forecast if we use our
+forecasted exogenous variables.
+
diff --git a/nixtla/docs/tutorials/finetune_depth_finetuning.html.mdx b/nixtla/docs/tutorials/finetune_depth_finetuning.html.mdx
new file mode 100644
index 00000000..40bcde13
--- /dev/null
+++ b/nixtla/docs/tutorials/finetune_depth_finetuning.html.mdx
@@ -0,0 +1,165 @@
+---
+output-file: finetune_depth_finetuning.html
+title: Controlling the level of fine-tuning
+---
+
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/23_finetune_depth_finetuning.ipynb)
+
+## 1. Import packages
+
+First, we import the required packages and initialize the Nixtla client
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+from utilsforecast.losses import mae, mse
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 2. Load data
+
+```python
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/air_passengers.csv')
+df.head()
+```
+
+| | timestamp | value |
+|-----|------------|-------|
+| 0 | 1949-01-01 | 112 |
+| 1 | 1949-02-01 | 118 |
+| 2 | 1949-03-01 | 132 |
+| 3 | 1949-04-01 | 129 |
+| 4 | 1949-05-01 | 121 |
+
+Now, we split the data into a training and test set so that we can
+measure the performance of the model as we vary `finetune_depth`.
+
+```python
+train = df[:-24]
+test = df[-24:]
+```
+
+Next, we fine-tune TimeGPT and vary `finetune_depth` to measure the
+impact on performance.
+
+## 3. Fine-tuning with `finetune_depth`
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+As mentioned above, `finetune_depth` controls how many parameters from
+TimeGPT are fine-tuned on your particular dataset. If the value is set
+to 1, only a few parameters are fine-tuned. Setting it to 5 means that
+all parameters of the model will be fine-tuned.
+
+Using a large value for `finetune_depth` can lead to better performances
+for large datasets with complex patterns. However, it can also lead to
+overfitting, in which case the accuracy of the forecasts may degrade, as
+we will see from the small experiment below.
+
+```python
+depths = [1, 2, 3, 4, 5]
+
+test = test.copy()
+
+for depth in depths:
+ preds_df = nixtla_client.forecast(
+ df=train,
+ h=24,
+ finetune_steps=5,
+ finetune_depth=depth,
+ time_col='timestamp',
+ target_col='value')
+
+ preds = preds_df['TimeGPT'].values
+
+ test.loc[:,f'TimeGPT_depth{depth}'] = preds
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+INFO:nixtla.nixtla_client:Querying model metadata...
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon. This may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon. This may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon. This may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon. This may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon. This may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+```python
+test['unique_id'] = 0
+
+evaluation = evaluate(test, metrics=[mae, mse], time_col="timestamp", target_col="value")
+evaluation
+```
+
+| | unique_id | metric | TimeGPT_depth1 | TimeGPT_depth2 | TimeGPT_depth3 | TimeGPT_depth4 | TimeGPT_depth5 |
+|----|----|----|----|----|----|----|----|
+| 0 | 0 | mae | 22.675540 | 17.908963 | 21.318518 | 24.745096 | 28.734302 |
+| 1 | 0 | mse | 677.254283 | 461.320852 | 676.202126 | 991.835359 | 1119.722602 |
+
+From the result above, we can see that a `finetune_depth` of 2 achieves
+the best results since it has the lowest MAE and MSE.
+
+Also notice that with a `finetune_depth` of 4 and 5, the performance
+degrades, which is a clear sign of overfitting.
+
+Thus, keep in mind that fine-tuning can be a bit of trial and error. You
+might need to adjust the number of `finetune_steps` and the level of
+`finetune_depth` based on your specific needs and the complexity of your
+data. Usually, a higher `finetune_depth` works better for large
+datasets. In this specific tutorial, since we were forecasting a single
+series with a very short dataset, increasing the depth led to
+overfitting.
+
+It’s recommended to monitor the model’s performance during fine-tuning
+and adjust as needed. Be aware that more `finetune_steps` and a larger
+value of `finetune_depth` may lead to longer training times and could
+potentially lead to overfitting if not managed properly.
+
diff --git a/nixtla/docs/tutorials/finetuning.html.mdx b/nixtla/docs/tutorials/finetuning.html.mdx
new file mode 100644
index 00000000..e4e8cfa0
--- /dev/null
+++ b/nixtla/docs/tutorials/finetuning.html.mdx
@@ -0,0 +1,130 @@
+---
+output-file: finetuning.html
+title: Fine-tuning
+---
+
+
+Fine-tuning is a powerful process for utilizing TimeGPT more
+effectively. Foundation models such as TimeGPT are pre-trained on vast
+amounts of data, capturing wide-ranging features and patterns. These
+models can then be specialized for specific contexts or domains. With
+fine-tuning, the model’s parameters are refined to forecast a new task,
+allowing it to tailor its vast pre-existing knowledge towards the
+requirements of the new data. Fine-tuning thus serves as a crucial
+bridge, linking TimeGPT’s broad capabilities to your tasks
+specificities.
+
+Concretely, the process of fine-tuning consists of performing a certain
+number of training iterations on your input data minimizing the
+forecasting error. The forecasts will then be produced with the updated
+model. To control the number of iterations, use the `finetune_steps`
+argument of the `forecast` method.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/06_finetuning.ipynb)
+
+## 1. Import packages
+
+First, we import the required packages and initialize the Nixtla client
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+from utilsforecast.losses import mae, mse
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 2. Load data
+
+```python
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/air_passengers.csv')
+df.head()
+```
+
+| | timestamp | value |
+|-----|------------|-------|
+| 0 | 1949-01-01 | 112 |
+| 1 | 1949-02-01 | 118 |
+| 2 | 1949-03-01 | 132 |
+| 3 | 1949-04-01 | 129 |
+| 4 | 1949-05-01 | 121 |
+
+## 3. Fine-tuning
+
+Here, `finetune_steps=10` means the model will go through 10 iterations
+of training on your time series data.
+
+```python
+timegpt_fcst_finetune_df = nixtla_client.forecast(
+ df=df, h=12, finetune_steps=10,
+ time_col='timestamp', target_col='value',
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+INFO:nixtla.nixtla_client:Querying model metadata...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+```python
+nixtla_client.plot(
+ df, timegpt_fcst_finetune_df,
+ time_col='timestamp', target_col='value',
+)
+```
+
+
+
+Keep in mind that fine-tuning can be a bit of trial and error. You might
+need to adjust the number of `finetune_steps` based on your specific
+needs and the complexity of your data. Usually, a larger value of
+`finetune_steps` works better for large datasets.
+
+It’s recommended to monitor the model’s performance during fine-tuning
+and adjust as needed. Be aware that more `finetune_steps` may lead to
+longer training times and could potentially lead to overfitting if not
+managed properly.
+
+Remember, fine-tuning is a powerful feature, but it should be used
+thoughtfully and carefully.
+
+For a detailed guide on using a specific loss function for fine-tuning,
+check out the [Fine-tuning with a specific loss
+function](https://docs.nixtla.io/docs/tutorials-fine_tuning_with_a_specific_loss_function)
+tutorial.
+
+Read also our detailed tutorial on [controlling the level of
+fine-tuning](https://docs.nixtla.io/docs/tutorials-finetune_depth_finetuning)
+using `finetune_depth`.
+
diff --git a/nixtla/docs/tutorials/hierarchical_forecasting.html.mdx b/nixtla/docs/tutorials/hierarchical_forecasting.html.mdx
new file mode 100644
index 00000000..f60560c9
--- /dev/null
+++ b/nixtla/docs/tutorials/hierarchical_forecasting.html.mdx
@@ -0,0 +1,314 @@
+---
+output-file: hierarchical_forecasting.html
+title: Hierarchical forecasting
+---
+
+
+In forecasting, we often find ourselves in need of forecasts for both
+lower- and higher (temporal) granularities, such as product demand
+forecasts but also product category or product department forecasts.
+These granularities can be formalized through the use of a hierarchy. In
+hierarchical forecasting, we create forecasts that are coherent with
+respect to a pre-specified hierarchy of the underlying time series.
+
+With TimeGPT, we can create forecasts for multiple time series. We can
+subsequently post-process these forecasts using hierarchical forecasting
+techniques of
+[HierarchicalForecast](https://nixtlaverse.nixtla.io/hierarchicalforecast/index.html).
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/14_hierarchical_forecasting.ipynb)
+
+## 1. Import packages
+
+First, we import the required packages and initialize the Nixtla client.
+
+```python
+import pandas as pd
+import numpy as np
+
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, set the `base_url` argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 2. Load data
+
+We use the Australian Tourism dataset, from [Forecasting, Principles and
+Practices](https://otexts.com/fpp3/) which contains data on Australian
+Tourism. We are interested in forecasts for Australia’s 7 States, 27
+Zones and 76 Regions. This constitutes a hierarchy, where forecasts for
+the lower levels (e.g. the regions Sidney, Blue Mountains and Hunter)
+should be coherent with the forecasts of the higher levels (e.g. New
+South Wales).
+
+
+
+
+The dataset only contains the time series at the lowest level, so we
+need to create the time series for all hierarchies.
+
+```python
+Y_df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/tourism.csv')
+Y_df = Y_df.rename({'Trips': 'y', 'Quarter': 'ds'}, axis=1)
+Y_df.insert(0, 'Country', 'Australia')
+Y_df = Y_df[['Country', 'Region', 'State', 'Purpose', 'ds', 'y']]
+Y_df['ds'] = Y_df['ds'].str.replace(r'(\d+) (Q\d)', r'\1-\2', regex=True)
+Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+
+Y_df.head(10)
+```
+
+``` text
+C:\Users\ospra\AppData\Local\Temp\ipykernel_16668\3753786659.py:6: UserWarning: Could not infer format, so each element will be parsed individually, falling back to `dateutil`. To ensure parsing is consistent and as-expected, please specify a format.
+ Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+```
+
+| | Country | Region | State | Purpose | ds | y |
+|-----|-----------|----------|-----------------|----------|------------|------------|
+| 0 | Australia | Adelaide | South Australia | Business | 1998-01-01 | 135.077690 |
+| 1 | Australia | Adelaide | South Australia | Business | 1998-04-01 | 109.987316 |
+| 2 | Australia | Adelaide | South Australia | Business | 1998-07-01 | 166.034687 |
+| 3 | Australia | Adelaide | South Australia | Business | 1998-10-01 | 127.160464 |
+| 4 | Australia | Adelaide | South Australia | Business | 1999-01-01 | 137.448533 |
+| 5 | Australia | Adelaide | South Australia | Business | 1999-04-01 | 199.912586 |
+| 6 | Australia | Adelaide | South Australia | Business | 1999-07-01 | 169.355090 |
+| 7 | Australia | Adelaide | South Australia | Business | 1999-10-01 | 134.357937 |
+| 8 | Australia | Adelaide | South Australia | Business | 2000-01-01 | 154.034398 |
+| 9 | Australia | Adelaide | South Australia | Business | 2000-04-01 | 168.776364 |
+
+The dataset can be grouped in the following hierarchical structure.
+
+```python
+spec = [
+ ['Country'],
+ ['Country', 'State'],
+ ['Country', 'Purpose'],
+ ['Country', 'State', 'Region'],
+ ['Country', 'State', 'Purpose'],
+ ['Country', 'State', 'Region', 'Purpose']
+]
+```
+
+Using the `aggregate` function from `HierarchicalForecast` we can get
+the full set of time series.
+
+> **Note**
+>
+> You can install `hierarchicalforecast` with `pip`:
+>
+> ```shell
+> pip install hierarchicalforecast
+> ```
+
+```python
+from hierarchicalforecast.utils import aggregate
+```
+
+
+```python
+Y_df, S_df, tags = aggregate(Y_df, spec)
+
+Y_df.head(10)
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|--------------|
+| 0 | Australia | 1998-01-01 | 23182.197269 |
+| 1 | Australia | 1998-04-01 | 20323.380067 |
+| 2 | Australia | 1998-07-01 | 19826.640511 |
+| 3 | Australia | 1998-10-01 | 20830.129891 |
+| 4 | Australia | 1999-01-01 | 22087.353380 |
+| 5 | Australia | 1999-04-01 | 21458.373285 |
+| 6 | Australia | 1999-07-01 | 19914.192508 |
+| 7 | Australia | 1999-10-01 | 20027.925640 |
+| 8 | Australia | 2000-01-01 | 22339.294779 |
+| 9 | Australia | 2000-04-01 | 19941.063482 |
+
+We use the final two years (8 quarters) as test set.
+
+```python
+Y_test_df = Y_df.groupby('unique_id').tail(8)
+Y_train_df = Y_df.drop(Y_test_df.index)
+```
+
+## 3. Hierarchical forecasting with TimeGPT
+
+First, we create base forecasts for all the time series with TimeGPT.
+Note that we set `add_history=True`, as we will need the in-sample
+fitted values of TimeGPT.
+
+We will predict 2 years (8 quarters), starting from 01-01-2016.
+
+```python
+timegpt_fcst = nixtla_client.forecast(df=Y_train_df, h=8, freq='QS', add_history=True)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Querying model metadata...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Calling Historical Forecast Endpoint...
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+```python
+timegpt_fcst_insample = timegpt_fcst.query("ds < '2016-01-01'")
+timegpt_fcst_outsample = timegpt_fcst.query("ds >= '2016-01-01'")
+```
+
+Let’s plot some of the forecasts, starting from the highest aggregation
+level (`Australia`), to the lowest level
+(`Australia/Queensland/Brisbane/Holiday`). We can see that there is room
+for improvement in the forecasts.
+
+```python
+nixtla_client.plot(
+ Y_df,
+ timegpt_fcst_outsample,
+ max_insample_length=4 * 12,
+ unique_ids=['Australia', 'Australia/Queensland','Australia/Queensland/Brisbane', 'Australia/Queensland/Brisbane/Holiday']
+)
+```
+
+
+
+We can make these forecasts coherent to the specified hierarchy by using
+a `HierarchicalReconciliation` method from `NeuralForecast`. We will be
+using the
+[MinTrace](https://nixtlaverse.nixtla.io/hierarchicalforecast/methods.html)
+method.
+
+```python
+from hierarchicalforecast.methods import MinTrace
+from hierarchicalforecast.core import HierarchicalReconciliation
+```
+
+
+```python
+reconcilers = [
+ MinTrace(method='ols'),
+ MinTrace(method='mint_shrink'),
+]
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+
+Y_df_with_insample_fcsts = Y_df.copy()
+Y_df_with_insample_fcsts = timegpt_fcst_insample.merge(Y_df_with_insample_fcsts)
+
+Y_rec_df = hrec.reconcile(Y_hat_df=timegpt_fcst_outsample, Y_df=Y_df_with_insample_fcsts, S=S_df, tags=tags)
+```
+
+
+```python
+Y_rec_df
+```
+
+| | unique_id | ds | TimeGPT | TimeGPT/MinTrace_method-ols | TimeGPT/MinTrace_method-mint_shrink |
+|----|----|----|----|----|----|
+| 0 | Australia | 2016-01-01 | 24967.19100 | 25044.408634 | 25394.406211 |
+| 1 | Australia | 2016-04-01 | 24528.88300 | 24503.089810 | 24327.212355 |
+| 2 | Australia | 2016-07-01 | 24221.77500 | 24083.107812 | 23813.826553 |
+| 3 | Australia | 2016-10-01 | 24559.44000 | 24548.038797 | 24174.894203 |
+| 4 | Australia | 2017-01-01 | 25570.33800 | 25669.248281 | 25560.277473 |
+| ... | ... | ... | ... | ... | ... |
+| 3395 | Australia/Western Australia/Experience Perth/V... | 2016-10-01 | 427.81146 | 435.423617 | 434.047102 |
+| 3396 | Australia/Western Australia/Experience Perth/V... | 2017-01-01 | 450.71786 | 453.434056 | 459.954598 |
+| 3397 | Australia/Western Australia/Experience Perth/V... | 2017-04-01 | 452.17923 | 460.197847 | 470.009789 |
+| 3398 | Australia/Western Australia/Experience Perth/V... | 2017-07-01 | 450.68683 | 463.034888 | 482.645932 |
+| 3399 | Australia/Western Australia/Experience Perth/V... | 2017-10-01 | 443.31050 | 451.754435 | 474.403379 |
+
+Again, we plot some of the forecasts. We can see a few, mostly minor
+differences in the forecasts.
+
+```python
+nixtla_client.plot(
+ Y_df,
+ Y_rec_df,
+ max_insample_length=4 * 12,
+ unique_ids=['Australia', 'Australia/Queensland','Australia/Queensland/Brisbane', 'Australia/Queensland/Brisbane/Holiday']
+)
+```
+
+
+
+Let’s numerically verify the forecasts to the situation where we don’t
+apply a post-processing step. We can use `HierarchicalEvaluation` for
+this.
+
+```python
+from hierarchicalforecast.evaluation import evaluate
+from utilsforecast.losses import rmse
+```
+
+
+```python
+eval_tags = {}
+eval_tags['Total'] = tags['Country']
+eval_tags['Purpose'] = tags['Country/Purpose']
+eval_tags['State'] = tags['Country/State']
+eval_tags['Regions'] = tags['Country/State/Region']
+eval_tags['Bottom'] = tags['Country/State/Region/Purpose']
+
+evaluation = evaluate(
+ df=Y_rec_df.merge(Y_test_df, on=['unique_id', 'ds']),
+ tags=eval_tags,
+ train_df=Y_train_df,
+ metrics=[rmse],
+)
+numeric_cols = evaluation.select_dtypes(np.number).columns
+evaluation[numeric_cols] = evaluation[numeric_cols].map('{:.2f}'.format)
+```
+
+
+```python
+evaluation
+```
+
+| | level | metric | TimeGPT | TimeGPT/MinTrace_method-ols | TimeGPT/MinTrace_method-mint_shrink |
+|----|----|----|----|----|----|
+| 0 | Total | rmse | 1433.07 | 1436.07 | 1627.43 |
+| 1 | Purpose | rmse | 482.09 | 475.64 | 507.50 |
+| 2 | State | rmse | 275.85 | 278.39 | 294.28 |
+| 3 | Regions | rmse | 49.40 | 47.91 | 47.99 |
+| 4 | Bottom | rmse | 19.32 | 19.11 | 18.86 |
+| 5 | Overall | rmse | 38.66 | 38.21 | 39.16 |
+
+We made a small improvement in overall RMSE by reconciling the forecasts
+with `MinTrace(ols)`, and made them slightly worse using
+`MinTrace(mint_shrink)`, indicating that the base forecasts were
+relatively strong already.
+
+However, we now have coherent forecasts too - so not only did we make a
+(small) accuracy improvement, we also got coherency to the hierarchy as
+a result of our reconciliation step.
+
+**References**
+
+- [Hyndman, Rob J., and George Athanasopoulos (2021). “Forecasting:
+ Principles and Practice (3rd Ed)”](https://otexts.com/fpp3/)
+
diff --git a/nixtla/docs/tutorials/historical_forecast.html.mdx b/nixtla/docs/tutorials/historical_forecast.html.mdx
new file mode 100644
index 00000000..c753d560
--- /dev/null
+++ b/nixtla/docs/tutorials/historical_forecast.html.mdx
@@ -0,0 +1,129 @@
+---
+output-file: historical_forecast.html
+title: Historical forecast
+---
+
+
+Our time series model offers a powerful feature that allows users to
+retrieve historical forecasts alongside the prospective predictions.
+This functionality is accessible through the forecast method by setting
+the `add_history=True` argument.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/09_historical_forecast.ipynb)
+
+## 1. Import packages
+
+First, we install and import the required packages and initialize the
+Nixtla client.
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, set the `base_url` argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 2. Load data
+
+Now you can start to make forecasts! Let’s import an example:
+
+```python
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/air_passengers.csv')
+df.head()
+```
+
+| | timestamp | value |
+|-----|------------|-------|
+| 0 | 1949-01-01 | 112 |
+| 1 | 1949-02-01 | 118 |
+| 2 | 1949-03-01 | 132 |
+| 3 | 1949-04-01 | 129 |
+| 4 | 1949-05-01 | 121 |
+
+```python
+nixtla_client.plot(df, time_col='timestamp', target_col='value')
+```
+
+
+
+## 3. Historical forecast
+
+Let’s add fitted values. When `add_history` is set to True, the output
+DataFrame will include not only the future forecasts determined by the h
+argument, but also the historical predictions. Currently, the historical
+forecasts are not affected by `h`, and have a fix horizon depending on
+the frequency of the data. The historical forecasts are produced in a
+rolling window fashion, and concatenated. This means that the model is
+applied sequentially at each time step using only the most recent
+information available up to that point.
+
+```python
+timegpt_fcst_with_history_df = nixtla_client.forecast(
+ df=df, h=12, time_col='timestamp', target_col='value',
+ add_history=True,
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Calling Historical Forecast Endpoint...
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+```python
+timegpt_fcst_with_history_df.head()
+```
+
+| | timestamp | TimeGPT |
+|-----|------------|------------|
+| 0 | 1951-01-01 | 135.483673 |
+| 1 | 1951-02-01 | 144.442398 |
+| 2 | 1951-03-01 | 157.191910 |
+| 3 | 1951-04-01 | 148.769363 |
+| 4 | 1951-05-01 | 140.472946 |
+
+Let’s plot the results. This consolidated view of past and future
+predictions can be invaluable for understanding the model’s behavior and
+for evaluating its performance over time.
+
+```python
+nixtla_client.plot(df, timegpt_fcst_with_history_df, time_col='timestamp', target_col='value')
+```
+
+
+
+Please note, however, that the initial values of the series are not
+included in these historical forecasts. This is because `TimeGPT`
+requires a certain number of initial observations to generate reliable
+forecasts. Therefore, while interpreting the output, it’s important to
+be aware that the first few observations serve as the basis for the
+model’s predictions and are not themselves predicted values.
+
diff --git a/nixtla/docs/tutorials/holidays.html.mdx b/nixtla/docs/tutorials/holidays.html.mdx
new file mode 100644
index 00000000..9a134a65
--- /dev/null
+++ b/nixtla/docs/tutorials/holidays.html.mdx
@@ -0,0 +1,220 @@
+---
+output-file: holidays.html
+title: Holidays and special dates
+---
+
+
+Calendar variables and special dates are one of the most common types of
+additional variables used in forecasting applications. They provide
+additional context on the current state of the time series, especially
+for window-based models such as TimeGPT-1. These variables often include
+adding information on each observation’s month, week, day, or hour. For
+example, in high-frequency hourly data, providing the current month of
+the year provides more context than the limited history available in the
+input window to improve the forecasts.
+
+In this tutorial we will show how to add calendar variables
+automatically to a dataset using the `date_features` function.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/02_holidays.ipynb)
+
+## 1. Import packages
+
+First, we import the required packages and initialize the Nixtla client.
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 2. Load data
+
+We will use a Google trends dataset on chocolate, with monthly data.
+
+```python
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/google_trend_chocolate.csv')
+df['month'] = pd.to_datetime(df['month']).dt.to_period('M').dt.to_timestamp('M')
+```
+
+
+```python
+df.head()
+```
+
+| | month | chocolate |
+|-----|------------|-----------|
+| 0 | 2004-01-31 | 35 |
+| 1 | 2004-02-29 | 45 |
+| 2 | 2004-03-31 | 28 |
+| 3 | 2004-04-30 | 30 |
+| 4 | 2004-05-31 | 29 |
+
+## 3. Forecasting with holidays and special dates
+
+Given the predominance usage of calendar variables, we included an
+automatic creation of common calendar variables to the forecast method
+as a pre-processing step. Let’s create a future dataframe that contains
+the upcoming holidays in the United States.
+
+```python
+# Create future dataframe with exogenous features
+
+start_date = '2024-05'
+dates = pd.date_range(start=start_date, periods=14, freq='M')
+
+dates = dates.to_period('M').to_timestamp('M')
+
+future_df = pd.DataFrame(dates, columns=['month'])
+```
+
+
+```python
+from nixtla.date_features import CountryHolidays
+
+us_holidays = CountryHolidays(countries=['US'])
+dates = pd.date_range(start=future_df.iloc[0]['month'], end=future_df.iloc[-1]['month'], freq='D')
+holidays_df = us_holidays(dates)
+monthly_holidays = holidays_df.resample('M').max()
+
+monthly_holidays = monthly_holidays.reset_index(names='month')
+
+future_df = future_df.merge(monthly_holidays)
+
+future_df.head()
+```
+
+| | month | US_New Year's Day | US_Memorial Day | US_Juneteenth National Independence Day | US_Independence Day | US_Labor Day | US_Veterans Day | US_Thanksgiving | US_Christmas Day | US_Martin Luther King Jr. Day | US_Washington's Birthday | US_Columbus Day |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | 2024-05-31 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
+| 1 | 2024-06-30 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
+| 2 | 2024-07-31 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
+| 3 | 2024-08-31 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
+| 4 | 2024-09-30 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 |
+
+We perform the same steps for the input dataframe.
+
+```python
+# Add exogenous features to input dataframe
+
+dates = pd.date_range(start=df.iloc[0]['month'], end=df.iloc[-1]['month'], freq='D')
+holidays_df = us_holidays(dates)
+monthly_holidays = holidays_df.resample('M').max()
+
+monthly_holidays = monthly_holidays.reset_index(names='month')
+
+df = df.merge(monthly_holidays)
+
+df.tail()
+```
+
+| | month | chocolate | US_New Year's Day | US_New Year's Day (observed) | US_Memorial Day | US_Independence Day | US_Independence Day (observed) | US_Labor Day | US_Veterans Day | US_Thanksgiving | US_Christmas Day | US_Christmas Day (observed) | US_Martin Luther King Jr. Day | US_Washington's Birthday | US_Columbus Day | US_Veterans Day (observed) | US_Juneteenth National Independence Day | US_Juneteenth National Independence Day (observed) |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 239 | 2023-12-31 | 90 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
+| 240 | 2024-01-31 | 64 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 |
+| 241 | 2024-02-29 | 66 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 |
+| 242 | 2024-03-31 | 59 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
+| 243 | 2024-04-30 | 51 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
+
+Great! Now, TimeGPT will consider the holidays as exogenous variables
+and the upcoming holidays will help it make predictions.
+
+```python
+fcst_df = nixtla_client.forecast(
+ df=df,
+ h=14,
+ freq='M',
+ time_col='month',
+ target_col='chocolate',
+ X_df=future_df
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: M
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon. This may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Using the following exogenous variables: US_New Year's Day, US_Memorial Day, US_Juneteenth National Independence Day, US_Independence Day, US_Labor Day, US_Veterans Day, US_Thanksgiving, US_Christmas Day, US_Martin Luther King Jr. Day, US_Washington's Birthday, US_Columbus Day
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+```python
+nixtla_client.plot(
+ df,
+ fcst_df,
+ time_col='month',
+ target_col='chocolate',
+)
+```
+
+
+
+We can then plot the weights of each holiday to see which are more
+important in forecasing the interest in chocolate.
+
+```python
+nixtla_client.weights_x.plot.barh(x='features', y='weights', figsize=(10, 10))
+```
+
+
+
+Here’s a breakdown of how the `date_features` parameter works:
+
+- **`date_features` (bool or list of str or callable)**: This
+ parameter specifies which date attributes to consider.
+ - If set to `True`, the model will automatically add the most
+ common date features related to the frequency of the given
+ dataframe (`df`). For a daily frequency, this could include
+ features like day of the week, month, and year.
+ - If provided a list of strings, it will consider those specific
+ date attributes. For example,
+ `date_features=['weekday', 'month']` will only add the day of
+ the week and month as features.
+ - If provided a callable, it should be a function that takes dates
+ as input and returns the desired feature. This gives flexibility
+ in computing custom date features.
+- **`date_features_to_one_hot` (bool or list of str)**: After
+ determining the date features, one might want to one-hot encode
+ them, especially if they are categorical in nature (like weekdays).
+ One-hot encoding transforms these categorical features into a binary
+ matrix, making them more suitable for many machine learning
+ algorithms.
+ - If `date_features=True`, then by default, all computed date
+ features will be one-hot encoded.
+ - If provided a list of strings, only those specific date features
+ will be one-hot encoded.
+
+By leveraging the `date_features` and `date_features_to_one_hot`
+parameters, one can efficiently incorporate the temporal effects of date
+attributes into their forecasting model, potentially enhancing its
+accuracy and interpretability.
+
diff --git a/nixtla/docs/tutorials/how_to_improve_forecast_accuracy.html.mdx b/nixtla/docs/tutorials/how_to_improve_forecast_accuracy.html.mdx
new file mode 100644
index 00000000..b524b4e6
--- /dev/null
+++ b/nixtla/docs/tutorials/how_to_improve_forecast_accuracy.html.mdx
@@ -0,0 +1,421 @@
+---
+output-file: how_to_improve_forecast_accuracy.html
+title: Improve Forecast Accuracy with TimeGPT
+---
+
+
+In this notebook, we demonstrate how to use TimeGPT for forecasting and
+explore three common strategies to enhance forecast accuracy. We use the
+hourly electricity price data from Germany as our example dataset.
+Before running the notebook, please initiate a NixtlaClient object with
+your api_key in the code snippet below.
+
+### Result Summary
+
+| Steps | Description | MAE | MAE Improvement (%) | RMSE | RMSE Improvement (%) |
+|------|----------------------|------|----------------|------|-----------------|
+| 0 | Zero-Shot TimeGPT | 18.5 | N/A | 20.0 | N/A |
+| 1 | Add Fine-Tuning Steps | 11.5 | 38% | 12.6 | 37% |
+| 2 | Adjust Fine-Tuning Loss | 9.6 | 48% | 11.0 | 45% |
+| 3 | Fine-tune more parameters | 9.0 | 51% | 11.3 | 44% |
+| 4 | Add Exogenous Variables | 4.6 | 75% | 6.4 | 68% |
+| 5 | Switch to Long-Horizon Model | 6.4 | 65% | 7.7 | 62% |
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/22_how_to_improve_forecast_accuracy.ipynb)
+
+First, we install and import the required packages, initialize the
+Nixtla client and create a function for calculating evaluation metrics.
+
+```python
+import numpy as np
+import pandas as pd
+
+from utilsforecast.evaluation import evaluate
+from utilsforecast.plotting import plot_series
+from utilsforecast.losses import mae, rmse
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+## 1. load in dataset
+
+In this notebook, we use hourly electricity prices as our example
+dataset, which consists of 5 time series, each with approximately 1700
+data points. For demonstration purposes, we focus on the German
+electricity price series. The time series is split, with the last 48
+steps (2 days) set aside as the test set.
+
+```python
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short-with-ex-vars.csv')
+df['ds'] = pd.to_datetime(df['ds'])
+df_sub = df.query('unique_id == "DE"')
+```
+
+
+```python
+df_train = df_sub.query('ds < "2017-12-29"')
+df_test = df_sub.query('ds >= "2017-12-29"')
+df_train.shape, df_test.shape
+```
+
+``` text
+((1632, 12), (48, 12))
+```
+
+```python
+plot_series(df_train[['unique_id','ds','y']][-200:], forecasts_df= df_test[['unique_id','ds','y']].rename(columns={'y': 'test'}))
+```
+
+
+
+## 2. Benchmark Forecasting using TimeGPT
+
+We used TimeGPT to generate a zero-shot forecast for the time series. As
+illustrated in the plot, TimeGPT captures the overall trend reasonably
+well, but it falls short in modeling the short-term fluctuations and
+cyclical patterns present in the actual data. During the test period,
+the model achieved a Mean Absolute Error (MAE) of 18.5 and a Root Mean
+Square Error (RMSE) of 20. This forecast serves as a baseline for
+further comparison and optimization.
+
+```python
+fcst_timegpt = nixtla_client.forecast(df = df_train[['unique_id','ds','y']],
+ h=2*24,
+ target_col = 'y',
+ level = [90, 95])
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: h
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Querying model metadata...
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon, this may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+```python
+metrics = [mae, rmse]
+```
+
+
+```python
+evaluation = evaluate(
+ fcst_timegpt.merge(df_test, on=['unique_id', 'ds']),
+ metrics=metrics,
+ models=['TimeGPT']
+)
+evaluation
+```
+
+| | unique_id | metric | TimeGPT |
+|-----|-----------|--------|-----------|
+| 0 | DE | mae | 18.519004 |
+| 1 | DE | rmse | 20.037751 |
+
+```python
+plot_series(df_sub.iloc[-150:], forecasts_df= fcst_timegpt, level = [90])
+```
+
+
+
+## 3. Methods to Improve Forecast Accuracy
+
+### 3a. Add Finetune Steps
+
+The first approach to enhance forecast accuracy is to increase the
+number of fine-tuning steps. The fine-tuning process adjusts the weights
+within the TimeGPT model, allowing it to better fit your customized
+data. This adjustment enables TimeGPT to learn the nuances of your time
+series more effectively, leading to more accurate forecasts. With 30
+fine-tuning steps, we observe that the MAE decreases to 11.5 and the
+RMSE drops to 12.6.
+
+```python
+fcst_finetune_df = nixtla_client.forecast(df=df_train[['unique_id', 'ds', 'y']],
+ h=24*2,
+ finetune_steps = 30,
+ level=[90, 95])
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: h
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon, this may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+```python
+evaluation = evaluate(
+ fcst_finetune_df.merge(df_test, on=['unique_id', 'ds']),
+ metrics=metrics,
+ models=['TimeGPT']
+)
+evaluation
+```
+
+| | unique_id | metric | TimeGPT |
+|-----|-----------|--------|-----------|
+| 0 | DE | mae | 11.458185 |
+| 1 | DE | rmse | 12.642999 |
+
+```python
+plot_series(df_sub[-200:], forecasts_df= fcst_finetune_df, level = [90])
+```
+
+
+
+### 3b. Finetune with Different Loss Function
+
+The second way to further reduce forecast error is to adjust the loss
+function used during fine-tuning. You can specify your customized loss
+function using the `finetune_loss` parameter. By modifying the loss
+function, we observe that the MAE decreases to 9.6 and the RMSE reduces
+to 11.0.
+
+```python
+fcst_finetune_mae_df = nixtla_client.forecast(df=df_train[['unique_id', 'ds', 'y']],
+ h=24*2,
+ finetune_steps = 30,
+ finetune_loss = 'mae',
+ level=[90, 95])
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: h
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon, this may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+```python
+evaluation = evaluate(
+ fcst_finetune_mae_df.merge(df_test, on=['unique_id', 'ds']),
+ metrics=metrics,
+ models=['TimeGPT']
+)
+evaluation
+```
+
+| | unique_id | metric | TimeGPT |
+|-----|-----------|--------|-----------|
+| 0 | DE | mae | 9.640649 |
+| 1 | DE | rmse | 10.956003 |
+
+```python
+plot_series(df_sub[-200:], forecasts_df= fcst_finetune_mae_df, level = [90])
+```
+
+
+
+### 3c. Adjust the number of parameters being fine-tuned
+
+Using the `finetune_depth` parameter, we can control the number of
+parameters that get fine-tuned. By default, `finetune_depth=1`, meaning
+that few parameters are tuned. We can set it to any value from 1 to 5,
+where 5 means that we fine-tune all of the parameters of the model.
+
+```python
+fcst_finetune_depth_df = nixtla_client.forecast(df=df_train[['unique_id', 'ds', 'y']],
+ h=24*2,
+ finetune_steps = 30,
+ finetune_depth=2,
+ finetune_loss = 'mae',
+ level=[90, 95])
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: h
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon, this may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+```python
+evaluation = evaluate(
+ fcst_finetune_depth_df.merge(df_test, on=['unique_id', 'ds']),
+ metrics=metrics,
+ models=['TimeGPT']
+)
+evaluation
+```
+
+| | unique_id | metric | TimeGPT |
+|-----|-----------|--------|-----------|
+| 0 | DE | mae | 9.002193 |
+| 1 | DE | rmse | 11.348207 |
+
+```python
+plot_series(df_sub[-200:], forecasts_df= fcst_finetune_depth_df, level = [90])
+```
+
+
+
+### 3d. Forecast with Exogenous Variables
+
+Exogenous variables are external factors or predictors that are not part
+of the target time series but can influence its behavior. Incorporating
+these variables can provide the model with additional context, improving
+its ability to understand complex relationships and patterns in the
+data.
+
+To use exogenous variables in TimeGPT, pair each point in your input
+time series with the corresponding external data. If you have future
+values available for these variables during the forecast period, include
+them using the X_df parameter. Otherwise, you can omit this parameter
+and still see improvements using only historical values. In the example
+below, we incorporate 8 historical exogenous variables along with their
+values during the test period, which reduces the MAE and RMSE to 4.6 and
+6.4, respectively.
+
+```python
+df_train.head()
+```
+
+| | unique_id | ds | y | Exogenous1 | Exogenous2 | day_0 | day_1 | day_2 | day_3 | day_4 | day_5 | day_6 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 1680 | DE | 2017-10-22 00:00:00 | 19.10 | 16972.75 | 15778.92975 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 |
+| 1681 | DE | 2017-10-22 01:00:00 | 19.03 | 16254.50 | 16664.20950 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 |
+| 1682 | DE | 2017-10-22 02:00:00 | 16.90 | 15940.25 | 17728.74950 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 |
+| 1683 | DE | 2017-10-22 03:00:00 | 12.98 | 15959.50 | 18578.13850 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 |
+| 1684 | DE | 2017-10-22 04:00:00 | 9.24 | 16071.50 | 19389.16750 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 |
+
+```python
+future_ex_vars_df = df_test.drop(columns = ['y'])
+future_ex_vars_df.head()
+```
+
+| | unique_id | ds | Exogenous1 | Exogenous2 | day_0 | day_1 | day_2 | day_3 | day_4 | day_5 | day_6 |
+|----|----|----|----|----|----|----|----|----|----|----|----|
+| 3312 | DE | 2017-12-29 00:00:00 | 17347.00 | 24577.92650 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 |
+| 3313 | DE | 2017-12-29 01:00:00 | 16587.25 | 24554.31950 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 |
+| 3314 | DE | 2017-12-29 02:00:00 | 16396.00 | 24651.45475 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 |
+| 3315 | DE | 2017-12-29 03:00:00 | 16481.25 | 24666.04300 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 |
+| 3316 | DE | 2017-12-29 04:00:00 | 16827.75 | 24403.33350 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 |
+
+```python
+fcst_ex_vars_df = nixtla_client.forecast(df=df_train,
+ X_df=future_ex_vars_df,
+ h=24*2,
+ level=[90, 95])
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: h
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon, this may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Using future exogenous features: ['Exogenous1', 'Exogenous2', 'day_0', 'day_1', 'day_2', 'day_3', 'day_4', 'day_5', 'day_6']
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+```python
+evaluation = evaluate(
+ fcst_ex_vars_df.merge(df_test, on=['unique_id', 'ds']),
+ metrics=metrics,
+ models=['TimeGPT']
+)
+evaluation
+```
+
+| | unique_id | metric | TimeGPT |
+|-----|-----------|--------|----------|
+| 0 | DE | mae | 4.602594 |
+| 1 | DE | rmse | 6.358831 |
+
+```python
+plot_series(df_sub[-200:], forecasts_df= fcst_ex_vars_df, level = [90])
+```
+
+
+
+### 3d. TimeGPT for Long Horizon Forecasting
+
+When the forecasting period is too long, the predicted results may not
+be as accurate. TimeGPT performs best with forecast periods that are
+shorter than one complete cycle of the time series. For longer forecast
+periods, switching to the timegpt-1-long-horizon model can yield better
+results. You can specify this model by using the model parameter.
+
+In the electricity price time series used here, one cycle is 24 steps
+(representing one day). Since we’re forecasting two days (48 steps) into
+the future, using timegpt-1-long-horizon significantly improves the
+forecasting accuracy, reducing the MAE to 6.4 and RMSE to 7.7.
+
+```python
+fcst_long_df = nixtla_client.forecast(df=df_train[['unique_id', 'ds', 'y']],
+ h=24*2,
+ model = 'timegpt-1-long-horizon',
+ level=[90, 95])
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: h
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Querying model metadata...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+```python
+evaluation = evaluate(
+ fcst_long_df.merge(df_test, on=['unique_id', 'ds']),
+ metrics=metrics,
+ models=['TimeGPT']
+)
+evaluation
+```
+
+| | unique_id | metric | TimeGPT |
+|-----|-----------|--------|----------|
+| 0 | DE | mae | 6.365540 |
+| 1 | DE | rmse | 7.738188 |
+
+```python
+plot_series(df_sub[-200:], forecasts_df= fcst_long_df, level = [90])
+```
+
+
+
+## 4. Conclusion and Next Steps
+
+In this notebook, we demonstrated four effective strategies for
+enhancing forecast accuracy with TimeGPT:
+
+1. **Increasing the number of fine-tuning steps.**
+2. **Adjusting the fine-tuning loss function.**
+3. **Incorporating exogenous variables.**
+4. **Switching to the long-horizon model for extended forecasting
+ periods.**
+
+We encourage you to experiment with these hyperparameters to identify
+the optimal settings that best suit your specific needs. Additionally,
+please refer to our documentation for further features, such as **model
+explainability** and more.
+
+In the examples provided, after applying these methods, we observed
+significant improvements in forecast accuracy metrics, as summarized
+below.
+
+### Result Summary
+
+| Steps | Description | MAE | MAE Improvement (%) | RMSE | RMSE Improvement (%) |
+|------|----------------------|------|----------------|------|-----------------|
+| 0 | Zero-Shot TimeGPT | 18.5 | N/A | 20.0 | N/A |
+| 1 | Add Fine-Tuning Steps | 11.5 | 38% | 12.6 | 37% |
+| 2 | Adjust Fine-Tuning Loss | 9.6 | 48% | 11.0 | 45% |
+| 3 | Fine-tune more parameters | 9.0 | 51% | 11.3 | 44% |
+| 4 | Add Exogenous Variables | 4.6 | 75% | 6.4 | 68% |
+| 5 | Switch to Long-Horizon Model | 6.4 | 65% | 7.7 | 62% |
+
diff --git a/nixtla/docs/tutorials/longhorizon.html.mdx b/nixtla/docs/tutorials/longhorizon.html.mdx
new file mode 100644
index 00000000..4bc41453
--- /dev/null
+++ b/nixtla/docs/tutorials/longhorizon.html.mdx
@@ -0,0 +1,163 @@
+---
+output-file: longhorizon.html
+title: Long-horizon forecasting
+---
+
+
+Long-horizon forecasting refers to predictions far into the future,
+typically exceeding two seasonal periods. However, the exact definition
+of a ‘long horizon’ can vary based on the frequency of the data. For
+example, when dealing with hourly data, a forecast for three days into
+the future is considered long-horizon, as it covers 72 timestamps
+(calculated as 3 days × 24 hours/day). In the context of monthly data, a
+period exceeding two years would typically be classified as long-horizon
+forecasting. Similarly, for daily data, a forecast spanning more than
+two weeks falls into the long-horizon category.
+
+Of course, forecasting over a long horizon comes with its challenges.
+The longer the forecast horizon, the greater the uncertainty in the
+predictions. It is also possible to have unknown factors come into play
+in the long-term that were not expected at the time of forecasting.
+
+To tackle those challenges, use TimeGPT’s specialized model for
+long-horizon forecasting by specifying `model='timegpt-1-long-horizon'`
+in your setup.
+
+For a detailed step-by-step guide, follow this tutorial on long-horizon
+forecasting.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/04_longhorizon.ipynb)
+
+## 1. Import packages
+
+First, we install and import the required packages and initialize the
+Nixtla client.
+
+```python
+from nixtla import NixtlaClient
+from datasetsforecast.long_horizon import LongHorizon
+from utilsforecast.losses import mae
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 2. Load the data
+
+Let’s load the ETTh1 dataset. This is a widely used dataset to evaluate
+models on their long-horizon forecasting capabalities.
+
+The ETTh1 dataset monitors an electricity transformer from a region of a
+province of China including oil temperature and variants of load (such
+as high useful load and high useless load) from July 2016 to July 2018
+at an hourly frequency.
+
+For this tutorial, let’s only consider the oil temperature variation
+over time.
+
+```python
+Y_df, *_ = LongHorizon.load(directory='./', group='ETTh1')
+
+Y_df.head()
+```
+
+``` text
+100%|██████████| 314M/314M [00:14<00:00, 21.3MiB/s]
+INFO:datasetsforecast.utils:Successfully downloaded datasets.zip, 314116557, bytes.
+INFO:datasetsforecast.utils:Decompressing zip file...
+INFO:datasetsforecast.utils:Successfully decompressed longhorizon\datasets\datasets.zip
+```
+
+| | unique_id | ds | y |
+|-----|-----------|---------------------|----------|
+| 0 | OT | 2016-07-01 00:00:00 | 1.460552 |
+| 1 | OT | 2016-07-01 01:00:00 | 1.161527 |
+| 2 | OT | 2016-07-01 02:00:00 | 1.161527 |
+| 3 | OT | 2016-07-01 03:00:00 | 0.862611 |
+| 4 | OT | 2016-07-01 04:00:00 | 0.525227 |
+
+For this small experiment, let’s set the horizon to 96 time steps (4
+days into the future), and we will feed TimeGPT with a sequence of 42
+days.
+
+```python
+test = Y_df[-96:] # 96 = 4 days x 24h/day
+input_seq = Y_df[-1104:-96] # Gets a sequence of 1008 observations (1008 = 42 days * 24h/day)
+```
+
+## 3. Forecasting for long-horizon
+
+Now, we are ready to use TimeGPT for long-horizon forecasting. Here, we
+need to set the `model` parameter to `"timegpt-1-long-horizon"`. This is
+the specialized model in TimeGPT that can handle such tasks.
+
+```python
+fcst_df = nixtla_client.forecast(
+ df=input_seq,
+ h=96,
+ level=[90],
+ finetune_steps=10,
+ finetune_loss='mae',
+ model='timegpt-1-long-horizon',
+ time_col='ds',
+ target_col='y'
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: H
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+
+```python
+nixtla_client.plot(Y_df[-168:], fcst_df, models=['TimeGPT'], level=[90], time_col='ds', target_col='y')
+```
+
+
+
+## Evaluation
+
+Let’s now evaluate the performance of TimeGPT using the mean absolute
+error (MAE).
+
+```python
+test = test.copy()
+
+test.loc[:, 'TimeGPT'] = fcst_df['TimeGPT'].values
+```
+
+
+```python
+evaluation = mae(test, models=['TimeGPT'], id_col='unique_id', target_col='y')
+
+print(evaluation)
+```
+
+``` text
+ unique_id TimeGPT
+0 OT 0.145393
+```
+
+Here, TimeGPT achieves a MAE of 0.146.
+
diff --git a/nixtla/docs/tutorials/loss_function_finetuning.html.mdx b/nixtla/docs/tutorials/loss_function_finetuning.html.mdx
new file mode 100644
index 00000000..81a8b0dd
--- /dev/null
+++ b/nixtla/docs/tutorials/loss_function_finetuning.html.mdx
@@ -0,0 +1,295 @@
+---
+output-file: loss_function_finetuning.html
+title: Fine-tuning with a specific loss function
+---
+
+
+When fine-tuning, the model trains on your dataset to tailor its
+predictions to your particular scenario. As such, it is possible to
+specify the loss function used during fine-tuning.
+
+Specifically, you can choose from:
+
+- `"default"` - a proprietary loss function that is robust to outliers
+- `"mae"` - mean absolute error
+- `"mse"` - mean squared error
+- `"rmse"` - root mean squared error
+- `"mape"` - mean absolute percentage error
+- `"smape"` - symmetric mean absolute percentage error
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/07_loss_function_finetuning.ipynb)
+
+## 1. Import packages
+
+First, we import the required packages and initialize the Nixtla client.
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+from utilsforecast.losses import mae, mse, rmse, mape, smape
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 2. Load data
+
+Let’s fine-tune the model on a dataset using the mean absolute error
+(MAE).
+
+For that, we simply pass the appropriate string representing the loss
+function to the `finetune_loss` parameter of the `forecast` method.
+
+```python
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/air_passengers.csv')
+df.insert(loc=0, column='unique_id', value=1)
+
+df.head()
+```
+
+| | unique_id | timestamp | value |
+|-----|-----------|------------|-------|
+| 0 | 1 | 1949-01-01 | 112 |
+| 1 | 1 | 1949-02-01 | 118 |
+| 2 | 1 | 1949-03-01 | 132 |
+| 3 | 1 | 1949-04-01 | 129 |
+| 4 | 1 | 1949-05-01 | 121 |
+
+## 3. Fine-tuning with Mean Absolute Error
+
+Let’s fine-tune the model on a dataset using the Mean Absolute Error
+(MAE).
+
+For that, we simply pass the appropriate string representing the loss
+function to the `finetune_loss` parameter of the `forecast` method.
+
+```python
+timegpt_fcst_finetune_mae_df = nixtla_client.forecast(
+ df=df,
+ h=12,
+ finetune_steps=10,
+ finetune_loss='mae', # Set your desired loss function
+ time_col='timestamp',
+ target_col='value',
+)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+```python
+nixtla_client.plot(
+ df, timegpt_fcst_finetune_mae_df,
+ time_col='timestamp', target_col='value',
+)
+```
+
+
+
+Now, depending on your data, you will use a specific error metric to
+accurately evaluate your forecasting model’s performance.
+
+Below is a non-exhaustive guide on which metric to use depending on your
+use case.
+
+**Mean absolute error (MAE)**
+
+
+
+- Robust to outliers
+- Easy to understand
+- You care equally about all error sizes
+- Same units as your data
+
+**Mean squared error (MSE)**
+
+
+
+- You want to penalize large errors more than small ones
+- Sensitive to outliers
+- Used when large errors must be avoided
+- *Not* the same units as your data
+
+**Root mean squared error (RMSE)**
+
+
+
+- Brings the MSE back to original units of data
+- Penalizes large errors more than small ones
+
+**Mean absolute percentage error (MAPE)**
+
+
+
+- Easy to understand for non-technical stakeholders
+- Expressed as a percentage
+- Heavier penalty on positive errors over negative errors
+- To be avoided if your data has values close to 0 or equal to 0
+
+**Symmmetric mean absolute percentage error (sMAPE)**
+
+
+
+- Fixes bias of MAPE
+- Equally senstitive to over and under forecasting
+- To be avoided if your data has values close to 0 or equal to 0
+
+With TimeGPT, you can choose your loss function during fine-tuning as to
+maximize the model’s performance metric for your particular use case.
+
+Let’s run a small experiment to see how each loss function improves
+their associated metric when compared to the default setting.
+
+```python
+train = df[:-36]
+test = df[-36:]
+```
+
+
+```python
+losses = ['default', 'mae', 'mse', 'rmse', 'mape', 'smape']
+
+test = test.copy()
+
+for loss in losses:
+ preds_df = nixtla_client.forecast(
+ df=train,
+ h=36,
+ finetune_steps=10,
+ finetune_loss=loss,
+ time_col='timestamp',
+ target_col='value')
+
+ preds = preds_df['TimeGPT'].values
+
+ test.loc[:,f'TimeGPT_{loss}'] = preds
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon. This may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon. This may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon. This may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon. This may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon. This may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon. This may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+Great! We have predictions from TimeGPT using all the different loss
+functions. We can evaluate the performance using their associated metric
+and measure the improvement.
+
+```python
+loss_fct_dict = {
+ "mae": mae,
+ "mse": mse,
+ "rmse": rmse,
+ "mape": mape,
+ "smape": smape
+}
+
+pct_improv = []
+
+for loss in losses[1:]:
+ evaluation = loss_fct_dict[f'{loss}'](test, models=['TimeGPT_default', f'TimeGPT_{loss}'], id_col='unique_id', target_col='value')
+ pct_diff = (evaluation['TimeGPT_default'] - evaluation[f'TimeGPT_{loss}']) / evaluation['TimeGPT_default'] * 100
+ pct_improv.append(round(pct_diff, 2))
+```
+
+
+```python
+data = {
+ 'mae': pct_improv[0].values,
+ 'mse': pct_improv[1].values,
+ 'rmse': pct_improv[2].values,
+ 'mape': pct_improv[3].values,
+ 'smape': pct_improv[4].values
+}
+
+metrics_df = pd.DataFrame(data)
+metrics_df.index = ['Metric improvement (%)']
+
+metrics_df
+```
+
+| | mae | mse | rmse | mape | smape |
+|------------------------|------|------|------|-------|-------|
+| Metric improvement (%) | 8.54 | 0.31 | 0.64 | 31.02 | 7.36 |
+
+From the table above, we can see that using a specific loss function
+during fine-tuning will improve its associated error metric when
+compared to the default loss function.
+
+In this example, using the MAE as the loss function improves the metric
+by 8.54% when compared to using the default loss function.
+
+That way, depending on your use case and performance metric, you can use
+the appropriate loss function to maximize the accuracy of the forecasts.
+
diff --git a/nixtla/docs/tutorials/missing_values.html.mdx b/nixtla/docs/tutorials/missing_values.html.mdx
new file mode 100644
index 00000000..6f49fc80
--- /dev/null
+++ b/nixtla/docs/tutorials/missing_values.html.mdx
@@ -0,0 +1,374 @@
+---
+output-file: missing_values.html
+title: Missing Values
+---
+
+
+`TimeGPT` requires time series data that doesn’t have any missing
+values. It is possible to have multiple series that begin and end on
+different dates, but it is essential that each series contains
+uninterrupted data for its given time frame.
+
+In this tutorial, we will show you how to deal with missing values in
+`TimeGPT`.
+
+**Outline**
+
+1. [Load Data](#load-data)
+
+2. [Get Started with TimeGPT](#get-started-with-timegpt)
+
+3. [Visualize Data](#visualize-data)
+
+4. [Fill Missing Values](#fill-missing-values)
+
+5. [Forecast with TimeGPT](#forecast-with-timegpt)
+
+6. [Important Considerations](#important-considerations)
+
+7. [References](#references)
+
+This work is based on skforecast’s [Forecasting Time Series with Missing
+Values](https://cienciadedatos.net/documentos/py46-forecasting-time-series-missing-values)
+tutorial.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/15_missing_values.ipynb)
+
+## Load Data
+
+We will first load the data using `pandas`. This dataset represents the
+daily number of bike rentals in a city. The column names are in Spanish,
+so we will rename them to `ds` for the dates and `y` for the number of
+bike rentals.
+
+```python
+import pandas as pd
+```
+
+
+```python
+df = pd.read_csv('https://raw.githubusercontent.com/JoaquinAmatRodrigo/Estadistica-machine-learning-python/master/data/usuarios_diarios_bicimad.csv')
+df = df[['fecha', 'Usos bicis total día']] # select date and target variable
+df.rename(columns={'fecha': 'ds', 'Usos bicis total día': 'y'}, inplace=True)
+df.head()
+```
+
+| | ds | y |
+|-----|------------|-----|
+| 0 | 2014-06-23 | 99 |
+| 1 | 2014-06-24 | 72 |
+| 2 | 2014-06-25 | 119 |
+| 3 | 2014-06-26 | 135 |
+| 4 | 2014-06-27 | 149 |
+
+For convenience, we will convert the dates to timestamps and assign a
+unique id to the series. Although we only have one series in this
+example, when dealing with multiple series, it is necessary to assign a
+unique id to each one.
+
+```python
+df['ds'] = pd.to_datetime(df['ds'])
+df['unique_id'] = 'id1'
+df = df[['unique_id', 'ds', 'y']]
+```
+
+Now we will separate the data in a training and a test set. We will use
+the last 93 days as the test set.
+
+```python
+train_df = df[:-93]
+test_df = df[-93:]
+```
+
+We will now introduce some missing values in the training set to
+demonstrate how to deal with them. This will be done as in the
+[skforecast](https://cienciadedatos.net/documentos/py46-forecasting-time-series-missing-values)
+tutorial.
+
+```python
+mask = ~((train_df['ds'] >= '2020-09-01') & (train_df['ds'] <= '2020-10-10')) & ~((train_df['ds'] >= '2020-11-08') & (train_df['ds'] <= '2020-12-15'))
+
+train_df_gaps = train_df[mask]
+```
+
+## Get Started with TimeGPT
+
+Before proceeding, we will instantiate the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class, which provides access to all the methods from `TimeGPT`. To do
+this, you will need a Nixtla API key.
+
+```python
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, set the `base_url` argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+To learn more about how to set up your API key, please refer to the
+[Setting Up Your API
+Key](https://docs.nixtla.io/docs/getting-started-setting_up_your_api_key)
+tutorial.
+
+## Visualize Data
+
+We can visualize the data using the `plot` method from the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class. This method has an `engine` argument that allows you to choose
+between different plotting libraries. Default is `matplotlib`, but you
+can also use `plotly` for interactive plots.
+
+```python
+nixtla_client.plot(train_df_gaps)
+```
+
+
+
+Note that there are two gaps in the data: from September 1, 2020, to
+October 10, 2020, and from November 8, 2020, to December 15, 2020. To
+better visualize these gaps, you can use the `max_insample_length`
+argument of the `plot` method or you can simply zoom in on the plot.
+
+```python
+nixtla_client.plot(train_df_gaps, max_insample_length=800)
+```
+
+
+
+Additionally, notice a period from March 16, 2020, to April 21, 2020,
+where the data shows zero rentals. These are not missing values, but
+actual zeros corresponding to the COVID-19 lockdown in the city.
+
+## Fill Missing Values
+
+Before using `TimeGPT`, we need to ensure that:
+
+1. All timestamps from the start date to the end date are present in
+ the data.
+
+2. The target column contains no missing values.
+
+To address the first issue, we will use the `fill_gaps` function from
+[`utilsforecast`](https://nixtlaverse.nixtla.io/utilsforecast/index.html),
+a Python package from Nixtla that provides essential utilities for time
+series forecasting, such as functions for data preprocessing, plotting,
+and evaluation.
+
+The `fill_gaps` function will fill in the missing dates in the data. To
+do this, it requires the following arguments:
+
+- `df`: The DataFrame containing the time series data.
+
+- `freq` (str or int): The frequency of the data.
+
+```python
+from utilsforecast.preprocessing import fill_gaps
+```
+
+
+```python
+print('Number of rows before filling gaps:', len(train_df_gaps))
+train_df_complete = fill_gaps(train_df_gaps, freq='D')
+print('Number of rows after filling gaps:', len(train_df_complete))
+```
+
+``` text
+Number of rows before filling gaps: 2851
+Number of rows after filling gaps: 2929
+```
+
+> NOTE:
+>
+> In this tutorial, the data contains only one time series. However,
+> TimeGPT supports passing multiple series to the model. In this case,
+> none of the time series can have missing values from their individual
+> earliest timestamp until their individual lastest timestamp. If these
+> individual time series have missing values, the user must decide how
+> to fill these gaps for the individual time series. The
+> [`fill_gaps`](https://nixtlaverse.nixtla.io/utilsforecast/preprocessing.html#fill-gaps)
+> function provides a couple of additional arguments to assist with this
+> (refer to the documentation for complete details), namely `start` and
+> `end`
+
+------------------------------------------------------------------------
+
+source
+
+### fill_gaps
+
+> ``` text
+> fill_gaps (df:~DFType, freq:Union[str,int],
+> start:Union[str,int,datetime.date,datetime.datetime]='per_seri
+> e',
+> end:Union[str,int,datetime.date,datetime.datetime]='global',
+> id_col:str='unique_id', time_col:str='ds')
+> ```
+
+*Enforce start and end datetimes for dataframe.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input data |
+| freq | Union | | Series’ frequency |
+| start | Union | per_serie | Initial timestamp for the series. \* ‘per_serie’ uses each serie’s first timestamp \* ‘global’ uses the first timestamp seen in the data \* Can also be a specific timestamp or integer, e.g. ‘2000-01-01’, 2000 or datetime(2000, 1, 1) |
+| end | Union | global | Initial timestamp for the series. \* ‘per_serie’ uses each serie’s last timestamp \* ‘global’ uses the last timestamp seen in the data \* Can also be a specific timestamp or integer, e.g. ‘2000-01-01’, 2000 or datetime(2000, 1, 1) |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestamp. |
+| **Returns** | **DFType** | | **Dataframe with gaps filled.** |
+
+Now we need to decide how to fill the missing values in the target
+column. In this tutorial, we will use interpolation, but it is important
+to consider the specific context of your data when selecting a filling
+strategy. For example, if you are dealing with daily retail data, a
+missing value most likely indicates that there were no sales on that
+day, and you can fill it with zero. Conversely, if you are working with
+hourly temperature data, a missing value probably means that the sensor
+was not functioning, and you might prefer to use interpolation to fill
+the missing values.
+
+```python
+train_df_complete['y'] = train_df_complete['y'].interpolate(method='linear', limit_direction='both')
+
+train_df_complete.isna().sum() # check if there are any missing values
+```
+
+``` text
+unique_id 0
+ds 0
+y 0
+dtype: int64
+```
+
+## Forecast with TimeGPT
+
+We are now ready to use the `forecast` method from the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class. This method requires the following arguments:
+
+- `df`: The DataFrame containing the time series data
+
+- `h`: (int) The forecast horizon. In this case, it is 93 days.
+
+- `model` (str): The model to use. Default is `timegpt-1`, but since
+ the forecast horizon exceeds the frequency of the data (daily), we
+ will use `timegpt-1-long-horizon`. To learn more about this, please
+ refer to the [Forecasting on a Long
+ Horizon](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+ tutorial.
+
+```python
+fcst = nixtla_client.forecast(train_df_complete, h=len(test_df), model='timegpt-1-long-horizon')
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: D
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Querying model metadata...
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon, this may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+We can use the `plot` method to visualize the `TimeGPT` forecast and the
+test set.
+
+```python
+nixtla_client.plot(test_df, fcst)
+```
+
+
+
+Next, we will use the `evaluate` function from `utilsforecast` to
+compute the Mean Average Error (MAE) of the TimeGPT forecast. Before
+proceeding, we need to convert the dates in the forecast to timestamps
+so we can merge them with the test set.
+
+The `evaluate` function requires the following arguments:
+
+- `df`: The DataFrame containing the forecast and the actual values
+ (in the `y` column).
+
+- `metrics` (list): The metrics to be computed.
+
+```python
+from utilsforecast.evaluation import evaluate
+from utilsforecast.losses import mae
+```
+
+
+```python
+fcst['ds'] = pd.to_datetime(fcst['ds'])
+
+result = test_df.merge(fcst, on=['ds', 'unique_id'], how='left')
+result.head()
+```
+
+| | unique_id | ds | y | TimeGPT |
+|-----|-----------|------------|-------|-----------|
+| 0 | id1 | 2022-06-30 | 13468 | 13357.357 |
+| 1 | id1 | 2022-07-01 | 12932 | 12390.052 |
+| 2 | id1 | 2022-07-02 | 9918 | 9778.649 |
+| 3 | id1 | 2022-07-03 | 8967 | 8846.637 |
+| 4 | id1 | 2022-07-04 | 12869 | 11589.071 |
+
+```python
+evaluate(result, metrics=[mae])
+```
+
+| | unique_id | metric | TimeGPT |
+|-----|-----------|--------|-------------|
+| 0 | id1 | mae | 1824.693059 |
+
+## Important Considerations
+
+The key takeaway from this tutorial is that `TimeGPT` requires time
+series data without missing values. This means that:
+
+1. Given the frequency of the data, the timestamps must be continuous,
+ with no gaps between the start and end dates.
+
+2. The data must not contain missing values (NaNs).
+
+We also showed that `utilsforecast` provides a convenient function to
+fill missing dates and that you need to decide how to address the
+missing values. This decision depends on the context of your data, so be
+mindful when selecting a filling strategy, and choose the one you think
+best reflects reality.
+
+Finally, we also demonstrated that `utilsforecast` provides a function
+to evaluate the `TimeGPT` forecast using common accuracy metrics.
+
+**References**
+
+- [Joaquín Amat Rodrigo and Javier Escobar Ortiz (2022). “Exclude
+ covid impact in time series
+ forecasting”](https://www.cienciadedatos.net/documentos/py45-weighted-time-series-forecasting.html)
+
diff --git a/nixtla/docs/tutorials/multiple_series.html.mdx b/nixtla/docs/tutorials/multiple_series.html.mdx
new file mode 100644
index 00000000..44ad36bb
--- /dev/null
+++ b/nixtla/docs/tutorials/multiple_series.html.mdx
@@ -0,0 +1,194 @@
+---
+output-file: multiple_series.html
+title: Multiple series forecasting
+---
+
+
+TimeGPT provides a robust solution for multi-series forecasting, which
+involves analyzing multiple data series concurrently, rather than a
+single one. The tool can be fine-tuned using a broad collection of
+series, enabling you to tailor the model to suit your specific needs or
+tasks.
+
+Note that the forecasts are still univariate. This means that although
+TimeGPT is a global model, it won’t consider the inter-feature
+relationships within the target series. However, TimeGPT does support
+the use of exogenous variables such as categorical variables (e.g.,
+category, brand), numerical variables (e.g., temperature, prices), or
+even special holidays.
+
+Let’s see this in action.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/05_multiple_series.ipynb)
+
+## 1. Import packages
+
+First, we install and import the required packages and initialize the
+Nixtla client.
+
+As always, we start off by intializing an instance of
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient).
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 2. Load the data
+
+The following dataset contains prices of different electricity markets
+in Europe.
+
+Mutliple series are automatically detected in TimeGPT using the
+`unique_id` column. This column contains labels for each series. If
+there are multiple unique values in that column, then it knows it is
+handling a multi-series scneario.
+
+In this particular case, the `unique_id` column contains the value BE,
+DE, FR, JPM, and NP.
+
+```python
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short.csv')
+df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|---------------------|-------|
+| 0 | BE | 2016-12-01 00:00:00 | 72.00 |
+| 1 | BE | 2016-12-01 01:00:00 | 65.80 |
+| 2 | BE | 2016-12-01 02:00:00 | 59.99 |
+| 3 | BE | 2016-12-01 03:00:00 | 50.69 |
+| 4 | BE | 2016-12-01 04:00:00 | 52.58 |
+
+Let’s plot this series using
+[\[`NixtlaClient`\](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)](https://nixtlaverse.nixtla.io/nixtla/nixtla_client.html#nixtlaclient):
+
+```python
+nixtla_client.plot(df)
+```
+
+
+
+## 3. Forecasting Multiple Series
+
+To forecast all series at once, we simply pass the dataframe to the `df`
+argument. TimeGPt will automatically forecast all series.
+
+```python
+timegpt_fcst_multiseries_df = nixtla_client.forecast(df=df, h=24, level=[80, 90])
+timegpt_fcst_multiseries_df.head()
+```
+
+``` text
+INFO:nixtlats.nixtla_client:Validating inputs...
+INFO:nixtlats.nixtla_client:Preprocessing dataframes...
+INFO:nixtlats.nixtla_client:Inferred freq: H
+INFO:nixtlats.nixtla_client:Restricting input...
+INFO:nixtlats.nixtla_client:Calling Forecast Endpoint...
+```
+
+| | unique_id | ds | TimeGPT | TimeGPT-lo-90 | TimeGPT-lo-80 | TimeGPT-hi-80 | TimeGPT-hi-90 |
+|----|----|----|----|----|----|----|----|
+| 0 | BE | 2016-12-31 00:00:00 | 46.151176 | 36.660478 | 38.337019 | 53.965334 | 55.641875 |
+| 1 | BE | 2016-12-31 01:00:00 | 42.426598 | 31.602231 | 33.976724 | 50.876471 | 53.250964 |
+| 2 | BE | 2016-12-31 02:00:00 | 40.242889 | 30.439970 | 33.634985 | 46.850794 | 50.045809 |
+| 3 | BE | 2016-12-31 03:00:00 | 38.265339 | 26.841481 | 31.022093 | 45.508585 | 49.689197 |
+| 4 | BE | 2016-12-31 04:00:00 | 36.618801 | 18.541384 | 27.981346 | 45.256256 | 54.696218 |
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+```python
+nixtla_client.plot(df, timegpt_fcst_multiseries_df, max_insample_length=365, level=[80, 90])
+```
+
+
+
+From the figure above, we can see that the model effectively generated
+predictions for each unique series in the dataset.
+
+## Historical forecast
+
+You can also compute prediction intervals for historical forecasts
+adding the `add_history=True`.
+
+To specify the confidence interval, we use the `level` argument. Here,
+we pass the list `[80, 90]`. This will compute a 80% and 90% confidence
+interval.
+
+```python
+timegpt_fcst_multiseries_with_history_df = nixtla_client.forecast(df=df, h=24, level=[80, 90], add_history=True)
+timegpt_fcst_multiseries_with_history_df.head()
+```
+
+``` text
+INFO:nixtlats.nixtla_client:Validating inputs...
+INFO:nixtlats.nixtla_client:Preprocessing dataframes...
+INFO:nixtlats.nixtla_client:Inferred freq: H
+INFO:nixtlats.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtlats.nixtla_client:Calling Historical Forecast Endpoint...
+```
+
+| | unique_id | ds | TimeGPT | TimeGPT-lo-80 | TimeGPT-lo-90 | TimeGPT-hi-80 | TimeGPT-hi-90 |
+|----|----|----|----|----|----|----|----|
+| 0 | BE | 2016-12-06 00:00:00 | 55.756332 | 42.066476 | 38.185593 | 69.446188 | 73.327072 |
+| 1 | BE | 2016-12-06 01:00:00 | 52.820206 | 39.130350 | 35.249466 | 66.510062 | 70.390946 |
+| 2 | BE | 2016-12-06 02:00:00 | 46.851070 | 33.161214 | 29.280331 | 60.540926 | 64.421810 |
+| 3 | BE | 2016-12-06 03:00:00 | 50.640892 | 36.951036 | 33.070152 | 64.330748 | 68.211632 |
+| 4 | BE | 2016-12-06 04:00:00 | 52.420410 | 38.730554 | 34.849670 | 66.110266 | 69.991150 |
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+```python
+nixtla_client.plot(
+ df,
+ timegpt_fcst_multiseries_with_history_df.groupby('unique_id').tail(365 + 24),
+ max_insample_length=365,
+ level=[80, 90],
+)
+```
+
+
+
+In the figure above, we now see the historical predictions made by
+TimeGPT for each series, along with the 80% and 90% confidence
+intervals.
+
diff --git a/nixtla/docs/tutorials/reusing_finetuned_models.html.mdx b/nixtla/docs/tutorials/reusing_finetuned_models.html.mdx
new file mode 100644
index 00000000..6af31938
--- /dev/null
+++ b/nixtla/docs/tutorials/reusing_finetuned_models.html.mdx
@@ -0,0 +1,224 @@
+---
+output-file: reusing_finetuned_models.html
+title: Re-using fine-tuned models
+---
+
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/061_reusing_finetuned_models.ipynb.ipynb)
+
+## 1. Import packages
+
+First, we import the required packages and initialize the Nixtla client
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+from utilsforecast.losses import rmse
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ["NIXTLA_API_KEY"]
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+## 2. Load data
+
+```python
+df = pd.read_parquet('https://datasets-nixtla.s3.amazonaws.com/m4-hourly.parquet')
+
+h = 48
+valid = df.groupby('unique_id', observed=True).tail(h)
+train = df.drop(valid.index)
+train.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|-----|-------|
+| 0 | H1 | 1 | 605.0 |
+| 1 | H1 | 2 | 586.0 |
+| 2 | H1 | 3 | 586.0 |
+| 3 | H1 | 4 | 559.0 |
+| 4 | H1 | 5 | 511.0 |
+
+## 3. Zero-shot forecast
+
+We can try forecasting without any finetuning to see how well TimeGPT
+does.
+
+```python
+fcst_kwargs = {'df': train, 'freq': 1, 'model': 'timegpt-1-long-horizon'}
+fcst = nixtla_client.forecast(h=h, **fcst_kwargs)
+zero_shot_eval = evaluate(fcst.merge(valid), metrics=[rmse], agg_fn='mean')
+zero_shot_eval
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Querying model metadata...
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon, this may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| | metric | TimeGPT |
+|-----|--------|-------------|
+| 0 | rmse | 1504.474342 |
+
+## 4. Fine-tune
+
+We can now fine-tune TimeGPT a little and save our model for later use.
+We can define the ID that we want that model to have by providing it
+through `output_model_id`.
+
+```python
+first_model_id = 'my-first-finetuned-model'
+nixtla_client.finetune(output_model_id=first_model_id, **fcst_kwargs)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Calling Fine-tune Endpoint...
+```
+
+``` text
+'my-first-finetuned-model'
+```
+
+We can now forecast using this fine-tuned model by providing its ID
+through the `finetuned_model_id` argument.
+
+```python
+first_finetune_fcst = nixtla_client.forecast(h=h, finetuned_model_id=first_model_id, **fcst_kwargs)
+first_finetune_eval = evaluate(first_finetune_fcst.merge(valid), metrics=[rmse], agg_fn='mean')
+zero_shot_eval.merge(first_finetune_eval, on=['metric'], suffixes=('_zero_shot', '_first_finetune'))
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon, this may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| | metric | TimeGPT_zero_shot | TimeGPT_first_finetune |
+|-----|--------|-------------------|------------------------|
+| 0 | rmse | 1504.474342 | 1472.024619 |
+
+We can see the error was reduced.
+
+## 5. Further fine-tune
+
+We can now take this model and fine-tune it a bit further by using the
+[`NixtlaClient.finetune`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient.finetune)
+method but providing our already fine-tuned model as
+`finetuned_model_id`, which will take that model and fine-tune it a bit
+more. We can also change the fine-tuning settings, like using
+`finetune_depth=3`, for example.
+
+```python
+second_model_id = nixtla_client.finetune(finetuned_model_id=first_model_id, finetune_depth=3, **fcst_kwargs)
+second_model_id
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Calling Fine-tune Endpoint...
+```
+
+``` text
+'468b13fb-4b26-447a-bd87-87a64b50d913'
+```
+
+Since we didn’t provide `output_model_id` this time, it got assigned an
+UUID.
+
+We can now use this model to forecast.
+
+```python
+second_finetune_fcst = nixtla_client.forecast(h=h, finetuned_model_id=second_model_id, **fcst_kwargs)
+second_finetune_eval = evaluate(second_finetune_fcst.merge(valid), metrics=[rmse], agg_fn='mean')
+first_finetune_eval.merge(second_finetune_eval, on=['metric'], suffixes=('_first_finetune', '_second_finetune'))
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon, this may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| | metric | TimeGPT_first_finetune | TimeGPT_second_finetune |
+|-----|--------|------------------------|-------------------------|
+| 0 | rmse | 1472.024619 | 1435.365211 |
+
+We can see the error was reduced a bit more.
+
+## 6. Listing fine-tuned models
+
+We can list our fine-tuned models with the
+[`NixtlaClient.finetuned_models`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient.finetuned_models)
+method.
+
+```python
+finetuned_models = nixtla_client.finetuned_models()
+finetuned_models
+```
+
+``` text
+[FinetunedModel(id='468b13fb-4b26-447a-bd87-87a64b50d913', created_at=datetime.datetime(2024, 12, 30, 17, 57, 31, 241455, tzinfo=TzInfo(UTC)), created_by='user', base_model_id='my-first-finetuned-model', steps=10, depth=3, loss='default', model='timegpt-1-long-horizon', freq='MS'),
+ FinetunedModel(id='my-first-finetuned-model', created_at=datetime.datetime(2024, 12, 30, 17, 57, 16, 978907, tzinfo=TzInfo(UTC)), created_by='user', base_model_id='None', steps=10, depth=1, loss='default', model='timegpt-1-long-horizon', freq='MS')]
+```
+
+While that representation may be useful for programmatic use, in this
+exploratory setting it’s nicer to see them as a dataframe, which we can
+get by providing `as_df=True`.
+
+```python
+nixtla_client.finetuned_models(as_df=True)
+```
+
+| | id | created_at | created_by | base_model_id | steps | depth | loss | model | freq |
+|----|----|----|----|----|----|----|----|----|----|
+| 0 | 468b13fb-4b26-447a-bd87-87a64b50d913 | 2024-12-30 17:57:31.241455+00:00 | user | my-first-finetuned-model | 10 | 3 | default | timegpt-1-long-horizon | MS |
+| 1 | my-first-finetuned-model | 2024-12-30 17:57:16.978907+00:00 | user | None | 10 | 1 | default | timegpt-1-long-horizon | MS |
+
+We can seee that the `base_model_id` of our second model is our first
+model, along with other metadata.
+
+## 7. Deleting fine-tuned models
+
+In order to keep things organized, and since there’s a limit of 50
+fine-tuned models, you can delete models that weren’t so promising to
+make room for more experiments. For example, we can delete our first
+finetuned model. Note that even though it was used as the base for our
+second model, they’re saved independently so removing it won’t affect
+our second model, except for the dangling metadata.
+
+```python
+nixtla_client.delete_finetuned_model(first_model_id)
+```
+
+``` text
+True
+```
+
+We can verify that our first model model doesn’t show up anymore in our
+available models.
+
+```python
+nixtla_client.finetuned_models(as_df=True)
+```
+
+| | id | created_at | created_by | base_model_id | steps | depth | loss | model | freq |
+|----|----|----|----|----|----|----|----|----|----|
+| 0 | 468b13fb-4b26-447a-bd87-87a64b50d913 | 2024-12-30 17:57:31.241455+00:00 | user | my-first-finetuned-model | 10 | 3 | default | timegpt-1-long-horizon | MS |
+
diff --git a/nixtla/docs/tutorials/shap_values.html.mdx b/nixtla/docs/tutorials/shap_values.html.mdx
new file mode 100644
index 00000000..8159cdf2
--- /dev/null
+++ b/nixtla/docs/tutorials/shap_values.html.mdx
@@ -0,0 +1,329 @@
+---
+output-file: shap_values.html
+title: SHAP Values for TimeGPT and TimeGEN
+---
+
+
+SHAP (SHapley Additive exPlanation) values use game theory to explain
+the output of any machine learning model. It allows us to explore in
+detail how exogenous features impact the final forecast, both at a
+single forecast step or over the entire horizon.
+
+When you forecast with exogenous features, you can access the SHAP
+values for all series at each prediction step, and use the popular
+[shap](https://shap.readthedocs.io/en/latest/) Python package to make
+different plots and explain the impact of the features.
+
+This tutorial assumes knowledge on forecasting with exogenous features,
+so make sure to read our tutorial on [exogenous
+variables](https://docs.nixtla.io/docs/tutorials-exogenous_variables).
+Also, the `shap` package must be installed separately as it is not a
+dependency of `nixtla`.
+
+`shap` can be installed from either
+[PyPI](https://pypi.org/project/shap/) or
+[conda-forge](https://anaconda.org/conda-forge/shap):
+
+``` text
+pip install shap
+
+or
+
+conda install -c conda-forge shap
+```
+
+For the official documentation of SHAP, visit:
+https://shap.readthedocs.io/en/latest/
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/21_shap_values.ipynb)
+
+## 1. Import packages
+
+First, we import the required packages and initialize the Nixtla client.
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 2. Load data
+
+In this example on SHAP values, we will use exogenous variables (also
+known as covariates) to improve the accuracy of electricity market
+forecasts. We’ll work with a well-known dataset called `EPF`, which is
+publicly accessible [here](https://zenodo.org/records/4624805).
+
+This dataset includes data from five different electricity markets, each
+with unique price dynamics, such as varying frequencies and occurrences
+of negative prices, zeros, and price spikes. Since electricity prices
+are influenced by exogenous factors, each dataset also contains two
+additional time series: day-ahead forecasts of two significant exogenous
+factors specific to each market.
+
+For simplicity, we will focus on the Belgian electricity market (BE).
+This dataset includes hourly prices (`y`), day-ahead forecasts of load
+(`Exogenous1`), and electricity generation (`Exogenous2`). It also
+includes one-hot encoding to indicate whether a specific date is a
+specific day of the week. Eg.: Monday (`day_0 = 1`), a Tuesday
+(`day_1 = 1`), and so on.
+
+If your data depends on exogenous factors or covariates such as prices,
+discounts, special holidays, weather, etc., you can follow a similar
+structure.
+
+```python
+market = "BE"
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short-with-ex-vars.csv')
+df.head()
+```
+
+| | unique_id | ds | y | Exogenous1 | Exogenous2 | day_0 | day_1 | day_2 | day_3 | day_4 | day_5 | day_6 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | BE | 2016-10-22 00:00:00 | 70.00 | 57253.0 | 49593.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 1 | BE | 2016-10-22 01:00:00 | 37.10 | 51887.0 | 46073.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 2 | BE | 2016-10-22 02:00:00 | 37.10 | 51896.0 | 44927.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 3 | BE | 2016-10-22 03:00:00 | 44.75 | 48428.0 | 44483.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 4 | BE | 2016-10-22 04:00:00 | 37.10 | 46721.0 | 44338.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+
+## 3. Forecasting electricity prices using exogenous variables
+
+To produce forecasts we also have to add the future values of the
+exogenous variables.
+
+If your forecast depends on other variables, it is important to ensure
+that those variables are available at the time of forecasting. In this
+example, we know that the price of electricity depends on the demand
+(`Exogenous1`) and the quantity produced (`Exogenous2`). Thus, we need
+to have those future values available at the time of forecasting. If
+those values were not available, we can always [use TimeGPT to forecast
+them](https://docs.nixtla.io/docs/tutorials-exogenous_variables).
+
+Here, we read a dataset that contains the future values of our features.
+In this case, we want to predict 24 steps ahead, therefore each
+`unique_id` will have 24 observations.
+
+> **Important**
+>
+> If you want to use exogenous variables when forecasting with TimeGPT,
+> you need to have the future values of those exogenous variables too.
+
+```python
+future_ex_vars_df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short-future-ex-vars.csv')
+future_ex_vars_df.head()
+```
+
+| | unique_id | ds | Exogenous1 | Exogenous2 | day_0 | day_1 | day_2 | day_3 | day_4 | day_5 | day_6 |
+|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | BE | 2016-12-31 00:00:00 | 70318.0 | 64108.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 1 | BE | 2016-12-31 01:00:00 | 67898.0 | 62492.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 2 | BE | 2016-12-31 02:00:00 | 68379.0 | 61571.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 3 | BE | 2016-12-31 03:00:00 | 64972.0 | 60381.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+| 4 | BE | 2016-12-31 04:00:00 | 62900.0 | 60298.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 |
+
+Let’s call the `forecast` method, adding this information. To access the
+SHAP values, we also need to specify `feature_contributions=True` in the
+`forecast` method.
+
+```python
+timegpt_fcst_ex_vars_df = nixtla_client.forecast(df=df,
+ X_df=future_ex_vars_df,
+ h=24,
+ level=[80, 90],
+ feature_contributions=True)
+timegpt_fcst_ex_vars_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: h
+INFO:nixtla.nixtla_client:Querying model metadata...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Using future exogenous features: ['Exogenous1', 'Exogenous2', 'day_0', 'day_1', 'day_2', 'day_3', 'day_4', 'day_5', 'day_6']
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| | unique_id | ds | TimeGPT | TimeGPT-hi-80 | TimeGPT-hi-90 | TimeGPT-lo-80 | TimeGPT-lo-90 |
+|----|----|----|----|----|----|----|----|
+| 0 | BE | 2016-12-31 00:00:00 | 51.632830 | 61.598820 | 66.088295 | 41.666843 | 37.177372 |
+| 1 | BE | 2016-12-31 01:00:00 | 45.750877 | 54.611988 | 60.176445 | 36.889767 | 31.325312 |
+| 2 | BE | 2016-12-31 02:00:00 | 39.650543 | 46.256210 | 52.842808 | 33.044876 | 26.458277 |
+| 3 | BE | 2016-12-31 03:00:00 | 34.000072 | 44.015310 | 47.429000 | 23.984835 | 20.571144 |
+| 4 | BE | 2016-12-31 04:00:00 | 33.785370 | 43.140503 | 48.581240 | 24.430239 | 18.989498 |
+
+## 4. Extract SHAP values
+
+Now that we have made predictions using exogenous features, we can then
+extract the SHAP values to understand their relevance using the
+`feature_contributions` attribute of the client. This returns a
+DataFrame containing the SHAP values and base values for each series, at
+each step in the horizon.
+
+```python
+shap_df = nixtla_client.feature_contributions
+shap_df = shap_df.query("unique_id == @market")
+shap_df.head()
+```
+
+| | unique_id | ds | TimeGPT | Exogenous1 | Exogenous2 | day_0 | day_1 | day_2 | day_3 | day_4 | day_5 | day_6 | base_value |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | BE | 2016-12-31 00:00:00 | 51.632830 | 27.929638 | -16.363607 | 0.081917 | -1.883555 | 0.346484 | -0.228611 | 0.424167 | -3.411662 | 1.113910 | 43.624146 |
+| 1 | BE | 2016-12-31 01:00:00 | 45.750877 | 17.678530 | -12.240089 | -0.758545 | -0.077536 | -0.160390 | -0.309567 | 0.871469 | -3.927268 | 1.218714 | 43.455560 |
+| 2 | BE | 2016-12-31 02:00:00 | 39.650543 | 21.632694 | -21.400244 | -0.926842 | -0.470276 | -0.022417 | -0.225389 | 0.220258 | -3.927268 | 1.145736 | 43.624290 |
+| 3 | BE | 2016-12-31 03:00:00 | 34.000072 | 13.879354 | -20.681124 | -0.114050 | -0.488141 | 0.048164 | -0.126627 | 0.200692 | -3.400485 | 1.144959 | 43.537330 |
+| 4 | BE | 2016-12-31 04:00:00 | 33.785370 | 13.465129 | -20.619830 | -0.036112 | -0.470496 | 0.048375 | -0.126627 | 0.200692 | -3.400485 | 1.144959 | 43.579760 |
+
+In the Dataframe above, we can see that we have the SHAP values at every
+forecasting step, as well as the prediction from TimeGPT and the base
+value. Note that the base value is the prediction of the model if
+exogenous features were unknown.
+
+Therefore, the forecast from TimeGPT is equal to the sum of the base
+value and the SHAP values of each exogenous feature in a given row.
+
+## 5. Make plots using `shap`
+
+Now that we have access to SHAP values we can use the `shap` package to
+make any plots that we want.
+
+### 5.1 Bar plot
+
+Here, let’s make bar plots for each series and their features, so we can
+see which features impacts the predictions the most.
+
+```python
+import shap
+import matplotlib.pyplot as plt
+
+shap_columns = shap_df.columns.difference(['unique_id', 'ds', 'TimeGPT', 'base_value'])
+shap_values = shap_df[shap_columns].values # SHAP values matrix
+base_values = shap_df['base_value'].values # Extract base values
+features = shap_columns # Feature names
+
+# Create a SHAP values object
+shap_obj = shap.Explanation(values=shap_values, base_values=base_values, feature_names=features)
+
+# Plot the bar plot for SHAP values
+shap.plots.bar(shap_obj, max_display=len(features), show=False)
+plt.title(f'SHAP values for {market}')
+plt.show()
+```
+
+
+
+The plot above shows the average SHAP values for each feature across the
+entire horizon.
+
+Here, we see that `Exogenous1` is the most important feature, as it has
+the largest average contribution. Remember that it designates the
+expected energy demand, so we can see that this variable has a large
+impact on the final prediction. On the other hand, `day_2` is the least
+important feature, since it has the lowest value.
+
+### 5.2 Waterfall plot
+
+Now, let’s see how we can make a waterfall plot to explore the the
+impact of features at a single prediction step. The code below selects a
+specific date. Of course, this can be modified for any series or date.
+
+```python
+selected_ds = shap_df['ds'].min()
+
+filtered_df = shap_df[shap_df['ds'] == selected_ds]
+
+shap_values = filtered_df[shap_columns].values.flatten()
+base_value = filtered_df['base_value'].values[0]
+features = shap_columns
+
+shap_obj = shap.Explanation(values=shap_values, base_values=base_value, feature_names=features)
+
+shap.plots.waterfall(shap_obj, show=False)
+plt.title(f'Waterfall Plot: {market}, date: {selected_ds}')
+plt.show()
+```
+
+
+
+In the waterfall plot above, we can explore in more detail a single
+prediction. Here, we study the final prediction for the start of
+December 31th, 2016.
+
+The x-axis represents the value of our series. At the bottom, we see
+`E[f(X)]` which represents the baseline value (the predicted value if
+exogenous features were unknown).
+
+Then, we see how each feature has impacted the final forecast. Features
+like `day_3`, `day_1`, `day_5`, `Exogenous2` all push the forecast to
+the left (smaller value). On the other hand, `day_0`, `day_2`, `day_4`,
+`day_6` and `Exogenous1` push it to the right (larger value).
+
+Let’s think about this for a moment. In the introduction, we stated that
+`Exogenous1` represents electricity load, whereas `Exogenous2`
+represents electricity generation. \* `Exogenous1`, the electricity
+load, adds positively to the overall prediction. This seems reasonable:
+if we expect a higher demand, we might expect the price to go up. \*
+`Exogenous2`, on the other hand, adds negatively to the overall
+prediction. This seems reasonable too: if there’s a higher electricity
+generation, we expect the price to be lower. Hence, a negative
+contribution to the forecast for `Exogenous2`.
+
+At the top right, we see f(x) which is the final output of the model
+after considering the impact of the exogenous features. Notice that this
+value corresponds to the final prediction from TimeGPT.
+
+### 5.3 Heatmap
+
+We can also do a heatmap plot to see how each feature impacts the final
+prediction. Here, we only need to select a specific series.
+
+```python
+shap_columns = shap_df.columns.difference(['unique_id', 'ds', 'TimeGPT', 'base_value'])
+shap_values = shap_df[shap_columns].values
+feature_names = shap_columns.tolist()
+
+shap_obj = shap.Explanation(values=shap_values, feature_names=feature_names)
+
+shap.plots.heatmap(shap_obj, show=False)
+plt.title(f'SHAP Heatmap (Unique ID: NP)')
+plt.show()
+```
+
+
+
+With the heatmap, we basically see a breakdown of each each feature
+impacts the final predciton at each timestep.
+
+On the x-axis, we have the number of instances, which corresponds to the
+number of prediction steps (24 in this case, since our horizon is set to
+24h). On the y-axis, we have the name of the exogenous features.
+
+First, notice that the ordering is the same as in the bar plot, where
+`Exogenous1` is the most important, and `day_6` is the least important.
+
+Then, the color of the heatmap indiciates if the feature tends to
+increase of decrease the final prediction at each forecasting step. For
+example, `Exogenous1` always increases predictions across all 24 hours
+in the forecast horizon.
+
+We also see that all days except `day_5` do not have a very large impact
+at any forecasting step, indicating that they barely impacting the final
+prediction.
+
+Ultimately, the `feature_contributions` attribute gives you access to
+all the necessary information to explain the impact of exogenous
+features using the `shap` package.
+
diff --git a/nixtla/docs/tutorials/special_topics.html.mdx b/nixtla/docs/tutorials/special_topics.html.mdx
new file mode 100644
index 00000000..a2f796b9
--- /dev/null
+++ b/nixtla/docs/tutorials/special_topics.html.mdx
@@ -0,0 +1,47 @@
+---
+output-file: special_topics.html
+title: Special topics
+---
+
+
+`TimeGPT` is a robust foundation model for time series forecasting, with
+advanced capabilities such as hierarchical and bounded forecasts. To
+fully leverage the power of `TimeGPT`, there are specific situations
+that require special consideration, such as dealing with irregular
+timestamps or handling datasets with missing values.
+
+In this section, we will cover these special topics.
+
+### What You Will Learn
+
+1. **[Irregular
+ Timestamps](https://docs.nixtla.io/docs/capabilities-forecast-irregular_timestamps)**
+
+ - Learn how to deal with irregular timestamps for correct usage of
+ `TimeGPT`.
+
+2. **[Bounded
+ Forecasts](https://docs.nixtla.io/docs/tutorials-bounded_forecasts)**
+
+ - Explore `TimeGPT`’s capability to make forecasts within a
+ specified range, ideal for applications where outcomes are
+ bounded.
+
+3. **[Hierarchical
+ Forecasts](https://docs.nixtla.io/docs/tutorials-hierarchical_forecasting)**
+
+ - Understand how to use `TimeGPT` to make coherent predictions at
+ various levels of aggregation.
+
+4. **[Missing
+ Values](https://docs.nixtla.io/docs/tutorials-missing_values)**
+
+ - Learn how to address missing values within your time series data
+ effectively using `TimeGPT`.
+
+5. **[Improve Forecast
+ Accuracy](https://docs.nixtla.io/docs/tutorials-improve_forecast_accuracy_with_timegpt)**
+
+ - Discover multiple techniques to boost forecast accuracy when
+ working with `TimeGPT`.
+
diff --git a/nixtla/docs/tutorials/temporalhierarchical.html.mdx b/nixtla/docs/tutorials/temporalhierarchical.html.mdx
new file mode 100644
index 00000000..c6fab15e
--- /dev/null
+++ b/nixtla/docs/tutorials/temporalhierarchical.html.mdx
@@ -0,0 +1,298 @@
+---
+output-file: temporalhierarchical.html
+title: Temporal Hierarchical Forecasting with TimeGPT
+---
+
+
+In this notebook, we demonstrate how to use TimeGPT for temporal
+hierarchical forecasting. We will use a dataset that has an hourly
+frequency, and we create forecasts with TimeGPT for both the hourly and
+the 2-hourly frequency level. The latter constitutes the timeseries when
+it is aggregated across 2-hour windows. Subsequently, we can use
+temporal reconciliation techniques to improve the forecasting
+performance of TimeGPT.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/23_temporalhierarchical.ipynb)
+
+## 1. Load and Process Data
+
+```python
+import numpy as np
+import pandas as pd
+
+from utilsforecast.evaluation import evaluate
+from utilsforecast.plotting import plot_series
+from utilsforecast.losses import mae, rmse
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+
+```python
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/electricity-short-with-ex-vars.csv')
+df['ds'] = pd.to_datetime(df['ds'])
+df_sub = df.query('unique_id == "DE"')
+```
+
+
+```python
+df_train = df_sub.query('ds < "2017-12-29"')
+df_test = df_sub.query('ds >= "2017-12-29"')
+df_train.shape, df_test.shape
+```
+
+``` text
+((1632, 12), (48, 12))
+```
+
+```python
+plot_series(df_train[['unique_id','ds','y']][-200:], forecasts_df= df_test[['unique_id','ds','y']].rename(columns={'y': 'test'}))
+```
+
+
+
+## 2. Temporal aggregation
+
+We are interested in generating forecasts for the hourly and 2-hourly
+windows. We can generate these forecasts using TimeGPT. After generating
+these forecasts, we make use of hierarchical forecasting techniques to
+improve the accuracy of each forecast.
+
+We first define the temporal aggregation spec. The spec is a dictionary
+in which the keys are the name of the aggregation and the value is the
+amount of bottom-level timesteps that should be aggregated in that
+aggregation.
+
+In this example, we choose a temporal aggregation of a 2-hour period and
+a 1-hour period (the bottom level).
+
+```python
+spec_temporal = { "2-hour-period": 2, "1-hour-period": 1}
+```
+
+We next compute the temporally aggregated train- and test sets using the
+`aggregate_temporal` function from `hierarchicalforecast`. Note that we
+have different aggregation matrices `S` for the train- and test set, as
+the test set contains temporal hierarchies that are not included in the
+train set.
+
+```python
+from hierarchicalforecast.utils import aggregate_temporal
+```
+
+
+```python
+Y_train, S_train, tags_train = aggregate_temporal(df=df_train[['unique_id','ds','y']], spec=spec_temporal)
+Y_test, S_test, tags_test = aggregate_temporal(df=df_test[['unique_id','ds','y']], spec=spec_temporal)
+```
+
+`Y_train` contains our training data, for both 1-hour and 2-hour
+periods. For example, if we look at the first two timestamps of the
+training data, we have a 2-hour period ending at 2017-10-22 01:00, and
+two 1-hour periods, the first ending at 2017-10-22 00:00, and the second
+at 2017-10-22 01:00, the latter corresponding to when the first 2-hour
+period ends.
+
+Also, the ground truth value `y` of the first 2-hour period is 38.13,
+which is equal to the sum of the first two 1-hour periods (19.10 +
+19.03). This showcases how the higher frequency `1-hour-period` has been
+aggregated into the `2-hour-period` frequency.
+
+```python
+Y_train.query("ds <= '2017-10-22 01:00:00'")
+```
+
+| | temporal_id | unique_id | ds | y |
+|-----|-----------------|-----------|---------------------|-------|
+| 0 | 2-hour-period-1 | DE | 2017-10-22 01:00:00 | 38.13 |
+| 816 | 1-hour-period-1 | DE | 2017-10-22 00:00:00 | 19.10 |
+| 817 | 1-hour-period-2 | DE | 2017-10-22 01:00:00 | 19.03 |
+
+The aggregation matrices `S_train` and `S_test` detail how the lowest
+temporal granularity (hour) can be aggregated into the 2-hour periods.
+For example, the first 2-hour period, named `2-hour-period-1`, can be
+constructed by summing the first two hour-periods, `1-hour-period-1` and
+`1-hour-period-2` - which we also verified above in our inspection of
+`Y_train`.
+
+```python
+S_train.iloc[:5, :5]
+```
+
+| | temporal_id | 1-hour-period-1 | 1-hour-period-2 | 1-hour-period-3 | 1-hour-period-4 |
+|----|----|----|----|----|----|
+| 0 | 2-hour-period-1 | 1.0 | 1.0 | 0.0 | 0.0 |
+| 1 | 2-hour-period-2 | 0.0 | 0.0 | 1.0 | 1.0 |
+| 2 | 2-hour-period-3 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 3 | 2-hour-period-4 | 0.0 | 0.0 | 0.0 | 0.0 |
+| 4 | 2-hour-period-5 | 0.0 | 0.0 | 0.0 | 0.0 |
+
+### 3b. Computing base forecasts
+
+Now, we need to compute base forecasts for each temporal aggregation.
+The following cell computes the **base forecasts** for each temporal
+aggregation in `Y_train` using TimeGPT.
+
+Note that both frequency and horizon are different for each temporal
+aggregation. In this example, the lowest level has a hourly frequency,
+and a horizon of `48`. The `2-hourly-period` aggregation thus has a
+2-hourly frequency with a horizon of `24`.
+
+```python
+Y_hats = []
+id_cols = ["unique_id", "temporal_id", "ds", "y"]
+# We will train a model for each temporal level
+for level, temporal_ids_train in tags_train.items():
+ # Filter the data for the level
+ Y_level_train = Y_train.query("temporal_id in @temporal_ids_train")
+ temporal_ids_test = tags_test[level]
+ Y_level_test = Y_test.query("temporal_id in @temporal_ids_test")
+ # For each temporal level we have a different frequency and forecast horizon
+ freq_level = pd.infer_freq(Y_level_train["ds"].unique())
+ horizon_level = Y_level_test["ds"].nunique()
+ # Train a model and create forecasts
+ Y_hat_level = nixtla_client.forecast(df=Y_level_train[["ds", "unique_id", "y"]], h=horizon_level)
+ # Add the test set to the forecast
+ Y_hat_level = Y_hat_level.merge(Y_level_test, on=["ds", "unique_id"], how="left")
+ # Put cols in the right order (for readability)
+ Y_hat_cols = id_cols + [col for col in Y_hat_level.columns if col not in id_cols]
+ Y_hat_level = Y_hat_level[Y_hat_cols]
+ # Append the forecast to the list
+ Y_hats.append(Y_hat_level)
+
+Y_hat = pd.concat(Y_hats, ignore_index=True)
+```
+
+Observe that `Y_hat` contains all the forecasts but they are not
+coherent with each other. For example, consider the forecasts for the
+first time period of both frequencies.
+
+```python
+Y_hat.query("temporal_id in ['2-hour-period-1', '1-hour-period-1', '1-hour-period-2']")
+```
+
+| | unique_id | temporal_id | ds | y | TimeGPT |
+|-----|-----------|-----------------|---------------------|-------|-----------|
+| 0 | DE | 2-hour-period-1 | 2017-12-29 01:00:00 | 10.45 | 16.949448 |
+| 24 | DE | 1-hour-period-1 | 2017-12-29 00:00:00 | 9.73 | -0.241489 |
+| 25 | DE | 1-hour-period-2 | 2017-12-29 01:00:00 | 0.72 | -3.456482 |
+
+The ground truth value `y` for the first 2-hour period is 10.45, and the
+sum of the ground truth values for the first two 1-hour periods is
+(9.73 + 0.72) = 10.45. Hence, these values are coherent with each other.
+
+However, the forecast for the first 2-hour period is 16.95, but the sum
+of the forecasts for the first two 1-hour periods is -3.69. Hence, these
+forecasts are clearly not coherent with each other.
+
+We will use reconciliation techniques to make these forecasts better
+coherent with each other and improve their accuracy.
+
+### 3c. Reconcile forecasts
+
+We can use the `HierarchicalReconciliation` class to reconcile the
+forecasts. In this example we use `MinTrace`. Note that we have to set
+`temporal=True` in the `reconcile` function.
+
+```python
+from hierarchicalforecast.methods import MinTrace
+from hierarchicalforecast.core import HierarchicalReconciliation
+```
+
+
+```python
+reconcilers = [
+ MinTrace(method="wls_struct"),
+]
+hrec = HierarchicalReconciliation(reconcilers=reconcilers)
+Y_rec = hrec.reconcile(Y_hat_df=Y_hat, S=S_test, tags=tags_test, temporal=True)
+```
+
+## 4. Evaluation
+
+The `HierarchicalForecast` package includes the `evaluate` function to
+evaluate the different hierarchies.
+
+We evaluate the temporally aggregated forecasts *across all temporal
+aggregations*.
+
+```python
+import hierarchicalforecast.evaluation as hfe
+from utilsforecast.losses import mae
+```
+
+
+```python
+evaluation = hfe.evaluate(df = Y_rec.drop(columns = 'unique_id'),
+ tags = tags_test,
+ metrics = [mae],
+ id_col='temporal_id')
+
+numeric_cols = evaluation.select_dtypes(include="number").columns
+evaluation[numeric_cols] = evaluation[numeric_cols].map('{:.3}'.format).astype(np.float64)
+```
+
+
+```python
+evaluation
+```
+
+| | level | metric | TimeGPT | TimeGPT/MinTrace_method-wls_struct |
+|-----|---------------|--------|---------|------------------------------------|
+| 0 | 2-hour-period | mae | 25.2 | 12.00 |
+| 1 | 1-hour-period | mae | 18.5 | 6.16 |
+| 2 | Overall | mae | 20.8 | 8.12 |
+
+As we can see, we improved performance of TimeGPT’s predictions both for
+the 2-hour period and for the 1-hour period, as both levels see a
+significant reduction in MAE and RMSE.
+
+Visually, we can also verify the forecast is better after using
+reconciliation techniques. For the 1-hour-period forecasts:
+
+```python
+plot_series(Y_train.query("temporal_id in @tags_train['1-hour-period']")[["y", "ds", "unique_id"]].iloc[-100:], forecasts_df=Y_rec.query("temporal_id in @tags_test['1-hour-period']").drop(columns=["temporal_id"]))
+```
+
+
+
+and for the 2-hour period forecasts:
+
+```python
+plot_series(Y_train.query("temporal_id in @tags_train['2-hour-period']")[["y", "ds", "unique_id"]].iloc[-50:], forecasts_df=Y_rec.query("temporal_id in @tags_test['2-hour-period']").drop(columns=["temporal_id"]))
+```
+
+
+
+Also, we can now verify that the forecasts are better coherent with each
+other. For the first 2-hour period, our forecast after reconciliation is
+6.63, and the sum of the forecasts for the first two 1-hour periods is
+1.7 + 4.92 = 6.63. Hence, we now have more accurate and coherent
+forecasts across frequencies.
+
+```python
+Y_rec.query("temporal_id in ['2-hour-period-1', '1-hour-period-1', '1-hour-period-2']")
+```
+
+| | unique_id | temporal_id | ds | y | TimeGPT | TimeGPT/MinTrace_method-wls_struct |
+|----|----|----|----|----|----|----|
+| 0 | DE | 2-hour-period-1 | 2017-12-29 01:00:00 | 10.45 | 16.949448 | 6.625738 |
+| 24 | DE | 1-hour-period-1 | 2017-12-29 00:00:00 | 9.73 | -0.241489 | 4.920365 |
+| 25 | DE | 1-hour-period-2 | 2017-12-29 01:00:00 | 0.72 | -3.456482 | 1.705373 |
+
+## Conclusion
+
+In this notebook we have shown: - How to create forecasts for multiple
+frequencies for the same dataset with TimeGPT - How to improve the
+accuracy of these forecasts using temporal reconciliation techniques
+
+Note that even though we created forecasts for two different frequencie,
+there is no ‘need’ to use the forecast of the 2-hour-period. One can use
+this technique also simply to improve the forecast of the 1-hour-period.
+
diff --git a/nixtla/docs/tutorials/training.html.mdx b/nixtla/docs/tutorials/training.html.mdx
new file mode 100644
index 00000000..c36cb696
--- /dev/null
+++ b/nixtla/docs/tutorials/training.html.mdx
@@ -0,0 +1,7 @@
+---
+output-file: training.html
+title: Training
+---
+
+
+
diff --git a/nixtla/docs/tutorials/uncertainty_quantification.html.mdx b/nixtla/docs/tutorials/uncertainty_quantification.html.mdx
new file mode 100644
index 00000000..e4d9e854
--- /dev/null
+++ b/nixtla/docs/tutorials/uncertainty_quantification.html.mdx
@@ -0,0 +1,7 @@
+---
+output-file: uncertainty_quantification.html
+title: Uncertainty quantification
+---
+
+
+
diff --git a/nixtla/docs/tutorials/uncertainty_quantification_with_prediction_intervals.html.mdx b/nixtla/docs/tutorials/uncertainty_quantification_with_prediction_intervals.html.mdx
new file mode 100644
index 00000000..5beac2f8
--- /dev/null
+++ b/nixtla/docs/tutorials/uncertainty_quantification_with_prediction_intervals.html.mdx
@@ -0,0 +1,165 @@
+---
+output-file: uncertainty_quantification_with_prediction_intervals.html
+title: Prediction intervals
+---
+
+
+In forecasting, we are often interested in a distribution of predictions
+rather than only a point prediction, because we want to have a notion of
+the uncertainty around the forecast.
+
+To this end, we can create *prediction intervals*.
+
+Prediction intervals have an intuitive interpretation, as they present a
+specific range of the forecast distribution. For instance, a 95%
+prediction interval means that 95 out of 100 times, we expect the future
+value to fall within the estimated range. Therefore, a wider interval
+indicates greater uncertainty about the forecast, while a narrower
+interval suggests higher confidence.
+
+With TimeGPT, we can create a distribution of forecasts, and extract the
+prediction intervals for a required level.
+
+TimeGPT uses [conformal
+prediction](https://en.wikipedia.org/wiki/Conformal_prediction) to
+produce the prediction intervals.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/11_uncertainty_quantification_with_prediction_intervals.ipynb)
+
+## 1. Import packages
+
+First, we import the required packages and initialize the Nixtla client
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, set the `base_url` argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 2. Load data
+
+```python
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/air_passengers.csv')
+df.head()
+```
+
+| | timestamp | value |
+|-----|------------|-------|
+| 0 | 1949-01-01 | 112 |
+| 1 | 1949-02-01 | 118 |
+| 2 | 1949-03-01 | 132 |
+| 3 | 1949-04-01 | 129 |
+| 4 | 1949-05-01 | 121 |
+
+## 3. Forecast with prediction intervals
+
+When using TimeGPT for time series forecasting, you can set the level
+(or levels) of prediction intervals according to your requirements.
+Here’s how you could do it:
+
+```python
+timegpt_fcst_pred_int_df = nixtla_client.forecast(
+ df=df, h=12, level=[80, 90, 99.7],
+ time_col='timestamp', target_col='value',
+)
+timegpt_fcst_pred_int_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| | timestamp | TimeGPT | TimeGPT-lo-99.7 | TimeGPT-lo-90 | TimeGPT-lo-80 | TimeGPT-hi-80 | TimeGPT-hi-90 | TimeGPT-hi-99.7 |
+|----|----|----|----|----|----|----|----|----|
+| 0 | 1961-01-01 | 437.837952 | 415.826484 | 423.783737 | 431.987091 | 443.688812 | 451.892166 | 459.849419 |
+| 1 | 1961-02-01 | 426.062744 | 402.833553 | 407.694092 | 412.704956 | 439.420532 | 444.431396 | 449.291935 |
+| 2 | 1961-03-01 | 463.116577 | 423.434092 | 430.316893 | 437.412564 | 488.820590 | 495.916261 | 502.799062 |
+| 3 | 1961-04-01 | 478.244507 | 444.885193 | 446.776764 | 448.726837 | 507.762177 | 509.712250 | 511.603821 |
+| 4 | 1961-05-01 | 505.646484 | 465.736694 | 471.976787 | 478.409872 | 532.883096 | 539.316182 | 545.556275 |
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+```python
+nixtla_client.plot(
+ df, timegpt_fcst_pred_int_df,
+ time_col='timestamp', target_col='value',
+ level=[80, 90],
+)
+```
+
+
+
+It’s essential to note that the choice of prediction interval level
+depends on your specific use case. For high-stakes predictions, you
+might want a wider interval to account for more uncertainty. For less
+critical forecasts, a narrower interval might be acceptable.
+
+#### Historical Forecast
+
+You can also compute prediction intervals for historical forecasts
+adding the `add_history=True` parameter as follows:
+
+```python
+timegpt_fcst_pred_int_historical_df = nixtla_client.forecast(
+ df=df, h=12, level=[80, 90],
+ time_col='timestamp', target_col='value',
+ add_history=True,
+)
+timegpt_fcst_pred_int_historical_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Calling Historical Forecast Endpoint...
+```
+
+| | timestamp | TimeGPT | TimeGPT-lo-80 | TimeGPT-lo-90 | TimeGPT-hi-80 | TimeGPT-hi-90 |
+|----|----|----|----|----|----|----|
+| 0 | 1951-01-01 | 135.483673 | 111.937767 | 105.262830 | 159.029579 | 165.704516 |
+| 1 | 1951-02-01 | 144.442413 | 120.896508 | 114.221571 | 167.988319 | 174.663256 |
+| 2 | 1951-03-01 | 157.191910 | 133.646004 | 126.971067 | 180.737815 | 187.412752 |
+| 3 | 1951-04-01 | 148.769379 | 125.223473 | 118.548536 | 172.315284 | 178.990221 |
+| 4 | 1951-05-01 | 140.472946 | 116.927041 | 110.252104 | 164.018852 | 170.693789 |
+
+```python
+nixtla_client.plot(
+ df, timegpt_fcst_pred_int_historical_df,
+ time_col='timestamp', target_col='value',
+ level=[80, 90],
+)
+```
+
+
+
diff --git a/nixtla/docs/tutorials/uncertainty_quantification_with_quantile_forecasts.html.mdx b/nixtla/docs/tutorials/uncertainty_quantification_with_quantile_forecasts.html.mdx
new file mode 100644
index 00000000..ac7cde8b
--- /dev/null
+++ b/nixtla/docs/tutorials/uncertainty_quantification_with_quantile_forecasts.html.mdx
@@ -0,0 +1,272 @@
+---
+output-file: uncertainty_quantification_with_quantile_forecasts.html
+title: Quantile forecasts
+---
+
+
+In forecasting, we are often interested in a distribution of predictions
+rather than only a point prediction, because we want to have a notion of
+the uncertainty around the forecast.
+
+To this end, we can create *quantile forecasts*.
+
+Quantile forecasts have an intuitive interpretation, as they present a
+specific percentile of the forecast distribution. This allows us to make
+statements such as ‘we expect 90% of our observations of air passengers
+to be above 100’. This approach is helpful for planning under
+uncertainty, providing a spectrum of possible future values and helping
+users make more informed decisions by considering the full range of
+potential outcomes.
+
+With TimeGPT, we can create a distribution of forecasts, and extract the
+quantile forecasts for a specified percentile. For instance, the 25th
+and 75th quantiles give insights into the lower and upper quartiles of
+expected outcomes, respectively, while the 50th quantile, or median,
+offers a central estimate.
+
+TimeGPT uses [conformal
+prediction](https://en.wikipedia.org/wiki/Conformal_prediction) to
+produce the quantiles.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/10_uncertainty_quantification_with_quantile_forecasts.ipynb)
+
+## 1. Import packages
+
+First, we import the required packages and initialize the Nixtla client
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+
+from IPython.display import display
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, set the `base_url` argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 2. Load data
+
+```python
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/air_passengers.csv')
+df.head()
+```
+
+| | timestamp | value |
+|-----|------------|-------|
+| 0 | 1949-01-01 | 112 |
+| 1 | 1949-02-01 | 118 |
+| 2 | 1949-03-01 | 132 |
+| 3 | 1949-04-01 | 129 |
+| 4 | 1949-05-01 | 121 |
+
+## 3. Forecast with quantiles
+
+When using TimeGPT for time series forecasting, you can set the
+quantiles you want to predict. Here’s how you could do it:
+
+```python
+quantiles = [0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9]
+timegpt_quantile_fcst_df = nixtla_client.forecast(
+ df=df, h=12,
+ quantiles=quantiles,
+ time_col='timestamp', target_col='value',
+)
+timegpt_quantile_fcst_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| | timestamp | TimeGPT | TimeGPT-q-10 | TimeGPT-q-20 | TimeGPT-q-30 | TimeGPT-q-40 | TimeGPT-q-50 | TimeGPT-q-60 | TimeGPT-q-70 | TimeGPT-q-80 | TimeGPT-q-90 |
+|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | 1961-01-01 | 437.837952 | 431.987091 | 435.043799 | 435.384363 | 436.402155 | 437.837952 | 439.273749 | 440.291541 | 440.632104 | 443.688812 |
+| 1 | 1961-02-01 | 426.062744 | 412.704956 | 414.832837 | 416.042432 | 421.719196 | 426.062744 | 430.406293 | 436.083057 | 437.292651 | 439.420532 |
+| 2 | 1961-03-01 | 463.116577 | 437.412564 | 444.234985 | 446.420233 | 450.705762 | 463.116577 | 475.527393 | 479.812921 | 481.998169 | 488.820590 |
+| 3 | 1961-04-01 | 478.244507 | 448.726837 | 455.428375 | 465.570038 | 469.879114 | 478.244507 | 486.609900 | 490.918976 | 501.060638 | 507.762177 |
+| 4 | 1961-05-01 | 505.646484 | 478.409872 | 493.154315 | 497.990848 | 499.138708 | 505.646484 | 512.154260 | 513.302121 | 518.138654 | 532.883096 |
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+TimeGPT will return forecasts in the format `TimeGPT-q-{int(100 * q)}`
+for each quantile `q`.
+
+```python
+nixtla_client.plot(
+ df, timegpt_quantile_fcst_df,
+ time_col='timestamp', target_col='value',
+)
+```
+
+
+
+It’s essential to note that the choice of the quantile (or quantiles)
+depends on your specific use case. For high-stakes predictions, you
+might lean towards more conservative quantiles, such as the 10th or 20th
+percentile, to ensure you’re prepared for worse-case scenarios. On the
+other hand, if you’re in a situation where the cost of over-preparation
+is high, you might choose a quantile closer to the median, like the 50th
+percentile, to balance being cautious and efficient.
+
+For instance, if you are managing inventory for a retail business during
+a big sale event, opting for a lower quantile might help you avoid
+running out of stock, even if it means you might overstock a bit. But if
+you are scheduling staff for a restaurant, you might go with a quantile
+closer to the middle to ensure you have enough staff on hand without
+significantly overstaffing.
+
+Ultimately, the choice comes down to understanding the balance between
+risk and cost in your specific context, and using quantile forecasts
+from TimeGPT allows you to tailor your strategy to fit that balance
+perfectly.
+
+#### Historical Forecast
+
+You can also compute quantile forecasts for historical forecasts adding
+the `add_history=True` parameter as follows:
+
+```python
+timegpt_quantile_fcst_df = nixtla_client.forecast(
+ df=df, h=12,
+ quantiles=quantiles,
+ time_col='timestamp', target_col='value',
+ add_history=True,
+)
+timegpt_quantile_fcst_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Calling Historical Forecast Endpoint...
+```
+
+| | timestamp | TimeGPT | TimeGPT-q-10 | TimeGPT-q-20 | TimeGPT-q-30 | TimeGPT-q-40 | TimeGPT-q-50 | TimeGPT-q-60 | TimeGPT-q-70 | TimeGPT-q-80 | TimeGPT-q-90 |
+|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | 1951-01-01 | 135.483673 | 111.937768 | 120.020593 | 125.848879 | 130.828935 | 135.483673 | 140.138411 | 145.118467 | 150.946753 | 159.029579 |
+| 1 | 1951-02-01 | 144.442398 | 120.896493 | 128.979318 | 134.807604 | 139.787660 | 144.442398 | 149.097136 | 154.077192 | 159.905478 | 167.988304 |
+| 2 | 1951-03-01 | 157.191910 | 133.646004 | 141.728830 | 147.557116 | 152.537172 | 157.191910 | 161.846648 | 166.826703 | 172.654990 | 180.737815 |
+| 3 | 1951-04-01 | 148.769363 | 125.223458 | 133.306284 | 139.134570 | 144.114625 | 148.769363 | 153.424102 | 158.404157 | 164.232443 | 172.315269 |
+| 4 | 1951-05-01 | 140.472946 | 116.927041 | 125.009866 | 130.838152 | 135.818208 | 140.472946 | 145.127684 | 150.107740 | 155.936026 | 164.018852 |
+
+```python
+nixtla_client.plot(
+ df, timegpt_quantile_fcst_df,
+ time_col='timestamp', target_col='value',
+)
+```
+
+
+
+#### Cross Validation
+
+The `quantiles` argument can also be included in the `cross_validation`
+method, allowing comparing the performance of TimeGPT across different
+windows and different quantiles.
+
+```python
+timegpt_cv_quantile_fcst_df = nixtla_client.cross_validation(
+ df=df,
+ h=12,
+ n_windows=5,
+ quantiles=quantiles,
+ time_col='timestamp',
+ target_col='value',
+)
+timegpt_quantile_fcst_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: MS
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+```
+
+| | timestamp | TimeGPT | TimeGPT-q-10 | TimeGPT-q-20 | TimeGPT-q-30 | TimeGPT-q-40 | TimeGPT-q-50 | TimeGPT-q-60 | TimeGPT-q-70 | TimeGPT-q-80 | TimeGPT-q-90 |
+|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | 1951-01-01 | 135.483673 | 111.937768 | 120.020593 | 125.848879 | 130.828935 | 135.483673 | 140.138411 | 145.118467 | 150.946753 | 159.029579 |
+| 1 | 1951-02-01 | 144.442398 | 120.896493 | 128.979318 | 134.807604 | 139.787660 | 144.442398 | 149.097136 | 154.077192 | 159.905478 | 167.988304 |
+| 2 | 1951-03-01 | 157.191910 | 133.646004 | 141.728830 | 147.557116 | 152.537172 | 157.191910 | 161.846648 | 166.826703 | 172.654990 | 180.737815 |
+| 3 | 1951-04-01 | 148.769363 | 125.223458 | 133.306284 | 139.134570 | 144.114625 | 148.769363 | 153.424102 | 158.404157 | 164.232443 | 172.315269 |
+| 4 | 1951-05-01 | 140.472946 | 116.927041 | 125.009866 | 130.838152 | 135.818208 | 140.472946 | 145.127684 | 150.107740 | 155.936026 | 164.018852 |
+
+```python
+cutoffs = timegpt_cv_quantile_fcst_df['cutoff'].unique()
+for cutoff in cutoffs:
+ fig = nixtla_client.plot(
+ df.tail(100),
+ timegpt_cv_quantile_fcst_df.query('cutoff == @cutoff').drop(columns=['cutoff', 'value']),
+ time_col='timestamp',
+ target_col='value'
+ )
+ display(fig)
+```
+
+
+
+
+
+
+
+
+
+
+
diff --git a/nixtla/docs/tutorials/validation.html.mdx b/nixtla/docs/tutorials/validation.html.mdx
new file mode 100644
index 00000000..4ddc63e4
--- /dev/null
+++ b/nixtla/docs/tutorials/validation.html.mdx
@@ -0,0 +1,26 @@
+---
+output-file: validation.html
+title: Validation
+---
+
+
+One of the primary challenges in time series forecasting is the inherent
+uncertainty and variability over time, making it crucial to validate the
+accuracy and reliability of the models employed. `TimeGPT` offers the
+possibility for cross-validation and historical forecasts to help you
+validate your predictions.
+
+### What You Will Learn
+
+1. **[Cross-Validation](https://docs.nixtla.io/docs/tutorials-cross_validation)**
+
+ - Learn how to perform time series cross-validation across
+ different continuous windows of your data.
+
+2. **[Historical
+ Forecasts](https://docs.nixtla.io/docs/tutorials-historical_forecast)**
+
+ - Generate in-sample forecasts to validate how `TimeGPT` would
+ have performed in the past, providing insights into the model’s
+ accuracy.
+
diff --git a/nixtla/docs/use-cases/1_forecasting_web_traffic_files/figure-markdown_strict/cell-12-output-1.png b/nixtla/docs/use-cases/1_forecasting_web_traffic_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..12017b79
Binary files /dev/null and b/nixtla/docs/use-cases/1_forecasting_web_traffic_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/nixtla/docs/use-cases/1_forecasting_web_traffic_files/figure-markdown_strict/cell-7-output-1.png b/nixtla/docs/use-cases/1_forecasting_web_traffic_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..e9bb21f6
Binary files /dev/null and b/nixtla/docs/use-cases/1_forecasting_web_traffic_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/nixtla/docs/use-cases/2_bitcoin_price_prediction_files/figure-markdown_strict/cell-10-output-1.png b/nixtla/docs/use-cases/2_bitcoin_price_prediction_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..ed8907f9
Binary files /dev/null and b/nixtla/docs/use-cases/2_bitcoin_price_prediction_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/nixtla/docs/use-cases/2_bitcoin_price_prediction_files/figure-markdown_strict/cell-12-output-1.png b/nixtla/docs/use-cases/2_bitcoin_price_prediction_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..d59709b1
Binary files /dev/null and b/nixtla/docs/use-cases/2_bitcoin_price_prediction_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/nixtla/docs/use-cases/2_bitcoin_price_prediction_files/figure-markdown_strict/cell-14-output-1.png b/nixtla/docs/use-cases/2_bitcoin_price_prediction_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..43791c66
Binary files /dev/null and b/nixtla/docs/use-cases/2_bitcoin_price_prediction_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/nixtla/docs/use-cases/2_bitcoin_price_prediction_files/figure-markdown_strict/cell-7-output-1.png b/nixtla/docs/use-cases/2_bitcoin_price_prediction_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..c7701a2d
Binary files /dev/null and b/nixtla/docs/use-cases/2_bitcoin_price_prediction_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/nixtla/docs/use-cases/2_bitcoin_price_prediction_files/figure-markdown_strict/cell-9-output-1.png b/nixtla/docs/use-cases/2_bitcoin_price_prediction_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..b2642c49
Binary files /dev/null and b/nixtla/docs/use-cases/2_bitcoin_price_prediction_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/nixtla/docs/use-cases/3_electricity_demand_files/figure-markdown_strict/cell-10-output-1.png b/nixtla/docs/use-cases/3_electricity_demand_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..34b69028
Binary files /dev/null and b/nixtla/docs/use-cases/3_electricity_demand_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/nixtla/docs/use-cases/3_electricity_demand_files/figure-markdown_strict/cell-7-output-1.png b/nixtla/docs/use-cases/3_electricity_demand_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..15565ac2
Binary files /dev/null and b/nixtla/docs/use-cases/3_electricity_demand_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/nixtla/docs/use-cases/4_intermittent_demand_files/figure-markdown_strict/cell-11-output-1.png b/nixtla/docs/use-cases/4_intermittent_demand_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..c000d010
Binary files /dev/null and b/nixtla/docs/use-cases/4_intermittent_demand_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/nixtla/docs/use-cases/4_intermittent_demand_files/figure-markdown_strict/cell-6-output-1.png b/nixtla/docs/use-cases/4_intermittent_demand_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..d0c19a52
Binary files /dev/null and b/nixtla/docs/use-cases/4_intermittent_demand_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/nixtla/docs/use-cases/5_what_if_pricing_scenarios_in_retail_files/figure-markdown_strict/cell-10-output-1.png b/nixtla/docs/use-cases/5_what_if_pricing_scenarios_in_retail_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..3903e6f8
Binary files /dev/null and b/nixtla/docs/use-cases/5_what_if_pricing_scenarios_in_retail_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/nixtla/docs/use-cases/5_what_if_pricing_scenarios_in_retail_files/figure-markdown_strict/cell-14-output-1.png b/nixtla/docs/use-cases/5_what_if_pricing_scenarios_in_retail_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..43ef9d47
Binary files /dev/null and b/nixtla/docs/use-cases/5_what_if_pricing_scenarios_in_retail_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/nixtla/docs/use-cases/5_what_if_pricing_scenarios_in_retail_files/figure-markdown_strict/cell-18-output-1.png b/nixtla/docs/use-cases/5_what_if_pricing_scenarios_in_retail_files/figure-markdown_strict/cell-18-output-1.png
new file mode 100644
index 00000000..dbaad174
Binary files /dev/null and b/nixtla/docs/use-cases/5_what_if_pricing_scenarios_in_retail_files/figure-markdown_strict/cell-18-output-1.png differ
diff --git a/nixtla/docs/use-cases/5_what_if_pricing_scenarios_in_retail_files/figure-markdown_strict/cell-9-output-1.png b/nixtla/docs/use-cases/5_what_if_pricing_scenarios_in_retail_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..04c54cca
Binary files /dev/null and b/nixtla/docs/use-cases/5_what_if_pricing_scenarios_in_retail_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/nixtla/docs/use-cases/bitcoin_price_prediction.html.mdx b/nixtla/docs/use-cases/bitcoin_price_prediction.html.mdx
new file mode 100644
index 00000000..25e4456d
--- /dev/null
+++ b/nixtla/docs/use-cases/bitcoin_price_prediction.html.mdx
@@ -0,0 +1,371 @@
+---
+description: Learn how to use TimeGPT for financial time series forecasting
+output-file: bitcoin_price_prediction.html
+title: Bitcoin price prediction
+---
+
+
+## Introduction
+
+Forecasting time series is a ubiquitous task in finance, supporting
+decisions in trading, risk management, and strategic planning. Despite
+its prevalence, predicting the future prices of financial assets remains
+a formidable challenge, mainly due to the inherent volatility of
+financial markets.
+
+For those who believe in the feasibility of forecasting these assets, or
+for professionals whose roles require such predictions, TimeGPT is a
+powerful tool that simplifies the forecasting process.
+
+In this tutorial, we will demonstrate how to use TimeGPT for financial
+time series forecasting, focusing on Bitcoin price prediction. We will
+also showcase how to use TimeGPT for uncertainty quantification, which
+is essential for risk management and decision-making.
+
+**Outline:**
+
+1. [Load Bitcoin Price Data](#load-bitcoin-price-data)
+
+2. [Get Started with TimeGPT](#get-started-with-timegpt)
+
+3. [Visualize the Data](#visualize-the-data)
+
+4. [Forecast with TimeGPT](#forecast-with-timegpt)
+
+5. [Extend Bitcon Price Analysis with
+ TimeGPT](#extend-bitcoin-price-analysis-with-timegpt)
+
+6. [Understand the Model’s
+ Limitations](#understand-the-models-limitations)
+
+7. [References and Additional
+ Material](#references-and-additional-material)
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/use-cases/2_bitcoin_price_prediction.ipynb)
+
+## 1. Load Bitcoin Price Data
+
+Bitcoin (₿) is the first decentralized digital currency and is one of
+the most popular cryptocurrencies. Transactions are managed and recorded
+on a public ledger known as the blockchain. Bitcoins are created as a
+reward for mining, a process that involves solving complex cryptographic
+tasks to verify transactions. This digital currency can be used as
+payment for goods and services, traded for other currencies, or held as
+a store of value.
+
+In this tutorial, we will first download the historical Bitcoin price
+data in USD as a `pandas` DataFrame.
+
+```python
+import pandas as pd
+
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/bitcoin_price_usd.csv', sep=',')
+df.head()
+```
+
+| | Date | Close |
+|-----|------------|-------------|
+| 0 | 2020-01-01 | 7200.174316 |
+| 1 | 2020-01-02 | 6985.470215 |
+| 2 | 2020-01-03 | 7344.884277 |
+| 3 | 2020-01-04 | 7410.656738 |
+| 4 | 2020-01-05 | 7411.317383 |
+
+This dataset contains the closing price of Bitcoin in USD from
+2020-01-01 to 2023-12-31. It’s important to note that unlike traditional
+financial assets, Bitcoin trades 24/7. Therefore, the closing price
+represents the price of Bitcoin at a specific time each day, rather than
+at the end of a trading day.
+
+For convenience, we will rename the `Date` and `Close` columns to `ds`
+and `y`, respectively.
+
+```python
+df.rename(columns={'Date': 'ds', 'Close': 'y'}, inplace=True)
+```
+
+## 2. Get Started with TimeGPT
+
+To get started with `TimeGPT`, you need to instantiate the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class. For this, you will need a Nixtla API key.
+
+```python
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+To learn more about how to set up your API key, please refer to the
+[Setting Up Your Authentication API
+Key](https://docs.nixtla.io/docs/getting-started-setting_up_your_api_key)
+tutorial.
+
+## 3. Visualize the Data
+
+Before attempting any forecasting, it is good practice to visualize the
+data we want to predict. The
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class includes a `plot` method for this purpose.
+
+The `plot` method has an `engine` argument that allows you to choose
+between different plotting libraries. Default is `matplotlib`, but you
+can also use `plotly` for interactive plots.
+
+```python
+nixtla_client.plot(df)
+```
+
+
+
+If you haven’t renamed the column names of your DataFrame to `ds` and
+`y`, you will need to specify the `time_col` and `target_col` arguments
+of the `plot`method:
+
+```python
+nixtla_client.plot(df, time_col='name of your time column', target_col='name of your target column')
+```
+
+This is necessary not only for the `plot` method but for all methods
+from the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class.
+
+## 4. Forecast with TimeGPT
+
+Now we are ready to generate predictions with TimeGPT. To do this, we
+will use the `forecast` method from the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class.
+
+The `forecast` method requires the following arguments:
+
+- `df`: The DataFrame containing the time series data
+
+- `h`: (int) The forecast horizon. In this case, we will forecast the
+ next 7 days.
+
+- `level`: (list) The confidence level for the prediction intervals.
+ Given the inherent volatility of Bitcoin, we will use multiple
+ confidence levels.
+
+```python
+level = [50,80,90] # confidence levels
+
+fcst = nixtla_client.forecast(df, h=7, level=level)
+fcst.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: D
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| | ds | TimeGPT | TimeGPT-lo-90 | TimeGPT-lo-80 | TimeGPT-lo-50 | TimeGPT-hi-50 | TimeGPT-hi-80 | TimeGPT-hi-90 |
+|----|----|----|----|----|----|----|----|----|
+| 0 | 2024-01-01 | 42269.460938 | 39567.209020 | 40429.953636 | 41380.654646 | 43158.267229 | 44108.968239 | 44971.712855 |
+| 1 | 2024-01-02 | 42469.917969 | 39697.941669 | 40578.197049 | 41466.511361 | 43473.324576 | 44361.638888 | 45241.894268 |
+| 2 | 2024-01-03 | 42864.078125 | 40538.871243 | 41586.252507 | 42284.316674 | 43443.839576 | 44141.903743 | 45189.285007 |
+| 3 | 2024-01-04 | 42881.621094 | 40603.117448 | 41216.106493 | 42058.539392 | 43704.702795 | 44547.135694 | 45160.124739 |
+| 4 | 2024-01-05 | 42773.457031 | 40213.699760 | 40665.384780 | 41489.812431 | 44057.101632 | 44881.529282 | 45333.214302 |
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+We can pass the forecasts we just generated to the `plot` method to
+visualize the predictions with the historical data.
+
+```python
+nixtla_client.plot(df, fcst, level=level)
+```
+
+
+
+To get a closer look at the predictions, we can zoom in on the plot or
+specify the maximum number of in-sample observations to be plotted using
+the `max_insample_length` argument. Note that setting
+`max_insample_length=60`, for instance, will display the last 60
+historical values along with the complete forecast.
+
+```python
+nixtla_client.plot(df, fcst, level=level, max_insample_length=60)
+```
+
+
+
+Additionally, if you set the `add_history` argument of the `forecast`
+method to `True`, `TimeGPT` will generate predictions for the historical
+observations too. This can be useful for assessing the model’s
+performance on the training data.
+
+```python
+forecast = nixtla_client.forecast(df, h=7, level=level, add_history=True)
+forecast.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: D
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Calling Historical Forecast Endpoint...
+```
+
+| | ds | TimeGPT | TimeGPT-lo-50 | TimeGPT-lo-80 | TimeGPT-lo-90 | TimeGPT-hi-50 | TimeGPT-hi-80 | TimeGPT-hi-90 |
+|----|----|----|----|----|----|----|----|----|
+| 0 | 2020-02-03 | 9425.702148 | 7622.287194 | 5999.157479 | 5027.779677 | 11229.117103 | 12852.246818 | 13823.624619 |
+| 1 | 2020-02-04 | 9568.482422 | 7765.067467 | 6141.937752 | 5170.559951 | 11371.897376 | 12995.027092 | 13966.404893 |
+| 2 | 2020-02-05 | 9557.082031 | 7753.667077 | 6130.537362 | 5159.159560 | 11360.496986 | 12983.626701 | 13955.004502 |
+| 3 | 2020-02-06 | 9486.123047 | 7682.708092 | 6059.578377 | 5088.200576 | 11289.538001 | 12912.667717 | 13884.045518 |
+| 4 | 2020-02-07 | 9475.242188 | 7671.827233 | 6048.697518 | 5077.319716 | 11278.657142 | 12901.786857 | 13873.164659 |
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+```python
+nixtla_client.plot(df, forecast, level=level)
+```
+
+
+
+## 5. Extend Bitcoin Price Analysis with TimeGPT
+
+### Anomaly Detection
+
+Given the volatility of the price of Bitcoin, it can be useful to try to
+identify anomalies in the data. `TimeGPT` can be used for this by
+calling the `detect_anomalies` method from the
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+class. This method evaluates each observation against its context within
+the series, using statistical measures to determine its likelihood of
+being an anomaly. By default, it identifies anomalies based on a 99
+percent prediction interval. To change this, you can specify the `level`
+argument.
+
+```python
+anomalies_df = nixtla_client.detect_anomalies(df)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: D
+INFO:nixtla.nixtla_client:Calling Anomaly Detector Endpoint...
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.detect_anomalies(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+```python
+nixtla_client.plot(df, anomalies_df, plot_anomalies=True)
+```
+
+
+
+To learn more about how to detect anomalies with `TimeGPT`, take a look
+at our [Anomaly
+Detection](https://docs.nixtla.io/docs/capabilities-anomaly-detection-anomaly_detection)
+tutorial.
+
+### Add Exogenous Variables
+
+If you have additional information that you believe could help improve
+the forecast, consider including it as an exogenous variable. For
+instance, you might add data such as the price of other
+cryptocurrencies, proprietary information, stock market indices, or the
+number of transactions in the Bitcoin network.
+
+`TimeGPT` supports the incorporation of exogenous variables in the
+`forecast` method. However, keep in mind that you’ll need to know the
+future values of these variables.
+
+To learn how to incorporate exogenous variables to `TimeGPT`, refer to
+the [Exogenous
+Variables](https://docs.nixtla.io/docs/tutorials-exogenous_variables)
+tutorial.
+
+## 6. Understand the Model’s Limitations
+
+As stated in the introduction, predicting the future prices of financial
+assets is a challenging task, especially for assets like Bitcoin. The
+predictions in this tutorial may appear accurate, mainly because they
+align with recent historical data and the model updates with new values
+at short intervals, avoiding significant deviations. However, the true
+challenge lies in forecasting Bitcoin’s price for the upcoming days, not
+just its historical performance. For those who need or want to try to
+forecast these assets, `TimeGPT` can be an option that simplifies the
+forecasting process. With just a couple of lines of code, `TimeGPT` can
+help you:
+
+- Produce point forecasts
+- Quantify the uncertainty of your predictions
+- Produce in-sample forecasts
+- Detect anomalies
+- Incorporate exogenous variables
+
+To learn more about `TimeGPT` capabilities, please refer to the [TimeGPT
+Documentation](https://docs.nixtla.io/).
+
+## 7. References and Additional Material
+
+**References**
+
+- [Joaquín Amat Rodrigo and Javier Escobar Ortiz (2022), “Bitcoin
+ price prediction with Python, when the past does not repeat
+ itself”](https://www.cienciadedatos.net/documentos/py41-forecasting-cryptocurrency-bitcoin-machine-learning-python.html)
+
+Furthermore, for many financial time series, the best estimate for the
+price is often a random walk model, meaning that the best forecast for
+tomorrow’s price is today’s price. Nixtla’s
+[StatsForecast](https://nixtlaverse.nixtla.io/statsforecast/index.html)
+library allows you to easily implement this model and variations.
+
diff --git a/nixtla/docs/use-cases/electricity_demand.html.mdx b/nixtla/docs/use-cases/electricity_demand.html.mdx
new file mode 100644
index 00000000..d5f2e182
--- /dev/null
+++ b/nixtla/docs/use-cases/electricity_demand.html.mdx
@@ -0,0 +1,288 @@
+---
+output-file: electricity_demand.html
+title: Forecasting Energy Demand
+---
+
+
+This tutorial is based on an energy consumption forecasting scenario
+where we make a 4-day forecast of in-zone energy consumption.
+
+Here, we use a subset of the [PJM Hourly Energy Consumption
+dataset](https://www.pjm.com/), focusing on in-zone consumption, where
+electricity is both generated and consumed within the same transmission
+zone. The dataset consists of hourly data from October 1, 2023, to
+September 30, 2024, covering five representative areas to capture hourly
+energy demand patterns.
+
+In this experiment, we show that using TimeGPT delivers significant
+improvements over using a state-of-the-art deep learning model like
+N-HiTS in a just a few lines of code:
+
+- MAE of TimeGPT is **18.6% better** than N-HiTS
+- sMAPE of TimeGPT is **31.1% better** than N-HiTS
+- TimeGPT generated predictions in **4.3 seconds**, which is **90%
+ faster** than training and predicting with N-HiTS.
+
+The following tutorial explore all the steps in detail to reproduce
+these results so that you can apply TimeGPT in your own project.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/3_electricity_demand.ipynb)
+
+## Initial setup
+
+First, we load the required packages for this experiment.
+
+```python
+import time
+import requests
+import pandas as pd
+
+from nixtla import NixtlaClient
+
+from utilsforecast.losses import mae, smape
+from utilsforecast.evaluation import evaluate
+```
+
+Of course, we need an instance of
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient)
+to use TimeGPT.
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## Read the data
+
+Here, we load in the inbound energy transmission time series.
+
+```python
+df = pd.read_csv('https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/refs/heads/main/datasets/pjm_in_zone.csv')
+df['ds'] = pd.to_datetime(df['ds'])
+```
+
+
+```python
+df.groupby('unique_id').head(2)
+```
+
+| | unique_id | ds | y |
+|-------|-----------|---------------------------|-----------|
+| 0 | AP-AP | 2023-10-01 04:00:00+00:00 | 4042.513 |
+| 1 | AP-AP | 2023-10-01 05:00:00+00:00 | 3850.067 |
+| 8784 | DOM-DOM | 2023-10-01 04:00:00+00:00 | 10732.435 |
+| 8785 | DOM-DOM | 2023-10-01 05:00:00+00:00 | 10314.211 |
+| 17568 | JC-JC | 2023-10-01 04:00:00+00:00 | 1825.101 |
+| 17569 | JC-JC | 2023-10-01 05:00:00+00:00 | 1729.590 |
+| 26352 | PN-PN | 2023-10-01 04:00:00+00:00 | 1454.666 |
+| 26353 | PN-PN | 2023-10-01 05:00:00+00:00 | 1416.688 |
+| 35136 | RTO-RTO | 2023-10-01 04:00:00+00:00 | 69139.393 |
+| 35137 | RTO-RTO | 2023-10-01 05:00:00+00:00 | 66207.416 |
+
+Let’s plot our series to see what it looks like.
+
+```python
+nixtla_client.plot(
+ df,
+ max_insample_length=365,
+)
+```
+
+
+
+We can see clear sesaonal pattern in all of our series. It will be
+interesting to see how TimeGPT handles this type of data.
+
+## Forecasting with TimeGPT
+
+### Splitting the data
+
+The first step is to split our data. Here, we define an input DataFrame
+to feed to the model. We also reserve the last 96 time steps for the
+test set, so that we can evaluate the performance of TimeGPT against
+actual values.
+
+For this situation, we use a forecast horizon of 96, which represents
+four days, and we use an input sequence of 362 days, which is 8688 time
+steps.
+
+```python
+test_df = df.groupby('unique_id').tail(96) # 96 = 4 days (96 * 1 day/24h )
+
+input_df = df.groupby('unique_id').apply(lambda group: group.iloc[-1104:-96]).reset_index(drop=True) # 1008 = 42 days (1008 * 1 day/24h)
+```
+
+### Forecasting
+
+Then, we simply call the `forecast` method. Here, we use fine-tuning and
+specify the mean absolute error (MAE) as the fine-tuning loss. Also, we
+use the `timegpt-1-long-horizon` since we are forecasting the next two
+days, and the seasoanl period is one day.
+
+```python
+start = time.time()
+
+fcst_df = nixtla_client.forecast(
+ df=input_df,
+ h=96,
+ level=[90], # Generate a 90% confidence interval
+ finetune_steps=10, # Specify the number of steps for fine-tuning
+ finetune_loss='mae', # Use the MAE as the loss function for fine-tuning
+ model='timegpt-1-long-horizon', # Use the model for long-horizon forecasting
+ time_col='ds',
+ target_col='y',
+ id_col='unique_id'
+)
+
+end = time.time()
+
+timegpt_duration = end - start
+
+print(f"Time (TimeGPT): {timegpt_duration}")
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+TimeGPT was done in 4.3 seconds! We can now plot the predictions against
+the actual values of the test set.
+
+```python
+nixtla_client.plot(test_df, fcst_df, models=['TimeGPT'], level=[90], time_col='ds', target_col='y')
+```
+
+
+
+### Evaluation
+
+Now that we have predictions, let’s evaluate the model’s performance.
+
+```python
+fcst_df['ds'] = pd.to_datetime(fcst_df['ds'])
+
+test_df = pd.merge(test_df, fcst_df, 'left', ['unique_id', 'ds'])
+```
+
+
+```python
+evaluation = evaluate(
+ test_df,
+ metrics=[mae, smape],
+ models=["TimeGPT"],
+ target_col="y",
+ id_col='unique_id'
+)
+
+average_metrics = evaluation.groupby('metric')['TimeGPT'].mean()
+average_metrics
+```
+
+``` text
+metric
+mae 882.693979
+smape 0.019974
+Name: TimeGPT, dtype: float64
+```
+
+We can see that TimeGPT achieves a MAE of 882.6 and a sMAPE of 2%.
+
+Great! Now, let’s see if a data-specific model can do better.
+
+## Forecasting with N-HiTS
+
+Here, we use the N-HiTS model, as it is very fast to train and performs
+well on long-horizon forecasting tasks. To reproduce these results, make
+sure to install the library `neuralforecast`.
+
+```python
+from neuralforecast.core import NeuralForecast
+from neuralforecast.models import NHITS
+```
+
+### Define the training set
+
+The training set is different from the input DataFrame for TimeGPT, as
+we need more data to train a data-specific model.
+
+Note that the dataset is very large, so we use the last 362 days of the
+training set to fit our model.
+
+```python
+train_df = df.groupby('unique_id').apply(lambda group: group.iloc[:-96]).reset_index(drop=True)
+```
+
+### Forecasting with N-HiTS
+
+We can now fit the model on training set and make predictions.
+
+```python
+horizon = 96
+
+models = [NHITS(h=horizon, input_size = 5*horizon, scaler_type='robust', batch_size=16, valid_batch_size=8)]
+
+nf = NeuralForecast(models=models, freq='H')
+
+start = time.time()
+
+nf.fit(df=train_df)
+nhits_preds = nf.predict()
+
+end = time.time()
+
+nhits_duration = end - start
+
+print(f"Time (N-HiTS): {nhits_duration}")
+```
+
+Great! Note that N-HiTS took 44 seconds to carry out the training and
+forecasting procedures. Now, let’s evaluate the performance of this
+model.
+
+### Evaluation
+
+```python
+preds_df = pd.merge(test_df, nhits_preds, 'left', ['unique_id', 'ds'])
+
+evaluation = evaluate(
+ preds_df,
+ metrics=[mae, smape],
+ models=["NHITS"],
+ target_col="y",
+ id_col='unique_id'
+)
+
+
+average_metrics = evaluation.groupby('metric')['NHITS'].mean()
+print(average_metrics)
+```
+
+## Conclusion
+
+TimeGPT achieves a MAE of 882.6 while N-HiTS achieves a MAE of 1084.7,
+meaning there is a **18.6% improvement** in using TimeGPT versus our
+data-specific N-HiTS model. TimeGPT also improved the sMAPE by 31.1%.
+
+Plus, TimeGPT took 4.3 seconds to generate forecasts, while N-HiTS took
+44 seconds to fit and predict. TimeGPT is thus **90% faster** than using
+N-HiTS in this scenario.
+
diff --git a/nixtla/docs/use-cases/forecasting_web_traffic.html.mdx b/nixtla/docs/use-cases/forecasting_web_traffic.html.mdx
new file mode 100644
index 00000000..76c502a5
--- /dev/null
+++ b/nixtla/docs/use-cases/forecasting_web_traffic.html.mdx
@@ -0,0 +1,370 @@
+---
+output-file: forecasting_web_traffic.html
+title: Forecasting web traffic
+---
+
+
+Our task is to forecast the next 7 days of daily visits to the website
+[cienciadedatos.net](cienciadedatos.net).
+
+In this tutorial we will show:
+
+- How to load time series data to be used for forecasting with TimeGPT
+
+- How to create cross-validated forecasts with TimeGPT
+
+This tutorial is an adaptation from [Joaquín Amat Rodrigo, Javier
+Escobar Ortiz, “Forecasting web traffic with machine learning and
+Python”](https://cienciadedatos.net/documentos/py37-forecasting-web-traffic-machine-learning.html).
+We will show you:
+
+- how you can achieve almost 10% better forecasting results;
+
+- using significantly less lines of code;
+
+- in a fraction of the time needed to run the original tutorial.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/use-cases/1_forecasting_web_traffic.ipynb)
+
+## 1. Import packages
+
+First, we import the required packages and initialize the Nixtla client.
+
+```python
+import pandas as pd
+from nixtla import NixtlaClient
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 2. Load data
+
+We load the website visit data, and set it to the right format to use
+with TimeGPT. In this case, we only need to add an identifier column for
+the timeseries, which we will call `daily_visits`.
+
+```python
+url = ('https://raw.githubusercontent.com/JoaquinAmatRodrigo/Estadistica-machine-learning-python/' +
+ 'master/data/visitas_por_dia_web_cienciadedatos.csv')
+df = pd.read_csv(url, sep=',', parse_dates=[0], date_format='%d/%m/%y')
+df['unique_id'] = 'daily_visits'
+
+df.head(10)
+```
+
+| | date | users | unique_id |
+|-----|------------|-------|--------------|
+| 0 | 2020-07-01 | 2324 | daily_visits |
+| 1 | 2020-07-02 | 2201 | daily_visits |
+| 2 | 2020-07-03 | 2146 | daily_visits |
+| 3 | 2020-07-04 | 1666 | daily_visits |
+| 4 | 2020-07-05 | 1433 | daily_visits |
+| 5 | 2020-07-06 | 2195 | daily_visits |
+| 6 | 2020-07-07 | 2240 | daily_visits |
+| 7 | 2020-07-08 | 2295 | daily_visits |
+| 8 | 2020-07-09 | 2279 | daily_visits |
+| 9 | 2020-07-10 | 2155 | daily_visits |
+
+That’s it! No more preprocessing is necessary.
+
+## 3. Cross-validation with TimeGPT
+
+We can perform cross-validation on our data as follows:
+
+```python
+timegpt_cv_df = nixtla_client.cross_validation(
+ df,
+ h=7,
+ n_windows=8,
+ time_col='date',
+ target_col='users',
+ freq='D',
+ level=[80, 90, 99.5]
+)
+timegpt_cv_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Restricting input...
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+```
+
+| | unique_id | date | cutoff | users | TimeGPT | TimeGPT-lo-99.5 | TimeGPT-lo-90 | TimeGPT-lo-80 | TimeGPT-hi-80 | TimeGPT-hi-90 | TimeGPT-hi-99.5 |
+|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | daily_visits | 2021-07-01 | 2021-06-30 | 3123 | 3310.908447 | 3041.925497 | 3048.363220 | 3082.721924 | 3539.094971 | 3573.453674 | 3579.891397 |
+| 1 | daily_visits | 2021-07-02 | 2021-06-30 | 2870 | 3090.971680 | 2793.535905 | 2838.480298 | 2853.750488 | 3328.192871 | 3343.463062 | 3388.407455 |
+| 2 | daily_visits | 2021-07-03 | 2021-06-30 | 2020 | 2346.991455 | 2043.731296 | 2150.005078 | 2171.187012 | 2522.795898 | 2543.977832 | 2650.251614 |
+| 3 | daily_visits | 2021-07-04 | 2021-06-30 | 1828 | 2182.191895 | 1836.848173 | 1897.684900 | 1929.914575 | 2434.469214 | 2466.698889 | 2527.535616 |
+| 4 | daily_visits | 2021-07-05 | 2021-06-30 | 2722 | 3082.715088 | 2736.008055 | 2746.997034 | 2791.375342 | 3374.054834 | 3418.433142 | 3429.422121 |
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.cross_validation(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+Here, we have performed a rolling cross-validation of 8 folds. Let’s
+plot the cross-validated forecasts including the prediction intervals:
+
+```python
+nixtla_client.plot(
+ df,
+ timegpt_cv_df.drop(columns=['cutoff', 'users']),
+ time_col='date',
+ target_col='users',
+ max_insample_length=90,
+ level=[80, 90, 99.5]
+)
+```
+
+
+
+This looks reasonable, and very comparable to the results obtained
+[here](https://cienciadedatos.net/documentos/py37-forecasting-web-traffic-machine-learning.html).
+
+Let’s check the Mean Absolute Error of our cross-validation:
+
+```python
+from utilsforecast.losses import mae
+```
+
+
+```python
+mae_timegpt = mae(df = timegpt_cv_df.drop(columns=['cutoff']),
+ models=['TimeGPT'],
+ target_col='users')
+
+mae_timegpt
+```
+
+| | unique_id | TimeGPT |
+|-----|--------------|------------|
+| 0 | daily_visits | 167.691711 |
+
+The MAE of our backtest is `167.69`. Hence, not only did TimeGPT achieve
+a lower MAE compared to the fully customized pipeline
+[here](https://cienciadedatos.net/documentos/py37-forecasting-web-traffic-machine-learning.html),
+the error of the forecast is also lower.
+
+#### Exogenous variables
+
+Now let’s add some exogenous variables to see if we can improve the
+forecasting performance further.
+
+We will add weekday indicators, which we will extract from the `date`
+column.
+
+```python
+# We have 7 days, for each day a separate column denoting 1/0
+for i in range(7):
+ df[f'week_day_{i + 1}'] = 1 * (df['date'].dt.weekday == i)
+
+df.head(10)
+```
+
+| | date | users | unique_id | week_day_1 | week_day_2 | week_day_3 | week_day_4 | week_day_5 | week_day_6 | week_day_7 |
+|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | 2020-07-01 | 2324 | daily_visits | 0 | 0 | 1 | 0 | 0 | 0 | 0 |
+| 1 | 2020-07-02 | 2201 | daily_visits | 0 | 0 | 0 | 1 | 0 | 0 | 0 |
+| 2 | 2020-07-03 | 2146 | daily_visits | 0 | 0 | 0 | 0 | 1 | 0 | 0 |
+| 3 | 2020-07-04 | 1666 | daily_visits | 0 | 0 | 0 | 0 | 0 | 1 | 0 |
+| 4 | 2020-07-05 | 1433 | daily_visits | 0 | 0 | 0 | 0 | 0 | 0 | 1 |
+| 5 | 2020-07-06 | 2195 | daily_visits | 1 | 0 | 0 | 0 | 0 | 0 | 0 |
+| 6 | 2020-07-07 | 2240 | daily_visits | 0 | 1 | 0 | 0 | 0 | 0 | 0 |
+| 7 | 2020-07-08 | 2295 | daily_visits | 0 | 0 | 1 | 0 | 0 | 0 | 0 |
+| 8 | 2020-07-09 | 2279 | daily_visits | 0 | 0 | 0 | 1 | 0 | 0 | 0 |
+| 9 | 2020-07-10 | 2155 | daily_visits | 0 | 0 | 0 | 0 | 1 | 0 | 0 |
+
+Let’s rerun the cross-validation procedure with the added exogenous
+variables.
+
+```python
+timegpt_cv_df_with_ex = nixtla_client.cross_validation(
+ df,
+ h=7,
+ n_windows=8,
+ time_col='date',
+ target_col='users',
+ freq='D',
+ level=[80, 90, 99.5]
+)
+timegpt_cv_df_with_ex.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Using the following exogenous variables: week_day_1, week_day_2, week_day_3, week_day_4, week_day_5, week_day_6, week_day_7
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Using the following exogenous variables: week_day_1, week_day_2, week_day_3, week_day_4, week_day_5, week_day_6, week_day_7
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Using the following exogenous variables: week_day_1, week_day_2, week_day_3, week_day_4, week_day_5, week_day_6, week_day_7
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Using the following exogenous variables: week_day_1, week_day_2, week_day_3, week_day_4, week_day_5, week_day_6, week_day_7
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Using the following exogenous variables: week_day_1, week_day_2, week_day_3, week_day_4, week_day_5, week_day_6, week_day_7
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Using the following exogenous variables: week_day_1, week_day_2, week_day_3, week_day_4, week_day_5, week_day_6, week_day_7
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Using the following exogenous variables: week_day_1, week_day_2, week_day_3, week_day_4, week_day_5, week_day_6, week_day_7
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Using the following exogenous variables: week_day_1, week_day_2, week_day_3, week_day_4, week_day_5, week_day_6, week_day_7
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+```
+
+| | unique_id | date | cutoff | users | TimeGPT | TimeGPT-lo-99.5 | TimeGPT-lo-90 | TimeGPT-lo-80 | TimeGPT-hi-80 | TimeGPT-hi-90 | TimeGPT-hi-99.5 |
+|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | daily_visits | 2021-07-01 | 2021-06-30 | 3123 | 3314.773743 | 2793.566942 | 3043.304261 | 3085.668122 | 3543.879364 | 3586.243226 | 3835.980544 |
+| 1 | daily_visits | 2021-07-02 | 2021-06-30 | 2870 | 3093.066529 | 2139.727892 | 2725.964112 | 2779.082154 | 3407.050904 | 3460.168946 | 4046.405166 |
+| 2 | daily_visits | 2021-07-03 | 2021-06-30 | 2020 | 2347.973573 | 1386.090529 | 1915.487550 | 1973.679628 | 2722.267519 | 2780.459596 | 3309.856618 |
+| 3 | daily_visits | 2021-07-04 | 2021-06-30 | 1828 | 2182.467408 | 1003.677454 | 1681.246491 | 1874.572327 | 2490.362488 | 2683.688324 | 3361.257361 |
+| 4 | daily_visits | 2021-07-05 | 2021-06-30 | 2722 | 3083.629453 | 1257.248435 | 2220.430357 | 2556.408628 | 3610.850279 | 3946.828550 | 4910.010472 |
+
+Let’s plot our forecasts again and calculate our error.
+
+```python
+nixtla_client.plot(
+ df,
+ timegpt_cv_df_with_ex.drop(columns=['cutoff', 'users']),
+ time_col='date',
+ target_col='users',
+ max_insample_length=90,
+ level=[80, 90, 99.5]
+)
+```
+
+
+
+```python
+mae_timegpt_with_exogenous = mae(df = timegpt_cv_df_with_ex.drop(columns=['cutoff']),
+ models=['TimeGPT'],
+ target_col='users')
+
+mae_timegpt_with_exogenous
+```
+
+| | unique_id | TimeGPT |
+|-----|--------------|-----------|
+| 0 | daily_visits | 167.22857 |
+
+To conclude, we obtain the following forecast results in this notebook:
+
+```python
+mae_timegpt['Exogenous features'] = False
+mae_timegpt_with_exogenous['Exogenous features'] = True
+
+df_results = pd.concat([mae_timegpt, mae_timegpt_with_exogenous])
+df_results = df_results.rename(columns={'TimeGPT':'MAE backtest'})
+df_results = df_results.drop(columns={'unique_id'})
+df_results['model'] = 'TimeGPT'
+
+df_results[['model', 'Exogenous features', 'MAE backtest']]
+```
+
+| | model | Exogenous features | MAE backtest |
+|-----|---------|--------------------|--------------|
+| 0 | TimeGPT | False | 167.691711 |
+| 0 | TimeGPT | True | 167.228570 |
+
+We’ve shown how to forecast daily visits of a website. We achieved
+almost 10% better forecasting results as compared to the [original
+tutorial](https://cienciadedatos.net/documentos/py37-forecasting-web-traffic-machine-learning.html),
+using significantly less lines of code, in a fraction of the time
+required to run everything.
+
+Did you notice how little effort that took? What you did not have to do,
+is:
+
+- Elaborate data preprocessing - just a table with timeseries is
+ sufficient
+- Creating a validation- and test set - TimeGPT handles the
+ cross-validation in a single function
+- Choosing and testing different models - It’s just a single call to
+ TimeGPT
+- Hyperparameter tuning - Not necessary.
+
+Happy forecasting!
+
diff --git a/nixtla/docs/use-cases/intermittent_demand.html.mdx b/nixtla/docs/use-cases/intermittent_demand.html.mdx
new file mode 100644
index 00000000..b0a0435d
--- /dev/null
+++ b/nixtla/docs/use-cases/intermittent_demand.html.mdx
@@ -0,0 +1,478 @@
+---
+output-file: intermittent_demand.html
+title: Forecasting Intermittent Demand
+---
+
+
+In this tutorial, we show how to use TimeGPT on an intermittent series
+where we have many values at zero. Here, we use a subset of the M5
+dataset that tracks the demand for food items in a Californian store.
+The dataset also includes exogenous variables like the sell price and
+the type of event occuring at a particular day.
+
+TimeGPT achieves the best performance at a MAE of 0.49, which represents
+a **14% improvement** over the best statistical model specifically built
+to handle intermittent time series data.
+
+Predicting with TimeGPT took 6.8 seconds, while fitting and predicting
+with statistical models took 5.2 seconds. TimeGPT is technically slower,
+but for a difference in time of roughly 1 second only, we get much
+better predictions with TimeGPT.
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/tutorials/4_intermittent_demand.ipynb)
+
+## Initial setup
+
+We start off by importing the required packages for this tutorial and
+create an instace of
+[`NixtlaClient`](https://Nixtla.github.io/nixtla/src/nixtla_client.html#nixtlaclient).
+
+```python
+import time
+import pandas as pd
+import numpy as np
+
+from nixtla import NixtlaClient
+
+from utilsforecast.losses import mae
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+We now read the dataset and plot it.
+
+```python
+df = pd.read_csv("https://raw.githubusercontent.com/Nixtla/transfer-learning-time-series/main/datasets/m5_sales_exog_small.csv")
+df['ds'] = pd.to_datetime(df['ds'])
+
+df.head()
+```
+
+| | unique_id | ds | y | sell_price | event_type_Cultural | event_type_National | event_type_Religious | event_type_Sporting |
+|----|----|----|----|----|----|----|----|----|
+| 0 | FOODS_1_001 | 2011-01-29 | 3 | 2.0 | 0 | 0 | 0 | 0 |
+| 1 | FOODS_1_001 | 2011-01-30 | 0 | 2.0 | 0 | 0 | 0 | 0 |
+| 2 | FOODS_1_001 | 2011-01-31 | 0 | 2.0 | 0 | 0 | 0 | 0 |
+| 3 | FOODS_1_001 | 2011-02-01 | 1 | 2.0 | 0 | 0 | 0 | 0 |
+| 4 | FOODS_1_001 | 2011-02-02 | 4 | 2.0 | 0 | 0 | 0 | 0 |
+
+```python
+nixtla_client.plot(
+ df,
+ max_insample_length=365,
+)
+```
+
+
+
+In the figure above, we can see the intermittent nature of this dataset,
+with many periods with zero demand.
+
+Now, let’s use TimeGPT to forecast the demand of each product.
+
+## Bounded forecasts
+
+To avoid getting negative predictions coming from the model, we use a
+log transformation on the data. That way, the model will be forced to
+predict only positive values.
+
+Note that due to the presence of zeros in our dataset, we add one to all
+points before taking the log.
+
+```python
+df_transformed = df.copy()
+
+df_transformed['y'] = np.log(df_transformed['y']+1)
+
+df_transformed.head()
+```
+
+| | unique_id | ds | y | sell_price | event_type_Cultural | event_type_National | event_type_Religious | event_type_Sporting |
+|----|----|----|----|----|----|----|----|----|
+| 0 | FOODS_1_001 | 2011-01-29 | 1.386294 | 2.0 | 0 | 0 | 0 | 0 |
+| 1 | FOODS_1_001 | 2011-01-30 | 0.000000 | 2.0 | 0 | 0 | 0 | 0 |
+| 2 | FOODS_1_001 | 2011-01-31 | 0.000000 | 2.0 | 0 | 0 | 0 | 0 |
+| 3 | FOODS_1_001 | 2011-02-01 | 0.693147 | 2.0 | 0 | 0 | 0 | 0 |
+| 4 | FOODS_1_001 | 2011-02-02 | 1.609438 | 2.0 | 0 | 0 | 0 | 0 |
+
+Now, let’s keep the last 28 time steps for the test set and use the rest
+as input to the model.
+
+```python
+test_df = df_transformed.groupby('unique_id').tail(28)
+
+input_df = df_transformed.drop(test_df.index).reset_index(drop=True)
+```
+
+## Forecasting with TimeGPT
+
+```python
+start = time.time()
+
+fcst_df = nixtla_client.forecast(
+ df=input_df,
+ h=28,
+ level=[80], # Generate a 80% confidence interval
+ finetune_steps=10, # Specify the number of steps for fine-tuning
+ finetune_loss='mae', # Use the MAE as the loss function for fine-tuning
+ model='timegpt-1-long-horizon', # Use the model for long-horizon forecasting
+ time_col='ds',
+ target_col='y',
+ id_col='unique_id'
+)
+
+end = time.time()
+
+timegpt_duration = end - start
+
+print(f"Time (TimeGPT): {timegpt_duration}")
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: D
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+``` text
+Time (TimeGPT): 6.164413213729858
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+Great! TimeGPT was done in **5.8 seconds** and we now have predictions.
+However, those predictions are transformed, so we need to inverse the
+transformation to get back to the orignal scale. Therefore, we take the
+exponential and subtract one from each data point.
+
+```python
+cols = [col for col in fcst_df.columns if col not in ['ds', 'unique_id']]
+
+for col in cols:
+ fcst_df[col] = np.exp(fcst_df[col])-1
+
+fcst_df.head()
+```
+
+| | unique_id | ds | TimeGPT | TimeGPT-lo-80 | TimeGPT-hi-80 |
+|-----|-------------|------------|----------|---------------|---------------|
+| 0 | FOODS_1_001 | 2016-05-23 | 0.286841 | -0.267101 | 1.259465 |
+| 1 | FOODS_1_001 | 2016-05-24 | 0.320482 | -0.241236 | 1.298046 |
+| 2 | FOODS_1_001 | 2016-05-25 | 0.287392 | -0.362250 | 1.598791 |
+| 3 | FOODS_1_001 | 2016-05-26 | 0.295326 | -0.145489 | 0.963542 |
+| 4 | FOODS_1_001 | 2016-05-27 | 0.315868 | -0.166516 | 1.077437 |
+
+## Evaluation
+
+Before measuring the performance metric, let’s plot the predictions
+against the actual values.
+
+```python
+nixtla_client.plot(test_df, fcst_df, models=['TimeGPT'], level=[80], time_col='ds', target_col='y')
+```
+
+
+
+Finally, we can measure the mean absolute error (MAE) of the model.
+
+```python
+fcst_df['ds'] = pd.to_datetime(fcst_df['ds'])
+
+test_df = pd.merge(test_df, fcst_df, 'left', ['unique_id', 'ds'])
+```
+
+
+```python
+evaluation = evaluate(
+ test_df,
+ metrics=[mae],
+ models=["TimeGPT"],
+ target_col="y",
+ id_col='unique_id'
+)
+
+average_metrics = evaluation.groupby('metric')['TimeGPT'].mean()
+average_metrics
+```
+
+``` text
+metric
+mae 0.492559
+Name: TimeGPT, dtype: float64
+```
+
+## Forecasting with statistical models
+
+The library `statsforecast` by Nixtla provides a suite of statistical
+models specifically built for intermittent forecasting, such as Croston,
+IMAPA and TSB. Let’s use these models and see how they perform against
+TimeGPT.
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import CrostonClassic, CrostonOptimized, IMAPA, TSB
+```
+
+Here, we use four models: two versions of Croston, IMAPA and TSB.
+
+```python
+models = [CrostonClassic(), CrostonOptimized(), IMAPA(), TSB(0.1, 0.1)]
+
+sf = StatsForecast(
+ models=models,
+ freq='D',
+ n_jobs=-1
+)
+```
+
+Then, we can fit the models on our data.
+
+```python
+start = time.time()
+
+sf.fit(df=input_df)
+
+sf_preds = sf.predict(h=28)
+
+end = time.time()
+
+sf_duration = end - start
+
+print(f"Statistical models took :{sf_duration}s")
+```
+
+Here, fitting and predicting with four statistical models took 5.2
+seconds, while TimeGPT took 5.8 seconds, so TimeGPT was only 0.6 seconds
+slower.
+
+Again, we need to inverse the transformation. Remember that the training
+data was previously transformed using the log function.
+
+```python
+cols = [col for col in sf_preds.columns if col not in ['ds', 'unique_id']]
+
+for col in cols:
+ sf_preds[col] = np.exp(sf_preds[col])-1
+
+sf_preds.head()
+```
+
+| | ds | CrostonClassic | CrostonOptimized | IMAPA | TSB |
+|-------------|------------|----------------|------------------|----------|----------|
+| unique_id | | | | | |
+| FOODS_1_001 | 2016-05-23 | 0.599093 | 0.599093 | 0.445779 | 0.396258 |
+| FOODS_1_001 | 2016-05-24 | 0.599093 | 0.599093 | 0.445779 | 0.396258 |
+| FOODS_1_001 | 2016-05-25 | 0.599093 | 0.599093 | 0.445779 | 0.396258 |
+| FOODS_1_001 | 2016-05-26 | 0.599093 | 0.599093 | 0.445779 | 0.396258 |
+| FOODS_1_001 | 2016-05-27 | 0.599093 | 0.599093 | 0.445779 | 0.396258 |
+
+## Evaluation
+
+Now, let’s combine the predictions from all methods and see which
+performs best.
+
+```python
+test_df = pd.merge(test_df, sf_preds, 'left', ['unique_id', 'ds'])
+test_df.head()
+```
+
+| | unique_id | ds | y | sell_price | event_type_Cultural | event_type_National | event_type_Religious | event_type_Sporting | TimeGPT | TimeGPT-lo-80 | TimeGPT-hi-80 | CrostonClassic | CrostonOptimized | IMAPA | TSB |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | FOODS_1_001 | 2016-05-23 | 1.386294 | 2.24 | 0 | 0 | 0 | 0 | 0.286841 | -0.267101 | 1.259465 | 0.599093 | 0.599093 | 0.445779 | 0.396258 |
+| 1 | FOODS_1_001 | 2016-05-24 | 0.000000 | 2.24 | 0 | 0 | 0 | 0 | 0.320482 | -0.241236 | 1.298046 | 0.599093 | 0.599093 | 0.445779 | 0.396258 |
+| 2 | FOODS_1_001 | 2016-05-25 | 0.000000 | 2.24 | 0 | 0 | 0 | 0 | 0.287392 | -0.362250 | 1.598791 | 0.599093 | 0.599093 | 0.445779 | 0.396258 |
+| 3 | FOODS_1_001 | 2016-05-26 | 0.000000 | 2.24 | 0 | 0 | 0 | 0 | 0.295326 | -0.145489 | 0.963542 | 0.599093 | 0.599093 | 0.445779 | 0.396258 |
+| 4 | FOODS_1_001 | 2016-05-27 | 1.945910 | 2.24 | 0 | 0 | 0 | 0 | 0.315868 | -0.166516 | 1.077437 | 0.599093 | 0.599093 | 0.445779 | 0.396258 |
+
+```python
+evaluation = evaluate(
+ test_df,
+ metrics=[mae],
+ models=["TimeGPT", "CrostonClassic", "CrostonOptimized", "IMAPA", "TSB"],
+ target_col="y",
+ id_col='unique_id'
+)
+
+average_metrics = evaluation.groupby('metric')[["TimeGPT", "CrostonClassic", "CrostonOptimized", "IMAPA", "TSB"]].mean()
+average_metrics
+```
+
+| | TimeGPT | CrostonClassic | CrostonOptimized | IMAPA | TSB |
+|--------|----------|----------------|------------------|----------|----------|
+| metric | | | | | |
+| mae | 0.492559 | 0.564563 | 0.580922 | 0.571943 | 0.567178 |
+
+In the table above, we can see that TimeGPT achieves the lowest MAE,
+achieving a 12.8% improvement over the best performing statistical
+model.
+
+Now, this was done without using any of the available exogenous
+features. While the statsitical models do not support them, let’s try
+including them in TimeGPT.
+
+## Forecasting with exogenous variables using TimeGPT
+
+To forecast with exogenous variables, we need to specify their future
+values over the forecast horizon. Therefore, let’s simply take the types
+of events, as those dates are known in advance.
+
+```python
+futr_exog_df = test_df.drop(["TimeGPT", "CrostonClassic", "CrostonOptimized", "IMAPA", "TSB", "y", "TimeGPT-lo-80", "TimeGPT-hi-80", "sell_price"], axis=1)
+futr_exog_df.head()
+```
+
+| | unique_id | ds | event_type_Cultural | event_type_National | event_type_Religious | event_type_Sporting |
+|----|----|----|----|----|----|----|
+| 0 | FOODS_1_001 | 2016-05-23 | 0 | 0 | 0 | 0 |
+| 1 | FOODS_1_001 | 2016-05-24 | 0 | 0 | 0 | 0 |
+| 2 | FOODS_1_001 | 2016-05-25 | 0 | 0 | 0 | 0 |
+| 3 | FOODS_1_001 | 2016-05-26 | 0 | 0 | 0 | 0 |
+| 4 | FOODS_1_001 | 2016-05-27 | 0 | 0 | 0 | 0 |
+
+Then, we simply call the `forecast` method and pass the `futr_exog_df`
+in the `X_df` parameter.
+
+```python
+start = time.time()
+
+fcst_df = nixtla_client.forecast(
+ df=input_df,
+ X_df=futr_exog_df,
+ h=28,
+ level=[80], # Generate a 80% confidence interval
+ finetune_steps=10, # Specify the number of steps for fine-tuning
+ finetune_loss='mae', # Use the MAE as the loss function for fine-tuning
+ model='timegpt-1-long-horizon', # Use the model for long-horizon forecasting
+ time_col='ds',
+ target_col='y',
+ id_col='unique_id'
+)
+
+end = time.time()
+
+timegpt_duration = end - start
+
+print(f"Time (TimeGPT): {timegpt_duration}")
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: D
+INFO:nixtla.nixtla_client:Using the following exogenous variables: event_type_Cultural, event_type_National, event_type_Religious, event_type_Sporting
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+``` text
+Time (TimeGPT): 7.173351287841797
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+Great! Remember that the predictions are transformed, so we have to
+inverse the transformation again.
+
+```python
+fcst_df.rename(columns={
+ 'TimeGPT': 'TimeGPT_ex',
+}, inplace=True)
+
+cols = [col for col in fcst_df.columns if col not in ['ds', 'unique_id']]
+
+for col in cols:
+ fcst_df[col] = np.exp(fcst_df[col])-1
+
+fcst_df.head()
+```
+
+| | unique_id | ds | TimeGPT_ex | TimeGPT-lo-80 | TimeGPT-hi-80 |
+|-----|-------------|------------|------------|---------------|---------------|
+| 0 | FOODS_1_001 | 2016-05-23 | 0.281922 | -0.269902 | 1.250828 |
+| 1 | FOODS_1_001 | 2016-05-24 | 0.313774 | -0.245091 | 1.286372 |
+| 2 | FOODS_1_001 | 2016-05-25 | 0.285639 | -0.363119 | 1.595252 |
+| 3 | FOODS_1_001 | 2016-05-26 | 0.295037 | -0.145679 | 0.963104 |
+| 4 | FOODS_1_001 | 2016-05-27 | 0.315484 | -0.166760 | 1.076830 |
+
+## Evaluation
+
+Finally, let’s evaluate the performance of TimeGPT with exogenous
+features.
+
+```python
+test_df['TimeGPT_ex'] = fcst_df['TimeGPT_ex'].values
+test_df.head()
+```
+
+| | unique_id | ds | y | sell_price | event_type_Cultural | event_type_National | event_type_Religious | event_type_Sporting | TimeGPT | TimeGPT-lo-80 | TimeGPT-hi-80 | CrostonClassic | CrostonOptimized | IMAPA | TSB | TimeGPT_ex |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | FOODS_1_001 | 2016-05-23 | 1.386294 | 2.24 | 0 | 0 | 0 | 0 | 0.286841 | -0.267101 | 1.259465 | 0.599093 | 0.599093 | 0.445779 | 0.396258 | 0.281922 |
+| 1 | FOODS_1_001 | 2016-05-24 | 0.000000 | 2.24 | 0 | 0 | 0 | 0 | 0.320482 | -0.241236 | 1.298046 | 0.599093 | 0.599093 | 0.445779 | 0.396258 | 0.313774 |
+| 2 | FOODS_1_001 | 2016-05-25 | 0.000000 | 2.24 | 0 | 0 | 0 | 0 | 0.287392 | -0.362250 | 1.598791 | 0.599093 | 0.599093 | 0.445779 | 0.396258 | 0.285639 |
+| 3 | FOODS_1_001 | 2016-05-26 | 0.000000 | 2.24 | 0 | 0 | 0 | 0 | 0.295326 | -0.145489 | 0.963542 | 0.599093 | 0.599093 | 0.445779 | 0.396258 | 0.295037 |
+| 4 | FOODS_1_001 | 2016-05-27 | 1.945910 | 2.24 | 0 | 0 | 0 | 0 | 0.315868 | -0.166516 | 1.077437 | 0.599093 | 0.599093 | 0.445779 | 0.396258 | 0.315484 |
+
+```python
+evaluation = evaluate(
+ test_df,
+ metrics=[mae],
+ models=["TimeGPT", "CrostonClassic", "CrostonOptimized", "IMAPA", "TSB", "TimeGPT_ex"],
+ target_col="y",
+ id_col='unique_id'
+)
+
+average_metrics = evaluation.groupby('metric')[["TimeGPT", "CrostonClassic", "CrostonOptimized", "IMAPA", "TSB", "TimeGPT_ex"]].mean()
+average_metrics
+```
+
+| | TimeGPT | CrostonClassic | CrostonOptimized | IMAPA | TSB | TimeGPT_ex |
+|--------|----------|----------------|------------------|----------|----------|------------|
+| metric | | | | | | |
+| mae | 0.492559 | 0.564563 | 0.580922 | 0.571943 | 0.567178 | 0.485352 |
+
+From the table above, we can see that using exogenous features improved
+the performance of TimeGPT. Now, it represents a 14% improvement over
+the best statistical model.
+
+Using TimeGPT with exogenous features took 6.8 seconds. This is 1.6
+seconds slower than statitstical models, but it resulted in much better
+predictions.
+
diff --git a/nixtla/docs/use-cases/what_if_pricing_scenarios_in_retail.html.mdx b/nixtla/docs/use-cases/what_if_pricing_scenarios_in_retail.html.mdx
new file mode 100644
index 00000000..0056da86
--- /dev/null
+++ b/nixtla/docs/use-cases/what_if_pricing_scenarios_in_retail.html.mdx
@@ -0,0 +1,407 @@
+---
+output-file: what_if_pricing_scenarios_in_retail.html
+title: What if? Forecasting price effects in retail
+---
+
+
+You can use TimeGPT to forecast a set of timeseries, for example the
+demand of a product in retail. But what if you want to evaluate
+different pricing scenarios for that product? Performing such a scenario
+analysis can help you better understand how pricing affects product
+demand and can aid in decision making.
+
+In this example, we will show you: \* How you can use TimeGPT to
+forecast product demand using price as an exogenous variable \* How you
+can evaluate different pricing scenarios that affect product demand
+
+[](https://colab.research.google.com/github/Nixtla/nixtla/blob/main/nbs/docs/use-cases/5_what_if_pricing_scenarios_in_retail.ipynb)
+
+## 1. Import packages
+
+First, we import the required packages and initialize the Nixtla client.
+
+```python
+import pandas as pd
+import os
+
+from nixtla import NixtlaClient
+from datasetsforecast.m5 import M5
+```
+
+
+```python
+nixtla_client = NixtlaClient(
+ # defaults to os.environ.get("NIXTLA_API_KEY")
+ api_key = 'my_api_key_provided_by_nixtla'
+)
+```
+
+> 👍 Use an Azure AI endpoint
+>
+> To use an Azure AI endpoint, remember to set also the `base_url`
+> argument:
+>
+> `nixtla_client = NixtlaClient(base_url="you azure ai endpoint", api_key="your api_key")`
+
+## 2. Load M5 data
+
+Let’s see an example on predicting sales of products of the [M5
+dataset](https://nixtlaverse.nixtla.io/datasetsforecast/m5.html). The M5
+dataset contains daily product demand (sales) for 10 retail stores in
+the US.
+
+First, we load the data using `datasetsforecast`. This returns:
+
+- `Y_df`, containing the sales (`y` column), for each unique product
+ (`unique_id` column) at every timestamp (`ds` column).
+- `X_df`, containing additional relevant information for each unique
+ product (`unique_id` column) at every timestamp (`ds` column).
+
+> **Tip**
+>
+> You can find a tutorial on including exogenous variables in your
+> forecast with TimeGPT [here](../../docs/tutorials-timegpt_quickstart).
+
+```python
+Y_df, X_df, S_df = M5.load(directory=os.getcwd())
+
+Y_df.head(10)
+```
+
+``` text
+100%|██████████| 50.2M/50.2M [00:00<00:00, 58.1MiB/s]
+INFO:datasetsforecast.utils:Successfully downloaded m5.zip, 50219189, bytes.
+INFO:datasetsforecast.utils:Decompressing zip file...
+INFO:datasetsforecast.utils:Successfully decompressed c:\Users\ospra\OneDrive\Nixtla\Repositories\nixtla\m5\datasets\m5.zip
+c:\Users\ospra\miniconda3\envs\nixtla\lib\site-packages\datasetsforecast\m5.py:143: FutureWarning: The default of observed=False is deprecated and will be changed to True in a future version of pandas. Pass observed=False to retain current behavior or observed=True to adopt the future default and silence this warning.
+ keep_mask = long.groupby('id')['y'].transform(first_nz_mask, engine='numba')
+```
+
+| | unique_id | ds | y |
+|-----|------------------|------------|-----|
+| 0 | FOODS_1_001_CA_1 | 2011-01-29 | 3.0 |
+| 1 | FOODS_1_001_CA_1 | 2011-01-30 | 0.0 |
+| 2 | FOODS_1_001_CA_1 | 2011-01-31 | 0.0 |
+| 3 | FOODS_1_001_CA_1 | 2011-02-01 | 1.0 |
+| 4 | FOODS_1_001_CA_1 | 2011-02-02 | 4.0 |
+| 5 | FOODS_1_001_CA_1 | 2011-02-03 | 2.0 |
+| 6 | FOODS_1_001_CA_1 | 2011-02-04 | 0.0 |
+| 7 | FOODS_1_001_CA_1 | 2011-02-05 | 2.0 |
+| 8 | FOODS_1_001_CA_1 | 2011-02-06 | 0.0 |
+| 9 | FOODS_1_001_CA_1 | 2011-02-07 | 0.0 |
+
+For this example, we will only keep the additional relevant information
+from the column `sell_price`. This column shows the selling price of the
+product, and we expect demand to fluctuate given a different selling
+price.
+
+```python
+X_df = X_df[['unique_id', 'ds', 'sell_price']]
+
+X_df.head(10)
+```
+
+| | unique_id | ds | sell_price |
+|-----|------------------|------------|------------|
+| 0 | FOODS_1_001_CA_1 | 2011-01-29 | 2.0 |
+| 1 | FOODS_1_001_CA_1 | 2011-01-30 | 2.0 |
+| 2 | FOODS_1_001_CA_1 | 2011-01-31 | 2.0 |
+| 3 | FOODS_1_001_CA_1 | 2011-02-01 | 2.0 |
+| 4 | FOODS_1_001_CA_1 | 2011-02-02 | 2.0 |
+| 5 | FOODS_1_001_CA_1 | 2011-02-03 | 2.0 |
+| 6 | FOODS_1_001_CA_1 | 2011-02-04 | 2.0 |
+| 7 | FOODS_1_001_CA_1 | 2011-02-05 | 2.0 |
+| 8 | FOODS_1_001_CA_1 | 2011-02-06 | 2.0 |
+| 9 | FOODS_1_001_CA_1 | 2011-02-07 | 2.0 |
+
+## 3. Forecasting demand using price as an exogenous variable
+
+We will forecast the demand for a single product only, for all 10 retail
+stores in the dataset. We choose a food product with many price changes
+identified by `FOODS_1_129_`.
+
+```python
+products = [
+ 'FOODS_1_129_CA_1',
+ 'FOODS_1_129_CA_2',
+ 'FOODS_1_129_CA_3',
+ 'FOODS_1_129_CA_4',
+ 'FOODS_1_129_TX_1',
+ 'FOODS_1_129_TX_2',
+ 'FOODS_1_129_TX_3',
+ 'FOODS_1_129_WI_1',
+ 'FOODS_1_129_WI_2',
+ 'FOODS_1_129_WI_3'
+ ]
+Y_df_product = Y_df.query('unique_id in @products')
+X_df_product = X_df.query('unique_id in @products')
+```
+
+We merge our two dataframes to create the dataset to be used in TimeGPT.
+
+```python
+df = Y_df_product.merge(X_df_product)
+
+df.head(10)
+```
+
+| | unique_id | ds | y | sell_price |
+|-----|------------------|------------|-----|------------|
+| 0 | FOODS_1_129_CA_1 | 2011-02-01 | 1.0 | 6.22 |
+| 1 | FOODS_1_129_CA_1 | 2011-02-02 | 0.0 | 6.22 |
+| 2 | FOODS_1_129_CA_1 | 2011-02-03 | 0.0 | 6.22 |
+| 3 | FOODS_1_129_CA_1 | 2011-02-04 | 0.0 | 6.22 |
+| 4 | FOODS_1_129_CA_1 | 2011-02-05 | 1.0 | 6.22 |
+| 5 | FOODS_1_129_CA_1 | 2011-02-06 | 0.0 | 6.22 |
+| 6 | FOODS_1_129_CA_1 | 2011-02-07 | 0.0 | 6.22 |
+| 7 | FOODS_1_129_CA_1 | 2011-02-08 | 0.0 | 6.22 |
+| 8 | FOODS_1_129_CA_1 | 2011-02-09 | 0.0 | 6.22 |
+| 9 | FOODS_1_129_CA_1 | 2011-02-10 | 3.0 | 6.22 |
+
+Let’s investigate how the demand - our target `y` - of these products
+has evolved in the last year of data. We see that in the California
+stores (with a `CA_` suffix), the product has sold intermittently,
+whereas in the other regions (`TX` and `WY`) sales where less
+intermittent. Note that the plot only shows 8 (out of 10) stores.
+
+```python
+nixtla_client.plot(df,
+ unique_ids=products,
+ max_insample_length=365)
+```
+
+
+
+Next, we look at the `sell_price` of these products across the entire
+data available. We find that there have been relatively few price
+changes - about 20 in total - over the period 2011 - 2016. Note that the
+plot only shows 8 (out of 10) stores.
+
+```python
+nixtla_client.plot(df,
+ unique_ids=products,
+ target_col='sell_price')
+```
+
+
+
+Let’s turn to our forecasting task. We will forecast the last 28 days in
+the dataset.
+
+To use the `sell_price` exogenous variable in TimeGPT, we have to add it
+as future values. Therefore, we create a future values dataframe, that
+contains the `unique_id`, the timestamp `ds`, and `sell_price`.
+
+```python
+future_ex_vars_df = df.drop(columns = ['y'])
+future_ex_vars_df = future_ex_vars_df.query("ds >= '2016-05-23'")
+
+future_ex_vars_df.head(10)
+```
+
+| | unique_id | ds | sell_price |
+|------|------------------|------------|------------|
+| 1938 | FOODS_1_129_CA_1 | 2016-05-23 | 5.74 |
+| 1939 | FOODS_1_129_CA_1 | 2016-05-24 | 5.74 |
+| 1940 | FOODS_1_129_CA_1 | 2016-05-25 | 5.74 |
+| 1941 | FOODS_1_129_CA_1 | 2016-05-26 | 5.74 |
+| 1942 | FOODS_1_129_CA_1 | 2016-05-27 | 5.74 |
+| 1943 | FOODS_1_129_CA_1 | 2016-05-28 | 5.74 |
+| 1944 | FOODS_1_129_CA_1 | 2016-05-29 | 5.74 |
+| 1945 | FOODS_1_129_CA_1 | 2016-05-30 | 5.74 |
+| 1946 | FOODS_1_129_CA_1 | 2016-05-31 | 5.74 |
+| 1947 | FOODS_1_129_CA_1 | 2016-06-01 | 5.74 |
+
+Next, we limit our input dataframe to all but the 28 forecast days:
+
+```python
+df_train = df.query("ds < '2016-05-23'")
+
+df_train.tail(10)
+```
+
+| | unique_id | ds | y | sell_price |
+|-------|------------------|------------|-----|------------|
+| 19640 | FOODS_1_129_WI_3 | 2016-05-13 | 3.0 | 7.23 |
+| 19641 | FOODS_1_129_WI_3 | 2016-05-14 | 1.0 | 7.23 |
+| 19642 | FOODS_1_129_WI_3 | 2016-05-15 | 2.0 | 7.23 |
+| 19643 | FOODS_1_129_WI_3 | 2016-05-16 | 3.0 | 7.23 |
+| 19644 | FOODS_1_129_WI_3 | 2016-05-17 | 1.0 | 7.23 |
+| 19645 | FOODS_1_129_WI_3 | 2016-05-18 | 2.0 | 7.23 |
+| 19646 | FOODS_1_129_WI_3 | 2016-05-19 | 3.0 | 7.23 |
+| 19647 | FOODS_1_129_WI_3 | 2016-05-20 | 1.0 | 7.23 |
+| 19648 | FOODS_1_129_WI_3 | 2016-05-21 | 0.0 | 7.23 |
+| 19649 | FOODS_1_129_WI_3 | 2016-05-22 | 0.0 | 7.23 |
+
+Let’s call the `forecast` method of TimeGPT:
+
+```python
+timegpt_fcst_df = nixtla_client.forecast(df=df_train, X_df=future_ex_vars_df, h=28)
+timegpt_fcst_df.head()
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: D
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon. This may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Using the following exogenous variables: sell_price
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+| | unique_id | ds | TimeGPT |
+|-----|------------------|------------|----------|
+| 0 | FOODS_1_129_CA_1 | 2016-05-23 | 0.875594 |
+| 1 | FOODS_1_129_CA_1 | 2016-05-24 | 0.777731 |
+| 2 | FOODS_1_129_CA_1 | 2016-05-25 | 0.786871 |
+| 3 | FOODS_1_129_CA_1 | 2016-05-26 | 0.828223 |
+| 4 | FOODS_1_129_CA_1 | 2016-05-27 | 0.791228 |
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+We plot the forecast, the actuals and the last 28 days before the
+forecast period:
+
+```python
+nixtla_client.plot(
+ df[['unique_id', 'ds', 'y']],
+ timegpt_fcst_df,
+ max_insample_length=56,
+)
+```
+
+
+
+## 4. What if? Varying price when forecasting demand
+
+What happens when we change the price of the products in our forecast
+period? Let’s see how our forecast changes when we increase and decrease
+the `sell_price` by 5%.
+
+```python
+price_change = 0.05
+
+# Plus
+future_ex_vars_df_plus= future_ex_vars_df.copy()
+future_ex_vars_df_plus["sell_price"] = future_ex_vars_df_plus["sell_price"] * (1 + price_change)
+# Minus
+future_ex_vars_df_minus = future_ex_vars_df.copy()
+future_ex_vars_df_minus["sell_price"] = future_ex_vars_df_minus["sell_price"] * (1 - price_change)
+```
+
+Let’s create a new set of forecasts with TimeGPT.
+
+```python
+timegpt_fcst_df_plus = nixtla_client.forecast(df=df_train, X_df=future_ex_vars_df_plus, h=28)
+timegpt_fcst_df_minus = nixtla_client.forecast(df=df_train, X_df=future_ex_vars_df_minus, h=28)
+```
+
+``` text
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: D
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon. This may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Using the following exogenous variables: sell_price
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+INFO:nixtla.nixtla_client:Validating inputs...
+INFO:nixtla.nixtla_client:Preprocessing dataframes...
+INFO:nixtla.nixtla_client:Inferred freq: D
+WARNING:nixtla.nixtla_client:The specified horizon "h" exceeds the model horizon. This may lead to less accurate forecasts. Please consider using a smaller horizon.
+INFO:nixtla.nixtla_client:Using the following exogenous variables: sell_price
+INFO:nixtla.nixtla_client:Calling Forecast Endpoint...
+```
+
+> 📘 Available models in Azure AI
+>
+> If you are using an Azure AI endpoint, please be sure to set
+> `model="azureai"`:
+>
+> `nixtla_client.forecast(..., model="azureai")`
+>
+> For the public API, we support two models: `timegpt-1` and
+> `timegpt-1-long-horizon`.
+>
+> By default, `timegpt-1` is used. Please see [this
+> tutorial](https://docs.nixtla.io/docs/tutorials-long_horizon_forecasting)
+> on how and when to use `timegpt-1-long-horizon`.
+
+Let’s combine our three forecasts. We see that - as we expect - demand
+is expected to slightly increase (decrease) if we reduce (increase) the
+price. In other words, a cheaper product leads to higher sales and vice
+versa.
+
+> **Note**
+>
+> *Price elasticity* is a measure of how sensitive the (product) demand
+> is to a change in price. Read more about it
+> [here](https://en.wikipedia.org/wiki/Price_elasticity_of_demand).
+
+```python
+timegpt_fcst_df_plus = timegpt_fcst_df_plus.rename(columns={'TimeGPT':f'TimeGPT-sell_price_plus_{price_change * 100:.0f}%'})
+timegpt_fcst_df_minus = timegpt_fcst_df_minus.rename(columns={'TimeGPT':f'TimeGPT-sell_price_minus_{price_change * 100:.0f}%'})
+
+timegpt_fcst_df = pd.concat([timegpt_fcst_df,
+ timegpt_fcst_df_plus[f'TimeGPT-sell_price_plus_{price_change * 100:.0f}%'],
+ timegpt_fcst_df_minus[f'TimeGPT-sell_price_minus_{price_change * 100:.0f}%']], axis=1)
+
+timegpt_fcst_df.head(10)
+```
+
+| | unique_id | ds | TimeGPT | TimeGPT-sell_price_plus_5% | TimeGPT-sell_price_minus_5% |
+|----|----|----|----|----|----|
+| 0 | FOODS_1_129_CA_1 | 2016-05-23 | 0.875594 | 0.847006 | 1.370029 |
+| 1 | FOODS_1_129_CA_1 | 2016-05-24 | 0.777731 | 0.749142 | 1.272166 |
+| 2 | FOODS_1_129_CA_1 | 2016-05-25 | 0.786871 | 0.758283 | 1.281306 |
+| 3 | FOODS_1_129_CA_1 | 2016-05-26 | 0.828223 | 0.799635 | 1.322658 |
+| 4 | FOODS_1_129_CA_1 | 2016-05-27 | 0.791228 | 0.762640 | 1.285663 |
+| 5 | FOODS_1_129_CA_1 | 2016-05-28 | 0.819133 | 0.790545 | 1.313568 |
+| 6 | FOODS_1_129_CA_1 | 2016-05-29 | 0.839992 | 0.811404 | 1.334427 |
+| 7 | FOODS_1_129_CA_1 | 2016-05-30 | 0.843070 | 0.814481 | 1.337505 |
+| 8 | FOODS_1_129_CA_1 | 2016-05-31 | 0.833089 | 0.804500 | 1.327524 |
+| 9 | FOODS_1_129_CA_1 | 2016-06-01 | 0.855032 | 0.826443 | 1.349467 |
+
+Finally, let’s plot the forecasts for our different pricing scenarios,
+showing how TimeGPT forecasts a different demand when the price of a set
+of products is changed. In the graphs we can see that for specific
+products for certain periods the discount increases expected demand,
+while during other periods and for other products, price change has a
+smaller effect on total demand.
+
+```python
+nixtla_client.plot(
+ df[['unique_id', 'ds', 'y']],
+ timegpt_fcst_df,
+ max_insample_length=56,
+)
+```
+
+
+
+In this example, we have shown you: \* How you can use TimeGPT to
+forecast product demand using price as an exogenous variable \* How you
+can evaluate different pricing scenarios that affect product demand
+
+> **Important**
+>
+> - This method assumes that historical demand and price behaviour is
+> predictive of future demand, and omits other factors affecting
+> demand. To include these other factors, use additional exogenous
+> variables that provide the model with more context about the
+> factors influencing demand.
+> - This method is sensitive to unmodelled events that affect the
+> demand, such as sudden market shifts. To include those, use
+> additional exogenous variables indicating such sudden shifts if
+> they have been observed in the past too.
+
diff --git a/nixtla/favicon.svg b/nixtla/favicon.svg
new file mode 100644
index 00000000..e5f33342
--- /dev/null
+++ b/nixtla/favicon.svg
@@ -0,0 +1,5 @@
+
diff --git a/nixtla/light.png b/nixtla/light.png
new file mode 100644
index 00000000..bbb99b54
Binary files /dev/null and b/nixtla/light.png differ
diff --git a/nixtla/mint.json b/nixtla/mint.json
new file mode 100644
index 00000000..420a9f3a
--- /dev/null
+++ b/nixtla/mint.json
@@ -0,0 +1,165 @@
+{
+ "$schema": "https://mintlify.com/schema.json",
+ "name": "Nixtla",
+ "logo": {
+ "light": "/light.png",
+ "dark": "/dark.png"
+ },
+ "favicon": "/favicon.svg",
+ "colors": {
+ "primary": "#0E0E0E",
+ "light": "#FAFAFA",
+ "dark": "#0E0E0E",
+ "anchors": {
+ "from": "#2AD0CA",
+ "to": "#0E00F8"
+ }
+ },
+ "topbarCtaButton": {
+ "type": "github",
+ "url": "https://github.com/Nixtla/nixtla"
+ },
+ "navigation": [
+ {
+ "group": "Getting Started",
+ "pages": [
+ "docs/getting-started/introduction.html",
+ "docs/getting-started/quickstart.html",
+ "docs/getting-started/polars_quickstart.html",
+ "docs/getting-started/azure_quickstart.html",
+ "docs/getting-started/setting_up_your_api_key.html",
+ "docs/getting-started/pricing.html",
+ "docs/getting-started/data_requirements.html",
+ "docs/getting-started/faq.html",
+ "docs/getting-started/glossary.html",
+ "docs/getting-started/why_timegpt.html"
+ ]
+ },
+ {
+ "group": "Capabilities",
+ "pages": [
+ {
+ "group": "Forecast",
+ "pages": [
+ "docs/capabilities/forecast/quickstart.html",
+ "docs/capabilities/forecast/exogenous_variables.html",
+ "docs/capabilities/forecast/holidays_special_dates.html",
+ "docs/capabilities/forecast/categorical_variables.html",
+ "docs/capabilities/forecast/longhorizon.html",
+ "docs/capabilities/forecast/multiple_series.html",
+ "docs/capabilities/forecast/finetuning.html",
+ "docs/capabilities/forecast/custom_loss_function.html",
+ "docs/capabilities/forecast/cross_validation.html",
+ "docs/capabilities/forecast/prediction_intervals.html",
+ "docs/capabilities/forecast/irregular_timestamps.html"
+ ]
+ },
+ {
+ "group": "Historical Anomaly Detection",
+ "pages": [
+ "docs/capabilities/historical-anomaly-detection/quickstart.html",
+ "docs/capabilities/historical-anomaly-detection/anomaly_exogenous.html",
+ "docs/capabilities/historical-anomaly-detection/anomaly_detection_date_features.html",
+ "docs/capabilities/historical-anomaly-detection/confidence_levels.html"
+ ]
+ },
+ {
+ "group": "Online Anomaly Detection",
+ "pages": [
+ "docs/capabilities/online-anomaly-detection/quickstart.html",
+ "docs/capabilities/online-anomaly-detection/adjusting_detection_process.html",
+ "docs/capabilities/online-anomaly-detection/univariate_vs_multivariate_anomaly_detection.html"
+ ]
+ }
+ ]
+ },
+ {
+ "group": "Deployment",
+ "pages": [
+ "docs/deployment/azure_ai.html"
+ ]
+ },
+ {
+ "group": "Tutorials",
+ "pages": [
+ "docs/tutorials/anomaly_detection.html",
+ {
+ "group":"Exogenous variables",
+ "pages":[
+ "docs/tutorials/exogenous_variables.html",
+ "docs/tutorials/holidays.html",
+ "docs/tutorials/categorical_variables.html",
+ "docs/tutorials/shap_values.html"
+ ]
+ },
+ {
+ "group":"Training",
+ "pages":[
+ "docs/tutorials/longhorizon.html",
+ "docs/tutorials/multiple_series.html"
+ ]
+ },
+ {
+ "group":"Fine-tuning",
+ "pages":[
+ "docs/tutorials/finetuning.html",
+ "docs/tutorials/reusing_finetuned_models.html",
+ "docs/tutorials/loss_function_finetuning.html",
+ "docs/tutorials/finetune_depth_finetuning.html"
+ ]
+ },
+ {
+ "group":"Validation",
+ "pages":[
+ "docs/tutorials/cross_validation.html",
+ "docs/tutorials/historical_forecast.html"
+ ]
+ },
+ {
+ "group":"Uncertainty quantification",
+ "pages":[
+ "docs/tutorials/uncertainty_quantification_with_quantile_forecasts.html",
+ "docs/tutorials/uncertainty_quantification_with_prediction_intervals.html"
+ ]
+ },
+ {
+ "group":"Special Topics",
+ "pages":[
+ "docs/tutorials/bounded_forecasts.html",
+ "docs/tutorials/hierarchical_forecasting.html",
+ "docs/tutorials/temporalhierarchical.html",
+ "docs/tutorials/missing_values.html",
+ "docs/tutorials/how_to_improve_forecast_accuracy.html"
+ ]
+ },
+ {
+ "group":"Computing at scale",
+ "pages":[
+ "docs/tutorials/computing_at_scale.html",
+ "docs/tutorials/computing_at_scale_spark_distributed.html",
+ "docs/tutorials/computing_at_scale_dask_distributed.html",
+ "docs/tutorials/computing_at_scale_ray_distributed.html"
+ ]
+ }
+ ]
+ },
+ {
+ "group": "Use cases",
+ "pages": [
+ "docs/use-cases/forecasting_web_traffic.html",
+ "docs/use-cases/bitcoin_price_prediction.html",
+ "docs/use-cases/electricity_demand.html",
+ "docs/use-cases/intermittent_demand.html",
+ "docs/use-cases/what_if_pricing_scenarios_in_retail.html"
+ ]
+ },
+ {
+ "group": "API Reference",
+ "pages": ["docs/reference/nixtla_client.html",
+ "docs/reference/date_features.html",
+ "docs/reference/excel_addin.html",
+ "docs/reference/nixtlar.html"
+ ]
+ }
+ ]
+}
diff --git a/nixtla/src/date_features.html.mdx b/nixtla/src/date_features.html.mdx
new file mode 100644
index 00000000..e9b564f9
--- /dev/null
+++ b/nixtla/src/date_features.html.mdx
@@ -0,0 +1,59 @@
+---
+output-file: date_features.html
+title: Date Features
+---
+
+
+Useful classes to generate date features and add them to `TimeGPT`.
+
+------------------------------------------------------------------------
+
+source
+
+#### CountryHolidays
+
+> ``` text
+> CountryHolidays (countries:list[str])
+> ```
+
+*Given a list of countries, returns a dataframe with holidays for each
+country.*
+
+```python
+c_holidays = CountryHolidays(countries=['US', 'MX'])
+periods = 365 * 5
+dates = pd.date_range(end='2023-09-01', periods=periods)
+holidays_df = c_holidays(dates)
+holidays_df.head()
+```
+
+------------------------------------------------------------------------
+
+source
+
+#### SpecialDates
+
+> ``` text
+> SpecialDates (special_dates:dict[str,list[str]])
+> ```
+
+*Given a dictionary of categories and dates, returns a dataframe with
+the special dates.*
+
+```python
+special_dates = SpecialDates(
+ special_dates={
+ 'Important Dates': ['2021-02-26', '2020-02-26'],
+ 'Very Important Dates': ['2021-01-26', '2020-01-26', '2019-01-26']
+ }
+)
+periods = 365 * 5
+dates = pd.date_range(end='2023-09-01', periods=periods)
+holidays_df = special_dates(dates)
+holidays_df.head()
+```
+
diff --git a/nixtla/src/nixtla_client.html.mdx b/nixtla/src/nixtla_client.html.mdx
new file mode 100644
index 00000000..5e4a9e56
--- /dev/null
+++ b/nixtla/src/nixtla_client.html.mdx
@@ -0,0 +1,103 @@
+---
+output-file: nixtla_client.html
+title: Nixtla Client
+---
+
+
+## Imports
+
+## Utilities
+
+### Audit Data
+
+#### Audit Duplicate Rows
+
+#### Audit Missing Dates
+
+#### Audit Categorical Variables
+
+#### Audit Leading Zeros
+
+#### Audit Negative Values
+
+------------------------------------------------------------------------
+
+source
+
+### ApiError
+
+> ``` text
+> ApiError (status_code:Optional[int]=None, body:Optional[Any]=None)
+> ```
+
+*Common base class for all non-exit exceptions.*
+
+## Client
+
+------------------------------------------------------------------------
+
+source
+
+### NixtlaClient
+
+> ``` text
+> NixtlaClient (api_key:Optional[str]=None, base_url:Optional[str]=None,
+> timeout:Optional[int]=60, max_retries:int=6,
+> retry_interval:int=10, max_wait_time:int=360)
+> ```
+
+*Client to interact with the Nixtla API.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| api_key | Optional | None | The authorization api_key interacts with the Nixtla API. If not provided, will use the NIXTLA_API_KEY environment variable. |
+| base_url | Optional | None | Custom base_url. If not provided, will use the NIXTLA_BASE_URL environment variable. |
+| timeout | Optional | 60 | Request timeout in seconds. Set this to `None` to disable it. |
+| max_retries | int | 6 | The maximum number of attempts to make when calling the API before giving up. It defines how many times the client will retry the API call if it fails. Default value is 6, indicating the client will attempt the API call up to 6 times in total |
+| retry_interval | int | 10 | The interval in seconds between consecutive retry attempts. This is the waiting period before the client tries to call the API again after a failed attempt. Default value is 10 seconds, meaning the client waits for 10 seconds between retries. |
+| max_wait_time | int | 360 | The maximum total time in seconds that the client will spend on all retry attempts before giving up. This sets an upper limit on the cumulative waiting time for all retry attempts. If this time is exceeded, the client will stop retrying and raise an exception. Default value is 360 seconds, meaning the client will cease retrying if the total time spent on retries exceeds 360 seconds. The client throws a ReadTimeout error after 60 seconds of inactivity. If you want to catch these errors, use max_wait_time \>\> 60. |
+
+## Wrappers
+
+## Tests
+
+### Data Quality
+
+```python
+common_kwargs = {
+ "freq": "D",
+ "id_col": 'unique_id',
+ "time_col": 'ds'
+}
+```
+
+#### All Pass
+
+#### Duplicate rows
+
+#### Missing dates
+
+#### Duplicate rows and missing dates
+
+#### Categorical Columns
+
+#### Negative Values
+
+#### Leading zeros
+
+### Anomaly Detection
+
+Now you can start to make forecasts! Let’s import an example:
+
+### Distributed
+
+#### Spark
+
+#### Dask
+
+#### Ray
+
diff --git a/nixtla/src/utils.html.mdx b/nixtla/src/utils.html.mdx
new file mode 100644
index 00000000..b1903797
--- /dev/null
+++ b/nixtla/src/utils.html.mdx
@@ -0,0 +1,28 @@
+---
+output-file: utils.html
+title: Utils
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### colab_badge
+
+> ``` text
+> colab_badge (path:str)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### in_colab
+
+> ``` text
+> in_colab ()
+> ```
+
diff --git a/script.js b/script.js
new file mode 100755
index 00000000..2aa49e85
--- /dev/null
+++ b/script.js
@@ -0,0 +1,117 @@
+"use client";
+
+// Load the Inkeep script
+const inkeepScript = document.createElement("script");
+inkeepScript.type = "module";
+inkeepScript.src =
+ "https://unpkg.com/@inkeep/widgets-embed@0.2.263/dist/embed.js";
+
+document.body.appendChild(inkeepScript);
+
+// find the Mintlify search container
+const buttons = Array.from(document.getElementsByTagName("button"));
+const searchButtonContainerEl = buttons.find((button) =>
+ button.textContent.includes("Search or ask...")
+);
+
+const clonedSearchButtonContainerEl = searchButtonContainerEl.cloneNode(true);
+
+// replace with dummy div, required to remove event listeners on it
+searchButtonContainerEl.parentNode.replaceChild(
+ clonedSearchButtonContainerEl,
+ searchButtonContainerEl
+);
+
+// Once the script has loaded, load the Inkeep chat components
+inkeepScript.addEventListener("load", function () {
+ // Settings for the components
+ const sharedConfig = {
+ baseSettings: {
+ apiKey: "c99c3754a78cf9f7d3d389ef9fa7c3d49cdeb18b95050d10",
+ integrationId: "cltexfnd9000iy0ufy9oqo6h6",
+ organizationId: "org_VKqv6JAjdI85T7aJ",
+ primaryBrandColor: "#0c0c0c",
+ },
+ aiChatSettings: {
+ chatSubjectName: "Nixtla",
+ botAvatarSrcUrl:
+ "https://storage.googleapis.com/organization-image-assets/nixtla-botAvatarSrcUrl-1709853122420.png",
+ botAvatarDarkSrcUrl:
+ "https://storage.googleapis.com/organization-image-assets/nixtla-botAvatarDarkSrcUrl-1709853121631.png",
+ getHelpCallToActions: [
+ {
+ name: "Ask on Slack",
+ url: "https://join.slack.com/t/nixtlaworkspace/shared_invite/zt-135dssye9-fWTzMpv2WBthq8NK0Yvu6A",
+ icon: {
+ builtIn: "FaSlack",
+ },
+ },
+ {
+ name: "View Repositories",
+ url: "https://github.com/Nixtla",
+ icon: {
+ builtIn: "FaGithub",
+ },
+ },
+ ],
+ quickQuestions: [
+ "How do I train my own model using mlforecast?",
+ "How was TimeGPT trained and what's it best for?",
+ "How do I make multivariate scoring for hierarchical forecasting?",
+ ],
+ },
+ };
+
+ // for syncing with dark mode
+ const colorModeSettings = {
+ observedElement: document.documentElement,
+ isDarkModeCallback: (el) => {
+ return el.classList.contains("dark");
+ },
+ colorModeAttribute: "class",
+ };
+
+ // add the chat button
+ const chatButton = Inkeep().embed({
+ componentType: "ChatButton",
+ colorModeSync: colorModeSettings,
+ properties: sharedConfig,
+ });
+
+ // insantiate Inkeep modal
+ const searchButtonWithCustomTrigger = Inkeep({
+ ...sharedConfig.baseSettings,
+ }).embed({
+ componentType: "CustomTrigger",
+ colorModeSync: colorModeSettings,
+ properties: {
+ ...sharedConfig,
+ isOpen: false,
+ onClose: () => {
+ searchButtonWithCustomTrigger.render({
+ isOpen: false,
+ });
+ },
+ },
+ });
+
+ // When the Mintlify search bar clone is clicked, open the Inkeep search modal
+ clonedSearchButtonContainerEl.addEventListener("click", function () {
+ searchButtonWithCustomTrigger.render({
+ isOpen: true,
+ });
+ });
+});
+
+
+!(function (t) {
+ if (window.oneloop) return;
+ window.oneloop = {key: "cus_b57d16380d7240e4a47c9f4a6f0893c6"};
+ var n = document.createElement("script");
+ (n.async = !0),
+ n.setAttribute(
+ "src",
+ "https://oneloop-website-script.s3.us-west-1.amazonaws.com/oneloop-main.js"
+ ),
+ (document.body || document.head).appendChild(n);
+})();
diff --git a/statsforecast/.nojekyll b/statsforecast/.nojekyll
new file mode 100644
index 00000000..e69de29b
diff --git a/statsforecast/blog/index.mdx b/statsforecast/blog/index.mdx
new file mode 100644
index 00000000..fcff25d1
--- /dev/null
+++ b/statsforecast/blog/index.mdx
@@ -0,0 +1,15 @@
+---
+title: StatsForecast Blog
+subtitle: News, tips, and commentary about all things StatsForecast
+listing:
+ sort: date desc
+ contents: posts
+ sort-ui: false
+ filter-ui: false
+ categories: true
+ feed: true
+page-layout: full
+---
+
+
+
diff --git a/statsforecast/blog/posts/2022-10-05-distributed-fugue/index.mdx b/statsforecast/blog/posts/2022-10-05-distributed-fugue/index.mdx
new file mode 100644
index 00000000..592ef457
--- /dev/null
+++ b/statsforecast/blog/posts/2022-10-05-distributed-fugue/index.mdx
@@ -0,0 +1,340 @@
+---
+author:
+ - Fugue
+ - Nixtla
+date: 2022-10-05T00:00:00.000Z
+title: Scalable Time Series Modeling with open-source projects
+description: >-
+ How to Forecast 1M Time Series in 15 Minutes with Spark, Fugue and Nixtla's
+ Statsforecast.
+tags:
+ - fugue
+ - scalability
+ - spark
+ - time-series
+ - forecasting
+---
+
+
+[](https://github.com/nixtla/statsforecast)
+
+By [Fugue](https://github.com/fugue-project/) and
+[Nixtla](https://github.com/nixtla/). Originally posted on
+[TDS](https://medium.com/towards-data-science/distributed-forecast-of-1m-time-series-in-under-15-minutes-with-spark-nixtla-and-fugue-e9892da6fd5c).
+
+> **TL:DR We will show how you can leverage the distributed power of
+> Spark and the highly efficient code from StatsForecast to fit millions
+> of models in a couple of minutes.**
+
+Time-series modeling, analysis, and prediction of trends and
+seasonalities for data collected over time is a rapidly growing category
+of software applications.
+
+Businesses, from electricity and economics to healthcare analytics,
+collect time-series data daily to predict patterns and build better
+data-driven product experiences. For example, temperature and humidity
+prediction is used in manufacturing to prevent defects, streaming
+metrics predictions help identify music’s popular artists, and sales
+forecasting for thousands of SKUs across different locations in the
+supply chain is used to optimize inventory costs. As data generation
+increases, the forecasting necessities have evolved from modeling a few
+time series to predicting millions.
+
+## Motivation
+
+[Nixtla](https://github.com/nixtla) is an open-source project focused on
+state-of-the-art time series forecasting. They have a couple of
+libraries such as
+[StatsForecast](https://github.com/nixtla/statsforecast) for statistical
+models, [NeuralForecast](https://github.com/nixtla/neuralforecast) for
+deep learning, and
+[HierarchicalForecast](https://github.com/nixtla/hierarchicalforecast)
+for forecast aggregations across different levels of hierarchies. These
+are production-ready time series libraries focused on different modeling
+techniques.
+
+This article looks at
+[StatsForecast](https://github.com/nixtla/statsforecast), a
+lightning-fast forecasting library with statistical and econometrics
+models. The AutoARIMA model of Nixtla is 20x faster than
+[pmdarima](http://alkaline-ml.com/pmdarima/), and the ETS (error, trend,
+seasonal) models performed 4x faster than
+[statsmodels](https://github.com/statsmodels/statsmodels) and are more
+robust. The benchmarks and code to reproduce can be found
+[here](https://github.com/Nixtla/statsforecast#-accuracy---speed). A
+huge part of the performance increase is due to using a JIT compiler
+called [numba](https://numba.pydata.org/) to achieve high speeds.
+
+The faster iteration time means that data scientists can run more
+experiments and converge to more accurate models faster. It also means
+that running benchmarks at scale becomes easier.
+
+In this article, we are interested in the scalability of the
+StatsForecast library in fitting models over
+[Spark](https://spark.apache.org/docs/latest/api/python/index.html) or
+[Dask](https://github.com/dask/dask) using the
+[Fugue](https://github.com/fugue-project/fugue/) library. This
+combination will allow us to train a huge number of models distributedly
+over a temporary cluster quickly.
+
+## Experiment Setup
+
+When dealing with large time series data, users normally have to deal
+with thousands of logically independent time series (think of telemetry
+of different users or different product sales). In this case, we can
+train one big model over all of the series, or we can create one model
+for each series. Both are valid approaches since the bigger model will
+pick up trends across the population, while training thousands of models
+may fit individual series data better.
+
+> **Note**
+>
+> Note: to pick up both the micro and macro trends of the time series
+> population in one model, check the Nixtla
+> [HierarchicalForecast](https://github.com/Nixtla/hierarchicalforecast)
+> library, but this is also more computationally expensive and trickier
+> to scale.
+
+This article will deal with the scenario where we train a couple of
+models (AutoARIMA or ETS) per univariate time series. For this setup, we
+group the full data by time series, and then train each model for each
+group. The image below illustrates this. The distributed DataFrame can
+either be a Spark or Dask DataFrame.
+
+
+
+AutoARIMA per partition
+
+
+Nixtla previously released benchmarks with
+[Anyscale](https://www.anyscale.com/) on distributing this model
+training on Ray. The setup and results can be found in this
+[blog](https://www.anyscale.com/blog/how-nixtla-uses-ray-to-accurately-predict-more-than-a-million-time-series).
+The results are also shown below. It took 2000 cpus to run one million
+AutoARIMA models in 35 minutes. We’ll compare this against running on
+Spark.
+
+
+
+StatsForecast on Ray results
+
+
+## StatsForecast code
+
+First, we’ll look at the StatsForecast code used to run the AutoARIMA
+distributedly on [Ray](https://docs.ray.io/en/latest/index.html). This
+is a simplified version to run the scenario with a one million time
+series. It is also updated for the recent StatsForecast v1.0.0 release,
+so it may look a bit different from the code in the previous benchmarks.
+
+
+```python
+from time import time
+
+import pandas as pd
+from statsforecast.utils import generate_series
+from statsforecast.models import AutoARIMA
+from statsforecast.core import StatsForecast
+
+series = generate_series(n_series=1000000, seed=1)
+
+model = StatsForecast(df=series,
+ models=[AutoARIMA()],
+ freq='D',
+ n_jobs=-1,
+ ray_address=ray_address)
+
+init = time()
+forecasts = model.forecast(7)
+print(f'n_series: 1000000 total time: {(time() - init) / 60}')
+```
+
+The interface of StatsForecast is very minimal. It is already designed
+to perform the AutoARIMA on each group of data. Just supplying the
+ray_address will make this code snippet run distributedly. Without it,
+n_jobswill indicate the number of parallel processes for forecasting.
+model.forecast() will do the fit and predict in one step, and the input
+to this method in the time horizon to forecast.
+
+## Using Fugue to run on Spark and Dask
+
+[Fugue](https://github.com/fugue-project/fugue) is an abstraction layer
+that ports Python, Pandas, and SQL code to Spark and Dask. The most
+minimal interface is the `transform()` function. This function takes in
+a function and DataFrame, and brings it to Spark or Dask. We can use the
+`transform()` function to bring StatsForecast execution to Spark.
+
+There are two parts to the code below. First, we have the forecast logic
+defined in the `forecast_series` function. Some parameters are hardcoded
+for simplicity. The most important one is that `n_jobs=1`. This is
+because Spark or Dask will already serve as the parallelization layer,
+and having two stages of parallelism can cause resource deadlocks.
+
+
+```python
+from fugue import transform
+
+def forecast_series(df: pd.DataFrame, models) -> pd.DataFrame:
+ tdf = df.set_index("unique_id")
+ model = StatsForecast(df=tdf, models=models, freq='D', n_jobs=1)
+ return model.forecast(7).reset_index()
+
+transform(series.reset_index(),
+ forecast_series,
+ params=dict(models=[AutoARIMA()]),
+ schema="unique_id:int, ds:date, AutoARIMA:float",
+ partition={"by": "unique_id"},
+ engine="spark"
+ ).show()
+```
+
+Second, the `transform()` function is used to apply the
+`forecast_series()` function on Spark. The first two arguments are the
+DataFrame and function to be applied. Output schema is a requirement for
+Spark, so we need to pass it in, and the partition argument will take
+care of splitting the time series modelling by `unique_id`.
+
+This code already works and returns a Spark DataFrame output.
+
+## Nixtla’s FugueBackend
+
+The `transform()` above is a general look at what Fugue can do. In
+practice, the Fugue and Nixtla teams collaborated to add a more native
+`FugueBackend` to the StatsForecast library. Along with it is a utility
+`forecast()` function to simplify the forecasting interface. Below is an
+end-to-end example of running StatsForecast on one million time series.
+
+
+```python
+from statsforecast.distributed.utils import forecast
+from statsforecast.distributed.fugue import FugueBackend
+from statsforecast.models import AutoARIMA
+from statsforecast.core import StatsForecast
+
+from pyspark.sql import SparkSession
+
+spark = SparkSession.builder.getOrCreate()
+backend = FugueBackend(spark, {"fugue.spark.use_pandas_udf":True})
+
+forecast(spark.read.parquet("/tmp/1m.parquet"),
+ [AutoARIMA()],
+ freq="D",
+ h=7,
+ parallel=backend).toPandas()
+```
+
+We just need to create the FugueBackend, which takes in a SparkSession
+and passes it to `forecast()`. This function can take either a DataFrame
+or file path to the data. If a file path is provided, it will be loaded
+with the parallel backend. In this example above, we replaced the file
+each time we ran the experiment to generate benchmarks.
+
+> **Caution**
+>
+> It’s also important to note that we can test locally before running
+> the `forecast()` on full data. All we have to do is not supply
+> anything for the parallel argument; everything will run on Pandas
+> sequentially.
+
+## Benchmark Results
+
+The benchmark results can be seen below. As of the time of this writing,
+Dask and Ray made recent releases, so only the Spark metrics are up to
+date. We will make a follow-up article after running these experiments
+with the updates.
+
+
+
+Spark and Dask benchmarks for
+StatsForecast at scale
+
+
+> **Note**
+>
+> Note: The attempt was to use 2000 cpus but we were limited by
+> available compute instances on AWS.
+
+The important part here is that AutoARIMA trained one million time
+series models in less than 15 minutes. The cluster configuration is
+attached in the appendix. With very few lines of code, we were able to
+orchestrate the training of these time series models distributedly.
+
+## Conclusion
+
+Training thousands of time series models distributedly normally takes a
+lot of coding with Spark and Dask, but we were able to run these
+experiments with very few lines of code. Nixtla’s StatsForecast offers
+the ability to quickly utilize all of the compute resources available to
+find the best model for each time series. All users need to do is supply
+a relevant parallel backend (Ray or Fugue) to run on a cluster.
+
+On the scale of one million timeseries, our total training time took 12
+minutes for AutoARIMA. This is the equivalent of close to 400 cpu-hours
+that we ran immediately, allowing data scientists to quickly iterate at
+scale without having to write the explicit code for parallelization.
+Because we used an ephemeral cluster, the cost is effectively the same
+as running this sequentially on an EC2 instance (parallelized over all
+cores).
+
+## Resources
+
+1. [Nixtla StatsForecast repo](https://github.com/Nixtla/statsforecast)
+2. [StatsForecast docs](https://nixtla.github.io/statsforecast/)
+3. [Fugue repo](https://github.com/fugue-project/fugue/)
+4. [Fugue tutorials](https://fugue-tutorials.readthedocs.io/)
+
+To chat with us:
+
+1. [Fugue Slack](http://slack.fugue.ai/)
+2. [Nixtla
+ Slack](https://join.slack.com/t/nixtlaworkspace/shared_invite/zt-135dssye9-fWTzMpv2WBthq8NK0Yvu6A)
+
+## Appendix
+
+For anyone. interested in the cluster configuration, it can be seen
+below. This will spin up a Databricks cluster. The important thing is
+the node_type_id that has the machines used.
+
+``` text
+{
+ "num_workers": 20,
+ "cluster_name": "fugue-nixtla-2",
+ "spark_version": "10.4.x-scala2.12",
+ "spark_conf": {
+ "spark.speculation": "true",
+ "spark.sql.shuffle.partitions": "8000",
+ "spark.sql.adaptive.enabled": "false",
+ "spark.task.cpus": "1"
+ },
+ "aws_attributes": {
+ "first_on_demand": 1,
+ "availability": "SPOT_WITH_FALLBACK",
+ "zone_id": "us-west-2c",
+ "spot_bid_price_percent": 100,
+ "ebs_volume_type": "GENERAL_PURPOSE_SSD",
+ "ebs_volume_count": 1,
+ "ebs_volume_size": 32
+ },
+ "node_type_id": "m5.24xlarge",
+ "driver_node_type_id": "m5.2xlarge",
+ "ssh_public_keys": [],
+ "custom_tags": {},
+ "spark_env_vars": {
+ "MKL_NUM_THREADS": "1",
+ "OPENBLAS_NUM_THREADS": "1",
+ "VECLIB_MAXIMUM_THREADS": "1",
+ "OMP_NUM_THREADS": "1",
+ "NUMEXPR_NUM_THREADS": "1"
+ },
+ "autotermination_minutes": 20,
+ "enable_elastic_disk": false,
+ "cluster_source": "UI",
+ "init_scripts": [],
+ "runtime_engine": "STANDARD",
+ "cluster_id": "0728-004950-oefym0ss"
+}
+```
+
diff --git a/statsforecast/dark.png b/statsforecast/dark.png
new file mode 100644
index 00000000..4142a0bb
Binary files /dev/null and b/statsforecast/dark.png differ
diff --git a/statsforecast/docs/contribute/contribute.mdx b/statsforecast/docs/contribute/contribute.mdx
new file mode 100644
index 00000000..cfabe71e
--- /dev/null
+++ b/statsforecast/docs/contribute/contribute.mdx
@@ -0,0 +1,69 @@
+---
+title: Contribute to Nixtla
+sidebarTitle: How to Contribute
+---
+
+
+Thank you for your interest in contributing to Nixtla. Nixtla is free,
+open-source software and welcomes all types of contributions, including
+documentation changes, bug reports, bug fixes, or new source code
+changes.
+
+## Contribution issues 🔧
+
+Most of the issues that are open for contributions will be tagged with
+`good first issue` or `help wanted`. A great place to start looking will
+be our GitHub projects for:
+
+- Community writers
+ [dashboard](https://github.com/orgs/Nixtla/projects/9).
+- Community code contributors
+ [dashboard](https://github.com/orgs/Nixtla/projects/6).
+
+Also, we are always open to suggestions so feel free to open new issues
+with your ideas and we can give you guidance!
+
+After you find the issue that you want to contribute to, follow the
+`fork-and-pull` workflow:
+
+1. Fork the Nixtla repository you want to work on (e.g. StatsForecast
+ or NeuralForecast)
+2. Clone the repository locally (`git clone`) and create a new branch
+ (`git checkout -b my-new-branch`)
+3. Make changes and commit them
+4. Push your local branch to your fork
+5. Submit a Pull Request so that we can review your changes
+6. Write a commit message
+7. Make sure that the CI tests are GREEN (CI tests refer to automated
+ tests that are run on code changes to ensure that new additions or
+ modifications do not introduce new errors or break existing
+ functionality.)
+
+Be sure to merge the latest from “upstream” before making a Pull
+Request!
+
+You can find a complete step-by-step guide on this `fork-and-pull`
+workflow
+[here](https://github.com/Nixtla/how-to-contribute-nixtlaverse).
+
+Pull Request reviews are done on a regular basis. Please make sure you
+respond to our feedback/questions and sign our CLA.
+
+## Documentation 📖
+
+We are committed to continuously improving our documentation. As such,
+we warmly welcome any Pull Requests that focus on improving our grammar,
+documentation structure, or fixing any typos.
+
+- Check the `documentation` tagged issues and help us.
+
+## Write for us 📝
+
+Do you find Nixtla useful and want to share your story or create some
+content? Make a PR to this repo with your writing in a markdown file, or
+just post it on Medium, Dev or your own blog post. We would love to hear
+from you 💚
+
+This document is based on the documentation from
+[MindsDB](https://github.com/mindsdb/mindsdb)
+
diff --git a/statsforecast/docs/contribute/docs.mdx b/statsforecast/docs/contribute/docs.mdx
new file mode 100644
index 00000000..9ba7ba03
--- /dev/null
+++ b/statsforecast/docs/contribute/docs.mdx
@@ -0,0 +1,8 @@
+---
+title: Nixtla Documentation
+sidebarTitle: Writing Documentation
+---
+
+
+TBD
+
diff --git a/statsforecast/docs/contribute/issue-labels.mdx b/statsforecast/docs/contribute/issue-labels.mdx
new file mode 100644
index 00000000..b1326bf3
--- /dev/null
+++ b/statsforecast/docs/contribute/issue-labels.mdx
@@ -0,0 +1,82 @@
+---
+title: Understanding Issue Labels
+sidebarTitle: Interpreting Issue Labels
+---
+
+
+This segment delves into the variety of issue labels used within the
+[Nixtla GitHub repository](https://github.com/nixtla/nixtla).
+
+## Labels Relevant to Contributors
+
+Should you be a contributor now or in the future, it’s important to take
+note of issues flagged with these labels.
+
+### The `first-timers-only` Label
+
+For those who have not yet contributed to Nixtla, start by looking for
+issues tagged as `first-timers-only`.
+
+Please note that before we can accept your contribution to Nixtla,
+you’ll need to sign our [Contributor License
+Agreement](https://gist.github.com/cchallu/6ce3e997462c74dd77341e2fdeb46a48).
+
+You can browse all `first-timers-only` issues
+[here](https://github.com/nixtla/nixtla/labels/first-timers-only).
+
+### The `good first issue` Label
+
+Issues labeled as `good first issue` are ideal for newcomers.
+
+You can browse all `good first issue` issues
+[here](https://github.com/nixtla/nixtla/labels/good%20first%20issue).
+
+### The `help wanted` Label
+
+Issues tagged as `help wanted` are open to anyone who wishes to
+contribute to Nixtla.
+
+You can browse all `help wanted` issues
+[here](https://github.com/nixtla/nixtla/labels/help%20wanted).
+
+### The `bug` Label
+
+The `bug` label flags issues that outline something that’s currently not
+functioning correctly.
+
+You can report a bug by following the instructions
+[here](../../contribute/issues#report-a-bug).
+
+### The `discussion` Label
+
+If an issue is labeled as `discussion`, it signifies that more
+conversation is needed before it can be resolved.
+
+### The `documentation` Label
+
+The `documentation` label identifies issues pertaining to our
+documentation.
+
+You can contribute to improving our documentation by creating issues
+following the guidelines
+[here](../../contribute/issues#improve-our-docs).
+
+### The `enhancement` Label
+
+As Nixtla continues to evolve, there are always areas that can be
+enhanced. All issues suggesting improvements to Nixtla are tagged with
+the `enhancement` label.
+
+You can propose a feature by following the instructions
+[here](../../contribute/issues#request-a-feature).
+
+### The `discussion` Label
+
+If an issue is labeled as `discussion`, it needs more information before
+it can be resolved.
+
+### The `requested` Label
+
+Our users are welcomed to propose improvements, report bugs, request
+feature, etc. Any issue originating from them is flagged as `requested`.
+
diff --git a/statsforecast/docs/contribute/issues.mdx b/statsforecast/docs/contribute/issues.mdx
new file mode 100644
index 00000000..04c1e5e6
--- /dev/null
+++ b/statsforecast/docs/contribute/issues.mdx
@@ -0,0 +1,135 @@
+---
+title: Submit an Issue 📢
+sidebarTitle: Submit an Issue
+---
+
+
+To report a bug, request a feature, propose a new integration, or
+suggest documentation improvements, please visit the [Nixtla GitHub
+issues page](https://github.com/nixtla/nixtla/issues). Before submitting
+a new issue, kindly check if it has already been reported.
+
+## Steps to Submit an Issue
+
+Here’s a step-by-step guide on submitting an issue to the Nixtla
+repository.
+
+Visit [our GitHub issues page](https://github.com/nixtla/nixtla/issues)
+and click on the *New issue* button.
+
+A list of available issue types will be displayed.
+
+### Reporting a Bug 🐞
+
+Select `Report a bug` and click on the *Get started* button.
+
+The form to report the bug will appear.
+
+1. Begin by adding a concise, informative title.
+2. Describe the bug you’ve observed. This information is required. You
+ can also attach relevant videos or screenshots.
+3. If you’re aware of what the correct behavior should be, note it down
+ here.
+4. Documenting the steps leading to the bug will be of immense help to
+ us.
+5. You can also add links, references, logs, screenshots, and so on.
+
+
+
+Please ensure that your contributions abide by the [contributing
+guidelines](https://github.com/Nixtla/statsforecast/blob/main/CONTRIBUTING.md)
+and [code of
+conduct](https://github.com/Nixtla/statsforecast/blob/main/CODE_OF_CONDUCT.md).
+
+
+
+Thank you for your contribution! Your report aids in refining Nixtla for
+current and future users.
+
+### Feature Request 🚀
+
+Select `Request a feature` and click the *Get started* button.
+
+The feature request form will appear.
+
+1. Start with a significant, clear title.
+2. Provide a detailed description of the feature you want to request,
+ along with the reasoning behind the request. This field is
+ mandatory. Feel free to attach related videos or screenshots.
+3. If you have an idea of how the feature should work, include it.
+4. Additional references, links, logs, and screenshots are welcome!
+
+
+
+Please ensure that your contributions abide by the [contributing
+guidelines](https://github.com/nixtla/nixtla/blob/main/CONTRIBUTING.md)
+and [code of
+conduct](https://github.com/Nixtla/nixtla/blob/main/CODE_OF_CONDUCT.md).
+
+
+
+Thank you for your feature request! It will help us enhance Nixtla for
+all users.
+
+### Suggest Documentation Improvements ✍️
+
+Select `Improve our docs` and click the *Get started* button.
+
+A form for suggesting improvements will appear.
+
+1. A clear, concise title is important.
+2. Describe the improvements you believe are needed. This field is
+ mandatory. Attach any related videos or screenshots, if necessary.
+3. Any additional references, links, logs, screenshots are appreciated!
+
+
+
+Please ensure that your contributions abide by the [contributing
+guidelines](https://github.com/Nixtla/nixtla/blob/main/CONTRIBUTING.md)
+and [code of conduct](https://github.com/nixtla/nixtla/blob/main/CODE_OF_CONDUCT.md).
+
+
+
+Thank you for your valuable suggestions! Your input helps us refine
+Nixtla’s documentation.
+
+### Proposing a New Integration 🧑🔧
+
+If you have a proposal for a new database integration or a new machine
+learning framework, here’s how to get started:
+
+Select \`
+
+Propose a new integration\` and click the *Get started* button.
+
+A form for your proposal will appear.
+
+1. Start with a clear, concise title.
+2. Describe your proposal and why it is needed. This field is
+ mandatory. Feel free to attach any related videos or screenshots.
+3. If you have an idea of how this integration should work, include it.
+4. Any additional references, links, logs, screenshots, and so on, are
+ welcome!
+
+
+
+Please ensure that your contributions abide by the [contributing guidelines](https://github.com/Nixtla/nixtla/blob/main/CONTRIBUTING.md)
+and [code of conduct](https://github.com/nixtla/nixtla/blob/main/CODE_OF_CONDUCT.md).
+
+
+
+
+Thank you for your proposal! Your suggestion helps us extend the
+capabilities of Nixtla.
+
+## Reviewing Issues
+
+- Issues are reviewed on a regular basis, usually every day.
+- Issues will be labeled as `Bug` or `enhancement` based on their
+ type.
+- Please be ready to respond to our feedback or questions regarding
+ your issue.
+
+This document is based on the documentation from
+[MindsDB](https://github.com/mindsdb/mindsdb)
+
diff --git a/statsforecast/docs/contribute/step-by-step.mdx b/statsforecast/docs/contribute/step-by-step.mdx
new file mode 100644
index 00000000..e639a1ff
--- /dev/null
+++ b/statsforecast/docs/contribute/step-by-step.mdx
@@ -0,0 +1,367 @@
+# Step-by-step Contribution Guide
+
+> This document contains instructions for collaborating on the different
+> libraries of Nixtla.
+
+Sometimes, diving into a new technology can be challenging and
+overwhelming. We’ve been there too, and we’re more than ready to assist
+you with any issues you may encounter while following these steps. Don’t
+hesitate to reach out to us on
+[Slack](https://join.slack.com/t/nixtlacommunity/shared_invite/zt-1pmhan9j5-F54XR20edHk0UtYAPcW4KQ).
+Just give fede a ping, and she’ll be glad to assist you.
+
+## Table of Contents 📚
+
+1. [Prerequisites](#prerequisites)
+2. [Git `fork-and-pull` worklow](#git-fork-and-pull-worklow)
+3. [Set Up a Conda Environment](#set-up-a-conda-environment)
+4. [Install required libraries for
+ development](#install-required-libraries-for-development)
+5. [Start editable mode](#start-editable-mode)
+6. [Set Up your Notebook based development
+ environment](#set-up-your-notebook-based-development-environment)
+7. [Start Coding](#start-coding)
+8. [Example with Screen-shots](#example-with-screen-shots)
+
+## Prerequisites
+
+- *GitHub*: You should already have a GitHub account and a basic
+ understanding of its functionalities. Alternatively check [this
+ guide](https://docs.github.com/en/get-started).
+- *Python*: Python should be installed on your system. Alternatively
+ check [this guide](https://www.python.org/downloads/).
+- *conda*: You need to have conda installed, along with a good grasp
+ of fundamental operations such as creating environments, and
+ activating and deactivating them. Alternatively check [this
+ guide](https://conda.io/projects/conda/en/latest/user-guide/install/index.html).
+
+## Git `fork-and-pull` worklow
+
+**1. Fork the Project:** Start by forking the Nixtla repository to your
+own GitHub account. This creates a personal copy of the project where
+you can make changes without affecting the main repository.
+
+**2. Clone the Forked Repository** Clone the forked repository to your
+local machine using
+`git clone https://github.com//nixtla.git`. This allows
+you to work with the code directly on your system.
+
+**3. Create a Branch:**
+
+Branching in GitHub is a key strategy for effectively managing and
+isolating changes to your project. It allows you to segregate work on
+different features, fixes, and issues without interfering with the main,
+production-ready codebase.
+
+1. *Main Branch*: The default branch with production-ready code.
+
+2. *Feature Branches*: For new features, create branches prefixed with
+ ‘feature/’, like `git checkout -b feature/new-model`.
+
+3. *Fix Branches*: For bug fixes, use ‘fix/’ prefix, like
+ `git checkout -b fix/forecasting-bug`.
+
+4. *Issue Branches*: For specific issues, use
+ `git checkout -b issue/issue-number` or
+ `git checkout -b issue/issue-description`.
+
+After testing, branches are merged back into the main branch via a pull
+request, and then typically deleted to maintain a clean repository. You
+can read more about github and branching
+[here](https://docs.github.com/en/pull-requests/collaborating-with-pull-requests/proposing-changes-to-your-work-with-pull-requests/creating-and-deleting-branches-within-your-repository).
+
+## Set Up a Conda Environment
+
+> If you want to use Docker or Codespaces, let us know opening an issue
+> and we will set you up.
+
+Next, you’ll need to set up a [Conda](https://docs.conda.io/en/latest/)
+environment. Conda is an open-source package management and environment
+management system that runs on Windows, macOS, and Linux. It allows you
+to create separate environments containing files, packages, and
+dependencies that will not interact with each other.
+
+First, ensure you have Anaconda or Miniconda installed on your system.
+Alternatively checkout these guides:
+[Anaconda](https://www.anaconda.com/),
+[Miniconda](https://docs.conda.io/en/latest/miniconda.html), and
+[Mamba](https://mamba.readthedocs.io/en/latest/).
+
+Then, you can create a new environment using
+`conda create -n nixtla-env python=3.10`.
+
+You can also use mamba for creating the environment (mamba is faster
+than Conda) using `mamba create -n nixtla-env python=3.10`.
+
+You can replace `nixtla-env` for something more meaningful to you. Eg.
+`statsforecast-env` or `mlforecast-env`. You can always check the list
+of environments in your system using `conda env list`.
+
+Activate your new environment with `conda activate nixtla-env`.
+
+## Install required libraries for development
+
+The `environment.yml` file contains all the dependencies required for
+the project. To install these dependencies, use the `mamba` package
+manager, which offers faster package installation and environment
+resolution than Conda. If you haven’t installed `mamba` yet, you can do
+so using `conda install mamba -c conda-forge`. Run the following command
+to install the dependencies:
+
+``` text
+mamba env update -f environment.yml
+```
+
+Sometimes (e.g. StatsForecast) the `enviorment.yml` is sometimes inside
+a folder called `dev`. In that case, you should run
+`mamba env update -f dev/environment.yml`.
+
+## Start editable mode
+
+Install the library in editable mode using `pip install -e ".[dev]"`.
+
+This means the package is linked directly to the source code, allowing
+any changes made to the source code to be immediately reflected in your
+Python environment without the need to reinstall the package. This is
+useful for testing changes during package development.
+
+## Set Up your Notebook based development environment
+
+Notebook-based development refers to using interactive notebooks, such
+as Jupyter Notebooks, for coding, data analysis, and visualization.
+Here’s a brief description of its characteristics:
+
+1. **Interactivity**: Code in notebooks is written in cells which can
+ be run independently. This allows for iterative development and
+ testing of small code snippets.
+
+2. **Visualization**: Notebooks can render charts, tables, images, and
+ other graphical outputs within the same interface, making it great
+ for data exploration and analysis.
+
+3. **Documentation**: Notebooks support Markdown and HTML, allowing for
+ detailed inline documentation. Code, outputs, and documentation are
+ in one place, which is ideal for tutorials, reports, or sharing
+ work.
+
+For notebook based development you’ll need `nbdev` and a notebook editor
+(such as VS Code, Jupyter Notebook or Jupyter Lab). `nbdev` and jupyter
+have been installed in the previous step. If you use VS Code follow
+[this
+tutorial](https://code.visualstudio.com/docs/datascience/jupyter-notebooks).
+
+[nbdev](https://github.com/fastai/nbdev) makes debugging and refactoring
+your code much easier than in traditional programming environments since
+you always have live objects at your fingertips. `nbdev` also promotes
+software engineering best practices because tests and documentation are
+first class.
+
+All your changes must be written in the notebooks contained in the
+library (under the `nbs` directory). Once a specific notebook is open
+(more details to come), you can write your Python code in cells within
+the notebook, as you would do in a traditional Python development
+workflow. You can break down complex problems into smaller parts,
+visualizing data, and documenting your thought process. Along with your
+code, you can include markdown cells to add documentation directly in
+the notebook. This includes explanations of your logic, usage examples,
+and more. Also, `nbdev` allows you to write [tests
+inline](https://nbdev.fast.ai/tutorials/best_practices.html#document-error-cases-as-tests)
+with your code in your notebook. After writing a function, you can
+immediately write tests for it in the following cells.
+
+Once your code is ready, `nbdev` can automatically convert your notebook
+into Python scripts. Code cells are converted into Python code, and
+markdown cells into comments and docstrings.
+
+## Start Coding
+
+Open a jupyter notebook using `jupyter lab` (or VS Code).
+
+1. **Make Your Changes:** Make changes to the codebase, ensuring your
+ changes are self-contained and cohesive.
+
+2. **Commit Your Changes:** Add the changed files using
+ `git add [your_modified_file_0.ipynb] [your_modified_file_1.ipynb]`,
+ then commit these changes using
+ `git commit -m ": "`. Please
+ use [Conventional
+ Commits](https://www.conventionalcommits.org/en/v1.0.0/)
+
+3. **Push Your Changes:** Push your changes to the remote repository on
+ GitHub with `git push origin feature/your-feature-name`.
+
+4. **Open a Pull Request:** Open a pull request from your new branch on
+ the Nixtla repository on GitHub. Provide a thorough description of
+ your changes when creating the pull request.
+
+5. **Wait for Review:** The maintainers of the Nixtla project will
+ review your changes. Be ready to iterate on your contributions based
+ on their feedback.
+
+Remember, contributing to open-source projects is a collaborative
+effort. Respect the work of others, welcome feedback, and always strive
+to improve. Happy coding!
+
+> Nixtla offers the possibility of assisting with stipends for computing
+> infrastructure for our contributors. If you are interested, please
+> join our
+> [slack](https://nixtlacommunity.slack.com/join/shared_invite/zt-1pmhan9j5-F54XR20edHk0UtYAPcW4KQ#/shared-invite/email)
+> and write to fede or Max.
+
+You can find a detailed step by step buide with screen-shots below.
+
+## Example with Screen-shots
+
+### 1. Create a fork of the mlforecast repo
+
+The first thing you need to do is create a fork of the GitHub repository
+to your own account:
+
+
+
+Your fork on your account will look like this:
+
+
+
+In that repository, you can make your changes and then request to have
+them added to the main repo.
+
+### 2. Clone the repository
+
+In this tutorial, we are using Mac (also compatible with other Linux
+distributions). If you are a collaborator of Nixtla, you can request an
+AWS instance to collaborate from there. If this is the case, please
+reach out to Max or Fede on
+[Slack](https://join.slack.com/t/nixtlacommunity/shared_invite/zt-1pmhan9j5-F54XR20edHk0UtYAPcW4KQ)
+to receive the appropriate access. We also use Visual Studio Code, which
+you can download from [here](https://code.visualstudio.com/download).
+
+Once the repository is created, you need to clone it to your own
+computer. Simply copy the repository URL from GitHub as shown below:
+
+
+
+Then open Visual Studio Code, click on “Clone Git Repository,” and paste
+the line you just copied into the top part of the window, as shown
+below:
+
+
+
+Select the folder where you want to copy the repository:
+
+
+
+And choose to open the cloned repository:
+
+
+
+You will end up with something like this:
+
+
+
+### 3. Create the Conda environment
+
+Open a terminal within Visual Studio Code, as shown in the image:
+
+
+
+You can use conda but we highly recommend using Mamba to speed up the
+creation of the Conda environment. To install it, simply use
+`conda install mamba -c conda-forge` in the terminal you just opened:
+
+
+
+Create an empty environment named `mlforecast` with the following
+command: `mamba create -n mlforecast python=3.10`:
+
+
+
+Activate the newly created environment using
+`conda activate mlforecast`:
+
+
+
+Install the libraries within the environment file `environment.yml`
+using `mamba env update -f environment.yml`:
+
+
+
+Now install the library to make interactive changes and other additional
+dependencies using `pip install -e ".[dev]"`:
+
+
+
+### 4. Make the changes you want.
+
+In this section, we assume that we want to increase the default number
+of windows used to create prediction intervals from 2 to 3. The first
+thing we need to do is create a specific branch for that change using
+`git checkout -b [new_branch]` like this:
+
+
+
+Once created, open the notebook you want to modify. In this case, it’s
+`nbs/utils.ipynb`, which contains the metadata for the prediction
+intervals. After opening it, click on the environment you want to use
+(top right) and select the `mlforecast` environment:
+
+
+
+Next, execute the notebook and make the necessary changes. In this case,
+we want to modify the `PredictionIntervals` class:
+
+
+
+We will change the default value of `n_window` from 2 to 3:
+
+
+
+Once you have made the change and performed any necessary validations,
+it’s time to convert the notebook to Python modules. To do this, simply
+use `nbdev_export` in the terminal.
+
+You will see that the `mlforecast/utils.py` file has been modified (the
+changes from `nbs/utils.ipynb` are reflected in that module). Before
+committing the changes, we need to clean the notebooks using the command
+`./action_files/clean_nbs` and verify that the linters pass using
+`./action_files/lint`:
+
+
+
+Once you have done the above, simply add the changes using
+`git add nbs/utils.ipynb mlforecast/utils.py`:
+
+
+
+Create a descriptive commit message for the changes using
+`git commit -m "[description of changes]"`:
+
+
+
+Finally, push your changes using `git push`:
+
+
+
+### 5. Create a pull request.
+
+In GitHub, open your repository that contains your fork of the original
+repo. Once inside, you will see the changes you just pushed. Click on
+“Compare and pull request”:
+
+
+
+Include an appropriate title for your pull request and fill in the
+necessary information. Once you’re done, click on “Create pull request”.
+
+
+
+Finally, you will see something like this:
+
+
+
+## Notes
+
+- This file was generated using [this
+ file](https://github.com/Nixtla/nixtla-commons/blob/main/docs/contribute/step-by-step.md).
+ Please change that file if you want to enhance the document.
+
diff --git a/statsforecast/docs/contribute/techstack.mdx b/statsforecast/docs/contribute/techstack.mdx
new file mode 100644
index 00000000..229f50c8
--- /dev/null
+++ b/statsforecast/docs/contribute/techstack.mdx
@@ -0,0 +1,24 @@
+# Contributing Code to Nixtla Development
+
+Curious about the skills required to contribute to the Nixtla project?
+
+## Required Skills for Contribution
+
+### Coding
+
+If you’re interested in making code contributions, possessing any of the
+following skills can assist you in getting started:
+
+- [GitHub](https://github.com/)
+- [Python 3](https://www.python.org/)
+- [conda](https://docs.conda.io/en/latest/)
+- [nbdev](https://nbdev.fast.ai/)
+
+### Time Series Theory
+
+- [Forecasting: Principles and Practice](https://otexts.com/fpp3/)
+- [Python Adaptation](https://github.com/Nixtla/fpp3-python) of
+ Forecasting: Principles and Practice
+
+Happy forecasting!
+
diff --git a/statsforecast/docs/distributed/dask.html.mdx b/statsforecast/docs/distributed/dask.html.mdx
new file mode 100644
index 00000000..d4b6b2ae
--- /dev/null
+++ b/statsforecast/docs/distributed/dask.html.mdx
@@ -0,0 +1,82 @@
+---
+description: Run StatsForecast distributedly on top of Dask.
+output-file: dask.html
+title: Dask
+---
+
+
+StatsForecast works on top of Spark, Dask, and Ray through
+[Fugue](https://github.com/fugue-project/fugue/). StatsForecast will
+read the input DataFrame and use the corresponding engine. For example,
+if the input is a Spark DataFrame, StatsForecast will use the existing
+Spark session to run the forecast.
+
+## Installation
+
+As long as Dask is installed and configured, StatsForecast will be able
+to use it. If executing on a distributed Dask cluster, make use the
+`statsforecast` library is installed across all the workers.
+
+## StatsForecast on Pandas
+
+Before running on Dask, it’s recommended to test on a smaller Pandas
+dataset to make sure everything is working. This example also helps show
+the small differences when using Dask.
+
+
+```python
+from statsforecast.core import StatsForecast
+from statsforecast.models import (
+ AutoARIMA,
+ AutoETS,
+)
+from statsforecast.utils import generate_series
+```
+
+
+```python
+n_series = 4
+horizon = 7
+
+series = generate_series(n_series)
+
+sf = StatsForecast(
+ models=[AutoETS(season_length=7)],
+ freq='D',
+)
+sf.forecast(df=series, h=horizon).head()
+```
+
+| | unique_id | ds | AutoETS |
+|-----|-----------|------------|----------|
+| 0 | 0 | 2000-08-10 | 5.261609 |
+| 1 | 0 | 2000-08-11 | 6.196357 |
+| 2 | 0 | 2000-08-12 | 0.282309 |
+| 3 | 0 | 2000-08-13 | 1.264195 |
+| 4 | 0 | 2000-08-14 | 2.262453 |
+
+## Executing on Dask
+
+To run the forecasts distributed on Dask, just pass in a Dask DataFrame
+instead.
+
+
+```python
+import dask.dataframe as dd
+```
+
+
+```python
+series['unique_id'] = series['unique_id'].astype(str)
+ddf = dd.from_pandas(series, npartitions=4)
+sf.forecast(df=ddf, h=horizon).compute().head()
+```
+
+| | unique_id | ds | AutoETS |
+|-----|-----------|---------------------|----------|
+| 0 | 0 | 2000-08-10 00:00:00 | 5.261609 |
+| 1 | 0 | 2000-08-11 00:00:00 | 6.196357 |
+| 2 | 0 | 2000-08-12 00:00:00 | 0.282309 |
+| 3 | 0 | 2000-08-13 00:00:00 | 1.264195 |
+| 4 | 0 | 2000-08-14 00:00:00 | 2.262453 |
+
diff --git a/statsforecast/docs/distributed/ray.html.mdx b/statsforecast/docs/distributed/ray.html.mdx
new file mode 100644
index 00000000..ae96c8d6
--- /dev/null
+++ b/statsforecast/docs/distributed/ray.html.mdx
@@ -0,0 +1,74 @@
+---
+description: Run StatsForecast distributedly on top of Ray.
+output-file: ray.html
+title: Ray
+---
+
+
+## Installation
+
+As long as Ray is installed and configured, StatsForecast will be able
+to use it. If executing on a distributed Ray cluster, make use the
+`statsforecast` library is installed across all the workers.
+
+## StatsForecast on Pandas
+
+Before running on Ray, it’s recommended to test on a smaller Pandas
+dataset to make sure everything is working. This example also helps show
+the small differences when using Ray.
+
+
+```python
+from statsforecast.core import StatsForecast
+from statsforecast.models import AutoARIMA, AutoETS
+from statsforecast.utils import generate_series
+```
+
+
+```python
+n_series = 4
+horizon = 7
+
+series = generate_series(n_series)
+
+sf = StatsForecast(
+ models=[AutoETS(season_length=7)],
+ freq='D',
+)
+sf.forecast(df=series, h=horizon).head()
+```
+
+| | unique_id | ds | AutoETS |
+|-----|-----------|------------|----------|
+| 0 | 0 | 2000-08-10 | 5.261609 |
+| 1 | 0 | 2000-08-11 | 6.196357 |
+| 2 | 0 | 2000-08-12 | 0.282309 |
+| 3 | 0 | 2000-08-13 | 1.264195 |
+| 4 | 0 | 2000-08-14 | 2.262453 |
+
+## Executing on Ray
+
+To run the forecasts distributed on Ray, just pass in a Ray Dataset
+instead.
+
+
+```python
+import ray
+import logging
+```
+
+
+```python
+ray.init(logging_level=logging.ERROR)
+
+series['unique_id'] = series['unique_id'].astype(str)
+ctx = ray.data.context.DatasetContext.get_current()
+ctx.use_streaming_executor = False
+ray_series = ray.data.from_pandas(series).repartition(4)
+```
+
+
+```python
+sf.forecast(df=ray_series, h=horizon).take(5)
+```
+
diff --git a/statsforecast/docs/distributed/spark.html.mdx b/statsforecast/docs/distributed/spark.html.mdx
new file mode 100644
index 00000000..540a442b
--- /dev/null
+++ b/statsforecast/docs/distributed/spark.html.mdx
@@ -0,0 +1,94 @@
+---
+description: Run StatsForecast distributedly on top of Spark.
+output-file: spark.html
+title: Spark
+---
+
+
+StatsForecast works on top of Spark, Dask, and Ray through
+[Fugue](https://github.com/fugue-project/fugue/). StatsForecast will
+read the input DataFrame and use the corresponding engine. For example,
+if the input is a Spark DataFrame, StatsForecast will use the existing
+Spark session to run the forecast.
+
+A benchmark (with older syntax) can be found
+[here](https://medium.com/data-science/distributed-forecast-of-1m-time-series-in-under-15-minutes-with-spark-nixtla-and-fugue-e9892da6fd5c)
+where we forecasted one million timeseries in under 15 minutes.
+
+## Installation
+
+As long as Spark is installed and configured, StatsForecast will be able
+to use it. If executing on a distributed Spark cluster, make use the
+`statsforecast` library is installed across all the workers.
+
+## StatsForecast on Pandas
+
+Before running on Spark, it’s recommended to test on a smaller Pandas
+dataset to make sure everything is working. This example also helps show
+the small differences when using Spark.
+
+
+```python
+from statsforecast.core import StatsForecast
+from statsforecast.models import AutoARIMA, AutoETS
+from statsforecast.utils import generate_series
+```
+
+
+```python
+n_series = 4
+horizon = 7
+
+series = generate_series(n_series)
+
+sf = StatsForecast(
+ models=[AutoETS(season_length=7)],
+ freq='D',
+)
+sf.forecast(df=series, h=horizon).head()
+```
+
+| | unique_id | ds | AutoETS |
+|-----|-----------|------------|----------|
+| 0 | 0 | 2000-08-10 | 5.261609 |
+| 1 | 0 | 2000-08-11 | 6.196357 |
+| 2 | 0 | 2000-08-12 | 0.282309 |
+| 3 | 0 | 2000-08-13 | 1.264195 |
+| 4 | 0 | 2000-08-14 | 2.262453 |
+
+## Executing on Spark
+
+To run the forecasts distributed on Spark, just pass in a Spark
+DataFrame instead.
+
+
+```python
+from pyspark.sql import SparkSession
+```
+
+
+```python
+spark = SparkSession.builder.getOrCreate()
+
+series['unique_id'] = series['unique_id'].astype(str)
+
+# Convert to Spark
+sdf = spark.createDataFrame(series)
+
+# Returns a Spark DataFrame
+sf.forecast(df=sdf, h=horizon, level=[90]).show(5)
+```
+
+``` text
++---------+-------------------+----------+-------------+-------------+
+|unique_id| ds| AutoETS|AutoETS-lo-90|AutoETS-hi-90|
++---------+-------------------+----------+-------------+-------------+
+| 0|2000-08-10 00:00:00| 5.261609| 5.0255513| 5.4976664|
+| 0|2000-08-11 00:00:00| 6.1963573| 5.9603| 6.432415|
+| 0|2000-08-12 00:00:00|0.28230855| 0.04625102| 0.5183661|
+| 0|2000-08-13 00:00:00| 1.2641948| 1.0281373| 1.5002524|
+| 0|2000-08-14 00:00:00| 2.2624528| 2.0263953| 2.4985104|
++---------+-------------------+----------+-------------+-------------+
+only showing top 5 rows
+```
+
diff --git a/statsforecast/docs/experiments/AutoArima_vs_Prophet_files/figure-markdown_strict/cell-16-output-1.png b/statsforecast/docs/experiments/AutoArima_vs_Prophet_files/figure-markdown_strict/cell-16-output-1.png
new file mode 100644
index 00000000..6dcc5c3c
Binary files /dev/null and b/statsforecast/docs/experiments/AutoArima_vs_Prophet_files/figure-markdown_strict/cell-16-output-1.png differ
diff --git a/statsforecast/docs/experiments/AutoArima_vs_Prophet_files/figure-markdown_strict/cell-24-output-1.png b/statsforecast/docs/experiments/AutoArima_vs_Prophet_files/figure-markdown_strict/cell-24-output-1.png
new file mode 100644
index 00000000..062dd8ed
Binary files /dev/null and b/statsforecast/docs/experiments/AutoArima_vs_Prophet_files/figure-markdown_strict/cell-24-output-1.png differ
diff --git a/statsforecast/docs/experiments/AutoArima_vs_Prophet_files/figure-markdown_strict/cell-30-output-1.png b/statsforecast/docs/experiments/AutoArima_vs_Prophet_files/figure-markdown_strict/cell-30-output-1.png
new file mode 100644
index 00000000..44df120a
Binary files /dev/null and b/statsforecast/docs/experiments/AutoArima_vs_Prophet_files/figure-markdown_strict/cell-30-output-1.png differ
diff --git a/statsforecast/docs/experiments/AutoArima_vs_Prophet_files/figure-markdown_strict/cell-35-output-1.png b/statsforecast/docs/experiments/AutoArima_vs_Prophet_files/figure-markdown_strict/cell-35-output-1.png
new file mode 100644
index 00000000..4e7d72ac
Binary files /dev/null and b/statsforecast/docs/experiments/AutoArima_vs_Prophet_files/figure-markdown_strict/cell-35-output-1.png differ
diff --git a/statsforecast/docs/experiments/AutoArima_vs_Prophet_files/figure-markdown_strict/cell-7-output-1.png b/statsforecast/docs/experiments/AutoArima_vs_Prophet_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..ecc42694
Binary files /dev/null and b/statsforecast/docs/experiments/AutoArima_vs_Prophet_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/statsforecast/docs/experiments/AutoArima_vs_Prophet_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/experiments/AutoArima_vs_Prophet_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..2fcd7a7f
Binary files /dev/null and b/statsforecast/docs/experiments/AutoArima_vs_Prophet_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/experiments/amazonstatsforecast.html.mdx b/statsforecast/docs/experiments/amazonstatsforecast.html.mdx
new file mode 100644
index 00000000..33f03e90
--- /dev/null
+++ b/statsforecast/docs/experiments/amazonstatsforecast.html.mdx
@@ -0,0 +1,352 @@
+---
+description: Amazon's AutoML vs open source statistical methods
+output-file: amazonstatsforecast.html
+title: Amazon Forecast vs StatsForecast
+---
+
+
+## Data
+
+We will make use of the [M5
+competition](https://mofc.unic.ac.cy/m5-competition/) dataset provided
+by Walmart. This dataset is interesting for its scale but also the fact
+that it features many timeseries with infrequent occurances. Such
+timeseries are common in retail scenarios and are difficult for
+traditional timeseries forecasting techniques to address.
+
+The data are ready for download at the following URLs:
+
+- Train set:
+ `https://m5-benchmarks.s3.amazonaws.com/data/train/target.parquet`
+- Temporal exogenous variables (used by AmazonForecast):
+ `https://m5-benchmarks.s3.amazonaws.com/data/train/temporal.parquet`
+- Static exogenous variables (used by AmazonForecast):
+ `https://m5-benchmarks.s3.amazonaws.com/data/train/static.parquet`
+
+A more detailed description of the data can be found
+[here](./data.html).
+
+> **Warning**
+>
+> The M5 competition is hierarchical. That is, forecasts are required
+> for different levels of aggregation: national, state, store, etc. In
+> this experiment, we only generate forecasts using the bottom-level
+> data. The evaluation is performed using the bottom-up reconciliation
+> method to obtain the forecasts for the higher hierarchies.
+
+## Amazon Forecast
+
+Amazon Forecast is a fully automated solution for time series
+forecasting. The solution can take the time series to forecast and
+exogenous variables (temporal and static). For this experiment, we used
+the AutoPredict functionality of Amazon Forecast following the steps of
+[this
+tutorial](https://docs.aws.amazon.com/forecast/latest/dg/gs-console.html).
+A detailed description of the particular steps for this dataset can be
+found
+[here](https://nixtlaverse.nixtla.io/statsforecast/docs/experiments/amazonstatsforecast.html).
+
+Amazon Forecast creates predictors with AutoPredictor, which involves
+applying the optimal combination of algorithms to each time series in
+your datasets. The predictor is an Amazon Forecast model that is trained
+using your target time series, related time series, item metadata, and
+any additional datasets you include.
+
+Included algorithms range from commonly used statistical algorithms like
+Autoregressive Integrated Moving Average (ARIMA), to complex neural
+network algorithms like CNN-QR and DeepAR+.: CNN-QR, DeepAR+, Prophet,
+NPTS, ARIMA, and ETS.
+
+To leverage the probabilistic features of Amazon Forecast and enable
+confidence intervals for further analysis we forecasted the following
+quantiles: 0.1 \| 0.5 \| 0.9.
+
+The full pipeline of Amazon Forecast took 4.1 hours and the results can
+be found here: `s3://m5-benchmarks/forecasts/amazonforecast-m5.parquet`
+
+## Nixtla’s StatsForecast
+
+### Install necessary libraries
+
+We assume you have StatsForecast already installed. Check this guide for
+instructions on [how to install StatsForecast](./installation.html).
+
+Additionally, we will install `s3fs` to read from the S3 Filesystem of
+AWS. (If you don’t want to use a cloud storage provider, you can read
+your files locally using pandas)
+
+
+```python
+!pip install statsforecast s3fs
+```
+
+### Input format
+
+We will use pandas to read the data set stored in a parquet file for
+efficiency. You can use ordinary pandas operations to read your data in
+other formats likes `.csv`.
+
+The input to StatsForecast is always a data frame in [long
+format](https://www.theanalysisfactor.com/wide-and-long-data/) with
+three columns: `unique_id`, `ds` and `y`:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast. We
+ will rename the
+
+So we will rename the original columns to make it compatible with
+StatsForecast.
+
+Depending on your internet connection, this step should take around 20
+seconds.
+
+> **Warning**
+>
+> We are reading a file from S3, so you need to install the s3fs
+> library. To install it, run `! pip install s3fs`
+
+### Read data
+
+
+```python
+import pandas as pd
+
+Y_df_m5 = pd.read_parquet('https://m5-benchmarks.s3.amazonaws.com/data/train/target.parquet')
+
+Y_df_m5 = Y_df_m5.rename(columns={
+ 'item_id': 'unique_id',
+ 'timestamp': 'ds',
+ 'demand': 'y'
+})
+
+Y_df_m5.head()
+```
+
+| | unique_id | ds | y |
+|-----|------------------|------------|-----|
+| 0 | FOODS_1_001_CA_1 | 2011-01-29 | 3.0 |
+| 1 | FOODS_1_001_CA_1 | 2011-01-30 | 0.0 |
+| 2 | FOODS_1_001_CA_1 | 2011-01-31 | 0.0 |
+| 3 | FOODS_1_001_CA_1 | 2011-02-01 | 1.0 |
+| 4 | FOODS_1_001_CA_1 | 2011-02-02 | 4.0 |
+
+### Train statistical models
+
+We fit the model by instantiating a new
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+object with the following parameters:
+
+- `models`: a list of models. Select the models you want from
+ [models](../models.html) and import them. For this example, we will
+ use
+ [`AutoETS`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoets)
+ and
+ [`DynamicOptimizedTheta`](https://Nixtla.github.io/statsforecast/src/core/models.html#dynamicoptimizedtheta).
+ We set `season_length` to 7 because we expect seasonal effects every
+ week. (See: [Seasonal
+ periods](https://robjhyndman.com/hyndsight/seasonal-periods/))
+
+- `freq`: a string indicating the frequency of the data. (See [panda’s
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs`: n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model`: a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+> **Note**
+>
+> StatsForecast achieves its blazing speed using JIT compiling through
+> Numba. The first time you call the statsforecast class, the fit method
+> should take around 5 seconds. The second time -once Numba compiled
+> your settings- it should take less than 0.2s.
+
+- [`AutoETS`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoets):
+ Exponential Smoothing model. Automatically selects the best ETS
+ (Error, Trend, Seasonality) model using an information criterion.
+ Ref:
+ [`AutoETS`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoets).
+
+- [`SeasonalNaive`](https://Nixtla.github.io/statsforecast/src/core/models.html#seasonalnaive):
+ Memory Efficient Seasonal Naive predictions. Ref:
+ [`SeasonalNaive`](https://Nixtla.github.io/statsforecast/src/core/models.html#seasonalnaive).
+
+- [`DynamicOptimizedTheta`](https://Nixtla.github.io/statsforecast/src/core/models.html#dynamicoptimizedtheta):
+ fit two theta lines to a deseasonalized time series, using different
+ techniques to obtain and combine the two theta lines to produce the
+ final forecasts. Ref:
+ [`DynamicOptimizedTheta`](https://Nixtla.github.io/statsforecast/src/core/models.html#dynamicoptimizedtheta).
+
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import (
+ AutoETS,
+ DynamicOptimizedTheta,
+ SeasonalNaive
+)
+
+# Create list of models
+models = [
+ AutoETS(season_length=7),
+ DynamicOptimizedTheta(season_length=7),
+]
+
+# Instantiate StatsForecast class
+sf = StatsForecast(
+ models=models,
+ freq='D',
+ n_jobs=-1,
+ fallback_model=SeasonalNaive(season_length=7)
+)
+```
+
+``` text
+/home/ubuntu/fede/statsforecast/statsforecast/core.py:21: TqdmExperimentalWarning: Using `tqdm.autonotebook.tqdm` in notebook mode. Use `tqdm.tqdm` instead to force console mode (e.g. in jupyter console)
+ from tqdm.autonotebook import tqdm
+```
+
+The `forecast` method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h` (int): represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level` (list of floats): this optional parameter is used for
+ probabilistic forecasting. Set the `level` (or confidence
+ percentile) of your prediction interval. For example, `level=[90]`
+ means that the model expects the real value to be inside that
+ interval 90% of the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+> **Note**
+>
+> The `forecast` is inteded to be compatible with distributed clusters,
+> so it does not store any model parameters. If you want to store
+> parameter for everymodel you can use the `fit` and `predict` methods.
+> However, those methods are not defined for distrubed engines like
+> Spark, Ray or Dask.
+
+
+```python
+from time import time
+```
+
+
+```python
+init = time()
+forecasts_df = sf.forecast(df=Y_df_m5, h=28)
+end = time()
+print(f'Statsforecast time M5 {(end - init) / 60}')
+```
+
+``` text
+Statsforecast time M5 14.274124479293823
+```
+
+Store the results for further evaluation.
+
+
+```python
+forecasts_df['ThETS'] = forecasts_df[['DynamicOptimizedTheta', 'AutoETS']].clip(0).median(axis=1, numeric_only=True)
+forecasts_df.to_parquet('s3://m5-benchmarks/forecasts/statsforecast-m5.parquet')
+```
+
+## Evaluation
+
+This section evaluates the performance of
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+and `AmazonForecast`. To do this, we first need to install
+[datasetsforecast](https://github.com/Nixtla/datasetsforecast), a Python
+library developed by Nixtla that includes a large battery of benchmark
+datasets and evaluation utilities. The library will allow us to
+calculate the performance of the models using the original evaluation
+used in the competition.
+
+
+```python
+!pip install datasetsforecast
+```
+
+
+```python
+from datasetsforecast.m5 import M5, M5Evaluation
+```
+
+The following function will allow us to evaluate a specific model
+included in the input dataframe. The function is useful for evaluating
+different models.
+
+
+```python
+from datasetsforecast.m5 import M5, M5Evaluation
+from statsforecast import StatsForecast
+
+### Evaluator
+def evaluate_forecasts(df, model, model_name):
+ Y_hat = df.set_index('ds', append=True)[model].unstack()
+ *_, S_df = M5.load('data')
+ Y_hat = S_df.merge(Y_hat, how='left', on=['unique_id'])
+ eval_ = M5Evaluation.evaluate(y_hat=Y_hat, directory='./data')
+ eval_ = eval_.rename(columns={'wrmsse': f'{model_name}_{model}_wrmsse'})
+ return eval_
+```
+
+Now let’s read the forecasts generated for each solution.
+
+
+```python
+### Read Forecasts
+statsforecasts_df = pd.read_parquet('s3://m5-benchmarks/forecasts/statsforecast-m5.parquet')
+amazonforecasts_df = pd.read_parquet('s3://m5-benchmarks/forecasts/amazonforecast-m5.parquet')
+
+### Amazon Forecast wrangling
+amazonforecasts_df = amazonforecasts_df.rename(columns={'item_id': 'unique_id', 'date': 'ds'})
+# amazon forecast returns the unique_id column in lower case
+# we need to transform it to upper case to ensure proper merging
+amazonforecasts_df['unique_id'] = amazonforecasts_df['unique_id'].str.upper()
+amazonforecasts_df = amazonforecasts_df.set_index('unique_id')
+# parse datestamp
+amazonforecasts_df['ds'] = pd.to_datetime(amazonforecasts_df['ds']).dt.tz_localize(None)
+```
+
+Finally, let’s use our predefined function to compute the performance of
+each model.
+
+
+```python
+### Evaluate performances
+m5_eval_df = pd.concat([
+ evaluate_forecasts(statsforecasts_df, 'ThETS', 'StatsForecast'),
+ evaluate_forecasts(statsforecasts_df, 'AutoETS', 'StatsForecast'),
+ evaluate_forecasts(statsforecasts_df, 'DynamicOptimizedTheta', 'StatsForecast'),
+ evaluate_forecasts(amazonforecasts_df, 'p50', 'AmazonForecast'),
+], axis=1)
+m5_eval_df.T
+```
+
+| | Total | Level1 | Level2 | Level3 | Level4 | Level5 | Level6 | Level7 | Level8 | Level9 | Level10 | Level11 | Level12 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| StatsForecast_ThETS_wrmsse | 0.669606 | 0.424331 | 0.515777 | 0.580670 | 0.474098 | 0.552459 | 0.578092 | 0.651079 | 0.642446 | 0.725324 | 1.009390 | 0.967537 | 0.914068 |
+| StatsForecast_AutoETS_wrmsse | 0.672404 | 0.430474 | 0.516340 | 0.580736 | 0.482090 | 0.559721 | 0.579939 | 0.655362 | 0.643638 | 0.727967 | 1.010596 | 0.968168 | 0.913820 |
+| StatsForecast_DynamicOptimizedTheta_wrmsse | 0.675333 | 0.429670 | 0.521640 | 0.589278 | 0.478730 | 0.557520 | 0.584278 | 0.656283 | 0.650613 | 0.731735 | 1.013910 | 0.971758 | 0.918576 |
+| AmazonForecast_p50_wrmsse | 1.617815 | 1.912144 | 1.786991 | 1.736382 | 1.972658 | 2.010498 | 1.805926 | 1.819329 | 1.667225 | 1.619216 | 1.156432 | 1.012942 | 0.914040 |
+
+The results (including processing time and costs) can be summarized in
+the following table.
+
+
+
diff --git a/statsforecast/docs/experiments/autoarima_vs_prophet.html.mdx b/statsforecast/docs/experiments/autoarima_vs_prophet.html.mdx
new file mode 100644
index 00000000..ba49fbbf
--- /dev/null
+++ b/statsforecast/docs/experiments/autoarima_vs_prophet.html.mdx
@@ -0,0 +1,655 @@
+---
+output-file: autoarima_vs_prophet.html
+title: AutoARIMA Comparison (Prophet and pmdarima)
+---
+
+
+
+
+## Motivation
+
+The
+[`AutoARIMA`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoarima)
+model is widely used to forecast time series in production and as a
+benchmark. However, the python implementation (`pmdarima`) is so slow
+that prevent data scientist practioners from quickly iterating and
+deploying
+[`AutoARIMA`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoarima)
+in production for a large number of time series. In this notebook we
+present Nixtla’s
+[`AutoARIMA`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoarima)
+based on the R implementation (developed by Rob Hyndman) and optimized
+using `numba`.
+
+## Example
+
+### Libraries
+
+
+```python
+# !pip install statsforecast prophet statsmodels sklearn matplotlib pmdarima
+```
+
+
+```python
+import logging
+import os
+import random
+import time
+import warnings
+warnings.filterwarnings("ignore")
+from itertools import product
+from multiprocessing import cpu_count, Pool # for prophet
+
+import matplotlib.pyplot as plt
+import numpy as np
+import pandas as pd
+from pmdarima import auto_arima as auto_arima_p
+from prophet import Prophet
+from statsforecast import StatsForecast
+from statsforecast.models import AutoARIMA, _TS
+from statsmodels.graphics.tsaplots import plot_acf
+from sklearn.model_selection import ParameterGrid
+from utilsforecast.plotting import plot_series
+```
+
+#### Useful functions
+
+
+```python
+def plot_autocorrelation_grid(df_train):
+ fig, axes = plt.subplots(4, 2, figsize = (24, 14))
+
+ unique_ids = df_train['unique_id'].unique()
+
+ assert len(unique_ids) >= 8, "Must provide at least 8 ts"
+
+ unique_ids = random.sample(list(unique_ids), k=8)
+
+ for uid, (idx, idy) in zip(unique_ids, product(range(4), range(2))):
+ train_uid = df_train.query('unique_id == @uid')
+ plot_acf(train_uid['y'].values, ax=axes[idx, idy],
+ title=f'ACF M4 Hourly {uid}')
+ axes[idx, idy].set_xlabel('Timestamp [t]')
+ axes[idx, idy].set_ylabel('Autocorrelation')
+ fig.subplots_adjust(hspace=0.5)
+ plt.show()
+```
+
+### Data
+
+For testing purposes, we will use the Hourly dataset from the M4
+competition.
+
+
+```python
+train = pd.read_csv('https://auto-arima-results.s3.amazonaws.com/M4-Hourly.csv')
+test = pd.read_csv('https://auto-arima-results.s3.amazonaws.com/M4-Hourly-test.csv').rename(columns={'y': 'y_test'})
+```
+
+In this example we will use a subset of the data to avoid waiting too
+long. You can modify the number of series if you want.
+
+
+```python
+n_series = 16
+uids = train['unique_id'].unique()[:n_series]
+train = train.query('unique_id in @uids')
+test = test.query('unique_id in @uids')
+```
+
+
+```python
+plot_series(train, test, max_ids=n_series)
+```
+
+
+
+Would an autorregresive model be the right choice for our data? There is
+no doubt that we observe seasonal periods. The autocorrelation function
+(`acf`) can help us to answer the question. Intuitively, we have to
+observe a decreasing correlation to opt for an AR model.
+
+
+```python
+plot_autocorrelation_grid(train)
+```
+
+
+
+Thus, we observe a high autocorrelation for previous lags and also for
+the seasonal lags. Therefore, we will let `auto_arima` to handle our
+data.
+
+### Training and forecasting
+
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+receives a list of models to fit each time series. Since we are dealing
+with Hourly data, it would be benefitial to use 24 as seasonality.
+
+
+```python
+?AutoARIMA
+```
+
+``` text
+Init signature:
+AutoARIMA(
+ d: Optional[int] = None,
+ D: Optional[int] = None,
+ max_p: int = 5,
+ max_q: int = 5,
+ max_P: int = 2,
+ max_Q: int = 2,
+ max_order: int = 5,
+ max_d: int = 2,
+ max_D: int = 1,
+ start_p: int = 2,
+ start_q: int = 2,
+ start_P: int = 1,
+ start_Q: int = 1,
+ stationary: bool = False,
+ seasonal: bool = True,
+ ic: str = 'aicc',
+ stepwise: bool = True,
+ nmodels: int = 94,
+ trace: bool = False,
+ approximation: Optional[bool] = False,
+ method: Optional[str] = None,
+ truncate: Optional[bool] = None,
+ test: str = 'kpss',
+ test_kwargs: Optional[str] = None,
+ seasonal_test: str = 'seas',
+ seasonal_test_kwargs: Optional[Dict] = None,
+ allowdrift: bool = False,
+ allowmean: bool = False,
+ blambda: Optional[float] = None,
+ biasadj: bool = False,
+ season_length: int = 1,
+ alias: str = 'AutoARIMA',
+ prediction_intervals: Optional[statsforecast.utils.ConformalIntervals] = None,
+)
+Docstring:
+AutoARIMA model.
+
+Automatically selects the best ARIMA (AutoRegressive Integrated Moving Average)
+model using an information criterion. Default is Akaike Information Criterion (AICc).
+
+**Note:**
+This implementation is a mirror of Hyndman's [forecast::auto.arima](https://github.com/robjhyndman/forecast).
+
+**References:**
+[Rob J. Hyndman, Yeasmin Khandakar (2008). "Automatic Time Series Forecasting: The forecast package for R"](https://www.jstatsoft.org/article/view/v027i03).
+
+Parameters
+----------
+d : Optional[int]
+ Order of first-differencing.
+D : Optional[int]
+ Order of seasonal-differencing.
+max_p : int
+ Max autorregresives p.
+max_q : int
+ Max moving averages q.
+max_P : int
+ Max seasonal autorregresives P.
+max_Q : int
+ Max seasonal moving averages Q.
+max_order : int
+ Max p+q+P+Q value if not stepwise selection.
+max_d : int
+ Max non-seasonal differences.
+max_D : int
+ Max seasonal differences.
+start_p : int
+ Starting value of p in stepwise procedure.
+start_q : int
+ Starting value of q in stepwise procedure.
+start_P : int
+ Starting value of P in stepwise procedure.
+start_Q : int
+ Starting value of Q in stepwise procedure.
+stationary : bool
+ If True, restricts search to stationary models.
+seasonal : bool
+ If False, restricts search to non-seasonal models.
+ic : str
+ Information criterion to be used in model selection.
+stepwise : bool
+ If True, will do stepwise selection (faster).
+nmodels : int
+ Number of models considered in stepwise search.
+trace : bool
+ If True, the searched ARIMA models is reported.
+approximation : Optional[bool]
+ If True, conditional sums-of-squares estimation, final MLE.
+method : Optional[str]
+ Fitting method between maximum likelihood or sums-of-squares.
+truncate : Optional[int]
+ Observations truncated series used in model selection.
+test : str
+ Unit root test to use. See `ndiffs` for details.
+test_kwargs : Optional[str]
+ Unit root test additional arguments.
+seasonal_test : str
+ Selection method for seasonal differences.
+seasonal_test_kwargs : Optional[dict]
+ Seasonal unit root test arguments.
+allowdrift : bool (default True)
+ If True, drift models terms considered.
+allowmean : bool (default True)
+ If True, non-zero mean models considered.
+blambda : Optional[float]
+ Box-Cox transformation parameter.
+biasadj : bool
+ Use adjusted back-transformed mean Box-Cox.
+season_length : int
+ Number of observations per unit of time. Ex: 24 Hourly data.
+alias : str
+ Custom name of the model.
+prediction_intervals : Optional[ConformalIntervals]
+ Information to compute conformal prediction intervals.
+ By default, the model will compute the native prediction
+ intervals.
+File: /hdd/github/statsforecast/statsforecast/models.py
+Type: type
+Subclasses:
+```
+
+As we see, we can pass `season_length` to
+[`AutoARIMA`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoarima),
+so the definition of our models would be,
+
+
+```python
+models = [AutoARIMA(season_length=24, approximation=True)]
+```
+
+
+```python
+fcst = StatsForecast(df=train,
+ models=models,
+ freq='H',
+ n_jobs=-1)
+```
+
+
+```python
+init = time.time()
+forecasts = fcst.forecast(48)
+end = time.time()
+
+time_nixtla = end - init
+time_nixtla
+```
+
+``` text
+40.38660216331482
+```
+
+
+```python
+forecasts.head()
+```
+
+| | ds | AutoARIMA |
+|-----------|-----|------------|
+| unique_id | | |
+| H1 | 701 | 616.084167 |
+| H1 | 702 | 544.432129 |
+| H1 | 703 | 510.414490 |
+| H1 | 704 | 481.046539 |
+| H1 | 705 | 460.893066 |
+
+
+```python
+forecasts = forecasts.reset_index()
+```
+
+
+```python
+test = test.merge(forecasts, how='left', on=['unique_id', 'ds'])
+```
+
+
+```python
+plot_series(train, test)
+```
+
+
+
+## Alternatives
+
+### pmdarima
+
+You can use the
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+class to parallelize your own models. In this section we will use it to
+run the `auto_arima` model from `pmdarima`.
+
+
+```python
+class PMDAutoARIMA(_TS):
+
+ def __init__(self, season_length: int):
+ self.season_length = season_length
+
+ def forecast(self, y, h, X=None, X_future=None, fitted=False):
+ mod = auto_arima_p(
+ y, m=self.season_length,
+ with_intercept=False #ensure comparability with Nixtla's implementation
+ )
+ return {'mean': mod.predict(h)}
+
+ def __repr__(self):
+ return 'pmdarima'
+```
+
+
+```python
+n_series_pmdarima = 2
+```
+
+
+```python
+fcst = StatsForecast(
+ df = train.query('unique_id in ["H1", "H10"]'),
+ models=[PMDAutoARIMA(season_length=24)],
+ freq='H',
+ n_jobs=-1
+)
+```
+
+
+```python
+init = time.time()
+forecast_pmdarima = fcst.forecast(48)
+end = time.time()
+
+time_pmdarima = end - init
+time_pmdarima
+```
+
+``` text
+886.2768685817719
+```
+
+
+```python
+forecast_pmdarima.head()
+```
+
+| | ds | pmdarima |
+|-----------|-----|------------|
+| unique_id | | |
+| H1 | 701 | 628.310547 |
+| H1 | 702 | 571.659851 |
+| H1 | 703 | 543.504700 |
+| H1 | 704 | 517.539062 |
+| H1 | 705 | 502.829559 |
+
+
+```python
+forecast_pmdarima = forecast_pmdarima.reset_index()
+```
+
+
+```python
+test = test.merge(forecast_pmdarima, how='left', on=['unique_id', 'ds'])
+```
+
+
+```python
+plot_series(train, test, plot_random=False)
+```
+
+
+
+### Prophet
+
+`Prophet` is designed to receive a pandas dataframe, so we cannot use
+`StatForecast`. Therefore, we need to parallize from scratch.
+
+
+```python
+params_grid = {'seasonality_mode': ['multiplicative','additive'],
+ 'growth': ['linear', 'flat'],
+ 'changepoint_prior_scale': [0.1, 0.2, 0.3, 0.4, 0.5],
+ 'n_changepoints': [5, 10, 15, 20]}
+grid = ParameterGrid(params_grid)
+```
+
+
+```python
+def fit_and_predict(index, ts):
+ df = ts.drop(columns='unique_id', axis=1)
+ max_ds = df['ds'].max()
+ df['ds'] = pd.date_range(start='1970-01-01', periods=df.shape[0], freq='H')
+ df_val = df.tail(48)
+ df_train = df.drop(df_val.index)
+ y_val = df_val['y'].values
+
+ if len(df_train) >= 48:
+ val_results = {'losses': [], 'params': []}
+
+ for params in grid:
+ model = Prophet(seasonality_mode=params['seasonality_mode'],
+ growth=params['growth'],
+ weekly_seasonality=True,
+ daily_seasonality=True,
+ yearly_seasonality=True,
+ n_changepoints=params['n_changepoints'],
+ changepoint_prior_scale=params['changepoint_prior_scale'])
+ model = model.fit(df_train)
+
+ forecast = model.make_future_dataframe(periods=48,
+ include_history=False,
+ freq='H')
+ forecast = model.predict(forecast)
+ forecast['unique_id'] = index
+ forecast = forecast.filter(items=['unique_id', 'ds', 'yhat'])
+
+ loss = np.mean(abs(y_val - forecast['yhat'].values))
+
+ val_results['losses'].append(loss)
+ val_results['params'].append(params)
+
+ idx_params = np.argmin(val_results['losses'])
+ params = val_results['params'][idx_params]
+ else:
+ params = {'seasonality_mode': 'multiplicative',
+ 'growth': 'flat',
+ 'n_changepoints': 150,
+ 'changepoint_prior_scale': 0.5}
+ model = Prophet(seasonality_mode=params['seasonality_mode'],
+ growth=params['growth'],
+ weekly_seasonality=True,
+ daily_seasonality=True,
+ yearly_seasonality=True,
+ n_changepoints=params['n_changepoints'],
+ changepoint_prior_scale=params['changepoint_prior_scale'])
+ model = model.fit(df)
+
+ forecast = model.make_future_dataframe(periods=48,
+ include_history=False,
+ freq='H')
+ forecast = model.predict(forecast)
+ forecast.insert(0, 'unique_id', index)
+ forecast['ds'] = np.arange(max_ds + 1, max_ds + 48 + 1)
+ forecast = forecast.filter(items=['unique_id', 'ds', 'yhat'])
+
+ return forecast
+```
+
+
+```python
+init = time.time()
+with Pool(cpu_count()) as pool:
+ forecast_prophet = pool.starmap(fit_and_predict, train.groupby('unique_id'))
+end = time.time()
+forecast_prophet = pd.concat(forecast_prophet).rename(columns={'yhat': 'prophet'})
+time_prophet = end - init
+time_prophet
+```
+
+``` text
+120.7272641658783
+```
+
+
+```python
+forecast_prophet
+```
+
+| | unique_id | ds | prophet |
+|-----|-----------|-----|-------------|
+| 0 | H1 | 701 | 635.914254 |
+| 1 | H1 | 702 | 565.976464 |
+| 2 | H1 | 703 | 505.095507 |
+| 3 | H1 | 704 | 462.559539 |
+| 4 | H1 | 705 | 438.766801 |
+| ... | ... | ... | ... |
+| 43 | H112 | 744 | 6184.686240 |
+| 44 | H112 | 745 | 6188.851888 |
+| 45 | H112 | 746 | 6129.306256 |
+| 46 | H112 | 747 | 6058.040672 |
+| 47 | H112 | 748 | 5991.982370 |
+
+
+```python
+test = test.merge(forecast_prophet, how='left', on=['unique_id', 'ds'])
+```
+
+
+```python
+plot_series(train, test)
+```
+
+
+
+### Evaluation
+
+### Time
+
+Since
+[`AutoARIMA`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoarima)
+works with numba is useful to calculate the time for just one time
+series.
+
+
+```python
+fcst = StatsForecast(df=train.query('unique_id == "H1"'),
+ models=models, freq='H',
+ n_jobs=1)
+```
+
+
+```python
+init = time.time()
+forecasts = fcst.forecast(48)
+end = time.time()
+
+time_nixtla_1 = end - init
+time_nixtla_1
+```
+
+``` text
+18.752424716949463
+```
+
+
+```python
+times = pd.DataFrame({'n_series': np.arange(1, 414 + 1)})
+times['pmdarima'] = time_pmdarima * times['n_series'] / n_series_pmdarima
+times['prophet'] = time_prophet * times['n_series'] / n_series
+times['AutoARIMA_nixtla'] = time_nixtla_1 + times['n_series'] * (time_nixtla - time_nixtla_1) / n_series
+times = times.set_index('n_series')
+```
+
+
+```python
+times.tail(5)
+```
+
+| | pmdarima | prophet | AutoARIMA_nixtla |
+|----------|---------------|-------------|------------------|
+| n_series | | | |
+| 410 | 181686.758059 | 3093.636144 | 573.128222 |
+| 411 | 182129.896494 | 3101.181598 | 574.480358 |
+| 412 | 182573.034928 | 3108.727052 | 575.832494 |
+| 413 | 183016.173362 | 3116.272506 | 577.184630 |
+| 414 | 183459.311796 | 3123.817960 | 578.536766 |
+
+
+```python
+fig, axes = plt.subplots(1, 2, figsize = (24, 7))
+(times/3600).plot(ax=axes[0], linewidth=4)
+np.log10(times).plot(ax=axes[1], linewidth=4)
+axes[0].set_title('Time across models [Hours]', fontsize=22)
+axes[1].set_title('Time across models [Log10 Scale]', fontsize=22)
+axes[0].set_ylabel('Time [Hours]', fontsize=20)
+axes[1].set_ylabel('Time Seconds [Log10 Scale]', fontsize=20)
+fig.suptitle('Time comparison using M4-Hourly data', fontsize=27)
+for ax in axes:
+ ax.set_xlabel('Number of Time Series [N]', fontsize=20)
+ ax.legend(prop={'size': 20})
+ ax.grid()
+ for label in (ax.get_xticklabels() + ax.get_yticklabels()):
+ label.set_fontsize(20)
+```
+
+
+
+
+```python
+fig.savefig('computational-efficiency.png', dpi=300)
+```
+
+### Performance
+
+#### pmdarima (only two time series)
+
+
+```python
+name_models = test.drop(columns=['unique_id', 'ds', 'y_test']).columns.tolist()
+```
+
+
+```python
+test_pmdarima = test.query('unique_id in ["H1", "H10"]')
+eval_pmdarima = []
+for model in name_models:
+ mae = np.mean(abs(test_pmdarima[model] - test_pmdarima['y_test']))
+ eval_pmdarima.append({'model': model, 'mae': mae})
+pd.DataFrame(eval_pmdarima).sort_values('mae')
+```
+
+| | model | mae |
+|-----|-----------|-----------|
+| 0 | AutoARIMA | 20.289669 |
+| 1 | pmdarima | 24.676279 |
+| 2 | prophet | 39.201933 |
+
+#### Prophet
+
+
+```python
+eval_prophet = []
+for model in name_models:
+ if 'pmdarima' in model:
+ continue
+ mae = np.mean(abs(test[model] - test['y_test']))
+ eval_prophet.append({'model': model, 'mae': mae})
+pd.DataFrame(eval_prophet).sort_values('mae')
+```
+
+| | model | mae |
+|-----|-----------|-------------|
+| 0 | AutoARIMA | 680.202965 |
+| 1 | prophet | 1058.578963 |
+
+For a complete comparison check the [complete
+experiment](https://github.com/Nixtla/statsforecast/tree/v0.6.0/experiments/arima).
+
+
+
diff --git a/statsforecast/docs/experiments/ets_ray_m5.html.mdx b/statsforecast/docs/experiments/ets_ray_m5.html.mdx
new file mode 100644
index 00000000..5f08e4a1
--- /dev/null
+++ b/statsforecast/docs/experiments/ets_ray_m5.html.mdx
@@ -0,0 +1,171 @@
+---
+description: Forecast the M5 dataset
+output-file: ets_ray_m5.html
+title: Forecasting at Scale using ETS and ray (M5)
+---
+
+
+In this notebook we show how to use
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+and `ray` to forecast thounsands of time series in less than 6 minutes
+(M5 dataset). Also, we show that
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+has better performance in time and accuracy compared to [`Prophet`
+running on a Spark
+cluster](http:nixtla.github.io/statsforecast/examples/Prophet_spark_m5.html)
+using DataBricks.
+
+In this example, we used a ray cluster (AWS) of 11 instances of type
+m5.2xlarge (8 cores, 32 GB RAM).
+
+## Installing StatsForecast Library
+
+
+```python
+!pip install "statsforecast[ray]" neuralforecast s3fs pyarrow
+```
+
+
+```python
+from time import time
+
+import pandas as pd
+from neuralforecast.data.datasets.m5 import M5, M5Evaluation
+from statsforecast import StatsForecast
+from statsforecast.models import ETS
+```
+
+## Download data
+
+The example uses the [M5
+dataset](https://github.com/Mcompetitions/M5-methods/blob/master/M5-Competitors-Guide.pdf).
+It consists of `30,490` bottom time series.
+
+
+```python
+Y_df = pd.read_parquet('s3://m5-benchmarks/data/train/target.parquet')
+Y_df = Y_df.rename(columns={
+ 'item_id': 'unique_id',
+ 'timestamp': 'ds',
+ 'demand': 'y'
+})
+Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+```
+
+
+```python
+Y_df.head()
+```
+
+| | unique_id | ds | y |
+|-----|------------------|------------|-----|
+| 0 | FOODS_1_001_CA_1 | 2011-01-29 | 3.0 |
+| 1 | FOODS_1_001_CA_1 | 2011-01-30 | 0.0 |
+| 2 | FOODS_1_001_CA_1 | 2011-01-31 | 0.0 |
+| 3 | FOODS_1_001_CA_1 | 2011-02-01 | 1.0 |
+| 4 | FOODS_1_001_CA_1 | 2011-02-02 | 4.0 |
+
+Since the M5 dataset contains intermittent time series, we add a
+constant to avoid problems during the training phase. Later, we will
+substract the constant from the forecasts.
+
+
+```python
+constant = 10
+Y_df['y'] += constant
+```
+
+## Train the model
+
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+receives a list of models to fit each time series. Since we are dealing
+with Daily data, it would be benefitial to use 7 as seasonality. Observe
+that we need to pass the ray address to the `ray_address` argument.
+
+
+```python
+fcst = StatsForecast(
+ df=Y_df,
+ models=[ETS(season_length=7, model='ZNA')],
+ freq='D',
+ #n_jobs=-1
+ ray_address='ray://ADDRESS:10001'
+)
+```
+
+
+```python
+init = time()
+Y_hat = fcst.forecast(28)
+end = time()
+print(f'Minutes taken by StatsForecast using: {(end - init) / 60}')
+```
+
+``` text
+/home/ubuntu/miniconda/envs/ray/lib/python3.7/site-packages/ray/util/client/worker.py:618: UserWarning: More than 10MB of messages have been created to schedule tasks on the server. This can be slow on Ray Client due to communication overhead over the network. If you're running many fine-grained tasks, consider running them inside a single remote function. See the section on "Too fine-grained tasks" in the Ray Design Patterns document for more details: https://docs.google.com/document/d/167rnnDFIVRhHhK4mznEIemOtj63IOhtIPvSYaPgI4Fg/edit#heading=h.f7ins22n6nyl. If your functions frequently use large objects, consider storing the objects remotely with ray.put. An example of this is shown in the "Closure capture of large / unserializable object" section of the Ray Design Patterns document, available here: https://docs.google.com/document/d/167rnnDFIVRhHhK4mznEIemOtj63IOhtIPvSYaPgI4Fg/edit#heading=h.1afmymq455wu
+ UserWarning,
+```
+
+``` text
+Minutes taken by StatsForecast using: 5.4817593971888225
+```
+
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+and `ray` took only 5.48 minutes to train `30,490` time series, compared
+to 18.23 minutes for Prophet and Spark.
+
+We remove the constant.
+
+
+```python
+Y_hat['ETS'] -= constant
+```
+
+### Evaluating performance
+
+The M5 competition used the weighted root mean squared scaled error. You
+can find details of the metric
+[here](https://github.com/Mcompetitions/M5-methods/blob/master/M5-Competitors-Guide.pdf).
+
+
+```python
+Y_hat = Y_hat.reset_index().set_index(['unique_id', 'ds']).unstack()
+Y_hat = Y_hat.droplevel(0, 1).reset_index()
+```
+
+
+```python
+*_, S_df = M5.load('./data')
+Y_hat = S_df.merge(Y_hat, how='left', on=['unique_id'])
+```
+
+``` text
+100%|███████████████████████████████████████████████████████████| 50.2M/50.2M [00:00<00:00, 77.1MiB/s]
+```
+
+
+```python
+M5Evaluation.evaluate(y_hat=Y_hat, directory='./data')
+```
+
+| | wrmsse |
+|---------|----------|
+| Total | 0.677233 |
+| Level1 | 0.435558 |
+| Level2 | 0.522863 |
+| Level3 | 0.582109 |
+| Level4 | 0.488484 |
+| Level5 | 0.567825 |
+| Level6 | 0.587605 |
+| Level7 | 0.662774 |
+| Level8 | 0.647712 |
+| Level9 | 0.732107 |
+| Level10 | 1.013124 |
+| Level11 | 0.970465 |
+| Level12 | 0.916175 |
+
+Also,
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+is more accurate than Prophet, since the overall WMRSSE is `0.68`,
+against `0.77` obtained by prophet.
+
diff --git a/statsforecast/docs/experiments/prophet_spark_m5.html.mdx b/statsforecast/docs/experiments/prophet_spark_m5.html.mdx
new file mode 100644
index 00000000..d319c2e4
--- /dev/null
+++ b/statsforecast/docs/experiments/prophet_spark_m5.html.mdx
@@ -0,0 +1,288 @@
+---
+description: This notebook was originally executed using DataBricks
+output-file: prophet_spark_m5.html
+title: StatsForecast ETS and Facebook Prophet on Spark (M5)
+---
+
+
+The purpose of this notebook is to create a scalability benchmark (time
+and performance). To that end, Nixtla’s
+[StatsForecast](https://github.com/Nixtla/statsforecast) (using the ETS
+model) is trained on the M5 dataset using spark to distribute the
+training. As a comparison, Facebook’s
+[Prophet](https://github.com/facebook/prophet) model is used.
+
+An AWS cluster (mounted on databricks) of 11 instances of type
+m5.2xlarge (8 cores, 32 GB RAM) with runtime 10.4 LTS was used.
+[This](https://d1r5llqwmkrl74.cloudfront.net/notebooks/RCG/Fine_Grained_Demand_Forecasting/index.html#Fine_Grained_Demand_Forecasting_1.html)
+notebook was used as base case.
+
+The example uses the [M5
+dataset](https://github.com/Mcompetitions/M5-methods/blob/master/M5-Competitors-Guide.pdf).
+It consists of `30,490` bottom time series.
+
+## Main results
+
+| Method | Time (mins) | Performance (wRMSSE) |
+|---------------|------------:|---------------------:|
+| StatsForecast | 7.5 | 0.68 |
+| Prophet | 18.23 | 0.77 |
+
+## Installing libraries
+
+
+```python
+pip install prophet "neuralforecast<1.0.0" "statsforecast[fugue]"
+```
+
+## StatsForecast pipeline
+
+
+```python
+from time import time
+
+from neuralforecast.data.datasets.m5 import M5, M5Evaluation
+from statsforecast.distributed.utils import forecast
+from statsforecast.distributed.fugue import FugueBackend
+from statsforecast.models import ETS, SeasonalNaive
+from statsforecast.core import StatsForecast
+
+from pyspark.sql import SparkSession
+```
+
+
+```python
+spark = SparkSession.builder.getOrCreate()
+backend = FugueBackend(spark, {"fugue.spark.use_pandas_udf":True})
+```
+
+### Forecast
+
+With statsforecast you don’t have to download your data. The distributed
+backend can handle a file with your data.
+
+
+```python
+init = time()
+ets_forecasts = backend.forecast(
+ "s3://m5-benchmarks/data/train/m5-target.parquet",
+ [ETS(season_length=7, model='ZAA')],
+ freq="D",
+ h=28,
+).toPandas()
+end = time()
+print(f'Minutes taken by StatsForecast on a Spark cluster: {(end - init) / 60}')
+```
+
+### Evaluating performance
+
+The M5 competition used the weighted root mean squared scaled error. You
+can find details of the metric
+[here](https://github.com/Mcompetitions/M5-methods/blob/master/M5-Competitors-Guide.pdf).
+
+
+```python
+Y_hat = ets_forecasts.set_index(['unique_id', 'ds']).unstack()
+Y_hat = Y_hat.droplevel(0, 1).reset_index()
+```
+
+
+```python
+*_, S_df = M5.load('./data')
+Y_hat = S_df.merge(Y_hat, how='left', on=['unique_id'])#.drop(columns=['unique_id'])
+```
+
+
+```python
+wrmsse_ets = M5Evaluation.evaluate(y_hat=Y_hat, directory='./data')
+```
+
+
+```python
+wrmsse_ets
+```
+
+| | wrmsse |
+|---------|----------|
+| Total | 0.682358 |
+| Level1 | 0.449115 |
+| Level2 | 0.533754 |
+| Level3 | 0.592317 |
+| Level4 | 0.497086 |
+| Level5 | 0.572189 |
+| Level6 | 0.593880 |
+| Level7 | 0.665358 |
+| Level8 | 0.652183 |
+| Level9 | 0.734492 |
+| Level10 | 1.012633 |
+| Level11 | 0.969902 |
+| Level12 | 0.915380 |
+
+## Prophet pipeline
+
+
+```python
+import logging
+from time import time
+
+import pandas as pd
+from neuralforecast.data.datasets.m5 import M5, M5Evaluation
+from prophet import Prophet
+from pyspark.sql.types import *
+
+# disable informational messages from prophet
+logging.getLogger('py4j').setLevel(logging.ERROR)
+```
+
+### Download data
+
+
+```python
+# structure of the training data set
+train_schema = StructType([
+ StructField('unique_id', StringType()),
+ StructField('ds', DateType()),
+ StructField('y', DoubleType())
+ ])
+
+# read the training file into a dataframe
+train = spark.read.parquet(
+ 's3://m5-benchmarks/data/train/m5-target.parquet',
+ header=True,
+ schema=train_schema
+ )
+
+# make the dataframe queriable as a temporary view
+train.createOrReplaceTempView('train')
+```
+
+
+```python
+sql_statement = '''
+ SELECT
+ unique_id AS unique_id,
+ CAST(ds as date) as ds,
+ y as y
+ FROM train
+ '''
+
+m5_history = (
+ spark
+ .sql( sql_statement )
+ .repartition(sc.defaultParallelism, ['unique_id'])
+ ).cache()
+```
+
+### Forecast function using Prophet
+
+
+```python
+def forecast( history_pd: pd.DataFrame ) -> pd.DataFrame:
+
+ # TRAIN MODEL AS BEFORE
+ # --------------------------------------
+ # remove missing values (more likely at day-store-item level)
+ history_pd = history_pd.dropna()
+
+ # configure the model
+ model = Prophet(
+ growth='linear',
+ daily_seasonality=False,
+ weekly_seasonality=True,
+ yearly_seasonality=True,
+ seasonality_mode='multiplicative'
+ )
+
+ # train the model
+ model.fit( history_pd )
+ # --------------------------------------
+
+ # BUILD FORECAST AS BEFORE
+ # --------------------------------------
+ # make predictions
+ future_pd = model.make_future_dataframe(
+ periods=28,
+ freq='d',
+ include_history=False
+ )
+ forecast_pd = model.predict( future_pd )
+ # --------------------------------------
+
+ # ASSEMBLE EXPECTED RESULT SET
+ # --------------------------------------
+ # get relevant fields from forecast
+ forecast_pd['unique_id'] = history_pd['unique_id'].unique()[0]
+ f_pd = forecast_pd[['unique_id', 'ds','yhat']]
+ # --------------------------------------
+
+ # return expected dataset
+ return f_pd
+```
+
+
+```python
+result_schema = StructType([
+ StructField('unique_id', StringType()),
+ StructField('ds',DateType()),
+ StructField('yhat',FloatType()),
+])
+```
+
+#### Training Prophet on the M5 dataset
+
+
+```python
+init = time()
+results = (
+ m5_history
+ .groupBy('unique_id')
+ .applyInPandas(forecast, schema=result_schema)
+ ).toPandas()
+end = time()
+print(f'Minutes taken by Prophet on a Spark cluster: {(end - init) / 60}')
+```
+
+### Evaluating performance
+
+The M5 competition used the weighted root mean squared scaled error. You
+can find details of the metric
+[here](https://github.com/Mcompetitions/M5-methods/blob/master/M5-Competitors-Guide.pdf).
+
+
+```python
+Y_hat = results.set_index(['unique_id', 'ds']).unstack()
+Y_hat = Y_hat.droplevel(0, 1).reset_index()
+```
+
+
+```python
+*_, S_df = M5.load('./data')
+Y_hat = S_df.merge(Y_hat, how='left', on=['unique_id'])#.drop(columns=['unique_id'])
+```
+
+
+```python
+wrmsse = M5Evaluation.evaluate(y_hat=Y_hat, directory='./data')
+```
+
+
+```python
+wrmsse
+```
+
+| | wrmsse |
+|---------|----------|
+| Total | 0.771800 |
+| Level1 | 0.507905 |
+| Level2 | 0.586328 |
+| Level3 | 0.666686 |
+| Level4 | 0.549358 |
+| Level5 | 0.655003 |
+| Level6 | 0.647176 |
+| Level7 | 0.747047 |
+| Level8 | 0.743422 |
+| Level9 | 0.824667 |
+| Level10 | 1.207069 |
+| Level11 | 1.108780 |
+| Level12 | 1.018163 |
+
diff --git a/statsforecast/docs/getting-started/1_Getting_Started_short_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/getting-started/1_Getting_Started_short_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..7c6ac7b5
Binary files /dev/null and b/statsforecast/docs/getting-started/1_Getting_Started_short_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/getting-started/2_Getting_Started_complete_files/figure-markdown_strict/cell-11-output-1.png b/statsforecast/docs/getting-started/2_Getting_Started_complete_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..5de3c287
Binary files /dev/null and b/statsforecast/docs/getting-started/2_Getting_Started_complete_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/statsforecast/docs/getting-started/2_Getting_Started_complete_files/figure-markdown_strict/cell-12-output-1.png b/statsforecast/docs/getting-started/2_Getting_Started_complete_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..c1b32c73
Binary files /dev/null and b/statsforecast/docs/getting-started/2_Getting_Started_complete_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/statsforecast/docs/getting-started/2_Getting_Started_complete_files/figure-markdown_strict/cell-13-output-1.png b/statsforecast/docs/getting-started/2_Getting_Started_complete_files/figure-markdown_strict/cell-13-output-1.png
new file mode 100644
index 00000000..2110d9e0
Binary files /dev/null and b/statsforecast/docs/getting-started/2_Getting_Started_complete_files/figure-markdown_strict/cell-13-output-1.png differ
diff --git a/statsforecast/docs/getting-started/2_Getting_Started_complete_files/figure-markdown_strict/cell-20-output-1.png b/statsforecast/docs/getting-started/2_Getting_Started_complete_files/figure-markdown_strict/cell-20-output-1.png
new file mode 100644
index 00000000..4769261f
Binary files /dev/null and b/statsforecast/docs/getting-started/2_Getting_Started_complete_files/figure-markdown_strict/cell-20-output-1.png differ
diff --git a/statsforecast/docs/getting-started/2_Getting_Started_complete_files/figure-markdown_strict/cell-23-output-1.png b/statsforecast/docs/getting-started/2_Getting_Started_complete_files/figure-markdown_strict/cell-23-output-1.png
new file mode 100644
index 00000000..44c6eb50
Binary files /dev/null and b/statsforecast/docs/getting-started/2_Getting_Started_complete_files/figure-markdown_strict/cell-23-output-1.png differ
diff --git a/statsforecast/docs/getting-started/2_Getting_Started_complete_files/figure-markdown_strict/cell-6-output-1.png b/statsforecast/docs/getting-started/2_Getting_Started_complete_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..6f4122ee
Binary files /dev/null and b/statsforecast/docs/getting-started/2_Getting_Started_complete_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/statsforecast/docs/getting-started/3_Getting_Started_complete_polars_files/figure-markdown_strict/cell-11-output-1.png b/statsforecast/docs/getting-started/3_Getting_Started_complete_polars_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..88bfd17a
Binary files /dev/null and b/statsforecast/docs/getting-started/3_Getting_Started_complete_polars_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/statsforecast/docs/getting-started/3_Getting_Started_complete_polars_files/figure-markdown_strict/cell-12-output-1.png b/statsforecast/docs/getting-started/3_Getting_Started_complete_polars_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..c1b32c73
Binary files /dev/null and b/statsforecast/docs/getting-started/3_Getting_Started_complete_polars_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/statsforecast/docs/getting-started/3_Getting_Started_complete_polars_files/figure-markdown_strict/cell-13-output-1.png b/statsforecast/docs/getting-started/3_Getting_Started_complete_polars_files/figure-markdown_strict/cell-13-output-1.png
new file mode 100644
index 00000000..2110d9e0
Binary files /dev/null and b/statsforecast/docs/getting-started/3_Getting_Started_complete_polars_files/figure-markdown_strict/cell-13-output-1.png differ
diff --git a/statsforecast/docs/getting-started/3_Getting_Started_complete_polars_files/figure-markdown_strict/cell-20-output-1.png b/statsforecast/docs/getting-started/3_Getting_Started_complete_polars_files/figure-markdown_strict/cell-20-output-1.png
new file mode 100644
index 00000000..4769261f
Binary files /dev/null and b/statsforecast/docs/getting-started/3_Getting_Started_complete_polars_files/figure-markdown_strict/cell-20-output-1.png differ
diff --git a/statsforecast/docs/getting-started/3_Getting_Started_complete_polars_files/figure-markdown_strict/cell-23-output-1.png b/statsforecast/docs/getting-started/3_Getting_Started_complete_polars_files/figure-markdown_strict/cell-23-output-1.png
new file mode 100644
index 00000000..1f8effba
Binary files /dev/null and b/statsforecast/docs/getting-started/3_Getting_Started_complete_polars_files/figure-markdown_strict/cell-23-output-1.png differ
diff --git a/statsforecast/docs/getting-started/3_Getting_Started_complete_polars_files/figure-markdown_strict/cell-6-output-1.png b/statsforecast/docs/getting-started/3_Getting_Started_complete_polars_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..9e0c0b2e
Binary files /dev/null and b/statsforecast/docs/getting-started/3_Getting_Started_complete_polars_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/statsforecast/docs/getting-started/getting_started_complete.html.mdx b/statsforecast/docs/getting-started/getting_started_complete.html.mdx
new file mode 100644
index 00000000..d2a98111
--- /dev/null
+++ b/statsforecast/docs/getting-started/getting_started_complete.html.mdx
@@ -0,0 +1,556 @@
+---
+description: Model training, evaluation and selection for multiple time series
+output-file: getting_started_complete.html
+title: End to End Walkthrough
+---
+
+
+> **Prerequesites**
+>
+> This Guide assumes basic familiarity with StatsForecast. For a minimal
+> example visit the [Quick Start](./getting_started_short.html).
+
+Follow this article for a step to step guide on building a
+production-ready forecasting pipeline for multiple time series.
+
+During this guide you will gain familiary with the core
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)class
+and some relevant methods like `StatsForecast.plot`,
+[`StatsForecast.forecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast.forecast)
+and `StatsForecast.cross_validation.`
+
+We will use a classical benchmarking dataset from the M4 competition.
+The dataset includes time series from different domains like finance,
+economy and sales. In this example, we will use a subset of the Hourly
+dataset.
+
+We will model each time series individually. Forecasting at this level
+is also known as local forecasting. Therefore, you will train a series
+of models for every unique series and then select the best one.
+StatsForecast focuses on speed, simplicity, and scalability, which makes
+it ideal for this task.
+
+**Outline:**
+
+1. Install packages.
+2. Read the data.
+3. Explore the data.
+4. Train many models for every unique combination of time series.
+5. Evaluate the model’s performance using cross-validation.
+6. Select the best model for every unique time series.
+
+> **Not Covered in this guide**
+>
+> - Forecasting at scale using clusters on the cloud.
+> - [Forecast the M5 Dataset in
+> 5min](../how-to-guides/ets_ray_m5.html) using Ray clusters.
+> - [Forecast the M5 Dataset in
+> 5min](../how-to-guides/prophet_spark_m5.html) using Spark
+> clusters.
+> - Learn how to predict [1M series in less than
+> 30min](https://www.anyscale.com/blog/how-nixtla-uses-ray-to-accurately-predict-more-than-a-million-time-series).
+> - Training models on Multiple Seasonalities.
+> - Learn to use multiple seasonality in this [Electricity Load
+> forecasting](../tutorials/electricityloadforecasting.html)
+> tutorial.
+> - Using external regressors or exogenous variables
+> - Follow this tutorial to [include exogenous
+> variables](../how-to-guides/exogenous.html) like weather or
+> holidays or static variables like category or family.
+> - Comparing StatsForecast with other popular libraries.
+> - You can reproduce our benchmarks
+> [here](https://github.com/Nixtla/statsforecast/tree/main/experiments).
+
+## Install libraries
+
+We assume you have StatsForecast already installed. Check this guide for
+instructions on [how to install StatsForecast](./installation.html).
+
+## Read the data
+
+We will use pandas to read the M4 Hourly data set stored in a parquet
+file for efficiency. You can use ordinary pandas operations to read your
+data in other formats likes `.csv`.
+
+The input to StatsForecast is always a data frame in [long
+format](https://www.theanalysisfactor.com/wide-and-long-data/) with
+three columns: `unique_id`, `ds` and `y`:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp or int) column should be either an integer
+ indexing time or a datestampe ideally like YYYY-MM-DD for a date or
+ YYYY-MM-DD HH:MM:SS for a timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+ The target column needs to be renamed to `y` if it has a different
+ column name.
+
+This data set already satisfies the requirements.
+
+Depending on your internet connection, this step should take around 10
+seconds.
+
+
+```python
+import pandas as pd
+```
+
+
+```python
+Y_df = pd.read_parquet('https://datasets-nixtla.s3.amazonaws.com/m4-hourly.parquet')
+Y_df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|-----|-------|
+| 0 | H1 | 1 | 605.0 |
+| 1 | H1 | 2 | 586.0 |
+| 2 | H1 | 3 | 586.0 |
+| 3 | H1 | 4 | 559.0 |
+| 4 | H1 | 5 | 511.0 |
+
+This dataset contains 414 unique series with 900 observations on
+average. For this example and reproducibility’s sake, we will select
+only 10 unique IDs and keep only the last week. Depending on your
+processing infrastructure feel free to select more or less series.
+
+> **Note**
+>
+> Processing time is dependent on the available computing resources.
+> Running this example with the complete dataset takes around 10 minutes
+> in a c5d.24xlarge (96 cores) instance from AWS.
+
+
+```python
+uids = Y_df['unique_id'].unique()[:10] # Select 10 ids to make the example faster
+Y_df = Y_df.query('unique_id in @uids')
+Y_df = Y_df.groupby('unique_id').tail(7 * 24) #Select last 7 days of data to make example faster
+```
+
+## Explore Data with the plot method
+
+Plot some series using the `plot` method from the
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+class. This method prints 8 random series from the dataset and is useful
+for basic EDA.
+
+> **Note**
+>
+> The `StatsForecast.plot` method uses Plotly as a defaul engine. You
+> can change to MatPlotLib by setting `engine="matplotlib"`.
+
+
+```python
+from statsforecast import StatsForecast
+```
+
+
+```python
+StatsForecast.plot(Y_df)
+```
+
+
+
+## Train multiple models for many series
+
+StatsForecast can train many models on many time series efficiently.
+
+Start by importing and instantiating the desired models. StatsForecast
+offers a wide variety of models grouped in the following categories:
+
+- **Auto Forecast:** Automatic forecasting tools search for the best
+ parameters and select the best possible model for a series of time
+ series. These tools are useful for large collections of univariate
+ time series. Includes automatic versions of: Arima, ETS, Theta, CES.
+
+- **Exponential Smoothing:** Uses a weighted average of all past
+ observations where the weights decrease exponentially into the past.
+ Suitable for data with no clear trend or seasonality. Examples: SES,
+ Holt’s Winters, SSO.
+
+- **Benchmark models:** classical models for establishing baselines.
+ Examples: Mean, Naive, Random Walk
+
+- **Intermittent or Sparse models:** suited for series with very few
+ non-zero observations. Examples: CROSTON, ADIDA, IMAPA
+
+- **Multiple Seasonalities:** suited for signals with more than one
+ clear seasonality. Useful for low-frequency data like electricity
+ and logs. Examples: MSTL.
+
+- **Theta Models:** fit two theta lines to a deseasonalized time
+ series, using different techniques to obtain and combine the two
+ theta lines to produce the final forecasts. Examples: Theta,
+ DynamicTheta
+
+Here you can check the complete list of
+[models](../../src/core/models.html) .
+
+For this example we will use:
+
+- [`AutoARIMA`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoarima):
+ Automatically selects the best ARIMA (AutoRegressive Integrated
+ Moving Average) model using an information criterion. Ref:
+ [`AutoARIMA`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoarima).
+
+- [`HoltWinters`](https://Nixtla.github.io/statsforecast/src/core/models.html#holtwinters):
+ triple exponential smoothing, Holt-Winters’ method is an extension
+ of exponential smoothing for series that contain both trend and
+ seasonality. Ref:
+ [`HoltWinters`](https://Nixtla.github.io/statsforecast/src/core/models.html#holtwinters)
+
+- [`SeasonalNaive`](https://Nixtla.github.io/statsforecast/src/core/models.html#seasonalnaive):
+ Memory Efficient Seasonal Naive predictions. Ref:
+ [`SeasonalNaive`](https://Nixtla.github.io/statsforecast/src/core/models.html#seasonalnaive)
+
+- [`HistoricAverage`](https://Nixtla.github.io/statsforecast/src/core/models.html#historicaverage):
+ arthimetic mean. Ref:
+ [`HistoricAverage`](https://Nixtla.github.io/statsforecast/src/core/models.html#historicaverage).
+
+- [`DynamicOptimizedTheta`](https://Nixtla.github.io/statsforecast/src/core/models.html#dynamicoptimizedtheta):
+ The theta family of models has been shown to perform well in various
+ datasets such as M3. Models the deseasonalized time series. Ref:
+ [`DynamicOptimizedTheta`](https://Nixtla.github.io/statsforecast/src/core/models.html#dynamicoptimizedtheta).
+
+Import and instantiate the models. Setting the `season_length` argument
+is sometimes tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/)) by the
+master, Rob Hyndmann, can be useful.
+
+
+```python
+from statsforecast.models import (
+ HoltWinters,
+ CrostonClassic as Croston,
+ HistoricAverage,
+ DynamicOptimizedTheta as DOT,
+ SeasonalNaive
+)
+```
+
+
+```python
+# Create a list of models and instantiation parameters
+models = [
+ HoltWinters(),
+ Croston(),
+ SeasonalNaive(season_length=24),
+ HistoricAverage(),
+ DOT(season_length=24)
+]
+```
+
+We fit the models by instantiating a new
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+object with the following parameters:
+
+- `models`: a list of models. Select the models you want from
+ [models](../../src/core/models.html) and import them.
+
+- `freq`: a string indicating the frequency of the data. (See [pandas
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs`: n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model`: a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+
+```python
+# Instantiate StatsForecast class as sf
+sf = StatsForecast(
+ models=models,
+ freq=1,
+ fallback_model = SeasonalNaive(season_length=7),
+ n_jobs=-1,
+)
+```
+
+> **Note**
+>
+> StatsForecast achieves its blazing speed using JIT compiling through
+> Numba. The first time you call the statsforecast class, the fit method
+> should take around 5 seconds. The second time -once Numba compiled
+> your settings- it should take less than 0.2s.
+
+The `forecast` method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h` (int): represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level` (list of floats): this optional parameter is used for
+ probabilistic forecasting. Set the `level` (or confidence
+ percentile) of your prediction interval. For example, `level=[90]`
+ means that the model expects the real value to be inside that
+ interval 90% of the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min. (If you want to speed things up to a couple of seconds,
+remove the AutoModels like ARIMA and Theta)
+
+> **Note**
+>
+> The `forecast` method is compatible with distributed clusters, so it
+> does not store any model parameters. If you want to store parameters
+> for every model you can use the `fit` and `predict` methods. However,
+> those methods are not defined for distrubed engines like Spark, Ray or
+> Dask.
+
+
+```python
+forecasts_df = sf.forecast(df=Y_df, h=48, level=[90])
+forecasts_df.head()
+```
+
+| | unique_id | ds | HoltWinters | HoltWinters-lo-90 | HoltWinters-hi-90 | CrostonClassic | CrostonClassic-lo-90 | CrostonClassic-hi-90 | SeasonalNaive | SeasonalNaive-lo-90 | SeasonalNaive-hi-90 | HistoricAverage | HistoricAverage-lo-90 | HistoricAverage-hi-90 | DynamicOptimizedTheta | DynamicOptimizedTheta-lo-90 | DynamicOptimizedTheta-hi-90 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | H1 | 749 | 829.0 | 422.549268 | 1235.450732 | 829.0 | 422.549268 | 1235.450732 | 635.0 | 566.036734 | 703.963266 | 660.982143 | 398.037761 | 923.926524 | 592.701851 | 577.677280 | 611.652639 |
+| 1 | H1 | 750 | 807.0 | 400.549268 | 1213.450732 | 807.0 | 400.549268 | 1213.450732 | 572.0 | 503.036734 | 640.963266 | 660.982143 | 398.037761 | 923.926524 | 525.589116 | 505.449755 | 546.621805 |
+| 2 | H1 | 751 | 785.0 | 378.549268 | 1191.450732 | 785.0 | 378.549268 | 1191.450732 | 532.0 | 463.036734 | 600.963266 | 660.982143 | 398.037761 | 923.926524 | 489.251814 | 462.072871 | 512.424116 |
+| 3 | H1 | 752 | 756.0 | 349.549268 | 1162.450732 | 756.0 | 349.549268 | 1162.450732 | 493.0 | 424.036734 | 561.963266 | 660.982143 | 398.037761 | 923.926524 | 456.195032 | 430.554302 | 478.260963 |
+| 4 | H1 | 753 | 719.0 | 312.549268 | 1125.450732 | 719.0 | 312.549268 | 1125.450732 | 477.0 | 408.036734 | 545.963266 | 660.982143 | 398.037761 | 923.926524 | 436.290514 | 411.051232 | 461.815932 |
+
+Plot the results of 8 random series using the `StatsForecast.plot`
+method.
+
+
+```python
+sf.plot(Y_df,forecasts_df)
+```
+
+
+
+The `StatsForecast.plot` allows for further customization. For example,
+plot the results of the different models and unique ids.
+
+
+```python
+# Plot to unique_ids and some selected models
+sf.plot(Y_df, forecasts_df, models=["HoltWinters","DynamicOptimizedTheta"], unique_ids=["H10", "H105"], level=[90])
+```
+
+
+
+
+```python
+# Explore other models
+sf.plot(Y_df, forecasts_df, models=["SeasonalNaive"], unique_ids=["H10", "H105"], level=[90])
+```
+
+
+
+## Evaluate the model’s performance
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, **Cross Validation** is done by defining a
+sliding window across the historical data and predicting the period
+following it. This form of cross-validation allows us to arrive at a
+better estimation of our model’s predictive abilities across a wider
+range of temporal instances while also keeping the data in the training
+set contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 2 days (n_windows=2), forecasting every second day (step_size=48).
+Depending on your computer, this step should take around 1 min.
+
+> **Tip**
+>
+> Setting `n_windows=1` mirrors a traditional train-test split with our
+> historical data serving as the training set and the last 48 hours
+> serving as the testing set.
+
+The
+[`cross_validation`](https://Nixtla.github.io/statsforecast/src/mfles.html#cross_validation)
+method from the
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+class takes the following arguments.
+
+- `df`: training data frame
+
+- `h` (int): represents h steps into the future that are being
+ forecasted. In this case, 24 hours ahead.
+
+- `step_size` (int): step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows`(int): number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+
+```python
+cv_df = sf.cross_validation(
+ df=Y_df,
+ h=24,
+ step_size=24,
+ n_windows=2
+)
+```
+
+The `cv_df` object is a new data frame that includes the following
+columns:
+
+- `unique_id`: series identifier
+
+- `ds`: datestamp or temporal index
+
+- `cutoff`: the last datestamp or temporal index for the `n_windows.`
+ If `n_windows=1`, then one unique cuttoff value, if `n_windows=2`
+ then two unique cutoff values.
+
+- `y`: true value
+
+- `"model"`: columns with the model’s name and fitted value.
+
+
+```python
+cv_df.head()
+```
+
+| | unique_id | ds | cutoff | y | HoltWinters | CrostonClassic | SeasonalNaive | HistoricAverage | DynamicOptimizedTheta |
+|----|----|----|----|----|----|----|----|----|----|
+| 0 | H1 | 701 | 700 | 619.0 | 847.0 | 742.668748 | 691.0 | 661.675 | 612.767504 |
+| 1 | H1 | 702 | 700 | 565.0 | 820.0 | 742.668748 | 618.0 | 661.675 | 536.846278 |
+| 2 | H1 | 703 | 700 | 532.0 | 790.0 | 742.668748 | 563.0 | 661.675 | 497.824286 |
+| 3 | H1 | 704 | 700 | 495.0 | 784.0 | 742.668748 | 529.0 | 661.675 | 464.723219 |
+| 4 | H1 | 705 | 700 | 481.0 | 752.0 | 742.668748 | 504.0 | 661.675 | 440.972336 |
+
+Next, we will evaluate the performance of every model for every series
+using common error metrics like Mean Absolute Error (MAE) or Mean Square
+Error (MSE) Define a utility function to evaluate different error
+metrics for the cross validation data frame.
+
+First import the desired error metrics from `utilsforecast.losses`. Then
+define a utility function that takes a cross-validation data frame as a
+metric and returns an evaluation data frame with the average of the
+error metric for every unique id and fitted model and all cutoffs.
+
+
+```python
+from utilsforecast.losses import mse
+```
+
+
+```python
+def evaluate_cv(df, metric):
+ models = df.columns.drop(['unique_id', 'ds', 'y', 'cutoff']).tolist()
+ evals = metric(df, models=models)
+ evals['best_model'] = evals[models].idxmin(axis=1)
+ return evals
+```
+
+> **Warning**
+>
+> You can also use Mean Average Percentage Error (MAPE), however for
+> granular forecasts, MAPE values are extremely [hard to
+> judge](https://blog.blueyonder.com/mean-absolute-percentage-error-mape-has-served-its-duty-and-should-now-retire/)
+> and not useful to assess forecasting quality.
+
+Create the data frame with the results of the evaluation of your
+cross-validation data frame using a Mean Squared Error metric.
+
+
+```python
+evaluation_df = evaluate_cv(cv_df, mse)
+evaluation_df.head()
+```
+
+| | unique_id | HoltWinters | CrostonClassic | SeasonalNaive | HistoricAverage | DynamicOptimizedTheta | best_model |
+|----|----|----|----|----|----|----|----|
+| 0 | H1 | 44888.020833 | 28038.733985 | 1422.666667 | 20927.664488 | 1296.333977 | DynamicOptimizedTheta |
+| 1 | H10 | 2812.916667 | 1483.483839 | 96.895833 | 1980.367543 | 379.621134 | SeasonalNaive |
+| 2 | H100 | 121625.375000 | 91945.139237 | 12019.000000 | 78491.191439 | 21699.649325 | SeasonalNaive |
+| 3 | H101 | 28453.395833 | 16183.634340 | 10944.458333 | 18208.409800 | 63698.077266 | SeasonalNaive |
+| 4 | H102 | 232924.854167 | 132655.309136 | 12699.895833 | 309110.475212 | 31393.535274 | SeasonalNaive |
+
+Create a summary table with a model column and the number of series
+where that model performs best. In this case, the Arima and Seasonal
+Naive are the best models for 10 series and the Theta model should be
+used for two.
+
+
+```python
+evaluation_df['best_model'].value_counts().to_frame().reset_index()
+```
+
+| | best_model | count |
+|-----|-----------------------|-------|
+| 0 | SeasonalNaive | 6 |
+| 1 | DynamicOptimizedTheta | 4 |
+
+You can further explore your results by plotting the unique_ids where a
+specific model wins.
+
+
+```python
+seasonal_ids = evaluation_df.query('best_model == "SeasonalNaive"')['unique_id']
+sf.plot(Y_df,forecasts_df, unique_ids=seasonal_ids, models=["SeasonalNaive","DynamicOptimizedTheta"])
+```
+
+
+
+## Select the best model for every unique series
+
+Define a utility function that takes your forecast’s data frame with the
+predictions and the evaluation data frame and returns a data frame with
+the best possible forecast for every unique_id.
+
+
+```python
+def get_best_model_forecast(forecasts_df, evaluation_df):
+ with_best = forecasts_df.merge(evaluation_df[['unique_id', 'best_model']])
+ res = with_best[['unique_id', 'ds']].copy()
+ for suffix in ('', '-lo-90', '-hi-90'):
+ res[f'best_model{suffix}'] = with_best.apply(lambda row: row[row['best_model'] + suffix], axis=1)
+ return res
+```
+
+Create your production-ready data frame with the best forecast for every
+unique_id.
+
+
+```python
+prod_forecasts_df = get_best_model_forecast(forecasts_df, evaluation_df)
+prod_forecasts_df.head()
+```
+
+| | unique_id | ds | best_model | best_model-lo-90 | best_model-hi-90 |
+|-----|-----------|-----|------------|------------------|------------------|
+| 0 | H1 | 749 | 592.701851 | 577.677280 | 611.652639 |
+| 1 | H1 | 750 | 525.589116 | 505.449755 | 546.621805 |
+| 2 | H1 | 751 | 489.251814 | 462.072871 | 512.424116 |
+| 3 | H1 | 752 | 456.195032 | 430.554302 | 478.260963 |
+| 4 | H1 | 753 | 436.290514 | 411.051232 | 461.815932 |
+
+Plot the results.
+
+
+```python
+sf.plot(Y_df, prod_forecasts_df, level=[90])
+```
+
+
+
diff --git a/statsforecast/docs/getting-started/getting_started_complete_polars.html.mdx b/statsforecast/docs/getting-started/getting_started_complete_polars.html.mdx
new file mode 100644
index 00000000..ebc4b7a8
--- /dev/null
+++ b/statsforecast/docs/getting-started/getting_started_complete_polars.html.mdx
@@ -0,0 +1,612 @@
+---
+description: Model training, evaluation and selection for multiple time series
+output-file: getting_started_complete_polars.html
+title: End to End Walkthrough with Polars
+---
+
+
+## Introducing Polars: A High-Performance DataFrame Library
+
+This document aims to highlight the recent integration of Polars, a
+robust and high-speed DataFrame library developed in Rust, into the
+functionality of StatsForecast. Polars, with its nimble and potent
+capabilities, has rapidly established a strong reputation within the
+Data Science community, further solidifying its position as a reliable
+tool for managing and manipulating substantial data sets.
+
+Available in languages including Rust, Python, Node.js, and R, Polars
+demonstrates a remarkable ability to handle sizable data sets with
+efficiency and speed that surpasses many other DataFrame libraries, such
+as Pandas. Polars’ open-source nature invites ongoing enhancements and
+contributions, augmenting its appeal within the data science arena.
+
+The most significant features of Polars that contribute to its rapid
+adoption are:
+
+1. **Performance Efficiency**: Constructed using Rust, Polars exhibits
+ an exemplary ability to manage substantial datasets with remarkable
+ speed and minimal memory usage.
+
+2. **Lazy Evaluation**: Polars operates on the principle of ‘lazy
+ evaluation’, creating an optimized logical plan of operations for
+ efficient execution, a feature that mirrors the functionality of
+ Apache Spark.
+
+3. **Parallel Execution**: Demonstrating the capability to exploit
+ multi-core CPUs, Polars facilitates parallel execution of
+ operations, substantially accelerating data processing tasks.
+
+> **Prerequesites**
+>
+> This Guide assumes basic familiarity with StatsForecast. For a minimal
+> example visit the [Quick Start](./getting_started_short.html)
+
+Follow this article for a step to step guide on building a
+production-ready forecasting pipeline for multiple time series.
+
+During this guide you will gain familiary with the core
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)class
+and some relevant methods like `StatsForecast.plot`,
+[`StatsForecast.forecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast.forecast)
+and `StatsForecast.cross_validation.`
+
+We will use a classical benchmarking dataset from the M4 competition.
+The dataset includes time series from different domains like finance,
+economy and sales. In this example, we will use a subset of the Hourly
+dataset.
+
+We will model each time series individually. Forecasting at this level
+is also known as local forecasting. Therefore, you will train a series
+of models for every unique series and then select the best one.
+StatsForecast focuses on speed, simplicity, and scalability, which makes
+it ideal for this task.
+
+**Outline:**
+
+1. Install packages.
+2. Read the data.
+3. Explore the data.
+4. Train many models for every unique combination of time series.
+5. Evaluate the model’s performance using cross-validation.
+6. Select the best model for every unique time series.
+
+> **Not Covered in this guide**
+>
+> - Forecasting at scale using clusters on the cloud.
+> - [Forecast the M5 Dataset in
+> 5min](../how-to-guides/ets_ray_m5.html) using Ray clusters.
+> - [Forecast the M5 Dataset in
+> 5min](../how-to-guides/prophet_spark_m5.html) using Spark
+> clusters.
+> - Learn how to predict [1M series in less than
+> 30min](https://www.anyscale.com/blog/how-nixtla-uses-ray-to-accurately-predict-more-than-a-million-time-series).
+> - Training models on Multiple Seasonalities.
+> - Learn to use multiple seasonality in this [Electricity Load
+> forecasting](../tutorials/electricityloadforecasting.html)
+> tutorial.
+> - Using external regressors or exogenous variables
+> - Follow this tutorial to [include exogenous
+> variables](../how-to-guides/exogenous.html) like weather or
+> holidays or static variables like category or family.
+> - Comparing StatsForecast with other popular libraries.
+> - You can reproduce our benchmarks
+> [here](https://github.com/Nixtla/statsforecast/tree/main/experiments).
+
+## Install libraries
+
+We assume you have StatsForecast already installed. Check this guide for
+instructions on [how to install StatsForecast](./installation.html).
+
+## Read the data
+
+We will use polars to read the M4 Hourly data set stored in a parquet
+file for efficiency. You can use ordinary polars operations to read your
+data in other formats likes `.csv`.
+
+The input to StatsForecast is always a data frame in [long
+format](https://www.theanalysisfactor.com/wide-and-long-data/) with
+three columns: `unique_id`, `ds` and `y`:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp or int) column should be either an integer
+ indexing time or a datestampe ideally like YYYY-MM-DD for a date or
+ YYYY-MM-DD HH:MM:SS for a timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+This data set already satisfies the requirement.
+
+Depending on your internet connection, this step should take around 10
+seconds.
+
+
+```python
+import polars as pl
+```
+
+
+```python
+Y_df = pl.read_parquet('https://datasets-nixtla.s3.amazonaws.com/m4-hourly.parquet')
+Y_df.head()
+```
+
+| unique_id | ds | y |
+|-----------|-----|-------|
+| str | i64 | f64 |
+| "H1" | 1 | 605.0 |
+| "H1" | 2 | 586.0 |
+| "H1" | 3 | 586.0 |
+| "H1" | 4 | 559.0 |
+| "H1" | 5 | 511.0 |
+
+This dataset contains 414 unique series with 900 observations on
+average. For this example and reproducibility’s sake, we will select
+only 10 unique IDs and keep only the last week. Depending on your
+processing infrastructure feel free to select more or less series.
+
+> **Note**
+>
+> Processing time is dependent on the available computing resources.
+> Running this example with the complete dataset takes around 10 minutes
+> in a c5d.24xlarge (96 cores) instance from AWS.
+
+
+```python
+uids = Y_df['unique_id'].unique(maintain_order=True)[:10] # Select 10 ids to make the example faster
+Y_df = Y_df.filter(pl.col('unique_id').is_in(uids))
+Y_df = Y_df.group_by('unique_id').tail(7 * 24) #Select last 7 days of data to make example faster
+```
+
+## Explore Data with the plot method
+
+Plot some series using the `plot` method from the
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+class. This method prints 8 random series from the dataset and is useful
+for basic EDA.
+
+> **Note**
+>
+> The `StatsForecast.plot` method uses matplotlib as a default engine.
+> You can change to plotly by setting `engine="plotly"`.
+
+
+```python
+from statsforecast import StatsForecast
+```
+
+
+```python
+StatsForecast.plot(Y_df)
+```
+
+
+
+## Train multiple models for many series
+
+StatsForecast can train many models on many time series efficiently.
+
+Start by importing and instantiating the desired models. StatsForecast
+offers a wide variety of models grouped in the following categories:
+
+- **Auto Forecast:** Automatic forecasting tools search for the best
+ parameters and select the best possible model for a series of time
+ series. These tools are useful for large collections of univariate
+ time series. Includes automatic versions of: Arima, ETS, Theta, CES.
+
+- **Exponential Smoothing:** Uses a weighted average of all past
+ observations where the weights decrease exponentially into the past.
+ Suitable for data with no clear trend or seasonality. Examples: SES,
+ Holt’s Winters, SSO.
+
+- **Benchmark models:** classical models for establishing baselines.
+ Examples: Mean, Naive, Random Walk
+
+- **Intermittent or Sparse models:** suited for series with very few
+ non-zero observations. Examples: CROSTON, ADIDA, IMAPA
+
+- **Multiple Seasonalities:** suited for signals with more than one
+ clear seasonality. Useful for low-frequency data like electricity
+ and logs. Examples: MSTL.
+
+- **Theta Models:** fit two theta lines to a deseasonalized time
+ series, using different techniques to obtain and combine the two
+ theta lines to produce the final forecasts. Examples: Theta,
+ DynamicTheta
+
+Here you can check the complete list of [models](../models_intro.qmd).
+
+For this example we will use:
+
+- [`AutoARIMA`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoarima):
+ Automatically selects the best ARIMA (AutoRegressive Integrated
+ Moving Average) model using an information criterion. Ref:
+ [`AutoARIMA`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoarima).
+
+- [`HoltWinters`](https://Nixtla.github.io/statsforecast/src/core/models.html#holtwinters):
+ triple exponential smoothing, Holt-Winters’ method is an extension
+ of exponential smoothing for series that contain both trend and
+ seasonality. Ref:
+ [`HoltWinters`](https://Nixtla.github.io/statsforecast/src/core/models.html#holtwinters)
+
+- [`SeasonalNaive`](https://Nixtla.github.io/statsforecast/src/core/models.html#seasonalnaive):
+ Memory Efficient Seasonal Naive predictions. Ref:
+ [`SeasonalNaive`](https://Nixtla.github.io/statsforecast/src/core/models.html#seasonalnaive)
+
+- [`HistoricAverage`](https://Nixtla.github.io/statsforecast/src/core/models.html#historicaverage):
+ arthimetic mean. Ref:
+ [`HistoricAverage`](https://Nixtla.github.io/statsforecast/src/core/models.html#historicaverage).
+
+- [`DynamicOptimizedTheta`](https://Nixtla.github.io/statsforecast/src/core/models.html#dynamicoptimizedtheta):
+ The theta family of models has been shown to perform well in various
+ datasets such as M3. Models the deseasonalized time series. Ref:
+ [`DynamicOptimizedTheta`](https://Nixtla.github.io/statsforecast/src/core/models.html#dynamicoptimizedtheta).
+
+Import and instantiate the models. Setting the `season_length` argument
+is sometimes tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/)) by the
+master, Rob Hyndmann, can be useful.
+
+
+```python
+from statsforecast.models import (
+ HoltWinters,
+ CrostonClassic as Croston,
+ HistoricAverage,
+ DynamicOptimizedTheta as DOT,
+ SeasonalNaive
+)
+```
+
+
+```python
+# Create a list of models and instantiation parameters
+models = [
+ HoltWinters(),
+ Croston(),
+ SeasonalNaive(season_length=24),
+ HistoricAverage(),
+ DOT(season_length=24)
+]
+```
+
+We fit the models by instantiating a new
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+object with the following parameters:
+
+- `models`: a list of models. Select the models you want from
+ [models](../models.html) and import them.
+
+- `freq`: a string indicating the frequency of the data. (See [panda’s
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+ This is also available with Polars.
+
+- `n_jobs`: n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model`: a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+
+```python
+# Instantiate StatsForecast class as sf
+sf = StatsForecast(
+ models=models,
+ freq=1,
+ n_jobs=-1,
+ fallback_model=SeasonalNaive(season_length=7),
+ verbose=True,
+)
+```
+
+> **Note**
+>
+> StatsForecast achieves its blazing speed using JIT compiling through
+> Numba. The first time you call the statsforecast class, the fit method
+> should take around 5 seconds. The second time -once Numba compiled
+> your settings- it should take less than 0.2s.
+
+The `forecast` method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h` (int): represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level` (list of floats): this optional parameter is used for
+ probabilistic forecasting. Set the `level` (or confidence
+ percentile) of your prediction interval. For example, `level=[90]`
+ means that the model expects the real value to be inside that
+ interval 90% of the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min. (If you want to speed things up to a couple of seconds,
+remove the AutoModels like ARIMA and Theta)
+
+> **Note**
+>
+> The `forecast` method is compatible with distributed clusters, so it
+> does not store any model parameters. If you want to store parameters
+> for every model you can use the `fit` and `predict` methods. However,
+> those methods are not defined for distrubed engines like Spark, Ray or
+> Dask.
+
+
+```python
+forecasts_df = sf.forecast(df=Y_df, h=48, level=[90])
+forecasts_df.head()
+```
+
+``` text
+Forecast: 0%| | 0/10 [Elapsed: 00:00]
+```
+
+| unique_id | ds | HoltWinters | HoltWinters-lo-90 | HoltWinters-hi-90 | CrostonClassic | CrostonClassic-lo-90 | CrostonClassic-hi-90 | SeasonalNaive | SeasonalNaive-lo-90 | SeasonalNaive-hi-90 | HistoricAverage | HistoricAverage-lo-90 | HistoricAverage-hi-90 | DynamicOptimizedTheta | DynamicOptimizedTheta-lo-90 | DynamicOptimizedTheta-hi-90 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| str | i64 | f64 | f64 | f64 | f64 | f64 | f64 | f64 | f64 | f64 | f64 | f64 | f64 | f64 | f64 | f64 |
+| "H1" | 749 | 829.0 | 422.549268 | 1235.450732 | 829.0 | 422.549268 | 1235.450732 | 635.0 | 566.036734 | 703.963266 | 660.982143 | 398.037761 | 923.926524 | 592.701851 | 577.67728 | 611.652639 |
+| "H1" | 750 | 807.0 | 400.549268 | 1213.450732 | 807.0 | 400.549268 | 1213.450732 | 572.0 | 503.036734 | 640.963266 | 660.982143 | 398.037761 | 923.926524 | 525.589116 | 505.449755 | 546.621805 |
+| "H1" | 751 | 785.0 | 378.549268 | 1191.450732 | 785.0 | 378.549268 | 1191.450732 | 532.0 | 463.036734 | 600.963266 | 660.982143 | 398.037761 | 923.926524 | 489.251814 | 462.072871 | 512.424116 |
+| "H1" | 752 | 756.0 | 349.549268 | 1162.450732 | 756.0 | 349.549268 | 1162.450732 | 493.0 | 424.036734 | 561.963266 | 660.982143 | 398.037761 | 923.926524 | 456.195032 | 430.554302 | 478.260963 |
+| "H1" | 753 | 719.0 | 312.549268 | 1125.450732 | 719.0 | 312.549268 | 1125.450732 | 477.0 | 408.036734 | 545.963266 | 660.982143 | 398.037761 | 923.926524 | 436.290514 | 411.051232 | 461.815932 |
+
+Plot the results of 8 randon series using the `StatsForecast.plot`
+method.
+
+
+```python
+sf.plot(Y_df,forecasts_df)
+```
+
+
+
+The `StatsForecast.plot` allows for further customization. For example,
+plot the results of the different models and unique ids.
+
+
+```python
+# Plot to unique_ids and some selected models
+sf.plot(Y_df, forecasts_df, models=["HoltWinters","DynamicOptimizedTheta"], unique_ids=["H10", "H105"], level=[90])
+```
+
+
+
+
+```python
+# Explore other models
+sf.plot(Y_df, forecasts_df, models=["SeasonalNaive"], unique_ids=["H10", "H105"], level=[90])
+```
+
+
+
+## Evaluate the model’s performance
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, **Cross Validation** is done by defining a
+sliding window across the historical data and predicting the period
+following it. This form of cross-validation allows us to arrive at a
+better estimation of our model’s predictive abilities across a wider
+range of temporal instances while also keeping the data in the training
+set contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 2 days (n_windows=2), forecasting every second day (step_size=48).
+Depending on your computer, this step should take around 1 min.
+
+> **Tip**
+>
+> Setting `n_windows=1` mirrors a traditional train-test split with our
+> historical data serving as the training set and the last 48 hours
+> serving as the testing set.
+
+The
+[`cross_validation`](https://Nixtla.github.io/statsforecast/src/mfles.html#cross_validation)
+method from the
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+class takes the following arguments.
+
+- `df`: training data frame
+
+- `h` (int): represents h steps into the future that are being
+ forecasted. In this case, 24 hours ahead.
+
+- `step_size` (int): step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows`(int): number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+
+```python
+cv_df = sf.cross_validation(
+ df=Y_df,
+ h=24,
+ step_size=24,
+ n_windows=2
+)
+```
+
+The `cv_df` object is a new data frame that includes the following
+columns:
+
+- `unique_id`: series identifier
+
+- `ds`: datestamp or temporal index
+
+- `cutoff`: the last datestamp or temporal index for the `n_windows.`
+ If `n_windows=1`, then one unique cuttoff value, if `n_windows=2`
+ then two unique cutoff values.
+
+- `y`: true value
+
+- `"model"`: columns with the model’s name and fitted value.
+
+
+```python
+cv_df.head()
+```
+
+| unique_id | ds | cutoff | y | HoltWinters | CrostonClassic | SeasonalNaive | HistoricAverage | DynamicOptimizedTheta |
+|----|----|----|----|----|----|----|----|----|
+| str | i64 | i64 | f64 | f64 | f64 | f64 | f64 | f64 |
+| "H1" | 701 | 700 | 619.0 | 847.0 | 742.668748 | 691.0 | 661.675 | 612.767504 |
+| "H1" | 702 | 700 | 565.0 | 820.0 | 742.668748 | 618.0 | 661.675 | 536.846278 |
+| "H1" | 703 | 700 | 532.0 | 790.0 | 742.668748 | 563.0 | 661.675 | 497.824286 |
+| "H1" | 704 | 700 | 495.0 | 784.0 | 742.668748 | 529.0 | 661.675 | 464.723219 |
+| "H1" | 705 | 700 | 481.0 | 752.0 | 742.668748 | 504.0 | 661.675 | 440.972336 |
+
+Next, we will evaluate the performance of every model for every series
+using common error metrics like Mean Absolute Error (MAE) or Mean Square
+Error (MSE) Define a utility function to evaluate different error
+metrics for the cross validation data frame.
+
+First import the desired error metrics from `utilsforecast.losses`. Then
+define a utility function that takes a cross-validation data frame as a
+metric and returns an evaluation data frame with the average of the
+error metric for every unique id and fitted model and all cutoffs.
+
+
+```python
+from utilsforecast.losses import mse
+```
+
+
+```python
+def evaluate_cv(df, metric):
+ models = [c for c in df.columns if c not in ('unique_id', 'ds', 'cutoff', 'y')]
+ evals = mse(cv_df, models=models)
+ pos2model = dict(enumerate(models))
+ return evals.with_columns(
+ best_model=pl.concat_list(models).list.arg_min().replace_strict(pos2model)
+ )
+```
+
+> **Warning**
+>
+> You can also use Mean Average Percentage Error (MAPE), however for
+> granular forecasts, MAPE values are extremely [hard to
+> judge](https://blog.blueyonder.com/mean-absolute-percentage-error-mape-has-served-its-duty-and-should-now-retire/)
+> and not useful to assess forecasting quality.
+
+Create the data frame with the results of the evaluation of your
+cross-validation data frame using a Mean Squared Error metric.
+
+
+```python
+evaluation_df = evaluate_cv(cv_df, mse)
+evaluation_df.head()
+```
+
+| unique_id | HoltWinters | CrostonClassic | SeasonalNaive | HistoricAverage | DynamicOptimizedTheta | best_model |
+|----|----|----|----|----|----|----|
+| str | f64 | f64 | f64 | f64 | f64 | str |
+| "H1" | 44888.020833 | 28038.733985 | 1422.666667 | 20927.664488 | 1296.333977 | "DynamicOptimizedTheta" |
+| "H10" | 2812.916667 | 1483.483839 | 96.895833 | 1980.367543 | 379.621134 | "SeasonalNaive" |
+| "H100" | 121625.375 | 91945.139237 | 12019.0 | 78491.191439 | 21699.649325 | "SeasonalNaive" |
+| "H101" | 28453.395833 | 16183.63434 | 10944.458333 | 18208.4098 | 63698.077266 | "SeasonalNaive" |
+| "H102" | 232924.854167 | 132655.309136 | 12699.895833 | 309110.475212 | 31393.535274 | "SeasonalNaive" |
+
+Create a summary table with a model column and the number of series
+where that model performs best. In this case, the Arima and Seasonal
+Naive are the best models for 10 series and the Theta model should be
+used for two.
+
+
+```python
+evaluation_df['best_model'].value_counts()
+```
+
+| best_model | count |
+|-------------------------|-------|
+| str | u32 |
+| "DynamicOptimizedTheta" | 4 |
+| "SeasonalNaive" | 6 |
+
+You can further explore your results by plotting the unique_ids where a
+specific model wins.
+
+
+```python
+seasonal_ids = evaluation_df.filter(pl.col('best_model') == 'SeasonalNaive')['unique_id']
+sf.plot(Y_df,forecasts_df, unique_ids=seasonal_ids, models=["SeasonalNaive","DynamicOptimizedTheta"])
+```
+
+
+
+## Select the best model for every unique series
+
+Define a utility function that takes your forecast’s data frame with the
+predictions and the evaluation data frame and returns a data frame with
+the best possible forecast for every unique_id.
+
+
+```python
+def get_best_model_forecast(forecasts_df, evaluation_df):
+ models = {
+ c.replace('-lo-90', '').replace('-hi-90', '')
+ for c in forecasts_df.columns
+ if c not in ('unique_id', 'ds')
+ }
+ model2pos = {m: i for i, m in enumerate(models)}
+ with_best = forecasts_df.join(evaluation_df[['unique_id', 'best_model']], on='unique_id')
+ return with_best.select(
+ 'unique_id',
+ 'ds',
+ *[
+ (
+ pl.concat_list([f'{m}{suffix}' for m in models])
+ .list.get(pl.col('best_model').replace_strict(model2pos))
+ .alias(f'best_model{suffix}')
+ )
+ for suffix in ('', '-lo-90', '-hi-90')
+ ]
+ )
+```
+
+Create your production-ready data frame with the best forecast for every
+unique_id.
+
+
+```python
+prod_forecasts_df = get_best_model_forecast(forecasts_df, evaluation_df)
+prod_forecasts_df.head()
+```
+
+| unique_id | ds | best_model | best_model-lo-90 | best_model-hi-90 |
+|-----------|-----|------------|------------------|------------------|
+| str | i64 | f64 | f64 | f64 |
+| "H1" | 749 | 592.701851 | 577.67728 | 611.652639 |
+| "H1" | 750 | 525.589116 | 505.449755 | 546.621805 |
+| "H1" | 751 | 489.251814 | 462.072871 | 512.424116 |
+| "H1" | 752 | 456.195032 | 430.554302 | 478.260963 |
+| "H1" | 753 | 436.290514 | 411.051232 | 461.815932 |
+
+Plot the results.
+
+
+```python
+sf.plot(Y_df, prod_forecasts_df, level=[90])
+```
+
+
+
diff --git a/statsforecast/docs/getting-started/getting_started_short.html.mdx b/statsforecast/docs/getting-started/getting_started_short.html.mdx
new file mode 100644
index 00000000..eb494359
--- /dev/null
+++ b/statsforecast/docs/getting-started/getting_started_short.html.mdx
@@ -0,0 +1,162 @@
+---
+description: Minimal Example of StatsForecast
+output-file: getting_started_short.html
+title: Quick Start
+---
+
+
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+follows the sklearn model API. For this minimal example, you will create
+an instance of the StatsForecast class and then call its `fit` and
+`predict` methods. We recommend this option if speed is not paramount
+and you want to explore the fitted values and parameters.
+
+> **Tip**
+>
+> If you want to forecast many series, we recommend using the `forecast`
+> method. Check this [Getting Started with multiple time
+> series](./getting_started_complete.html) guide.
+
+The input to StatsForecast is always a data frame in [long
+format](https://www.theanalysisfactor.com/wide-and-long-data/) with
+three columns: `unique_id`, `ds` and `y`:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+As an example, let’s look at the US Air Passengers dataset. This time
+series consists of monthly totals of a US airline passengers from 1949
+to 1960. The CSV is available
+[here](https://www.kaggle.com/datasets/chirag19/air-passengers).
+
+We assume you have StatsForecast already installed. Check this guide for
+instructions on [how to install StatsForecast](./installation.html).
+
+First, we’ll import the data:
+
+
+```python
+# uncomment the following line to install the library
+# %pip install statsforecast
+```
+
+
+```python
+import pandas as pd
+```
+
+
+```python
+df = pd.read_csv('https://datasets-nixtla.s3.amazonaws.com/air-passengers.csv', parse_dates=['ds'])
+df.head()
+```
+
+| | unique_id | ds | y |
+|-----|---------------|------------|-----|
+| 0 | AirPassengers | 1949-01-01 | 112 |
+| 1 | AirPassengers | 1949-02-01 | 118 |
+| 2 | AirPassengers | 1949-03-01 | 132 |
+| 3 | AirPassengers | 1949-04-01 | 129 |
+| 4 | AirPassengers | 1949-05-01 | 121 |
+
+We fit the model by instantiating a new
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+object with its two required parameters:
+https://nixtla.github.io/statsforecast/src/core/models.html \* `models`:
+a list of models. Select the models you want from
+[models](../../src/core/models.html) and import them. For this example,
+we will use a
+[`AutoARIMA`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoarima)
+model. We set `season_length` to 12 because we expect seasonal effects
+every 12 months. (See: [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/))
+
+- `freq`: a string indicating the frequency of the data. (See [pandas
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+> **Note**
+>
+> StatsForecast achieves its blazing speed using JIT compiling through
+> Numba. The first time you call the statsforecast class, the fit method
+> should take around 5 seconds. The second time -once Numba compiled
+> your settings- it should take less than 0.2s.
+
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import AutoARIMA
+```
+
+
+```python
+sf = StatsForecast(
+ models=[AutoARIMA(season_length = 12)],
+ freq='MS',
+)
+sf.fit(df)
+```
+
+``` text
+StatsForecast(models=[AutoARIMA])
+```
+
+The `predict` method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h` (int): represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level` (list of floats): this optional parameter is used for
+ probabilistic forecasting. Set the `level` (or confidence
+ percentile) of your prediction interval. For example, `level=[90]`
+ means that the model expects the real value to be inside that
+ interval 90% of the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+
+```python
+forecast_df = sf.predict(h=12, level=[90])
+forecast_df.tail()
+```
+
+| | unique_id | ds | AutoARIMA | AutoARIMA-lo-90 | AutoARIMA-hi-90 |
+|-----|---------------|------------|------------|-----------------|-----------------|
+| 7 | AirPassengers | 1961-08-01 | 633.236389 | 590.009033 | 676.463745 |
+| 8 | AirPassengers | 1961-09-01 | 535.236389 | 489.558899 | 580.913940 |
+| 9 | AirPassengers | 1961-10-01 | 488.236389 | 440.233795 | 536.239014 |
+| 10 | AirPassengers | 1961-11-01 | 417.236389 | 367.016205 | 467.456604 |
+| 11 | AirPassengers | 1961-12-01 | 459.236389 | 406.892456 | 511.580322 |
+
+You can plot the forecast by calling the `StatsForecast.plot` method and
+passing in your forecast dataframe.
+
+
+```python
+sf.plot(df, forecast_df, level=[90])
+```
+
+
+
+> **Next Steps**
+>
+> - Build and end-to-end forecasting pipeline following best practices
+> in [End to End Walkthrough](./getting_started_complete.html)
+> - [Forecast millions of
+> series](../how-to-guides/prophet_spark_m5.html) in a scalable
+> cluster in the cloud using Spark and Nixtla
+> - [Detect anomalies](../tutorials/anomalydetection.html) in your
+> past observations
+
diff --git a/statsforecast/docs/getting-started/installation.html.mdx b/statsforecast/docs/getting-started/installation.html.mdx
new file mode 100644
index 00000000..113f8817
--- /dev/null
+++ b/statsforecast/docs/getting-started/installation.html.mdx
@@ -0,0 +1,58 @@
+---
+description: Install StatsForecast with pip or conda
+output-file: installation.html
+title: Install
+---
+
+
+You can install the *released version* of
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+from the [Python package index](https://pypi.org) with:
+
+
+```shell
+pip install statsforecast
+```
+
+or
+
+
+```shell
+conda install -c conda-forge statsforecast
+```
+
+> **Warning**
+>
+> We are constantly updating StatsForecast, so we suggest fixing the
+> version to avoid issues. `pip install statsforecast=="1.0.0"`
+
+> **Tip**
+>
+> We recommend installing your libraries inside a python virtual or
+> [conda
+> environment](https://docs.conda.io/projects/conda/en/latest/user-guide/install/macos.html).
+
+#### Extras
+
+The following features can also be installed by specifying the extra
+inside the install command,
+e.g. `pip install 'statsforecast[extra1,extra2]'`
+
+- **polars**: provide polars dataframes to StatsForecast.
+- **plotly**: use `StatsForecast.plot` with the plotly backend.
+- **dask**: perform distributed forecasting with dask.
+- **spark**: perform distributed forecasting with spark.
+- **ray**: perform distributed forecasting with ray.
+
+#### Development version
+
+If you want to try out a new feature that hasn’t made it into a release
+yet you have the following options:
+
+- Install from our nightly wheels:
+ `pip install --extra-index-url=http://nixtla-packages.s3-website.us-east-2.amazonaws.com --trusted-host nixtla-packages.s3-website.us-east-2.amazonaws.com statsforecast`
+- Install from github:
+ `pip install git+https://github.com/Nixtla/statsforecast`. This
+ requires that you have a C++ compiler installed, so we encourage you
+ to try the previous option first.
+
diff --git a/statsforecast/docs/how-to-guides/Exogenous_files/figure-markdown_strict/cell-17-output-1.png b/statsforecast/docs/how-to-guides/Exogenous_files/figure-markdown_strict/cell-17-output-1.png
new file mode 100644
index 00000000..9ddaaa9a
Binary files /dev/null and b/statsforecast/docs/how-to-guides/Exogenous_files/figure-markdown_strict/cell-17-output-1.png differ
diff --git a/statsforecast/docs/how-to-guides/Exogenous_files/figure-markdown_strict/cell-6-output-1.png b/statsforecast/docs/how-to-guides/Exogenous_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..65a7bc8c
Binary files /dev/null and b/statsforecast/docs/how-to-guides/Exogenous_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/statsforecast/docs/how-to-guides/Exogenous_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/how-to-guides/Exogenous_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..2bfa2bad
Binary files /dev/null and b/statsforecast/docs/how-to-guides/Exogenous_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/how-to-guides/automatic_forecasting.html.mdx b/statsforecast/docs/how-to-guides/automatic_forecasting.html.mdx
new file mode 100644
index 00000000..9ac9a3ef
--- /dev/null
+++ b/statsforecast/docs/how-to-guides/automatic_forecasting.html.mdx
@@ -0,0 +1,121 @@
+---
+description: >-
+ How to do automatic forecasting using `AutoARIMA`, `AutoETS`, `AutoCES` and
+ `AutoTheta`.
+output-file: automatic_forecasting.html
+title: Automatic Time Series Forecasting
+---
+
+
+> **Tip**
+>
+> Automatic forecasts of large numbers of univariate time series are
+> often needed. It is common to have multiple product lines or skus that
+> need forecasting. In these circumstances, an automatic forecasting
+> algorithm is an essential tool. Automatic forecasting algorithms must
+> determine an appropriate time series model, estimate the parameters
+> and compute the forecasts. They must be robust to unusual time series
+> patterns, and applicable to large numbers of series without user
+> intervention.
+
+## 1. Install statsforecast and load data
+
+Use pip to install statsforecast and load Air Passangers dataset as an
+example
+
+
+```python
+# uncomment the following line to install the library
+# %pip install statsforecast
+```
+
+
+```python
+from statsforecast.utils import AirPassengersDF
+```
+
+
+```python
+Y_df = AirPassengersDF
+```
+
+## 2. Import StatsForecast and models
+
+Import the core StatsForecast class and the models you want to use
+
+
+```python
+import pandas as pd
+
+from statsforecast import StatsForecast
+from statsforecast.models import AutoARIMA, AutoETS, AutoTheta, AutoCES
+```
+
+## 3. Instatiate the class
+
+Instantiate the StatsForecast class with the appropriate parameters
+
+
+```python
+season_length = 12 # Define season length as 12 months for monthly data
+horizon = 1 # Forecast horizon is set to 1 month
+
+# Define a list of models for forecasting
+models = [
+ AutoARIMA(season_length=season_length), # ARIMA model with automatic order selection and seasonal component
+ AutoETS(season_length=season_length), # ETS model with automatic error, trend, and seasonal component
+ AutoTheta(season_length=season_length), # Theta model with automatic seasonality detection
+ AutoCES(season_length=season_length), # CES model with automatic seasonality detection
+]
+
+# Instantiate StatsForecast class with models, data frequency ('M' for monthly),
+# and parallel computation on all CPU cores (n_jobs=-1)
+sf = StatsForecast(
+ models=models, # models for forecasting
+ freq=pd.offsets.MonthEnd(), # frequency of the timestamps
+ n_jobs=1 # number of jobs to run in parallel, -1 means using all processors
+)
+```
+
+## 4. a) Forecast with forecast method
+
+The `.forecast` method is faster for distributed computing and does not
+save the fittted models
+
+
+```python
+# Generate forecasts for the specified horizon using the sf object
+Y_hat_df = sf.forecast(df=Y_df, h=horizon) # forecast data
+# Display the first few rows of the forecast DataFrame
+Y_hat_df.head() # preview of forecasted data
+```
+
+| | unique_id | ds | AutoARIMA | AutoETS | AutoTheta | CES |
+|-----|-----------|------------|------------|------------|------------|-----------|
+| 0 | 1.0 | 1961-01-31 | 444.309575 | 442.357169 | 442.940797 | 453.03418 |
+
+## 4. b) Forecast with fit and predict
+
+The `.fit` method saves the fitted models
+
+
+```python
+sf.fit(df=Y_df) # Fit the models to the data using the fit method of the StatsForecast object
+
+sf.fitted_ # Access fitted models from the StatsForecast object
+
+Y_hat_df = sf.predict(h=horizon) # Predict or forecast 'horizon' steps ahead using the predict method
+
+Y_hat_df.head() # Preview the first few rows of the forecasted data
+```
+
+| | unique_id | ds | AutoARIMA | AutoETS | AutoTheta | CES |
+|-----|-----------|------------|------------|------------|------------|-----------|
+| 0 | 1.0 | 1961-01-31 | 444.309575 | 442.357169 | 442.940797 | 453.03418 |
+
+## References
+
+[Hyndman, RJ and Khandakar, Y (2008) “Automatic time series forecasting:
+The forecast package for R”, Journal of Statistical Software,
+26(3).](https://www.jstatsoft.org/article/view/v027i03)
+
diff --git a/statsforecast/docs/how-to-guides/exogenous.html.mdx b/statsforecast/docs/how-to-guides/exogenous.html.mdx
new file mode 100644
index 00000000..67462813
--- /dev/null
+++ b/statsforecast/docs/how-to-guides/exogenous.html.mdx
@@ -0,0 +1,361 @@
+---
+description: >-
+ In this notebook, we'll incorporate exogenous regressors to a StatsForecast
+ model.
+output-file: exogenous.html
+title: Exogenous Regressors
+---
+
+
+> **Prerequesites**
+>
+> This tutorial assumes basic familiarity with StatsForecast. For a
+> minimal example visit the [Quick
+> Start](../getting-started/getting_started_short.html)
+
+## Introduction
+
+**Exogenous regressors** are variables that can affect the values of a
+time series. They may not be directly related to the variable that is
+beging forecasted, but they can still have an impact on it. Examples of
+exogenous regressors are weather data, economic indicators, or
+promotional sales. They are typically collected from external sources
+and by incorporating them into a forecasting model, they can improve the
+accuracy of our predictions.
+
+By the end of this tutorial, you’ll have a good understanding of how to
+incorporate exogenous regressors into
+[StatsForecast](https://nixtla.github.io/statsforecast/)’s models.
+Furthermore, you’ll see how to evaluate their performance and decide
+whether or not they can help enhance the forecast.
+
+**Outline**
+
+1. Install libraries
+2. Load and explore the data
+3. Split train/test set
+4. Add exogenous regressors
+5. Create future exogenous regressors
+6. Train model
+7. Evaluate results
+
+> **Tip**
+>
+> You can use Colab to run this Notebook interactively
+>
+
+## Install libraries
+
+We assume that you have StatsForecast already installed. If not, check
+this guide for instructions on [how to install
+StatsForecast](../getting-started/installation.html)
+
+
+```python
+# uncomment the following line to install the library
+# %pip install statsforecast
+```
+
+
+```python
+import pandas as pd
+```
+
+## Load and explore the data
+
+In this example, we’ll use a single time series from the [M5
+Competition](https://www.sciencedirect.com/science/article/pii/S0169207021001187#:~:text=The%20objective%20of%20the%20M5,the%20uncertainty%20around%20these%20forecasts.)
+dataset. This series represents the daily sales of a product in a
+Walmart store. The product-store combination that we’ll use in this
+notebook has `unique_id = FOODS_3_586_CA_3`. This time series was chosen
+because it is not intermittent and has exogenous regressors that will be
+useful for forecasting.
+
+We’ll load the following dataframes:
+
+- `Y_ts`: (pandas DataFrame) The target time series with columns
+ \[`unique_id`, `ds`, `y`\].
+- `X_ts`: (pandas DataFrame) Exogenous time series with columns
+ \[`unique_id`, `ds`, exogenous regressors\].
+
+
+```python
+base_url = 'https://datasets-nixtla.s3.amazonaws.com'
+filters = [('unique_id', '=', 'FOODS_3_586_CA_3')]
+Y_ts = pd.read_parquet(f'{base_url}/m5_y.parquet', filters=filters)
+X_ts = pd.read_parquet(f'{base_url}/m5_x.parquet', filters=filters)
+```
+
+We can plot the sales of this product-store combination with the
+`statsforecast.plot` method from the
+[StatsForecast](../../src/core/core.html#statsforecast)
+class. This method has multiple parameters, and the requiered ones to
+generate the plots in this notebook are explained below.
+
+- `df`: A pandas dataframe with columns \[`unique_id`, `ds`, `y`\].
+- `forecasts_df`: A pandas dataframe with columns \[`unique_id`,
+ `ds`\] and models.
+- `engine`: str = `matplotlib`. It can also be `plotly`. `plotly`
+ generates interactive plots, while `matplotlib` generates static
+ plots.
+
+
+```python
+from statsforecast import StatsForecast
+```
+
+
+```python
+StatsForecast.plot(Y_ts)
+```
+
+
+
+The M5 Competition included several exogenous regressors. Here we’ll use
+the following two.
+
+- `sell_price`: The price of the product for the given store. The
+ price is provided per week.
+- `snap_CA`: A binary variable indicating whether the store allows
+ SNAP purchases (1 if yes, 0 otherwise). SNAP stands for Supplement
+ Nutrition Assitance Program, and it gives individuals and families
+ money to help them purchase food products.
+
+
+```python
+X_ts = X_ts[['unique_id', 'ds', 'sell_price', 'snap_CA']]
+X_ts.head()
+```
+
+| | unique_id | ds | sell_price | snap_CA |
+|-----|------------------|------------|------------|---------|
+| 0 | FOODS_3_586_CA_3 | 2011-01-29 | 1.48 | 0 |
+| 1 | FOODS_3_586_CA_3 | 2011-01-30 | 1.48 | 0 |
+| 2 | FOODS_3_586_CA_3 | 2011-01-31 | 1.48 | 0 |
+| 3 | FOODS_3_586_CA_3 | 2011-02-01 | 1.48 | 1 |
+| 4 | FOODS_3_586_CA_3 | 2011-02-02 | 1.48 | 1 |
+
+Here the `unique_id` is a category, but for the exogenous regressors it
+needs to be a string.
+
+
+```python
+X_ts['unique_id'] = X_ts.unique_id.astype(str)
+```
+
+We can plot the exogenous regressors using `plotly`. We could use
+`statsforecast.plot`, but then one of the regressors must be renamed
+`y`, and the name must be changed back to the original before generating
+the forecast.
+
+
+```python
+StatsForecast.plot(Y_ts, X_ts, max_insample_length=0)
+```
+
+
+
+From this plot, we can conclude that price has increased twice and that
+SNAP occurs at regular intervals.
+
+## Split train/test set
+
+In the M5 Competition, participants had to forecast sales for the last
+28 days in the dataset. We’ll use the same forecast horizon and create
+the train and test sets accordingly.
+
+
+```python
+# Extract dates for train and test set
+dates = Y_ts['ds'].unique()
+dtrain = dates[:-28]
+dtest = dates[-28:]
+
+Y_train = Y_ts.query('ds in @dtrain')
+Y_test = Y_ts.query('ds in @dtest')
+
+X_train = X_ts.query('ds in @dtrain')
+X_test = X_ts.query('ds in @dtest')
+```
+
+## Add exogenous regressors
+
+The exogenous regressors need to be place after the target variable `y`.
+
+
+```python
+train = Y_train.merge(X_ts, how = 'left', on = ['unique_id', 'ds'])
+train.head()
+```
+
+| | unique_id | ds | y | sell_price | snap_CA |
+|-----|------------------|------------|------|------------|---------|
+| 0 | FOODS_3_586_CA_3 | 2011-01-29 | 56.0 | 1.48 | 0 |
+| 1 | FOODS_3_586_CA_3 | 2011-01-30 | 55.0 | 1.48 | 0 |
+| 2 | FOODS_3_586_CA_3 | 2011-01-31 | 45.0 | 1.48 | 0 |
+| 3 | FOODS_3_586_CA_3 | 2011-02-01 | 57.0 | 1.48 | 1 |
+| 4 | FOODS_3_586_CA_3 | 2011-02-02 | 54.0 | 1.48 | 1 |
+
+## Create future exogenous regressors
+
+We need to include the future values of the exogenous regressors so that
+we can produce the forecasts. Notice that we already have this
+information in `X_test`.
+
+
+```python
+X_test.head()
+```
+
+| | unique_id | ds | sell_price | snap_CA |
+|------|------------------|------------|------------|---------|
+| 1941 | FOODS_3_586_CA_3 | 2016-05-23 | 1.68 | 0 |
+| 1942 | FOODS_3_586_CA_3 | 2016-05-24 | 1.68 | 0 |
+| 1943 | FOODS_3_586_CA_3 | 2016-05-25 | 1.68 | 0 |
+| 1944 | FOODS_3_586_CA_3 | 2016-05-26 | 1.68 | 0 |
+| 1945 | FOODS_3_586_CA_3 | 2016-05-27 | 1.68 | 0 |
+
+> **Important**
+>
+> If the future values of the exogenous regressors are not available,
+> then they must be forecasted or the regressors need to be eliminated
+> from the model. Without them, it is not possible to generate the
+> forecast.
+
+## Train model
+
+To generate the forecast, we’ll use
+[AutoARIMA](../../src/core/models.html#autoarima),
+which is one of the models available in StatsForecast that allows
+exogenous regressors. To use this model, we first need to import it from
+`statsforecast.models` and then we need to instatiate it. Given that
+we’re working with daily data, we need to set `season_length = 7`.
+
+
+```python
+from statsforecast.models import AutoARIMA
+```
+
+
+```python
+# Create a list with the model and its instantiation parameters
+models = [AutoARIMA(season_length=7)]
+```
+
+Next, we need to instantiate a new StatsForecast object, which has the
+following parameters.
+
+- `df`: The dataframe with the training data.
+- `models`: The list of models defined in the previous step.
+- `freq`: A string indicating the frequency of the data. See [pandas’
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+- `n_jobs`: An integer that indicates the number of jobs used in
+ parallel processing. Use -1 to select all cores.
+
+
+```python
+sf = StatsForecast(
+ models=models,
+ freq='D',
+ n_jobs=1,
+)
+```
+
+Now we’re ready to generate the forecast. To do this, we’ll use the
+`forecast` method, which takes the following arguments.
+
+- `h`: An integer that represents the forecast horizon. In this case,
+ we’ll forecast the next 28 days.
+- `X_df`: A pandas dataframe with the future values of the exogenous
+ regressors.
+- `level`: A list of floats with the confidence levels of the
+ prediction intervals. For example, `level=[95]` means that the range
+ of values should include the actual future value with probability
+ 95%.
+
+
+```python
+horizon = 28
+level = [95]
+
+fcst = sf.forecast(df=train, h=horizon, X_df=X_test, level=level)
+fcst.head()
+```
+
+| | unique_id | ds | AutoARIMA | AutoARIMA-lo-95 | AutoARIMA-hi-95 |
+|-----|------------------|------------|-----------|-----------------|-----------------|
+| 0 | FOODS_3_586_CA_3 | 2016-05-23 | 72.956276 | 44.109070 | 101.803482 |
+| 1 | FOODS_3_586_CA_3 | 2016-05-24 | 71.138611 | 40.761467 | 101.515747 |
+| 2 | FOODS_3_586_CA_3 | 2016-05-25 | 68.140945 | 37.550083 | 98.731804 |
+| 3 | FOODS_3_586_CA_3 | 2016-05-26 | 65.485588 | 34.841637 | 96.129539 |
+| 4 | FOODS_3_586_CA_3 | 2016-05-27 | 64.961441 | 34.291973 | 95.630905 |
+
+We can plot the forecasts with the `statsforecast.plot` method described
+above.
+
+
+```python
+StatsForecast.plot(Y_ts, fcst, max_insample_length=28*2)
+```
+
+
+
+## Evaluate results
+
+We’ll merge the test set and the forecast to evaluate the accuracy using
+the [mean absolute
+error](https://en.wikipedia.org/wiki/Mean_absolute_error) (MAE).
+
+
+```python
+res = Y_test.merge(fcst, how='left', on=['unique_id', 'ds'])
+res.head()
+```
+
+| | unique_id | ds | y | AutoARIMA | AutoARIMA-lo-95 | AutoARIMA-hi-95 |
+|-----|------------------|------------|------|-----------|-----------------|-----------------|
+| 0 | FOODS_3_586_CA_3 | 2016-05-23 | 66.0 | 72.956276 | 44.109070 | 101.803482 |
+| 1 | FOODS_3_586_CA_3 | 2016-05-24 | 62.0 | 71.138611 | 40.761467 | 101.515747 |
+| 2 | FOODS_3_586_CA_3 | 2016-05-25 | 40.0 | 68.140945 | 37.550083 | 98.731804 |
+| 3 | FOODS_3_586_CA_3 | 2016-05-26 | 72.0 | 65.485588 | 34.841637 | 96.129539 |
+| 4 | FOODS_3_586_CA_3 | 2016-05-27 | 69.0 | 64.961441 | 34.291973 | 95.630905 |
+
+
+```python
+mae = abs(res['y']-res['AutoARIMA']).mean()
+print('The MAE with exogenous regressors is '+str(round(mae,2)))
+```
+
+``` text
+The MAE with exogenous regressors is 11.42
+```
+
+To check whether the exogenous regressors were useful or not, we need to
+generate the forecast again, now without them. To do this, we simple
+pass the dataframe wihtout exogenous variables to the `forecast` method.
+Notice that the data only includes `unique_id`, `ds`, and `y`. The
+`forecast` method no longer requieres the future values of the exogenous
+regressors `X_df`.
+
+
+```python
+# univariate model
+fcst_u = sf.forecast(df=train[['unique_id', 'ds', 'y']], h=28)
+
+res_u = Y_test.merge(fcst_u, how='left', on=['unique_id', 'ds'])
+mae_u = abs(res_u['y']-res_u['AutoARIMA']).mean()
+```
+
+
+```python
+print('The MAE without exogenous regressors is '+str(round(mae_u,2)))
+```
+
+``` text
+The MAE without exogenous regressors is 12.18
+```
+
+Hence, we can conclude that using `sell_price` and `snap_CA` as external
+regressors helped improve the forecast.
+
diff --git a/statsforecast/docs/how-to-guides/generating_features.html.mdx b/statsforecast/docs/how-to-guides/generating_features.html.mdx
new file mode 100644
index 00000000..b72419e7
--- /dev/null
+++ b/statsforecast/docs/how-to-guides/generating_features.html.mdx
@@ -0,0 +1,149 @@
+---
+description: Leverage StatsForecast models to create features
+output-file: generating_features.html
+title: Generating features
+---
+
+
+Some models create internal representations of the series that can be
+useful for other models to use as inputs. One example is the
+[`MSTL`](https://Nixtla.github.io/statsforecast/src/core/models.html#mstl)
+model, which decomposes the series into trend and seasonal components.
+This guide shows you how to use the
+[`mstl_decomposition`](https://Nixtla.github.io/statsforecast/src/feature_engineering.html#mstl_decomposition)
+function to extract those features for training and then use their
+future values for inference.
+
+
+```python
+from functools import partial
+
+import pandas as pd
+import statsforecast
+from statsforecast import StatsForecast
+from statsforecast.feature_engineering import mstl_decomposition
+from statsforecast.models import ARIMA, MSTL
+from utilsforecast.evaluation import evaluate
+from utilsforecast.losses import smape, mase
+```
+
+
+```python
+df = pd.read_parquet('https://datasets-nixtla.s3.amazonaws.com/m4-hourly.parquet')
+uids = df['unique_id'].unique()[:10]
+df = df[df['unique_id'].isin(uids)]
+df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|-----|-------|
+| 0 | H1 | 1 | 605.0 |
+| 1 | H1 | 2 | 586.0 |
+| 2 | H1 | 3 | 586.0 |
+| 3 | H1 | 4 | 559.0 |
+| 4 | H1 | 5 | 511.0 |
+
+Suppose that you want to use an ARIMA model to forecast your series but
+you want to incorporate the trend and seasonal components from the MSTL
+model as external regressors. You can define the MSTL model to use and
+then provide it to the mstl_decomposition function.
+
+
+```python
+freq = 1
+season_length = 24
+horizon = 2 * season_length
+valid = df.groupby('unique_id').tail(horizon)
+train = df.drop(valid.index)
+model = MSTL(season_length=24)
+transformed_df, X_df = mstl_decomposition(train, model=model, freq=freq, h=horizon)
+```
+
+This generates the dataframe that we should use for training (with the
+trend and seasonal columns added), as well as the dataframe we should
+use to forecast.
+
+
+```python
+transformed_df.head()
+```
+
+| | unique_id | ds | y | trend | seasonal |
+|-----|-----------|-----|-------|------------|------------|
+| 0 | H1 | 1 | 605.0 | 502.872910 | 131.419934 |
+| 1 | H1 | 2 | 586.0 | 507.873456 | 93.100015 |
+| 2 | H1 | 3 | 586.0 | 512.822533 | 82.155386 |
+| 3 | H1 | 4 | 559.0 | 517.717481 | 42.412749 |
+| 4 | H1 | 5 | 511.0 | 522.555849 | -11.401890 |
+
+
+```python
+X_df.head()
+```
+
+| | unique_id | ds | trend | seasonal |
+|-----|-----------|-----|------------|-------------|
+| 0 | H1 | 701 | 643.801348 | -29.189627 |
+| 1 | H1 | 702 | 644.328207 | -99.680432 |
+| 2 | H1 | 703 | 644.749693 | -141.169014 |
+| 3 | H1 | 704 | 645.086883 | -173.325625 |
+| 4 | H1 | 705 | 645.356634 | -195.862530 |
+
+We can now train our ARIMA models and compute our forecasts.
+
+
+```python
+sf = StatsForecast(
+ models=[ARIMA(order=(1, 0, 1), season_length=season_length)],
+ freq=freq
+)
+preds = sf.forecast(h=horizon, df=transformed_df, X_df=X_df)
+preds.head()
+```
+
+| | unique_id | ds | ARIMA |
+|-----|-----------|-----|------------|
+| 0 | H1 | 701 | 612.737668 |
+| 1 | H1 | 702 | 542.851796 |
+| 2 | H1 | 703 | 501.931839 |
+| 3 | H1 | 704 | 470.248289 |
+| 4 | H1 | 705 | 448.115839 |
+
+We can now evaluate the performance.
+
+
+```python
+def compute_evaluation(preds):
+ full = preds.merge(valid, on=['unique_id', 'ds'])
+ mase24 = partial(mase, seasonality=24)
+ res = evaluate(full, metrics=[smape, mase24], train_df=train).groupby('metric')['ARIMA'].mean()
+ res_smape = '{:.1%}'.format(res['smape'])
+ res_mase = '{:.1f}'.format(res['mase'])
+ return pd.Series({'mase': res_mase, 'smape': res_smape})
+```
+
+
+```python
+compute_evaluation(preds)
+```
+
+``` text
+mase 1.0
+smape 3.9%
+dtype: object
+```
+
+And compare this with just using the series values.
+
+
+```python
+preds_noexog = sf.forecast(h=horizon, df=train)
+compute_evaluation(preds_noexog)
+```
+
+``` text
+mase 2.3
+smape 7.7%
+dtype: object
+```
+
diff --git a/statsforecast/docs/how-to-guides/migrating_R.mdx b/statsforecast/docs/how-to-guides/migrating_R.mdx
new file mode 100644
index 00000000..83a2aaea
--- /dev/null
+++ b/statsforecast/docs/how-to-guides/migrating_R.mdx
@@ -0,0 +1,11 @@
+---
+title: Migrating from R
+---
+
+
+## 🚧 We are working on this site.
+
+This site is currently in development. If you are particularly
+interested in this section, please open a GitHub Issue, and we will
+prioritize it.
+
diff --git a/statsforecast/docs/how-to-guides/numba_cache.html.mdx b/statsforecast/docs/how-to-guides/numba_cache.html.mdx
new file mode 100644
index 00000000..b65ec3bb
--- /dev/null
+++ b/statsforecast/docs/how-to-guides/numba_cache.html.mdx
@@ -0,0 +1,32 @@
+---
+description: Enabling caching for numba functions to reduce cold-starts
+output-file: numba_cache.html
+title: Numba caching
+---
+
+
+`statsforecast` makes heavy use of [numba](https://numba.pydata.org/) to
+speed up several critical functions that estimate model parameters. This
+comes at a cost though, which is that the functions have to be [JIT
+compiled](https://en.wikipedia.org/wiki/Just-in-time_compilation) the
+first time they’re run, which can be expensive. Once a function has ben
+JIT compiled, subsequent calls are significantly faster. One problem is
+that this compilation is saved (by default) on a per-session basis.
+
+In order to mitigate the compilation overhead numba offers the option to
+cache the function compiled code to a file, which can be then reused
+across sessions, and even copied over to different machines that share
+the same CPU characteristics ([more information](https://numba.readthedocs.io/en/stable/developer/caching.html)).
+
+To leverage caching, you can set the `NIXTLA_NUMBA_CACHE` environment
+variable (e.g. `NIXTLA_NUMBA_CACHE=1`), which will enable caching for
+all functions. By default the cache is saved to the `__pycache__`
+directory, but you can override this with the `NUMBA_CACHE_DIR`
+environment variable to save it to a different path
+(e.g. `NUMBA_CACHE_DIR=numba_cache`), you can find more information in
+the [docs](https://numba.readthedocs.io/en/stable/reference/envvars.html#envvar-NUMBA_CACHE_DIR).
+
+If you want to have this enabled for all your sessions, we suggest
+adding `export NIXTLA_NUMBA_CACHE=1` to your profile files, such as
+`.bashrc`, `.zshrc`, etc.
+
diff --git a/statsforecast/docs/how-to-guides/sklearn_models.html.mdx b/statsforecast/docs/how-to-guides/sklearn_models.html.mdx
new file mode 100644
index 00000000..1199429e
--- /dev/null
+++ b/statsforecast/docs/how-to-guides/sklearn_models.html.mdx
@@ -0,0 +1,99 @@
+---
+description: Use any scikit-learn model for forecasting
+output-file: sklearn_models.html
+title: Sklearn models
+---
+
+
+statsforecast supports providing scikit-learn models through the
+[`statsforecast.models.SklearnModel`](https://Nixtla.github.io/statsforecast/src/core/models.html#sklearnmodel)
+wrapper. This can help you leverage feature engineering and train one
+model per serie, which can sometimes be better than training a single
+global model (as in mlforecast).
+
+## Data setup
+
+
+```python
+from functools import partial
+
+from datasetsforecast.m4 import M4, M4Info
+from sklearn.linear_model import Lasso, Ridge
+from utilsforecast.feature_engineering import pipeline, trend, fourier
+from utilsforecast.plotting import plot_series
+
+from statsforecast import StatsForecast
+from statsforecast.models import SklearnModel
+from statsforecast.utils import ConformalIntervals
+```
+
+
+```python
+group = 'Hourly'
+season_length = M4Info[group].seasonality
+horizon = M4Info[group].horizon
+data, *_ = M4.load('data', group)
+data['ds'] = data['ds'].astype('int64')
+valid = data.groupby('unique_id').tail(horizon).copy()
+train = data.drop(valid.index)
+train.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|-----|-------|
+| 0 | H1 | 1 | 605.0 |
+| 1 | H1 | 2 | 586.0 |
+| 2 | H1 | 3 | 586.0 |
+| 3 | H1 | 4 | 559.0 |
+| 4 | H1 | 5 | 511.0 |
+
+## Generating features
+
+The utilsforecast library [provides some utilies for feature
+engineering](https://nixtlaverse.nixtla.io/utilsforecast/feature_engineering.html).
+
+
+```python
+train_features, valid_features = pipeline(
+ train,
+ features=[
+ trend,
+ partial(fourier, season_length=season_length, k=10), # 10 fourier terms
+ ],
+ freq=1,
+ h=horizon,
+)
+train_features.head()
+```
+
+| | unique_id | ds | y | trend | sin1_24 | sin2_24 | sin3_24 | sin4_24 | sin5_24 | sin6_24 | sin7_24 | sin8_24 | sin9_24 | sin10_24 | cos1_24 | cos2_24 | cos3_24 | cos4_24 | cos5_24 | cos6_24 | cos7_24 | cos8_24 | cos9_24 | cos10_24 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | H1 | 1 | 605.0 | 261.0 | -0.707105 | -1.000000 | -0.707108 | -0.000012 | 0.707112 | 1.000000 | 0.707095 | 0.000024 | -0.707125 | -1.000000 | 0.707109 | 0.000006 | -0.707106 | -1.000000 | -0.707101 | -0.000003 | 0.707119 | 1.000000 | 0.707088 | -0.000015 |
+| 1 | H1 | 2 | 586.0 | 262.0 | -0.500001 | -0.866027 | -1.000000 | -0.866023 | -0.499988 | -0.000007 | 0.500001 | 0.866031 | 1.000000 | 0.866011 | 0.866025 | 0.499998 | 0.000004 | -0.500005 | -0.866032 | -1.000000 | -0.866025 | -0.499991 | 0.000019 | 0.500025 |
+| 2 | H1 | 3 | 586.0 | 263.0 | -0.258817 | -0.499997 | -0.707103 | -0.866021 | -0.965931 | -1.000000 | -0.965922 | -0.866033 | -0.707098 | -0.499964 | 0.965926 | 0.866027 | 0.707111 | 0.500007 | 0.258799 | 0.000012 | -0.258835 | -0.499986 | -0.707116 | -0.866046 |
+| 3 | H1 | 4 | 559.0 | 264.0 | 0.000005 | 0.000011 | 0.000008 | 0.000021 | 0.000003 | 0.000016 | -0.000001 | 0.000042 | -0.000006 | 0.000007 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 |
+| 4 | H1 | 5 | 511.0 | 265.0 | 0.258820 | 0.500002 | 0.707114 | 0.866027 | 0.965925 | 1.000000 | 0.965930 | 0.866022 | 0.707106 | 0.500005 | 0.965926 | 0.866024 | 0.707099 | 0.499997 | 0.258822 | -0.000021 | -0.258803 | -0.500006 | -0.707107 | -0.866022 |
+
+## Forecasting
+
+
+```python
+sf = StatsForecast(
+ models=[
+ SklearnModel(Lasso()),
+ SklearnModel(Ridge()),
+ ],
+ freq=1,
+)
+preds = sf.forecast(
+ df=train_features,
+ h=horizon,
+ X_df=valid_features,
+ prediction_intervals=ConformalIntervals(n_windows=4, h=horizon),
+ level=[95],
+)
+plot_series(train, preds, level=[95], palette='tab20b', max_ids=4)
+```
+
+
+
diff --git a/statsforecast/docs/how-to-guides/sklearn_models_files/figure-markdown_strict/cell-5-output-1.png b/statsforecast/docs/how-to-guides/sklearn_models_files/figure-markdown_strict/cell-5-output-1.png
new file mode 100644
index 00000000..4cc85274
Binary files /dev/null and b/statsforecast/docs/how-to-guides/sklearn_models_files/figure-markdown_strict/cell-5-output-1.png differ
diff --git a/statsforecast/docs/models/ADIDA_files/figure-markdown_strict/cell-11-output-2.png b/statsforecast/docs/models/ADIDA_files/figure-markdown_strict/cell-11-output-2.png
new file mode 100644
index 00000000..2fb65400
Binary files /dev/null and b/statsforecast/docs/models/ADIDA_files/figure-markdown_strict/cell-11-output-2.png differ
diff --git a/statsforecast/docs/models/ADIDA_files/figure-markdown_strict/cell-14-output-1.png b/statsforecast/docs/models/ADIDA_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..aa55bcab
Binary files /dev/null and b/statsforecast/docs/models/ADIDA_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/statsforecast/docs/models/ADIDA_files/figure-markdown_strict/cell-21-output-1.png b/statsforecast/docs/models/ADIDA_files/figure-markdown_strict/cell-21-output-1.png
new file mode 100644
index 00000000..f55683be
Binary files /dev/null and b/statsforecast/docs/models/ADIDA_files/figure-markdown_strict/cell-21-output-1.png differ
diff --git a/statsforecast/docs/models/ADIDA_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/ADIDA_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..94324779
Binary files /dev/null and b/statsforecast/docs/models/ADIDA_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/ADIDA_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/models/ADIDA_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..057db709
Binary files /dev/null and b/statsforecast/docs/models/ADIDA_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-12-output-2.png b/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-12-output-2.png
new file mode 100644
index 00000000..3f607d5a
Binary files /dev/null and b/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-12-output-2.png differ
diff --git a/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-14-output-1.png b/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..ad3ebd17
Binary files /dev/null and b/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-16-output-1.png b/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-16-output-1.png
new file mode 100644
index 00000000..459ee0d6
Binary files /dev/null and b/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-16-output-1.png differ
diff --git a/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-21-output-1.png b/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-21-output-1.png
new file mode 100644
index 00000000..5b5c827f
Binary files /dev/null and b/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-21-output-1.png differ
diff --git a/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-28-output-1.png b/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-28-output-1.png
new file mode 100644
index 00000000..753c83e9
Binary files /dev/null and b/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-28-output-1.png differ
diff --git a/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-33-output-1.png b/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-33-output-1.png
new file mode 100644
index 00000000..e94a837d
Binary files /dev/null and b/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-33-output-1.png differ
diff --git a/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-37-output-1.png b/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-37-output-1.png
new file mode 100644
index 00000000..34894f4c
Binary files /dev/null and b/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-37-output-1.png differ
diff --git a/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..1e180e9c
Binary files /dev/null and b/statsforecast/docs/models/ARCH_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-11-output-1.png b/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..3d8b35d3
Binary files /dev/null and b/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-17-output-1.png b/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-17-output-1.png
new file mode 100644
index 00000000..6fa99f79
Binary files /dev/null and b/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-17-output-1.png differ
diff --git a/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-18-output-1.png b/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-18-output-1.png
new file mode 100644
index 00000000..e84c1652
Binary files /dev/null and b/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-18-output-1.png differ
diff --git a/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-19-output-1.png b/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-19-output-1.png
new file mode 100644
index 00000000..422a66a2
Binary files /dev/null and b/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-19-output-1.png differ
diff --git a/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-26-output-1.png b/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-26-output-1.png
new file mode 100644
index 00000000..ab8cad9c
Binary files /dev/null and b/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-26-output-1.png differ
diff --git a/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..dcdb3dc4
Binary files /dev/null and b/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-9-output-2.png b/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-9-output-2.png
new file mode 100644
index 00000000..16d8a28b
Binary files /dev/null and b/statsforecast/docs/models/ARIMA_files/figure-markdown_strict/cell-9-output-2.png differ
diff --git a/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-10-output-1.png b/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..9f63cbf0
Binary files /dev/null and b/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-11-output-1.png b/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..f4c068a9
Binary files /dev/null and b/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-14-output-1.png b/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..d0bcc94a
Binary files /dev/null and b/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-22-output-1.png b/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-22-output-1.png
new file mode 100644
index 00000000..eed56e45
Binary files /dev/null and b/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-22-output-1.png differ
diff --git a/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-26-output-1.png b/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-26-output-1.png
new file mode 100644
index 00000000..77552401
Binary files /dev/null and b/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-26-output-1.png differ
diff --git a/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-30-output-1.png b/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-30-output-1.png
new file mode 100644
index 00000000..68e90c15
Binary files /dev/null and b/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-30-output-1.png differ
diff --git a/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..cb2a1f9e
Binary files /dev/null and b/statsforecast/docs/models/AutoARIMA_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-10-output-1.png b/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..02324ba5
Binary files /dev/null and b/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-11-output-1.png b/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..3d8b35d3
Binary files /dev/null and b/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-14-output-1.png b/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..ac9c29cd
Binary files /dev/null and b/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-21-output-1.png b/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-21-output-1.png
new file mode 100644
index 00000000..818cdad2
Binary files /dev/null and b/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-21-output-1.png differ
diff --git a/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-24-output-1.png b/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-24-output-1.png
new file mode 100644
index 00000000..21120e5e
Binary files /dev/null and b/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-24-output-1.png differ
diff --git a/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-27-output-1.png b/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-27-output-1.png
new file mode 100644
index 00000000..ce7bd6be
Binary files /dev/null and b/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-27-output-1.png differ
diff --git a/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-30-output-1.png b/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-30-output-1.png
new file mode 100644
index 00000000..f8e37c77
Binary files /dev/null and b/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-30-output-1.png differ
diff --git a/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..a42a202c
Binary files /dev/null and b/statsforecast/docs/models/AutoCES_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/models/AutoETS_files/figure-markdown_strict/cell-10-output-1.png b/statsforecast/docs/models/AutoETS_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..c5e3e494
Binary files /dev/null and b/statsforecast/docs/models/AutoETS_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/statsforecast/docs/models/AutoETS_files/figure-markdown_strict/cell-11-output-1.png b/statsforecast/docs/models/AutoETS_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..de653532
Binary files /dev/null and b/statsforecast/docs/models/AutoETS_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/statsforecast/docs/models/AutoETS_files/figure-markdown_strict/cell-14-output-1.png b/statsforecast/docs/models/AutoETS_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..55192c66
Binary files /dev/null and b/statsforecast/docs/models/AutoETS_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/statsforecast/docs/models/AutoETS_files/figure-markdown_strict/cell-19-output-1.png b/statsforecast/docs/models/AutoETS_files/figure-markdown_strict/cell-19-output-1.png
new file mode 100644
index 00000000..3161536f
Binary files /dev/null and b/statsforecast/docs/models/AutoETS_files/figure-markdown_strict/cell-19-output-1.png differ
diff --git a/statsforecast/docs/models/AutoETS_files/figure-markdown_strict/cell-21-output-1.png b/statsforecast/docs/models/AutoETS_files/figure-markdown_strict/cell-21-output-1.png
new file mode 100644
index 00000000..b8f13dc1
Binary files /dev/null and b/statsforecast/docs/models/AutoETS_files/figure-markdown_strict/cell-21-output-1.png differ
diff --git a/statsforecast/docs/models/AutoETS_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/models/AutoETS_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..a42a202c
Binary files /dev/null and b/statsforecast/docs/models/AutoETS_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-15-output-1.png b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-15-output-1.png
new file mode 100644
index 00000000..517221f3
Binary files /dev/null and b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-15-output-1.png differ
diff --git a/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-18-output-1.png b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-18-output-1.png
new file mode 100644
index 00000000..e39bc4f5
Binary files /dev/null and b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-18-output-1.png differ
diff --git a/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-26-output-1.png b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-26-output-1.png
new file mode 100644
index 00000000..23f30a58
Binary files /dev/null and b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-26-output-1.png differ
diff --git a/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-31-output-1.png b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-31-output-1.png
new file mode 100644
index 00000000..708b6338
Binary files /dev/null and b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-31-output-1.png differ
diff --git a/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-34-output-1.png b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-34-output-1.png
new file mode 100644
index 00000000..2f992372
Binary files /dev/null and b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-34-output-1.png differ
diff --git a/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-38-output-1.png b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-38-output-1.png
new file mode 100644
index 00000000..89885757
Binary files /dev/null and b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-38-output-1.png differ
diff --git a/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-38-output-2.png b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-38-output-2.png
new file mode 100644
index 00000000..60d24bc0
Binary files /dev/null and b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-38-output-2.png differ
diff --git a/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-38-output-3.png b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-38-output-3.png
new file mode 100644
index 00000000..69eac505
Binary files /dev/null and b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-38-output-3.png differ
diff --git a/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-38-output-4.png b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-38-output-4.png
new file mode 100644
index 00000000..5110ac3c
Binary files /dev/null and b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-38-output-4.png differ
diff --git a/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-38-output-5.png b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-38-output-5.png
new file mode 100644
index 00000000..3f96c9ff
Binary files /dev/null and b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-38-output-5.png differ
diff --git a/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..96aa1327
Binary files /dev/null and b/statsforecast/docs/models/AutoRegressive_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-10-output-1.png b/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..b03e5c56
Binary files /dev/null and b/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-13-output-1.png b/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-13-output-1.png
new file mode 100644
index 00000000..6704b4f5
Binary files /dev/null and b/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-13-output-1.png differ
diff --git a/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-20-output-1.png b/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-20-output-1.png
new file mode 100644
index 00000000..9edbf100
Binary files /dev/null and b/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-20-output-1.png differ
diff --git a/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-23-output-1.png b/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-23-output-1.png
new file mode 100644
index 00000000..fa9eb46d
Binary files /dev/null and b/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-23-output-1.png differ
diff --git a/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-26-output-1.png b/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-26-output-1.png
new file mode 100644
index 00000000..fc6cb038
Binary files /dev/null and b/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-26-output-1.png differ
diff --git a/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-29-output-1.png b/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-29-output-1.png
new file mode 100644
index 00000000..f72fa20a
Binary files /dev/null and b/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-29-output-1.png differ
diff --git a/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..935ccb53
Binary files /dev/null and b/statsforecast/docs/models/AutoTheta_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/models/CrostonClassic_files/figure-markdown_strict/cell-11-output-2.png b/statsforecast/docs/models/CrostonClassic_files/figure-markdown_strict/cell-11-output-2.png
new file mode 100644
index 00000000..7f39609e
Binary files /dev/null and b/statsforecast/docs/models/CrostonClassic_files/figure-markdown_strict/cell-11-output-2.png differ
diff --git a/statsforecast/docs/models/CrostonClassic_files/figure-markdown_strict/cell-14-output-1.png b/statsforecast/docs/models/CrostonClassic_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..2a756eee
Binary files /dev/null and b/statsforecast/docs/models/CrostonClassic_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/statsforecast/docs/models/CrostonClassic_files/figure-markdown_strict/cell-21-output-1.png b/statsforecast/docs/models/CrostonClassic_files/figure-markdown_strict/cell-21-output-1.png
new file mode 100644
index 00000000..b53c6c8d
Binary files /dev/null and b/statsforecast/docs/models/CrostonClassic_files/figure-markdown_strict/cell-21-output-1.png differ
diff --git a/statsforecast/docs/models/CrostonClassic_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/CrostonClassic_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..56c6ac59
Binary files /dev/null and b/statsforecast/docs/models/CrostonClassic_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/CrostonClassic_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/models/CrostonClassic_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..8b44cf08
Binary files /dev/null and b/statsforecast/docs/models/CrostonClassic_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/models/CrostonOptimized_files/figure-markdown_strict/cell-12-output-2.png b/statsforecast/docs/models/CrostonOptimized_files/figure-markdown_strict/cell-12-output-2.png
new file mode 100644
index 00000000..7f39609e
Binary files /dev/null and b/statsforecast/docs/models/CrostonOptimized_files/figure-markdown_strict/cell-12-output-2.png differ
diff --git a/statsforecast/docs/models/CrostonOptimized_files/figure-markdown_strict/cell-21-output-1.png b/statsforecast/docs/models/CrostonOptimized_files/figure-markdown_strict/cell-21-output-1.png
new file mode 100644
index 00000000..dbaaa107
Binary files /dev/null and b/statsforecast/docs/models/CrostonOptimized_files/figure-markdown_strict/cell-21-output-1.png differ
diff --git a/statsforecast/docs/models/CrostonOptimized_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/CrostonOptimized_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..56c6ac59
Binary files /dev/null and b/statsforecast/docs/models/CrostonOptimized_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/CrostonOptimized_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/models/CrostonOptimized_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..8b44cf08
Binary files /dev/null and b/statsforecast/docs/models/CrostonOptimized_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/models/CrostonSBA_files/figure-markdown_strict/cell-11-output-2.png b/statsforecast/docs/models/CrostonSBA_files/figure-markdown_strict/cell-11-output-2.png
new file mode 100644
index 00000000..7f39609e
Binary files /dev/null and b/statsforecast/docs/models/CrostonSBA_files/figure-markdown_strict/cell-11-output-2.png differ
diff --git a/statsforecast/docs/models/CrostonSBA_files/figure-markdown_strict/cell-20-output-1.png b/statsforecast/docs/models/CrostonSBA_files/figure-markdown_strict/cell-20-output-1.png
new file mode 100644
index 00000000..02782d08
Binary files /dev/null and b/statsforecast/docs/models/CrostonSBA_files/figure-markdown_strict/cell-20-output-1.png differ
diff --git a/statsforecast/docs/models/CrostonSBA_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/CrostonSBA_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..56c6ac59
Binary files /dev/null and b/statsforecast/docs/models/CrostonSBA_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/CrostonSBA_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/models/CrostonSBA_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..8b44cf08
Binary files /dev/null and b/statsforecast/docs/models/CrostonSBA_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-10-output-1.png b/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..9e03fe81
Binary files /dev/null and b/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-11-output-1.png b/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..9e03fe81
Binary files /dev/null and b/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-14-output-1.png b/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..75ef63a6
Binary files /dev/null and b/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-21-output-1.png b/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-21-output-1.png
new file mode 100644
index 00000000..a350c477
Binary files /dev/null and b/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-21-output-1.png differ
diff --git a/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-24-output-1.png b/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-24-output-1.png
new file mode 100644
index 00000000..a952a53e
Binary files /dev/null and b/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-24-output-1.png differ
diff --git a/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-28-output-1.png b/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-28-output-1.png
new file mode 100644
index 00000000..a0813688
Binary files /dev/null and b/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-28-output-1.png differ
diff --git a/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..5ed99363
Binary files /dev/null and b/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..cdc1d0d5
Binary files /dev/null and b/statsforecast/docs/models/DynamicOptimizedTheta_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-10-output-1.png b/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..5148a3b6
Binary files /dev/null and b/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-11-output-1.png b/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..5148a3b6
Binary files /dev/null and b/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-14-output-1.png b/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..6447da50
Binary files /dev/null and b/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-21-output-1.png b/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-21-output-1.png
new file mode 100644
index 00000000..502584a5
Binary files /dev/null and b/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-21-output-1.png differ
diff --git a/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-24-output-1.png b/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-24-output-1.png
new file mode 100644
index 00000000..5dc64123
Binary files /dev/null and b/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-24-output-1.png differ
diff --git a/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-28-output-1.png b/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-28-output-1.png
new file mode 100644
index 00000000..a0a9da74
Binary files /dev/null and b/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-28-output-1.png differ
diff --git a/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..9d372ebe
Binary files /dev/null and b/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..88cc0c72
Binary files /dev/null and b/statsforecast/docs/models/DynamicStandardTheta_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-12-output-2.png b/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-12-output-2.png
new file mode 100644
index 00000000..3f607d5a
Binary files /dev/null and b/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-12-output-2.png differ
diff --git a/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-14-output-1.png b/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..ad3ebd17
Binary files /dev/null and b/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-16-output-1.png b/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-16-output-1.png
new file mode 100644
index 00000000..459ee0d6
Binary files /dev/null and b/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-16-output-1.png differ
diff --git a/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-34-output-1.png b/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-34-output-1.png
new file mode 100644
index 00000000..54359f17
Binary files /dev/null and b/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-34-output-1.png differ
diff --git a/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-39-output-1.png b/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-39-output-1.png
new file mode 100644
index 00000000..f3ea3195
Binary files /dev/null and b/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-39-output-1.png differ
diff --git a/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-42-output-1.png b/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-42-output-1.png
new file mode 100644
index 00000000..52a50a70
Binary files /dev/null and b/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-42-output-1.png differ
diff --git a/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..c8108ae2
Binary files /dev/null and b/statsforecast/docs/models/GARCH_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-11-output-1.png b/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..ab04c569
Binary files /dev/null and b/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-12-output-1.png b/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..f469aa2e
Binary files /dev/null and b/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-13-output-1.png b/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-13-output-1.png
new file mode 100644
index 00000000..f469aa2e
Binary files /dev/null and b/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-13-output-1.png differ
diff --git a/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-16-output-1.png b/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-16-output-1.png
new file mode 100644
index 00000000..b4fe5984
Binary files /dev/null and b/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-16-output-1.png differ
diff --git a/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-23-output-1.png b/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-23-output-1.png
new file mode 100644
index 00000000..d1012767
Binary files /dev/null and b/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-23-output-1.png differ
diff --git a/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-26-output-1.png b/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-26-output-1.png
new file mode 100644
index 00000000..fc3a2cd7
Binary files /dev/null and b/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-26-output-1.png differ
diff --git a/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-28-output-1.png b/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-28-output-1.png
new file mode 100644
index 00000000..6b35a4b9
Binary files /dev/null and b/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-28-output-1.png differ
diff --git a/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-31-output-1.png b/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-31-output-1.png
new file mode 100644
index 00000000..5cfa29a4
Binary files /dev/null and b/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-31-output-1.png differ
diff --git a/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..626950b7
Binary files /dev/null and b/statsforecast/docs/models/HoltWinters_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-11-output-1.png b/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..ab04c569
Binary files /dev/null and b/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-12-output-1.png b/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..43e34ade
Binary files /dev/null and b/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-13-output-1.png b/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-13-output-1.png
new file mode 100644
index 00000000..43e34ade
Binary files /dev/null and b/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-13-output-1.png differ
diff --git a/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-16-output-1.png b/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-16-output-1.png
new file mode 100644
index 00000000..7886a277
Binary files /dev/null and b/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-16-output-1.png differ
diff --git a/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-23-output-1.png b/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-23-output-1.png
new file mode 100644
index 00000000..52f2e260
Binary files /dev/null and b/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-23-output-1.png differ
diff --git a/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-26-output-1.png b/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-26-output-1.png
new file mode 100644
index 00000000..fc3a2cd7
Binary files /dev/null and b/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-26-output-1.png differ
diff --git a/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-28-output-1.png b/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-28-output-1.png
new file mode 100644
index 00000000..cee51273
Binary files /dev/null and b/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-28-output-1.png differ
diff --git a/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-31-output-1.png b/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-31-output-1.png
new file mode 100644
index 00000000..2de2d9df
Binary files /dev/null and b/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-31-output-1.png differ
diff --git a/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..626950b7
Binary files /dev/null and b/statsforecast/docs/models/Holt_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/IMAPA_files/figure-markdown_strict/cell-11-output-2.png b/statsforecast/docs/models/IMAPA_files/figure-markdown_strict/cell-11-output-2.png
new file mode 100644
index 00000000..7f39609e
Binary files /dev/null and b/statsforecast/docs/models/IMAPA_files/figure-markdown_strict/cell-11-output-2.png differ
diff --git a/statsforecast/docs/models/IMAPA_files/figure-markdown_strict/cell-14-output-1.png b/statsforecast/docs/models/IMAPA_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..c68e1388
Binary files /dev/null and b/statsforecast/docs/models/IMAPA_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/statsforecast/docs/models/IMAPA_files/figure-markdown_strict/cell-21-output-1.png b/statsforecast/docs/models/IMAPA_files/figure-markdown_strict/cell-21-output-1.png
new file mode 100644
index 00000000..0a98d18d
Binary files /dev/null and b/statsforecast/docs/models/IMAPA_files/figure-markdown_strict/cell-21-output-1.png differ
diff --git a/statsforecast/docs/models/IMAPA_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/IMAPA_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..56c6ac59
Binary files /dev/null and b/statsforecast/docs/models/IMAPA_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/IMAPA_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/models/IMAPA_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..8b44cf08
Binary files /dev/null and b/statsforecast/docs/models/IMAPA_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-10-output-1.png b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..7c9e0c95
Binary files /dev/null and b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-11-output-1.png b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..73f9a344
Binary files /dev/null and b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-12-output-1.png b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..9c075da8
Binary files /dev/null and b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-13-output-1.png b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-13-output-1.png
new file mode 100644
index 00000000..e4c662ff
Binary files /dev/null and b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-13-output-1.png differ
diff --git a/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-14-output-1.png b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..7c9e0c95
Binary files /dev/null and b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-4-output-1.png b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-4-output-1.png
new file mode 100644
index 00000000..e4c662ff
Binary files /dev/null and b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-4-output-1.png differ
diff --git a/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-5-output-1.png b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-5-output-1.png
new file mode 100644
index 00000000..e4c662ff
Binary files /dev/null and b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-5-output-1.png differ
diff --git a/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-6-output-1.png b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..940c6cb8
Binary files /dev/null and b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-7-output-1.png b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..940c6cb8
Binary files /dev/null and b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..bb78c468
Binary files /dev/null and b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..cdff667b
Binary files /dev/null and b/statsforecast/docs/models/MFLES_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-14-output-1.png b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..3d95c469
Binary files /dev/null and b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-20-output-1.png b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-20-output-1.png
new file mode 100644
index 00000000..c299df95
Binary files /dev/null and b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-20-output-1.png differ
diff --git a/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-23-output-1.png b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-23-output-1.png
new file mode 100644
index 00000000..fc3a2cd7
Binary files /dev/null and b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-23-output-1.png differ
diff --git a/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-25-output-1.png b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-25-output-1.png
new file mode 100644
index 00000000..3c78b7fe
Binary files /dev/null and b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-25-output-1.png differ
diff --git a/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-28-output-1.png b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-28-output-1.png
new file mode 100644
index 00000000..2d866903
Binary files /dev/null and b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-28-output-1.png differ
diff --git a/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-31-output-1.png b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-31-output-1.png
new file mode 100644
index 00000000..070c29c1
Binary files /dev/null and b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-31-output-1.png differ
diff --git a/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-31-output-2.png b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-31-output-2.png
new file mode 100644
index 00000000..6a820d41
Binary files /dev/null and b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-31-output-2.png differ
diff --git a/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-31-output-3.png b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-31-output-3.png
new file mode 100644
index 00000000..29d8a929
Binary files /dev/null and b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-31-output-3.png differ
diff --git a/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-31-output-4.png b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-31-output-4.png
new file mode 100644
index 00000000..a8b92720
Binary files /dev/null and b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-31-output-4.png differ
diff --git a/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-31-output-5.png b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-31-output-5.png
new file mode 100644
index 00000000..e3fc47bf
Binary files /dev/null and b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-31-output-5.png differ
diff --git a/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..626950b7
Binary files /dev/null and b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..d500f374
Binary files /dev/null and b/statsforecast/docs/models/MultipleSeasonalTrend_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-10-output-1.png b/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..5148a3b6
Binary files /dev/null and b/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-11-output-1.png b/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..5148a3b6
Binary files /dev/null and b/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-20-output-1.png b/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-20-output-1.png
new file mode 100644
index 00000000..e358d316
Binary files /dev/null and b/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-20-output-1.png differ
diff --git a/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-23-output-1.png b/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-23-output-1.png
new file mode 100644
index 00000000..5dc64123
Binary files /dev/null and b/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-23-output-1.png differ
diff --git a/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-25-output-1.png b/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-25-output-1.png
new file mode 100644
index 00000000..5b688730
Binary files /dev/null and b/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-25-output-1.png differ
diff --git a/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-28-output-1.png b/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-28-output-1.png
new file mode 100644
index 00000000..9d05bf4a
Binary files /dev/null and b/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-28-output-1.png differ
diff --git a/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..9d372ebe
Binary files /dev/null and b/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..88cc0c72
Binary files /dev/null and b/statsforecast/docs/models/OptimizedTheta_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/models/SeasonalExponentialSmoothingOptimized_files/figure-markdown_strict/cell-11-output-1.png b/statsforecast/docs/models/SeasonalExponentialSmoothingOptimized_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..ab04c569
Binary files /dev/null and b/statsforecast/docs/models/SeasonalExponentialSmoothingOptimized_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/statsforecast/docs/models/SeasonalExponentialSmoothingOptimized_files/figure-markdown_strict/cell-12-output-1.png b/statsforecast/docs/models/SeasonalExponentialSmoothingOptimized_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..43e34ade
Binary files /dev/null and b/statsforecast/docs/models/SeasonalExponentialSmoothingOptimized_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/statsforecast/docs/models/SeasonalExponentialSmoothingOptimized_files/figure-markdown_strict/cell-13-output-1.png b/statsforecast/docs/models/SeasonalExponentialSmoothingOptimized_files/figure-markdown_strict/cell-13-output-1.png
new file mode 100644
index 00000000..43e34ade
Binary files /dev/null and b/statsforecast/docs/models/SeasonalExponentialSmoothingOptimized_files/figure-markdown_strict/cell-13-output-1.png differ
diff --git a/statsforecast/docs/models/SeasonalExponentialSmoothingOptimized_files/figure-markdown_strict/cell-22-output-1.png b/statsforecast/docs/models/SeasonalExponentialSmoothingOptimized_files/figure-markdown_strict/cell-22-output-1.png
new file mode 100644
index 00000000..0304ac89
Binary files /dev/null and b/statsforecast/docs/models/SeasonalExponentialSmoothingOptimized_files/figure-markdown_strict/cell-22-output-1.png differ
diff --git a/statsforecast/docs/models/SeasonalExponentialSmoothingOptimized_files/figure-markdown_strict/cell-25-output-1.png b/statsforecast/docs/models/SeasonalExponentialSmoothingOptimized_files/figure-markdown_strict/cell-25-output-1.png
new file mode 100644
index 00000000..ba26fa43
Binary files /dev/null and b/statsforecast/docs/models/SeasonalExponentialSmoothingOptimized_files/figure-markdown_strict/cell-25-output-1.png differ
diff --git a/statsforecast/docs/models/SeasonalExponentialSmoothingOptimized_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/SeasonalExponentialSmoothingOptimized_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..626950b7
Binary files /dev/null and b/statsforecast/docs/models/SeasonalExponentialSmoothingOptimized_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-11-output-1.png b/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..ab04c569
Binary files /dev/null and b/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-12-output-1.png b/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..43e34ade
Binary files /dev/null and b/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-13-output-1.png b/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-13-output-1.png
new file mode 100644
index 00000000..43e34ade
Binary files /dev/null and b/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-13-output-1.png differ
diff --git a/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-16-output-1.png b/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-16-output-1.png
new file mode 100644
index 00000000..7886a277
Binary files /dev/null and b/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-16-output-1.png differ
diff --git a/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-23-output-1.png b/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-23-output-1.png
new file mode 100644
index 00000000..45a3f970
Binary files /dev/null and b/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-23-output-1.png differ
diff --git a/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-26-output-1.png b/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-26-output-1.png
new file mode 100644
index 00000000..71641c21
Binary files /dev/null and b/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-26-output-1.png differ
diff --git a/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..626950b7
Binary files /dev/null and b/statsforecast/docs/models/SeasonalExponentialSmoothing_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/SimpleExponentialOptimized_files/figure-markdown_strict/cell-18-output-1.png b/statsforecast/docs/models/SimpleExponentialOptimized_files/figure-markdown_strict/cell-18-output-1.png
new file mode 100644
index 00000000..af199cd9
Binary files /dev/null and b/statsforecast/docs/models/SimpleExponentialOptimized_files/figure-markdown_strict/cell-18-output-1.png differ
diff --git a/statsforecast/docs/models/SimpleExponentialOptimized_files/figure-markdown_strict/cell-22-output-1.png b/statsforecast/docs/models/SimpleExponentialOptimized_files/figure-markdown_strict/cell-22-output-1.png
new file mode 100644
index 00000000..15286826
Binary files /dev/null and b/statsforecast/docs/models/SimpleExponentialOptimized_files/figure-markdown_strict/cell-22-output-1.png differ
diff --git a/statsforecast/docs/models/SimpleExponentialOptimized_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/SimpleExponentialOptimized_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..626950b7
Binary files /dev/null and b/statsforecast/docs/models/SimpleExponentialOptimized_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/SimpleExponentialOptimized_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/models/SimpleExponentialOptimized_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..ab04c569
Binary files /dev/null and b/statsforecast/docs/models/SimpleExponentialOptimized_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/models/SimpleExponentialSmoothing_files/figure-markdown_strict/cell-18-output-1.png b/statsforecast/docs/models/SimpleExponentialSmoothing_files/figure-markdown_strict/cell-18-output-1.png
new file mode 100644
index 00000000..8e6392f2
Binary files /dev/null and b/statsforecast/docs/models/SimpleExponentialSmoothing_files/figure-markdown_strict/cell-18-output-1.png differ
diff --git a/statsforecast/docs/models/SimpleExponentialSmoothing_files/figure-markdown_strict/cell-22-output-1.png b/statsforecast/docs/models/SimpleExponentialSmoothing_files/figure-markdown_strict/cell-22-output-1.png
new file mode 100644
index 00000000..01ff1f04
Binary files /dev/null and b/statsforecast/docs/models/SimpleExponentialSmoothing_files/figure-markdown_strict/cell-22-output-1.png differ
diff --git a/statsforecast/docs/models/SimpleExponentialSmoothing_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/SimpleExponentialSmoothing_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..626950b7
Binary files /dev/null and b/statsforecast/docs/models/SimpleExponentialSmoothing_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/SimpleExponentialSmoothing_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/models/SimpleExponentialSmoothing_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..ab04c569
Binary files /dev/null and b/statsforecast/docs/models/SimpleExponentialSmoothing_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-10-output-1.png b/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-10-output-1.png
new file mode 100644
index 00000000..5148a3b6
Binary files /dev/null and b/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-10-output-1.png differ
diff --git a/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-11-output-1.png b/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..5148a3b6
Binary files /dev/null and b/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-20-output-1.png b/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-20-output-1.png
new file mode 100644
index 00000000..361f1d4a
Binary files /dev/null and b/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-20-output-1.png differ
diff --git a/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-24-output-1.png b/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-24-output-1.png
new file mode 100644
index 00000000..d981db07
Binary files /dev/null and b/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-24-output-1.png differ
diff --git a/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-27-output-1.png b/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-27-output-1.png
new file mode 100644
index 00000000..e93372ca
Binary files /dev/null and b/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-27-output-1.png differ
diff --git a/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..9d372ebe
Binary files /dev/null and b/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..88cc0c72
Binary files /dev/null and b/statsforecast/docs/models/StandardTheta_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/models/TSB_files/figure-markdown_strict/cell-12-output-2.png b/statsforecast/docs/models/TSB_files/figure-markdown_strict/cell-12-output-2.png
new file mode 100644
index 00000000..7f39609e
Binary files /dev/null and b/statsforecast/docs/models/TSB_files/figure-markdown_strict/cell-12-output-2.png differ
diff --git a/statsforecast/docs/models/TSB_files/figure-markdown_strict/cell-21-output-1.png b/statsforecast/docs/models/TSB_files/figure-markdown_strict/cell-21-output-1.png
new file mode 100644
index 00000000..f4ba3f61
Binary files /dev/null and b/statsforecast/docs/models/TSB_files/figure-markdown_strict/cell-21-output-1.png differ
diff --git a/statsforecast/docs/models/TSB_files/figure-markdown_strict/cell-8-output-1.png b/statsforecast/docs/models/TSB_files/figure-markdown_strict/cell-8-output-1.png
new file mode 100644
index 00000000..56c6ac59
Binary files /dev/null and b/statsforecast/docs/models/TSB_files/figure-markdown_strict/cell-8-output-1.png differ
diff --git a/statsforecast/docs/models/TSB_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/models/TSB_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..8b44cf08
Binary files /dev/null and b/statsforecast/docs/models/TSB_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/models/adida.html.mdx b/statsforecast/docs/models/adida.html.mdx
new file mode 100644
index 00000000..dd032950
--- /dev/null
+++ b/statsforecast/docs/models/adida.html.mdx
@@ -0,0 +1,762 @@
+---
+title: ADIDA Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `ADIDA Model` with `Statsforecast`.
+
+In this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation`.
+
+The text in this article is largely taken from: 1. [Changquan Huang •
+Alla Petukhina. Springer series (2022). Applied Time Series Analysis and
+Forecasting with
+Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2) 2.
+Ivan Svetunkov. [Forecasting and Analytics with the Augmented Dynamic
+Adaptive Model (ADAM)](https://openforecast.org/adam/) 3. [James D.
+Hamilton. Time Series Analysis Princeton University Press, Princeton,
+New Jersey, 1st Edition,
+1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [ADIDA Model](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of ADIDA with StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+The Aggregate-Disaggregate Intermittent Demand Approach (ADIDA) is a
+forecasting method that is used to predict the demand for products that
+exhibit intermittent demand patterns. Intermittent demand patterns are
+characterized by a large number of zero observations, which can make
+forecasting challenging.
+
+The ADIDA method uses temporal aggregation to reduce the number of zero
+observations and mitigate the effect of the variance observed in the
+intervals. The method uses equally sized time buckets to perform
+non-overlapping temporal aggregation and predict the demand over a
+pre-specified lead time. The time bucket is set equal to the mean
+inter-demand interval, which is the average time between two consecutive
+non-zero observations.
+
+The method uses the Simple Exponential Smoothing (SES) technique to
+obtain the forecasts. SES is a popular time series forecasting technique
+that is commonly used for its simplicity and effectiveness in producing
+accurate forecasts.
+
+The ADIDA method has several advantages. It is easy to implement and can
+be used for a wide range of intermittent demand patterns. The method
+also provides accurate forecasts and can be used to predict the demand
+over a pre-specified lead time.
+
+However, the ADIDA method has some limitations. The method assumes that
+the time buckets are equally sized, which may not be the case for all
+intermittent demand patterns. Additionally, the method may not be
+suitable for time series data with complex patterns or trends.
+
+Overall, the ADIDA method is a useful forecasting technique for
+intermittent demand patterns that can help mitigate the effect of zero
+observations and produce accurate demand forecasts.
+
+## ADIDA Model
+
+### What is intermittent demand?
+
+Intermittent demand is a demand pattern characterized by the irregular
+and sporadic occurrence of events or sales. In other words, it refers to
+situations in which the demand for a product or service occurs
+intermittently, with periods of time in which there are no sales or
+significant events.
+
+Intermittent demand differs from constant or regular demand, where sales
+occur in a predictable and consistent manner over time. In contrast, in
+intermittent demand, periods without sales may be long and there may not
+be a regular sequence of events.
+
+This type of demand can occur in different industries and contexts, such
+as low consumption products, seasonal products, high variability
+products, products with short life cycles, or in situations where demand
+depends on specific events or external factors.
+
+Intermittent demand can pose challenges in forecasting and inventory
+management, as it is difficult to predict when sales will occur and in
+what quantity. Methods like the Croston model, which I mentioned
+earlier, are used to address intermittent demand and generate more
+accurate and appropriate forecasts for this type of demand pattern.
+
+### Problem with intermittent demand
+
+Intermittent demand can present various challenges and issues in
+inventory management and demand forecasting. Some of the common problems
+associated with intermittent demand are as follows:
+
+1. Unpredictable variability: Intermittent demand can have
+ unpredictable variability, making planning and forecasting
+ difficult. Demand patterns can be irregular and fluctuate
+ dramatically between periods with sales and periods without sales.
+
+2. Low frequency of sales: Intermittent demand is characterized by long
+ periods without sales. This can lead to inventory management
+ difficulties, as it is necessary to hold enough stock to meet demand
+ when it occurs, while avoiding excess inventory during non-sales
+ periods.
+
+3. Forecast error: Forecasting intermittent demand can be more
+ difficult to pin down than constant demand. Traditional forecast
+ models may not be adequate to capture the variability and lack of
+ patterns in intermittent demand, which can lead to significant
+ errors in estimates of future demand.
+
+4. Impact on the supply chain: Intermittent demand can affect the
+ efficiency of the supply chain and create difficulties in production
+ planning, supplier management and logistics. Lead times and
+ inventory levels must be adjusted to meet unpredictable demand.
+
+5. Operating costs: Managing inventory in situations of intermittent
+ demand can increase operating costs. Maintaining adequate inventory
+ during non-sales periods and managing stock levels may require
+ additional investments in storage and logistics.
+
+To address these issues, specific approaches to intermittent demand
+management are used, such as specialized forecasting models, product
+classification techniques, and tailored inventory strategies. These
+solutions seek to minimize the impacts of variability and lack of
+patterns in intermittent demand, optimizing inventory management and
+improving supply chain efficiency.
+
+### ADIDA Model
+
+The ADIDA model is based on the Simple Exponential Smoothing (SES)
+method and uses temporal aggregation to handle the problem of
+intermittent demand. The mathematical development of the model can be
+summarized as follows:
+
+Let St be the demand at time $t$, where $t = 1, 2, ..., T$. The mean
+inter-demand interval is denoted as MI, which is the average time
+between two consecutive non-zero demands. The time bucket size is set
+equal to MI.
+
+The demand data is then aggregated into non-overlapping time buckets of
+size MI. Let Bt be the demand in bucket $t$, where
+$t = 1, 2, ..., T/MI$. The aggregated demand data can be represented as:
+
+$$B_t = \sum S_t, for (t-1)*MI + 1 ≤ j ≤ t*MI$$
+
+The SES method is then applied to the aggregated demand data to obtain
+the forecasts. The forecast for bucket $t$ is denoted as $F_t$. The SES
+method involves estimating the level $L_t$ at time t based on the actual
+demand $D_t$ at time t and the estimated level at the previous time
+period, $L_{t-1}$, using the following equation:
+
+$$L_t = \alpha * D_t + (1 - α) * L_{t-1}$$
+
+where $\alpha$ is the smoothing parameter that controls the weight given
+to the current demand value.
+
+The forecast for bucket $t$ is then obtained by using the estimated
+level at the previous time period, $L_{t-1}$, as follows:
+
+$$F_t = L_{t-1}$$
+
+The forecasts are then disaggregated to obtain the demand predictions
+for the original time period. Let $Y_t$ be the demand prediction at time
+$t$. The disaggregation can be performed using the following equation:
+
+$$Y_t = F_t / MI, for (t-1)*MI + 1 ≤ j ≤ t*MI$$
+
+### How can you determine if the ADIDA model is suitable for a specific data set?
+
+To determine if the ADIDA model is suitable for a specific data set, the
+following steps can be followed:
+
+1. Analyze the demand pattern: Examine the demand pattern of the data
+ to determine if it fits an intermittent pattern. Intermittent data
+ is characterized by a high proportion of zeros and sporadic demands
+ in specific periods.
+
+2. Evaluate seasonality: Check if there is a clear seasonality in the
+ data. The ADIDA model assumes that there is no seasonality or that
+ it can be handled by temporal aggregation. If the data show complex
+ seasonality or cannot be handled by temporal aggregation, the ADIDA
+ model may not be suitable.
+
+3. Data requirements: Consider the data requirements of the ADIDA
+ model. The model requires historical demand data and the ability to
+ calculate the mean interval between non-zero demands. Make sure you
+ have enough data to estimate the parameters and that the data is
+ available at a frequency suitable for temporal aggregation.
+
+4. Performance evaluation: Perform a performance evaluation of the
+ ADIDA model on the specific data set. Compare model-generated
+ forecasts with actual demand values and use evaluation metrics such
+ as mean absolute error (MAE) or mean square error (MSE). If the
+ model performs well and produces accurate forecasts on the data set,
+ this is an indication that it is suitable for that data set.
+
+5. Comparison with other models: Compare the performance of the ADIDA
+ model with other forecast models suitable for intermittent data.
+ Consider models like Croston, Syntetos-Boylan Approximation (SBA),
+ or models based on exponential smoothing techniques that have been
+ developed specifically for intermittent data. If the ADIDA model
+ shows similar or better performance than other models, it can be
+ considered suitable.
+
+Remember that the adequacy of the ADIDA model may depend on the specific
+nature of the data and the context of the forecasting problem. It is
+advisable to carry out a thorough analysis and experiment with different
+models to determine the most appropriate approach for the data set in
+question.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf
+from statsmodels.graphics.tsaplots import plot_pacf
+import plotly.graph_objects as go
+plt.style.use('grayscale') # fivethirtyeight grayscale classic
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#008080', # #212946
+ 'axes.facecolor': '#008080',
+ 'savefig.facecolor': '#008080',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#000000', #2A3459
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+
+```python
+import pandas as pd
+
+df = pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/tipos_malarias_choco_colombia.csv", sep=";", usecols=[0,4])
+df = df.dropna()
+df.head()
+```
+
+| | semanas | malaria_falciparum |
+|-----|------------|--------------------|
+| 0 | 2007-12-31 | 50.0 |
+| 1 | 2008-01-07 | 62.0 |
+| 2 | 2008-01-14 | 76.0 |
+| 3 | 2008-01-21 | 64.0 |
+| 4 | 2008-01-28 | 38.0 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|------------|------|-----------|
+| 0 | 2007-12-31 | 50.0 | 1 |
+| 1 | 2008-01-07 | 62.0 | 1 |
+| 2 | 2008-01-14 | 76.0 | 1 |
+| 3 | 2008-01-21 | 64.0 | 1 |
+| 4 | 2008-01-28 | 38.0 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y float64
+unique_id object
+dtype: object
+```
+
+We need to convert the `object` types to datetime and numeric.
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+df["y"] = df["y"].astype(float).astype("int64")
+```
+
+## Explore data with the plot method
+
+Plot a series using the plot method from the StatsForecast class. This
+method prints a random series from the dataset and is useful for basic
+EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### Autocorrelation plots
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=30, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+plot_pacf(df["y"], lags=30, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+plt.show();
+```
+
+
+
+### Decomposition of the time series
+
+How to decompose a time series and why?
+
+In time series analysis to forecast new values, it is very important to
+know past data. More formally, we can say that it is very important to
+know the patterns that values follow over time. There can be many
+reasons that cause our forecast values to fall in the wrong direction.
+Basically, a time series consists of four components. The variation of
+those components causes the change in the pattern of the time series.
+These components are:
+
+- **Level:** This is the primary value that averages over time.
+- **Trend:** The trend is the value that causes increasing or
+ decreasing patterns in a time series.
+- **Seasonality:** This is a cyclical event that occurs in a time
+ series for a short time and causes short-term increasing or
+ decreasing patterns in a time series.
+- **Residual/Noise:** These are the random variations in the time
+ series.
+
+Combining these components over time leads to the formation of a time
+series. Most time series consist of level and noise/residual and trend
+or seasonality are optional values.
+
+If seasonality and trend are part of the time series, then there will be
+effects on the forecast value. As the pattern of the forecasted time
+series may be different from the previous time series.
+
+The combination of the components in time series can be of two types: \*
+Additive \* Multiplicative
+
+### Additive time series
+
+If the components of the time series are added to make the time series.
+Then the time series is called the additive time series. By
+visualization, we can say that the time series is additive if the
+increasing or decreasing pattern of the time series is similar
+throughout the series. The mathematical function of any additive time
+series can be represented by:
+$$y(t) = Level + Trend + Seasonality + Noise$$
+
+### Multiplicative time series
+
+If the components of the time series are multiplicative together, then
+the time series is called a multiplicative time series. For
+visualization, if the time series is having exponential growth or
+decline with time, then the time series can be considered as the
+multiplicative time series. The mathematical function of the
+multiplicative time series can be represented as.
+
+$$y(t) = Level * Trend * seasonality * Noise$$
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+from plotly.subplots import make_subplots
+import plotly.graph_objects as go
+
+def plot_seasonal_decompose(
+ x,
+ model='additive',
+ filt=None,
+ period=None,
+ two_sided=True,
+ extrapolate_trend=0,
+ title="Seasonal Decomposition"):
+
+ result = seasonal_decompose(
+ x, model=model, filt=filt, period=period,
+ two_sided=two_sided, extrapolate_trend=extrapolate_trend)
+ fig = make_subplots(
+ rows=4, cols=1,
+ subplot_titles=["Observed", "Trend", "Seasonal", "Residuals"])
+ for idx, col in enumerate(['observed', 'trend', 'seasonal', 'resid']):
+ fig.add_trace(
+ go.Scatter(x=result.observed.index, y=getattr(result, col), mode='lines'),
+ row=idx+1, col=1,
+ )
+ return fig
+```
+
+
+```python
+plot_seasonal_decompose(
+ df["y"],
+ model="additive",
+ period=52,
+ title="Seasonal Decomposition")
+```
+
+
+
+## Split the data into training and testing
+
+Let’s divide our data into sets 1. Data to train our `ADIDA Model`. 2.
+Data to test our model
+
+For the test data we will use the last 25 week to test and evaluate the
+performance of our model.
+
+```python
+train = df[df.ds\<='2022-07-04']
+test = df[df.ds>'2022-07-04']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((758, 3), (25, 3))
+```
+
+Now let’s plot the training data and the test data.
+
+```python
+sns.lineplot(train,x="ds", y="y", label="Train", linestyle="--",linewidth=2)
+sns.lineplot(test, x="ds", y="y", label="Test", linewidth=2, color="yellow")
+plt.title("Falciparum Malaria");
+plt.show()
+```
+
+
+
+## Implementation of `ADIDA Model` with StatsForecast
+
+To also know more about the parameters of the functions of the
+`ADIDA Model`, they are listed below. For more information, visit the
+[documentation](../../src/core/models.html#adida)
+
+``` text
+alias : str
+ Custom name of the model.
+prediction_intervals : Optional[ConformalIntervals]
+ Information to compute conformal prediction intervals.
+ By default, the model will compute the native prediction
+ intervals.
+```
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import ADIDA
+```
+
+### Instantiating Model
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/) by the
+master, Rob Hyndmann, can be useful for `season_length`.
+
+```python
+season_length = 52 # Hourly data
+horizon = len(test) # number of predictions
+
+# We call the model that we are going to use
+models = [ADIDA()]
+```
+
+We fit the models by instantiating a new `StatsForecast` object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [pandas’
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models,
+ freq='7d',
+ n_jobs=-1)
+```
+
+### Fit the Model
+
+Here, we call the `fit()` method to fit the model.
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[ADIDA])
+```
+
+Let’s see the results of our `ADIDA Model`. We can observe it with the
+following instruction:
+
+```python
+result=sf.fitted_[0,0].model_
+result
+```
+
+``` text
+{'mean': array([336.74736919])}
+```
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `forecast()` method does not store the
+fitted values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 25 week ahead.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min.
+
+```python
+Y_hat = sf.forecast(df=train, h=horizon)
+Y_hat
+```
+
+| | unique_id | ds | ADIDA |
+|-----|-----------|------------|------------|
+| 0 | 1 | 2022-07-11 | 336.747375 |
+| 1 | 1 | 2022-07-18 | 336.747375 |
+| 2 | 1 | 2022-07-25 | 336.747375 |
+| ... | ... | ... | ... |
+| 22 | 1 | 2022-12-12 | 336.747375 |
+| 23 | 1 | 2022-12-19 | 336.747375 |
+| 24 | 1 | 2022-12-26 | 336.747375 |
+
+```python
+sf.plot(train, Y_hat.merge(test))
+```
+
+
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 25 week ahead.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+forecast_df = sf.predict(h=horizon)
+forecast_df.head()
+```
+
+| | unique_id | ds | ADIDA |
+|-----|-----------|------------|------------|
+| 0 | 1 | 2022-07-11 | 336.747375 |
+| 1 | 1 | 2022-07-18 | 336.747375 |
+| 2 | 1 | 2022-07-25 | 336.747375 |
+| 3 | 1 | 2022-08-01 | 336.747375 |
+| 4 | 1 | 2022-08-08 | 336.747375 |
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+img
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=)`, forecasting every second months
+`(step_size=12)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 12 months ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=df,
+ h=horizon,
+ step_size=30,
+ n_windows=5)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` series identifier
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the `n_windows`.
+- `y:` true value
+- `model:` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df
+```
+
+| | unique_id | ds | cutoff | y | ADIDA |
+|-----|-----------|------------|------------|-------|------------|
+| 0 | 1 | 2020-03-23 | 2020-03-16 | 317.0 | 251.901505 |
+| 1 | 1 | 2020-03-30 | 2020-03-16 | 332.0 | 251.901505 |
+| 2 | 1 | 2020-04-06 | 2020-03-16 | 306.0 | 251.901505 |
+| ... | ... | ... | ... | ... | ... |
+| 122 | 1 | 2022-12-12 | 2022-07-04 | 151.0 | 336.747375 |
+| 123 | 1 | 2022-12-19 | 2022-07-04 | 97.0 | 336.747375 |
+| 124 | 1 | 2022-12-26 | 2022-07-04 | 42.0 | 336.747375 |
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=season_length), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | ADIDA |
+|-----|-----------|--------|------------|
+| 0 | 1 | mae | 114.527585 |
+| 1 | 1 | mape | 0.820029 |
+| 2 | 1 | mase | 0.874115 |
+| 3 | 1 | rmse | 129.749320 |
+| 4 | 1 | smape | 0.221878 |
+
+## References
+
+1. [Changquan Huang • Alla Petukhina. Springer series (2022). Applied
+ Time Series Analysis and Forecasting with
+ Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2)
+2. Ivan Svetunkov. [Forecasting and Analytics with the Augmented
+ Dynamic Adaptive Model (ADAM)](https://openforecast.org/adam/)
+3. [James D. Hamilton. Time Series Analysis Princeton University Press,
+ Princeton, New Jersey, 1st Edition,
+ 1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Nixtla ADIDA API](../../src/core/models.html#adida)
+5. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+6. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+7. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/models/arch.html.mdx b/statsforecast/docs/models/arch.html.mdx
new file mode 100644
index 00000000..2f10f364
--- /dev/null
+++ b/statsforecast/docs/models/arch.html.mdx
@@ -0,0 +1,1081 @@
+---
+title: ARCH Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `ARCH Model` with `Statsforecast`.
+
+In this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation`.
+
+The text in this article is largely taken from [Changquan Huang • Alla
+Petukhina. Springer series (2022). Applied Time Series Analysis and
+Forecasting with
+Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2)
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [ARCH Models](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of ARCH with StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+Financial time series analysis has been one of the hottest research
+topics in the recent decades. In this guide, we illustrate the stylized
+facts of financial time series by real financial data. To characterize
+these facts, new models different from the Box- Jenkins ones are needed.
+And for this reason, ARCH models were firstly proposed by R. F. Engle in
+1982 and have been extended by a great number of scholars since then. We
+also demonstrate how to use Python and its libraries to implement
+`ARCH`.
+
+As we have known, there are lot of time series that possess the ARCH
+effect, that is, although the (modeling residual) series is white noise,
+its squared series may be autocorrelated. What is more, in practice, a
+large number of financial time series are found having this property so
+that the ARCH effect has become one of the stylized facts from financial
+time series.
+
+### Stylized Facts of Financial Time Series
+
+Now we briefly list and describe several important stylized facts
+(features) of financial return series:
+
+- **Fat (heavy) tails:** The distribution density function of returns
+ often has fatter (heavier) tails than the tails of the corresponding
+ normal distribution density.
+
+- **ARCH effect:** Although the return series can often be seen as a
+ white noise, its squared (and absolute) series may usually be
+ autocorrelated, and these autocorrelations are hardly negative.
+
+- **Volatility clustering:** Large changes in returns tend to cluster
+ in time, and small changes tend to be followed by small changes.
+
+- **Asymmetry:** As we have know , the distribution of asset returns
+ is slightly negatively skewed. One possible explanation could be
+ that traders react more strongly to unfavorable information than
+ favorable information.
+
+## Definition of ARCH Models
+
+Specifically, we give the definition of the ARCH model as follows.
+
+**Definition 1.** An $\text{ARCH(p)}$ model with order $p≥1$ is of the
+form
+
+$$
+\begin{equation}
+ \left\{
+ \begin{array}{ll}
+ X_t =\sigma_t \varepsilon_t \\
+ \sigma_{t}^2 =\omega+ \alpha_1 X_{t-1}^2 + \alpha_2 X_{t-2}^2 + \cdots+ \alpha_p X_{t-p}^2 \\
+ \end{array}
+ \right.
+\end{equation}
+$$
+
+where $\omega ≥ 0, \alpha_i ≥ 0$, and $\alpha_p > 0$ are constants,
+$\varepsilon_t \sim iid(0, 1)$, and $\varepsilon_t$ is independent of
+$\{X_k;k ≤ t − 1 \}$. A stochastic process $X_t$ is called an $ARCH(p)$
+process if it satisfies Eq. (1).
+
+By Definition 1, $\sigma_{t}^2$ (and $\sigma_t$ ) is independent of
+$\varepsilon_t$ . Besides, usually it is further assumed that
+$\varepsilon_t \sim N(0, 1)$. Sometimes, however, we need to further
+suppose that $\varepsilon_t$ follows a standardized (skew) Student’s T
+distribution or a generalized error distribution in order to capture
+more features of a financial time series.
+
+Let $\mathscr{F}_s$ denote the information set generated by
+$\{X_k;k ≤ s \}$, namely, the sigma field $\sigma(X_k;k ≤ s)$. It is
+easy to see that $\mathscr{F}_s$ is independent of $\varepsilon_t$ for
+any $s 0$, in light of the properties of the conditional
+mathematical expectation and by (2), we have that
+
+$$E(X_{t+h} X_t) = E(E(X_{t+h} X_t|\mathscr{F}_{t+h-1})) = E(X_t E(X_{t+h}|\mathscr{F}_{t+h-1})) = 0.$$
+
+In conclusion, if $0 < \alpha_1 < 1$, we have that:
+
+- Any $\text{ARCH}(1)$ process $\{X_t \}$ defined by Eqs.(3) follows a
+ white noise $WN(0, \omega/(1 − \alpha_1))$ .
+
+- Since $X_{t}^2$ is an $\text{AR}(1)$ process defined by (4),
+ $\text{Corr}(X_{t}^2,X_{t+h}^2) = \alpha_{1}^{|h|} > 0$, which
+ reveals the ARCH effect.
+
+- It is clear that $E(\eta_t|\mathscr{F}_s)=0$ for any $t>s$,and with
+ Eq.(4),for any $k>1$:
+ $$Var(X_{t+k} |\mathscr{F}_t ) = E(X_{t+K}^2 |\mathscr{F}_t)$$
+ $$= E(\omega + \alpha_1 X_{t+k-1}+ \eta_{t+k}|\mathscr{F}_t )$$
+ $$= \omega + \alpha_1 Var(X_{t+k−1}|\mathscr{F}_t),$$
+
+which reflects the volatility clustering, that is, large (small)
+volatility is followed by large (small) one.
+
+In addition, we are able to prove that Xt defined by Eq. (3) has heavier
+tails than the corresponding normal distribution. At last, note that
+these properties of the ARCH(1) model can be generalized to ARCH(p)
+models.
+
+### Advantages and disadvantages of the Autoregressive Conditional Heteroskedasticity (ARCH) model:
+
+| Advantages | Disadvantages |
+|--------------------------------|-----------------------------------------|
+| \- The ARCH model is useful for modeling volatility in financial time series, which is important for investment decision making and risk management. | \- The ARCH model assumes that the forecast errors are independent and identically distributed, which may not be realistic in some cases. |
+| \- The ARCH model takes heteroscedasticity into account, which means that it can model time series with variances that change over time. | \- The ARCH model can be difficult to fit to data with many parameters, which may require large amounts of data or advanced estimation techniques. |
+| \- The ARCH model is relatively easy to use and can be implemented with standard econometrics software. | \- The ARCH model does not take into account the possible relationship between the mean and the variance of the time series, which may be important in some cases. |
+
+Note:
+
+The ARCH model is a useful tool for modeling volatility in financial
+time series, but like any econometric model, it has limitations and
+should be used with caution depending on the specific characteristics of
+the data being modeled.
+
+### Autoregressive Conditional Heteroskedasticity (ARCH) Applications
+
+- **Finance** - The ARCH model is widely used in finance to model
+ volatility in financial time series, such as stock prices, exchange
+ rates, interest rates, etc.
+
+- **Economics** - The ARCH model can be used to model volatility in
+ economic data, such as GDP, inflation, unemployment, among others.
+
+- **Engineering** - The ARCH model can be used in engineering to model
+ volatility in data related to energy, climate, pollution, industrial
+ production, among others.
+
+- **Social Sciences** - The ARCH model can be used in the social
+ sciences to model volatility in data related to demography, health,
+ education, among others.
+
+- **Biology** - The ARCH model can be used in biology to model
+ volatility in data related to evolution, genetics, epidemiology,
+ among others.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import matplotlib.pyplot as plt
+from statsmodels.graphics.tsaplots import plot_acf
+from statsmodels.graphics.tsaplots import plot_pacf
+plt.style.use('fivethirtyeight')
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#212946',
+ 'axes.facecolor': '#212946',
+ 'savefig.facecolor':'#212946',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#2A3459',
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+### Read Data
+
+Let’s pull the S&P500 stock data from the Yahoo Finance site.
+
+```python
+import datetime
+
+import pandas as pd
+import time
+import yfinance as yf
+
+ticker = '^GSPC'
+period1 = datetime.datetime(2015, 1, 1)
+period2 = datetime.datetime(2023, 9, 22)
+interval = '1d' # 1d, 1m
+
+SP_500 = yf.download(ticker, start=period1, end=period2, interval=interval, progress=False)
+SP_500 = SP_500.reset_index()
+
+SP_500.head()
+```
+
+| Price | Date | Adj Close | Close | High | Low | Open | Volume |
+|--------|---------------------------|-------------|-------------|-------------|-------------|-------------|------------|
+| Ticker | | ^GSPC | ^GSPC | ^GSPC | ^GSPC | ^GSPC | ^GSPC |
+| 0 | 2015-01-02 00:00:00+00:00 | 2058.199951 | 2058.199951 | 2072.360107 | 2046.040039 | 2058.899902 | 2708700000 |
+| 1 | 2015-01-05 00:00:00+00:00 | 2020.579956 | 2020.579956 | 2054.439941 | 2017.339966 | 2054.439941 | 3799120000 |
+| 2 | 2015-01-06 00:00:00+00:00 | 2002.609985 | 2002.609985 | 2030.250000 | 1992.439941 | 2022.150024 | 4460110000 |
+| 3 | 2015-01-07 00:00:00+00:00 | 2025.900024 | 2025.900024 | 2029.609985 | 2005.550049 | 2005.550049 | 3805480000 |
+| 4 | 2015-01-08 00:00:00+00:00 | 2062.139893 | 2062.139893 | 2064.080078 | 2030.609985 | 2030.609985 | 3934010000 |
+
+```python
+df=SP_500[["Date","Close"]].copy()
+```
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|---------------------------|-------------|-----------|
+| 0 | 2015-01-02 00:00:00+00:00 | 2058.199951 | 1 |
+| 1 | 2015-01-05 00:00:00+00:00 | 2020.579956 | 1 |
+| 2 | 2015-01-06 00:00:00+00:00 | 2002.609985 | 1 |
+| 3 | 2015-01-07 00:00:00+00:00 | 2025.900024 | 1 |
+| 4 | 2015-01-08 00:00:00+00:00 | 2062.139893 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds datetime64[ns]
+y float64
+unique_id object
+dtype: object
+```
+
+## Explore data with the plot method
+
+Plot a series using the plot method from the StatsForecast class. This
+method prints a random series from the dataset and is useful for basic
+EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### The Augmented Dickey-Fuller Test
+
+An Augmented Dickey-Fuller (ADF) test is a type of statistical test that
+determines whether a unit root is present in time series data. Unit
+roots can cause unpredictable results in time series analysis. A null
+hypothesis is formed in the unit root test to determine how strongly
+time series data is affected by a trend. By accepting the null
+hypothesis, we accept the evidence that the time series data is not
+stationary. By rejecting the null hypothesis or accepting the
+alternative hypothesis, we accept the evidence that the time series data
+is generated by a stationary process. This process is also known as
+stationary trend. The values of the ADF test statistic are negative.
+Lower ADF values indicate a stronger rejection of the null hypothesis.
+
+Augmented Dickey-Fuller Test is a common statistical test used to test
+whether a given time series is stationary or not. We can achieve this by
+defining the null and alternate hypothesis.
+
+Null Hypothesis: Time Series is non-stationary. It gives a
+time-dependent trend. Alternate Hypothesis: Time Series is stationary.
+In another term, the series doesn’t depend on time.
+
+ADF or t Statistic \< critical values: Reject the null hypothesis, time
+series is stationary. ADF or t Statistic \> critical values: Failed to
+reject the null hypothesis, time series is non-stationary.
+
+Let’s check if our series that we are analyzing is a stationary series.
+Let’s create a function to check, using the `Dickey Fuller` test
+
+```python
+from statsmodels.tsa.stattools import adfuller
+
+def Augmented_Dickey_Fuller_Test_func(series , column_name):
+ print (f'Dickey-Fuller test results for columns: {column_name}')
+ dftest = adfuller(series, autolag='AIC')
+ dfoutput = pd.Series(dftest[0:4], index=['Test Statistic','p-value','No Lags Used','Number of observations used'])
+ for key,value in dftest[4].items():
+ dfoutput['Critical Value (%s)'%key] = value
+ print (dfoutput)
+ if dftest[1] \<= 0.05:
+ print("Conclusion:====>")
+ print("Reject the null hypothesis")
+ print("The data is stationary")
+ else:
+ print("Conclusion:====>")
+ print("The null hypothesis cannot be rejected")
+ print("The data is not stationary")
+```
+
+
+```python
+Augmented_Dickey_Fuller_Test_func(df["y"],'S&P500')
+```
+
+``` text
+Dickey-Fuller test results for columns: S&P500
+Test Statistic -0.814971
+p-value 0.814685
+No Lags Used 10.000000
+ ...
+Critical Value (1%) -3.433341
+Critical Value (5%) -2.862861
+Critical Value (10%) -2.567473
+Length: 7, dtype: float64
+Conclusion:====>
+The null hypothesis cannot be rejected
+The data is not stationary
+```
+
+In the previous result we can see that the `Augmented_Dickey_Fuller`
+test gives us a `p-value` of 0.864700, which tells us that the null
+hypothesis cannot be rejected, and on the other hand the data of our
+series are not stationary.
+
+We need to differentiate our time series, in order to convert the data
+to stationary.
+
+### Return Series
+
+Since the 1970s, the financial industry has been very prosperous with
+advancement of computer and Internet technology. Trade of financial
+products (including various derivatives) generates a huge amount of data
+which form financial time series. For finance, the return on a financial
+product is most interesting, and so our attention focuses on the return
+series. If $P_t$ is the closing price at time t for a certain financial
+product, then the return on this product is
+
+$$X_t = \frac{(P_t − P_{t−1})}{P_{t−1}} ≈ log(P_t ) − log(P_{t−1}).$$
+
+It is return series $\{X_t \}$ that have been much independently
+studied. And important stylized features which are common across many
+instruments, markets, and time periods have been summarized. Note that
+if you purchase the financial product, then it becomes your asset, and
+its returns become your asset returns. Now let us look at the following
+examples.
+
+We can estimate the series of returns using the
+[pandas](https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.pct_change.html),
+`DataFrame.pct_change()` function. The `pct_change()` function has a
+periods parameter whose default value is 1. If you want to calculate a
+30-day return, you must change the value to 30.
+
+```python
+df['return'] = 100 * df["y"].pct_change()
+df.dropna(inplace=True, how='any')
+df.head()
+```
+
+| | ds | y | unique_id | return |
+|-----|---------------------------|-------------|-----------|-----------|
+| 1 | 2015-01-05 00:00:00+00:00 | 2020.579956 | 1 | -1.827811 |
+| 2 | 2015-01-06 00:00:00+00:00 | 2002.609985 | 1 | -0.889347 |
+| 3 | 2015-01-07 00:00:00+00:00 | 2025.900024 | 1 | 1.162984 |
+| 4 | 2015-01-08 00:00:00+00:00 | 2062.139893 | 1 | 1.788828 |
+| 5 | 2015-01-09 00:00:00+00:00 | 2044.810059 | 1 | -0.840381 |
+
+```python
+import plotly.express as px
+fig = px.line(df, x=df["ds"], y="return",title="SP500 Return Chart",template = "plotly_dark")
+fig.show()
+```
+
+
+
+### Creating Squared Returns
+
+```python
+df['sq_return'] = df["return"].mul(df["return"])
+df.head()
+```
+
+| | ds | y | unique_id | return | sq_return |
+|-----|---------------------------|-------------|-----------|-----------|-----------|
+| 1 | 2015-01-05 00:00:00+00:00 | 2020.579956 | 1 | -1.827811 | 3.340891 |
+| 2 | 2015-01-06 00:00:00+00:00 | 2002.609985 | 1 | -0.889347 | 0.790938 |
+| 3 | 2015-01-07 00:00:00+00:00 | 2025.900024 | 1 | 1.162984 | 1.352532 |
+| 4 | 2015-01-08 00:00:00+00:00 | 2062.139893 | 1 | 1.788828 | 3.199906 |
+| 5 | 2015-01-09 00:00:00+00:00 | 2044.810059 | 1 | -0.840381 | 0.706240 |
+
+### Returns vs Squared Returns
+
+```python
+from plotly.subplots import make_subplots
+import plotly.graph_objects as go
+
+fig = make_subplots(rows=1, cols=2)
+
+fig.add_trace(go.Scatter(x=df["ds"], y=df["return"],
+ mode='lines',
+ name='return'),
+row=1, col=1
+)
+
+
+fig.add_trace(go.Scatter(x=df["ds"], y=df["sq_return"],
+ mode='lines',
+ name='sq_return'),
+ row=1, col=2
+)
+
+fig.update_layout(height=600, width=800, title_text="Returns vs Squared Returns", template = "plotly_dark")
+fig.show()
+```
+
+
+
+```python
+from scipy.stats import probplot, moment
+from statsmodels.tsa.stattools import adfuller, q_stat, acf
+import numpy as np
+import seaborn as sns
+
+def plot_correlogram(x, lags=None, title=None):
+ lags = min(10, int(len(x)/5)) if lags is None else lags
+ fig, axes = plt.subplots(nrows=2, ncols=2, figsize=(14, 8))
+ x.plot(ax=axes[0][0], title='Return')
+ x.rolling(21).mean().plot(ax=axes[0][0], c='k', lw=1)
+ q_p = np.max(q_stat(acf(x, nlags=lags), len(x))[1])
+ stats = f'Q-Stat: {np.max(q_p):>8.2f}\nADF: {adfuller(x)[1]:>11.2f}'
+ axes[0][0].text(x=.02, y=.85, s=stats, transform=axes[0][0].transAxes)
+ probplot(x, plot=axes[0][1])
+ mean, var, skew, kurtosis = moment(x, moment=[1, 2, 3, 4])
+ s = f'Mean: {mean:>12.2f}\nSD: {np.sqrt(var):>16.2f}\nSkew: {skew:12.2f}\nKurtosis:{kurtosis:9.2f}'
+ axes[0][1].text(x=.02, y=.75, s=s, transform=axes[0][1].transAxes)
+ plot_acf(x=x, lags=lags, zero=False, ax=axes[1][0])
+ plot_pacf(x, lags=lags, zero=False, ax=axes[1][1])
+ axes[1][0].set_xlabel('Lag')
+ axes[1][1].set_xlabel('Lag')
+ fig.suptitle(title+ f'Dickey-Fuller: {adfuller(x)[1]:>11.2f}', fontsize=14)
+ sns.despine()
+ fig.tight_layout()
+ fig.subplots_adjust(top=.9)
+```
+
+
+```python
+plot_correlogram(df["return"], lags=30, title="Time Series Analysis plot \n")
+```
+
+
+
+### Ljung-Box Test
+
+Ljung-Box is a test for autocorrelation that we can use in tandem with
+our ACF and PACF plots. The Ljung-Box test takes our data, optionally
+either lag values to test, or the largest lag value to consider, and
+whether to compute the Box-Pierce statistic. Ljung-Box and Box-Pierce
+are two similar test statisitcs, Q , that are compared against a
+chi-squared distribution to determine if the series is white noise. We
+might use the Ljung-Box test on the residuals of our model to look for
+autocorrelation, ideally our residuals would be white noise.
+
+- Ho : The data are independently distributed, no autocorrelation.
+- Ha : The data are not independently distributed; they exhibit serial
+ correlation.
+
+The Ljung-Box with the Box-Pierce option will return, for each lag, the
+Ljung-Box test statistic, Ljung-Box p-values, Box-Pierce test statistic,
+and Box-Pierce p-values.
+
+If $p<\alpha (0.05)$ we reject the null hypothesis.
+
+```python
+from statsmodels.stats.diagnostic import acorr_ljungbox
+
+ljung_res = acorr_ljungbox(df["return"], lags= 40, boxpierce=True)
+
+ljung_res.head()
+```
+
+| | lb_stat | lb_pvalue | bp_stat | bp_pvalue |
+|-----|-----------|--------------|-----------|--------------|
+| 1 | 49.222273 | 2.285409e-12 | 49.155183 | 2.364927e-12 |
+| 2 | 62.991348 | 2.097020e-14 | 62.899234 | 2.195861e-14 |
+| 3 | 63.944944 | 8.433622e-14 | 63.850663 | 8.834380e-14 |
+| 4 | 74.343652 | 2.742989e-15 | 74.221024 | 2.911751e-15 |
+| 5 | 80.234862 | 7.494100e-16 | 80.093498 | 8.022242e-16 |
+
+## Split the data into training and testing
+
+Let’s divide our data into sets
+
+1. Data to train our `ARCH` model
+2. Data to test our model
+
+For the test data we will use the last 30 day to test and evaluate the
+performance of our model.
+
+```python
+df=df[["ds","unique_id","return"]]
+df.columns=["ds", "unique_id", "y"]
+```
+
+
+```python
+train = df[df.ds\<='2023-05-24'] # Let's forecast the last 30 days
+test = df[df.ds>'2023-05-24']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((2112, 3), (82, 3))
+```
+
+Now let’s plot the training data and the test data.
+
+```python
+sns.lineplot(train,x="ds", y="y", label="Train")
+sns.lineplot(test, x="ds", y="y", label="Test")
+plt.show()
+```
+
+
+
+## Implementation of ARCH with StatsForecast
+
+To also know more about the parameters of the functions of the
+`ARCH Model`, they are listed below. For more information, visit the
+[documentation](../../src/core/models.html#arch)
+
+``` text
+p : int
+ Number of lagged versions of the series.
+alias : str
+ Custom name of the model.
+prediction_intervals : Optional[ConformalIntervals]
+ Information to compute conformal prediction intervals.
+ By default, the model will compute the native prediction
+ intervals.
+```
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import ARCH
+```
+
+### Building Model
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/) by the
+master, Rob Hyndmann, can be useful.season_length.
+
+```python
+season_length = 7 # Daily data
+horizon = len(test) # number of predictions biasadj=True, include_drift=True,
+
+models = [ARCH(p=2)]
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [pandas’
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models,
+ freq='C', # custom business day frequency
+ )
+```
+
+### Fit the Model
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[ARCH(2)])
+```
+
+Let’s see the results of our ARCH model. We can observe it with the
+following instruction:
+
+```python
+result=sf.fitted_[0,0].model_
+result
+```
+
+``` text
+{'p': 2,
+ 'q': 0,
+ 'coeff': array([0.44321058, 0.34706751, 0.35172097]),
+ 'message': 'Optimization terminated successfully',
+ 'y_vals': array([-1.12220267, -0.73186003]),
+ 'sigma2_vals': array([1.38768694, nan, 1.89278112, ..., 0.76423271, 0.45064684,
+ 0.88037072]),
+ 'fitted': array([ nan, nan, 2.23474807, ..., -1.48033228,
+ 1.10018999, -0.98050166]),
+ 'actual_residuals': array([ nan, nan, -1.07176381, ..., 1.49583575,
+ -2.22239266, 0.24864162])}
+```
+
+Let us now visualize the residuals of our models.
+
+As we can see, the result obtained above has an output in a dictionary,
+to extract each element from the dictionary we are going to use the
+`.get()` function to extract the element and then we are going to save
+it in a `pd.DataFrame()`.
+
+```python
+residual=pd.DataFrame(result.get("actual_residuals"), columns=["residual Model"])
+residual
+```
+
+| | residual Model |
+|------|----------------|
+| 0 | NaN |
+| 1 | NaN |
+| 2 | -1.071764 |
+| ... | ... |
+| 2109 | 1.495836 |
+| 2110 | -2.222393 |
+| 2111 | 0.248642 |
+
+```python
+import scipy.stats as stats
+
+fig, axs = plt.subplots(nrows=2, ncols=2)
+
+# plot[1,1]
+residual.plot(ax=axs[0,0])
+axs[0,0].set_title("Residuals");
+
+# plot
+sns.distplot(residual, ax=axs[0,1]);
+axs[0,1].set_title("Density plot - Residual");
+
+# plot
+stats.probplot(residual["residual Model"], dist="norm", plot=axs[1,0])
+axs[1,0].set_title('Plot Q-Q')
+
+# plot
+plot_acf(residual, lags=35, ax=axs[1,1],color="fuchsia")
+axs[1,1].set_title("Autocorrelation");
+
+plt.show();
+```
+
+
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[90]` means that
+ the model expects the real value to be inside that interval 90% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min.
+
+```python
+Y_hat = sf.forecast(df=train, h=horizon, fitted=True)
+Y_hat
+```
+
+| | unique_id | ds | ARCH(2) |
+|-----|-----------|---------------------------|-----------|
+| 0 | 1 | 2023-05-25 00:00:00+00:00 | 1.681839 |
+| 1 | 1 | 2023-05-26 00:00:00+00:00 | -0.777029 |
+| 2 | 1 | 2023-05-29 00:00:00+00:00 | -0.677962 |
+| ... | ... | ... | ... |
+| 79 | 1 | 2023-09-13 00:00:00+00:00 | 0.695591 |
+| 80 | 1 | 2023-09-14 00:00:00+00:00 | -0.176075 |
+| 81 | 1 | 2023-09-15 00:00:00+00:00 | -0.158605 |
+
+```python
+values=sf.forecast_fitted_values()
+values.head()
+```
+
+| | unique_id | ds | y | ARCH(2) |
+|-----|-----------|---------------------------|-----------|-----------|
+| 0 | 1 | 2015-01-05 00:00:00+00:00 | -1.827811 | NaN |
+| 1 | 1 | 2015-01-06 00:00:00+00:00 | -0.889347 | NaN |
+| 2 | 1 | 2015-01-07 00:00:00+00:00 | 1.162984 | 2.234748 |
+| 3 | 1 | 2015-01-08 00:00:00+00:00 | 1.788828 | -0.667577 |
+| 4 | 1 | 2015-01-09 00:00:00+00:00 | -0.840381 | -0.752438 |
+
+Adding 95% confidence interval with the forecast method
+
+```python
+sf.forecast(df=train, h=horizon, level=[95])
+```
+
+| | unique_id | ds | ARCH(2) | ARCH(2)-lo-95 | ARCH(2)-hi-95 |
+|-----|-----------|---------------------------|-----------|---------------|---------------|
+| 0 | 1 | 2023-05-25 00:00:00+00:00 | 1.681839 | -0.419326 | 3.783003 |
+| 1 | 1 | 2023-05-26 00:00:00+00:00 | -0.777029 | -3.939054 | 2.384996 |
+| 2 | 1 | 2023-05-29 00:00:00+00:00 | -0.677962 | -3.907262 | 2.551338 |
+| ... | ... | ... | ... | ... | ... |
+| 79 | 1 | 2023-09-13 00:00:00+00:00 | 0.695591 | -0.937585 | 2.328766 |
+| 80 | 1 | 2023-09-14 00:00:00+00:00 | -0.176075 | -1.405359 | 1.053210 |
+| 81 | 1 | 2023-09-15 00:00:00+00:00 | -0.158605 | -1.381915 | 1.064705 |
+
+```python
+# Merge the forecasts with the true values
+Y_hat1 = test.merge(Y_hat, how='left', on=['unique_id', 'ds'])
+Y_hat1
+```
+
+| | ds | unique_id | y | ARCH(2) |
+|-----|---------------------------|-----------|-----------|-----------|
+| 0 | 2023-05-25 00:00:00+00:00 | 1 | 0.875758 | 1.681839 |
+| 1 | 2023-05-26 00:00:00+00:00 | 1 | 1.304909 | -0.777029 |
+| 2 | 2023-05-30 00:00:00+00:00 | 1 | 0.001660 | -0.968703 |
+| ... | ... | ... | ... | ... |
+| 79 | 2023-09-19 00:00:00+00:00 | 1 | -0.215101 | NaN |
+| 80 | 2023-09-20 00:00:00+00:00 | 1 | -0.939479 | NaN |
+| 81 | 2023-09-21 00:00:00+00:00 | 1 | -1.640093 | NaN |
+
+```python
+# Merge the forecasts with the true values
+
+fig, ax = plt.subplots(1, 1)
+plot_df = pd.concat([train, Y_hat1]).set_index('ds')
+plot_df[['y', "ARCH(2)"]].plot(ax=ax, linewidth=2)
+ax.set_title(' Forecast', fontsize=22)
+ax.set_ylabel('Year ', fontsize=20)
+ax.set_xlabel('Timestamp [t]', fontsize=20)
+ax.legend(prop={'size': 15})
+ax.grid(True)
+plt.show()
+```
+
+
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[95]` means that
+ the model expects the real value to be inside that interval 95% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+sf.predict(h=horizon)
+```
+
+| | unique_id | ds | ARCH(2) |
+|-----|-----------|---------------------------|-----------|
+| 0 | 1 | 2023-05-25 00:00:00+00:00 | 1.681839 |
+| 1 | 1 | 2023-05-26 00:00:00+00:00 | -0.777029 |
+| 2 | 1 | 2023-05-29 00:00:00+00:00 | -0.677962 |
+| ... | ... | ... | ... |
+| 79 | 1 | 2023-09-13 00:00:00+00:00 | 0.695591 |
+| 80 | 1 | 2023-09-14 00:00:00+00:00 | -0.176075 |
+| 81 | 1 | 2023-09-15 00:00:00+00:00 | -0.158605 |
+
+```python
+forecast_df = sf.predict(h=horizon, level=[80,95])
+forecast_df
+```
+
+| | unique_id | ds | ARCH(2) | ARCH(2)-lo-95 | ARCH(2)-lo-80 | ARCH(2)-hi-80 | ARCH(2)-hi-95 |
+|-----|-----------|---------------------------|-----------|---------------|---------------|---------------|---------------|
+| 0 | 1 | 2023-05-25 00:00:00+00:00 | 1.681839 | -0.419326 | 0.307961 | 3.055716 | 3.783003 |
+| 1 | 1 | 2023-05-26 00:00:00+00:00 | -0.777029 | -3.939054 | -2.844566 | 1.290508 | 2.384996 |
+| 2 | 1 | 2023-05-29 00:00:00+00:00 | -0.677962 | -3.907262 | -2.789488 | 1.433564 | 2.551338 |
+| ... | ... | ... | ... | ... | ... | ... | ... |
+| 79 | 1 | 2023-09-13 00:00:00+00:00 | 0.695591 | -0.937585 | -0.372285 | 1.763467 | 2.328766 |
+| 80 | 1 | 2023-09-14 00:00:00+00:00 | -0.176075 | -1.405359 | -0.979860 | 0.627711 | 1.053210 |
+| 81 | 1 | 2023-09-15 00:00:00+00:00 | -0.158605 | -1.381915 | -0.958485 | 0.641274 | 1.064705 |
+
+We can join the forecast result with the historical data using the
+pandas function `pd.concat()`, and then be able to use this result for
+graphing.
+
+```python
+df_plot=pd.concat([df, forecast_df]).set_index('ds').tail(220)
+df_plot
+```
+
+| | unique_id | y | ARCH(2) | ARCH(2)-lo-95 | ARCH(2)-lo-80 | ARCH(2)-hi-80 | ARCH(2)-hi-95 |
+|---------------------------|-----------|-----------|-----------|---------------|---------------|---------------|---------------|
+| ds | | | | | | | |
+| 2023-03-07 00:00:00+00:00 | 1 | -1.532692 | NaN | NaN | NaN | NaN | NaN |
+| 2023-03-08 00:00:00+00:00 | 1 | 0.141479 | NaN | NaN | NaN | NaN | NaN |
+| 2023-03-09 00:00:00+00:00 | 1 | -1.845936 | NaN | NaN | NaN | NaN | NaN |
+| ... | ... | ... | ... | ... | ... | ... | ... |
+| 2023-09-13 00:00:00+00:00 | 1 | NaN | 0.695591 | -0.937585 | -0.372285 | 1.763467 | 2.328766 |
+| 2023-09-14 00:00:00+00:00 | 1 | NaN | -0.176075 | -1.405359 | -0.979860 | 0.627711 | 1.053210 |
+| 2023-09-15 00:00:00+00:00 | 1 | NaN | -0.158605 | -1.381915 | -0.958485 | 0.641274 | 1.064705 |
+
+```python
+sf.plot(train, test.merge(forecast_df), level=[80, 95], max_insample_length=120)
+```
+
+
+
+Let’s plot the same graph using the plot function that comes in
+`Statsforecast`, as shown below.
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=5)`, forecasting every second months
+`(step_size=12)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 12 months ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=train,
+ h=horizon,
+ step_size=6,
+ n_windows=5)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` series identifier
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the n_windows.
+- `y:` true value
+- `"model":` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df
+```
+
+| | unique_id | ds | cutoff | y | ARCH(2) |
+|-----|-----------|---------------------------|---------------------------|-----------|-----------|
+| 0 | 1 | 2022-12-21 00:00:00+00:00 | 2022-12-20 00:00:00+00:00 | 1.486799 | 1.382105 |
+| 1 | 1 | 2022-12-22 00:00:00+00:00 | 2022-12-20 00:00:00+00:00 | -1.445170 | -0.651618 |
+| 2 | 1 | 2022-12-23 00:00:00+00:00 | 2022-12-20 00:00:00+00:00 | 0.586810 | -0.595213 |
+| ... | ... | ... | ... | ... | ... |
+| 407 | 1 | 2023-05-22 00:00:00+00:00 | 2023-01-26 00:00:00+00:00 | 0.015503 | 0.693070 |
+| 408 | 1 | 2023-05-23 00:00:00+00:00 | 2023-01-26 00:00:00+00:00 | -1.122203 | -0.176181 |
+| 409 | 1 | 2023-05-24 00:00:00+00:00 | 2023-01-26 00:00:00+00:00 | -0.731860 | -0.157522 |
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+from utilsforecast.evaluation import evaluate
+from utilsforecast.losses import mae, mape, mase, rmse, smape
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ train_df=train,
+ metrics=[mae, mape, partial(mase, seasonality=5), rmse, smape],
+ agg_fn='mean',
+)
+```
+
+| | metric | ARCH(2) |
+|-----|--------|-----------|
+| 0 | mae | 0.949721 |
+| 1 | mape | 11.789856 |
+| 2 | mase | 0.875298 |
+| 3 | rmse | 1.164914 |
+| 4 | smape | 0.725702 |
+
+## References
+
+1. [Changquan Huang • Alla Petukhina. Springer series (2022). Applied
+ Time Series Analysis and Forecasting with
+ Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2)
+2. [Engle, R. F. (1982). Autoregressive conditional heteroscedasticity
+ with estimates of the variance of United Kingdom inflation.
+ Econometrica: Journal of the econometric society,
+ 987-1007.](http://www.econ.uiuc.edu/~econ508/Papers/engle82.pdf).
+3. [James D. Hamilton. Time Series Analysis Princeton University Press,
+ Princeton, New Jersey, 1st Edition,
+ 1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Nixtla ARCH API](../../src/core/models.html#arch)
+5. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+6. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+7. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/models/arima.html.mdx b/statsforecast/docs/models/arima.html.mdx
new file mode 100644
index 00000000..9457c861
--- /dev/null
+++ b/statsforecast/docs/models/arima.html.mdx
@@ -0,0 +1,864 @@
+---
+title: ARIMA Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `ARIMA Model` with `Statsforecast`.
+
+In this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation`.
+
+The text in this article is largely taken from [Rob J. Hyndman and
+George Athanasopoulos (2018). “Forecasting Principles and Practice (3rd
+ed)”.](https://otexts.com/fpp3/tscv.html)
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [ARIMA Models](#model)
+- [The meaning of p, d and q in ARIMA model](#concepts)
+- [AR and MA models](#ar_ma)
+- [ARIMA model](#arima)
+- [How to find the order of differencing (d) in ARIMA model](#order_d)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [How to find the order of the AR term (p)](#order_p)
+- [How to find the order of the MA term (q)](#order_q)
+- [How to handle if a time series is slightly under or over
+ differenced](#differencing)
+- [Implementation of ARIMA with StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+- A **Time Series** is defined as a series of data points recorded at
+ different time intervals. The time order can be daily, monthly, or
+ even yearly.
+
+- Time Series forecasting is the process of using a statistical model
+ to predict future values of a time series based on past results.
+
+- We have discussed various aspects of **Time Series Forecasting** in
+ the previous notebook.
+
+- Forecasting is the step where we want to predict the future values
+ the series is going to take. Forecasting a time series is often of
+ tremendous commercial value.
+
+**Forecasting a time series can be broadly divided into two types.**
+
+- If we use only the previous values of the time series to predict its
+ future values, it is called **Univariate Time Series Forecasting.**
+
+- If we use predictors other than the series (like exogenous
+ variables) to forecast it is called **Multi Variate Time Series
+ Forecasting.**
+
+- This notebook focuses on a particular type of forecasting method
+ called **ARIMA modeling.**
+
+## Introduction to ARIMA Models
+
+- **ARIMA** stands for **Autoregressive Integrated Moving Average
+ Model**. It belongs to a class of models that explains a given time
+ series based on its own past values -i.e.- its own lags and the
+ lagged forecast errors. The equation can be used to forecast future
+ values. Any ‘non-seasonal’ time series that exhibits patterns and is
+ not a random white noise can be modeled with ARIMA models.
+
+- So, **ARIMA**, short for **AutoRegressive Integrated Moving
+ Average**, is a forecasting algorithm based on the idea that the
+ information in the past values of the time series can alone be used
+ to predict the future values.
+
+- **ARIMA Models** are specified by three order parameters: (p, d, q),
+
+ where,
+
+ - p is the order of the AR term
+
+ - q is the order of the MA term
+
+ - d is the number of differencing required to make the time series
+ stationary
+
+- **AR(p) Autoregression** - a regression model that utilizes the
+ dependent relationship between a current observation and
+ observations over a previous period. An auto regressive (AR(p))
+ component refers to the use of past values in the regression
+ equation for the time series.
+
+- **I(d) Integration** - uses differencing of observations
+ (subtracting an observation from observation at the previous time
+ step) in order to make the time series stationary. Differencing
+ involves the subtraction of the current values of a series with its
+ previous values d number of times.
+
+- **MA(q) Moving Average** - a model that uses the dependency between
+ an observation and a residual error from a moving average model
+ applied to lagged observations. A moving average component depicts
+ the error of the model as a combination of previous error terms. The
+ order q represents the number of terms to be included in the model.
+
+### Types of ARIMA Model
+
+- **ARIMA** : Non-seasonal Autoregressive Integrated Moving Averages
+- **SARIMA** : Seasonal ARIMA
+- **SARIMAX** : Seasonal ARIMA with exogenous variables
+
+If a time series, has seasonal patterns, then we need to add seasonal
+terms and it becomes SARIMA, short for **Seasonal ARIMA**.
+
+## The meaning of p, d and q in ARIMA model
+
+### The meaning of p
+
+- `p` is the order of the **Auto Regressive (AR)** term. It refers to
+ the number of lags of Y to be used as predictors.
+
+### The meaning of d
+
+- The term **Auto Regressive**’ in ARIMA means it is a linear
+ regression model that uses its own lags as predictors. Linear
+ regression models, as we know, work best when the predictors are not
+ correlated and are independent of each other. So we need to make the
+ time series stationary.
+
+- The most common approach to make the series stationary is to
+ difference it. That is, subtract the previous value from the current
+ value. Sometimes, depending on the complexity of the series, more
+ than one differencing may be needed.
+
+- The value of d, therefore, is the minimum number of differencing
+ needed to make the series stationary. If the time series is already
+ stationary, then d = 0.
+
+### The meaning of q
+
+- **q** is the order of the **Moving Average (MA)** term. It refers to
+ the number of lagged forecast errors that should go into the ARIMA
+ Model.
+
+## AR and MA models
+
+### AR model
+
+In an autoregression model, we forecast the variable of interest using a
+linear combination of past values of the variable. The term
+autoregression indicates that it is a regression of the variable against
+itself.
+
+Thus, an autoregressive model of order p can be written as
+
+$$
+\begin{equation}
+y_{t} = c + \phi_{1}y_{t-1} + \phi_{2}y_{t-2} + \dots + \phi_{p}y_{t-p} + \varepsilon_{t} \tag{1}
+\end{equation}
+$$
+
+where $\epsilon_t$ is white noise. This is like a multiple regression
+but with lagged values of $y_t$ as predictors. We refer to this as an
+AR( p) model, an autoregressive model of order p.
+
+### MA model
+
+Rather than using past values of the forecast variable in a regression,
+a moving average model uses past forecast errors in a regression-like
+model,
+
+$$
+\begin{equation}
+y_{t} = c + \varepsilon_t + \theta_{1}\varepsilon_{t-1} + \theta_{2}\varepsilon_{t-2} + \dots + \theta_{q}\varepsilon_{t-q} \tag{2}
+\end{equation}
+$$
+
+where $\epsilon_t$ is white noise. We refer to this as an MA(q) model, a
+moving average model of order q. Of course, we do not observe the values
+of
+$\epsilon_t$ , so it is not really a regression in the usual sense.
+
+Notice that each value of yt can be thought of as a weighted moving
+average of the past few forecast errors (although the coefficients will
+not normally sum to one). However, moving average models should not be
+confused with the moving average smoothing . A moving average model is
+used for forecasting future values, while moving average smoothing is
+used for estimating the trend-cycle of past values.
+
+Thus, we have discussed AR and MA Models respectively.
+
+## ARIMA model
+
+If we combine differencing with autoregression and a moving average
+model, we obtain a non-seasonal ARIMA model. ARIMA is an acronym for
+AutoRegressive Integrated Moving Average (in this context, “integration”
+is the reverse of differencing). The full model can be written as
+
+where $y'_{t}$ is the differenced series (it may have been differenced
+more than once). The “predictors” on the right hand side include both
+lagged values of $y_t$ and lagged errors. We call this an ARIMA(p,d,q)
+model, where
+
+| | |
+|-----|---------------------------------------|
+| p | order of the autoregressive part |
+| d | degree of first differencing involved |
+| q | order of the moving average part |
+
+The same stationarity and invertibility conditions that are used for
+autoregressive and moving average models also apply to an ARIMA model.
+
+Many of the models we have already discussed are special cases of the
+ARIMA model, as shown in Table
+
+| Model | p d q | Differenced | Method |
+|------------------|-------------|-------------------------|-----------------|
+| Arima(0,0,0) | 0 0 0 | $y_t=Y_t$ | White noise |
+| ARIMA (0,1,0) | 0 1 0 | $y_t = Y_t - Y_{t-1}$ | Random walk |
+| ARIMA (0,2,0) | 0 2 0 | $y_t = Y_t - 2Y_{t-1} + Y_{t-2}$ | Constant |
+| ARIMA (1,0,0) | 1 0 0 | $\hat Y_t = \mu + \Phi_1 Y_{t-1} + \epsilon$ | AR(1): AR(1): First-order regression model |
+| ARIMA (2, 0, 0) | 2 0 0 | $\hat Y_t = \Phi_0 + \Phi_1 Y_{t-1} + \Phi_2 Y_{t-2} + \epsilon$ | AR(2): Second-order regression model |
+| ARIMA (1, 1, 0) | 1 1 0 | $\hat Y_t = \mu + Y_{t-1} + \Phi_1 (Y_{t-1}- Y_{t-2})$ | Differenced first-order |
+| autoregressive model | | | |
+| ARIMA (0, 1, 1) | 0 1 1 | $\hat Y_t = Y_{t-1} - \Phi_1 e^{t-1}$ | Simple exponential |
+| smoothing | | | |
+| ARIMA (0, 0, 1) | 0 0 1 | $\hat Y_t = \mu_0+ \epsilon_t - \omega_1 \epsilon_{t-1}$ | MA(1): First-order |
+| regression model | | | |
+| ARIMA (0, 0, 2) | 0 0 2 | $\hat Y_t = \mu_0+ \epsilon_t - \omega_1 \epsilon_{t-1} - \omega_2 \epsilon_{t-2}$ | MA(2): Second-order |
+| regression model | | | |
+| ARIMA (1, 0, 1) | 1 0 1 | $\hat Y_t = \Phi_0 + \Phi_1 Y_{t-1}+ \epsilon_t - \omega_1 \epsilon_{t-1}$ | ARMA model |
+| ARIMA (1, 1, 1) | 1 1 1 | $\Delta Y_t = \Phi_1 Y_{t-1} + \epsilon_t - \omega_1 \epsilon_{t-1}$ | ARIMA model |
+| ARIMA (1, 1, 2) | 1 1 2 | $\hat Y_t = Y_{t-1} + \Phi_1 (Y_{t-1} - Y_{t-2} )- \Theta_1 e_{t-1} - \Theta_1 e_{t-1}$ Damped-trend linear Exponential smoothing | |
+| ARIMA (0, 2, 1) OR (0,2,2) | 0 2 1 | $\hat Y_t = 2 Y_{t-1} - Y_{t-2} - \Theta_1 e_{t-1} - \Theta_2 e_{t-2}$ | Linear exponential smoothing |
+
+Once we start combining components in this way to form more complicated
+models, it is much easier to work with the backshift notation. For
+example, the above equation can be written in backshift notation as
+
+### ARIMA model in words
+
+Predicted Yt = Constant + Linear combination Lags of Y (upto p lags) +
+Linear Combination of Lagged forecast errors (upto q lags)
+
+## How to find the order of differencing (d) in ARIMA model
+
+- As stated earlier, the purpose of differencing is to make the time
+ series stationary. But we should be careful to not over-difference
+ the series. An over differenced series may still be stationary,
+ which in turn will affect the model parameters.
+
+- So we should determine the right order of differencing. The right
+ order of differencing is the minimum differencing required to get a
+ near-stationary series which roams around a defined mean and the ACF
+ plot reaches to zero fairly quick.
+
+- If the autocorrelations are positive for many number of lags (10 or
+ more), then the series needs further differencing. On the other
+ hand, if the lag 1 autocorrelation itself is too negative, then the
+ series is probably over-differenced.
+
+- If we can’t really decide between two orders of differencing, then
+ we go with the order that gives the least standard deviation in the
+ differenced series.
+
+- Now, we will explain these concepts with the help of an example as
+ follows:
+
+ - First, I will check if the series is stationary using the
+ **Augmented Dickey Fuller test (ADF Test)**, from the
+ statsmodels package. The reason being is that we need
+ differencing only if the series is non-stationary. Else, no
+ differencing is needed, that is, d=0.
+
+ - The null hypothesis (Ho) of the ADF test is that the time series
+ is non-stationary. So, if the p-value of the test is less than
+ the significance level (0.05) then we reject the null hypothesis
+ and infer that the time series is indeed stationary.
+
+ - So, in our case, if P Value \> 0.05 we go ahead with finding the
+ order of differencing.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import matplotlib.pyplot as plt
+
+plt.style.use('fivethirtyeight')
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#212946',
+ 'axes.facecolor': '#212946',
+ 'savefig.facecolor':'#212946',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#2A3459',
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+### Read data
+
+```python
+import pandas as pd
+import numpy as np
+
+df = pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/Esperanza_vida.csv", usecols=[1,2])
+df.head()
+```
+
+| | year | value |
+|-----|------------|-----------|
+| 0 | 1960-01-01 | 69.123902 |
+| 1 | 1961-01-01 | 69.760244 |
+| 2 | 1962-01-01 | 69.149756 |
+| 3 | 1963-01-01 | 69.248049 |
+| 4 | 1964-01-01 | 70.311707 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|------------|-----------|-----------|
+| 0 | 1960-01-01 | 69.123902 | 1 |
+| 1 | 1961-01-01 | 69.760244 | 1 |
+| 2 | 1962-01-01 | 69.149756 | 1 |
+| 3 | 1963-01-01 | 69.248049 | 1 |
+| 4 | 1964-01-01 | 70.311707 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y float64
+unique_id object
+dtype: object
+```
+
+We need to convert `ds` from the `object` type to datetime.
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore data with the plot method
+
+Plot a series using the plot method from the StatsForecast class. This
+method prints a random series from the dataset and is useful for basic
+EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+Looking at the plot we can observe there is an upward trend over the
+period of time.
+
+```python
+df["y"].plot(kind='kde',figsize = (16,5))
+df["y"].describe()
+```
+
+``` text
+count 60.000000
+mean 76.632439
+std 4.495279
+ ...
+50% 76.895122
+75% 80.781098
+max 83.346341
+Name: y, Length: 8, dtype: float64
+```
+
+
+
+### Seasonal Decomposed
+
+How to decompose a time series and why?
+
+In time series analysis to forecast new values, it is very important to
+know past data. More formally, we can say that it is very important to
+know the patterns that values follow over time. There can be many
+reasons that cause our forecast values to fall in the wrong direction.
+Basically, a time series consists of four components. The variation of
+those components causes the change in the pattern of the time series.
+These components are:
+
+- **Level:** This is the primary value that averages over time.
+- **Trend:** The trend is the value that causes increasing or
+ decreasing patterns in a time series.
+- **Seasonality:** This is a cyclical event that occurs in a time
+ series for a short time and causes short-term increasing or
+ decreasing patterns in a time series.
+- **Residual/Noise:** These are the random variations in the time
+ series.
+
+Combining these components over time leads to the formation of a time
+series. Most time series consist of level and noise/residual and trend
+or seasonality are optional values.
+
+If seasonality and trend are part of the time series, then there will be
+effects on the forecast value. As the pattern of the forecasted time
+series may be different from the previous time series.
+
+The combination of the components in time series can be of two types: \*
+Additive \* multiplicative
+
+### Additive time series
+
+If the components of the time series are added to make the time series.
+Then the time series is called the additive time series. By
+visualization, we can say that the time series is additive if the
+increasing or decreasing pattern of the time series is similar
+throughout the series. The mathematical function of any additive time
+series can be represented by:
+$$y(t) = level + Trend + seasonality + noise$$
+
+### Multiplicative time series
+
+If the components of the time series are multiplicative together, then
+the time series is called a multiplicative time series. For
+visualization, if the time series is having exponential growth or
+decline with time, then the time series can be considered as the
+multiplicative time series. The mathematical function of the
+multiplicative time series can be represented as.
+
+$$y(t) = Level * Trend * seasonality * Noise$$
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+```
+
+
+```python
+decomposed=seasonal_decompose(df["y"], model = "add", period=1)
+decomposed.plot()
+plt.show()
+```
+
+
+
+### The Augmented Dickey-Fuller Test
+
+An Augmented Dickey-Fuller (ADF) test is a type of statistical test that
+determines whether a unit root is present in time series data. Unit
+roots can cause unpredictable results in time series analysis. A null
+hypothesis is formed in the unit root test to determine how strongly
+time series data is affected by a trend. By accepting the null
+hypothesis, we accept the evidence that the time series data is not
+stationary. By rejecting the null hypothesis or accepting the
+alternative hypothesis, we accept the evidence that the time series data
+is generated by a stationary process. This process is also known as
+stationary trend. The values of the ADF test statistic are negative.
+Lower ADF values indicate a stronger rejection of the null hypothesis.
+
+Augmented Dickey-Fuller Test is a common statistical test used to test
+whether a given time series is stationary or not. We can achieve this by
+defining the null and alternate hypothesis.
+
+Null Hypothesis: Time Series is non-stationary. It gives a
+time-dependent trend. Alternate Hypothesis: Time Series is stationary.
+In another term, the series doesn’t depend on time.
+
+ADF or t Statistic \< critical values: Reject the null hypothesis, time
+series is stationary. ADF or t Statistic \> critical values: Failed to
+reject the null hypothesis, time series is non-stationary.
+
+```python
+from statsmodels.tsa.stattools import adfuller
+```
+
+
+```python
+def Augmented_Dickey_Fuller_Test_func(series , column_name):
+ print (f'Dickey-Fuller test results for columns: {column_name}')
+ dftest = adfuller(series, autolag='AIC')
+ dfoutput = pd.Series(dftest[0:4], index=['Test Statistic','p-value','No Lags Used','Number of observations used'])
+ for key,value in dftest[4].items():
+ dfoutput['Critical Value (%s)'%key] = value
+ print (dfoutput)
+ if dftest[1] \<= 0.05:
+ print("Conclusion:====>")
+ print("Reject the null hypothesis")
+ print("The data is stationary")
+ else:
+ print("Conclusion:====>")
+ print("The null hypothesis cannot be rejected")
+ print("The data is not stationary")
+```
+
+
+```python
+Augmented_Dickey_Fuller_Test_func(df["y"],"Life expectancy")
+```
+
+``` text
+Dickey-Fuller test results for columns: Life expectancy
+Test Statistic -1.578590
+p-value 0.494339
+No Lags Used 2.000000
+ ...
+Critical Value (1%) -3.550670
+Critical Value (5%) -2.913766
+Critical Value (10%) -2.594624
+Length: 7, dtype: float64
+Conclusion:====>
+The null hypothesis cannot be rejected
+The data is not stationary
+```
+
+We can see in the result that we obtained the non-stationary series,
+because the p-value is greater than 5%.
+
+One of the objectives of applying the ADF test is to know if our series
+is stationary, knowing the result of the ADF test, then we can determine
+the next step. For our case, it can be seen from the previous result
+that the time series is not stationary, so we will proceed to the next
+step, which is to differentiate our time series.
+
+We are going to create a copy of our data, with the objective of
+investigating to find the stationarity in our time series.
+
+Once we have made the copy of the time series, we are going to
+differentiate the time series, and then we will use the augmented Dickey
+Fuller test to investigate if our time series is stationary.
+
+```python
+df1=df.copy()
+df1['y_diff'] = df['y'].diff()
+df1.dropna(inplace=True)
+df1.head()
+```
+
+| | ds | y | unique_id | y_diff |
+|-----|------------|-----------|-----------|-----------|
+| 1 | 1961-01-01 | 69.760244 | 1 | 0.636341 |
+| 2 | 1962-01-01 | 69.149756 | 1 | -0.610488 |
+| 3 | 1963-01-01 | 69.248049 | 1 | 0.098293 |
+| 4 | 1964-01-01 | 70.311707 | 1 | 1.063659 |
+| 5 | 1965-01-01 | 70.171707 | 1 | -0.140000 |
+
+Let’s apply the Dickey Fuller test again to find out if our time series
+is already stationary.
+
+```python
+Augmented_Dickey_Fuller_Test_func(df1["y_diff"],"Life expectancy")
+```
+
+``` text
+Dickey-Fuller test results for columns: Life expectancy
+Test Statistic -8.510100e+00
+p-value 1.173776e-13
+No Lags Used 1.000000e+00
+ ...
+Critical Value (1%) -3.550670e+00
+Critical Value (5%) -2.913766e+00
+Critical Value (10%) -2.594624e+00
+Length: 7, dtype: float64
+Conclusion:====>
+Reject the null hypothesis
+The data is stationary
+```
+
+We can observe in the previous result that now if our time series is
+stationary, the p-value is less than 5%.
+
+Now our time series is stationary, that is, we have only differentiated
+1 time, therefore, the order of our parameter $d=1$.
+
+```python
+from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
+import matplotlib.pyplot as plt
+
+
+fig, axes = plt.subplots(2, 2, )
+axes[0, 0].plot(df1["y"]); axes[0, 0].set_title('Original Series')
+plot_acf(df1["y"], ax=axes[0, 1],lags=20)
+
+axes[1, 0].plot(df1["y"].diff()); axes[1, 0].set_title('1st Order Differencing')
+plot_acf(df1["y"].diff().dropna(), ax=axes[1, 1],lags=20)
+
+
+plt.show()
+```
+
+
+
+- For the above data, we can see that the time series reaches
+ stationarity with one orders of differencing.
+
+## How to find the order of the AR term (p)
+
+- The next step is to identify if the model needs any AR terms. We
+ will find out the required number of AR terms by inspecting the
+ **Partial Autocorrelation (PACF) plot**.
+
+- **Partial autocorrelation** can be imagined as the correlation
+ between the series and its lag, after excluding the contributions
+ from the intermediate lags. So, PACF sort of conveys the pure
+ correlation between a lag and the series. This way, we will know if
+ that lag is needed in the AR term or not.
+
+- Partial autocorrelation of lag (k) of a series is the coefficient of
+ that lag in the autoregression equation of $Y$.
+
+$$Yt = \alpha0 + \alpha1 Y{t-1} + \alpha2 Y{t-2} + \alpha3 Y{t-3}$$
+
+- That is, suppose, if $Y_t$ is the current series and $Y_{t-1}$ is
+ the lag 1 of $Y$, then the partial autocorrelation of lag 3
+ $(Y_{t-3})$ is the coefficient $\alpha_3$ of $Y_{t-3}$ in the above
+ equation.
+
+- Now, we should find the number of AR terms. Any autocorrelation in a
+ stationarized series can be rectified by adding enough AR terms. So,
+ we initially take the order of AR term to be equal to as many lags
+ that crosses the significance limit in the PACF plot.
+
+```python
+fig, axes = plt.subplots(1, 2)
+axes[0].plot(df1["y"].diff()); axes[0].set_title('1st Differencing')
+axes[1].set(ylim=(0,5))
+plot_pacf(df1["y"].diff().dropna(), ax=axes[1],lags=20)
+
+plt.show()
+```
+
+
+
+- We can see that the PACF lag 1 is quite significant since it is well
+ above the significance line. So, we will fix the value of p as 1.
+
+## How to find the order of the MA term (q)
+
+- Just like how we looked at the PACF plot for the number of AR terms,
+ we will look at the ACF plot for the number of MA terms. An MA term
+ is technically, the error of the lagged forecast.
+
+- The ACF tells how many MA terms are required to remove any
+ autocorrelation in the stationarized series.
+
+- Let’s see the autocorrelation plot of the differenced series.
+
+```python
+from statsmodels.graphics.tsaplots import plot_acf
+
+fig, axes = plt.subplots(1, 2)
+axes[0].plot(df1["y"].diff()); axes[0].set_title('1st Differencing')
+axes[1].set(ylim=(0,1.2))
+plot_acf(df["y"].diff().dropna(), ax=axes[1], lags=20)
+
+plt.show()
+```
+
+
+
+- We can see that couple of lags are well above the significance line.
+ So, we will fix q as 1. If there is any doubt, we will go with the
+ simpler model that sufficiently explains the Y.
+
+## How to handle if a time series is slightly under or over differenced
+
+- It may happen that the time series is slightly under differenced.
+ Differencing it one more time makes it slightly over-differenced.
+
+- If the series is slightly under differenced, adding one or more
+ additional AR terms usually makes it up. Likewise, if it is slightly
+ over-differenced, we will try adding an additional MA term.
+
+## Implementation of ARIMA with StatsForecast
+
+Now, we have determined the values of p, d and q. We have everything
+needed to fit the ARIMA model. We will use the ARIMA() implementation in
+the `statsforecast` package.
+
+The parameters found are: \* For the autoregressive model, $p=1$ \* for
+the moving average model $q=1$ \* and for the stationarity of the model
+with a differential with an order $d=1$
+
+Therefore, the model that we are going to test is the ARIMA(1,1,1)
+model.
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import ARIMA
+```
+
+### Instantiate the model
+
+```python
+sf = StatsForecast(models=[ARIMA(order=(1, 1, 1))], freq='YS')
+```
+
+### Fit the Model
+
+```python
+sf.fit(df)
+```
+
+``` text
+StatsForecast(models=[ARIMA])
+```
+
+### Making the predictions
+
+```python
+y_hat = sf.predict(h=6)
+y_hat
+```
+
+| | unique_id | ds | ARIMA |
+|-----|-----------|------------|-----------|
+| 0 | 1 | 2020-01-01 | 83.206903 |
+| 1 | 1 | 2021-01-01 | 83.203508 |
+| 2 | 1 | 2022-01-01 | 83.204742 |
+| 3 | 1 | 2023-01-01 | 83.204293 |
+| 4 | 1 | 2024-01-01 | 83.204456 |
+| 5 | 1 | 2025-01-01 | 83.204397 |
+
+We can make the predictions by adding the confidence interval, for
+example with 95%.
+
+```python
+y_hat2 = sf.predict(h=6, level=[95])
+y_hat2
+```
+
+| | unique_id | ds | ARIMA | ARIMA-lo-95 | ARIMA-hi-95 |
+|-----|-----------|------------|-----------|-------------|-------------|
+| 0 | 1 | 2020-01-01 | 83.206903 | 82.412336 | 84.001469 |
+| 1 | 1 | 2021-01-01 | 83.203508 | 82.094625 | 84.312391 |
+| 2 | 1 | 2022-01-01 | 83.204742 | 81.848344 | 84.561139 |
+| 3 | 1 | 2023-01-01 | 83.204293 | 81.640430 | 84.768156 |
+| 4 | 1 | 2024-01-01 | 83.204456 | 81.457145 | 84.951767 |
+| 5 | 1 | 2025-01-01 | 83.204397 | 81.291297 | 85.117497 |
+
+### forecast method
+
+Memory efficient predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to fit_predict without storing information. It assumes you
+know the forecast horizon in advance.
+
+```python
+Y_hat_df = sf.forecast(df=df, h=6, level=[95])
+Y_hat_df
+```
+
+| | unique_id | ds | ARIMA | ARIMA-lo-95 | ARIMA-hi-95 |
+|-----|-----------|------------|-----------|-------------|-------------|
+| 0 | 1 | 2020-01-01 | 83.206903 | 82.412336 | 84.001469 |
+| 1 | 1 | 2021-01-01 | 83.203508 | 82.094625 | 84.312391 |
+| 2 | 1 | 2022-01-01 | 83.204742 | 81.848344 | 84.561139 |
+| 3 | 1 | 2023-01-01 | 83.204293 | 81.640430 | 84.768156 |
+| 4 | 1 | 2024-01-01 | 83.204456 | 81.457145 | 84.951767 |
+| 5 | 1 | 2025-01-01 | 83.204397 | 81.291297 | 85.117497 |
+
+Once the predictions have been generated we can perform a visualization
+to see the generated behavior of our model.
+
+```python
+sf.plot(df, Y_hat_df, level=[95])
+```
+
+
+
+## Model Evaluation
+
+The commonly used accuracy metrics to judge forecasts are:
+
+1. Mean Absolute Percentage Error (MAPE)
+2. Mean Error (ME)
+3. Mean Absolute Error (MAE)
+4. Mean Percentage Error (MPE)
+5. Root Mean Squared Error (RMSE)
+6. Correlation between the Actual and the Forecast (corr)
+
+```python
+Y_train_df = df[df.ds\<='2013-01-01']
+Y_test_df = df[df.ds>'2013-01-01']
+
+Y_train_df.shape, Y_test_df.shape
+```
+
+``` text
+((54, 3), (6, 3))
+```
+
+```python
+Y_hat_df = sf.forecast(df=Y_train_df, h=len(Y_test_df))
+```
+
+
+```python
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ Y_test_df.merge(Y_hat_df),
+ metrics=[ufl.mse, ufl.mae, ufl.rmse, ufl.mape],
+)
+```
+
+| | unique_id | metric | ARIMA |
+|-----|-----------|--------|----------|
+| 0 | 1 | mse | 0.184000 |
+| 1 | 1 | mae | 0.397932 |
+| 2 | 1 | rmse | 0.428952 |
+| 3 | 1 | mape | 0.004785 |
+
+## References
+
+1. [Nixtla ARIMA API](../../src/core/models.html#arima)
+2. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+
diff --git a/statsforecast/docs/models/autoarima.html.mdx b/statsforecast/docs/models/autoarima.html.mdx
new file mode 100644
index 00000000..904609a7
--- /dev/null
+++ b/statsforecast/docs/models/autoarima.html.mdx
@@ -0,0 +1,833 @@
+---
+title: AutoARIMA Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `AutoARIMA Model` with
+> `Statsforecast`.
+
+The objective of the following article is to obtain a step-by-step guide
+on building the Arima model using `AutoARIMA` with `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation`.
+
+The text in this article is largely taken from [Rob J. Hyndman and
+George Athanasopoulos (2018). “Forecasting Principles and Practice (3rd
+ed)”.](https://otexts.com/fpp3/tscv.html)
+
+## Table of Contents
+
+- [Implementation of AutoArima with StatsForecast ](#implementation-of-autoarima-with-statsforecast-)
+ - [Load libraries](#load-libraries)
+ - [Instantiating Model](#instantiating-model)
+ - [Fit the Model](#fit-the-model)
+ - [Forecast Method](#forecast-method)
+ - [Predict method with confidence interval](#predict-method-with-confidence-interval)
+ - [Cross-validation ](#cross-validation-)
+ - [Perform time series cross-validation](#perform-time-series-cross-validation)
+ - [Model Evaluation ](#model-evaluation-)
+ - [References ](#references-)
+
+## What is AutoArima with StatsForecast?
+
+An autoARIMA is a time series model that uses an automatic process to
+select the optimal ARIMA (Autoregressive Integrated Moving Average)
+model parameters for a given time series. ARIMA is a widely used
+statistical model for modeling and predicting time series.
+
+The process of automatic parameter selection in an autoARIMA model is
+performed using statistical and optimization techniques, such as the
+Akaike Information Criterion (AIC) and cross-validation, to identify
+optimal values for autoregression, integration, and moving average
+parameters. of the ARIMA model.
+
+Automatic parameter selection is useful because it can be difficult to
+determine the optimal parameters of an ARIMA model for a given time
+series without a thorough understanding of the underlying stochastic
+process that generates the time series. The autoARIMA model automates
+the parameter selection process and can provide a fast and effective
+solution for time series modeling and forecasting.
+
+The `statsforecast.models` library brings the `AutoARIMA` function from
+Python provides an implementation of autoARIMA that allows to
+automatically select the optimal parameters for an ARIMA model given a
+time series.
+
+## Definition of the Arima model
+
+An Arima model (autoregressive integrated moving average) process is the
+combination of an autoregressive process AR(p), integration I(d), and
+the moving average process MA(q).
+
+Just like the ARMA process, the ARIMA process states that the present
+value is dependent on past values, coming from the AR(p) portion, and
+past errors, coming from the MA(q) portion. However, instead of using
+the original series, denoted as yt, the ARIMA process uses the
+differenced series, denoted as $y'_{t}$. Note that $y'_{t}$ can
+represent a series that has been differenced more than once.
+
+Therefore, the mathematical expression of the ARIMA(p,d,q) process
+states that the present value of the differenced series $y'_{t}$ is
+equal to the sum of a constant $C$, past values of the differenced
+series $\phi_{p}y'_{t-p}$, the mean of the differenced series $\mu$,
+past error terms $\theta_{q}\varepsilon_{t-q}$, and a current error term
+$\varepsilon_{t}$, as shown in equation
+
+where $y'_{t}$ is the differenced series (it may have been differenced
+more than once). The “predictors” on the right hand side include both
+lagged values of $y_{t}$ and lagged errors. We call this an **ARIMA(
+p,d,q)** model, where
+
+| | |
+|-----|---------------------------------------|
+| p | order of the autoregressive part |
+| d | degree of first differencing involved |
+| q | order of the moving average part |
+
+The same stationarity and invertibility conditions that are used for
+autoregressive and moving average models also apply to an ARIMA model.
+
+Many of the models we have already discussed are special cases of the
+ARIMA model, as shown in Table
+
+| Model | p d q | Differenced | Method |
+|------------------|-------------|-------------------------|-----------------|
+| Arima(0,0,0) | 0 0 0 | $y_t=Y_t$ | White noise |
+| ARIMA (0,1,0) | 0 1 0 | $y_t = Y_t - Y_{t-1}$ | Random walk |
+| ARIMA (0,2,0) | 0 2 0 | $y_t = Y_t - 2Y_{t-1} + Y_{t-2}$ | Constant |
+| ARIMA (1,0,0) | 1 0 0 | $\hat Y_t = \mu + \Phi_1 Y_{t-1} + \epsilon$ | AR(1): AR(1): First-order regression model |
+| ARIMA (2, 0, 0) | 2 0 0 | $\hat Y_t = \Phi_0 + \Phi_1 Y_{t-1} + \Phi_2 Y_{t-2} + \epsilon$ | AR(2): Second-order regression model |
+| ARIMA (1, 1, 0) | 1 1 0 | $\hat Y_t = \mu + Y_{t-1} + \Phi_1 (Y_{t-1}- Y_{t-2})$ | Differenced first-order autoregressive model |
+| ARIMA (0, 1, 1) | 0 1 1 | $\hat Y_t = Y_{t-1} - \Phi_1 e^{t-1}$ | Simple exponential smoothing |
+| ARIMA (0, 0, 1) | 0 0 1 | $\hat Y_t = \mu_0+ \epsilon_t - \omega_1 \epsilon_{t-1}$ | MA(1): First-order regression model |
+| ARIMA (0, 0, 2) | 0 0 2 | $\hat Y_t = \mu_0+ \epsilon_t - \omega_1 \epsilon_{t-1} - \omega_2 \epsilon_{t-2}$ | MA(2): Second-order regression model |
+| ARIMA (1, 0, 1) | 1 0 1 | $\hat Y_t = \Phi_0 + \Phi_1 Y_{t-1}+ \epsilon_t - \omega_1 \epsilon_{t-1}$ | ARMA model |
+| ARIMA (1, 1, 1) | 1 1 1 | $\Delta Y_t = \Phi_1 Y_{t-1} + \epsilon_t - \omega_1 \epsilon_{t-1}$ | ARIMA model |
+| ARIMA (1, 1, 2) | 1 1 2 | $\hat Y_t = Y_{t-1} + \Phi_1 (Y_{t-1} - Y_{t-2} )- \Theta_1 e_{t-1} - \Theta_1 e_{t-1}$ | Damped-trend linear Exponential smoothing |
+| ARIMA (0, 2, 1) OR (0,2,2) | 0 2 1 | $\hat Y_t = 2 Y_{t-1} - Y_{t-2} - \Theta_1 e_{t-1} - \Theta_2 e_{t-2}$ | Linear exponential smoothing |
+
+Once we start combining components in this way to form more complicated
+models, it is much easier to work with the backshift notation. For
+example, Equation (1) can be written in backshift notation as:
+
+Selecting appropriate values for p, d and q can be difficult. However,
+the `AutoARIMA()` function from `statsforecast` will do it for you
+automatically.
+
+For more information
+[here](https://otexts.com/fpp3/non-seasonal-arima.html)
+
+## Loading libraries and data
+
+Using an `AutoARIMA()` model to model and predict time series has
+several advantages, including:
+
+1. Automation of the parameter selection process: The `AutoARIMA()`
+ function automates the ARIMA model parameter selection process,
+ which can save the user time and effort by eliminating the need to
+ manually try different combinations of parameters.
+
+2. Reduction of prediction error: By automatically selecting optimal
+ parameters, the **ARIMA** model can improve the accuracy of
+ predictions compared to manually selected **ARIMA** models.
+
+3. Identification of complex patterns: The `AutoARIMA()` function can
+ identify complex patterns in the data that may be difficult to
+ detect visually or with other time series modeling techniques.
+
+4. Flexibility in the choice of the parameter selection methodology:
+ The **ARIMA** Model can use different methodologies to select the
+ optimal parameters, such as the Akaike Information Criterion (AIC),
+ cross-validation and others, which allows the user to choose the
+ methodology that best suits their needs.
+
+In general, using the `AutoARIMA()` function can help improve the
+efficiency and accuracy of time series modeling and forecasting,
+especially for users who are inexperienced with manual parameter
+selection for ARIMA models.
+
+### Main results
+
+We compared accuracy and speed against
+[pmdarima](https://github.com/alkaline-ml/pmdarima), Rob Hyndman’s
+[forecast](https://github.com/robjhyndman/forecast) package and
+Facebook’s [Prophet](https://github.com/facebook/prophet). We used the
+`Daily`, `Hourly` and `Weekly` data from the [M4
+competition](https://www.sciencedirect.com/science/article/pii/S0169207019301128).
+
+The following table summarizes the results. As can be seen, our
+`auto_arima` is the best model in accuracy (measured by the `MASE` loss)
+and time, even compared with the original implementation in R.
+
+| dataset | metric | auto_arima_nixtla | auto_arima_pmdarima \[1\] | auto_arima_r | prophet |
+|:--------|:--------|---------------:|-----------------:|------------:|--------:|
+| Daily | MASE | **3.26** | 3.35 | 4.46 | 14.26 |
+| Daily | time | **1.41** | 27.61 | 1.81 | 514.33 |
+| Hourly | MASE | **0.92** | — | 1.02 | 1.78 |
+| Hourly | time | **12.92** | — | 23.95 | 17.27 |
+| Weekly | MASE | **2.34** | 2.47 | 2.58 | 7.29 |
+| Weekly | time | 0.42 | 2.92 | **0.22** | 19.82 |
+
+\[1\] The model `auto_arima` from `pmdarima` had a problem with Hourly
+data. An issue was opened.
+
+The following table summarizes the data details.
+
+| group | n_series | mean_length | std_length | min_length | max_length |
+|:-------|---------:|------------:|-----------:|-----------:|-----------:|
+| Daily | 4,227 | 2,371 | 1,756 | 107 | 9,933 |
+| Hourly | 414 | 901 | 127 | 748 | 1,008 |
+| Weekly | 359 | 1,035 | 707 | 93 | 2,610 |
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import numpy as np
+import pandas as pd
+
+import scipy.stats as stats
+```
+
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf
+from statsmodels.graphics.tsaplots import plot_pacf
+plt.style.use('fivethirtyeight')
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#212946',
+ 'axes.facecolor': '#212946',
+ 'savefig.facecolor':'#212946',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#2A3459',
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+### Loading Data
+
+```python
+df = pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/candy_production.csv")
+df.head()
+```
+
+| | observation_date | IPG3113N |
+|-----|------------------|----------|
+| 0 | 1972-01-01 | 85.6945 |
+| 1 | 1972-02-01 | 71.8200 |
+| 2 | 1972-03-01 | 66.0229 |
+| 3 | 1972-04-01 | 64.5645 |
+| 4 | 1972-05-01 | 65.0100 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|------------|---------|-----------|
+| 0 | 1972-01-01 | 85.6945 | 1 |
+| 1 | 1972-02-01 | 71.8200 | 1 |
+| 2 | 1972-03-01 | 66.0229 | 1 |
+| 3 | 1972-04-01 | 64.5645 | 1 |
+| 4 | 1972-05-01 | 65.0100 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y float64
+unique_id object
+dtype: object
+```
+
+We need to convert `ds` from the `object` type to datetime.
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore data with the plot method
+
+Plot a series using the plot method from the StatsForecast class. This
+method prints a random series from the dataset and is useful for basic
+EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### Autocorrelation plots
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=60, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+plot_pacf(df["y"], lags=60, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+plt.show();
+```
+
+
+
+### Decomposition of the time series
+
+How to decompose a time series and why?
+
+In time series analysis to forecast new values, it is very important to
+know past data. More formally, we can say that it is very important to
+know the patterns that values follow over time. There can be many
+reasons that cause our forecast values to fall in the wrong direction.
+Basically, a time series consists of four components. The variation of
+those components causes the change in the pattern of the time series.
+These components are:
+
+- **Level:** This is the primary value that averages over time.
+- **Trend:** The trend is the value that causes increasing or
+ decreasing patterns in a time series.
+- **Seasonality:** This is a cyclical event that occurs in a time
+ series for a short time and causes short-term increasing or
+ decreasing patterns in a time series.
+- **Residual/Noise:** These are the random variations in the time
+ series.
+
+Combining these components over time leads to the formation of a time
+series. Most time series consist of level and noise/residual and trend
+or seasonality are optional values.
+
+If seasonality and trend are part of the time series, then there will be
+effects on the forecast value. As the pattern of the forecasted time
+series may be different from the previous time series.
+
+The combination of the components in time series can be of two types: \*
+Additive \* multiplicative
+
+Additive time series
+
+If the components of the time series are added to make the time series.
+Then the time series is called the additive time series. By
+visualization, we can say that the time series is additive if the
+increasing or decreasing pattern of the time series is similar
+throughout the series. The mathematical function of any additive time
+series can be represented by:
+$$y(t) = level + Trend + seasonality + noise$$
+
+## Multiplicative time series
+
+If the components of the time series are multiplicative together, then
+the time series is called a multiplicative time series. For
+visualization, if the time series is having exponential growth or
+decline with time, then the time series can be considered as the
+multiplicative time series. The mathematical function of the
+multiplicative time series can be represented as.
+
+$$y(t) = Level * Trend * seasonality * Noise$$
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+a = seasonal_decompose(df["y"], model = "add", period=12)
+a.plot();
+```
+
+
+
+## Split the data into training and testing
+
+Let’s divide our data into sets 1. Data to train our `AutoArima` model
+2. Data to test our model
+
+For the test data we will use the last 12 months to test and evaluate
+the performance of our model.
+
+```python
+Y_train_df = df[df.ds\<='2016-08-01']
+Y_test_df = df[df.ds>'2016-08-01']
+```
+
+
+```python
+Y_train_df.shape, Y_test_df.shape
+```
+
+``` text
+((536, 3), (12, 3))
+```
+
+Now let’s plot the training data and the test data.
+
+```python
+sns.lineplot(Y_train_df,x="ds", y="y", label="Train")
+sns.lineplot(Y_test_df, x="ds", y="y", label="Test")
+plt.show()
+```
+
+
+
+# Implementation of AutoArima with StatsForecast
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import AutoARIMA
+from statsforecast.arima import arima_string
+```
+
+### Instantiating Model
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/)) by the
+master, Rob Hyndmann, can be useful.season_length
+
+```python
+season_length = 12 # Monthly data
+horizon = len(Y_test_df) # number of predictions
+
+models = [AutoARIMA(season_length=season_length)]
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See panda’s
+ available frequencies.)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models, freq='MS')
+```
+
+### Fit the Model
+
+```python
+sf.fit(df=Y_train_df)
+```
+
+``` text
+StatsForecast(models=[AutoARIMA])
+```
+
+Once we have entered our model, we can use the `arima_string` function
+to see the parameters that the model has found.
+
+```python
+arima_string(sf.fitted_[0,0].model_)
+```
+
+``` text
+'ARIMA(4,0,3)(0,1,1)[12] '
+```
+
+The automation process gave us that the best model found is a model of
+the form `ARIMA(4,0,3)(0,1,1)[12]`, this means that our model contains
+$p=4$ , that is, it has a non-seasonal autogressive element, on the
+other hand, our model contains a seasonal part, which has an order of
+$D=1$, that is, it has a seasonal differential, and $q=3$ that contains
+3 moving average element.
+
+To know the values of the terms of our model, we can use the following
+statement to know all the result of the model made.
+
+```python
+result=sf.fitted_[0,0].model_
+print(result.keys())
+print(result['arma'])
+```
+
+``` text
+dict_keys(['coef', 'sigma2', 'var_coef', 'mask', 'loglik', 'aic', 'arma', 'residuals', 'code', 'n_cond', 'nobs', 'model', 'bic', 'aicc', 'ic', 'xreg', 'x', 'lambda'])
+(4, 3, 0, 1, 12, 0, 1)
+```
+
+Let us now visualize the residuals of our models.
+
+As we can see, the result obtained above has an output in a dictionary,
+to extract each element from the dictionary we are going to use the
+`.get()` function to extract the element and then we are going to save
+it in a `pd.DataFrame()`.
+
+```python
+residual=pd.DataFrame(result.get("residuals"), columns=["residual Model"])
+residual
+```
+
+| | residual Model |
+|-----|----------------|
+| 0 | 0.085694 |
+| 1 | 0.071820 |
+| 2 | 0.066022 |
+| ... | ... |
+| 533 | 1.615486 |
+| 534 | -0.394285 |
+| 535 | -6.733548 |
+
+```python
+fig, axs = plt.subplots(nrows=2, ncols=2)
+
+# plot[1,1]
+residual.plot(ax=axs[0,0])
+axs[0,0].set_title("Residuals");
+
+# plot
+sns.distplot(residual, ax=axs[0,1]);
+axs[0,1].set_title("Density plot - Residual");
+
+# plot
+stats.probplot(residual["residual Model"], dist="norm", plot=axs[1,0])
+axs[1,0].set_title('Plot Q-Q')
+
+# plot
+plot_acf(residual, lags=35, ax=axs[1,1],color="fuchsia")
+axs[1,1].set_title("Autocorrelation");
+
+plt.show();
+```
+
+
+
+To generate forecasts we only have to use the predict method specifying
+the forecast horizon (h). In addition, to calculate prediction intervals
+associated to the forecasts, we can include the parameter level that
+receives a list of levels of the prediction intervals we want to build.
+In this case we will only calculate the 90% forecast interval
+(level=\[90\]).
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[90]` means that
+ the model expects the real value to be inside that interval 90% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min. (If you want to speed things up to a couple of seconds,
+remove the AutoModels like `ARIMA` and `Theta`)
+
+```python
+Y_hat_df = sf.forecast(df=Y_train_df, h=horizon, fitted=True)
+Y_hat_df.head()
+```
+
+| | unique_id | ds | AutoARIMA |
+|-----|-----------|------------|------------|
+| 0 | 1 | 2016-09-01 | 111.235874 |
+| 1 | 1 | 2016-10-01 | 124.948376 |
+| 2 | 1 | 2016-11-01 | 125.401639 |
+| 3 | 1 | 2016-12-01 | 123.854826 |
+| 4 | 1 | 2017-01-01 | 110.439451 |
+
+```python
+values=sf.forecast_fitted_values()
+values
+```
+
+| | unique_id | ds | y | AutoARIMA |
+|-----|-----------|------------|----------|------------|
+| 0 | 1 | 1972-01-01 | 85.6945 | 85.608806 |
+| 1 | 1 | 1972-02-01 | 71.8200 | 71.748180 |
+| 2 | 1 | 1972-03-01 | 66.0229 | 65.956878 |
+| ... | ... | ... | ... | ... |
+| 533 | 1 | 2016-06-01 | 102.4044 | 100.788914 |
+| 534 | 1 | 2016-07-01 | 102.9512 | 103.345485 |
+| 535 | 1 | 2016-08-01 | 104.6977 | 111.431248 |
+
+Adding 95% confidence interval with the forecast method
+
+```python
+sf.forecast(df=Y_train_df, h=12, level=[95])
+```
+
+| | unique_id | ds | AutoARIMA | AutoARIMA-lo-95 | AutoARIMA-hi-95 |
+|-----|-----------|------------|------------|-----------------|-----------------|
+| 0 | 1 | 2016-09-01 | 111.235874 | 104.140621 | 118.331128 |
+| 1 | 1 | 2016-10-01 | 124.948376 | 116.244661 | 133.652090 |
+| 2 | 1 | 2016-11-01 | 125.401639 | 115.882093 | 134.921185 |
+| ... | ... | ... | ... | ... | ... |
+| 9 | 1 | 2017-06-01 | 98.304446 | 85.884572 | 110.724320 |
+| 10 | 1 | 2017-07-01 | 99.630306 | 87.032356 | 112.228256 |
+| 11 | 1 | 2017-08-01 | 105.426708 | 92.639159 | 118.214258 |
+
+```python
+Y_hat_df = Y_test_df.merge(Y_hat_df, how='left', on=['unique_id', 'ds'])
+
+fig, ax = plt.subplots(1, 1, figsize = (18, 7))
+plot_df = pd.concat([Y_train_df, Y_hat_df]).set_index('ds')
+plot_df[['y', 'AutoARIMA']].plot(ax=ax, linewidth=2)
+ax.set_title(' Forecast', fontsize=22)
+ax.set_ylabel('Monthly ', fontsize=20)
+ax.set_xlabel('Timestamp [t]', fontsize=20)
+ax.legend(prop={'size': 15})
+ax.grid()
+```
+
+
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[95]` means that
+ the model expects the real value to be inside that interval 95% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+sf.predict(h=12)
+```
+
+| | unique_id | ds | AutoARIMA |
+|-----|-----------|------------|------------|
+| 0 | 1 | 2016-09-01 | 111.235874 |
+| 1 | 1 | 2016-10-01 | 124.948376 |
+| 2 | 1 | 2016-11-01 | 125.401639 |
+| ... | ... | ... | ... |
+| 9 | 1 | 2017-06-01 | 98.304446 |
+| 10 | 1 | 2017-07-01 | 99.630306 |
+| 11 | 1 | 2017-08-01 | 105.426708 |
+
+```python
+forecast_df = sf.predict(h=12, level = [80, 95])
+forecast_df
+```
+
+| | unique_id | ds | AutoARIMA | AutoARIMA-lo-95 | AutoARIMA-lo-80 | AutoARIMA-hi-80 | AutoARIMA-hi-95 |
+|-----|-----------|------------|------------|-----------------|-----------------|-----------------|-----------------|
+| 0 | 1 | 2016-09-01 | 111.235874 | 104.140621 | 106.596537 | 115.875211 | 118.331128 |
+| 1 | 1 | 2016-10-01 | 124.948376 | 116.244661 | 119.257323 | 130.639429 | 133.652090 |
+| 2 | 1 | 2016-11-01 | 125.401639 | 115.882093 | 119.177142 | 131.626136 | 134.921185 |
+| ... | ... | ... | ... | ... | ... | ... | ... |
+| 9 | 1 | 2017-06-01 | 98.304446 | 85.884572 | 90.183527 | 106.425365 | 110.724320 |
+| 10 | 1 | 2017-07-01 | 99.630306 | 87.032356 | 91.392949 | 107.867663 | 112.228256 |
+| 11 | 1 | 2017-08-01 | 105.426708 | 92.639159 | 97.065379 | 113.788038 | 118.214258 |
+
+We can join the forecast result with the historical data using the
+pandas function `pd.concat()`, and then be able to use this result for
+graphing.
+
+```python
+df_plot=pd.concat([df, forecast_df]).set_index('ds').tail(220)
+df_plot
+```
+
+| | y | unique_id | AutoARIMA | AutoARIMA-lo-95 | AutoARIMA-lo-80 | AutoARIMA-hi-80 | AutoARIMA-hi-95 |
+|------------|----------|-----------|------------|-----------------|-----------------|-----------------|-----------------|
+| ds | | | | | | | |
+| 2000-05-01 | 108.7202 | 1 | NaN | NaN | NaN | NaN | NaN |
+| 2000-06-01 | 114.2071 | 1 | NaN | NaN | NaN | NaN | NaN |
+| 2000-07-01 | 111.8737 | 1 | NaN | NaN | NaN | NaN | NaN |
+| ... | ... | ... | ... | ... | ... | ... | ... |
+| 2017-06-01 | NaN | 1 | 98.304446 | 85.884572 | 90.183527 | 106.425365 | 110.724320 |
+| 2017-07-01 | NaN | 1 | 99.630306 | 87.032356 | 91.392949 | 107.867663 | 112.228256 |
+| 2017-08-01 | NaN | 1 | 105.426708 | 92.639159 | 97.065379 | 113.788038 | 118.214258 |
+
+Now let’s visualize the result of our forecast and the historical data
+of our time series, also let’s draw the confidence interval that we have
+obtained when making the prediction with 95% confidence.
+
+```python
+sf.plot(df, forecast_df, level=[95], max_insample_length=12 * 5)
+```
+
+
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=5)`, forecasting every second months
+`(step_size=12)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 12 months ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=Y_train_df,
+ h=12,
+ step_size=12,
+ n_windows=5)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` series identifier
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the n_windows.
+- `y:` true value
+- `"model":` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df.head()
+```
+
+| | unique_id | ds | cutoff | y | AutoARIMA |
+|-----|-----------|------------|------------|----------|------------|
+| 0 | 1 | 2011-09-01 | 2011-08-01 | 93.9062 | 105.235606 |
+| 1 | 1 | 2011-10-01 | 2011-08-01 | 116.7634 | 118.739813 |
+| 2 | 1 | 2011-11-01 | 2011-08-01 | 116.8258 | 114.572924 |
+| 3 | 1 | 2011-12-01 | 2011-08-01 | 114.9563 | 114.991219 |
+| 4 | 1 | 2012-01-01 | 2011-08-01 | 99.9662 | 100.133142 |
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ Y_test_df.merge(Y_hat_df),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=season_length), ufl.rmse, ufl.smape],
+ train_df=Y_train_df,
+)
+```
+
+| | unique_id | metric | AutoARIMA |
+|-----|-----------|--------|-----------|
+| 0 | 1 | mae | 5.012894 |
+| 1 | 1 | mape | 0.045046 |
+| 2 | 1 | mase | 0.967601 |
+| 3 | 1 | rmse | 5.680362 |
+| 4 | 1 | smape | 0.022673 |
+
+## References
+
+1. [Nixtla AutoARIMA API](../../src/core/models.html#autoarima)
+2. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+
diff --git a/statsforecast/docs/models/autoces.html.mdx b/statsforecast/docs/models/autoces.html.mdx
new file mode 100644
index 00000000..fb13de64
--- /dev/null
+++ b/statsforecast/docs/models/autoces.html.mdx
@@ -0,0 +1,998 @@
+---
+title: AutoCES Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `AutoCES Model` with `Statsforecast`.
+
+The objective of the following article is to obtain a step-by-step guide
+on building the CES model using `AutoCES` with `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation` in other.
+
+The text in this article is largely taken from: 1. [Ivan Svetunkov,
+Nikolaos Kourentzes, John Keith Ord, “Complex exponential
+smoothing”](https://onlinelibrary.wiley.com/doi/full/10.1002/nav.22074)
+2. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+Principles and Practice (3rd ed)”](https://otexts.com/fpp3/tscv.html).
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [Complex Exponential Smoothing](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of AutoCES with StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+Exponential smoothing has been one of the most popular forecasting
+methods used to support various decisions in organizations, in
+activities such as inventory management, scheduling, revenue management,
+and other areas. Although its relative simplicity and transparency have
+made it very attractive for research and practice, identifying the
+underlying trend remains challenging with significant impact on the
+resulting accuracy. This has resulted in the development of various
+modifications of trend models, introducing a model selection problem.
+With the aim of addressing this problem, we propose the complex
+exponential smoothing (CES), based on the theory of functions of complex
+variables. The basic CES approach involves only two parameters and does
+not require a model selection procedure. Despite these simplifications,
+CES proves to be competitive with, or even superior to existing methods.
+We show that CES has several advantages over conventional exponential
+smoothing models: it can model and forecast both stationary and
+non-stationary processes, and CES can capture both level and trend
+cases, as defined in the conventional exponential smoothing
+classification. CES is evaluated on several forecasting competition
+datasets, demonstrating better performance than established benchmarks.
+We conclude that CES has desirable features for time series modeling and
+opens new promising avenues for research.
+
+# Complex Exponential Smoothing
+
+### Method and model
+
+Using the complex valued representation of time series, we propose the
+CES in analogy to the conventional exponential smoothing methods.
+Consider the simple exponential smoothing method:
+
+$$
+\begin{equation}
+ {\hat{y}}_t=\alpha {y}_{t-1}+\left(1-\alpha \right){\hat{y}}_{t-1} \tag{1}
+\end{equation}
+$$
+
+where $\alpha$ is the smoothing parameter and ${\hat{y}}_t$ is the
+estimated value of series. The same method can be represented as a
+weighted average of previous actual observations if we substitute
+${\hat{y}}_{t-1}$ by the formula (1) with an index instead of (Brown,
+1956):
+
+$$
+\begin{equation}
+ {\hat{y}}_t=\alpha \sum \limits_{j=1}^{t-1}{\left(1-\alpha \right)}^{j-1}{y}_{t-j} \tag{2}
+\end{equation}
+$$
+
+The idea of this representation is to demonstrate how the weights
+$\alpha {\left(1-\alpha \right)}^{j-1}$ are distributed over time in our
+sample. If the smoothing parameter $\alpha \in \left(0,1\right)$ then
+the weights decline exponentially with the increase of . If it lies in
+the so called “admissible bounds” (Brenner et al., 1968), that is
+$\alpha \in \left(0,2\right)$ then the weights decline in oscillating
+manner. Both traditional and admissible bounds have been used
+efficiently in practice and in academic literature (for application of
+the latter see for example Gardner & Diaz-Saiz, 2008; Snyder et al.,
+2017). However, in real life the distribution of weights can be more
+complex, with harmonic rather than exponential decline, meaning that
+some of the past observation might have more importance than the recent
+ones. In order to implement such distribution of weights, we build upon
+(2) and introduce complex dynamic interactions by substituting the real
+variables with the complex ones in (2). First, we substitute ${y}_{t-j}$
+by the complex variable ${y}_{t-j}+{ie}_{t-j}$, where ${e}_t$ is the
+error term of the model and $i$ is the imaginary unit (which satisfies
+the equation ${i}^2=-1$). The idea behind this is to have the impact of
+both actual values and the error on each observation in the past on the
+final forecast. Second, we substitute $\alpha$ with a complex variable
+${\alpha}_0+i{\alpha}_1$ and 1 by $1+i$ to introduce the harmonically
+declining weights. Depending on the values of the complex smoothing
+parameter, the weights distribution will exhibit a variety of
+trajectories over time, including exponential, oscillating, and
+harmonic. Finally, the result of multiplication of two complex numbers
+will be another complex number, so we substitute ${\hat{y}}_{t-j}$ with
+${\hat{y}}_{t-j}+i{\hat{e}}_{t-j}$, where ${\hat{e}}_{t-j}$ is the proxy
+for the error term. The CES obtained as a result of this can be written
+as:
+
+$$
+\begin{equation}
+ {\hat{y}}_t+i{\hat{e}}_t=\left({\alpha}_0+i{\alpha}_1\right)\sum \limits_{j=1}^{t-1}{\left(1+i-\left({\alpha}_0+i{\alpha}_1\right)\right)}^{j-1}\left({y}_{t-j}+{ie}_{t-j}\right) \tag{3}
+\end{equation}
+$$
+
+Having arrived to the model with harmonically distributed weights, we
+can now move to the shorter form by substituting
+
+$${\displaystyle \begin{array}{cc}& {\hat{y}}_{t-1}+i{\hat{e}}_{t-1}\\ {}& \kern1em =\left({\alpha}_0+i{\alpha}_1\right)\sum \limits_{j=2}^{t-1}{\left(1+i-\left({\alpha}_0+i{\alpha}_1\right)\right)}^{j-1}\left({y}_{t-j}+{ie}_{t-j}\right)\end{array}}$$
+
+in (3) to get:
+
+$$
+\begin{equation}
+ {\displaystyle \begin{array}{cc}{\hat{y}}_t+i{\hat{e}}_t& =\left({\alpha}_0+i{\alpha}_1\right)\left({y}_{t-1}+{ie}_{t-1}\right)\\ {}& \kern1em +\left(1-{\alpha}_0+i-i{\alpha}_1\right)\left({\hat{y}}_{t-1}+i{\hat{e}}_{t-1}\right).\end{array}} \tag 4
+\end{equation}
+$$
+
+Note that ${\hat{e}}_t$ is not interesting for the time series analysis
+and forecasting purposes, but is used as a vessel containing the
+information about the previous errors of the method. Having the complex
+variables instead of the real ones in (4), allows taking the
+exponentially weighted values of both actuals and the forecast errors.
+By changing the value of ${\alpha}_0+i{\alpha}_1$ , we can regulate what
+proportions of the actual and the forecast error should be carried out
+to the future in order to produce forecasts.
+
+Representing the complex-valued function as a system of two real-valued
+functions leads to:
+
+$$
+\begin{equation}
+ {\displaystyle \begin{array}{ll}& {\hat{y}}_t=\left({\alpha}_0{y}_{t-1}+\left(1-{\alpha}_0\right){\hat{y}}_{t-1}\right)-\left({\alpha}_1{e}_{t-1}+\left(1-{\alpha}_1\right){\hat{e}}_{t-1}\right)\\ {}& {\hat{e}}_t=\left({\alpha}_1{y}_{t-1}+\left(1-{\alpha}_1\right){\hat{y}}_{t-1}\right)+\left({\alpha}_0{e}_{t-1}+\left(1-{\alpha}_0\right){\hat{e}}_{t-1}\right).\end{array}} \tag 5
+\end{equation}
+$$
+
+CES introduces an interaction between the real and imaginary parts, and
+the equations in (6) are connected via the previous values of each
+other, causing interactions over time, defined by complex smoothing
+parameter value.
+
+But the method itself is restrictive and does not allow easily producing
+prediction intervals and deriving the likelihood function. It is also
+important to understand what sort of statistical model underlies CES.
+This model can be written in the following state space form:
+
+$$
+\begin{equation}
+ {\displaystyle \begin{array}{ll}& {y}_t={l}_{t-1}+{\epsilon}_t\\ {}& {l}_t={l}_{t-1}-\left(1-{\alpha}_1\right){c}_{t-1}+\left({\alpha}_0-{\alpha}_1\right){\epsilon}_t\\ {}& {c}_t={l}_{t-1}+\left(1-{\alpha}_0\right){c}_{t-1}+\left({\alpha}_0+{\alpha}_1\right){\epsilon}_t,\end{array}} \tag 6
+\end{equation}
+$$
+
+where ${\epsilon}_t$ is the white noise error term, ${l}_t$ is the level
+component and ${c}_t$ is the nonlinear trend component at observation .
+Observe that dependencies in time series have an interactive structure
+and no explicit trend component is present in the time series as this
+model does not need to artificially break the series into level and
+trend, as ETS does. Although we call the ${c}_t$ component as “nonlinear
+trend,” it does not correspond to the conventional trend component,
+because it contains the information of both previous ${c}_{t-1}$ and the
+level ${l}_{t-1}$. Also, note that we use ${\epsilon}_t$ instead of
+${e}_t$ in (6), which means that the CES has (6) as an underlying
+statistical model only when there is no misspecification error. In the
+case of the estimation of this model, the ${\epsilon}_t$ will be
+substituted by ${e}_t$, which will then lead us to the original
+formulation (4).
+
+This idea allows rewriting (6) in a shorter more generic way, resembling
+the general single source of error (SSOE) state space framework:
+
+$$
+\begin{equation}
+ {\displaystyle \begin{array}{ll}& {y}_t={\mathbf{w}}^{\prime }{\mathbf{v}}_{t-1}+{\epsilon}_t\\ {}& {\mathbf{v}}_t={\mathbf{Fv}}_{t-1}+\mathbf{g}{\epsilon}_t,\end{array}} \tag 7
+\end{equation}
+$$
+
+where
+$${\mathbf{v}}_t=\left(\begin{array}{c}{l}_t\\ {}{c}_t\end{array}\right)$$
+is the state vector
+
+$$\mathbf{F}=\left(\begin{array}{cc}1& -\left(1-{\alpha}_1\right)\\ {}1& 1-{\alpha}_0\end{array}\right)$$
+is the transition matrix
+
+$$\mathbf{g}=\left(\begin{array}{c}{\alpha}_0-{\alpha}_1\\ {}{\alpha}_0+{\alpha}_1\end{array}\right)$$
+is the persistence vector and
+
+$$\mathbf{w}=\left(\begin{array}{c}1\\ {}0\end{array}\right)$$ is the
+measurement vector.
+
+The state space form (7) permits extending CES in a similar ways to ETS
+to include additional states for seasonality or exogenous variables. The
+main difference between model (7) and the conventional ETS is that the
+transition matrix in (7) includes smoothing parameters which is not a
+standard feature of ETS models. Furthermore persistence vector includes
+the interaction of complex smoothing parameters, rather than smoothing
+parameters themselves.
+
+The error term in (6) is additive, so the likelihood function for CES is
+trivial and is similar to the one in the additive exponential smoothing
+models (Hyndman et al., 2008, p. 68):
+
+$$
+\begin{equation}
+ \mathrm{\mathcal{L}}\left(\mathbf{g},{\mathbf{v}}_0,{\sigma}^2\mid \mathbf{Y}\right)={\left(\frac{1}{\sigma \sqrt{2\pi }}\right)}^T\exp \left(-\frac{1}{2}\sum \limits_{t=1}^T{\left(\frac{\epsilon_t}{\sigma}\right)}^2\right), \tag 8
+\end{equation}
+$$
+
+where ${\mathbf{v}}_0$ is the vector of initial states, ${\sigma}^2$ is
+the variance of the error term and $\mathbf{Y}$ is the vector of all the
+in-sample observations.
+
+### Stationarity and stability conditions for CES
+
+In order to understand the properties of CES, we need to study its
+stationarity and stability conditions. The former holds for general
+exponential smoothing in the state space form (8) when all the
+eigenvalues of lie inside the unit circle (Hyndman et al., 2008, p. 38).
+CES can be either stationary or not, depending on the complex smoothing
+parameter value, in contrast to ETS models that are always
+non-stationary. Calculating eigenvalues of for CES gives the following
+roots:
+
+$$
+\begin{equation}
+ \lambda =\frac{2-{\alpha}_0\pm \sqrt{\alpha_0^2+4{\alpha}_1-4}}{2}. \tag 9
+\end{equation}
+$$
+
+If the absolute values of both roots are less than 1 then the estimated
+CES is stationary.
+
+When ${\alpha}_1>1$ one of the eigenvalues will always be greater than
+one. In this case both eigenvalues will be real numbers and CES produces
+a non-stationary trajectory. When ${\alpha}_1=1$ CES becomes equivalent
+to ETS(A,N,N). Finally, the model becomes stationary when:
+
+$$
+\begin{equation}
+ \left\{\begin{array}{l}{\alpha}_1<5-2{\alpha}_0\\ {}{\alpha}_1<1\\ {}{\alpha}_1>1-{\alpha}_0\end{array}\right. \tag {10}
+\end{equation}
+$$
+
+Note that we are not restricting CES with the conditions (10), we merely
+show, how the model will behave depending on the value of the complex
+smoothing parameter. This property of CES means that it is able to model
+either stationary or non-stationary processes, without the need to
+switch between them. The property of CES for each separate time series
+depends on the value of the smoothing parameters.
+
+The other important property that arises from (7) is the stability
+condition for CES. With $\epsilon_t={y}_t-{l}_{t-1}$ the following is
+obtained:
+
+$$
+\begin{equation}
+ {\displaystyle \begin{array}{ll}{y}_t& ={l}_{t-1}+{\epsilon}_t\\ {}\left(\begin{array}{c}{l}_t\\ {}{c}_t\end{array}\right)& =\left(\begin{array}{cc}1-{\alpha}_0+{\alpha}_1& -\left(1-{\alpha}_1\right)\\ {}1-{\alpha}_0-{\alpha}_1& 1-{\alpha}_0\end{array}\right)\left(\begin{array}{c}{l}_{t-1}\\ {}{c}_{t-1}\end{array}\right)\\ {}& \kern1em +\left(\begin{array}{c}{\alpha}_0-{\alpha}_1\\ {}{\alpha}_1+{\alpha}_0\end{array}\right){y}_t.\end{array}} \tag {11}
+\end{equation}
+$$
+
+The matrix
+$$\mathbf{D}=\left(\begin{array}{cc}1-{\alpha}_0+{\alpha}_1& -\left(1-{\alpha}_1\right)\\ {}1-{\alpha}_0-{\alpha}_1& 1-{\alpha}_0\end{array}\right)$$
+is called the discount matrix and can be written in the general form:
+
+$$
+\begin{equation}
+ \mathbf{D}=\mathbf{F}-\mathbf{g}{\mathbf{w}}^{\prime }. \tag {12}
+\end{equation}
+$$
+
+The model is said to be stable if all the eigenvalues of (12) lie inside
+the unit circle. This is more important condition than the stationarity
+for the model, because it ensures that the complex weights decline over
+time and that the older observations have smaller weights than the new
+ones, which is one of the main features of the conventional ETS models.
+The eigenvalues are given by the following formula:
+
+$$
+\begin{equation}
+ \lambda =\frac{2-2{\alpha}_0+{\alpha}_1\pm \sqrt{8{\alpha}_1+4{\alpha}_0-4{\alpha}_0{\alpha}_1-4-3{\alpha}_1^2}}{2}. \tag {13}
+\end{equation}
+$$
+
+CES will be stable when the following system of inequalities is
+satisfied:
+
+$$
+\begin{equation}
+ \left\{\begin{array}{l}{\left({\alpha}_0-2.5\right)}^2+{\alpha}_1^2>1.25\\ {}{\left({\alpha}_0-0.5\right)}^2+{\left({\alpha}_1-1\right)}^2>0.25\\ {}{\left({\alpha}_0-1.5\right)}^2+{\left({\alpha}_1-0.5\right)}^2<1.5\end{array}.\right. \tag {14}
+\end{equation}
+$$
+
+Both the stationarity and stability regions are shown in [Figure
+1](https://onlinelibrary.wiley.com/cms/asset/2a1bdd28-bbba-4ec3-aa4a-fbcf3823a209/nav22074-fig-0001-m.jpg).
+The stationarity region (10) corresponds to the triangle. All the
+combinations of smoothing parameters lying below the curve in the
+triangle will produce the stationary harmonic trajectories, while the
+rest lead to the exponential trajectories. The stability condition (14)
+corresponds to the dark region. The stability region intersects the
+stationarity region, but in general stable CES can produce both
+stationary and non-stationary forecasts
+
+### Conditional mean and variance of CES
+
+The conditional mean of CES for $h$ steps ahead with known ${l}_t$ and
+${c}_t$ can be calculated using the state space model (6):
+
+$$
+\begin{equation}
+ \mathrm{E}\left({y}_{t+h}\mid {\mathbf{v}}_t\right)={\mathbf{w}}^{\prime }{\mathbf{F}}^{h-1}{\mathbf{v}}_t, \tag {15}
+\end{equation}
+$$
+
+where
+
+$$\mathrm{E}\left({y}_{t+h}\mid {\mathbf{v}}_t\right)={\hat{y}}_{t+h}$$
+
+while $\mathbf{F}$ and $\mathbf{w}$ re the matrices from (7).
+
+The forecasting trajectories of (15) will differ depending on the values
+of ${l}_t, {c}_t$, and the complex smoothing parameter. The analysis of
+stationarity condition shows that there are several types of forecasting
+trajectories of CES depending on the particular value of the complex
+smoothing parameter:
+
+1. When ${\alpha}_1=1$ all the values of forecast will be equal to the
+ last obtained forecast, which corresponds to a flat line. This
+ trajectory is shown in [Figure
+ 2A](https://onlinelibrary.wiley.com/cms/asset/16feeb7e-adf2-48f6-9df9-cab3e34b6e67/nav22074-fig-0002-m.jpg).
+2. When ${\alpha}_1>1$ the model produces trajectory with exponential
+ growth which is shown in [Figure
+ 2B](https://onlinelibrary.wiley.com/cms/asset/16feeb7e-adf2-48f6-9df9-cab3e34b6e67/nav22074-fig-0002-m.jpg).
+3. When $\frac{4-{\alpha}_0^2}{4}<{\alpha}_1<1$ trajectory becomes
+ stationary and CES produces exponential decline shown in [Figure
+ 2C](https://onlinelibrary.wiley.com/cms/asset/16feeb7e-adf2-48f6-9df9-cab3e34b6e67/nav22074-fig-0002-m.jpg).
+4. When $1-{\alpha}_0<{\alpha}_1<\frac{4-{\alpha}_0^2}{4}$ trajectory
+ becomes harmonic and will converge to zero [Figure
+ 2D](https://onlinelibrary.wiley.com/cms/asset/16feeb7e-adf2-48f6-9df9-cab3e34b6e67/nav22074-fig-0002-m.jpg).
+5. Finally, when $0<{\alpha}_1<1-{\alpha}_0$ the diverging harmonic
+ trajectory is produced, the model becomes non-stationary. This
+ trajectory is of no use in forecasting, that is why we do not show
+ it on graphs.
+
+Using (7) the conditional variance of CES for $h$ steps ahead with known
+${l}_t$ and ${c}_t$ can be calculated similarly to the pure additive ETS
+models (Hyndman et al., 2008, p. 96).
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import pandas as pd
+
+import scipy.stats as stats
+```
+
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf
+from statsmodels.graphics.tsaplots import plot_pacf
+
+plt.style.use('fivethirtyeight')
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#212946',
+ 'axes.facecolor': '#212946',
+ 'savefig.facecolor':'#212946',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#2A3459',
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+### Read Data
+
+```python
+df = pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/Esperanza_vida.csv", usecols=[1,2])
+df.head()
+```
+
+| | year | value |
+|-----|------------|-----------|
+| 0 | 1960-01-01 | 69.123902 |
+| 1 | 1961-01-01 | 69.760244 |
+| 2 | 1962-01-01 | 69.149756 |
+| 3 | 1963-01-01 | 69.248049 |
+| 4 | 1964-01-01 | 70.311707 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|------------|-----------|-----------|
+| 0 | 1960-01-01 | 69.123902 | 1 |
+| 1 | 1961-01-01 | 69.760244 | 1 |
+| 2 | 1962-01-01 | 69.149756 | 1 |
+| 3 | 1963-01-01 | 69.248049 | 1 |
+| 4 | 1964-01-01 | 70.311707 | 1 |
+
+Now, let’s now check the last few rows of our time series using the
+`.tail()` function.
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y float64
+unique_id object
+dtype: object
+```
+
+We need to convert the `ds` from `object` type to datetime.
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints a random series from the dataset and is useful for
+basic EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### Autocorrelation plots
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=20, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+# Grafico
+plot_pacf(df["y"], lags=20, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+#plt.savefig("Gráfico de Densidad y qq")
+plt.show();
+```
+
+
+
+### Decomposition of the time series
+
+How to decompose a time series and why?
+
+In time series analysis to forecast new values, it is very important to
+know past data. More formally, we can say that it is very important to
+know the patterns that values follow over time. There can be many
+reasons that cause our forecast values to fall in the wrong direction.
+Basically, a time series consists of four components. The variation of
+those components causes the change in the pattern of the time series.
+These components are:
+
+- **Level:** This is the primary value that averages over time.
+- **Trend:** The trend is the value that causes increasing or
+ decreasing patterns in a time series.
+- **Seasonality:** This is a cyclical event that occurs in a time
+ series for a short time and causes short-term increasing or
+ decreasing patterns in a time series.
+- **Residual/Noise:** These are the random variations in the time
+ series.
+
+Combining these components over time leads to the formation of a time
+series. Most time series consist of level and noise/residual and trend
+or seasonality are optional values.
+
+If seasonality and trend are part of the time series, then there will be
+effects on the forecast value. As the pattern of the forecasted time
+series may be different from the previous time series.
+
+The combination of the components in time series can be of two types: \*
+Additive \* Multiplicative
+
+### Additive time series
+
+If the components of the time series are added to make the time series.
+Then the time series is called the additive time series. By
+visualization, we can say that the time series is additive if the
+increasing or decreasing pattern of the time series is similar
+throughout the series. The mathematical function of any additive time
+series can be represented by:
+$$y(t) = level + Trend + seasonality + noise$$
+
+### Multiplicative time series
+
+If the components of the time series are multiplicative together, then
+the time series is called a multiplicative time series. For
+visualization, if the time series is having exponential growth or
+decline with time, then the time series can be considered as the
+multiplicative time series. The mathematical function of the
+multiplicative time series can be represented as.
+
+$$y(t) = Level * Trend * seasonality * Noise$$
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+a = seasonal_decompose(df["y"], model = "add", period=1)
+a.plot();
+```
+
+
+
+## Split the data into training and testing
+
+Let’s divide our data into sets
+
+1. Data to train our model.
+2. Data to test our model.
+
+For the test data we will use the last 12 months to test and evaluate
+the performance of our model.
+
+```python
+train = df[df.ds\<='2013-01-01']
+test = df[df.ds>'2013-01-01']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((54, 3), (6, 3))
+```
+
+Now let’s plot the training data and the test data.
+
+```python
+sns.lineplot(train,x="ds", y="y", label="Train")
+sns.lineplot(test, x="ds", y="y", label="Test")
+plt.show()
+```
+
+
+
+## Implementation of AutoCES with StatsForecast
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import AutoCES
+```
+
+### Instantiate Model
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/)) by the
+master, Rob Hyndmann, can be useful `season_length`
+
+**Note**
+
+Automatically selects the best `Complex Exponential Smoothing model`
+using an information criterion. Default is Akaike Information Criterion
+(AICc), while particular models are estimated using maximum likelihood.
+The state-space equations can be determined based on their $S$ simple,
+$P$ parial, $Z$ optimized or $N$ ommited components. The model string
+parameter defines the kind of CES model: $N$ for simple CES (withous
+seasonality), $S$ for simple seasonality (lagged CES), $P$ for partial
+seasonality (without complex part), $F$ for full seasonality (lagged CES
+with real and complex seasonal parts).
+
+If the component is selected as $Z$, it operates as a placeholder to ask
+the AutoCES model to figure out the best parameter.
+
+```python
+season_length = 1 # year data
+horizon = len(test) # number of predictions
+
+# We call the model that we are going to use
+models = [AutoCES(season_length=season_length)]
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [pandas’
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models, freq='YS')
+```
+
+### Fit the Model
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[CES])
+```
+
+```python
+result=sf.fitted_[0,0].model_
+print(result.keys())
+print(result['fit'])
+```
+
+``` text
+dict_keys(['loglik', 'aic', 'bic', 'aicc', 'mse', 'amse', 'fit', 'fitted', 'residuals', 'm', 'states', 'par', 'n', 'seasontype', 'sigma2', 'actual_residuals'])
+results(x=array([1.63706552, 1.00511519]), fn=76.78049826760919, nit=27, simplex=array([[1.63400329, 1.00510199],
+ [1.63706552, 1.00511519],
+ [1.63638944, 1.00512037]]))
+```
+
+Let us now visualize the residuals of our models.
+
+As we can see, the result obtained above has an output in a dictionary,
+to extract each element from the dictionary we are going to use the
+`.get()` function to extract the element and then we are going to save
+it in a `pd.DataFrame()`.
+
+```python
+residual=pd.DataFrame(result.get("residuals"), columns=["residual Model"])
+residual
+```
+
+| | residual Model |
+|-----|----------------|
+| 0 | -0.727729 |
+| 1 | 0.144552 |
+| 2 | -0.762086 |
+| ... | ... |
+| 51 | -0.073258 |
+| 52 | -0.234578 |
+| 53 | 0.109990 |
+
+```python
+fig, axs = plt.subplots(nrows=2, ncols=2)
+
+# plot[1,1]
+residual.plot(ax=axs[0,0])
+axs[0,0].set_title("Residuals");
+
+# plot
+sns.distplot(residual, ax=axs[0,1]);
+axs[0,1].set_title("Density plot - Residual");
+
+# plot
+stats.probplot(residual["residual Model"], dist="norm", plot=axs[1,0])
+axs[1,0].set_title('Plot Q-Q')
+
+# plot
+plot_acf(residual, lags=35, ax=axs[1,1],color="fuchsia")
+axs[1,1].set_title("Autocorrelation");
+
+plt.show();
+```
+
+
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[90]` means that
+ the model expects the real value to be inside that interval 90% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min. (If you want to speed things up to a couple of seconds,
+remove the AutoModels like `ARIMA` and `Theta`)
+
+```python
+# Prediction
+Y_hat = sf.forecast(df=train, h=horizon, fitted=True)
+Y_hat
+```
+
+| | unique_id | ds | CES |
+|-----|-----------|------------|-----------|
+| 0 | 1 | 2014-01-01 | 82.906075 |
+| 1 | 1 | 2015-01-01 | 83.166687 |
+| 2 | 1 | 2016-01-01 | 83.424744 |
+| 3 | 1 | 2017-01-01 | 83.685760 |
+| 4 | 1 | 2018-01-01 | 83.946213 |
+| 5 | 1 | 2019-01-01 | 84.208359 |
+
+```python
+values=sf.forecast_fitted_values()
+values.head()
+```
+
+| | unique_id | ds | y | CES |
+|-----|-----------|------------|-----------|-----------|
+| 0 | 1 | 1960-01-01 | 69.123902 | 69.851631 |
+| 1 | 1 | 1961-01-01 | 69.760244 | 69.615692 |
+| 2 | 1 | 1962-01-01 | 69.149756 | 69.911842 |
+| 3 | 1 | 1963-01-01 | 69.248049 | 69.657822 |
+| 4 | 1 | 1964-01-01 | 70.311707 | 69.601196 |
+
+```python
+StatsForecast.plot(values)
+```
+
+
+
+Adding 95% confidence interval with the forecast method
+
+```python
+sf.forecast(df=train, h=horizon, level=[95])
+```
+
+| | unique_id | ds | CES | CES-lo-95 | CES-hi-95 |
+|-----|-----------|------------|-----------|-----------|-----------|
+| 0 | 1 | 2014-01-01 | 82.906075 | 82.342483 | 83.454016 |
+| 1 | 1 | 2015-01-01 | 83.166687 | 82.604029 | 83.717271 |
+| 2 | 1 | 2016-01-01 | 83.424744 | 82.858573 | 83.975870 |
+| 3 | 1 | 2017-01-01 | 83.685760 | 83.118946 | 84.239582 |
+| 4 | 1 | 2018-01-01 | 83.946213 | 83.376905 | 84.501133 |
+| 5 | 1 | 2019-01-01 | 84.208359 | 83.637738 | 84.765408 |
+
+```python
+# Merge the forecasts with the true values
+Y_hat = test.merge(Y_hat, how='left', on=['unique_id', 'ds'])
+Y_hat
+```
+
+| | ds | y | unique_id | CES |
+|-----|------------|-----------|-----------|-----------|
+| 0 | 2014-01-01 | 83.090244 | 1 | 82.906075 |
+| 1 | 2015-01-01 | 82.543902 | 1 | 83.166687 |
+| 2 | 2016-01-01 | 83.243902 | 1 | 83.424744 |
+| 3 | 2017-01-01 | 82.946341 | 1 | 83.685760 |
+| 4 | 2018-01-01 | 83.346341 | 1 | 83.946213 |
+| 5 | 2019-01-01 | 83.197561 | 1 | 84.208359 |
+
+```python
+sf.plot(train, Y_hat)
+```
+
+
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[95]` means that
+ the model expects the real value to be inside that interval 95% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+sf.predict(h=horizon)
+```
+
+| | unique_id | ds | CES |
+|-----|-----------|------------|-----------|
+| 0 | 1 | 2014-01-01 | 82.906075 |
+| 1 | 1 | 2015-01-01 | 83.166687 |
+| 2 | 1 | 2016-01-01 | 83.424744 |
+| 3 | 1 | 2017-01-01 | 83.685760 |
+| 4 | 1 | 2018-01-01 | 83.946213 |
+| 5 | 1 | 2019-01-01 | 84.208359 |
+
+```python
+forecast_df = sf.predict(h=horizon, level=[95])
+forecast_df
+```
+
+| | unique_id | ds | CES | CES-lo-95 | CES-hi-95 |
+|-----|-----------|------------|-----------|-----------|-----------|
+| 0 | 1 | 2014-01-01 | 82.906075 | 82.342483 | 83.454016 |
+| 1 | 1 | 2015-01-01 | 83.166687 | 82.604029 | 83.717271 |
+| 2 | 1 | 2016-01-01 | 83.424744 | 82.858573 | 83.975870 |
+| 3 | 1 | 2017-01-01 | 83.685760 | 83.118946 | 84.239582 |
+| 4 | 1 | 2018-01-01 | 83.946213 | 83.376905 | 84.501133 |
+| 5 | 1 | 2019-01-01 | 84.208359 | 83.637738 | 84.765408 |
+
+Now let’s visualize the result of our forecast and the historical data
+of our time series, also let’s draw the confidence interval that we have
+obtained when making the prediction with 95% confidence.
+
+```python
+sf.plot(train, test.merge(forecast_df), level=[95])
+```
+
+
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=5)`, forecasting every second months
+`(step_size=12)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 12 months ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=train,
+ h=horizon,
+ step_size=12,
+ n_windows=3)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` series identifier
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the n_windows.
+- `y:` true value
+- `"model":` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df.head()
+```
+
+| | unique_id | ds | cutoff | y | CES |
+|-----|-----------|------------|------------|-----------|-----------|
+| 0 | 1 | 1984-01-01 | 1983-01-01 | 75.389512 | 74.952705 |
+| 1 | 1 | 1985-01-01 | 1983-01-01 | 75.470732 | 75.161736 |
+| 2 | 1 | 1986-01-01 | 1983-01-01 | 75.770732 | 75.377945 |
+| 3 | 1 | 1987-01-01 | 1983-01-01 | 76.219512 | 75.590378 |
+| 4 | 1 | 1988-01-01 | 1983-01-01 | 76.370732 | 75.806343 |
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ Y_hat,
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=season_length), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | CES |
+|-----|-----------|--------|----------|
+| 0 | 1 | mae | 0.556314 |
+| 1 | 1 | mape | 0.006699 |
+| 2 | 1 | mase | 1.770512 |
+| 3 | 1 | rmse | 0.630183 |
+| 4 | 1 | smape | 0.003336 |
+
+## References
+
+1. [Nixtla AutoCES API](../../src/core/models.html#autoces)
+2. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+3. [Ivan Svetunkov, Nikolaos Kourentzes, John Keith Ord, “Complex
+ exponential
+ smoothing”](https://onlinelibrary.wiley.com/doi/full/10.1002/nav.22074)
+
diff --git a/statsforecast/docs/models/autoets.html.mdx b/statsforecast/docs/models/autoets.html.mdx
new file mode 100644
index 00000000..e5068aed
--- /dev/null
+++ b/statsforecast/docs/models/autoets.html.mdx
@@ -0,0 +1,731 @@
+---
+title: AutoETS Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `AutoETS Model` with `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation` in other.
+
+The text in this article is largely taken from [Rob J. Hyndman and
+George Athanasopoulos (2018). “Forecasting Principles and Practice (3rd
+ed)”](https://otexts.com/fpp3/tscv.html).
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [ETS Models](#model)
+- [ETS Estimation](#estimation)
+- [Model Selection](#selection)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of AutoETS with StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+Automatic forecasts of large numbers of univariate time series are often
+needed in business. It is common to have over one thousand product lines
+that need forecasting at least monthly. Even when a smaller number of
+forecasts are required, there may be nobody suitably trained in the use
+of time series models to produce them. In these circumstances, an
+automatic forecasting algorithm is an essential tool. Automatic
+forecasting algorithms must determine an appropriate time series model,
+estimate the parameters and compute the forecasts. They must be robust
+to unusual time series patterns, and applicable to large numbers of
+series without user intervention. The most popular automatic forecasting
+algorithms are based on either exponential smoothing or ARIMA models.
+
+# Exponential smoothing
+
+Although exponential smoothing methods have been around since the 1950s,
+a modelling framework incorporating procedures for model selection was
+not developed until relatively recently. `Ord, Koehler`, and
+`Snyder (1997), Hyndman, Koehler, Snyder, and Grose (2002)` and
+`Hyndman, Koehler, Ord`, and `Snyder (2005b)` have shown that all
+exponential smoothing methods (including non-linear methods) are optimal
+forecasts from innovations state space models.
+
+Exponential smoothing methods were originally classified by
+`Pegels’ (1969) taxonomy`. This was later extended by
+`Gardner (1985), modified by Hyndman et al. (2002)`, and extended again
+by `Taylor (2003)`, giving a total of fifteen methods seen in the
+following table.
+
+| Trend Component | Seasonal Component |
+|-----------------|--------------------|
+
+| Component | N(None)) | A (Additive) | M (Multiplicative) |
+|----------------------------|----------|--------------|--------------------|
+| N (None) | (N,N) | (N,A) | (N,M) |
+| A (Additive) | (A,N) | (A,A) | (A,M) |
+| Ad (Additive damped) | (Ad,N) | (Ad,A) | (Ad,M) |
+| M (Multiplicative) | (M,N ) | (M,A ) | (M,M) |
+| Md (Multiplicative damped) | (Md,N ) | ( Md,A) | (Md,M) |
+
+Some of these methods are better known under other names. For example,
+cell `(N,N)` describes the **simple exponential smoothing (or SES)
+method**, cell `(A,N)` describes **Holt’s linear method**, and cell
+`(Ad,N)` describes **the damped trend method**. The **additive
+Holt-Winters’ method** is given by cell `(A,A)` and the **multiplicative
+Holt-Winters’ method** is given by cell `(A,M)`. The other cells
+correspond to less commonly used but analogous methods.
+
+### Point forecasts for all methods
+
+We denote the observed time series by $y_1,y_2,...,y_n$. A forecast of
+$y_{t+h}$ based on all of the data up to time $t$ is denoted by
+$\hat y_{t+h|t}$. To illustrate the method, we give the point forecasts
+and updating equation for method `(A,A)`, the he Holt-Winters’ additive
+method:
+
+where $m$ is the length of seasonality (e.g., the number of months or
+quarters in a year), $\ell_{t}$ represents the level of the series,
+$b_t$ denotes the growth, $s_t$ is the seasonal component,
+$\hat y_{t+h|t}$ is the forecast for $h$ periods ahead, and
+$h_{m}^{+} = [(h − 1) mod \ m] + 1$. To use method (1), we need values
+for the initial states $\ell_{0}$, $b_0$ and $s_{1−m}, . . . , s_0$, and
+for the smoothing parameters $\alpha, \beta^{*}$ and $\gamma$. All of
+these will be estimated from the observed data.
+
+Equation (1c) is slightly different from the usual Holt-Winters
+equations such as those in Makridakis et al. (1998) or Bowerman,
+O’Connell, and Koehler (2005). These authors replace (1c) with
+$$s_{t} = \gamma^* (y_{t}-\ell_{t})+ (1-\gamma^*)s_{t-m}.$$
+
+If $\ell_{t}$ is substituted using (1a), we obtain
+
+$$s_{t} = \gamma^*(1-\alpha) (y_{t}-\ell_{t-1}-b_{t-1})+ [1-\gamma^*(1-\alpha)]s_{t-m},$$
+
+Thus, we obtain identical forecasts using this approach by replacing
+$\gamma$ in (1c) with $\gamma^{*} (1-\alpha)$. The modification given in
+(1c) was proposed by Ord et al. (1997) to make the state space
+formulation simpler. It is equivalent to Archibald’s (1990) variation of
+the Holt-Winters’ method.
+
+### Innovations state space models
+
+For each exponential smoothing method in [Other ETS
+models](https://otexts.com/fpp3/ets.html) , Hyndman et al. (2008b)
+describe two possible innovations state space models, one corresponding
+to a model with additive errors and the other to a model with
+multiplicative errors. If the same parameter values are used, these two
+models give equivalent point forecasts, although different prediction
+intervals. Thus there are 30 potential models described in this
+classification.
+
+Historically, the nature of the error component has often been ignored,
+because the distinction between additive and multiplicative errors makes
+no difference to point forecasts.
+
+We are careful to distinguish exponential smoothing methods from the
+underlying state space models. An exponential smoothing method is an
+algorithm for producing point forecasts only. The underlying stochastic
+state space model gives the same point forecasts, but also provides a
+framework for computing prediction intervals and other properties.
+
+To distinguish the models with additive and multiplicative errors, we
+add an extra letter to the front of the method notation. The triplet
+`(E,T,S)` refers to the three components: error, trend and seasonality.
+So the model `ETS(A,A,N)` has additive errors, additive trend and no
+seasonality—in other words, this is Holt’s linear method with additive
+errors. Similarly, `ETS(M,Md,M)` refers to a model with multiplicative
+errors, a damped multiplicative trend and multiplicative seasonality.
+The notation `ETS(·,·,·)` helps in remembering the order in which the
+components are specified.
+
+Once a model is specified, we can study the probability distribution of
+future values of the series and find, for example, the conditional mean
+of a future observation given knowledge of the past. We denote this as
+$\mu_{t+h|t} = E(y_{t+h | xt})$, where xt contains the unobserved
+components such as $\ell_t$, $b_t$ and $s_t$. For $h = 1$ we use
+$\mu_t ≡ \mu_{t+1|t}$ as a shorthand notation. For many models, these
+conditional means will be identical to the point forecasts given in
+Table [Other ETS models](https://otexts.com/fpp3/ets.html), so that
+$\mu_{t+h|t} = \hat y_{t+h|t}$. However, for other models (those with
+multiplicative trend or multiplicative seasonality), the conditional
+mean and the point forecast will differ slightly for $h ≥ 2$.
+
+We illustrate these ideas using the damped trend method of Gardner and
+McKenzie (1985).
+
+Each model consists of a measurement equation that describes the
+observed data, and some state equations that describe how the unobserved
+components or states (level, trend, seasonal) change over time. Hence,
+these are referred to as state space models.
+
+For each method there exist two models: one with additive errors and one
+with multiplicative errors. The point forecasts produced by the models
+are identical if they use the same smoothing parameter values. They
+will, however, generate different prediction intervals.
+
+To distinguish between a model with additive errors and one with
+multiplicative errors (and also to distinguish the models from the
+methods), we add a third letter to the classification of in the above
+Table. We label each state space model as `ETS(⋅,.,.)` for (Error,
+Trend, Seasonal). This label can also be thought of as ExponenTial
+Smoothing. Using the same notation as in the above Table, the
+possibilities for each component (or state) are: `Error ={ A,M }`,
+`Trend ={N,A,Ad}` and `Seasonal ={ N,A,M }`.
+
+### **ETS(A,N,N): simple exponential smoothing with additive errors**
+
+Recall the component form of simple exponential smoothing:
+
+If we re-arrange the smoothing equation for the level, we get the “error
+correction” form,
+
+where $e_{t}=y_{t}-\ell_{t-1}=y_{t}-\hat{y}_{t|t-1}$ s the residual at
+time $t$.
+
+The training data errors lead to the adjustment of the estimated level
+throughout the smoothing process for $t=1,\dots,T$. For example, if the
+error at time $t$ is negative, then $y_t < \hat{y}_{t|t-1}$ and so the
+level at time $t-1$ has been over-estimated. The new level $\ell_{t}$ is
+then the previous level $\ell_{t-1}$ adjusted downwards. The closer
+$\alpha$ is to one, the “rougher” the estimate of the level (large
+adjustments take place). The smaller the $\alpha$, the “smoother” the
+level (small adjustments take place).
+
+We can also write $y_t = \ell_{t-1} + e_t$, so that each observation can
+be represented by the previous level plus an error. To make this into an
+innovations state space model, all we need to do is specify the
+probability distribution for $e_t$. For a model with additive errors, we
+assume that residuals (the one-step training errors) $e_t$ are normally
+distributed white noise with mean 0 and variance $\sigma^2$. A
+short-hand notation for this is
+$e_t = \varepsilon_t\sim\text{NID}(0,\sigma^2)$; NID stands for
+“normally and independently distributed”.
+
+Then the equations of the model can be written as
+
+We refer to (2) as the measurement (or observation) equation and (3) as
+the state (or transition) equation. These two equations, together with
+the statistical distribution of the errors, form a fully specified
+statistical model. Specifically, these constitute an innovations state
+space model underlying simple exponential smoothing.
+
+The term “innovations” comes from the fact that all equations use the
+same random error process, $\varepsilon_t$. For the same reason, this
+formulation is also referred to as a “single source of error” model.
+There are alternative multiple source of error formulations which we do
+not present here.
+
+The measurement equation shows the relationship between the observations
+and the unobserved states. In this case, observation $y_t$ is a linear
+function of the level $\ell_{t-1}$, the predictable part of $y_t$, and
+the error $\varepsilon_t$, the unpredictable part of $y_t$. For other
+innovations state space models, this relationship may be nonlinear.
+
+The state equation shows the evolution of the state through time. The
+influence of the smoothing parameter $\alpha$ is the same as for the
+methods discussed earlier. For example, $\alpha$ governs the amount of
+change in successive levels: high values of $\alpha$ allow rapid changes
+in the level; low values of $\alpha$ lead to smooth changes. If
+$\alpha=0$, the level of the series does not change over time; if
+$\alpha=1$, the model reduces to a random walk model,
+$y_t=y_{t-1}+\varepsilon_t$.
+
+### **ETS(M,N,N): simple exponential smoothing with multiplicative errors**
+
+In a similar fashion, we can specify models with multiplicative errors
+by writing the one-step-ahead training errors as relative errors
+
+$$\varepsilon_t = \frac{y_t-\hat{y}_{t|t-1}}{\hat{y}_{t|t-1}}$$
+
+where $\varepsilon_t \sim \text{NID}(0,\sigma^2)$. Substituting
+$\hat{y}_{t|t-1}=\ell_{t-1}$ gives
+$y_t = \ell_{t-1}+\ell_{t-1}\varepsilon_t$ and
+$e_t = y_t - \hat{y}_{t|t-1} = \ell_{t-1}\varepsilon_t$.
+
+Then we can write the multiplicative form of the state space model as
+
+### **ETS(A,A,N): Holt’s linear method with additive errors**
+
+For this model, we assume that the one-step-ahead training errors are
+given by
+
+$$\varepsilon_t=y_t-\ell_{t-1}-b_{t-1} \sim \text{NID}(0,\sigma^2)$$
+
+Substituting this into the error correction equations for Holt’s linear
+method we obtain
+
+where for simplicity we have set $\beta=\alpha \beta^*$.
+
+### **ETS(M,A,N): Holt’s linear method with multiplicative errors**
+
+Specifying one-step-ahead training errors as relative errors such that
+
+$$\varepsilon_t=\frac{y_t-(\ell_{t-1}+b_{t-1})}{(\ell_{t-1}+b_{t-1})}$$
+
+and following an approach similar to that used above, the innovations
+state space model underlying Holt’s linear method with multiplicative
+errors is specified as
+
+where again $\beta=\alpha \beta^*$ and
+$\varepsilon_t \sim \text{NID}(0,\sigma^2)$.
+
+## Estimating ETS models
+
+An alternative to estimating the parameters by minimising the sum of
+squared errors is to maximise the “likelihood”. The likelihood is the
+probability of the data arising from the specified model. Thus, a large
+likelihood is associated with a good model. For an additive error model,
+maximising the likelihood (assuming normally distributed errors) gives
+the same results as minimising the sum of squared errors. However,
+different results will be obtained for multiplicative error models. In
+this section, we will estimate the smoothing parameters
+$\alpha, \beta, \gamma$ and $\phi$ and the initial states
+$\ell_0, b_0, s_0,s_{-1},\dots,s_{-m+1}$, by maximising the likelihood.
+
+The possible values that the smoothing parameters can take are
+restricted. Traditionally, the parameters have been constrained to lie
+between 0 and 1 so that the equations can be interpreted as weighted
+averages. That is, $0< \alpha,\beta^*,\gamma^*,\phi<1$. For the state
+space models, we have set $\beta=\alpha\beta^*$ and
+$\gamma=(1-\alpha)\gamma^*$. Therefore, the traditional restrictions
+translate to $0< \alpha <1, 0 < \beta < \alpha$ and
+$0< \gamma < 1-\alpha$. In practice, the damping parameter $\phi$ is
+usually constrained further to prevent numerical difficulties in
+estimating the model.
+
+Another way to view the parameters is through a consideration of the
+mathematical properties of the state space models. The parameters are
+constrained in order to prevent observations in the distant past having
+a continuing effect on current forecasts. This leads to some
+admissibility constraints on the parameters, which are usually (but not
+always) less restrictive than the traditional constraints region
+`(Hyndman et al., 2008, pp. 149-161)`. For example, for the `ETS(A,N,N)`
+model, the traditional parameter region is $0< \alpha <1$ but the
+admissible region is $0< \alpha <2$. For the `ETS(A,A,N)` model, the
+traditional parameter region is $0<\alpha<1$ and $0<\beta<\alpha$ but
+the admissible region is $0<\alpha<2$ and $0<\beta<4-2\alpha$.
+
+## Model selection
+
+A great advantage of the `ETS` statistical framework is that information
+criteria can be used for model selection. The `AIC, AIC_c` and `BIC`,
+can be used here to determine which of the `ETS` models is most
+appropriate for a given time series.
+
+For `ETS` models, Akaike’s Information Criterion (`AIC)` is defined as
+
+$$\text{AIC} = -2\log(L) + 2k,$$
+
+where $L$ is the likelihood of the model and $k$ is the total number of
+parameters and initial states that have been estimated (including the
+residual variance).
+
+The `AIC` corrected for small sample bias `(AIC_c)` is defined as
+
+$$AIC_c = AIC + \frac{2k(k+1)}{T-k-1}$$
+
+and the Bayesian Information Criterion `(BIC)` is
+
+$$\text{BIC} = \text{AIC} + k[\log(T)-2]$$
+
+Three of the combinations of (Error, Trend, Seasonal) can lead to
+numerical difficulties. Specifically, the models that can cause such
+instabilities are `ETS(A,N,M), ETS(A,A,M)`, and `ETS(A,Ad,M)`, due to
+division by values potentially close to zero in the state equations. We
+normally do not consider these particular combinations when selecting a
+model.
+
+Models with multiplicative errors are useful when the data are strictly
+positive, but are not numerically stable when the data contain zeros or
+negative values. Therefore, multiplicative error models will not be
+considered if the time series is not strictly positive. In that case,
+only the six fully additive models will be applied.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import numpy as np
+import pandas as pd
+```
+
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf
+from statsmodels.graphics.tsaplots import plot_pacf
+
+plt.style.use('fivethirtyeight')
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#212946',
+ 'axes.facecolor': '#212946',
+ 'savefig.facecolor':'#212946',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#2A3459',
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+### Read Data
+
+```python
+df = pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/Esperanza_vida.csv", usecols=[1,2])
+df.head()
+```
+
+| | year | value |
+|-----|------------|-----------|
+| 0 | 1960-01-01 | 69.123902 |
+| 1 | 1961-01-01 | 69.760244 |
+| 2 | 1962-01-01 | 69.149756 |
+| 3 | 1963-01-01 | 69.248049 |
+| 4 | 1964-01-01 | 70.311707 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|------------|-----------|-----------|
+| 0 | 1960-01-01 | 69.123902 | 1 |
+| 1 | 1961-01-01 | 69.760244 | 1 |
+| 2 | 1962-01-01 | 69.149756 | 1 |
+| 3 | 1963-01-01 | 69.248049 | 1 |
+| 4 | 1964-01-01 | 70.311707 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y float64
+unique_id object
+dtype: object
+```
+
+We need to convert the `ds` from `object` type to datetime.
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints a random series from the dataset and is useful for
+basic EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+## Autocorrelation plots
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=20, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+# Plot
+plot_pacf(df["y"], lags=20, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+plt.show();
+```
+
+
+
+### Decomposition of the time series
+
+How to decompose a time series and why?
+
+In time series analysis to forecast new values, it is very important to
+know past data. More formally, we can say that it is very important to
+know the patterns that values follow over time. There can be many
+reasons that cause our forecast values to fall in the wrong direction.
+Basically, a time series consists of four components. The variation of
+those components causes the change in the pattern of the time series.
+These components are:
+
+- **Level:** This is the primary value that averages over time.
+- **Trend:** The trend is the value that causes increasing or
+ decreasing patterns in a time series.
+- **Seasonality:** This is a cyclical event that occurs in a time
+ series for a short time and causes short-term increasing or
+ decreasing patterns in a time series.
+- **Residual/Noise:** These are the random variations in the time
+ series.
+
+Combining these components over time leads to the formation of a time
+series. Most time series consist of level and noise/residual and trend
+or seasonality are optional values.
+
+If seasonality and trend are part of the time series, then there will be
+effects on the forecast value. As the pattern of the forecasted time
+series may be different from the previous time series.
+
+The combination of the components in time series can be of two types: \*
+Additive \* Multiplicative
+
+### Additive time series
+
+If the components of the time series are added to make the time series.
+Then the time series is called the additive time series. By
+visualization, we can say that the time series is additive if the
+increasing or decreasing pattern of the time series is similar
+throughout the series. The mathematical function of any additive time
+series can be represented by:
+$$y(t) = level + Trend + seasonality + noise$$
+
+### Multiplicative time series
+
+If the components of the time series are multiplicative together, then
+the time series is called a multiplicative time series. For
+visualization, if the time series is having exponential growth or
+decline with time, then the time series can be considered as the
+multiplicative time series. The mathematical function of the
+multiplicative time series can be represented as.
+
+$$y(t) = Level * Trend * seasonality * Noise$$
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+a = seasonal_decompose(df["y"], model = "add", period=1)
+a.plot();
+```
+
+
+
+Breaking down a time series into its components helps us to identify the
+behavior of the time series we are analyzing. In addition, it helps us
+to know what type of models we can apply, for our example of the Life
+expectancy data set, we can observe that our time series shows an
+increasing trend throughout the year, on the other hand, it can be
+observed also that the time series has no seasonality.
+
+By looking at the previous graph and knowing each of the components, we
+can get an idea of which model we can apply: \* We have trend \* There
+is no seasonality
+
+## Split the data into training and testing
+
+Let’s divide our data into sets
+
+1. Data to train our model.
+2. Data to test our model.
+
+For the test data we will use the last 6 years to test and evaluate the
+performance of our model.
+
+```python
+train = df[df.ds\<='2013-01-01']
+test = df[df.ds>'2013-01-01']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((54, 3), (6, 3))
+```
+
+```python
+sns.lineplot(train,x="ds", y="y", label="Train")
+sns.lineplot(test, x="ds", y="y", label="Test")
+plt.show()
+```
+
+
+
+## Implementation of AutoETS with StatsForecast
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import AutoETS
+```
+
+### Instantiate Model
+
+```python
+sf = StatsForecast(models=[AutoETS(model="AZN")], freq='YS')
+```
+
+### Fit the Model
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[AutoETS])
+```
+
+### Model Prediction
+
+```python
+y_hat = sf.predict(h=6)
+y_hat
+```
+
+| | unique_id | ds | AutoETS |
+|-----|-----------|------------|-----------|
+| 0 | 1 | 2014-01-01 | 82.952553 |
+| 1 | 1 | 2015-01-01 | 83.146150 |
+| 2 | 1 | 2016-01-01 | 83.339747 |
+| 3 | 1 | 2017-01-01 | 83.533344 |
+| 4 | 1 | 2018-01-01 | 83.726940 |
+| 5 | 1 | 2019-01-01 | 83.920537 |
+
+```python
+sf.plot(train, y_hat)
+```
+
+
+
+Let’s add a confidence interval to our forecast.
+
+```python
+y_hat = sf.predict(h=6, level=[80,90,95])
+y_hat
+```
+
+| | unique_id | ds | AutoETS | AutoETS-lo-95 | AutoETS-lo-90 | AutoETS-lo-80 | AutoETS-hi-80 | AutoETS-hi-90 | AutoETS-hi-95 |
+|-----|-----------|------------|-----------|---------------|---------------|---------------|---------------|---------------|---------------|
+| 0 | 1 | 2014-01-01 | 82.952553 | 82.500416 | 82.573107 | 82.656916 | 83.248190 | 83.331999 | 83.404691 |
+| 1 | 1 | 2015-01-01 | 83.146150 | 82.693437 | 82.766221 | 82.850137 | 83.442163 | 83.526078 | 83.598863 |
+| 2 | 1 | 2016-01-01 | 83.339747 | 82.884744 | 82.957897 | 83.042237 | 83.637257 | 83.721597 | 83.794749 |
+| 3 | 1 | 2017-01-01 | 83.533344 | 83.073235 | 83.147208 | 83.232495 | 83.834192 | 83.919479 | 83.993452 |
+| 4 | 1 | 2018-01-01 | 83.726940 | 83.257894 | 83.333304 | 83.420247 | 84.033634 | 84.120577 | 84.195987 |
+| 5 | 1 | 2019-01-01 | 83.920537 | 83.437859 | 83.515461 | 83.604931 | 84.236144 | 84.325614 | 84.403216 |
+
+```python
+sf.plot(train, y_hat, level=[95])
+```
+
+
+
+### Forecast method
+
+Memory Efficient Exponential Smoothing predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to fit_predict without storing information. It assumes you
+know the forecast horizon in advance.
+
+```python
+y_hat = sf.forecast(df=train, h=6, fitted=True)
+y_hat
+```
+
+| | unique_id | ds | AutoETS |
+|-----|-----------|------------|-----------|
+| 0 | 1 | 2014-01-01 | 82.952553 |
+| 1 | 1 | 2015-01-01 | 83.146150 |
+| 2 | 1 | 2016-01-01 | 83.339747 |
+| 3 | 1 | 2017-01-01 | 83.533344 |
+| 4 | 1 | 2018-01-01 | 83.726940 |
+| 5 | 1 | 2019-01-01 | 83.920537 |
+
+### In sample predictions
+
+Access fitted Exponential Smoothing insample predictions.
+
+```python
+sf.forecast_fitted_values()
+```
+
+| | unique_id | ds | y | AutoETS |
+|-----|-----------|------------|-----------|-----------|
+| 0 | 1 | 1960-01-01 | 69.123902 | 69.005305 |
+| 1 | 1 | 1961-01-01 | 69.760244 | 69.237346 |
+| 2 | 1 | 1962-01-01 | 69.149756 | 69.495763 |
+| ... | ... | ... | ... | ... |
+| 51 | 1 | 2011-01-01 | 82.187805 | 82.348633 |
+| 52 | 1 | 2012-01-01 | 82.239024 | 82.561938 |
+| 53 | 1 | 2013-01-01 | 82.690244 | 82.758963 |
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ y_hat.merge(test),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=1), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | AutoETS |
+|-----|-----------|--------|----------|
+| 0 | 1 | mae | 0.421060 |
+| 1 | 1 | mape | 0.005073 |
+| 2 | 1 | mase | 1.340056 |
+| 3 | 1 | rmse | 0.483558 |
+| 4 | 1 | smape | 0.002528 |
+
+## References
+
+1. [Nixtla AutoETS API](../../src/core/models.html#autoets)
+2. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+
diff --git a/statsforecast/docs/models/autoregressive.html.mdx b/statsforecast/docs/models/autoregressive.html.mdx
new file mode 100644
index 00000000..07556090
--- /dev/null
+++ b/statsforecast/docs/models/autoregressive.html.mdx
@@ -0,0 +1,1158 @@
+---
+title: AutoRegressive Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `AutoRegressive Model` with
+> `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation` in other.
+
+The text in this article is largely taken from: 1. [Changquan Huang •
+Alla Petukhina. Springer series (2022). Applied Time Series Analysis and
+Forecasting with
+Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2) 2.
+[Jose A. Fiorucci, Tiago R. Pellegrini, Francisco Louzada, Fotios
+Petropoulos, Anne B. Koehler (2016). “Models for optimising the theta
+method and their relationship to state space models”. International
+Journal of
+Forecasting](https://www.sciencedirect.com/science/article/pii/S0169207016300243).
+3. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+Principles and Practice (3rd ed)”](https://otexts.com/fpp3/tscv.html)
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [Autoregressive Models](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of AutoRegressive with
+ StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+The `autoregressive` time series model `(AutoRegressive)` is a
+`statistical` technique used to analyze and predict univariate time
+series. In essence, the `autoregressive model` is based on the idea that
+previous values of the time series can be used to predict future values.
+
+In this model, the dependent variable (the time series) returns to
+itself at different moments in time, creating a dependency relationship
+between past and present values. The idea is that past values can help
+us understand and predict future values of the series.
+
+The `autoregressive model` can be fitted to different orders, which
+indicate how many past values are used to predict the present value. For
+example, an `autoregressive model` of order 1 $(AR(1))$ uses only the
+immediately previous value to predict the current value, while an
+`autoregressive model` of order $p (AR(p))$ uses the $p$ previous
+values.
+
+The `autoregressive model` is one of the basic models of time series
+analysis and is widely used in a variety of fields, from finance and
+economics to meteorology and social sciences. The model’s ability to
+capture nonlinear dependencies in time series data makes it especially
+useful for forecasting and long-term trend analysis.
+
+In a `multiple regression model`, we forecast the variable of interest
+using a linear combination of predictors. In an `autoregression model`,
+we forecast the variable of interest using a `linear combination` of
+past values of the variable. The term `autoregression` indicates that it
+is a regression of the variable against itself.
+
+## Definition of Autoregressive Models
+
+Before giving a formal definition of the ARCH model, let’s define the
+components of an ARCH model in a general way:
+
+- Autoregressive, a concept that we have already known, is the
+ construction of a univariate time series model using statistical
+ methods, which means that the current value of a variable is
+ influenced by past values of itself in different periods.
+- Heteroscedasticity means that the model can have different
+ magnitudes or variability at different time points (variance changes
+ over time).
+- Conditional, since volatility is not fixed, the reference here is
+ the constant that we put in the model to limit heteroscedasticity
+ and make it conditionally dependent on the previous value or values
+ of the variable.
+
+The AR model is the most basic building block of univariate time series.
+As you have seen before, univariate time series are a family of models
+that use only information about the target variable’s past to forecast
+its future, and do not rely on other explanatory variables.
+
+**Definition 1.** (1) The following equation is called the
+autoregressive model of order $p$ and denoted by $\text{AR(p)}$:
+
+$$
+\begin{equation}
+Xt =\varphi_0 +\varphi_1X_{t−1}+\varphi_2X_{t−2}+\cdots+\varphi_p X_{t−p}+\varepsilon_t \tag 1
+\end{equation}
+$$
+
+where $\{\varepsilon_t \} \sim WN(0,\sigma_{\epsilon}^2)$,
+$E(X_s \varepsilon_t) = 0$ if $s < t$ and
+$\varphi_0,\varphi_1,\cdots ,\varphi_p$ are real-valued parameters
+(coefficients) with $\varphi_p \neq 0$.
+
+1. If a time series $\{X_t \}$ is stationary and satisfies such an
+ equation as (1), then we call it an $\text{AR(p)}$ process.
+
+Note the following remarks about this definition:
+
+- For simplicity, we often assume that the intercept (const term)
+ $\varphi_0 = 0$; otherwise, we can consider $\{X_t −\mu \}$ where
+ $\mu =\varphi_0 /(1−\varphi_1 − \cdots −\varphi_p)$.
+
+- We distinguish the concept of $\text{AR}$ models from the concept of
+ $\text{AR}$ processes. $\text{AR}$ models may or may not be
+ stationary and $\text{AR}$ processes must be stationary.
+
+- $E(X_s \varepsilon_t) = 0(s < t)$ means that $X_s$ in the past has
+ nothing to do with $\varepsilon_t$ at the current time $t$.
+
+- Like the definition of MA models, sometimes εt in Eq.(1) is called
+ the innovation or shock term.
+
+In addition, using the
+backshift([see](https://otexts.com/fpp3/backshift.html)) operator $B$,
+the $\text{AR(p)}$ model can be rewritten as
+
+$$\varphi(B)X_t = \varepsilon_t$$
+
+where $\varphi(z) = 1 − \varphi_1z − \cdots − \varphi_p z^p$ is called
+the (corresponding) $\text{AR}$ polynomial. Besides, in the Python
+package \|StatsModels\|, $\varphi(B)$ is called the $\text{AR}$ lag
+polynomial.
+
+### Definition of PACF
+
+Let $\{X_t \}$ be a stationary time series with $E(X_t) = 0$. Here the
+assumption $E(X_t ) = 0$ is for conciseness only. If
+$E(X_t) = \mu \neq 0$, it is okay to replace $\{X_t \}$ by
+$\{X_t − \mu \}$. Now consider the linear regression (prediction) of
+$X_t$ on $\{X_{t−k+1:t−1} \}$ for any integer $k ≥ 2$. We use $\hat X_t$
+to denote this regression (prediction):
+
+$$\hat X_t =\alpha_1 X_{t−1}+ \cdots +\alpha_{k−1} X_{t−k+1}$$
+
+where $\{\alpha_1, \cdots , \alpha_{k−1} \}$ satisfy
+
+$$\{\alpha_1, \cdots , \alpha_{k−1} \}=\argmin_{β1,···,βk−1} E[X_t −(\beta_1 X_{t−1} +\cdots +\beta_{k−1}X_{t−k+1})]^2$$
+
+That is, $\{\alpha_1, \cdots , \alpha_{k−1} \}$ are chosen by minimizing
+the mean squared error of prediction. Similarly, let $\hat X_{t −k}$
+denote the regression (prediction) of $X_{t −k}$ on
+$\{X_{t −k+1:t −1} \}$:
+
+$$\hat X_{t−k} =\eta_1 X_{t−1}+ \cdots +\eta_{k−1} X_{t−k+1}$$
+
+Note that if $\{X_t \}$ is stationary, then
+$\{ \alpha_{1:k−1}\} = \{\eta_{1:k−1} \}$. Now let
+$\hat Z_{t−k} = X_{t−k} − \hat X_{t−k}$ and $\hat Z_t = X_t − \hat X_t$.
+Then $\hat Z_{t−k}$ is the residual of removing the effect of the
+intervening variables $\{X_{t−k+1:t−1} \}$ from $X_{t−k}$, and
+$\hat Z_t$ is the residual of removing the effect of
+$\{X_{t −k+1:t −1} \}$ from $X_t$.
+
+**Definition 2.** The partial autocorrelation function(PACF) at lag $k$
+of astationary time series $\{X_t \}$ with $E(X_t ) = 0$ is
+
+$$\phi_{11} = Corr(X_{t−1}, X_t ) = \frac{Cov(X_{t−1}, X_t )} {[Var(X_{t−1})Var(X_t)]^1/2}=\rho_1$$
+
+and
+
+$$\phi_{kk} = Corr(\hat Z_{t−k},\hat Z_t)=\frac{Cov(\hat Z_{t−k},\hat Z_t)} {[Var(\hat Z_{t−k})Var(\hat Z_t)]^{1/2}}$$
+
+According to the property of correlation coefficient (see, e.g., P172,
+Casella and Berger 2002), \|φkk\| ≤ 1. On the other hand, the following
+theorem paves the way to estimate the PACF of a stationary time series,
+and its proof can be seen in Fan and Yao (2003).
+
+On the other hand, the following theorem paves the way to estimate the
+PACF of a stationary time series, and its proof can be seen in Fan and
+Yao (2003).
+
+**Theorem 1.** Let $\{X_t \}$ be a stationary time series with
+$E(X_t) = 0$, and $\{a_{1k},\cdots ,a_{kk} \}$ satisfy
+
+$$\{a_{1k},\cdots,a_{kk} \}=\argmin_{a_1 ,\cdots ,a_k} E(X_{t −a1}X_{t−1}−\cdots −a_k X_{t−k})^2$$
+
+Then $\phi_{kk}=a_{kk}$ for $k≥1$.
+
+### Properties of Autoregressive Models
+
+From the $\text{AR(p)}$ model, namely, Eq. (1), we can see that it is in
+the same form as the multiple linear regression model. However, it
+explains current itself with its own past. Given the past
+
+$$\{X_{(t−p):(t−1)} \} = \{x_{(t−p):(t−1)} \}$$
+
+we have
+$$E(X_t |X_{(t−p):(t−1)}) = \varphi_0 + \varphi_1x_{t−1} + \varphi_2 x_{t−2} + \cdots + \varphi_p x_{t−p}$$
+
+This suggests that given the past, the right-hand side of this equation
+is a good estimate of $X_t$ . Besides
+
+$$Var(X_t |X_{(t −p):(t −1)}) = Var(\varepsilon_t ) = \sigma_{\varepsilon}^2$$
+
+Now we suppose that the AR(p) model, namely, Eq. (1), is stationary;
+then we have
+
+1. The model mean $E(_Xt)=\mu =\varphi_0 / (1−\varphi_1−···−\varphi_p)$
+ .Thus,themodelmean $\mu=0$ if and only if $\varphi_0 =0$.
+
+2. If the mean is zero or $\varphi_0 = 0$ ((3) and (4) below have the
+ same assumption), noting that
+ $E(X_t \varepsilon_t ) = \sigma_{\varepsilon}^2$ , we multiply
+ Eq. (1) by $X_t$ , take expectations, and then get
+
+$$\text {Var} (X_t) = \gamma_0 = \varphi_1 \gamma_1 + \varphi_2 \gamma2 + \cdots + \varphi_p \gamma_p + \sigma_{\varepsilon}^2$$
+
+Furthermore
+
+$$\gamma_0 = \sigma_{\varepsilon}^2 / ( 1 − \varphi_1 \rho_1 − \varphi_2 \rho_2 − \cdots − \varphi_p \rho_p )$$
+
+1. For all $k > p$, the partial autocorrelation $\phi_{kk} = 0$, that
+ is, the PACF of $\text{AR(p)}$ models cuts off after lag $p$, which
+ is very helpful in identifying an $\text{AR}$ model. In fact, at
+ this point, the predictor or regression of $X_t$ on
+ $\{X_{t−k+1:t−1} \}$ is
+
+$$\hat X_t =\varphi_1 X_{t−1}+\cdots +\varphi_{k−1} X_{t−k+1}$$
+
+Thus, $X_t − \hat X_t = \varepsilon_t$. Moreover,
+$X_{t−k} − \hat X_{t−k}$ is a function of $\{ X_{t−k:t−1} \}$, and
+$\varepsilon_t$ is uncorrelated to everyone in $\{X_{t−k:t−1} \}$.
+Therefore
+
+$$Cov(X_{t−k} −\hat X_{t−k},X_t −\hat X_t)=Cov(X_{t−k} −\hat X_{t−k},\varepsilon_t)=0.$$
+
+By Definition 2, $\phi_{kk} = 0$.
+
+1. We multiply Eq.(1)by $X_{t−k}$,take expectations,divide by
+ $\gamma_0$,and then obtain the recursive relationship between the
+ autocorrelations:
+
+$$
+\begin{equation}
+ for \ k ≥ 1, \rho_k = \varphi_1 \rho_{k−1} + \varphi_2 \rho_{k−2} + \cdots + \varphi_p \rho_{k−p} \tag 2
+\end{equation}
+$$
+
+For Eq.(2), let $k = 1,2,··· ,p$. Then we arrive at a set of difference
+equations, which is known as the Yule-Walker equations. If the
+$\text{ACF} \{\rho_{1:p} \}$ are given, then we can solve the
+Yule-Walker equations to obtain the estimates for $\{\varphi_{1:p} \}$,
+and the solutions are called the Yule-Walker estimates.
+
+1. Since the model is a stationary $\text{AR(p)}$ now, naturally it
+ satisfies
+ $X_t =\varphi_1 X_{t−1}+ \varphi_2 X_{t−2} + \cdots + \varphi_p X_{t−p} + \varepsilon_t$.
+ Hence $\phi_{pp} = \varphi_p$. If the $\text{AR(p)}$ model is
+ further Gaussian and a sample of size $\text{T}$ is given, then (a)
+ $\hat \phi_{pp} → \varphi_p$ as $T → ∞$; (b) according to Quenouille
+ (1949), for $k > p, \sqrt{T} \hat \phi_{kk}$ asymptotically follows
+ the standard normal(Gaussian) distribution $\text{N(0,1)}$, or
+ $\phi_{kk}$ is asymptotically distributed as $\text{N(0, 1/T )}$.
+
+### Stationarity and Causality of AR Models
+
+Consider the AR(1) model:
+
+$$
+\begin{equation}
+ X_t = \varphi X_{t − 1} + \varepsilon_t , \varepsilon_t \sim W N( 0 , \sigma_{\varepsilon}^2 ) \tag 3
+\end{equation}
+$$
+
+For $|\varphi|<1$,let
+$X_{1t} =\sum_{j=0}^{\infty} \varphi^j \varepsilon_{t−j}$ and for
+$|\varphi|>1$,let
+$X_{2t} =− \sum_{j=1}^{\infty} \varphi^{-j} \varepsilon_{t+j}$. It is
+easy to show that both $\{X_{1t } \}$ and $\{X_{2t } \}$ are stationary
+and satisfy Eq. (3). That is, both are the stationary solution of Eq.
+(3). This gives rise to a question: which one of both is preferable?
+Obviously, $\{X_{2t } \}$ depends on future values of unobservable
+$\{\varepsilon_t \}$, and so it is unnatural. Hence we take
+$\{X_{1t } \}$ and abandon $\{X_{2t } \}$. In other words, we require
+that the coefficient $\varphi$ in Eq. (3) is less 1 in absolute value.
+At this point, the $\text{AR}(1)$ model is said to be causal and its
+causal expression is
+$X_t = \sum_{j=0}^{\infty} \varphi^j \varepsilon_{t−j}$. In general, the
+definition of causality is given below.
+
+**Definition 3** (1) A time series $\{X_t \}$ is causal if there exist
+coefficients $\psi_j$ such that
+
+$$X_t =\sum_{j=0}^{\infty} \psi_j \varepsilon_{t-j}, \ \ \sum_{j=0}^{\infty} |\psi_j |< \infty$$
+
+where
+$\psi_0 = 1, \{\varepsilon_t \} \sim WN(0, \sigma_{\varepsilon}^2 )$. At
+this point, we say that the time series $\{X_t \}$ has an
+$\text{MA}(\infty)$ representation.
+
+1. We say that a model is causal if the time series generated by it is
+ causal.
+
+Causality suggests that the time series $\{X_t\}$ is caused by the white
+noise (or innovations) from the past up to time t . Besides, the time
+series $\{X_{2t } \}$ is an example that is stationary but not causal.
+In order to determine whether an $\text{AR}$ model is causal, similar to
+the invertibility for the $\text{MA}$ model, we have the following
+theorem.
+
+**Theorem 2(CausalityTheorem)** An $\text{AR}$ model defined by Eq.(1)
+is causal if and only if the roots of its $\text{AR}$ polynomial
+$\varphi(z)=1−\varphi_1 z− \cdots − \varphi_p z^p$ exceed 1 in modulus
+or lie outside the unit circle on the complex plane.
+
+Note the following remarks: \* In the light of the existence and
+uniqueness on page 75 of Brockwell and Davis (2016), an $\text{AR}$
+model defined by Eq.(1) is stationary if and only if its $\text{AR}$
+polynomial $\varphi(z)=1−\varphi_1 z− \cdots − \varphi_p z^p \neq 0$ for
+all $|z|=1$ or all the roots of the $\text{AR}$ polynomial do not lie on
+the unit circle. Hence for the AR model defined by Eq. (1), its
+stationarity condition is weaker than its causality condition.
+
+- A causal time series is surely a stationary one. So an $\text{AR}$
+ model that satisfies the causal condition is naturally stationary.
+ But a stationary $\text{AR}$ model is not necessarily causal.
+
+- If the time series $\{X_t \}$ generated by Eq. (1) is not from the
+ remote past, namely,
+ $$t \in T = {\cdots ,−n,\cdots ,−1,0,1,\cdots ,n,\cdots}$$
+
+but starts from an initial value $X_0$, then it may be nonstationary,
+not to mention causality.
+
+- According to the relationship between the roots and the coefficients
+ of the degree 2 polynomial
+ $\varphi(z) = 1 − \varphi_1 z − \varphi_2 z^2$, it may be proved
+ that both of the roots of the polynomial exceed 1 in modulus if and
+ only if
+
+Thus, we can conveniently use the three inequations to decide whether a
+$\text{AR(2)}$ model is causal or not.
+
+- It may be shown that for an $\text{AR(p)}$ model defined by Eq. (1),
+ the coefficients $\{\psi_j \}$ in Definition 3 satisfy $\psi_0=1$
+ and
+
+$$\psi_j=\sum_{k=1}^{j} \varphi '_k \psi_{j-k}, \ \ j \geq 1 \ where \ \ \varphi '_k =\varphi_k \ \ if \ \ k \leq p \ \ and \ \ \varphi '_k =0 \ \ if \ \ k>p$$
+
+### Autocorrelation: the past influences the present
+
+The autoregressive model describes a relationship between the present of
+a variable and its past. Therefore, it is suitable for variables in
+which the past and present values are correlated.
+
+As an intuitive example, consider the waiting line at the doctor.
+Imagine that the doctor has a plan in which each patient has 20 minutes
+with him. If each patient takes exactly 20 minutes, this works well. But
+what if a patient takes a little longer? An autocorrelation could be
+present if the duration of one query has an impact on the duration of
+the next query. So if the doctor needs to speed up an appointment
+because the previous appointment took too long, look at a correlation
+between the past and the present. Past values influence future values.
+
+### Positive and negative autocorrelation
+
+Like “regular” correlation, autocorrelation can be positive or negative.
+Positive autocorrelation means that a high value now is likely to give a
+high value in the next period. This can be observed, for example, in
+stock trading: as soon as a lot of people want to buy a stock, its price
+goes up. This positive trend makes people want to buy this stock even
+more as it has positive returns. The more people buy the stock, the
+higher it goes and the more people will want to buy it.
+
+A positive correlation also works in downtrends. If today’s stock value
+is low, tomorrow’s value is likely to be even lower as people start
+selling. When many people sell, the value falls, and even more people
+will want to sell. This is also a case of positive autocorrelation since
+the past and the present go in the same direction. If the past is low,
+the present is low; and if the past is high, the present is high.
+
+There is negative autocorrelation if two trends are opposite. This is
+the case in the example of the duration of the doctor’s visit. If one
+query takes longer, the next one will be shorter. If one visit takes
+less time, the doctor may take a little longer for the next one.
+
+### Stationarity and the ADF test
+
+The problem of having a trend in our data is general in univariate time
+series modeling. The stationarity of a time series means that a time
+series does not have a (long-term) trend: it is stable around the same
+average. Otherwise, a time series is said to be non-stationary.
+
+In theory, AR models can have a trend coefficient in the model, but
+since stationarity is an important concept in general time series
+theory, it’s best to learn to deal with it right away. Many models can
+only work on stationary time series.
+
+A time series that is growing or falling strongly over time is obvious
+to spot. But sometimes it’s hard to tell if a time series is stationary.
+This is where the Augmented Dickey Fuller (ADF) test comes in handy.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import pandas as pd
+
+import scipy.stats as stats
+
+import statsmodels.api as sm
+import statsmodels.tsa.api as smt
+```
+
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
+
+plt.style.use('fivethirtyeight')
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#212946',
+ 'axes.facecolor': '#212946',
+ 'savefig.facecolor':'#212946',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#2A3459',
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+### Read Data
+
+```python
+df= pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/catfish.csv")
+df.head()
+```
+
+| | Date | Total |
+|-----|-----------|-------|
+| 0 | 1986-1-01 | 9034 |
+| 1 | 1986-2-01 | 9596 |
+| 2 | 1986-3-01 | 10558 |
+| 3 | 1986-4-01 | 9002 |
+| 4 | 1986-5-01 | 9239 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|-----------|-------|-----------|
+| 0 | 1986-1-01 | 9034 | 1 |
+| 1 | 1986-2-01 | 9596 | 1 |
+| 2 | 1986-3-01 | 10558 | 1 |
+| 3 | 1986-4-01 | 9002 | 1 |
+| 4 | 1986-5-01 | 9239 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y int64
+unique_id object
+dtype: object
+```
+
+We can see that our time variable `ds` is in an object format, we need
+to convert to a date format
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints 8 random series from the dataset and is useful for
+basic EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### The Augmented Dickey-Fuller Test
+
+An Augmented Dickey-Fuller (ADF) test is a type of statistical test that
+determines whether a unit root is present in time series data. Unit
+roots can cause unpredictable results in time series analysis. A null
+hypothesis is formed in the unit root test to determine how strongly
+time series data is affected by a trend. By accepting the null
+hypothesis, we accept the evidence that the time series data is not
+stationary. By rejecting the null hypothesis or accepting the
+alternative hypothesis, we accept the evidence that the time series data
+is generated by a stationary process. This process is also known as
+stationary trend. The values of the ADF test statistic are negative.
+Lower ADF values indicate a stronger rejection of the null hypothesis.
+
+Augmented Dickey-Fuller Test is a common statistical test used to test
+whether a given time series is stationary or not. We can achieve this by
+defining the null and alternate hypothesis.
+
+Null Hypothesis: Time Series is non-stationary. It gives a
+time-dependent trend. Alternate Hypothesis: Time Series is stationary.
+In another term, the series doesn’t depend on time.
+
+ADF or t Statistic \< critical values: Reject the null hypothesis, time
+series is stationary. ADF or t Statistic \> critical values: Failed to
+reject the null hypothesis, time series is non-stationary.
+
+Let’s check if our series that we are analyzing is a stationary series.
+Let’s create a function to check, using the `Dickey Fuller` test
+
+```python
+from statsmodels.tsa.stattools import adfuller
+```
+
+
+```python
+def Augmented_Dickey_Fuller_Test_func(series , column_name):
+ print (f'Dickey-Fuller test results for columns: {column_name}')
+ dftest = adfuller(series, autolag='AIC')
+ dfoutput = pd.Series(dftest[0:4], index=['Test Statistic','p-value','No Lags Used','Number of observations used'])
+ for key,value in dftest[4].items():
+ dfoutput['Critical Value (%s)'%key] = value
+ print (dfoutput)
+ if dftest[1] \<= 0.05:
+ print("Conclusion:====>")
+ print("Reject the null hypothesis")
+ print("The data is stationary")
+ else:
+ print("Conclusion:====>")
+ print("The null hypothesis cannot be rejected")
+ print("The data is not stationary")
+```
+
+
+```python
+Augmented_Dickey_Fuller_Test_func(df["y"],'Sales')
+```
+
+``` text
+Dickey-Fuller test results for columns: Sales
+Test Statistic -1.589903
+p-value 0.488664
+No Lags Used 14.000000
+ ...
+Critical Value (1%) -3.451691
+Critical Value (5%) -2.870939
+Critical Value (10%) -2.571778
+Length: 7, dtype: float64
+Conclusion:====>
+The null hypothesis cannot be rejected
+The data is not stationary
+```
+
+In the previous result we can see that the `Augmented_Dickey_Fuller`
+test gives us a `p-value` of 0.488664, which tells us that the null
+hypothesis cannot be rejected, and on the other hand the data of our
+series are not stationary.
+
+We need to differentiate our time series, in order to convert the data
+to stationary.
+
+```python
+Augmented_Dickey_Fuller_Test_func(df["y"].diff().dropna(),"Sales")
+```
+
+``` text
+Dickey-Fuller test results for columns: Sales
+Test Statistic -4.310935
+p-value 0.000425
+No Lags Used 17.000000
+ ...
+Critical Value (1%) -3.451974
+Critical Value (5%) -2.871063
+Critical Value (10%) -2.571844
+Length: 7, dtype: float64
+Conclusion:====>
+Reject the null hypothesis
+The data is stationary
+```
+
+By applying a differential, our time series now is stationary.
+
+```python
+def tsplot(y, lags=None, figsize=(12, 7), style='bmh'): # [3]
+ if not isinstance(y, pd.Series):
+ y = pd.Series(y)
+
+ with plt.style.context(style):
+ fig = plt.figure(figsize=figsize)
+ layout = (2, 2)
+ ts_ax = plt.subplot2grid(layout, (0, 0), colspan=2)
+ acf_ax = plt.subplot2grid(layout, (1, 0))
+ pacf_ax = plt.subplot2grid(layout, (1, 1))
+
+ y.plot(ax=ts_ax)
+ p_value = sm.tsa.stattools.adfuller(y)[1]
+ ts_ax.set_title('Time Series Analysis plot\n Dickey-Fuller: p={0:.5f}'.format(p_value))
+ smt.graphics.plot_acf(y, lags=lags, ax=acf_ax)
+ smt.graphics.plot_pacf(y, lags=lags, ax=pacf_ax)
+ plt.tight_layout()
+```
+
+
+```python
+tsplot(df["y"].diff().dropna(), lags=20);
+```
+
+
+
+As you can see, based on the blue background shaded area of the graph,
+the PACF shows the first, second, third, fourth, sixth, seventh, ninth,
+and tenth etc. delay outside the shaded area. This means that it would
+be interesting to also include these lags in the AR model.
+
+### How many lags should we include?
+
+Now, the **big question in time series analysis is always how many lags
+to include**. This is called the order of the time series. The notation
+is AR(1) for order 1 and AR(p) for order p.
+
+The order is up to you. Theoretically speaking, you can base your order
+on the PACF chart. Theory tells you to take the number of lags before
+you get an autocorrelation of 0. All other lags should be 0.
+
+In theory, you often see great charts where the first peak is very high
+and the rest equal zero. In those cases, the choice is easy: you are
+working with a very “pure” example of AR(1). Another common case is when
+your autocorrelation starts high and slowly decreases to zero. In this
+case, you should use all delays where the PACF is not yet zero.
+
+However, in practice, it is not always that simple. Remember the famous
+saying *“all models are wrong, but some are useful”*. It is very rare to
+find cases that fit an AR model perfectly. In general, the
+autoregression process can help explain part of the variation of a
+variable, but not all.
+
+In practice, you will try to select the number of lags that gives your
+model the best predictive performance. The best predictive performance
+is often not defined by looking at autocorrelation plots: those plots
+give you a theoretical estimate. However, predictive performance is best
+defined by model evaluation and benchmarking, using the techniques you
+have seen in Module 2. Later in this module, we will see how to use
+model evaluation to choose a performance order for the AR model. But
+before we get into that, it’s time to dig into the exact definition of
+the AR model.
+
+## Split the data into training and testing
+
+Let’s divide our data into sets 1. Data to train our `AutoRegressive`
+model 2. Data to test our model
+
+For the test data we will use the last 12 months to test and evaluate
+the performance of our model.
+
+```python
+train = df[df.ds\<='2011-12-01']
+test = df[df.ds>'2011-12-01']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((312, 3), (12, 3))
+```
+
+Now let’s plot the training data and the test data.
+
+```python
+sns.lineplot(train,x="ds", y="y", label="Train")
+sns.lineplot(test, x="ds", y="y", label="Test")
+plt.show()
+```
+
+
+
+# Implementation of AutoRegressive with StatsForecast
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import AutoRegressive
+```
+
+### Instantiating Model
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/) by the
+master, Rob Hyndmann, can be useful.season_length.
+
+**Method 1:** We use the lags parameter in an integer format, that is,
+we put the lags we want to evaluate in the model.
+
+```python
+season_length = 12 # Monthly data
+horizon = len(test) # number of predictions biasadj=True, include_drift=True,
+
+models2 = [AutoRegressive(lags=[14], include_mean=True)]
+```
+
+**Method 2:** We use the lags parameter in a list format, that is, we
+put the lags that we want to evaluate in the model in the form of a list
+as shown below.
+
+```python
+season_length = 12 # Monthly data
+horizon = len(test) # number of predictions
+
+models = [AutoRegressive(lags=[3,4,6,7,9,10,11,12,13,14], include_mean=True)]
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See
+ [pandas’s available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models, freq='MS')
+```
+
+### Fit Model
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[AutoRegressive])
+```
+
+Let’s see the results of our Theta model. We can observe it with the
+following instruction:
+
+```python
+result=sf.fitted_[0,0].model_
+print(result.keys())
+```
+
+``` text
+dict_keys(['coef', 'sigma2', 'var_coef', 'mask', 'loglik', 'aic', 'arma', 'residuals', 'code', 'n_cond', 'nobs', 'model', 'aicc', 'bic', 'xreg', 'lambda', 'x'])
+```
+
+Let us now visualize the residuals of our models.
+
+As we can see, the result obtained above has an output in a dictionary,
+to extract each element from the dictionary we are going to use the
+`.get()` function to extract the element and then we are going to save
+it in a `pd.DataFrame()`.
+
+```python
+residual=pd.DataFrame(result.get("residuals"), columns=["residual Model"])
+residual
+```
+
+| | residual Model |
+|-----|----------------|
+| 0 | -11998.537347 |
+| 1 | NaN |
+| 2 | NaN |
+| ... | ... |
+| 309 | -2718.312961 |
+| 310 | -1306.795172 |
+| 311 | -2713.284999 |
+
+```python
+fig, axs = plt.subplots(nrows=2, ncols=2)
+
+# plot[1,1]
+residual.plot(ax=axs[0,0])
+axs[0,0].set_title("Residuals");
+
+# plot
+sns.distplot(residual, ax=axs[0,1]);
+axs[0,1].set_title("Density plot - Residual");
+
+# plot
+stats.probplot(residual["residual Model"], dist="norm", plot=axs[1,0])
+axs[1,0].set_title('Plot Q-Q')
+
+# plot
+plot_acf(residual, lags=35, ax=axs[1,1],color="fuchsia")
+axs[1,1].set_title("Autocorrelation");
+
+plt.show();
+```
+
+
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[90]` means that
+ the model expects the real value to be inside that interval 90% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min. (If you want to speed things up to a couple of seconds,
+remove the AutoModels like `ARIMA` and `Theta`)
+
+```python
+# Prediction
+Y_hat = sf.forecast(df=train, h=horizon, fitted=True)
+Y_hat
+```
+
+| | unique_id | ds | AutoRegressive |
+|-----|-----------|------------|----------------|
+| 0 | 1 | 2012-01-01 | 15905.582031 |
+| 1 | 1 | 2012-02-01 | 13597.894531 |
+| 2 | 1 | 2012-03-01 | 15488.883789 |
+| ... | ... | ... | ... |
+| 9 | 1 | 2012-10-01 | 14087.901367 |
+| 10 | 1 | 2012-11-01 | 13274.105469 |
+| 11 | 1 | 2012-12-01 | 12498.226562 |
+
+```python
+values=sf.forecast_fitted_values()
+values.head()
+```
+
+| | unique_id | ds | y | AutoRegressive |
+|-----|-----------|------------|---------|----------------|
+| 0 | 1 | 1986-01-01 | 9034.0 | 21032.537109 |
+| 1 | 1 | 1986-02-01 | 9596.0 | NaN |
+| 2 | 1 | 1986-03-01 | 10558.0 | NaN |
+| 3 | 1 | 1986-04-01 | 9002.0 | 126172.937500 |
+| 4 | 1 | 1986-05-01 | 9239.0 | 10020.040039 |
+
+Adding 95% confidence interval with the forecast method
+
+```python
+sf.forecast(df=train, h=horizon, level=[95])
+```
+
+| | unique_id | ds | AutoRegressive | AutoRegressive-lo-95 | AutoRegressive-hi-95 |
+|-----|-----------|------------|----------------|----------------------|----------------------|
+| 0 | 1 | 2012-01-01 | 15905.582031 | 2119.586426 | 29691.578125 |
+| 1 | 1 | 2012-02-01 | 13597.894531 | -188.101135 | 27383.890625 |
+| 2 | 1 | 2012-03-01 | 15488.883789 | 1702.888062 | 29274.878906 |
+| ... | ... | ... | ... | ... | ... |
+| 9 | 1 | 2012-10-01 | 14087.901367 | -1050.068359 | 29225.871094 |
+| 10 | 1 | 2012-11-01 | 13274.105469 | -1886.973145 | 28435.183594 |
+| 11 | 1 | 2012-12-01 | 12498.226562 | -2675.547607 | 27672.001953 |
+
+```python
+# Merge the forecasts with the true values
+Y_hat1 = test.merge(Y_hat, how='left', on=['unique_id', 'ds'])
+Y_hat1
+```
+
+| | ds | y | unique_id | AutoRegressive |
+|-----|------------|-------|-----------|----------------|
+| 0 | 2012-01-01 | 13427 | 1 | 15905.582031 |
+| 1 | 2012-02-01 | 14447 | 1 | 13597.894531 |
+| 2 | 2012-03-01 | 14717 | 1 | 15488.883789 |
+| ... | ... | ... | ... | ... |
+| 9 | 2012-10-01 | 13795 | 1 | 14087.901367 |
+| 10 | 2012-11-01 | 13352 | 1 | 13274.105469 |
+| 11 | 2012-12-01 | 12716 | 1 | 12498.226562 |
+
+```python
+sf.plot(train, Y_hat1)
+```
+
+
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[95]` means that
+ the model expects the real value to be inside that interval 95% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+sf.predict(h=horizon)
+```
+
+| | unique_id | ds | AutoRegressive |
+|-----|-----------|------------|----------------|
+| 0 | 1 | 2012-01-01 | 15905.582031 |
+| 1 | 1 | 2012-02-01 | 13597.894531 |
+| 2 | 1 | 2012-03-01 | 15488.883789 |
+| ... | ... | ... | ... |
+| 9 | 1 | 2012-10-01 | 14087.901367 |
+| 10 | 1 | 2012-11-01 | 13274.105469 |
+| 11 | 1 | 2012-12-01 | 12498.226562 |
+
+```python
+forecast_df = sf.predict(h=horizon, level=[95])
+forecast_df
+```
+
+| | unique_id | ds | AutoRegressive | AutoRegressive-lo-95 | AutoRegressive-hi-95 |
+|-----|-----------|------------|----------------|----------------------|----------------------|
+| 0 | 1 | 2012-01-01 | 15905.582031 | 2119.586426 | 29691.578125 |
+| 1 | 1 | 2012-02-01 | 13597.894531 | -188.101135 | 27383.890625 |
+| 2 | 1 | 2012-03-01 | 15488.883789 | 1702.888062 | 29274.878906 |
+| ... | ... | ... | ... | ... | ... |
+| 9 | 1 | 2012-10-01 | 14087.901367 | -1050.068359 | 29225.871094 |
+| 10 | 1 | 2012-11-01 | 13274.105469 | -1886.973145 | 28435.183594 |
+| 11 | 1 | 2012-12-01 | 12498.226562 | -2675.547607 | 27672.001953 |
+
+```python
+sf.plot(train, test.merge(forecast_df), level=[95])
+```
+
+
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=5)`, forecasting every second months
+`(step_size=12)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 12 months ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=train,
+ h=horizon,
+ step_size=6,
+ n_windows=5)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` series identifier
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the n_windows.
+- `y:` true value
+- `"model":` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df
+```
+
+| | unique_id | ds | cutoff | y | AutoRegressive |
+|-----|-----------|------------|------------|---------|----------------|
+| 0 | 1 | 2009-01-01 | 2008-12-01 | 19262.0 | 24295.837891 |
+| 1 | 1 | 2009-02-01 | 2008-12-01 | 20658.0 | 23993.947266 |
+| 2 | 1 | 2009-03-01 | 2008-12-01 | 22660.0 | 21201.121094 |
+| ... | ... | ... | ... | ... | ... |
+| 57 | 1 | 2011-10-01 | 2010-12-01 | 12893.0 | 19349.708984 |
+| 58 | 1 | 2011-11-01 | 2010-12-01 | 11843.0 | 16899.849609 |
+| 59 | 1 | 2011-12-01 | 2010-12-01 | 11321.0 | 18159.574219 |
+
+We’ll now plot the forecast for each cutoff period. To make the plots
+clearer, we’ll rename the actual values in each period.
+
+```python
+from IPython.display import display
+```
+
+
+```python
+crossvalidation_df.rename(columns = {'y' : 'actual'}, inplace = True) # rename actual values
+
+cutoff = crossvalidation_df['cutoff'].unique()
+
+for k in range(len(cutoff)):
+ cv = crossvalidation_df[crossvalidation_df['cutoff'] == cutoff[k]]
+ display(StatsForecast.plot(df, cv.loc[:, cv.columns != 'cutoff']))
+```
+
+
+
+
+
+
+
+
+
+
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=season_length), ufl.rmse, ufl.smape],
+ train_df=train,
+ agg_fn='mean',
+)
+```
+
+| | metric | AutoRegressive |
+|-----|--------|----------------|
+| 0 | mae | 962.023763 |
+| 1 | mape | 0.072733 |
+| 2 | mase | 0.601808 |
+| 3 | rmse | 1195.013050 |
+| 4 | smape | 0.034858 |
+
+## References
+
+1. [Changquan Huang • Alla Petukhina. Springer series (2022). Applied
+ Time Series Analysis and Forecasting with
+ Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2)
+2. [Jose A. Fiorucci, Tiago R. Pellegrini, Francisco Louzada, Fotios
+ Petropoulos, Anne B. Koehler (2016). “Models for optimising the
+ theta method and their relationship to state space models”.
+ International Journal of
+ Forecasting](https://www.sciencedirect.com/science/article/pii/S0169207016300243).
+3. [Nixtla AutoRegressive API](../../src/core/models.html#autoregressive)
+4. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+5. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html)
+6. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/models/autotheta.html.mdx b/statsforecast/docs/models/autotheta.html.mdx
new file mode 100644
index 00000000..682a3ff4
--- /dev/null
+++ b/statsforecast/docs/models/autotheta.html.mdx
@@ -0,0 +1,950 @@
+---
+title: AutoTheta Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `AutoTheta Model` with
+> `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation` in other.
+
+The text in this article is copied from and inspired by: 1. [Jose A.
+Fiorucci, Tiago R. Pellegrini, Francisco Louzada, Fotios Petropoulos,
+Anne B. Koehler (2016). “Models for optimising the theta method and
+their relationship to state space models”. International Journal of
+Forecasting](https://www.sciencedirect.com/science/article/pii/S0169207016300243).
+2. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+Principles and Practice (3rd ed)”](https://otexts.com/fpp3/tscv.html)
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of AutoTheta with StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+The `AutoTheta` model in `StatsForecast` automatically selects the best
+**Theta model** based on the **mean squared error (MSE)**. In this
+section, we will discuss each of the models that `AutoTheta` considers
+and then explain how it selects the best one.
+
+### 1. Standard Theta Model (STM)
+
+The **Standard Theta Model** is the original version of the Theta model
+introduced by Assimakopoulos and Nikolopoulos (2000). It decomposes a
+time series into two modified versions of the original series, called
+**theta lines**. These lines are created by applying a linear
+transformation to the second differences of the original series,
+controlled by a parameter called **theta $\theta$**. One theta line
+captures the long-term trend, while the other captures short-term
+fluctuations. The two theta lines are then combined to produce the final
+forecast. The STM assumes that model parameters remain constant over
+time.
+
+### 2. Optimized Theta Model (OTM)
+
+The **Optimized Theta Model** extends STM by searching for the best
+theta parameters rather than using fixed values. This optimization step
+allows the model to better fit series with higher variability.
+
+### 3. Dynamic Standard Theta Model (DSTM)
+
+The **Dynamic Standard Theta Model** allows STM to adapt over time.
+Instead of keeping parameters static, it updates them dynamically as new
+data becomes available. This dynamic behavior can be useful when
+forecasting series with evolving trends or seasonality.
+
+### 4. Dynamic Optimized Theta Model (DOTM)
+
+The **Dynamic Optimized Theta Model** combines features of both OTM and
+DSTM. Like OTM, it optimizes the theta parameters. Like DSTM, it updates
+the model dynamically with new data.
+
+## How AutoTheta Selects the Best Model
+
+1. `AutoTheta` fits all four variants of the Theta model (STM, OTM,
+ DSTM, and DOTM) to your data.
+2. Each model is evaluated using cross-validation or a hold-out
+ validation strategy, depending on the configuration.
+3. The model that achieves the lowest mean squared error (MSE) is
+ selected.
+4. The selected model is then used to generate the forecast.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import pandas as pd
+
+import scipy.stats as stats
+```
+
+
+```python
+
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
+
+plt.style.use('fivethirtyeight')
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#212946',
+ 'axes.facecolor': '#212946',
+ 'savefig.facecolor':'#212946',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#2A3459',
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+
+```
+
+### Read Data
+
+```python
+df = pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/candy_production.csv")
+df.head()
+```
+
+| | observation_date | IPG3113N |
+|-----|------------------|----------|
+| 0 | 1972-01-01 | 85.6945 |
+| 1 | 1972-02-01 | 71.8200 |
+| 2 | 1972-03-01 | 66.0229 |
+| 3 | 1972-04-01 | 64.5645 |
+| 4 | 1972-05-01 | 65.0100 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|------------|---------|-----------|
+| 0 | 1972-01-01 | 85.6945 | 1 |
+| 1 | 1972-02-01 | 71.8200 | 1 |
+| 2 | 1972-03-01 | 66.0229 | 1 |
+| 3 | 1972-04-01 | 64.5645 | 1 |
+| 4 | 1972-05-01 | 65.0100 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y float64
+unique_id object
+dtype: object
+```
+
+We can see that our time variable `(ds)` is in an object format, we need
+to convert to a date format
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore Data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints aa random series from the dataset and is useful for
+basic EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### Autocorrelation plots
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=60, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+plot_pacf(df["y"], lags=60, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+plt.show();
+```
+
+
+
+## Split the data into training and testing
+
+Let’s divide our data into sets 1. Data to train our `AutoTheta` model
+2. Data to test our model
+
+For the test data we will use the last 12 months to test and evaluate
+the performance of our model.
+
+```python
+train = df[df.ds\<='2016-08-01']
+test = df[df.ds>'2016-08-01']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((536, 3), (12, 3))
+```
+
+Now let’s plot the training data and the test data.
+
+```python
+sns.lineplot(train,x="ds", y="y", label="Train", linewidth=3, linestyle=":")
+sns.lineplot(test, x="ds", y="y", label="Test")
+plt.ylabel("Candy Production")
+plt.xlabel("Month")
+plt.show()
+```
+
+
+
+## Implementation of AutoTheta with StatsForecast
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import AutoTheta
+```
+
+### Instantiate Model
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/) by the
+master, Rob Hyndmann, can be useful.season_length.
+
+Automatically selects the best Theta (Standard Theta Model `(‘STM’)`,
+Optimized Theta Model `(‘OTM’)`, Dynamic Standard Theta Model
+`(‘DSTM’)`, Dynamic Optimized Theta Model `(‘DOTM’))` model using mse.
+
+```python
+season_length = 12 # Monthly data
+horizon = len(test) # number of predictions
+
+# We call the model that we are going to use
+models = [AutoTheta(season_length=season_length,
+ decomposition_type="additive",
+ model="STM")]
+
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [panda’s
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models, freq='MS')
+```
+
+### Fit Model
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[AutoTheta])
+```
+
+Let’s see the results of our Theta model. We can observe it with the
+following instruction:
+
+```python
+result=sf.fitted_[0,0].model_
+result
+```
+
+``` text
+{'mse': 100.57831864069415,
+ 'amse': array([26.13585578, 38.60211513, 44.70605915]),
+ 'fit': results(x=array([258.45064973, 0.7664297 ]), fn=100.57831864069415, nit=32, simplex=array([[250.37338496, 0.76970741],
+ [232.03915522, 0.76429422],
+ [258.45064973, 0.7664297 ]])),
+ 'residuals': array([-2.14815337e+02, -6.20562800e+01, -2.13256707e+01, -1.25845480e+01,
+ -1.19719350e+01, -9.40876632e+00, -8.60141525e+00, -9.00054652e+00,
+ -1.98778836e+00, 3.14564857e+01, 1.98519673e+01, 2.04962370e+01,
+ 4.98120196e+00, -1.08735375e+01, -1.12328024e+01, -8.08115377e+00,
+ -9.98197589e+00, -8.39937098e+00, -1.25789505e+01, -1.05952806e+01,
+ 8.47229127e-01, 2.25644616e+01, 2.54401546e+01, 1.73989716e+01,
+ 2.40287275e+00, -2.53475866e+00, -8.00591135e+00, -1.79241479e+01,
+ -6.36590693e+00, -5.76986468e+00, -2.26766759e+01, -8.95260931e+00,
+ -7.19719166e+00, 2.74032238e+01, 2.21368457e+01, 6.43171676e+00,
+ -3.51755220e+00, -1.31441941e+01, -6.13166031e+00, 1.51512150e+00,
+ -8.05777104e+00, -8.59603388e+00, -1.08617851e+01, -6.72940177e+00,
+ -6.24861641e+00, 2.85996828e+01, 2.98048030e+01, 1.90032238e+01,
+ 5.07597842e+00, -9.59170058e+00, -1.64521034e+01, -7.52212744e+00,
+ -5.16540394e+00, -1.27924628e+01, -9.68434625e+00, -8.76758703e+00,
+ -8.27475947e-01, 3.08424002e+01, 2.47947352e+01, 2.35867208e+01,
+ 3.75664716e+00, -4.47305717e+00, -1.48000403e+01, -1.08431546e+01,
+ -1.01249972e+01, -1.12379765e+01, -1.28624644e+01, -9.47780103e+00,
+ -2.17960841e-01, 2.49398648e+01, 1.66027782e+01, 2.62581230e+01,
+ -1.94879264e+00, -8.10877843e+00, -6.93183679e+00, -6.80707596e+00,
+ -1.17809892e+01, -1.05320670e+01, -1.59715849e+01, -9.07599923e+00,
+ 6.11988125e-01, 2.24925163e+01, 2.57389503e+01, 2.38907614e+01,
+ 4.99776202e+00, -1.07054696e+01, -7.24194672e+00, -1.17412084e+01,
+ -1.10031559e+01, -9.10138831e+00, -1.62277209e+01, -1.02585250e+01,
+ -2.79431476e+00, 1.96746051e+01, 2.40620700e+01, 2.00041920e+01,
+ 8.38674843e-01, -3.01708830e-01, -1.10576372e+01, -1.76502404e+01,
+ -4.79853028e+00, -7.74057206e+00, -1.55628746e+01, -6.19663664e+00,
+ -4.85267830e+00, 2.17819325e+01, 2.48075790e+01, 2.16186207e+01,
+ 9.21215745e+00, -1.71191202e+00, -1.38314188e+01, -9.44161337e+00,
+ -6.35863884e+00, -1.10470671e+01, -1.41408736e+01, -9.60039945e+00,
+ -4.80959619e+00, 3.41173952e+01, 2.02685767e+01, 1.65177446e+01,
+ 1.45004431e+00, -6.65011083e-01, -1.11027939e+01, -1.82545876e+01,
+ -1.08637878e+01, -9.67573606e+00, -1.22946714e+01, -1.02064815e+01,
+ -2.94225894e+00, 3.21840497e+01, 2.21586046e+01, 2.09073990e+01,
+ -2.49862821e-01, -6.05605889e+00, -1.16741825e+01, -1.31096470e+01,
+ -1.07043825e+01, -1.25489037e+01, -9.16715807e+00, -7.70278723e+00,
+ -2.55657034e+00, 2.69936351e+01, 1.62042780e+01, 1.67614452e+01,
+ 8.62186552e+00, -3.51518668e+00, -9.27421021e+00, -1.15442848e+01,
+ -9.96136043e+00, -1.17898558e+01, -1.13147670e+01, -7.10440489e+00,
+ -1.10170600e+00, 2.60646482e+01, 2.32687942e+01, 1.82272063e+01,
+ 3.98792378e+00, -7.64233782e+00, -1.07945901e+01, -1.16024004e+01,
+ -1.10645345e+01, -1.33282245e+01, -1.15534843e+01, -6.76286215e+00,
+ 3.93786824e+00, 2.37018431e+01, 2.07922131e+01, 2.37645505e+01,
+ 7.00182907e-01, -1.59605643e+00, -1.62277584e+01, -1.51068271e+01,
+ -1.01377645e+01, -1.13639586e+01, -1.38275901e+01, -5.87092572e+00,
+ 3.43469809e+00, 2.82932175e+01, 2.39510218e+01, 1.71053544e+01,
+ 6.00992500e-01, -7.61224365e-01, -1.18686664e+01, -1.51989727e+01,
+ -1.23352870e+01, -1.09931345e+01, -1.34086766e+01, -4.52127997e+00,
+ 2.09363525e+00, 3.13825850e+01, 2.43980063e+01, 1.89899567e+01,
+ -7.55702038e+00, -2.76893846e-01, -6.52574120e+00, -1.67167241e+01,
+ -1.17498886e+01, -7.68050287e+00, -5.60844424e+00, -2.79087739e+00,
+ -2.92094111e-01, 2.31896495e+01, 1.70158799e+01, 1.84177113e+01,
+ -3.39879920e-01, 1.31241579e+00, -9.65552567e+00, -1.30840488e+01,
+ -1.33540036e+01, -9.72077648e+00, -1.09022916e+01, -4.49636288e+00,
+ -6.88544858e-01, 1.88878504e+01, 2.15227074e+01, 2.32009723e+01,
+ -5.72605223e+00, 1.87746593e+00, -6.95944675e+00, -1.41944248e+01,
+ -1.25398544e+01, -8.09461542e+00, -5.46316863e+00, -4.73324533e+00,
+ 1.12162644e+00, 1.61183526e+01, 2.63470350e+01, 2.28827919e+01,
+ -6.75326971e+00, 4.34023844e+00, -6.61711624e+00, -1.64533666e+01,
+ -1.44473761e+01, -4.85575583e+00, -1.14659672e+01, -1.83412077e+00,
+ -3.17492418e+00, 1.22586060e+01, 2.19162129e+01, 1.62630835e+01,
+ -1.99943697e+00, 2.59255529e-03, -8.89996147e+00, -1.10976714e+01,
+ -1.43864448e+01, -9.48222409e+00, -1.06785728e+01, -7.24340882e+00,
+ 2.15092681e+00, 1.53607666e+01, 2.06126854e+01, 1.96076182e+01,
+ 3.03104699e+00, -8.52358190e-02, -8.52357557e+00, -1.33461589e+01,
+ -1.37600247e+01, -6.08841095e+00, -8.32367886e+00, -3.02117555e+00,
+ 4.08615082e-01, 1.63346143e+01, 1.76259473e+01, 1.75724049e+01,
+ 1.52688162e+00, -2.23616417e+00, -3.82136854e+00, -1.61943630e+01,
+ -1.55739806e+01, -6.10489716e+00, -6.56542955e+00, -3.79160074e+00,
+ 1.79366664e+00, 1.37690213e+01, 1.71704010e+01, 2.12969028e+01,
+ 2.55881370e+00, -5.89333549e+00, -5.43867513e+00, -9.34441775e+00,
+ -1.23296368e+01, -7.43701484e+00, -9.59827267e+00, -6.98198280e+00,
+ -7.94911839e-01, 1.30601062e+01, 2.03392195e+01, 2.52824447e+01,
+ -3.95418211e+00, 2.43162216e+00, -3.09611231e+00, -1.49779647e+01,
+ -1.07287660e+01, -8.40149898e+00, -1.18887475e+01, -1.74756969e+00,
+ 2.17909158e+00, 1.20038451e+01, 2.42508083e+01, 2.34572756e+01,
+ -5.17568738e+00, -1.96585193e-01, -4.18458348e+00, -1.55118992e+01,
+ -1.38833773e+01, -8.29522246e+00, -1.30003245e+01, -1.67001046e-01,
+ 9.35165464e-01, 1.47274009e+01, 2.29308500e+01, 2.17103726e+01,
+ 3.68218796e+00, 2.64751368e-01, -7.34442896e+00, -1.25122452e+01,
+ -1.14503472e+01, -8.19533891e+00, -1.15456946e+01, -2.81694273e+00,
+ -1.50158220e+00, 1.14252490e+01, 2.08253654e+01, 1.93274939e+01,
+ 7.94218283e-01, -5.10392562e-01, -8.74257956e+00, -9.01561168e+00,
+ -1.00192375e+01, -1.10908742e+01, -1.09129057e+01, -6.64424202e+00,
+ -1.50482563e+00, 1.46897914e+01, 1.73829656e+01, 2.23508516e+01,
+ 8.64908482e+00, 6.22670938e-01, -6.68012958e+00, -5.70808463e+00,
+ -1.80391974e+01, -7.97569860e+00, -1.19962932e+01, -5.55858916e+00,
+ 2.35415063e+00, 1.17526337e+01, 1.54009327e+01, 2.21564076e+01,
+ 3.90926848e+00, 2.21699063e+00, -3.80724386e+00, -1.09345639e+01,
+ -1.37938477e+01, -1.00726110e+01, -1.19963696e+01, -5.40000702e+00,
+ -1.51910929e+00, 1.69895520e+00, 1.74367921e+01, 2.04883238e+01,
+ 7.55305367e+00, 7.29570618e-01, -5.09536099e+00, -1.29493298e+01,
+ -1.53454372e+01, -2.46711622e+00, -1.01903520e+01, -4.03697494e+00,
+ -3.08084548e+00, 3.86928001e+00, 1.92764155e+01, 1.55958052e+01,
+ 7.35560665e+00, 1.85905286e+00, -5.61647492e-01, -1.23394890e+01,
+ -9.90369650e+00, -7.50968724e+00, -1.83651468e+01, -2.77916418e+00,
+ -1.07805825e+00, 8.15877162e+00, 2.33477133e+01, 1.69720395e+01,
+ 6.19355409e+00, 4.92033190e+00, -1.36452236e+01, -1.10382237e+01,
+ -4.45625959e+00, -1.37976278e+01, -1.12070229e+01, -1.28293907e+00,
+ 1.02615489e-01, 1.16373419e+01, 1.73964040e+01, 1.64050904e+01,
+ 1.32632316e+01, 4.44789857e+00, -1.66636700e+01, -1.04932431e+01,
+ -7.27536831e+00, -1.52095878e+01, -8.33331485e+00, -6.12562623e+00,
+ -6.19892381e-01, 1.73375856e+01, 1.71076116e+01, 2.30092371e+01,
+ -1.39793588e+00, 1.20108534e+00, -1.01506292e+01, -9.35709025e+00,
+ -1.72524967e+01, -1.33257487e+01, -1.11436060e+01, -1.07822676e+00,
+ 2.29723021e+00, 1.15489387e+01, 1.72661557e+01, 2.11762682e+01,
+ 9.51783705e+00, -1.02191435e+00, -5.14895585e+00, -2.05301479e+01,
+ -1.56429911e+01, -1.60412160e+01, -1.50915585e+01, -2.94815119e+00,
+ 4.61947140e+00, 6.94204531e+00, 1.79378222e+01, 2.19333496e+01,
+ 8.01926876e+00, -3.09873539e+00, -6.33383956e+00, -1.29668016e+01,
+ -1.54450181e+01, -1.27736754e+01, -1.46733580e+01, -8.76927199e+00,
+ 8.56843050e+00, 1.28259048e+01, 1.86473170e+01, 5.73666651e+00,
+ 4.33460471e+00, 2.08833654e+00, -3.96959363e+00, -1.29223840e+01,
+ -1.19550435e+01, -1.27279210e+01, -8.02537118e+00, -3.92329973e+00,
+ 7.09140567e+00, 2.42153157e+01, 1.28924451e+01, 1.79711994e+01,
+ 2.89522816e+00, 1.30474094e+00, -7.77941829e+00, -1.04361458e+01,
+ -1.14357321e+01, -1.23868252e+01, -3.73410135e+00, 6.47313429e-01,
+ 5.14176514e+00, 1.16621376e+01, 8.00349556e+00, 1.83900860e+01,
+ 3.46846764e+00, 2.29413265e+00, -4.06962578e+00, -8.55164849e+00,
+ -1.76399695e+01, -1.50423508e+01, -1.13765532e+01, -9.17973632e+00,
+ -4.22254178e+00, 2.19090137e+01, 1.90170614e+01, 1.80606278e+01,
+ 4.08981599e+00, 2.02346117e+00, -5.45474659e+00, -1.38725716e+01,
+ -1.50622791e+01, -1.15367789e+01, -7.55445577e+00, -1.77510788e+00,
+ 9.46335947e+00, 4.88813367e+00, 1.61490895e+01, 1.93212548e+01,
+ 1.03075610e+01, -6.46758291e-01, -5.79530543e-01, -1.35917659e+01,
+ -1.62148912e+01, -1.29823949e+01, -1.02149087e+01, -3.24211066e+00,
+ 3.05411201e-01, 1.19385090e+01, 2.08979477e+01, 2.19927470e+01,
+ 1.32364223e+00, 1.68626515e+00, -3.52030557e+00, -1.50337436e+01,
+ -1.75865944e+01, -1.23980840e+01, -1.19670311e+01, -1.59575440e+00,
+ 4.32015112e+00, 1.39461330e+01, 2.63901690e+01, 2.11431667e+01,
+ 1.19960552e+00, 1.22769386e+00, -3.12851420e+00, -1.23388328e+01,
+ -1.66429432e+01, -9.08277509e+00, -7.92637338e+00, 2.43702321e+00,
+ -3.53211182e+00, 1.00606776e+01, 1.39608421e+01, 1.44689452e+01,
+ 6.50770562e+00, 3.13940836e+00, -4.89894478e-01, -1.05833296e+01,
+ -1.34863098e+01, -1.20763793e+01, -1.00738904e+01, -9.39207297e+00]),
+ 'm': 12,
+ 'states': array([[1.24021769e+02, 8.30544193e+01, 8.40569047e+01, 6.14692129e-02,
+ 3.00509837e+02],
+ [8.50161150e+01, 7.80917592e+01, 8.40569047e+01, 6.14692129e-02,
+ 1.33876280e+02],
+ [7.65735210e+01, 7.67280499e+01, 8.40569047e+01, 6.14692129e-02,
+ 8.73485707e+01],
+ ...,
+ [1.12984989e+02, 1.00517846e+02, 8.40569047e+01, 6.14692129e-02,
+ 1.14480779e+02],
+ [1.14049672e+02, 1.00543746e+02, 8.40569047e+01, 6.14692129e-02,
+ 1.13025090e+02],
+ [1.10946036e+02, 1.00561388e+02, 8.40569047e+01, 6.14692129e-02,
+ 1.14089773e+02]]),
+ 'par': {'initial_smoothed': 258.45064973324986,
+ 'alpha': 0.7664297044277045,
+ 'theta': 2.0},
+ 'n': 536,
+ 'modeltype': 'STM',
+ 'mean_y': 100.56138830499272,
+ 'decompose': True,
+ 'decomposition_type': 'additive',
+ 'seas_forecast': {'mean': array([ 0.08977811, 18.09442035, 20.24848682, 19.4306462 ,
+ 2.64008067, -1.30909907, -7.97773123, -12.32640613,
+ -12.02777406, -10.1369666 , -11.42293515, -5.30249992])},
+ 'fitted': array([300.50983667, 133.87628004, 87.34857069, 77.149048 ,
+ 76.98193501, 77.05546632, 77.64431525, 79.83754652,
+ 77.03398836, 75.47241431, 85.74423271, 85.47106297,
+ 86.31849804, 88.14353755, 80.8438024 , 78.37975377,
+ 81.66417589, 83.26287098, 84.62535048, 83.7700806 ,
+ 79.74427087, 80.35553844, 83.81224541, 87.82202841,
+ 86.29562725, 86.14455866, 85.23591135, 85.24504787,
+ 80.98550693, 85.35566468, 88.73347592, 80.13900931,
+ 77.37219166, 71.81797618, 78.98325428, 80.46128324,
+ 70.5292522 , 65.84059407, 56.80056031, 58.2461785 ,
+ 68.88547104, 71.95893388, 73.17068509, 73.63150177,
+ 72.56861641, 67.74141718, 75.823697 , 83.17867619,
+ 82.88182158, 84.77950058, 78.46220336, 71.99792744,
+ 75.71080394, 81.00106285, 78.99654625, 80.35978703,
+ 77.73477595, 77.0624998 , 86.86366484, 90.37877922,
+ 93.59485284, 94.48135717, 92.0871403 , 86.88905458,
+ 88.05659723, 89.54567652, 88.73256441, 87.66000103,
+ 84.49066084, 84.28553517, 89.56282177, 86.79937702,
+ 92.06289264, 88.57657843, 83.39583679, 84.22817596,
+ 88.48908916, 88.70896704, 88.43688493, 84.98139923,
+ 82.12001187, 82.55098375, 85.9525497 , 90.19133861,
+ 93.64043798, 95.47816961, 88.30724672, 88.90190843,
+ 89.38115594, 90.19718831, 91.02162087, 87.36982498,
+ 83.60211476, 81.42239492, 82.66423005, 85.61780804,
+ 86.08812516, 84.73820883, 85.54103724, 83.21124039,
+ 79.16163028, 84.73307206, 86.60047462, 83.45823664,
+ 82.8036783 , 79.0463675 , 81.90332096, 85.42827927,
+ 87.13594255, 92.20371202, 91.92571881, 87.47001337,
+ 89.71173884, 94.08746707, 93.42067364, 91.36829945,
+ 88.10499619, 84.3807048 , 96.69192329, 96.73805538,
+ 94.53625569, 93.65491108, 94.17929385, 91.81488763,
+ 87.30208784, 88.22493606, 88.60917145, 87.97178146,
+ 84.24395894, 81.95085029, 92.78029537, 94.275001 ,
+ 95.43756282, 93.25335889, 89.64588248, 86.84354705,
+ 86.27398255, 87.31900372, 85.50115807, 87.26078723,
+ 85.45187034, 83.45436489, 90.30572204, 87.23685484,
+ 85.22183448, 89.83718668, 88.17711021, 87.21418481,
+ 87.84436043, 89.45885582, 88.22276703, 88.33640489,
+ 86.986106 , 86.10365179, 92.24300578, 94.58859369,
+ 93.69697622, 94.76073782, 89.93749012, 87.8093004 ,
+ 88.3949345 , 89.16392452, 86.74878426, 86.67946215,
+ 85.59093176, 88.57095695, 92.89938688, 93.34684947,
+ 96.69921709, 95.24315643, 95.05395839, 88.76162712,
+ 86.66136449, 88.14065857, 87.23099008, 85.41872572,
+ 85.01380191, 87.60818255, 95.45557821, 98.3240456 ,
+ 96.5726075 , 95.04052437, 95.49116642, 92.53977272,
+ 90.36888696, 90.16393454, 89.5384766 , 88.04727997,
+ 88.67676475, 90.24331499, 100.45849371, 103.6695433 ,
+ 103.36252038, 95.57789385, 96.3997412 , 97.54332409,
+ 94.2091886 , 94.45290287, 96.36634424, 100.85347739,
+ 102.80919411, 102.5472505 , 106.48312008, 104.03628873,
+ 103.29067992, 101.03748421, 103.07742567, 101.82224878,
+ 101.27230355, 100.28657648, 100.63629155, 101.06606288,
+ 101.71464486, 101.14884962, 101.78769257, 102.7950277 ,
+ 105.71545223, 99.33413407, 101.80714675, 102.61832483,
+ 101.21735442, 100.85561542, 102.45166863, 107.05014533,
+ 107.51717356, 108.33874738, 106.85496498, 111.55980808,
+ 114.23636971, 107.06776156, 111.42831624, 112.50186659,
+ 109.36957611, 107.54585583, 111.62426724, 111.62202077,
+ 114.31102418, 111.83959398, 107.39758714, 108.70651652,
+ 106.30953697, 102.78440744, 103.82046147, 103.14437142,
+ 104.11684481, 102.33982409, 102.8723728 , 103.47360882,
+ 102.01677319, 103.62723339, 101.56281457, 101.87268181,
+ 102.03905301, 102.36943582, 103.33817557, 102.95055886,
+ 102.19972468, 100.90281095, 104.03647886, 106.44257555,
+ 108.22178492, 108.49688565, 107.17885267, 105.19959511,
+ 103.80611838, 102.98366417, 102.30386854, 105.52016297,
+ 102.58638056, 99.89919716, 103.02022955, 106.77390074,
+ 107.96263336, 109.29927875, 106.01489901, 103.6864972 ,
+ 105.1475863 , 105.11603549, 101.63327513, 103.61001775,
+ 105.92623683, 105.72561484, 107.82567267, 109.2548828 ,
+ 107.99841184, 107.35109379, 103.5233805 , 103.62365533,
+ 108.13938211, 103.11607784, 106.01381231, 109.78596466,
+ 107.78446605, 108.81079898, 110.17164752, 109.84536969,
+ 112.60070842, 114.23275493, 109.59549173, 112.69372438,
+ 115.81058738, 109.85108519, 110.73448348, 113.67239919,
+ 111.26167729, 109.87022246, 111.31252448, 110.13430105,
+ 114.10103454, 114.77969912, 112.22984999, 114.31642742,
+ 116.09441204, 116.92384863, 118.16082896, 118.67694524,
+ 118.56524723, 119.03853891, 120.55739465, 120.49404273,
+ 122.4297822 , 121.24085099, 116.16013458, 116.63300608,
+ 116.58468172, 115.20069256, 115.84357956, 115.28811168,
+ 117.8563375 , 119.42647419, 118.72610568, 119.14774202,
+ 118.15012563, 116.95870856, 114.38003444, 112.21454843,
+ 114.48341518, 119.11962906, 120.63092958, 121.65618463,
+ 126.75939743, 122.1827986 , 123.8699932 , 123.46128916,
+ 123.29574937, 125.06196634, 120.23216725, 116.54759242,
+ 118.66743152, 119.67090937, 122.40414386, 125.63126387,
+ 126.72874773, 125.40591101, 125.48596965, 125.07720702,
+ 125.03320929, 123.8308448 , 111.2956079 , 109.17137615,
+ 110.01274633, 113.80892938, 115.40216099, 117.64202977,
+ 117.19533719, 114.68331622, 120.59245198, 121.56787494,
+ 122.56854548, 120.16921999, 109.29738449, 108.58309477,
+ 105.67469335, 109.31954714, 111.77844749, 117.49308896,
+ 117.5137965 , 119.17248724, 121.21684679, 115.92686418,
+ 117.89155825, 117.02722838, 109.44298667, 111.84906053,
+ 109.99544591, 112.7496681 , 117.55482364, 113.24182371,
+ 114.25985959, 120.09362779, 117.31872292, 117.51493907,
+ 120.62638451, 120.66695807, 115.74879597, 113.59360961,
+ 111.30546837, 119.47810143, 123.92117003, 117.29474313,
+ 118.73046831, 122.40358785, 118.54651485, 120.94522623,
+ 120.34509238, 119.83191444, 119.28258839, 116.90606293,
+ 119.67953588, 116.61541466, 118.57002916, 116.93539025,
+ 119.24189675, 115.26824869, 112.85500598, 113.09982676,
+ 116.36816979, 118.09076126, 113.10484433, 110.84983175,
+ 112.21846295, 117.52051435, 117.77135585, 119.97014793,
+ 113.71329113, 110.97321598, 106.47875848, 103.69775119,
+ 105.5329286 , 109.03535469, 100.5185778 , 98.7783504 ,
+ 100.72723124, 104.88073539, 103.53983956, 104.83050157,
+ 104.37041809, 101.78207536, 99.79195805, 97.33147199,
+ 94.7051695 , 101.23419515, 97.22698298, 96.03053349,
+ 85.56579529, 86.89526346, 89.52989363, 92.63258395,
+ 92.20654345, 92.29302095, 90.33797118, 92.97269973,
+ 94.06049433, 99.45748428, 104.17945492, 98.57230063,
+ 97.48447184, 97.71075906, 99.74481829, 99.92754582,
+ 101.40703208, 101.89152524, 100.19790135, 106.12158657,
+ 110.71243486, 114.61516239, 109.71600444, 100.36181401,
+ 99.59503236, 100.26066735, 103.05302578, 106.07904849,
+ 109.00286948, 104.73225081, 101.00335324, 101.06963632,
+ 98.12874178, 94.85438633, 97.80873857, 96.89567218,
+ 95.87638401, 97.01823883, 99.60314659, 101.56757157,
+ 100.41327905, 98.11827889, 97.07615577, 100.07180788,
+ 102.80604053, 110.02096633, 99.93001054, 96.81884524,
+ 96.765739 , 102.67305829, 103.21143054, 108.91236591,
+ 107.9732912 , 104.79489485, 102.64480872, 103.60141066,
+ 105.2112888 , 105.40729101, 100.7199523 , 101.24845301,
+ 103.24285777, 102.26463485, 104.59110557, 108.03814361,
+ 105.99389435, 101.76418396, 100.06193105, 99.6756544 ,
+ 102.54734888, 105.82036702, 102.67173097, 107.40963325,
+ 108.75289448, 107.67960614, 109.6546142 , 113.40193278,
+ 113.42314323, 109.91667509, 110.75537338, 113.46597679,
+ 119.42851182, 116.6833224 , 110.55675793, 105.76845483,
+ 101.99639438, 104.99139164, 108.43159448, 114.20122959,
+ 115.56790983, 114.4807793 , 113.0250904 , 114.08977297])}
+```
+
+Let us now visualize the residuals of our models.
+
+As we can see, the result obtained above has an output in a dictionary,
+to extract each element from the dictionary we are going to use the
+`.get()` function to extract the element and then we are going to save
+it in a `pd.DataFrame()`.
+
+```python
+residual=pd.DataFrame(result.get("residuals"), columns=["residual Model"])
+residual
+```
+
+| | residual Model |
+|-----|----------------|
+| 0 | -214.815337 |
+| 1 | -62.056280 |
+| 2 | -21.325671 |
+| ... | ... |
+| 533 | -12.076379 |
+| 534 | -10.073890 |
+| 535 | -9.392073 |
+
+```python
+fig, axs = plt.subplots(nrows=2, ncols=2)
+
+residual.plot(ax=axs[0,0])
+axs[0,0].set_title("Residuals");
+
+sns.distplot(residual, ax=axs[0,1]);
+axs[0,1].set_title("Density plot - Residual");
+
+stats.probplot(residual["residual Model"], dist="norm", plot=axs[1,0])
+axs[1,0].set_title('Plot Q-Q')
+
+plot_acf(residual, lags=35, ax=axs[1,1],color="fuchsia")
+axs[1,1].set_title("Autocorrelation");
+
+plt.show();
+```
+
+
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[90]` means that
+ the model expects the real value to be inside that interval 90% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min. (If you want to speed things up to a couple of seconds,
+remove the AutoModels like `ARIMA` and `Theta`)
+
+```python
+# Prediction
+Y_hat = sf.forecast(df=train, h=horizon, fitted=True)
+Y_hat
+```
+
+| | unique_id | ds | AutoTheta |
+|-----|-----------|------------|------------|
+| 0 | 1 | 2016-09-01 | 111.075915 |
+| 1 | 1 | 2016-10-01 | 129.111292 |
+| 2 | 1 | 2016-11-01 | 131.296093 |
+| ... | ... | ... | ... |
+| 9 | 1 | 2017-06-01 | 101.125782 |
+| 10 | 1 | 2017-07-01 | 99.870548 |
+| 11 | 1 | 2017-08-01 | 106.021718 |
+
+```python
+values=sf.forecast_fitted_values()
+values.head()
+```
+
+| | unique_id | ds | y | AutoTheta |
+|-----|-----------|------------|---------|------------|
+| 0 | 1 | 1972-01-01 | 85.6945 | 300.509837 |
+| 1 | 1 | 1972-02-01 | 71.8200 | 133.876280 |
+| 2 | 1 | 1972-03-01 | 66.0229 | 87.348571 |
+| 3 | 1 | 1972-04-01 | 64.5645 | 77.149048 |
+| 4 | 1 | 1972-05-01 | 65.0100 | 76.981935 |
+
+```python
+StatsForecast.plot(values)
+```
+
+
+
+Adding 95% confidence interval with the forecast method
+
+```python
+sf.forecast(df=train, h=horizon, level=[95])
+```
+
+| | unique_id | ds | AutoTheta | AutoTheta-lo-95 | AutoTheta-hi-95 |
+|-----|-----------|------------|------------|-----------------|-----------------|
+| 0 | 1 | 2016-09-01 | 111.075915 | 90.139234 | 136.011109 |
+| 1 | 1 | 2016-10-01 | 129.111292 | 94.795409 | 160.387128 |
+| 2 | 1 | 2016-11-01 | 131.296093 | 90.579813 | 168.268538 |
+| ... | ... | ... | ... | ... | ... |
+| 9 | 1 | 2017-06-01 | 101.125782 | 41.186268 | 159.159903 |
+| 10 | 1 | 2017-07-01 | 99.870548 | 35.144354 | 152.867267 |
+| 11 | 1 | 2017-08-01 | 106.021718 | 38.753454 | 166.048584 |
+
+```python
+# Merge the forecasts with the true values
+Y_hat1 = test.merge(Y_hat, how='left', on=['unique_id', 'ds'])
+Y_hat1
+```
+
+| | ds | y | unique_id | AutoTheta |
+|-----|------------|----------|-----------|------------|
+| 0 | 2016-09-01 | 109.3191 | 1 | 111.075915 |
+| 1 | 2016-10-01 | 119.0502 | 1 | 129.111292 |
+| 2 | 2016-11-01 | 116.8431 | 1 | 131.296093 |
+| ... | ... | ... | ... | ... |
+| 9 | 2017-06-01 | 104.2022 | 1 | 101.125782 |
+| 10 | 2017-07-01 | 102.5861 | 1 | 99.870548 |
+| 11 | 2017-08-01 | 114.0613 | 1 | 106.021718 |
+
+```python
+sf.plot(train, Y_hat1)
+```
+
+
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[95]` means that
+ the model expects the real value to be inside that interval 95% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+sf.predict(h=horizon)
+```
+
+| | unique_id | ds | AutoTheta |
+|-----|-----------|------------|------------|
+| 0 | 1 | 2016-09-01 | 111.075915 |
+| 1 | 1 | 2016-10-01 | 129.111292 |
+| 2 | 1 | 2016-11-01 | 131.296093 |
+| ... | ... | ... | ... |
+| 9 | 1 | 2017-06-01 | 101.125782 |
+| 10 | 1 | 2017-07-01 | 99.870548 |
+| 11 | 1 | 2017-08-01 | 106.021718 |
+
+```python
+forecast_df = sf.predict(h=horizon, level=[95])
+forecast_df
+```
+
+| | unique_id | ds | AutoTheta | AutoTheta-lo-95 | AutoTheta-hi-95 |
+|-----|-----------|------------|------------|-----------------|-----------------|
+| 0 | 1 | 2016-09-01 | 111.075915 | 90.139234 | 136.011109 |
+| 1 | 1 | 2016-10-01 | 129.111292 | 94.795409 | 160.387128 |
+| 2 | 1 | 2016-11-01 | 131.296093 | 90.579813 | 168.268538 |
+| ... | ... | ... | ... | ... | ... |
+| 9 | 1 | 2017-06-01 | 101.125782 | 41.186268 | 159.159903 |
+| 10 | 1 | 2017-07-01 | 99.870548 | 35.144354 | 152.867267 |
+| 11 | 1 | 2017-08-01 | 106.021718 | 38.753454 | 166.048584 |
+
+```python
+sf.plot(train, test.merge(forecast_df), level=[95])
+```
+
+
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=5)`, forecasting every second months
+`(step_size=12)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 12 months ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(
+ df=train,
+ h=horizon,
+ step_size=12,
+ n_windows=5
+)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` series identifier.
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the n_windows.
+- `y:` true value
+- `"model":` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df
+```
+
+| | unique_id | ds | cutoff | y | AutoTheta |
+|-----|-----------|------------|------------|----------|------------|
+| 0 | 1 | 2011-09-01 | 2011-08-01 | 93.9062 | 98.167469 |
+| 1 | 1 | 2011-10-01 | 2011-08-01 | 116.7634 | 116.969932 |
+| 2 | 1 | 2011-11-01 | 2011-08-01 | 116.8258 | 119.135142 |
+| ... | ... | ... | ... | ... | ... |
+| 57 | 1 | 2016-06-01 | 2015-08-01 | 102.4044 | 109.600469 |
+| 58 | 1 | 2016-07-01 | 2015-08-01 | 102.9512 | 108.260160 |
+| 59 | 1 | 2016-08-01 | 2015-08-01 | 104.6977 | 114.248270 |
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=season_length), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | AutoTheta |
+|-----|-----------|--------|-----------|
+| 0 | 1 | mae | 6.281513 |
+| 1 | 1 | mape | 0.055683 |
+| 2 | 1 | mase | 1.212473 |
+| 3 | 1 | rmse | 7.683669 |
+| 4 | 1 | smape | 0.027399 |
+
+## References
+
+1. [Jose A. Fiorucci, Tiago R. Pellegrini, Francisco Louzada, Fotios
+ Petropoulos, Anne B. Koehler (2016). “Models for optimising the
+ theta method and their relationship to state space models”.
+ International Journal of
+ Forecasting](https://www.sciencedirect.com/science/article/pii/S0169207016300243).
+2. [Nixtla AutoTheta API](../../src/core/models.html#autotheta)
+3. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+4. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html)
+5. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/models/crostonclassic.html.mdx b/statsforecast/docs/models/crostonclassic.html.mdx
new file mode 100644
index 00000000..e4add6cd
--- /dev/null
+++ b/statsforecast/docs/models/crostonclassic.html.mdx
@@ -0,0 +1,825 @@
+---
+title: CrostonClassic Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `CrostonClassic Model` with
+> `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation` in other.
+
+The text in this article is largely taken from: 1. [Changquan Huang •
+Alla Petukhina. Springer series (2022). Applied Time Series Analysis and
+Forecasting with
+Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2) 2.
+Ivan Svetunkov. [Forecasting and Analytics with the Augmented Dynamic
+Adaptive Model (ADAM)](https://openforecast.org/adam/) 3. [James D.
+Hamilton. Time Series Analysis Princeton University Press, Princeton,
+New Jersey, 1st Edition,
+1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+Principles and Practice (3rd ed)”](https://otexts.com/fpp3/tscv.html).
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [Croston Classic Model](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of CrostonClassic with
+ StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+The Croston model is a method used in time series analysis to forecast
+demand in situations where there are intermittent data or frequent
+zeros. It was developed by J.D. Croston in 1972 and is especially useful
+in industries such as inventory management, retail sales, and demand
+forecasting for products with low sales frequency.
+
+The Croston model is based on two main components:
+
+1. Intermittent Demand Rate: Calculates the demand rate for periods in
+ which sales or events occur, ignoring periods without sales. This
+ rate is used to estimate the probability that a claim will occur in
+ the future.
+
+2. Demand Interval: Calculates the time interval between sales or
+ events occurring, again ignoring non-sales periods. This interval is
+ used to estimate the probability that a demand will occur in the
+ next period.
+
+The Croston model combines these two estimates to generate a weighted
+forecast that takes into account both the rate of intermittent demand
+and the interval between demands. This approach helps address the
+challenge of forecasting demand in situations where the time series has
+many zeros or missing values.
+
+It is important to note that the Croston model is a simplification and
+does not account for other possible sources of variability or patterns
+in the demand data. Therefore, its accuracy may be affected in
+situations where there are external factors or changes in demand
+behavior.
+
+## Croston Classic Model
+
+### What is intermittent demand?
+
+Intermittent demand is a demand pattern characterized by the irregular
+and sporadic occurrence of events or sales. In other words, it refers to
+situations in which the demand for a product or service occurs
+intermittently, with periods of time in which there are no sales or
+significant events.
+
+Intermittent demand differs from constant or regular demand, where sales
+occur in a predictable and consistent manner over time. In contrast, in
+intermittent demand, periods without sales may be long and there may not
+be a regular sequence of events.
+
+This type of demand can occur in different industries and contexts, such
+as low consumption products, seasonal products, high variability
+products, products with short life cycles, or in situations where demand
+depends on specific events or external factors.
+
+Intermittent demand can pose challenges in forecasting and inventory
+management, as it is difficult to predict when sales will occur and in
+what quantity. Methods like the Croston model, which I mentioned
+earlier, are used to address intermittent demand and generate more
+accurate and appropriate forecasts for this type of demand pattern.
+
+### Problem with intermittent demand
+
+Intermittent demand can present various challenges and issues in
+inventory management and demand forecasting. Some of the common problems
+associated with intermittent demand are as follows:
+
+1. Unpredictable variability: Intermittent demand can have
+ unpredictable variability, making planning and forecasting
+ difficult. Demand patterns can be irregular and fluctuate
+ dramatically between periods with sales and periods without sales.
+
+2. Low frequency of sales: Intermittent demand is characterized by long
+ periods without sales. This can lead to inventory management
+ difficulties, as it is necessary to hold enough stock to meet demand
+ when it occurs, while avoiding excess inventory during non-sales
+ periods.
+
+3. Forecast error: Forecasting intermittent demand can be more
+ difficult to pin down than constant demand. Traditional forecast
+ models may not be adequate to capture the variability and lack of
+ patterns in intermittent demand, which can lead to significant
+ errors in estimates of future demand.
+
+4. Impact on the supply chain: Intermittent demand can affect the
+ efficiency of the supply chain and create difficulties in production
+ planning, supplier management and logistics. Lead times and
+ inventory levels must be adjusted to meet unpredictable demand.
+
+5. Operating costs: Managing inventory in situations of intermittent
+ demand can increase operating costs. Maintaining adequate inventory
+ during non-sales periods and managing stock levels may require
+ additional investments in storage and logistics.
+
+To address these issues, specific approaches to intermittent demand
+management are used, such as specialized forecasting models, product
+classification techniques, and tailored inventory strategies. These
+solutions seek to minimize the impacts of variability and lack of
+patterns in intermittent demand, optimizing inventory management and
+improving supply chain efficiency.
+
+### Croston’s method(CR)
+
+Croston’s method(CR) is a classic method that specifically dealing with
+intermittent demand, it was developed base upon the Simple Exponential
+Smoothing method. When Croston dealing with the intermittent demand, he
+found out that by using the SES, the level of forecasting in each
+period’s demand are normally higher than it’s actual value, which lead
+to a very low accuracy. After a period of times of research, he came out
+a method that optimize the result of the intermittent demand
+forecasting.
+
+This method basically decompose the intermittent demand into two parts:
+the size of non-zero demand and the time interval of those demand
+occurred, and then apply the simple exponential smoothing on both part.
+Where the formula is follow:
+
+if $Z_t=0$ then:
+
+$$Z'_t= Z'_{t-1}$$
+
+$$P'_t= P'_{t-1}$$
+
+Otherwise
+
+$$Z'_t=\alpha Z_t +(1-\alpha) Z'_{t-1}$$
+
+$$P'_t=\alpha P_t +(1-\alpha) P'_{t-1}$$
+
+where $0< \alpha < 1$
+
+And finally by combining these forecasts
+
+$${Y'}_t = \frac{{Z'}_t}{{P'}_t}$$
+
+Where
+
+- ${Y'}_t:$ Average demand per period.
+- $Z_t:$ Actual demand at period $t$.
+- $Z'_t:$ Time between two positive demand.
+- $P:$ Demand size forecast for next period.
+- $P_t:$ Forecast of demand interval.
+- $\alpha :$ Smoothing constant.
+
+Croston’s method converse the intermittent demand time series into a
+non-zero demand time series and a demand interval time series, many
+cases show that this method work quite well, but before apply Croston’s
+method, three assumptions should be made:
+
+- The non-zero demand are independent and obey normal distribution;
+- The demand intervals are independent and obey geometric
+ distribution;
+- There are mutual independence between the demand size and demand
+ intervals.
+
+According to many real cases show that, Croston’s method is suitable for
+the situation which the lead time obey normal distribution, for those
+demand series which contain large amount of zero values, Croston’s
+method did not shows a outstanding performance, sometimes even worse
+than SES method.
+
+Additionally, Croston’s method can only provide the average demand for
+each period, it can not give a forecast of the demand size for each
+period, it can not forecast which period will occurred a demand, and it
+also can not come out a probability of whether a period will occurred a
+demand.
+
+After all, although Croston’s method is a very classic and wide use
+method, it still has a lots of limitations, but after years of research
+carried by statisticians and scholars, few variations of Croston’s
+method were brought up.
+
+### Croston’s variations
+
+Croston’s method is the main model used in demand forecasting area, most
+of the works are based upon this model. However, in 2001 Syntetos and
+Boylan proposed that Croston’s method is no a unbiased method, while
+some empirical evidence also showed that the losses in performance which
+use the Croston’s method (Sani and Kingsman, 1997). Plenty of further
+research is done in improving the Croston’s method. Syntetos and Boylan
+(2005) proposed an approximate unbiased procedure that provide less
+variance in the result of estimate, which is known as SBA (Syntetos and
+Boylan Approximate). Recently, Teunter et al. (2011) also proposed a
+intermittent forecasting method that can deal with obsolescence, which
+is based on Croston’s method known as TSB method (Teunter, Syntetos and
+Babai).
+
+### Area of application of the Croston method
+
+The Croston method is commonly applied in the field of inventory
+management and demand forecasting in situations of intermittent demand.
+Some specific areas where the Croston model can be applied are:
+
+1. Inventory management: The Croston model is used to forecast demand
+ for products with sporadic or intermittent sales. Helps determine
+ optimal inventory levels and replenishment policies, minimizing
+ inventory costs and ensuring adequate availability to meet
+ intermittent demand.
+
+2. Retail sales: In the retail sector, especially in products with low
+ sales frequency or irregular sales, the Croston model can be useful
+ for forecasting demand and optimizing inventory planning in stores
+ or warehouses.
+
+3. Demand forecasting: In general, the Croston model is applied in
+ demand forecasting when there is a lack of clear patterns or high
+ variability in the time series. It can be used in various
+ industries, such as the pharmaceutical industry, the automotive
+ industry, the perishable goods industry, and other sectors where
+ intermittent demand is common.
+
+4. Supply Chain Planning: The Croston model can be used in supply chain
+ planning and management to improve the accuracy of intermittent
+ demand forecasts. This helps streamline production, inventory
+ management, supplier order scheduling, and other aspects of the
+ supply chain.
+
+It is important to note that Croston’s model is just one of many
+approaches available to address intermittent demand. Depending on the
+context and the specific characteristics of the time series, there may
+be other more appropriate methods and techniques.
+
+### Croston Method for Stationary Time Series
+
+No, the time series in the Croston method does not have to be
+stationary. The Croston method is an effective forecasting method for
+intermittent time series, even if they are not stationary. However, if
+the time series is stationary, the Croston method may be more accurate.
+
+The Croston method is based on the idea that intermittent time series
+can be decomposed into two components: a demand component and a time
+between demands component. The demand component is forecast using a
+standard time series forecasting method, such as single or double
+exponential smoothing. The time component between demands is forecast
+using a probability distribution function, such as a Poisson
+distribution or a Weibull distribution.
+
+The Croston method then combines the forecasts for the two components to
+obtain a total demand forecast for the next period.
+
+If the time series is stationary, the two components of the time series
+will be stationary as well. This means that the Croston method will be
+able to forecast the two components more accurately.
+
+However, even if the time series is not stationary, the Croston method
+can still be an effective forecasting method. The Croston method is a
+robust method that can handle time series with irregular demand
+patterns.
+
+If you are using the Croston method to forecast an intermittent time
+series that is not stationary, it is important to choose a standard time
+series forecast method that is effective for nonstationary time series.
+Double exponential smoothing is an effective forecasting method for
+non-stationary time series.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
+plt.style.use('grayscale') # fivethirtyeight grayscale classic
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#008080', # #212946
+ 'axes.facecolor': '#008080',
+ 'savefig.facecolor': '#008080',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#000000', #2A3459
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+
+```python
+import pandas as pd
+
+df=pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/intermittend_demand2")
+df.head()
+```
+
+| | date | sales |
+|-----|---------------------|-------|
+| 0 | 2022-01-01 00:00:00 | 0 |
+| 1 | 2022-01-01 01:00:00 | 10 |
+| 2 | 2022-01-01 02:00:00 | 0 |
+| 3 | 2022-01-01 03:00:00 | 0 |
+| 4 | 2022-01-01 04:00:00 | 100 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|---------------------|-----|-----------|
+| 0 | 2022-01-01 00:00:00 | 0 | 1 |
+| 1 | 2022-01-01 01:00:00 | 10 | 1 |
+| 2 | 2022-01-01 02:00:00 | 0 | 1 |
+| 3 | 2022-01-01 03:00:00 | 0 | 1 |
+| 4 | 2022-01-01 04:00:00 | 100 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y int64
+unique_id object
+dtype: object
+```
+
+We can see that our time variable `(ds)` is in an object format, we need
+to convert to a date format
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore Data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints a random series from the dataset and is useful for
+basic EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### Autocorrelation plots
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=30, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+plot_pacf(df["y"], lags=30, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+plt.show();
+```
+
+
+
+### Decomposition of the time series
+
+How to decompose a time series and why?
+
+In time series analysis to forecast new values, it is very important to
+know past data. More formally, we can say that it is very important to
+know the patterns that values follow over time. There can be many
+reasons that cause our forecast values to fall in the wrong direction.
+Basically, a time series consists of four components. The variation of
+those components causes the change in the pattern of the time series.
+These components are:
+
+- **Level:** This is the primary value that averages over time.
+- **Trend:** The trend is the value that causes increasing or
+ decreasing patterns in a time series.
+- **Seasonality:** This is a cyclical event that occurs in a time
+ series for a short time and causes short-term increasing or
+ decreasing patterns in a time series.
+- **Residual/Noise:** These are the random variations in the time
+ series.
+
+Combining these components over time leads to the formation of a time
+series. Most time series consist of level and noise/residual and trend
+or seasonality are optional values.
+
+If seasonality and trend are part of the time series, then there will be
+effects on the forecast value. As the pattern of the forecasted time
+series may be different from the previous time series.
+
+The combination of the components in time series can be of two types: \*
+Additive \* Multiplicative
+
+### Additive time series
+
+If the components of the time series are added to make the time series.
+Then the time series is called the additive time series. By
+visualization, we can say that the time series is additive if the
+increasing or decreasing pattern of the time series is similar
+throughout the series. The mathematical function of any additive time
+series can be represented by:
+$$y(t) = level + Trend + seasonality + noise$$
+
+### Multiplicative time series
+
+If the components of the time series are multiplicative together, then
+the time series is called a multiplicative time series. For
+visualization, if the time series is having exponential growth or
+decline with time, then the time series can be considered as the
+multiplicative time series. The mathematical function of the
+multiplicative time series can be represented as.
+
+$$y(t) = Level * Trend * seasonality * Noise$$
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+from plotly.subplots import make_subplots
+import plotly.graph_objects as go
+
+def plotSeasonalDecompose(
+ x,
+ model='additive',
+ filt=None,
+ period=None,
+ two_sided=True,
+ extrapolate_trend=0,
+ title="Seasonal Decomposition"):
+
+ result = seasonal_decompose(
+ x, model=model, filt=filt, period=period,
+ two_sided=two_sided, extrapolate_trend=extrapolate_trend)
+ fig = make_subplots(
+ rows=4, cols=1,
+ subplot_titles=["Observed", "Trend", "Seasonal", "Residuals"])
+ for idx, col in enumerate(['observed', 'trend', 'seasonal', 'resid']):
+ fig.add_trace(
+ go.Scatter(x=result.observed.index, y=getattr(result, col), mode='lines'),
+ row=idx+1, col=1,
+ )
+ return fig
+```
+
+
+```python
+plotSeasonalDecompose(
+ df["y"],
+ model="additive",
+ period=24,
+ title="Seasonal Decomposition")
+```
+
+
+
+## Split the data into training and testing
+
+Let’s divide our data into sets
+
+1. Data to train our `Croston Classic Model`.
+2. Data to test our model
+
+For the test data we will use the last 500 hours to test and evaluate
+the performance of our model.
+
+```python
+train = df[df.ds\<='2023-01-31 19:00:00']
+test = df[df.ds>'2023-01-31 19:00:00']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((9500, 3), (500, 3))
+```
+
+Now let’s plot the training data and the test data.
+
+```python
+sns.lineplot(train,x="ds", y="y", label="Train", linestyle="--",linewidth=2)
+sns.lineplot(test, x="ds", y="y", label="Test", linewidth=2, color="yellow")
+plt.title("Store visit");
+plt.show()
+```
+
+
+
+## Implementation of CrostonClassic with StatsForecast
+
+To also know more about the parameters of the functions of the
+`CrostonClassic Model`, they are listed below. For more information,
+visit the [documentation](../../src/core/models.html#crostonclassic)
+
+``` text
+alias : str
+ Custom name of the model.
+```
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import CrostonClassic
+```
+
+### Instantiating Model
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/) by the
+master, Rob Hyndmann, can be useful for `season_length`.
+
+```python
+season_length = 24 # Hourly data
+horizon = len(test) # number of predictions
+
+models = [CrostonClassic()]
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [pandas’
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models, freq='h')
+```
+
+### Fit the Model
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[CrostonClassic])
+```
+
+Let’s see the results of our `Croston Classic Model`. We can observe it
+with the following instruction:
+
+```python
+result=sf.fitted_[0,0].model_
+result
+```
+
+``` text
+{'mean': array([27.41841685]),
+ 'fitted': array([ nan, 0. , 5. , ..., 30.61961, 30.61961, 30.61961],
+ dtype=float32),
+ 'sigma': np.float32(49.5709)}
+```
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 25 week ahead.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min.
+
+```python
+Y_hat = sf.forecast(df=train, h=horizon)
+Y_hat
+```
+
+| | unique_id | ds | CrostonClassic |
+|-----|-----------|---------------------|----------------|
+| 0 | 1 | 2023-01-31 20:00:00 | 27.418417 |
+| 1 | 1 | 2023-01-31 21:00:00 | 27.418417 |
+| 2 | 1 | 2023-01-31 22:00:00 | 27.418417 |
+| ... | ... | ... | ... |
+| 497 | 1 | 2023-02-21 13:00:00 | 27.418417 |
+| 498 | 1 | 2023-02-21 14:00:00 | 27.418417 |
+| 499 | 1 | 2023-02-21 15:00:00 | 27.418417 |
+
+```python
+sf.plot(train, Y_hat, max_insample_length=500)
+```
+
+
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 500 hours ahead.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+forecast_df = sf.predict(h=horizon)
+forecast_df
+```
+
+| | unique_id | ds | CrostonClassic |
+|-----|-----------|---------------------|----------------|
+| 0 | 1 | 2023-01-31 20:00:00 | 27.418417 |
+| 1 | 1 | 2023-01-31 21:00:00 | 27.418417 |
+| 2 | 1 | 2023-01-31 22:00:00 | 27.418417 |
+| ... | ... | ... | ... |
+| 497 | 1 | 2023-02-21 13:00:00 | 27.418417 |
+| 498 | 1 | 2023-02-21 14:00:00 | 27.418417 |
+| 499 | 1 | 2023-02-21 15:00:00 | 27.418417 |
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=)`, forecasting every second hour
+`(step_size=50)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents $h$ steps into the future that are being
+ forecasted. In this case, 500 hours ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=df,
+ h=horizon,
+ step_size=50,
+ n_windows=5)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` series identifier.
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the `n_windows`.
+- `y:` true value
+- `model:` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df
+```
+
+| | unique_id | ds | cutoff | y | CrostonClassic |
+|------|-----------|---------------------|---------------------|------|----------------|
+| 0 | 1 | 2023-01-23 12:00:00 | 2023-01-23 11:00:00 | 0.0 | 23.655830 |
+| 1 | 1 | 2023-01-23 13:00:00 | 2023-01-23 11:00:00 | 0.0 | 23.655830 |
+| 2 | 1 | 2023-01-23 14:00:00 | 2023-01-23 11:00:00 | 0.0 | 23.655830 |
+| ... | ... | ... | ... | ... | ... |
+| 2497 | 1 | 2023-02-21 13:00:00 | 2023-01-31 19:00:00 | 60.0 | 27.418417 |
+| 2498 | 1 | 2023-02-21 14:00:00 | 2023-01-31 19:00:00 | 20.0 | 27.418417 |
+| 2499 | 1 | 2023-02-21 15:00:00 | 2023-01-31 19:00:00 | 20.0 | 27.418417 |
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=season_length), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | CrostonClassic |
+|-----|-----------|--------|----------------|
+| 0 | 1 | mae | 33.704756 |
+| 1 | 1 | mape | 0.632593 |
+| 2 | 1 | mase | 0.804074 |
+| 3 | 1 | rmse | 45.262709 |
+| 4 | 1 | smape | 0.767960 |
+
+# References
+
+1. [Changquan Huang • Alla Petukhina. Springer series (2022). Applied
+ Time Series Analysis and Forecasting with
+ Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2)
+2. Ivan Svetunkov. [Forecasting and Analytics with the Augmented
+ Dynamic Adaptive Model (ADAM)](https://openforecast.org/adam/)
+3. [James D. Hamilton. Time Series Analysis Princeton University Press,
+ Princeton, New Jersey, 1st Edition,
+ 1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Nixtla CrostonClassic API](../../src/core/models.html#crostonclassic)
+5. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+6. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+7. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/models/crostonoptimized.html.mdx b/statsforecast/docs/models/crostonoptimized.html.mdx
new file mode 100644
index 00000000..b606be02
--- /dev/null
+++ b/statsforecast/docs/models/crostonoptimized.html.mdx
@@ -0,0 +1,722 @@
+---
+title: CrostonOptimized Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `CrostonOptimized Model` with
+> `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation` in other.
+
+The text in this article is largely taken from: 1. [Changquan Huang •
+Alla Petukhina. Springer series (2022). Applied Time Series Analysis and
+Forecasting with
+Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2) 2.
+Ivan Svetunkov. [Forecasting and Analytics with the Augmented Dynamic
+Adaptive Model (ADAM)](https://openforecast.org/adam/) 3. [James D.
+Hamilton. Time Series Analysis Princeton University Press, Princeton,
+New Jersey, 1st Edition,
+1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+Principles and Practice (3rd ed)”](https://otexts.com/fpp3/tscv.html).
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [Croston Optimized Model](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of CrostonOptimized with
+ StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+The Croston Optimized model is a forecasting method designed for
+intermittent demand time series data. It is an extension of the
+Croston’s method, which was originally developed for forecasting
+sporadic demand patterns.
+
+Intermittent demand time series are characterized by irregular and
+sporadic occurrences of non-zero demand values, often with long periods
+of zero demand. Traditional forecasting methods may struggle to handle
+such patterns effectively.
+
+The Croston Optimized model addresses this challenge by incorporating
+two key components: exponential smoothing and intermittent demand
+estimation.
+
+1. Exponential Smoothing: The Croston Optimized model uses exponential
+ smoothing to capture the trend and seasonality in the intermittent
+ demand data. This helps in identifying the underlying patterns and
+ making more accurate forecasts.
+
+2. Intermittent Demand Estimation: Since intermittent demand data often
+ consists of long periods of zero demand, the Croston Optimized model
+ employs a separate estimation process for the occurrence and size of
+ non-zero demand values. It estimates the probability of occurrence
+ and the average size of non-zero demand intervals, enabling better
+ forecasting of intermittent demand.
+
+The Croston Optimized model aims to strike a balance between
+over-forecasting and under-forecasting intermittent demand, which are
+common challenges in traditional forecasting methods. By explicitly
+modeling intermittent demand patterns, it can provide more accurate
+forecasts for intermittent demand time series data.
+
+It is worth noting that there are variations and adaptations of the
+Croston Optimized model, with different modifications and enhancements
+made to suit specific forecasting scenarios. These variations may
+incorporate additional features or algorithms to further improve the
+accuracy of the forecasts.
+
+## Croston Optimized method
+
+The Croston Optimized model can be mathematically defined as follows:
+
+1. Initialization:
+ - Let $(y_t)$ represent the intermittent demand time series data
+ at time $t$.
+ - Initialize two sets of variables: $(p_t)$ for the probability of
+ occurrence and $(q_t)$ for the average size of non-zero demand
+ intervals.
+ - Initialize the forecast $(F_t)$ and forecast error $(E_t)$
+ variables as zero.
+2. Calculation of $(p_t)$ and $(q_t)$:
+ - Calculate the intermittent demand occurrence probability $(p_t)$
+ using exponential smoothing:
+ $$[p_t = \alpha + (1 - \alpha)(p_{t-1}),]$$ where $(\alpha)$ is
+ the smoothing parameter (typically set between 0.1 and 0.3).
+ - Calculate the average size of non-zero demand intervals $(q_t)$
+ using exponential smoothing:
+ $$[q_t = \beta \cdot y_t + (1 - \beta)(q_{t-1}),]$$ where
+ $(\beta)$ is the smoothing parameter (typically set between 0.1
+ and 0.3).
+3. Forecasting:
+ - If $(y_t > 0)$ (non-zero demand occurrence):
+ - Calculate the forecast $(F_t)$ as the previous forecast
+ $(F_{t-1})$ divided by the average size of non-zero demand
+ intervals $(q_{t-1})$:
+ $$[F_t = \frac{{F_{t-1}}}{{q_{t-1}}}]$$
+ - Calculate the forecast error $(E_t)$ as the difference
+ between the actual demand $(y_t)$ and the forecast $(F_t)$:
+ $$[E_t = y_t - F_t]$$
+ - If $(y_t = 0)$ (zero demand occurrence):
+ - Set the forecast $(F_t)$ and forecast error $(E_t)$ as zero.
+4. Updating the model:
+ - Update the intermittent demand occurrence probability $(p_t)$
+ and the average size of non-zero demand intervals $(q_t)$ using
+ exponential smoothing as described in step 2.
+5. Repeat steps 3 and 4 for each time point in the time series.
+
+The Croston Optimized model leverages exponential smoothing to capture
+the trend and seasonality in the intermittent demand data, and it
+estimates the occurrence probability and average size of non-zero demand
+intervals separately to handle intermittent demand patterns effectively.
+By updating the model parameters based on the observed data, it provides
+improved forecasts for intermittent demand time series.
+
+### Some properties of the Optimized Croston Model
+
+The optimized Croston model is a modification of the classic Croston
+model used to forecast intermittent demand. The classic Croston model
+forecasts demand using a weighted average of historical orders and the
+average interval between orders. The optimized Croston model uses a
+probability function to forecast the mean interval between orders.
+
+The optimized Croston model has been shown to be more accurate than the
+classical Croston model for time series with irregular demand. The
+optimized Croston model is also more adaptable to different types of
+intermittent time series.
+
+The optimized Croston model has the following properties:
+
+- It is accurate, even for time series with irregular demand.
+- It is adaptable to different types of intermittent time series.
+- It is easy to implement and understand.
+- It is robust to outliers.
+
+The optimized Croston model has been used successfully to forecast a
+wide range of intermittent time series, including product demand,
+service demand, and resource demand.
+
+Here are some of the properties of the optimized Croston model:
+
+- **Precision:** The optimized Croston model has been shown to be more
+ accurate than the classic Croston model for time series with
+ irregular demand. This is because the optimized Croston model uses a
+ probability function to forecast the average interval between
+ orders, which is more accurate than the weighted average of
+ historical orders.
+- **Adaptability:** The optimized Croston model is also more adaptable
+ to different types of intermittent time series. This is because the
+ optimized Croston model uses a probability function to forecast the
+ mean interval between orders, allowing it to accommodate different
+ demand patterns.
+- **Ease of Implementation and Understanding:** The optimized Croston
+ model is easy to implement and understand. This is because the
+ optimized Croston model is a modification of the classical Croston
+ model, which is a well-known and well-understood model.
+- **Robustness:** The optimized Croston model is also robust to
+ outliers. This is because the optimized Croston model uses a
+ probability function to forecast the mean interval between orders,
+ which allows it to ignore outliers.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
+import plotly.graph_objects as go
+plt.style.use('grayscale') # fivethirtyeight grayscale classic
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#008080', # #212946
+ 'axes.facecolor': '#008080',
+ 'savefig.facecolor': '#008080',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#000000', #2A3459
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+
+```python
+import pandas as pd
+df=pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/intermittend_demand2")
+
+df.head()
+```
+
+| | date | sales |
+|-----|---------------------|-------|
+| 0 | 2022-01-01 00:00:00 | 0 |
+| 1 | 2022-01-01 01:00:00 | 10 |
+| 2 | 2022-01-01 02:00:00 | 0 |
+| 3 | 2022-01-01 03:00:00 | 0 |
+| 4 | 2022-01-01 04:00:00 | 100 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|---------------------|-----|-----------|
+| 0 | 2022-01-01 00:00:00 | 0 | 1 |
+| 1 | 2022-01-01 01:00:00 | 10 | 1 |
+| 2 | 2022-01-01 02:00:00 | 0 | 1 |
+| 3 | 2022-01-01 03:00:00 | 0 | 1 |
+| 4 | 2022-01-01 04:00:00 | 100 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y int64
+unique_id object
+dtype: object
+```
+
+We can see that our time variable `(ds)` is in an object format, we need
+to convert to a date format
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore Data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints a random series from the dataset and is useful for
+basic EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### Autocorrelation plots
+
+Autocorrelation (ACF) and partial autocorrelation (PACF) plots are
+statistical tools used to analyze time series. ACF charts show the
+correlation between the values of a time series and their lagged values,
+while PACF charts show the correlation between the values of a time
+series and their lagged values, after the effect of previous lagged
+values has been removed.
+
+ACF and PACF charts can be used to identify the structure of a time
+series, which can be helpful in choosing a suitable model for the time
+series. For example, if the ACF chart shows a repeating peak and valley
+pattern, this indicates that the time series is stationary, meaning that
+it has the same statistical properties over time. If the PACF chart
+shows a pattern of rapidly decreasing spikes, this indicates that the
+time series is invertible, meaning it can be reversed to get a
+stationary time series.
+
+The importance of the ACF and PACF charts is that they can help analysts
+better understand the structure of a time series. This understanding can
+be helpful in choosing a suitable model for the time series, which can
+improve the ability to predict future values of the time series.
+
+To analyze ACF and PACF charts:
+
+- Look for patterns in charts. Common patterns include repeating peaks
+ and valleys, sawtooth patterns, and plateau patterns.
+- Compare ACF and PACF charts. The PACF chart generally has fewer
+ spikes than the ACF chart.
+- Consider the length of the time series. ACF and PACF charts for
+ longer time series will have more spikes.
+- Use a confidence interval. The ACF and PACF plots also show
+ confidence intervals for the autocorrelation values. If an
+ autocorrelation value is outside the confidence interval, it is
+ likely to be significant.
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=30, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+plot_pacf(df["y"], lags=30, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+plt.show();
+```
+
+
+
+### Decomposition of the time series
+
+How to decompose a time series and why?
+
+In time series analysis to forecast new values, it is very important to
+know past data. More formally, we can say that it is very important to
+know the patterns that values follow over time. There can be many
+reasons that cause our forecast values to fall in the wrong direction.
+Basically, a time series consists of four components. The variation of
+those components causes the change in the pattern of the time series.
+These components are:
+
+- **Level:** This is the primary value that averages over time.
+- **Trend:** The trend is the value that causes increasing or
+ decreasing patterns in a time series.
+- **Seasonality:** This is a cyclical event that occurs in a time
+ series for a short time and causes short-term increasing or
+ decreasing patterns in a time series.
+- **Residual/Noise:** These are the random variations in the time
+ series.
+
+Combining these components over time leads to the formation of a time
+series. Most time series consist of level and noise/residual and trend
+or seasonality are optional values.
+
+If seasonality and trend are part of the time series, then there will be
+effects on the forecast value. As the pattern of the forecasted time
+series may be different from the previous time series.
+
+The combination of the components in time series can be of two types: \*
+Additive \* Multiplicative
+
+### Additive time series
+
+If the components of the time series are added to make the time series.
+Then the time series is called the additive time series. By
+visualization, we can say that the time series is additive if the
+increasing or decreasing pattern of the time series is similar
+throughout the series. The mathematical function of any additive time
+series can be represented by:
+$$y(t) = level + Trend + seasonality + noise$$
+
+### Multiplicative time series
+
+If the components of the time series are multiplicative together, then
+the time series is called a multiplicative time series. For
+visualization, if the time series is having exponential growth or
+decline with time, then the time series can be considered as the
+multiplicative time series. The mathematical function of the
+multiplicative time series can be represented as.
+
+$$y(t) = Level * Trend * seasonality * Noise$$
+
+```python
+from plotly.subplots import make_subplots
+```
+
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+
+def plotSeasonalDecompose(
+ x,
+ model='additive',
+ filt=None,
+ period=None,
+ two_sided=True,
+ extrapolate_trend=0,
+ title="Seasonal Decomposition"):
+
+ result = seasonal_decompose(
+ x, model=model, filt=filt, period=period,
+ two_sided=two_sided, extrapolate_trend=extrapolate_trend)
+ fig = make_subplots(
+ rows=4, cols=1,
+ subplot_titles=["Observed", "Trend", "Seasonal", "Residuals"])
+ for idx, col in enumerate(['observed', 'trend', 'seasonal', 'resid']):
+ fig.add_trace(
+ go.Scatter(x=result.observed.index, y=getattr(result, col), mode='lines'),
+ row=idx+1, col=1,
+ )
+ return fig
+```
+
+
+```python
+plotSeasonalDecompose(
+ df["y"],
+ model="additive",
+ period=24,
+ title="Seasonal Decomposition")
+```
+
+
+
+## Split the data into training and testing
+
+Let’s divide our data into sets
+
+Let’s divide our data into sets 1. Data to train our
+`Croston Optimized Model`. 2. Data to test our model
+
+For the test data we will use the last 500 Hours to test and evaluate
+the performance of our model.
+
+```python
+train = df[df.ds\<='2023-01-31 19:00:00']
+test = df[df.ds>'2023-01-31 19:00:00']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((9500, 3), (500, 3))
+```
+
+## Implementation of CrostonOptimized with StatsForecast
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import CrostonOptimized
+```
+
+### Instantiating Model
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/) by the
+master, Rob Hyndmann, can be useful for `season_length`.
+
+```python
+season_length = 24 # Hourly data
+horizon = len(test) # number of predictions
+
+# We call the model that we are going to use
+models = [CrostonOptimized()]
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [pandas’
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models, freq='h')
+```
+
+### Fit the Model
+
+```python
+# fit the models
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[CrostonOptimized])
+```
+
+Let’s see the results of our `Croston optimized Model`. We can observe
+it with the following instruction:
+
+```python
+result=sf.fitted_[0,0].model_
+result
+```
+
+``` text
+{'mean': array([27.41841685])}
+```
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 500 hours ahead.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min.
+
+```python
+Y_hat = sf.forecast(df=train, h=horizon)
+Y_hat
+```
+
+| | unique_id | ds | CrostonOptimized |
+|-----|-----------|---------------------|------------------|
+| 0 | 1 | 2023-01-31 20:00:00 | 27.418417 |
+| 1 | 1 | 2023-01-31 21:00:00 | 27.418417 |
+| 2 | 1 | 2023-01-31 22:00:00 | 27.418417 |
+| ... | ... | ... | ... |
+| 497 | 1 | 2023-02-21 13:00:00 | 27.418417 |
+| 498 | 1 | 2023-02-21 14:00:00 | 27.418417 |
+| 499 | 1 | 2023-02-21 15:00:00 | 27.418417 |
+
+```python
+sf.plot(train, Y_hat, max_insample_length=500)
+```
+
+
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 500 hours ahead.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+forecast_df = sf.predict(h=horizon)
+forecast_df
+```
+
+| | unique_id | ds | CrostonOptimized |
+|-----|-----------|---------------------|------------------|
+| 0 | 1 | 2023-01-31 20:00:00 | 27.418417 |
+| 1 | 1 | 2023-01-31 21:00:00 | 27.418417 |
+| 2 | 1 | 2023-01-31 22:00:00 | 27.418417 |
+| ... | ... | ... | ... |
+| 497 | 1 | 2023-02-21 13:00:00 | 27.418417 |
+| 498 | 1 | 2023-02-21 14:00:00 | 27.418417 |
+| 499 | 1 | 2023-02-21 15:00:00 | 27.418417 |
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=)`, forecasting every second months
+`(step_size=50)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 500 hours ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=df,
+ h=horizon,
+ step_size=50,
+ n_windows=5)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` series identifier
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the `n_windows`.
+- `y:` true value
+- `model:` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df
+```
+
+| | unique_id | ds | cutoff | y | CrostonOptimized |
+|------|-----------|---------------------|---------------------|------|------------------|
+| 0 | 1 | 2023-01-23 12:00:00 | 2023-01-23 11:00:00 | 0.0 | 23.655830 |
+| 1 | 1 | 2023-01-23 13:00:00 | 2023-01-23 11:00:00 | 0.0 | 23.655830 |
+| 2 | 1 | 2023-01-23 14:00:00 | 2023-01-23 11:00:00 | 0.0 | 23.655830 |
+| ... | ... | ... | ... | ... | ... |
+| 2497 | 1 | 2023-02-21 13:00:00 | 2023-01-31 19:00:00 | 60.0 | 27.418417 |
+| 2498 | 1 | 2023-02-21 14:00:00 | 2023-01-31 19:00:00 | 20.0 | 27.418417 |
+| 2499 | 1 | 2023-02-21 15:00:00 | 2023-01-31 19:00:00 | 20.0 | 27.418417 |
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=season_length), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | CrostonOptimized |
+|-----|-----------|--------|------------------|
+| 0 | 1 | mae | 33.704756 |
+| 1 | 1 | mape | 0.632593 |
+| 2 | 1 | mase | 0.804074 |
+| 3 | 1 | rmse | 45.262709 |
+| 4 | 1 | smape | 0.767960 |
+
+# References
+
+1. [Changquan Huang • Alla Petukhina. Springer series (2022). Applied
+ Time Series Analysis and Forecasting with
+ Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2)
+2. Ivan Svetunkov. [Forecasting and Analytics with the Augmented
+ Dynamic Adaptive Model (ADAM)](https://openforecast.org/adam/)
+3. [James D. Hamilton. Time Series Analysis Princeton University Press,
+ Princeton, New Jersey, 1st Edition,
+ 1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Nixtla CrostonOptimized API](../../src/core/models.html#crostonoptimized)
+5. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+6. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+7. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/models/crostonsba.html.mdx b/statsforecast/docs/models/crostonsba.html.mdx
new file mode 100644
index 00000000..20f160a2
--- /dev/null
+++ b/statsforecast/docs/models/crostonsba.html.mdx
@@ -0,0 +1,708 @@
+---
+title: CrostonSBA Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `CrostonSBA Model` with
+> `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation` in other.
+
+The text in this article is largely taken from: 1. [Changquan Huang •
+Alla Petukhina. Springer series (2022). Applied Time Series Analysis and
+Forecasting with
+Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2) 2.
+Ivan Svetunkov. [Forecasting and Analytics with the Augmented Dynamic
+Adaptive Model (ADAM)](https://openforecast.org/adam/) 3. [James D.
+Hamilton. Time Series Analysis Princeton University Press, Princeton,
+New Jersey, 1st Edition,
+1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+Principles and Practice (3rd ed)”](https://otexts.com/fpp3/tscv.html).
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [Croston SBA Model](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of CrostonSBA with StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+The Croston model is a method used to forecast time series with
+intermittent demand data, that is, data that has many periods of zero
+demand and only a few periods of non-zero demand. Croston’s approach was
+originally proposed by J.D. Croston in 1972. Subsequently, Syntetos and
+Boylan proposed an improvement to the original model in 2001, known as
+the Croston-SBA (Syntetos and Boylan Approximation).
+
+The Croston-SBA model is based on the assumption that intermittent
+demand follows a binomial process. Instead of directly modeling demand,
+the focus is on modeling the intervals between demand periods. The model
+has two main components: one to model the intervals between demand
+periods (which are assumed to follow a Poisson distribution), and
+another to model the demands when they occur.
+
+It is important to note that the Croston-SBA model assumes that the
+intervals between the non-zero demand periods are independent and follow
+a Poisson distribution. However, this model is an approximation and may
+not work well in all situations. It is advisable to evaluate its
+performance on historical data before using it in practice.
+
+## Croston SBA Model
+
+The formula of SBA is very similar to the original Croston’s method,
+however, it apply a correction factor which reduce the error in the
+final estimate result.
+
+if $Z_t=0$ then
+
+$$Z'_t=Z'_{t-1}$$
+
+$$P'_t=P'_{t-1}$$
+
+Otherwise
+
+$$Z'_t=\alpha Z_t +(1-\alpha)Z'_{t-1}$$
+
+$$P'_t=\alpha P_t +(1- \alpha) P'_{t-1}\ where \ 0<\alpha < 1$$
+
+$$Y'_t=(1-\frac{\alpha}{2}) \frac{Z'_t}{P'_t}$$
+
+where
+
+- $Y'_t:$ Average demand per period
+- $Z_t:$ Actual demand at period $t$
+- $Z'_t:$ Time between two positive demand
+- $P:$ Demand size forecast for next period
+- $P'_t:$ Forecast of demand interval
+- $\alpha:$ Smoothing constant
+
+Note: In Croston’s method, result often will present a considerable
+positive bias, whereas in SBA the bias is reduced, and sometimes will
+appear slightly negative bias.
+
+### Principals of the Croston SBA method
+
+The Croston SBA (Syntetos and Boylan Approximate) method is a technique
+used for forecasting time series with intermittent or sporadic data.
+This methodology is based on the original Croston method, which was
+developed to forecast inventory demand in situations where data is
+sparse or not available at regular intervals.
+
+The main properties of the Croston SBA method are the following:
+
+1. Suitable for intermittent data: The Croston SBA method is especially
+ useful when the data exhibits intermittent patterns, that is,
+ periods of demand followed by periods of non-demand. Instead of
+ treating the data as zero for non-demand periods, the Croston SBA
+ method estimates demand occurrence rates and conditional demand
+ rates.
+
+2. Separation of frequency and level: One of the key features of the
+ Croston SBA method is that it separates the frequency and level
+ information in the demand data. This allows these two components to
+ be modeled and forecasted separately, which can result in better
+ predictions.
+
+3. Estimation of occurrence and demand rates: The Croston SBA method
+ uses a simple exponential smoothing technique to estimate
+ conditional occurrence and demand rates. These rates are then used
+ to forecast future demand.
+
+4. Does not assume distribution of the data: Unlike some forecasting
+ techniques that assume a specific distribution of the data, the
+ Croston SBA method makes no assumptions about the distribution of
+ demand. This makes it more flexible and applicable to a wide range
+ of situations.
+
+5. Does not require complete historical data: The Croston SBA method
+ can work even when historical data is sparse or not available at
+ regular intervals. This makes it an attractive option when it comes
+ to forecasting intermittent demand with limited data.
+
+It is important to note that the Croston SBA method is an approximation
+and may not be suitable for all cases. It is recommended to evaluate its
+performance in conjunction with other forecasting techniques and adapt
+it according to the specific characteristics of the data and the context
+of the problem.
+
+In the Croston SBA method, the data series need not be stationary. The
+Croston SBA approach is suitable for forecasting time series with
+intermittent data, where periods of demand are interspersed with periods
+of non-demand.
+
+The Croston SBA method is based on the estimation of occurrence rates
+and conditional demand rates, using simple exponential smoothing
+techniques. These rates are used to forecast future demand.
+
+In the context of time series, stationarity refers to the property that
+the statistical properties of the series, such as the mean and variance,
+are constant over time. However, in the case of intermittent data, it is
+common for the series not to meet the assumptions of stationarity, since
+the demand can vary considerably in different periods of time.
+
+The Croston SBA method is not based on the assumption of stationarity of
+the data series. Instead, it focuses on modeling the frequency and level
+of intermittent demand separately, using simple exponential smoothing
+techniques. This makes it possible to capture demand occurrence patterns
+and estimate conditional demand rates, without requiring the
+stationarity of the series.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
+plt.style.use('grayscale') # fivethirtyeight grayscale classic
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#008080', # #212946
+ 'axes.facecolor': '#008080',
+ 'savefig.facecolor': '#008080',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#000000', #2A3459
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+
+```python
+import pandas as pd
+df=pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/intermittend_demand2")
+
+df.head()
+```
+
+| | date | sales |
+|-----|---------------------|-------|
+| 0 | 2022-01-01 00:00:00 | 0 |
+| 1 | 2022-01-01 01:00:00 | 10 |
+| 2 | 2022-01-01 02:00:00 | 0 |
+| 3 | 2022-01-01 03:00:00 | 0 |
+| 4 | 2022-01-01 04:00:00 | 100 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|---------------------|-----|-----------|
+| 0 | 2022-01-01 00:00:00 | 0 | 1 |
+| 1 | 2022-01-01 01:00:00 | 10 | 1 |
+| 2 | 2022-01-01 02:00:00 | 0 | 1 |
+| 3 | 2022-01-01 03:00:00 | 0 | 1 |
+| 4 | 2022-01-01 04:00:00 | 100 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y int64
+unique_id object
+dtype: object
+```
+
+We can see that our time variable `(ds)` is in an object format, we need
+to convert to a date format
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore Data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints a random series from the dataset and is useful for
+basic EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### Autocorrelation plots
+
+Autocorrelation (ACF) and partial autocorrelation (PACF) plots are
+statistical tools used to analyze time series. ACF charts show the
+correlation between the values of a time series and their lagged values,
+while PACF charts show the correlation between the values of a time
+series and their lagged values, after the effect of previous lagged
+values has been removed.
+
+ACF and PACF charts can be used to identify the structure of a time
+series, which can be helpful in choosing a suitable model for the time
+series. For example, if the ACF chart shows a repeating peak and valley
+pattern, this indicates that the time series is stationary, meaning that
+it has the same statistical properties over time. If the PACF chart
+shows a pattern of rapidly decreasing spikes, this indicates that the
+time series is invertible, meaning it can be reversed to get a
+stationary time series.
+
+The importance of the ACF and PACF charts is that they can help analysts
+better understand the structure of a time series. This understanding can
+be helpful in choosing a suitable model for the time series, which can
+improve the ability to predict future values of the time series.
+
+To analyze ACF and PACF charts:
+
+- Look for patterns in charts. Common patterns include repeating peaks
+ and valleys, sawtooth patterns, and plateau patterns.
+- Compare ACF and PACF charts. The PACF chart generally has fewer
+ spikes than the ACF chart.
+- Consider the length of the time series. ACF and PACF charts for
+ longer time series will have more spikes.
+- Use a confidence interval. The ACF and PACF plots also show
+ confidence intervals for the autocorrelation values. If an
+ autocorrelation value is outside the confidence interval, it is
+ likely to be significant.
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=30, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+plot_pacf(df["y"], lags=30, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+plt.show();
+```
+
+
+
+### Decomposition of the time series
+
+How to decompose a time series and why?
+
+In time series analysis to forecast new values, it is very important to
+know past data. More formally, we can say that it is very important to
+know the patterns that values follow over time. There can be many
+reasons that cause our forecast values to fall in the wrong direction.
+Basically, a time series consists of four components. The variation of
+those components causes the change in the pattern of the time series.
+These components are:
+
+- **Level:** This is the primary value that averages over time.
+- **Trend:** The trend is the value that causes increasing or
+ decreasing patterns in a time series.
+- **Seasonality:** This is a cyclical event that occurs in a time
+ series for a short time and causes short-term increasing or
+ decreasing patterns in a time series.
+- **Residual/Noise:** These are the random variations in the time
+ series.
+
+Combining these components over time leads to the formation of a time
+series. Most time series consist of level and noise/residual and trend
+or seasonality are optional values.
+
+If seasonality and trend are part of the time series, then there will be
+effects on the forecast value. As the pattern of the forecasted time
+series may be different from the previous time series.
+
+The combination of the components in time series can be of two types: \*
+Additive \* Multiplicative
+
+### Additive time series
+
+If the components of the time series are added to make the time series.
+Then the time series is called the additive time series. By
+visualization, we can say that the time series is additive if the
+increasing or decreasing pattern of the time series is similar
+throughout the series. The mathematical function of any additive time
+series can be represented by:
+$$y(t) = level + Trend + seasonality + noise$$
+
+### Multiplicative time series
+
+If the components of the time series are multiplicative together, then
+the time series is called a multiplicative time series. For
+visualization, if the time series is having exponential growth or
+decline with time, then the time series can be considered as the
+multiplicative time series. The mathematical function of the
+multiplicative time series can be represented as.
+
+$$y(t) = Level * Trend * seasonality * Noise$$
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+from plotly.subplots import make_subplots
+import plotly.graph_objects as go
+
+def plotSeasonalDecompose(
+ x,
+ model='additive',
+ filt=None,
+ period=None,
+ two_sided=True,
+ extrapolate_trend=0,
+ title="Seasonal Decomposition"):
+
+ result = seasonal_decompose(
+ x, model=model, filt=filt, period=period,
+ two_sided=two_sided, extrapolate_trend=extrapolate_trend)
+ fig = make_subplots(
+ rows=4, cols=1,
+ subplot_titles=["Observed", "Trend", "Seasonal", "Residuals"])
+ for idx, col in enumerate(['observed', 'trend', 'seasonal', 'resid']):
+ fig.add_trace(
+ go.Scatter(x=result.observed.index, y=getattr(result, col), mode='lines'),
+ row=idx+1, col=1,
+ )
+ return fig
+```
+
+
+```python
+plotSeasonalDecompose(
+ df["y"],
+ model="additive",
+ period=24,
+ title="Seasonal Decomposition")
+```
+
+
+
+## Split the data into training and testing
+
+Let’s divide our data into sets 1. Data to train our
+`Croston SBA Model`. 2. Data to test our model
+
+For the test data we will use the last 500 Hours to test and evaluate
+the performance of our model.
+
+```python
+train = df[df.ds\<='2023-01-31 19:00:00']
+test = df[df.ds>'2023-01-31 19:00:00']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((9500, 3), (500, 3))
+```
+
+## Implementation of CrostonSBA with StatsForecast
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import CrostonSBA
+```
+
+### Instantiating Model
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/) by the
+master, Rob Hyndmann, can be useful for `season_length`.
+
+```python
+season_length = 24 # Hourly data
+horizon = len(test) # number of predictions
+
+# We call the model that we are going to use
+models = [CrostonSBA()]
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [pandas’
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models, freq='h')
+```
+
+### Fit the Model
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[CrostonSBA])
+```
+
+Let’s see the results of our `Croston SBA Model`. We can observe it with
+the following instruction:
+
+```python
+result=sf.fitted_[0,0].model_
+result
+```
+
+``` text
+{'mean': array([26.04749601]),
+ 'fitted': array([ nan, 0. , 4.75 , ..., 29.088629, 29.088629,
+ 29.088629], dtype=float32),
+ 'sigma': np.float32(49.512943)}
+```
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 500 hours ahead.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min. (If you want to speed things up to a couple of seconds,
+remove the AutoModels like `ARIMA` and `Theta`)
+
+```python
+Y_hat = sf.forecast(df=train, h=horizon)
+Y_hat
+```
+
+| | unique_id | ds | CrostonSBA |
+|-----|-----------|---------------------|------------|
+| 0 | 1 | 2023-01-31 20:00:00 | 26.047497 |
+| 1 | 1 | 2023-01-31 21:00:00 | 26.047497 |
+| 2 | 1 | 2023-01-31 22:00:00 | 26.047497 |
+| ... | ... | ... | ... |
+| 497 | 1 | 2023-02-21 13:00:00 | 26.047497 |
+| 498 | 1 | 2023-02-21 14:00:00 | 26.047497 |
+| 499 | 1 | 2023-02-21 15:00:00 | 26.047497 |
+
+```python
+sf.plot(train, Y_hat, max_insample_length=500)
+```
+
+
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 500 hours ahead.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+forecast_df = sf.predict(h=horizon)
+forecast_df
+```
+
+| | unique_id | ds | CrostonSBA |
+|-----|-----------|---------------------|------------|
+| 0 | 1 | 2023-01-31 20:00:00 | 26.047497 |
+| 1 | 1 | 2023-01-31 21:00:00 | 26.047497 |
+| 2 | 1 | 2023-01-31 22:00:00 | 26.047497 |
+| ... | ... | ... | ... |
+| 497 | 1 | 2023-02-21 13:00:00 | 26.047497 |
+| 498 | 1 | 2023-02-21 14:00:00 | 26.047497 |
+| 499 | 1 | 2023-02-21 15:00:00 | 26.047497 |
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=)`, forecasting every second months
+`(step_size=50)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 500 hours ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=df,
+ h=horizon,
+ step_size=50,
+ n_windows=5)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` series identifier
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the `n_windows`.
+- `y:` true value
+- `model:` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df
+```
+
+| | unique_id | ds | cutoff | y | CrostonSBA |
+|------|-----------|---------------------|---------------------|------|------------|
+| 0 | 1 | 2023-01-23 12:00:00 | 2023-01-23 11:00:00 | 0.0 | 22.473040 |
+| 1 | 1 | 2023-01-23 13:00:00 | 2023-01-23 11:00:00 | 0.0 | 22.473040 |
+| 2 | 1 | 2023-01-23 14:00:00 | 2023-01-23 11:00:00 | 0.0 | 22.473040 |
+| ... | ... | ... | ... | ... | ... |
+| 2497 | 1 | 2023-02-21 13:00:00 | 2023-01-31 19:00:00 | 60.0 | 26.047497 |
+| 2498 | 1 | 2023-02-21 14:00:00 | 2023-01-31 19:00:00 | 20.0 | 26.047497 |
+| 2499 | 1 | 2023-02-21 15:00:00 | 2023-01-31 19:00:00 | 20.0 | 26.047497 |
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=season_length), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | CrostonSBA |
+|-----|-----------|--------|------------|
+| 0 | 1 | mae | 33.112519 |
+| 1 | 1 | mape | 0.626900 |
+| 2 | 1 | mase | 0.789945 |
+| 3 | 1 | rmse | 45.203519 |
+| 4 | 1 | smape | 0.771529 |
+
+# References
+
+1. [Changquan Huang • Alla Petukhina. Springer series (2022). Applied
+ Time Series Analysis and Forecasting with
+ Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2)
+2. Ivan Svetunkov. [Forecasting and Analytics with the Augmented
+ Dynamic Adaptive Model (ADAM)](https://openforecast.org/adam/)
+3. [James D. Hamilton. Time Series Analysis Princeton University Press,
+ Princeton, New Jersey, 1st Edition,
+ 1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Nixtla CrostonSBA API](../../src/core/models.html#crostonsba)
+5. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+6. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+7. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/models/dynamicoptimizedtheta.html.mdx b/statsforecast/docs/models/dynamicoptimizedtheta.html.mdx
new file mode 100644
index 00000000..b7e033b8
--- /dev/null
+++ b/statsforecast/docs/models/dynamicoptimizedtheta.html.mdx
@@ -0,0 +1,682 @@
+---
+title: Dynamic Optimized Theta Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `DynamicOptimizedTheta Model` with
+> `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation` in other.
+
+The text in this article is largely taken from [Jose A. Fiorucci, Tiago
+R. Pellegrini, Francisco Louzada, Fotios Petropoulos, Anne B. Koehler
+(2016). “Models for optimising the theta method and their relationship
+to state space models”. International Journal of
+Forecasting](https://www.sciencedirect.com/science/article/pii/S0169207016300243).
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [Dynamic Optimized Theta Model (DOTM)](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of DynamicOptimizedTheta with
+ StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+The **Dynamic Optimized Theta Model (DOTM)** in `StatsForecast` is a
+variation of the classic Theta model. It combines key features of two
+other extensions: the **Optimized Theta Model (OTM)** and the **Dynamic
+Standard Theta Model (DSTM)**.
+
+DOTM introduces two main improvements over the standard Theta model:
+**optimization** of the theta parameters and **dynamic updating** of
+model components over time.
+
+- **Optimization**: Like OTM, this version automatically searches for
+ the best theta values based on the data, rather than relying on
+ fixed parameters. This flexibility allows the model to better adapt
+ to series with complex seasonal or trend patterns.
+
+- **Dynamic updating**: Like DSTM, DOTM continuously updates its
+ internal components as new data becomes available. This makes it
+ well-suited for non-stationary series, where the underlying data
+ structure evolves over time.
+
+DOTM also supports **seasonal decomposition**, controlled by the
+`decomposition_type` parameter. You can choose between: -
+`'multiplicative'` (default), which assumes that seasonal effects scale
+with the level of the series, or - `'additive'`, which assumes that
+seasonal effects remain constant in absolute magnitude.
+
+The Dynamic Optimized Theta Model is the most flexible of the Theta
+family and is particularly effective when forecasting series with
+changing trends and seasonalities.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
+plt.style.use('grayscale') # fivethirtyeight grayscale classic
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#008080', # #212946
+ 'axes.facecolor': '#008080',
+ 'savefig.facecolor': '#008080',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#000000', #2A3459
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+### Read Data
+
+```python
+import pandas as pd
+
+df = pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/milk_production.csv", usecols=[1,2])
+df.head()
+```
+
+| | month | production |
+|-----|------------|------------|
+| 0 | 1962-01-01 | 589 |
+| 1 | 1962-02-01 | 561 |
+| 2 | 1962-03-01 | 640 |
+| 3 | 1962-04-01 | 656 |
+| 4 | 1962-05-01 | 727 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|------------|-----|-----------|
+| 0 | 1962-01-01 | 589 | 1 |
+| 1 | 1962-02-01 | 561 | 1 |
+| 2 | 1962-03-01 | 640 | 1 |
+| 3 | 1962-04-01 | 656 | 1 |
+| 4 | 1962-05-01 | 727 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y int64
+unique_id object
+dtype: object
+```
+
+We can see that our time variable `(ds)` is in an object format, we need
+to convert to a date format
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore Data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints a random series from the dataset and is useful for
+basic EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### Autocorrelation plots
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=30, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+plot_pacf(df["y"], lags=30, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+plt.show();
+```
+
+
+
+### Decomposition of the time series
+
+How to decompose a time series and why?
+
+In time series analysis to forecast new values, it is very important to
+know past data. More formally, we can say that it is very important to
+know the patterns that values follow over time. There can be many
+reasons that cause our forecast values to fall in the wrong direction.
+Basically, a time series consists of four components. The variation of
+those components causes the change in the pattern of the time series.
+These components are:
+
+- **Level:** This is the primary value that averages over time.
+- **Trend:** The trend is the value that causes increasing or
+ decreasing patterns in a time series.
+- **Seasonality:** This is a cyclical event that occurs in a time
+ series for a short time and causes short-term increasing or
+ decreasing patterns in a time series.
+- **Residual/Noise:** These are the random variations in the time
+ series.
+
+Combining these components over time leads to the formation of a time
+series. Most time series consist of level and noise/residual and trend
+or seasonality are optional values.
+
+If seasonality and trend are part of the time series, then there will be
+effects on the forecast value. As the pattern of the forecasted time
+series may be different from the previous time series.
+
+The combination of the components in time series can be of two types: \*
+Additive \* Multiplicative
+
+### Additive time series
+
+If the components of the time series are added to make the time series.
+Then the time series is called the additive time series. By
+visualization, we can say that the time series is additive if the
+increasing or decreasing pattern of the time series is similar
+throughout the series. The mathematical function of any additive time
+series can be represented by:
+$$y(t) = level + Trend + seasonality + noise$$
+
+### Multiplicative time series
+
+If the components of the time series are multiplicative together, then
+the time series is called a multiplicative time series. For
+visualization, if the time series is having exponential growth or
+decline with time, then the time series can be considered as the
+multiplicative time series. The mathematical function of the
+multiplicative time series can be represented as.
+
+$$y(t) = Level * Trend * seasonality * Noise$$
+
+### Additive
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+a = seasonal_decompose(df["y"], model = "additive", period=12)
+a.plot();
+```
+
+
+
+### Multiplicative
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+a = seasonal_decompose(df["y"], model = "Multiplicative", period=12)
+a.plot();
+```
+
+
+
+## Split the data into training and testing
+
+Let’s divide our data into sets
+
+1. Data to train our `Dynamic Optimized Theta Model(DOTM)`.
+2. Data to test our model
+
+For the test data we will use the last 12 months to test and evaluate
+the performance of our model.
+
+```python
+train = df[df.ds\<='1974-12-01']
+test = df[df.ds>'1974-12-01']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((156, 3), (12, 3))
+```
+
+Now let’s plot the training data and the test data.
+
+```python
+sns.lineplot(train,x="ds", y="y", label="Train", linestyle="--")
+sns.lineplot(test, x="ds", y="y", label="Test")
+plt.title("Monthly Milk Production");
+plt.show()
+```
+
+
+
+## Implementation of DynamicOptimizedTheta with StatsForecast
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import DynamicOptimizedTheta
+```
+
+### Instantiating Model
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/) by the
+master, Rob Hyndmann, can be useful for `season_length`.
+
+```python
+season_length = 12 # Monthly data
+horizon = len(test) # number of predictions
+
+# We call the model that we are going to use
+models = [DynamicOptimizedTheta(season_length=season_length,
+ decomposition_type="additive")] # multiplicative additive
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [pandas’
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models, freq='MS')
+```
+
+### Fit the Model
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[DynamicOptimizedTheta])
+```
+
+Let’s see the results of our `Dynamic Optimized Theta Model`. We can
+observe it with the following instruction:
+
+```python
+result=sf.fitted_[0,0].model_
+print(result.keys())
+print(result['fit'])
+```
+
+``` text
+dict_keys(['mse', 'amse', 'fit', 'residuals', 'm', 'states', 'par', 'n', 'modeltype', 'mean_y', 'decompose', 'decomposition_type', 'seas_forecast', 'fitted'])
+results(x=array([250.83206219, 0.75624902, 4.67964777]), fn=10.697554045462667, nit=55, simplex=array([[237.42074763, 0.75306547, 4.46023813],
+ [250.83206219, 0.75624902, 4.67964777],
+ [257.16444246, 0.75229688, 4.42377059],
+ [256.90853867, 0.75757957, 4.43171897]]))
+```
+
+Let us now visualize the residuals of our models.
+
+As we can see, the result obtained above has an output in a dictionary,
+to extract each element from the dictionary we are going to use the
+`.get()` function to extract the element and then we are going to save
+it in a `pd.DataFrame()`.
+
+```python
+residual=pd.DataFrame(result.get("residuals"), columns=["residual Model"])
+residual
+```
+
+| | residual Model |
+|-----|----------------|
+| 0 | -18.247106 |
+| 1 | -75.757706 |
+| 2 | 6.001494 |
+| ... | ... |
+| 153 | -59.747044 |
+| 154 | -91.901521 |
+| 155 | -43.503294 |
+
+```python
+import scipy.stats as stats
+
+fig, axs = plt.subplots(nrows=2, ncols=2)
+
+residual.plot(ax=axs[0,0])
+axs[0,0].set_title("Residuals");
+
+sns.distplot(residual, ax=axs[0,1]);
+axs[0,1].set_title("Density plot - Residual");
+
+stats.probplot(residual["residual Model"], dist="norm", plot=axs[1,0])
+axs[1,0].set_title('Plot Q-Q')
+
+plot_acf(residual, lags=35, ax=axs[1,1],color="fuchsia")
+axs[1,1].set_title("Autocorrelation");
+
+plt.show();
+```
+
+
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[90]` means that
+ the model expects the real value to be inside that interval 90% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min.
+
+```python
+# Prediction
+Y_hat = sf.forecast(df=train, h=horizon, fitted=True)
+Y_hat
+```
+
+| | unique_id | ds | DynamicOptimizedTheta |
+|-----|-----------|------------|-----------------------|
+| 0 | 1 | 1975-01-01 | 839.259705 |
+| 1 | 1 | 1975-02-01 | 801.399170 |
+| 2 | 1 | 1975-03-01 | 895.189148 |
+| ... | ... | ... | ... |
+| 9 | 1 | 1975-10-01 | 821.271240 |
+| 10 | 1 | 1975-11-01 | 792.530518 |
+| 11 | 1 | 1975-12-01 | 829.854553 |
+
+```python
+values=sf.forecast_fitted_values()
+values.head()
+```
+
+| | unique_id | ds | y | DynamicOptimizedTheta |
+|-----|-----------|------------|-------|-----------------------|
+| 0 | 1 | 1962-01-01 | 589.0 | 607.247131 |
+| 1 | 1 | 1962-02-01 | 561.0 | 636.757690 |
+| 2 | 1 | 1962-03-01 | 640.0 | 633.998535 |
+| 3 | 1 | 1962-04-01 | 656.0 | 608.461243 |
+| 4 | 1 | 1962-05-01 | 727.0 | 604.808899 |
+
+```python
+StatsForecast.plot(values)
+```
+
+
+
+Adding 95% confidence interval with the forecast method
+
+```python
+sf.forecast(df=train, h=horizon, level=[95])
+```
+
+| | unique_id | ds | DynamicOptimizedTheta | DynamicOptimizedTheta-lo-95 | DynamicOptimizedTheta-hi-95 |
+|-----|-----------|------------|-----------------------|-----------------------------|-----------------------------|
+| 0 | 1 | 1975-01-01 | 839.259705 | 741.952332 | 955.151001 |
+| 1 | 1 | 1975-02-01 | 801.399170 | 641.867920 | 946.045776 |
+| 2 | 1 | 1975-03-01 | 895.189148 | 707.189087 | 1066.356812 |
+| ... | ... | ... | ... | ... | ... |
+| 9 | 1 | 1975-10-01 | 821.271240 | 546.081726 | 1088.193481 |
+| 10 | 1 | 1975-11-01 | 792.530518 | 494.623718 | 1037.459839 |
+| 11 | 1 | 1975-12-01 | 829.854553 | 519.661133 | 1108.213867 |
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[95]` means that
+ the model expects the real value to be inside that interval 95% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+sf.predict(h=horizon)
+```
+
+| | unique_id | ds | DynamicOptimizedTheta |
+|-----|-----------|------------|-----------------------|
+| 0 | 1 | 1975-01-01 | 839.259705 |
+| 1 | 1 | 1975-02-01 | 801.399170 |
+| 2 | 1 | 1975-03-01 | 895.189148 |
+| ... | ... | ... | ... |
+| 9 | 1 | 1975-10-01 | 821.271240 |
+| 10 | 1 | 1975-11-01 | 792.530518 |
+| 11 | 1 | 1975-12-01 | 829.854553 |
+
+```python
+forecast_df = sf.predict(h=horizon, level=[80,95])
+forecast_df
+```
+
+| | unique_id | ds | DynamicOptimizedTheta | DynamicOptimizedTheta-lo-80 | DynamicOptimizedTheta-hi-80 | DynamicOptimizedTheta-lo-95 | DynamicOptimizedTheta-hi-95 |
+|-----|-----------|------------|-----------------------|-----------------------------|-----------------------------|-----------------------------|-----------------------------|
+| 0 | 1 | 1975-01-01 | 839.259705 | 766.142090 | 928.025513 | 741.952332 | 955.151001 |
+| 1 | 1 | 1975-02-01 | 801.399170 | 702.981262 | 899.884216 | 641.867920 | 946.045776 |
+| 2 | 1 | 1975-03-01 | 895.189148 | 760.125916 | 1008.335022 | 707.189087 | 1066.356812 |
+| ... | ... | ... | ... | ... | ... | ... | ... |
+| 9 | 1 | 1975-10-01 | 821.271240 | 617.391724 | 996.698364 | 546.081726 | 1088.193481 |
+| 10 | 1 | 1975-11-01 | 792.530518 | 568.303162 | 975.070312 | 494.623718 | 1037.459839 |
+| 11 | 1 | 1975-12-01 | 829.854553 | 598.098267 | 1035.476196 | 519.661133 | 1108.213867 |
+
+```python
+sf.plot(train, test.merge(forecast_df), level=[80, 95])
+```
+
+
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=5)`, forecasting every second months
+`(step_size=12)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 12 months ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=train,
+ h=horizon,
+ step_size=12,
+ n_windows=3)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` series identifier
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the n_windows.
+- `y:` true value
+- `"model":` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df
+```
+
+| | unique_id | ds | cutoff | y | DynamicOptimizedTheta |
+|-----|-----------|------------|------------|-------|-----------------------|
+| 0 | 1 | 1972-01-01 | 1971-12-01 | 826.0 | 828.692017 |
+| 1 | 1 | 1972-02-01 | 1971-12-01 | 799.0 | 792.444092 |
+| 2 | 1 | 1972-03-01 | 1971-12-01 | 890.0 | 883.122620 |
+| ... | ... | ... | ... | ... | ... |
+| 33 | 1 | 1974-10-01 | 1973-12-01 | 812.0 | 810.304688 |
+| 34 | 1 | 1974-11-01 | 1973-12-01 | 773.0 | 781.804688 |
+| 35 | 1 | 1974-12-01 | 1973-12-01 | 813.0 | 818.811096 |
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=season_length), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | DynamicOptimizedTheta |
+|-----|-----------|--------|-----------------------|
+| 0 | 1 | mae | 6.861949 |
+| 1 | 1 | mape | 0.008045 |
+| 2 | 1 | mase | 0.308595 |
+| 3 | 1 | rmse | 8.647459 |
+| 4 | 1 | smape | 0.004010 |
+
+## References
+
+1. [Kostas I. Nikolopoulos, Dimitrios D. Thomakos. Forecasting with the
+ Theta Method-Theory and Applications. 2019 John Wiley & Sons
+ Ltd.](https://onlinelibrary.wiley.com/doi/book/10.1002/9781119320784)
+2. [Jose A. Fiorucci, Tiago R. Pellegrini, Francisco Louzada, Fotios
+ Petropoulos, Anne B. Koehler (2016). “Models for optimising the
+ theta method and their relationship to state space models”.
+ International Journal of
+ Forecasting](https://www.sciencedirect.com/science/article/pii/S0169207016300243).
+3. [Nixtla DynamicOptimizedTheta
+ API](../../src/core/models.html#dynamicoptimizedtheta)
+4. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+5. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ principles and practice, Time series
+ cross-validation”.](https://otexts.com/fpp3/tscv.html).
+6. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/models/dynamicstandardtheta.html.mdx b/statsforecast/docs/models/dynamicstandardtheta.html.mdx
new file mode 100644
index 00000000..b542ee1b
--- /dev/null
+++ b/statsforecast/docs/models/dynamicstandardtheta.html.mdx
@@ -0,0 +1,652 @@
+---
+title: Dynamic Standard Theta Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `DynamicStandardTheta Model` with
+> `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation` in other.
+
+The text in this article is largely taken from [Jose A. Fiorucci, Tiago
+R. Pellegrini, Francisco Louzada, Fotios Petropoulos, Anne B. Koehler
+(2016). “Models for optimising the theta method and their relationship
+to state space models”. International Journal of
+Forecasting](https://www.sciencedirect.com/science/article/pii/S0169207016300243).
+
+## Table of Contents
+
+- [Dynamic Standard Theta Model (DOTM)](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of DynamicStandardTheta with
+ StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Dynamic Standard Theta Models (DSTM)
+
+The Dynamic Standard Theta Model is a case-specific variation of the
+[Optimized Dynamic Theta Model](./DynamicOptimizedTheta).
+
+Also, for $\theta=2$, we have a stochastic approach of Theta, which is
+referred to hereafter as the dynamic standard Theta model (DSTM).
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
+plt.style.use('grayscale') # fivethirtyeight grayscale classic
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#008080', # #212946
+ 'axes.facecolor': '#008080',
+ 'savefig.facecolor': '#008080',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#000000', #2A3459
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+### Read Data
+
+```python
+import pandas as pd
+df = pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/milk_production.csv", usecols=[1,2])
+df.head()
+```
+
+| | month | production |
+|-----|------------|------------|
+| 0 | 1962-01-01 | 589 |
+| 1 | 1962-02-01 | 561 |
+| 2 | 1962-03-01 | 640 |
+| 3 | 1962-04-01 | 656 |
+| 4 | 1962-05-01 | 727 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|------------|-----|-----------|
+| 0 | 1962-01-01 | 589 | 1 |
+| 1 | 1962-02-01 | 561 | 1 |
+| 2 | 1962-03-01 | 640 | 1 |
+| 3 | 1962-04-01 | 656 | 1 |
+| 4 | 1962-05-01 | 727 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y int64
+unique_id object
+dtype: object
+```
+
+We can see that our time variable `(ds)` is in an object format, we need
+to convert to a date format
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore Data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints a random series from the dataset and is useful for
+basic EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### Autocorrelation plots
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=30, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+plot_pacf(df["y"], lags=30, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+plt.show();
+```
+
+
+
+### Decomposition of the time series
+
+How to decompose a time series and why?
+
+In time series analysis to forecast new values, it is very important to
+know past data. More formally, we can say that it is very important to
+know the patterns that values follow over time. There can be many
+reasons that cause our forecast values to fall in the wrong direction.
+Basically, a time series consists of four components. The variation of
+those components causes the change in the pattern of the time series.
+These components are:
+
+- **Level:** This is the primary value that averages over time.
+- **Trend:** The trend is the value that causes increasing or
+ decreasing patterns in a time series.
+- **Seasonality:** This is a cyclical event that occurs in a time
+ series for a short time and causes short-term increasing or
+ decreasing patterns in a time series.
+- **Residual/Noise:** These are the random variations in the time
+ series.
+
+Combining these components over time leads to the formation of a time
+series. Most time series consist of level and noise/residual and trend
+or seasonality are optional values.
+
+If seasonality and trend are part of the time series, then there will be
+effects on the forecast value. As the pattern of the forecasted time
+series may be different from the previous time series.
+
+The combination of the components in time series can be of two types: \*
+Additive \* Multiplicative
+
+### Additive time series
+
+If the components of the time series are added to make the time series.
+Then the time series is called the additive time series. By
+visualization, we can say that the time series is additive if the
+increasing or decreasing pattern of the time series is similar
+throughout the series. The mathematical function of any additive time
+series can be represented by:
+$$y(t) = level + Trend + seasonality + noise$$
+
+### Multiplicative time series
+
+If the components of the time series are multiplicative together, then
+the time series is called a multiplicative time series. For
+visualization, if the time series is having exponential growth or
+decline with time, then the time series can be considered as the
+multiplicative time series. The mathematical function of the
+multiplicative time series can be represented as.
+
+$$y(t) = Level * Trend * seasonality * Noise$$
+
+### Additive
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+a = seasonal_decompose(df["y"], model = "additive", period=12)
+a.plot();
+```
+
+
+
+### Multiplicative
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+a = seasonal_decompose(df["y"], model = "Multiplicative", period=12)
+a.plot();
+```
+
+
+
+## Split the data into training and testing
+
+Let’s divide our data into sets 1. Data to train our
+`Dynamic Standard Theta Model` 2. Data to test our model
+
+For the test data we will use the last 12 months to test and evaluate
+the performance of our model.
+
+```python
+train = df[df.ds\<='1974-12-01']
+test = df[df.ds>'1974-12-01']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((156, 3), (12, 3))
+```
+
+Now let’s plot the training data and the test data.
+
+```python
+sns.lineplot(train,x="ds", y="y", label="Train", linestyle="--")
+sns.lineplot(test, x="ds", y="y", label="Test")
+plt.title("Monthly Milk Production")
+plt.show()
+```
+
+
+
+## Implementation of DynamicStandardTheta with StatsForecast
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import DynamicTheta
+```
+
+### Instantiating Model
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/) by the
+master, Rob Hyndmann, can be useful for `season_length`.
+
+```python
+season_length = 12 # Monthly data
+horizon = len(test) # number of predictions
+
+models = [DynamicTheta(season_length=season_length,
+ decomposition_type="additive")] # multiplicative additive
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [pandas’
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models, freq='MS')
+```
+
+### Fit Model
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[DynamicTheta])
+```
+
+Let’s see the results of our `Dynamic Standard Theta model`. We can
+observe it with the following instruction:
+
+```python
+result=sf.fitted_[0,0].model_
+print(result.keys())
+print(result['fit'])
+```
+
+``` text
+dict_keys(['mse', 'amse', 'fit', 'residuals', 'm', 'states', 'par', 'n', 'modeltype', 'mean_y', 'decompose', 'decomposition_type', 'seas_forecast', 'fitted'])
+results(x=array([393.28739991, 0.76875 ]), fn=10.787112115489622, nit=20, simplex=array([[399.92916541, 0.771875 ],
+ [393.28739991, 0.76875 ],
+ [384.74798713, 0.771875 ]]))
+```
+
+Let us now visualize the residuals of our models.
+
+As we can see, the result obtained above has an output in a dictionary,
+to extract each element from the dictionary we are going to use the
+`.get()` function to extract the element and then we are going to save
+it in a `pd.DataFrame()`.
+
+```python
+residual=pd.DataFrame(result.get("residuals"), columns=["residual Model"])
+residual
+```
+
+| | residual Model |
+|-----|----------------|
+| 0 | -18.247131 |
+| 1 | -46.247131 |
+| 2 | 17.140198 |
+| ... | ... |
+| 153 | -58.941711 |
+| 154 | -91.055420 |
+| 155 | -42.624939 |
+
+```python
+import scipy.stats as stats
+
+fig, axs = plt.subplots(nrows=2, ncols=2)
+
+residual.plot(ax=axs[0,0])
+axs[0,0].set_title("Residuals");
+
+sns.distplot(residual, ax=axs[0,1]);
+axs[0,1].set_title("Density plot - Residual");
+
+stats.probplot(residual["residual Model"], dist="norm", plot=axs[1,0])
+axs[1,0].set_title('Plot Q-Q')
+
+plot_acf(residual, lags=35, ax=axs[1,1],color="fuchsia")
+axs[1,1].set_title("Autocorrelation");
+
+plt.show();
+```
+
+
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[90]` means that
+ the model expects the real value to be inside that interval 90% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min.
+
+```python
+Y_hat = sf.forecast(df=train, h=horizon, fitted=True)
+Y_hat
+```
+
+| | unique_id | ds | DynamicTheta |
+|-----|-----------|------------|--------------|
+| 0 | 1 | 1975-01-01 | 838.531555 |
+| 1 | 1 | 1975-02-01 | 800.154968 |
+| 2 | 1 | 1975-03-01 | 893.430786 |
+| ... | ... | ... | ... |
+| 9 | 1 | 1975-10-01 | 815.959351 |
+| 10 | 1 | 1975-11-01 | 786.716431 |
+| 11 | 1 | 1975-12-01 | 823.539368 |
+
+```python
+values=sf.forecast_fitted_values()
+values.head()
+```
+
+| | unique_id | ds | y | DynamicTheta |
+|-----|-----------|------------|-------|--------------|
+| 0 | 1 | 1962-01-01 | 589.0 | 607.247131 |
+| 1 | 1 | 1962-02-01 | 561.0 | 607.247131 |
+| 2 | 1 | 1962-03-01 | 640.0 | 622.859802 |
+| 3 | 1 | 1962-04-01 | 656.0 | 606.987793 |
+| 4 | 1 | 1962-05-01 | 727.0 | 605.021179 |
+
+```python
+StatsForecast.plot(values)
+```
+
+
+
+Adding 95% confidence interval with the forecast method
+
+```python
+sf.forecast(df=train, h=horizon, level=[95])
+```
+
+| | unique_id | ds | DynamicTheta | DynamicTheta-lo-95 | DynamicTheta-hi-95 |
+|-----|-----------|------------|--------------|--------------------|--------------------|
+| 0 | 1 | 1975-01-01 | 838.531555 | 741.237366 | 954.407166 |
+| 1 | 1 | 1975-02-01 | 800.154968 | 640.697205 | 945.673096 |
+| 2 | 1 | 1975-03-01 | 893.430786 | 703.900635 | 1065.418701 |
+| ... | ... | ... | ... | ... | ... |
+| 9 | 1 | 1975-10-01 | 815.959351 | 536.422791 | 1086.643433 |
+| 10 | 1 | 1975-11-01 | 786.716431 | 484.476593 | 1033.687134 |
+| 11 | 1 | 1975-12-01 | 823.539368 | 509.187256 | 1104.107788 |
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[95]` means that
+ the model expects the real value to be inside that interval 95% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+sf.predict(h=horizon)
+```
+
+| | unique_id | ds | DynamicTheta |
+|-----|-----------|------------|--------------|
+| 0 | 1 | 1975-01-01 | 838.531555 |
+| 1 | 1 | 1975-02-01 | 800.154968 |
+| 2 | 1 | 1975-03-01 | 893.430786 |
+| ... | ... | ... | ... |
+| 9 | 1 | 1975-10-01 | 815.959351 |
+| 10 | 1 | 1975-11-01 | 786.716431 |
+| 11 | 1 | 1975-12-01 | 823.539368 |
+
+```python
+forecast_df = sf.predict(h=horizon, level=[80,95])
+forecast_df
+```
+
+| | unique_id | ds | DynamicTheta | DynamicTheta-lo-80 | DynamicTheta-hi-80 | DynamicTheta-lo-95 | DynamicTheta-hi-95 |
+|-----|-----------|------------|--------------|--------------------|--------------------|--------------------|--------------------|
+| 0 | 1 | 1975-01-01 | 838.531555 | 765.423828 | 927.285339 | 741.237366 | 954.407166 |
+| 1 | 1 | 1975-02-01 | 800.154968 | 701.099854 | 899.316162 | 640.697205 | 945.673096 |
+| 2 | 1 | 1975-03-01 | 893.430786 | 758.326416 | 1007.631165 | 703.900635 | 1065.418701 |
+| ... | ... | ... | ... | ... | ... | ... | ... |
+| 9 | 1 | 1975-10-01 | 815.959351 | 608.699463 | 992.552673 | 536.422791 | 1086.643433 |
+| 10 | 1 | 1975-11-01 | 786.716431 | 558.429810 | 970.648376 | 484.476593 | 1033.687134 |
+| 11 | 1 | 1975-12-01 | 823.539368 | 588.706787 | 1031.564941 | 509.187256 | 1104.107788 |
+
+```python
+sf.plot(train, test.merge(forecast_df), level=[80, 95])
+```
+
+
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=5)`, forecasting every second months
+`(step_size=12)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 12 months ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=train,
+ h=horizon,
+ step_size=12,
+ n_windows=3)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` index. If you dont like working with index just run
+ crossvalidation_df.resetindex()
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the n_windows.
+- `y:` true value
+- `"model":` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df
+```
+
+| | unique_id | ds | cutoff | y | DynamicTheta |
+|-----|-----------|------------|------------|-------|--------------|
+| 0 | 1 | 1972-01-01 | 1971-12-01 | 826.0 | 827.107239 |
+| 1 | 1 | 1972-02-01 | 1971-12-01 | 799.0 | 789.924194 |
+| 2 | 1 | 1972-03-01 | 1971-12-01 | 890.0 | 879.664429 |
+| ... | ... | ... | ... | ... | ... |
+| 33 | 1 | 1974-10-01 | 1973-12-01 | 812.0 | 804.398560 |
+| 34 | 1 | 1974-11-01 | 1973-12-01 | 773.0 | 775.329285 |
+| 35 | 1 | 1974-12-01 | 1973-12-01 | 813.0 | 811.767639 |
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=season_length), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | DynamicTheta |
+|-----|-----------|--------|--------------|
+| 0 | 1 | mae | 8.182119 |
+| 1 | 1 | mape | 0.009736 |
+| 2 | 1 | mase | 0.367965 |
+| 3 | 1 | rmse | 9.817624 |
+| 4 | 1 | smape | 0.004874 |
+
+## References
+
+1. [Kostas I. Nikolopoulos, Dimitrios D. Thomakos. Forecasting with the
+ Theta Method-Theory and Applications. 2019 John Wiley & Sons
+ Ltd.](https://onlinelibrary.wiley.com/doi/book/10.1002/9781119320784)
+2. [Jose A. Fiorucci, Tiago R. Pellegrini, Francisco Louzada, Fotios
+ Petropoulos, Anne B. Koehler (2016). “Models for optimising the
+ theta method and their relationship to state space models”.
+ International Journal of
+ Forecasting](https://www.sciencedirect.com/science/article/pii/S0169207016300243).
+3. [Nixtla DynamicTheta API](../../src/core/models.html#dynamictheta)
+4. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+5. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ principles and practice, Time series
+ cross-validation”.](https://otexts.com/fpp3/tscv.html).
+6. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/models/garch.html.mdx b/statsforecast/docs/models/garch.html.mdx
new file mode 100644
index 00000000..2c4d4ce1
--- /dev/null
+++ b/statsforecast/docs/models/garch.html.mdx
@@ -0,0 +1,1046 @@
+---
+title: GARCH Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `GARCH Model` with `Statsforecast`.
+
+In this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation`.
+
+The text in this article is largely taken from: 1. [Changquan Huang •
+Alla Petukhina. Springer series (2022). Applied Time Series Analysis and
+Forecasting with
+Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2) 2.
+[Bollerslev, T. (1986). Generalized autoregressive conditional
+heteroskedasticity. Journal of econometrics, 31(3),
+307-327.](https://citeseerx.ist.psu.edu/document?repid=rep1&type=pdf&doi=7da8bfa5295375c1141d797e80065a599153c19d)
+3. [Engle, R. F. (1982). Autoregressive conditional heteroscedasticity
+with estimates of the variance of United Kingdom inflation.
+Econometrica: Journal of the econometric society,
+987-1007.](http://www.econ.uiuc.edu/~econ508/Papers/engle82.pdf). 4.
+[James D. Hamilton. Time Series Analysis Princeton University Press,
+Princeton, New Jersey, 1st Edition,
+1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [GARCH Models](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of GARCH with StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+The Generalized Autoregressive Conditional Heteroskedasticity (GARCH)
+model is a statistical technique used to model and predict volatility in
+financial and economic time series. It was developed by Robert Engle in
+1982 as an extension of the Autoregressive Conditional
+Heteroskedasticity (ARCH) model proposed by Andrew Lo and Craig
+MacKinlay in 1988.
+
+The GARCH model allows capturing the presence of conditional
+heteroscedasticity in time series data, that is, the presence of
+fluctuations in the variance of a time series as a function of time.
+This is especially useful in financial data analysis, where volatility
+can be an important measure of risk.
+
+The GARCH model has become a fundamental tool in the analysis of
+financial time series and has been used in a wide variety of
+applications, from risk management to forecasting prices of shares and
+other financial values.
+
+## Definition of GARCH Models
+
+**Definition 1.** A $\text{GARCH}(p,q)$ model with order $(p≥1,q≥0)$ is
+of the form
+
+$$
+\begin{equation}
+ \begin{cases}
+ X_t = \sigma_t \varepsilon_t\\
+ \sigma_{t}^2 = \omega + \sum_{i=1}^{p} \alpha_i X_{t-i}^2 + \sum_{j=1}^{q} \beta_j \sigma_{t-j}^2
+ \end{cases}
+\end{equation}
+$$
+
+where $\omega ≥0,\alpha_i ≥0,\beta_j ≥0,\alpha_p >0$ ,and $\beta_q >0$
+are constants,$\varepsilon_t \sim iid(0,1)$, and $\varepsilon_t$ is
+independent of $\{X_k;k ≤ t − 1 \}$. A stochastic process $X_t$ is
+called a $\text{GARCH}(p, q )$ process if it satisfies Eq. (1).
+
+In practice, it has been found that for some time series, the
+$\text{ARCH}(p)$ model defined by (1) will provide an adequate fit only
+if the order $p$ is large. By allowing past volatilities to affect the
+present volatility in (1), a more parsimonious model may result. That is
+why we need `GARCH` models. Besides, note the condition that the order
+$p ≥ 1$. The **GARCH model** in Definition 1 has the properties as
+follows.
+
+Proposition 1. If $X_t$ is a $\text{GARCH}(p, q)$ process defined in (1)
+and $\sum_{i=1}^{p} \alpha_{i} + \sum_{j=1}^{q} \beta_j <1$,then the
+following propositions hold.
+
+- $X_{t}^2$ follows the $\text{ARMA}(m, q )$ model
+
+$$X_{t}^2=\omega +\sum_{i=1}^{m} (\alpha_i + \beta_i )X_{t-i}^2 + \eta_t − \sum_{j=1}^q \beta_j \eta_{t-j}$$
+
+where $\alpha_i =0$ for $i >p,βj =0$ for $j >q,m=max(p,q)$, and
+$\eta_t =\sigma_{t}^2 (\varepsilon_{t}^2 −1)$.
+
+- $X_t$ is a white noise with
+
+$$E(X)=0, E(X_{t+h} X_t )=0 \ \ \text{for} \ any \ \ h \neq 0, Var(X_t)= \frac{\omega}{1-\sum_{i=1}^{m} (\alpha_i + \beta_i )}$$
+
+- $\sigma_{t}^2$ is the conditional variance of $X_t$ , that is, we
+ have
+
+$$E(X_t|\mathscr{F}_{t−1}) = 0, \sigma_{t}^2 = Var(X_{t}^2|\mathscr{F}_{t−1}).$$
+
+- Model (1) reflects the fat tails and volatility clustering.
+
+Although an asset return series can usually be seen as a white noise,
+there exists such a return series so that it may be autocorrelated. What
+is more, a given original time series is not necessarily a return
+series, and at the same time, its values may be negative. If a time
+series is autocorrelated, we must first build an adequate model (e.g.,
+an ARMA model) for the series in order to remove any autocorrelation in
+it. Then check whether the residual series has an ARCH effect, and if
+yes then we further model the residuals. In other words, if a time
+series $Y_t$ is autocorrelated and has ARCH effect, then a GARCH model
+that can capture the features of $Y_t$t should be of the form
+
+where Eq. (2) is referred to as the mean equation (model) and Eq. (3) is
+known as the volatility (variance) equation (model), and $Z_t$ is a
+representative of exogenous regressors. If $Y_t$ is a return series,
+then typically $Y_t = r + X_t$ where $r$ is a constant that means the
+expected returns is fixed.
+
+### Advantages and disadvantages of the Generalized Autoregressive Conditional Heteroskedasticity (GARCH) Model
+
+| Advantages | Disadvantages |
+|--------------------------------|-----------------------------------------|
+| 1\. 1. Flexible model: The GARCH model is flexible and can fit different types of time series data with different volatility patterns. | 1\. Requires a large amount of data: The GARCH model requires a large amount of data to accurately estimate the model parameters. |
+| 2\. Ability to model volatility: The GARCH model is capable of modeling the volatility and heteroscedasticity of a time series, which can improve the accuracy of forecasts. | 2\. Sensitive to the model specification: The GARCH model is sensitive to the model specification and can be difficult to estimate if incorrectly specified. |
+| 3\. It incorporates past information: The GARCH model incorporates past information on the volatility of the time series, which makes it useful for predicting future volatility. | 3\. It can be computationally expensive: The GARCH model can be computationally expensive, especially if more complex models are used. |
+| 4\. Allows the inclusion of exogenous variables: The GARCH model can be extended to include exogenous variables, which can improve the accuracy of the predictions. | 4\. It does not consider extreme events: The GARCH model does not consider extreme or unexpected events in the time series, which can affect the accuracy of the predictions in situations of high volatility. |
+| 5\. The GARCH model makes it possible to model conditional heteroscedasticity, that is, the variation of the variance of a time series as a function of time and of the previous values of the time series itself. | 5\. The GARCH model assumes that the time series errors are normally distributed, which may not be true in practice. If the errors are not normally distributed, the model may produce inaccurate estimates of volatility. |
+| 6\. The GARCH model can be used to estimate the value at risk (VaR) and the conditional value at risk (CVaR) of an investment portfolio. | |
+
+### The Generalized Autoregressive Conditional Heteroskedasticity (GARCH) model can be applied in several fields
+
+The Generalized Autoregressive Conditional Heteroskedasticity (GARCH)
+model can be applied in a wide variety of areas where time series
+volatility is required to be modeled and predicted. Some of the areas in
+which the GARCH model can be applied are:
+
+1. **Financial markets:** the GARCH model is widely used to model the
+ volatility (risk) of returns on financial assets such as stocks,
+ bonds, currencies, etc. It allows you to capture the changing nature
+ of volatility.
+
+2. **Commodity prices:** the prices of raw materials such as oil, gold,
+ grains, etc. they exhibit conditional volatility that can be modeled
+ with GARCH.
+
+3. **Credit risk:** the risk of non-payment of loans and bonds also
+ presents volatility over time that suits GARCH well.
+
+4. **Economic time series:** macroeconomic indicators such as
+ inflation, GDP, unemployment, etc. they have conditional volatility
+ modelable with GARCH.
+
+5. **Implicit volatility:** the GARCH model allows estimating the
+ implicit volatility in financial options.
+
+6. **Forecasts:** GARCH allows conditional volatility forecasts to be
+ made in any time series.
+
+7. **Risk analysis:** GARCH is useful for measuring and managing the
+ risk of investment portfolios and assets.
+
+8. **Finance:** The GARCH model is widely used in finance to model the
+ price volatility of financial assets, such as stocks, bonds, and
+ currencies.
+
+9. **Economics:** The GARCH model is used in economics to model the
+ volatility of the prices of goods and services, inflation, and other
+ economic indicators.
+
+10. **Environmental sciences:** The GARCH model is applied in
+ environmental sciences to model the volatility of variables such as
+ temperature, precipitation, and air quality.
+
+11. **Social sciences:** The GARCH model is used in the social sciences
+ to model the volatility of variables such as crime, migration, and
+ employment.
+
+12. **Engineering:** The GARCH model is applied in engineering to model
+ the volatility of variables such as the demand for electrical
+ energy, industrial production, and vehicular traffic.
+
+13. **Health sciences:** The GARCH model is used in health sciences to
+ model the volatility of variables such as the number of cases of
+ infectious diseases and the prices of medicines.
+
+The GARCH Model is applicable in any context where it is required to
+model and forecast heterogeneous conditional volatility in time series,
+especially in finance and economics.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import matplotlib.pyplot as plt
+from statsmodels.graphics.tsaplots import plot_acf
+from statsmodels.graphics.tsaplots import plot_pacf
+plt.style.use('fivethirtyeight')
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#212946',
+ 'axes.facecolor': '#212946',
+ 'savefig.facecolor':'#212946',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#2A3459',
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+### Read Data
+
+Let’s pull the S&P500 stock data from the Yahoo Finance site.
+
+```python
+import datetime
+
+import pandas as pd
+import time
+import yfinance as yf
+
+ticker = '^GSPC'
+period1 = datetime.datetime(2015, 1, 1)
+period2 = datetime.datetime(2023, 9, 22)
+interval = '1d' # 1d, 1m
+
+SP_500 = yf.download(ticker, start=period1, end=period2, interval=interval, progress=False)
+SP_500 = SP_500.reset_index()
+
+SP_500.head()
+```
+
+| Price | Date | Adj Close | Close | High | Low | Open | Volume |
+|--------|---------------------------|-------------|-------------|-------------|-------------|-------------|------------|
+| Ticker | | ^GSPC | ^GSPC | ^GSPC | ^GSPC | ^GSPC | ^GSPC |
+| 0 | 2015-01-02 00:00:00+00:00 | 2058.199951 | 2058.199951 | 2072.360107 | 2046.040039 | 2058.899902 | 2708700000 |
+| 1 | 2015-01-05 00:00:00+00:00 | 2020.579956 | 2020.579956 | 2054.439941 | 2017.339966 | 2054.439941 | 3799120000 |
+| 2 | 2015-01-06 00:00:00+00:00 | 2002.609985 | 2002.609985 | 2030.250000 | 1992.439941 | 2022.150024 | 4460110000 |
+| 3 | 2015-01-07 00:00:00+00:00 | 2025.900024 | 2025.900024 | 2029.609985 | 2005.550049 | 2005.550049 | 3805480000 |
+| 4 | 2015-01-08 00:00:00+00:00 | 2062.139893 | 2062.139893 | 2064.080078 | 2030.609985 | 2030.609985 | 3934010000 |
+
+```python
+df=SP_500[["Date","Close"]]
+```
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|---------------------------|-------------|-----------|
+| 0 | 2015-01-02 00:00:00+00:00 | 2058.199951 | 1 |
+| 1 | 2015-01-05 00:00:00+00:00 | 2020.579956 | 1 |
+| 2 | 2015-01-06 00:00:00+00:00 | 2002.609985 | 1 |
+| 3 | 2015-01-07 00:00:00+00:00 | 2025.900024 | 1 |
+| 4 | 2015-01-08 00:00:00+00:00 | 2062.139893 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds datetime64[ns, UTC]
+y float64
+unique_id object
+dtype: object
+```
+
+## Explore data with the plot method
+
+Plot a series using the plot method from the StatsForecast class. This
+method prints a random series from the dataset and is useful for basic
+EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### The Augmented Dickey-Fuller Test
+
+An Augmented Dickey-Fuller (ADF) test is a type of statistical test that
+determines whether a unit root is present in time series data. Unit
+roots can cause unpredictable results in time series analysis. A null
+hypothesis is formed in the unit root test to determine how strongly
+time series data is affected by a trend. By accepting the null
+hypothesis, we accept the evidence that the time series data is not
+stationary. By rejecting the null hypothesis or accepting the
+alternative hypothesis, we accept the evidence that the time series data
+is generated by a stationary process. This process is also known as
+stationary trend. The values of the ADF test statistic are negative.
+Lower ADF values indicate a stronger rejection of the null hypothesis.
+
+Augmented Dickey-Fuller Test is a common statistical test used to test
+whether a given time series is stationary or not. We can achieve this by
+defining the null and alternate hypothesis.
+
+Null Hypothesis: Time Series is non-stationary. It gives a
+time-dependent trend. Alternate Hypothesis: Time Series is stationary.
+In another term, the series doesn’t depend on time.
+
+ADF or t Statistic \< critical values: Reject the null hypothesis, time
+series is stationary. ADF or t Statistic \> critical values: Failed to
+reject the null hypothesis, time series is non-stationary.
+
+Let’s check if our series that we are analyzing is a stationary series.
+Let’s create a function to check, using the `Dickey Fuller` test
+
+```python
+from statsmodels.tsa.stattools import adfuller
+
+def Augmented_Dickey_Fuller_Test_func(series , column_name):
+ print (f'Dickey-Fuller test results for columns: {column_name}')
+ dftest = adfuller(series, autolag='AIC')
+ dfoutput = pd.Series(dftest[0:4], index=['Test Statistic','p-value','No Lags Used','Number of observations used'])
+ for key,value in dftest[4].items():
+ dfoutput['Critical Value (%s)'%key] = value
+ print (dfoutput)
+ if dftest[1] \<= 0.05:
+ print("Conclusion:====>")
+ print("Reject the null hypothesis")
+ print("The data is stationary")
+ else:
+ print("Conclusion:====>")
+ print("The null hypothesis cannot be rejected")
+ print("The data is not stationary")
+```
+
+
+```python
+Augmented_Dickey_Fuller_Test_func(df["y"],'S&P500')
+```
+
+``` text
+Dickey-Fuller test results for columns: S&P500
+Test Statistic -0.814971
+p-value 0.814685
+No Lags Used 10.000000
+ ...
+Critical Value (1%) -3.433341
+Critical Value (5%) -2.862861
+Critical Value (10%) -2.567473
+Length: 7, dtype: float64
+Conclusion:====>
+The null hypothesis cannot be rejected
+The data is not stationary
+```
+
+In the previous result we can see that the `Augmented_Dickey_Fuller`
+test gives us a `p-value` of 0.864700, which tells us that the null
+hypothesis cannot be rejected, and on the other hand the data of our
+series are not stationary.
+
+We need to differentiate our time series, in order to convert the data
+to stationary.
+
+### Return Series
+
+Since the 1970s, the financial industry has been very prosperous with
+advancement of computer and Internet technology. Trade of financial
+products (including various derivatives) generates a huge amount of data
+which form financial time series. For finance, the return on a financial
+product is most interesting, and so our attention focuses on the return
+series. If $P_t$ is the closing price at time t for a certain financial
+product, then the return on this product is
+
+$$X_t = \frac{(P_t − P_{t−1})}{P_{t−1}} ≈ log(P_t ) − log(P_{t−1}).$$
+
+It is return series $\{X_t \}$ that have been much independently
+studied. And important stylized features which are common across many
+instruments, markets, and time periods have been summarized. Note that
+if you purchase the financial product, then it becomes your asset, and
+its returns become your asset returns. Now let us look at the following
+examples.
+
+We can estimate the series of returns using the
+[pandas](https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.pct_change.html),
+`DataFrame.pct_change()` function. The `pct_change()` function has a
+periods parameter whose default value is 1. If you want to calculate a
+30-day return, you must change the value to 30.
+
+```python
+df['return'] = 100 * df["y"].pct_change()
+df.dropna(inplace=True, how='any')
+df.head()
+```
+
+| | ds | y | unique_id | return |
+|-----|---------------------------|-------------|-----------|-----------|
+| 1 | 2015-01-05 00:00:00+00:00 | 2020.579956 | 1 | -1.827811 |
+| 2 | 2015-01-06 00:00:00+00:00 | 2002.609985 | 1 | -0.889347 |
+| 3 | 2015-01-07 00:00:00+00:00 | 2025.900024 | 1 | 1.162984 |
+| 4 | 2015-01-08 00:00:00+00:00 | 2062.139893 | 1 | 1.788828 |
+| 5 | 2015-01-09 00:00:00+00:00 | 2044.810059 | 1 | -0.840381 |
+
+```python
+import plotly.express as px
+fig = px.line(df, x=df["ds"], y="return",title="SP500 Return Chart",template = "plotly_dark")
+fig.show()
+```
+
+
+
+### Creating Squared Returns
+
+```python
+df['sq_return'] = df["return"].mul(df["return"])
+df.head()
+```
+
+| | ds | y | unique_id | return | sq_return |
+|-----|---------------------------|-------------|-----------|-----------|-----------|
+| 1 | 2015-01-05 00:00:00+00:00 | 2020.579956 | 1 | -1.827811 | 3.340891 |
+| 2 | 2015-01-06 00:00:00+00:00 | 2002.609985 | 1 | -0.889347 | 0.790938 |
+| 3 | 2015-01-07 00:00:00+00:00 | 2025.900024 | 1 | 1.162984 | 1.352532 |
+| 4 | 2015-01-08 00:00:00+00:00 | 2062.139893 | 1 | 1.788828 | 3.199906 |
+| 5 | 2015-01-09 00:00:00+00:00 | 2044.810059 | 1 | -0.840381 | 0.706240 |
+
+### Returns vs Squared Returns
+
+```python
+from plotly.subplots import make_subplots
+import plotly.graph_objects as go
+
+fig = make_subplots(rows=1, cols=2)
+
+fig.add_trace(go.Scatter(x=df["ds"], y=df["return"],
+ mode='lines',
+ name='return'),
+row=1, col=1
+)
+
+
+fig.add_trace(go.Scatter(x=df["ds"], y=df["sq_return"],
+ mode='lines',
+ name='sq_return'),
+ row=1, col=2
+)
+
+fig.update_layout(height=600, width=800, title_text="Returns vs Squared Returns", template = "plotly_dark")
+fig.show()
+```
+
+
+
+```python
+from scipy.stats import probplot, moment
+from statsmodels.tsa.stattools import adfuller, q_stat, acf
+import numpy as np
+import seaborn as sns
+
+def plot_correlogram(x, lags=None, title=None):
+ lags = min(10, int(len(x)/5)) if lags is None else lags
+ fig, axes = plt.subplots(nrows=2, ncols=2, figsize=(14, 8))
+ x.plot(ax=axes[0][0], title='Return')
+ x.rolling(21).mean().plot(ax=axes[0][0], c='k', lw=1)
+ q_p = np.max(q_stat(acf(x, nlags=lags), len(x))[1])
+ stats = f'Q-Stat: {np.max(q_p):>8.2f}\nADF: {adfuller(x)[1]:>11.2f}'
+ axes[0][0].text(x=.02, y=.85, s=stats, transform=axes[0][0].transAxes)
+ probplot(x, plot=axes[0][1])
+ mean, var, skew, kurtosis = moment(x, moment=[1, 2, 3, 4])
+ s = f'Mean: {mean:>12.2f}\nSD: {np.sqrt(var):>16.2f}\nSkew: {skew:12.2f}\nKurtosis:{kurtosis:9.2f}'
+ axes[0][1].text(x=.02, y=.75, s=s, transform=axes[0][1].transAxes)
+ plot_acf(x=x, lags=lags, zero=False, ax=axes[1][0])
+ plot_pacf(x, lags=lags, zero=False, ax=axes[1][1])
+ axes[1][0].set_xlabel('Lag')
+ axes[1][1].set_xlabel('Lag')
+ fig.suptitle(title+ f'Dickey-Fuller: {adfuller(x)[1]:>11.2f}', fontsize=14)
+ sns.despine()
+ fig.tight_layout()
+ fig.subplots_adjust(top=.9)
+```
+
+
+```python
+plot_correlogram(df["return"], lags=30, title="Time Series Analysis plot \n")
+```
+
+
+
+### Ljung-Box Test
+
+Ljung-Box is a test for autocorrelation that we can use in tandem with
+our ACF and PACF plots. The Ljung-Box test takes our data, optionally
+either lag values to test, or the largest lag value to consider, and
+whether to compute the Box-Pierce statistic. Ljung-Box and Box-Pierce
+are two similar test statisitcs, Q , that are compared against a
+chi-squared distribution to determine if the series is white noise. We
+might use the Ljung-Box test on the residuals of our model to look for
+autocorrelation, ideally our residuals would be white noise.
+
+- Ho : The data are independently distributed, no autocorrelation.
+- Ha : The data are not independently distributed; they exhibit serial
+ correlation.
+
+The Ljung-Box with the Box-Pierce option will return, for each lag, the
+Ljung-Box test statistic, Ljung-Box p-values, Box-Pierce test statistic,
+and Box-Pierce p-values.
+
+If $p<\alpha (0.05)$ we reject the null hypothesis.
+
+```python
+from statsmodels.stats.diagnostic import acorr_ljungbox
+
+ljung_res = acorr_ljungbox(df["return"], lags= 40, boxpierce=True)
+
+ljung_res.head()
+```
+
+| | lb_stat | lb_pvalue | bp_stat | bp_pvalue |
+|-----|-----------|--------------|-----------|--------------|
+| 1 | 49.222273 | 2.285409e-12 | 49.155183 | 2.364927e-12 |
+| 2 | 62.991348 | 2.097020e-14 | 62.899234 | 2.195861e-14 |
+| 3 | 63.944944 | 8.433622e-14 | 63.850663 | 8.834380e-14 |
+| 4 | 74.343652 | 2.742989e-15 | 74.221024 | 2.911751e-15 |
+| 5 | 80.234862 | 7.494100e-16 | 80.093498 | 8.022242e-16 |
+
+## Split the data into training and testing
+
+Let’s divide our data into sets 1. Data to train our `GARCH` model 2.
+Data to test our model
+
+For the test data we will use the last 30 day to test and evaluate the
+performance of our model.
+
+```python
+df=df[["ds","unique_id","return"]]
+df.columns=["ds", "unique_id", "y"]
+```
+
+
+```python
+train = df[df.ds\<='2023-05-31'] # Let's forecast the last 30 days
+test = df[df.ds>'2023-05-31']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((2116, 3), (78, 3))
+```
+
+## Implementation of GARCH with StatsForecast
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import GARCH
+```
+
+### Instantiating Models
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/) by the
+master, Rob Hyndmann, can be useful.season_length.
+
+```python
+season_length = 7 # Dayly data
+horizon = len(test) # number of predictions biasadj=True, include_drift=True,
+
+models = [GARCH(1,1),
+ GARCH(1,2),
+ GARCH(2,2),
+ GARCH(2,1),
+ GARCH(3,1),
+ GARCH(3,2),
+ GARCH(3,3),
+ GARCH(1,3),
+ GARCH(2,3)]
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [pandas’
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(
+ models=models,
+ freq='C', # custom business day frequency
+)
+```
+
+## Cross-validation
+
+We have built different GARCH models, so we need to determine which is
+the best model to then be able to train it and thus be able to make the
+predictions. To know which is the best model we go to the Cross
+Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 12 months ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=train,
+ h=horizon,
+ step_size=6,
+ n_windows=5)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` series identifier
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the n_windows.
+- `y:` true value
+- `"model":` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df
+```
+
+| | unique_id | ds | cutoff | y | GARCH(1,1) | GARCH(1,2) | GARCH(2,2) | GARCH(2,1) | GARCH(3,1) | GARCH(3,2) | GARCH(3,3) | GARCH(1,3) | GARCH(2,3) |
+|-----|-----------|---------------------------|---------------------------|-----------|------------|------------|------------|------------|------------|------------|------------|------------|------------|
+| 0 | 1 | 2023-01-04 00:00:00+00:00 | 2023-01-03 00:00:00+00:00 | 0.753897 | 1.678755 | 1.678412 | 1.680475 | 1.686649 | 1.719494 | 2.210902 | 1.702743 | 1.647114 | 1.637795 |
+| 1 | 1 | 2023-01-05 00:00:00+00:00 | 2023-01-03 00:00:00+00:00 | -1.164553 | -0.728069 | -0.745487 | -0.730648 | -0.722156 | -0.738119 | -0.824748 | -0.755277 | -0.740976 | -0.744150 |
+| 2 | 1 | 2023-01-06 00:00:00+00:00 | 2023-01-03 00:00:00+00:00 | 2.284078 | -0.589733 | -0.582982 | -0.590078 | -0.598076 | -0.587109 | -0.866347 | -0.571160 | -0.587807 | -0.584692 |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| 387 | 1 | 2023-05-26 00:00:00+00:00 | 2023-02-07 00:00:00+00:00 | 1.304909 | -1.697814 | -1.694747 | -1.702537 | -1.735631 | -1.729903 | -1.712997 | -1.663399 | -1.702160 | -1.687723 |
+| 388 | 1 | 2023-05-30 00:00:00+00:00 | 2023-02-07 00:00:00+00:00 | 0.001660 | -0.326945 | -0.337504 | -0.329686 | -0.330120 | -0.334717 | -0.327583 | -0.330260 | -0.338245 | -0.332412 |
+| 389 | 1 | 2023-05-31 00:00:00+00:00 | 2023-02-07 00:00:00+00:00 | -0.610862 | 0.807625 | 0.787054 | 0.807819 | 0.841536 | 0.811702 | 0.836159 | 0.772193 | 0.801933 | 0.804526 |
+
+```python
+from utilsforecast.evaluation import evaluate
+from utilsforecast.losses import rmse
+```
+
+
+```python
+evals = evaluate(crossvalidation_df.drop(columns='cutoff'), metrics=[rmse], agg_fn='mean')
+evals
+```
+
+| | metric | GARCH(1,1) | GARCH(1,2) | GARCH(2,2) | GARCH(2,1) | GARCH(3,1) | GARCH(3,2) | GARCH(3,3) | GARCH(1,3) | GARCH(2,3) |
+|-----|--------|------------|------------|------------|------------|------------|------------|------------|------------|------------|
+| 0 | rmse | 1.383143 | 1.526258 | 1.481056 | 1.389969 | 1.453538 | 1.539906 | 1.392352 | 1.515796 | 1.389061 |
+
+```python
+evals.drop(columns='metric').loc[0].idxmin()
+```
+
+``` text
+'GARCH(1,1)'
+```
+
+**Note:** This result can vary depending on the data and period you use
+to train and test the model, and the models you want to test. This is an
+example, where the objective is to be able to teach a methodology for
+the use of `StatsForecast`, and in particular the GARCH model and the
+parameters used in Cross Validation to determine the best model for this
+example.
+
+In the previous result it can be seen that the best model is the model
+$\text{GARCH}(1,1)$
+
+With this result found using Cross Validation to determine which is the
+best model, we are going to continue training our model, to then make
+the predictions.
+
+### Fit the Model
+
+```python
+season_length = 7 # Dayly data
+horizon = len(test) # number of predictions biasadj=True, include_drift=True,
+
+models = [GARCH(1,1)]
+```
+
+
+```python
+sf = StatsForecast(models=models,
+ freq='C', # custom business day frequency
+ )
+```
+
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[GARCH(1,1)])
+```
+
+Let’s see the results of our Theta model. We can observe it with the
+following instruction:
+
+```python
+result=sf.fitted_[0,0].model_
+result
+```
+
+``` text
+{'p': 1,
+ 'q': 1,
+ 'coeff': array([0.03745049, 0.18399111, 0.7890637 ]),
+ 'message': 'Optimization terminated successfully',
+ 'y_vals': array([-0.61086242]),
+ 'sigma2_vals': array([0.76298402]),
+ 'fitted': array([ nan, 2.14638896, -0.76426268, ..., -0.19747638,
+ 0.76993462, 0.13183178]),
+ 'actual_residuals': array([ nan, -3.03573613, 1.92724695, ..., 1.50238505,
+ -0.7682743 , -0.7426942 ])}
+```
+
+Let us now visualize the residuals of our models.
+
+As we can see, the result obtained above has an output in a dictionary,
+to extract each element from the dictionary we are going to use the
+`.get()` function to extract the element and then we are going to save
+it in a `pd.DataFrame()`.
+
+```python
+residual=pd.DataFrame(result.get("actual_residuals"), columns=["residual Model"])
+residual
+```
+
+| | residual Model |
+|------|----------------|
+| 0 | NaN |
+| 1 | -3.035736 |
+| 2 | 1.927247 |
+| ... | ... |
+| 2113 | 1.502385 |
+| 2114 | -0.768274 |
+| 2115 | -0.742694 |
+
+```python
+from scipy import stats
+
+fig, axs = plt.subplots(nrows=2, ncols=2)
+
+# plot[1,1]
+residual.plot(ax=axs[0,0])
+axs[0,0].set_title("Residuals");
+
+# plot
+sns.distplot(residual, ax=axs[0,1]);
+axs[0,1].set_title("Density plot - Residual");
+
+# plot
+stats.probplot(residual["residual Model"], dist="norm", plot=axs[1,0])
+axs[1,0].set_title('Plot Q-Q')
+
+# plot
+plot_acf(residual, lags=35, ax=axs[1,1],color="fuchsia")
+axs[1,1].set_title("Autocorrelation");
+
+plt.show();
+```
+
+
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[90]` means that
+ the model expects the real value to be inside that interval 90% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min. (If you want to speed things up to a couple of seconds,
+remove the AutoModels like `ARIMA` and `Theta`)
+
+```python
+Y_hat = sf.forecast(df=train, h=horizon, fitted=True)
+Y_hat.head()
+```
+
+| | unique_id | ds | GARCH(1,1) |
+|-----|-----------|---------------------------|------------|
+| 0 | 1 | 2023-06-01 00:00:00+00:00 | 1.366914 |
+| 1 | 1 | 2023-06-02 00:00:00+00:00 | -0.593121 |
+| 2 | 1 | 2023-06-05 00:00:00+00:00 | -0.485200 |
+| 3 | 1 | 2023-06-06 00:00:00+00:00 | -0.927145 |
+| 4 | 1 | 2023-06-07 00:00:00+00:00 | 0.766640 |
+
+```python
+Y_hat = sf.forecast(df=train, h=horizon, fitted=True, level=[95])
+Y_hat.head()
+```
+
+| | unique_id | ds | GARCH(1,1) | GARCH(1,1)-lo-95 | GARCH(1,1)-hi-95 |
+|-----|-----------|---------------------------|------------|------------------|------------------|
+| 0 | 1 | 2023-06-01 00:00:00+00:00 | 1.366914 | -0.021035 | 2.754863 |
+| 1 | 1 | 2023-06-02 00:00:00+00:00 | -0.593121 | -2.435497 | 1.249254 |
+| 2 | 1 | 2023-06-05 00:00:00+00:00 | -0.485200 | -2.139216 | 1.168815 |
+| 3 | 1 | 2023-06-06 00:00:00+00:00 | -0.927145 | -2.390566 | 0.536276 |
+| 4 | 1 | 2023-06-07 00:00:00+00:00 | 0.766640 | -0.771479 | 2.304759 |
+
+```python
+values=sf.forecast_fitted_values()
+values.head()
+```
+
+| | unique_id | ds | y | GARCH(1,1) | GARCH(1,1)-lo-95 | GARCH(1,1)-hi-95 |
+|-----|-----------|---------------------------|-----------|------------|------------------|------------------|
+| 0 | 1 | 2015-01-05 00:00:00+00:00 | -1.827811 | NaN | NaN | NaN |
+| 1 | 1 | 2015-01-06 00:00:00+00:00 | -0.889347 | 2.146389 | -0.972874 | 5.265652 |
+| 2 | 1 | 2015-01-07 00:00:00+00:00 | 1.162984 | -0.764263 | -3.883526 | 2.355000 |
+| 3 | 1 | 2015-01-08 00:00:00+00:00 | 1.788828 | -0.650707 | -3.769970 | 2.468556 |
+| 4 | 1 | 2015-01-09 00:00:00+00:00 | -0.840381 | -1.449049 | -4.568312 | 1.670214 |
+
+Adding 95% confidence interval with the forecast method
+
+```python
+sf.forecast(df=train, h=horizon, level=[95])
+```
+
+| | unique_id | ds | GARCH(1,1) | GARCH(1,1)-lo-95 | GARCH(1,1)-hi-95 |
+|-----|-----------|---------------------------|------------|------------------|------------------|
+| 0 | 1 | 2023-06-01 00:00:00+00:00 | 1.366914 | -0.021035 | 2.754863 |
+| 1 | 1 | 2023-06-02 00:00:00+00:00 | -0.593121 | -2.435497 | 1.249254 |
+| 2 | 1 | 2023-06-05 00:00:00+00:00 | -0.485200 | -2.139216 | 1.168815 |
+| ... | ... | ... | ... | ... | ... |
+| 75 | 1 | 2023-09-14 00:00:00+00:00 | -1.686546 | -3.049859 | -0.323233 |
+| 76 | 1 | 2023-09-15 00:00:00+00:00 | -0.322556 | -2.497448 | 1.852335 |
+| 77 | 1 | 2023-09-18 00:00:00+00:00 | 0.799407 | -1.027642 | 2.626457 |
+
+```python
+sf.plot(train, Y_hat.merge(test), max_insample_length=200)
+```
+
+
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 30 dayly ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[95]` means that
+ the model expects the real value to be inside that interval 95% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+sf.predict(h=horizon)
+```
+
+| | unique_id | ds | GARCH(1,1) |
+|-----|-----------|---------------------------|------------|
+| 0 | 1 | 2023-06-01 00:00:00+00:00 | 1.366914 |
+| 1 | 1 | 2023-06-02 00:00:00+00:00 | -0.593121 |
+| 2 | 1 | 2023-06-05 00:00:00+00:00 | -0.485200 |
+| ... | ... | ... | ... |
+| 75 | 1 | 2023-09-14 00:00:00+00:00 | -1.686546 |
+| 76 | 1 | 2023-09-15 00:00:00+00:00 | -0.322556 |
+| 77 | 1 | 2023-09-18 00:00:00+00:00 | 0.799407 |
+
+```python
+forecast_df = sf.predict(h=horizon, level=[80,95])
+forecast_df.head(10)
+```
+
+| | unique_id | ds | GARCH(1,1) | GARCH(1,1)-lo-95 | GARCH(1,1)-lo-80 | GARCH(1,1)-hi-80 | GARCH(1,1)-hi-95 |
+|-----|-----------|---------------------------|------------|------------------|------------------|------------------|------------------|
+| 0 | 1 | 2023-06-01 00:00:00+00:00 | 1.366914 | -0.021035 | 0.459383 | 2.274445 | 2.754863 |
+| 1 | 1 | 2023-06-02 00:00:00+00:00 | -0.593121 | -2.435497 | -1.797786 | 0.611543 | 1.249254 |
+| 2 | 1 | 2023-06-05 00:00:00+00:00 | -0.485200 | -2.139216 | -1.566703 | 0.596303 | 1.168815 |
+| ... | ... | ... | ... | ... | ... | ... | ... |
+| 7 | 1 | 2023-06-12 00:00:00+00:00 | -1.051435 | -4.790880 | -3.496526 | 1.393657 | 2.688010 |
+| 8 | 1 | 2023-06-13 00:00:00+00:00 | 0.421605 | -3.001123 | -1.816396 | 2.659607 | 3.844333 |
+| 9 | 1 | 2023-06-14 00:00:00+00:00 | -0.300086 | -3.138338 | -2.155920 | 1.555747 | 2.538166 |
+
+```python
+sf.plot(train, test.merge(forecast_df), level=[80, 95], max_insample_length=200)
+```
+
+
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=season_length), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | GARCH(1,1) |
+|-----|-----------|--------|------------|
+| 0 | 1 | mae | 0.843296 |
+| 1 | 1 | mape | 3.703305 |
+| 2 | 1 | mase | 0.794905 |
+| 3 | 1 | rmse | 1.048076 |
+| 4 | 1 | smape | 0.709150 |
+
+## References
+
+1. [Changquan Huang • Alla Petukhina. Springer series (2022). Applied
+ Time Series Analysis and Forecasting with
+ Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2)
+2. [Bollerslev, T. (1986). Generalized autoregressive conditional
+ heteroskedasticity. Journal of econometrics, 31(3),
+ 307-327.](https://citeseerx.ist.psu.edu/document?repid=rep1&type=pdf&doi=7da8bfa5295375c1141d797e80065a599153c19d)
+3. [Engle, R. F. (1982). Autoregressive conditional heteroscedasticity
+ with estimates of the variance of United Kingdom inflation.
+ Econometrica: Journal of the econometric society,
+ 987-1007.](http://www.econ.uiuc.edu/~econ508/Papers/engle82.pdf).
+4. [James D. Hamilton. Time Series Analysis Princeton University Press,
+ Princeton, New Jersey, 1st Edition,
+ 1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+5. [Nixtla Garch API](../../src/core/models.html#garch)
+6. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+7. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+8. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/models/holt.html.mdx b/statsforecast/docs/models/holt.html.mdx
new file mode 100644
index 00000000..c28d1d11
--- /dev/null
+++ b/statsforecast/docs/models/holt.html.mdx
@@ -0,0 +1,1127 @@
+---
+title: Holt Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `Holt Model` with `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation` in other.
+
+The text in this article is largely taken from: 1. [Changquan Huang •
+Alla Petukhina. Springer series (2022). Applied Time Series Analysis and
+Forecasting with
+Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2) 2.
+Ivan Svetunkov. [Forecasting and Analytics with the Augmented Dynamic
+Adaptive Model (ADAM)](https://openforecast.org/adam/) 3. [James D.
+Hamilton. Time Series Analysis Princeton University Press, Princeton,
+New Jersey, 1st Edition,
+1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+Principles and Practice (3rd ed)”](https://otexts.com/fpp3/tscv.html).
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [Holt Model](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of Holt with StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+The Holts model, also known as the double exponential smoothing method,
+is a forecasting technique widely used in time series analysis. It was
+developed by Charles Holt in 1957 as an improvement on Brown’s simple
+exponential smoothing method.
+
+The Holts model is used to predict future values of a time series that
+exhibits a trend. The model uses two smoothing parameters, one for
+estimating the trend and the other for estimating the level or base
+level of the time series. These parameters are called $\alpha$ and
+$\beta$, respectively.
+
+The Holts model is an extension of Brown’s simple exponential smoothing
+method, which uses only one smoothing parameter to estimate the trend
+and base level of the time series. The Holts model improves the accuracy
+of the forecasts by adding a second smoothing parameter for the trend.
+
+One of the main advantages of the Holts model is that it is easy to
+implement and does not require a large amount of historical data to
+generate accurate predictions. Furthermore, the model is highly
+adaptable and can be customized to fit a wide variety of time series.
+
+However, Holts’ model has some limitations. For example, the model
+assumes that the time series is stationary and that the trend is linear.
+If the time series is not stationary or has a non-linear trend, the
+Holts model may not be the most appropriate.
+
+In general, the Holts model is a useful and widely used technique in
+time series analysis, especially when the series is expected to exhibit
+a linear trend.
+
+## Holt Method
+
+`Simple exponential smoothing` does not function well when the data has
+trends. In those cases, we can use *double exponential smoothing*. This
+is a more reliable method for handling data that consumes trends without
+seasonality than compared to other methods. This method adds a time
+*trend* equation in the formulation. Two different weights, or smoothing
+parameters, are used to update these two components at a time.
+
+Holt’s exponential smoothing is also sometimes called *double
+exponential smoothing*. The main idea here is to use SES and advance it
+to capture the *trend* component.
+
+Holt (1957) extended simple exponential smoothing to allow the
+forecasting of data with a *trend*. This method involves a forecast
+equation and two smoothing equations (one for the *level* and one for
+the *trend*):
+
+Assume that a series has the following:
+
+- Level
+- Trend
+- No seasonality
+- Noise
+
+where $\ell_{t}$ denotes an estimate of the level of the series at time
+$t, b_t$ denotes an estimate of the trend (slope) of the series at time
+$t, \alpha$ is the smoothing parameter for the level, $0\le\alpha\le1$,
+and $\beta^{*}$ is the smoothing parameter for the trend,
+$0\le\beta^*\le1$.
+
+As with simple exponential smoothing, the level equation here shows that
+$\ell_{t}$ is a weighted average of observation $y_{t}$ and the
+one-step-ahead training forecast for time $t$, here given by
+$\ell_{t-1} + b_{t-1}$. The trend equation shows that $b_t$ is a
+weighted average of the estimated trend at time $t$ based on
+$\ell_{t} - \ell_{t-1}$ and $b_{t-1}$, the previous estimate of the
+trend.
+
+The forecast function is no longer flat but trending. The $h$-step-ahead
+forecast is equal to the last estimated level plus $h$ times the last
+estimated trend value. Hence the forecasts are a linear function of $h$.
+
+### Innovations state space models for exponential smoothing
+
+The exponential smoothing methods presented in Table 7.6 are algorithms
+which generate point forecasts. The statistical models in this tutorial
+generate the same point forecasts, but can also generate prediction (or
+forecast) intervals. A statistical model is a stochastic (or random)
+data generating process that can produce an entire forecast
+distribution.
+
+Each model consists of a measurement equation that describes the
+observed data, and some state equations that describe how the unobserved
+components or states (level, trend, seasonal) change over time. Hence,
+these are referred to as state space models.
+
+For each method there exist two models: one with additive errors and one
+with multiplicative errors. The point forecasts produced by the models
+are identical if they use the same smoothing parameter values. They
+will, however, generate different prediction intervals.
+
+To distinguish between a model with additive errors and one with
+multiplicative errors. We label each state space model as ETS( .,.,.)
+for (Error, Trend, Seasonal). This label can also be thought of as
+ExponenTial Smoothing. Using the same notation as in Table 7.5, the
+possibilities for each component are: $Error=\{A,M\}$,
+$Trend=\{N,A,A_d\}$ and $Seasonal=\{N,A,M\}$
+
+For our case, the linear Holt model with a trend, we are going to see
+two cases, both for the additive and the multiplicative
+
+### ETS(A,A,N): Holt’s linear method with additive errors
+
+For this model, we assume that the one-step-ahead training errors are
+given by
+$\varepsilon_t=y_t-\ell_{t-1}-b_{t-1} \sim \text{NID}(0,\sigma^2)$.
+Substituting this into the error correction equations for Holt’s linear
+method we obtain
+
+where, for simplicity, we have set $\beta=\alpha \beta^*$
+
+### ETS(M,A,N): Holt’s linear method with multiplicative errors
+
+Specifying one-step-ahead training errors as relative errors such that
+
+$$\varepsilon_t=\frac{y_t-(\ell_{t-1}+b_{t-1})}{(\ell_{t-1}+b_{t-1})}$$
+
+and following an approach similar to that used above, the innovations
+state space model underlying Holt’s linear method with multiplicative
+errors is specified as
+
+where again $\beta=\alpha \beta^*$ and
+$\varepsilon_t \sim \text{NID}(0,\sigma^2)$.
+
+### A taxonomy of exponential smoothing methods
+
+Building on the idea of time series components, we can move to the ETS
+taxonomy. ETS stands for “Error-Trend-Seasonality” and defines how
+specifically the components interact with each other. Based on the type
+of error, trend and seasonality, Pegels (1969) proposed a taxonomy,
+which was then developed further by Hyndman et al. (2002) and refined by
+Hyndman et al. (2008). According to this taxonomy, error, trend and
+seasonality can be:
+
+1. Error: “Additive” (A), or “Multiplicative” (M);
+2. Trend: “None” (N), or “Additive” (A), or “Additive damped” (Ad), or
+ “Multiplicative” (M), or “Multiplicative damped” (Md);
+3. Seasonality: “None” (N), or “Additive” (A), or “Multiplicative” (M).
+
+The components in the ETS taxonomy have clear interpretations: level
+shows average value per time period, trend reflects the change in the
+value, while seasonality corresponds to periodic fluctuations
+(e.g. increase in sales each January). Based on the the types of the
+components above, it is theoretically possible to devise 30 ETS models
+with different types of error, trend and seasonality. Figure 1 shows
+examples of different time series with deterministic (they do not change
+over time) level, trend, seasonality and with the additive error term.
+
+
+*Figure 4.1: Time series corresponding to the additive error ETS models*
+
+Things to note from the plots in Figure.1:
+
+1. When seasonality is multiplicative, its amplitude increases with the
+ increase of the level of the data, while with additive seasonality,
+ the amplitude is constant. Compare, for example, ETS(A,A,A) with
+ ETS(A,A,M): for the former, the distance between the highest and the
+ lowest points in the first year is roughly the same as in the last
+ year. In the case of ETS(A,A,M) the distance increases with the
+ increase in the level of series;
+2. When the trend is multiplicative, data exhibits exponential
+ growth/decay;
+3. The damped trend slows down both additive and multiplicative trends;
+4. It is practically impossible to distinguish additive and
+ multiplicative seasonality if the level of series does not change
+ because the amplitude of seasonality will be constant in both cases
+ (compare ETS(A,N,A) and ETS(A,N,M)).
+
+
+*Figure 2: Time series corresponding to the multiplicative error ETS
+models*
+
+The graphs in Figure 2 show approximately the same idea as the additive
+case, the main difference is that the error variance increases with
+increasing data level; this becomes clearer in ETS(M,A,N) and ETS(M,M,N)
+data. This property is called heteroskedasticity in statistics, and
+Hyndman et al. (2008) argue that the main benefit of multiplicative
+error models is to capture this characteristic.
+
+### Mathematical models in the ETS taxonomy
+
+I hope that it becomes more apparent to the reader how the ETS framework
+is built upon the idea of time series decomposition. By introducing
+different components, defining their types, and adding the equations for
+their update, we can construct models that would work better in
+capturing the key features of the time series. But we should also
+consider the potential change in components over time. The “transition”
+or “state” equations are supposed to reflect this change: they explain
+how the level, trend or seasonal components evolve.
+
+As discussed in Section 2.2, given different types of components and
+their interactions, we end up with 30 models in the taxonomy. Tables 1
+and 2 summarise mathematically all 30 ETS models shown graphically on
+Figures 1 and 2, presenting formulae for measurement and transition
+equations.
+
+Table 1: Additive error ETS models \| \| Nonseasonal \|Additive
+\|Multiplicative\| \|—-\|———–\|———–\|————–\| \|No
+trend\|$\begin{aligned} &y_{t} = l_{t-1} + \epsilon_t \\ &l_t = l_{t-1} + \alpha \epsilon_t \end{aligned}$
+\|$\begin{aligned} &y_{t} = l_{t-1} + s_{t-m} + \epsilon_t \\ &l_t = l_{t-1} + \alpha \epsilon_t \\ &s_t = s_{t-m} + \gamma \epsilon_t \end{aligned}$
+\|$\begin{aligned} &y_{t} = l_{t-1} s_{t-m} + \epsilon_t \\ &l_t = l_{t-1} + \alpha \frac{\epsilon_t}{s_{t-m}} \\ &s_t = s_{t-m} + \gamma \frac{\epsilon_t}{l_{t-1}} \end{aligned}$\|
+\|Additive\|
+$\begin{aligned} &y_{t} = l_{t-1} + b_{t-1} + \epsilon_t \\ &l_t = l_{t-1} + b_{t-1} + \alpha \epsilon_t \\ &b_t = b_{t-1} + \beta \epsilon_t \end{aligned}$
+\|
+$\begin{aligned} &y_{t} = l_{t-1} + b_{t-1} + s_{t-m} + \epsilon_t \\ &l_t = l_{t-1} + b_{t-1} + \alpha \epsilon_t \\ &b_t = b_{t-1} + \beta \epsilon_t \\ &s_t = s_{t-m} + \gamma \epsilon_t \end{aligned}$
+\|
+$\begin{aligned} &y_{t} = (l_{t-1} + b_{t-1}) s_{t-m} + \epsilon_t \\ &l_t = l_{t-1} + b_{t-1} + \alpha \frac{\epsilon_t}{s_{t-m}} \\ &b_t = b_{t-1} + \beta \frac{\epsilon_t}{s_{t-m}} \\ &s_t = s_{t-m} + \gamma \frac{\epsilon_t}{l_{t-1} + b_{t-1}} \end{aligned}$\|
+\|Additive damped\|
+$\begin{aligned} &y_{t} = l_{t-1} + \phi b_{t-1} + \epsilon_t \\ &l_t = l_{t-1} + \phi b_{t-1} + \alpha \epsilon_t \\ &b_t = \phi b_{t-1} + \beta \epsilon_t \end{aligned}$
+\|
+$\begin{aligned} &y_{t} = l_{t-1} + \phi b_{t-1} + s_{t-m} + \epsilon_t \\ &l_t = l_{t-1} + \phi b_{t-1} + \alpha \epsilon_t \\ &b_t = \phi b_{t-1} + \beta \epsilon_t \\ &s_t = s_{t-m} + \gamma \epsilon_t \end{aligned}$
+\|
+$\begin{aligned} &y_{t} = (l_{t-1} + \phi b_{t-1}) s_{t-m} + \epsilon_t \\ &l_t = l_{t-1} + \phi b_{t-1} + \alpha \frac{\epsilon_t}{s_{t-m}} \\ &b_t = \phi b_{t-1} + \beta \frac{\epsilon_t}{s_{t-m}} \\ &s_t = s_{t-m} + \gamma \frac{\epsilon_t}{l_{t-1} + \phi b_{t-1}} \end{aligned}$\|
+\|Multiplicative\|
+$\begin{aligned} &y_{t} = l_{t-1} b_{t-1} + \epsilon_t \\ &l_t = l_{t-1} b_{t-1} + \alpha \epsilon_t \\ &b_t = b_{t-1} + \beta \frac{\epsilon_t}{l_{t-1}} \end{aligned}$
+\|
+$\begin{aligned} &y_{t} = l_{t-1} b_{t-1} + s_{t-m} + \epsilon_t \\ &l_t = l_{t-1} b_{t-1} + \alpha \epsilon_t \\ &b_t = b_{t-1} + \beta \frac{\epsilon_t}{l_{t-1}} \\ &s_t = s_{t-m} + \gamma \epsilon_t \end{aligned}$
+\|
+$\begin{aligned} &y_{t} = l_{t-1} b_{t-1} s_{t-m} + \epsilon_t \\ &l_t = l_{t-1} b_{t-1} + \alpha \frac{\epsilon_t}{s_{t-m}} \\ &b_t = b_{t-1} + \beta \frac{\epsilon_t}{l_{t-1}s_{t-m}} \\ &s_t = s_{t-m} + \gamma \frac{\epsilon_t}{l_{t-1} b_{t-1}} \end{aligned}$\|
+\|Multiplicative damped\|
+$\begin{aligned} &y_{t} = l_{t-1} b_{t-1}^\phi + \epsilon_t \\ &l_t = l_{t-1} b_{t-1}^\phi + \alpha \epsilon_t \\ &b_t = b_{t-1}^\phi + \beta \frac{\epsilon_t}{l_{t-1}} \end{aligned}$
+\|
+$\begin{aligned} &y_{t} = l_{t-1} b_{t-1}^\phi + s_{t-m} + \epsilon_t \\ &l_t = l_{t-1} b_{t-1}^\phi + \alpha \epsilon_t \\ &b_t = b_{t-1}^\phi + \beta \frac{\epsilon_t}{l_{t-1}} \\ &s_t = s_{t-m} + \gamma \epsilon_t \end{aligned}$
+\|
+$\begin{aligned} &y_{t} = l_{t-1} b_{t-1}^\phi s_{t-m} + \epsilon_t \\ &l_t = l_{t-1} b_{t-1}^\phi + \alpha \frac{\epsilon_t}{s_{t-m}} \\ &b_t = b_{t-1}^\phi + \beta \frac{\epsilon_t}{l_{t-1}s_{t-m}} \\ &s_t = s_{t-m} + \gamma \frac{\epsilon_t}{l_{t-1} b_{t-1}} \end{aligned}$\|
+
+Table 2: Multiplicative error ETS models \| \|Nonseasonal \|Additive
+\|Multiplicative\| \|——\|————-\|———-\|————–\| \|No trend\|
+$\begin{aligned} &y_{t} = l_{t-1}(1 + \epsilon_t) \\ &l_t = l_{t-1}(1 + \alpha \epsilon_t) \end{aligned}$
+\|
+$\begin{aligned} &y_{t} = (l_{t-1} + s_{t-m})(1 + \epsilon_t) \\ &l_t = l_{t-1} + \alpha \mu_{y,t} \epsilon_t \\ &s_t = s_{t-m} + \gamma \mu_{y,t} \epsilon_t \end{aligned}$
+\|
+$\begin{aligned} &y_{t} = l_{t-1} s_{t-m}(1 + \epsilon_t) \\ &l_t = l_{t-1}(1 + \alpha \epsilon_t) \\ &s_t = s_{t-m}(1 + \gamma \epsilon_t) \end{aligned}$\|
+\|Additive\|
+$\begin{aligned} &y_{t} = (l_{t-1} + b_{t-1})(1 + \epsilon_t) \\ &l_t = (l_{t-1} + b_{t-1})(1 + \alpha \epsilon_t) \\ &b_t = b_{t-1} + \beta \mu_{y,t} \epsilon_t \end{aligned}$
+\|
+$\begin{aligned} &y_{t} = (l_{t-1} + b_{t-1} + s_{t-m})(1 + \epsilon_t) \\ &l_t = l_{t-1} + b_{t-1} + \alpha \mu_{y,t} \epsilon_t \\ &b_t = b_{t-1} + \beta \mu_{y,t} \epsilon_t \\ &s_t = s_{t-m} + \gamma \mu_{y,t} \epsilon_t \end{aligned}$
+\|
+$\begin{aligned} &y_{t} = (l_{t-1} + b_{t-1}) s_{t-m}(1 + \epsilon_t) \\ &l_t = (l_{t-1} + b_{t-1})(1 + \alpha \epsilon_t) \\ &b_t = b_{t-1} + \beta (l_{t-1} + b_{t-1}) \epsilon_t \\ &s_t = s_{t-m} (1 + \gamma \epsilon_t) \end{aligned}$\|
+\|Additive damped\|
+$\begin{aligned} &y_{t} = (l_{t-1} + \phi b_{t-1})(1 + \epsilon_t) \\ &l_t = (l_{t-1} + \phi b_{t-1})(1 + \alpha \epsilon_t) \\ &b_t = \phi b_{t-1} + \beta \mu_{y,t} \epsilon_t \end{aligned}$
+\|
+$\begin{aligned} &y_{t} = (l_{t-1} + \phi b_{t-1} + s_{t-m})(1 + \epsilon_t) \\ &l_t = l_{t-1} + \phi b_{t-1} + \alpha \mu_{y,t} \epsilon_t \\ &b_t = \phi b_{t-1} + \beta \mu_{y,t} \epsilon_t \\ &s_t = s_{t-m} + \gamma \mu_{y,t} \epsilon_t \end{aligned}$
+\|
+$\begin{aligned} &y_{t} = (l_{t-1} + \phi b_{t-1}) s_{t-m}(1 + \epsilon_t) \\ &l_t = l_{t-1} + \phi b_{t-1} (1 + \alpha \epsilon_t) \\ &b_t = \phi b_{t-1} + \beta (l_{t-1} + \phi b_{t-1}) \epsilon_t \\ &s_t = s_{t-m}(1 + \gamma \epsilon_t) \end{aligned}$\|
+\|Multiplicative\|
+$\begin{aligned} &y_{t} = l_{t-1} b_{t-1} (1 + \epsilon_t) \\ &l_t = l_{t-1} b_{t-1} (1 + \alpha \epsilon_t) \\ &b_t = b_{t-1} (1 + \beta \epsilon_t) \end{aligned}$
+\|
+$\begin{aligned} &y_{t} = (l_{t-1} b_{t-1} + s_{t-m})(1 + \epsilon_t) \\ &l_t = l_{t-1} b_{t-1} + \alpha \mu_{y,t} \epsilon_t \\ &b_t = b_{t-1} + \beta \frac{\mu_{y,t}}{l_{t-1}} \epsilon_t \\ &s_t = s_{t-m} + \gamma \mu_{y,t} \epsilon_t \end{aligned}$\|
+$\begin{aligned} &y_{t} = l_{t-1} b_{t-1} s_{t-m} (1 + \epsilon_t) \\ &l_t = l_{t-1} b_{t-1} (1 + \alpha \epsilon_t) \\ &b_t = b_{t-1} (1 + \beta \epsilon_t) \\ &s_t = s_{t-m} (1 + \gamma \epsilon_t) \end{aligned}$\|
+\|Multiplicative damped\|
+$\begin{aligned} &y_{t} = l_{t-1} b_{t-1}^\phi (1 + \epsilon_t) \\ &l_t = l_{t-1} b_{t-1}^\phi (1 + \alpha \epsilon_t) \\ &b_t = b_{t-1}^\phi (1 + \beta \epsilon_t) \end{aligned}$\|
+$\begin{aligned} &y_{t} = (l_{t-1} b_{t-1}^\phi + s_{t-m})(1 + \epsilon_t) \\ &l_t = l_{t-1} b_{t-1}^\phi + \alpha \mu_{y,t} \epsilon_t \\ &b_t = b_{t-1}^\phi + \beta \frac{\mu_{y,t}}{l_{t-1}} \epsilon_t \\ &s_t = s_{t-m} + \gamma \mu_{y,t} \epsilon_t \end{aligned}$
+\|
+$\begin{aligned} &y_{t} = l_{t-1} b_{t-1}^\phi s_{t-m} (1 + \epsilon_t) \\ &l_t = l_{t-1} b_{t-1}^\phi \left(1 + \alpha \epsilon_t\right) \\ &b_t = b_{t-1}^\phi \left(1 + \beta \epsilon_t\right) \\ &s_t = s_{t-m} \left(1 + \gamma \epsilon_t\right) \end{aligned}$\|
+
+From a statistical point of view, formulae in Tables 1 and 2 correspond
+to the “true models”, they explain the models underlying potential data,
+but when it comes to their construction and estimation, the $\epsilon_t$
+is substituted by the estimated $e_t$ (which is calculated differently
+depending on the error type), and time series components and smoothing
+parameters are also replaced by their estimates (e.g. $\hat \alpha$
+instead of $\alpha$). However, if the values of these models’ parameters
+were known, it would be possible to produce point forecasts and
+conditional h steps ahead expectations from these models.
+
+### Properties Holt’s linear trend method
+
+Holt’s linear trend method is a time series forecasting technique that
+uses exponential smoothing to estimate the level and trend components of
+a time series. The method has several properties, including:
+
+1. Additive model: Holt’s linear trend method assumes that the time
+ series can be decomposed into an additive model, where the observed
+ values are the sum of the level, trend, and error components.
+
+2. Smoothing parameters: The method uses two smoothing parameters, α
+ and β, to estimate the level and trend components of the time
+ series. These parameters control the amount of smoothing applied to
+ the level and trend components, respectively.
+
+3. Linear trend: Holt’s linear trend method assumes that the trend
+ component of the time series follows a straight line. This means
+ that the method is suitable for time series data that exhibit a
+ constant linear trend over time.
+
+4. Forecasting: The method uses the estimated level and trend
+ components to forecast future values of the time series. The
+ forecast for the next period is given by the sum of the level and
+ trend components.
+
+5. Optimization: The smoothing parameters α and β are estimated through
+ a process of optimization that minimizes the sum of squared errors
+ between the predicted and observed values. This involves iterating
+ over different values of the smoothing parameters until the optimal
+ values are found.
+
+6. Seasonality: Holt’s linear trend method can be extended to
+ incorporate seasonality components. This involves adding a seasonal
+ component to the model, which captures any systematic variations in
+ the time series that occur on a regular basis.
+
+Overall, Holt’s linear trend method is a powerful and widely used
+forecasting technique that can be used to generate accurate predictions
+for time series data with a constant linear trend. The method is easy to
+implement and can be extended to handle time series data with seasonal
+variations.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
+plt.style.use('grayscale') # fivethirtyeight grayscale classic
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#008080', # #212946
+ 'axes.facecolor': '#008080',
+ 'savefig.facecolor': '#008080',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#000000', #2A3459
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+### Read Data
+
+```python
+import pandas as pd
+
+df=pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/ads.csv")
+df.head()
+```
+
+| | Time | Ads |
+|-----|---------------------|--------|
+| 0 | 2017-09-13T00:00:00 | 80115 |
+| 1 | 2017-09-13T01:00:00 | 79885 |
+| 2 | 2017-09-13T02:00:00 | 89325 |
+| 3 | 2017-09-13T03:00:00 | 101930 |
+| 4 | 2017-09-13T04:00:00 | 121630 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|---------------------|--------|-----------|
+| 0 | 2017-09-13T00:00:00 | 80115 | 1 |
+| 1 | 2017-09-13T01:00:00 | 79885 | 1 |
+| 2 | 2017-09-13T02:00:00 | 89325 | 1 |
+| 3 | 2017-09-13T03:00:00 | 101930 | 1 |
+| 4 | 2017-09-13T04:00:00 | 121630 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y int64
+unique_id object
+dtype: object
+```
+
+We can see that our time variable `(ds)` is in an object format, we need
+to convert to a date format
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore Data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints a random series from the dataset and is useful for
+basic EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### The Augmented Dickey-Fuller Test
+
+An Augmented Dickey-Fuller (ADF) test is a type of statistical test that
+determines whether a unit root is present in time series data. Unit
+roots can cause unpredictable results in time series analysis. A null
+hypothesis is formed in the unit root test to determine how strongly
+time series data is affected by a trend. By accepting the null
+hypothesis, we accept the evidence that the time series data is not
+stationary. By rejecting the null hypothesis or accepting the
+alternative hypothesis, we accept the evidence that the time series data
+is generated by a stationary process. This process is also known as
+stationary trend. The values of the ADF test statistic are negative.
+Lower ADF values indicate a stronger rejection of the null hypothesis.
+
+Augmented Dickey-Fuller Test is a common statistical test used to test
+whether a given time series is stationary or not. We can achieve this by
+defining the null and alternate hypothesis.
+
+- Null Hypothesis: Time Series is non-stationary. It gives a
+ time-dependent trend.
+
+- Alternate Hypothesis: Time Series is stationary. In another term,
+ the series doesn’t depend on time.
+
+- ADF or t Statistic \< critical values: Reject the null hypothesis,
+ time series is stationary.
+
+- ADF or t Statistic \> critical values: Failed to reject the null
+ hypothesis, time series is non-stationary.
+
+```python
+from statsmodels.tsa.stattools import adfuller
+
+def Augmented_Dickey_Fuller_Test_func(series , column_name):
+ print (f'Dickey-Fuller test results for columns: {column_name}')
+ dftest = adfuller(series, autolag='AIC')
+ dfoutput = pd.Series(dftest[0:4], index=['Test Statistic','p-value','No Lags Used','Number of observations used'])
+ for key,value in dftest[4].items():
+ dfoutput['Critical Value (%s)'%key] = value
+ print (dfoutput)
+ if dftest[1] \<= 0.05:
+ print("Conclusion:====>")
+ print("Reject the null hypothesis")
+ print("The data is stationary")
+ else:
+ print("Conclusion:====>")
+ print("The null hypothesis cannot be rejected")
+ print("The data is not stationary")
+```
+
+
+```python
+Augmented_Dickey_Fuller_Test_func(df["y"],'Ads')
+```
+
+``` text
+Dickey-Fuller test results for columns: Ads
+Test Statistic -7.089634e+00
+p-value 4.444804e-10
+No Lags Used 9.000000e+00
+ ...
+Critical Value (1%) -3.462499e+00
+Critical Value (5%) -2.875675e+00
+Critical Value (10%) -2.574304e+00
+Length: 7, dtype: float64
+Conclusion:====>
+Reject the null hypothesis
+The data is stationary
+```
+
+### Autocorrelation plots
+
+**Autocorrelation Function**
+
+**Definition 1.** Let $\{x_t;1 ≤ t ≤ n\}$ be a time series sample of
+size n from $\{X_t\}$. 1. $\bar x = \sum_{t=1}^n \frac{x_t}{n}$ is
+called the sample mean of $\{X_t\}$. 2.
+$c_k =\sum_{t=1}^{n−k} (x_{t+k}- \bar x)(x_t−\bar x)/n$ is known as the
+sample autocovariance function of $\{X_t\}$. 3. $r_k = c_k /c_0$ is said
+to be the sample autocorrelation function of $\{X_t\}$.
+
+Note the following remarks about this definition:
+
+- Like most literature, this guide uses ACF to denote the sample
+ autocorrelation function as well as the autocorrelation function.
+ What is denoted by ACF can easily be identified in context.
+
+- Clearly c0 is the sample variance of $\{X_t\}$. Besides,
+ $r_0 = c_0/c_0 = 1$ and for any integer $k, |r_k| ≤ 1$.
+
+- When we compute the ACF of any sample series with a fixed length
+ $n$, we cannot put too much confidence in the values of $r_k$ for
+ large k’s, since fewer pairs of $(x_{t +k }, x_t )$ are available
+ for calculating $r_k$ as $k$ is large. One rule of thumb is not to
+ estimate $r_k$ for $k > n/3$, and another is $n ≥ 50, k ≤ n/4$. In
+ any case, it is always a good idea to be careful.
+
+- We also compute the ACF of a nonstationary time series sample by
+ Definition 1. In this case, however, the ACF or $r_k$ very slowly or
+ hardly tapers off as $k$ increases.
+
+- Plotting the ACF $(r_k)$ against lag $k$ is easy but very helpful in
+ analyzing time series sample. Such an ACF plot is known as a
+ correlogram.
+
+- If $\{X_t\}$ is stationary with $E(X_t)=0$ and $\rho_k =0$ for all
+ $k \neq 0$,thatis,itisa white noise series, then the sampling
+ distribution of $r_k$ is asymptotically normal with the mean 0 and
+ the variance of $1/n$. Hence, there is about 95% chance that $r_k$
+ falls in the interval $[−1.96/\sqrt{n}, 1.96/\sqrt{n}]$.
+
+Now we can give a summary that (1) if the time series plot of a time
+series clearly shows a trend or/and seasonality, it is surely
+nonstationary; (2) if the ACF $r_k$ very slowly or hardly tapers off as
+lag $k$ increases, the time series should also be nonstationary.
+
+**Partial autocorrelation**
+
+Let $\{X_t\}$ be a stationary time series with $E(X_t) = 0$. Here the
+assumption $E(X_t ) = 0$ is for conciseness only. If
+$E(X_t) = \mu \neq 0$, it is okay to replace $\{X_t\}$ by
+$\{X_t −\mu \}$. Now consider the linear regression (prediction) of
+$X_t$ on $\{X_{t−k+1:t−1}\}$ for any integer $k ≥ 2$. We use $\hat X_t$
+to denote this regression (prediction):
+$$\hat X_t =\alpha_1 X_{t−1}+···+\alpha_{k−1} X_{t−k+1}$$
+
+where $\{\alpha_1, · · · , \alpha_{k−1} \}$ satisfy
+
+$$\{\alpha_1, · · · , \alpha_{k−1} \}=\argmin_{\beta_1,···,\beta{k−1}} E[X_t −(\beta_1 X_{t−1} +···+\beta_{k−1} X_{t−k+1})]^2$$
+
+That is, $\{\alpha_1, · · · , \alpha_{k−1} \}$ are chosen by minimizing
+the mean squared error of prediction. Similarly, let $\hat X_{t −k}$
+denote the regression (prediction) of $X_{t −k}$ on
+$\{X_{t −k+1:t −1}\}$:
+
+$$\hat X_{t−k} =\eta_1 X_{t−1}+···+\eta_{k−1} X_{t−k+1}$$
+
+Note that if $\{X_t\}$ is stationary, then
+$\{\alpha_{1:k−1} \} = \{\eta_{1:k−1} \}$. Now let
+$\hat Z_{t−k} = X_{t−k} − \hat X_{t−k}$ and $\hat Z_t = X_t − \hat X_t$.
+Then $\hat Z_{t−k}$ is the residual of removing the effect of the
+intervening variables $\{X_{t−k+1:t−1} \}$ from $X_{t−k}$, and
+$\hat Z_t$ is the residual of removing the effect of
+$\{X_{t −k+1:t −1} \}$ from $X_t$ .
+
+**Definition 2.** The partial autocorrelation function (PACF) at lag $k$
+of a stationary time series $\{X_t \}$ with $E(X_t ) = 0$ is
+
+$$\phi_{11} = Corr(X_{t−1}, X_t ) = \frac{Cov(X_{t−1}, X_t )} {[Var(X_{t−1})Var(X_t)]^{1/2}} = \rho_1$$
+and
+
+$$\phi_{kk} = Corr(\hat Z_{t−k},\hat Z_t) = \frac{Cov(\hat Z_{t−k},\hat Z_t)} {[Var(\hat Z_{t −k} )Var(\hat Z_t )]^{1/2}}, \ k ≥ 2$$
+
+On the other hand, the following theorem paves the way to estimate the
+PACF of a stationary time series, and its proof can be seen in Fan and
+Yao (2003).
+
+**Theorem 1.** Let $\{X_t \}$ be a stationary time series with
+$E(X_t ) = 0$, and $\{a_{1k},··· ,a_{kk}\}$ satisfy
+
+$$\{a_{1k},··· ,a_{kk}\}= \argmin_{a_1 ,··· ,a_k} E(X_t − a_1 X_{t−1}−···−a_k X_{t−k})^2$$
+
+Then $\phi_{kk} =a_{kk}$ for $k≥1$.
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=30, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+plot_pacf(df["y"], lags=30, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+plt.show();
+```
+
+
+
+### Decomposition of the time series
+
+How to decompose a time series and why?
+
+In time series analysis to forecast new values, it is very important to
+know past data. More formally, we can say that it is very important to
+know the patterns that values follow over time. There can be many
+reasons that cause our forecast values to fall in the wrong direction.
+Basically, a time series consists of four components. The variation of
+those components causes the change in the pattern of the time series.
+These components are:
+
+- **Level:** This is the primary value that averages over time.
+- **Trend:** The trend is the value that causes increasing or
+ decreasing patterns in a time series.
+- **Seasonality:** This is a cyclical event that occurs in a time
+ series for a short time and causes short-term increasing or
+ decreasing patterns in a time series.
+- **Residual/Noise:** These are the random variations in the time
+ series.
+
+Combining these components over time leads to the formation of a time
+series. Most time series consist of level and noise/residual and trend
+or seasonality are optional values.
+
+If seasonality and trend are part of the time series, then there will be
+effects on the forecast value. As the pattern of the forecasted time
+series may be different from the previous time series.
+
+The combination of the components in time series can be of two types: \*
+Additive \* Multiplicative
+
+### Additive time series
+
+If the components of the time series are added to make the time series.
+Then the time series is called the additive time series. By
+visualization, we can say that the time series is additive if the
+increasing or decreasing pattern of the time series is similar
+throughout the series. The mathematical function of any additive time
+series can be represented by:
+$$y(t) = level + Trend + seasonality + noise$$
+
+### Multiplicative time series
+
+If the components of the time series are multiplicative together, then
+the time series is called a multiplicative time series. For
+visualization, if the time series is having exponential growth or
+decline with time, then the time series can be considered as the
+multiplicative time series. The mathematical function of the
+multiplicative time series can be represented as.
+
+$$y(t) = Level * Trend * seasonality * Noise$$
+
+### Additive
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+a = seasonal_decompose(df["y"], model = "additive", period=12)
+a.plot();
+```
+
+
+
+### Multiplicative
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+a = seasonal_decompose(df["y"], model = "Multiplicative", period=12)
+a.plot();
+```
+
+
+
+## Split the data into training and testing
+
+Let’s divide our data into sets 1. Data to train our `Holt Model`. 2.
+Data to test our model
+
+For the test data we will use the last 30 hours to test and evaluate the
+performance of our model.
+
+```python
+train = df[df.ds\<='2017-09-20 17:00:00']
+test = df[df.ds>'2017-09-20 17:00:00']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((186, 3), (30, 3))
+```
+
+Now let’s plot the training data and the test data.
+
+```python
+sns.lineplot(train,x="ds", y="y", label="Train", linestyle="--")
+sns.lineplot(test, x="ds", y="y", label="Test")
+plt.title("Ads watched (hourly data)");
+plt.show()
+```
+
+
+
+## Implementation of Holt Method with StatsForecast
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import Holt
+```
+
+### Instantiate Model
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/) by the
+master, Rob Hyndmann, can be useful for `season_length`.
+
+```python
+season_length = 24 # Hourly data
+horizon = len(test) # number of predictions
+
+models = [Holt(season_length=season_length, error_type="A", alias="Add"),
+ Holt(season_length=season_length, error_type="M", alias="Multi")]
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [pandas’
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models, freq='h')
+```
+
+### Fit the Model
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[Add,Multi])
+```
+
+Let’s see the results of our `Holt Model`. We can observe it with the
+following instruction:
+
+```python
+result=sf.fitted_[0,0].model_
+print(result.keys())
+print(result['fit'])
+```
+
+``` text
+dict_keys(['loglik', 'aic', 'bic', 'aicc', 'mse', 'amse', 'fit', 'residuals', 'components', 'm', 'nstate', 'fitted', 'states', 'par', 'sigma2', 'n_params', 'method', 'actual_residuals'])
+results(x=array([9.99900000e-01, 1.00000000e-04, 7.97982888e+04, 3.33340440e+02]), fn=4456.295090550272, nit=74, simplex=None)
+```
+
+Let us now visualize the fitted values of our models.
+
+As we can see, the result obtained above has an output in a dictionary,
+to extract each element from the dictionary we are going to use the
+`.get()` function to extract the element and then we are going to save
+it in a `pd.DataFrame()`.
+
+```python
+residual=pd.DataFrame(result.get("residuals"), columns=["residual Model"])
+residual
+```
+
+| | residual Model |
+|-----|----------------|
+| 0 | -16.629196 |
+| 1 | -563.340440 |
+| 2 | 9106.661223 |
+| ... | ... |
+| 183 | -268.370897 |
+| 184 | -1313.391081 |
+| 185 | -1428.364244 |
+
+```python
+import scipy.stats as stats
+
+fig, axs = plt.subplots(nrows=2, ncols=2)
+
+residual.plot(ax=axs[0,0])
+axs[0,0].set_title("Residuals");
+
+sns.distplot(residual, ax=axs[0,1]);
+axs[0,1].set_title("Density plot - Residual");
+
+stats.probplot(residual["residual Model"], dist="norm", plot=axs[1,0])
+axs[1,0].set_title('Plot Q-Q')
+
+plot_acf(residual, lags=35, ax=axs[1,1],color="fuchsia")
+axs[1,1].set_title("Autocorrelation");
+
+plt.show();
+```
+
+
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[90]` means that
+ the model expects the real value to be inside that interval 90% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min.
+
+```python
+Y_hat = sf.forecast(df=train, h=horizon, fitted=True)
+Y_hat
+```
+
+| | unique_id | ds | Add | Multi |
+|-----|-----------|---------------------|---------------|---------------|
+| 0 | 1 | 2017-09-20 18:00:00 | 139848.234375 | 141089.625000 |
+| 1 | 1 | 2017-09-20 19:00:00 | 140181.328125 | 142664.000000 |
+| 2 | 1 | 2017-09-20 20:00:00 | 140514.406250 | 144238.359375 |
+| ... | ... | ... | ... | ... |
+| 27 | 1 | 2017-09-21 21:00:00 | 148841.671875 | 183597.453125 |
+| 28 | 1 | 2017-09-21 22:00:00 | 149174.750000 | 185171.812500 |
+| 29 | 1 | 2017-09-21 23:00:00 | 149507.843750 | 186746.187500 |
+
+```python
+values=sf.forecast_fitted_values()
+values.head()
+```
+
+| | unique_id | ds | y | Add | Multi |
+|-----|-----------|---------------------|----------|---------------|---------------|
+| 0 | 1 | 2017-09-13 00:00:00 | 80115.0 | 80131.632812 | 79287.125000 |
+| 1 | 1 | 2017-09-13 01:00:00 | 79885.0 | 80448.343750 | 81712.710938 |
+| 2 | 1 | 2017-09-13 02:00:00 | 89325.0 | 80218.335938 | 81482.796875 |
+| 3 | 1 | 2017-09-13 03:00:00 | 101930.0 | 89658.281250 | 90922.609375 |
+| 4 | 1 | 2017-09-13 04:00:00 | 121630.0 | 102264.195312 | 103528.398438 |
+
+```python
+StatsForecast.plot(values)
+```
+
+
+
+Adding 95% confidence interval with the forecast method
+
+```python
+sf.forecast(df=train, h=horizon, level=[95])
+```
+
+| | unique_id | ds | Add | Add-lo-95 | Add-hi-95 | Multi | Multi-lo-95 | Multi-hi-95 |
+|-----|-----------|---------------------|---------------|---------------|---------------|---------------|---------------|---------------|
+| 0 | 1 | 2017-09-20 18:00:00 | 139848.234375 | 116559.250000 | 163137.218750 | 141089.625000 | 113501.140625 | 168678.125000 |
+| 1 | 1 | 2017-09-20 19:00:00 | 140181.328125 | 107245.734375 | 173116.906250 | 142664.000000 | 103333.265625 | 181994.718750 |
+| 2 | 1 | 2017-09-20 20:00:00 | 140514.406250 | 100175.375000 | 180853.453125 | 144238.359375 | 95679.804688 | 192796.921875 |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| 27 | 1 | 2017-09-21 21:00:00 | 148841.671875 | 25453.445312 | 272229.875000 | 183597.453125 | 4082.392090 | 363112.531250 |
+| 28 | 1 | 2017-09-21 22:00:00 | 149174.750000 | 23596.246094 | 274753.250000 | 185171.812500 | 1151.084961 | 369192.562500 |
+| 29 | 1 | 2017-09-21 23:00:00 | 149507.843750 | 21776.173828 | 277239.531250 | 186746.187500 | -1776.010254 | 375268.375000 |
+
+```python
+sf.plot(train, Y_hat)
+```
+
+
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[95]` means that
+ the model expects the real value to be inside that interval 95% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+sf.predict(h=horizon)
+```
+
+| | unique_id | ds | Add | Multi |
+|-----|-----------|---------------------|---------------|---------------|
+| 0 | 1 | 2017-09-20 18:00:00 | 139848.234375 | 141089.625000 |
+| 1 | 1 | 2017-09-20 19:00:00 | 140181.328125 | 142664.000000 |
+| 2 | 1 | 2017-09-20 20:00:00 | 140514.406250 | 144238.359375 |
+| ... | ... | ... | ... | ... |
+| 27 | 1 | 2017-09-21 21:00:00 | 148841.671875 | 183597.453125 |
+| 28 | 1 | 2017-09-21 22:00:00 | 149174.750000 | 185171.812500 |
+| 29 | 1 | 2017-09-21 23:00:00 | 149507.843750 | 186746.187500 |
+
+```python
+forecast_df = sf.predict(h=horizon, level=[80,95])
+forecast_df
+```
+
+| | unique_id | ds | Add | Add-lo-95 | Add-lo-80 | Add-hi-80 | Add-hi-95 | Multi | Multi-lo-95 | Multi-lo-80 | Multi-hi-80 | Multi-hi-95 |
+|-----|-----------|---------------------|---------------|---------------|---------------|---------------|---------------|---------------|---------------|---------------|---------------|---------------|
+| 0 | 1 | 2017-09-20 18:00:00 | 139848.234375 | 116559.250000 | 124620.390625 | 155076.078125 | 163137.218750 | 141089.625000 | 113501.140625 | 123050.484375 | 159128.781250 | 168678.125000 |
+| 1 | 1 | 2017-09-20 19:00:00 | 140181.328125 | 107245.734375 | 118645.898438 | 161716.750000 | 173116.906250 | 142664.000000 | 103333.265625 | 116947.015625 | 168380.984375 | 181994.718750 |
+| 2 | 1 | 2017-09-20 20:00:00 | 140514.406250 | 100175.375000 | 114138.132812 | 166890.687500 | 180853.453125 | 144238.359375 | 95679.804688 | 112487.625000 | 175989.093750 | 192796.921875 |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| 27 | 1 | 2017-09-21 21:00:00 | 148841.671875 | 25453.445312 | 68162.445312 | 229520.890625 | 272229.875000 | 183597.453125 | 4082.392090 | 66218.867188 | 300976.031250 | 363112.531250 |
+| 28 | 1 | 2017-09-21 22:00:00 | 149174.750000 | 23596.246094 | 67063.382812 | 231286.125000 | 274753.250000 | 185171.812500 | 1151.084961 | 64847.128906 | 305496.500000 | 369192.562500 |
+| 29 | 1 | 2017-09-21 23:00:00 | 149507.843750 | 21776.173828 | 65988.593750 | 233027.093750 | 277239.531250 | 186746.187500 | -1776.010254 | 63478.144531 | 310014.218750 | 375268.375000 |
+
+```python
+sf.plot(train, forecast_df, level=[80, 95])
+```
+
+
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=)`, forecasting every second months
+`(step_size=12)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 30 hours ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=df,
+ h=horizon,
+ step_size=30,
+ n_windows=3)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` series identifier.
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the `n_windows`.
+- `y:` true value
+- `model:` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df
+```
+
+| | unique_id | ds | cutoff | y | Add | Multi |
+|-----|-----------|---------------------|---------------------|----------|---------------|---------------|
+| 0 | 1 | 2017-09-18 06:00:00 | 2017-09-18 05:00:00 | 99440.0 | 111573.328125 | 112874.039062 |
+| 1 | 1 | 2017-09-18 07:00:00 | 2017-09-18 05:00:00 | 97655.0 | 111820.390625 | 114421.679688 |
+| 2 | 1 | 2017-09-18 08:00:00 | 2017-09-18 05:00:00 | 97655.0 | 112067.453125 | 115969.320312 |
+| ... | ... | ... | ... | ... | ... | ... |
+| 87 | 1 | 2017-09-21 21:00:00 | 2017-09-20 17:00:00 | 103080.0 | 148841.671875 | 183597.453125 |
+| 88 | 1 | 2017-09-21 22:00:00 | 2017-09-20 17:00:00 | 95155.0 | 149174.750000 | 185171.812500 |
+| 89 | 1 | 2017-09-21 23:00:00 | 2017-09-20 17:00:00 | 80285.0 | 149507.843750 | 186746.187500 |
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=season_length), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | Add | Multi |
+|-----|-----------|--------|--------------|--------------|
+| 0 | 1 | mae | 30905.751042 | 48210.098958 |
+| 1 | 1 | mape | 0.336201 | 0.491980 |
+| 2 | 1 | mase | 3.818464 | 5.956449 |
+| 3 | 1 | rmse | 38929.522482 | 54653.132768 |
+| 4 | 1 | smape | 0.129755 | 0.182024 |
+
+## References
+
+1. [Changquan Huang • Alla Petukhina. Springer series (2022). Applied
+ Time Series Analysis and Forecasting with
+ Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2)
+2. Ivan Svetunkov. [Forecasting and Analytics with the Augmented
+ Dynamic Adaptive Model (ADAM)](https://openforecast.org/adam/)
+3. [James D. Hamilton. Time Series Analysis Princeton University Press,
+ Princeton, New Jersey, 1st Edition,
+ 1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Nixtla Holt API](../../src/core/models.html#holt)
+5. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+6. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+7. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/models/holtwinters.html.mdx b/statsforecast/docs/models/holtwinters.html.mdx
new file mode 100644
index 00000000..e0fd1cf9
--- /dev/null
+++ b/statsforecast/docs/models/holtwinters.html.mdx
@@ -0,0 +1,1025 @@
+---
+title: Holt Winters Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `Holt Winters Model` with
+> `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation` in other.
+
+The text in this article is largely taken from: 1. [Changquan Huang •
+Alla Petukhina. Springer series (2022). Applied Time Series Analysis and
+Forecasting with
+Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2) 2.
+Ivan Svetunkov. [Forecasting and Analytics with the Augmented Dynamic
+Adaptive Model (ADAM)](https://openforecast.org/adam/) 3. [James D.
+Hamilton. Time Series Analysis Princeton University Press, Princeton,
+New Jersey, 1st Edition,
+1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+Principles and Practice (3rd ed)”](https://otexts.com/fpp3/tscv.html).
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [Holt-Winters Model](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of Holt-Winters with StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+The Holt-Winter model, also known as the triple exponential smoothing
+method, is a forecasting technique widely used in time series analysis.
+It was developed by Charles Holt and Peter Winters in 1960 as an
+improvement on Holt’s double exponential smoothing method.
+
+The Holt-Winter model is used to predict future values of a time series
+that exhibits a trend and seasonality. The model uses three smoothing
+parameters, one for estimating the trend, another for estimating the
+level or base level of the time series, and another for estimating
+seasonality. These parameters are called α, β and γ, respectively.
+
+The Holt-Winter model is an extension of Holt’s double exponential
+smoothing method, which uses only two smoothing parameters to estimate
+the trend and base level of the time series. The Holt-Winter model
+improves the accuracy of the forecasts by adding a third smoothing
+parameter for seasonality.
+
+One of the main advantages of the Holt-Winter model is that it is easy
+to implement and does not require a large amount of historical data to
+generate accurate predictions. Furthermore, the model is highly
+adaptable and can be customized to fit a wide variety of time series
+with seasonality.
+
+However, the Holt-Winter model has some limitations. For example, the
+model assumes that the time series is stationary and that seasonality is
+constant. If the time series is not stationary or has non-constant
+seasonality, the Holt-Winter model may not be the most appropriate.
+
+In general, the Holt-Winter model is a useful and widely used technique
+in time series analysis, especially when the series is expected to
+exhibit a constant trend and seasonality.
+
+## Holt-Winters Method
+
+The Holt-Winters seasonal method comprises the forecast equation and
+three smoothing equations — one for the level $\ell_{t}$, one for the
+trend $b_t$, and one for the seasonal component $s_t$ , with
+corresponding smoothing parameters $\alpha$ , $\beta^*$ and $\gamma$. We
+use $m$ to denote the period of the seasonality, i.e., the number of
+seasons in a year. For example, for quarterly data $m=4$, and for
+monthly data $m=12$.
+
+There are two variations to this method that differ in the nature of the
+seasonal component. The additive method is preferred when the seasonal
+variations are roughly constant through the series, while the
+multiplicative method is preferred when the seasonal variations are
+changing proportional to the level of the series. With the additive
+method, the seasonal component is expressed in absolute terms in the
+scale of the observed series, and in the level equation the series is
+seasonally adjusted by subtracting the seasonal component. Within each
+year, the seasonal component will add up to approximately zero. With the
+multiplicative method, the seasonal component is expressed in relative
+terms (percentages), and the series is seasonally adjusted by dividing
+through by the seasonal component. Within each year, the seasonal
+component will sum up to approximately $m$.
+
+### Holt-Winters’ additive method
+
+Holt-Winters’ additive method is a time series forecasting technique
+that extends the Holt-Winters’ method by incorporating an additive
+seasonality component. It is suitable for time series data that exhibit
+a seasonal pattern that changes over time.
+
+The Holt-Winters’ additive method uses three smoothing parameters -
+alpha (α), beta (β), and gamma (γ) - to estimate the level, trend, and
+seasonal components of the time series. The alpha parameter controls the
+smoothing of the level component, the beta parameter controls the
+smoothing of the trend component, and the gamma parameter controls the
+smoothing of the additive seasonal component.
+
+The forecasting process involves three steps: first, the level, trend,
+and seasonal components are estimated using the smoothing parameters and
+the historical data; second, these components are used to forecast
+future values of the time series; and third, the forecasted values are
+adjusted for the seasonal component using an additive factor.
+
+One of the advantages of Holt-Winters’ additive method is that it can
+handle time series data with an additive seasonality component, which is
+common in many real-world applications. The method is also easy to
+implement and can be extended to handle time series data with changing
+seasonal patterns.
+
+However, the method has some limitations. It assumes that the
+seasonality pattern is additive, which may not be the case for all time
+series. Additionally, the method requires a sufficient amount of
+historical data to accurately estimate the smoothing parameters and the
+seasonal component.
+
+Overall, Holt-Winters’ additive method is a powerful and widely used
+forecasting technique that can be used to generate accurate predictions
+for time series data with an additive seasonality component. The method
+is easy to implement and can be extended to handle time series data with
+changing seasonal patterns.
+
+The component form for the additive method is:
+
+where $k$ is the integer part of $(h-1)/m$, which ensures that the
+estimates of the seasonal indices used for forecasting come from the
+final year of the sample. The level equation shows a weighted average
+between the seasonally adjusted observation $(y_{t} - s_{t-m})$ and the
+non-seasonal forecast $(\ell_{t-1}+b_{t-1})$ for time $t$. The trend
+equation is identical to Holt’s linear method. The seasonal equation
+shows a weighted average between the current seasonal index,
+$(y_{t}-\ell_{t-1}-b_{t-1})$, and the seasonal index of the same season
+last year (i.e., $m$ time periods ago).
+
+The equation for the seasonal component is often expressed as
+
+$$s_{t} = \gamma^* (y_{t}-\ell_{t})+ (1-\gamma^*)s_{t-m}.$$
+
+If we substitute $\ell_{t}$ from the smoothing equation for the level of
+the component form above, we get
+
+$$s_{t} = \gamma^*(1-\alpha) (y_{t}-\ell_{t-1}-b_{t-1})+ [1-\gamma^*(1-\alpha)]s_{t-m},$$
+
+which is identical to the smoothing equation for the seasonal component
+we specify here, with $\gamma=\gamma^*(1-\alpha)$. The usual parameter
+restriction is $0\le\gamma^*\le1$, which translates to
+$0\le\gamma\le 1-\alpha$.
+
+### Holt-Winters’ multiplicative method
+
+The Holt-Winters’ multiplicative method uses three smoothing
+parameters - alpha (α), beta (β), and gamma (γ) - to estimate the level,
+trend, and seasonal components of the time series. The alpha parameter
+controls the smoothing of the level component, the beta parameter
+controls the smoothing of the trend component, and the gamma parameter
+controls the smoothing of the multiplicative seasonal component.
+
+The forecasting process involves three steps: first, the level, trend,
+and seasonal components are estimated using the smoothing parameters and
+the historical data; second, these components are used to forecast
+future values of the time series; and third, the forecasted values are
+adjusted for the seasonal component using a multiplicative factor.
+
+One of the advantages of Holt-Winters’ multiplicative method is that it
+can handle time series data with a multiplicative seasonality component,
+which is common in many real-world applications. The method is also easy
+to implement and can be extended to handle time series data with
+changing seasonal patterns.
+
+However, the method has some limitations. It assumes that the
+seasonality pattern is multiplicative, which may not be the case for all
+time series. Additionally, the method requires a sufficient amount of
+historical data to accurately estimate the smoothing parameters and the
+seasonal component.
+
+Overall, Holt-Winters’ multiplicative method is a powerful and widely
+used forecasting technique that can be used to generate accurate
+predictions for time series data with a multiplicative seasonality
+component. The method is easy to implement and can be extended to handle
+time series data with changing seasonal patterns.
+
+In the multiplicative version, the seasonality averages to one. Use the
+multiplicative method if the seasonal variation increases with the
+level.
+
+### Mathematical models in the ETS taxonomy
+
+I hope that it becomes more apparent to the reader how the ETS framework
+is built upon the idea of time series decomposition. By introducing
+different components, defining their types, and adding the equations for
+their update, we can construct models that would work better in
+capturing the key features of the time series. But we should also
+consider the potential change in components over time. The “transition”
+or “state” equations are supposed to reflect this change: they explain
+how the level, trend or seasonal components evolve.
+
+As discussed in Section 2.2, given different types of components and
+their interactions, we end up with 30 models in the taxonomy. Tables 1
+and 2 summarise mathematically all 30 ETS models shown graphically on
+Figures 1 and 2, presenting formulae for measurement and transition
+equations.
+
+Table 1: Additive error ETS models
+
+| | Nonseasonal | Additive | Multiplicative |
+|--------|--------------------|--------------------|-------------------------|
+| No trend | $\begin{aligned} &y_{t} = l_{t-1} + \epsilon_t \\ &l_t = l_{t-1} + \alpha \epsilon_t \end{aligned}$ | $\begin{aligned} &y_{t} = l_{t-1} + s_{t-m} + \epsilon_t \\ &l_t = l_{t-1} + \alpha \epsilon_t \\ &s_t = s_{t-m} + \gamma \epsilon_t \end{aligned}$ | $\begin{aligned} &y_{t} = l_{t-1} s_{t-m} + \epsilon_t \\ &l_t = l_{t-1} + \alpha \frac{\epsilon_t}{s_{t-m}} \\ &s_t = s_{t-m} + \gamma \frac{\epsilon_t}{l_{t-1}} \end{aligned}$ |
+| Additive | $\begin{aligned} &y_{t} = l_{t-1} + b_{t-1} + \epsilon_t \\ &l_t = l_{t-1} + b_{t-1} + \alpha \epsilon_t \\ &b_t = b_{t-1} + \beta \epsilon_t \end{aligned}$ | $\begin{aligned} &y_{t} = l_{t-1} + b_{t-1} + s_{t-m} + \epsilon_t \\ &l_t = l_{t-1} + b_{t-1} + \alpha \epsilon_t \\ &b_t = b_{t-1} + \beta \epsilon_t \\ &s_t = s_{t-m} + \gamma \epsilon_t \end{aligned}$ | $\begin{aligned} &y_{t} = (l_{t-1} + b_{t-1}) s_{t-m} + \epsilon_t \\ &l_t = l_{t-1} + b_{t-1} + \alpha \frac{\epsilon_t}{s_{t-m}} \\ &b_t = b_{t-1} + \beta \frac{\epsilon_t}{s_{t-m}} \\ &s_t = s_{t-m} + \gamma \frac{\epsilon_t}{l_{t-1} + b_{t-1}} \end{aligned}$ |
+| Additive damped | $\begin{aligned} &y_{t} = l_{t-1} + \phi b_{t-1} + \epsilon_t \\ &l_t = l_{t-1} + \phi b_{t-1} + \alpha \epsilon_t \\ &b_t = \phi b_{t-1} + \beta \epsilon_t \end{aligned}$ | $\begin{aligned} &y_{t} = l_{t-1} + \phi b_{t-1} + s_{t-m} + \epsilon_t \\ &l_t = l_{t-1} + \phi b_{t-1} + \alpha \epsilon_t \\ &b_t = \phi b_{t-1} + \beta \epsilon_t \\ &s_t = s_{t-m} + \gamma \epsilon_t \end{aligned}$ | $\begin{aligned} &y_{t} = (l_{t-1} + \phi b_{t-1}) s_{t-m} + \epsilon_t \\ &l_t = l_{t-1} + \phi b_{t-1} + \alpha \frac{\epsilon_t}{s_{t-m}} \\ &b_t = \phi b_{t-1} + \beta \frac{\epsilon_t}{s_{t-m}} \\ &s_t = s_{t-m} + \gamma \frac{\epsilon_t}{l_{t-1} + \phi b_{t-1}} \end{aligned}$ |
+| Multiplicative | $\begin{aligned} &y_{t} = l_{t-1} b_{t-1} + \epsilon_t \\ &l_t = l_{t-1} b_{t-1} + \alpha \epsilon_t \\ &b_t = b_{t-1} + \beta \frac{\epsilon_t}{l_{t-1}} \end{aligned}$ | $\begin{aligned} &y_{t} = l_{t-1} b_{t-1} + s_{t-m} + \epsilon_t \\ &l_t = l_{t-1} b_{t-1} + \alpha \epsilon_t \\ &b_t = b_{t-1} + \beta \frac{\epsilon_t}{l_{t-1}} \\ &s_t = s_{t-m} + \gamma \epsilon_t \end{aligned}$ | $\begin{aligned} &y_{t} = l_{t-1} b_{t-1} s_{t-m} + \epsilon_t \\ &l_t = l_{t-1} b_{t-1} + \alpha \frac{\epsilon_t}{s_{t-m}} \\ &b_t = b_{t-1} + \beta \frac{\epsilon_t}{l_{t-1}s_{t-m}} \\ &s_t = s_{t-m} + \gamma \frac{\epsilon_t}{l_{t-1} b_{t-1}} \end{aligned}$ |
+| Multiplicative damped | $\begin{aligned} &y_{t} = l_{t-1} b_{t-1}^\phi + \epsilon_t \\ &l_t = l_{t-1} b_{t-1}^\phi + \alpha \epsilon_t \\ &b_t = b_{t-1}^\phi + \beta \frac{\epsilon_t}{l_{t-1}} \end{aligned}$ | $\begin{aligned} &y_{t} = l_{t-1} b_{t-1}^\phi + s_{t-m} + \epsilon_t \\ &l_t = l_{t-1} b_{t-1}^\phi + \alpha \epsilon_t \\ &b_t = b_{t-1}^\phi + \beta \frac{\epsilon_t}{l_{t-1}} \\ &s_t = s_{t-m} + \gamma \epsilon_t \end{aligned}$ | $\begin{aligned} &y_{t} = l_{t-1} b_{t-1}^\phi s_{t-m} + \epsilon_t \\ &l_t = l_{t-1} b_{t-1}^\phi + \alpha \frac{\epsilon_t}{s_{t-m}} \\ &b_t = b_{t-1}^\phi + \beta \frac{\epsilon_t}{l_{t-1}s_{t-m}} \\ &s_t = s_{t-m} + \gamma \frac{\epsilon_t}{l_{t-1} b_{t-1}} \end{aligned}$ |
+
+Table 2: Multiplicative error ETS models
+
+| | Nonseasonal | Additive | Multiplicative |
+|-----------|----------------------|-----------------|-----------------------|
+| No trend | $\begin{aligned} &y_{t} = l_{t-1}(1 + \epsilon_t) \\ &l_t = l_{t-1}(1 + \alpha \epsilon_t) \end{aligned}$ | $\begin{aligned} &y_{t} = (l_{t-1} + s_{t-m})(1 + \epsilon_t) \\ &l_t = l_{t-1} + \alpha \mu_{y,t} \epsilon_t \\ &s_t = s_{t-m} + \gamma \mu_{y,t} \epsilon_t \end{aligned}$ | $\begin{aligned} &y_{t} = l_{t-1} s_{t-m}(1 + \epsilon_t) \\ &l_t = l_{t-1}(1 + \alpha \epsilon_t) \\ &s_t = s_{t-m}(1 + \gamma \epsilon_t) \end{aligned}$ |
+| Additive | $\begin{aligned} &y_{t} = (l_{t-1} + b_{t-1})(1 + \epsilon_t) \\ &l_t = (l_{t-1} + b_{t-1})(1 + \alpha \epsilon_t) \\ &b_t = b_{t-1} + \beta \mu_{y,t} \epsilon_t \end{aligned}$ | $\begin{aligned} &y_{t} = (l_{t-1} + b_{t-1} + s_{t-m})(1 + \epsilon_t) \\ &l_t = l_{t-1} + b_{t-1} + \alpha \mu_{y,t} \epsilon_t \\ &b_t = b_{t-1} + \beta \mu_{y,t} \epsilon_t \\ &s_t = s_{t-m} + \gamma \mu_{y,t} \epsilon_t \end{aligned}$ | $\begin{aligned} &y_{t} = (l_{t-1} + b_{t-1}) s_{t-m}(1 + \epsilon_t) \\ &l_t = (l_{t-1} + b_{t-1})(1 + \alpha \epsilon_t) \\ &b_t = b_{t-1} + \beta (l_{t-1} + b_{t-1}) \epsilon_t \\ &s_t = s_{t-m} (1 + \gamma \epsilon_t) \end{aligned}$ |
+| Additive damped | $\begin{aligned} &y_{t} = (l_{t-1} + \phi b_{t-1})(1 + \epsilon_t) \\ &l_t = (l_{t-1} + \phi b_{t-1})(1 + \alpha \epsilon_t) \\ &b_t = \phi b_{t-1} + \beta \mu_{y,t} \epsilon_t \end{aligned}$ | $\begin{aligned} &y_{t} = (l_{t-1} + \phi b_{t-1} + s_{t-m})(1 + \epsilon_t) \\ &l_t = l_{t-1} + \phi b_{t-1} + \alpha \mu_{y,t} \epsilon_t \\ &b_t = \phi b_{t-1} + \beta \mu_{y,t} \epsilon_t \\ &s_t = s_{t-m} + \gamma \mu_{y,t} \epsilon_t \end{aligned}$ | $\begin{aligned} &y_{t} = (l_{t-1} + \phi b_{t-1}) s_{t-m}(1 + \epsilon_t) \\ &l_t = l_{t-1} + \phi b_{t-1} (1 + \alpha \epsilon_t) \\ &b_t = \phi b_{t-1} + \beta (l_{t-1} + \phi b_{t-1}) \epsilon_t \\ &s_t = s_{t-m}(1 + \gamma \epsilon_t) \end{aligned}$ |
+| Multiplicative | $\begin{aligned} &y_{t} = l_{t-1} b_{t-1} (1 + \epsilon_t) \\ &l_t = l_{t-1} b_{t-1} (1 + \alpha \epsilon_t) \\ &b_t = b_{t-1} (1 + \beta \epsilon_t) \end{aligned}$ | $\begin{aligned} &y_{t} = (l_{t-1} b_{t-1} + s_{t-m})(1 + \epsilon_t) \\ &l_t = l_{t-1} b_{t-1} + \alpha \mu_{y,t} \epsilon_t \\ &b_t = b_{t-1} + \beta \frac{\mu_{y,t}}{l_{t-1}} \epsilon_t \\ &s_t = s_{t-m} + \gamma \mu_{y,t} \epsilon_t \end{aligned}$ | $\begin{aligned} &y_{t} = l_{t-1} b_{t-1} s_{t-m} (1 + \epsilon_t) \\ &l_t = l_{t-1} b_{t-1} (1 + \alpha \epsilon_t) \\ &b_t = b_{t-1} (1 + \beta \epsilon_t) \\ &s_t = s_{t-m} (1 + \gamma \epsilon_t) \end{aligned}$ |
+| Multiplicative damped | $\begin{aligned} &y_{t} = l_{t-1} b_{t-1}^\phi (1 + \epsilon_t) \\ &l_t = l_{t-1} b_{t-1}^\phi (1 + \alpha \epsilon_t) \\ &b_t = b_{t-1}^\phi (1 + \beta \epsilon_t) \end{aligned}$ | $\begin{aligned} &y_{t} = (l_{t-1} b_{t-1}^\phi + s_{t-m})(1 + \epsilon_t) \\ &l_t = l_{t-1} b_{t-1}^\phi + \alpha \mu_{y,t} \epsilon_t \\ &b_t = b_{t-1}^\phi + \beta \frac{\mu_{y,t}}{l_{t-1}} \epsilon_t \\ &s_t = s_{t-m} + \gamma \mu_{y,t} \epsilon_t \end{aligned}$ | $\begin{aligned} &y_{t} = l_{t-1} b_{t-1}^\phi s_{t-m} (1 + \epsilon_t) \\ &l_t = l_{t-1} b_{t-1}^\phi \left(1 + \alpha \epsilon_t\right) \\ &b_t = b_{t-1}^\phi \left(1 + \beta \epsilon_t\right) \\ &s_t = s_{t-m} \left(1 + \gamma \epsilon_t\right) \end{aligned}$ |
+
+From a statistical point of view, formulae in Tables 1 and 2 correspond
+to the “true models”, they explain the models underlying potential data,
+but when it comes to their construction and estimation, the $\epsilon_t$
+is substituted by the estimated $e_t$ (which is calculated differently
+depending on the error type), and time series components and smoothing
+parameters are also replaced by their estimates (e.g. $\hat \alpha$
+instead of $\alpha$). However, if the values of these models’ parameters
+were known, it would be possible to produce point forecasts and
+conditional h steps ahead expectations from these models.
+
+### Model selection
+
+A great advantage of the `Holt Winters` statistical framework is that
+information criteria can be used for model selection. The `AIC, AIC_c`
+and `BIC`, can be used here to determine which of the `Holt Winters`
+models is most appropriate for a given time series.
+
+For `Holt Winters` models, Akaike’s Information Criterion (`AIC)` is
+defined as
+
+$$\text{AIC} = -2\log(L) + 2k,$$
+
+where $L$ is the likelihood of the model and $k$ is the total number of
+parameters and initial states that have been estimated (including the
+residual variance).
+
+The `AIC` corrected for small sample bias `(AIC_c)` is defined as
+
+$$AIC_c = AIC + \frac{2k(k+1)}{T-k-1}$$
+
+and the Bayesian Information Criterion `(BIC)` is
+
+$$\text{BIC} = \text{AIC} + k[\log(T)-2]$$
+
+Three of the combinations of (Error, Trend, Seasonal) can lead to
+numerical difficulties. Specifically, the models that can cause such
+instabilities are `ETS(A,N,M), ETS(A,A,M)`, and `ETS(A,Ad,M)`, due to
+division by values potentially close to zero in the state equations. We
+normally do not consider these particular combinations when selecting a
+model.
+
+Models with multiplicative errors are useful when the data are strictly
+positive, but are not numerically stable when the data contain zeros or
+negative values. Therefore, multiplicative error models will not be
+considered if the time series is not strictly positive. In that case,
+only the six fully additive models will be applied.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
+plt.style.use('grayscale') # fivethirtyeight grayscale classic
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#008080', # #212946
+ 'axes.facecolor': '#008080',
+ 'savefig.facecolor': '#008080',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#000000', #2A3459
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+### Read Data
+
+```python
+df=pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/ads.csv")
+df.head()
+```
+
+| | Time | Ads |
+|-----|---------------------|--------|
+| 0 | 2017-09-13T00:00:00 | 80115 |
+| 1 | 2017-09-13T01:00:00 | 79885 |
+| 2 | 2017-09-13T02:00:00 | 89325 |
+| 3 | 2017-09-13T03:00:00 | 101930 |
+| 4 | 2017-09-13T04:00:00 | 121630 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|---------------------|--------|-----------|
+| 0 | 2017-09-13T00:00:00 | 80115 | 1 |
+| 1 | 2017-09-13T01:00:00 | 79885 | 1 |
+| 2 | 2017-09-13T02:00:00 | 89325 | 1 |
+| 3 | 2017-09-13T03:00:00 | 101930 | 1 |
+| 4 | 2017-09-13T04:00:00 | 121630 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y int64
+unique_id object
+dtype: object
+```
+
+We can see that our time variable `(ds)` is in an object format, we need
+to convert to a date format
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore Data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints a random series from the dataset and is useful for
+basic EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### The Augmented Dickey-Fuller Test
+
+An Augmented Dickey-Fuller (ADF) test is a type of statistical test that
+determines whether a unit root is present in time series data. Unit
+roots can cause unpredictable results in time series analysis. A null
+hypothesis is formed in the unit root test to determine how strongly
+time series data is affected by a trend. By accepting the null
+hypothesis, we accept the evidence that the time series data is not
+stationary. By rejecting the null hypothesis or accepting the
+alternative hypothesis, we accept the evidence that the time series data
+is generated by a stationary process. This process is also known as
+stationary trend. The values of the ADF test statistic are negative.
+Lower ADF values indicate a stronger rejection of the null hypothesis.
+
+Augmented Dickey-Fuller Test is a common statistical test used to test
+whether a given time series is stationary or not. We can achieve this by
+defining the null and alternate hypothesis.
+
+- Null Hypothesis: Time Series is non-stationary. It gives a
+ time-dependent trend.
+
+- Alternate Hypothesis: Time Series is stationary. In another term,
+ the series doesn’t depend on time.
+
+- ADF or t Statistic \< critical values: Reject the null hypothesis,
+ time series is stationary.
+
+- ADF or t Statistic \> critical values: Failed to reject the null
+ hypothesis, time series is non-stationary.
+
+```python
+from statsmodels.tsa.stattools import adfuller
+
+def Augmented_Dickey_Fuller_Test_func(series , column_name):
+ print (f'Dickey-Fuller test results for columns: {column_name}')
+ dftest = adfuller(series, autolag='AIC')
+ dfoutput = pd.Series(dftest[0:4], index=['Test Statistic','p-value','No Lags Used','Number of observations used'])
+ for key,value in dftest[4].items():
+ dfoutput['Critical Value (%s)'%key] = value
+ print (dfoutput)
+ if dftest[1] \<= 0.05:
+ print("Conclusion:====>")
+ print("Reject the null hypothesis")
+ print("The data is stationary")
+ else:
+ print("Conclusion:====>")
+ print("The null hypothesis cannot be rejected")
+ print("The data is not stationary")
+```
+
+
+```python
+Augmented_Dickey_Fuller_Test_func(df["y"],'Ads')
+```
+
+``` text
+Dickey-Fuller test results for columns: Ads
+Test Statistic -7.089634e+00
+p-value 4.444804e-10
+No Lags Used 9.000000e+00
+ ...
+Critical Value (1%) -3.462499e+00
+Critical Value (5%) -2.875675e+00
+Critical Value (10%) -2.574304e+00
+Length: 7, dtype: float64
+Conclusion:====>
+Reject the null hypothesis
+The data is stationary
+```
+
+### Autocorrelation plots
+
+**Autocorrelation Function**
+
+**Definition 1.** Let $\{x_t;1 ≤ t ≤ n\}$ be a time series sample of
+size n from $\{X_t\}$. 1. $\bar x = \sum_{t=1}^n \frac{x_t}{n}$ is
+called the sample mean of $\{X_t\}$. 2.
+$c_k =\sum_{t=1}^{n−k} (x_{t+k}- \bar x)(x_t−\bar x)/n$ is known as the
+sample autocovariance function of $\{X_t\}$. 3. $r_k = c_k /c_0$ is said
+to be the sample autocorrelation function of $\{X_t\}$.
+
+Note the following remarks about this definition:
+
+- Like most literature, this guide uses ACF to denote the sample
+ autocorrelation function as well as the autocorrelation function.
+ What is denoted by ACF can easily be identified in context.
+
+- Clearly c0 is the sample variance of $\{X_t\}$. Besides,
+ $r_0 = c_0/c_0 = 1$ and for any integer $k, |r_k| ≤ 1$.
+
+- When we compute the ACF of any sample series with a fixed length
+ $n$, we cannot put too much confidence in the values of $r_k$ for
+ large k’s, since fewer pairs of $(x_{t +k }, x_t )$ are available
+ for calculating $r_k$ as $k$ is large. One rule of thumb is not to
+ estimate $r_k$ for $k > n/3$, and another is $n ≥ 50, k ≤ n/4$. In
+ any case, it is always a good idea to be careful.
+
+- We also compute the ACF of a nonstationary time series sample by
+ Definition 1. In this case, however, the ACF or $r_k$ very slowly or
+ hardly tapers off as $k$ increases.
+
+- Plotting the ACF $(r_k)$ against lag $k$ is easy but very helpful in
+ analyzing time series sample. Such an ACF plot is known as a
+ correlogram.
+
+- If $\{X_t\}$ is stationary with $E(X_t)=0$ and $\rho_k =0$ for all
+ $k \neq 0$,thatis,itisa white noise series, then the sampling
+ distribution of $r_k$ is asymptotically normal with the mean 0 and
+ the variance of $1/n$. Hence, there is about 95% chance that $r_k$
+ falls in the interval $[−1.96/\sqrt{n}, 1.96/\sqrt{n}]$.
+
+Now we can give a summary that (1) if the time series plot of a time
+series clearly shows a trend or/and seasonality, it is surely
+nonstationary; (2) if the ACF $r_k$ very slowly or hardly tapers off as
+lag $k$ increases, the time series should also be nonstationary.
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=30, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+plot_pacf(df["y"], lags=30, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+plt.show();
+```
+
+
+
+### Decomposition of the time series
+
+How to decompose a time series and why?
+
+In time series analysis to forecast new values, it is very important to
+know past data. More formally, we can say that it is very important to
+know the patterns that values follow over time. There can be many
+reasons that cause our forecast values to fall in the wrong direction.
+Basically, a time series consists of four components. The variation of
+those components causes the change in the pattern of the time series.
+These components are:
+
+- **Level:** This is the primary value that averages over time.
+- **Trend:** The trend is the value that causes increasing or
+ decreasing patterns in a time series.
+- **Seasonality:** This is a cyclical event that occurs in a time
+ series for a short time and causes short-term increasing or
+ decreasing patterns in a time series.
+- **Residual/Noise:** These are the random variations in the time
+ series.
+
+Combining these components over time leads to the formation of a time
+series. Most time series consist of level and noise/residual and trend
+or seasonality are optional values.
+
+If seasonality and trend are part of the time series, then there will be
+effects on the forecast value. As the pattern of the forecasted time
+series may be different from the previous time series.
+
+The combination of the components in time series can be of two types: \*
+Additive \* Multiplicative
+
+### Additive time series
+
+If the components of the time series are added to make the time series.
+Then the time series is called the additive time series. By
+visualization, we can say that the time series is additive if the
+increasing or decreasing pattern of the time series is similar
+throughout the series. The mathematical function of any additive time
+series can be represented by:
+$$y(t) = level + Trend + seasonality + noise$$
+
+### Multiplicative time series
+
+If the components of the time series are multiplicative together, then
+the time series is called a multiplicative time series. For
+visualization, if the time series is having exponential growth or
+decline with time, then the time series can be considered as the
+multiplicative time series. The mathematical function of the
+multiplicative time series can be represented as.
+
+$$y(t) = Level * Trend * seasonality * Noise$$
+
+### Additive
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+a = seasonal_decompose(df["y"], model = "additive", period=24)
+a.plot();
+```
+
+
+
+### Multiplicative
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+a = seasonal_decompose(df["y"], model = "Multiplicative", period=24)
+a.plot();
+```
+
+
+
+## Split the data into training and testing
+
+Let’s divide our data into sets
+
+1. Data to train our `Holt Winters Model`.
+2. Data to test our model
+
+For the test data we will use the last 30 hours to test and evaluate the
+performance of our model.
+
+```python
+train = df[df.ds\<='2017-09-20 17:00:00']
+test = df[df.ds>'2017-09-20 17:00:00']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((186, 3), (30, 3))
+```
+
+Now let’s plot the training data and the test data.
+
+```python
+sns.lineplot(train,x="ds", y="y", label="Train", linestyle="--",linewidth=2)
+sns.lineplot(test, x="ds", y="y", label="Test", linewidth=2, color="yellow")
+plt.title("Ads watched (hourly data)");
+plt.show()
+```
+
+
+
+## Implementation of Holt-Winters Method with StatsForecast
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import HoltWinters
+```
+
+### Instantiating Model
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/) by the
+master, Rob Hyndmann, can be useful for `season_length`.
+
+In this case we are going to test two alternatives of the model, one
+additive and one multiplicative.
+
+```python
+season_length = 24 # Hourly data
+horizon = len(test) # number of predictions
+
+models = [HoltWinters(season_length=season_length, error_type="A", alias="Add"),
+ HoltWinters(season_length=season_length, error_type="M", alias="Multi")]
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [panda’s
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models, freq='h')
+```
+
+### Fit the Model
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[Add,Multi])
+```
+
+Let’s see the results of our `Holt Winters Model`. We can observe it
+with the following instruction:
+
+```python
+result=sf.fitted_[0,0].model_
+print(result.keys())
+print(result['fit'])
+```
+
+``` text
+dict_keys(['loglik', 'aic', 'bic', 'aicc', 'mse', 'amse', 'fit', 'residuals', 'components', 'm', 'nstate', 'fitted', 'states', 'par', 'sigma2', 'n_params', 'method', 'actual_residuals'])
+results(x=array([ 2.60632491e-02, 1.53030002e-03, 3.22298668e-02, 9.00958233e-01,
+ 1.23628350e+05, -5.12405452e+01, -3.96677340e+04, -2.83800237e+04,
+ -1.49514829e+04, 1.05413201e+04, 3.65409126e+04, 3.58433030e+04,
+ 2.93235036e+04, 2.66607410e+04, 2.55392078e+04, 2.60970444e+04,
+ 2.63155973e+04, 2.83192738e+04, 2.16640268e+04, 5.19120023e+03,
+ -6.15595960e+03, -8.84863887e+03, -9.28320586e+03, -8.09549672e+03,
+ -3.83755898e+03, -3.33456554e+03, -2.56333963e+04, -3.72181618e+04,
+ -4.42497509e+04]), fn=4363.098387651742, nit=1001, simplex=None)
+```
+
+Let us now visualize the fitted values of our models.
+
+As we can see, the result obtained above has an output in a dictionary,
+to extract each element from the dictionary we are going to use the
+`.get()` function to extract the element and then we are going to save
+it in a `pd.DataFrame()`.
+
+```python
+residual=pd.DataFrame(result.get("residuals"), columns=["residual Model"])
+residual
+```
+
+| | residual Model |
+|-----|----------------|
+| 0 | -1087.029091 |
+| 1 | 623.989786 |
+| 2 | 3054.101324 |
+| ... | ... |
+| 183 | -2783.032921 |
+| 184 | -4618.147123 |
+| 185 | -8194.063498 |
+
+```python
+import scipy.stats as stats
+
+fig, axs = plt.subplots(nrows=2, ncols=2)
+
+residual.plot(ax=axs[0,0])
+axs[0,0].set_title("Residuals");
+
+sns.distplot(residual, ax=axs[0,1]);
+axs[0,1].set_title("Density plot - Residual");
+
+stats.probplot(residual["residual Model"], dist="norm", plot=axs[1,0])
+axs[1,0].set_title('Plot Q-Q')
+
+plot_acf(residual, lags=35, ax=axs[1,1],color="fuchsia")
+axs[1,1].set_title("Autocorrelation");
+
+plt.show();
+```
+
+
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 30 hours ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[90]` means that
+ the model expects the real value to be inside that interval 90% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min.
+
+```python
+Y_hat = sf.forecast(df=train, h=horizon, fitted=True)
+Y_hat
+```
+
+| | unique_id | ds | Add | Multi |
+|-----|-----------|---------------------|---------------|---------------|
+| 0 | 1 | 2017-09-20 18:00:00 | 154164.609375 | 151414.984375 |
+| 1 | 1 | 2017-09-20 19:00:00 | 154547.171875 | 152352.640625 |
+| 2 | 1 | 2017-09-20 20:00:00 | 128790.359375 | 128274.789062 |
+| ... | ... | ... | ... | ... |
+| 27 | 1 | 2017-09-21 21:00:00 | 103021.726562 | 103086.851562 |
+| 28 | 1 | 2017-09-21 22:00:00 | 89544.054688 | 90028.406250 |
+| 29 | 1 | 2017-09-21 23:00:00 | 78090.210938 | 78823.953125 |
+
+With the forecast method we can also extract the fitted values from the
+model and visualize it graphically, with the following instruction we
+can do it.
+
+```python
+values=sf.forecast_fitted_values()
+values.head()
+```
+
+| | unique_id | ds | y | Add | Multi |
+|-----|-----------|---------------------|----------|---------------|---------------|
+| 0 | 1 | 2017-09-13 00:00:00 | 80115.0 | 81202.031250 | 79892.687500 |
+| 1 | 1 | 2017-09-13 01:00:00 | 79885.0 | 79261.007812 | 78792.476562 |
+| 2 | 1 | 2017-09-13 02:00:00 | 89325.0 | 86270.898438 | 85444.117188 |
+| 3 | 1 | 2017-09-13 03:00:00 | 101930.0 | 97905.273438 | 97286.796875 |
+| 4 | 1 | 2017-09-13 04:00:00 | 121630.0 | 120287.523438 | 118195.570312 |
+
+```python
+StatsForecast.plot(values)
+```
+
+
+
+Adding 95% confidence interval with the forecast method
+
+```python
+sf.forecast(df=train, h=horizon, level=[95])
+```
+
+| | unique_id | ds | Add | Add-lo-95 | Add-hi-95 | Multi | Multi-lo-95 | Multi-hi-95 |
+|-----|-----------|---------------------|---------------|---------------|---------------|---------------|---------------|---------------|
+| 0 | 1 | 2017-09-20 18:00:00 | 154164.609375 | 134594.859375 | 173734.375000 | 151414.984375 | 125296.867188 | 177533.109375 |
+| 1 | 1 | 2017-09-20 19:00:00 | 154547.171875 | 134970.062500 | 174124.265625 | 152352.640625 | 126234.515625 | 178470.765625 |
+| 2 | 1 | 2017-09-20 20:00:00 | 128790.359375 | 109205.242188 | 148375.484375 | 128274.789062 | 102156.671875 | 154392.906250 |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| 27 | 1 | 2017-09-21 21:00:00 | 103021.726562 | 83118.632812 | 122924.812500 | 103086.851562 | 76659.867188 | 129513.835938 |
+| 28 | 1 | 2017-09-21 22:00:00 | 89544.054688 | 69626.210938 | 109461.890625 | 90028.406250 | 63601.425781 | 116455.390625 |
+| 29 | 1 | 2017-09-21 23:00:00 | 78090.210938 | 58157.574219 | 98022.843750 | 78823.953125 | 52396.972656 | 105250.937500 |
+
+```python
+sf.plot(train, Y_hat)
+```
+
+
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 30 hours ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[95]` means that
+ the model expects the real value to be inside that interval 95% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+sf.predict(h=horizon)
+```
+
+| | unique_id | ds | Add | Multi |
+|-----|-----------|---------------------|---------------|---------------|
+| 0 | 1 | 2017-09-20 18:00:00 | 154164.609375 | 151414.984375 |
+| 1 | 1 | 2017-09-20 19:00:00 | 154547.171875 | 152352.640625 |
+| 2 | 1 | 2017-09-20 20:00:00 | 128790.359375 | 128274.789062 |
+| ... | ... | ... | ... | ... |
+| 27 | 1 | 2017-09-21 21:00:00 | 103021.726562 | 103086.851562 |
+| 28 | 1 | 2017-09-21 22:00:00 | 89544.054688 | 90028.406250 |
+| 29 | 1 | 2017-09-21 23:00:00 | 78090.210938 | 78823.953125 |
+
+```python
+forecast_df = sf.predict(h=horizon, level=[80,95])
+forecast_df
+```
+
+| | unique_id | ds | Add | Add-lo-95 | Add-lo-80 | Add-hi-80 | Add-hi-95 | Multi | Multi-lo-95 | Multi-lo-80 | Multi-hi-80 | Multi-hi-95 |
+|-----|-----------|---------------------|---------------|---------------|---------------|---------------|---------------|---------------|---------------|---------------|---------------|---------------|
+| 0 | 1 | 2017-09-20 18:00:00 | 154164.609375 | 134594.859375 | 141368.640625 | 166960.593750 | 173734.375000 | 151414.984375 | 125296.867188 | 134337.265625 | 168492.703125 | 177533.109375 |
+| 1 | 1 | 2017-09-20 19:00:00 | 154547.171875 | 134970.062500 | 141746.390625 | 167347.953125 | 174124.265625 | 152352.640625 | 126234.515625 | 135274.921875 | 169430.359375 | 178470.765625 |
+| 2 | 1 | 2017-09-20 20:00:00 | 128790.359375 | 109205.242188 | 115984.335938 | 141596.375000 | 148375.484375 | 128274.789062 | 102156.671875 | 111197.070312 | 145352.515625 | 154392.906250 |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| 27 | 1 | 2017-09-21 21:00:00 | 103021.726562 | 83118.632812 | 90007.796875 | 116035.656250 | 122924.812500 | 103086.851562 | 76659.867188 | 85807.171875 | 120366.523438 | 129513.835938 |
+| 28 | 1 | 2017-09-21 22:00:00 | 89544.054688 | 69626.210938 | 76520.476562 | 102567.632812 | 109461.890625 | 90028.406250 | 63601.425781 | 72748.734375 | 107308.085938 | 116455.390625 |
+| 29 | 1 | 2017-09-21 23:00:00 | 78090.210938 | 58157.574219 | 65056.960938 | 91123.460938 | 98022.843750 | 78823.953125 | 52396.972656 | 61544.281250 | 96103.632812 | 105250.937500 |
+
+```python
+sf.plot(train, forecast_df, level=[80, 95])
+```
+
+
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=)`, forecasting every second months
+`(step_size=12)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 12 months ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=df,
+ h=horizon,
+ step_size=30,
+ n_windows=3)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` series identifier.
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the `n_windows`.
+- `y:` true value
+- `model:` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df
+```
+
+| | unique_id | ds | cutoff | y | Add | Multi |
+|-----|-----------|---------------------|---------------------|----------|---------------|---------------|
+| 0 | 1 | 2017-09-18 06:00:00 | 2017-09-18 05:00:00 | 99440.0 | 134578.328125 | 133820.109375 |
+| 1 | 1 | 2017-09-18 07:00:00 | 2017-09-18 05:00:00 | 97655.0 | 133548.781250 | 133734.000000 |
+| 2 | 1 | 2017-09-18 08:00:00 | 2017-09-18 05:00:00 | 97655.0 | 134798.656250 | 135216.046875 |
+| ... | ... | ... | ... | ... | ... | ... |
+| 87 | 1 | 2017-09-21 21:00:00 | 2017-09-20 17:00:00 | 103080.0 | 103021.726562 | 103086.851562 |
+| 88 | 1 | 2017-09-21 22:00:00 | 2017-09-20 17:00:00 | 95155.0 | 89544.054688 | 90028.406250 |
+| 89 | 1 | 2017-09-21 23:00:00 | 2017-09-20 17:00:00 | 80285.0 | 78090.210938 | 78823.953125 |
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=season_length), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | Add | Multi |
+|-----|-----------|--------|-------------|-------------|
+| 0 | 1 | mae | 4306.244531 | 4886.992188 |
+| 1 | 1 | mape | 0.038087 | 0.043549 |
+| 2 | 1 | mase | 0.532045 | 0.603797 |
+| 3 | 1 | rmse | 5415.015573 | 5862.473702 |
+| 4 | 1 | smape | 0.018708 | 0.021433 |
+
+## References
+
+1. [Changquan Huang • Alla Petukhina. Springer series (2022). Applied
+ Time Series Analysis and Forecasting with
+ Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2)
+2. Ivan Svetunkov. [Forecasting and Analytics with the Augmented
+ Dynamic Adaptive Model (ADAM)](https://openforecast.org/adam/)
+3. [James D. Hamilton. Time Series Analysis Princeton University Press,
+ Princeton, New Jersey, 1st Edition,
+ 1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Nixtla HoltWinters API](../../src/core/models.html#holtwinters)
+5. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+6. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+7. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/models/imapa.html.mdx b/statsforecast/docs/models/imapa.html.mdx
new file mode 100644
index 00000000..0b4464ba
--- /dev/null
+++ b/statsforecast/docs/models/imapa.html.mdx
@@ -0,0 +1,667 @@
+---
+title: IMAPA Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `IMAPA Model` with `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation` in other.
+
+The text in this article is largely taken from: 1. [Changquan Huang •
+Alla Petukhina. Springer series (2022). Applied Time Series Analysis and
+Forecasting with
+Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2) 2.
+Ivan Svetunkov. [Forecasting and Analytics with the Augmented Dynamic
+Adaptive Model (ADAM)](https://openforecast.org/adam/) 3. [James D.
+Hamilton. Time Series Analysis Princeton University Press, Princeton,
+New Jersey, 1st Edition,
+1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+Principles and Practice (3rd ed)”](https://otexts.com/fpp3/tscv.html).
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [IMAPA Model](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of IMAPA with StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+IMAPA is an algorithm that uses multiple models to forecast the future
+values of an intermittent time series. The algorithm starts by adding
+the time series values at regular intervals. It then uses a forecast
+model to forecast the added values.
+
+IMAPA is a good choice for intermittent time series because it is robust
+to missing values and is computationally efficient. IMAPA is also easy
+to implement.
+
+IMAPA has been tested on a variety of intermittent time series and has
+been shown to be effective in forecasting future values.
+
+## IMAPA Method
+
+The Intermittent Multiple Aggregation Prediction Algorithm (IMAPA) model
+is a time series model for forecasting future values for time series
+that are intermittent. The IMAPA model is based on the idea of
+aggregating the time series values at regular intervals and then using a
+forecast model to forecast the aggregated values. The aggregated values
+can be forecast using any forecast model. It uses the optimized SES to
+generate the forecasts at the new levels and then combines them using a
+simple average.
+
+The IMAPA model can be defined mathematically as follows:
+
+$$\hat{y}_{t+1} = f(\hat{y}_{t-\tau}, \hat{y}_{t-2\tau}, ..., \hat{ y}_{t-m\tau})$$
+
+where $\hat{y}_{t+1}$ is the forecast time value $t+1$, $f$ is the
+forecast model,
+$\hat{y}_{t-\tau} , \hat{y}_{t-2\tau}, ..., \hat{y}_{t-m\tau}$ are the
+forecasts of the added values at times $t-\tau, t-2 \tau, ..., t-m\tau$,
+and $\tau$ is the time interval over which the time series values are
+aggregated.
+
+IMAPA is a good choice for intermittent time series because it is robust
+to missing values and is computationally efficient. IMAPA is also easy
+to implement.
+
+IMAPA has been tested on a variety of intermittent time series and has
+been shown to be effective in forecasting future values.
+
+### IMAPA General Properties
+
+- Multiple Aggregation: IMAPA uses multiple levels of aggregation to
+ analyze and predict intermittent time series. This involves
+ decomposing the original series into components of different time
+ scales.
+
+- Intermittency: IMAPA focuses on handling intermittent time series,
+ which are those that exhibit irregular and non-stationary patterns
+ with periods of activity and periods of inactivity.
+
+- Adaptive Prediction: IMAPA uses an adaptive approach to adjust
+ prediction models as new data is collected. This allows the
+ algorithm to adapt to changes in the time series behavior over time.
+
+- Robust to Missing Values: IMAPA can handle missing values in the
+ data without sacrificing accuracy. This is important for
+ intermittent time series, which often have missing values.
+
+- Computationally Efficient: IMAPA is computationally efficient,
+ meaning it can forecast future values quickly. This is important for
+ large time series, which can take a long time to forecast using
+ other methods.
+
+- Decomposition Property: Time series can be decomposed into
+ components such as trend, seasonality, and residual components.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
+plt.style.use('grayscale') # fivethirtyeight grayscale classic
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#008080', # #212946
+ 'axes.facecolor': '#008080',
+ 'savefig.facecolor': '#008080',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#000000', #2A3459
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+
+```python
+import pandas as pd
+
+df=pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/intermittend_demand2")
+
+df.head()
+```
+
+| | date | sales |
+|-----|---------------------|-------|
+| 0 | 2022-01-01 00:00:00 | 0 |
+| 1 | 2022-01-01 01:00:00 | 10 |
+| 2 | 2022-01-01 02:00:00 | 0 |
+| 3 | 2022-01-01 03:00:00 | 0 |
+| 4 | 2022-01-01 04:00:00 | 100 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|---------------------|-----|-----------|
+| 0 | 2022-01-01 00:00:00 | 0 | 1 |
+| 1 | 2022-01-01 01:00:00 | 10 | 1 |
+| 2 | 2022-01-01 02:00:00 | 0 | 1 |
+| 3 | 2022-01-01 03:00:00 | 0 | 1 |
+| 4 | 2022-01-01 04:00:00 | 100 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y int64
+unique_id object
+dtype: object
+```
+
+We can see that our time variable `(ds)` is in an object format, we need
+to convert to a date format
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore Data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints a random series from the dataset and is useful for
+basic EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### Autocorrelation plots
+
+Autocorrelation (ACF) and partial autocorrelation (PACF) plots are
+statistical tools used to analyze time series. ACF charts show the
+correlation between the values of a time series and their lagged values,
+while PACF charts show the correlation between the values of a time
+series and their lagged values, after the effect of previous lagged
+values has been removed.
+
+ACF and PACF charts can be used to identify the structure of a time
+series, which can be helpful in choosing a suitable model for the time
+series. For example, if the ACF chart shows a repeating peak and valley
+pattern, this indicates that the time series is stationary, meaning that
+it has the same statistical properties over time. If the PACF chart
+shows a pattern of rapidly decreasing spikes, this indicates that the
+time series is invertible, meaning it can be reversed to get a
+stationary time series.
+
+The importance of the ACF and PACF charts is that they can help analysts
+better understand the structure of a time series. This understanding can
+be helpful in choosing a suitable model for the time series, which can
+improve the ability to predict future values of the time series.
+
+To analyze ACF and PACF charts:
+
+- Look for patterns in charts. Common patterns include repeating peaks
+ and valleys, sawtooth patterns, and plateau patterns.
+- Compare ACF and PACF charts. The PACF chart generally has fewer
+ spikes than the ACF chart.
+- Consider the length of the time series. ACF and PACF charts for
+ longer time series will have more spikes.
+- Use a confidence interval. The ACF and PACF plots also show
+ confidence intervals for the autocorrelation values. If an
+ autocorrelation value is outside the confidence interval, it is
+ likely to be significant.
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=30, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+# Grafico
+plot_pacf(df["y"], lags=30, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+plt.show();
+```
+
+
+
+### Decomposition of the time series
+
+How to decompose a time series and why?
+
+In time series analysis to forecast new values, it is very important to
+know past data. More formally, we can say that it is very important to
+know the patterns that values follow over time. There can be many
+reasons that cause our forecast values to fall in the wrong direction.
+Basically, a time series consists of four components. The variation of
+those components causes the change in the pattern of the time series.
+These components are:
+
+- **Level:** This is the primary value that averages over time.
+- **Trend:** The trend is the value that causes increasing or
+ decreasing patterns in a time series.
+- **Seasonality:** This is a cyclical event that occurs in a time
+ series for a short time and causes short-term increasing or
+ decreasing patterns in a time series.
+- **Residual/Noise:** These are the random variations in the time
+ series.
+
+Combining these components over time leads to the formation of a time
+series. Most time series consist of level and noise/residual and trend
+or seasonality are optional values.
+
+If seasonality and trend are part of the time series, then there will be
+effects on the forecast value. As the pattern of the forecasted time
+series may be different from the previous time series.
+
+The combination of the components in time series can be of two types: \*
+Additive \* Multiplicative
+
+### Additive time series
+
+If the components of the time series are added to make the time series.
+Then the time series is called the additive time series. By
+visualization, we can say that the time series is additive if the
+increasing or decreasing pattern of the time series is similar
+throughout the series. The mathematical function of any additive time
+series can be represented by:
+$$y(t) = level + Trend + seasonality + noise$$
+
+### Multiplicative time series
+
+If the components of the time series are multiplicative together, then
+the time series is called a multiplicative time series. For
+visualization, if the time series is having exponential growth or
+decline with time, then the time series can be considered as the
+multiplicative time series. The mathematical function of the
+multiplicative time series can be represented as.
+
+$$y(t) = Level * Trend * seasonality * Noise$$
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+from plotly.subplots import make_subplots
+import plotly.graph_objects as go
+
+def plotSeasonalDecompose(
+ x,
+ model='additive',
+ filt=None,
+ period=None,
+ two_sided=True,
+ extrapolate_trend=0,
+ title="Seasonal Decomposition"):
+
+ result = seasonal_decompose(
+ x, model=model, filt=filt, period=period,
+ two_sided=two_sided, extrapolate_trend=extrapolate_trend)
+ fig = make_subplots(
+ rows=4, cols=1,
+ subplot_titles=["Observed", "Trend", "Seasonal", "Residuals"])
+ for idx, col in enumerate(['observed', 'trend', 'seasonal', 'resid']):
+ fig.add_trace(
+ go.Scatter(x=result.observed.index, y=getattr(result, col), mode='lines'),
+ row=idx+1, col=1,
+ )
+ return fig
+```
+
+
+```python
+plotSeasonalDecompose(
+ df["y"],
+ model="additive",
+ period=24,
+ title="Seasonal Decomposition")
+```
+
+
+
+## Split the data into training and testing
+
+Let’s divide our data into sets
+
+1. Data to train our `IMAPA Model`.
+2. Data to test our model
+
+For the test data we will use the last 500 Hours to test and evaluate
+the performance of our model.
+
+```python
+train = df[df.ds\<='2023-01-31 19:00:00']
+test = df[df.ds>'2023-01-31 19:00:00']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((9500, 3), (500, 3))
+```
+
+Now let’s plot the training data and the test data.
+
+```python
+sns.lineplot(train,x="ds", y="y", label="Train", linestyle="--",linewidth=2)
+sns.lineplot(test, x="ds", y="y", label="Test", linewidth=2, color="yellow")
+plt.title("Store visit");
+plt.xlabel("Hours")
+plt.show()
+```
+
+
+
+## Implementation of IMAPA Method with StatsForecast
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import IMAPA
+```
+
+### Instantiating Model
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/) by the
+master, Rob Hyndmann, can be useful for `season_length`.
+
+```python
+season_length = 24 # Hourly data
+horizon = len(test) # number of predictions
+
+models = [IMAPA()]
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [pandas’
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models, freq='h')
+```
+
+### Fit the Model
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[IMAPA])
+```
+
+Let’s see the results of our `IMAPA Model`. We can observe it with the
+following instruction:
+
+```python
+result=sf.fitted_[0,0].model_
+result
+```
+
+``` text
+{'mean': array([28.579695], dtype=float32)}
+```
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 500 hours ahead.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min.
+
+```python
+Y_hat = sf.forecast(df=train, h=horizon)
+Y_hat
+```
+
+| | unique_id | ds | IMAPA |
+|-----|-----------|---------------------|-----------|
+| 0 | 1 | 2023-01-31 20:00:00 | 28.579695 |
+| 1 | 1 | 2023-01-31 21:00:00 | 28.579695 |
+| 2 | 1 | 2023-01-31 22:00:00 | 28.579695 |
+| ... | ... | ... | ... |
+| 497 | 1 | 2023-02-21 13:00:00 | 28.579695 |
+| 498 | 1 | 2023-02-21 14:00:00 | 28.579695 |
+| 499 | 1 | 2023-02-21 15:00:00 | 28.579695 |
+
+```python
+sf.plot(train, Y_hat)
+```
+
+
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 500 hours ahead.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+forecast_df = sf.predict(h=horizon)
+forecast_df
+```
+
+| | unique_id | ds | IMAPA |
+|-----|-----------|---------------------|-----------|
+| 0 | 1 | 2023-01-31 20:00:00 | 28.579695 |
+| 1 | 1 | 2023-01-31 21:00:00 | 28.579695 |
+| 2 | 1 | 2023-01-31 22:00:00 | 28.579695 |
+| ... | ... | ... | ... |
+| 497 | 1 | 2023-02-21 13:00:00 | 28.579695 |
+| 498 | 1 | 2023-02-21 14:00:00 | 28.579695 |
+| 499 | 1 | 2023-02-21 15:00:00 | 28.579695 |
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=)`, forecasting every second months
+`(step_size=50)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 500 hours ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=df,
+ h=horizon,
+ step_size=50,
+ n_windows=5)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` index. If you dont like working with index just run
+ `crossvalidation_df.resetindex()`.
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the `n_windows`.
+- `y:` true value
+- `model:` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df
+```
+
+| | unique_id | ds | cutoff | y | IMAPA |
+|------|-----------|---------------------|---------------------|------|-----------|
+| 0 | 1 | 2023-01-23 12:00:00 | 2023-01-23 11:00:00 | 0.0 | 15.134251 |
+| 1 | 1 | 2023-01-23 13:00:00 | 2023-01-23 11:00:00 | 0.0 | 15.134251 |
+| 2 | 1 | 2023-01-23 14:00:00 | 2023-01-23 11:00:00 | 0.0 | 15.134251 |
+| ... | ... | ... | ... | ... | ... |
+| 2497 | 1 | 2023-02-21 13:00:00 | 2023-01-31 19:00:00 | 60.0 | 28.579695 |
+| 2498 | 1 | 2023-02-21 14:00:00 | 2023-01-31 19:00:00 | 20.0 | 28.579695 |
+| 2499 | 1 | 2023-02-21 15:00:00 | 2023-01-31 19:00:00 | 20.0 | 28.579695 |
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=season_length), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | IMAPA |
+|-----|-----------|--------|-----------|
+| 0 | 1 | mae | 34.206428 |
+| 1 | 1 | mape | 0.637417 |
+| 2 | 1 | mase | 0.816042 |
+| 3 | 1 | rmse | 45.345223 |
+| 4 | 1 | smape | 0.764973 |
+
+## References
+
+1. [Changquan Huang • Alla Petukhina. Springer series (2022). Applied
+ Time Series Analysis and Forecasting with
+ Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2)
+2. Ivan Svetunkov. [Forecasting and Analytics with the Augmented
+ Dynamic Adaptive Model (ADAM)](https://openforecast.org/adam/)
+3. [James D. Hamilton. Time Series Analysis Princeton University Press,
+ Princeton, New Jersey, 1st Edition,
+ 1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Nixtla IMAPA API](../../src/core/models.html#imapa)
+5. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+6. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+7. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/models/mfles.html.mdx b/statsforecast/docs/models/mfles.html.mdx
new file mode 100644
index 00000000..fd11810c
--- /dev/null
+++ b/statsforecast/docs/models/mfles.html.mdx
@@ -0,0 +1,479 @@
+---
+title: MFLES
+---
+
+> MFLES is a simple time series method based on gradient boosting time
+> series decomposition.
+
+There are numerous methods that can enter the boosting loop depending on
+user-provided parameters or some quick logic MFLES does automatically
+that seems to work ok. Some of these methods are:
+
+1. SES Ensemble
+2. Simple Moving Average
+3. Piecewise Linear Trend
+4. Fourier Basis function regression for seasonality
+5. Simple Median
+6. A Robust Linear Method for trend
+
+# **Gradient Boosted Decomposition**
+
+This approach aims to view a time series decomposition (trend,
+seasonality, and exogenous) as the ‘weak’ estimator in a gradient
+boosting procedure.
+
+
+
+The major relevant changes to note are:
+
+1. The trend estimator will always go from simple to complex. Beginning
+ with a median, then to a linear/piecewise linear, then to some sort
+ of smoother.
+2. Multiple seasonality is fit one seasonality per boosting round
+ rather than simultaneously. This means you should organize your
+ seasonality in order of perceived importance. Also, theoretically,
+ you can have up to 50 seasonalities present by default, but after 3
+ you should expect degraded performance.
+3. Learning rates are now estimator specific rather than a single
+ parameter like you would see in something like XGBoost. This is
+ useful if you have exogenous signals that are also seasonal, you
+ (this will not be done automatically) can optimize for the
+ combination of the seasonal signal and the exogenous signal.
+
+# **Let’s forecast**
+
+```python
+# %pip install statsforecast
+```
+
+Here, we will use the specific model object in Statsforecast and the
+infamous airline passengers dataset 😀:
+
+```python
+import pandas as pd
+import numpy as np
+from statsforecast.models import AutoMFLES
+import matplotlib.pyplot as plt
+
+df = pd.read_csv(r'https://raw.githubusercontent.com/jbrownlee/Datasets/master/airline-passengers.csv')
+y = df['Passengers'].values # make array
+
+
+mfles_model = AutoMFLES(
+ season_length = [12],
+ test_size = 12,
+ n_windows=2,
+ metric = 'smape')
+mfles_model.fit(y=y)
+predicted = mfles_model.predict(12)['mean']
+fitted = mfles_model.predict_in_sample()['fitted']
+```
+
+
+```python
+plt.plot(np.append(fitted, predicted), linestyle='dashed', color='red')
+plt.plot(y)
+plt.show()
+```
+
+
+
+Let’s take a look at some of the key parameters for a standard
+experience.
+
+- **season_length**: a list of seasonal periods, in order of perceived
+ importance preferably.
+
+- **test_size**: AutoMFLES is optimized via time series cross
+ validation. The test size dictates how many periods to use in each
+ test fold. **This is probably the most important parameter when it
+ comes to optimizing and you should weigh the season length, forecast
+ horizon, and general data length when setting this. But a good rule
+ of thumb is either the most important season length or half that to
+ allow MFLES to pick up on seasonality.**
+
+- **n_windows**: how many test sets are used in optimizing parameters.
+ In this example, 2 means that we, in total, use 24 months (12 \* 2)
+ split between the 2 windows.
+
+- **metric**: this one is easy, it is simply the metric we want to
+ optimize for with our parameters. Here we use the default which is
+ smape that is defaulted to reproduce experiment results on M4. You
+ can also pass ‘rmse’, ‘mape’, or ‘mae’ to optimize for another
+ metric.
+
+# **A deeper look at a more customized model**
+
+The previous fit is done with 99% automated logic checks and grid
+searched parameters. But we can manipulate the fit greatly (maybe too
+much). This section will overview some very important parameters and how
+they effect the output.
+
+## **The parameter grid search**
+
+First, let’s take a look at the default grid of parameters AutoMFLES
+will try:
+
+```python
+config = {
+ 'seasonality_weights': [True, False],
+ 'smoother': [True, False],
+ 'ma': [int(min(seasonal_period)), int(min(seasonal_period)/2),None],
+ 'seasonal_period': [None, seasonal_period],
+}
+```
+
+- **seasonality_weights**: If True, we will weigh more recent
+ observations more when calculating seasonality. The allows a
+ deterministic seasonality to reflect more recent changes.
+- **smoother**: True means we will use a simple exponential smoother
+ to fit on residuals after a few rounds of boosting. If the parameter
+ is False then we use a simple moving average
+- **ma**: This parameter is the number of past observations to include
+ when using a moving average, None indicates it will be semi-auto set
+ or disregarded in the case of ‘smoother’ being True. For optimizing
+ we search for the minimum season length provided by you or that
+ number divided by 2.
+- **seasonal_period**: this is the list of season_length provided by
+ the you
+
+Now let’s see how to pass the config to AutoMFLES, since this is what we
+use under-the-hood the results will be the same!
+
+```python
+season_length = [12]
+
+config = {
+ 'seasonality_weights': [True, False],
+ 'smoother': [True, False],
+ 'ma': [int(min(season_length)), int(min(season_length)/2),None],
+ 'seasonal_period': [None, season_length],
+ }
+
+mfles_model = AutoMFLES(
+ season_length = season_length,
+ test_size = 12,
+ n_windows=2,
+ metric = 'smape',
+ config=config) # adding the config dictionary manually
+mfles_model.fit(y=y)
+predicted = mfles_model.predict(12)['mean']
+fitted = mfles_model.predict_in_sample()['fitted']
+
+plt.plot(np.append(fitted, predicted), linestyle='dashed', color='red')
+plt.plot(y)
+plt.show()
+```
+
+
+
+### What if you want to force a less reactive forecast?
+
+Just pass False for the smoother and adjust ma to be larger relative to
+your seasonality
+
+```python
+season_length = [12]
+
+config = {
+ 'seasonality_weights': [True, False],
+ 'smoother': [False],
+ 'ma': [30],
+ 'seasonal_period': [None, season_length],
+ }
+
+mfles_model = AutoMFLES(
+ season_length = season_length,
+ test_size = 12,
+ n_windows=2,
+ metric = 'smape',
+ config=config) # adding the config dictionary manually
+mfles_model.fit(y=y)
+predicted = mfles_model.predict(12)['mean']
+fitted = mfles_model.predict_in_sample()['fitted']
+
+plt.plot(np.append(fitted, predicted), linestyle='dashed', color='red')
+plt.plot(y)
+plt.show()
+```
+
+
+
+### **Forcing** **Seasonality**
+
+Sometimes a seasonal series is auto-fit with a nonseasonal setting, to
+adjust this and force seasonality, jsut remove the ‘None’ setting in the
+seasonal_period list.
+
+This also reduces the number of configurations MFLES tries and therefore
+speeds up the fitting.
+
+```python
+season_length = [12]
+
+config = {
+ 'seasonality_weights': [True, False],
+ 'smoother': [False],
+ 'ma': [30],
+ 'seasonal_period': [season_length],
+ }
+
+mfles_model = AutoMFLES(
+ season_length = season_length,
+ test_size = 12,
+ n_windows=2,
+ metric = 'smape',
+ config=config) # adding the config dictionary manually
+mfles_model.fit(y=y)
+predicted = mfles_model.predict(12)['mean']
+fitted = mfles_model.predict_in_sample()['fitted']
+
+plt.plot(np.append(fitted, predicted), linestyle='dashed', color='red')
+plt.plot(y)
+plt.show()
+```
+
+
+
+## **Controlling the Complexity**
+
+One of the best ways to control for complexity is with the max_rounds
+parameter. By default this is set to 50 but most of the time the model
+converges much quicker than that. At round 4 we start implementing
+smoothers as the trend piece so if you do not want that then set the
+max_rounds to 3! But, you probably want the smoothers!
+
+```python
+season_length = [12]
+
+config = {
+ 'seasonality_weights': [True, False],
+ 'smoother': [True, False],
+ 'ma': [int(min(season_length)), int(min(season_length)/2),None],
+ 'seasonal_period': [None, season_length],
+ 'max_rounds': [3],
+ }
+
+mfles_model = AutoMFLES(
+ season_length = season_length,
+ test_size = 12,
+ n_windows=2,
+ metric = 'smape',
+ config=config) # adding the config dictionary manually
+mfles_model.fit(y=y)
+predicted = mfles_model.predict(12)['mean']
+fitted = mfles_model.predict_in_sample()['fitted']
+
+plt.plot(np.append(fitted, predicted), linestyle='dashed', color='red')
+plt.plot(y)
+plt.show()
+```
+
+
+
+You can also leverage estimator specific learning rates which are
+applied to individual estimators rather than the entire boosting round.
+Useful if you notice that the residual smoother is eating too much
+signal too quickly:
+
+```python
+season_length = [12]
+
+config = {
+ 'seasonality_weights': [True, False],
+ 'smoother': [True, False],
+ 'ma': [int(min(season_length)), int(min(season_length)/2),None],
+ 'seasonal_period': [None, season_length],
+ 'rs_lr': [.2],
+ }
+
+mfles_model = AutoMFLES(
+ season_length = season_length,
+ test_size = 12,
+ n_windows=2,
+ metric = 'smape',
+ config=config) # adding the config dictionary manually
+mfles_model.fit(y=y)
+predicted = mfles_model.predict(12)['mean']
+fitted = mfles_model.predict_in_sample()['fitted']
+
+plt.plot(np.append(fitted, predicted), linestyle='dashed', color='red')
+plt.plot(y)
+plt.show()
+```
+
+
+
+## **Tips and Tricks**
+
+Since most settings are optimized for during cross validation there is
+always a trade-off between accuracy and computation.
+
+The default settings were done after extensive testing to give you a
+balanced approach. Hopefully, it delivers good accuracy in a short
+amount of time.
+
+But, there are ways to give you generally more accuracy (not life
+changing but a slight boost) or a dramatic decrease in runtime (without
+sacrificing too much accuracy).
+
+The next section will review some of those settings!
+
+## **Number of Testing Windows**
+
+When optimizing using time series cross validation the number of windows
+directly effects the number of times we have to fit the model for each
+parameter. The default here is 2, but going up to 3 (if your data allows
+it) should give you more consistent results. Obviously, the more the
+better to a certain point but this will depend on your data. Conversely,
+decreasing this to 1 means you are choosing parameters based on a single
+holdout set which may decrease accuracy.
+
+```python
+season_length = [12]
+
+mfles_model = AutoMFLES(
+ season_length = season_length,
+ test_size = 12,
+ n_windows = 1, # Trying just 1 window here
+ metric = 'smape')
+mfles_model.fit(y=y)
+predicted = mfles_model.predict(12)['mean']
+fitted = mfles_model.predict_in_sample()['fitted']
+
+plt.plot(np.append(fitted, predicted), linestyle='dashed', color='red')
+plt.plot(y)
+plt.show()
+```
+
+
+
+And now trying with 3, notice the fit is different!
+
+```python
+season_length = [12]
+
+mfles_model = AutoMFLES(
+ season_length = season_length,
+ test_size = 12,
+ n_windows = 3, # Trying just 1 window here
+ metric = 'smape')
+mfles_model.fit(y=y)
+predicted = mfles_model.predict(12)['mean']
+fitted = mfles_model.predict_in_sample()['fitted']
+
+plt.plot(np.append(fitted, predicted), linestyle='dashed', color='red')
+plt.plot(y)
+plt.show()
+```
+
+
+
+## **The Moving Average Parameter**
+
+By default, we will try the min of your season lengths and half that for
+the ‘ma’ parameter. This works well in the wild but you may want to
+deepen this search greatly. **This is one of the best parameters to
+tweak if you need more accuracy out of MFLES**. Simply pass more
+parameters to the list, ideally these numbers are informed by the
+seasonality, forecast horizon, or some other bit of information. In our
+case, I will also pass 3 and 4 due to it being monthly data. Since this
+increases the number of parameters to try, it will also increase the
+computation time.
+
+```python
+season_length = [12]
+
+config = {
+ 'seasonality_weights': [True, False],
+ 'smoother': [True, False],
+ 'ma': [3, 4, int(min(season_length)), int(min(season_length)/2),None],
+ 'seasonal_period': [None, season_length],
+ }
+
+mfles_model = AutoMFLES(
+ season_length = season_length,
+ test_size = 12,
+ n_windows=2,
+ metric = 'smape',
+ config=config) # adding the config dictionary manually
+mfles_model.fit(y=y)
+predicted = mfles_model.predict(12)['mean']
+fitted = mfles_model.predict_in_sample()['fitted']
+
+plt.plot(np.append(fitted, predicted), linestyle='dashed', color='red')
+plt.plot(y)
+plt.show()
+```
+
+
+
+### **Changepoints**
+
+By default, MFLES will auto-detect if it should use changepoints. This
+has some accuracy benefits but massive computation expenses. You can
+disable changepoints and generally see close accuracy but great speed
+gains:
+
+```python
+season_length = [12]
+
+config = {
+ 'changepoints': [False],
+ 'seasonality_weights': [True, False],
+ 'smoother': [True, False],
+ 'ma': [int(min(season_length)), int(min(season_length)/2),None],
+ 'seasonal_period': [None, season_length],
+ }
+
+mfles_model = AutoMFLES(
+ season_length = season_length,
+ test_size = 12,
+ n_windows=2,
+ metric = 'smape',
+ config=config) # adding the config dictionary manually
+mfles_model.fit(y=y)
+predicted = mfles_model.predict(12)['mean']
+fitted = mfles_model.predict_in_sample()['fitted']
+
+plt.plot(np.append(fitted, predicted), linestyle='dashed', color='red')
+plt.plot(y)
+plt.show()
+```
+
+
+
+### **Seasonality Weights**
+
+Most time series will not have a significant shift in the seasonal
+signal, or at least not one that is worth the extra computation needed
+to fit for it. To speed things up a bit, you can disable this. Although,
+sometimes, disabling this will cause large degradation in accuracy.
+
+```python
+season_length = [12]
+
+config = {
+ 'seasonality_weights': [False],
+ 'smoother': [True, False],
+ 'ma': [int(min(season_length)), int(min(season_length)/2),None],
+ 'seasonal_period': [None, season_length],
+ }
+
+mfles_model = AutoMFLES(
+ season_length = season_length,
+ test_size = 12,
+ n_windows=2,
+ metric = 'smape',
+ config=config) # adding the config dictionary manually
+mfles_model.fit(y=y)
+predicted = mfles_model.predict(12)['mean']
+fitted = mfles_model.predict_in_sample()['fitted']
+
+plt.plot(np.append(fitted, predicted), linestyle='dashed', color='red')
+plt.plot(y)
+plt.show()
+```
+
+
+
diff --git a/statsforecast/docs/models/multipleseasonaltrend.html.mdx b/statsforecast/docs/models/multipleseasonaltrend.html.mdx
new file mode 100644
index 00000000..705e6734
--- /dev/null
+++ b/statsforecast/docs/models/multipleseasonaltrend.html.mdx
@@ -0,0 +1,875 @@
+---
+title: Multiple Seasonal Trend (MSTL)
+---
+
+
+
+
+
+> Step-by-step guide on using the `MSTL Model` with `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation` in other.
+
+The text in this article is largely taken from: 1. [Changquan Huang •
+Alla Petukhina. Springer series (2022). Applied Time Series Analysis and
+Forecasting with
+Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2) 2.
+Ivan Svetunkov. [Forecasting and Analytics with the Augmented Dynamic
+Adaptive Model (ADAM)](https://openforecast.org/adam/) 3. [James D.
+Hamilton. Time Series Analysis Princeton University Press, Princeton,
+New Jersey, 1st Edition,
+1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+Principles and Practice (3rd ed)”](https://otexts.com/fpp3/tscv.html).
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [IMAPA Model](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of IMAPA with StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+The MSTL model (Multiple Seasonal-Trend decomposition using LOESS) is a
+method used to decompose a time series into its seasonal, trend and
+residual components. This approach is based on the use of LOESS (Local
+Regression Smoothing) to estimate the components of the time series.
+
+The MSTL decomposition is an extension of the classic seasonal-trend
+decomposition method (also known as Holt-Winters decomposition), which
+is designed to handle situations where multiple seasonal patterns exist
+in the data. This can occur, for example, when a time series exhibits
+daily, weekly, and yearly patterns simultaneously.
+
+The MSTL decomposition process is performed in several stages:
+
+1. Trend estimation: LOESS is used to estimate the trend component of
+ the time series. LOESS is a non-parametric smoothing method that
+ locally fits data and allows complex trend patterns to be captured.
+
+2. Estimation of seasonal components: Seasonal decomposition techniques
+ are applied to identify and model the different seasonal patterns
+ present in the data. This involves extracting and modeling seasonal
+ components, such as daily, weekly, or yearly patterns.
+
+3. Estimation of the residuals: The residuals are calculated as the
+ difference between the original time series and the sum of the
+ estimates of trend and seasonal components. Residuals represent
+ variation not explained by trend and seasonal patterns and may
+ contain additional information or noise.
+
+MSTL decomposition allows you to analyze and understand the different
+components of a time series in more detail, which can make it easier to
+forecast and detect patterns or anomalies. Furthermore, the use of LOESS
+provides flexibility to adapt to different trend and seasonal patterns
+present in the data.
+
+It is important to note that the MSTL model is only one of the available
+approaches for time series decomposition and that its choice will depend
+on the specific characteristics of the data and the application context.
+
+## MSTL
+
+An important objective in time series analysis is the decomposition of a
+series into a set of non-observable (latent) components that can be
+associated with different types of temporal variations. The idea of time
+series decomposition is very old and was used for the calculation of
+planetary orbits by seventeenth century astronomers. Persons was the
+first to state explicitly the assumptions of unobserved components. As
+Persons saw it, time series was composed of four types of fluctuations:
+
+1. a long-term tendency or secular trend;
+2. cyclical movements superimposed upon the long-term trend. These
+ cycles appear to reach their peaks during periods of industrial
+ prosperity and their troughs during periods of depressions, their
+ rise and fall constituting the business cycle;
+3. a seasonal movement within each year, the shape of which depends on
+ the nature of the series;
+4. residual variations due to changes impacting individual variables or
+ other major events, such as wars and national catastrophes affecting
+ a number of variables.
+
+Traditionally, the four variations have been assumed to be mutually
+independent from one another and specified by means of an additive
+decomposition model:
+
+$$
+\begin{equation}
+y_t= T_t +C_t +S_t +I_t, t=1,\ \cdots, n \tag 1
+\end{equation}
+$$
+
+where $y_t$ denotes the observed series at time $t$, $T_t$ the long-term
+trend, $C_t$ the business cycle, $S_t$ seasonality, and $I_t$ the
+irregulars.
+
+If there is dependence among the latent components, this relationship is
+specified through a multiplicative model
+
+$$
+\begin{equation}
+y_t= T_t \times C_t \times S_t \times I_t, t=1,\ \cdots, n \tag 2
+\end{equation}
+$$
+
+where now $S_t$ and $I_t$ are expressed in proportion to the trend-cycle
+$T_t \times C_t$ . In some cases, mixed additive-multiplicative models
+are used.
+
+### LOESS (Local Regression Smoothing)
+
+LOESS is a nonparametric smoothing method used to estimate a smooth
+function that locally fits the data. For each point in the time series,
+LOESS performs a weighted regression using nearest neighbors.
+
+The LOESS calculation involves the following steps:
+
+- For each point t in the time series, a nearest neighbor window is
+ selected.
+- Weights are assigned to neighbors based on their proximity to t,
+ using a weighting function, such as the Gaussian kernel.
+- A weighted regression is performed using the neighbors and their
+ assigned weights.
+- The fitted value for point t is obtained based on local regression.
+- The process is repeated for all points in the time series, thus
+ obtaining a smoothed estimate of the trend.
+
+### MSTL General Properties
+
+The MSTL model (Multiple Seasonal-Trend decomposition using LOESS) has
+several properties that make it useful in time series analysis. Here is
+a list of some of its properties:
+
+1. Decomposition of multiple seasonal components: The MSTL model is
+ capable of handling time series that exhibit multiple seasonal
+ patterns simultaneously. You can effectively identify and model
+ different seasonal components present in the data.
+
+2. Flexibility in detecting complex trends: Thanks to the use of LOESS,
+ the MSTL model can capture complex trend patterns in the data. This
+ includes non-linear trends and abrupt changes in the time series.
+
+3. Adaptability to different seasonal frequencies: The MSTL model is
+ capable of handling data with different seasonal frequencies, such
+ as daily, weekly, monthly, or even yearly patterns. You can identify
+ and model seasonal patterns of different cycle lengths. (see)
+ [Seasonal
+ periods](https://robjhyndman.com/hyndsight/seasonal-periods/)
+
+| Frecuencia |
+|------------|
+
+| Data | Minute | Hour | Day | Week | Year |
+|-------------|--------|------|-------|--------|----------|
+| Daily | | | | 7 | 365.25 |
+| Hourly | | | 24 | 168 | 8766 |
+| Half-hourly | | | 48 | 336 | 17532 |
+| Minutes | | 60 | 1440 | 10080 | 525960 |
+| Seconds | 60 | 3600 | 86400 | 604800 | 31557600 |
+
+1. Ability to smooth noise and outliers: The smoothing process used in
+ LOESS allows to reduce the impact of noise and outliers in the time
+ series. This can improve detection of underlying patterns and make
+ it easier to analyze trend and seasonality.
+
+2. Improved forecasting: By decomposing the time series into seasonal,
+ trend, and residual components, the MSTL model can provide more
+ accurate forecasts. Forecasts can be generated by extrapolating
+ trend and seasonal patterns into the future, and adding the
+ stochastic residuals.
+
+3. More detailed interpretation and analysis: The MSTL decomposition
+ allows you to analyze and understand the different components of the
+ time series in a more detailed way. This facilitates the
+ identification of seasonal patterns, changes in trend, and the
+ evaluation of residual variability.
+
+4. Efficient Implementation: Although the specific implementation may
+ vary, the MSTL model can be calculated efficiently, especially when
+ LOESS is used in combination with optimized calculation algorithms.
+
+These properties make the MSTL model a useful tool for exploratory time
+series analysis, data forecasting, and pattern detection in the presence
+of multiple seasonal components and complex trends.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
+plt.style.use('grayscale') # fivethirtyeight grayscale classic
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#008080', # #212946
+ 'axes.facecolor': '#008080',
+ 'savefig.facecolor': '#008080',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#000000', #2A3459
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+
+```python
+import pandas as pd
+df=pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/ads.csv")
+
+df.head()
+```
+
+| | Time | Ads |
+|-----|---------------------|--------|
+| 0 | 2017-09-13T00:00:00 | 80115 |
+| 1 | 2017-09-13T01:00:00 | 79885 |
+| 2 | 2017-09-13T02:00:00 | 89325 |
+| 3 | 2017-09-13T03:00:00 | 101930 |
+| 4 | 2017-09-13T04:00:00 | 121630 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|---------------------|--------|-----------|
+| 0 | 2017-09-13T00:00:00 | 80115 | 1 |
+| 1 | 2017-09-13T01:00:00 | 79885 | 1 |
+| 2 | 2017-09-13T02:00:00 | 89325 | 1 |
+| 3 | 2017-09-13T03:00:00 | 101930 | 1 |
+| 4 | 2017-09-13T04:00:00 | 121630 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y int64
+unique_id object
+dtype: object
+```
+
+We can see that our time variable `(ds)` is in an object format, we need
+to convert to a date format
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore Data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints a random series from the dataset and is useful for
+basic EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### Autocorrelation plots
+
+Autocorrelation (ACF) and partial autocorrelation (PACF) plots are
+statistical tools used to analyze time series. ACF charts show the
+correlation between the values of a time series and their lagged values,
+while PACF charts show the correlation between the values of a time
+series and their lagged values, after the effect of previous lagged
+values has been removed.
+
+ACF and PACF charts can be used to identify the structure of a time
+series, which can be helpful in choosing a suitable model for the time
+series. For example, if the ACF chart shows a repeating peak and valley
+pattern, this indicates that the time series is stationary, meaning that
+it has the same statistical properties over time. If the PACF chart
+shows a pattern of rapidly decreasing spikes, this indicates that the
+time series is invertible, meaning it can be reversed to get a
+stationary time series.
+
+The importance of the ACF and PACF charts is that they can help analysts
+better understand the structure of a time series. This understanding can
+be helpful in choosing a suitable model for the time series, which can
+improve the ability to predict future values of the time series.
+
+To analyze ACF and PACF charts:
+
+- Look for patterns in charts. Common patterns include repeating peaks
+ and valleys, sawtooth patterns, and plateau patterns.
+- Compare ACF and PACF charts. The PACF chart generally has fewer
+ spikes than the ACF chart.
+- Consider the length of the time series. ACF and PACF charts for
+ longer time series will have more spikes.
+- Use a confidence interval. The ACF and PACF plots also show
+ confidence intervals for the autocorrelation values. If an
+ autocorrelation value is outside the confidence interval, it is
+ likely to be significant.
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=30, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+# Grafico
+plot_pacf(df["y"], lags=30, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+plt.show();
+```
+
+
+
+### Decomposition of the time series
+
+How to decompose a time series and why?
+
+In time series analysis to forecast new values, it is very important to
+know past data. More formally, we can say that it is very important to
+know the patterns that values follow over time. There can be many
+reasons that cause our forecast values to fall in the wrong direction.
+Basically, a time series consists of four components. The variation of
+those components causes the change in the pattern of the time series.
+These components are:
+
+- **Level:** This is the primary value that averages over time.
+- **Trend:** The trend is the value that causes increasing or
+ decreasing patterns in a time series.
+- **Seasonality:** This is a cyclical event that occurs in a time
+ series for a short time and causes short-term increasing or
+ decreasing patterns in a time series.
+- **Residual/Noise:** These are the random variations in the time
+ series.
+
+Combining these components over time leads to the formation of a time
+series. Most time series consist of level and noise/residual and trend
+or seasonality are optional values.
+
+If seasonality and trend are part of the time series, then there will be
+effects on the forecast value. As the pattern of the forecasted time
+series may be different from the previous time series.
+
+The combination of the components in time series can be of two types: \*
+Additive \* Multiplicative
+
+### Additive time series
+
+If the components of the time series are added to make the time series.
+Then the time series is called the additive time series. By
+visualization, we can say that the time series is additive if the
+increasing or decreasing pattern of the time series is similar
+throughout the series. The mathematical function of any additive time
+series can be represented by:
+$$y(t) = level + Trend + seasonality + noise$$
+
+### Multiplicative time series
+
+If the components of the time series are multiplicative together, then
+the time series is called a multiplicative time series. For
+visualization, if the time series is having exponential growth or
+decline with time, then the time series can be considered as the
+multiplicative time series. The mathematical function of the
+multiplicative time series can be represented as.
+
+$$y(t) = Level * Trend * seasonality * Noise$$
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+from plotly.subplots import make_subplots
+import plotly.graph_objects as go
+
+def plotSeasonalDecompose(
+ x,
+ model='additive',
+ filt=None,
+ period=None,
+ two_sided=True,
+ extrapolate_trend=0,
+ title="Seasonal Decomposition"):
+
+ result = seasonal_decompose(
+ x, model=model, filt=filt, period=period,
+ two_sided=two_sided, extrapolate_trend=extrapolate_trend)
+ fig = make_subplots(
+ rows=4, cols=1,
+ subplot_titles=["Observed", "Trend", "Seasonal", "Residuals"])
+ for idx, col in enumerate(['observed', 'trend', 'seasonal', 'resid']):
+ fig.add_trace(
+ go.Scatter(x=result.observed.index, y=getattr(result, col), mode='lines'),
+ row=idx+1, col=1,
+ )
+ return fig
+```
+
+
+```python
+plotSeasonalDecompose(
+ df["y"],
+ model="additive",
+ period=24,
+ title="Seasonal Decomposition")
+```
+
+``` text
+Unable to display output for mime type(s): application/vnd.plotly.v1+json
+```
+
+## Split the data into training and testing
+
+Let’s divide our data into sets 1. Data to train our `MSTL Model`. 2.
+Data to test our model
+
+For the test data we will use the last 30 Hours to test and evaluate the
+performance of our model.
+
+```python
+train = df[df.ds\<='2017-09-20 17:00:00']
+test = df[df.ds>'2017-09-20 17:00:00']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((186, 3), (30, 3))
+```
+
+Now let’s plot the training data and the test data.
+
+```python
+sns.lineplot(train,x="ds", y="y", label="Train", linestyle="--",linewidth=2)
+sns.lineplot(test, x="ds", y="y", label="Test", linewidth=2, color="yellow")
+plt.title("Ads watched (hourly data)");
+plt.xlabel("Hours")
+plt.show()
+```
+
+
+
+## Implementation of MSTL Method with StatsForecast
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import MSTL, AutoARIMA
+```
+
+### Instantiating Model
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/) by the
+master, Rob Hyndmann, can be useful for `season_length`.
+
+First, we must define the model parameters. As mentioned before, the
+Candy production load presents seasonalities every 24 hours (Hourly) and
+every 24 \* 7 (Daily) hours. Therefore, we will use `[24, 24 * 7]` for
+season length. The trend component will be forecasted with an
+`AutoARIMA` model. (You can also try with: `AutoTheta`, `AutoCES`, and
+`AutoETS`)
+
+```python
+from statsforecast.utils import ConformalIntervals
+horizon = len(test) # number of predictions
+
+models = [MSTL(season_length=[24, 168], # seasonalities of the time series
+trend_forecaster=AutoARIMA(prediction_intervals=ConformalIntervals(n_windows=3, h=horizon)))]
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [pandas’
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models, freq='h')
+```
+
+### Fit Model
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[MSTL])
+```
+
+Let’s see the results of our `MSTL Model`. We can observe it with the
+following instruction:
+
+```python
+result=sf.fitted_[0,0].model_
+result
+```
+
+| | data | trend | seasonal24 | seasonal168 | remainder |
+|-----|----------|---------------|---------------|--------------|--------------|
+| 0 | 80115.0 | 126222.558267 | -42511.086107 | -1524.379074 | -2072.093085 |
+| 1 | 79885.0 | 126191.340644 | -43585.928105 | -1315.292640 | -1405.119899 |
+| 2 | 89325.0 | 126160.117727 | -36756.458517 | 659.187427 | -737.846637 |
+| ... | ... | ... | ... | ... | ... |
+| 183 | 141590.0 | 120314.325647 | 25363.015190 | -2808.715638 | -1278.625199 |
+| 184 | 140610.0 | 120280.850692 | 26306.688690 | -6221.712712 | 244.173330 |
+| 185 | 139515.0 | 120247.361703 | 27571.777796 | -5745.053631 | -2559.085868 |
+
+```python
+sf.fitted_[0, 0].model_.tail(24 * 28).plot(subplots=True, grid=True)
+plt.tight_layout()
+plt.show()
+```
+
+
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 30 hours ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[90]` means that
+ the model expects the real value to be inside that interval 90% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min. (If you want to speed things up to a couple of seconds,
+remove the AutoModels like `ARIMA` and `Theta`)
+
+```python
+Y_hat = sf.forecast(df=train, h=horizon, fitted=True)
+Y_hat
+```
+
+| | unique_id | ds | MSTL |
+|-----|-----------|---------------------|---------------|
+| 0 | 1 | 2017-09-20 18:00:00 | 157848.500000 |
+| 1 | 1 | 2017-09-20 19:00:00 | 159790.328125 |
+| 2 | 1 | 2017-09-20 20:00:00 | 133002.281250 |
+| ... | ... | ... | ... |
+| 27 | 1 | 2017-09-21 21:00:00 | 98109.875000 |
+| 28 | 1 | 2017-09-21 22:00:00 | 86342.015625 |
+| 29 | 1 | 2017-09-21 23:00:00 | 76815.976562 |
+
+```python
+values=sf.forecast_fitted_values()
+values.head()
+```
+
+| | unique_id | ds | y | MSTL |
+|-----|-----------|---------------------|----------|---------------|
+| 0 | 1 | 2017-09-13 00:00:00 | 80115.0 | 79990.851562 |
+| 1 | 1 | 2017-09-13 01:00:00 | 79885.0 | 79329.132812 |
+| 2 | 1 | 2017-09-13 02:00:00 | 89325.0 | 88401.179688 |
+| 3 | 1 | 2017-09-13 03:00:00 | 101930.0 | 102109.929688 |
+| 4 | 1 | 2017-09-13 04:00:00 | 121630.0 | 123543.671875 |
+
+```python
+StatsForecast.plot(values)
+```
+
+
+
+Adding 95% confidence interval with the forecast method
+
+```python
+sf.forecast(df=train, h=horizon, level=[95])
+```
+
+| | unique_id | ds | MSTL | MSTL-lo-95 | MSTL-hi-95 |
+|-----|-----------|---------------------|---------------|---------------|---------------|
+| 0 | 1 | 2017-09-20 18:00:00 | 157848.500000 | 157796.406250 | 157900.593750 |
+| 1 | 1 | 2017-09-20 19:00:00 | 159790.328125 | 159714.218750 | 159866.437500 |
+| 2 | 1 | 2017-09-20 20:00:00 | 133002.281250 | 132893.937500 | 133110.609375 |
+| ... | ... | ... | ... | ... | ... |
+| 27 | 1 | 2017-09-21 21:00:00 | 98109.875000 | 95957.031250 | 100262.726562 |
+| 28 | 1 | 2017-09-21 22:00:00 | 86342.015625 | 85410.578125 | 87273.460938 |
+| 29 | 1 | 2017-09-21 23:00:00 | 76815.976562 | 73476.195312 | 80155.757812 |
+
+```python
+sf.plot(train, Y_hat)
+```
+
+
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 30 hours ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[95]` means that
+ the model expects the real value to be inside that interval 95% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+sf.predict(h=horizon)
+```
+
+| | unique_id | ds | MSTL |
+|-----|-----------|---------------------|---------------|
+| 0 | 1 | 2017-09-20 18:00:00 | 157848.500000 |
+| 1 | 1 | 2017-09-20 19:00:00 | 159790.328125 |
+| 2 | 1 | 2017-09-20 20:00:00 | 133002.281250 |
+| ... | ... | ... | ... |
+| 27 | 1 | 2017-09-21 21:00:00 | 98109.875000 |
+| 28 | 1 | 2017-09-21 22:00:00 | 86342.015625 |
+| 29 | 1 | 2017-09-21 23:00:00 | 76815.976562 |
+
+```python
+forecast_df = sf.predict(h=horizon, level=[80,95])
+forecast_df
+```
+
+| | unique_id | ds | MSTL | MSTL-lo-95 | MSTL-lo-80 | MSTL-hi-80 | MSTL-hi-95 |
+|-----|-----------|---------------------|---------------|---------------|---------------|---------------|---------------|
+| 0 | 1 | 2017-09-20 18:00:00 | 157848.500000 | 157796.406250 | 157798.484375 | 157898.531250 | 157900.593750 |
+| 1 | 1 | 2017-09-20 19:00:00 | 159790.328125 | 159714.218750 | 159716.187500 | 159864.468750 | 159866.437500 |
+| 2 | 1 | 2017-09-20 20:00:00 | 133002.281250 | 132893.937500 | 132894.515625 | 133110.031250 | 133110.609375 |
+| ... | ... | ... | ... | ... | ... | ... | ... |
+| 27 | 1 | 2017-09-21 21:00:00 | 98109.875000 | 95957.031250 | 96493.921875 | 99725.828125 | 100262.726562 |
+| 28 | 1 | 2017-09-21 22:00:00 | 86342.015625 | 85410.578125 | 85411.835938 | 87272.195312 | 87273.460938 |
+| 29 | 1 | 2017-09-21 23:00:00 | 76815.976562 | 73476.195312 | 74494.546875 | 79137.406250 | 80155.757812 |
+
+```python
+sf.plot(train, forecast_df, level=[80, 95])
+```
+
+
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=)`, forecasting every second months
+`(step_size=50)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 500 hours ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=df,
+ h=horizon,
+ step_size=30,
+ n_windows=5)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` series identifier
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the `n_windows`.
+- `y:` true value
+- `model:` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df
+```
+
+| | unique_id | ds | cutoff | y | MSTL |
+|-----|-----------|---------------------|---------------------|----------|---------------|
+| 0 | 1 | 2017-09-15 18:00:00 | 2017-09-15 17:00:00 | 159725.0 | 158384.250000 |
+| 1 | 1 | 2017-09-15 19:00:00 | 2017-09-15 17:00:00 | 161085.0 | 162015.171875 |
+| 2 | 1 | 2017-09-15 20:00:00 | 2017-09-15 17:00:00 | 135520.0 | 138495.093750 |
+| ... | ... | ... | ... | ... | ... |
+| 147 | 1 | 2017-09-21 21:00:00 | 2017-09-20 17:00:00 | 103080.0 | 98109.875000 |
+| 148 | 1 | 2017-09-21 22:00:00 | 2017-09-20 17:00:00 | 95155.0 | 86342.015625 |
+| 149 | 1 | 2017-09-21 23:00:00 | 2017-09-20 17:00:00 | 80285.0 | 76815.976562 |
+
+We’ll now plot the forecast for each cutoff period. To make the plots
+clearer, we’ll rename the actual values in each period.
+
+```python
+from IPython.display import display
+
+cross_validation=crossvalidation_df.copy()
+cross_validation.rename(columns = {'y' : 'actual'}, inplace = True) # rename actual values
+
+cutoff = cross_validation['cutoff'].unique()
+
+for k in range(len(cutoff)):
+ cv = cross_validation[cross_validation['cutoff'] == cutoff[k]]
+ display(StatsForecast.plot(df, cv.loc[:, cv.columns != 'cutoff']))
+```
+
+
+
+
+
+
+
+
+
+
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=24), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | MSTL |
+|-----|-----------|--------|-------------|
+| 0 | 1 | mae | 4932.395052 |
+| 1 | 1 | mape | 0.040514 |
+| 2 | 1 | mase | 0.609407 |
+| 3 | 1 | rmse | 6495.207028 |
+| 4 | 1 | smape | 0.020267 |
+
+## References
+
+1. [Changquan Huang • Alla Petukhina. Springer series (2022). Applied
+ Time Series Analysis and Forecasting with
+ Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2)
+2. Ivan Svetunkov. [Forecasting and Analytics with the Augmented
+ Dynamic Adaptive Model (ADAM)](https://openforecast.org/adam/)
+3. [James D. Hamilton. Time Series Analysis Princeton University Press,
+ Princeton, New Jersey, 1st Edition,
+ 1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Nixtla MultipleSeasonalTrend API](../../src/core/models.html#mstl)
+5. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+6. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+7. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/models/optimizedtheta.html.mdx b/statsforecast/docs/models/optimizedtheta.html.mdx
new file mode 100644
index 00000000..e5e226f3
--- /dev/null
+++ b/statsforecast/docs/models/optimizedtheta.html.mdx
@@ -0,0 +1,862 @@
+---
+title: Optimized Theta Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `OptimizedTheta Model` with
+> `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation` in other.
+
+The text in this article is largely taken from: 1. [Kostas I.
+Nikolopoulos, Dimitrios D. Thomakos. Forecasting with the Theta
+Method-Theory and Applications. 2019 John Wiley & Sons
+Ltd.](https://onlinelibrary.wiley.com/doi/book/10.1002/9781119320784) 2.
+[Jose A. Fiorucci, Tiago R. Pellegrini, Francisco Louzada, Fotios
+Petropoulos, Anne B. Koehler (2016). “Models for optimising the theta
+method and their relationship to state space models”. International
+Journal of
+Forecasting](https://www.sciencedirect.com/science/article/pii/S0169207016300243).
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [Optimized Theta Model (OTM)](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of OptimizedTheta with
+ StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+The optimized Theta model is a time series forecasting method that is
+based on the decomposition of the time series into three components:
+trend, seasonality and noise. The model then forecasts the long-term
+trend and seasonality, and uses the noise to adjust the short-term
+forecasts. The optimized Theta model has been shown to be more accurate
+than other time series forecasting methods, especially for time series
+with complex trends and seasonality.
+
+The optimized Theta model was developed by Athanasios N. Antoniadis and
+Nikolaos D. Tsonis in 2013. The model is based on the Theta forecasting
+method, which was developed by George E. P. Box and Gwilym M. Jenkins in
+1976. Theta method is a time series forecasting method that is based on
+the decomposition of the time series into three components: trend,
+seasonality, and noise. The Theta model then forecasts the long-term
+trend and seasonality, and uses the noise to adjust the short-term
+forecasts.
+
+The Theta Optimized model improves on the Theta method by using an
+optimization algorithm to find the best parameters for the model. The
+optimization algorithm is based on the Akaike loss function (AIC), which
+is a measure of the goodness of fit of a model to the data. The
+optimization algorithm looks for the parameters that minimize the AIC
+function.
+
+The optimized Theta model has been shown to be more accurate than other
+time series forecasting methods, especially for time series with complex
+trends and seasonality. The model has been used to forecast a variety of
+time series, including sales, production, prices, and weather.
+
+Below are some of the benefits of the optimized Theta model:
+
+- It is more accurate than other time series forecasting methods.
+- It’s easy to use.
+- Can be used to forecast a variety of time series.
+- It is flexible and can be adapted to different scenarios.
+
+If you are looking for an easy-to-use and accurate time series
+forecasting method, the Optimized Theta model is a good choice.
+
+The optimized Theta model can be applied in a variety of areas,
+including:
+
+- **Sales:** The optimized Theta model can be used to forecast sales
+ of products or services. This can help companies make decisions
+ about production, inventory, and marketing.
+- **Production:** The optimized Theta model can be used to forecast
+ the production of goods or services. This can help companies ensure
+ they have the capacity to meet demand and avoid overproduction.
+- **Prices:** The optimized Theta model can be used to forecast the
+ prices of goods or services. This can help companies make decisions
+ about pricing and marketing strategy.
+- **Weather:** The optimized Theta model can be used to forecast the
+ weather. This can help companies make decisions about agricultural
+ production, travel planning and risk management.
+- **Other:** The optimized Theta model can also be used to forecast
+ other types of time series, including traffic, energy demand, and
+ population.
+
+The Optimized Theta model is a powerful tool that can be used to improve
+the accuracy of time series forecasts. It is easy to use and can be
+applied to a variety of areas. If you are looking for a tool to improve
+your time series forecasts, the Optimized Theta model is a good choice.
+
+## Optimized Theta Model (OTM)
+
+Assume that either the time series $Y_1, \cdots Y_n$ is non-seasonal or
+it has been seasonally adjusted using the multiplicative classical
+decomposition approach.
+
+Let $X_t$ be the linear combination of two theta lines,
+
+$$
+\begin{equation}
+X_t=\omega \text{Z}_t (\theta_1) +(1-\omega) \text{Z}_t (\theta_2) \tag 1
+\end{equation}
+$$
+
+where $\omega \in [0,1]$ is the weight parameter. Assuming that
+$\theta_1 <1$ and $\theta_2 \geq 1$, the weight $\omega$ can be derived
+as
+
+$$
+\begin{equation}
+\omega:=\omega(\theta_1, \theta_2)=\frac{\theta_2 -1}{\theta_2 -\theta_1} \tag 2
+\end{equation}
+$$
+
+It is straightforward to see from Eqs. (1), (2) that
+$X_t=Y_t, \ t=1, \cdots n$ i.e., the weights are calculated properly in
+such a way that Eq. (1) reproduces the original series.
+
+**Theorem 1:** Let $\theta_1 <1$ and $\theta_2 \geq 1$. We will prove
+that
+
+1. the linear system given by $X_t=Y_t$ for all $t=1, \cdots, n$, where
+ $X_t$ is given by Eq.(4), has the single solution
+
+$$\omega= (\theta_2 -1)/(\theta_2 - \theta_1)$$
+
+1. the error of choosing a non-optimal weight
+ $\omega_{\delta} =\omega + \delta$ is proportional to the error for
+ a simple linear regression model.
+
+In Theorem 1 , we prove that the solution is unique and that the error
+from not choosing the optimal weights ($\omega$ and $1-\omega$) s
+proportional to the error of a linear regression model. As a
+consequence, the STheta method is given simply by setting $\theta_1=0$
+and $\theta_2=2$ while from Eq. (2) we get $\omega=0.5$. Thus, Eqs. (1),
+(2) allow us to construct a generalisation of the Theta model that
+maintains the re-composition propriety of the original time series for
+any theta lines $\text{Z}_t (\theta_1)$ and $\text{Z}_t (\theta_2)$.
+
+In order to maintain the modelling of the long-term component and retain
+a fair comparison with the STheta method, in this work we fix
+$\theta_1=0$ and focus on the optimisation of the short-term component,
+$\theta_2=0$ with $\theta \geq 1$. Thus, $\theta$ is the only parameter
+that requires estimation so far. The theta decomposition is now given by
+
+$$Y_t=(1-\frac{1}{\theta}) (\text{A}_n+\text{B}_n t)+ \frac{1}{\theta} \text{Z}_t (\theta), \ t=1, \cdots , n$$
+
+The $h$ -step-ahead forecasts calculated at origin are given by
+
+$$
+\begin{equation}
+\hat Y_{n+h|n} = (1-\frac{1}{\theta}) [\text{A}_n+\text{B}_n (n+h)]+ \frac{1}{\theta} \tilde {\text{Z}}_{n+h|n} (\theta) \tag 3
+\end{equation}
+$$
+
+where
+$\tilde {\text{Z}}_{n+h|n} (\theta)=\tilde {\text{Z}}_{n+1|n} (\theta)=\alpha \sum_{i=0}^{n-1}(1-\alpha)^i \text{Z}_{n-i}(\theta)+(1-\alpha)^n \ell_{0}^{*}$
+is the extrapolation of $\text{Z}_t(\theta)$ by an SES model with
+$\ell_{0}^{*} \in \mathbb{R}$ as the initial level parameter and
+$\alpha \in (0,1)$ as the smoothing parameter. Note that for $\theta=2$
+Eq. (3) corresponds to Step 4 of the STheta algorithm. After some
+algebra, we can write
+
+$$
+\begin{equation}
+\tilde {\text{Z}}_{n+1|n} (\theta)=\theta \ell{n}+(1-\theta) \{ \text{A}_n [1-(1-\alpha)^n] + \text{B}_n [n+(1-\frac{1}{\alpha}) [1-(1-\alpha)^n] ] \} \tag 4
+\end{equation}
+$$
+
+where $\ell_{t}=\alpha Y_t +(1-\alpha) \ell_{t-1}$ for $t=1, \cdots, n$
+and $\ell_{0}=\ell_{0}^{*}/\theta$.
+
+In the light of Eqs. (3), (4), we suggest four stochastic approaches.
+These approaches differ due to the parameter $\theta$ which may be
+either fixed at two or optimised, and the coefficients $\text{A}_n$ and
+$\text{B}_n$, which can be either fixed or dynamic functions. To
+formulate the state space models, it is helpful to adopt $\mu_{t}$ as
+the one-step-ahead forecast at origin $t-1$ and $\varepsilon_{t}$ as the
+respective additive error, i.e., $\varepsilon_{t}=Y_t - \mu_{t}$ if
+$\mu_{t}= \hat Y_{t|t-1}$. We assume $\{ \varepsilon_{t} \}$ to be a
+Gaussian white noise process with mean zero and variance $\sigma^2$.
+
+### More on Optimised Theta models
+
+Let $\text{A}_n$ and $\text{B}_n$ be fixed coefficients for all
+$t=1, \cdots, n$ so that Eqs. (3), (4) configure the state space model
+given by
+
+$$
+\begin{equation}
+Y_t=\mu_{t}+\varepsilon_{t} \tag 5
+\end{equation}
+$$
+
+$$
+\begin{equation}
+\mu_{t}=\ell_{t-1}+(1-\frac{1}{\theta}) \{(1-\alpha)^{t-1} \text{A}_n +[\frac{1-(1-\alpha)^t}{\alpha} \text{B}_n] \tag 6
+\end{equation}
+$$
+
+$$
+\begin{equation}
+\ell_{t}=\alpha Y_t +(1-\alpha)\ell_{t-1} \tag 7
+\end{equation}
+$$
+
+with parameters $\ell_{0} \in \mathbb{R}$, $\alpha \in (0,1)$ and
+$\theta \in [1,\infty)$ . The parameter $\theta$ is to be estimated
+along with $\alpha$ and $\ell_{0}$ We call this the optimised Theta
+model (OTM).
+
+The $h$-step-ahead forecast at origin $n$ is given by
+
+$$\hat Y_{n+h|n}=E[Y_{n+h}|Y_1,\cdots, Y_n]=\ell_{n}+(1-\frac{1}{\theta}) \{(1-\alpha)^n \text{A}_n +[(h-1) + \frac{1-(1-\alpha)^{n+1}}{\alpha}] \text{B}_n \}$$
+
+which is equivalent to Eq. (3). The conditional variance
+$\text{Var}[Y_{n+h}|Y_1, \cdots, Y_n]=[1+(h-1)\alpha^2]\sigma^2$ can be
+computed easily from the state space model. Thus, the $(1-\alpha)\%$
+prediction interval for $Y_{n+h}$ is given by
+$$\hat Y_{n+h|n} \ \pm \ q_{1-\alpha/2} \sqrt{[1+(h-1)\alpha^2 ]\sigma^2 }$$
+
+For $\theta=2$ OTM reproduces the forecasts of the STheta method;
+hereafter, we will refer to this particular case as the standard Theta
+model (STM).
+
+**Theorem 2:** The SES-d $(\ell_{0}^{**}, \alpha, b)$ model, where
+$\ell_{0}^{**} \in \mathbb{R}, \alpha \in (0,1)$ and $b \in \mathbb{R}$
+is equivalent to $\text{OTM} (\ell_{0}, \alpha, \theta )$ where
+$\ell_{0} \in \mathbb{R}$ and $\theta \geq 1$, if
+
+$$\ell_{0}^{**} = \ell_{0} + (1- \frac{1}{\theta} )A_n \ \ and \ \ b=(1-\frac{1}{\theta} )B_n$$
+
+In Theorem 2, we show that OTM is mathematically equivalent to the SES-d
+model. As a corollary of Theorem 2, STM is mathematically equivalent to
+SES-d with $b=\frac{1}{2} \text{B}_n$. Therefore, for $\theta=2$ the
+corollary also re-confirms the H&B result on the relationship between
+STheta and the SES-d model.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
+plt.style.use('grayscale') # fivethirtyeight grayscale classic
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#008080', # #212946
+ 'axes.facecolor': '#008080',
+ 'savefig.facecolor': '#008080',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#000000', #2A3459
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+### Read Data
+
+```python
+import pandas as pd
+
+df = pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/milk_production.csv", usecols=[1,2])
+df.head()
+```
+
+| | month | production |
+|-----|------------|------------|
+| 0 | 1962-01-01 | 589 |
+| 1 | 1962-02-01 | 561 |
+| 2 | 1962-03-01 | 640 |
+| 3 | 1962-04-01 | 656 |
+| 4 | 1962-05-01 | 727 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|------------|-----|-----------|
+| 0 | 1962-01-01 | 589 | 1 |
+| 1 | 1962-02-01 | 561 | 1 |
+| 2 | 1962-03-01 | 640 | 1 |
+| 3 | 1962-04-01 | 656 | 1 |
+| 4 | 1962-05-01 | 727 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y int64
+unique_id object
+dtype: object
+```
+
+We can see that our time variable `(ds)` is in an object format, we need
+to convert to a date format
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore Data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints a random series from the dataset and is useful for
+basic EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### Autocorrelation plots
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=30, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+plot_pacf(df["y"], lags=30, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+plt.show();
+```
+
+
+
+### Decomposition of the time series
+
+How to decompose a time series and why?
+
+In time series analysis to forecast new values, it is very important to
+know past data. More formally, we can say that it is very important to
+know the patterns that values follow over time. There can be many
+reasons that cause our forecast values to fall in the wrong direction.
+Basically, a time series consists of four components. The variation of
+those components causes the change in the pattern of the time series.
+These components are:
+
+- **Level:** This is the primary value that averages over time.
+- **Trend:** The trend is the value that causes increasing or
+ decreasing patterns in a time series.
+- **Seasonality:** This is a cyclical event that occurs in a time
+ series for a short time and causes short-term increasing or
+ decreasing patterns in a time series.
+- **Residual/Noise:** These are the random variations in the time
+ series.
+
+Combining these components over time leads to the formation of a time
+series. Most time series consist of level and noise/residual and trend
+or seasonality are optional values.
+
+If seasonality and trend are part of the time series, then there will be
+effects on the forecast value. As the pattern of the forecasted time
+series may be different from the previous time series.
+
+The combination of the components in time series can be of two types: \*
+Additive \* Multiplicative
+
+### Additive time series
+
+If the components of the time series are added to make the time series.
+Then the time series is called the additive time series. By
+visualization, we can say that the time series is additive if the
+increasing or decreasing pattern of the time series is similar
+throughout the series. The mathematical function of any additive time
+series can be represented by:
+$$y(t) = level + Trend + seasonality + noise$$
+
+### Multiplicative time series
+
+If the components of the time series are multiplicative together, then
+the time series is called a multiplicative time series. For
+visualization, if the time series is having exponential growth or
+decline with time, then the time series can be considered as the
+multiplicative time series. The mathematical function of the
+multiplicative time series can be represented as.
+
+$$y(t) = Level * Trend * seasonality * Noise$$
+
+### Additive
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+a = seasonal_decompose(df["y"], model = "additive", period=12)
+a.plot();
+```
+
+
+
+### Multiplicative
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+a = seasonal_decompose(df["y"], model = "Multiplicative", period=12)
+a.plot();
+```
+
+
+
+## Split the data into training and testing
+
+Let’s divide our data into sets 1. Data to train our
+`Optimized Theta model`. 2. Data to test our model
+
+For the test data we will use the last 12 months to test and evaluate
+the performance of our model.
+
+```python
+train = df[df.ds\<='1974-12-01']
+test = df[df.ds>'1974-12-01']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((156, 3), (12, 3))
+```
+
+## Implementation of OptimizedTheta with StatsForecast
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import OptimizedTheta
+```
+
+### Instantiating Model
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/) by the
+master, Rob Hyndmann, can be useful for `season_length`.
+
+```python
+season_length = 12 # Monthly data
+horizon = len(test) # number of predictions
+
+models = [OptimizedTheta(season_length=season_length,
+ decomposition_type="additive")] # multiplicative additive
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [pandas’
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models, freq='MS')
+```
+
+### Fit the Model
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[OptimizedTheta])
+```
+
+Let’s see the results of our `Optimized Theta Model (OTM)`. We can
+observe it with the following instruction:
+
+```python
+result=sf.fitted_[0,0].model_
+print(result.keys())
+print(result['fit'])
+```
+
+``` text
+dict_keys(['mse', 'amse', 'fit', 'residuals', 'm', 'states', 'par', 'n', 'modeltype', 'mean_y', 'decompose', 'decomposition_type', 'seas_forecast', 'fitted'])
+results(x=array([-83.14191626, 0.73681394, 12.45013763]), fn=10.448217519858636, nit=47, simplex=array([[-58.73988124, 0.7441127 , 11.69842922],
+ [-49.97233449, 0.73580297, 11.41787513],
+ [-83.14191626, 0.73681394, 12.45013763],
+ [-77.04867427, 0.73498431, 11.99254037]]))
+```
+
+Let us now visualize the residuals of our models.
+
+As we can see, the result obtained above has an output in a dictionary,
+to extract each element from the dictionary we are going to use the
+`.get()` function to extract the element and then we are going to save
+it in a `pd.DataFrame()`.
+
+```python
+residual=pd.DataFrame(result.get("residuals"), columns=["residual Model"])
+residual
+```
+
+| | residual Model |
+|-----|----------------|
+| 0 | -271.899414 |
+| 1 | -114.671692 |
+| 2 | 4.768066 |
+| ... | ... |
+| 153 | -60.233887 |
+| 154 | -92.472839 |
+| 155 | -44.143982 |
+
+```python
+import scipy.stats as stats
+
+fig, axs = plt.subplots(nrows=2, ncols=2)
+
+residual.plot(ax=axs[0,0])
+axs[0,0].set_title("Residuals");
+
+sns.distplot(residual, ax=axs[0,1]);
+axs[0,1].set_title("Density plot - Residual");
+
+stats.probplot(residual["residual Model"], dist="norm", plot=axs[1,0])
+axs[1,0].set_title('Plot Q-Q')
+
+plot_acf(residual, lags=35, ax=axs[1,1],color="fuchsia")
+axs[1,1].set_title("Autocorrelation");
+
+plt.show();
+```
+
+
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[90]` means that
+ the model expects the real value to be inside that interval 90% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min. (If you want to speed things up to a couple of seconds,
+remove the AutoModels like `ARIMA` and `Theta`)
+
+```python
+Y_hat = sf.forecast(df=train, h=horizon, fitted=True)
+Y_hat
+```
+
+| | unique_id | ds | OptimizedTheta |
+|-----|-----------|------------|----------------|
+| 0 | 1 | 1975-01-01 | 839.682800 |
+| 1 | 1 | 1975-02-01 | 802.071838 |
+| 2 | 1 | 1975-03-01 | 896.117126 |
+| ... | ... | ... | ... |
+| 9 | 1 | 1975-10-01 | 824.135498 |
+| 10 | 1 | 1975-11-01 | 795.691223 |
+| 11 | 1 | 1975-12-01 | 833.316345 |
+
+Let’s visualize the fitted values
+
+```python
+values=sf.forecast_fitted_values()
+values.head()
+```
+
+| | unique_id | ds | y | OptimizedTheta |
+|-----|-----------|------------|-------|----------------|
+| 0 | 1 | 1962-01-01 | 589.0 | 860.899414 |
+| 1 | 1 | 1962-02-01 | 561.0 | 675.671692 |
+| 2 | 1 | 1962-03-01 | 640.0 | 635.231934 |
+| 3 | 1 | 1962-04-01 | 656.0 | 614.731323 |
+| 4 | 1 | 1962-05-01 | 727.0 | 609.770752 |
+
+```python
+StatsForecast.plot(values)
+```
+
+
+
+Adding 95% confidence interval with the forecast method
+
+```python
+sf.forecast(df=train, h=horizon, level=[95])
+```
+
+| | unique_id | ds | OptimizedTheta | OptimizedTheta-lo-95 | OptimizedTheta-hi-95 |
+|-----|-----------|------------|----------------|----------------------|----------------------|
+| 0 | 1 | 1975-01-01 | 839.682800 | 742.509583 | 955.414307 |
+| 1 | 1 | 1975-02-01 | 802.071838 | 643.581360 | 945.119202 |
+| 2 | 1 | 1975-03-01 | 896.117126 | 710.785095 | 1065.057495 |
+| ... | ... | ... | ... | ... | ... |
+| 9 | 1 | 1975-10-01 | 824.135498 | 555.948669 | 1084.320190 |
+| 10 | 1 | 1975-11-01 | 795.691223 | 503.147858 | 1036.519531 |
+| 11 | 1 | 1975-12-01 | 833.316345 | 530.259705 | 1106.636597 |
+
+```python
+sf.plot(train, Y_hat)
+```
+
+
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[95]` means that
+ the model expects the real value to be inside that interval 95% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+sf.predict(h=horizon)
+```
+
+| | unique_id | ds | OptimizedTheta |
+|-----|-----------|------------|----------------|
+| 0 | 1 | 1975-01-01 | 839.682800 |
+| 1 | 1 | 1975-02-01 | 802.071838 |
+| 2 | 1 | 1975-03-01 | 896.117126 |
+| ... | ... | ... | ... |
+| 9 | 1 | 1975-10-01 | 824.135498 |
+| 10 | 1 | 1975-11-01 | 795.691223 |
+| 11 | 1 | 1975-12-01 | 833.316345 |
+
+```python
+forecast_df = sf.predict(h=horizon, level=[80,95])
+forecast_df
+```
+
+| | unique_id | ds | OptimizedTheta | OptimizedTheta-lo-80 | OptimizedTheta-hi-80 | OptimizedTheta-lo-95 | OptimizedTheta-hi-95 |
+|-----|-----------|------------|----------------|----------------------|----------------------|----------------------|----------------------|
+| 0 | 1 | 1975-01-01 | 839.682800 | 766.665955 | 928.326172 | 742.509583 | 955.414307 |
+| 1 | 1 | 1975-02-01 | 802.071838 | 704.290039 | 899.335815 | 643.581360 | 945.119202 |
+| 2 | 1 | 1975-03-01 | 896.117126 | 761.334778 | 1007.408447 | 710.785095 | 1065.057495 |
+| ... | ... | ... | ... | ... | ... | ... | ... |
+| 9 | 1 | 1975-10-01 | 824.135498 | 623.903992 | 996.567200 | 555.948669 | 1084.320190 |
+| 10 | 1 | 1975-11-01 | 795.691223 | 576.546570 | 975.490784 | 503.147858 | 1036.519531 |
+| 11 | 1 | 1975-12-01 | 833.316345 | 606.713623 | 1033.885742 | 530.259705 | 1106.636597 |
+
+```python
+sf.plot(train, test.merge(forecast_df), level=[80, 95])
+```
+
+
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=5)`, forecasting every second months
+`(step_size=12)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 12 months ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=train,
+ h=horizon,
+ step_size=12,
+ n_windows=3)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` index. If you dont like working with index just run
+ crossvalidation_df.resetindex()
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the n_windows.
+- `y:` true value
+- `"model":` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df
+```
+
+| | unique_id | ds | cutoff | y | OptimizedTheta |
+|-----|-----------|------------|------------|-------|----------------|
+| 0 | 1 | 1972-01-01 | 1971-12-01 | 826.0 | 828.836365 |
+| 1 | 1 | 1972-02-01 | 1971-12-01 | 799.0 | 792.592346 |
+| 2 | 1 | 1972-03-01 | 1971-12-01 | 890.0 | 883.269592 |
+| ... | ... | ... | ... | ... | ... |
+| 33 | 1 | 1974-10-01 | 1973-12-01 | 812.0 | 812.183838 |
+| 34 | 1 | 1974-11-01 | 1973-12-01 | 773.0 | 783.898376 |
+| 35 | 1 | 1974-12-01 | 1973-12-01 | 813.0 | 821.124329 |
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=season_length), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | OptimizedTheta |
+|-----|-----------|--------|----------------|
+| 0 | 1 | mae | 6.740204 |
+| 1 | 1 | mape | 0.007828 |
+| 2 | 1 | mase | 0.303120 |
+| 3 | 1 | rmse | 8.701501 |
+| 4 | 1 | smape | 0.003893 |
+
+## References
+
+1. [Kostas I. Nikolopoulos, Dimitrios D. Thomakos. Forecasting with the
+ Theta Method-Theory and Applications. 2019 John Wiley & Sons
+ Ltd.](https://onlinelibrary.wiley.com/doi/book/10.1002/9781119320784)
+2. [Jose A. Fiorucci, Tiago R. Pellegrini, Francisco Louzada, Fotios
+ Petropoulos, Anne B. Koehler (2016). “Models for optimising the
+ theta method and their relationship to state space models”.
+ International Journal of
+ Forecasting](https://www.sciencedirect.com/science/article/pii/S0169207016300243).
+3. [Nixtla OptimizedTheta API](../../src/core/models.html#optimizedtheta)
+4. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+5. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+6. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/models/seasonalexponentialsmoothing.html.mdx b/statsforecast/docs/models/seasonalexponentialsmoothing.html.mdx
new file mode 100644
index 00000000..8278a36c
--- /dev/null
+++ b/statsforecast/docs/models/seasonalexponentialsmoothing.html.mdx
@@ -0,0 +1,888 @@
+---
+title: Seasonal Exponential Smoothing Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `SeasonalExponentialSmoothing Model`
+> with `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation` in other.
+
+The text in this article is largely taken from: 1. [Changquan Huang •
+Alla Petukhina. Springer series (2022). Applied Time Series Analysis and
+Forecasting with
+Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2) 2.
+Ivan Svetunkov. [Forecasting and Analytics with the Augmented Dynamic
+Adaptive Model (ADAM)](https://openforecast.org/adam/) 3. [James D.
+Hamilton. Time Series Analysis Princeton University Press, Princeton,
+New Jersey, 1st Edition,
+1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+Principles and Practice (3rd ed)”](https://otexts.com/fpp3/tscv.html).
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [Seasonal Exponential Smoothing](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of SeasonalExponentialSmoothing with
+ StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+Simple Exponential Smoothing (SES) is a forecasting method that uses a
+weighted average of historical values to predict the next value. The
+weight is assigned to the most recent values, and the oldest values
+receive a lower weight. This is because SES assumes that more recent
+values are more relevant to predicting the future than older values.
+
+SES is implemented by a simple formula:
+
+$$\hat{y}_{T+1|T} = \alpha y_T + \alpha(1-\alpha) y_{T-1} + \alpha(1-\alpha)^2 y_{T-2}+ \cdots,$$
+
+The smoothing factor controls the amount of weight that is assigned to
+the most recent values. A higher α value means more weight will be
+assigned to newer values, while a lower α value means more weight will
+be assigned to older values.
+
+Seasonality in time series refers to the regular, repeating pattern of
+variation in a time series over a specified period of time.
+
+Seasonality can be a challenge to deal with in time series analysis, as
+it can obscure the underlying trend in the data.
+
+Seasonality is an important factor to consider when analyzing time
+series data. By understanding the seasonal patterns in the data, it is
+possible to make more accurate forecasts and better decisions.
+
+## Seasonal Exponential Smoothing Model
+
+The simplest of the exponentially smoothing methods is naturally called
+simple exponential smoothing (SES). This method is suitable for
+forecasting data with no clear trend or seasonal pattern.
+
+Using the naïve method, all forecasts for the future are equal to the
+last observed value of the series, $$\hat{y}_{T+h|T} = y_{T},$$
+
+for $h=1,2,\dots$. Hence, the naïve method assumes that the most recent
+observation is the only important one, and all previous observations
+provide no information for the future. This can be thought of as a
+weighted average where all of the weight is given to the last
+observation.
+
+Using the average method, all future forecasts are equal to a simple
+average of the observed data,
+$$\hat{y}_{T+h|T} = \frac1T \sum_{t=1}^T y_t,$$
+
+for $h=1,2,\dots$ Hence, the average method assumes that all
+observations are of equal importance, and gives them equal weights when
+generating forecasts.
+
+We often want something between these two extremes. For example, it may
+be sensible to attach larger weights to more recent observations than to
+observations from the distant past. This is exactly the concept behind
+simple exponential smoothing. Forecasts are calculated using weighted
+averages, where the weights decrease exponentially as observations come
+from further in the past — the smallest weights are associated with the
+oldest observations:
+
+where $0 \le \alpha \le 1$ is the smoothing parameter. The
+one-step-ahead forecast for time $T+1$ is a weighted average of all of
+the observations in the series $y_1,\dots,y_T$. The rate at which the
+weights decrease is controlled by the parameter $\alpha$.
+
+For any $\alpha$ between 0 and 1, the weights attached to the
+observations decrease exponentially as we go back in time, hence the
+name “exponential smoothing”. If $\alpha$ is small (i.e., close to 0),
+more weight is given to observations from the more distant past. If
+$\alpha$ is large (i.e., close to 1), more weight is given to the more
+recent observations. For the extreme case where $\alpha=1$,
+$\hat{y}_{T+1|T}=y_T$ and the forecasts are equal to the naïve
+forecasts.
+
+### How do you know the value of the seasonal parameters?
+
+To determine the value of the seasonal parameter s in the Seasonally
+Adjusted `Simple Exponential Smoothing (SES Seasonally Adjusted) model`,
+different methods can be used, depending on the nature of the data and
+the objective of the analysis.
+
+Here are some common methods to determine the value of the seasonal
+parameter $s$:
+
+1. **Visual Analysis:** A visual analysis of the time series data can
+ be performed to identify any seasonal patterns. If a clear seasonal
+ pattern is observed in the data, the length of the seasonal period
+ can be used as the value of $s$.
+
+2. **Statistical methods:** Statistical techniques, such as
+ autocorrelation, can be used to identify seasonal patterns in the
+ data. The value of $s$ can be the number of periods in which a
+ significant peak in the autocorrelation function is observed.
+
+3. **Frequency Analysis:** A frequency analysis of the data can be
+ performed to identify seasonal patterns. The value of $s$ can be the
+ number of periods in which a significant peak in the frequency
+ spectrum is observed.
+ [see](https://robjhyndman.com/hyndsight/seasonal-periods/)
+
+4. **Trial and error:** You can try different values of $s$ and select
+ the value that results in the best fit of the model to the data.
+
+It is important to note that the choice of the value of $s$ can
+significantly affect the `accuracy` of the seasonally adjusted SES model
+predictions. Therefore, it is recommended to test different values of
+$s$ and evaluate the performance of the model using appropriate
+evaluation measures before selecting the final value of $s$.
+
+### How can we validate the simple exponential smoothing model with seasonal adjustment?
+
+To validate the Seasonally Adjusted Simple Exponential Smoothing (SES
+Seasonally Adjusted) model, different theorems and evaluation measures
+can be used, depending on the objective of the analysis and the nature
+of the data.
+
+Here are some common theorems used to validate the seasonally adjusted
+SES model:
+
+1. Gauss-Markov Theorem: This theorem states that, if certain
+ conditions are met, the least squares estimator is the best linear
+ unbiased estimator. In the case of the seasonally adjusted SES, the
+ model parameters are estimated using least squares, so the
+ Gauss-Markov theorem can be used to assess the quality of model fit.
+
+2. Unit Root Theorem: This theorem is used to determine if a time
+ series is stationary or not. If a time series is non-stationary, the
+ seasonally adjusted SES model is not appropriate, since it assumes
+ that the time series is stationary. Therefore, the unit root theorem
+ is used to assess the stationarity of the time series and determine
+ whether the seasonally adjusted SES model is appropriate.
+
+3. Ljung-Box Theorem: This theorem is used to assess the goodness of
+ fit of the model and to determine if the model residuals are white
+ noise. If the residuals are white noise, the model fits the data
+ well and the model predictions are accurate. The Ljung-Box theorem
+ is used to test whether the model residuals are independent and
+ uncorrelated.
+
+In addition to these theorems, various evaluation measures, such as root
+mean square error (MSE), mean absolute error (MAE), and coefficient of
+determination (R²), can be used to evaluate the performance of the
+seasonally adjusted SES model and compare it with other forecast models.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
+plt.style.use('grayscale') # fivethirtyeight grayscale classic
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#008080', # #212946
+ 'axes.facecolor': '#008080',
+ 'savefig.facecolor': '#008080',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#000000', #2A3459
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+### Read Data
+
+```python
+import pandas as pd
+
+df=pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/ads.csv")
+df.head()
+```
+
+| | Time | Ads |
+|-----|---------------------|--------|
+| 0 | 2017-09-13T00:00:00 | 80115 |
+| 1 | 2017-09-13T01:00:00 | 79885 |
+| 2 | 2017-09-13T02:00:00 | 89325 |
+| 3 | 2017-09-13T03:00:00 | 101930 |
+| 4 | 2017-09-13T04:00:00 | 121630 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|---------------------|--------|-----------|
+| 0 | 2017-09-13T00:00:00 | 80115 | 1 |
+| 1 | 2017-09-13T01:00:00 | 79885 | 1 |
+| 2 | 2017-09-13T02:00:00 | 89325 | 1 |
+| 3 | 2017-09-13T03:00:00 | 101930 | 1 |
+| 4 | 2017-09-13T04:00:00 | 121630 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y int64
+unique_id object
+dtype: object
+```
+
+We can see that our time variable `(ds)` is in an object format, we need
+to convert to a date format
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore Data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints a random series from the dataset and is useful for
+basic EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### The Augmented Dickey-Fuller Test
+
+An Augmented Dickey-Fuller (ADF) test is a type of statistical test that
+determines whether a unit root is present in time series data. Unit
+roots can cause unpredictable results in time series analysis. A null
+hypothesis is formed in the unit root test to determine how strongly
+time series data is affected by a trend. By accepting the null
+hypothesis, we accept the evidence that the time series data is not
+stationary. By rejecting the null hypothesis or accepting the
+alternative hypothesis, we accept the evidence that the time series data
+is generated by a stationary process. This process is also known as
+stationary trend. The values of the ADF test statistic are negative.
+Lower ADF values indicate a stronger rejection of the null hypothesis.
+
+Augmented Dickey-Fuller Test is a common statistical test used to test
+whether a given time series is stationary or not. We can achieve this by
+defining the null and alternate hypothesis.
+
+Null Hypothesis: Time Series is non-stationary. It gives a
+time-dependent trend. Alternate Hypothesis: Time Series is stationary.
+In another term, the series doesn’t depend on time.
+
+ADF or t Statistic \< critical values: Reject the null hypothesis, time
+series is stationary. ADF or t Statistic \> critical values: Failed to
+reject the null hypothesis, time series is non-stationary.
+
+```python
+from statsmodels.tsa.stattools import adfuller
+
+def Augmented_Dickey_Fuller_Test_func(series , column_name):
+ print (f'Dickey-Fuller test results for columns: {column_name}')
+ dftest = adfuller(series, autolag='AIC')
+ dfoutput = pd.Series(dftest[0:4], index=['Test Statistic','p-value','No Lags Used','Number of observations used'])
+ for key,value in dftest[4].items():
+ dfoutput['Critical Value (%s)'%key] = value
+ print (dfoutput)
+ if dftest[1] \<= 0.05:
+ print("Conclusion:====>")
+ print("Reject the null hypothesis")
+ print("The data is stationary")
+ else:
+ print("Conclusion:====>")
+ print("The null hypothesis cannot be rejected")
+ print("The data is not stationary")
+```
+
+
+```python
+Augmented_Dickey_Fuller_Test_func(df["y"],'Ads')
+```
+
+``` text
+Dickey-Fuller test results for columns: Ads
+Test Statistic -7.089634e+00
+p-value 4.444804e-10
+No Lags Used 9.000000e+00
+ ...
+Critical Value (1%) -3.462499e+00
+Critical Value (5%) -2.875675e+00
+Critical Value (10%) -2.574304e+00
+Length: 7, dtype: float64
+Conclusion:====>
+Reject the null hypothesis
+The data is stationary
+```
+
+### Autocorrelation plots
+
+The important characteristics of Autocorrelation (ACF) and Partial
+Autocorrelation (PACF) are as follows:
+
+Autocorrelation (ACF): 1. Identify patterns of temporal dependence: The
+ACF shows the correlation between an observation and its lagged values
+at different time intervals. Helps identify patterns of temporal
+dependency in a time series, such as the presence of trends or
+seasonality.
+
+1. Indicates the “memory” of the series: The ACF allows us to determine
+ how much past observations influence future ones. If the ACF shows
+ significant autocorrelations in several lags, it indicates that the
+ series has a long-term memory and that past observations are
+ relevant to predict future ones.
+
+2. Helps identify MA (moving average) models: The shape of the ACF can
+ reveal the presence of moving average components in the time series.
+ Lags where the ACF shows a significant correlation may indicate the
+ order of an MA model.
+
+Partial Autocorrelation (PACF): 1. Identify direct dependence: Unlike
+the ACF, the PACF eliminates the indirect effects of intermediate lags
+and measures the direct correlation between an observation and its
+lagged values. It helps to identify the direct dependence between an
+observation and its lag values, without the influence of intermediate
+lags.
+
+1. Helps to identify AR (autoregressive) models: The shape of the PACF
+ can reveal the presence of autoregressive components in the time
+ series. Lags in which the PACF shows a significant correlation may
+ indicate the order of an AR model.
+
+2. Used in conjunction with the ACF: The PACF is used in conjunction
+ with the ACF to determine the order of an AR or MA model. By
+ analyzing both the ACF and the PACF, significant lags can be
+ identified and a model suitable for time series analysis and
+ forecasting can be built.
+
+In summary, the ACF and the PACF are complementary tools in time series
+analysis that provide information on time dependence and help identify
+the appropriate components to build forecast models.
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=30, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+plot_pacf(df["y"], lags=30, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+plt.show();
+```
+
+
+
+### Decomposition of the time series
+
+How to decompose a time series and why?
+
+In time series analysis to forecast new values, it is very important to
+know past data. More formally, we can say that it is very important to
+know the patterns that values follow over time. There can be many
+reasons that cause our forecast values to fall in the wrong direction.
+Basically, a time series consists of four components. The variation of
+those components causes the change in the pattern of the time series.
+These components are:
+
+- **Level:** This is the primary value that averages over time.
+- **Trend:** The trend is the value that causes increasing or
+ decreasing patterns in a time series.
+- **Seasonality:** This is a cyclical event that occurs in a time
+ series for a short time and causes short-term increasing or
+ decreasing patterns in a time series.
+- **Residual/Noise:** These are the random variations in the time
+ series.
+
+Combining these components over time leads to the formation of a time
+series. Most time series consist of level and noise/residual and trend
+or seasonality are optional values.
+
+If seasonality and trend are part of the time series, then there will be
+effects on the forecast value. As the pattern of the forecasted time
+series may be different from the previous time series.
+
+The combination of the components in time series can be of two types: \*
+Additive \* Multiplicative
+
+### Additive time series
+
+If the components of the time series are added to make the time series.
+Then the time series is called the additive time series. By
+visualization, we can say that the time series is additive if the
+increasing or decreasing pattern of the time series is similar
+throughout the series. The mathematical function of any additive time
+series can be represented by:
+$$y(t) = level + Trend + seasonality + noise$$
+
+### Multiplicative time series
+
+If the components of the time series are multiplicative together, then
+the time series is called a multiplicative time series. For
+visualization, if the time series is having exponential growth or
+decline with time, then the time series can be considered as the
+multiplicative time series. The mathematical function of the
+multiplicative time series can be represented as.
+
+$$y(t) = Level * Trend * seasonality * Noise$$
+
+### Additive
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+a = seasonal_decompose(df["y"], model = "additive", period=12)
+a.plot();
+```
+
+
+
+### Multiplicative
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+a = seasonal_decompose(df["y"], model = "Multiplicative", period=12)
+a.plot();
+```
+
+
+
+## Split the data into training and testing
+
+Let’s divide our data into sets
+
+1. Data to train our `Seasonal Exponential Smoothing Model`.
+2. Data to test our model
+
+For the test data we will use the last 30 hourly to test and evaluate
+the performance of our model.
+
+```python
+train = df[df.ds\<='2017-09-20 17:00:00']
+test = df[df.ds>'2017-09-20 17:00:00']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((186, 3), (30, 3))
+```
+
+Now let’s plot the training data and the test data.
+
+```python
+sns.lineplot(train,x="ds", y="y", label="Train", linestyle="--")
+sns.lineplot(test, x="ds", y="y", label="Test")
+plt.title("Ads watched (hourly data)");
+plt.show()
+```
+
+
+
+## Implementation of SeasonalExponentialSmoothing with StatsForecast
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import SeasonalExponentialSmoothing
+```
+
+### Instantiating Model
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/) by the
+master, Rob Hyndmann, can be useful for `season_length`.
+
+```python
+season_length = 24 # Hourly data
+horizon = len(test) # number of predictions
+
+models = [SeasonalExponentialSmoothing(alpha=0.8, season_length=season_length)]
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [pandas’
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models, freq='h')
+```
+
+### Fit the Model
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[SeasonalES])
+```
+
+Let’s see the results of our `Seasonal Exponential Smoothing Model`. We
+can observe it with the following instruction:
+
+```python
+result=sf.fitted_[0,0].model_
+result
+```
+
+``` text
+{'mean': array([161567.6 , 163186.56 , 134410.94 , 106145.6 , 93383.164,
+ 79489.72 , 79769. , 77651.984, 85288.33 , 99665.31 ,
+ 123067.336, 115759.51 , 103556.234, 100510.09 , 97411.65 ,
+ 107672.88 , 121150.84 , 140041.22 , 140075.19 , 140903.34 ,
+ 142615.28 , 142360.75 , 142615.6 , 142658.62 ], dtype=float32),
+ 'fitted': array([ nan, nan, nan, nan, nan,
+ nan, nan, nan, nan, nan,
+ nan, nan, nan, nan, nan,
+ nan, nan, nan, nan, nan,
+ nan, nan, nan, nan, nan,
+ nan, nan, nan, nan, nan,
+ nan, nan, nan, nan, nan,
+ nan, nan, nan, nan, nan,
+ nan, nan, 163840. , 166235. , 139520. ,
+ 105895. , 96780. , 82520. , 80125. , 75335. ,
+ 85105. , 102080. , 125135. , 118030. , 109225. ,
+ 102475. , 102240. , 115840. , 130540. , 144325. ,
+ 148970. , 149150. , 148040. , 148810. , 149830. ,
+ 150570. , 152320. , 153663. , 131208. , 104231. ,
+ 93096. , 82716. , 77077. , 75171. , 83133. ,
+ 91452. , 119771. , 115758. , 110597. , 99583. ,
+ 103796. , 110100. , 127420. , 141213. , 151770. ,
+ 146850. , 148024. , 147950. , 146566. , 149542. ,
+ 158244. , 159600.6 , 134657.6 , 111202.2 , 98779.2 ,
+ 86635.2 , 85683.4 , 86110.2 , 90506.6 , 101862.4 ,
+ 116706.2 , 126311.6 , 135227.4 , 135468.6 , 135359.2 ,
+ 128572. , 130476. , 142226.6 , 156254. , 151370. ,
+ 152592.8 , 150546. , 149829.2 , 147856.4 , 153668.8 ,
+ 149420.12 , 127127.52 , 116580.44 , 97115.84 , 92215.04 ,
+ 88384.68 , 88698.04 , 90561.32 , 99004.48 , 113397.24 ,
+ 128838.32 , 140169.48 , 149141.72 , 149135.84 , 138650.4 ,
+ 144135.2 , 157333.31 , 164318.8 , 163698. , 161030.56 ,
+ 155953.2 , 157209.84 , 157587.28 , 165409.77 , 165804.03 ,
+ 139593.5 , 113680.086, 97299.17 , 83783.01 , 81284.94 ,
+ 80419.61 , 88548.266, 99632.9 , 121703.445, 114827.664,
+ 107585.9 , 107952.34 , 107951.17 , 109782.08 , 124771.04 ,
+ 140070.66 , 144959.77 , 146123.6 , 145982.11 , 147478.64 ,
+ 147709.97 , 151845.45 , 162297.95 , 155892.81 , 135694.7 ,
+ 108388.016, 95495.836, 80368.6 , 78924.984, 75819.92 ,
+ 83301.66 , 98286.58 , 119816.69 , 113457.53 , 100621.18 ,
+ 96790.47 , 96518.234, 105304.414, 120754.21 , 136806.12 ,
+ 146155.95 , 140556.72 , 146976.42 , 145443.73 , 150638. ,
+ 155233.1 ], dtype=float32)}
+```
+
+Let us now visualize the fitted values of our models.
+
+As we can see, the result obtained above has an output in a dictionary,
+to extract each element from the dictionary we are going to use the
+`.get()` function to extract the element and then we are going to save
+it in a `pd.DataFrame()`.
+
+```python
+fitted=pd.DataFrame(result.get("fitted"), columns=["fitted"])
+fitted["ds"]=df["ds"]
+fitted
+```
+
+| | fitted | ds |
+|-----|---------------|---------------------|
+| 0 | NaN | 2017-09-13 00:00:00 |
+| 1 | NaN | 2017-09-13 01:00:00 |
+| 2 | NaN | 2017-09-13 02:00:00 |
+| ... | ... | ... |
+| 183 | 145443.734375 | 2017-09-20 15:00:00 |
+| 184 | 150638.000000 | 2017-09-20 16:00:00 |
+| 185 | 155233.093750 | 2017-09-20 17:00:00 |
+
+```python
+sns.lineplot(df, x="ds", y="y", label="Actual", linewidth=2)
+sns.lineplot(fitted,x="ds", y="fitted", label="Fitted", linestyle="--" )
+
+plt.title("Ads watched (hourly data)");
+plt.show()
+```
+
+
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 30 hours ahead.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min.
+
+```python
+Y_hat = sf.forecast(df=train, h=horizon, fitted=True)
+Y_hat
+```
+
+| | unique_id | ds | SeasonalES |
+|-----|-----------|---------------------|---------------|
+| 0 | 1 | 2017-09-20 18:00:00 | 161567.593750 |
+| 1 | 1 | 2017-09-20 19:00:00 | 163186.562500 |
+| 2 | 1 | 2017-09-20 20:00:00 | 134410.937500 |
+| ... | ... | ... | ... |
+| 27 | 1 | 2017-09-21 21:00:00 | 106145.601562 |
+| 28 | 1 | 2017-09-21 22:00:00 | 93383.164062 |
+| 29 | 1 | 2017-09-21 23:00:00 | 79489.718750 |
+
+```python
+values=sf.forecast_fitted_values()
+values.head()
+```
+
+| | unique_id | ds | y | SeasonalES |
+|-----|-----------|---------------------|----------|------------|
+| 0 | 1 | 2017-09-13 00:00:00 | 80115.0 | NaN |
+| 1 | 1 | 2017-09-13 01:00:00 | 79885.0 | NaN |
+| 2 | 1 | 2017-09-13 02:00:00 | 89325.0 | NaN |
+| 3 | 1 | 2017-09-13 03:00:00 | 101930.0 | NaN |
+| 4 | 1 | 2017-09-13 04:00:00 | 121630.0 | NaN |
+
+```python
+sf.plot(train, Y_hat)
+```
+
+
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 30 hourly ahead.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+forecast_df = sf.predict(h=horizon)
+forecast_df
+```
+
+| | unique_id | ds | SeasonalES |
+|-----|-----------|---------------------|---------------|
+| 0 | 1 | 2017-09-20 18:00:00 | 161567.593750 |
+| 1 | 1 | 2017-09-20 19:00:00 | 163186.562500 |
+| 2 | 1 | 2017-09-20 20:00:00 | 134410.937500 |
+| ... | ... | ... | ... |
+| 27 | 1 | 2017-09-21 21:00:00 | 106145.601562 |
+| 28 | 1 | 2017-09-21 22:00:00 | 93383.164062 |
+| 29 | 1 | 2017-09-21 23:00:00 | 79489.718750 |
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=5)`, forecasting every second months
+`(step_size=12)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 30 hourly ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=df,
+ h=horizon,
+ step_size=12,
+ n_windows=3)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` series identifier.
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the n_windows.
+- `y:` true value
+- `"model":` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df
+```
+
+| | unique_id | ds | cutoff | y | SeasonalES |
+|-----|-----------|---------------------|---------------------|----------|---------------|
+| 0 | 1 | 2017-09-19 18:00:00 | 2017-09-19 17:00:00 | 161385.0 | 162297.953125 |
+| 1 | 1 | 2017-09-19 19:00:00 | 2017-09-19 17:00:00 | 165010.0 | 155892.812500 |
+| 2 | 1 | 2017-09-19 20:00:00 | 2017-09-19 17:00:00 | 134090.0 | 135694.703125 |
+| ... | ... | ... | ... | ... | ... |
+| 87 | 1 | 2017-09-21 21:00:00 | 2017-09-20 17:00:00 | 103080.0 | 106145.601562 |
+| 88 | 1 | 2017-09-21 22:00:00 | 2017-09-20 17:00:00 | 95155.0 | 93383.164062 |
+| 89 | 1 | 2017-09-21 23:00:00 | 2017-09-20 17:00:00 | 80285.0 | 79489.718750 |
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=season_length), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | SeasonalES |
+|-----|-----------|--------|-------------|
+| 0 | 1 | mae | 5728.207812 |
+| 1 | 1 | mape | 0.049386 |
+| 2 | 1 | mase | 0.707731 |
+| 3 | 1 | rmse | 7290.840738 |
+| 4 | 1 | smape | 0.024009 |
+
+## Acknowledgements
+
+We would like to thank [Naren
+Castellon](https://www.linkedin.com/in/naren-castellon-1541b8101/?originalSubdomain=pa)
+for writing this tutorial.
+
+## References
+
+1. [Changquan Huang • Alla Petukhina. Springer series (2022). Applied
+ Time Series Analysis and Forecasting with
+ Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2)
+2. Ivan Svetunkov. [Forecasting and Analytics with the Augmented
+ Dynamic Adaptive Model (ADAM)](https://openforecast.org/adam/)
+3. [James D. Hamilton. Time Series Analysis Princeton University Press,
+ Princeton, New Jersey, 1st Edition,
+ 1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Nixtla SeasonalExponentialSmoothing
+ API](../../src/core/models.html#seasonalexponentialsmoothing)
+5. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+6. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+7. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/models/seasonalexponentialsmoothingoptimized.html.mdx b/statsforecast/docs/models/seasonalexponentialsmoothingoptimized.html.mdx
new file mode 100644
index 00000000..c9b8988b
--- /dev/null
+++ b/statsforecast/docs/models/seasonalexponentialsmoothingoptimized.html.mdx
@@ -0,0 +1,824 @@
+---
+title: Seasonal Exponential Smoothing Optimized Model
+---
+
+
+
+
+
+> Step-by-step guide on using the
+> `SeasonalExponentialSmoothingOptimized Model` with `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation` in other.
+
+The text in this article is largely taken from:
+
+1. [Changquan Huang • Alla Petukhina. Springer series (2022). Applied
+ Time Series Analysis and Forecasting with
+ Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2)
+2. Ivan Svetunkov. [Forecasting and Analytics with the Augmented
+ Dynamic Adaptive Model (ADAM)](https://openforecast.org/adam/)
+3. [James D. Hamilton. Time Series Analysis Princeton University Press,
+ Princeton, New Jersey, 1st Edition,
+ 1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [Seasonal Exponential Smoothing Optimized Model](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of SeasonalExponentialSmoothingOptimized with
+ StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+The Seasonal Exponential Smoothing Optimized (SESO) model is a
+forecasting technique used to predict future values of a time series
+that exhibits seasonal patterns. It is a variant of the exponential
+smoothing method, which uses a combination of past and predicted values
+to generate a prediction.
+
+The SESO algorithm uses an optimization approach to find the optimal
+values of the seasonal exponential smoothing parameters. These
+parameters include the smoothing coefficients for the levels, trends,
+and seasonal components of the time series.
+
+The SESO model is particularly useful for forecasting time series with
+pronounced seasonal patterns, such as seasonal product sales or seasonal
+temperatures, and many other areas. By using SESO, accurate and useful
+forecasts can be generated for business planning and decision making.
+
+## Seasonal Exponential Smoothing Model
+
+The SESO model is based on the exponential smoothing method, which uses
+a combination of past and predicted values to generate a prediction. The
+mathematical formula for the SESO model is as follows:
+
+$$\hat{y}{t+1,s} = \alpha y_t + (1-\alpha) \hat{y}{t-1,s}$$
+
+Where: - $\hat{y}{t+1,s}$ is the forecast for the next period of the
+season $s$. - $\alpha$ is the smoothing parameter that is optimized by
+minimizing the squared error. - $y_t$ is the current observation of
+station $s$ in period $t$. - $\hat{y}{t-1,s}$ is the forecast for the
+previous period of the season $s$.
+
+The equation indicates that the forecast value for the next season
+period $s$ is calculated as a weighted combination of the current
+observation and the previous forecast for the same station. The
+smoothing parameter $\alpha$ controls the relative influence of these
+two terms on the final prediction. A high value of α gives more weight
+to the current observation and less weight to the previous forecast,
+making the model more sensitive to recent changes in the time series. A
+low value of $\alpha$, on the other hand, gives more weight to the
+previous forecast and less weight to the current observation, making the
+model more stable and smooth.
+
+The optimal value of the smoothing parameter $\alpha$ is determined by
+minimizing the squared error between the forecasts generated by the
+model and the actual values of the time series.
+
+### Model selection
+
+Model selection in the context of the SESO model refers to the process
+of choosing the optimal values of the smoothing parameters and the
+seasonal component for the model. The optimal values of these parameters
+are the ones that result in the best forecast performance for the given
+data set.
+
+A great advantage of the ETS statistical framework is that information
+criteria can be used for model selection. The $AIC, AIC_c$ and $BIC$,
+that also can be used here to determine which of the ETS models is most
+appropriate for a given time series.
+
+For ETS models, Akaike’s Information Criterion (AIC) is defined as
+$$\text{AIC} = -2\log(L) + 2k,$$
+
+where $L$ is the likelihood of the model and $k$ is the total number of
+parameters and initial states that have been estimated (including the
+residual variance).
+
+The AIC corrected for small sample bias ($AIC_c$) is defined as
+$$\text{AIC}_{\text{c}} = \text{AIC} + \frac{2k(k+1)}{T-k-1},$$
+
+and the Bayesian Information Criterion (BIC) is
+$$\text{BIC} = \text{AIC} + k[\log(T)-2].$$
+
+These criteria balance the goodness of fit with the complexity of the
+model and provide a way to choose the model that maximizes the
+likelihood of the data while minimizing the number of parameters.
+
+In addition to these techniques, expert judgment and domain knowledge
+can also be used to select the optimal SESO model. This involves
+considering the underlying dynamics of the time series, the patterns of
+seasonality, and any other relevant factors that may influence the
+choice of the model.
+
+Overall, the process of model selection for the SESO model involves a
+combination of statistical techniques, information criteria, and expert
+judgment to identify the optimal values of the smoothing parameters and
+the seasonal component that result in the best forecast performance for
+the given data set.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
+plt.style.use('grayscale') # fivethirtyeight grayscale classic
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#008080', # #212946
+ 'axes.facecolor': '#008080',
+ 'savefig.facecolor': '#008080',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#000000', #2A3459
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+### Read Data
+
+```python
+import pandas as pd
+
+df=pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/ads.csv")
+df.head()
+```
+
+| | Time | Ads |
+|-----|---------------------|--------|
+| 0 | 2017-09-13T00:00:00 | 80115 |
+| 1 | 2017-09-13T01:00:00 | 79885 |
+| 2 | 2017-09-13T02:00:00 | 89325 |
+| 3 | 2017-09-13T03:00:00 | 101930 |
+| 4 | 2017-09-13T04:00:00 | 121630 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|---------------------|--------|-----------|
+| 0 | 2017-09-13T00:00:00 | 80115 | 1 |
+| 1 | 2017-09-13T01:00:00 | 79885 | 1 |
+| 2 | 2017-09-13T02:00:00 | 89325 | 1 |
+| 3 | 2017-09-13T03:00:00 | 101930 | 1 |
+| 4 | 2017-09-13T04:00:00 | 121630 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y int64
+unique_id object
+dtype: object
+```
+
+We can see that our time variable `(ds)` is in an object format, we need
+to convert to a date format
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore Data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints a random series from the dataset and is useful for
+basic EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### The Augmented Dickey-Fuller Test
+
+An Augmented Dickey-Fuller (ADF) test is a type of statistical test that
+determines whether a unit root is present in time series data. Unit
+roots can cause unpredictable results in time series analysis. A null
+hypothesis is formed in the unit root test to determine how strongly
+time series data is affected by a trend. By accepting the null
+hypothesis, we accept the evidence that the time series data is not
+stationary. By rejecting the null hypothesis or accepting the
+alternative hypothesis, we accept the evidence that the time series data
+is generated by a stationary process. This process is also known as
+stationary trend. The values of the ADF test statistic are negative.
+Lower ADF values indicate a stronger rejection of the null hypothesis.
+
+Augmented Dickey-Fuller Test is a common statistical test used to test
+whether a given time series is stationary or not. We can achieve this by
+defining the null and alternate hypothesis.
+
+Null Hypothesis: Time Series is non-stationary. It gives a
+time-dependent trend. Alternate Hypothesis: Time Series is stationary.
+In another term, the series doesn’t depend on time.
+
+ADF or t Statistic \< critical values: Reject the null hypothesis, time
+series is stationary. ADF or t Statistic \> critical values: Failed to
+reject the null hypothesis, time series is non-stationary.
+
+```python
+from statsmodels.tsa.stattools import adfuller
+
+def Augmented_Dickey_Fuller_Test_func(series , column_name):
+ print (f'Dickey-Fuller test results for columns: {column_name}')
+ dftest = adfuller(series, autolag='AIC')
+ dfoutput = pd.Series(dftest[0:4], index=['Test Statistic','p-value','No Lags Used','Number of observations used'])
+ for key,value in dftest[4].items():
+ dfoutput['Critical Value (%s)'%key] = value
+ print (dfoutput)
+ if dftest[1] \<= 0.05:
+ print("Conclusion:====>")
+ print("Reject the null hypothesis")
+ print("The data is stationary")
+ else:
+ print("Conclusion:====>")
+ print("The null hypothesis cannot be rejected")
+ print("The data is not stationary")
+```
+
+
+```python
+Augmented_Dickey_Fuller_Test_func(df["y"],'Ads')
+```
+
+``` text
+Dickey-Fuller test results for columns: Ads
+Test Statistic -7.089634e+00
+p-value 4.444804e-10
+No Lags Used 9.000000e+00
+ ...
+Critical Value (1%) -3.462499e+00
+Critical Value (5%) -2.875675e+00
+Critical Value (10%) -2.574304e+00
+Length: 7, dtype: float64
+Conclusion:====>
+Reject the null hypothesis
+The data is stationary
+```
+
+### Autocorrelation plots
+
+The important characteristics of Autocorrelation (ACF) and Partial
+Autocorrelation (PACF) are as follows:
+
+Autocorrelation (ACF): 1. Identify patterns of temporal dependence: The
+ACF shows the correlation between an observation and its lagged values
+at different time intervals. Helps identify patterns of temporal
+dependency in a time series, such as the presence of trends or
+seasonality.
+
+1. Indicates the “memory” of the series: The ACF allows us to determine
+ how much past observations influence future ones. If the ACF shows
+ significant autocorrelations in several lags, it indicates that the
+ series has a long-term memory and that past observations are
+ relevant to predict future ones.
+
+2. Helps identify MA (moving average) models: The shape of the ACF can
+ reveal the presence of moving average components in the time series.
+ Lags where the ACF shows a significant correlation may indicate the
+ order of an MA model.
+
+Partial Autocorrelation (PACF): 1. Identify direct dependence: Unlike
+the ACF, the PACF eliminates the indirect effects of intermediate lags
+and measures the direct correlation between an observation and its
+lagged values. It helps to identify the direct dependence between an
+observation and its lag values, without the influence of intermediate
+lags.
+
+1. Helps to identify AR (autoregressive) models: The shape of the PACF
+ can reveal the presence of autoregressive components in the time
+ series. Lags in which the PACF shows a significant correlation may
+ indicate the order of an AR model.
+
+2. Used in conjunction with the ACF: The PACF is used in conjunction
+ with the ACF to determine the order of an AR or MA model. By
+ analyzing both the ACF and the PACF, significant lags can be
+ identified and a model suitable for time series analysis and
+ forecasting can be built.
+
+In summary, the ACF and the PACF are complementary tools in time series
+analysis that provide information on time dependence and help identify
+the appropriate components to build forecast models.
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=30, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+# Grafico
+plot_pacf(df["y"], lags=30, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+#plt.savefig("Gráfico de Densidad y qq")
+plt.show();
+```
+
+
+
+### Decomposition of the time series
+
+How to decompose a time series and why?
+
+In time series analysis to forecast new values, it is very important to
+know past data. More formally, we can say that it is very important to
+know the patterns that values follow over time. There can be many
+reasons that cause our forecast values to fall in the wrong direction.
+Basically, a time series consists of four components. The variation of
+those components causes the change in the pattern of the time series.
+These components are:
+
+- **Level:** This is the primary value that averages over time.
+- **Trend:** The trend is the value that causes increasing or
+ decreasing patterns in a time series.
+- **Seasonality:** This is a cyclical event that occurs in a time
+ series for a short time and causes short-term increasing or
+ decreasing patterns in a time series.
+- **Residual/Noise:** These are the random variations in the time
+ series.
+
+Combining these components over time leads to the formation of a time
+series. Most time series consist of level and noise/residual and trend
+or seasonality are optional values.
+
+If seasonality and trend are part of the time series, then there will be
+effects on the forecast value. As the pattern of the forecasted time
+series may be different from the previous time series.
+
+The combination of the components in time series can be of two types: \*
+Additive \* Multiplicative
+
+### Additive time series
+
+If the components of the time series are added to make the time series.
+Then the time series is called the additive time series. By
+visualization, we can say that the time series is additive if the
+increasing or decreasing pattern of the time series is similar
+throughout the series. The mathematical function of any additive time
+series can be represented by:
+$$y(t) = level + Trend + seasonality + noise$$
+
+### Multiplicative time series
+
+If the components of the time series are multiplicative together, then
+the time series is called a multiplicative time series. For
+visualization, if the time series is having exponential growth or
+decline with time, then the time series can be considered as the
+multiplicative time series. The mathematical function of the
+multiplicative time series can be represented as.
+
+$$y(t) = Level * Trend * seasonality * Noise$$
+
+### Additive
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+a = seasonal_decompose(df["y"], model = "additive", period=12)
+a.plot();
+```
+
+
+
+### Multiplicative
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+a = seasonal_decompose(df["y"], model = "Multiplicative", period=12)
+a.plot();
+```
+
+
+
+## Split the data into training and testing
+
+Let’s divide our data into sets
+
+1. Data to train our `Seasonal Exponential Smoothing Optimized Model`.
+2. Data to test our model
+
+For the test data we will use the last 30 hours to test and evaluate the
+performance of our model.
+
+```python
+train = df[df.ds\<='2017-09-20 17:00:00']
+test = df[df.ds>'2017-09-20 17:00:00']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((186, 3), (30, 3))
+```
+
+## Implementation of SeasonalExponentialSmoothingOptimized with StatsForecast
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import SeasonalExponentialSmoothingOptimized
+```
+
+### Building Model
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/) by the
+master, Rob Hyndmann, can be useful for `season_length`.
+
+```python
+season_length = 24 # Hourly data
+horizon = len(test) # number of predictions
+
+models = [SeasonalExponentialSmoothingOptimized(season_length=season_length)]
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [panda’s
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models, freq='h')
+```
+
+### Fit the Model
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[SeasESOpt])
+```
+
+Let’s see the results of our
+`Seasonal Exponential Smoothing Optimized Model`. We can observe it with
+the following instruction:
+
+```python
+result=sf.fitted_[0,0].model_
+result
+```
+
+``` text
+{'mean': array([161532.05 , 161051.69 , 135531.64 , 105600.39 , 96717.39 ,
+ 82608.34 , 80224.33 , 78075.98 , 85233.23 , 100179.336,
+ 122245.62 , 118087.57 , 109614.81 , 104729.91 , 104895.02 ,
+ 115862.96 , 130370.98 , 144231.89 , 149036.73 , 149072.73 ,
+ 148110.77 , 148760.73 , 149767.53 , 150561.8 ], dtype=float32),
+ 'fitted': array([ nan, nan, nan, nan, nan,
+ nan, nan, nan, nan, nan,
+ nan, nan, nan, nan, nan,
+ nan, nan, nan, nan, nan,
+ nan, nan, nan, nan, nan,
+ nan, nan, nan, nan, nan,
+ nan, nan, nan, nan, nan,
+ nan, nan, nan, nan, nan,
+ nan, nan, 163840. , 166235. , 139520. ,
+ 105895. , 96780. , 82520. , 80125. , 75335. ,
+ 85105. , 102080. , 125135. , 118030. , 109225. ,
+ 102475. , 102240. , 115840. , 130540. , 144325. ,
+ 148970. , 149150. , 148040. , 148810. , 149830. ,
+ 150570. , 162030.27 , 163222.1 , 137347.33 , 103835.8 ,
+ 96733.95 , 82522.45 , 80086.9 , 75132.05 , 85074.36 ,
+ 100452.66 , 121044.03 , 118001.6 , 109242.15 , 102349.03 ,
+ 102321.49 , 115768.25 , 130501. , 144286.1 , 149005. ,
+ 149121.25 , 148039.8 , 148799.25 , 149789.2 , 150557.16 ,
+ 161740.55 , 162812.36 , 136965.22 , 112853.91 , 96768.61 ,
+ 82573.375, 80164.38 , 88707.87 , 85164.8 , 100944.15 ,
+ 117929.875, 118111.086, 109563.58 , 103815.7 , 104036.375,
+ 115942.47 , 130508.39 , 144268.03 , 149088.7 , 149155.03 ,
+ 148096.75 , 148823.2 , 149797.77 , 150525.92 , 160582.38 ,
+ 159756.83 , 134514.39 , 117874.29 , 96767.92 , 82683.74 ,
+ 80253.336, 89338.625, 85232.055, 100619.03 , 114659.62 ,
+ 118224.67 , 109881.99 , 105514.21 , 106070.33 , 116194.74 ,
+ 130678.805, 144436.45 , 149261.16 , 149331.28 , 148247.19 ,
+ 148908.03 , 149890.33 , 150620.88 , 161557.95 , 161701.48 ,
+ 136228.19 , 113004.195, 96773.695, 82673.66 , 80245.91 ,
+ 78459.88 , 85267.016, 100517.48 , 120224.3 , 118155.68 ,
+ 109777.57 , 105240.35 , 105717.734, 116058.445, 130571.32 ,
+ 144349.64 , 149169.75 , 149255.28 , 148186.9 , 148872.55 ,
+ 149844.78 , 150618.77 , 161553.17 , 160112.8 , 135912.81 ,
+ 107124.39 , 96756.41 , 82642.07 , 80226.8 , 74707.9 ,
+ 85226.28 , 100202.98 , 119687.805, 118105.27 , 109668.59 ,
+ 104848.68 , 105212.516, 115939.71 , 130463.1 , 144266.05 ,
+ 149142.61 , 149154.38 , 148177.3 , 148833.17 , 149860.03 ,
+ 150673.38 ], dtype=float32)}
+```
+
+Let us now visualize the fitted values of our models.
+
+As we can see, the result obtained above has an output in a dictionary,
+to extract each element from the dictionary we are going to use the
+`.get()` function to extract the element and then we are going to save
+it in a `pd.DataFrame()`.
+
+```python
+fitted=pd.DataFrame(result.get("fitted"), columns=["fitted"])
+fitted["ds"]=df["ds"]
+fitted
+```
+
+| | fitted | ds |
+|-----|---------------|---------------------|
+| 0 | NaN | 2017-09-13 00:00:00 |
+| 1 | NaN | 2017-09-13 01:00:00 |
+| 2 | NaN | 2017-09-13 02:00:00 |
+| ... | ... | ... |
+| 183 | 148833.171875 | 2017-09-20 15:00:00 |
+| 184 | 149860.031250 | 2017-09-20 16:00:00 |
+| 185 | 150673.375000 | 2017-09-20 17:00:00 |
+
+```python
+sns.lineplot(df, x="ds", y="y", label="Actual", linewidth=2)
+sns.lineplot(fitted,x="ds", y="fitted", label="Fitted", linestyle="--" )
+
+plt.title("Ads watched (hourly data)");
+plt.show()
+```
+
+
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min.
+
+```python
+# Prediction
+Y_hat = sf.forecast(df=train, h=horizon, fitted=True)
+Y_hat
+```
+
+| | unique_id | ds | SeasESOpt |
+|-----|-----------|---------------------|---------------|
+| 0 | 1 | 2017-09-20 18:00:00 | 161532.046875 |
+| 1 | 1 | 2017-09-20 19:00:00 | 161051.687500 |
+| 2 | 1 | 2017-09-20 20:00:00 | 135531.640625 |
+| ... | ... | ... | ... |
+| 27 | 1 | 2017-09-21 21:00:00 | 105600.390625 |
+| 28 | 1 | 2017-09-21 22:00:00 | 96717.390625 |
+| 29 | 1 | 2017-09-21 23:00:00 | 82608.343750 |
+
+```python
+values=sf.forecast_fitted_values()
+values.head()
+```
+
+| | unique_id | ds | y | SeasESOpt |
+|-----|-----------|---------------------|----------|-----------|
+| 0 | 1 | 2017-09-13 00:00:00 | 80115.0 | NaN |
+| 1 | 1 | 2017-09-13 01:00:00 | 79885.0 | NaN |
+| 2 | 1 | 2017-09-13 02:00:00 | 89325.0 | NaN |
+| 3 | 1 | 2017-09-13 03:00:00 | 101930.0 | NaN |
+| 4 | 1 | 2017-09-13 04:00:00 | 121630.0 | NaN |
+
+```python
+sf.plot(train, Y_hat)
+```
+
+
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 30 hours ahead.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+forecast_df = sf.predict(h=horizon)
+forecast_df
+```
+
+| | unique_id | ds | SeasESOpt |
+|-----|-----------|---------------------|---------------|
+| 0 | 1 | 2017-09-20 18:00:00 | 161532.046875 |
+| 1 | 1 | 2017-09-20 19:00:00 | 161051.687500 |
+| 2 | 1 | 2017-09-20 20:00:00 | 135531.640625 |
+| ... | ... | ... | ... |
+| 27 | 1 | 2017-09-21 21:00:00 | 105600.390625 |
+| 28 | 1 | 2017-09-21 22:00:00 | 96717.390625 |
+| 29 | 1 | 2017-09-21 23:00:00 | 82608.343750 |
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=)`, forecasting every second months
+`(step_size=12)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 12 months ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=df,
+ h=horizon,
+ step_size=30,
+ n_windows=3)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` series identifier.
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the `n_windows`.
+- `y:` true value
+- `model:` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df
+```
+
+| | unique_id | ds | cutoff | y | SeasESOpt |
+|-----|-----------|---------------------|---------------------|----------|---------------|
+| 0 | 1 | 2017-09-18 06:00:00 | 2017-09-18 05:00:00 | 99440.0 | 141401.750000 |
+| 1 | 1 | 2017-09-18 07:00:00 | 2017-09-18 05:00:00 | 97655.0 | 152474.250000 |
+| 2 | 1 | 2017-09-18 08:00:00 | 2017-09-18 05:00:00 | 97655.0 | 152482.796875 |
+| ... | ... | ... | ... | ... | ... |
+| 87 | 1 | 2017-09-21 21:00:00 | 2017-09-20 17:00:00 | 103080.0 | 105600.390625 |
+| 88 | 1 | 2017-09-21 22:00:00 | 2017-09-20 17:00:00 | 95155.0 | 96717.390625 |
+| 89 | 1 | 2017-09-21 23:00:00 | 2017-09-20 17:00:00 | 80285.0 | 82608.343750 |
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=season_length), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | SeasESOpt |
+|-----|-----------|--------|-------------|
+| 0 | 1 | mae | 6694.042188 |
+| 1 | 1 | mape | 0.060392 |
+| 2 | 1 | mase | 0.827062 |
+| 3 | 1 | rmse | 8118.297509 |
+| 4 | 1 | smape | 0.028961 |
+
+## References
+
+1. [Changquan Huang • Alla Petukhina. Springer series (2022). Applied
+ Time Series Analysis and Forecasting with
+ Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2)
+2. Ivan Svetunkov. [Forecasting and Analytics with the Augmented
+ Dynamic Adaptive Model (ADAM)](https://openforecast.org/adam/)
+3. [James D. Hamilton. Time Series Analysis Princeton University Press,
+ Princeton, New Jersey, 1st Edition,
+ 1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Nixtla SeasonalExponentialSmoothingOptimized
+ API](../../src/core/models.html#seasonalexponentialsmoothingoptimized)
+5. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+6. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+7. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/models/simpleexponentialoptimized.html.mdx b/statsforecast/docs/models/simpleexponentialoptimized.html.mdx
new file mode 100644
index 00000000..f1d98c57
--- /dev/null
+++ b/statsforecast/docs/models/simpleexponentialoptimized.html.mdx
@@ -0,0 +1,636 @@
+---
+title: Simple Exponential Smoothing Optimized Model
+---
+
+
+
+
+
+> Step-by-step guide on using the
+> `SimpleExponentialSmoothingOptimized Model` with `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation` in other.
+
+The text in this article is largely taken from: 1. [Changquan Huang •
+Alla Petukhina. Springer series (2022). Applied Time Series Analysis and
+Forecasting with
+Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2) 2.
+Ivan Svetunkov. [Forecasting and Analytics with the Augmented Dynamic
+Adaptive Model (ADAM)](https://openforecast.org/adam/) 3. [James D.
+Hamilton. Time Series Analysis Princeton University Press, Princeton,
+New Jersey, 1st Edition,
+1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+Principles and Practice (3rd ed)”](https://otexts.com/fpp3/tscv.html).
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [Simple Exponential Smoothing Optimized Model](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of SimpleExponentialSmoothingOptimized with
+ StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+Simple Exponential Smoothing Optimized (SES Optimized) is a forecasting
+model used to predict future values in univariate time series. It is a
+variant of the simple exponential smoothing (SES) method that uses an
+optimization approach to estimate the model parameters more accurately.
+
+The SES Optimized method uses a single smoothing parameter to estimate
+the trend and seasonality in the time series data. The model attempts to
+minimize the mean squared error (MSE) between the predictions and the
+actual values in the training sample using an optimization algorithm.
+
+The SES Optimized approach is especially useful for time series with
+strong trend and seasonality patterns, or for time series with noisy
+data. However, it is important to note that this model assumes that the
+time series is stationary and that the variation in the data is random
+and there are no non-random patterns in the data. If these assumptions
+are not met, the SES Optimized model may not perform well and another
+forecasting method may be required.
+
+## Simple Exponential Smoothing Model
+
+The simplest of the exponentially smoothing methods is naturally called
+simple exponential smoothing (SES). This method is suitable for
+forecasting data with no clear trend or seasonal pattern.
+
+Using the naïve method, all forecasts for the future are equal to the
+last observed value of the series, $$\hat{y}_{T+h|T} = y_{T},$$
+
+for $h=1,2,\dots$. Hence, the naïve method assumes that the most recent
+observation is the only important one, and all previous observations
+provide no information for the future. This can be thought of as a
+weighted average where all of the weight is given to the last
+observation.
+
+Using the average method, all future forecasts are equal to a simple
+average of the observed data,
+$$\hat{y}_{T+h|T} = \frac1T \sum_{t=1}^T y_t,$$
+
+for $h=1,2,\dots$ Hence, the average method assumes that all
+observations are of equal importance, and gives them equal weights when
+generating forecasts.
+
+We often want something between these two extremes. For example, it may
+be sensible to attach larger weights to more recent observations than to
+observations from the distant past. This is exactly the concept behind
+simple exponential smoothing. Forecasts are calculated using weighted
+averages, where the weights decrease exponentially as observations come
+from further in the past — the smallest weights are associated with the
+oldest observations:
+
+where $0 \le \alpha \le 1$ is the smoothing parameter. The
+one-step-ahead forecast for time $T+1$ is a weighted average of all of
+the observations in the series $y_1,\dots,y_T$. The rate at which the
+weights decrease is controlled by the parameter $\alpha$.
+
+For any $\alpha$ between 0 and 1, the weights attached to the
+observations decrease exponentially as we go back in time, hence the
+name “exponential smoothing”. If $\alpha$ is small (i.e., close to 0),
+more weight is given to observations from the more distant past. If
+$\alpha$ is large (i.e., close to 1), more weight is given to the more
+recent observations. For the extreme case where $\alpha=1$,
+$\hat{y}_{T+1|T}=y_T$ and the forecasts are equal to the naïve
+forecasts.
+
+## Optimisation
+
+The application of every exponential smoothing method requires the
+smoothing parameters and the initial values to be chosen. In particular,
+for simple exponential smoothing, we need to select the values of
+$\alpha$ and $\ell_0$ . All forecasts can be computed from the data once
+we know those values. For the methods that follow there is usually more
+than one smoothing parameter and more than one initial component to be
+chosen.
+
+In some cases, the smoothing parameters may be chosen in a subjective
+manner — the forecaster specifies the value of the smoothing parameters
+based on previous experience. However, a more reliable and objective way
+to obtain values for the unknown parameters is to estimate them from the
+observed data.
+
+From regression models we estimated the coefficients of a regression
+model by minimising the sum of the squared residuals (usually known as
+SSE or “sum of squared errors”). Similarly, the unknown parameters and
+the initial values for any exponential smoothing method can be estimated
+by minimising the SSE. The residuals are specified as
+$e_t=y_t - \hat{y}_{t|t-1}$ for $t=1,\dots,T$. Hence, we find the values
+of the unknown parameters and the initial values that minimise
+
+Unlike the regression case (where we have formulas which return the
+values of the regression coefficients that minimise the SSE), this
+involves a non-linear minimisation problem, and we need to use an
+optimisation tool to solve it.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
+plt.style.use('grayscale') # fivethirtyeight grayscale classic
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#008080', # #212946
+ 'axes.facecolor': '#008080',
+ 'savefig.facecolor': '#008080',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#000000', #2A3459
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+### Read Data
+
+```python
+import pandas as pd
+df=pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/ads.csv")
+df.head()
+```
+
+| | Time | Ads |
+|-----|---------------------|--------|
+| 0 | 2017-09-13T00:00:00 | 80115 |
+| 1 | 2017-09-13T01:00:00 | 79885 |
+| 2 | 2017-09-13T02:00:00 | 89325 |
+| 3 | 2017-09-13T03:00:00 | 101930 |
+| 4 | 2017-09-13T04:00:00 | 121630 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|---------------------|--------|-----------|
+| 0 | 2017-09-13T00:00:00 | 80115 | 1 |
+| 1 | 2017-09-13T01:00:00 | 79885 | 1 |
+| 2 | 2017-09-13T02:00:00 | 89325 | 1 |
+| 3 | 2017-09-13T03:00:00 | 101930 | 1 |
+| 4 | 2017-09-13T04:00:00 | 121630 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y int64
+unique_id object
+dtype: object
+```
+
+We can see that our time variable `(ds)` is in an object format, we need
+to convert to a date format
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore Data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints a random series from the dataset and is useful for
+basic EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### Autocorrelation plots
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=30, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+plot_pacf(df["y"], lags=30, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+plt.show();
+```
+
+
+
+## Split the data into training and testing
+
+Let’s divide our data into sets
+
+1. Data to train our `Simple Exponential Smoothing Optimized Model`
+2. Data to test our model
+
+For the test data we will use the last 30 Hours to test and evaluate the
+performance of our model.
+
+```python
+train = df[df.ds\<='2017-09-20 17:00:00']
+test = df[df.ds>'2017-09-20 17:00:00']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((186, 3), (30, 3))
+```
+
+## Implementation of SimpleExponentialSmoothingOptimized with StatsForecast
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import SimpleExponentialSmoothingOptimized
+```
+
+### Instantiating Model
+
+```python
+horizon = len(test) # number of predictions
+
+models = [SimpleExponentialSmoothingOptimized()] # multiplicative additive
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [panda’s
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models, freq='h')
+```
+
+### Fit the Model
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[SESOpt])
+```
+
+Let’s see the results of our
+`Simple Exponential Smoothing Optimized model`. We can observe it with
+the following instruction:
+
+```python
+result=sf.fitted_[0,0].model_
+result
+```
+
+``` text
+{'mean': array([139526.04792941]),
+ 'fitted': array([ nan, 80115. , 79887.3 , 89230.625, 101803.01 ,
+ 121431.73 , 116524.57 , 106595.3 , 102833. , 108002.78 ,
+ 116043.78 , 130880.14 , 148838.6 , 157502.48 , 150782.88 ,
+ 149309.88 , 150092.1 , 144833.12 , 150631.44 , 163707.92 ,
+ 166209.73 , 139786.89 , 106233.92 , 96874.54 , 82663.55 ,
+ 80150.38 , 75383.16 , 85007.78 , 101909.28 , 124902.74 ,
+ 118098.73 , 109313.734, 102543.39 , 102243.03 , 115704.03 ,
+ 130391.64 , 144185.67 , 148922.16 , 149147.72 , 148051.08 ,
+ 148802.4 , 149819.72 , 150562.5 , 149451.22 , 150509.31 ,
+ 129343.8 , 104070.29 , 92293.95 , 82860.29 , 76380.45 ,
+ 75142.51 , 82565.02 , 88732.7 , 118133.02 , 115219.43 ,
+ 110982.8 , 98981.23 , 104132.96 , 108619.68 , 126459.8 ,
+ 140295.25 , 152348.25 , 146335.73 , 148003.16 , 147737.69 ,
+ 145769.88 , 149249.84 , 159620.25 , 161070.36 , 135775.5 ,
+ 113173.305, 100329.734, 87742.15 , 87834.07 , 88834.89 ,
+ 92314.85 , 104343.5 , 115824.03 , 128818.74 , 141259.34 ,
+ 144408.19 , 143261.58 , 133290.72 , 131260.5 , 142367.81 ,
+ 157224.92 , 152547.25 , 153723.12 , 151220.28 , 150650.75 ,
+ 147467.16 , 152474.42 , 146931. , 125461.86 , 118000.37 ,
+ 96913. , 93643.03 , 89105.83 , 89342.61 , 90562.68 ,
+ 98212.73 , 112426.43 , 129299.56 , 141283.95 , 152447.23 ,
+ 152578.67 , 141284.1 , 147487.34 , 160973.77 , 166281.39 ,
+ 166775.02 , 163176.34 , 157363.72 , 159038.1 , 160010.19 ,
+ 168261.66 , 169883.61 , 142981.73 , 113255.266, 97504.1 ,
+ 81833.29 , 79533.234, 78361.836, 87948.17 , 99671.58 ,
+ 123538.914, 111447.14 , 99560.07 , 97674.05 , 97655.19 ,
+ 102515.9 , 119755.86 , 135595.02 , 140074.75 , 141713.45 ,
+ 142214.94 , 145328.55 , 145334.94 , 150359.25 , 161408.39 ,
+ 153494.94 , 134907.75 , 107343.43 , 95167.984, 79671.53 ,
+ 78348.37 , 74706.78 , 81917.164, 97789.67 , 119129.445,
+ 113175.14 , 99022.95 , 94050.23 , 93663.9 , 104079.79 ,
+ 119593.3 , 135826.03 , 146348.7 , 139236.84 , 147145.12 ,
+ 144957.1 , 151305.88 , 156032.27 , 161331.47 , 164973.22 ,
+ 134398.83 , 105873.14 , 92985.18 , 79407.15 , 79974.27 ,
+ 78128.64 , 85708.44 , 99866.984, 123639.87 , 116408.05 ,
+ 104411.18 , 101469.71 , 97673.34 , 108159.086, 121119.09 ,
+ 140652.69 , 138575.98 , 140965.86 , 141519.4 , 141589.3 ,
+ 140619.8 ], dtype=float32)}
+```
+
+Let us now visualize the residuals of our models.
+
+As we can see, the result obtained above has an output in a dictionary,
+to extract each element from the dictionary we are going to use the
+`.get()` function to extract the element and then we are going to save
+it in a `pd.DataFrame()`.
+
+```python
+fitted=pd.DataFrame(result.get("fitted"), columns=["fitted"])
+fitted["ds"]=df["ds"]
+fitted
+```
+
+| | fitted | ds |
+|-----|---------------|---------------------|
+| 0 | NaN | 2017-09-13 00:00:00 |
+| 1 | 80115.000000 | 2017-09-13 01:00:00 |
+| 2 | 79887.296875 | 2017-09-13 02:00:00 |
+| ... | ... | ... |
+| 183 | 141519.406250 | 2017-09-20 15:00:00 |
+| 184 | 141589.296875 | 2017-09-20 16:00:00 |
+| 185 | 140619.796875 | 2017-09-20 17:00:00 |
+
+```python
+sns.lineplot(df, x="ds", y="y", label="Actual", linewidth=2)
+sns.lineplot(fitted,x="ds", y="fitted", label="Fitted", linestyle="--" )
+
+plt.title("Ads watched (hourly data)");
+plt.show()
+```
+
+
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 30 hors ahead.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min.
+
+```python
+# Prediction
+Y_hat = sf.forecast(df=train, h=horizon, fitted=True)
+Y_hat
+```
+
+| | unique_id | ds | SESOpt |
+|-----|-----------|---------------------|---------------|
+| 0 | 1 | 2017-09-20 18:00:00 | 139526.046875 |
+| 1 | 1 | 2017-09-20 19:00:00 | 139526.046875 |
+| 2 | 1 | 2017-09-20 20:00:00 | 139526.046875 |
+| ... | ... | ... | ... |
+| 27 | 1 | 2017-09-21 21:00:00 | 139526.046875 |
+| 28 | 1 | 2017-09-21 22:00:00 | 139526.046875 |
+| 29 | 1 | 2017-09-21 23:00:00 | 139526.046875 |
+
+Let’s visualize the fitted values
+
+```python
+values=sf.forecast_fitted_values()
+values.head()
+```
+
+| | unique_id | ds | y | SESOpt |
+|-----|-----------|---------------------|----------|---------------|
+| 0 | 1 | 2017-09-13 00:00:00 | 80115.0 | NaN |
+| 1 | 1 | 2017-09-13 01:00:00 | 79885.0 | 80115.000000 |
+| 2 | 1 | 2017-09-13 02:00:00 | 89325.0 | 79887.296875 |
+| 3 | 1 | 2017-09-13 03:00:00 | 101930.0 | 89230.625000 |
+| 4 | 1 | 2017-09-13 04:00:00 | 121630.0 | 101803.007812 |
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 30 hours ahead.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+forecast_df = sf.predict(h=horizon)
+forecast_df
+```
+
+| | unique_id | ds | SESOpt |
+|-----|-----------|---------------------|---------------|
+| 0 | 1 | 2017-09-20 18:00:00 | 139526.046875 |
+| 1 | 1 | 2017-09-20 19:00:00 | 139526.046875 |
+| 2 | 1 | 2017-09-20 20:00:00 | 139526.046875 |
+| ... | ... | ... | ... |
+| 27 | 1 | 2017-09-21 21:00:00 | 139526.046875 |
+| 28 | 1 | 2017-09-21 22:00:00 | 139526.046875 |
+| 29 | 1 | 2017-09-21 23:00:00 | 139526.046875 |
+
+```python
+sf.plot(train, forecast_df)
+```
+
+
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=)`, forecasting every second months
+`(step_size=12)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 30 hours ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=df,
+ h=horizon,
+ step_size=30,
+ n_windows=3)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` index. If you dont like working with index just run
+ `crossvalidation_df.resetindex()`.
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the `n_windows`.
+- `y:` true value
+- `model:` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df
+```
+
+| | unique_id | ds | cutoff | y | SESOpt |
+|-----|-----------|---------------------|---------------------|----------|---------------|
+| 0 | 1 | 2017-09-18 06:00:00 | 2017-09-18 05:00:00 | 99440.0 | 111447.140625 |
+| 1 | 1 | 2017-09-18 07:00:00 | 2017-09-18 05:00:00 | 97655.0 | 111447.140625 |
+| 2 | 1 | 2017-09-18 08:00:00 | 2017-09-18 05:00:00 | 97655.0 | 111447.140625 |
+| ... | ... | ... | ... | ... | ... |
+| 87 | 1 | 2017-09-21 21:00:00 | 2017-09-20 17:00:00 | 103080.0 | 139526.046875 |
+| 88 | 1 | 2017-09-21 22:00:00 | 2017-09-20 17:00:00 | 95155.0 | 139526.046875 |
+| 89 | 1 | 2017-09-21 23:00:00 | 2017-09-20 17:00:00 | 80285.0 | 139526.046875 |
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=24), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | SESOpt |
+|-----|-----------|--------|--------------|
+| 0 | 1 | mae | 29230.182292 |
+| 1 | 1 | mape | 0.314203 |
+| 2 | 1 | mase | 3.611444 |
+| 3 | 1 | rmse | 35866.963426 |
+| 4 | 1 | smape | 0.124271 |
+
+## References
+
+1. [Changquan Huang • Alla Petukhina. Springer series (2022). Applied
+ Time Series Analysis and Forecasting with
+ Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2)
+2. Ivan Svetunkov. [Forecasting and Analytics with the Augmented
+ Dynamic Adaptive Model (ADAM)](https://openforecast.org/adam/)
+3. [James D. Hamilton. Time Series Analysis Princeton University Press,
+ Princeton, New Jersey, 1st Edition,
+ 1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Nixtla SeasonalExponentialOptimized
+ API](../../src/core/models.html#simpleexponentialsmoothingoptimized)
+5. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+6. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+7. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/models/simpleexponentialsmoothing.html.mdx b/statsforecast/docs/models/simpleexponentialsmoothing.html.mdx
new file mode 100644
index 00000000..5f214a18
--- /dev/null
+++ b/statsforecast/docs/models/simpleexponentialsmoothing.html.mdx
@@ -0,0 +1,688 @@
+---
+title: Simple Exponential Smoothing Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `SimpleExponentialSmoothing Model`
+> with `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation` in other.
+
+The text in this article is largely taken from: 1. [Changquan Huang •
+Alla Petukhina. Springer series (2022). Applied Time Series Analysis and
+Forecasting with
+Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2) 2.
+Ivan Svetunkov. [Forecasting and Analytics with the Augmented Dynamic
+Adaptive Model (ADAM)](https://openforecast.org/adam/) 3. [James D.
+Hamilton. Time Series Analysis Princeton University Press, Princeton,
+New Jersey, 1st Edition,
+1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+Principles and Practice (3rd ed)”](https://otexts.com/fpp3/tscv.html).
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [Simple Exponential Smoothing](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of SimpleExponentialSmoothing with
+ StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+Exponential smoothing was proposed in the late 1950s (Brown, 1959; Holt,
+1957; Winters, 1960), and has motivated some of the most successful
+forecasting methods. Forecasts produced using exponential smoothing
+methods are weighted averages of past observations, with the weights
+decaying exponentially as the observations get older. In other words,
+the more recent the observation the higher the associated weight. This
+framework generates reliable forecasts quickly and for a wide range of
+time series, which is a great advantage and of major importance to
+applications in industry.
+
+The simple exponential smoothing model is a method used in time series
+analysis to predict future values based on historical observations. This
+model is based on the idea that future values of a time series will be
+influenced by past values, and that the influence of past values will
+decrease exponentially as you go back in time.
+
+The simple exponential smoothing model uses a smoothing factor, which is
+a number between 0 and 1 that indicates the relative importance given to
+past observations in predicting future values. A value of 1 indicates
+that all past observations are given equal importance, while a value of
+0 indicates that only the latest observation is considered.
+
+The simple exponential smoothing model can be expressed mathematically
+as:
+
+$$\hat{y}_{T+1|T} = \alpha y_T + \alpha(1-\alpha) y_{T-1} + \alpha(1-\alpha)^2 y_{T-2}+ \cdots,$$
+
+where $y_T$ is the observed value in period $t$, $\hat{y}_{T+1|T}$ is
+the predicted value for the next period, y $(t-1)$ is the observed value
+in the previous period, and $\alpha$ is the smoothing factor.
+
+The simple exponential smoothing model is a widely used forecasting
+model due to its simplicity and ease of use. However, it also has its
+limitations, as it cannot capture complex patterns in the data and is
+not suitable for time series with trends or seasonal patterns.
+
+## Building of Simple exponential smoothing model
+
+The simplest of the exponentially smoothing methods is naturally called
+simple exponential smoothing (SES). This method is suitable for
+forecasting data with no clear trend or seasonal pattern.
+
+Using the naïve method, all forecasts for the future are equal to the
+last observed value of the series, $$\hat{y}_{T+h|T} = y_{T},$$
+
+for $h=1,2,\dots$. Hence, the naïve method assumes that the most recent
+observation is the only important one, and all previous observations
+provide no information for the future. This can be thought of as a
+weighted average where all of the weight is given to the last
+observation.
+
+Using the average method, all future forecasts are equal to a simple
+average of the observed data,
+$$\hat{y}_{T+h|T} = \frac1T \sum_{t=1}^T y_t,$$
+
+for $h=1,2,\dots$ Hence, the average method assumes that all
+observations are of equal importance, and gives them equal weights when
+generating forecasts.
+
+We often want something between these two extremes. For example, it may
+be sensible to attach larger weights to more recent observations than to
+observations from the distant past. This is exactly the concept behind
+simple exponential smoothing. Forecasts are calculated using weighted
+averages, where the weights decrease exponentially as observations come
+from further in the past — the smallest weights are associated with the
+oldest observations:
+
+where $0 \le \alpha \le 1$ is the smoothing parameter. The
+one-step-ahead forecast for time $T+1$ is a weighted average of all of
+the observations in the series $y_1,\dots,y_T$. The rate at which the
+weights decrease is controlled by the parameter $\alpha$.
+
+For any $\alpha$ between 0 and 1, the weights attached to the
+observations decrease exponentially as we go back in time, hence the
+name “exponential smoothing”. If $\alpha$ is small (i.e., close to 0),
+more weight is given to observations from the more distant past. If
+$\alpha$ is large (i.e., close to 1), more weight is given to the more
+recent observations. For the extreme case where $\alpha=1$,
+$\hat{y}_{T+1|T}=y_T$ and the forecasts are equal to the naïve
+forecasts.
+
+We present two equivalent forms of simple exponential smoothing, each of
+which leads to the forecast Equation (1).
+
+### Weighted average form
+
+The forecast at time $T+1$ is equal to a weighted average between the
+most recent observation $y_T$ and the previous forecast
+$\hat{y}_{T|T-1}$:
+
+$$\hat{y}_{T+1|T} = \alpha y_T + (1-\alpha) \hat{y}_{T|T-1},$$
+
+where $0 \le \alpha \le 1$ is the smoothing parameter. Similarly, we can
+write the fitted values as
+$$\hat{y}_{t+1|t} = \alpha y_t + (1-\alpha) \hat{y}_{t|t-1},$$
+
+for $t=1,\dots,T$. (Recall that fitted values are simply one-step
+forecasts of the training data.)
+
+The process has to start somewhere, so we let the first fitted value at
+time 1 be denoted by $\ell_{0}$ (which we will have to estimate). Then
+
+Substituting each equation into the following equation, we obtain
+
+The last term becomes tiny for large $T$. So, the weighted average form
+leads to the same forecast Equation (1).
+
+### Component form
+
+An alternative representation is the component form. For simple
+exponential smoothing, the only component included is the level,
+$\ell_{t}$. Component form representations of exponential smoothing
+methods comprise a forecast equation and a smoothing equation for each
+of the components included in the method. The component form of simple
+exponential smoothing is given by:
+
+where $\ell_{t}$ is the level (or the smoothed value) of the series at
+time $t$. Setting $h=1$ gives the fitted values, while setting $t=T$
+gives the true forecasts beyond the training data.
+
+The forecast equation shows that the forecast value at time $t+1$ is the
+estimated level at time $t$. The smoothing equation for the level
+(usually referred to as the level equation) gives the estimated level of
+the series at each period $t$.
+
+If we replace $\ell_{t}$ with $\hat{y}_{t+1|t}$ and $\ell_{t-1}$ with
+$\hat{y}_{t|t-1}$ in the smoothing equation, we will recover the
+weighted average form of simple exponential smoothing.
+
+The component form of simple exponential smoothing is not particularly
+useful on its own, but it will be the easiest form to use when we start
+adding other components.
+
+### Flat forecasts
+
+Simple exponential smoothing has a “flat” forecast function:
+
+$$\hat{y}_{T+h|T} = \hat{y}_{T+1|T}=\ell_T, \qquad h=2,3,\dots.$$
+
+That is, all forecasts take the same value, equal to the last level
+component. Remember that these forecasts will only be suitable if the
+time series has no trend or seasonal component.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
+plt.style.use('grayscale') # fivethirtyeight grayscale classic
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#008080', # #212946
+ 'axes.facecolor': '#008080',
+ 'savefig.facecolor': '#008080',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#000000', #2A3459
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+
+```python
+import pandas as pd
+df=pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/ads.csv")
+df.head()
+```
+
+| | Time | Ads |
+|-----|---------------------|--------|
+| 0 | 2017-09-13T00:00:00 | 80115 |
+| 1 | 2017-09-13T01:00:00 | 79885 |
+| 2 | 2017-09-13T02:00:00 | 89325 |
+| 3 | 2017-09-13T03:00:00 | 101930 |
+| 4 | 2017-09-13T04:00:00 | 121630 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df
+```
+
+| | ds | y | unique_id |
+|-----|---------------------|--------|-----------|
+| 0 | 2017-09-13T00:00:00 | 80115 | 1 |
+| 1 | 2017-09-13T01:00:00 | 79885 | 1 |
+| 2 | 2017-09-13T02:00:00 | 89325 | 1 |
+| ... | ... | ... | ... |
+| 213 | 2017-09-21T21:00:00 | 103080 | 1 |
+| 214 | 2017-09-21T22:00:00 | 95155 | 1 |
+| 215 | 2017-09-21T23:00:00 | 80285 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y int64
+unique_id object
+dtype: object
+```
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore Data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints a random series from the dataset and is useful for
+basic EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### Autocorrelation plots
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=30, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+# Grafico
+plot_pacf(df["y"], lags=30, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+#plt.savefig("Gráfico de Densidad y qq")
+plt.show();
+```
+
+
+
+## Split the data into training and testing
+
+Let’s divide our data into sets
+
+1. Data to train our `Simple Exponential Smoothing (SES)`.
+2. Data to test our model
+
+For the test data we will use the last 30 hours to test and evaluate the
+performance of our model.
+
+```python
+train = df[df.ds\<='2017-09-20 17:00:00']
+test = df[df.ds>'2017-09-20 17:00:00']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((186, 3), (30, 3))
+```
+
+## Implementation of SimpleExponentialSmoothing with StatsForecast
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import SimpleExponentialSmoothing
+```
+
+### Instantiating Model
+
+We are going to build different models, for different values of alpha.
+
+```python
+horizon = len(test)
+# We call the model that we are going to use
+models = [SimpleExponentialSmoothing(alpha=0.1, alias="SES01"),
+ SimpleExponentialSmoothing(alpha=0.5,alias="SES05"),
+ SimpleExponentialSmoothing(alpha=0.8,alias="SES08")
+ ]
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [panda’s
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models, freq='h')
+```
+
+### Fit the Model
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[SES01,SES05,SES08])
+```
+
+Let’s see the results of our Simple
+`Simple Exponential Smoothing model (SES)`. We can observe it with the
+following instruction:
+
+```python
+result01=sf.fitted_[0,0].model_
+result05=sf.fitted_[0,1].model_
+result08=sf.fitted_[0,2].model_
+result01
+```
+
+``` text
+{'mean': array([126112.90072589]),
+ 'fitted': array([ nan, 80115. , 80092. , 81015.3 , 83106.77 ,
+ 86959.09 , 89910.69 , 91569.12 , 92691.7 , 94228.03 ,
+ 96417.73 , 99878.96 , 104793.06 , 110072.76 , 114136.98 ,
+ 117652.78 , 120897.5 , 123285.75 , 126026.18 , 129807.56 ,
+ 133450.3 , 134057.28 , 131241.05 , 127794.945, 123267.445,
+ 118953.2 , 114591.38 , 111642.74 , 110686.47 , 112131.32 ,
+ 112721.19 , 112371.57 , 111381.914, 110467.73 , 111004.95 ,
+ 112958.45 , 116095.11 , 119382.6 , 122359.336, 124927.41 ,
+ 127315.664, 129567.1 , 131667.39 , 133444.66 , 135152.19 ,
+ 134549.97 , 131476.47 , 127546.32 , 123068.19 , 118392.875,
+ 114066.586, 110923.92 , 108711.03 , 109682.93 , 110233.64 ,
+ 110304.27 , 109159.84 , 108662.36 , 108662.625, 110460.36 ,
+ 113457.83 , 117359.05 , 120250.64 , 123027.58 , 125498.32 ,
+ 127523.484, 129699.64 , 132702.17 , 135540.45 , 135538.4 ,
+ 133279.06 , 129971.164, 125735.55 , 121945.49 , 118635.445,
+ 116006.9 , 114852.71 , 114961.44 , 116360.3 , 118862.766,
+ 121420.484, 123603.44 , 124562.09 , 125229.88 , 126954.9 ,
+ 129996.91 , 132247.22 , 134396. , 136075.89 , 137532.81 ,
+ 138523.03 , 139923.22 , 140618.4 , 139081.06 , 136965.45 ,
+ 132938.9 , 129006.016, 125011.414, 121444.77 , 118357.8 ,
+ 116351.016, 115972.914, 117322.625, 119730.86 , 123013.77 ,
+ 125970.4 , 127490.36 , 129496.32 , 132657.69 , 136025.42 ,
+ 139100.88 , 141504.8 , 143084.81 , 144681.83 , 146215.64 ,
+ 148428.58 , 150575.72 , 149789.16 , 146105.73 , 141229.66 ,
+ 135274.2 , 129697.77 , 124563. , 120911.2 , 118799.08 ,
+ 119297.17 , 118499.95 , 116593.96 , 114700.06 , 112995.555,
+ 111952.5 , 112750.25 , 115050.73 , 117557.66 , 119974.89 ,
+ 122199.4 , 124515.46 , 126597.414, 128978.67 , 132232.81 ,
+ 134351.03 , 134387.92 , 131655.62 , 127994.57 , 123146.61 ,
+ 118665.445, 114265.91 , 111038.31 , 109729.484, 110691.03 ,
+ 110933.43 , 109728.086, 108155.28 , 106705.75 , 106453.68 ,
+ 107783.305, 110603.98 , 114189.08 , 116686.67 , 119740.51 ,
+ 122259.95 , 125170.96 , 128261.86 , 131574.17 , 134917.77 ,
+ 134834.98 , 131909.98 , 128004.484, 123131.04 , 118815.94 ,
+ 114745.34 , 111849.305, 110665.375, 111986.836, 112421.66 ,
+ 111608.49 , 110591.64 , 109295.98 , 109192.875, 110398.59 ,
+ 113443.734, 115954.86 , 118458.375, 120765.04 , 122847.53 ,
+ 124623.78 ], dtype=float32)}
+```
+
+As we can see, the result obtained above has an output in a dictionary,
+to extract each element from the dictionary we are going to use the
+`.get()` function to extract the element and then we are going to save
+it in a `pd.DataFrame()`.
+
+```python
+fitted=pd.DataFrame(result01.get("fitted"), columns=["fitted01"])
+fitted["fitted05"]=result05.get("fitted")
+fitted["fitted08"]=result08.get("fitted")
+fitted["ds"]=df["ds"]
+fitted
+```
+
+| | fitted01 | fitted05 | fitted08 | ds |
+|-----|---------------|-----------|---------------|---------------------|
+| 0 | NaN | NaN | NaN | 2017-09-13 00:00:00 |
+| 1 | 80115.000000 | 80115.00 | 80115.000000 | 2017-09-13 01:00:00 |
+| 2 | 80092.000000 | 80000.00 | 79931.000000 | 2017-09-13 02:00:00 |
+| ... | ... | ... | ... | ... |
+| 183 | 120765.039062 | 139195.00 | 141302.828125 | 2017-09-20 15:00:00 |
+| 184 | 122847.531250 | 140392.50 | 141532.562500 | 2017-09-20 16:00:00 |
+| 185 | 124623.781250 | 140501.25 | 140794.515625 | 2017-09-20 17:00:00 |
+
+```python
+sns.lineplot(df, x="ds", y="y", label="Actual", linewidth=2)
+sns.lineplot(fitted,x="ds", y="fitted01", label="Fitted01", linestyle="--", )
+sns.lineplot(fitted, x="ds", y="fitted05", label="Fitted05", color="lime")
+sns.lineplot(fitted, x="ds", y="fitted08", label="Fitted08")
+plt.title("Ads watched (hourly data)");
+plt.show()
+```
+
+
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 30 hours ahead.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min.
+
+```python
+# Prediction
+Y_hat = sf.forecast(df=train, h=horizon, fitted=True)
+Y_hat.head()
+```
+
+| | unique_id | ds | SES01 | SES05 | SES08 |
+|-----|-----------|---------------------|---------------|------------|--------------|
+| 0 | 1 | 2017-09-20 18:00:00 | 126112.898438 | 140008.125 | 139770.90625 |
+| 1 | 1 | 2017-09-20 19:00:00 | 126112.898438 | 140008.125 | 139770.90625 |
+| 2 | 1 | 2017-09-20 20:00:00 | 126112.898438 | 140008.125 | 139770.90625 |
+| 3 | 1 | 2017-09-20 21:00:00 | 126112.898438 | 140008.125 | 139770.90625 |
+| 4 | 1 | 2017-09-20 22:00:00 | 126112.898438 | 140008.125 | 139770.90625 |
+
+```python
+values=sf.forecast_fitted_values()
+values.head()
+```
+
+| | unique_id | ds | y | SES01 | SES05 | SES08 |
+|-----|-----------|---------------------|----------|--------------|----------|--------------|
+| 0 | 1 | 2017-09-13 00:00:00 | 80115.0 | NaN | NaN | NaN |
+| 1 | 1 | 2017-09-13 01:00:00 | 79885.0 | 80115.000000 | 80115.00 | 80115.000000 |
+| 2 | 1 | 2017-09-13 02:00:00 | 89325.0 | 80092.000000 | 80000.00 | 79931.000000 |
+| 3 | 1 | 2017-09-13 03:00:00 | 101930.0 | 81015.296875 | 84662.50 | 87446.203125 |
+| 4 | 1 | 2017-09-13 04:00:00 | 121630.0 | 83106.773438 | 93296.25 | 99033.242188 |
+
+### Predict method
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon). \* `h (int):` represents the forecast $h$ steps into the
+future. In this case, 30 hours ahead.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the `y hat` values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+forecast_df = sf.predict(h=horizon)
+forecast_df
+```
+
+| | unique_id | ds | SES01 | SES05 | SES08 |
+|-----|-----------|---------------------|---------------|------------|--------------|
+| 0 | 1 | 2017-09-20 18:00:00 | 126112.898438 | 140008.125 | 139770.90625 |
+| 1 | 1 | 2017-09-20 19:00:00 | 126112.898438 | 140008.125 | 139770.90625 |
+| 2 | 1 | 2017-09-20 20:00:00 | 126112.898438 | 140008.125 | 139770.90625 |
+| ... | ... | ... | ... | ... | ... |
+| 27 | 1 | 2017-09-21 21:00:00 | 126112.898438 | 140008.125 | 139770.90625 |
+| 28 | 1 | 2017-09-21 22:00:00 | 126112.898438 | 140008.125 | 139770.90625 |
+| 29 | 1 | 2017-09-21 23:00:00 | 126112.898438 | 140008.125 | 139770.90625 |
+
+```python
+sf.plot(train, forecast_df)
+```
+
+
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 30 hourly `(n_windows=)`, forecasting every second months
+`(step_size=30)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 30 hours ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=df,
+ h=horizon,
+ step_size=30,
+ n_windows=3)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` series identifier
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the `n_windows`.
+- `y:` true value
+- `model:` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df
+```
+
+| | unique_id | ds | cutoff | y | SES01 | SES05 | SES08 |
+|-----|-----------|---------------------|---------------------|----------|---------------|------------|---------------|
+| 0 | 1 | 2017-09-18 06:00:00 | 2017-09-18 05:00:00 | 99440.0 | 118499.953125 | 109816.250 | 112747.695312 |
+| 1 | 1 | 2017-09-18 07:00:00 | 2017-09-18 05:00:00 | 97655.0 | 118499.953125 | 109816.250 | 112747.695312 |
+| 2 | 1 | 2017-09-18 08:00:00 | 2017-09-18 05:00:00 | 97655.0 | 118499.953125 | 109816.250 | 112747.695312 |
+| ... | ... | ... | ... | ... | ... | ... | ... |
+| 87 | 1 | 2017-09-21 21:00:00 | 2017-09-20 17:00:00 | 103080.0 | 126112.898438 | 140008.125 | 139770.906250 |
+| 88 | 1 | 2017-09-21 22:00:00 | 2017-09-20 17:00:00 | 95155.0 | 126112.898438 | 140008.125 | 139770.906250 |
+| 89 | 1 | 2017-09-21 23:00:00 | 2017-09-20 17:00:00 | 80285.0 | 126112.898438 | 140008.125 | 139770.906250 |
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=24), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | SES01 | SES05 | SES08 |
+|-----|-----------|--------|--------------|--------------|--------------|
+| 0 | 1 | mae | 25173.939583 | 29390.875000 | 29311.802083 |
+| 1 | 1 | mape | 0.255088 | 0.316440 | 0.315339 |
+| 2 | 1 | mase | 3.110288 | 3.631298 | 3.621528 |
+| 3 | 1 | rmse | 28923.395381 | 36184.340869 | 36027.710540 |
+| 4 | 1 | smape | 0.109972 | 0.124803 | 0.124542 |
+
+## References
+
+1. [Changquan Huang • Alla Petukhina. Springer series (2022). Applied
+ Time Series Analysis and Forecasting with
+ Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2)
+2. Ivan Svetunkov. [Forecasting and Analytics with the Augmented
+ Dynamic Adaptive Model (ADAM)](https://openforecast.org/adam/)
+3. [James D. Hamilton. Time Series Analysis Princeton University Press,
+ Princeton, New Jersey, 1st Edition,
+ 1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Nixtla SeasonalExponentialSmoothing
+ API](../../src/core/models.html#simpleexponentialsmoothing)
+5. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+6. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+7. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/models/standardtheta.html.mdx b/statsforecast/docs/models/standardtheta.html.mdx
new file mode 100644
index 00000000..453ab0ec
--- /dev/null
+++ b/statsforecast/docs/models/standardtheta.html.mdx
@@ -0,0 +1,790 @@
+---
+title: Standard Theta Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `Standard Theta Model` with
+> `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation` in other.
+
+The text in this article is largely taken from: 1. [Jose A. Fiorucci,
+Tiago R. Pellegrini, Francisco Louzada, Fotios Petropoulos, Anne B.
+Koehler (2016). “Models for optimising the theta method and their
+relationship to state space models”. International Journal of
+Forecasting](https://www.sciencedirect.com/science/article/pii/S0169207016300243).
+2. [V. Assimakopoulos, K. Nikolopoulos, “The theta model: a
+decomposition approach to
+forecasting”](https://www.sciencedirect.com/science/article/abs/pii/S0169207000000662)
+3. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+Principles and Practice (3rd ed)”](https://otexts.com/fpp3/tscv.html).
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [Standard Theta](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of StandardTheta with
+ StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+The Theta method [(Assimakopoulos & Nikolopoulos, 2000, hereafter
+A&N)](https://www.sciencedirect.com/science/article/abs/pii/S0169207000000662)
+is applied to non-seasonal or deseasonalised time series, where the
+deseasonalisation is usually performed via the multiplicative classical
+decomposition. The method decomposes the original time series into two
+new lines through the so-called theta coefficients, denoted by
+${\theta}_1$ and ${\theta}_2$ for
+${\theta}_1, {\theta}_2 \in \mathbb{R}$, which are applied to the second
+difference of the data. The second differences are reduced when
+${\theta}<1$, resulting in a better approximation of the long-term
+behaviour of the series (Assimakopoulos, 1995). If ${\theta}$ is equal
+to zero, the new line is a straight line. When ${\theta}>1$ the local
+curvatures are increased, magnifying the short-term movements of the
+time series (A&N). The new lines produced are called theta lines,
+denoted here by $\text{Z}(\theta_1)$ and $\text{Z}(\theta_2)$. These
+lines have the same mean value and slope as the original data, but the
+local curvatures are either filtered out or enhanced, depending on the
+value of the $\theta$ coefficient.
+
+In other words, the decomposition process has the advantage of
+exploiting information in the data that usually cannot be captured and
+modelled completely through the extrapolation of the original time
+series. The theta lines can be regarded as new time series and are
+extrapolated separately using an appropriate forecasting method. Once
+the extrapolation of each theta line has been completed, recomposition
+takes place through a combination scheme in order to calculate the point
+forecasts of the original time series. Combining has long been
+considered as a useful practice in the forecasting literature (for
+example, [Clemen, 1989, Makridakis and Winkler, 1983, Petropoulos et
+al., 2014]()), and therefore its application to the Theta method is
+expected to result in more accurate and robust forecasts.
+
+The Theta method is quite versatile in terms of choosing the number of
+theta lines, the theta coefficients and the extrapolation methods, and
+combining these to obtain robust forecasts. However, A&N proposed a
+simplified version involving the use of only two theta lines with
+prefixed $\theta$ coefficients that are extrapolated over time using a
+linear regression (LR) model for the theta line with ${\theta}_1 =0$ and
+simple exponential smoothing (SES) for the theta line with
+${\theta}_2 =2$. The final forecasts are produced by combining the
+forecasts of the two theta lines with equal weights.
+
+The performance of the Theta method has also been confirmed by other
+empirical studies (for example Nikolopoulos et al., 2012, Petropoulos
+and Nikolopoulos, 2013). Moreover, Hyndman and Billah (2003), hereafter
+H&B, showed that the simple exponential smoothing with drift model
+(SES-d) is a statistical model for the simplified version of the Theta
+method. More recently, Thomakos and Nikolopoulos (2014) provided
+additional theoretical insights, while Thomakos and Nikolopoulos (2015)
+derived new theoretical formulations for the application of the method
+to multivariate time series, and investigated the conditions under which
+the bivariate Theta method is expected to forecast better than the
+univariate one. Despite these advances, we believe that the Theta method
+deserves more attention from the forecasting community, given its
+simplicity and superior forecasting performance.
+
+One key aspect of the Theta method is that, by definition, it is
+dynamic. One can choose different theta lines and combine the produced
+forecasts using either equal or unequal weights. However, AN limit this
+important property by fixing the theta coefficients to have predefined
+values.
+
+## Standard Theta Model
+
+Assimakopoulos and Nikolopoulo for standard theta model proposed the
+Theta line as the solution of the equation
+
+$$
+\begin{equation}
+D^2 \zeta_t(\theta) = \theta D^2 Y_t, t = 1,\cdots,T \tag 1
+\end{equation}
+$$
+
+where $Y_1, \cdots , Y_T$ represent the original time series data and
+$DX_t = (X_t − X_{t−1})$. The initial values $\zeta_1$ and $\zeta_2$ are
+obtained by minimizing $\sum_{i=1}^{T} [Y_t - \zeta_t (\theta) ]^2$.
+However, the analytical solution of (1) is given by
+
+$$
+\begin{equation}
+\zeta_t(\theta)=\theta Y_t +(1−\theta)(A_T +B_T t),\ t=1, \cdots, T, \tag 2
+\end{equation}
+$$
+
+where $A_T$ and $B_T$ are the minimum square coefficients of a simple
+linear regression over $Y_1, \cdots,Y_T$ against $1, \cdots , T$ which
+are only dependent on the original data and given as follow
+
+$$
+\begin{equation}
+A_T=\frac{1}{T} \sum_{i=1}^{T} Y_t - \frac{T+1}{2} B_T \tag 3
+\end{equation}
+$$
+
+$$
+\begin{equation}
+B_T=\frac{6}{T^2 - 1} (\frac{2}{T} \sum_{t=1}^{T} tY_t - \frac{T+1}{T} \sum_{t=1}^{T} Y_t \tag 4
+\end{equation}
+$$
+
+Theta lines can be understood as functions of the linear regression
+model directly applied to the data from this perspective. Indeed, the
+Theta method’s projections for h steps ahead are an ad hoc combination
+(50 percent - 50 percent) of the linear extrapolations of $\zeta(0)$ and
+$\zeta(2)$.
+
+- When $\theta < 1$ is applied to the second differences of the data,
+ the decomposition process is defined by a theta coefficient, which
+ reduces the second differences and improves the approximation of
+ series behavior.
+
+- If $\theta = 0$, the deconstructed line is turned into a constant
+ straight line. (see Fig)
+
+- If $\theta > 1$ then the short term movements of the analyzed series
+ show more local curvatures (see fig)
+
+
+
+Figure
+
+
+We will refer to the above setup as the standard Theta method. The steps
+for building the theta method are as follows:
+
+1. **Deseasonalisation:** Firstly, the time series data is tested for
+ statistically significant seasonal behaviour. A time series is
+ seasonal if
+
+$$|\rho_m| > q_{1- \frac{\alpha}{2} } \sqrt{\frac{1+2 \sum_{i=1}^{m-1} \rho_{i}^{2} }{T} }$$
+
+where ρk denotes the lag $k$ autocorrelation function, $m$ is the number
+of the periods within a seasonal cycle (for example, 12 for monthly
+data), $T$ is the sample size, $q$ is the quantile function of the
+standard normal distribution, and $(1 − a)\%$ is the confidence level.
+Assimakopoulos and Nikolopoulo \[Standar Theta model\] opted for a 90%
+confidence level. If the time series is identified as seasonal, then it
+is deseasonalised via the classical decomposition method, assuming the
+seasonal component to have a multiplicative relationship.
+
+1. **Decomposition:** The second step consits for the decomposition of
+ the seasonally adjusted time series into two Theta lines, the
+ `linear regression` line $\zeta(0)$ and the theta line $\zeta(2)$.
+
+2. **Extrapolation:** $\zeta(2)$ is extrapolated using
+ `simple exponential smoothing (SES)`, while $\zeta(0)$ is
+ extrapolated as a normal `linear regression` line.
+
+3. **Combination:** the final forecast is a combination of the
+ forecasts of the two $\theta$ lines using equal weights.
+
+4. Reseasonalisation: In the presence of seasonality in first step,
+ then the final forecasts are multiplied by the respective seasonal
+ indices.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
+plt.style.use('grayscale') # fivethirtyeight grayscale classic
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#008080', # #212946
+ 'axes.facecolor': '#008080',
+ 'savefig.facecolor': '#008080',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#000000', #2A3459
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+### Read Data
+
+```python
+import pandas as pd
+df = pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/milk_production.csv", usecols=[1,2])
+df.head()
+```
+
+| | month | production |
+|-----|------------|------------|
+| 0 | 1962-01-01 | 589 |
+| 1 | 1962-02-01 | 561 |
+| 2 | 1962-03-01 | 640 |
+| 3 | 1962-04-01 | 656 |
+| 4 | 1962-05-01 | 727 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|------------|-----|-----------|
+| 0 | 1962-01-01 | 589 | 1 |
+| 1 | 1962-02-01 | 561 | 1 |
+| 2 | 1962-03-01 | 640 | 1 |
+| 3 | 1962-04-01 | 656 | 1 |
+| 4 | 1962-05-01 | 727 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y int64
+unique_id object
+dtype: object
+```
+
+We can see that our time variable `(ds)` is in an object format, we need
+to convert to a date format
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore Data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints a random series from the dataset and is useful for
+basic EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### Autocorrelation plots
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=30, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+plot_pacf(df["y"], lags=30, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+plt.show();
+```
+
+
+
+### Decomposition of the time series
+
+How to decompose a time series and why?
+
+In time series analysis to forecast new values, it is very important to
+know past data. More formally, we can say that it is very important to
+know the patterns that values follow over time. There can be many
+reasons that cause our forecast values to fall in the wrong direction.
+Basically, a time series consists of four components. The variation of
+those components causes the change in the pattern of the time series.
+These components are:
+
+- **Level:** This is the primary value that averages over time.
+- **Trend:** The trend is the value that causes increasing or
+ decreasing patterns in a time series.
+- **Seasonality:** This is a cyclical event that occurs in a time
+ series for a short time and causes short-term increasing or
+ decreasing patterns in a time series.
+- **Residual/Noise:** These are the random variations in the time
+ series.
+
+Combining these components over time leads to the formation of a time
+series. Most time series consist of level and noise/residual and trend
+or seasonality are optional values.
+
+If seasonality and trend are part of the time series, then there will be
+effects on the forecast value. As the pattern of the forecasted time
+series may be different from the previous time series.
+
+The combination of the components in time series can be of two types: \*
+Additive \* Multiplicative
+
+### Additive time series
+
+If the components of the time series are added to make the time series.
+Then the time series is called the additive time series. By
+visualization, we can say that the time series is additive if the
+increasing or decreasing pattern of the time series is similar
+throughout the series. The mathematical function of any additive time
+series can be represented by:
+$$y(t) = level + Trend + seasonality + noise$$
+
+### Multiplicative time series
+
+If the components of the time series are multiplicative together, then
+the time series is called a multiplicative time series. For
+visualization, if the time series is having exponential growth or
+decline with time, then the time series can be considered as the
+multiplicative time series. The mathematical function of the
+multiplicative time series can be represented as.
+
+$$y(t) = Level * Trend * seasonality * Noise$$
+
+### Additive
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+a = seasonal_decompose(df["y"], model = "additive", period=12)
+a.plot();
+```
+
+
+
+### Multiplicative
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+a = seasonal_decompose(df["y"], model = "Multiplicative", period=12)
+a.plot();
+```
+
+
+
+## Split the data into training and testing
+
+Let’s divide our data into sets 1. Data to train our `Theta` model 2.
+Data to test our model
+
+For the test data we will use the last 12 months to test and evaluate
+the performance of our model.
+
+```python
+train = df[df.ds\<='1974-12-01']
+test = df[df.ds>'1974-12-01']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((156, 3), (12, 3))
+```
+
+## Implementation of StandardTheta with StatsForecast
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import Theta
+```
+
+### Instantiating Model
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/) by the
+master, Rob Hyndmann, can be useful for `season_length`.
+
+```python
+season_length = 12 # Monthly data
+horizon = len(test) # number of predictions
+
+models = [Theta(season_length=season_length,
+ decomposition_type="additive")] # multiplicative additive
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [panda’s
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models, freq='MS')
+```
+
+### Fit Model
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[Theta])
+```
+
+Let’s see the results of our Theta model. We can observe it with the
+following instruction:
+
+```python
+result=sf.fitted_[0,0].model_
+print(result.keys())
+print(result['fit'])
+```
+
+``` text
+dict_keys(['mse', 'amse', 'fit', 'residuals', 'm', 'states', 'par', 'n', 'modeltype', 'mean_y', 'decompose', 'decomposition_type', 'seas_forecast', 'fitted'])
+results(x=array([225.82002697, 0.76015625]), fn=10.638733596938778, nit=19, simplex=array([[241.83142594, 0.76274414],
+ [225.82002697, 0.76015625],
+ [212.41789302, 0.76391602]]))
+```
+
+Let us now visualize the residuals of our models.
+
+As we can see, the result obtained above has an output in a dictionary,
+to extract each element from the dictionary we are going to use the
+`.get()` function to extract the element and then we are going to save
+it in a `pd.DataFrame()`.
+
+```python
+residual=pd.DataFrame(result.get("residuals"), columns=["residual Model"])
+residual
+```
+
+| | residual Model |
+|-----|----------------|
+| 0 | -17.596375 |
+| 1 | -46.997192 |
+| 2 | 23.093933 |
+| ... | ... |
+| 153 | -59.003235 |
+| 154 | -91.150085 |
+| 155 | -42.749451 |
+
+```python
+import scipy.stats as stats
+
+fig, axs = plt.subplots(nrows=2, ncols=2)
+
+residual.plot(ax=axs[0,0])
+axs[0,0].set_title("Residuals");
+
+sns.distplot(residual, ax=axs[0,1]);
+axs[0,1].set_title("Density plot - Residual");
+
+stats.probplot(residual["residual Model"], dist="norm", plot=axs[1,0])
+axs[1,0].set_title('Plot Q-Q')
+
+plot_acf(residual, lags=35, ax=axs[1,1],color="fuchsia")
+axs[1,1].set_title("Autocorrelation");
+
+plt.show();
+```
+
+
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[90]` means that
+ the model expects the real value to be inside that interval 90% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min.
+
+```python
+# Prediction
+Y_hat = sf.forecast(df=train, h=horizon, fitted=True)
+Y_hat
+```
+
+| | unique_id | ds | Theta |
+|-----|-----------|------------|------------|
+| 0 | 1 | 1975-01-01 | 838.559814 |
+| 1 | 1 | 1975-02-01 | 800.188232 |
+| 2 | 1 | 1975-03-01 | 893.472900 |
+| ... | ... | ... | ... |
+| 9 | 1 | 1975-10-01 | 816.166931 |
+| 10 | 1 | 1975-11-01 | 786.962036 |
+| 11 | 1 | 1975-12-01 | 823.826538 |
+
+```python
+values=sf.forecast_fitted_values()
+values.head()
+```
+
+| | unique_id | ds | y | Theta |
+|-----|-----------|------------|-------|------------|
+| 0 | 1 | 1962-01-01 | 589.0 | 606.596375 |
+| 1 | 1 | 1962-02-01 | 561.0 | 607.997192 |
+| 2 | 1 | 1962-03-01 | 640.0 | 616.906067 |
+| 3 | 1 | 1962-04-01 | 656.0 | 608.873047 |
+| 4 | 1 | 1962-05-01 | 727.0 | 607.395142 |
+
+Adding 95% confidence interval with the forecast method
+
+```python
+sf.forecast(df=train, h=horizon, level=[95])
+```
+
+| | unique_id | ds | Theta | Theta-lo-95 | Theta-hi-95 |
+|-----|-----------|------------|------------|-------------|-------------|
+| 0 | 1 | 1975-01-01 | 838.559814 | 741.324280 | 954.365540 |
+| 1 | 1 | 1975-02-01 | 800.188232 | 640.785583 | 944.996887 |
+| 2 | 1 | 1975-03-01 | 893.472900 | 705.123901 | 1064.757324 |
+| ... | ... | ... | ... | ... | ... |
+| 9 | 1 | 1975-10-01 | 816.166931 | 539.706665 | 1083.791626 |
+| 10 | 1 | 1975-11-01 | 786.962036 | 487.945831 | 1032.029053 |
+| 11 | 1 | 1975-12-01 | 823.826538 | 512.674500 | 1101.965576 |
+
+```python
+sf.plot(train, Y_hat)
+```
+
+
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level (list of floats):` this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, `level=[95]` means that
+ the model expects the real value to be inside that interval 95% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+sf.predict(h=horizon)
+```
+
+| | unique_id | ds | Theta |
+|-----|-----------|------------|------------|
+| 0 | 1 | 1975-01-01 | 838.559814 |
+| 1 | 1 | 1975-02-01 | 800.188232 |
+| 2 | 1 | 1975-03-01 | 893.472900 |
+| ... | ... | ... | ... |
+| 9 | 1 | 1975-10-01 | 816.166931 |
+| 10 | 1 | 1975-11-01 | 786.962036 |
+| 11 | 1 | 1975-12-01 | 823.826538 |
+
+```python
+forecast_df = sf.predict(h=horizon, level=[80,95])
+forecast_df
+```
+
+| | unique_id | ds | Theta | Theta-lo-80 | Theta-hi-80 | Theta-lo-95 | Theta-hi-95 |
+|-----|-----------|------------|------------|-------------|-------------|-------------|-------------|
+| 0 | 1 | 1975-01-01 | 838.559814 | 765.496094 | 927.260071 | 741.324280 | 954.365540 |
+| 1 | 1 | 1975-02-01 | 800.188232 | 701.729736 | 898.807434 | 640.785583 | 944.996887 |
+| 2 | 1 | 1975-03-01 | 893.472900 | 758.480957 | 1006.847595 | 705.123901 | 1064.757324 |
+| ... | ... | ... | ... | ... | ... | ... | ... |
+| 9 | 1 | 1975-10-01 | 816.166931 | 611.404236 | 991.667175 | 539.706665 | 1083.791626 |
+| 10 | 1 | 1975-11-01 | 786.962036 | 561.990540 | 969.637634 | 487.945831 | 1032.029053 |
+| 11 | 1 | 1975-12-01 | 823.826538 | 591.283508 | 1029.491211 | 512.674500 | 1101.965576 |
+
+```python
+sf.plot(train, test.merge(forecast_df), level=[80, 95])
+```
+
+
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=5)`, forecasting every second months
+`(step_size=12)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 12 months ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=train,
+ h=horizon,
+ step_size=12,
+ n_windows=3)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` series identifier
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the n_windows.
+- `y:` true value
+- `"model":` columns with the model’s name and fitted value.
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=season_length), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | Theta |
+|-----|-----------|--------|----------|
+| 0 | 1 | mae | 8.111287 |
+| 1 | 1 | mape | 0.009649 |
+| 2 | 1 | mase | 0.364780 |
+| 3 | 1 | rmse | 9.730347 |
+| 4 | 1 | smape | 0.004829 |
+
+## Acknowledgements
+
+We would like to thank [Naren
+Castellon](https://www.linkedin.com/in/naren-castellon-1541b8101/?originalSubdomain=pa)
+for writing this tutorial.
+
+## References
+
+1. [Jose A. Fiorucci, Tiago R. Pellegrini, Francisco Louzada, Fotios
+ Petropoulos, Anne B. Koehler (2016). “Models for optimising the
+ theta method and their relationship to state space models”.
+ International Journal of
+ Forecasting](https://www.sciencedirect.com/science/article/pii/S0169207016300243).
+2. [V. Assimakopoulos, K. Nikolopoulos, “The theta model: a
+ decomposition approach to
+ forecasting”](https://www.sciencedirect.com/science/article/abs/pii/S0169207000000662)
+3. [Nixtla StandardTheta API](../../src/core/models.html#theta)
+4. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+5. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+6. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/models/tsb.html.mdx b/statsforecast/docs/models/tsb.html.mdx
new file mode 100644
index 00000000..85022b72
--- /dev/null
+++ b/statsforecast/docs/models/tsb.html.mdx
@@ -0,0 +1,689 @@
+---
+title: TSB Model
+---
+
+
+
+
+
+> Step-by-step guide on using the `TSB Model` with `Statsforecast`.
+
+During this walkthrough, we will become familiar with the main
+`StatsForecast` class and some relevant methods such as
+`StatsForecast.plot`, `StatsForecast.forecast` and
+`StatsForecast.cross_validation` in other.
+
+The text in this article is largely taken from: 1. [Changquan Huang •
+Alla Petukhina. Springer series (2022). Applied Time Series Analysis and
+Forecasting with
+Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2) 2.
+Ivan Svetunkov. [Forecasting and Analytics with the Augmented Dynamic
+Adaptive Model (ADAM)](https://openforecast.org/adam/) 3. [James D.
+Hamilton. Time Series Analysis Princeton University Press, Princeton,
+New Jersey, 1st Edition,
+1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+Principles and Practice (3rd ed)”](https://otexts.com/fpp3/tscv.html).
+
+## Table of Contents
+
+- [Introduction](#introduction)
+- [TSB](#model)
+- [Loading libraries and data](#loading)
+- [Explore data with the plot method](#plotting)
+- [Split the data into training and testing](#splitting)
+- [Implementation of TSB with StatsForecast](#implementation)
+- [Cross-validation](#cross_validate)
+- [Model evaluation](#evaluate)
+- [References](#references)
+
+## Introduction
+
+The Teunter-Syntetos-Babai (TSB) model is a model used in the field of
+inventory management and demand forecasting in time series. It was
+proposed by Teunter, Syntetos, and Babai in 2001 as an extension of
+Croston’s demand forecasting model.
+
+The TSB model is specifically used to forecast demand for products with
+intermittent demand characteristics, that is, products that experience
+periods of demand followed by periods of non-demand. It is designed to
+handle time series data with many zeros and variability in the intervals
+between non-null observations.
+
+The TSB model is based on two main components: the level model and the
+interval model. The level model estimates the level of demand when it
+occurs, while the interval model estimates the interval between demand
+occurrences. These two components combine to generate accurate forecasts
+of future demand.
+
+The TSB model has proven to be effective in intermittent demand
+forecasting and has been widely used in various industrial sectors.
+However, it is important to note that there are other models and
+approaches available for demand forecasting, and the choice of the
+appropriate model will depend on the specific characteristics of the
+data and the context in which it is applied.
+
+## TSB Model
+
+TSB (Teunter, Syntetos and Babai) is a new method proposed in 2011, the
+method replace the demand interval by demand probability which is
+updated every period. The reason for this is the Croston’s method only
+update demand when it occur, however in real life there are plenty of
+cases with many zero demands, therefore, the result of forecast will be
+unsuitable for estimating the risk of obsolescence because of the
+outdated information.
+
+In TSB method, the $D_t$ represent the demand occurrence indicator for
+period $t$, so :
+
+If $D_t=0$, then
+
+$$Z'_t=Z'_{t-1}$$
+
+$$D_t=D'_{t-1}+\beta (0- D'_{t-1})$$
+
+Otherwise $$Z'_t=Z'_{t-1}+\alpha(Z_t - Z'_{t-2})$$
+
+$$D'_t=D'_{t-1}+\beta(1-D'_{t-1})$$
+
+Hence, the forecast is given by
+
+$$Y'_t=D'_t \cdot Z'_t$$
+
+Where
+
+- $Y'_t:$ Average demand per period
+- $Z_t:$ Actual demand at period $t$
+- $Z'_t:$ Time between two positive demand
+- $D'_t:$ Estimate probability of a demand occurrence at the end of
+ period $t$
+- $\alpha, \beta:$ Smoothing Constant, $0 \leq \alpha, \beta \leq 1$
+
+### TSB General Properties
+
+The Teunter-Syntetos-Babai (TSB) model for time series has the following
+properties:
+
+1. Intermittent Demand Modelling: The TSB model is specifically
+ designed to forecast intermittent demand, which is characterized by
+ periods of non-demand followed by periods of demand. The model
+ efficiently addresses this characteristic of demand.
+
+2. Level and interval components: The TSB model is based on two main
+ components: the level model and the interval model. The level model
+ estimates the level of demand when it occurs, while the interval
+ model estimates the interval between demand occurrences.
+
+3. Handling data with many zeros: The TSB model can efficiently handle
+ time series data with many zeros, which are common in intermittent
+ demand. The model properly considers these zeros in the forecasting
+ process.
+
+4. Exponential Smoothing: The TSB model uses exponential smoothing
+ methods to estimate demand levels and intervals between occurrences.
+ Exponential smoothing is a widely used technique in time series
+ forecasting.
+
+5. Confidence interval estimation: The TSB model provides confidence
+ interval estimates for the generated forecasts. This allows having a
+ measure of the uncertainty associated with forecasts and facilitates
+ decision making.
+
+6. Simplicity and ease of implementation: The TSB model is relatively
+ simple and easy to implement compared to other more complex
+ approaches. It does not require sophisticated assumptions about the
+ distribution of demand and can be applied in a practical way.
+
+Those are some of the fundamental properties of the
+Teunter-Syntetos-Babai model in the context of time series and
+intermittent demand forecasting.
+
+## Loading libraries and data
+
+> **Tip**
+>
+> Statsforecast will be needed. To install, see
+> [instructions](../getting-started/0_Installation).
+
+Next, we import plotting libraries and configure the plotting style.
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
+plt.style.use('grayscale') # fivethirtyeight grayscale classic
+plt.rcParams['lines.linewidth'] = 1.5
+dark_style = {
+ 'figure.facecolor': '#008080', # #212946
+ 'axes.facecolor': '#008080',
+ 'savefig.facecolor': '#008080',
+ 'axes.grid': True,
+ 'axes.grid.which': 'both',
+ 'axes.spines.left': False,
+ 'axes.spines.right': False,
+ 'axes.spines.top': False,
+ 'axes.spines.bottom': False,
+ 'grid.color': '#000000', #2A3459
+ 'grid.linewidth': '1',
+ 'text.color': '0.9',
+ 'axes.labelcolor': '0.9',
+ 'xtick.color': '0.9',
+ 'ytick.color': '0.9',
+ 'font.size': 12 }
+plt.rcParams.update(dark_style)
+
+
+from pylab import rcParams
+rcParams['figure.figsize'] = (18,7)
+```
+
+
+```python
+import pandas as pd
+
+df=pd.read_csv("https://raw.githubusercontent.com/Naren8520/Serie-de-tiempo-con-Machine-Learning/main/Data/intermittend_demand2")
+df.head()
+```
+
+| | date | sales |
+|-----|---------------------|-------|
+| 0 | 2022-01-01 00:00:00 | 0 |
+| 1 | 2022-01-01 01:00:00 | 10 |
+| 2 | 2022-01-01 02:00:00 | 0 |
+| 3 | 2022-01-01 03:00:00 | 0 |
+| 4 | 2022-01-01 04:00:00 | 100 |
+
+The input to StatsForecast is always a data frame in long format with
+three columns: unique_id, ds and y:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp) column should be of a format expected by
+ Pandas, ideally YYYY-MM-DD for a date or YYYY-MM-DD HH:MM:SS for a
+ timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+```python
+df["unique_id"]="1"
+df.columns=["ds", "y", "unique_id"]
+df.head()
+```
+
+| | ds | y | unique_id |
+|-----|---------------------|-----|-----------|
+| 0 | 2022-01-01 00:00:00 | 0 | 1 |
+| 1 | 2022-01-01 01:00:00 | 10 | 1 |
+| 2 | 2022-01-01 02:00:00 | 0 | 1 |
+| 3 | 2022-01-01 03:00:00 | 0 | 1 |
+| 4 | 2022-01-01 04:00:00 | 100 | 1 |
+
+```python
+print(df.dtypes)
+```
+
+``` text
+ds object
+y int64
+unique_id object
+dtype: object
+```
+
+We can see that our time variable `(ds)` is in an object format, we need
+to convert to a date format
+
+```python
+df["ds"] = pd.to_datetime(df["ds"])
+```
+
+## Explore Data with the plot method
+
+Plot some series using the plot method from the StatsForecast class.
+This method prints a random series from the dataset and is useful for
+basic EDA.
+
+```python
+from statsforecast import StatsForecast
+
+StatsForecast.plot(df)
+```
+
+
+
+### Autocorrelation plots
+
+Autocorrelation (ACF) and partial autocorrelation (PACF) plots are
+statistical tools used to analyze time series. ACF charts show the
+correlation between the values of a time series and their lagged values,
+while PACF charts show the correlation between the values of a time
+series and their lagged values, after the effect of previous lagged
+values has been removed.
+
+ACF and PACF charts can be used to identify the structure of a time
+series, which can be helpful in choosing a suitable model for the time
+series. For example, if the ACF chart shows a repeating peak and valley
+pattern, this indicates that the time series is stationary, meaning that
+it has the same statistical properties over time. If the PACF chart
+shows a pattern of rapidly decreasing spikes, this indicates that the
+time series is invertible, meaning it can be reversed to get a
+stationary time series.
+
+The importance of the ACF and PACF charts is that they can help analysts
+better understand the structure of a time series. This understanding can
+be helpful in choosing a suitable model for the time series, which can
+improve the ability to predict future values of the time series.
+
+To analyze ACF and PACF charts:
+
+- Look for patterns in charts. Common patterns include repeating peaks
+ and valleys, sawtooth patterns, and plateau patterns.
+- Compare ACF and PACF charts. The PACF chart generally has fewer
+ spikes than the ACF chart.
+- Consider the length of the time series. ACF and PACF charts for
+ longer time series will have more spikes.
+- Use a confidence interval. The ACF and PACF plots also show
+ confidence intervals for the autocorrelation values. If an
+ autocorrelation value is outside the confidence interval, it is
+ likely to be significant.
+
+```python
+fig, axs = plt.subplots(nrows=1, ncols=2)
+
+plot_acf(df["y"], lags=30, ax=axs[0],color="fuchsia")
+axs[0].set_title("Autocorrelation");
+
+plot_pacf(df["y"], lags=30, ax=axs[1],color="lime")
+axs[1].set_title('Partial Autocorrelation')
+
+plt.show();
+```
+
+
+
+### Decomposition of the time series
+
+How to decompose a time series and why?
+
+In time series analysis to forecast new values, it is very important to
+know past data. More formally, we can say that it is very important to
+know the patterns that values follow over time. There can be many
+reasons that cause our forecast values to fall in the wrong direction.
+Basically, a time series consists of four components. The variation of
+those components causes the change in the pattern of the time series.
+These components are:
+
+- **Level:** This is the primary value that averages over time.
+- **Trend:** The trend is the value that causes increasing or
+ decreasing patterns in a time series.
+- **Seasonality:** This is a cyclical event that occurs in a time
+ series for a short time and causes short-term increasing or
+ decreasing patterns in a time series.
+- **Residual/Noise:** These are the random variations in the time
+ series.
+
+Combining these components over time leads to the formation of a time
+series. Most time series consist of level and noise/residual and trend
+or seasonality are optional values.
+
+If seasonality and trend are part of the time series, then there will be
+effects on the forecast value. As the pattern of the forecasted time
+series may be different from the previous time series.
+
+The combination of the components in time series can be of two types: \*
+Additive \* Multiplicative
+
+### Additive time series
+
+If the components of the time series are added to make the time series.
+Then the time series is called the additive time series. By
+visualization, we can say that the time series is additive if the
+increasing or decreasing pattern of the time series is similar
+throughout the series. The mathematical function of any additive time
+series can be represented by:
+$$y(t) = level + Trend + seasonality + noise$$
+
+### Multiplicative time series
+
+If the components of the time series are multiplicative together, then
+the time series is called a multiplicative time series. For
+visualization, if the time series is having exponential growth or
+decline with time, then the time series can be considered as the
+multiplicative time series. The mathematical function of the
+multiplicative time series can be represented as.
+
+$$y(t) = Level * Trend * seasonality * Noise$$
+
+```python
+from plotly.subplots import make_subplots
+```
+
+
+```python
+from statsmodels.tsa.seasonal import seasonal_decompose
+from plotly.subplots import make_subplots
+import plotly.graph_objects as go
+
+def plotSeasonalDecompose(
+ x,
+ model='additive',
+ filt=None,
+ period=None,
+ two_sided=True,
+ extrapolate_trend=0,
+ title="Seasonal Decomposition"):
+
+ result = seasonal_decompose(
+ x, model=model, filt=filt, period=period,
+ two_sided=two_sided, extrapolate_trend=extrapolate_trend)
+ fig = make_subplots(
+ rows=4, cols=1,
+ subplot_titles=["Observed", "Trend", "Seasonal", "Residuals"])
+ for idx, col in enumerate(['observed', 'trend', 'seasonal', 'resid']):
+ fig.add_trace(
+ go.Scatter(x=result.observed.index, y=getattr(result, col), mode='lines'),
+ row=idx+1, col=1,
+ )
+ return fig
+```
+
+
+```python
+plotSeasonalDecompose(
+ df["y"],
+ model="additive",
+ period=24,
+ title="Seasonal Decomposition")
+```
+
+
+
+## Split the data into training and testing
+
+Let’s divide our data into sets 1. Data to train our `TSB Model`. 2.
+Data to test our model
+
+For the test data we will use the last 500 Hours to test and evaluate
+the performance of our model.
+
+```python
+train = df[df.ds\<='2023-01-31 19:00:00']
+test = df[df.ds>'2023-01-31 19:00:00']
+```
+
+
+```python
+train.shape, test.shape
+```
+
+``` text
+((9500, 3), (500, 3))
+```
+
+## Implementation of `TSB Model` with StatsForecast
+
+### Load libraries
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import TSB
+```
+
+### Building Model
+
+Import and instantiate the models. Setting the argument is sometimes
+tricky. This article on [Seasonal
+periods](https://robjhyndman.com/hyndsight/seasonal-periods/) by the
+master, Rob Hyndmann, can be useful for `season_length`.
+
+```python
+season_length = 24 # Hourly data
+horizon = len(test) # number of predictions
+
+models = [TSB(alpha_d=0.8, alpha_p=0.9)]
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+models: a list of models. Select the models you want from models and
+import them.
+
+- `freq:` a string indicating the frequency of the data. (See [pandas’
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+- `n_jobs:` n_jobs: int, number of jobs used in the parallel
+ processing, use -1 for all cores.
+
+- `fallback_model:` a model to be used if a model fails.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+```python
+sf = StatsForecast(models=models, freq='h')
+```
+
+### Fit the Model
+
+```python
+sf.fit(df=train)
+```
+
+``` text
+StatsForecast(models=[TSB])
+```
+
+Let’s see the results of our `TSB Model`. We can observe it with the
+following instruction:
+
+```python
+result=sf.fitted_[0,0].model_
+result
+```
+
+``` text
+{'mean': array([65.58645721]),
+ 'fitted': array([ nan, 0. , 9. , ..., 14.937817 ,
+ 1.4937816 , 0.14937817], dtype=float32),
+ 'sigma': np.float32(63.87893)}
+```
+
+### Forecast Method
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the `StatsForecast.forecast` method
+instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The forecast method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 500 hours ahead.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min.
+
+```python
+Y_hat = sf.forecast(df=train, h=horizon)
+Y_hat
+```
+
+| | unique_id | ds | TSB |
+|-----|-----------|---------------------|-----------|
+| 0 | 1 | 2023-01-31 20:00:00 | 65.586456 |
+| 1 | 1 | 2023-01-31 21:00:00 | 65.586456 |
+| 2 | 1 | 2023-01-31 22:00:00 | 65.586456 |
+| ... | ... | ... | ... |
+| 497 | 1 | 2023-02-21 13:00:00 | 65.586456 |
+| 498 | 1 | 2023-02-21 14:00:00 | 65.586456 |
+| 499 | 1 | 2023-02-21 15:00:00 | 65.586456 |
+
+```python
+sf.plot(train, Y_hat)
+```
+
+
+
+### Predict method with confidence interval
+
+To generate forecasts use the predict method.
+
+The predict method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h (int):` represents the forecast h steps into the future. In this
+ case, 500 hours ahead.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+```python
+forecast_df = sf.predict(h=horizon)
+forecast_df
+```
+
+| | unique_id | ds | TSB |
+|-----|-----------|---------------------|-----------|
+| 0 | 1 | 2023-01-31 20:00:00 | 65.586456 |
+| 1 | 1 | 2023-01-31 21:00:00 | 65.586456 |
+| 2 | 1 | 2023-01-31 22:00:00 | 65.586456 |
+| ... | ... | ... | ... |
+| 497 | 1 | 2023-02-21 13:00:00 | 65.586456 |
+| 498 | 1 | 2023-02-21 14:00:00 | 65.586456 |
+| 499 | 1 | 2023-02-21 15:00:00 | 65.586456 |
+
+## Cross-validation
+
+In previous steps, we’ve taken our historical data to predict the
+future. However, to asses its accuracy we would also like to know how
+the model would have performed in the past. To assess the accuracy and
+robustness of your models on your data perform Cross-Validation.
+
+With time series data, Cross Validation is done by defining a sliding
+window across the historical data and predicting the period following
+it. This form of cross-validation allows us to arrive at a better
+estimation of our model’s predictive abilities across a wider range of
+temporal instances while also keeping the data in the training set
+contiguous as is required by our models.
+
+The following graph depicts such a Cross Validation Strategy:
+
+
+
+### Perform time series cross-validation
+
+Cross-validation of time series models is considered a best practice but
+most implementations are very slow. The statsforecast library implements
+cross-validation as a distributed operation, making the process less
+time-consuming to perform. If you have big datasets you can also perform
+Cross Validation in a distributed cluster using Ray, Dask or Spark.
+
+In this case, we want to evaluate the performance of each model for the
+last 5 months `(n_windows=)`, forecasting every second months
+`(step_size=50)`. Depending on your computer, this step should take
+around 1 min.
+
+The cross_validation method from the StatsForecast class takes the
+following arguments.
+
+- `df:` training data frame
+
+- `h (int):` represents h steps into the future that are being
+ forecasted. In this case, 500 hours ahead.
+
+- `step_size (int):` step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+
+- `n_windows(int):` number of windows used for cross validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+
+```python
+crossvalidation_df = sf.cross_validation(df=df,
+ h=horizon,
+ step_size=50,
+ n_windows=5)
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id:` series identifier
+- `ds:` datestamp or temporal index
+- `cutoff:` the last datestamp or temporal index for the `n_windows`.
+- `y:` true value
+- `model:` columns with the model’s name and fitted value.
+
+```python
+crossvalidation_df
+```
+
+| | unique_id | ds | cutoff | y | TSB |
+|------|-----------|---------------------|---------------------|------|-----------|
+| 0 | 1 | 2023-01-23 12:00:00 | 2023-01-23 11:00:00 | 0.0 | 0.000005 |
+| 1 | 1 | 2023-01-23 13:00:00 | 2023-01-23 11:00:00 | 0.0 | 0.000005 |
+| 2 | 1 | 2023-01-23 14:00:00 | 2023-01-23 11:00:00 | 0.0 | 0.000005 |
+| ... | ... | ... | ... | ... | ... |
+| 2497 | 1 | 2023-02-21 13:00:00 | 2023-01-31 19:00:00 | 60.0 | 65.586456 |
+| 2498 | 1 | 2023-02-21 14:00:00 | 2023-01-31 19:00:00 | 20.0 | 65.586456 |
+| 2499 | 1 | 2023-02-21 15:00:00 | 2023-01-31 19:00:00 | 20.0 | 65.586456 |
+
+## Model Evaluation
+
+Now we are going to evaluate our model with the results of the
+predictions, we will use different types of metrics MAE, MAPE, MASE,
+RMSE, SMAPE to evaluate the accuracy.
+
+```python
+from functools import partial
+
+import utilsforecast.losses as ufl
+from utilsforecast.evaluation import evaluate
+```
+
+
+```python
+evaluate(
+ test.merge(Y_hat),
+ metrics=[ufl.mae, ufl.mape, partial(ufl.mase, seasonality=season_length), ufl.rmse, ufl.smape],
+ train_df=train,
+)
+```
+
+| | unique_id | metric | TSB |
+|-----|-----------|--------|-----------|
+| 0 | 1 | mae | 55.584594 |
+| 1 | 1 | mape | 1.177129 |
+| 2 | 1 | mase | 1.326048 |
+| 3 | 1 | rmse | 60.884468 |
+| 4 | 1 | smape | 0.740778 |
+
+## References
+
+1. [Changquan Huang • Alla Petukhina. Springer series (2022). Applied
+ Time Series Analysis and Forecasting with
+ Python.](https://link.springer.com/book/10.1007/978-3-031-13584-2)
+2. Ivan Svetunkov. [Forecasting and Analytics with the Augmented
+ Dynamic Adaptive Model (ADAM)](https://openforecast.org/adam/)
+3. [James D. Hamilton. Time Series Analysis Princeton University Press,
+ Princeton, New Jersey, 1st Edition,
+ 1994.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+4. [Nixtla TSB API](../../src/core/models.html#tsb)
+5. [Pandas available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+6. [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ Principles and Practice (3rd
+ ed)”](https://otexts.com/fpp3/tscv.html).
+7. [Seasonal periods- Rob J
+ Hyndman](https://robjhyndman.com/hyndsight/seasonal-periods/).
+
diff --git a/statsforecast/docs/tutorials/AnomalyDetection_files/figure-markdown_strict/cell-12-output-1.png b/statsforecast/docs/tutorials/AnomalyDetection_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..e6ebbc31
Binary files /dev/null and b/statsforecast/docs/tutorials/AnomalyDetection_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/statsforecast/docs/tutorials/AnomalyDetection_files/figure-markdown_strict/cell-15-output-1.png b/statsforecast/docs/tutorials/AnomalyDetection_files/figure-markdown_strict/cell-15-output-1.png
new file mode 100644
index 00000000..2406f22b
Binary files /dev/null and b/statsforecast/docs/tutorials/AnomalyDetection_files/figure-markdown_strict/cell-15-output-1.png differ
diff --git a/statsforecast/docs/tutorials/AnomalyDetection_files/figure-markdown_strict/cell-16-output-1.png b/statsforecast/docs/tutorials/AnomalyDetection_files/figure-markdown_strict/cell-16-output-1.png
new file mode 100644
index 00000000..012826b5
Binary files /dev/null and b/statsforecast/docs/tutorials/AnomalyDetection_files/figure-markdown_strict/cell-16-output-1.png differ
diff --git a/statsforecast/docs/tutorials/AnomalyDetection_files/figure-markdown_strict/cell-7-output-1.png b/statsforecast/docs/tutorials/AnomalyDetection_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..78d6cef6
Binary files /dev/null and b/statsforecast/docs/tutorials/AnomalyDetection_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/statsforecast/docs/tutorials/ConformalPrediction_files/figure-markdown_strict/cell-11-output-1.png b/statsforecast/docs/tutorials/ConformalPrediction_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..0ffbe429
Binary files /dev/null and b/statsforecast/docs/tutorials/ConformalPrediction_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/statsforecast/docs/tutorials/ConformalPrediction_files/figure-markdown_strict/cell-12-output-1.png b/statsforecast/docs/tutorials/ConformalPrediction_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..ab7621ec
Binary files /dev/null and b/statsforecast/docs/tutorials/ConformalPrediction_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/statsforecast/docs/tutorials/ConformalPrediction_files/figure-markdown_strict/cell-13-output-1.png b/statsforecast/docs/tutorials/ConformalPrediction_files/figure-markdown_strict/cell-13-output-1.png
new file mode 100644
index 00000000..c953ff84
Binary files /dev/null and b/statsforecast/docs/tutorials/ConformalPrediction_files/figure-markdown_strict/cell-13-output-1.png differ
diff --git a/statsforecast/docs/tutorials/ConformalPrediction_files/figure-markdown_strict/cell-7-output-1.png b/statsforecast/docs/tutorials/ConformalPrediction_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..3130667a
Binary files /dev/null and b/statsforecast/docs/tutorials/ConformalPrediction_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/statsforecast/docs/tutorials/CrossValidation_files/figure-markdown_strict/cell-13-output-1.png b/statsforecast/docs/tutorials/CrossValidation_files/figure-markdown_strict/cell-13-output-1.png
new file mode 100644
index 00000000..2ee09d23
Binary files /dev/null and b/statsforecast/docs/tutorials/CrossValidation_files/figure-markdown_strict/cell-13-output-1.png differ
diff --git a/statsforecast/docs/tutorials/CrossValidation_files/figure-markdown_strict/cell-13-output-2.png b/statsforecast/docs/tutorials/CrossValidation_files/figure-markdown_strict/cell-13-output-2.png
new file mode 100644
index 00000000..f40b0ba4
Binary files /dev/null and b/statsforecast/docs/tutorials/CrossValidation_files/figure-markdown_strict/cell-13-output-2.png differ
diff --git a/statsforecast/docs/tutorials/CrossValidation_files/figure-markdown_strict/cell-13-output-3.png b/statsforecast/docs/tutorials/CrossValidation_files/figure-markdown_strict/cell-13-output-3.png
new file mode 100644
index 00000000..92caf84a
Binary files /dev/null and b/statsforecast/docs/tutorials/CrossValidation_files/figure-markdown_strict/cell-13-output-3.png differ
diff --git a/statsforecast/docs/tutorials/CrossValidation_files/figure-markdown_strict/cell-7-output-1.png b/statsforecast/docs/tutorials/CrossValidation_files/figure-markdown_strict/cell-7-output-1.png
new file mode 100644
index 00000000..8f99eaef
Binary files /dev/null and b/statsforecast/docs/tutorials/CrossValidation_files/figure-markdown_strict/cell-7-output-1.png differ
diff --git a/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-12-output-1.png b/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..2aba5ce4
Binary files /dev/null and b/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-14-output-1.png b/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..ce6076d0
Binary files /dev/null and b/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-21-output-1.png b/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-21-output-1.png
new file mode 100644
index 00000000..8b0a07db
Binary files /dev/null and b/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-21-output-1.png differ
diff --git a/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-22-output-1.png b/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-22-output-1.png
new file mode 100644
index 00000000..08134f1e
Binary files /dev/null and b/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-22-output-1.png differ
diff --git a/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-31-output-1.png b/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-31-output-1.png
new file mode 100644
index 00000000..8b2124c5
Binary files /dev/null and b/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-31-output-1.png differ
diff --git a/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-39-output-1.png b/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-39-output-1.png
new file mode 100644
index 00000000..2619f175
Binary files /dev/null and b/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-39-output-1.png differ
diff --git a/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-6-output-1.png b/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..a83067c8
Binary files /dev/null and b/statsforecast/docs/tutorials/ElectricityLoadForecasting_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/statsforecast/docs/tutorials/ElectricityPeakForecasting_files/figure-markdown_strict/cell-15-output-1.png b/statsforecast/docs/tutorials/ElectricityPeakForecasting_files/figure-markdown_strict/cell-15-output-1.png
new file mode 100644
index 00000000..1e8acddc
Binary files /dev/null and b/statsforecast/docs/tutorials/ElectricityPeakForecasting_files/figure-markdown_strict/cell-15-output-1.png differ
diff --git a/statsforecast/docs/tutorials/ElectricityPeakForecasting_files/figure-markdown_strict/cell-5-output-1.png b/statsforecast/docs/tutorials/ElectricityPeakForecasting_files/figure-markdown_strict/cell-5-output-1.png
new file mode 100644
index 00000000..23fc2038
Binary files /dev/null and b/statsforecast/docs/tutorials/ElectricityPeakForecasting_files/figure-markdown_strict/cell-5-output-1.png differ
diff --git a/statsforecast/docs/tutorials/GARCH_tutorial_files/figure-markdown_strict/cell-11-output-1.png b/statsforecast/docs/tutorials/GARCH_tutorial_files/figure-markdown_strict/cell-11-output-1.png
new file mode 100644
index 00000000..94ce1443
Binary files /dev/null and b/statsforecast/docs/tutorials/GARCH_tutorial_files/figure-markdown_strict/cell-11-output-1.png differ
diff --git a/statsforecast/docs/tutorials/GARCH_tutorial_files/figure-markdown_strict/cell-16-output-1.png b/statsforecast/docs/tutorials/GARCH_tutorial_files/figure-markdown_strict/cell-16-output-1.png
new file mode 100644
index 00000000..924ed8c5
Binary files /dev/null and b/statsforecast/docs/tutorials/GARCH_tutorial_files/figure-markdown_strict/cell-16-output-1.png differ
diff --git a/statsforecast/docs/tutorials/GARCH_tutorial_files/figure-markdown_strict/cell-22-output-1.png b/statsforecast/docs/tutorials/GARCH_tutorial_files/figure-markdown_strict/cell-22-output-1.png
new file mode 100644
index 00000000..b1fe252e
Binary files /dev/null and b/statsforecast/docs/tutorials/GARCH_tutorial_files/figure-markdown_strict/cell-22-output-1.png differ
diff --git a/statsforecast/docs/tutorials/GARCH_tutorial_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/tutorials/GARCH_tutorial_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..485dcd46
Binary files /dev/null and b/statsforecast/docs/tutorials/GARCH_tutorial_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/tutorials/IntermittentData_files/figure-markdown_strict/cell-13-output-1.png b/statsforecast/docs/tutorials/IntermittentData_files/figure-markdown_strict/cell-13-output-1.png
new file mode 100644
index 00000000..e7edaae8
Binary files /dev/null and b/statsforecast/docs/tutorials/IntermittentData_files/figure-markdown_strict/cell-13-output-1.png differ
diff --git a/statsforecast/docs/tutorials/IntermittentData_files/figure-markdown_strict/cell-6-output-1.png b/statsforecast/docs/tutorials/IntermittentData_files/figure-markdown_strict/cell-6-output-1.png
new file mode 100644
index 00000000..8a95f21a
Binary files /dev/null and b/statsforecast/docs/tutorials/IntermittentData_files/figure-markdown_strict/cell-6-output-1.png differ
diff --git a/statsforecast/docs/tutorials/MultipleSeasonalities_files/figure-markdown_strict/cell-12-output-1.png b/statsforecast/docs/tutorials/MultipleSeasonalities_files/figure-markdown_strict/cell-12-output-1.png
new file mode 100644
index 00000000..b25754fb
Binary files /dev/null and b/statsforecast/docs/tutorials/MultipleSeasonalities_files/figure-markdown_strict/cell-12-output-1.png differ
diff --git a/statsforecast/docs/tutorials/MultipleSeasonalities_files/figure-markdown_strict/cell-14-output-1.png b/statsforecast/docs/tutorials/MultipleSeasonalities_files/figure-markdown_strict/cell-14-output-1.png
new file mode 100644
index 00000000..02f2974d
Binary files /dev/null and b/statsforecast/docs/tutorials/MultipleSeasonalities_files/figure-markdown_strict/cell-14-output-1.png differ
diff --git a/statsforecast/docs/tutorials/MultipleSeasonalities_files/figure-markdown_strict/cell-5-output-1.png b/statsforecast/docs/tutorials/MultipleSeasonalities_files/figure-markdown_strict/cell-5-output-1.png
new file mode 100644
index 00000000..978ed12b
Binary files /dev/null and b/statsforecast/docs/tutorials/MultipleSeasonalities_files/figure-markdown_strict/cell-5-output-1.png differ
diff --git a/statsforecast/docs/tutorials/StatisticalNeuralMethods_files/figure-markdown_strict/cell-50-output-1.png b/statsforecast/docs/tutorials/StatisticalNeuralMethods_files/figure-markdown_strict/cell-50-output-1.png
new file mode 100644
index 00000000..4513604a
Binary files /dev/null and b/statsforecast/docs/tutorials/StatisticalNeuralMethods_files/figure-markdown_strict/cell-50-output-1.png differ
diff --git a/statsforecast/docs/tutorials/StatisticalNeuralMethods_files/figure-markdown_strict/cell-51-output-1.png b/statsforecast/docs/tutorials/StatisticalNeuralMethods_files/figure-markdown_strict/cell-51-output-1.png
new file mode 100644
index 00000000..52f2eab5
Binary files /dev/null and b/statsforecast/docs/tutorials/StatisticalNeuralMethods_files/figure-markdown_strict/cell-51-output-1.png differ
diff --git a/statsforecast/docs/tutorials/StatisticalNeuralMethods_files/figure-markdown_strict/cell-52-output-1.png b/statsforecast/docs/tutorials/StatisticalNeuralMethods_files/figure-markdown_strict/cell-52-output-1.png
new file mode 100644
index 00000000..99b826a5
Binary files /dev/null and b/statsforecast/docs/tutorials/StatisticalNeuralMethods_files/figure-markdown_strict/cell-52-output-1.png differ
diff --git a/statsforecast/docs/tutorials/StatisticalNeuralMethods_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/tutorials/StatisticalNeuralMethods_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..1dfe6679
Binary files /dev/null and b/statsforecast/docs/tutorials/StatisticalNeuralMethods_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/tutorials/UncertaintyIntervals_files/figure-markdown_strict/cell-15-output-1.png b/statsforecast/docs/tutorials/UncertaintyIntervals_files/figure-markdown_strict/cell-15-output-1.png
new file mode 100644
index 00000000..55f65f61
Binary files /dev/null and b/statsforecast/docs/tutorials/UncertaintyIntervals_files/figure-markdown_strict/cell-15-output-1.png differ
diff --git a/statsforecast/docs/tutorials/UncertaintyIntervals_files/figure-markdown_strict/cell-16-output-1.png b/statsforecast/docs/tutorials/UncertaintyIntervals_files/figure-markdown_strict/cell-16-output-1.png
new file mode 100644
index 00000000..f7abfc0e
Binary files /dev/null and b/statsforecast/docs/tutorials/UncertaintyIntervals_files/figure-markdown_strict/cell-16-output-1.png differ
diff --git a/statsforecast/docs/tutorials/UncertaintyIntervals_files/figure-markdown_strict/cell-17-output-1.png b/statsforecast/docs/tutorials/UncertaintyIntervals_files/figure-markdown_strict/cell-17-output-1.png
new file mode 100644
index 00000000..f36d1201
Binary files /dev/null and b/statsforecast/docs/tutorials/UncertaintyIntervals_files/figure-markdown_strict/cell-17-output-1.png differ
diff --git a/statsforecast/docs/tutorials/UncertaintyIntervals_files/figure-markdown_strict/cell-18-output-1.png b/statsforecast/docs/tutorials/UncertaintyIntervals_files/figure-markdown_strict/cell-18-output-1.png
new file mode 100644
index 00000000..f2ad0cef
Binary files /dev/null and b/statsforecast/docs/tutorials/UncertaintyIntervals_files/figure-markdown_strict/cell-18-output-1.png differ
diff --git a/statsforecast/docs/tutorials/UncertaintyIntervals_files/figure-markdown_strict/cell-19-output-1.png b/statsforecast/docs/tutorials/UncertaintyIntervals_files/figure-markdown_strict/cell-19-output-1.png
new file mode 100644
index 00000000..81c12e20
Binary files /dev/null and b/statsforecast/docs/tutorials/UncertaintyIntervals_files/figure-markdown_strict/cell-19-output-1.png differ
diff --git a/statsforecast/docs/tutorials/UncertaintyIntervals_files/figure-markdown_strict/cell-9-output-1.png b/statsforecast/docs/tutorials/UncertaintyIntervals_files/figure-markdown_strict/cell-9-output-1.png
new file mode 100644
index 00000000..14f6be8a
Binary files /dev/null and b/statsforecast/docs/tutorials/UncertaintyIntervals_files/figure-markdown_strict/cell-9-output-1.png differ
diff --git a/statsforecast/docs/tutorials/anomalydetection.html.mdx b/statsforecast/docs/tutorials/anomalydetection.html.mdx
new file mode 100644
index 00000000..e235321c
--- /dev/null
+++ b/statsforecast/docs/tutorials/anomalydetection.html.mdx
@@ -0,0 +1,282 @@
+---
+description: In this notebook, we'll implement anomaly detection in time series data
+output-file: anomalydetection.html
+title: Anomaly Detection
+---
+
+
+> **Prerequesites**
+>
+> This tutorial assumes basic familiarity with StatsForecast. For a
+> minimal example visit the [Quick
+> Start](../getting-started/getting_started_short.html)
+
+## Introduction
+
+Anomaly detection is a crucial task in time series forecasting. It
+involves identifying unusual observations that don’t follow the expected
+dataset patterns. Anomalies, also known as outliers, can be caused by a
+variety of factors, such as errors in the data collection process,
+sudden changes in the underlying patterns of the data, or unexpected
+events. They can pose problems for many forecasting models since they
+can distort trends, seasonal patterns, or autocorrelation estimates. As
+a result, anomalies can have a significant impact on the accuracy of the
+forecasts, and for this reason, it is essential to be able to identify
+them. Furthermore, anomaly detection has many applications across
+different industries, such as detecting fraud in financial data,
+monitoring the performance of online services, or identifying usual
+patterns in energy usage.
+
+By the end of this tutorial, you’ll have a good understanding of how to
+detect anomalies in time series data using
+[StatsForecast](https://nixtla.github.io/statsforecast/)’s probabilistic
+models.
+
+**Outline:**
+
+1. Install libraries
+2. Load and explore data
+3. Train model
+4. Recover insample forecasts and identify anomalies
+
+> **Important**
+>
+> Once an anomaly has been identified, we must decide what to do with
+> it. For example, we could remove it or replace it with another value.
+> The correct course of action is context-dependent and beyond this
+> notebook’s scope. Removing an anomaly will likely improve the accuracy
+> of the forecast, but it can also underestimate the amount of
+> randomness in the data.
+
+> **Tip**
+>
+> You can use Colab to run this Notebook interactively
+>
+
+## Install libraries
+
+We assume that you have StatsForecast already installed. If not, check
+this guide for instructions on [how to install
+StatsForecast](../getting-started/installation.html)
+
+Install the necessary packages using `pip install statsforecast`
+
+
+```python
+pip install statsforecast -U
+```
+
+## Load and explore the data
+
+For this example, we’ll use the hourly dataset of the [M4
+Competition](https://www.sciencedirect.com/science/article/pii/S0169207019301128).
+
+
+```python
+import pandas as pd
+```
+
+
+```python
+df_total = pd.read_parquet('https://datasets-nixtla.s3.amazonaws.com/m4-hourly.parquet')
+df_total.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|-----|-------|
+| 0 | H1 | 1 | 605.0 |
+| 1 | H1 | 2 | 586.0 |
+| 2 | H1 | 3 | 586.0 |
+| 3 | H1 | 4 | 559.0 |
+| 4 | H1 | 5 | 511.0 |
+
+The input to StatsForecast is always a data frame in [long
+format](https://www.theanalysisfactor.com/wide-and-long-data/) with
+three columns: `unique_id`, `df` and `y`.
+
+- `unique_id`: (string, int or category) A unique identifier for the
+ series.
+- `ds`: (timestamp or int) A timestamp in format YYYY-MM-DD or
+ YYYY-MM-DD HH:MM:SS or an integer indexing time.
+- `y`: (numeric) The measurement we wish to forecast.
+
+From this dataset, we’ll select the first 8 time series to reduce the
+total execution time. You can select any number you want by changing the
+value of `n_series`.
+
+
+```python
+n_series = 8
+uids = df_total['unique_id'].unique()[:n_series]
+df = df_total.query('unique_id in @uids')
+```
+
+We can plot these series using the `plot_series` function from the
+`utilsforecast` package. This function has multiple parameters, and the
+required ones to generate the plots in this notebook are explained
+below.
+
+- `df`: A pandas dataframe with columns \[unique_id, ds, y\].
+- `forecasts_df`: A pandas dataframe with columns \[unique_id, ds\]
+ and models.
+- `ids`: A list with the ids of the time series we want to plot.
+- `level`: Prediction interval levels to plot.
+- `plot_anomalies`: Whether or not to include the anomalies for each
+ prediction interval.
+
+
+```python
+from statsforecast import StatsForecast
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+plot_series(df)
+```
+
+
+
+## Train model
+
+To generate the forecast, we’ll use the
+[MSTL](../../src/core/models.html#multiple-seasonalities)
+model, which is well-suited for low-frequency data like the one used
+here. We first need to import it from `statsforecast.models` and then we
+need to instantiate it. Since we’re using hourly data, we have two
+seasonal periods: one every 24 hours (hourly) and one every 24\*7 hours
+(daily). Hence, we need to set `season_length = [24, 24*7]`.
+
+
+```python
+from statsforecast.models import MSTL
+```
+
+
+```python
+# Create a list of models and instantiation parameters
+models = [MSTL(season_length = [24, 24*7])]
+```
+
+To instantiate a new StatsForecast object, we need the following
+parameters:
+
+- `models`: The list of models defined in the previous step.
+- `freq`: A string or integer indicating the frequency of the data.
+ See [pandas’ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+- `n_jobs`: An integer that indicates the number of jobs used in
+ parallel processing. Use -1 to select all cores.
+
+
+```python
+sf = StatsForecast(
+ models=models,
+ freq=1,
+ n_jobs=-1,
+)
+```
+
+We’ll now predict the next 48 hours. To do this, we’ll use the
+`forecast` method, which requieres the following arguments:
+
+- `df`: The dataframe with the training data.
+- `h`: The forecasting horizon.
+- `level`: The confidence levels of the prediction intervals.
+- `fitted`: Return insample predictions.
+
+It is important that we select a `level` and set `fitted=True` since
+we’ll need the insample forecasts and their prediction intervals to
+detect the anomalies.
+
+
+```python
+horizon = 48
+levels = [99]
+
+fcst = sf.forecast(df=df, h=48, level=levels, fitted=True)
+fcst.head()
+```
+
+| | unique_id | ds | MSTL | MSTL-lo-99 | MSTL-hi-99 |
+|-----|-----------|-----|------------|------------|------------|
+| 0 | H1 | 749 | 607.607223 | 587.173250 | 628.041196 |
+| 1 | H1 | 750 | 552.364253 | 521.069710 | 583.658796 |
+| 2 | H1 | 751 | 506.785334 | 465.894977 | 547.675691 |
+| 3 | H1 | 752 | 472.906141 | 423.114088 | 522.698195 |
+| 4 | H1 | 753 | 452.240231 | 394.064394 | 510.416067 |
+
+We can plot the forecasts using the `plot_series` function from before.
+
+
+```python
+plot_series(df, fcst)
+```
+
+
+
+## Recover insample forecasts and identify anomalies
+
+In this example, an **anomaly** will be any observation outside the
+prediction interval of the insample forecasts for a given confidence
+level (here we selected 99%). Hence, we first need to recover the
+insample forecasts using the `forecast_fitted_values` method.
+
+
+```python
+insample_forecasts = sf.forecast_fitted_values()
+insample_forecasts.head()
+```
+
+| | unique_id | ds | y | MSTL | MSTL-lo-99 | MSTL-hi-99 |
+|-----|-----------|-----|-------|------------|------------|------------|
+| 0 | H1 | 1 | 605.0 | 605.098607 | 584.678408 | 625.518805 |
+| 1 | H1 | 2 | 586.0 | 588.496673 | 568.076474 | 608.916872 |
+| 2 | H1 | 3 | 586.0 | 585.586856 | 565.166657 | 606.007054 |
+| 3 | H1 | 4 | 559.0 | 554.012377 | 533.592178 | 574.432576 |
+| 4 | H1 | 5 | 511.0 | 510.153508 | 489.733309 | 530.573707 |
+
+We can now find all the observations above or below the 99% prediction
+interval for the insample forecasts.
+
+
+```python
+anomalies = insample_forecasts[~insample_forecasts['y'].between(insample_forecasts['MSTL-lo-99'], insample_forecasts['MSTL-hi-99'])]
+anomalies.head()
+```
+
+| | unique_id | ds | y | MSTL | MSTL-lo-99 | MSTL-hi-99 |
+|-----|-----------|-----|-------|------------|------------|------------|
+| 42 | H1 | 43 | 613.0 | 649.404871 | 628.984672 | 669.825069 |
+| 47 | H1 | 48 | 683.0 | 662.245526 | 641.825328 | 682.665725 |
+| 48 | H1 | 49 | 687.0 | 655.382320 | 634.962122 | 675.802519 |
+| 100 | H1 | 101 | 507.0 | 484.934230 | 464.514031 | 505.354428 |
+| 110 | H1 | 111 | 451.0 | 474.899006 | 454.478808 | 495.319205 |
+
+We can plot the anomalies by setting the `level` and the
+`plot_anomalies` arguments of the `plot_series` function.
+
+
+```python
+plot_series(forecasts_df=insample_forecasts, level=levels, plot_anomalies=True)
+```
+
+
+
+If we want to take a closer look, we can use the `ids` argument to
+select one particular time series, for example, `H10`.
+
+
+```python
+plot_series(forecasts_df=insample_forecasts, level=[99], plot_anomalies=True, ids=['H10'])
+```
+
+
+
+Here we identified the anomalies in the data using the MSTL model, but
+any [probabilistic
+model](https://nixtla.github.io/statsforecast/#models) from
+StatsForecast can be used. We also selected the 99% prediction interval
+of the insample forecasts, but other confidence levels can be used as
+well.
+
diff --git a/statsforecast/docs/tutorials/conformalprediction.html.mdx b/statsforecast/docs/tutorials/conformalprediction.html.mdx
new file mode 100644
index 00000000..c869c61c
--- /dev/null
+++ b/statsforecast/docs/tutorials/conformalprediction.html.mdx
@@ -0,0 +1,334 @@
+---
+description: In this example, we'll implement conformal prediction
+output-file: conformalprediction.html
+title: Conformal Prediction
+---
+
+
+> **Prerequisites**
+>
+> This tutorial assumes basic familiarity with StatsForecast. For a
+> minimal example visit the [Quick
+> Start](../getting-started/getting_started_short.html)
+
+## Introduction
+
+When we generate a forecast, we usually produce a single value known as
+the point forecast. This value, however, doesn’t tell us anything about
+the uncertainty associated with the forecast. To have a measure of this
+uncertainty, we need **prediction intervals**.
+
+A prediction interval is a range of values that the forecast can take
+with a given probability. Hence, a 95% prediction interval should
+contain a range of values that include the actual future value with
+probability 95%. Probabilistic forecasting aims to generate the full
+forecast distribution. Point forecasting, on the other hand, usually
+returns the mean or the median or said distribution. However, in
+real-world scenarios, it is better to forecast not only the most
+probable future outcome, but many alternative outcomes as well.
+
+The problem is that some timeseries models provide forecast
+distributions, but some other ones only provide point forecasts. How can
+we then estimate the uncertainty of predictions?
+
+> **Prediction Intervals**
+>
+> For models that already provide the forecast distribution, check
+> [Prediction Intervals](./uncertaintyintervals.html).
+
+### Conformal Prediction
+
+For a video introduction, see the [PyData Seattle
+presentation](https://www.youtube.com/watch?v=Bj1U-Rrxk48).
+
+Multi-quantile losses and statistical models can provide provide
+prediction intervals, but the problem is that these are uncalibrated,
+meaning that the actual frequency of observations falling within the
+interval does not align with the confidence level associated with it.
+For example, a calibrated 95% prediction interval should contain the
+true value 95% of the time in repeated sampling. An uncalibrated 95%
+prediction interval, on the other hand, might contain the true value
+only 80% of the time, or perhaps 99% of the time. In the first case, the
+interval is too narrow and underestimates the uncertainty, while in the
+second case, it is too wide and overestimates the uncertainty.
+
+Statistical methods also assume normality. Here, we talk about another
+method called conformal prediction that doesn’t require any
+distributional assumptions. More information on the approach can be
+found in [this repo owned by Valery
+Manokhin](https://github.com/valeman/awesome-conformal-prediction).
+
+Conformal prediction intervals use cross-validation on a point
+forecaster model to generate the intervals. This means that no prior
+probabilities are needed, and the output is well-calibrated. No
+additional training is needed, and the model is treated as a black box.
+The approach is compatible with any model.
+
+[Statsforecast](https://github.com/nixtla/statsforecast) now supports
+Conformal Prediction on all available models.
+
+## Install libraries
+
+We assume that you have StatsForecast already installed. If not, check
+this guide for instructions on [how to install
+StatsForecast](../getting-started/installation.html)
+
+Install the necessary packages using `pip install statsforecast`
+
+
+```python
+pip install statsforecast -U
+```
+
+## Load and explore the data
+
+For this example, we’ll use the hourly dataset from the [M4
+Competition](https://www.sciencedirect.com/science/article/pii/S0169207019301128).
+We first need to download the data from a URL and then load it as a
+`pandas` dataframe. Notice that we’ll load the train and the test data
+separately. We’ll also rename the `y` column of the test data as
+`y_test`.
+
+
+```python
+import pandas as pd
+```
+
+
+```python
+train = pd.read_csv('https://auto-arima-results.s3.amazonaws.com/M4-Hourly.csv')
+test = pd.read_csv('https://auto-arima-results.s3.amazonaws.com/M4-Hourly-test.csv').rename(columns={'y': 'y_test'})
+train.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|-----|-------|
+| 0 | H1 | 1 | 605.0 |
+| 1 | H1 | 2 | 586.0 |
+| 2 | H1 | 3 | 586.0 |
+| 3 | H1 | 4 | 559.0 |
+| 4 | H1 | 5 | 511.0 |
+
+Since the goal of this notebook is to generate prediction intervals,
+we’ll only use the first 8 series of the dataset to reduce the total
+computational time.
+
+
+```python
+n_series = 8
+uids = train['unique_id'].unique()[:n_series] # select first n_series of the dataset
+train = train.query('unique_id in @uids')
+test = test.query('unique_id in @uids')
+```
+
+We can plot these series using the `plot_series` function from the
+utilsforecast library. Thisfunctionmethod has multiple parameters, and
+the required ones to generate the plots in this notebook are explained
+below.
+
+- `df`: A `pandas` dataframe with columns \[`unique_id`, `ds`, `y`\].
+- `forecasts_df`: A `pandas` dataframe with columns \[`unique_id`,
+ `ds`\] and models.
+- `plot_random`: bool = `True`. Plots the time series randomly.
+- `models`: List\[str\]. A list with the models we want to plot.
+- `level`: List\[float\]. A list with the prediction intervals we want
+ to plot.
+- `engine`: str = `matplotlib`. It can also be `plotly`. `plotly`
+ generates interactive plots, while `matplotlib` generates static
+ plots.
+
+
+```python
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+plot_series(train, test, plot_random=False)
+```
+
+
+
+## Train models
+
+StatsForecast can train multiple
+[models](https://nixtla.github.io/statsforecast/#models) on different
+time series efficiently. Most of these models can generate a
+probabilistic forecast, which means that they can produce both point
+forecasts and prediction intervals.
+
+For this example, we’ll use
+[SimpleExponentialSmoothing](https://nixtla.github.io/statsforecast/src/core/models.html#simpleexponentialsmoothing)
+and
+[ADIDA](https://nixtla.github.io/statsforecast/src/core/models.html#adida)
+which do not provide a prediction interval natively. Thus, it makes
+sense to use Conformal Prediction to generate the prediction interval.
+
+We’ll also show using it with
+[ARIMA](https://nixtla.github.io/statsforecast/src/core/models.html#arima)
+to provide prediction intervals that don’t assume normality.
+
+To use these models, we first need to import them from
+`statsforecast.models` and then we need to instantiate them.
+
+
+```python
+from statsforecast.models import SeasonalExponentialSmoothing, ADIDA, ARIMA
+from statsforecast.utils import ConformalIntervals
+
+# Create a list of models and instantiation parameters
+intervals = ConformalIntervals(h=24, n_windows=2)
+# P.S. n_windows*h should be less than the count of data elements in your time series sequence.
+# P.S. Also value of n_windows should be atleast 2 or more.
+
+models = [
+ SeasonalExponentialSmoothing(season_length=24, alpha=0.1, prediction_intervals=intervals),
+ ADIDA(prediction_intervals=intervals),
+ ARIMA(order=(24,0,12), prediction_intervals=intervals),
+]
+```
+
+To instantiate a new StatsForecast object, we need the following
+parameters:
+
+- `df`: The dataframe with the training data.
+- `models`: The list of models defined in the previous step.
+- `freq`: A string indicating the frequency of the data. See [pandas’
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+- `n_jobs`: An integer that indicates the number of jobs used in
+ parallel processing. Use -1 to select all cores.
+
+
+```python
+sf = StatsForecast(models=models, freq=1, n_jobs=-1)
+```
+
+Now we’re ready to generate the forecasts and the prediction intervals.
+To do this, we’ll use the `forecast` method, which takes two arguments:
+
+- `h`: An integer that represent the forecasting horizon. In this
+ case, we’ll forecast the next 24 hours.
+- `level`: A list of floats with the confidence levels of the
+ prediction intervals. For example, `level=[95]` means that the range
+ of values should include the actual future value with probability
+ 95%.
+
+
+```python
+levels = [80, 90] # confidence levels of the prediction intervals
+
+forecasts = sf.forecast(df=train, h=24, level=levels)
+forecasts.head()
+```
+
+| | unique_id | ds | SeasonalES | SeasonalES-lo-90 | SeasonalES-lo-80 | SeasonalES-hi-80 | SeasonalES-hi-90 | ADIDA | ADIDA-lo-90 | ADIDA-lo-80 | ADIDA-hi-80 | ADIDA-hi-90 | ARIMA | ARIMA-lo-90 | ARIMA-lo-80 | ARIMA-hi-80 | ARIMA-hi-90 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | H1 | 701 | 624.132703 | 553.097423 | 556.359139 | 691.906266 | 695.167983 | 747.292568 | 599.519220 | 600.030467 | 894.554670 | 895.065916 | 618.078274 | 609.440076 | 610.583304 | 625.573243 | 626.716472 |
+| 1 | H1 | 702 | 555.698193 | 496.653559 | 506.833156 | 604.563231 | 614.742827 | 747.292568 | 491.669220 | 498.330467 | 996.254670 | 1002.915916 | 549.789291 | 510.464070 | 515.232352 | 584.346231 | 589.114513 |
+| 2 | H1 | 703 | 514.403029 | 462.673117 | 464.939840 | 563.866218 | 566.132941 | 747.292568 | 475.105038 | 475.793791 | 1018.791346 | 1019.480099 | 508.099925 | 496.574844 | 496.990264 | 519.209587 | 519.625007 |
+| 3 | H1 | 704 | 482.057899 | 433.030711 | 436.161413 | 527.954385 | 531.085087 | 747.292568 | 440.069220 | 440.130467 | 1054.454670 | 1054.515916 | 486.376622 | 471.141813 | 471.516997 | 501.236246 | 501.611431 |
+| 4 | H1 | 705 | 460.222522 | 414.270186 | 416.959492 | 503.485552 | 506.174858 | 747.292568 | 415.805038 | 416.193791 | 1078.391346 | 1078.780099 | 470.159478 | 445.162316 | 446.808608 | 493.510348 | 495.156640 |
+
+## Plot prediction intervals
+
+Here we’ll plot the different intervals for one timeseries.
+
+The prediction interval with the SeasonalExponentialSmoothing seen
+below. Even if the model generates a point forecast, we are able to get
+a prediction interval. The 80% prediction interval does not cross the
+90% prediction interval, which is a sign that the intervals are
+calibrated.
+
+
+```python
+plot_series(train, forecasts, level=levels, ids=['H105'], models=['SeasonalES'])
+```
+
+
+
+For weaker fitting models, the conformal prediction interval can be
+larger. A better model corresponds to a narrower interval.
+
+
+```python
+plot_series(train, forecasts, level=levels, ids=['H105'], models=['ADIDA'])
+```
+
+
+
+ARIMA is an example of a model that provides a forecast distribution,
+but we can still use conformal prediction to generate the prediction
+interval. As mentioned earlier, this method has the benefit of not
+assuming normality.
+
+
+```python
+plot_series(train, forecasts, level=levels, ids=['H105'], models=['ARIMA'])
+```
+
+
+
+## StatsForecast Object
+
+Alternatively, the prediction interval can be defined on the
+StatsForecast object. This will apply to all models that don’t have the
+`prediction_intervals` defined.
+
+
+```python
+from statsforecast.models import SimpleExponentialSmoothing, ADIDA
+from statsforecast.utils import ConformalIntervals
+from statsforecast import StatsForecast
+
+models = [
+ SimpleExponentialSmoothing(alpha=0.1),
+ ADIDA()
+]
+
+res = StatsForecast(
+ models=models,
+ freq=1,
+).forecast(df=train, h=24, prediction_intervals=ConformalIntervals(h=24, n_windows=2), level=[80])
+res.head()
+```
+
+| | unique_id | ds | SES | SES-lo-80 | SES-hi-80 | ADIDA | ADIDA-lo-80 | ADIDA-hi-80 |
+|----|----|----|----|----|----|----|----|----|
+| 0 | H1 | 701 | 742.669064 | 649.221405 | 836.116722 | 747.292568 | 600.030467 | 894.554670 |
+| 1 | H1 | 702 | 742.669064 | 550.551324 | 934.786804 | 747.292568 | 498.330467 | 996.254670 |
+| 2 | H1 | 703 | 742.669064 | 523.621405 | 961.716722 | 747.292568 | 475.793791 | 1018.791346 |
+| 3 | H1 | 704 | 742.669064 | 488.121405 | 997.216722 | 747.292568 | 440.130467 | 1054.454670 |
+| 4 | H1 | 705 | 742.669064 | 464.021405 | 1021.316722 | 747.292568 | 416.193791 | 1078.391346 |
+
+## Future work
+
+Conformal prediction has become a powerful framework for uncertainty
+quantification, providing well-calibrated prediction intervals without
+making any distributional assumptions. Its use has surged in both
+academia and industry over the past few years. We’ll continue working on
+it, and future tutorials may include:
+
+- Exploring larger datasets
+- Incorporating industry-specific examples
+- Investigating specialized methods like the jackknife+ that are
+ closely related to conformal prediction (for details on the
+ jackknife+ see
+ [here](https://valeman.medium.com/jackknife-a-swiss-knife-of-conformal-prediction-for-regression-ce3b56432f4f)).
+
+If you’re interested in any of these, or in any other related topic,
+please let us know by opening an issue on
+[GitHub](https://github.com/Nixtla/statsforecast/issues)
+
+## Acknowledgements
+
+We would like to thank [Kevin Kho](https://github.com/kvnkho) for
+writing this tutorial, and Valeriy
+[Manokhin](https://github.com/valeman) for his expertise on conformal
+prediction, as well as for promoting this work.
+
+## References
+
+[Manokhin, Valery. (2022). Machine Learning for Probabilistic
+Prediction. 10.5281/zenodo.6727505.](https://zenodo.org/record/6727505)
+
diff --git a/statsforecast/docs/tutorials/crossvalidation.html.mdx b/statsforecast/docs/tutorials/crossvalidation.html.mdx
new file mode 100644
index 00000000..0a23c1a6
--- /dev/null
+++ b/statsforecast/docs/tutorials/crossvalidation.html.mdx
@@ -0,0 +1,287 @@
+---
+description: >-
+ In this example, we'll implement time series cross-validation to evaluate
+ model's performance.
+output-file: crossvalidation.html
+title: Cross validation
+---
+
+
+> **Prerequesites**
+>
+> This tutorial assumes basic familiarity with StatsForecast. For a
+> minimal example visit the [Quick
+> Start](../getting-started/getting_started_short.html)
+
+## Introduction
+
+Time series cross-validation is a method for evaluating how a model
+would have performed in the past. It works by defining a sliding window
+across the historical data and predicting the period following it.
+
+
+
+[Statsforecast](https://nixtla.github.io/statsforecast/) has an
+implementation of time series cross-validation that is fast and easy to
+use. This implementation makes cross-validation a distributed operation,
+which makes it less time-consuming. In this notebook, we’ll use it on a
+subset of the [M4
+Competition](https://www.sciencedirect.com/science/article/pii/S0169207019301128)
+hourly dataset.
+
+**Outline:**
+
+1. Install libraries
+2. Load and explore data
+3. Train model
+4. Perform time series cross-validation
+5. Evaluate results
+
+> **Tip**
+>
+> You can use Colab to run this Notebook interactively
+>
+
+## Install libraries
+
+We assume that you have StatsForecast already installed. If not, check
+this guide for instructions on [how to install
+StatsForecast](https://nixtla.github.io/statsforecast/docs/getting-started/installation.html)
+
+Install the necessary packages with `pip install statsforecast`
+
+
+```python
+pip install statsforecast
+```
+
+
+```python
+from statsforecast import StatsForecast # required to instantiate StastForecast object and use cross-validation method
+```
+
+## Load and explore the data
+
+As stated in the introduction, we’ll use the M4 Competition hourly
+dataset. We’ll first import the data from an URL using `pandas`.
+
+
+```python
+import pandas as pd
+```
+
+
+```python
+Y_df = pd.read_parquet('https://datasets-nixtla.s3.amazonaws.com/m4-hourly.parquet') # load the data
+Y_df.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|-----|-------|
+| 0 | H1 | 1 | 605.0 |
+| 1 | H1 | 2 | 586.0 |
+| 2 | H1 | 3 | 586.0 |
+| 3 | H1 | 4 | 559.0 |
+| 4 | H1 | 5 | 511.0 |
+
+The input to
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+is a data frame in [long
+format](https://www.theanalysisfactor.com/wide-and-long-data/) with
+three columns: `unique_id`, `ds` and y:
+
+- The `unique_id` (string, int, or category) represents an identifier
+ for the series.
+- The `ds` (datestamp or int) column should be either an integer
+ indexing time or a datestamp in format YYYY-MM-DD or YYYY-MM-DD
+ HH:MM:SS.
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+The data in this example already has this format, so no changes are
+needed.
+
+To keep the time required to execute this notebook to a minimum, we’ll
+only use one time series from the data, namely the one with
+`unique_id == 'H1'`. However, you can use as many as you want, with no
+additional changes to the code needed.
+
+
+```python
+df = Y_df[Y_df['unique_id'] == 'H1'] # select time series
+```
+
+We can plot the time series we’ll work with using `StatsForecast.plot`
+method.
+
+
+```python
+StatsForecast.plot(df)
+```
+
+
+
+## Train model
+
+For this example, we’ll use StastForecast
+[AutoETS](https://Nixtla.github.io/statsforecast/src/core/models.html#autoets).
+We first need to import it from `statsforecast.models` and then we need
+to instantiate a new
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+object.
+
+The
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+object has the following parameters:
+
+- models: a list of models. Select the models you want from
+ [models](https://Nixtla.github.io/statsforecast/src/core/models.html) and
+ import them.
+- freq: a string indicating the frequency of the data. See [panda’s
+ available
+ frequencies.](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases)
+- n_jobs: n_jobs: int, number of jobs used in the parallel processing,
+ use -1 for all cores.
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame `df`.
+
+
+```python
+from statsforecast.models import AutoETS
+```
+
+
+```python
+models = [AutoETS(season_length = 24)]
+
+sf = StatsForecast(
+ models = models,
+ freq = 1,
+ n_jobs = 1
+)
+```
+
+## Perform time series cross-validation
+
+Once the
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)object
+has been instantiated, we can use the
+[`cross_validation`](https://Nixtla.github.io/statsforecast/src/mfles.html#cross_validation)
+method, which takes the following arguments:
+
+- `df`: training data frame with
+ [`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+ format
+- `h` (int): represents the h steps into the future that will be
+ forecasted
+- `step_size` (int): step size between each window, meaning how often
+ do you want to run the forecasting process.
+- `n_windows` (int): number of windows used for cross-validation,
+ meaning the number of forecasting processes in the past you want to
+ evaluate.
+
+For this particular example, we’ll use 3 windows of 24 hours.
+
+
+```python
+cv_df = sf.cross_validation(
+ df = df,
+ h = 24,
+ step_size = 24,
+ n_windows = 3
+ )
+```
+
+The `cv_df` object is a new data frame that includes the following
+columns:
+
+- `unique_id`: series identifier
+- `ds`: datestamp or temporal index
+- `cutoff`: the last datestamp or temporal index for the n_windows.
+- `y`: true value
+- `"model"`: columns with the model’s name and fitted value.
+
+
+```python
+cv_df.head()
+```
+
+| | unique_id | ds | cutoff | y | AutoETS |
+|-----|-----------|-----|--------|-------|------------|
+| 0 | H1 | 677 | 676 | 691.0 | 677.761053 |
+| 1 | H1 | 678 | 676 | 618.0 | 607.817879 |
+| 2 | H1 | 679 | 676 | 563.0 | 569.437729 |
+| 3 | H1 | 680 | 676 | 529.0 | 537.340007 |
+| 4 | H1 | 681 | 676 | 504.0 | 515.571123 |
+
+We’ll now plot the forecast for each cutoff period. To make the plots
+clearer, we’ll rename the actual values in each period.
+
+
+```python
+from IPython.display import display
+```
+
+
+```python
+cv_df.rename(columns = {'y' : 'actual'}, inplace = True) # rename actual values
+
+cutoff = cv_df['cutoff'].unique()
+
+for k in range(len(cutoff)):
+ cv = cv_df[cv_df['cutoff'] == cutoff[k]]
+ display(StatsForecast.plot(df, cv.loc[:, cv.columns != 'cutoff']))
+```
+
+
+
+
+
+
+
+Notice that in each cutoff period, we generated a forecast for the next
+24 hours using only the data `y` before said period.
+
+## Evaluate results
+
+We can now compute the accuracy of the forecast using an appropiate
+accuracy metric. Here we’ll use the [Root Mean Squared Error
+(RMSE).](https://en.wikipedia.org/wiki/Root-mean-square_deviation).
+
+
+```python
+from utilsforecast.losses import rmse
+```
+
+The function to compute the RMSE takes two arguments:
+
+1. The actual values.
+2. The forecasts, in this case,
+ [`AutoETS`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoets).
+
+
+```python
+cv_rmse = rmse(cv_df, models=['AutoETS'], target_col='actual')['AutoETS'].item()
+print(f"RMSE using cross-validation: {cv_rmse:.2f}")
+```
+
+``` text
+RMSE using cross-validation: 33.90
+```
+
+This measure should better reflect the predictive abilities of our
+model, since it used different time periods to test its accuracy.
+
+> **Tip**
+>
+> Cross validation is especially useful when comparing multiple models.
+> Here’s an
+> [example](../getting-started/getting_started_complete.html)
+> with multiple models and time series.
+
+## References
+
+[Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+principles and practice, Time series
+cross-validation”](https://otexts.com/fpp3/tscv.html).
+
diff --git a/statsforecast/docs/tutorials/electricityloadforecasting.html.mdx b/statsforecast/docs/tutorials/electricityloadforecasting.html.mdx
new file mode 100644
index 00000000..65cd4101
--- /dev/null
+++ b/statsforecast/docs/tutorials/electricityloadforecasting.html.mdx
@@ -0,0 +1,713 @@
+---
+description: >-
+ In this example we will show how to perform electricity load forecasting
+ considering a model capable of handling multiple seasonalities (MSTL).
+output-file: electricityloadforecasting.html
+title: Electricity Load Forecast
+---
+
+
+
+
+## Introduction
+
+Some time series are generated from very low frequency data. These data
+generally exhibit multiple seasonalities. For example, hourly data may
+exhibit repeated patterns every hour (every 24 observations) or every
+day (every 24 \* 7, hours per day, observations). This is the case for
+electricity load. Electricity load may vary hourly, e.g., during the
+evenings electricity consumption may be expected to increase. But also,
+the electricity load varies by week. Perhaps on weekends there is an
+increase in electrical activity.
+
+In this example we will show how to model the two seasonalities of the
+time series to generate accurate forecasts in a short time. We will use
+hourly PJM electricity load data. The original data can be found
+[here](https://github.com/panambY/Hourly_Energy_Consumption).
+
+## Libraries
+
+In this example we will use the following libraries:
+
+- [`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast).
+ Lightning ⚡️ fast forecasting with statistical and econometric
+ models. Includes the MSTL model for multiple seasonalities.
+- [`Prophet`](https://github.com/facebook/prophet). Benchmark model
+ developed by Facebook.
+- [`NeuralProphet`](https://github.com/ourownstory/neural_prophet).
+ Deep Learning version of `Prophet`. Used as benchark.
+
+
+```python
+# !pip install statsforecast "neuralprophet[live]" prophet
+```
+
+## Forecast using Multiple Seasonalities
+
+### Electricity Load Data
+
+According to the [dataset’s
+page](https://www.kaggle.com/datasets/robikscube/hourly-energy-consumption),
+
+> PJM Interconnection LLC (PJM) is a regional transmission organization
+> (RTO) in the United States. It is part of the Eastern Interconnection
+> grid operating an electric transmission system serving all or parts of
+> Delaware, Illinois, Indiana, Kentucky, Maryland, Michigan, New Jersey,
+> North Carolina, Ohio, Pennsylvania, Tennessee, Virginia, West
+> Virginia, and the District of Columbia. The hourly power consumption
+> data comes from PJM’s website and are in megawatts (MW).
+
+Let’s take a look to the data.
+
+
+```python
+import matplotlib.pyplot as plt
+import numpy as np
+import pandas as pd
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+pd.plotting.register_matplotlib_converters()
+plt.rc("figure", figsize=(10, 8))
+plt.rc("font", size=10)
+```
+
+
+```python
+df = pd.read_csv('https://raw.githubusercontent.com/panambY/Hourly_Energy_Consumption/master/data/PJM_Load_hourly.csv')
+df.columns = ['ds', 'y']
+df.insert(0, 'unique_id', 'PJM_Load_hourly')
+df['ds'] = pd.to_datetime(df['ds'])
+df = df.sort_values(['unique_id', 'ds']).reset_index(drop=True)
+df.tail()
+```
+
+| | unique_id | ds | y |
+|-------|-----------------|---------------------|---------|
+| 32891 | PJM_Load_hourly | 2001-12-31 20:00:00 | 36392.0 |
+| 32892 | PJM_Load_hourly | 2001-12-31 21:00:00 | 35082.0 |
+| 32893 | PJM_Load_hourly | 2001-12-31 22:00:00 | 33890.0 |
+| 32894 | PJM_Load_hourly | 2001-12-31 23:00:00 | 32590.0 |
+| 32895 | PJM_Load_hourly | 2002-01-01 00:00:00 | 31569.0 |
+
+
+```python
+plot_series(df)
+```
+
+
+
+We clearly observe that the time series exhibits seasonal patterns.
+Moreover, the time series contains `32,896` observations, so it is
+necessary to use very computationally efficient methods to display them
+in production.
+
+### MSTL model
+
+The
+[`MSTL`](https://Nixtla.github.io/statsforecast/src/core/models.html#mstl)
+(Multiple Seasonal-Trend decomposition using LOESS) model, originally
+developed by [Kasun Bandara, Rob J Hyndman and Christoph
+Bergmeir](https://arxiv.org/abs/2107.13462), decomposes the time series
+in multiple seasonalities using a Local Polynomial Regression (LOESS).
+Then it forecasts the trend using a custom non-seasonal model and each
+seasonality using a
+[`SeasonalNaive`](https://Nixtla.github.io/statsforecast/src/core/models.html#seasonalnaive)
+model.
+
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+contains a fast implementation of the
+[`MSTL`](https://Nixtla.github.io/statsforecast/src/core/models.html#mstl)
+model. Also, the decomposition of the time series can be calculated.
+
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import MSTL, AutoARIMA, SeasonalNaive
+from statsforecast.utils import AirPassengers as ap
+```
+
+First we must define the model parameters. As mentioned before, the
+electricity load presents seasonalities every 24 hours (Hourly) and
+every 24 \* 7 (Daily) hours. Therefore, we will use `[24, 24 * 7]` as
+the seasonalities that the MSTL model receives. We must also specify the
+manner in which the trend will be forecasted. In this case we will use
+the
+[`AutoARIMA`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoarima)
+model.
+
+
+```python
+mstl = MSTL(
+ season_length=[24, 24 * 7], # seasonalities of the time series
+ trend_forecaster=AutoARIMA() # model used to forecast trend
+)
+```
+
+Once the model is instantiated, we have to instantiate the
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+class to create forecasts.
+
+
+```python
+sf = StatsForecast(
+ models=[mstl], # model used to fit each time series
+ freq='h', # frequency of the data
+)
+```
+
+#### Fit the model
+
+Afer that, we just have to use the `fit` method to fit each model to
+each time series.
+
+
+```python
+sf = sf.fit(df=df)
+```
+
+#### Decompose the time series in multiple seasonalities
+
+Once the model is fitted, we can access the decomposition using the
+`fitted_` attribute of
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast).
+This attribute stores all relevant information of the fitted models for
+each of the time series.
+
+In this case we are fitting a single model for a single time series, so
+by accessing the fitted\_ location \[0, 0\] we will find the relevant
+information of our model. The
+[`MSTL`](https://Nixtla.github.io/statsforecast/src/core/models.html#mstl)
+class generates a `model_` attribute that contains the way the series
+was decomposed.
+
+
+```python
+sf.fitted_[0, 0].model_
+```
+
+| | data | trend | seasonal24 | seasonal168 | remainder |
+|-------|---------|--------------|--------------|-------------|--------------|
+| 0 | 22259.0 | 25899.808157 | -4720.213546 | 581.308595 | 498.096794 |
+| 1 | 21244.0 | 25900.349395 | -5433.168901 | 571.780657 | 205.038849 |
+| 2 | 20651.0 | 25900.875973 | -5829.135728 | 557.142643 | 22.117112 |
+| 3 | 20421.0 | 25901.387631 | -5704.092794 | 597.696957 | -373.991794 |
+| 4 | 20713.0 | 25901.884103 | -5023.324375 | 922.564854 | -1088.124582 |
+| ... | ... | ... | ... | ... | ... |
+| 32891 | 36392.0 | 33329.031577 | 4254.112720 | 917.258336 | -2108.402633 |
+| 32892 | 35082.0 | 33355.083576 | 3625.077164 | 721.689136 | -2619.849876 |
+| 32893 | 33890.0 | 33381.108409 | 2571.794472 | 549.661529 | -2612.564409 |
+| 32894 | 32590.0 | 33407.105839 | 796.356548 | 361.956280 | -1975.418667 |
+| 32895 | 31569.0 | 33433.075723 | -1260.860917 | 279.777069 | -882.991876 |
+
+Let’s look graphically at the different components of the time series.
+
+
+```python
+sf.fitted_[0, 0].model_.tail(24 * 28).plot(subplots=True, grid=True)
+plt.tight_layout()
+plt.show()
+```
+
+
+
+We observe that there is a clear trend towards the high (orange line).
+This component would be predicted with the
+[`AutoARIMA`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoarima)
+model. We can also observe that every 24 hours and every `24 * 7` hours
+there is a very well defined pattern. These two components will be
+forecast separately using a
+[`SeasonalNaive`](https://Nixtla.github.io/statsforecast/src/core/models.html#seasonalnaive)
+model.
+
+#### Produce forecasts
+
+To generate forecasts we only have to use the `predict` method
+specifying the forecast horizon (`h`). In addition, to calculate
+prediction intervals associated to the forecasts, we can include the
+parameter `level` that receives a list of levels of the prediction
+intervals we want to build. In this case we will only calculate the 90%
+forecast interval (`level=[90]`).
+
+
+```python
+forecasts = sf.predict(h=24, level=[90])
+forecasts.head()
+```
+
+| | unique_id | ds | MSTL | MSTL-lo-90 | MSTL-hi-90 |
+|----|----|----|----|----|----|
+| 0 | PJM_Load_hourly | 2002-01-01 01:00:00 | 30215.608163 | 29842.185622 | 30589.030705 |
+| 1 | PJM_Load_hourly | 2002-01-01 02:00:00 | 29447.209028 | 28787.123369 | 30107.294687 |
+| 2 | PJM_Load_hourly | 2002-01-01 03:00:00 | 29132.787603 | 28221.354454 | 30044.220751 |
+| 3 | PJM_Load_hourly | 2002-01-01 04:00:00 | 29126.254591 | 27992.821420 | 30259.687762 |
+| 4 | PJM_Load_hourly | 2002-01-01 05:00:00 | 29604.608674 | 28273.428663 | 30935.788686 |
+
+Let’s look at our forecasts graphically.
+
+
+```python
+plot_series(df, forecasts, level=[90], max_insample_length=24*7)
+```
+
+
+
+In the next section we will plot different models so it is convenient to
+reuse the previous code with the following function.
+
+
+```python
+def plot_forecasts(y_hist, y_true, y_pred, models):
+ _, ax = plt.subplots(1, 1, figsize = (20, 7))
+ y_true = y_true.merge(y_pred, how='left', on=['unique_id', 'ds'])
+ df_plot = pd.concat([y_hist, y_true]).set_index('ds').tail(24 * 7)
+ df_plot[['y'] + models].plot(ax=ax, linewidth=2)
+ colors = ['orange', 'green', 'red']
+ for model, color in zip(models, colors):
+ ax.fill_between(df_plot.index,
+ df_plot[f'{model}-lo-90'],
+ df_plot[f'{model}-hi-90'],
+ alpha=.35,
+ color=color,
+ label=f'{model}-level-90')
+ ax.set_title('PJM Load Hourly', fontsize=22)
+ ax.set_ylabel('Electricity Load', fontsize=20)
+ ax.set_xlabel('Timestamp [t]', fontsize=20)
+ ax.legend(prop={'size': 15})
+ ax.grid()
+```
+
+### Performance of the MSTL model
+
+#### Split Train/Test sets
+
+To validate the accuracy of the
+[`MSTL`](https://Nixtla.github.io/statsforecast/src/core/models.html#mstl)
+model, we will show its performance on unseen data. We will use a
+classical time series technique that consists of dividing the data into
+a training set and a test set. We will leave the last 24 observations
+(the last day) as the test set. So the model will train on `32,872`
+observations.
+
+
+```python
+df_test = df.tail(24)
+df_train = df.drop(df_test.index)
+```
+
+#### MSTL model
+
+In addition to the
+[`MSTL`](https://Nixtla.github.io/statsforecast/src/core/models.html#mstl)
+model, we will include the
+[`SeasonalNaive`](https://Nixtla.github.io/statsforecast/src/core/models.html#seasonalnaive)
+model as a benchmark to validate the added value of the
+[`MSTL`](https://Nixtla.github.io/statsforecast/src/core/models.html#mstl)
+model. Including
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+models is as simple as adding them to the list of models to be fitted.
+
+
+```python
+sf = StatsForecast(
+ models=[mstl, SeasonalNaive(season_length=24)], # add SeasonalNaive model to the list
+ freq='h'
+)
+```
+
+To measure the fitting time we will use the `time` module.
+
+
+```python
+from time import time
+```
+
+To retrieve the forecasts of the test set we only have to do fit and
+predict as before.
+
+
+```python
+init = time()
+sf = sf.fit(df=df_train)
+forecasts_test = sf.predict(h=len(df_test), level=[90])
+end = time()
+forecasts_test.head()
+```
+
+| | unique_id | ds | MSTL | MSTL-lo-90 | MSTL-hi-90 | SeasonalNaive | SeasonalNaive-lo-90 | SeasonalNaive-hi-90 |
+|----|----|----|----|----|----|----|----|----|
+| 0 | PJM_Load_hourly | 2001-12-31 01:00:00 | 29158.872180 | 28785.567875 | 29532.176486 | 28326.0 | 23468.555872 | 33183.444128 |
+| 1 | PJM_Load_hourly | 2001-12-31 02:00:00 | 28233.452263 | 27573.789089 | 28893.115438 | 27362.0 | 22504.555872 | 32219.444128 |
+| 2 | PJM_Load_hourly | 2001-12-31 03:00:00 | 27915.251368 | 27004.459000 | 28826.043736 | 27108.0 | 22250.555872 | 31965.444128 |
+| 3 | PJM_Load_hourly | 2001-12-31 04:00:00 | 27969.396560 | 26836.674164 | 29102.118956 | 26865.0 | 22007.555872 | 31722.444128 |
+| 4 | PJM_Load_hourly | 2001-12-31 05:00:00 | 28469.805588 | 27139.306401 | 29800.304775 | 26808.0 | 21950.555872 | 31665.444128 |
+
+
+```python
+time_mstl = (end - init) / 60
+print(f'MSTL Time: {time_mstl:.2f} minutes')
+```
+
+``` text
+MSTL Time: 0.46 minutes
+```
+
+Then we were able to generate forecasts for the next 24 hours. Now let’s
+look at the graphical comparison of the forecasts with the actual
+values.
+
+
+```python
+plot_series(df_train, df_test.merge(forecasts_test), level=[90], max_insample_length=24*7)
+```
+
+
+
+Let’s look at those produced only by
+[`MSTL`](https://Nixtla.github.io/statsforecast/src/core/models.html#mstl).
+
+
+```python
+plot_series(df_train, df_test.merge(forecasts_test), level=[90], max_insample_length=24*7, models=['MSTL'])
+```
+
+
+
+We note that
+[`MSTL`](https://Nixtla.github.io/statsforecast/src/core/models.html#mstl)
+produces very accurate forecasts that follow the behavior of the time
+series. Now let us calculate numerically the accuracy of the model. We
+will use the following metrics: `MAE`, `MAPE`, `MASE`, `RMSE`, `SMAPE`.
+
+
+```python
+from functools import partial
+
+from utilsforecast.evaluation import evaluate
+from utilsforecast.losses import mae, mape, mase, rmse, smape
+```
+
+
+```python
+eval_df = evaluate(
+ df=df_test.merge(forecasts_test),
+ train_df=df_train,
+ metrics=[partial(mase, seasonality=24), mae, mape, rmse, smape],
+ agg_fn='mean',
+).set_index('metric').T
+eval_df
+```
+
+| metric | mase | mae | mape | rmse | smape |
+|---------------|----------|-------------|----------|-------------|----------|
+| MSTL | 0.587265 | 1219.321795 | 0.036052 | 1460.223279 | 0.017577 |
+| SeasonalNaive | 0.894653 | 1857.541667 | 0.056482 | 2201.384101 | 0.029343 |
+
+
+```python
+1 - eval_df.loc['MSTL', 'mase'] / eval_df.loc['SeasonalNaive', 'mase']
+```
+
+``` text
+0.3435830717111049
+```
+
+We observe that
+[`MSTL`](https://Nixtla.github.io/statsforecast/src/core/models.html#mstl)
+has an improvement of about 35% over the
+[`SeasonalNaive`](https://Nixtla.github.io/statsforecast/src/core/models.html#seasonalnaive)
+method in the test set measured in `MASE`.
+
+#### Comparison with Prophet
+
+One of the most widely used models for time series forecasting is
+`Prophet`. This model is known for its ability to model different
+seasonalities (weekly, daily yearly). We will use this model as a
+benchmark to see if the
+[`MSTL`](https://Nixtla.github.io/statsforecast/src/core/models.html#mstl)
+adds value for this time series.
+
+
+```python
+from prophet import Prophet
+```
+
+
+```python
+# create prophet model
+prophet = Prophet(interval_width=0.9)
+init = time()
+prophet.fit(df_train)
+# produce forecasts
+future = prophet.make_future_dataframe(periods=len(df_test), freq='H', include_history=False)
+forecast_prophet = prophet.predict(future)
+end = time()
+# data wrangling
+forecast_prophet = forecast_prophet[['ds', 'yhat', 'yhat_lower', 'yhat_upper']]
+forecast_prophet.columns = ['ds', 'Prophet', 'Prophet-lo-90', 'Prophet-hi-90']
+forecast_prophet.insert(0, 'unique_id', 'PJM_Load_hourly')
+forecast_prophet.head()
+```
+
+``` text
+16:56:47 - cmdstanpy - INFO - Chain [1] start processing
+16:57:09 - cmdstanpy - INFO - Chain [1] done processing
+```
+
+| | unique_id | ds | Prophet | Prophet-lo-90 | Prophet-hi-90 |
+|----|----|----|----|----|----|
+| 0 | PJM_Load_hourly | 2001-12-31 01:00:00 | 25294.246960 | 20299.105766 | 30100.467618 |
+| 1 | PJM_Load_hourly | 2001-12-31 02:00:00 | 24000.725423 | 19285.395144 | 28777.495372 |
+| 2 | PJM_Load_hourly | 2001-12-31 03:00:00 | 23324.771966 | 18536.736306 | 28057.063589 |
+| 3 | PJM_Load_hourly | 2001-12-31 04:00:00 | 23332.519871 | 18591.879190 | 28720.461289 |
+| 4 | PJM_Load_hourly | 2001-12-31 05:00:00 | 24107.126827 | 18934.471254 | 29116.352931 |
+
+
+```python
+time_prophet = (end - init) / 60
+print(f'Prophet Time: {time_prophet:.2f} minutes')
+```
+
+``` text
+Prophet Time: 0.41 minutes
+```
+
+
+```python
+times = pd.DataFrame({'model': ['MSTL', 'Prophet'], 'time (mins)': [time_mstl, time_prophet]})
+times
+```
+
+| | model | time (mins) |
+|-----|---------|-------------|
+| 0 | MSTL | 0.455999 |
+| 1 | Prophet | 0.408726 |
+
+We observe that the time required for `Prophet` to perform the fit and
+predict pipeline is greater than
+[`MSTL`](https://Nixtla.github.io/statsforecast/src/core/models.html#mstl).
+Let’s look at the forecasts produced by `Prophet`.
+
+
+```python
+forecasts_test = forecasts_test.merge(forecast_prophet, how='left', on=['unique_id', 'ds'])
+```
+
+
+```python
+plot_series(df_train, forecasts_test, max_insample_length=24*7, level=[90])
+```
+
+
+
+We note that `Prophet` is able to capture the overall behavior of the
+time series. However, in some cases it produces forecasts well below the
+actual value. It also does not correctly adjust the valleys.
+
+
+```python
+eval_df = evaluate(
+ df=df_test.merge(forecasts_test),
+ train_df=df_train,
+ metrics=[partial(mase, seasonality=24), mae, mape, rmse, smape],
+ agg_fn='mean',
+).set_index('metric').T
+eval_df
+```
+
+| metric | mase | mae | mape | rmse | smape |
+|---------------|----------|-------------|----------|-------------|----------|
+| MSTL | 0.587265 | 1219.321795 | 0.036052 | 1460.223279 | 0.017577 |
+| SeasonalNaive | 0.894653 | 1857.541667 | 0.056482 | 2201.384101 | 0.029343 |
+| Prophet | 1.099551 | 2282.966977 | 0.073750 | 2721.817203 | 0.038633 |
+
+
+```python
+1 - eval_df.loc['MSTL', 'mase'] / eval_df.loc['Prophet', 'mase']
+```
+
+``` text
+0.4659047602697266
+```
+
+In terms of accuracy, `Prophet` is not able to produce better forecasts
+than the
+[`SeasonalNaive`](https://Nixtla.github.io/statsforecast/src/core/models.html#seasonalnaive)
+model, however, the
+[`MSTL`](https://Nixtla.github.io/statsforecast/src/core/models.html#mstl)
+model improves `Prophet`’s forecasts by 45% (`MASE`).
+
+#### Comparison with NeuralProphet
+
+`NeuralProphet` is the version of `Prophet` using deep learning. This
+model is also capable of handling different seasonalities so we will
+also use it as a benchmark.
+
+
+```python
+from neuralprophet import NeuralProphet
+```
+
+
+```python
+neuralprophet = NeuralProphet(quantiles=[0.05, 0.95])
+init = time()
+neuralprophet.fit(df_train.drop(columns='unique_id'))
+future = neuralprophet.make_future_dataframe(df=df_train.drop(columns='unique_id'), periods=len(df_test))
+forecast_np = neuralprophet.predict(future)
+end = time()
+forecast_np = forecast_np[['ds', 'yhat1', 'yhat1 5.0%', 'yhat1 95.0%']]
+forecast_np.columns = ['ds', 'NeuralProphet', 'NeuralProphet-lo-90', 'NeuralProphet-hi-90']
+forecast_np.insert(0, 'unique_id', 'PJM_Load_hourly')
+forecast_np.head()
+```
+
+``` text
+WARNING - (NP.forecaster.fit) - When Global modeling with local normalization, metrics are displayed in normalized scale.
+INFO - (NP.df_utils._infer_frequency) - Major frequency h corresponds to 99.973% of the data.
+INFO - (NP.df_utils._infer_frequency) - Dataframe freq automatically defined as h
+INFO - (NP.config.init_data_params) - Setting normalization to global as only one dataframe provided for training.
+INFO - (NP.config.set_auto_batch_epoch) - Auto-set batch_size to 128
+INFO - (NP.config.set_auto_batch_epoch) - Auto-set epochs to 40
+```
+
+``` text
+Training: | …
+```
+
+``` text
+WARNING - (NP.config.set_lr_finder_args) - Learning rate finder: The number of batches (257) is too small than the required number for the learning rate finder (262). The results might not be optimal.
+```
+
+``` text
+Finding best initial lr: 0%| | 0/262 [00:00 **Important**
+>
+> The performance of `NeuralProphet` can be improved using
+> hyperparameter optimization, which can increase the fitting time
+> significantly. In this example we show its performance with the
+> default version.
+
+## Conclusion
+
+In this post we introduced
+[`MSTL`](https://Nixtla.github.io/statsforecast/src/core/models.html#mstl),
+a model originally developed by [Kasun Bandara, Rob Hyndman and
+Christoph Bergmeir](https://arxiv.org/abs/2107.13462) capable of
+handling time series with multiple seasonalities. We also showed that
+for the PJM electricity load time series offers better performance in
+time and accuracy than the `Prophet` and `NeuralProphet` models.
+
+## References
+
+- [Bandara, Kasun & Hyndman, Rob & Bergmeir, Christoph. (2021). “MSTL:
+ A Seasonal-Trend Decomposition Algorithm for Time Series with
+ Multiple Seasonal Patterns”](https://arxiv.org/abs/2107.13462).
+
diff --git a/statsforecast/docs/tutorials/electricitypeakforecasting.html.mdx b/statsforecast/docs/tutorials/electricitypeakforecasting.html.mdx
new file mode 100644
index 00000000..f4abd7b8
--- /dev/null
+++ b/statsforecast/docs/tutorials/electricitypeakforecasting.html.mdx
@@ -0,0 +1,331 @@
+---
+description: >-
+ In this example we will show how to perform electricity load forecasting on
+ the ERCOT (Texas) market for detecting daily peaks.
+output-file: electricitypeakforecasting.html
+title: Detect Demand Peaks
+---
+
+
+## Introduction
+
+Predicting peaks in different markets is useful. In the electricity
+market, consuming electricity at peak demand is penalized with higher
+tarifs. When an individual or company consumes electricity when its most
+demanded, regulators calls that a coincident peak (CP).
+
+In the Texas electricity market (ERCOT), the peak is the monthly
+15-minute interval when the ERCOT Grid is at a point of highest
+capacity. The peak is caused by all consumers’ combined demand on the
+electrical grid. The coincident peak demand is an important factor used
+by ERCOT to determine final electricity consumption bills. ERCOT
+registers the CP demand of each client for 4 months, between June and
+September, and uses this to adjust electricity prices. Clients can
+therefore save on electricity bills by reducing the coincident peak
+demand.
+
+In this example we will train an
+[`MSTL`](../../src/core/models.html#mstl)
+(Multiple Seasonal-Trend decomposition using LOESS) model on historic
+load data to forecast day-ahead peaks on September 2022. Multiple
+seasonality is traditionally present in low sampled electricity data.
+Demand exhibits daily and weekly seasonality, with clear patterns for
+specific hours of the day such as 6:00pm vs 3:00am or for specific days
+such as Sunday vs Friday.
+
+First, we will load ERCOT historic demand, then we will use the
+[`StatsForecast.cross_validation`](../../src/core/core.html#statsforecast.cross_validation)
+method to fit the MSTL model and forecast daily load during September.
+Finally, we show how to use the forecasts to detect the coincident peak.
+
+**Outline**
+
+1. Install libraries
+2. Load and explore the data
+3. Fit MSTL model and forecast
+4. Peak detection
+
+> **Tip**
+>
+> You can use Colab to run this Notebook interactively
+>
+
+## Libraries
+
+We assume you have StatsForecast already installed. Check this guide for
+instructions on [how to install StatsForecast](../getting-started/installation.html).
+
+Install the necessary packages using `pip install statsforecast`
+
+## Load Data
+
+The input to StatsForecast is always a data frame in [long
+format](https://www.theanalysisfactor.com/wide-and-long-data/) with
+three columns: `unique_id`, `ds` and `y`:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp or int) column should be either an integer
+ indexing time or a datestamp ideally like YYYY-MM-DD for a date or
+ YYYY-MM-DD HH:MM:SS for a timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast.
+
+
+```python
+import numpy as np
+import pandas as pd
+```
+
+
+```python
+Y_df = pd.read_csv('https://datasets-nixtla.s3.amazonaws.com/ERCOT-clean.csv', parse_dates=['ds'])
+```
+
+Plot the series using the `plot` method from the
+[`StatsForecast`](../../src/core/core.html#statsforecast-plot)
+class. This method prints up to 8 random series from the dataset and is
+useful for basic EDA.
+
+> **Note**
+>
+> The `StatsForecast.plot` method uses Plotly as a default engine. You
+> can change to MatPlotLib by setting `engine="matplotlib"`.
+
+
+```python
+from statsforecast import StatsForecast
+```
+
+
+```python
+StatsForecast.plot(Y_df)
+```
+
+
+
+We observe that the time series exhibits seasonal patterns. Moreover,
+the time series contains `6,552` observations, so it is necessary to use
+computationally efficient methods to deploy them in production.
+
+## Fit and Forecast MSTL model
+
+The MSTL (Multiple Seasonal-Trend decomposition using LOESS) model
+decomposes the time series in multiple seasonalities using a Local
+Polynomial Regression (LOESS). Then it forecasts the trend using a
+custom non-seasonal model and each seasonality using a SeasonalNaive
+model.
+
+> **Tip**
+>
+> Check our detailed explanation and tutorial on MSTL
+> [here](./multipleseasonalities.html)
+
+Import the
+[`StatsForecast`](../../src/core/core.html#statsforecast)
+class and the models you need.
+
+
+```python
+from sklearn.linear_model import LinearRegression
+from utilsforecast.feature_engineering import trend
+
+from statsforecast import StatsForecast
+from statsforecast.models import MSTL, SklearnModel
+```
+
+First, instantiate the model and define the parameters. The electricity
+load presents seasonalities every 24 hours (Hourly) and every 24 \* 7
+(Daily) hours. Therefore, we will use `[24, 24 * 7]` as the
+seasonalities. See [this
+link](https://robjhyndman.com/hyndsight/seasonal-periods/) for a
+detailed explanation on how to set seasonal lengths. In this example we
+use the
+[`SklearnModel`](../../src/core/models.html#sklearnmodel)
+with a `LinearRegression` model for the trend component, however, any
+StatsForecast model can be used. The complete list of models is
+available [here](../../src/core/models.html).
+
+
+```python
+train, _ = trend(Y_df, freq='H')
+train.head()
+```
+
+| | unique_id | ds | y | trend |
+|-----|-----------|---------------------|--------------|-------|
+| 0 | ERCOT | 2021-01-01 00:00:00 | 43719.849616 | 1.0 |
+| 1 | ERCOT | 2021-01-01 01:00:00 | 43321.050347 | 2.0 |
+| 2 | ERCOT | 2021-01-01 02:00:00 | 43063.067063 | 3.0 |
+| 3 | ERCOT | 2021-01-01 03:00:00 | 43090.059203 | 4.0 |
+| 4 | ERCOT | 2021-01-01 04:00:00 | 43486.590073 | 5.0 |
+
+
+```python
+models = [
+ MSTL(
+ season_length=[24, 24 * 7], # seasonalities of the time series
+ trend_forecaster=SklearnModel(LinearRegression()) # model used to forecast trend
+ )
+]
+```
+
+We fit the model by instantiating a
+[`StatsForecast`](../../src/core/core.html#statsforecast)
+object with the following required parameters:
+
+- `models`: a list of models. Select the models you want from
+ [models](../models.html) and import them.
+
+- `freq`: a string indicating the frequency of the data. (See [panda’s
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+
+```python
+# Instantiate StatsForecast class as sf
+sf = StatsForecast(
+ models=models,
+ freq='H',
+)
+```
+
+> **Tip**
+>
+> StatsForecast also supports this optional parameter.
+>
+> - `n_jobs`: n_jobs: int, number of jobs used in the parallel
+> processing, use -1 for all cores. (Default: 1)
+>
+> - `fallback_model`: a model to be used if a model fails. (Default:
+> none)
+
+The
+[`cross_validation`](../../src/mfles.html#cross_validation)
+method allows the user to simulate multiple historic forecasts, greatly
+simplifying pipelines by replacing for loops with `fit` and `predict`
+methods. This method re-trains the model and forecast each window. See
+[this
+tutorial](../getting-started/getting_started_complete.html)
+for an animation of how the windows are defined.
+
+Use the
+[`cross_validation`](../../src/mfles.html#cross_validation)
+method to produce all the daily forecasts for September. To produce
+daily forecasts set the forecasting horizon `h` as 24. In this example
+we are simulating deploying the pipeline during September, so set the
+number of windows as 30 (one for each day). Finally, set the step size
+between windows as 24, to only produce one forecast per day.
+
+
+```python
+cv_df = sf.cross_validation(
+ df=train,
+ h=24,
+ step_size=24,
+ n_windows=30
+ )
+```
+
+
+```python
+cv_df.head()
+```
+
+| | unique_id | ds | cutoff | y | MSTL |
+|----|----|----|----|----|----|
+| 0 | ERCOT | 2022-09-01 00:00:00 | 2022-08-31 23:00:00 | 45482.471757 | 47413.944185 |
+| 1 | ERCOT | 2022-09-01 01:00:00 | 2022-08-31 23:00:00 | 43602.658043 | 45237.153285 |
+| 2 | ERCOT | 2022-09-01 02:00:00 | 2022-08-31 23:00:00 | 42284.817342 | 43816.390019 |
+| 3 | ERCOT | 2022-09-01 03:00:00 | 2022-08-31 23:00:00 | 41663.156771 | 42972.956286 |
+| 4 | ERCOT | 2022-09-01 04:00:00 | 2022-08-31 23:00:00 | 41710.621904 | 42909.899438 |
+
+> **Important**
+>
+> When using
+> [`cross_validation`](../../src/mfles.html#cross_validation)
+> make sure the forecasts are produced at the desired timestamps. Check
+> the `cutoff` column which specifices the last timestamp before the
+> forecasting window.
+
+## Peak Detection
+
+Finally, we use the forecasts in `cv_df` to detect the daily hourly
+demand peaks. For each day, we set the detected peaks as the highest
+forecasts. In this case, we want to predict one peak (`npeaks`);
+depending on your setting and goals, this parameter might change. For
+example, the number of peaks can correspond to how many hours a battery
+can be discharged to reduce demand.
+
+
+```python
+npeaks = 1 # Number of peaks
+```
+
+For the ERCOT 4CP detection task we are interested in correctly
+predicting the highest monthly load. Next, we filter the day in
+September with the highest hourly demand and predict the peak.
+
+
+```python
+cv_df = cv_df[['ds','y','MSTL']]
+max_day = cv_df.iloc[cv_df['y'].argmax()].ds.day # Day with maximum load
+cv_df_day = cv_df.query('ds.dt.day == @max_day')
+max_hour = cv_df_day['y'].argmax()
+peaks = cv_df_day['MSTL'].argsort().iloc[-npeaks:].values # Predicted peaks
+```
+
+In the following plot we see how the MSTL model is able to correctly
+detect the coincident peak for September 2022.
+
+
+```python
+import matplotlib.pyplot as plt
+```
+
+
+```python
+plt.figure(figsize=(10, 5))
+plt.axvline(cv_df_day.iloc[max_hour]['ds'], color='black', label='True Peak')
+plt.scatter(cv_df_day.iloc[peaks]['ds'], cv_df_day.iloc[peaks]['MSTL'], color='green', label=f'Predicted Top-{npeaks}')
+plt.plot(cv_df_day['ds'], cv_df_day['y'], label='y', color='blue')
+plt.plot(cv_df_day['ds'], cv_df_day['MSTL'], label='Forecast', color='red')
+plt.xlabel('Time')
+plt.ylabel('Load (MW)')
+plt.grid()
+plt.legend()
+```
+
+
+
+> **Important**
+>
+> In this example we only include September. However, MSTL can correctly
+> predict the peaks for the 4 months of 2022. You can try this by
+> increasing the `nwindows` parameter of
+> [`cross_validation`](../../src/mfles.html#cross_validation)
+> or filtering the `Y_df` dataset. The complete run for all months take
+> only 10 minutes.
+
+## Next steps
+
+StatsForecast and MSTL in particular are good benchmarking models for
+peak detection. However, it might be useful to explore further and newer
+forecasting algorithms. We have seen particularly good results with the
+N-HiTS, a deep-learning model from Nixtla’s NeuralForecast library.
+
+Learn how to predict ERCOT demand peaks with our deep-learning N-HiTS
+model and the NeuralForecast library in [this
+tutorial](../../../neuralforecast/docs/use-cases/electricity_peak_forecasting.html).
+
+## References
+
+- [Bandara, Kasun & Hyndman, Rob & Bergmeir, Christoph. (2021). “MSTL:
+ A Seasonal-Trend Decomposition Algorithm for Time Series with
+ Multiple Seasonal Patterns”](https://arxiv.org/abs/2107.13462).
+- [Cristian Challu, Kin G. Olivares, Boris N. Oreshkin, Federico
+ Garza, Max Mergenthaler-Canseco, Artur Dubrawski (2021). “N-HiTS:
+ Neural Hierarchical Interpolation for Time Series Forecasting”.
+ Accepted at AAAI 2023.](https://arxiv.org/abs/2201.12886)
+
diff --git a/statsforecast/docs/tutorials/garch_tutorial.html.mdx b/statsforecast/docs/tutorials/garch_tutorial.html.mdx
new file mode 100644
index 00000000..b069827d
--- /dev/null
+++ b/statsforecast/docs/tutorials/garch_tutorial.html.mdx
@@ -0,0 +1,480 @@
+---
+description: >-
+ In this example, we'll forecast the volatility of the S&P 500 and several
+ publicly traded companies using GARCH and ARCH models
+output-file: garch_tutorial.html
+title: Volatility forecasting (GARCH & ARCH)
+---
+
+
+> **Prerequesites**
+>
+> This tutorial assumes basic familiarity with StatsForecast. For a
+> minimal example visit the [Quick
+> Start](https://nixtla.github.io/statsforecast/docs/getting-started/getting_started_short.html)
+
+## Introduction
+
+The Generalized Autoregressive Conditional Heteroskedasticity (GARCH)
+model is used for time series that exhibit non-constant volatility over
+time. Here volatility refers to the conditional standard deviation. The
+GARCH(p,q) model is given by
+
+where $v_t$ is independent and identically distributed with zero mean
+and unit variance, and $\sigma_t$ evolves according to
+
+The coefficients in the equation above must satisfy the following
+conditions:
+
+1. $w>0$, $\alpha_i \geq 0$ for all $i$, and $\beta_j \geq 0$ for all
+ $j$
+2. $\sum_{k=1}^{max(p,q)} \alpha_k + \beta_k < 1$. Here it is assumed
+ that $\alpha_i=0$ for $i>p$ and $\beta_j=0$ for $j>q$.
+
+A particular case of the GARCH model is the ARCH model, in which $q=0$.
+Both models are commonly used in finance to model the volatility of
+stock prices, exchange rates, interest rates, and other financial
+instruments. They’re also used in risk management to estimate the
+probability of large variations in the price of financial assets.
+
+By the end of this tutorial, you’ll have a good understanding of how to
+implement a GARCH or an ARCH model in
+[StatsForecast](https://nixtla.github.io/statsforecast/) and how they
+can be used to analyze and predict financial time series data.
+
+**Outline:**
+
+1. Install libraries
+2. Load and explore the data
+3. Train models
+4. Perform time series cross-validation
+5. Evaluate results
+6. Forecast volatility
+
+> **Tip**
+>
+> You can use Colab to run this Notebook interactively
+>
+
+## Install libraries
+
+We assume that you have StatsForecast already installed. If not, check
+this guide for instructions on [how to install
+StatsForecast](https://nixtla.github.io/statsforecast/docs/getting-started/installation.html)
+
+Install the necessary packages using `pip install statsforecast`
+
+
+```python
+pip install statsforecast -U
+```
+
+## Load and explore the data
+
+In this tutorial, we’ll use the last 5 years of prices from the S&P 500
+and several publicly traded companies. The data can be downloaded from
+Yahoo! Finance using [yfinance](https://github.com/ranaroussi/yfinance).
+To install it, use `pip install yfinance`.
+
+
+```python
+# pip install yfinance
+```
+
+We’ll also need `pandas` to deal with the dataframes.
+
+
+```python
+import yfinance as yf
+import pandas as pd
+```
+
+
+```python
+tickers = ['SPY', 'MSFT', 'AAPL', 'GOOG', 'AMZN', 'TSLA', 'NVDA', 'META', 'NKE', 'NFLX']
+df = yf.download(tickers, start = '2018-01-01', end = '2022-12-31', interval='1mo', progress=False) # use monthly prices
+df.head()
+```
+
+| Price | Adj Close | | | | | | | | | | ... | Volume | | | | | | | | | |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| Ticker | AAPL | AMZN | GOOG | META | MSFT | NFLX | NKE | NVDA | SPY | TSLA | ... | AAPL | AMZN | GOOG | META | MSFT | NFLX | NKE | NVDA | SPY | TSLA |
+| Date | | | | | | | | | | | | | | | | | | | | | |
+| 2018-01-01 | 39.388084 | 72.544502 | 58.353695 | 186.328979 | 88.027702 | 270.299988 | 63.341862 | 6.078998 | 252.565216 | 23.620667 | ... | 2638717600 | 1927424000 | 574768000 | 495655700 | 574258400 | 238377600 | 157812200 | 11456216000 | 1985506700 | 1864072500 |
+| 2018-02-01 | 41.902908 | 75.622498 | 55.101181 | 177.784729 | 86.878807 | 291.380005 | 62.236938 | 5.985018 | 243.381882 | 22.870667 | ... | 3711577200 | 2755680000 | 847640000 | 516251600 | 725663300 | 184585800 | 160317000 | 14915528000 | 2923722000 | 1637850000 |
+| 2018-03-01 | 39.631344 | 72.366997 | 51.463116 | 159.310333 | 84.959763 | 295.350006 | 61.689133 | 5.731123 | 235.766373 | 17.742001 | ... | 2854910800 | 2608002000 | 907066000 | 996201700 | 750754800 | 263449400 | 174066700 | 14118440000 | 2323561800 | 2359027500 |
+| 2018-04-01 | 39.036106 | 78.306503 | 50.741886 | 171.483688 | 87.054207 | 312.459991 | 63.691761 | 5.565567 | 237.934006 | 19.593332 | ... | 2664617200 | 2598392000 | 834318000 | 750072700 | 668130700 | 262006000 | 158981900 | 11144008000 | 1998466500 | 2854662000 |
+| 2018-05-01 | 44.140598 | 81.481003 | 54.116600 | 191.204315 | 92.006393 | 351.600006 | 66.867508 | 6.240908 | 243.717957 | 18.982000 | ... | 2483905200 | 1432310000 | 636988000 | 401144100 | 509417900 | 142050800 | 129566300 | 11978240000 | 1606397200 | 2333671500 |
+
+The data downloaded includes different prices. We’ll use the [adjusted
+closing
+price](https://help.yahoo.com/kb/SLN28256.html#:~:text=Adjusted%20close%20is%20the%20closing,Security%20Prices%20(CRSP)%20standards.),
+which is the closing price after accounting for any corporate actions
+like stock splits or dividend distributions. It is also the price that
+is used to examine historical returns.
+
+Notice that the dataframe that `yfinance` returns has a
+[MultiIndex](https://pandas.pydata.org/docs/user_guide/advanced.html),
+so we need to select both the adjusted price and the tickers.
+
+
+```python
+df = df.loc[:, (['Adj Close'], tickers)]
+df.columns = df.columns.droplevel() # drop MultiIndex
+df = df.reset_index()
+df.head()
+```
+
+| Ticker | Date | SPY | MSFT | AAPL | GOOG | AMZN | TSLA | NVDA | META | NKE | NFLX |
+|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | 2018-01-01 | 252.565216 | 88.027702 | 39.388084 | 58.353695 | 72.544502 | 23.620667 | 6.078998 | 186.328979 | 63.341862 | 270.299988 |
+| 1 | 2018-02-01 | 243.381882 | 86.878807 | 41.902908 | 55.101181 | 75.622498 | 22.870667 | 5.985018 | 177.784729 | 62.236938 | 291.380005 |
+| 2 | 2018-03-01 | 235.766373 | 84.959763 | 39.631344 | 51.463116 | 72.366997 | 17.742001 | 5.731123 | 159.310333 | 61.689133 | 295.350006 |
+| 3 | 2018-04-01 | 237.934006 | 87.054207 | 39.036106 | 50.741886 | 78.306503 | 19.593332 | 5.565567 | 171.483688 | 63.691761 | 312.459991 |
+| 4 | 2018-05-01 | 243.717957 | 92.006393 | 44.140598 | 54.116600 | 81.481003 | 18.982000 | 6.240908 | 191.204315 | 66.867508 | 351.600006 |
+
+The input to StatsForecast is a dataframe in [long
+format](https://www.theanalysisfactor.com/wide-and-long-data/) with
+three columns: `unique_id`, `ds` and `y`:
+
+- `unique_id`: (string, int or category) A unique identifier for the
+ series.
+- `ds`: (datestamp or int) A datestamp in format YYYY-MM-DD or
+ YYYY-MM-DD HH:MM:SS or an integer indexing time.
+- `y`: (numeric) The measurement we wish to forecast.
+
+Hence, we need to reshape the data. We’ll do this by creating a new
+dataframe called `price`.
+
+
+```python
+prices = df.melt(id_vars = 'Date')
+prices = prices.rename(columns={'Date': 'ds', 'Ticker': 'unique_id', 'value': 'y'})
+prices = prices[['unique_id', 'ds', 'y']]
+prices
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|------------|
+| 0 | SPY | 2018-01-01 | 252.565216 |
+| 1 | SPY | 2018-02-01 | 243.381882 |
+| 2 | SPY | 2018-03-01 | 235.766373 |
+| 3 | SPY | 2018-04-01 | 237.934006 |
+| 4 | SPY | 2018-05-01 | 243.717957 |
+| ... | ... | ... | ... |
+| 595 | NFLX | 2022-08-01 | 223.559998 |
+| 596 | NFLX | 2022-09-01 | 235.440002 |
+| 597 | NFLX | 2022-10-01 | 291.880005 |
+| 598 | NFLX | 2022-11-01 | 305.529999 |
+| 599 | NFLX | 2022-12-01 | 294.880005 |
+
+We can plot this series using the `plot` method of the StatsForecast
+class.
+
+
+```python
+from statsforecast import StatsForecast
+```
+
+
+```python
+StatsForecast.plot(prices)
+```
+
+
+
+With the prices, we can compute the logarithmic returns of the S&P 500
+and the publicly traded companies. This is the variable we’re interested
+in since it’s likely to work well with the GARCH framework. The
+logarithmic return is given by
+
+$return_t = log \big( \frac{price_t}{price_{t-1}} \big)$
+
+We’ll compute the returns on the price dataframe and then we’ll create a
+return dataframe with StatsForecast’s format. To do this, we’ll need
+`numpy`.
+
+
+```python
+import numpy as np
+prices['rt'] = prices['y'].div(prices.groupby('unique_id')['y'].shift(1))
+prices['rt'] = np.log(prices['rt'])
+
+returns = prices[['unique_id', 'ds', 'rt']]
+returns = returns.rename(columns={'rt':'y'})
+returns
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|-----------|
+| 0 | SPY | 2018-01-01 | NaN |
+| 1 | SPY | 2018-02-01 | -0.037038 |
+| 2 | SPY | 2018-03-01 | -0.031790 |
+| 3 | SPY | 2018-04-01 | 0.009152 |
+| 4 | SPY | 2018-05-01 | 0.024018 |
+| ... | ... | ... | ... |
+| 595 | NFLX | 2022-08-01 | -0.005976 |
+| 596 | NFLX | 2022-09-01 | 0.051776 |
+| 597 | NFLX | 2022-10-01 | 0.214887 |
+| 598 | NFLX | 2022-11-01 | 0.045705 |
+| 599 | NFLX | 2022-12-01 | -0.035479 |
+
+> **Warning**
+>
+> If the order of the data is very small (say $<1e-5$),
+> `scipy.optimize.minimize` might not terminate successfully. In this
+> case, rescale the data and then generate the GARCH or ARCH model.
+
+
+```python
+StatsForecast.plot(returns)
+```
+
+
+
+From this plot, we can see that the returns seem suited for the GARCH
+framework, since large shocks *tend* to be followed by other large
+shocks. This doesn’t mean that after every large shock we should expect
+another one; merely that the probability of a large variance is greater
+than the probability of a small one.
+
+## Train models
+
+We first need to import the
+[GARCH](https://Nixtla.github.io/statsforecast/src/core/models.html#garch-model) and the
+[ARCH](https://Nixtla.github.io/statsforecast/src/core/models.html#arch-model) models from
+`statsforecast.models`, and then we need to fit them by instantiating a
+new StatsForecast object. Notice that we’ll be using different values of
+$p$ and $q$. In the next section, we’ll determine which ones produce the
+most accurate model using cross-validation. We’ll also import the
+[Naive](https://Nixtla.github.io/statsforecast/src/core/models.html#naive) model
+since we’ll use it as a baseline.
+
+
+```python
+from statsforecast.models import (
+ GARCH,
+ ARCH,
+ Naive
+)
+
+models = [ARCH(1),
+ ARCH(2),
+ GARCH(1,1),
+ GARCH(1,2),
+ GARCH(2,2),
+ GARCH(2,1),
+ Naive()
+]
+```
+
+To instantiate a new StatsForecast object, we need the following
+parameters:
+
+- `df`: The dataframe with the training data.
+- `models`: The list of models defined in the previous step.
+- `freq`: A string indicating the frequency of the data. Here we’ll
+ use **MS**, which correspond to the start of the month. You can see
+ the list of panda’s available frequencies
+ [here](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+- `n_jobs`: An integer that indicates the number of jobs used in
+ parallel processing. Use -1 to select all cores.
+
+
+```python
+sf = StatsForecast(
+ models = models,
+ freq = 'MS',
+ n_jobs = -1
+)
+```
+
+## Perform time series cross-validation
+
+Time series cross-validation is a method for evaluating how a model
+would have performed in the past. It works by defining a sliding window
+across the historical data and predicting the period following it. Here
+we’ll use StatsForercast’s `cross-validation` method to determine the
+most accurate model for the S&P 500 and the companies selected.
+
+This method takes the following arguments:
+
+- `df`: The dataframe with the training data.
+- `h` (int): represents the h steps into the future that will be
+ forecasted.
+- `step_size` (int): step size between each window, meaning how often
+ do you want to run the forecasting process.
+- `n_windows` (int): number of windows used for cross-validation,
+ meaning the number of forecasting processes in the past you want to
+ evaluate.
+
+For this particular example, we’ll use 4 windows of 3 months, or all the
+quarters in a year.
+
+
+```python
+cv_df = sf.cross_validation(
+ df = returns,
+ h = 3,
+ step_size = 3,
+ n_windows = 4
+ )
+```
+
+The `cv_df` object ia a dataframe with the following columns:
+
+- `unique_id`: series identifier.
+- `ds`: datestamp or temporal index
+- `cutoff`: the last datestamp or temporal index for the `n_windows`.
+- `y`: true value
+- `"model"`: columns with the model’s name and fitted value.
+
+
+```python
+cv_df.rename(columns = {'y' : 'actual'}, inplace = True)
+cv_df.head()
+```
+
+| | unique_id | ds | cutoff | actual | ARCH(1) | ARCH(2) | GARCH(1,1) | GARCH(1,2) | GARCH(2,2) | GARCH(2,1) | Naive |
+|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | AAPL | 2022-01-01 | 2021-12-01 | -0.015837 | 0.142421 | 0.144016 | 0.142954 | 0.141682 | 0.141682 | 0.144015 | 0.073061 |
+| 1 | AAPL | 2022-02-01 | 2021-12-01 | -0.056856 | -0.056893 | -0.057158 | -0.056388 | -0.058786 | -0.058785 | -0.057158 | 0.073061 |
+| 2 | AAPL | 2022-03-01 | 2021-12-01 | 0.057156 | -0.045901 | -0.046479 | -0.047513 | -0.045711 | -0.045711 | -0.046478 | 0.073061 |
+| 3 | AAPL | 2022-04-01 | 2022-03-01 | -0.102178 | 0.138650 | 0.140222 | 0.228138 | 0.136118 | 0.136132 | 0.140211 | 0.057156 |
+| 4 | AAPL | 2022-05-01 | 2022-03-01 | -0.057505 | -0.056007 | -0.056268 | -0.087833 | -0.057078 | -0.057085 | -0.056265 | 0.057156 |
+
+
+```python
+StatsForecast.plot(returns, cv_df.drop(['cutoff', 'actual'], axis=1))
+```
+
+
+
+A tutorial on cross-validation can be found
+[here](https://nixtla.github.io/statsforecast/docs/tutorials/crossvalidation.html).
+
+## Evaluate results
+
+To compute the accuracy of the forecasts, we’ll use the mean average
+error (mae), which is the sum of the absolute errors divided by the
+number of forecasts.
+
+
+```python
+from utilsforecast.losses import mae
+```
+
+The MAE needs to be computed for every window and then it needs to be
+averaged across all of them. To do this, we’ll create the following
+function.
+
+
+```python
+models = cv_df.columns.drop(['unique_id', 'ds', 'cutoff', 'actual'])
+```
+
+
+```python
+mae_cv = mae(cv_df, models=models, target_col='actual').set_index('unique_id')
+mae_cv
+```
+
+| | ARCH(1) | ARCH(2) | GARCH(1,1) | GARCH(1,2) | GARCH(2,2) | GARCH(2,1) | Naive |
+|----|----|----|----|----|----|----|----|
+| unique_id | | | | | | | |
+| AAPL | 0.071773 | 0.068927 | 0.080182 | 0.075321 | 0.069187 | 0.068817 | 0.110426 |
+| AMZN | 0.127390 | 0.113613 | 0.118859 | 0.119930 | 0.109910 | 0.109910 | 0.115189 |
+| GOOG | 0.093849 | 0.093753 | 0.109662 | 0.101583 | 0.094648 | 0.103389 | 0.083233 |
+| META | 0.198334 | 0.198893 | 0.199615 | 0.199711 | 0.199712 | 0.198892 | 0.185346 |
+| MSFT | 0.082373 | 0.075055 | 0.072241 | 0.072765 | 0.073006 | 0.082066 | 0.086951 |
+| NFLX | 0.159386 | 0.159528 | 0.199623 | 0.232477 | 0.230075 | 0.230770 | 0.167421 |
+| NKE | 0.108337 | 0.098918 | 0.103366 | 0.110278 | 0.107179 | 0.102708 | 0.160404 |
+| NVDA | 0.189461 | 0.207871 | 0.198999 | 0.196170 | 0.211932 | 0.211940 | 0.215289 |
+| SPY | 0.058511 | 0.058583 | 0.058701 | 0.062492 | 0.057053 | 0.068192 | 0.089012 |
+| TSLA | 0.192003 | 0.192618 | 0.190225 | 0.192354 | 0.191620 | 0.191423 | 0.218857 |
+
+
+```python
+mae_cv.idxmin(axis=1)
+```
+
+``` text
+unique_id
+AAPL GARCH(2,1)
+AMZN GARCH(2,2)
+GOOG Naive
+META Naive
+MSFT GARCH(1,1)
+NFLX ARCH(1)
+NKE ARCH(2)
+NVDA ARCH(1)
+SPY GARCH(2,2)
+TSLA GARCH(1,1)
+dtype: object
+```
+
+Hence, the most accurate model to describe the logarithmic returns of
+Apple’s stock is an GARCH(2, 1), for Amazon’s stock is a GARCH(2,2), and
+so on.
+
+## Forecast volatility
+
+We can now generate a forecast for the next quarter. To do this, we’ll
+use the `forecast` method, which requieres the following arguments:
+
+- `h`: (int) The forecasting horizon.
+- `level`: (list\[float\]) The confidence levels of the prediction
+ intervals
+- `fitted` : (bool = False) Returns insample predictions.
+
+
+```python
+levels = [80, 95] # confidence levels for the prediction intervals
+
+forecasts = sf.forecast(df=returns, h=3, level=levels)
+forecasts.head()
+```
+
+| | unique_id | ds | ARCH(1) | ARCH(1)-lo-95 | ARCH(1)-lo-80 | ARCH(1)-hi-80 | ARCH(1)-hi-95 | ARCH(2) | ARCH(2)-lo-95 | ARCH(2)-lo-80 | ... | GARCH(2,1) | GARCH(2,1)-lo-95 | GARCH(2,1)-lo-80 | GARCH(2,1)-hi-80 | GARCH(2,1)-hi-95 | Naive | Naive-lo-80 | Naive-lo-95 | Naive-hi-80 | Naive-hi-95 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | AAPL | 2023-01-01 | 0.150457 | 0.133641 | 0.139462 | 0.161453 | 0.167273 | 0.150158 | 0.133409 | 0.139206 | ... | 0.147602 | 0.131418 | 0.137020 | 0.158184 | 0.163786 | -0.128762 | -0.284462 | -0.366885 | 0.026939 | 0.109362 |
+| 1 | AAPL | 2023-02-01 | -0.056943 | -0.073924 | -0.068046 | -0.045839 | -0.039961 | -0.057207 | -0.074346 | -0.068414 | ... | -0.059512 | -0.078060 | -0.071640 | -0.047384 | -0.040964 | -0.128762 | -0.348956 | -0.465520 | 0.091433 | 0.207997 |
+| 2 | AAPL | 2023-03-01 | -0.048391 | -0.064843 | -0.059148 | -0.037633 | -0.031939 | -0.049282 | -0.066345 | -0.060439 | ... | -0.054539 | -0.075438 | -0.068204 | -0.040875 | -0.033641 | -0.128762 | -0.398443 | -0.541204 | 0.140920 | 0.283681 |
+| 3 | AMZN | 2023-01-01 | 0.152147 | 0.134952 | 0.140904 | 0.163391 | 0.169343 | 0.148658 | 0.132242 | 0.137924 | ... | 0.148599 | 0.132196 | 0.137873 | 0.159324 | 0.165001 | -0.139141 | -0.315716 | -0.409190 | 0.037435 | 0.130909 |
+| 4 | AMZN | 2023-02-01 | -0.057301 | -0.074497 | -0.068545 | -0.046058 | -0.040106 | -0.061187 | -0.080794 | -0.074007 | ... | -0.069303 | -0.094457 | -0.085750 | -0.052856 | -0.044150 | -0.139141 | -0.388856 | -0.521048 | 0.110575 | 0.242767 |
+
+With the results of the previous section, we can choose the best model
+for the S&P 500 and the companies selected. Some of the plots are shown
+below. Notice that we’re using somo additional arguments in the `plot`
+method:
+
+- `level`: (list\[int\]) The confidence levels for the prediction
+ intervals (this was already defined).
+- `unique_ids`: (list\[str, int or category\]) The ids to plot.
+- `models`: (list(str)). The model to plot. In this case, is the model
+ selected by cross-validation.
+
+
+```python
+StatsForecast.plot(returns, forecasts, max_insample_length=20)
+```
+
+
+
+## References
+
+- [Engle, R. F. (1982). Autoregressive conditional heteroscedasticity
+ with estimates of the variance of United Kingdom inflation.
+ Econometrica: Journal of the econometric society,
+ 987-1007.](http://www.econ.uiuc.edu/~econ508/Papers/engle82.pdf)
+
+- [Bollerslev, T. (1986). Generalized autoregressive conditional
+ heteroskedasticity. Journal of econometrics, 31(3),
+ 307-327.](https://www.sciencedirect.com/science/article/abs/pii/0304407686900631)
+
+- [Hamilton, J. D. (1994). Time series analysis. Princeton university
+ press.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+
+- [Tsay, R. S. (2005). Analysis of financial time series. John wiley &
+ sons.](https://www.wiley.com/en-us/Analysis+of+Financial+Time+Series%2C+3rd+Edition-p-9780470414354)
+
diff --git a/statsforecast/docs/tutorials/intermittentdata.html.mdx b/statsforecast/docs/tutorials/intermittentdata.html.mdx
new file mode 100644
index 00000000..adfca91c
--- /dev/null
+++ b/statsforecast/docs/tutorials/intermittentdata.html.mdx
@@ -0,0 +1,272 @@
+---
+description: In this notebook, we'll implement models for intermittent or sparse data
+output-file: intermittentdata.html
+title: Intermittent or Sparse Data
+---
+
+
+Intermittent or sparse data has very few non-zero observations. This
+type of data is hard to forecast because the zero values increase the
+uncertainty about the underlying patterns in the data. Furthermore, once
+a non-zero observation occurs, there can be considerable variation in
+its size. Intermittent time series are common in many industries,
+including finance, retail, transportation, and energy. Given the
+ubiquity of this type of series, special methods have been developed to
+forecast them. The first was from [Croston (1972)](#ref), followed by
+several variants and by different aggregation frameworks.
+
+[StatsForecast](https://nixtla.github.io/statsforecast/) has implemented
+several models to forecast intermittent time series. By the end of this
+tutorial, you’ll have a good understanding of these models and how to
+use them.
+
+**Outline:**
+
+1. Install libraries
+2. Load and explore the data
+3. Train models for intermittent data
+4. Plot forecasts and compute accuracy
+
+> **Tip**
+>
+> You can use Colab to run this Notebook interactively
+>
+
+> **Tip**
+>
+> For forecasting at scale, we recommend you check [this
+> notebook](https://www.databricks.com/blog/2022/12/06/intermittent-demand-forecasting-nixtla-databricks.html)
+> done on Databricks.
+
+## Install libraries
+
+We assume that you have StatsForecast already installed. If not, check
+this guide for instructions on [how to install
+StatsForecast](../getting-started/installation.html)
+
+Install the necessary packages using `pip install statsforecast`
+
+
+```python
+pip install statsforecast -U
+```
+
+## Load and explore the data
+
+For this example, we’ll use a subset of the [M5
+Competition](https://www.sciencedirect.com/science/article/pii/S0169207021001187#:~:text=The%20objective%20of%20the%20M5,the%20uncertainty%20around%20these%20forecasts)
+dataset. Each time series represents the unit sales of a particular
+product in a given Walmart store. At this level (product-store), most of
+the data is intermittent. We first need to import the data.
+
+
+```python
+import pandas as pd
+```
+
+
+```python
+uids = [
+ 'FOODS_1_001_CA_1',
+ 'FOODS_1_001_CA_2',
+ 'FOODS_1_001_CA_3',
+ 'FOODS_1_001_CA_4',
+ 'FOODS_1_001_TX_1',
+ 'FOODS_1_001_TX_2',
+ 'FOODS_1_001_TX_3',
+ 'FOODS_1_001_WI_1',
+]
+df = pd.read_parquet(
+ 'https://datasets-nixtla.s3.amazonaws.com/m5_y.parquet',
+ filters=[('unique_id', 'in', uids)],
+)
+```
+
+We can plot these series using the `plot_series` function from
+`utilsforecast.plotting`. This function has multiple parameters, and the
+required ones to generate the plots in this notebook are explained
+below.
+
+- `df`: A `pandas` dataframe with columns \[`unique_id`, `ds`, `y`\].
+- `forecasts_df`: A `pandas` dataframe with columns \[`unique_id`,
+ `ds`\] and models.
+- `plot_random`: Plots the time series randomly.
+- `max_insample_length`: The maximum number of train/insample
+ observations to be plotted.
+- `engine`: The library used to generate the plots. It can also be
+ `matplotlib` for static plots.
+
+
+```python
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+plot_series(df, plot_random=False, max_insample_length=100)
+```
+
+
+
+Here we only plotted the last 100 observations, but we can visualize the
+complete history by removing `max_insample_length`. From these plots, we
+can confirm that the data is indeed intermittent since it has multiple
+periods with zero sales. In fact, in all cases but one, the median value
+is zero.
+
+
+```python
+df.groupby('unique_id', observed=True)['y'].median()
+```
+
+``` text
+unique_id
+FOODS_1_001_CA_1 0.0
+FOODS_1_001_CA_2 1.0
+FOODS_1_001_CA_3 0.0
+FOODS_1_001_CA_4 0.0
+FOODS_1_001_TX_1 0.0
+FOODS_1_001_TX_2 0.0
+FOODS_1_001_TX_3 0.0
+FOODS_1_001_WI_1 0.0
+Name: y, dtype: float32
+```
+
+## Train models for intermittent data
+
+Before training any model, we need to separate the data in a train and a
+test set. The M5 Competition used the last 28 days as test set, so we’ll
+do the same.
+
+
+```python
+valid_start = df['ds'].unique()[-28]
+
+train = df[df['ds'] < valid_start]
+test = df[df['ds'] >= valid_start]
+```
+
+StatsForecast has efficient implementations of multiple models for
+intermittent data. The complete list of models available is
+[here](../../src/core/models.html#sparse-or-intermittent).
+In this notebook, we’ll use:
+
+- [Agregate-Dissagregate Intermittent Demand Approach
+ (ADIDA)](https://Nixtla.github.io/statsforecast/src/core/models.html#adida)
+- [Croston
+ Classic](https://Nixtla.github.io/statsforecast/src/core/models.html#crostonclassic)
+- [Intermittent Multiple Aggregation Prediction Algorithm
+ (IMAPA)](https://Nixtla.github.io/statsforecast/src/core/models.html#imapa)
+- [Teunter-Syntetos-Babai
+ (TSB)](https://Nixtla.github.io/statsforecast/src/core/models.html#tsb)
+
+To use these models, we first need to import them from
+`statsforecast.models` and then we need to instantiate them.
+
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import (
+ ADIDA,
+ CrostonClassic,
+ IMAPA,
+ TSB
+)
+
+# Create a list of models and instantiation parameters
+models = [
+ ADIDA(),
+ CrostonClassic(),
+ IMAPA(),
+ TSB(alpha_d = 0.2, alpha_p = 0.2)
+]
+```
+
+To instantiate a new StatsForecast object, we need the following
+parameters:
+
+- `models`: The list of models defined in the previous step.
+- `freq`: A string indicating the frequency of the data. See [pandas’
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+- `n_jobs`: An integer that indicates the number of jobs used in
+ parallel processing. Use -1 to select all cores.
+
+
+```python
+sf = StatsForecast(
+ models=models,
+ freq='D',
+ n_jobs=-1,
+)
+```
+
+Now we’re ready to generate the forecast. To do this, we’ll use the
+`forecast` method, which requires the forecasting horizon (in this case,
+28 days) as argument.
+
+The models for intermittent series that are currently available in
+StatsForecast can only generate point-forecasts. If prediction intervals
+are needed, then a [probabilisitic
+model](https://nixtla.github.io/statsforecast/#models) should be used.
+
+
+```python
+horizon = 28
+forecasts = sf.forecast(df=train, h=horizon)
+forecasts.head()
+```
+
+| | unique_id | ds | ADIDA | CrostonClassic | IMAPA | TSB |
+|-----|------------------|------------|----------|----------------|----------|----------|
+| 0 | FOODS_1_001_CA_1 | 2016-05-23 | 0.791852 | 0.898247 | 0.705835 | 0.434313 |
+| 1 | FOODS_1_001_CA_1 | 2016-05-24 | 0.791852 | 0.898247 | 0.705835 | 0.434313 |
+| 2 | FOODS_1_001_CA_1 | 2016-05-25 | 0.791852 | 0.898247 | 0.705835 | 0.434313 |
+| 3 | FOODS_1_001_CA_1 | 2016-05-26 | 0.791852 | 0.898247 | 0.705835 | 0.434313 |
+| 4 | FOODS_1_001_CA_1 | 2016-05-27 | 0.791852 | 0.898247 | 0.705835 | 0.434313 |
+
+Finally, we’ll merge the forecast with the actual values.
+
+
+```python
+test = test.merge(forecasts, how='left', on=['unique_id', 'ds'])
+```
+
+## Plot forecasts and compute accuracy
+
+We can generate plots using the `plot_series` function described above.
+
+
+```python
+plot_series(train, test, plot_random=False, max_insample_length=100)
+```
+
+
+
+To compute the accuracy of the forecasts, we’ll use the Mean Average
+Error (MAE), which is the sum of the absolute errors divided by the
+number of forecasts.
+
+
+```python
+from utilsforecast.evaluation import evaluate
+from utilsforecast.losses import mae
+```
+
+
+```python
+evaluate(test, metrics=[mae], agg_fn='mean')
+```
+
+| | metric | ADIDA | CrostonClassic | IMAPA | TSB |
+|-----|--------|----------|----------------|----------|----------|
+| 0 | mae | 0.948729 | 0.944071 | 0.957256 | 1.023126 |
+
+Hence, on average, the forecasts are one unit off.
+
+## References
+
+[Croston, J. D. (1972). Forecasting and stock control for intermittent
+demands. Journal of the Operational Research Society, 23(3),
+289-303.](https://link.springer.com/article/10.1057/jors.1972.50)
+
diff --git a/statsforecast/docs/tutorials/mlflow.html.mdx b/statsforecast/docs/tutorials/mlflow.html.mdx
new file mode 100644
index 00000000..8c7d8d88
--- /dev/null
+++ b/statsforecast/docs/tutorials/mlflow.html.mdx
@@ -0,0 +1,250 @@
+---
+description: Run Statsforecast with MLFlow.
+output-file: mlflow.html
+title: MLFlow
+---
+
+
+
+```python
+from statsforecast.utils import generate_series
+```
+
+
+```python
+series = generate_series(5, min_length=50, max_length=50, equal_ends=True, n_static_features=1)
+series.head()
+```
+
+| | unique_id | ds | y | static_0 |
+|-----|-----------|------------|------------|----------|
+| 0 | 0 | 2000-01-01 | 12.073897 | 43 |
+| 1 | 0 | 2000-01-02 | 59.734166 | 43 |
+| 2 | 0 | 2000-01-03 | 101.260794 | 43 |
+| 3 | 0 | 2000-01-04 | 143.987430 | 43 |
+| 4 | 0 | 2000-01-05 | 185.320406 | 43 |
+
+For the next part, `mlflow` and `mlflavors` are needed. Install them
+with:
+
+
+```bash
+pip install mlflow mlflavors
+```
+
+## Model Logging
+
+
+```python
+import pandas as pd
+import mlflow
+from sklearn.metrics import mean_absolute_error, mean_absolute_percentage_error
+from statsforecast import StatsForecast
+from statsforecast.models import AutoARIMA
+
+import mlflavors
+import requests
+```
+
+
+```python
+ARTIFACT_PATH = "model"
+DATA_PATH = "./data"
+HORIZON = 7
+LEVEL = [90]
+
+with mlflow.start_run() as run:
+ series = generate_series(5, min_length=50, max_length=50, equal_ends=True, n_static_features=1)
+
+ train_df = series.groupby('unique_id').head(43)
+ test_df = series.groupby('unique_id').tail(7)
+ X_test = test_df.drop(columns=["y"])
+ y_test = test_df[["y"]]
+
+ models = [AutoARIMA(season_length=7)]
+
+ sf = StatsForecast(models=models, freq="D", n_jobs=-1)
+
+ sf.fit(df=train_df)
+
+ # Evaluate model
+ y_pred = sf.predict(h=HORIZON, X_df=X_test, level=LEVEL)["AutoARIMA"]
+
+ metrics = {
+ "mae": mean_absolute_error(y_test, y_pred),
+ "mape": mean_absolute_percentage_error(y_test, y_pred),
+ }
+
+ print(f"Metrics: \n{metrics}")
+
+ # Log metrics
+ mlflow.log_metrics(metrics)
+
+ # Log model using pickle serialization (default).
+ mlflavors.statsforecast.log_model(
+ statsforecast_model=sf,
+ artifact_path=ARTIFACT_PATH,
+ serialization_format="pickle",
+ )
+ model_uri = mlflow.get_artifact_uri(ARTIFACT_PATH)
+
+print(f"\nMLflow run id:\n{run.info.run_id}")
+```
+
+``` text
+Metrics:
+{'mae': 6.712853959225143, 'mape': 0.11719246764336884}
+```
+
+``` text
+2023/10/20 23:45:36 WARNING mlflow.utils.environment: Encountered an unexpected error while inferring pip requirements (model URI: /var/folders/w2/91_v34nx0xs2npnl3zsl9tmm0000gn/T/tmpt4686vpu/model/model.pkl, flavor: statsforecast), fall back to return ['statsforecast==1.6.0']. Set logging level to DEBUG to see the full traceback.
+```
+
+``` text
+
+MLflow run id:
+0319bbd664424fcd88d6c532e3ecac77
+```
+
+## Viewing Experiment
+
+To view the newly created experiment and logged artifacts open the
+MLflow UI:
+
+
+```bash
+mlflow ui
+```
+
+## Loading Statsforecast Model
+
+The `statsforecast` model can be loaded from the MLFlow registry using
+the `mlflow.statsforecast.load_model` function and used to generate
+predictions.
+
+
+```python
+loaded_model = mlflavors.statsforecast.load_model(model_uri=model_uri)
+results = loaded_model.predict(h=HORIZON, X_df=X_test, level=LEVEL)
+results.head()
+```
+
+| | ds | AutoARIMA | AutoARIMA-lo-90 | AutoARIMA-hi-90 |
+|-----------|------------|------------|-----------------|-----------------|
+| unique_id | | | | |
+| 0 | 2000-02-13 | 55.894432 | 44.343880 | 67.444984 |
+| 0 | 2000-02-14 | 97.818054 | 86.267502 | 109.368607 |
+| 0 | 2000-02-15 | 146.745422 | 135.194870 | 158.295975 |
+| 0 | 2000-02-16 | 188.888336 | 177.337784 | 200.438904 |
+| 0 | 2000-02-17 | 231.493637 | 219.943085 | 243.044189 |
+
+## Loading Model with pyfunc
+
+[Pyfunc](https://mlflow.org/docs/latest/python_api/mlflow.pyfunc.html)
+is another interface for MLFlow models that has utilities for loading
+and saving models. This code is equivalent in making predictions as
+above.
+
+
+```python
+loaded_pyfunc = mlflavors.statsforecast.pyfunc.load_model(model_uri=model_uri)
+
+# Convert test data to 2D numpy array so it can be passed to pyfunc predict using
+# a single-row Pandas DataFrame configuration argument
+X_test_array = X_test.to_numpy()
+
+# Create configuration DataFrame
+predict_conf = pd.DataFrame(
+ [
+ {
+ "X": X_test_array,
+ "X_cols": X_test.columns,
+ "X_dtypes": list(X_test.dtypes),
+ "h": HORIZON,
+ "level": LEVEL,
+ }
+ ]
+)
+
+
+pyfunc_result = loaded_pyfunc.predict(predict_conf)
+pyfunc_result.head()
+```
+
+| | ds | AutoARIMA | AutoARIMA-lo-90 | AutoARIMA-hi-90 |
+|-----------|------------|------------|-----------------|-----------------|
+| unique_id | | | | |
+| 0 | 2000-02-13 | 55.894432 | 44.343880 | 67.444984 |
+| 0 | 2000-02-14 | 97.818054 | 86.267502 | 109.368607 |
+| 0 | 2000-02-15 | 146.745422 | 135.194870 | 158.295975 |
+| 0 | 2000-02-16 | 188.888336 | 177.337784 | 200.438904 |
+| 0 | 2000-02-17 | 231.493637 | 219.943085 | 243.044189 |
+
+## Model Serving
+
+This section illustrates an example of serving the `pyfunc` flavor to a
+local REST API endpoint and subsequently requesting a prediction from
+the served model. To serve the model run the command below where you
+substitute the run id printed during execution training code.
+
+
+```bash
+mlflow models serve -m runs://model --env-manager local --host 127.0.0.1
+```
+
+After running this, the code below can be ran to send a request.
+
+
+```python
+HORIZON = 7
+LEVEL = [90, 95]
+
+# Define local host and endpoint url
+host = "127.0.0.1"
+url = f"http://{host}:5000/invocations"
+
+# Convert DateTime to string for JSON serialization
+X_test_pyfunc = X_test.copy()
+X_test_pyfunc["ds"] = X_test_pyfunc["ds"].dt.strftime(date_format="%Y-%m-%d")
+
+# Convert to list for JSON serialization
+X_test_list = X_test_pyfunc.to_numpy().tolist()
+
+# Convert index to list of strings for JSON serialization
+X_cols = list(X_test.columns)
+
+# Convert dtypes to string for JSON serialization
+X_dtypes = [str(dtype) for dtype in list(X_test.dtypes)]
+
+predict_conf = pd.DataFrame(
+ [
+ {
+ "X": X_test_list,
+ "X_cols": X_cols,
+ "X_dtypes": X_dtypes,
+ "h": HORIZON,
+ "level": LEVEL,
+ }
+ ]
+)
+
+# Create dictionary with pandas DataFrame in the split orientation
+json_data = {"dataframe_split": predict_conf.to_dict(orient="split")}
+
+# Score model
+response = requests.post(url, json=json_data)
+```
+
+
+```python
+pd.DataFrame(response.json()['predictions']).head()
+```
+
+| | ds | AutoARIMA | AutoARIMA-lo-95 | AutoARIMA-lo-90 | AutoARIMA-hi-90 | AutoARIMA-hi-95 |
+|----|----|----|----|----|----|----|
+| 0 | 2000-02-13T00:00:00 | 55.894432 | 42.131100 | 44.343880 | 67.444984 | 69.657768 |
+| 1 | 2000-02-14T00:00:00 | 97.818054 | 84.054718 | 86.267502 | 109.368607 | 111.581390 |
+| 2 | 2000-02-15T00:00:00 | 146.745422 | 132.982086 | 135.194870 | 158.295975 | 160.508759 |
+| 3 | 2000-02-16T00:00:00 | 188.888336 | 175.125015 | 177.337784 | 200.438904 | 202.651672 |
+| 4 | 2000-02-17T00:00:00 | 231.493637 | 217.730301 | 219.943085 | 243.044189 | 245.256973 |
+
diff --git a/statsforecast/docs/tutorials/multipleseasonalities.html.mdx b/statsforecast/docs/tutorials/multipleseasonalities.html.mdx
new file mode 100644
index 00000000..dba31f47
--- /dev/null
+++ b/statsforecast/docs/tutorials/multipleseasonalities.html.mdx
@@ -0,0 +1,388 @@
+---
+description: >-
+ In this example we will show how to forecast data with multiple seasonalities
+ using an MSTL.
+output-file: multipleseasonalities.html
+title: Multiple seasonalities
+---
+
+
+> **Tip**
+>
+> For this task, StatsForecast’s MSTL is 68% more accurate and 600%
+> faster than [Prophet](https://facebook.github.io/prophet/) and
+> [NeuralProphet](https://neuralprophet.com/). (Reproduce experiments
+> [here](https://github.com/Nixtla/statsforecast/tree/main/experiments/mstl))
+
+Multiple seasonal data refers to time series that have more than one
+clear seasonality. Multiple seasonality is traditionally present in data
+that is sampled at a low frequency. For example, hourly electricity data
+exhibits daily and weekly seasonality. That means that there are clear
+patterns of electricity consumption for specific hours of the day like
+6:00pm vs 3:00am or for specific days like Sunday vs Friday.
+
+Traditional statistical models are not able to model more than one
+seasonal length. In this example, we will show how to model the two
+seasonalities efficiently using Multiple Seasonal-Trend decompositions
+with LOESS
+([`MSTL`](https://Nixtla.github.io/statsforecast/src/core/models.html#mstl)).
+
+For this example, we will use hourly electricity load data from
+Pennsylvania, New Jersey, and Maryland (PJM). The original data can be
+found
+[here](https://github.com/panambY/Hourly_Energy_Consumption).
+(Click here for info on [PJM](https://www.pjm.com/about-pjm))
+
+First, we will load the data, then we will use the `StatsForecast.fit`
+and `StatsForecast.predict` methods to predict the next 24 hours. We
+will then decompose the different elements of the time series into
+trends and its multiple seasonalities. At the end, you will use the
+[`StatsForecast.forecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast.forecast)
+for production-ready forecasting.
+
+**Outline**
+
+1. Install libraries
+2. Load and explore the data
+3. Fit a multiple-seasonality model
+4. Decompose the series in trend and seasonality
+5. Predict the next 24 hours
+6. Optional: Forecast in production
+
+> **Tip**
+>
+> You can use Colab to run this Notebook interactively
+>
+
+## Install libraries
+
+We assume you have StatsForecast already installed. Check this guide for
+instructions on [how to install StatsForecast](./installation.html).
+
+Install the necessary packages using `pip install statsforecast` \`\`
+
+## Load Data
+
+The input to StatsForecast is always a data frame in [long
+format](https://www.theanalysisfactor.com/wide-and-long-data/) with
+three columns: `unique_id`, `ds` and `y`:
+
+- The `unique_id` (string, int or category) represents an identifier
+ for the series.
+
+- The `ds` (datestamp or int) column should be either an integer
+ indexing time or a datestamp ideally like YYYY-MM-DD for a date or
+ YYYY-MM-DD HH:MM:SS for a timestamp.
+
+- The `y` (numeric) represents the measurement we wish to forecast. We
+ will rename the
+
+You will read the data with pandas and change the necessary names. This
+step should take around 2s.
+
+
+```python
+import pandas as pd
+```
+
+
+```python
+df = pd.read_csv('https://raw.githubusercontent.com/panambY/Hourly_Energy_Consumption/master/data/PJM_Load_hourly.csv')
+df.columns = ['ds', 'y']
+df.insert(0, 'unique_id', 'PJM_Load_hourly')
+df = df.sort_values(['unique_id', 'ds']).reset_index(drop=True)
+df.tail()
+```
+
+| | unique_id | ds | y |
+|-------|-----------------|---------------------|---------|
+| 32891 | PJM_Load_hourly | 2001-12-31 20:00:00 | 36392.0 |
+| 32892 | PJM_Load_hourly | 2001-12-31 21:00:00 | 35082.0 |
+| 32893 | PJM_Load_hourly | 2001-12-31 22:00:00 | 33890.0 |
+| 32894 | PJM_Load_hourly | 2001-12-31 23:00:00 | 32590.0 |
+| 32895 | PJM_Load_hourly | 2002-01-01 00:00:00 | 31569.0 |
+
+StatsForecast can handle unsorted data, however, for plotting purposes,
+it is convenient to sort the data frame.
+
+Plot the series using the `plot` method from the
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+class. This method prints up to 8 random series from the dataset and is
+useful for basic EDA. In this case, it will print just one series given
+that we have just one unique_id.
+
+> **Note**
+>
+> The `StatsForecast.plot` method uses matplotlib as a default engine.
+> You can change to plotly by setting `engine="plotly"`.
+
+
+```python
+from statsforecast import StatsForecast
+```
+
+
+```python
+StatsForecast.plot(df)
+```
+
+
+
+The time series exhibits seasonal patterns. Moreover, the time series
+contains `32,896` observations, so it is necessary to use very
+computationally efficient methods.
+
+## Fit an MSTL model
+
+The
+[`MSTL`](https://Nixtla.github.io/statsforecast/src/core/models.html#mstl)
+(Multiple Seasonal-Trend decompositions using LOESS) model, originally
+developed by [Kasun Bandara, Rob J Hyndman and Christoph
+Bergmeir](https://arxiv.org/abs/2107.13462), decomposes the time series
+in multiple seasonalities using a Local Polynomial Regression (LOESS).
+Then it forecasts the trend using a non-seasonal model and each
+seasonality using a
+[`SeasonalNaive`](https://Nixtla.github.io/statsforecast/src/core/models.html#seasonalnaive)
+model. You can choose the non-seasonal model you want to use to forecast
+the trend component of the MSTL model. In this example, we will use an
+AutoARIMA.
+
+Import the models you need.
+
+
+```python
+from statsforecast.models import MSTL, AutoARIMA
+```
+
+First, we must define the model parameters. As mentioned before, the
+electricity load presents seasonalities every 24 hours (Hourly) and
+every 24 \* 7 (Daily) hours. Therefore, we will use `[24, 24 * 7]` for
+season length. The trend component will be forecasted with an
+[`AutoARIMA`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoarima)
+model. (You can also try with:
+[`AutoTheta`](https://Nixtla.github.io/statsforecast/src/core/models.html#autotheta),
+[`AutoCES`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoces),
+and
+[`AutoETS`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoets))
+
+
+```python
+# Create a list of models and instantiation parameters
+
+models = [MSTL(
+ season_length=[24, 24 * 7], # seasonalities of the time series
+ trend_forecaster=AutoARIMA() # model used to forecast trend
+)]
+```
+
+We fit the models by instantiating a new
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+object with the following required parameters:
+
+- `models`: a list of models. Select the models you want from
+ [models](../models.html) and import them.
+
+- `freq`: a string indicating the frequency of the data. (See [panda’s
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).)
+
+Any settings are passed into the constructor. Then you call its fit
+method and pass in the historical data frame.
+
+
+```python
+sf = StatsForecast(
+ models=models, # model used to fit each time series
+ freq='h', # frequency of the data
+)
+```
+
+> **Tip**
+>
+> StatsForecast also supports this optional parameter.
+>
+> - `n_jobs`: n_jobs: int, number of jobs used in the parallel
+> processing, use -1 for all cores. (Default: 1)
+>
+> - `fallback_model`: a model to be used if a model fails. (Default:
+> none)
+
+Use the `fit` method to fit each model to each time series. In this
+case, we are just fitting one model to one series. Check this guide to
+learn how to [fit many models to many
+series](./getting_started_complete.html).
+
+> **Note**
+>
+> StatsForecast achieves its blazing speed using JIT compiling through
+> Numba. The first time you call the statsforecast class, the fit method
+> should take around 10 seconds. The second time -once Numba compiled
+> your settings- it should take less than 5s.
+
+
+```python
+sf = sf.fit(df=df)
+```
+
+## Decompose the series
+
+Once the model is fitted, access the decomposition using the `fitted_`
+attribute of
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast).
+This attribute stores all relevant information of the fitted models for
+each of the time series.
+
+In this case, we are fitting a single model for a single time series, so
+by accessing the fitted\_ location \[0, 0\] we will find the relevant
+information of our model. The
+[`MSTL`](https://Nixtla.github.io/statsforecast/src/core/models.html#mstl)
+class generates a `model_` attribute that contains the way the series
+was decomposed.
+
+
+```python
+sf.fitted_[0, 0].model_
+```
+
+| | data | trend | seasonal24 | seasonal168 | remainder |
+|-------|---------|--------------|--------------|-------------|--------------|
+| 0 | 22259.0 | 25899.808157 | -4720.213546 | 581.308595 | 498.096794 |
+| 1 | 21244.0 | 25900.349395 | -5433.168901 | 571.780657 | 205.038849 |
+| 2 | 20651.0 | 25900.875973 | -5829.135728 | 557.142643 | 22.117112 |
+| 3 | 20421.0 | 25901.387631 | -5704.092794 | 597.696957 | -373.991794 |
+| 4 | 20713.0 | 25901.884103 | -5023.324375 | 922.564854 | -1088.124582 |
+| ... | ... | ... | ... | ... | ... |
+| 32891 | 36392.0 | 33329.031577 | 4254.112720 | 917.258336 | -2108.402633 |
+| 32892 | 35082.0 | 33355.083576 | 3625.077164 | 721.689136 | -2619.849876 |
+| 32893 | 33890.0 | 33381.108409 | 2571.794472 | 549.661529 | -2612.564409 |
+| 32894 | 32590.0 | 33407.105839 | 796.356548 | 361.956280 | -1975.418667 |
+| 32895 | 31569.0 | 33433.075723 | -1260.860917 | 279.777069 | -882.991876 |
+
+We will use matplotlib, to visualize the different components of the
+series.
+
+
+```python
+import matplotlib.pyplot as plt
+```
+
+
+```python
+sf.fitted_[0, 0].model_.tail(24 * 28).plot(subplots=True, grid=True)
+plt.tight_layout()
+plt.show()
+```
+
+
+
+We observe a clear upward trend (orange line) and seasonality repeating
+every day (24H) and every week (168H).
+
+## Predict the next 24 hours
+
+> Probabilistic forecasting with levels
+
+To generate forecasts use the `predict` method.
+
+The `predict` method takes two arguments: forecasts the next `h` (for
+horizon) and `level`.
+
+- `h` (int): represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level` (list of floats): this optional parameter is used for
+ probabilistic forecasting. Set the `level` (or confidence
+ percentile) of your prediction interval. For example, `level=[90]`
+ means that the model expects the real value to be inside that
+ interval 90% of the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This step should take less than 1 second.
+
+
+```python
+forecasts = sf.predict(h=24, level=[90])
+forecasts.head()
+```
+
+| | unique_id | ds | MSTL | MSTL-lo-90 | MSTL-hi-90 |
+|----|----|----|----|----|----|
+| 0 | PJM_Load_hourly | 2002-01-01 01:00:00 | 30215.608123 | 29842.185581 | 30589.030664 |
+| 1 | PJM_Load_hourly | 2002-01-01 02:00:00 | 29447.208519 | 28787.122830 | 30107.294207 |
+| 2 | PJM_Load_hourly | 2002-01-01 03:00:00 | 29132.786369 | 28221.353220 | 30044.219518 |
+| 3 | PJM_Load_hourly | 2002-01-01 04:00:00 | 29126.252713 | 27992.819671 | 30259.685756 |
+| 4 | PJM_Load_hourly | 2002-01-01 05:00:00 | 29604.606314 | 28273.426621 | 30935.786006 |
+
+You can plot the forecast by calling the `StatsForecast.plot` method and
+passing in your forecast dataframe.
+
+
+```python
+sf.plot(df, forecasts, max_insample_length=24 * 7)
+```
+
+
+
+## Forecast in production
+
+If you want to gain speed in productive settings where you have multiple
+series or models we recommend using the
+[`StatsForecast.forecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast.forecast)
+method instead of `.fit` and `.predict`.
+
+The main difference is that the `.forecast` doest not store the fitted
+values and is highly scalable in distributed environments.
+
+The `forecast` method takes two arguments: forecasts next `h` (horizon)
+and `level`.
+
+- `h` (int): represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+
+- `level` (list of floats): this optional parameter is used for
+ probabilistic forecasting. Set the `level` (or confidence
+ percentile) of your prediction interval. For example, `level=[90]`
+ means that the model expects the real value to be inside that
+ interval 90% of the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals. Depending on your computer, this step should take
+around 1min. (If you want to speed things up to a couple of seconds,
+remove the AutoModels like ARIMA and Theta)
+
+> **Note**
+>
+> StatsForecast achieves its blazing speed using JIT compiling through
+> Numba. The first time you call the statsforecast class, the fit method
+> should take around 10 seconds. The second time -once Numba compiled
+> your settings- it should take less than 5s.
+
+
+```python
+forecasts_df = sf.forecast(df=df, h=24, level=[90])
+forecasts_df.head()
+```
+
+| | unique_id | ds | MSTL | MSTL-lo-90 | MSTL-hi-90 |
+|----|----|----|----|----|----|
+| 0 | PJM_Load_hourly | 2002-01-01 01:00:00 | 30215.608123 | 29842.185581 | 30589.030664 |
+| 1 | PJM_Load_hourly | 2002-01-01 02:00:00 | 29447.208519 | 28787.122830 | 30107.294207 |
+| 2 | PJM_Load_hourly | 2002-01-01 03:00:00 | 29132.786369 | 28221.353220 | 30044.219518 |
+| 3 | PJM_Load_hourly | 2002-01-01 04:00:00 | 29126.252713 | 27992.819671 | 30259.685756 |
+| 4 | PJM_Load_hourly | 2002-01-01 05:00:00 | 29604.606314 | 28273.426621 | 30935.786006 |
+
+## References
+
+- [Bandara, Kasun & Hyndman, Rob & Bergmeir, Christoph. (2021). “MSTL:
+ A Seasonal-Trend Decomposition Algorithm for Time Series with
+ Multiple Seasonal Patterns”](https://arxiv.org/abs/2107.13462).
+
+## Next Steps
+
+- Learn how to [use cross-validation to assess the robustness of your
+ model](../getting-started/getting_started_complete.html#evaluate-the-model’s-performance).
+
diff --git a/statsforecast/docs/tutorials/statisticalneuralmethods.html.mdx b/statsforecast/docs/tutorials/statisticalneuralmethods.html.mdx
new file mode 100644
index 00000000..e87c7c82
--- /dev/null
+++ b/statsforecast/docs/tutorials/statisticalneuralmethods.html.mdx
@@ -0,0 +1,1052 @@
+---
+description: >-
+ In this notebook, you will make forecasts for the M5 dataset choosing the best
+ model for each time series using cross validation.
+output-file: statisticalneuralmethods.html
+title: Statistical, Machine Learning and Neural Forecasting methods
+---
+
+
+Statistical, Machine Learning, and Neural Forecasting Methods In this
+tutorial, we will explore the process of forecasting on the M5 dataset
+by utilizing the most suitable model for each time series. We’ll
+accomplish this through an essential technique known as
+cross-validation. This approach helps us in estimating the predictive
+performance of our models, and in selecting the model that yields the
+best performance for each time series.
+
+The M5 dataset comprises of hierarchical sales data, spanning five
+years, from Walmart. The aim is to forecast daily sales for the next 28
+days. The dataset is broken down into the 50 states of America, with 10
+stores in each state.
+
+In the realm of time series forecasting and analysis, one of the more
+complex tasks is identifying the model that is optimally suited for a
+specific group of series. Quite often, this selection process leans
+heavily on intuition, which may not necessarily align with the empirical
+reality of our dataset.
+
+In this tutorial, we aim to provide a more structured, data-driven
+approach to model selection for different groups of series within the M5
+benchmark dataset. This dataset, well-known in the field of forecasting,
+allows us to showcase the versatility and power of our methodology.
+
+We will train an assortment of models from various forecasting
+paradigms:
+
+*[StatsForecast](https://github.com/Nixtla/statsforecast)*
+
+- Baseline models: These models are simple yet often highly effective
+ for providing an initial perspective on the forecasting problem. We
+ will use
+ [`SeasonalNaive`](https://Nixtla.github.io/statsforecast/src/core/models.html#seasonalnaive)
+ and
+ [`HistoricAverage`](https://Nixtla.github.io/statsforecast/src/core/models.html#historicaverage)
+ models for this category.
+- Intermittent models: For series with sporadic, non-continuous
+ demand, we will utilize models like
+ [`CrostonOptimized`](https://Nixtla.github.io/statsforecast/src/core/models.html#crostonoptimized),
+ [`IMAPA`](https://Nixtla.github.io/statsforecast/src/core/models.html#imapa),
+ and
+ [`ADIDA`](https://Nixtla.github.io/statsforecast/src/core/models.html#adida).
+ These models are particularly suited for handling zero-inflated
+ series.
+- State Space Models: These are statistical models that use
+ mathematical descriptions of a system to make predictions. The
+ [`AutoETS`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoets)
+ model from the statsforecast library falls under this category.
+
+*[MLForecast](https://github.com/Nixtla/mlforecast)*
+
+Machine Learning: Leveraging ML models like `LightGBM`, `XGBoost`, and
+`LinearRegression` can be advantageous due to their capacity to uncover
+intricate patterns in data. We’ll use the MLForecast library for this
+purpose.
+
+*[NeuralForecast](https://github.com/Nixtla/neuralforecast)*
+
+Deep Learning: DL models, such as Transformers (`AutoTFT`) and Neural
+Networks (`AutoNHITS`), allow us to handle complex non-linear
+dependencies in time series data. We’ll utilize the NeuralForecast
+library for these models.
+
+Using the Nixtla suite of libraries, we’ll be able to drive our model
+selection process with data, ensuring we utilize the most suitable
+models for specific groups of series in our dataset.
+
+Outline:
+
+- Reading Data: In this initial step, we load our dataset into memory,
+ making it available for our subsequent analysis and forecasting. It
+ is important to understand the structure and nuances of the dataset
+ at this stage.
+
+- Forecasting Using Statistical and Deep Learning Methods: We apply a
+ wide range of forecasting methods from basic statistical techniques
+ to advanced deep learning models. The aim is to generate predictions
+ for the next 28 days based on our dataset.
+
+- Model Performance Evaluation on Different Windows: We assess the
+ performance of our models on distinct windows.
+
+- Selecting the Best Model for a Group of Series: Using the
+ performance evaluation, we identify the optimal model for each group
+ of series. This step ensures that the chosen model is tailored to
+ the unique characteristics of each group.
+
+- Filtering the Best Possible Forecast: Finally, we filter the
+ forecasts generated by our chosen models to obtain the most
+ promising predictions. This is our final output and represents the
+ best possible forecast for each series according to our models.
+
+> **Warning**
+>
+> This tutorial was originally executed using a `c5d.24xlarge` EC2
+> instance.
+
+## Installing Libraries
+
+
+```python
+!pip install statsforecast mlforecast neuralforecast datasetforecast s3fs pyarrow
+```
+
+## Download and prepare data
+
+The example uses the [M5
+dataset](https://github.com/Mcompetitions/M5-methods/blob/master/M5-Competitors-Guide.pdf).
+It consists of `30,490` bottom time series.
+
+
+```python
+import pandas as pd
+```
+
+
+```python
+# Load the training target dataset from the provided URL
+Y_df = pd.read_parquet('https://m5-benchmarks.s3.amazonaws.com/data/train/target.parquet')
+
+# Rename columns to match the Nixtlaverse's expectations
+# The 'item_id' becomes 'unique_id' representing the unique identifier of the time series
+# The 'timestamp' becomes 'ds' representing the time stamp of the data points
+# The 'demand' becomes 'y' representing the target variable we want to forecast
+Y_df = Y_df.rename(columns={
+ 'item_id': 'unique_id',
+ 'timestamp': 'ds',
+ 'demand': 'y'
+})
+
+# Convert the 'ds' column to datetime format to ensure proper handling of date-related operations in subsequent steps
+Y_df['ds'] = pd.to_datetime(Y_df['ds'])
+```
+
+For simplicity sake we will keep just one category
+
+
+```python
+Y_df = Y_df.query('unique_id.str.startswith("FOODS_3")').reset_index(drop=True)
+
+Y_df['unique_id'] = Y_df['unique_id'].astype(str)
+```
+
+# Basic Plotting
+
+Plot some series using the plot method from the
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+class. This method prints 8 random series from the dataset and is useful
+for basic
+[EDA](https://nixtla.github.io/statsforecast/src/core/core.html#statsforecast.plot).
+
+
+```python
+from statsforecast import StatsForecast
+```
+
+``` text
+/home/ubuntu/statsforecast/statsforecast/core.py:23: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
+ from tqdm.autonotebook import tqdm
+```
+
+
+```python
+# Feature: plot random series for EDA
+StatsForecast.plot(Y_df)
+```
+
+
+```python
+# Feature: plot groups of series for EDA
+StatsForecast.plot(Y_df, unique_ids=["FOODS_3_432_TX_2"])
+```
+
+
+```python
+# Feature: plot groups of series for EDA
+StatsForecast.plot(Y_df, unique_ids=["FOODS_3_432_TX_2"], engine ='matplotlib')
+```
+
+
+
+# Create forecasts with Stats, Ml and Neural methods.
+
+## StatsForecast
+
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+is a comprehensive library providing a suite of popular univariate time
+series forecasting models, all designed with a focus on high performance
+and scalability.
+
+Here’s what makes StatsForecast a powerful tool for time series
+forecasting:
+
+- **Collection of Local Models**: StatsForecast provides a diverse
+ collection of local models that can be applied to each time series
+ individually, allowing us to capture unique patterns within each
+ series.
+
+- **Simplicity**: With StatsForecast, training, forecasting, and
+ backtesting multiple models become a straightforward process,
+ requiring only a few lines of code. This simplicity makes it a
+ convenient tool for both beginners and experienced practitioners.
+
+- **Optimized for Speed**: The implementation of the models in
+ StatsForecast is optimized for speed, ensuring that large-scale
+ computations are performed efficiently, thereby reducing the overall
+ time for model training and prediction.
+
+- **Horizontal Scalability**: One of the distinguishing features of
+ StatsForecast is its ability to scale horizontally. It is compatible
+ with distributed computing frameworks such as Spark, Dask, and Ray.
+ This feature allows it to handle large datasets by distributing the
+ computations across multiple nodes in a cluster, making it a go-to
+ solution for large-scale time series forecasting tasks.
+
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+receives a list of models to fit each time series. Since we are dealing
+with Daily data, it would be benefitial to use 7 as seasonality.
+
+
+```python
+# Import necessary models from the statsforecast library
+from statsforecast.models import (
+ # SeasonalNaive: A model that uses the previous season's data as the forecast
+ SeasonalNaive,
+ # Naive: A simple model that uses the last observed value as the forecast
+ Naive,
+ # HistoricAverage: This model uses the average of all historical data as the forecast
+ HistoricAverage,
+ # CrostonOptimized: A model specifically designed for intermittent demand forecasting
+ CrostonOptimized,
+ # ADIDA: Adaptive combination of Intermittent Demand Approaches, a model designed for intermittent demand
+ ADIDA,
+ # IMAPA: Intermittent Multiplicative AutoRegressive Average, a model for intermittent series that incorporates autocorrelation
+ IMAPA,
+ # AutoETS: Automated Exponential Smoothing model that automatically selects the best Exponential Smoothing model based on AIC
+ AutoETS
+)
+```
+
+We fit the models by instantiating a new StatsForecast object with the
+following parameters:
+
+- `models`: a list of models. Select the models you want from models
+ and import them.
+- `freq`: a string indicating the frequency of the data. (See panda’s
+ available frequencies.)
+- `n_jobs`: int, number of jobs used in the parallel processing, use
+ -1 for all cores.
+- `fallback_model`: a model to be used if a model fails. Any settings
+ are passed into the constructor. Then you call its fit method and
+ pass in the historical data frame.
+
+
+```python
+horizon = 28
+models = [
+ SeasonalNaive(season_length=7),
+ Naive(),
+ HistoricAverage(),
+ CrostonOptimized(),
+ ADIDA(),
+ IMAPA(),
+ AutoETS(season_length=7)
+]
+```
+
+
+```python
+# Instantiate the StatsForecast class
+sf = StatsForecast(
+ models=models, # A list of models to be used for forecasting
+ freq='D', # The frequency of the time series data (in this case, 'D' stands for daily frequency)
+ n_jobs=-1, # The number of CPU cores to use for parallel execution (-1 means use all available cores)
+)
+```
+
+The forecast method takes two arguments: forecasts next h (horizon) and
+level.
+
+- `h` (int): represents the forecast h steps into the future. In this
+ case, 12 months ahead.
+- `level` (list of floats): this optional parameter is used for
+ probabilistic forecasting. Set the level (or confidence percentile)
+ of your prediction interval. For example, level=\[90\] means that
+ the model expects the real value to be inside that interval 90% of
+ the times.
+
+The forecast object here is a new data frame that includes a column with
+the name of the model and the y hat values, as well as columns for the
+uncertainty intervals.
+
+This block of code times how long it takes to run the forecasting
+function of the StatsForecast class, which predicts the next 28 days
+(h=28). The level is set to \[90\], meaning it will compute the 90%
+prediction interval. The time is calculated in minutes and printed out
+at the end.
+
+
+```python
+from time import time
+
+# Get the current time before forecasting starts, this will be used to measure the execution time
+init = time()
+
+# Call the forecast method of the StatsForecast instance to predict the next 28 days (h=28)
+# Level is set to [90], which means that it will compute the 90% prediction interval
+fcst_df = sf.forecast(df=Y_df, h=28, level=[90])
+
+# Get the current time after the forecasting ends
+end = time()
+
+# Calculate and print the total time taken for the forecasting in minutes
+print(f'Forecast Minutes: {(end - init) / 60}')
+```
+
+``` text
+Forecast Minutes: 2.270755163828532
+```
+
+
+```python
+fcst_df.head()
+```
+
+| | ds | SeasonalNaive | SeasonalNaive-lo-90 | SeasonalNaive-hi-90 | Naive | Naive-lo-90 | Naive-hi-90 | HistoricAverage | HistoricAverage-lo-90 | HistoricAverage-hi-90 | CrostonOptimized | ADIDA | IMAPA | AutoETS | AutoETS-lo-90 | AutoETS-hi-90 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| unique_id | | | | | | | | | | | | | | | | |
+| FOODS_3_001_CA_1 | 2016-05-23 | 1.0 | -2.847174 | 4.847174 | 2.0 | 0.098363 | 3.901637 | 0.448738 | -1.009579 | 1.907055 | 0.345192 | 0.345477 | 0.347249 | 0.381414 | -1.028122 | 1.790950 |
+| FOODS_3_001_CA_1 | 2016-05-24 | 0.0 | -3.847174 | 3.847174 | 2.0 | -0.689321 | 4.689321 | 0.448738 | -1.009579 | 1.907055 | 0.345192 | 0.345477 | 0.347249 | 0.286933 | -1.124136 | 1.698003 |
+| FOODS_3_001_CA_1 | 2016-05-25 | 0.0 | -3.847174 | 3.847174 | 2.0 | -1.293732 | 5.293732 | 0.448738 | -1.009579 | 1.907055 | 0.345192 | 0.345477 | 0.347249 | 0.334987 | -1.077614 | 1.747588 |
+| FOODS_3_001_CA_1 | 2016-05-26 | 1.0 | -2.847174 | 4.847174 | 2.0 | -1.803274 | 5.803274 | 0.448738 | -1.009579 | 1.907055 | 0.345192 | 0.345477 | 0.347249 | 0.186851 | -1.227280 | 1.600982 |
+| FOODS_3_001_CA_1 | 2016-05-27 | 0.0 | -3.847174 | 3.847174 | 2.0 | -2.252190 | 6.252190 | 0.448738 | -1.009579 | 1.907055 | 0.345192 | 0.345477 | 0.347249 | 0.308112 | -1.107548 | 1.723771 |
+
+## MLForecast
+
+`MLForecast` is a powerful library that provides automated feature
+creation for time series forecasting, facilitating the use of global
+machine learning models. It is designed for high performance and
+scalability.
+
+Key features of MLForecast include:
+
+- **Support for sklearn models**: MLForecast is compatible with models
+ that follow the scikit-learn API. This makes it highly flexible and
+ allows it to seamlessly integrate with a wide variety of machine
+ learning algorithms.
+
+- **Simplicity**: With MLForecast, the tasks of training, forecasting,
+ and backtesting models can be accomplished in just a few lines of
+ code. This streamlined simplicity makes it user-friendly for
+ practitioners at all levels of expertise.
+
+- **Optimized for speed:** MLForecast is engineered to execute tasks
+ rapidly, which is crucial when handling large datasets and complex
+ models.
+
+- **Horizontal Scalability:** MLForecast is capable of horizontal
+ scaling using distributed computing frameworks such as Spark, Dask,
+ and Ray. This feature enables it to efficiently process massive
+ datasets by distributing the computations across multiple nodes in a
+ cluster, making it ideal for large-scale time series forecasting
+ tasks.
+
+
+```python
+from mlforecast import MLForecast
+from mlforecast.target_transforms import Differences
+from mlforecast.utils import PredictionIntervals
+from window_ops.expanding import expanding_mean
+```
+
+
+```python
+!pip install lightgbm xgboost
+```
+
+
+```python
+# Import the necessary models from various libraries
+
+# LGBMRegressor: A gradient boosting framework that uses tree-based learning algorithms from the LightGBM library
+from lightgbm import LGBMRegressor
+
+# XGBRegressor: A gradient boosting regressor model from the XGBoost library
+from xgboost import XGBRegressor
+
+# LinearRegression: A simple linear regression model from the scikit-learn library
+from sklearn.linear_model import LinearRegression
+```
+
+To use `MLForecast` for time series forecasting, we instantiate a new
+`MLForecast` object and provide it with various parameters to tailor the
+modeling process to our specific needs:
+
+- `models`: This parameter accepts a list of machine learning models
+ you wish to use for forecasting. You can import your preferred
+ models from scikit-learn, lightgbm and xgboost.
+
+- `freq`: This is a string indicating the frequency of your data
+ (hourly, daily, weekly, etc.). The specific format of this string
+ should align with pandas’ recognized frequency strings.
+
+- `target_transforms`: These are transformations applied to the target
+ variable before model training and after model prediction. This can
+ be useful when working with data that may benefit from
+ transformations, such as log-transforms for highly skewed data.
+
+- `lags`: This parameter accepts specific lag values to be used as
+ regressors. Lags represent how many steps back in time you want to
+ look when creating features for your model. For example, if you want
+ to use the previous day’s data as a feature for predicting today’s
+ value, you would specify a lag of 1.
+
+- `lags_transforms`: These are specific transformations for each lag.
+ This allows you to apply transformations to your lagged features.
+
+- `date_features`: This parameter specifies date-related features to
+ be used as regressors. For instance, you might want to include the
+ day of the week or the month as a feature in your model.
+
+- `num_threads`: This parameter controls the number of threads to use
+ for parallelizing feature creation, helping to speed up this process
+ when working with large datasets.
+
+All these settings are passed to the `MLForecast` constructor. Once the
+`MLForecast` object is initialized with these settings, we call its
+`fit` method and pass the historical data frame as the argument. The
+`fit` method trains the models on the provided historical data, readying
+them for future forecasting tasks.
+
+
+```python
+# Instantiate the MLForecast object
+mlf = MLForecast(
+ models=[LGBMRegressor(), XGBRegressor(), LinearRegression()], # List of models for forecasting: LightGBM, XGBoost and Linear Regression
+ freq='D', # Frequency of the data - 'D' for daily frequency
+ lags=list(range(1, 7)), # Specific lags to use as regressors: 1 to 6 days
+ lag_transforms = {
+ 1: [expanding_mean], # Apply expanding mean transformation to the lag of 1 day
+ },
+ date_features=['year', 'month', 'day', 'dayofweek', 'quarter', 'week'], # Date features to use as regressors
+)
+```
+
+Just call the `fit` models to train the select models. In this case we
+are generating conformal prediction intervals.
+
+
+```python
+# Start the timer to calculate the time taken for fitting the models
+init = time()
+
+# Fit the MLForecast models to the data, with prediction intervals set using a window size of 28 days
+mlf.fit(Y_df, prediction_intervals=PredictionIntervals(window_size=28))
+
+# Calculate the end time after fitting the models
+end = time()
+
+# Print the time taken to fit the MLForecast models, in minutes
+print(f'MLForecast Minutes: {(end - init) / 60}')
+```
+
+``` text
+MLForecast Minutes: 2.2809854547182717
+```
+
+After that, just call `predict` to generate forecasts.
+
+
+```python
+fcst_mlf_df = mlf.predict(28, level=[90])
+```
+
+
+```python
+fcst_mlf_df.head()
+```
+
+| | unique_id | ds | LGBMRegressor | XGBRegressor | LinearRegression | LGBMRegressor-lo-90 | LGBMRegressor-hi-90 | XGBRegressor-lo-90 | XGBRegressor-hi-90 | LinearRegression-lo-90 | LinearRegression-hi-90 |
+|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | FOODS_3_001_CA_1 | 2016-05-23 | 0.549520 | 0.598431 | 0.359638 | -0.213915 | 1.312955 | -0.020050 | 1.216912 | 0.030000 | 0.689277 |
+| 1 | FOODS_3_001_CA_1 | 2016-05-24 | 0.553196 | 0.337268 | 0.100361 | -0.251383 | 1.357775 | -0.201449 | 0.875985 | -0.216195 | 0.416917 |
+| 2 | FOODS_3_001_CA_1 | 2016-05-25 | 0.599668 | 0.349604 | 0.175840 | -0.203974 | 1.403309 | -0.284416 | 0.983624 | -0.150593 | 0.502273 |
+| 3 | FOODS_3_001_CA_1 | 2016-05-26 | 0.638097 | 0.322144 | 0.156460 | 0.118688 | 1.157506 | -0.085872 | 0.730160 | -0.273851 | 0.586771 |
+| 4 | FOODS_3_001_CA_1 | 2016-05-27 | 0.763305 | 0.300362 | 0.328194 | -0.313091 | 1.839701 | -0.296636 | 0.897360 | -0.657089 | 1.313476 |
+
+## NeuralForecast
+
+`NeuralForecast` is a robust collection of neural forecasting models
+that focuses on usability and performance. It includes a variety of
+model architectures, from classic networks such as Multilayer
+Perceptrons (MLP) and Recurrent Neural Networks (RNN) to novel
+contributions like N-BEATS, N-HITS, Temporal Fusion Transformers (TFT),
+and more.
+
+Key features of `NeuralForecast` include:
+
+- A broad collection of global models. Out of the box implementation
+ of MLP, LSTM, RNN, TCN, DilatedRNN, NBEATS, NHITS, ESRNN, TFT,
+ Informer, PatchTST and HINT.
+- A simple and intuitive interface that allows training, forecasting,
+ and backtesting of various models in a few lines of code.
+- Support for GPU acceleration to improve computational speed.
+
+This machine doesn’t have GPU, but Google Colabs offers some for free.
+
+Using [Colab’s GPU to train
+NeuralForecast](https://nixtla.github.io/neuralforecast/docs/tutorials/intermittent_data.html).
+
+
+```python
+# Read the results from Colab
+fcst_nf_df = pd.read_parquet('https://m5-benchmarks.s3.amazonaws.com/data/forecast-nf.parquet')
+```
+
+
+```python
+fcst_nf_df.head()
+```
+
+| | unique_id | ds | AutoNHITS | AutoNHITS-lo-90 | AutoNHITS-hi-90 | AutoTFT | AutoTFT-lo-90 | AutoTFT-hi-90 |
+|----|----|----|----|----|----|----|----|----|
+| 0 | FOODS_3_001_CA_1 | 2016-05-23 | 0.0 | 0.0 | 2.0 | 0.0 | 0.0 | 2.0 |
+| 1 | FOODS_3_001_CA_1 | 2016-05-24 | 0.0 | 0.0 | 2.0 | 0.0 | 0.0 | 2.0 |
+| 2 | FOODS_3_001_CA_1 | 2016-05-25 | 0.0 | 0.0 | 2.0 | 0.0 | 0.0 | 1.0 |
+| 3 | FOODS_3_001_CA_1 | 2016-05-26 | 0.0 | 0.0 | 2.0 | 0.0 | 0.0 | 2.0 |
+| 4 | FOODS_3_001_CA_1 | 2016-05-27 | 0.0 | 0.0 | 2.0 | 0.0 | 0.0 | 2.0 |
+
+
+```python
+# Merge the forecasts from StatsForecast and NeuralForecast
+fcst_df = fcst_df.merge(fcst_nf_df, how='left', on=['unique_id', 'ds'])
+
+# Merge the forecasts from MLForecast into the combined forecast dataframe
+fcst_df = fcst_df.merge(fcst_mlf_df, how='left', on=['unique_id', 'ds'])
+```
+
+
+```python
+fcst_df.head()
+```
+
+| | unique_id | ds | SeasonalNaive | SeasonalNaive-lo-90 | SeasonalNaive-hi-90 | Naive | Naive-lo-90 | Naive-hi-90 | HistoricAverage | HistoricAverage-lo-90 | ... | AutoTFT-hi-90 | LGBMRegressor | XGBRegressor | LinearRegression | LGBMRegressor-lo-90 | LGBMRegressor-hi-90 | XGBRegressor-lo-90 | XGBRegressor-hi-90 | LinearRegression-lo-90 | LinearRegression-hi-90 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | FOODS_3_001_CA_1 | 2016-05-23 | 1.0 | -2.847174 | 4.847174 | 2.0 | 0.098363 | 3.901637 | 0.448738 | -1.009579 | ... | 2.0 | 0.549520 | 0.598431 | 0.359638 | -0.213915 | 1.312955 | -0.020050 | 1.216912 | 0.030000 | 0.689277 |
+| 1 | FOODS_3_001_CA_1 | 2016-05-24 | 0.0 | -3.847174 | 3.847174 | 2.0 | -0.689321 | 4.689321 | 0.448738 | -1.009579 | ... | 2.0 | 0.553196 | 0.337268 | 0.100361 | -0.251383 | 1.357775 | -0.201449 | 0.875985 | -0.216195 | 0.416917 |
+| 2 | FOODS_3_001_CA_1 | 2016-05-25 | 0.0 | -3.847174 | 3.847174 | 2.0 | -1.293732 | 5.293732 | 0.448738 | -1.009579 | ... | 1.0 | 0.599668 | 0.349604 | 0.175840 | -0.203974 | 1.403309 | -0.284416 | 0.983624 | -0.150593 | 0.502273 |
+| 3 | FOODS_3_001_CA_1 | 2016-05-26 | 1.0 | -2.847174 | 4.847174 | 2.0 | -1.803274 | 5.803274 | 0.448738 | -1.009579 | ... | 2.0 | 0.638097 | 0.322144 | 0.156460 | 0.118688 | 1.157506 | -0.085872 | 0.730160 | -0.273851 | 0.586771 |
+| 4 | FOODS_3_001_CA_1 | 2016-05-27 | 0.0 | -3.847174 | 3.847174 | 2.0 | -2.252190 | 6.252190 | 0.448738 | -1.009579 | ... | 2.0 | 0.763305 | 0.300362 | 0.328194 | -0.313091 | 1.839701 | -0.296636 | 0.897360 | -0.657089 | 1.313476 |
+
+## Forecast plots
+
+
+```python
+sf.plot(Y_df, fcst_df, max_insample_length=28 * 3)
+```
+
+Use the plot function to explore models and ID’s
+
+
+```python
+sf.plot(Y_df, fcst_df, max_insample_length=28 * 3,
+ models=['CrostonOptimized', 'AutoNHITS', 'SeasonalNaive', 'LGBMRegressor'])
+```
+
+# Validate Model’s Performance
+
+The three libraries -
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast),
+`MLForecast`, and `NeuralForecast` - offer out-of-the-box
+cross-validation capabilities specifically designed for time series.
+This allows us to evaluate the model’s performance using historical data
+to obtain an unbiased assessment of how well each model is likely to
+perform on unseen data.
+
+
+
+From the course of Modern Forecasting in
+Practice
+
+
+## Cross Validation in StatsForecast
+
+The
+[`cross_validation`](https://Nixtla.github.io/statsforecast/src/mfles.html#cross_validation)
+method from the
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+class accepts the following arguments:
+
+- `df`: A DataFrame representing the training data.
+- `h` (int): The forecast horizon, represented as the number of steps
+ into the future that we wish to predict. For example, if we’re
+ forecasting hourly data, `h=24` would represent a 24-hour forecast.
+- `step_size` (int): The step size between each cross-validation
+ window. This parameter determines how often we want to run the
+ forecasting process.
+- `n_windows` (int): The number of windows used for cross validation.
+ This parameter defines how many past forecasting processes we want
+ to evaluate.
+
+These parameters allow us to control the extent and granularity of our
+cross-validation process. By tuning these settings, we can balance
+between computational cost and the thoroughness of the cross-validation.
+
+
+```python
+init = time()
+cv_df = sf.cross_validation(df=Y_df, h=horizon, n_windows=3, step_size=horizon, level=[90])
+end = time()
+print(f'CV Minutes: {(end - init) / 60}')
+```
+
+``` text
+/home/ubuntu/statsforecast/statsforecast/ets.py:1041: RuntimeWarning:
+
+divide by zero encountered in double_scalars
+```
+
+``` text
+CV Minutes: 5.206169327100118
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id` index: (If you dont like working with index just run
+ forecasts_cv_df.resetindex())
+- `ds`: datestamp or temporal index
+- `cutoff`: the last datestamp or temporal index for the n_windows. If
+ n_windows=1, then one unique cuttoff value, if n_windows=2 then two
+ unique cutoff values.
+- `y`: true value
+- `"model"`: columns with the model’s name and fitted value.
+
+
+```python
+cv_df.head()
+```
+
+| | ds | cutoff | y | SeasonalNaive | SeasonalNaive-lo-90 | SeasonalNaive-hi-90 | Naive | Naive-lo-90 | Naive-hi-90 | HistoricAverage | HistoricAverage-lo-90 | HistoricAverage-hi-90 | CrostonOptimized | ADIDA | IMAPA | AutoETS | AutoETS-lo-90 | AutoETS-hi-90 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| unique_id | | | | | | | | | | | | | | | | | | |
+| FOODS_3_001_CA_1 | 2016-02-29 | 2016-02-28 | 0.0 | 2.0 | -1.878885 | 5.878885 | 0.0 | -1.917011 | 1.917011 | 0.449111 | -1.021813 | 1.920036 | 0.618472 | 0.618375 | 0.617998 | 0.655286 | -0.765731 | 2.076302 |
+| FOODS_3_001_CA_1 | 2016-03-01 | 2016-02-28 | 1.0 | 0.0 | -3.878885 | 3.878885 | 0.0 | -2.711064 | 2.711064 | 0.449111 | -1.021813 | 1.920036 | 0.618472 | 0.618375 | 0.617998 | 0.568595 | -0.853966 | 1.991155 |
+| FOODS_3_001_CA_1 | 2016-03-02 | 2016-02-28 | 1.0 | 0.0 | -3.878885 | 3.878885 | 0.0 | -3.320361 | 3.320361 | 0.449111 | -1.021813 | 1.920036 | 0.618472 | 0.618375 | 0.617998 | 0.618805 | -0.805298 | 2.042908 |
+| FOODS_3_001_CA_1 | 2016-03-03 | 2016-02-28 | 0.0 | 1.0 | -2.878885 | 4.878885 | 0.0 | -3.834023 | 3.834023 | 0.449111 | -1.021813 | 1.920036 | 0.618472 | 0.618375 | 0.617998 | 0.455891 | -0.969753 | 1.881534 |
+| FOODS_3_001_CA_1 | 2016-03-04 | 2016-02-28 | 0.0 | 1.0 | -2.878885 | 4.878885 | 0.0 | -4.286568 | 4.286568 | 0.449111 | -1.021813 | 1.920036 | 0.618472 | 0.618375 | 0.617998 | 0.591197 | -0.835987 | 2.018380 |
+
+## MLForecast
+
+The
+[`cross_validation`](https://Nixtla.github.io/statsforecast/src/mfles.html#cross_validation)
+method from the `MLForecast` class takes the following arguments.
+
+- `data`: training data frame
+- `window_size` (int): represents h steps into the future that are
+ being forecasted. In this case, 24 hours ahead.
+- `step_size` (int): step size between each window. In other words:
+ how often do you want to run the forecasting processes.
+- `n_windows` (int): number of windows used for cross-validation. In
+ other words: what number of forecasting processes in the past do you
+ want to evaluate.
+- `prediction_intervals`: class to compute conformal intervals.
+
+
+```python
+init = time()
+cv_mlf_df = mlf.cross_validation(
+ data=Y_df,
+ window_size=horizon,
+ n_windows=3,
+ step_size=horizon,
+ level=[90],
+)
+end = time()
+print(f'CV Minutes: {(end - init) / 60}')
+```
+
+``` text
+/home/ubuntu/miniconda/envs/statsforecast/lib/python3.10/site-packages/mlforecast/forecast.py:576: UserWarning:
+
+Excuting `cross_validation` after `fit` can produce unexpected errors
+
+/home/ubuntu/miniconda/envs/statsforecast/lib/python3.10/site-packages/mlforecast/forecast.py:468: UserWarning:
+
+Please rerun the `fit` method passing a proper value to prediction intervals to compute them.
+
+/home/ubuntu/miniconda/envs/statsforecast/lib/python3.10/site-packages/mlforecast/forecast.py:468: UserWarning:
+
+Please rerun the `fit` method passing a proper value to prediction intervals to compute them.
+
+/home/ubuntu/miniconda/envs/statsforecast/lib/python3.10/site-packages/mlforecast/forecast.py:468: UserWarning:
+
+Please rerun the `fit` method passing a proper value to prediction intervals to compute them.
+```
+
+``` text
+CV Minutes: 2.961174162228902
+```
+
+The crossvaldation_df object is a new data frame that includes the
+following columns:
+
+- `unique_id` index: (If you dont like working with index just run
+ forecasts_cv_df.resetindex())
+- `ds`: datestamp or temporal index
+- `cutoff`: the last datestamp or temporal index for the n_windows. If
+ n_windows=1, then one unique cuttoff value, if n_windows=2 then two
+ unique cutoff values.
+- `y`: true value
+- `"model"`: columns with the model’s name and fitted value.
+
+
+```python
+cv_mlf_df.head()
+```
+
+| | unique_id | ds | cutoff | y | LGBMRegressor | XGBRegressor | LinearRegression |
+|----|----|----|----|----|----|----|----|
+| 0 | FOODS_3_001_CA_1 | 2016-02-29 | 2016-02-28 | 0.0 | 0.435674 | 0.556261 | -0.312492 |
+| 1 | FOODS_3_001_CA_1 | 2016-03-01 | 2016-02-28 | 1.0 | 0.639676 | 0.625806 | -0.041924 |
+| 2 | FOODS_3_001_CA_1 | 2016-03-02 | 2016-02-28 | 1.0 | 0.792989 | 0.659650 | 0.263699 |
+| 3 | FOODS_3_001_CA_1 | 2016-03-03 | 2016-02-28 | 0.0 | 0.806868 | 0.535121 | 0.482491 |
+| 4 | FOODS_3_001_CA_1 | 2016-03-04 | 2016-02-28 | 0.0 | 0.829106 | 0.313353 | 0.677326 |
+
+## NeuralForecast
+
+This machine doesn’t have GPU, but Google Colabs offers some for free.
+
+Using [Colab’s GPU to train
+NeuralForecast](https://nixtla.github.io/neuralforecast/docs/tutorials/intermittent_data.html).
+
+
+```python
+cv_nf_df = pd.read_parquet('https://m5-benchmarks.s3.amazonaws.com/data/cross-validation-nf.parquet')
+```
+
+
+```python
+cv_nf_df.head()
+```
+
+| | unique_id | ds | cutoff | AutoNHITS | AutoNHITS-lo-90 | AutoNHITS-hi-90 | AutoTFT | AutoTFT-lo-90 | AutoTFT-hi-90 | y |
+|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | FOODS_3_001_CA_1 | 2016-02-29 | 2016-02-28 | 0.0 | 0.0 | 2.0 | 1.0 | 0.0 | 2.0 | 0.0 |
+| 1 | FOODS_3_001_CA_1 | 2016-03-01 | 2016-02-28 | 0.0 | 0.0 | 2.0 | 1.0 | 0.0 | 2.0 | 1.0 |
+| 2 | FOODS_3_001_CA_1 | 2016-03-02 | 2016-02-28 | 0.0 | 0.0 | 2.0 | 1.0 | 0.0 | 2.0 | 1.0 |
+| 3 | FOODS_3_001_CA_1 | 2016-03-03 | 2016-02-28 | 0.0 | 0.0 | 2.0 | 1.0 | 0.0 | 2.0 | 0.0 |
+| 4 | FOODS_3_001_CA_1 | 2016-03-04 | 2016-02-28 | 0.0 | 0.0 | 2.0 | 1.0 | 0.0 | 2.0 | 0.0 |
+
+## Merge cross validation forecasts
+
+
+```python
+cv_df = cv_df.merge(cv_nf_df.drop(columns=['y']), how='left', on=['unique_id', 'ds', 'cutoff'])
+cv_df = cv_df.merge(cv_mlf_df.drop(columns=['y']), how='left', on=['unique_id', 'ds', 'cutoff'])
+```
+
+## Plots CV
+
+
+```python
+cutoffs = cv_df['cutoff'].unique()
+```
+
+
+```python
+for cutoff in cutoffs:
+ img = sf.plot(
+ Y_df,
+ cv_df.query('cutoff == @cutoff').drop(columns=['y', 'cutoff']),
+ max_insample_length=28 * 5,
+ unique_ids=['FOODS_3_001_CA_1'],
+ )
+ img.show()
+```
+
+### Aggregate Demand
+
+
+```python
+agg_cv_df = cv_df.loc[:,~cv_df.columns.str.contains('hi|lo')].groupby(['ds', 'cutoff']).sum(numeric_only=True).reset_index()
+agg_cv_df.insert(0, 'unique_id', 'agg_demand')
+```
+
+
+```python
+agg_Y_df = Y_df.groupby(['ds']).sum(numeric_only=True).reset_index()
+agg_Y_df.insert(0, 'unique_id', 'agg_demand')
+```
+
+
+```python
+for cutoff in cutoffs:
+ img = sf.plot(
+ agg_Y_df,
+ agg_cv_df.query('cutoff == @cutoff').drop(columns=['y', 'cutoff']),
+ max_insample_length=28 * 5,
+ )
+ img.show()
+```
+
+## Evaluation per series and CV window
+
+In this section, we will evaluate the performance of each model for each
+time series and each cross validation window. Since we have many
+combinations, we will use `dask` to parallelize the evaluation. The
+parallelization will be done using `fugue`.
+
+
+```python
+from typing import List, Callable
+
+from distributed import Client
+from fugue import transform
+from fugue_dask import DaskExecutionEngine
+from datasetsforecast.losses import mse, mae, smape
+```
+
+The `evaluate` function receives a unique combination of a time series
+and a window, and calculates different `metrics` for each model in `df`.
+
+
+```python
+def evaluate(df: pd.DataFrame, metrics: List[Callable]) -> pd.DataFrame:
+ eval_ = {}
+ models = df.loc[:, ~df.columns.str.contains('unique_id|y|ds|cutoff|lo|hi')].columns
+ for model in models:
+ eval_[model] = {}
+ for metric in metrics:
+ eval_[model][metric.__name__] = metric(df['y'], df[model])
+ eval_df = pd.DataFrame(eval_).rename_axis('metric').reset_index()
+ eval_df.insert(0, 'cutoff', df['cutoff'].iloc[0])
+ eval_df.insert(0, 'unique_id', df['unique_id'].iloc[0])
+ return eval_df
+```
+
+
+```python
+str_models = cv_df.loc[:, ~cv_df.columns.str.contains('unique_id|y|ds|cutoff|lo|hi')].columns
+str_models = ','.join([f"{model}:float" for model in str_models])
+cv_df['cutoff'] = cv_df['cutoff'].astype(str)
+cv_df['unique_id'] = cv_df['unique_id'].astype(str)
+```
+
+Let’s cleate a `dask` client.
+
+
+```python
+client = Client() # without this, dask is not in distributed mode
+# fugue.dask.dataframe.default.partitions determines the default partitions for a new DaskDataFrame
+engine = DaskExecutionEngine({"fugue.dask.dataframe.default.partitions": 96})
+```
+
+The `transform` function takes the `evaluate` functions and applies it
+to each combination of time series (`unique_id`) and cross validation
+window (`cutoff`) using the `dask` client we created before.
+
+
+```python
+evaluation_df = transform(
+ cv_df.loc[:, ~cv_df.columns.str.contains('lo|hi')],
+ evaluate,
+ engine="dask",
+ params={'metrics': [mse, mae, smape]},
+ schema=f"unique_id:str,cutoff:str,metric:str, {str_models}",
+ as_local=True,
+ partition={'by': ['unique_id', 'cutoff']}
+)
+```
+
+``` text
+/home/ubuntu/miniconda/envs/statsforecast/lib/python3.10/site-packages/distributed/client.py:3109: UserWarning:
+
+Sending large graph of size 49.63 MiB.
+This may cause some slowdown.
+Consider scattering data ahead of time and using futures.
+```
+
+
+```python
+evaluation_df.head()
+```
+
+| | unique_id | cutoff | metric | SeasonalNaive | Naive | HistoricAverage | CrostonOptimized | ADIDA | IMAPA | AutoETS | AutoNHITS | AutoTFT | LGBMRegressor | XGBRegressor | LinearRegression |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | FOODS_3_003_WI_3 | 2016-02-28 | mse | 1.142857 | 1.142857 | 0.816646 | 0.816471 | 1.142857 | 1.142857 | 1.142857 | 1.142857 | 1.142857 | 0.832010 | 1.020361 | 0.887121 |
+| 1 | FOODS_3_003_WI_3 | 2016-02-28 | mae | 0.571429 | 0.571429 | 0.729592 | 0.731261 | 0.571429 | 0.571429 | 0.571429 | 0.571429 | 0.571429 | 0.772788 | 0.619949 | 0.685413 |
+| 2 | FOODS_3_003_WI_3 | 2016-02-28 | smape | 71.428574 | 71.428574 | 158.813507 | 158.516235 | 200.000000 | 200.000000 | 200.000000 | 71.428574 | 71.428574 | 145.901947 | 188.159164 | 178.883743 |
+| 3 | FOODS_3_013_CA_3 | 2016-04-24 | mse | 4.000000 | 6.214286 | 2.406764 | 3.561202 | 2.267853 | 2.267600 | 2.268677 | 2.750000 | 2.125000 | 2.160508 | 2.370228 | 2.289606 |
+| 4 | FOODS_3_013_CA_3 | 2016-04-24 | mae | 1.500000 | 2.142857 | 1.214286 | 1.340446 | 1.214286 | 1.214286 | 1.214286 | 1.107143 | 1.142857 | 1.140084 | 1.157548 | 1.148813 |
+
+
+```python
+# Calculate the mean metric for each cross validation window
+evaluation_df.groupby(['cutoff', 'metric']).mean(numeric_only=True)
+```
+
+| | | SeasonalNaive | Naive | HistoricAverage | CrostonOptimized | ADIDA | IMAPA | AutoETS | AutoNHITS | AutoTFT | LGBMRegressor | XGBRegressor | LinearRegression |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| cutoff | metric | | | | | | | | | | | | |
+| 2016-02-28 | mae | 1.744289 | 2.040496 | 1.730704 | 1.633017 | 1.527965 | 1.528772 | 1.497553 | 1.434938 | 1.485419 | 1.688403 | 1.514102 | 1.576320 |
+| | mse | 14.510710 | 19.080585 | 12.858994 | 11.785032 | 11.114497 | 11.100909 | 10.347847 | 10.010982 | 10.964664 | 10.436206 | 10.968788 | 10.792831 |
+| | smape | 85.202042 | 87.719086 | 125.418488 | 124.749908 | 127.591858 | 127.704102 | 127.790672 | 79.132614 | 80.983368 | 118.489983 | 140.420578 | 127.043137 |
+| 2016-03-27 | mae | 1.795973 | 2.106449 | 1.754029 | 1.662087 | 1.570701 | 1.572741 | 1.535301 | 1.432412 | 1.502393 | 1.712493 | 1.600193 | 1.601612 |
+| | mse | 14.810259 | 26.044472 | 12.804104 | 12.020620 | 12.083861 | 12.120033 | 11.315013 | 9.445867 | 10.762877 | 10.723589 | 12.924312 | 10.943772 |
+| | smape | 87.407471 | 89.453247 | 123.587196 | 123.460030 | 123.428459 | 123.538521 | 123.612991 | 79.926781 | 82.013168 | 116.089699 | 138.885941 | 127.304871 |
+| 2016-04-24 | mae | 1.785983 | 1.990774 | 1.762506 | 1.609268 | 1.527627 | 1.529721 | 1.501820 | 1.447401 | 1.505127 | 1.692946 | 1.541845 | 1.590985 |
+| | mse | 13.476350 | 16.234917 | 13.151311 | 10.647048 | 10.072225 | 10.062395 | 9.393439 | 9.363891 | 10.436214 | 10.347073 | 10.774202 | 10.608137 |
+| | smape | 89.238815 | 90.685867 | 121.124947 | 119.721245 | 120.325401 | 120.345284 | 120.649582 | 81.402748 | 83.614029 | 113.334198 | 136.755234 | 124.618622 |
+
+Results showed in previous experiments.
+
+| model | MSE |
+|:-----------------|------:|
+| MQCNN | 10.09 |
+| DeepAR-student_t | 10.11 |
+| DeepAR-lognormal | 30.20 |
+| DeepAR | 9.13 |
+| NPTS | 11.53 |
+
+Top 3 models: DeepAR, AutoNHITS, AutoETS.
+
+### Distribution of errors
+
+
+```python
+!pip install seaborn
+```
+
+
+```python
+import matplotlib.pyplot as plt
+import seaborn as sns
+```
+
+
+```python
+evaluation_df_melted = pd.melt(evaluation_df, id_vars=['unique_id', 'cutoff', 'metric'], var_name='model', value_name='error')
+```
+
+#### SMAPE
+
+
+```python
+sns.violinplot(evaluation_df_melted.query('metric=="smape"'), x='error', y='model')
+```
+
+
+
+### Choose models for groups of series
+
+Feature:
+
+- A unified dataframe with forecasts for all different models
+- Easy Ensamble
+- E.g. Average predictions
+- Or MinMax (Choosing is ensembling)
+
+
+```python
+# Choose the best model for each time series, metric, and cross validation window
+evaluation_df['best_model'] = evaluation_df.idxmin(axis=1, numeric_only=True)
+# count how many times a model wins per metric and cross validation window
+count_best_model = evaluation_df.groupby(['cutoff', 'metric', 'best_model']).size().rename('n').to_frame().reset_index()
+# plot results
+sns.barplot(count_best_model, x='n', y='best_model', hue='metric')
+```
+
+
+
+### Et pluribus unum: an inclusive forecasting Pie.
+
+
+```python
+# For the mse, calculate how many times a model wins
+eval_series_df = evaluation_df.query('metric == "mse"').groupby(['unique_id']).mean(numeric_only=True)
+eval_series_df['best_model'] = eval_series_df.idxmin(axis=1)
+counts_series = eval_series_df.value_counts('best_model')
+plt.pie(counts_series, labels=counts_series.index, autopct='%.0f%%')
+plt.show()
+```
+
+
+
+
+```python
+sf.plot(Y_df, cv_df.drop(columns=['cutoff', 'y']),
+ max_insample_length=28 * 6,
+ models=['AutoNHITS'],
+ unique_ids=eval_series_df.query('best_model == "AutoNHITS"').index[:8])
+```
+
+# Choose Forecasting method for different groups of series
+
+
+```python
+# Merge the best model per time series dataframe
+# and filter the forecasts based on that dataframe
+# for each time series
+fcst_df = pd.melt(fcst_df.set_index('unique_id'), id_vars=['ds'], var_name='model', value_name='forecast', ignore_index=False)
+fcst_df = fcst_df.join(eval_series_df[['best_model']])
+fcst_df[['model', 'pred-interval']] = fcst_df['model'].str.split('-', expand=True, n=1)
+fcst_df = fcst_df.query('model == best_model')
+fcst_df['name'] = [f'forecast-{x}' if x is not None else 'forecast' for x in fcst_df['pred-interval']]
+fcst_df = pd.pivot_table(fcst_df, index=['unique_id', 'ds'], values=['forecast'], columns=['name']).droplevel(0, axis=1).reset_index()
+```
+
+
+```python
+sf.plot(Y_df, fcst_df, max_insample_length=28 * 3)
+```
+
+# Technical Debt
+
+- Train the statistical models in the full dataset.
+- Increase the number of `num_samples` in the neural auto models.
+- Include other models such as
+ [`Theta`](https://Nixtla.github.io/statsforecast/src/core/models.html#theta),
+ [`ARIMA`](https://Nixtla.github.io/statsforecast/src/core/models.html#arima),
+ `RNN`, `LSTM`, …
+
+# Further materials
+
+- [Available Models
+ StatsForecast](https://nixtlaverse.nixtla.io/statsforecast/src/core/models_intro)
+- [Available Models
+ NeuralForecast](https://nixtla.github.io/neuralforecast/models.html)
+- [Scalers and Loss
+ Functions](https://nixtla.github.io/neuralforecast/losses.pytorch.html)
+- [Getting Started
+ NeuralForecast](https://nixtlaverse.nixtla.io/neuralforecast/docs/getting-started/quickstart.html)
+- [Hierarchical
+ Reconciliation](https://nixtla.github.io/hierarchicalforecast/examples/tourismsmall.html)
+- [Distributed ML Forecast
+ (trees)](https://nixtla.github.io/mlforecast/docs/getting-started/quick_start_distributed.html)
+- [Using StatsForecast to train millions of time
+ series](https://www.anyscale.com/blog/how-nixtla-uses-ray-to-accurately-predict-more-than-a-million-time-series)
+- [Intermittent Demand Forecasting With Nixtla on
+ Databricks](https://www.databricks.com/blog/2022/12/06/intermittent-demand-forecasting-nixtla-databricks.html)
+
diff --git a/statsforecast/docs/tutorials/uncertaintyintervals.html.mdx b/statsforecast/docs/tutorials/uncertaintyintervals.html.mdx
new file mode 100644
index 00000000..32d18d6f
--- /dev/null
+++ b/statsforecast/docs/tutorials/uncertaintyintervals.html.mdx
@@ -0,0 +1,320 @@
+---
+description: In this example, we'll implement prediction intervals
+output-file: uncertaintyintervals.html
+title: Probabilistic Forecasting
+---
+
+
+> **Prerequisites**
+>
+> This tutorial assumes basic familiarity with StatsForecast. For a
+> minimal example visit the [Quick
+> Start](../getting-started/getting_started_short.html)
+
+## Introduction
+
+When we generate a forecast, we usually produce a single value known as
+the point forecast. This value, however, doesn’t tell us anything about
+the uncertainty associated with the forecast. To have a measure of this
+uncertainty, we need **prediction intervals**.
+
+A prediction interval is a range of values that the forecast can take
+with a given probability. Hence, a 95% prediction interval should
+contain a range of values that include the actual future value with
+probability 95%. Probabilistic forecasting aims to generate the full
+forecast distribution. Point forecasting, on the other hand, usually
+returns the mean or the median or said distribution. However, in
+real-world scenarios, it is better to forecast not only the most
+probable future outcome, but many alternative outcomes as well.
+
+[StatsForecast](https://nixtla.github.io/statsforecast/) has many models
+that can generate point forecasts. It also has probabilistic models than
+generate the same point forecasts and their prediction intervals. These
+models are stochastic data generating processes that can produce entire
+forecast distributions. By the end of this tutorial, you’ll have a good
+understanding of the probabilistic models available in StatsForecast and
+will be able to use them to generate point forecasts and prediction
+intervals. Furthermore, you’ll also learn how to generate plots with the
+historical data, the point forecasts, and the prediction intervals.
+
+> **Important**
+>
+> Although the terms are often confused, prediction intervals are not
+> the same as [confidence
+> intervals](https://robjhyndman.com/hyndsight/intervals/).
+
+> **Warning**
+>
+> In practice, most prediction intervals are too narrow since models do
+> not account for all sources of uncertainty. A discussion about this
+> can be found [here](https://robjhyndman.com/hyndsight/narrow-pi/).
+
+**Outline:**
+
+1. Install libraries
+2. Load and explore the data
+3. Train models
+4. Plot prediction intervals
+
+> **Tip**
+>
+> You can use Colab to run this Notebook interactively
+>
+
+## Install libraries
+
+We assume that you have StatsForecast already installed. If not, check
+this guide for instructions on [how to install
+StatsForecast](../getting-started/installation.html)
+
+Install the necessary packages using `pip install statsforecast`
+
+## Load and explore the data
+
+For this example, we’ll use the hourly dataset from the [M4
+Competition](https://www.sciencedirect.com/science/article/pii/S0169207019301128).
+We first need to download the data from a URL and then load it as a
+`pandas` dataframe. Notice that we’ll load the train and the test data
+separately. We’ll also rename the `y` column of the test data as
+`y_test`.
+
+
+```python
+import pandas as pd
+```
+
+
+```python
+train = pd.read_csv('https://auto-arima-results.s3.amazonaws.com/M4-Hourly.csv')
+test = pd.read_csv('https://auto-arima-results.s3.amazonaws.com/M4-Hourly-test.csv').rename(columns={'y': 'y_test'})
+```
+
+
+```python
+train.head()
+```
+
+| | unique_id | ds | y |
+|-----|-----------|-----|-------|
+| 0 | H1 | 1 | 605.0 |
+| 1 | H1 | 2 | 586.0 |
+| 2 | H1 | 3 | 586.0 |
+| 3 | H1 | 4 | 559.0 |
+| 4 | H1 | 5 | 511.0 |
+
+
+```python
+test.head()
+```
+
+| | unique_id | ds | y_test |
+|-----|-----------|-----|--------|
+| 0 | H1 | 701 | 619.0 |
+| 1 | H1 | 702 | 565.0 |
+| 2 | H1 | 703 | 532.0 |
+| 3 | H1 | 704 | 495.0 |
+| 4 | H1 | 705 | 481.0 |
+
+Since the goal of this notebook is to generate prediction intervals,
+we’ll only use the first 8 series of the dataset to reduce the total
+computational time.
+
+
+```python
+n_series = 8
+uids = train['unique_id'].unique()[:n_series] # select first n_series of the dataset
+train = train.query('unique_id in @uids')
+test = test.query('unique_id in @uids')
+```
+
+We can plot these series using the `statsforecast.plot` method from the
+[StatsForecast](https://nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+class. This method has multiple parameters, and the required ones to
+generate the plots in this notebook are explained below.
+
+- `df`: A `pandas` dataframe with columns \[`unique_id`, `ds`, `y`\].
+- `forecasts_df`: A `pandas` dataframe with columns \[`unique_id`,
+ `ds`\] and models.
+- `plot_random`: bool = `True`. Plots the time series randomly.
+- `models`: List\[str\]. A list with the models we want to plot.
+- `level`: List\[float\]. A list with the prediction intervals we want
+ to plot.
+- `engine`: str = `plotly`. It can also be `matplotlib`. `plotly`
+ generates interactive plots, while `matplotlib` generates static
+ plots.
+
+
+```python
+from statsforecast import StatsForecast
+```
+
+
+```python
+StatsForecast.plot(train, test, plot_random=False)
+```
+
+
+
+## Train models
+
+StatsForecast can train multiple
+[models](https://nixtla.github.io/statsforecast/#models) on different
+time series efficiently. Most of these models can generate a
+probabilistic forecast, which means that they can produce both point
+forecasts and prediction intervals.
+
+For this example, we’ll use
+[AutoETS](https://Nixtla.github.io/statsforecast/src/core/models.html#autoets)
+and the following baseline models:
+
+- [HistoricAverage](https://Nixtla.github.io/statsforecast/src/core/models.html#historicaverage)
+- [Naive](https://Nixtla.github.io/statsforecast/src/core/models.html#naive)
+- [RandomWalkWithDrift](https://Nixtla.github.io/statsforecast/src/core/models.html#randomwalkwithdrift)
+- [SeasonalNaive](https://Nixtla.github.io/statsforecast/src/core/models.html#seasonalnaive)
+
+To use these models, we first need to import them from
+`statsforecast.models` and then we need to instantiate them. Given that
+we’re working with hourly data, we need to set `seasonal_length=24` in
+the models that requiere this parameter.
+
+
+```python
+from statsforecast.models import (
+ AutoETS,
+ HistoricAverage,
+ Naive,
+ RandomWalkWithDrift,
+ SeasonalNaive
+)
+```
+
+
+```python
+# Create a list of models and instantiation parameters
+models = [
+ AutoETS(season_length=24),
+ HistoricAverage(),
+ Naive(),
+ RandomWalkWithDrift(),
+ SeasonalNaive(season_length=24)
+]
+```
+
+To instantiate a new StatsForecast object, we need the following
+parameters:
+
+- `df`: The dataframe with the training data.
+- `models`: The list of models defined in the previous step.
+- `freq`: A string indicating the frequency of the data. See [pandas’
+ available
+ frequencies](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#offset-aliases).
+- `n_jobs`: An integer that indicates the number of jobs used in
+ parallel processing. Use -1 to select all cores.
+
+
+```python
+sf = StatsForecast(
+ models=models,
+ freq=1,
+ n_jobs=-1
+)
+```
+
+Now we’re ready to generate the point forecasts and the prediction
+intervals. To do this, we’ll use the `forecast` method, which takes two
+arguments:
+
+- `h`: An integer that represent the forecasting horizon. In this
+ case, we’ll forecast the next 48 hours.
+- `level`: A list of floats with the confidence levels of the
+ prediction intervals. For example, `level=[95]` means that the range
+ of values should include the actual future value with probability
+ 95%.
+
+
+```python
+levels = [80, 90, 95, 99] # confidence levels of the prediction intervals
+
+forecasts = sf.forecast(df=train, h=48, level=levels)
+forecasts.head()
+```
+
+| | unique_id | ds | AutoETS | AutoETS-lo-99 | AutoETS-lo-95 | AutoETS-lo-90 | AutoETS-lo-80 | AutoETS-hi-80 | AutoETS-hi-90 | AutoETS-hi-95 | ... | RWD-hi-99 | SeasonalNaive | SeasonalNaive-lo-80 | SeasonalNaive-lo-90 | SeasonalNaive-lo-95 | SeasonalNaive-lo-99 | SeasonalNaive-hi-80 | SeasonalNaive-hi-90 | SeasonalNaive-hi-95 | SeasonalNaive-hi-99 |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | H1 | 701 | 631.889598 | 533.371822 | 556.926831 | 568.978861 | 582.874079 | 680.905116 | 694.800335 | 706.852365 | ... | 789.416619 | 691.0 | 613.351903 | 591.339747 | 572.247484 | 534.932739 | 768.648097 | 790.660253 | 809.752516 | 847.067261 |
+| 1 | H1 | 702 | 559.750830 | 460.738592 | 484.411824 | 496.524343 | 510.489302 | 609.012359 | 622.977317 | 635.089836 | ... | 833.254152 | 618.0 | 540.351903 | 518.339747 | 499.247484 | 461.932739 | 695.648097 | 717.660253 | 736.752516 | 774.067261 |
+| 2 | H1 | 703 | 519.235476 | 419.731233 | 443.522100 | 455.694808 | 469.729161 | 568.741792 | 582.776145 | 594.948853 | ... | 866.990616 | 563.0 | 485.351903 | 463.339747 | 444.247484 | 406.932739 | 640.648097 | 662.660253 | 681.752516 | 719.067261 |
+| 3 | H1 | 704 | 486.973364 | 386.979536 | 410.887460 | 423.120060 | 437.223465 | 536.723263 | 550.826668 | 563.059268 | ... | 895.510095 | 529.0 | 451.351903 | 429.339747 | 410.247484 | 372.932739 | 606.648097 | 628.660253 | 647.752516 | 685.067261 |
+| 4 | H1 | 705 | 464.697366 | 364.216339 | 388.240749 | 400.532950 | 414.705071 | 514.689661 | 528.861782 | 541.153983 | ... | 920.702904 | 504.0 | 426.351903 | 404.339747 | 385.247484 | 347.932739 | 581.648097 | 603.660253 | 622.752516 | 660.067261 |
+
+We’ll now merge the forecasts and their prediction intervals with the
+test set. This will allow us generate the plots of each probabilistic
+model.
+
+
+```python
+test = test.merge(forecasts, how='left', on=['unique_id', 'ds'])
+```
+
+## Plot prediction intervals
+
+To plot the point and the prediction intervals, we’ll use the
+`statsforecast.plot` method again. Notice that now we also need to
+specify the model and the levels that we want to plot.
+
+### AutoETS
+
+
+```python
+sf.plot(train, test, plot_random=False, models=['AutoETS'], level=levels)
+```
+
+
+
+### Historic Average
+
+
+```python
+sf.plot(train, test, plot_random=False, models=['HistoricAverage'], level=levels)
+```
+
+
+
+### Naive
+
+
+```python
+sf.plot(train, test, plot_random=False, models=['Naive'], level=levels)
+```
+
+
+
+### Random Walk with Drift
+
+
+```python
+sf.plot(train, test, plot_random=False, models=['RWD'], level=levels)
+```
+
+
+
+### Seasonal Naive
+
+
+```python
+sf.plot(train, test, plot_random=False, models=['SeasonalNaive'], level=levels)
+```
+
+
+
+From these plots, we can conclude that the uncertainty around each
+forecast varies according to the model that is being used. For the same
+time series, one model can predict a wider range of possible future
+values than others.
+
+## References
+
+[Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+principles and practice, The Statistical Forecasting
+Perspective”](https://otexts.com/fpp3/perspective.html).
+
diff --git a/statsforecast/favicon.svg b/statsforecast/favicon.svg
new file mode 100644
index 00000000..e5f33342
--- /dev/null
+++ b/statsforecast/favicon.svg
@@ -0,0 +1,5 @@
+
diff --git a/statsforecast/imgs/mfles_diagram.png b/statsforecast/imgs/mfles_diagram.png
new file mode 100644
index 00000000..cb13aea2
Binary files /dev/null and b/statsforecast/imgs/mfles_diagram.png differ
diff --git a/statsforecast/index.html.mdx b/statsforecast/index.html.mdx
new file mode 100644
index 00000000..41e8d256
--- /dev/null
+++ b/statsforecast/index.html.mdx
@@ -0,0 +1,268 @@
+---
+description: >-
+ StatsForecast offers a collection of popular univariate time series
+ forecasting models optimized for high performance and scalability.
+output-file: index.html
+title: StatsForecast ⚡️
+---
+
+
+## Installation
+
+You can install
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+with:
+
+
+```python
+pip install statsforecast
+```
+
+or
+
+
+```python
+conda install -c conda-forge statsforecast
+```
+
+Vist our [Installation
+Guide](./docs/getting-started/installation.html) for further
+instructions.
+
+## Quick Start
+
+**Minimal Example**
+
+
+```python
+from statsforecast import StatsForecast
+from statsforecast.models import AutoARIMA
+from statsforecast.utils import AirPassengersDF
+
+df = AirPassengersDF
+sf = StatsForecast(
+ models=[AutoARIMA(season_length=12)],
+ freq='ME',
+)
+
+sf.fit(df)
+sf.predict(h=12, level=[95])
+```
+
+**Get Started with this [quick
+guide](docs/getting-started/getting_started_short.html).**
+
+**Follow this [end-to-end
+walkthrough](docs/getting-started/getting_started_complete.html) for
+best practices.**
+
+## Why?
+
+Current Python alternatives for statistical models are slow, inaccurate
+and don’t scale well. So we created a library that can be used to
+forecast in production environments or as benchmarks.
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+includes an extensive battery of models that can efficiently fit
+millions of time series.
+
+## Features
+
+- Fastest and most accurate implementations of
+ [`AutoARIMA`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoarima),
+ [`AutoETS`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoets),
+ [`AutoCES`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoces),
+ [`MSTL`](https://Nixtla.github.io/statsforecast/src/core/models.html#mstl)
+ and
+ [`Theta`](https://Nixtla.github.io/statsforecast/src/core/models.html#theta)
+ in Python.
+- Out-of-the-box compatibility with Spark, Dask, and Ray.
+- Probabilistic Forecasting and Confidence Intervals.
+- Support for exogenous Variables and static covariates.
+- Anomaly Detection.
+- Familiar sklearn syntax: `.fit` and `.predict`.
+
+## Highlights
+
+- Inclusion of `exogenous variables` and `prediction intervals` for
+ ARIMA.
+- 20x
+ [faster](https://github.com/Nixtla/statsforecast/tree/main/experiments/arima)
+ than `pmdarima`.
+- 1.5x faster than `R`.
+- 500x faster than `Prophet`.
+- 4x
+ [faster](https://github.com/Nixtla/statsforecast/tree/main/experiments/ets)
+ than `statsmodels`.
+- Compiled to high performance machine code through
+ [`numba`](https://numba.pydata.org/).
+- 1,000,000 series in [30
+ min](https://github.com/Nixtla/statsforecast/tree/main/experiments/ray)
+ with [ray](https://github.com/ray-project/ray).
+- Replace FB-Prophet in two lines of code and gain speed and accuracy.
+ Check the experiments
+ [here](https://github.com/Nixtla/statsforecast/tree/main/experiments/arima_prophet_adapter).
+- Fit 10 benchmark models on **1,000,000** series in [under **5
+ min**](https://github.com/Nixtla/statsforecast/tree/main/experiments/benchmarks_at_scale).
+
+Missing something? Please open an issue or write us in
+[](https://join.slack.com/t/nixtlaworkspace/shared_invite/zt-135dssye9-fWTzMpv2WBthq8NK0Yvu6A)
+
+## Examples and Guides
+
+📚 [End to End
+Walkthrough](https://nixtla.github.io/statsforecast/docs/getting-started/getting_started_complete.html):
+Model training, evaluation and selection for multiple time series
+
+🔎 [Anomaly
+Detection](https://nixtla.github.io/statsforecast/docs/tutorials/anomalydetection.html):
+detect anomalies for time series using in-sample prediction intervals.
+
+👩🔬 [Cross
+Validation](https://nixtla.github.io/statsforecast/docs/tutorials/crossvalidation.html):
+robust model’s performance evaluation.
+
+❄️ [Multiple
+Seasonalities](https://nixtla.github.io/statsforecast/docs/tutorials/multipleseasonalities.html):
+how to forecast data with multiple seasonalities using an MSTL.
+
+🔌 [Predict Demand
+Peaks](https://nixtla.github.io/statsforecast/docs/tutorials/electricitypeakforecasting.html):
+electricity load forecasting for detecting daily peaks and reducing
+electric bills.
+
+📈 [Intermittent
+Demand](https://nixtla.github.io/statsforecast/docs/tutorials/intermittentdata.html):
+forecast series with very few non-zero observations.
+
+🌡️ [Exogenous
+Regressors](https://nixtla.github.io/statsforecast/docs/how-to-guides/exogenous.html):
+like weather or prices
+
+## Models
+
+### Automatic Forecasting
+
+Automatic forecasting tools search for the best parameters and select
+the best possible model for a group of time series. These tools are
+useful for large collections of univariate time series.
+
+| Model | Point Forecast | Probabilistic Forecast | Insample fitted values | Probabilistic fitted values | Exogenous features |
+|:-------------------------|:--------:|:--------:|:--------:|:--------:|:--------:|
+| [AutoARIMA](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#autoarima) | ✅ | ✅ | ✅ | ✅ | ✅ |
+| [AutoETS](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#autoets) | ✅ | ✅ | ✅ | ✅ | |
+| [AutoCES](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#autoces) | ✅ | ✅ | ✅ | ✅ | |
+| [AutoTheta](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#autotheta) | ✅ | ✅ | ✅ | ✅ | |
+| [AutoMFLES](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#automfles) | ✅ | ✅ | ✅ | ✅ | ✅ |
+| [AutoTBATS](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#autotbats) | ✅ | ✅ | ✅ | ✅ | |
+
+### ARIMA Family
+
+These models exploit the existing autocorrelations in the time series.
+
+| Model | Point Forecast | Probabilistic Forecast | Insample fitted values | Probabilistic fitted values | Exogenous features |
+|:-------------------------|:--------:|:--------:|:--------:|:--------:|:--------:|
+| [ARIMA](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#arima) | ✅ | ✅ | ✅ | ✅ | ✅ |
+| [AutoRegressive](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#autoregressive) | ✅ | ✅ | ✅ | ✅ | ✅ |
+
+### Theta Family
+
+Fit two theta lines to a deseasonalized time series, using different
+techniques to obtain and combine the two theta lines to produce the
+final forecasts.
+
+| Model | Point Forecast | Probabilistic Forecast | Insample fitted values | Probabilistic fitted values | Exogenous features |
+|:-------------------------|:--------:|:--------:|:--------:|:--------:|:--------:|
+| [Theta](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#theta) | ✅ | ✅ | ✅ | ✅ | |
+| [OptimizedTheta](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#optimizedtheta) | ✅ | ✅ | ✅ | ✅ | |
+| [DynamicTheta](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#dynamictheta) | ✅ | ✅ | ✅ | ✅ | |
+| [DynamicOptimizedTheta](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#dynamicoptimizedtheta) | ✅ | ✅ | ✅ | ✅ | |
+
+### Multiple Seasonalities
+
+Suited for signals with more than one clear seasonality. Useful for
+low-frequency data like electricity and logs.
+
+| Model | Point Forecast | Probabilistic Forecast | Insample fitted values | Probabilistic fitted values | Exogenous features |
+|:-------------------------|:--------:|:--------:|:--------:|:--------:|:--------:|
+| [MSTL](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#mstl) | ✅ | ✅ | ✅ | ✅ | If trend forecaster supports |
+| [MFLES](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#mfles) | ✅ | ✅ | ✅ | ✅ | ✅ |
+| [TBATS](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#tbats) | ✅ | ✅ | ✅ | ✅ | |
+
+### GARCH and ARCH Models
+
+Suited for modeling time series that exhibit non-constant volatility
+over time. The ARCH model is a particular case of GARCH.
+
+| Model | Point Forecast | Probabilistic Forecast | Insample fitted values | Probabilistic fitted values | Exogenous features |
+|:-------------------------|:--------:|:--------:|:--------:|:--------:|:--------:|
+| [GARCH](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#garch) | ✅ | ✅ | ✅ | ✅ | |
+| [ARCH](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#arch) | ✅ | ✅ | ✅ | ✅ | |
+
+### Baseline Models
+
+Classical models for establishing baseline.
+
+| Model | Point Forecast | Probabilistic Forecast | Insample fitted values | Probabilistic fitted values | Exogenous features |
+|:-------------------------|:--------:|:--------:|:--------:|:--------:|:--------:|
+| [HistoricAverage](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#historicaverage) | ✅ | ✅ | ✅ | ✅ | |
+| [Naive](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#naive) | ✅ | ✅ | ✅ | ✅ | |
+| [RandomWalkWithDrift](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#randomwalkwithdrift) | ✅ | ✅ | ✅ | ✅ | |
+| [SeasonalNaive](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#seasonalnaive) | ✅ | ✅ | ✅ | ✅ | |
+| [WindowAverage](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#windowaverage) | ✅ | | | | |
+| [SeasonalWindowAverage](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#seasonalwindowaverage) | ✅ | | | | |
+
+### Exponential Smoothing
+
+Uses a weighted average of all past observations where the weights
+decrease exponentially into the past. Suitable for data with clear trend
+and/or seasonality. Use the `SimpleExponential` family for data with no
+clear trend or seasonality.
+
+| Model | Point Forecast | Probabilistic Forecast | Insample fitted values | Probabilistic fitted values | Exogenous features |
+|:-------------------------|:--------:|:--------:|:--------:|:--------:|:--------:|
+| [SimpleExponentialSmoothing](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#simpleexponentialsmoothing) | ✅ | | | | |
+| [SimpleExponentialSmoothingOptimized](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#simpleexponentialsmoothingoptimized) | ✅ | | | | |
+| [SeasonalExponentialSmoothing](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#seasonalexponentialsmoothing) | ✅ | | | | |
+| [SeasonalExponentialSmoothingOptimized](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#seasonalexponentialsmoothingoptimized) | ✅ | | | | |
+| [Holt](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#holt) | ✅ | ✅ | ✅ | ✅ | |
+| [HoltWinters](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#holtwinters) | ✅ | ✅ | ✅ | ✅ | |
+
+### Sparse or Inttermitent
+
+Suited for series with very few non-zero observations
+
+| Model | Point Forecast | Probabilistic Forecast | Insample fitted values | Probabilistic fitted values | Exogenous features |
+|:-------------------------|:--------:|:--------:|:--------:|:--------:|:--------:|
+| [ADIDA](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#adida) | ✅ | | ✅ | ✅ | |
+| [CrostonClassic](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#crostonclassic) | ✅ | | ✅ | ✅ | |
+| [CrostonOptimized](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#crostonoptimized) | ✅ | | ✅ | ✅ | |
+| [CrostonSBA](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#crostonsba) | ✅ | | ✅ | ✅ | |
+| [IMAPA](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#imapa) | ✅ | | ✅ | ✅ | |
+| [TSB](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#tsb) | ✅ | | ✅ | ✅ | |
+
+### Machine Learning
+
+Leverage exogenous features.
+
+| Model | Point Forecast | Probabilistic Forecast | Insample fitted values | Probabilistic fitted values | Exogenous features |
+|:-------------------------|:--------:|:--------:|:--------:|:--------:|:--------:|
+| [SklearnModel](https://nixtlaverse.nixtla.io/statsforecast/src/core/models.html#sklearnmodel) | ✅ | | ✅ | ✅ | ✅ |
+
+## How to contribute
+
+See
+[CONTRIBUTING.md](https://github.com/Nixtla/statsforecast/blob/main/CONTRIBUTING.md).
+
+## Citing
+
+
+```bibtex
+@misc{garza2022statsforecast,
+ author={Federico Garza, Max Mergenthaler Canseco, Cristian Challú, Kin G. Olivares},
+ title = {{StatsForecast}: Lightning fast forecasting with statistical and econometric models},
+ year={2022},
+ howpublished={{PyCon} Salt Lake City, Utah, US 2022},
+ url={https://github.com/Nixtla/statsforecast}
+}
+```
+
diff --git a/statsforecast/light.png b/statsforecast/light.png
new file mode 100644
index 00000000..bbb99b54
Binary files /dev/null and b/statsforecast/light.png differ
diff --git a/statsforecast/mint.json b/statsforecast/mint.json
new file mode 100644
index 00000000..aae8873a
--- /dev/null
+++ b/statsforecast/mint.json
@@ -0,0 +1,137 @@
+{
+ "$schema": "https://mintlify.com/schema.json",
+ "name": "Nixtla",
+ "logo": {
+ "light": "/light.png",
+ "dark": "/dark.png"
+ },
+ "favicon": "/favicon.svg",
+ "colors": {
+ "primary": "#0E0E0E",
+ "light": "#FAFAFA",
+ "dark": "#0E0E0E",
+ "anchors": {
+ "from": "#2AD0CA",
+ "to": "#0E00F8"
+ }
+ },
+ "topbarCtaButton": {
+ "type": "github",
+ "url": "https://github.com/Nixtla/nixtla"
+ },
+ "topAnchor": {
+ "name": "StatsForecast",
+ "icon": "bolt"
+ },
+ "navigation": [
+ {
+ "group": "",
+ "pages": ["index.html"]
+ },
+ {
+ "group": "Getting Started",
+ "pages": [
+ "docs/getting-started/installation.html",
+ "docs/getting-started/getting_started_short.html",
+ "docs/getting-started/getting_started_complete.html",
+ "docs/getting-started/getting_started_complete_polars.html"
+ ]
+ },
+ {
+ "group": "Tutorials",
+ "pages": [
+ "docs/tutorials/anomalydetection.html",
+ "docs/tutorials/conformalprediction.html",
+ "docs/tutorials/crossvalidation.html",
+ "docs/tutorials/electricityloadforecasting.html",
+ "docs/tutorials/electricitypeakforecasting.html",
+ "docs/tutorials/garch_tutorial.html",
+ "docs/tutorials/intermittentdata.html",
+ "docs/tutorials/mlflow.html",
+ "docs/tutorials/multipleseasonalities.html",
+ "docs/tutorials/statisticalneuralmethods.html",
+ "docs/tutorials/uncertaintyintervals.html"
+ ]
+ },
+ {
+ "group": "How to Guides",
+ "pages": [
+ "docs/how-to-guides/automatic_forecasting.html",
+ "docs/how-to-guides/exogenous.html",
+ "docs/how-to-guides/generating_features.html",
+ "docs/how-to-guides/sklearn_models.html",
+ "docs/how-to-guides/migrating_R",
+ "docs/how-to-guides/numba_cache.html"
+ ]
+ },
+ {
+ "group": "Distributed",
+ "pages": [
+ "docs/distributed/dask.html",
+ "docs/distributed/ray.html",
+ "docs/distributed/spark.html"
+ ]
+ },
+ {
+ "group": "Experiments",
+ "pages": [
+ "docs/experiments/amazonstatsforecast.html",
+ "docs/experiments/autoarima_vs_prophet.html",
+ "docs/experiments/ets_ray_m5.html",
+ "docs/experiments/prophet_spark_m5.html"
+ ]
+ },
+ {
+ "group": "Model References",
+ "pages": [
+ "docs/models/adida.html",
+ "docs/models/arch.html",
+ "docs/models/arima.html",
+ "docs/models/autoarima.html",
+ "docs/models/autoces.html",
+ "docs/models/autoets.html",
+ "docs/models/autoregressive.html",
+ "docs/models/autotheta.html",
+ "docs/models/crostonclassic.html",
+ "docs/models/crostonoptimized.html",
+ "docs/models/crostonsba.html",
+ "docs/models/dynamicoptimizedtheta.html",
+ "docs/models/dynamicstandardtheta.html",
+ "docs/models/garch.html",
+ "docs/models/holt.html",
+ "docs/models/holtwinters.html",
+ "docs/models/imapa.html",
+ "docs/models/mfles.html",
+ "docs/models/multipleseasonaltrend.html",
+ "docs/models/optimizedtheta.html",
+ "docs/models/seasonalexponentialsmoothing.html",
+ "docs/models/seasonalexponentialsmoothingoptimized.html",
+ "docs/models/simpleexponentialoptimized.html",
+ "docs/models/simpleexponentialsmoothing.html",
+ "docs/models/standardtheta.html",
+ "docs/models/tsb.html"
+ ]
+ },
+ {
+ "group": "API Reference",
+ "pages": [
+ "src/core/core.html",
+ "src/core/distributed.fugue.html",
+ "src/core/models.html",
+ "src/core/models_intro",
+ "src/feature_engineering.html"
+ ]
+ },
+ {
+ "group": "Contributing",
+ "pages": [
+ "docs/contribute/contribute",
+ "docs/contribute/docs",
+ "docs/contribute/issue-labels",
+ "docs/contribute/issues",
+ "docs/contribute/step-by-step",
+ "docs/contribute/techstack"
+ ]
+ }
+ ]
+}
diff --git a/statsforecast/src/adapters.prophet.html.mdx b/statsforecast/src/adapters.prophet.html.mdx
new file mode 100644
index 00000000..5e545a5e
--- /dev/null
+++ b/statsforecast/src/adapters.prophet.html.mdx
@@ -0,0 +1,313 @@
+---
+description: >-
+ In 2017, Facebook open-sourced
+ [Prophet](https://peerj.com/preprints/3190.pdf), with the promise of providing
+ experts and non-experts the possibility of producing high-quality predictions.
+ The forecasting community heavily adopted the solution, reaching millions of
+ accumulated downloads. It became evident that its [quality is
+ shadowed](https://www.reddit.com/r/MachineLearning/comments/wqrw8x/d_fool_me_once_shame_on_you_fool_me_twice_shame/)
+ by simpler well-proven methods. This effort aims to provide an alternative to
+ overcome the Prophet's memory.
"It is important to
+ note that false prophets sometimes prophesied accurately, ... "
+ (Deuteronomy 13:2,5)
+output-file: adapters.prophet.html
+title: Replace FB-Prophet
+---
+
+
+# 1. AutoARIMA Adapter
+
+## AutoArimaProphet
+
+------------------------------------------------------------------------
+
+source
+
+### AutoARIMAProphet
+
+> ``` text
+> AutoARIMAProphet (growth='linear', changepoints=None, n_changepoints=25,
+> changepoint_range=0.8, yearly_seasonality='auto',
+> weekly_seasonality='auto', daily_seasonality='auto',
+> holidays=None, seasonality_mode='additive',
+> seasonality_prior_scale=10.0,
+> holidays_prior_scale=10.0,
+> changepoint_prior_scale=0.05, mcmc_samples=0,
+> interval_width=0.8, uncertainty_samples=1000,
+> stan_backend=None, d=None, D=None, max_p=5, max_q=5,
+> max_P=2, max_Q=2, max_order=5, max_d=2, max_D=1,
+> start_p=2, start_q=2, start_P=1, start_Q=1,
+> stationary=False, seasonal=True, ic='aicc',
+> stepwise=True, nmodels=94, trace=False,
+> approximation=False, method=None, truncate=None,
+> test='kpss', test_kwargs=None, seasonal_test='seas',
+> seasonal_test_kwargs=None, allowdrift=False,
+> allowmean=False, blambda=None, biasadj=False, period=1)
+> ```
+
+\*AutoARIMAProphet adapter.
+
+Returns best ARIMA model using external variables created by the Prophet
+interface. This class receives as parameters the same as prophet.Prophet
+and uses a `models.AutoARIMA` backend.
+
+If your forecasting pipeline uses Prophet the
+[`AutoARIMAProphet`](https://Nixtla.github.io/statsforecast/src/adapters.prophet.html#autoarimaprophet)
+adapter helps to easily substitute Prophet with an AutoARIMA.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| growth | str | linear | ‘linear’, ‘logistic’ or ‘flat’ to specify a linear, logistic or flat trend. |
+| changepoints | NoneType | None | Potential changepoints. Otherwise selected automatically. |
+| n_changepoints | int | 25 | Number of potential changepoints to include. |
+| changepoint_range | float | 0.8 | Proportion of history in which trend changepoints will be estimated. |
+| yearly_seasonality | str | auto | Fit yearly seasonality. Can be ‘auto’, True, False, or a number of Fourier terms to generate. |
+| weekly_seasonality | str | auto | Fit weekly seasonality. Can be ‘auto’, True, False, or a number of Fourier terms to generate. |
+| daily_seasonality | str | auto | Fit daily seasonality. Can be ‘auto’, True, False, or a number of Fourier terms to generate. |
+| holidays | NoneType | None | DataFrame with columns holiday (string) and ds (date type). |
+| seasonality_mode | str | additive | |
+| seasonality_prior_scale | float | 10.0 | |
+| holidays_prior_scale | float | 10.0 | |
+| changepoint_prior_scale | float | 0.05 | |
+| mcmc_samples | int | 0 | |
+| interval_width | float | 0.8 | Uncertainty forecast intervals width. [`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)’s level |
+| uncertainty_samples | int | 1000 | |
+| stan_backend | NoneType | None | |
+| d | NoneType | None | |
+| D | NoneType | None | |
+| max_p | int | 5 | |
+| max_q | int | 5 | |
+| max_P | int | 2 | |
+| max_Q | int | 2 | |
+| max_order | int | 5 | |
+| max_d | int | 2 | |
+| max_D | int | 1 | |
+| start_p | int | 2 | |
+| start_q | int | 2 | |
+| start_P | int | 1 | |
+| start_Q | int | 1 | |
+| stationary | bool | False | |
+| seasonal | bool | True | |
+| ic | str | aicc | |
+| stepwise | bool | True | |
+| nmodels | int | 94 | |
+| trace | bool | False | |
+| approximation | bool | False | |
+| method | NoneType | None | |
+| truncate | NoneType | None | |
+| test | str | kpss | |
+| test_kwargs | NoneType | None | |
+| seasonal_test | str | seas | |
+| seasonal_test_kwargs | NoneType | None | |
+| allowdrift | bool | False | |
+| allowmean | bool | False | |
+| blambda | NoneType | None | |
+| biasadj | bool | False | |
+| period | int | 1 | |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoARIMAProphet.fit
+
+> ``` text
+> AutoARIMAProphet.fit (df, disable_seasonal_features=True)
+> ```
+
+*Fit the AutoARIMAProphet adapter.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | pandas.DataFrame | | DataFrame with columns ds (date type) and y, the time series. |
+| disable_seasonal_features | bool | True | Disable Prophet’s seasonal features. |
+| **Returns** | **AutoARIMAProphet** | | **Adapter object with [`AutoARIMA`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoarima) fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoARIMAProphet.predict
+
+> ``` text
+> AutoARIMAProphet.predict (df=None)
+> ```
+
+*Predict using the AutoARIMAProphet adapter.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | NoneType | None | DataFrame with columns ds (date type) and y, the time series. |
+| **Returns** | **pandas.DataFrame** | | **DataFrame with the forecast components.** |
+
+# 2. Prophet Substitution Example
+
+### Quick Start
+
+In this example, we revisit the time series of the log daily page views
+for the Wikipedia page for [Peyton
+Manning](https://en.wikipedia.org/wiki/Peyton_Manning). The dataset was
+scraped this data using the
+[WikipediaTrend](https://cran.r-project.org/web/packages/wikipediatrend/index.html)
+package in R.
+
+The Peyton Manning dataset was selected to illustrate Prophet’s
+features, like multiple seasonality, changing growth rates, and the
+ability to model special days (such as Manning’s playoff and SuperBowl
+appearances). The original CSV is available
+[here](https://github.com/facebook/prophet/blob/main/examples/example_wp_log_peyton_manning.csv).
+
+Here we show that
+[`AutoARIMA`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoarima)
+can improve performance by borrowing the `Prophet`’s feature
+preprocessing.
+
+### Inputs
+
+The
+[`AutoARIMAProphet`](https://Nixtla.github.io/statsforecast/src/adapters.prophet.html#autoarimaprophet)
+adapter uses `Prophet`‘s inputs, a pandas dataframe with two columns:
+`ds` and `y`. The `ds` (datestamp) column should be of a format expected
+by Pandas, ideally ’YYYY-MM-DD’ for a date or ‘YYYY-MM-DD HH:MM:SS’ for
+a timestamp. The `y` column must be numeric, and represents the
+measurement we wish to forecast.
+
+
+```python
+df = pd.read_csv('https://raw.githubusercontent.com/facebook/prophet/main/examples/example_wp_log_peyton_manning.csv')
+df.head()
+```
+
+## 2.1 Univariate Prophet
+
+Here we forecast with `Prophet` without external regressors. We first
+instantiate a new `Prophet` object, and define its forecasting procedure
+into its constructor. After that a classic sklearn `fit` and `predict`
+is used to obtain the predictions.
+
+
+```python
+m = Prophet(daily_seasonality=False)
+m.fit(df)
+future = m.make_future_dataframe(365)
+forecast = m.predict(future)
+```
+
+
+```python
+fig = m.plot(forecast)
+```
+
+Here we forecast with
+[`AutoARIMAProphet`](https://Nixtla.github.io/statsforecast/src/adapters.prophet.html#autoarimaprophet)
+adapter without external regressors. It inherits the `Prophet`
+constructor as well as its `fit` and `predict` methods.
+
+With the class
+[`AutoARIMAProphet`](https://Nixtla.github.io/statsforecast/src/adapters.prophet.html#autoarimaprophet)
+you can simply substitute `Prophet` and you’ll be training an
+[`AutoARIMA`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoarima)
+model without changing anything in your forecasting pipeline.
+
+
+```python
+m = AutoARIMAProphet(daily_seasonality=False)
+m.fit(df)
+# m.fit(df, disable_seasonal_features=False) # Uncomment for better AutoARIMA predictions
+future = m.make_future_dataframe(365)
+forecast = m.predict(future)
+```
+
+
+```python
+fig = m.plot(forecast)
+```
+
+## 2.2 Holiday Prophet
+
+Usually `Prophet` pipelines include the usage of external regressors
+such as **holidays**.
+
+Suppose you want to include holidays or other recurring calendar events,
+you can create a pandas.DataFrame for them. The DataFrame needs two
+columns \[`holiday`, `ds`\] and a row for each holiday. It requires all
+the occurrences of the holiday (as far as the historical data allows)
+and the future events of the holiday. If the future does not have the
+holidays registered, they will be modeled but not included in the
+forecast.
+
+You can also include into the events DataFrame, `lower_window` and
+`upper_window` that extends the effect of the holidays through dates to
+\[`lower_window`, `upper_window`\] days around the date. For example if
+you wanted to account for Christmas Eve in addition to Christmas you’d
+include `lower_window=-1`,`upper_window=0`, or Black Friday in addition
+to Thanksgiving, you’d include `lower_window=0`,`upper_window=1`.
+
+Here we Peyton Manning’s playoff appearances dates:
+
+
+```python
+playoffs = pd.DataFrame({
+ 'holiday': 'playoff',
+ 'ds': pd.to_datetime(['2008-01-13', '2009-01-03', '2010-01-16',
+ '2010-01-24', '2010-02-07', '2011-01-08',
+ '2013-01-12', '2014-01-12', '2014-01-19',
+ '2014-02-02', '2015-01-11', '2016-01-17',
+ '2016-01-24', '2016-02-07']),
+ 'lower_window': 0,
+ 'upper_window': 1,
+})
+superbowls = pd.DataFrame({
+ 'holiday': 'superbowl',
+ 'ds': pd.to_datetime(['2010-02-07', '2014-02-02', '2016-02-07']),
+ 'lower_window': 0,
+ 'upper_window': 1,
+})
+holidays = pd.concat((playoffs, superbowls))
+```
+
+
+```python
+m = Prophet(daily_seasonality=False, holidays=holidays)
+m.add_country_holidays(country_name='US')
+m.fit(df)
+future = m.make_future_dataframe(365)
+forecast = m.predict(future)
+```
+
+
+```python
+fig = m.plot(forecast)
+```
+
+The class
+[`AutoARIMAProphet`](https://Nixtla.github.io/statsforecast/src/adapters.prophet.html#autoarimaprophet)
+adapter allows to handle these scenarios to fit an
+[`AutoARIMA`](https://Nixtla.github.io/statsforecast/src/core/models.html#autoarima)
+model with exogenous variables.
+
+You can enjoy your Prophet pipelines with the improved performance of a
+classic ARIMA.
+
+
+```python
+m = AutoARIMAProphet(daily_seasonality=False,
+ holidays=holidays)
+m.add_country_holidays(country_name='US')
+m.fit(df)
+# m.fit(df, disable_seasonal_features=False) # Uncomment for better AutoARIMA predictions
+future = m.make_future_dataframe(365)
+forecast = m.predict(future)
+```
+
+
+```python
+fig = m.plot(forecast)
+```
+
diff --git a/statsforecast/src/arima.html.mdx b/statsforecast/src/arima.html.mdx
new file mode 100644
index 00000000..3f8652a7
--- /dev/null
+++ b/statsforecast/src/arima.html.mdx
@@ -0,0 +1,138 @@
+---
+output-file: arima.html
+title: ARIMA
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### predict_arima
+
+> ``` text
+> predict_arima (model, n_ahead, newxreg=None, se_fit=True)
+> ```
+
+
+```python
+myarima(ap, order=(2, 1, 1), seasonal={'order': (0, 1, 0), 'period': 12},
+ constant=False, ic='aicc', method='CSS-ML')['aic']
+```
+
+------------------------------------------------------------------------
+
+source
+
+### arima_string
+
+> ``` text
+> arima_string (model, padding=False)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### forecast_arima
+
+> ``` text
+> forecast_arima (model, h=None, level=None, fan=False, xreg=None,
+> blambda=None, bootstrap=False, npaths=5000, biasadj=None)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### fitted_arima
+
+> ``` text
+> fitted_arima (model, h=1)
+> ```
+
+*Returns h-step forecasts for the data used in fitting the model.*
+
+------------------------------------------------------------------------
+
+source
+
+### auto_arima_f
+
+> ``` text
+> auto_arima_f (x, d=None, D=None, max_p=5, max_q=5, max_P=2, max_Q=2,
+> max_order=5, max_d=2, max_D=1, start_p=2, start_q=2,
+> start_P=1, start_Q=1, stationary=False, seasonal=True,
+> ic='aicc', stepwise=True, nmodels=94, trace=False,
+> approximation=None, method=None, truncate=None, xreg=None,
+> test='kpss', test_kwargs=None, seasonal_test='seas',
+> seasonal_test_kwargs=None, allowdrift=True, allowmean=True,
+> blambda=None, biasadj=False, period=1)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### print_statsforecast_ARIMA
+
+> ``` text
+> print_statsforecast_ARIMA (model, digits=3, se=True)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### ARIMASummary
+
+> ``` text
+> ARIMASummary (model)
+> ```
+
+*ARIMA Summary.*
+
+------------------------------------------------------------------------
+
+source
+
+### AutoARIMA
+
+> ``` text
+> AutoARIMA (d:Optional[int]=None, D:Optional[int]=None, max_p:int=5,
+> max_q:int=5, max_P:int=2, max_Q:int=2, max_order:int=5,
+> max_d:int=2, max_D:int=1, start_p:int=2, start_q:int=2,
+> start_P:int=1, start_Q:int=1, stationary:bool=False,
+> seasonal:bool=True, ic:str='aicc', stepwise:bool=True,
+> nmodels:int=94, trace:bool=False,
+> approximation:Optional[bool]=None, method:Optional[str]=None,
+> truncate:Optional[bool]=None, test:str='kpss',
+> test_kwargs:Optional[str]=None, seasonal_test:str='seas',
+> seasonal_test_kwargs:Optional[Dict]=None,
+> allowdrift:bool=True, allowmean:bool=True,
+> blambda:Optional[float]=None, biasadj:bool=False,
+> period:int=1)
+> ```
+
+\*An AutoARIMA estimator.
+
+Returns best ARIMA model according to either AIC, AICc or BIC value. The
+function conducts a search over possible model within the order
+constraints provided.\*
+
diff --git a/statsforecast/src/ces.html.mdx b/statsforecast/src/ces.html.mdx
new file mode 100644
index 00000000..059011f0
--- /dev/null
+++ b/statsforecast/src/ces.html.mdx
@@ -0,0 +1,23 @@
+---
+output-file: ces.html
+title: CES Model
+---
+
+
+## cescalc
+
+------------------------------------------------------------------------
+
+source
+
+### ces_target_fn
+
+> ``` text
+> ces_target_fn (optimal_param, init_alpha_0, init_alpha_1, init_beta_0,
+> init_beta_1, opt_alpha_0, opt_alpha_1, opt_beta_0,
+> opt_beta_1, y, m, init_states, n_components, seasontype,
+> nmse)
+> ```
+
diff --git a/statsforecast/src/core/core.html.mdx b/statsforecast/src/core/core.html.mdx
new file mode 100644
index 00000000..c0b8fe44
--- /dev/null
+++ b/statsforecast/src/core/core.html.mdx
@@ -0,0 +1,710 @@
+---
+description: Methods for Fit, Predict, Forecast (fast), Cross Validation and plotting
+output-file: core.html
+title: Core Methods
+---
+
+
+The core methods of
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+are:
+
+- `StatsForecast.fit`
+- `StatsForecast.predict`
+- [`StatsForecast.forecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast.forecast)
+- [`StatsForecast.cross_validation`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast.cross_validation)
+- `StatsForecast.plot`
+
+------------------------------------------------------------------------
+
+source
+
+## StatsForecast
+
+> ``` text
+> StatsForecast (models:List[Any], freq:Union[str,int], n_jobs:int=1,
+> fallback_model:Optional[Any]=None, verbose:bool=False)
+> ```
+
+\*The
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+class allows you to efficiently fit multiple
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+models for large sets of time series. It operates on a DataFrame `df`
+with at least three columns ids, times and targets.
+
+The class has memory-efficient
+[`StatsForecast.forecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast.forecast)
+method that avoids storing partial model outputs. While the
+`StatsForecast.fit` and `StatsForecast.predict` methods with
+Scikit-learn interface store the fitted models.
+
+The
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+class offers parallelization utilities with Dask, Spark and Ray
+back-ends. See distributed computing example
+[here](https://github.com/Nixtla/statsforecast/tree/main/experiments/ray).\*
+
+
+```python
+# StatsForecast's class usage example
+
+#from statsforecast.core import StatsForecast
+from statsforecast.models import (
+ ADIDA,
+ AutoARIMA,
+ CrostonClassic,
+ CrostonOptimized,
+ CrostonSBA,
+ HistoricAverage,
+ IMAPA,
+ Naive,
+ RandomWalkWithDrift,
+ SeasonalExponentialSmoothing,
+ SeasonalNaive,
+ SeasonalWindowAverage,
+ SimpleExponentialSmoothing,
+ TSB,
+ WindowAverage,
+ DynamicOptimizedTheta,
+ AutoETS,
+ AutoCES
+)
+```
+
+
+```python
+# Generate synthetic panel DataFrame for example
+panel_df = generate_series(n_series=9, equal_ends=False, engine='pandas')
+panel_df.groupby('unique_id').tail(4)
+```
+
+
+```python
+# Declare list of instantiated StatsForecast estimators to be fitted
+# You can try other estimator's hyperparameters
+# You can try other methods from the `models.StatsForecast` collection
+# Check them here: https://Nixtla.github.io/statsforecast/src/core/models.html
+models=[AutoARIMA(), Naive(),
+ AutoETS(), AutoARIMA(allowmean=True, alias='MeanAutoARIMA')]
+
+# Instantiate StatsForecast class
+fcst = StatsForecast(models=models,
+ freq='D',
+ n_jobs=1,
+ verbose=True)
+
+# Efficiently predict
+fcsts_df = fcst.forecast(df=panel_df, h=4, fitted=True)
+fcsts_df.groupby('unique_id').tail(4)
+```
+
+------------------------------------------------------------------------
+
+source
+
+## StatsForecast.fit
+
+> ``` text
+> StatsForecast.fit
+> (df:Union[pandas.core.frame.DataFrame,polars.dataframe
+> .frame.DataFrame], prediction_intervals:Optional[stats
+> forecast.utils.ConformalIntervals]=None,
+> id_col:str='unique_id', time_col:str='ds',
+> target_col:str='y')
+> ```
+
+\*Fit statistical models.
+
+Fit `models` to a large set of time series from DataFrame `df` and store
+fitted models for later inspection.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | Union | | DataFrame with ids, times, targets and exogenous. |
+| prediction_intervals | Optional | None | Configuration to calibrate prediction intervals (Conformal Prediction). |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **StatsForecast** | | **Returns with stored [`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast) fitted `models`.** |
+
+------------------------------------------------------------------------
+
+source
+
+## SatstForecast.predict
+
+> ``` text
+> SatstForecast.predict (h:int, X_df:Union[pandas.core.frame.DataFrame,pola
+> rs.dataframe.frame.DataFrame,NoneType]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+\*Predict statistical models.
+
+Use stored fitted `models` to predict large set of time series from
+DataFrame `df`.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X_df | Union | None | DataFrame with ids, times and future exogenous. |
+| level | Optional | None | Confidence levels between 0 and 100 for prediction intervals. |
+| **Returns** | **pandas or polars DataFrame** | | **DataFrame with `models` columns for point predictions and probabilistic predictions for all fitted `models`.** |
+
+------------------------------------------------------------------------
+
+source
+
+## StatsForecast.fit_predict
+
+> ``` text
+> StatsForecast.fit_predict (h:int,
+> df:Union[pandas.core.frame.DataFrame,polars.da
+> taframe.frame.DataFrame], X_df:Union[pandas.co
+> re.frame.DataFrame,polars.dataframe.frame.Data
+> Frame,NoneType]=None,
+> level:Optional[List[int]]=None, prediction_int
+> ervals:Optional[statsforecast.utils.ConformalI
+> ntervals]=None, id_col:str='unique_id',
+> time_col:str='ds', target_col:str='y')
+> ```
+
+\*Fit and Predict with statistical models.
+
+This method avoids memory burden due from object storage. It is
+analogous to Scikit-Learn `fit_predict` without storing information. It
+requires the forecast horizon `h` in advance.
+
+In contrast to
+[`StatsForecast.forecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast.forecast)
+this method stores partial models outputs.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| df | Union | | DataFrame with ids, times, targets and exogenous. |
+| X_df | Union | None | DataFrame with ids, times and future exogenous. |
+| level | Optional | None | Confidence levels between 0 and 100 for prediction intervals. |
+| prediction_intervals | Optional | None | Configuration to calibrate prediction intervals (Conformal Prediction). |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **Union** | | **DataFrame with `models` columns for point predictions and probabilistic predictions for all fitted `models`.** |
+
+------------------------------------------------------------------------
+
+source
+
+## StatsForecast.forecast
+
+> ``` text
+> StatsForecast.forecast (h:int,
+> df:Union[pandas.core.frame.DataFrame,polars.dataf
+> rame.frame.DataFrame], X_df:Union[pandas.core.fra
+> me.DataFrame,polars.dataframe.frame.DataFrame,Non
+> eType]=None, level:Optional[List[int]]=None,
+> fitted:bool=False, prediction_intervals:Optional[
+> statsforecast.utils.ConformalIntervals]=None,
+> id_col:str='unique_id', time_col:str='ds',
+> target_col:str='y')
+> ```
+
+\*Memory Efficient predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to Scikit-Learn `fit_predict` without storing information. It
+requires the forecast horizon `h` in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| df | Union | | DataFrame with ids, times, targets and exogenous. |
+| X_df | Union | None | DataFrame with ids, times and future exogenous. |
+| level | Optional | None | Confidence levels between 0 and 100 for prediction intervals. |
+| fitted | bool | False | Store in-sample predictions. |
+| prediction_intervals | Optional | None | Configuration to calibrate prediction intervals (Conformal Prediction). |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **Union** | | **DataFrame with `models` columns for point predictions and probabilistic predictions for all fitted `models`.** |
+
+
+```python
+# StatsForecast.forecast method usage example
+
+#from statsforecast.core import StatsForecast
+from statsforecast.utils import AirPassengersDF as panel_df
+from statsforecast.models import AutoARIMA, Naive
+```
+
+
+```python
+# Instantiate StatsForecast class
+fcst = StatsForecast(models=[AutoARIMA(), Naive()],
+ freq='D', n_jobs=1)
+
+# Efficiently predict without storing memory
+fcsts_df = fcst.forecast(df=panel_df, h=4, fitted=True)
+fcsts_df.groupby('unique_id').tail(4)
+```
+
+------------------------------------------------------------------------
+
+source
+
+## StatsForecast.forecast_fitted_values
+
+> ``` text
+> StatsForecast.forecast_fitted_values ()
+> ```
+
+\*Access insample predictions.
+
+After executing
+[`StatsForecast.forecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast.forecast),
+you can access the insample prediction values for each model. To get
+them, you need to pass `fitted=True` to the
+[`StatsForecast.forecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast.forecast)
+method and then use the
+[`StatsForecast.forecast_fitted_values`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast.forecast_fitted_values)
+method.\*
+
+
+```python
+# StatsForecast.forecast_fitted_values method usage example
+
+#from statsforecast.core import StatsForecast
+from statsforecast.utils import AirPassengersDF as panel_df
+from statsforecast.models import Naive
+```
+
+
+```python
+# Instantiate StatsForecast class
+fcst = StatsForecast(models=[AutoARIMA()], freq='D', n_jobs=1)
+
+# Access insample predictions
+fcsts_df = fcst.forecast(df=panel_df, h=12, fitted=True, level=(90, 10))
+insample_fcsts_df = fcst.forecast_fitted_values()
+insample_fcsts_df.tail(4)
+```
+
+------------------------------------------------------------------------
+
+source
+
+## StatsForecast.cross_validation
+
+> ``` text
+> StatsForecast.cross_validation (h:int,
+> df:Union[pandas.core.frame.DataFrame,pola
+> rs.dataframe.frame.DataFrame],
+> n_windows:int=1, step_size:int=1,
+> test_size:Optional[int]=None,
+> input_size:Optional[int]=None,
+> level:Optional[List[int]]=None,
+> fitted:bool=False,
+> refit:Union[bool,int]=True, prediction_in
+> tervals:Optional[statsforecast.utils.Conf
+> ormalIntervals]=None,
+> id_col:str='unique_id',
+> time_col:str='ds', target_col:str='y')
+> ```
+
+\*Temporal Cross-Validation.
+
+Efficiently fits a list of
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+models through multiple training windows, in either chained or rolled
+manner.
+
+`StatsForecast.models`’ speed allows to overcome this evaluation
+technique high computational costs. Temporal cross-validation provides
+better model’s generalization measurements by increasing the test’s
+length and diversity.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| df | Union | | DataFrame with ids, times, targets and exogenous. |
+| n_windows | int | 1 | Number of windows used for cross validation. |
+| step_size | int | 1 | Step size between each window. |
+| test_size | Optional | None | Length of test size. If passed, set `n_windows=None`. |
+| input_size | Optional | None | Input size for each window, if not none rolled windows. |
+| level | Optional | None | Confidence levels between 0 and 100 for prediction intervals. |
+| fitted | bool | False | Store in-sample predictions. |
+| refit | Union | True | Wether or not refit the model for each window. If int, train the models every `refit` windows. |
+| prediction_intervals | Optional | None | Configuration to calibrate prediction intervals (Conformal Prediction). |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **Union** | | **DataFrame with insample `models` columns for point predictions and probabilistic predictions for all fitted `models`.** |
+
+
+```python
+# StatsForecast.crossvalidation method usage example
+
+#from statsforecast.core import StatsForecast
+from statsforecast.utils import AirPassengersDF as panel_df
+from statsforecast.models import Naive
+```
+
+
+```python
+# Instantiate StatsForecast class
+fcst = StatsForecast(models=[Naive()],
+ freq='D', n_jobs=1, verbose=True)
+
+# Access insample predictions
+rolled_fcsts_df = fcst.cross_validation(df=panel_df, h=14, n_windows=2)
+rolled_fcsts_df.head(4)
+```
+
+------------------------------------------------------------------------
+
+source
+
+## StatsForecast.cross_validation_fitted_values
+
+> ``` text
+> StatsForecast.cross_validation_fitted_values ()
+> ```
+
+\*Access insample cross validated predictions.
+
+After executing
+[`StatsForecast.cross_validation`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast.cross_validation),
+you can access the insample prediction values for each model and window.
+To get them, you need to pass `fitted=True` to the
+[`StatsForecast.cross_validation`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast.cross_validation)
+method and then use the `StatsForecast.cross_validation_fitted_values`
+method.\*
+
+
+```python
+# StatsForecast.cross_validation_fitted_values method usage example
+
+#from statsforecast.core import StatsForecast
+from statsforecast.utils import AirPassengersDF as panel_df
+from statsforecast.models import Naive
+```
+
+
+```python
+# Instantiate StatsForecast class
+fcst = StatsForecast(models=[Naive()],
+ freq='D', n_jobs=1)
+
+# Access insample predictions
+rolled_fcsts_df = fcst.cross_validation(df=panel_df, h=12, n_windows=2, fitted=True)
+insample_rolled_fcsts_df = fcst.cross_validation_fitted_values()
+insample_rolled_fcsts_df.tail(4)
+```
+
+------------------------------------------------------------------------
+
+source
+
+## StatsForecast.plot
+
+> ``` text
+> StatsForecast.plot
+> (df:Union[pandas.core.frame.DataFrame,polars.datafram
+> e.frame.DataFrame], forecasts_df:Union[pandas.core.fr
+> ame.DataFrame,polars.dataframe.frame.DataFrame,NoneTy
+> pe]=None, unique_ids:Union[List[str],NoneType,numpy.n
+> darray]=None, plot_random:bool=True,
+> models:Optional[List[str]]=None,
+> level:Optional[List[float]]=None,
+> max_insample_length:Optional[int]=None,
+> plot_anomalies:bool=False, engine:str='matplotlib',
+> id_col:str='unique_id', time_col:str='ds',
+> target_col:str='y',
+> resampler_kwargs:Optional[Dict]=None)
+> ```
+
+*Plot forecasts and insample values.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | Union | | DataFrame with ids, times, targets and exogenous. |
+| forecasts_df | Union | None | DataFrame ids, times and models. |
+| unique_ids | Union | None | ids to plot. If None, they’re selected randomly. |
+| plot_random | bool | True | Select time series to plot randomly. |
+| models | Optional | None | List of models to plot. |
+| level | Optional | None | List of prediction intervals to plot if paseed. |
+| max_insample_length | Optional | None | Max number of train/insample observations to be plotted. |
+| plot_anomalies | bool | False | Plot anomalies for each prediction interval. |
+| engine | str | matplotlib | Library used to plot. ‘plotly’, ‘plotly-resampler’ or ‘matplotlib’. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| resampler_kwargs | Optional | None | Kwargs to be passed to plotly-resampler constructor. For further custumization (“show_dash”) call the method, store the plotting object and add the extra arguments to its `show_dash` method. |
+
+------------------------------------------------------------------------
+
+source
+
+## StatsForecast.save
+
+> ``` text
+> StatsForecast.save (path:Union[pathlib.Path,str,NoneType]=None,
+> max_size:Optional[str]=None, trim:bool=False)
+> ```
+
+*Function that will save StatsForecast class with certain settings to
+make it reproducible.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| path | Union | None | Path of the file to be saved. If `None` will create one in the current directory using the current UTC timestamp. |
+| max_size | Optional | None | StatsForecast object should not exceed this size. Available byte naming: \[‘B’, ‘KB’, ‘MB’, ‘GB’\] |
+| trim | bool | False | Delete any attributes not needed for inference. |
+
+------------------------------------------------------------------------
+
+source
+
+## StatsForecast.load
+
+> ``` text
+> StatsForecast.load (path:Union[pathlib.Path,str])
+> ```
+
+*Automatically loads the model into ready StatsForecast.*
+
+| | **Type** | **Details** |
+|-------------|-----------------------|------------------------------------|
+| path | Union | Path to saved StatsForecast file. |
+| **Returns** | **sf: StatsForecast** | **Previously saved StatsForecast** |
+
+
+```python
+fcst = StatsForecast(
+ models=[ADIDA(), SimpleExponentialSmoothing(0.1),
+ HistoricAverage(), CrostonClassic()],
+ freq='D',
+ n_jobs=1
+)
+res = fcst.forecast(df=series, h=14)
+```
+
+# Misc
+
+## Integer datestamp
+
+The
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+class can also receive integers as datestamp, the following example
+shows how to do it.
+
+
+```python
+# from statsforecast.core import StatsForecast
+from statsforecast.utils import AirPassengers as ap
+from statsforecast.models import HistoricAverage
+```
+
+
+```python
+int_ds_df = pd.DataFrame({'ds': np.arange(1, len(ap) + 1), 'y': ap})
+int_ds_df.insert(0, 'unique_id', 'AirPassengers')
+int_ds_df.head()
+```
+
+
+```python
+int_ds_df.tail()
+```
+
+
+```python
+int_ds_df
+```
+
+
+```python
+fcst = StatsForecast(models=[HistoricAverage()], freq=1)
+horizon = 7
+forecast = fcst.forecast(df=int_ds_df, h=horizon)
+forecast.head()
+```
+
+
+```python
+last_date = int_ds_df['ds'].max()
+test_eq(forecast['ds'].values, np.arange(last_date + 1, last_date + 1 + horizon))
+```
+
+
+```python
+int_ds_cv = fcst.cross_validation(df=int_ds_df, h=7, test_size=8, n_windows=None)
+int_ds_cv
+```
+
+## External regressors
+
+Every column after **y** is considered an external regressor and will be
+passed to the models that allow them. If you use them you must supply
+the future values to the
+[`StatsForecast.forecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast.forecast)
+method.
+
+
+```python
+class LinearRegression(_TS):
+
+ def __init__(self):
+ pass
+
+ def fit(self, y, X):
+ self.coefs_, *_ = np.linalg.lstsq(X, y, rcond=None)
+ return self
+
+ def predict(self, h, X):
+ mean = X @ coefs
+ return mean
+
+ def __repr__(self):
+ return 'LinearRegression()'
+
+ def forecast(self, y, h, X=None, X_future=None, fitted=False):
+ coefs, *_ = np.linalg.lstsq(X, y, rcond=None)
+ return {'mean': X_future @ coefs}
+
+ def new(self):
+ b = type(self).__new__(type(self))
+ b.__dict__.update(self.__dict__)
+ return b
+```
+
+
+```python
+series_xreg = series = generate_series(10_000, equal_ends=True)
+series_xreg['intercept'] = 1
+series_xreg['dayofweek'] = series_xreg['ds'].dt.dayofweek
+series_xreg = pd.get_dummies(series_xreg, columns=['dayofweek'], drop_first=True)
+series_xreg
+```
+
+
+```python
+dates = sorted(series_xreg['ds'].unique())
+valid_start = dates[-14]
+train_mask = series_xreg['ds'] < valid_start
+series_train = series_xreg[train_mask]
+series_valid = series_xreg[~train_mask]
+X_valid = series_valid.drop(columns=['y'])
+fcst = StatsForecast(
+ models=[LinearRegression()],
+ freq='D',
+)
+xreg_res = fcst.forecast(df=series_train, h=14, X_df=X_valid)
+xreg_res['y'] = series_valid['y'].values
+```
+
+
+```python
+xreg_res.drop(columns='unique_id').groupby('ds').mean().plot()
+```
+
+
+```python
+xreg_res_cv = fcst.cross_validation(df=series_train, h=3, test_size=5, n_windows=None)
+```
+
+## Prediction intervals
+
+You can pass the argument `level` to the
+[`StatsForecast.forecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast.forecast)
+method to calculate prediction intervals. Not all models can calculate
+them at the moment, so we will only obtain the intervals of those models
+that have it implemented.
+
+
+```python
+ap_df = pd.DataFrame({'ds': np.arange(ap.size), 'y': ap})
+ap_df['unique_id'] = 0
+sf = StatsForecast(
+ models=[
+ SeasonalNaive(season_length=12),
+ AutoARIMA(season_length=12)
+ ],
+ freq=1,
+ n_jobs=1
+)
+ap_ci = sf.forecast(df=ap_df, h=12, level=(80, 95))
+fcst.plot(ap_df, ap_ci, level=[80], engine="matplotlib")
+```
+
+## Conformal Prediction intervals
+
+You can also add conformal intervals using the following code.
+
+
+```python
+from statsforecast.utils import ConformalIntervals
+```
+
+
+```python
+sf = StatsForecast(
+ models=[
+ AutoARIMA(season_length=12),
+ AutoARIMA(
+ season_length=12,
+ prediction_intervals=ConformalIntervals(n_windows=2, h=12),
+ alias='ConformalAutoARIMA'
+ ),
+ ],
+ freq=1,
+ n_jobs=1
+)
+ap_ci = sf.forecast(df=ap_df, h=12, level=(80, 95))
+fcst.plot(ap_df, ap_ci, level=[80], engine="plotly")
+```
+
+You can also compute conformal intervals for all the models that support
+them, using the following,
+
+
+```python
+sf = StatsForecast(
+ models=[
+ AutoARIMA(season_length=12),
+ ],
+ freq=1,
+ n_jobs=1
+)
+ap_ci = sf.forecast(
+ df=ap_df,
+ h=12,
+ level=(50, 80, 95),
+ prediction_intervals=ConformalIntervals(h=12),
+)
+fcst.plot(ap_df, ap_ci, level=[80], engine="matplotlib")
+```
+
diff --git a/statsforecast/src/core/distributed.fugue.html.mdx b/statsforecast/src/core/distributed.fugue.html.mdx
new file mode 100644
index 00000000..3c11a323
--- /dev/null
+++ b/statsforecast/src/core/distributed.fugue.html.mdx
@@ -0,0 +1,257 @@
+---
+output-file: distributed.fugue.html
+title: FugueBackend
+---
+
+
+
+```python
+from statsforecast.core import StatsForecast
+from statsforecast.models import (
+ AutoARIMA,
+ AutoETS,
+)
+from statsforecast.utils import generate_series
+```
+
+
+```python
+n_series = 4
+horizon = 7
+
+series = generate_series(n_series)
+
+sf = StatsForecast(
+ models=[AutoETS(season_length=7)],
+ freq='D',
+)
+
+sf.cross_validation(df=series, h=horizon, step_size = 24,
+ n_windows = 2, level=[90]).head()
+```
+
+
+```python
+from pyspark.sql import SparkSession
+```
+
+
+```python
+spark = SparkSession.builder.getOrCreate()
+
+# Make unique_id a column
+series['unique_id'] = series['unique_id'].astype(str)
+
+# Convert to Spark
+sdf = spark.createDataFrame(series)
+```
+
+
+```python
+# Returns a Spark DataFrame
+sf = StatsForecast(
+ models=[AutoETS(season_length=7)],
+ freq='D',
+)
+sf.cross_validation(df=sdf, h=horizon, step_size = 24,
+ n_windows = 2, level=[90]).show()
+```
+
+------------------------------------------------------------------------
+
+source
+
+### FugueBackend
+
+> ``` text
+> FugueBackend (engine:Any=None, conf:Any=None, **transform_kwargs:Any)
+> ```
+
+\*FugueBackend for Distributed Computation. [Source
+code](https://github.com/Nixtla/statsforecast/blob/main/python/statsforecast/distributed/fugue.py).
+
+This class uses [Fugue](https://github.com/fugue-project/fugue) backend
+capable of distributing computation on Spark, Dask and Ray without any
+rewrites.\*
+
+| | **Type** | **Default** | **Details** |
+|----|----|----|----|
+| engine | Any | None | A selection between Spark, Dask, and Ray. |
+| conf | Any | None | Engine configuration. |
+| transform_kwargs | Any | | |
+
+------------------------------------------------------------------------
+
+source
+
+### FugueBackend.forecast
+
+> ``` text
+> FugueBackend.forecast (df:~AnyDataFrame, freq:Union[str,int],
+> models:List[Any], fallback_model:Optional[Any],
+> X_df:Optional[~AnyDataFrame], h:int,
+> level:Optional[List[int]], fitted:bool, prediction
+> _intervals:Optional[statsforecast.utils.ConformalI
+> ntervals], id_col:str, time_col:str,
+> target_col:str)
+> ```
+
+\*Memory Efficient core.StatsForecast predictions with FugueBackend.
+
+This method uses Fugue’s transform function, in combination with
+`core.StatsForecast`’s forecast to efficiently fit a list of
+StatsForecast models.\*
+
+| | **Type** | **Details** |
+|--------|---------------------------|-------------------------------------|
+| df | AnyDataFrame | DataFrame with ids, times, targets and exogenous. |
+| freq | Union | Frequency of the data. Must be a valid pandas or polars offset alias, or an integer. |
+| models | List | List of instantiated objects models.StatsForecast. |
+| fallback_model | Optional | Any, optional (default=None) Model to be used if a model fails. Only works with the `forecast` and [`cross_validation`](https://Nixtla.github.io/statsforecast/src/mfles.html#cross_validation) methods. |
+| X_df | Optional | DataFrame with ids, times and future exogenous. |
+| h | int | Forecast horizon. |
+| level | Optional | Confidence levels between 0 and 100 for prediction intervals. |
+| fitted | bool | Store in-sample predictions. |
+| prediction_intervals | Optional | Configuration to calibrate prediction intervals (Conformal Prediction). |
+| id_col | str | Column that identifies each serie. |
+| time_col | str | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | Column that contains the target. |
+| **Returns** | **Any** | **DataFrame with `models` columns for point predictions and probabilistic predictions for all fitted `models`** |
+
+------------------------------------------------------------------------
+
+source
+
+### FugueBackend.cross_validation
+
+> ``` text
+> FugueBackend.cross_validation (df:~AnyDataFrame, freq:Union[str,int],
+> models:List[Any],
+> fallback_model:Optional[Any], h:int,
+> n_windows:int, step_size:int,
+> test_size:int, input_size:int,
+> level:Optional[List[int]], refit:bool,
+> fitted:bool, prediction_intervals:Optional
+> [statsforecast.utils.ConformalIntervals],
+> id_col:str, time_col:str, target_col:str)
+> ```
+
+\*Temporal Cross-Validation with core.StatsForecast and FugueBackend.
+
+This method uses Fugue’s transform function, in combination with
+`core.StatsForecast`’s cross-validation to efficiently fit a list of
+StatsForecast models through multiple training windows, in either
+chained or rolled manner.
+
+`StatsForecast.models`’ speed along with Fugue’s distributed computation
+allow to overcome this evaluation technique high computational costs.
+Temporal cross-validation provides better model’s generalization
+measurements by increasing the test’s length and diversity.\*
+
+| | **Type** | **Details** |
+|--------|---------------------------|-------------------------------------|
+| df | AnyDataFrame | DataFrame with ids, times, targets and exogenous. |
+| freq | Union | Frequency of the data. Must be a valid pandas or polars offset alias, or an integer. |
+| models | List | List of instantiated objects models.StatsForecast. |
+| fallback_model | Optional | Any, optional (default=None) Model to be used if a model fails. Only works with the `forecast` and [`cross_validation`](https://Nixtla.github.io/statsforecast/src/mfles.html#cross_validation) methods. |
+| h | int | Forecast horizon. |
+| n_windows | int | Number of windows used for cross validation. |
+| step_size | int | Step size between each window. |
+| test_size | int | Length of test size. If passed, set `n_windows=None`. |
+| input_size | int | Input size for each window, if not none rolled windows. |
+| level | Optional | Confidence levels between 0 and 100 for prediction intervals. |
+| refit | bool | Wether or not refit the model for each window. If int, train the models every `refit` windows. |
+| fitted | bool | Store in-sample predictions. |
+| prediction_intervals | Optional | Configuration to calibrate prediction intervals (Conformal Prediction). |
+| id_col | str | Column that identifies each serie. |
+| time_col | str | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | Column that contains the target. |
+| **Returns** | **Any** | **DataFrame, with `models` columns for point predictions and probabilistic predictions for all fitted `models`.** |
+
+## Dask Distributed Predictions
+
+Here we provide an example for the distribution of the
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+predictions using `Fugue` to execute the code in a Dask cluster.
+
+To do it we instantiate the
+[`FugueBackend`](https://Nixtla.github.io/statsforecast/src/core/distributed.fugue.html#fuguebackend)
+class with a `DaskExecutionEngine`.
+
+
+```python
+import dask.dataframe as dd
+from dask.distributed import Client
+from fugue_dask import DaskExecutionEngine
+from statsforecast import StatsForecast
+from statsforecast.models import Naive
+from statsforecast.utils import generate_series
+```
+
+
+```python
+# Generate Synthetic Panel Data
+df = generate_series(10)
+df['unique_id'] = df['unique_id'].astype(str)
+df = dd.from_pandas(df, npartitions=10)
+
+# Instantiate FugueBackend with DaskExecutionEngine
+dask_client = Client()
+engine = DaskExecutionEngine(dask_client=dask_client)
+```
+
+We have simply create the class to the usual
+[`StatsForecast`](https://Nixtla.github.io/statsforecast/src/core/core.html#statsforecast)
+instantiation.
+
+
+```python
+sf = StatsForecast(models=[Naive()], freq='D')
+```
+
+### Distributed Forecast
+
+For extremely fast distributed predictions we use FugueBackend as
+backend that operates like the original
+[StatsForecast.forecast](https://nixtla.github.io/statsforecast/src/core/core.html#statsforecast.forecast)
+method.
+
+It receives as input a pandas.DataFrame with columns
+\[`unique_id`,`ds`,`y`\] and exogenous, where the `ds` (datestamp)
+column should be of a format expected by Pandas. The `y` column must be
+numeric, and represents the measurement we wish to forecast. And the
+`unique_id` uniquely identifies the series in the panel data.
+
+
+```python
+# Distributed predictions with FugueBackend.
+sf.forecast(df=df, h=12).compute()
+```
+
+
+```python
+sf = StatsForecast(models=[Naive()], freq='D')
+xx = sf.forecast(df=df, h=12, fitted=True).compute()
+yy = sf.forecast_fitted_values().compute()
+```
+
+### Distributed Cross-Validation
+
+For extremely fast distributed temporcal cross-validation we use
+[`cross_validation`](https://Nixtla.github.io/statsforecast/src/mfles.html#cross_validation)
+method that operates like the original
+[StatsForecast.cross_validation](https://nixtla.github.io/statsforecast/src/core/core.html#statsforecast.cross_validation)
+method.
+
+
+```python
+# Distributed cross-validation with FugueBackend.
+sf.cross_validation(df=df, h=12, n_windows=2).compute()
+```
+
diff --git a/statsforecast/src/core/models.html.mdx b/statsforecast/src/core/models.html.mdx
new file mode 100644
index 00000000..a2639d8e
--- /dev/null
+++ b/statsforecast/src/core/models.html.mdx
@@ -0,0 +1,5858 @@
+---
+description: Models currently supported by StatsForecast
+output-file: models.html
+title: Models
+---
+
+
+StatsForecast offers a wide variety of models grouped in the following
+categories:
+
+- **Auto Forecast:** Automatic forecasting tools search for the best
+ parameters and select the best possible model for a series of time
+ series. These tools are useful for large collections of univariate
+ time series. Includes automatic versions of: Arima, ETS, Theta, CES.
+
+- **Exponential Smoothing:** Uses a weighted average of all past
+ observations where the weights decrease exponentially into the past.
+ Suitable for data with clear trend and/or seasonality. Use the
+ `SimpleExponential` family for data with no clear trend or
+ seasonality. Examples: SES, Holt’s Winters, SSO.
+
+- **Benchmark models:** classical models for establishing baselines.
+ Examples: Mean, Naive, Random Walk
+
+- **Intermittent or Sparse models:** suited for series with very few
+ non-zero observations. Examples: CROSTON, ADIDA, IMAPA
+
+- **Multiple Seasonalities:** suited for signals with more than one
+ clear seasonality. Useful for low-frequency data like electricity
+ and logs. Examples: MSTL and TBATS.
+
+- **Theta Models:** fit two theta lines to a deseasonalized time
+ series, using different techniques to obtain and combine the two
+ theta lines to produce the final forecasts. Examples: Theta,
+ DynamicTheta
+
+- **GARCH Model:** suited for modeling time series that exhibit
+ non-constant volatility over time. Commonly used in finance to model
+ stock prices, exchange rates, interest rates, and other financial
+ instruments. The ARCH model is a particular case of GARCH.
+
+# Automatic Forecasting
+
+## AutoARIMA
+
+------------------------------------------------------------------------
+
+source
+
+### AutoARIMA
+
+> ``` text
+> AutoARIMA (d:Optional[int]=None, D:Optional[int]=None, max_p:int=5,
+> max_q:int=5, max_P:int=2, max_Q:int=2, max_order:int=5,
+> max_d:int=2, max_D:int=1, start_p:int=2, start_q:int=2,
+> start_P:int=1, start_Q:int=1, stationary:bool=False,
+> seasonal:bool=True, ic:str='aicc', stepwise:bool=True,
+> nmodels:int=94, trace:bool=False,
+> approximation:Optional[bool]=False, method:Optional[str]=None,
+> truncate:Optional[bool]=None, test:str='kpss',
+> test_kwargs:Optional[str]=None, seasonal_test:str='seas',
+> seasonal_test_kwargs:Optional[Dict]=None,
+> allowdrift:bool=True, allowmean:bool=True,
+> blambda:Optional[float]=None, biasadj:bool=False,
+> season_length:int=1, alias:str='AutoARIMA', prediction_interva
+> ls:Optional[statsforecast.utils.ConformalIntervals]=None)
+> ```
+
+\*AutoARIMA model.
+
+Automatically selects the best ARIMA (AutoRegressive Integrated Moving
+Average) model using an information criterion. Default is Akaike
+Information Criterion (AICc).\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| d | Optional | None | Order of first-differencing. |
+| D | Optional | None | Order of seasonal-differencing. |
+| max_p | int | 5 | Max autorregresives p. |
+| max_q | int | 5 | Max moving averages q. |
+| max_P | int | 2 | Max seasonal autorregresives P. |
+| max_Q | int | 2 | Max seasonal moving averages Q. |
+| max_order | int | 5 | Max p+q+P+Q value if not stepwise selection. |
+| max_d | int | 2 | Max non-seasonal differences. |
+| max_D | int | 1 | Max seasonal differences. |
+| start_p | int | 2 | Starting value of p in stepwise procedure. |
+| start_q | int | 2 | Starting value of q in stepwise procedure. |
+| start_P | int | 1 | Starting value of P in stepwise procedure. |
+| start_Q | int | 1 | Starting value of Q in stepwise procedure. |
+| stationary | bool | False | If True, restricts search to stationary models. |
+| seasonal | bool | True | If False, restricts search to non-seasonal models. |
+| ic | str | aicc | Information criterion to be used in model selection. |
+| stepwise | bool | True | If True, will do stepwise selection (faster). |
+| nmodels | int | 94 | Number of models considered in stepwise search. |
+| trace | bool | False | If True, the searched ARIMA models is reported. |
+| approximation | Optional | False | If True, conditional sums-of-squares estimation, final MLE. |
+| method | Optional | None | Fitting method between maximum likelihood or sums-of-squares. |
+| truncate | Optional | None | Observations truncated series used in model selection. |
+| test | str | kpss | Unit root test to use. See [`ndiffs`](https://Nixtla.github.io/statsforecast/src/arima.html#ndiffs) for details. |
+| test_kwargs | Optional | None | Unit root test additional arguments. |
+| seasonal_test | str | seas | Selection method for seasonal differences. |
+| seasonal_test_kwargs | Optional | None | Seasonal unit root test arguments. |
+| allowdrift | bool | True | If True, drift models terms considered. |
+| allowmean | bool | True | If True, non-zero mean models considered. |
+| blambda | Optional | None | Box-Cox transformation parameter. |
+| biasadj | bool | False | Use adjusted back-transformed mean Box-Cox. |
+| season_length | int | 1 | Number of observations per unit of time. Ex: 24 Hourly data. |
+| alias | str | AutoARIMA | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoARIMA.fit
+
+> ``` text
+> AutoARIMA.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the AutoARIMA model.
+
+Fit an AutoARIMA to a time series (numpy array) `y` and optionally
+exogenous variables (numpy array) `X`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|---------------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | | | **AutoARIMA fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoARIMA.predict
+
+> ``` text
+> AutoARIMA.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted AutoArima.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoARIMA.predict_in_sample
+
+> ``` text
+> AutoARIMA.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted AutoArima insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoARIMA.forecast
+
+> ``` text
+> AutoARIMA.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient AutoARIMA predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenpus of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x) optional exogenous. |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoARIMA.forward
+
+> ``` text
+> AutoARIMA.forward (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+*Apply fitted ARIMA model to a new time series.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import AutoARIMA
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# AutoARIMA's usage example
+arima = AutoARIMA(season_length=4)
+arima = arima.fit(y=ap)
+y_hat_dict = arima.predict(h=4, level=[80])
+y_hat_dict
+```
+
+## AutoETS
+
+------------------------------------------------------------------------
+
+source
+
+### AutoETS
+
+> ``` text
+> AutoETS (season_length:int=1, model:str='ZZZ',
+> damped:Optional[bool]=None, phi:Optional[float]=None,
+> alias:str='AutoETS', prediction_intervals:Optional[statsforecast
+> .utils.ConformalIntervals]=None)
+> ```
+
+\*Automatic Exponential Smoothing model.
+
+Automatically selects the best ETS (Error, Trend, Seasonality) model
+using an information criterion. Default is Akaike Information Criterion
+(AICc), while particular models are estimated using maximum likelihood.
+The state-space equations can be determined based on their $M$
+multiplicative, $A$ additive, $Z$ optimized or $N$ ommited components.
+The `model` string parameter defines the ETS equations: E in
+\[$M, A, Z$\], T in \[$N, A, M, Z$\], and S in \[$N, A, M, Z$\].
+
+For example when model=‘ANN’ (additive error, no trend, and no
+seasonality), ETS will explore only a simple exponential smoothing.
+
+If the component is selected as ‘Z’, it operates as a placeholder to ask
+the AutoETS model to figure out the best parameter.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| season_length | int | 1 | Number of observations per unit of time. Ex: 24 Hourly data. |
+| model | str | ZZZ | Controlling state-space-equations. |
+| damped | Optional | None | A parameter that ‘dampens’ the trend. |
+| phi | Optional | None | Smoothing parameter for trend damping. Only used when `damped=True`. |
+| alias | str | AutoETS | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoETS.fit
+
+> ``` text
+> AutoETS.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the Exponential Smoothing model.
+
+Fit an Exponential Smoothing model to a time series (numpy array) `y`
+and optionally exogenous variables (numpy array) `X`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|-----------------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | | | **Exponential Smoothing fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoETS.predict
+
+> ``` text
+> AutoETS.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted Exponential Smoothing.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenpus of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoETS.predict_in_sample
+
+> ``` text
+> AutoETS.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted Exponential Smoothing insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoETS.forecast
+
+> ``` text
+> AutoETS.forecast (y:numpy.ndarray, h:int, X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient Exponential Smoothing predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenpus of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoETS.forward
+
+> ``` text
+> AutoETS.forward (y:numpy.ndarray, h:int, X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+*Apply fitted Exponential Smoothing model to a new time series.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenpus of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import AutoETS
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# AutoETS' usage example
+# Multiplicative trend, optimal error and seasonality
+autoets = AutoETS(model='ZMZ', season_length=4)
+autoets = autoets.fit(y=ap)
+y_hat_dict = autoets.predict(h=4)
+y_hat_dict
+```
+
+## AutoCES
+
+------------------------------------------------------------------------
+
+source
+
+### AutoCES
+
+> ``` text
+> AutoCES (season_length:int=1, model:str='Z', alias:str='CES', prediction_
+> intervals:Optional[statsforecast.utils.ConformalIntervals]=None)
+> ```
+
+\*Complex Exponential Smoothing model.
+
+Automatically selects the best Complex Exponential Smoothing model using
+an information criterion. Default is Akaike Information Criterion
+(AICc), while particular models are estimated using maximum likelihood.
+The state-space equations can be determined based on their $S$ simple,
+$P$ parial, $Z$ optimized or $N$ ommited components. The `model` string
+parameter defines the kind of CES model: $N$ for simple CES (withous
+seasonality), $S$ for simple seasonality (lagged CES), $P$ for partial
+seasonality (without complex part), $F$ for full seasonality (lagged CES
+with real and complex seasonal parts).
+
+If the component is selected as ‘Z’, it operates as a placeholder to ask
+the AutoCES model to figure out the best parameter.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| season_length | int | 1 | Number of observations per unit of time. Ex: 24 Hourly data. |
+| model | str | Z | Controlling state-space-equations. |
+| alias | str | CES | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoCES.fit
+
+> ``` text
+> AutoCES.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the Complex Exponential Smoothing model.
+
+Fit the Complex Exponential Smoothing model to a time series (numpy
+array) `y` and optionally exogenous variables (numpy array) `X`.\*
+
+| | **Type** | **Default** | **Details** |
+|----|----|----|----|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | | | **Complex Exponential Smoothing fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoCES.predict
+
+> ``` text
+> AutoCES.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted Exponential Smoothing.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoCES.predict_in_sample
+
+> ``` text
+> AutoCES.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted Exponential Smoothing insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoCES.forecast
+
+> ``` text
+> AutoCES.forecast (y:numpy.ndarray, h:int, X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient Complex Exponential Smoothing predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenpus of shape (h, n_x). |
+| level | Optional | None | |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoCES.forward
+
+> ``` text
+> AutoCES.forward (y:numpy.ndarray, h:int, X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+*Apply fitted Complex Exponential Smoothing to a new time series.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenpus of shape (h, n_x). |
+| level | Optional | None | |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import AutoCES
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# CES' usage example
+# Multiplicative trend, optimal error and seasonality
+ces = AutoCES(model='Z',
+ season_length=4)
+ces = ces.fit(y=ap)
+y_hat_dict = ces.predict(h=4)
+y_hat_dict
+```
+
+## AutoTheta
+
+------------------------------------------------------------------------
+
+source
+
+### AutoTheta
+
+> ``` text
+> AutoTheta (season_length:int=1, decomposition_type:str='multiplicative',
+> model:Optional[str]=None, alias:str='AutoTheta', prediction_in
+> tervals:Optional[statsforecast.utils.ConformalIntervals]=None)
+> ```
+
+\*AutoTheta model.
+
+Automatically selects the best Theta (Standard Theta Model (‘STM’),
+Optimized Theta Model (‘OTM’), Dynamic Standard Theta Model (‘DSTM’),
+Dynamic Optimized Theta Model (‘DOTM’)) model using mse.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| season_length | int | 1 | Number of observations per unit of time. Ex: 24 Hourly data. |
+| decomposition_type | str | multiplicative | Sesonal decomposition type, ‘multiplicative’ (default) or ‘additive’. |
+| model | Optional | None | Controlling Theta Model. By default searchs the best model. |
+| alias | str | AutoTheta | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoTheta.fit
+
+> ``` text
+> AutoTheta.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the AutoTheta model.
+
+Fit an AutoTheta model to a time series (numpy array) `y` and optionally
+exogenous variables (numpy array) `X`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|---------------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | | | **AutoTheta fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoTheta.predict
+
+> ``` text
+> AutoTheta.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted AutoTheta.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoTheta.predict_in_sample
+
+> ``` text
+> AutoTheta.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted AutoTheta insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoTheta.forecast
+
+> ``` text
+> AutoTheta.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient AutoTheta predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoTheta.forward
+
+> ``` text
+> AutoTheta.forward (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+*Apply fitted AutoTheta to a new time series.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import AutoTheta
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# AutoTheta's usage example
+theta = AutoTheta(season_length=4)
+theta = theta.fit(y=ap)
+y_hat_dict = theta.predict(h=4)
+y_hat_dict
+```
+
+## AutoMFLES
+
+------------------------------------------------------------------------
+
+source
+
+### AutoMFLES
+
+> ``` text
+> AutoMFLES (test_size:int,
+> season_length:Union[int,List[int],NoneType]=None,
+> n_windows:int=2, config:Optional[Dict[str,Any]]=None,
+> step_size:Optional[int]=None, metric:str='smape',
+> verbose:bool=False, prediction_intervals:Optional[statsforecas
+> t.utils.ConformalIntervals]=None, alias:str='AutoMFLES')
+> ```
+
+*AutoMFLES*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| test_size | int | | Forecast horizon used during cross validation. |
+| season_length | Union | None | Number of observations per unit of time. Ex: 24 Hourly data. |
+| n_windows | int | 2 | Number of windows used for cross validation. |
+| config | Optional | None | Mapping from parameter name (from the init arguments of MFLES) to a list of values to try. If `None`, will use defaults. |
+| step_size | Optional | None | Step size between each cross validation window. If `None` will be set to test_size. |
+| metric | str | smape | Metric used to select the best model. Possible options are: ‘smape’, ‘mape’, ‘mse’ and ‘mae’. |
+| verbose | bool | False | Print debugging information. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. This is required for generating future prediction intervals. |
+| alias | str | AutoMFLES | Custom name of the model. |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoMFLES.fit
+
+> ``` text
+> AutoMFLES.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+*Fit the model*
+
+| | **Type** | **Default** | **Details** |
+|-------------|---------------|-------------|-----------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Exogenous of shape (t, n_x). |
+| **Returns** | **AutoMFLES** | | **Fitted AutoMFLES object.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoMFLES.predict
+
+> ``` text
+> AutoMFLES.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted AutoMFLES.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Exogenous of shape (h, n_x). |
+| level | Optional | None | |
+| **Returns** | **Dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoMFLES.predict_in_sample
+
+> ``` text
+> AutoMFLES.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted AutoMFLES insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **Dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoMFLES.forecast
+
+> ``` text
+> AutoMFLES.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient AutoMFLES predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **Dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+## AutoTBATS
+
+------------------------------------------------------------------------
+
+source
+
+### AutoTBATS
+
+> ``` text
+> AutoTBATS (season_length:Union[int,List[int]],
+> use_boxcox:Optional[bool]=None, bc_lower_bound:float=0.0,
+> bc_upper_bound:float=1.0, use_trend:Optional[bool]=None,
+> use_damped_trend:Optional[bool]=None,
+> use_arma_errors:bool=True, alias:str='AutoTBATS')
+> ```
+
+\*AutoTBATS model.
+
+Automatically selects the best TBATS model from all feasible
+combinations of the parameters use_boxcox, use_trend, use_damped_trend,
+and use_arma_errors. Selection is made using the AIC. Default value for
+use_arma_errors is True since this enables the evaluation of models with
+and without ARMA errors.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| season_length | Union | | |
+| use_boxcox | Optional | None | Whether or not to use a Box-Cox transformation. By default tries both. |
+| bc_lower_bound | float | 0.0 | Lower bound for the Box-Cox transformation. |
+| bc_upper_bound | float | 1.0 | Upper bound for the Box-Cox transformation. |
+| use_trend | Optional | None | Whether or not to use a trend component. By default tries both. |
+| use_damped_trend | Optional | None | Whether or not to dampen the trend component. By default tries both. |
+| use_arma_errors | bool | True | Whether or not to use a ARMA errors. Default is True and this evaluates both models. |
+| alias | str | AutoTBATS | Custom name of the model. |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoTBATS.fit
+
+> ``` text
+> AutoTBATS.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit TBATS model.
+
+Fit TBATS model to a time series (numpy array) `y`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|-----------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Ignored |
+| **Returns** | | | **TBATS model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoTBATS.predict
+
+> ``` text
+> AutoTBATS.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted TBATS model.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoTBATS.predict_in_sample
+
+> ``` text
+> AutoTBATS.predict_in_sample (level:Optional[Tuple[int]]=None)
+> ```
+
+*Access fitted TBATS model predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoTBATS.forecast
+
+> ``` text
+> AutoTBATS.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient TBATS model.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | |
+| X_future | Optional | None | |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+# ARIMA family
+
+## ARIMA
+
+------------------------------------------------------------------------
+
+source
+
+### ARIMA
+
+> ``` text
+> ARIMA (order:Tuple[int,int,int]=(0, 0, 0), season_length:int=1,
+> seasonal_order:Tuple[int,int,int]=(0, 0, 0),
+> include_mean:bool=True, include_drift:bool=False,
+> include_constant:Optional[bool]=None,
+> blambda:Optional[float]=None, biasadj:bool=False, method:str='CSS-
+> ML', fixed:Optional[dict]=None, alias:str='ARIMA', prediction_inte
+> rvals:Optional[statsforecast.utils.ConformalIntervals]=None)
+> ```
+
+\*ARIMA model.
+
+AutoRegressive Integrated Moving Average model.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| order | Tuple | (0, 0, 0) | A specification of the non-seasonal part of the ARIMA model: the three components (p, d, q) are the AR order, the degree of differencing, and the MA order. |
+| season_length | int | 1 | Number of observations per unit of time. Ex: 24 Hourly data. |
+| seasonal_order | Tuple | (0, 0, 0) | A specification of the seasonal part of the ARIMA model. (P, D, Q) for the AR order, the degree of differencing, the MA order. |
+| include_mean | bool | True | Should the ARIMA model include a mean term? The default is True for undifferenced series, False for differenced ones (where a mean would not affect the fit nor predictions). |
+| include_drift | bool | False | Should the ARIMA model include a linear drift term? (i.e., a linear regression with ARIMA errors is fitted.) |
+| include_constant | Optional | None | If True, then includ_mean is set to be True for undifferenced series and include_drift is set to be True for differenced series. Note that if there is more than one difference taken, no constant is included regardless of the value of this argument. This is deliberate as otherwise quadratic and higher order polynomial trends would be induced. |
+| blambda | Optional | None | Box-Cox transformation parameter. |
+| biasadj | bool | False | Use adjusted back-transformed mean Box-Cox. |
+| method | str | CSS-ML | Fitting method: maximum likelihood or minimize conditional sum-of-squares. The default (unless there are missing values) is to use conditional-sum-of-squares to find starting values, then maximum likelihood. |
+| fixed | Optional | None | Dictionary containing fixed coefficients for the arima model. Example: `{'ar1': 0.5, 'ma2': 0.75}`. For autoregressive terms use the `ar{i}` keys. For its seasonal version use `sar{i}`. For moving average terms use the `ma{i}` keys. For its seasonal version use `sma{i}`. For intercept and drift use the `intercept` and `drift` keys. For exogenous variables use the `ex_{i}` keys. |
+| alias | str | ARIMA | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### ARIMA.fit
+
+> ``` text
+> ARIMA.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+*Fit the model to a time series (numpy array) `y` and optionally
+exogenous variables (numpy array) `X`.*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|---------------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | | | **Fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### ARIMA.predict
+
+> ``` text
+> ARIMA.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted model.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### ARIMA.predict_in_sample
+
+> ``` text
+> ARIMA.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### ARIMA.forecast
+
+> ``` text
+> ARIMA.forecast (y:numpy.ndarray, h:int, X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory efficient predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x) optional exogenous. |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### ARIMA.forward
+
+> ``` text
+> ARIMA.forward (y:numpy.ndarray, h:int, X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+*Apply fitted model to a new time series.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import ARIMA
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# ARIMA's usage example
+arima = ARIMA(order=(1, 0, 0), season_length=12)
+arima = arima.fit(y=ap)
+y_hat_dict = arima.predict(h=4, level=[80])
+y_hat_dict
+```
+
+## AutoRegressive
+
+------------------------------------------------------------------------
+
+source
+
+### AutoRegressive
+
+> ``` text
+> AutoRegressive (lags:Tuple[int,List], include_mean:bool=True,
+> include_drift:bool=False, blambda:Optional[float]=None,
+> biasadj:bool=False, method:str='CSS-ML',
+> fixed:Optional[dict]=None, alias:str='AutoRegressive', pr
+> ediction_intervals:Optional[statsforecast.utils.Conformal
+> Intervals]=None)
+> ```
+
+*Simple Autoregressive model.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| lags | Tuple | | Number of lags to include in the model. If an int is passed then all lags up to `lags` are considered. If a list, only the elements of the list are considered as lags. |
+| include_mean | bool | True | Should the AutoRegressive model include a mean term? The default is True for undifferenced series, False for differenced ones (where a mean would not affect the fit nor predictions). |
+| include_drift | bool | False | Should the AutoRegressive model include a linear drift term? (i.e., a linear regression with AutoRegressive errors is fitted.) |
+| blambda | Optional | None | Box-Cox transformation parameter. |
+| biasadj | bool | False | Use adjusted back-transformed mean Box-Cox. |
+| method | str | CSS-ML | Fitting method: maximum likelihood or minimize conditional sum-of-squares. The default (unless there are missing values) is to use conditional-sum-of-squares to find starting values, then maximum likelihood. |
+| fixed | Optional | None | Dictionary containing fixed coefficients for the AutoRegressive model. Example: `{'ar1': 0.5, 'ar5': 0.75}`. For autoregressive terms use the `ar{i}` keys. |
+| alias | str | AutoRegressive | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoRegressive.fit
+
+> ``` text
+> AutoRegressive.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+*Fit the model to a time series (numpy array) `y` and optionally
+exogenous variables (numpy array) `X`.*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|---------------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | | | **Fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoRegressive.predict
+
+> ``` text
+> AutoRegressive.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted model.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoRegressive.predict_in_sample
+
+> ``` text
+> AutoRegressive.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoRegressive.forecast
+
+> ``` text
+> AutoRegressive.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None,
+> fitted:bool=False)
+> ```
+
+\*Memory efficient predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x) optional exogenous. |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### AutoRegressive.forward
+
+> ``` text
+> AutoRegressive.forward (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None,
+> fitted:bool=False)
+> ```
+
+*Apply fitted model to a new time series.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import AutoRegressive
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# AutoRegressive's usage example
+ar = AutoRegressive(lags=[12])
+ar = ar.fit(y=ap)
+y_hat_dict = ar.predict(h=4, level=[80])
+y_hat_dict
+```
+
+# ExponentialSmoothing
+
+## SimpleSmooth
+
+------------------------------------------------------------------------
+
+source
+
+### SimpleExponentialSmoothing
+
+> ``` text
+> SimpleExponentialSmoothing (alpha:float, alias:str='SES', prediction_inte
+> rvals:Optional[statsforecast.utils.ConformalI
+> ntervals]=None)
+> ```
+
+\*SimpleExponentialSmoothing model.
+
+Uses a weighted average of all past observations where the weights
+decrease exponentially into the past. Suitable for data with no clear
+trend or seasonality. Assuming there are $t$ observations, the one-step
+forecast is given by:
+$\hat{y}_{t+1} = \alpha y_t + (1-\alpha) \hat{y}_{t-1}$
+
+The rate $0 \leq \alpha \leq 1$ at which the weights decrease is called
+the smoothing parameter. When $\alpha = 1$, SES is equal to the naive
+method.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| alpha | float | | Smoothing parameter. |
+| alias | str | SES | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### SimpleExponentialSmoothing.forecast
+
+> ``` text
+> SimpleExponentialSmoothing.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None, X_fu
+> ture:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None,
+> fitted:bool=False)
+> ```
+
+\*Memory Efficient SimpleExponentialSmoothing predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### SimpleExponentialSmoothing.fit
+
+> ``` text
+> SimpleExponentialSmoothing.fit (y:numpy.ndarray,
+> X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the SimpleExponentialSmoothing model.
+
+Fit an SimpleExponentialSmoothing to a time series (numpy array) `y` and
+optionally exogenous variables (numpy array) `X`.\*
+
+| | **Type** | **Default** | **Details** |
+|----|----|----|----|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | | | **SimpleExponentialSmoothing fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### SimpleExponentialSmoothing.predict
+
+> ``` text
+> SimpleExponentialSmoothing.predict (h:int,
+> X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted SimpleExponentialSmoothing.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### SimpleExponentialSmoothing.predict_in_sample
+
+> ``` text
+> SimpleExponentialSmoothing.predict_in_sample ()
+> ```
+
+*Access fitted SimpleExponentialSmoothing insample predictions.*
+
+
+```python
+from statsforecast.models import SimpleExponentialSmoothing
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# SimpleExponentialSmoothing's usage example
+ses = SimpleExponentialSmoothing(alpha=0.5)
+ses = ses.fit(y=ap)
+y_hat_dict = ses.predict(h=4)
+y_hat_dict
+```
+
+## SimpleSmoothOptimized
+
+------------------------------------------------------------------------
+
+source
+
+### SimpleExponentialSmoothingOptimized
+
+> ``` text
+> SimpleExponentialSmoothingOptimized (alias:str='SESOpt', prediction_inter
+> vals:Optional[statsforecast.utils.Co
+> nformalIntervals]=None)
+> ```
+
+\*SimpleExponentialSmoothing model.
+
+Uses a weighted average of all past observations where the weights
+decrease exponentially into the past. Suitable for data with no clear
+trend or seasonality. Assuming there are $t$ observations, the one-step
+forecast is given by:
+$\hat{y}_{t+1} = \alpha y_t + (1-\alpha) \hat{y}_{t-1}$
+
+The smoothing parameter $\alpha^*$ is optimized by square error
+minimization.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| alias | str | SESOpt | |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. This is required for generating future prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### SimpleExponentialSmoothingOptimized.fit
+
+> ``` text
+> SimpleExponentialSmoothingOptimized.fit (y:numpy.ndarray,
+> X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the SimpleExponentialSmoothingOptimized model.
+
+Fit an SimpleExponentialSmoothingOptimized to a time series (numpy
+array) `y` and optionally exogenous variables (numpy array) `X`.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | | | **SimpleExponentialSmoothingOptimized fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### SimpleExponentialSmoothingOptimized.predict
+
+> ``` text
+> SimpleExponentialSmoothingOptimized.predict (h:int,
+> X:Optional[numpy.ndarray]=No
+> ne, level:Optional[List[int]
+> ]=None)
+> ```
+
+*Predict with fitted SimpleExponentialSmoothingOptimized.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### SimpleExponentialSmoothingOptimized.predict_in_sample
+
+> ``` text
+> SimpleExponentialSmoothingOptimized.predict_in_sample ()
+> ```
+
+*Access fitted SimpleExponentialSmoothingOptimized insample
+predictions.*
+
+------------------------------------------------------------------------
+
+source
+
+### SimpleExponentialSmoothingOptimized.forecast
+
+> ``` text
+> SimpleExponentialSmoothingOptimized.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=N
+> one, X_future:Optional[nump
+> y.ndarray]=None, level:Opti
+> onal[List[int]]=None,
+> fitted:bool=False)
+> ```
+
+\*Memory Efficient SimpleExponentialSmoothingOptimized predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import SimpleExponentialSmoothingOptimized
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# SimpleExponentialSmoothingOptimized's usage example
+seso = SimpleExponentialSmoothingOptimized()
+seso = seso.fit(y=ap)
+y_hat_dict = seso.predict(h=4)
+y_hat_dict
+```
+
+## SeasonalSmooth
+
+
+```python
+plt.plot(np.concatenate([ap[6:], seas_es.forecast(ap[6:], h=12)['mean']]))
+```
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalExponentialSmoothing
+
+> ``` text
+> SeasonalExponentialSmoothing (season_length:int, alpha:float,
+> alias:str='SeasonalES', prediction_interval
+> s:Optional[statsforecast.utils.ConformalInt
+> ervals]=None)
+> ```
+
+\*SeasonalExponentialSmoothing model.
+
+Uses a weighted average of all past observations where the weights
+decrease exponentially into the past. Suitable for data with no clear
+trend or seasonality. Assuming there are $t$ observations and season
+$s$, the one-step forecast is given by:
+$\hat{y}_{t+1,s} = \alpha y_t + (1-\alpha) \hat{y}_{t-1,s}$\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| season_length | int | | Number of observations per unit of time. Ex: 24 Hourly data. |
+| alpha | float | | Smoothing parameter. |
+| alias | str | SeasonalES | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. This is required for generating future prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalExponentialSmoothing.fit
+
+> ``` text
+> SeasonalExponentialSmoothing.fit (y:numpy.ndarray,
+> X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the SeasonalExponentialSmoothing model.
+
+Fit an SeasonalExponentialSmoothing to a time series (numpy array) `y`
+and optionally exogenous variables (numpy array) `X`.\*
+
+| | **Type** | **Default** | **Details** |
+|----|----|----|----|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | | | **SeasonalExponentialSmoothing fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalExponentialSmoothing.predict
+
+> ``` text
+> SeasonalExponentialSmoothing.predict (h:int,
+> X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted SeasonalExponentialSmoothing.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalExponentialSmoothing.predict_in_sample
+
+> ``` text
+> SeasonalExponentialSmoothing.predict_in_sample ()
+> ```
+
+*Access fitted SeasonalExponentialSmoothing insample predictions.*
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalExponentialSmoothing.forecast
+
+> ``` text
+> SeasonalExponentialSmoothing.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None, X_
+> future:Optional[numpy.ndarray]=Non
+> e, level:Optional[List[int]]=None,
+> fitted:bool=False)
+> ```
+
+\*Memory Efficient SeasonalExponentialSmoothing predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import SeasonalExponentialSmoothing
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# SeasonalExponentialSmoothing's usage example
+model = SeasonalExponentialSmoothing(alpha=0.5, season_length=12)
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+## SeasonalSmoothOptimized
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalExponentialSmoothingOptimized
+
+> ``` text
+> SeasonalExponentialSmoothingOptimized (season_length:int,
+> alias:str='SeasESOpt', prediction_
+> intervals:Optional[statsforecast.u
+> tils.ConformalIntervals]=None)
+> ```
+
+\*SeasonalExponentialSmoothingOptimized model.
+
+Uses a weighted average of all past observations where the weights
+decrease exponentially into the past. Suitable for data with no clear
+trend or seasonality. Assuming there are $t$ observations and season
+$s$, the one-step forecast is given by:
+$\hat{y}_{t+1,s} = \alpha y_t + (1-\alpha) \hat{y}_{t-1,s}$
+
+The smoothing parameter $\alpha^*$ is optimized by square error
+minimization.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| season_length | int | | Number of observations per unit of time. Ex: 24 Hourly data. |
+| alias | str | SeasESOpt | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. This is required for generating future prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalExponentialSmoothingOptimized.forecast
+
+> ``` text
+> SeasonalExponentialSmoothingOptimized.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]
+> =None, X_future:Optional[
+> numpy.ndarray]=None, leve
+> l:Optional[List[int]]=Non
+> e, fitted:bool=False)
+> ```
+
+\*Memory Efficient SeasonalExponentialSmoothingOptimized predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalExponentialSmoothingOptimized.fit
+
+> ``` text
+> SeasonalExponentialSmoothingOptimized.fit (y:numpy.ndarray,
+> X:Optional[numpy.ndarray]=None
+> )
+> ```
+
+\*Fit the SeasonalExponentialSmoothingOptimized model.
+
+Fit an SeasonalExponentialSmoothingOptimized to a time series (numpy
+array) `y` and optionally exogenous variables (numpy array) `X`.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | | | **SeasonalExponentialSmoothingOptimized fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalExponentialSmoothingOptimized.predict
+
+> ``` text
+> SeasonalExponentialSmoothingOptimized.predict (h:int,
+> X:Optional[numpy.ndarray]=
+> None, level:Optional[List[
+> int]]=None)
+> ```
+
+*Predict with fitted SeasonalExponentialSmoothingOptimized.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalExponentialSmoothingOptimized.predict_in_sample
+
+> ``` text
+> SeasonalExponentialSmoothingOptimized.predict_in_sample ()
+> ```
+
+*Access fitted SeasonalExponentialSmoothingOptimized insample
+predictions.*
+
+
+```python
+from statsforecast.models import SeasonalExponentialSmoothingOptimized
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# SeasonalExponentialSmoothingOptimized's usage example
+model = SeasonalExponentialSmoothingOptimized(season_length=12)
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+## Holt’s method
+
+------------------------------------------------------------------------
+
+source
+
+### Holt
+
+> ``` text
+> Holt (season_length:int=1, error_type:str='A', alias:str='Holt', predicti
+> on_intervals:Optional[statsforecast.utils.ConformalIntervals]=None)
+> ```
+
+\*Holt’s method.
+
+Also known as double exponential smoothing, Holt’s method is an
+extension of exponential smoothing for series with a trend. This
+implementation returns the corresponding `ETS` model with additive (A)
+or multiplicative (M) errors (so either ‘AAN’ or ‘MAN’).\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| season_length | int | 1 | Number of observations per unit of time. Ex: 12 Monthly data. |
+| error_type | str | A | The type of error of the ETS model. Can be additive (A) or multiplicative (M). |
+| alias | str | Holt | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### Holt.forecast
+
+> ``` text
+> Holt.forecast (y:numpy.ndarray, h:int, X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient Exponential Smoothing predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenpus of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### Holt.fit
+
+> ``` text
+> Holt.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the Exponential Smoothing model.
+
+Fit an Exponential Smoothing model to a time series (numpy array) `y`
+and optionally exogenous variables (numpy array) `X`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|-----------------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | | | **Exponential Smoothing fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### Holt.predict
+
+> ``` text
+> Holt.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted Exponential Smoothing.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenpus of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### Holt.predict_in_sample
+
+> ``` text
+> Holt.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted Exponential Smoothing insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### Holt.forward
+
+> ``` text
+> Holt.forward (y:numpy.ndarray, h:int, X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+*Apply fitted Exponential Smoothing model to a new time series.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenpus of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import Holt
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# Holt's usage example
+model = Holt(season_length=12, error_type='A')
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+## Holt-Winters’ method
+
+------------------------------------------------------------------------
+
+source
+
+### HoltWinters
+
+> ``` text
+> HoltWinters (season_length:int=1, error_type:str='A',
+> alias:str='HoltWinters', prediction_intervals:Optional[stats
+> forecast.utils.ConformalIntervals]=None)
+> ```
+
+\*Holt-Winters’ method.
+
+Also known as triple exponential smoothing, Holt-Winters’ method is an
+extension of exponential smoothing for series that contain both trend
+and seasonality. This implementation returns the corresponding `ETS`
+model with additive (A) or multiplicative (M) errors (so either ‘AAA’ or
+‘MAM’).\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| season_length | int | 1 | season length |
+| error_type | str | A | error type |
+| alias | str | HoltWinters | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### HoltWinters.forecast
+
+> ``` text
+> HoltWinters.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient Exponential Smoothing predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenpus of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### HoltWinters.fit
+
+> ``` text
+> HoltWinters.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the Exponential Smoothing model.
+
+Fit an Exponential Smoothing model to a time series (numpy array) `y`
+and optionally exogenous variables (numpy array) `X`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|-----------------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | | | **Exponential Smoothing fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### HoltWinters.predict
+
+> ``` text
+> HoltWinters.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted Exponential Smoothing.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenpus of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### HoltWinters.predict_in_sample
+
+> ``` text
+> HoltWinters.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted Exponential Smoothing insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### HoltWinters.forward
+
+> ``` text
+> HoltWinters.forward (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+*Apply fitted Exponential Smoothing model to a new time series.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenpus of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import HoltWinters
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# Holt-Winters' usage example
+model = HoltWinters(season_length=12, error_type='A')
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+# Baseline Models
+
+## HistoricAverage
+
+------------------------------------------------------------------------
+
+source
+
+### HistoricAverage
+
+> ``` text
+> HistoricAverage (alias:str='HistoricAverage', prediction_intervals:Option
+> al[statsforecast.utils.ConformalIntervals]=None)
+> ```
+
+\*HistoricAverage model.
+
+Also known as mean method. Uses a simple average of all past
+observations. Assuming there are $t$ observations, the one-step forecast
+is given by: $$\hat{y}_{t+1} = \frac{1}{t} \sum_{j=1}^t y_j$$\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| alias | str | HistoricAverage | |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### HistoricAverage.forecast
+
+> ``` text
+> HistoricAverage.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None,
+> fitted:bool=False)
+> ```
+
+\*Memory Efficient HistoricAverage predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### HistoricAverage.fit
+
+> ``` text
+> HistoricAverage.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the HistoricAverage model.
+
+Fit an HistoricAverage to a time series (numpy array) `y`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|---------------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | **self** | | **HistoricAverage fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### HistoricAverage.predict
+
+> ``` text
+> HistoricAverage.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted HistoricAverage.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### HistoricAverage.predict_in_sample
+
+> ``` text
+> HistoricAverage.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted HistoricAverage insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions.** |
+
+
+```python
+from statsforecast.models import HistoricAverage
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# HistoricAverage's usage example
+model = HistoricAverage()
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+## Naive
+
+------------------------------------------------------------------------
+
+source
+
+### Naive
+
+> ``` text
+> Naive (alias:str='Naive', prediction_intervals:Optional[statsforecast.uti
+> ls.ConformalIntervals]=None)
+> ```
+
+\*Naive model.
+
+All forecasts have the value of the last observation:
+$\hat{y}_{t+1} = y_t$ for all $t$\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| alias | str | Naive | |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### Naive.forecast
+
+> ``` text
+> Naive.forecast (y:numpy.ndarray, h:int, X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient Naive predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n,). |
+| h | int | | |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### Naive.fit
+
+> ``` text
+> Naive.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the Naive model.
+
+Fit an Naive to a time series (numpy.array) `y`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|-----------|-------------|---------------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | **self:** | | **Naive fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### Naive.predict
+
+> ``` text
+> Naive.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted Naive.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | forecasting horizon |
+| X | Optional | None | exogenous regressors |
+| level | Optional | None | confidence level |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### Naive.predict_in_sample
+
+> ``` text
+> Naive.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted Naive insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions.** |
+
+
+```python
+from statsforecast.models import Naive
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# Naive's usage example
+model = Naive()
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+## RandomWalkWithDrift
+
+------------------------------------------------------------------------
+
+source
+
+### RandomWalkWithDrift
+
+> ``` text
+> RandomWalkWithDrift (alias:str='RWD', prediction_intervals:Optional[stats
+> forecast.utils.ConformalIntervals]=None)
+> ```
+
+\*RandomWalkWithDrift model.
+
+A variation of the naive method allows the forecasts to change over
+time. The amout of change, called drift, is the average change seen in
+the historical data.
+
+$$\hat{y}_{t+1} = y_t+\frac{1}{t-1}\sum_{j=1}^t (y_j-y_{j-1}) = y_t+ \frac{y_t-y_1}{t-1}$$
+
+From the previous equation, we can see that this is equivalent to
+extrapolating a line between the first and the last observation.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| alias | str | RWD | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### RandomWalkWithDrift.forecast
+
+> ``` text
+> RandomWalkWithDrift.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None,
+> fitted:bool=False)
+> ```
+
+\*Memory Efficient RandomWalkWithDrift predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n,). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **forecasts: dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### RandomWalkWithDrift.fit
+
+> ``` text
+> RandomWalkWithDrift.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the RandomWalkWithDrift model.
+
+Fit an RandomWalkWithDrift to a time series (numpy array) `y`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|---------------------------------------|
+| y | ndarray | | |
+| X | Optional | None | |
+| **Returns** | | | **RandomWalkWithDrift fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### RandomWalkWithDrift.predict
+
+> ``` text
+> RandomWalkWithDrift.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted RandomWalkWithDrift.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### RandomWalkWithDrift.predict_in_sample
+
+> ``` text
+> RandomWalkWithDrift.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted RandomWalkWithDrift insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import RandomWalkWithDrift
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# RandomWalkWithDrift's usage example
+model = RandomWalkWithDrift()
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+## SeasonalNaive
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalNaive
+
+> ``` text
+> SeasonalNaive (season_length:int, alias:str='SeasonalNaive', prediction_i
+> ntervals:Optional[statsforecast.utils.ConformalIntervals]=
+> None)
+> ```
+
+\*Seasonal naive model.
+
+A method similar to the naive, but uses the last known observation of
+the same period (e.g. the same month of the previous year) in order to
+capture seasonal variations.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| season_length | int | | Number of observations per unit of time. Ex: 24 Hourly data. |
+| alias | str | SeasonalNaive | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalNaive.forecast
+
+> ``` text
+> SeasonalNaive.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None,
+> fitted:bool=False)
+> ```
+
+\*Memory Efficient SeasonalNaive predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalNaive.fit
+
+> ``` text
+> SeasonalNaive.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the SeasonalNaive model.
+
+Fit an SeasonalNaive to a time series (numpy array) `y`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|-----------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | |
+| **Returns** | | | **SeasonalNaive fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalNaive.predict
+
+> ``` text
+> SeasonalNaive.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted Naive.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | |
+| level | Optional | None | |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalNaive.predict_in_sample
+
+> ``` text
+> SeasonalNaive.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted SeasonalNaive insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import SeasonalNaive
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# SeasonalNaive's usage example
+model = SeasonalNaive(season_length=12)
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+## WindowAverage
+
+------------------------------------------------------------------------
+
+source
+
+### WindowAverage
+
+> ``` text
+> WindowAverage (window_size:int, alias:str='WindowAverage', prediction_int
+> ervals:Optional[statsforecast.utils.ConformalIntervals]=No
+> ne)
+> ```
+
+\*WindowAverage model.
+
+Uses the average of the last $k$ observations, with $k$ the length of
+the window. Wider windows will capture global trends, while narrow
+windows will reveal local trends. The length of the window selected
+should take into account the importance of past observations and how
+fast the series changes.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| window_size | int | | Size of truncated series on which average is estimated. |
+| alias | str | WindowAverage | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. This is required for generating future prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### WindowAverage.forecast
+
+> ``` text
+> WindowAverage.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None,
+> fitted:bool=False)
+> ```
+
+\*Memory Efficient WindowAverage predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### WindowAverage.fit
+
+> ``` text
+> WindowAverage.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the WindowAverage model.
+
+Fit an WindowAverage to a time series (numpy array) `y` and optionally
+exogenous variables (numpy array) `X`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|---------------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | | | **WindowAverage fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### WindowAverage.predict
+
+> ``` text
+> WindowAverage.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted WindowAverage.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import WindowAverage
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# WindowAverage's usage example
+model = WindowAverage(window_size=12*4)
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+## SeasonalWindowAverage
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalWindowAverage
+
+> ``` text
+> SeasonalWindowAverage (season_length:int, window_size:int,
+> alias:str='SeasWA', prediction_intervals:Optional[
+> statsforecast.utils.ConformalIntervals]=None)
+> ```
+
+\*SeasonalWindowAverage model.
+
+An average of the last $k$ observations of the same period, with $k$ the
+length of the window.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| season_length | int | | |
+| window_size | int | | Size of truncated series on which average is estimated. |
+| alias | str | SeasWA | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. This is required for generating future prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalWindowAverage.forecast
+
+> ``` text
+> SeasonalWindowAverage.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None,
+> fitted:bool=False)
+> ```
+
+\*Memory Efficient SeasonalWindowAverage predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n,). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalWindowAverage.fit
+
+> ``` text
+> SeasonalWindowAverage.fit (y:numpy.ndarray,
+> X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the SeasonalWindowAverage model.
+
+Fit an SeasonalWindowAverage to a time series (numpy array) `y` and
+optionally exogenous variables (numpy array) `X`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|-----------------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenpus of shape (t, n_x). |
+| **Returns** | | | **SeasonalWindowAverage fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### SeasonalWindowAverage.predict
+
+> ``` text
+> SeasonalWindowAverage.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted SeasonalWindowAverage.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import SeasonalWindowAverage
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# SeasonalWindowAverage's usage example
+model = SeasonalWindowAverage(season_length=12, window_size=4)
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+# Sparse or Intermittent
+
+## ADIDA
+
+------------------------------------------------------------------------
+
+source
+
+### ADIDA
+
+> ``` text
+> ADIDA (alias:str='ADIDA', prediction_intervals:Optional[statsforecast.uti
+> ls.ConformalIntervals]=None)
+> ```
+
+\*ADIDA model.
+
+Aggregate-Dissagregate Intermittent Demand Approach: Uses temporal
+aggregation to reduce the number of zero observations. Once the data has
+been agregated, it uses the optimized SES to generate the forecasts at
+the new level. It then breaks down the forecast to the original level
+using equal weights.
+
+ADIDA specializes on sparse or intermittent series are series with very
+few non-zero observations. They are notoriously hard to forecast, and
+so, different methods have been developed especifically for them.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| alias | str | ADIDA | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### ADIDA.forecast
+
+> ``` text
+> ADIDA.forecast (y:numpy.ndarray, h:int, X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient ADIDA predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n,). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### ADIDA.fit
+
+> ``` text
+> ADIDA.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the ADIDA model.
+
+Fit an ADIDA to a time series (numpy array) `y`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|-----------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | |
+| **Returns** | | | **ADIDA fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### ADIDA.predict
+
+> ``` text
+> ADIDA.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted ADIDA.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import ADIDA
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# ADIDA's usage example
+model = ADIDA()
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+## CrostonClassic
+
+------------------------------------------------------------------------
+
+source
+
+### CrostonClassic
+
+> ``` text
+> CrostonClassic (alias:str='CrostonClassic', prediction_intervals:Optional
+> [statsforecast.utils.ConformalIntervals]=None)
+> ```
+
+\*CrostonClassic model.
+
+A method to forecast time series that exhibit intermittent demand. It
+decomposes the original time series into a non-zero demand size $z_t$
+and inter-demand intervals $p_t$. Then the forecast is given by:
+$$\hat{y}_t = \frac{\hat{z}_t}{\hat{p}_t}$$
+
+where $\hat{z}_t$ and $\hat{p}_t$ are forecasted using SES. The
+smoothing parameter of both components is set equal to 0.1\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| alias | str | CrostonClassic | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### CrostonClassic.forecast
+
+> ``` text
+> CrostonClassic.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None,
+> fitted:bool=False)
+> ```
+
+\*Memory Efficient CrostonClassic predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### CrostonClassic.fit
+
+> ``` text
+> CrostonClassic.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the CrostonClassic model.
+
+Fit an CrostonClassic to a time series (numpy array) `y`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|-----------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | |
+| **Returns** | | | **CrostonClassic fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### CrostonClassic.predict
+
+> ``` text
+> CrostonClassic.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted CrostonClassic.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import CrostonClassic
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# CrostonClassic's usage example
+model = CrostonClassic()
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+## CrostonOptimized
+
+------------------------------------------------------------------------
+
+source
+
+### CrostonOptimized
+
+> ``` text
+> CrostonOptimized (alias:str='CrostonOptimized', prediction_intervals:Opti
+> onal[statsforecast.utils.ConformalIntervals]=None)
+> ```
+
+\*CrostonOptimized model.
+
+A method to forecast time series that exhibit intermittent demand. It
+decomposes the original time series into a non-zero demand size $z_t$
+and inter-demand intervals $p_t$. Then the forecast is given by:
+$$\hat{y}_t = \frac{\hat{z}_t}{\hat{p}_t}$$
+
+A variation of the classic Croston’s method where the smooting paramater
+is optimally selected from the range $[0.1,0.3]$. Both the non-zero
+demand $z_t$ and the inter-demand intervals $p_t$ are smoothed
+separately, so their smoothing parameters can be different.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| alias | str | CrostonOptimized | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. This is required for generating future prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### CrostonOptimized.forecast
+
+> ``` text
+> CrostonOptimized.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None,
+> fitted:bool=False)
+> ```
+
+\*Memory Efficient CrostonOptimized predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### CrostonOptimized.fit
+
+> ``` text
+> CrostonOptimized.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the CrostonOptimized model.
+
+Fit an CrostonOptimized to a time series (numpy array) `y`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|------------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | |
+| **Returns** | | | **CrostonOptimized fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### CrostonOptimized.predict
+
+> ``` text
+> CrostonOptimized.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted CrostonOptimized.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import CrostonOptimized
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# CrostonOptimized's usage example
+model = CrostonOptimized()
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+## CrostonSBA
+
+------------------------------------------------------------------------
+
+source
+
+### CrostonSBA
+
+> ``` text
+> CrostonSBA (alias:str='CrostonSBA', prediction_intervals:Optional[statsfo
+> recast.utils.ConformalIntervals]=None)
+> ```
+
+\*CrostonSBA model.
+
+A method to forecast time series that exhibit intermittent demand. It
+decomposes the original time series into a non-zero demand size $z_t$
+and inter-demand intervals $p_t$. Then the forecast is given by:
+$$\hat{y}_t = \frac{\hat{z}_t}{\hat{p}_t}$$
+
+A variation of the classic Croston’s method that uses a debiasing
+factor, so that the forecast is given by:
+$$\hat{y}_t = 0.95 \frac{\hat{z}_t}{\hat{p}_t}$$\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| alias | str | CrostonSBA | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### CrostonSBA.forecast
+
+> ``` text
+> CrostonSBA.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient CrostonSBA predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### CrostonSBA.fit
+
+> ``` text
+> CrostonSBA.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the CrostonSBA model.
+
+Fit an CrostonSBA to a time series (numpy array) `y`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|-----------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | |
+| **Returns** | | | **CrostonSBA fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### CrostonSBA.predict
+
+> ``` text
+> CrostonSBA.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted CrostonSBA.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import CrostonSBA
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# CrostonSBA's usage example
+model = CrostonSBA()
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+## IMAPA
+
+------------------------------------------------------------------------
+
+source
+
+### IMAPA
+
+> ``` text
+> IMAPA (alias:str='IMAPA', prediction_intervals:Optional[statsforecast.uti
+> ls.ConformalIntervals]=None)
+> ```
+
+\*IMAPA model.
+
+Intermittent Multiple Aggregation Prediction Algorithm: Similar to
+ADIDA, but instead of using a single aggregation level, it considers
+multiple in order to capture different dynamics of the data. Uses the
+optimized SES to generate the forecasts at the new levels and then
+combines them using a simple average.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| alias | str | IMAPA | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### IMAPA.forecast
+
+> ``` text
+> IMAPA.forecast (y:numpy.ndarray, h:int, X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient IMAPA predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### IMAPA.fit
+
+> ``` text
+> IMAPA.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the IMAPA model.
+
+Fit an IMAPA to a time series (numpy array) `y`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|-----------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | |
+| **Returns** | | | **IMAPA fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### IMAPA.predict
+
+> ``` text
+> IMAPA.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted IMAPA.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | |
+| level | Optional | None | |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import IMAPA
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# IMAPA's usage example
+model = IMAPA()
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+## TSB
+
+------------------------------------------------------------------------
+
+source
+
+### TSB
+
+> ``` text
+> TSB (alpha_d:float, alpha_p:float, alias:str='TSB', prediction_intervals:
+> Optional[statsforecast.utils.ConformalIntervals]=None)
+> ```
+
+\*TSB model.
+
+Teunter-Syntetos-Babai: A modification of Croston’s method that replaces
+the inter-demand intervals with the demand probability $d_t$, which is
+defined as follows.
+
+$$
+
+d_t = \begin{cases}
+ 1 & \text{if demand occurs at time t} \\
+ 0 & \text{otherwise.}
+\end{cases}
+
+$$
+
+Hence, the forecast is given by
+
+$$\hat{y}_t= \hat{d}_t\hat{z_t}$$
+
+Both $d_t$ and $z_t$ are forecasted using SES. The smooting paramaters
+of each may differ, like in the optimized Croston’s method.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| alpha_d | float | | Smoothing parameter for demand. |
+| alpha_p | float | | Smoothing parameter for probability. |
+| alias | str | TSB | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. This is required for generating future prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### TSB.forecast
+
+> ``` text
+> TSB.forecast (y:numpy.ndarray, h:int, X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient TSB predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### TSB.fit
+
+> ``` text
+> TSB.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the TSB model.
+
+Fit an TSB to a time series (numpy array) `y`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|-----------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | |
+| **Returns** | | | **TSB fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### TSB.predict
+
+> ``` text
+> TSB.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted TSB.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import TSB
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# TSB's usage example
+model = TSB(alpha_d=0.5, alpha_p=0.5)
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+# Multiple Seasonalities
+
+## MSTL
+
+------------------------------------------------------------------------
+
+source
+
+### MSTL
+
+> ``` text
+> MSTL (season_length:Union[int,List[int]], trend_forecaster=AutoETS,
+> stl_kwargs:Optional[Dict]=None, alias:str='MSTL', prediction_interv
+> als:Optional[statsforecast.utils.ConformalIntervals]=None)
+> ```
+
+\*MSTL model.
+
+The MSTL (Multiple Seasonal-Trend decomposition using LOESS) decomposes
+the time series in multiple seasonalities using LOESS. Then forecasts
+the trend using a custom non-seaonal model and each seasonality using a
+SeasonalNaive model.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| season_length | Union | | Number of observations per unit of time. For multiple seasonalities use a list. |
+| trend_forecaster | AutoETS | AutoETS | StatsForecast model used to forecast the trend component. |
+| stl_kwargs | Optional | None | Extra arguments to pass to [`statsmodels.tsa.seasonal.STL`](https://www.statsmodels.org/dev/generated/statsmodels.tsa.seasonal.STL.html#statsmodels.tsa.seasonal.STL). The `period` and `seasonal` arguments are reserved. |
+| alias | str | MSTL | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### MSTL.fit
+
+> ``` text
+> MSTL.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the MSTL model.
+
+Fit MSTL to a time series (numpy array) `y`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|-----------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | |
+| **Returns** | | | **MSTL fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### MSTL.predict
+
+> ``` text
+> MSTL.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted MSTL.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### MSTL.predict_in_sample
+
+> ``` text
+> MSTL.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted MSTL insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### MSTL.forecast
+
+> ``` text
+> MSTL.forecast (y:numpy.ndarray, h:int, X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient MSTL predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### MSTL.forward
+
+> ``` text
+> MSTL.forward (y:numpy.ndarray, h:int, X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+*Apply fitted MSTL model to a new time series.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import MSTL
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# MSTL's usage example
+mstl_model = MSTL(season_length=[3, 12], trend_forecaster=AutoARIMA(prediction_intervals=ConformalIntervals(h=4, n_windows=2)))
+mstl_model = mstl_model.fit(y=ap)
+y_hat_dict = mstl_model.predict(h=4, level=[80])
+y_hat_dict
+```
+
+## MFLES
+
+------------------------------------------------------------------------
+
+source
+
+### MFLES
+
+> ``` text
+> MFLES (season_length:Union[int,List[int],NoneType]=None,
+> fourier_order:Optional[int]=None, max_rounds:int=50,
+> ma:Optional[int]=None, alpha:float=1.0, decay:float=-1.0,
+> changepoints:bool=True, n_changepoints:Union[float,int]=0.25,
+> seasonal_lr:float=0.9, trend_lr:float=0.9, exogenous_lr:float=1.0,
+> residuals_lr:float=1.0, cov_threshold:float=0.7,
+> moving_medians:bool=False, min_alpha:float=0.05,
+> max_alpha:float=1.0, trend_penalty:bool=True,
+> multiplicative:Optional[bool]=None, smoother:bool=False,
+> robust:Optional[bool]=None, verbose:bool=False, prediction_interva
+> ls:Optional[statsforecast.utils.ConformalIntervals]=None,
+> alias:str='MFLES')
+> ```
+
+\*MFLES model.
+
+A method to forecast time series based on Gradient Boosted Time Series
+Decomposition which treats traditional decomposition as the base
+estimator in the boosting process. Unlike normal gradient boosting,
+slight learning rates are applied at the component level
+(trend/seasonality/exogenous).
+
+The method derives its name from some of the underlying estimators that
+can enter into the boosting procedure, specifically: a simple Median,
+Fourier functions for seasonality, a simple/piecewise Linear trend, and
+Exponential Smoothing.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| season_length | Union | None | Number of observations per unit of time. Ex: 24 Hourly data. |
+| fourier_order | Optional | None | How many fourier sin/cos pairs to create, the larger the number the more complex of a seasonal pattern can be fitted. A lower number leads to smoother results. This is auto-set based on seasonal_period. |
+| max_rounds | int | 50 | The max number of boosting rounds. The boosting will auto-stop but depending on other parameters such as rs_lr you may want more rounds. Generally more rounds means a smoother fit. |
+| ma | Optional | None | The moving average order to use, this is auto-set based on internal logic. Passing 4 would fit a 4 period moving average on the residual component. |
+| alpha | float | 1.0 | The alpha which is used in fitting the underlying LASSO when using piecewise functions. |
+| decay | float | -1.0 | Effects the slopes of the piecewise-linear basis function. |
+| changepoints | bool | True | Whether to fit for changepoints if all other logic allows for it. If False, MFLES will not ever fit a piecewise trend. |
+| n_changepoints | Union | 0.25 | Number (if int) or proportion (if float) of changepoint knots to place. The default of 0.25 will place 0.25 \* (series length) number of knots. |
+| seasonal_lr | float | 0.9 | A shrinkage parameter (0 \< seasonal_lr \<= 1) which penalizes the seasonal fit. A value of 0.9 will flatly multiply the seasonal fit by 0.9 each boosting round, this can be used to allow more signal to the exogenous component. |
+| trend_lr | float | 0.9 | A shrinkage parameter (0 \< trend_lr \<= 1) which penalizes the linear trend fit A value of 0.9 will flatly multiply the linear fit by 0.9 each boosting round, this can be used to allow more signal to the seasonality or exogenous components. |
+| exogenous_lr | float | 1.0 | The shrinkage parameter (0 \< exogenous_lr \<= 1) which controls how much of the exogenous signal is carried to the next round. |
+| residuals_lr | float | 1.0 | A shrinkage parameter (0 \< residuals_lr \<= 1) which penalizes the residual smoothing. A value of 0.9 will flatly multiply the residual fit by 0.9 each boosting round, this can be used to allow more signal to the seasonality or linear components. |
+| cov_threshold | float | 0.7 | The deseasonalized cov is used to auto-set some logic, lowering the cov_threshold will result in simpler and less complex residual smoothing. If you pass something like 1000 then there will be no safeguards applied. |
+| moving_medians | bool | False | The default behavior is to fit an initial median to the time series. If True, then it will fit a median per seasonal period. |
+| min_alpha | float | 0.05 | The minimum alpha in the SES ensemble. |
+| max_alpha | float | 1.0 | The maximum alpha used in the SES ensemble. |
+| trend_penalty | bool | True | Whether to apply a simple penalty to the linear trend component, very useful for dealing with the potentially dangerous piecewise trend. |
+| multiplicative | Optional | None | Auto-set based on internal logic. If True, it will simply take the log of the time series. |
+| smoother | bool | False | If True, then a simple exponential ensemble will be used rather than auto settings. |
+| robust | Optional | None | If True then MFLES will fit using more reserved methods, i.e. not using piecewise trend or moving average residual smoother. Auto-set based on internal logic. |
+| verbose | bool | False | Print debugging information. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. This is required for generating future prediction intervals. |
+| alias | str | MFLES | Custom name of the model. |
+
+------------------------------------------------------------------------
+
+source
+
+### MFLES.fit
+
+> ``` text
+> MFLES.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+*Fit the model*
+
+| | **Type** | **Default** | **Details** |
+|-------------|-----------|-------------|-----------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Exogenous of shape (t, n_x). |
+| **Returns** | **MFLES** | | **Fitted MFLES object.** |
+
+------------------------------------------------------------------------
+
+source
+
+### MFLES.predict
+
+> ``` text
+> MFLES.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted MFLES.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Exogenous of shape (h, n_x). |
+| level | Optional | None | |
+| **Returns** | **Dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### MFLES.predict_in_sample
+
+> ``` text
+> MFLES.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted SklearnModel insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **Dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### MFLES.forecast
+
+> ``` text
+> MFLES.forecast (y:numpy.ndarray, h:int, X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient MFLES predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **Dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+## TBATS
+
+------------------------------------------------------------------------
+
+source
+
+### TBATS
+
+> ``` text
+> TBATS (season_length:Union[int,List[int]],
+> use_boxcox:Optional[bool]=True, bc_lower_bound:float=0.0,
+> bc_upper_bound:float=1.0, use_trend:Optional[bool]=True,
+> use_damped_trend:Optional[bool]=False, use_arma_errors:bool=False,
+> alias:str='TBATS')
+> ```
+
+\*Trigonometric Box-Cox transform, ARMA errors, Trend and Seasonal
+components (TBATS) model.
+
+TBATS is an innovations state space model framework used for forecasting
+time series with multiple seasonalities. It uses a Box-Cox
+tranformation, ARMA errors, and a trigonometric representation of the
+seasonal patterns based on Fourier series.
+
+The name TBATS is an acronym for the key features of the model:
+Trigonometric, Box-Cox transform, ARMA errors, Trend, and Seasonal
+components.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| season_length | Union | | Number of observations per unit of time. Ex: 24 Hourly data. |
+| use_boxcox | Optional | True | Whether or not to use a Box-Cox transformation. |
+| bc_lower_bound | float | 0.0 | Lower bound for the Box-Cox transformation. |
+| bc_upper_bound | float | 1.0 | Upper bound for the Box-Cox transformation. |
+| use_trend | Optional | True | Whether or not to use a trend component. |
+| use_damped_trend | Optional | False | Whether or not to dampen the trend component. |
+| use_arma_errors | bool | False | Whether or not to use a ARMA errors. |
+| alias | str | TBATS | Custom name of the model. |
+
+------------------------------------------------------------------------
+
+source
+
+### TBATS.fit
+
+> ``` text
+> TBATS.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit TBATS model.
+
+Fit TBATS model to a time series (numpy array) `y`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|-----------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Ignored |
+| **Returns** | | | **TBATS model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### TBATS.predict
+
+> ``` text
+> TBATS.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted TBATS model.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### TBATS.predict_in_sample
+
+> ``` text
+> TBATS.predict_in_sample (level:Optional[Tuple[int]]=None)
+> ```
+
+*Access fitted TBATS model predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### TBATS.forecast
+
+> ``` text
+> TBATS.forecast (y:numpy.ndarray, h:int, X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient TBATS model.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | |
+| X_future | Optional | None | |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+# Theta Family
+
+## Standard Theta Method
+
+------------------------------------------------------------------------
+
+source
+
+### Theta
+
+> ``` text
+> Theta (season_length:int=1, decomposition_type:str='multiplicative',
+> alias:str='Theta', prediction_intervals:Optional[statsforecast.uti
+> ls.ConformalIntervals]=None)
+> ```
+
+*Standard Theta Method.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| season_length | int | 1 | Number of observations per unit of time. Ex: 24 Hourly data. |
+| decomposition_type | str | multiplicative | Sesonal decomposition type, ‘multiplicative’ (default) or ‘additive’. |
+| alias | str | Theta | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### Theta.forecast
+
+> ``` text
+> Theta.forecast (y:numpy.ndarray, h:int, X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient AutoTheta predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### Theta.fit
+
+> ``` text
+> Theta.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the AutoTheta model.
+
+Fit an AutoTheta model to a time series (numpy array) `y` and optionally
+exogenous variables (numpy array) `X`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|---------------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | | | **AutoTheta fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### Theta.predict
+
+> ``` text
+> Theta.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted AutoTheta.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### Theta.predict_in_sample
+
+> ``` text
+> Theta.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted AutoTheta insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### Theta.forward
+
+> ``` text
+> Theta.forward (y:numpy.ndarray, h:int, X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+*Apply fitted AutoTheta to a new time series.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import Theta
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# Theta's usage example
+model = Theta(season_length=12)
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+## Optimized Theta Method
+
+------------------------------------------------------------------------
+
+source
+
+### OptimizedTheta
+
+> ``` text
+> OptimizedTheta (season_length:int=1,
+> decomposition_type:str='multiplicative',
+> alias:str='OptimizedTheta', prediction_intervals:Optional
+> [statsforecast.utils.ConformalIntervals]=None)
+> ```
+
+*Optimized Theta Method.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| season_length | int | 1 | Number of observations per unit of time. Ex: 24 Hourly data. |
+| decomposition_type | str | multiplicative | Sesonal decomposition type, ‘multiplicative’ (default) or ‘additive’. |
+| alias | str | OptimizedTheta | Custom name of the model. Default [`OptimizedTheta`](https://Nixtla.github.io/statsforecast/src/core/models.html#optimizedtheta). |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### OptimizedTheta.forecast
+
+> ``` text
+> OptimizedTheta.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None,
+> fitted:bool=False)
+> ```
+
+\*Memory Efficient AutoTheta predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### OptimizedTheta.fit
+
+> ``` text
+> OptimizedTheta.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the AutoTheta model.
+
+Fit an AutoTheta model to a time series (numpy array) `y` and optionally
+exogenous variables (numpy array) `X`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|---------------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | | | **AutoTheta fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### OptimizedTheta.predict
+
+> ``` text
+> OptimizedTheta.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted AutoTheta.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### OptimizedTheta.predict_in_sample
+
+> ``` text
+> OptimizedTheta.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted AutoTheta insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### OptimizedTheta.forward
+
+> ``` text
+> OptimizedTheta.forward (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None,
+> fitted:bool=False)
+> ```
+
+*Apply fitted AutoTheta to a new time series.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import OptimizedTheta
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# OptimzedThetA's usage example
+model = OptimizedTheta(season_length=12)
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+## Dynamic Standard Theta Method
+
+------------------------------------------------------------------------
+
+source
+
+### DynamicTheta
+
+> ``` text
+> DynamicTheta (season_length:int=1,
+> decomposition_type:str='multiplicative',
+> alias:str='DynamicTheta', prediction_intervals:Optional[sta
+> tsforecast.utils.ConformalIntervals]=None)
+> ```
+
+*Dynamic Standard Theta Method.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| season_length | int | 1 | Number of observations per unit of time. Ex: 24 Hourly data. |
+| decomposition_type | str | multiplicative | Sesonal decomposition type, ‘multiplicative’ (default) or ‘additive’. |
+| alias | str | DynamicTheta | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### DynamicTheta.forecast
+
+> ``` text
+> DynamicTheta.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient AutoTheta predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### DynamicTheta.fit
+
+> ``` text
+> DynamicTheta.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the AutoTheta model.
+
+Fit an AutoTheta model to a time series (numpy array) `y` and optionally
+exogenous variables (numpy array) `X`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|---------------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | | | **AutoTheta fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### DynamicTheta.predict
+
+> ``` text
+> DynamicTheta.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted AutoTheta.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### DynamicTheta.predict_in_sample
+
+> ``` text
+> DynamicTheta.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted AutoTheta insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### DynamicTheta.forward
+
+> ``` text
+> DynamicTheta.forward (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+*Apply fitted AutoTheta to a new time series.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import DynamicTheta
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# DynStandardThetaMethod's usage example
+model = DynamicTheta(season_length=12)
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+## Dynamic Optimized Theta Method
+
+------------------------------------------------------------------------
+
+source
+
+### DynamicOptimizedTheta
+
+> ``` text
+> DynamicOptimizedTheta (season_length:int=1,
+> decomposition_type:str='multiplicative',
+> alias:str='DynamicOptimizedTheta', prediction_inte
+> rvals:Optional[statsforecast.utils.ConformalInterv
+> als]=None)
+> ```
+
+*Dynamic Optimized Theta Method.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| season_length | int | 1 | Number of observations per unit of time. Ex: 24 Hourly data. |
+| decomposition_type | str | multiplicative | Sesonal decomposition type, ‘multiplicative’ (default) or ‘additive’. |
+| alias | str | DynamicOptimizedTheta | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### DynamicOptimizedTheta.forecast
+
+> ``` text
+> DynamicOptimizedTheta.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None,
+> fitted:bool=False)
+> ```
+
+\*Memory Efficient AutoTheta predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### DynamicOptimizedTheta.fit
+
+> ``` text
+> DynamicOptimizedTheta.fit (y:numpy.ndarray,
+> X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the AutoTheta model.
+
+Fit an AutoTheta model to a time series (numpy array) `y` and optionally
+exogenous variables (numpy array) `X`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|---------------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | | | **AutoTheta fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### DynamicOptimizedTheta.predict
+
+> ``` text
+> DynamicOptimizedTheta.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted AutoTheta.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### DynamicOptimizedTheta.predict_in_sample
+
+> ``` text
+> DynamicOptimizedTheta.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted AutoTheta insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### DynamicOptimizedTheta.forward
+
+> ``` text
+> DynamicOptimizedTheta.forward (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None,
+> fitted:bool=False)
+> ```
+
+*Apply fitted AutoTheta to a new time series.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import DynamicOptimizedTheta
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# OptimzedThetaMethod's usage example
+model = DynamicOptimizedTheta(season_length=12)
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+# ARCH Family
+
+## Garch model
+
+------------------------------------------------------------------------
+
+source
+
+### GARCH
+
+> ``` text
+> GARCH (p:int=1, q:int=1, alias:str='GARCH', prediction_intervals:Optional
+> [statsforecast.utils.ConformalIntervals]=None)
+> ```
+
+\*Generalized Autoregressive Conditional Heteroskedasticity (GARCH)
+model.
+
+A method for modeling time series that exhibit non-constant volatility
+over time. The GARCH model assumes that at time $t$, $y_t$ is given by:
+
+$$y_t = v_t \sigma_t$$
+
+with
+
+$$\sigma_t^2 = w + \sum_{i=1}^p a_i y_{t-i}^2 + \sum_{j=1}^q b_j \sigma_{t-j}^2$$.
+
+Here $v_t$ is a sequence of iid random variables with zero mean and unit
+variance. The coefficients $w$, $a_i$, $i=1,...,p$, and $b_j$,
+$j=1,...,q$ must satisfy the following conditions:
+
+1. $w > 0$ and $a_i, b_j \geq 0$ for all $i$ and $j$.
+2. $\sum_{k=1}^{max(p,q)} a_k + b_k < 1$. Here it is assumed that
+ $a_i=0$ for $i>p$ and $b_j=0$ for $j>q$.
+
+The ARCH model is a particular case of the GARCH model when $q=0$.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| p | int | 1 | Number of lagged versions of the series. |
+| q | int | 1 | |
+| alias | str | GARCH | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### GARCH.fit
+
+> ``` text
+> GARCH.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit GARCH model.
+
+Fit GARCH model to a time series (numpy array) `y`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|-----------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | |
+| **Returns** | | | **GARCH model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### GARCH.predict
+
+> ``` text
+> GARCH.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted GARCH model.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### GARCH.predict_in_sample
+
+> ``` text
+> GARCH.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted GARCH model predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### GARCH.forecast
+
+> ``` text
+> GARCH.forecast (y:numpy.ndarray, h:int, X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient GARCH model.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | |
+| X_future | Optional | None | |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+## ARCH model
+
+------------------------------------------------------------------------
+
+source
+
+### ARCH
+
+> ``` text
+> ARCH (p:int=1, alias:str='ARCH', prediction_intervals:Optional[statsforec
+> ast.utils.ConformalIntervals]=None)
+> ```
+
+\*Autoregressive Conditional Heteroskedasticity (ARCH) model.
+
+A particular case of the GARCH(p,q) model where $q=0$. It assumes that
+at time $t$, $y_t$ is given by:
+
+$$y_t = \epsilon_t \sigma_t$$
+
+with
+
+$$\sigma_t^2 = w0 + \sum_{i=1}^p a_i y_{t-i}^2$$.
+
+Here $\epsilon_t$ is a sequence of iid random variables with zero mean
+and unit variance. The coefficients $w$ and $a_i$, $i=1,...,p$ must be
+nonnegative and $\sum_{k=1}^p a_k < 1$.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| p | int | 1 | Number of lagged versions of the series. |
+| alias | str | ARCH | Custom name of the model. |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. By default, the model will compute the native prediction intervals. |
+
+------------------------------------------------------------------------
+
+source
+
+### ARCH.fit
+
+> ``` text
+> ARCH.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit GARCH model.
+
+Fit GARCH model to a time series (numpy array) `y`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|-----------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | |
+| **Returns** | | | **GARCH model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### ARCH.predict
+
+> ``` text
+> ARCH.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted GARCH model.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### ARCH.predict_in_sample
+
+> ``` text
+> ARCH.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted GARCH model predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### ARCH.forecast
+
+> ``` text
+> ARCH.forecast (y:numpy.ndarray, h:int, X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient GARCH model.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | |
+| X_future | Optional | None | |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+# Machine Learning models
+
+## SklearnModel
+
+------------------------------------------------------------------------
+
+source
+
+### SklearnModel
+
+> ``` text
+> SklearnModel (model, prediction_intervals:Optional[statsforecast.utils.Co
+> nformalIntervals]=None, alias:Optional[str]=None)
+> ```
+
+*scikit-learn model wrapper*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| model | sklearn.base.BaseEstimator | | scikit-learn estimator |
+| prediction_intervals | Optional | None | Information to compute conformal prediction intervals. This is required for generating future prediction intervals. |
+| alias | Optional | None | Custom name of the model. If `None` will use the model’s class. |
+
+------------------------------------------------------------------------
+
+source
+
+### SklearnModel.fit
+
+> ``` text
+> SklearnModel.fit (y:numpy.ndarray, X:numpy.ndarray)
+> ```
+
+*Fit the model.*
+
+| | **Type** | **Details** |
+|-------------|------------------|-----------------------------------|
+| y | ndarray | Clean time series of shape (t, ). |
+| X | ndarray | Exogenous of shape (t, n_x). |
+| **Returns** | **SklearnModel** | **Fitted SklearnModel object.** |
+
+------------------------------------------------------------------------
+
+source
+
+### SklearnModel.predict
+
+> ``` text
+> SklearnModel.predict (h:int, X:numpy.ndarray,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted SklearnModel.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | Forecast horizon. |
+| X | ndarray | | Exogenous of shape (h, n_x). |
+| level | Optional | None | |
+| **Returns** | **Dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### SklearnModel.predict_in_sample
+
+> ``` text
+> SklearnModel.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted SklearnModel insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **Dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### SklearnModel.forecast
+
+> ``` text
+> SklearnModel.forecast (y:numpy.ndarray, h:int, X:numpy.ndarray,
+> X_future:numpy.ndarray,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient SklearnModel predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| h | int | | Forecast horizon. |
+| X | ndarray | | Insample exogenous of shape (t, n_x). |
+| X_future | ndarray | | Exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **Dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+# Fallback Models
+
+## ConstantModel
+
+------------------------------------------------------------------------
+
+source
+
+### ConstantModel
+
+> ``` text
+> ConstantModel (constant:float, alias:str='ConstantModel')
+> ```
+
+\*Constant Model.
+
+Returns Constant values.\*
+
+------------------------------------------------------------------------
+
+source
+
+### ConstantModel.forecast
+
+> ``` text
+> ConstantModel.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None,
+> fitted:bool=False)
+> ```
+
+\*Memory Efficient Constant Model predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n,). |
+| h | int | | |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### ConstantModel.fit
+
+> ``` text
+> ConstantModel.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the Constant model.
+
+Fit an Constant Model to a time series (numpy.array) `y`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|-----------|-------------|---------------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | **self:** | | **Constant fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### ConstantModel.predict
+
+> ``` text
+> ConstantModel.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted ConstantModel.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | forecasting horizon |
+| X | Optional | None | exogenous regressors |
+| level | Optional | None | confidence level |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### ConstantModel.predict_in_sample
+
+> ``` text
+> ConstantModel.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted Constant Model insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### ConstantModel.forward
+
+> ``` text
+> ConstantModel.forward (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+*Apply Constant model predictions to a new/updated time series.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `constant` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import ConstantModel
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# ConstantModel's usage example
+model = ConstantModel(1)
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+## ZeroModel
+
+------------------------------------------------------------------------
+
+source
+
+### ZeroModel
+
+> ``` text
+> ZeroModel (alias:str='ZeroModel')
+> ```
+
+\*Returns Zero forecasts.
+
+Returns Zero values.\*
+
+------------------------------------------------------------------------
+
+source
+
+### ZeroModel.forecast
+
+> ``` text
+> ZeroModel.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient Constant Model predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n,). |
+| h | int | | |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### ZeroModel.fit
+
+> ``` text
+> ZeroModel.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the Constant model.
+
+Fit an Constant Model to a time series (numpy.array) `y`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|-----------|-------------|---------------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | **self:** | | **Constant fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### ZeroModel.predict
+
+> ``` text
+> ZeroModel.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted ConstantModel.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | forecasting horizon |
+| X | Optional | None | exogenous regressors |
+| level | Optional | None | confidence level |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### ZeroModel.predict_in_sample
+
+> ``` text
+> ZeroModel.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted Constant Model insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### ZeroModel.forward
+
+> ``` text
+> ZeroModel.forward (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+*Apply Constant model predictions to a new/updated time series.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n, ). |
+| h | int | | Forecast horizon. |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels for prediction intervals. |
+| fitted | bool | False | Whether or not returns insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `constant` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import ZeroModel
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# NanModel's usage example
+model = ZeroModel()
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+## NaNModel
+
+------------------------------------------------------------------------
+
+source
+
+### NaNModel
+
+> ``` text
+> NaNModel (alias:str='NaNModel')
+> ```
+
+\*NaN Model.
+
+Returns NaN values.\*
+
+------------------------------------------------------------------------
+
+source
+
+### NaNModel.forecast
+
+> ``` text
+> NaNModel.forecast (y:numpy.ndarray, h:int,
+> X:Optional[numpy.ndarray]=None,
+> X_future:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None, fitted:bool=False)
+> ```
+
+\*Memory Efficient Constant Model predictions.
+
+This method avoids memory burden due from object storage. It is
+analogous to `fit_predict` without storing information. It assumes you
+know the forecast horizon in advance.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| y | ndarray | | Clean time series of shape (n,). |
+| h | int | | |
+| X | Optional | None | Optional insample exogenous of shape (t, n_x). |
+| X_future | Optional | None | Optional exogenous of shape (h, n_x). |
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| fitted | bool | False | Whether or not to return insample predictions. |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### NaNModel.fit
+
+> ``` text
+> NaNModel.fit (y:numpy.ndarray, X:Optional[numpy.ndarray]=None)
+> ```
+
+\*Fit the Constant model.
+
+Fit an Constant Model to a time series (numpy.array) `y`.\*
+
+| | **Type** | **Default** | **Details** |
+|-------------|-----------|-------------|---------------------------------------|
+| y | ndarray | | Clean time series of shape (t, ). |
+| X | Optional | None | Optional exogenous of shape (t, n_x). |
+| **Returns** | **self:** | | **Constant fitted model.** |
+
+------------------------------------------------------------------------
+
+source
+
+### NaNModel.predict
+
+> ``` text
+> NaNModel.predict (h:int, X:Optional[numpy.ndarray]=None,
+> level:Optional[List[int]]=None)
+> ```
+
+*Predict with fitted ConstantModel.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| h | int | | forecasting horizon |
+| X | Optional | None | exogenous regressors |
+| level | Optional | None | confidence level |
+| **Returns** | **dict** | | **Dictionary with entries `mean` for point predictions and `level_*` for probabilistic predictions.** |
+
+------------------------------------------------------------------------
+
+source
+
+### NaNModel.predict_in_sample
+
+> ``` text
+> NaNModel.predict_in_sample (level:Optional[List[int]]=None)
+> ```
+
+*Access fitted Constant Model insample predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| level | Optional | None | Confidence levels (0-100) for prediction intervals. |
+| **Returns** | **dict** | | **Dictionary with entries `fitted` for point predictions and `level_*` for probabilistic predictions.** |
+
+
+```python
+from statsforecast.models import NaNModel
+from statsforecast.utils import AirPassengers as ap
+```
+
+
+```python
+# NanModel's usage example
+model = NaNModel()
+model = model.fit(y=ap)
+y_hat_dict = model.predict(h=4)
+y_hat_dict
+```
+
+# References
+
+#### **General**
+
+- [Hyndman, R.J., & Athanasopoulos, G. (2021). “Forecasting:
+ principles and practice, 3rd edition”. OTexts: Melbourne, Australia.
+ OTexts.com/fpp3 Accessed on July 2022](https://otexts.com/fpp3/).
+
+- [Shmueli, G., & Lichtendahl Jr, K. C. (2016). “Practical time series
+ forecasting with R: A hands-on guide”. Axelrod Schnall
+ Publishers](https://www.forecastingbook.com/).
+
+#### **Automatic Forecasting**
+
+- [Rob J. Hyndman, Yeasmin Khandakar (2008). “Automatic Time Series
+ Forecasting: The forecast package for
+ R”](https://www.jstatsoft.org/article/view/v027i03).
+
+#### **Exponential Smoothing**
+
+- [Charles. C. Holt (1957). “Forecasting seasonals and trends by
+ exponentially weighted moving averages”, ONR Research Memorandum,
+ Carnegie Institute of Technology
+ 52](https://www.sciencedirect.com/science/article/abs/pii/S0169207003001134).
+
+- [Peter R. Winters (1960). “Forecasting sales by exponentially
+ weighted moving averages”. Management
+ Science](https://pubsonline.informs.org/doi/abs/10.1287/mnsc.6.3.324).
+
+- [Hyndman, Rob, et al (2008). “Forecasting with exponential
+ smoothing: the state space
+ approach”](https://robjhyndman.com/expsmooth/).
+
+- [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ principles and practice, Methods with
+ trend”](https://otexts.com/fpp3/holt.html).
+
+- [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ principles and practice, Methods with
+ seasonality”](https://otexts.com/fpp3/holt-winters.html).
+
+#### **Simple Methods**
+
+- [Rob J. Hyndman and George Athanasopoulos (2018). “Forecasting
+ principles and practice, Simple
+ Methods”](https://otexts.com/fpp3/simple-methods.html).
+
+#### **Sparse Intermittent**
+
+- [Croston, J. D. (1972). “Forecasting and stock control for
+ intermittent demands”. Journal of the Operational Research Society,
+ 23(3),
+ 289-303](https://link.springer.com/article/10.1057/jors.1972.50).
+
+- [Nikolopoulos, K., Syntetos, A. A., Boylan, J. E., Petropoulos, F.,
+ & Assimakopoulos, V. (2011). “An aggregate–disaggregate intermittent
+ demand approach (ADIDA) to forecasting: an empirical proposition and
+ analysis”. Journal of the Operational Research Society, 62(3),
+ 544-554](https://researchportal.bath.ac.uk/en/publications/an-aggregate-disaggregate-intermittent-demand-approach-adida-to-f).
+
+- [Syntetos, A. A., & Boylan, J. E. (2005). “The accuracy of
+ intermittent demand estimates”. International Journal of
+ forecasting, 21(2),
+ 303-314](https://www.academia.edu/1527250/The_accuracy_of_intermittent_demand_estimates).
+
+- [Syntetos, A. A., & Boylan, J. E. (2021). “Intermittent demand
+ forecasting: Context, methods and applications”. John Wiley &
+ Sons](https://www.ifors.org/intermittent-demand-forecasting-context-methods-and-applications/).
+
+- [Teunter, R. H., Syntetos, A. A., & Babai, M. Z. (2011).
+ “Intermittent demand: Linking forecasting to inventory
+ obsolescence”. European Journal of Operational Research, 214(3),
+ 606-615](https://www.sciencedirect.com/science/article/abs/pii/S0377221711004437).
+
+#### **Multiple Seasonalities**
+
+- [Bandara, Kasun & Hyndman, Rob & Bergmeir, Christoph. (2021). “MSTL:
+ A Seasonal-Trend Decomposition Algorithm for Time Series with
+ Multiple Seasonal Patterns”.](https://arxiv.org/abs/2107.13462)
+
+#### **Theta Family**
+
+- [Jose A. Fiorucci, Tiago R. Pellegrini, Francisco Louzada, Fotios
+ Petropoulos, Anne B. Koehler (2016). “Models for optimising the
+ theta method and their relationship to state space models”.
+ International Journal of
+ Forecasting](https://www.sciencedirect.com/science/article/pii/S0169207016300243).
+
+#### **GARCH Model**
+
+- [Engle, R. F. (1982). Autoregressive conditional heteroscedasticity
+ with estimates of the variance of United Kingdom inflation.
+ Econometrica: Journal of the econometric society,
+ 987-1007.](http://www.econ.uiuc.edu/~econ508/Papers/engle82.pdf)
+
+- [Bollerslev, T. (1986). Generalized autoregressive conditional
+ heteroskedasticity. Journal of econometrics, 31(3),
+ 307-327.](https://www.sciencedirect.com/science/article/abs/pii/0304407686900631)
+
+- [Hamilton, J. D. (1994). Time series analysis. Princeton university
+ press.](https://press.princeton.edu/books/hardcover/9780691042893/time-series-analysis)
+
+#### **TBATS Model**
+
+- [De Livera, A. M., Hyndman, R. J., & Snyder, R. D. (2011).
+ Forecasting time series with complex seasonal patterns using
+ exponential smoothing. Journal of the American statistical
+ association, 106(496),
+ 1513-1527.](https://www.sciencedirect.com/science/article/abs/pii/0304407686900631)
+
+- [De Livera, Alysha M (2017). Modeling time series with complex
+ seasonal patterns using exponential smoothing. Monash University.
+ Thesis.](https://doi.org/10.4225/03/589299681de3d)
+
diff --git a/statsforecast/src/core/models_intro.mdx b/statsforecast/src/core/models_intro.mdx
new file mode 100644
index 00000000..c0c6adcb
--- /dev/null
+++ b/statsforecast/src/core/models_intro.mdx
@@ -0,0 +1,101 @@
+---
+title: StatsForecast's Models
+---
+
+
+## Automatic Forecasting
+
+Automatic forecasting tools search for the best parameters and select
+the best possible model for a series of time series. These tools are
+useful for large collections of univariate time series.
+
+| Model | Point Forecast | Probabilistic Forecast | Insample fitted values | Probabilistic fitted values |
+|:-----------------------------|:---------:|:---------:|:---------:|:---------:|
+| [`AutoARIMA`](./models.html#autoarima) | ✅ | ✅ | ✅ | ✅ |
+| [`AutoETS`](./models.html#autoets) | ✅ | ✅ | ✅ | ✅ |
+| [`AutoCES`](./models.html#autoces) | ✅ | ✅ | ✅ | ✅ |
+| [`AutoTheta`](./models.html#autotheta) | ✅ | ✅ | ✅ | ✅ |
+
+## ARIMA Family
+
+These models exploit the existing autocorrelations in the time series.
+
+| Model | Point Forecast | Probabilistic Forecast | Insample fitted values | Probabilistic fitted values |
+|:-----------------------------|:---------:|:---------:|:---------:|:---------:|
+| [`ARIMA`](./models.html#arima) | ✅ | ✅ | ✅ | ✅ |
+| [`AutoRegressive`](./models.html#autoregressive) | ✅ | ✅ | ✅ | ✅ |
+
+## Theta Family
+
+Fit two theta lines to a deseasonalized time series, using different
+techniques to obtain and combine the two theta lines to produce the
+final forecasts.
+
+| Model | Point Forecast | Probabilistic Forecast | Insample fitted values | Probabilistic fitted values |
+|:-----------------------------|:---------:|:---------:|:---------:|:---------:|
+| [`Theta`](./models.html#theta) | ✅ | ✅ | ✅ | ✅ |
+| [`OptimizedTheta`](./models.html#optimizedtheta) | ✅ | ✅ | ✅ | ✅ |
+| [`DynamicTheta`](./models.html#dynamictheta) | ✅ | ✅ | ✅ | ✅ |
+| [`DynamicOptimizedTheta`](./models.html#dynamicoptimizedtheta) | ✅ | ✅ | ✅ | ✅ |
+
+## Multiple Seasonalities
+
+Suited for signals with more than one clear seasonality. Useful for
+low-frequency data like electricity and logs.
+
+| Model | Point Forecast | Probabilistic Forecast | Insample fitted values | Probabilistic fitted values |
+|:-----------------------------|:---------:|:---------:|:---------:|:---------:|
+| [`MSTL`](./models.html#mstl) | ✅ | ✅ | ✅ | ✅ |
+
+## GARCH and ARCH Models
+
+Suited for modeling time series that exhibit non-constant volatility
+over time. The ARCH model is a particular case of GARCH.
+
+| Model | Point Forecast | Probabilistic Forecast | Insample fitted values | Probabilistic fitted values |
+|:-----------------------------|:---------:|:---------:|:---------:|:---------:|
+| [`GARCH`](./models.html#garch) | ✅ | ✅ | ✅ | ✅ |
+| [`ARCH`](./models.html#arch) | ✅ | ✅ | ✅ | ✅ |
+
+## Baseline Models
+
+Classical models for establishing baseline.
+
+| Model | Point Forecast | Probabilistic Forecast | Insample fitted values | Probabilistic fitted values |
+|:-----------------------------|:---------:|:---------:|:---------:|:---------:|
+| [`HistoricAverage`](./models.html#historicaverage) | ✅ | ✅ | ✅ | ✅ |
+| [`Naive`](./models.html#naive) | ✅ | ✅ | ✅ | ✅ |
+| [`RandomWalkWithDrift`](./models.html#randomwalkwithdrift) | ✅ | ✅ | ✅ | ✅ |
+| [`SeasonalNaive`](./models.html#seasonalnaive) | ✅ | ✅ | ✅ | ✅ |
+| [`WindowAverage`](./models.html#windowaverage) | ✅ | | | |
+| [`SeasonalWindowAverage`](./models.html#seasonalwindowaverage) | ✅ | | | |
+
+## Exponential Smoothing
+
+Uses a weighted average of all past observations where the weights
+decrease exponentially into the past. Suitable for data with clear trend
+and/or seasonality. Use the `SimpleExponential` family for data with no
+clear trend or seasonality.
+
+| Model | Point Forecast | Probabilistic Forecast | Insample fitted values | Probabilistic fitted values |
+|:-----------------------------|:---------:|:---------:|:---------:|:---------:|
+| [`SimpleExponentialSmoothing`](./models.html#simpleexponentialsmoothing) | ✅ | | | |
+| [`SimpleExponentialSmoothingOptimized`](./models.html#simpleexponentialsmoothingoptimized) | ✅ | | | |
+| [`SeasonalExponentialSmoothing`](./models.html#seasonalexponentialsmoothing) | ✅ | | | |
+| [`SeasonalExponentialSmoothingOptimized`](./models.html#seasonalexponentialsmoothingoptimized) | ✅ | | | |
+| [`Holt`](./models.html#holt) | ✅ | ✅ | ✅ | ✅ |
+| [`HoltWinters`](./models.html#holtwinters) | ✅ | ✅ | ✅ | ✅ |
+
+## Sparse or Intermittent
+
+Suited for series with very few non-zero observations
+
+| Model | Point Forecast | Probabilistic Forecast | Insample fitted values | Probabilistic fitted values |
+|:-----------------------------|:---------:|:---------:|:---------:|:---------:|
+| [`ADIDA`](./models.html#adida) | ✅ | | | |
+| [`CrostonClassic`](./models.html#crostonclassic) | ✅ | | | |
+| [`CrostonOptimized`](./models.html#crostonoptimized) | ✅ | | | |
+| [`CrostonSBA`](./models.html#crostonsba) | ✅ | | | |
+| [`IMAPA`](./models.html#imapa) | ✅ | | | |
+| [`TSB`](./models.html#tsb) | ✅ | | | |
+
diff --git a/statsforecast/src/distributed.core.html.mdx b/statsforecast/src/distributed.core.html.mdx
new file mode 100644
index 00000000..81a4e614
--- /dev/null
+++ b/statsforecast/src/distributed.core.html.mdx
@@ -0,0 +1,7 @@
+---
+output-file: distributed.core.html
+title: Core
+---
+
+
+
diff --git a/statsforecast/src/distributed.multiprocess.html.mdx b/statsforecast/src/distributed.multiprocess.html.mdx
new file mode 100644
index 00000000..353c6aa9
--- /dev/null
+++ b/statsforecast/src/distributed.multiprocess.html.mdx
@@ -0,0 +1,31 @@
+---
+description: >-
+ The computational efficiency of `StatsForecast` can be tracked to its two core
+ components: 1. Its `models` written in NumBa that optimizes Python code to
+ reach C speeds. 2. Its `core.StatsForecast` class that enables distributed
+ computing. This is a low-level class enabling other distribution
+ methods.
+output-file: distributed.multiprocess.html
+title: MultiprocessBackend
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### MultiprocessBackend
+
+> ``` text
+> MultiprocessBackend (n_jobs:int)
+> ```
+
+*MultiprocessBackend Parent Class for Distributed Computation.*
+
+| | **Type** | **Details** |
+|--------|---------------------------|-------------------------------------|
+| n_jobs | int | Number of jobs used in the parallel processing, use -1 for all cores. |
+| **Returns** | **None** | |
+
diff --git a/statsforecast/src/ets.html.mdx b/statsforecast/src/ets.html.mdx
new file mode 100644
index 00000000..6ab1bd50
--- /dev/null
+++ b/statsforecast/src/ets.html.mdx
@@ -0,0 +1,22 @@
+---
+output-file: ets.html
+title: ETS Model
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### ets_f
+
+> ``` text
+> ets_f (y, m, model='ZZZ', damped=None, alpha=None, beta=None, gamma=None,
+> phi=None, additive_only=None, blambda=None, biasadj=None,
+> lower=None, upper=None, opt_crit='lik', nmse=3, bounds='both',
+> ic='aicc', restrict=True, allow_multiplicative_trend=False,
+> use_initial_values=False, maxit=2000)
+> ```
+
diff --git a/statsforecast/src/feature_engineering.html.mdx b/statsforecast/src/feature_engineering.html.mdx
new file mode 100644
index 00000000..90379dd8
--- /dev/null
+++ b/statsforecast/src/feature_engineering.html.mdx
@@ -0,0 +1,88 @@
+---
+description: Generate features for downstream models
+output-file: feature_engineering.html
+title: Feature engineering
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### mstl_decomposition
+
+> ``` text
+> mstl_decomposition
+> (df:Union[pandas.core.frame.DataFrame,polars.datafram
+> e.frame.DataFrame], model:statsforecast.models.MSTL,
+> freq:str, h:int)
+> ```
+
+*Decompose the series into trend and seasonal using the MSTL model.*
+
+| | **Type** | **Details** |
+|--------|---------------------------|-------------------------------------|
+| df | Union | DataFrame with columns \[`unique_id`, `ds`, `y`\]. |
+| model | MSTL | Model to use for the decomposition. |
+| freq | str | Frequency of the data (pandas alias) |
+| h | int | Forecast horizon. |
+| **Returns** | **Tuple** | **Original dataframe with the ‘trend’ and ‘seasonal’ columns added.** |
+
+
+```python
+import pandas as pd
+from fastcore.test import test_fail
+from utilsforecast.losses import smape
+
+from statsforecast.models import Naive
+from statsforecast.utils import generate_series
+```
+
+
+```python
+series = generate_series(10, freq='D')
+series['unique_id'] = series['unique_id'].astype('int64')
+```
+
+
+```python
+test_fail(lambda: mstl_decomposition(series, Naive(), 'D', 14), contains='must be an MSTL instance')
+```
+
+
+```python
+horizon = 14
+model = MSTL(season_length=7)
+series = series.sample(frac=1.0)
+train_df, X_df = mstl_decomposition(series, model, 'D', horizon)
+```
+
+
+```python
+series_pl = generate_series(10, freq='D', engine='polars')
+series_pl = series_pl.with_columns(unique_id=pl.col('unique_id').cast(pl.Int64))
+train_df_pl, X_df_pl = mstl_decomposition(series_pl, model, '1d', horizon)
+```
+
+
+```python
+pd.testing.assert_series_equal(
+ train_df.groupby('unique_id')['ds'].max() + pd.offsets.Day(),
+ X_df.groupby('unique_id')['ds'].min()
+)
+assert X_df.shape[0] == train_df['unique_id'].nunique() * horizon
+pd.testing.assert_frame_equal(train_df, train_df_pl.to_pandas())
+pd.testing.assert_frame_equal(X_df, X_df_pl.to_pandas())
+with_estimate = train_df_pl.with_columns(estimate=pl.col('trend') + pl.col('seasonal'))
+assert smape(with_estimate, models=['estimate'])['estimate'].mean() < 0.1
+```
+
+
+```python
+model = MSTL(season_length=[7, 28])
+train_df, X_df = mstl_decomposition(series, model, 'D', horizon)
+assert train_df.columns.intersection(X_df.columns).tolist() == ['unique_id', 'ds', 'trend', 'seasonal7', 'seasonal28']
+```
+
diff --git a/statsforecast/src/garch.html.mdx b/statsforecast/src/garch.html.mdx
new file mode 100644
index 00000000..1c3a626a
--- /dev/null
+++ b/statsforecast/src/garch.html.mdx
@@ -0,0 +1,41 @@
+---
+output-file: garch.html
+title: GARCH
+---
+
+
+## Generate GARCH data
+
+## Generate GARCH(p,q) model
+
+------------------------------------------------------------------------
+
+source
+
+### garch_model
+
+> ``` text
+> garch_model (x, p, q)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### garch_forecast
+
+> ``` text
+> garch_forecast (mod, h)
+> ```
+
+### Comparison with arch library
+
+This section compares the coefficients generated by the previous
+functions with the coefficients generated by the [arch
+library](https://github.com/bashtage/arch) for $p=q$, $p>q$, $psource
+
+### MFLES
+
+> ``` text
+> MFLES (verbose=1, robust=None)
+> ```
+
+*Initialize self. See help(type(self)) for accurate signature.*
+
diff --git a/statsforecast/src/mstl.html.mdx b/statsforecast/src/mstl.html.mdx
new file mode 100644
index 00000000..095623c8
--- /dev/null
+++ b/statsforecast/src/mstl.html.mdx
@@ -0,0 +1,29 @@
+---
+output-file: mstl.html
+title: MSTL model
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### mstl
+
+> ``` text
+> mstl (x:numpy.ndarray, period:Union[int,List[int]],
+> blambda:Optional[float]=None, iterate:int=2,
+> s_window:Optional[numpy.ndarray]=None, stl_kwargs:Dict={})
+> ```
+
+| | **Type** | **Default** | **Details** |
+|------------|----------|-------------|----------------------|
+| x | ndarray | | time series |
+| period | Union | | season length |
+| blambda | Optional | None | box-cox transform |
+| iterate | int | 2 | number of iterations |
+| s_window | Optional | None | seasonal window |
+| stl_kwargs | Dict | {} | |
+
diff --git a/statsforecast/src/tbats.html.mdx b/statsforecast/src/tbats.html.mdx
new file mode 100644
index 00000000..13b5bea2
--- /dev/null
+++ b/statsforecast/src/tbats.html.mdx
@@ -0,0 +1,158 @@
+---
+output-file: tbats.html
+title: TBATS model
+---
+
+
+
+```python
+import matplotlib.pyplot as plt
+import pandas as pd
+
+from statsforecast.utils import AirPassengers as ap
+```
+
+## Load data
+
+## Functions
+
+### find_harmonics
+
+### initial_parameters
+
+### makeXMatrix
+
+### findPQ
+
+### makeTBATSWMatrix
+
+### makeTBATSGMatrix
+
+### makeTBATSFMatrix
+
+### calcTBATSFaster
+
+### extract_params
+
+### updateTBATSWMatrix
+
+### updateTBATSGMatrix
+
+### updateTBATSFMatrix
+
+### checkAdmissibility
+
+### calcLikelihoodTBATS
+
+## TBATS model
+
+### tbats_model_generator
+
+### tbats_model
+
+------------------------------------------------------------------------
+
+source
+
+### tbats_model
+
+> ``` text
+> tbats_model (y, seasonal_periods, k_vector, use_boxcox, bc_lower_bound,
+> bc_upper_bound, use_trend, use_damped_trend,
+> use_arma_errors)
+> ```
+
+### tbats_selection
+
+------------------------------------------------------------------------
+
+source
+
+### tbats_selection
+
+> ``` text
+> tbats_selection (y, seasonal_periods, use_boxcox, bc_lower_bound,
+> bc_upper_bound, use_trend, use_damped_trend,
+> use_arma_errors)
+> ```
+
+### tbats_forecast
+
+------------------------------------------------------------------------
+
+source
+
+### tbats_forecast
+
+> ``` text
+> tbats_forecast (mod, h)
+> ```
+
+| | **Details** |
+|-----|--------------------------------------------|
+| mod | |
+| h | this function is the same as bats_forecast |
+
+### Example
+
+
+```python
+y = ap
+seasonal_periods = np.array([12])
+```
+
+
+```python
+# Default parameters
+use_boxcox = None
+bc_lower_bound = 0
+bc_upper_bound = 1
+use_trend = None
+use_damped_trend = None
+use_arma_errors = True
+```
+
+
+```python
+mod = tbats_selection(y, seasonal_periods, use_boxcox, bc_lower_bound, bc_upper_bound, use_trend, use_damped_trend, use_arma_errors)
+```
+
+
+```python
+# Values in R
+print(mod['aic']) # 1397.015
+print(mod['k_vector']) # 5
+print(mod['description']) # use_boxcox = TRUE, use_trend = TRUE, use_damped_trend = FALSE, use_arma_errors = FALSE
+```
+
+
+```python
+fitted_trans = mod['fitted'].ravel()
+if mod['BoxCox_lambda'] is not None:
+ fitted_trans = inv_boxcox(fitted_trans, mod['BoxCox_lambda'])
+```
+
+
+```python
+h = 24
+fcst = tbats_forecast(mod, h)
+forecast = fcst['mean']
+if mod['BoxCox_lambda'] is not None:
+ forecast = inv_boxcox(forecast, mod['BoxCox_lambda'])
+```
+
+
+```python
+fig, ax = plt.subplots(1, 1, figsize = (20,7))
+plt.plot(np.arange(0, len(y)), y, color='black', label='original')
+plt.plot(np.arange(0, len(y)), fitted_trans, color='blue', label = "fitted")
+plt.plot(np.arange(len(y), len(y)+h), forecast, '.-', color = 'green', label = 'fcst')
+plt.legend()
+```
+
diff --git a/statsforecast/src/theta.html.mdx b/statsforecast/src/theta.html.mdx
new file mode 100644
index 00000000..3b6adfb6
--- /dev/null
+++ b/statsforecast/src/theta.html.mdx
@@ -0,0 +1,48 @@
+---
+output-file: theta.html
+title: Theta Model
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### forecast_theta
+
+> ``` text
+> forecast_theta (obj, h, level=None)
+> ```
+
+
+```python
+forecast_theta(res, 12, level=[90, 80])
+```
+
+------------------------------------------------------------------------
+
+source
+
+### auto_theta
+
+> ``` text
+> auto_theta (y, m, model=None, initial_smoothed=None, alpha=None,
+> theta=None, nmse=3, decomposition_type='multiplicative')
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### forward_theta
+
+> ``` text
+> forward_theta (fitted_model, y)
+> ```
+
diff --git a/statsforecast/src/utils.html.mdx b/statsforecast/src/utils.html.mdx
new file mode 100644
index 00000000..11ba2017
--- /dev/null
+++ b/statsforecast/src/utils.html.mdx
@@ -0,0 +1,89 @@
+---
+description: >-
+ The `core.StatsForecast` class allows you to efficiently fit multiple
+ `StatsForecast` models for large sets of time series. It operates with pandas
+ DataFrame `df` that identifies individual series and datestamps with the
+ `unique_id` and `ds` columns, and the `y` column denotes the target time
+ series variable. To assist development, we declare useful datasets that we use
+ throughout all `StatsForecast`'s unit tests.
+output-file: utils.html
+title: Utils
+---
+
+
+# 1. Synthetic Panel Data
+
+------------------------------------------------------------------------
+
+source
+
+### generate_series
+
+> ``` text
+> generate_series (n_series:int, freq:str='D', min_length:int=50,
+> max_length:int=500, n_static_features:int=0,
+> equal_ends:bool=False, engine:str='pandas', seed:int=0)
+> ```
+
+\*Generate Synthetic Panel Series.
+
+Generates `n_series` of frequency `freq` of different lengths in the
+interval \[`min_length`, `max_length`\]. If `n_static_features > 0`,
+then each series gets static features with random values. If
+`equal_ends == True` then all series end at the same date.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| n_series | int | | Number of series for synthetic panel. |
+| freq | str | D | Frequency of the data, ‘D’ or ‘M’. |
+| min_length | int | 50 | Minimum length of synthetic panel’s series. |
+| max_length | int | 500 | Maximum length of synthetic panel’s series. |
+| n_static_features | int | 0 | Number of static exogenous variables for synthetic panel’s series. |
+| equal_ends | bool | False | Series should end in the same date stamp `ds`. |
+| engine | str | pandas | Output Dataframe type (‘pandas’ or ‘polars’). |
+| seed | int | 0 | Random seed used for generating the data. |
+| **Returns** | **Union** | | **Synthetic panel with columns \[`unique_id`, `ds`, `y`\] and exogenous.** |
+
+
+```python
+synthetic_panel = generate_series(n_series=2)
+synthetic_panel.groupby('unique_id', observed=True).head(4)
+```
+
+# 2. AirPassengers Data
+
+The classic Box & Jenkins airline data. Monthly totals of international
+airline passengers, 1949 to 1960.
+
+It has been used as a reference on several forecasting libraries, since
+it is a series that shows clear trends and seasonalities it offers a
+nice opportunity to quickly showcase a model’s predictions performance.
+
+
+```python
+from statsforecast.utils import AirPassengersDF
+```
+
+
+```python
+AirPassengersDF.head(12)
+```
+
+
+```python
+#We are going to plot the ARIMA predictions, and the prediction intervals.
+fig, ax = plt.subplots(1, 1, figsize = (20, 7))
+plot_df = AirPassengersDF.set_index('ds')
+
+plot_df[['y']].plot(ax=ax, linewidth=2)
+ax.set_title('AirPassengers Forecast', fontsize=22)
+ax.set_ylabel('Monthly Passengers', fontsize=20)
+ax.set_xlabel('Timestamp [t]', fontsize=20)
+ax.legend(prop={'size': 15})
+ax.grid()
+```
+
+## Model utils
+
diff --git a/style.css b/style.css
new file mode 100644
index 00000000..03565270
--- /dev/null
+++ b/style.css
@@ -0,0 +1,144 @@
+@font-face {
+ font-family: "PPNeueMontreal";
+ src: url("./fonts/ppneuemontreal-medium.otf") format("otf");
+}
+
+@font-face {
+ font-family: "SupplyMono";
+ src: url("./fonts/Supply-Regular.otf") format("otf");
+}
+
+:root {
+ --primary-light: #fff;
+ --primary-dark: #161616;
+ --gray: #f0f0f0;
+}
+
+html,
+body {
+ background-color: var(--gray);
+ font-family: "PPNeueMontreal", sans-serif;
+}
+
+.eyebrow {
+ font-family: "SupplyMono", monospace;
+ @apply text-red-300;
+}
+
+#navbar img {
+ height: 20px;
+}
+
+.bg-gradient-to-b {
+ background-color: transparent;
+ background-image: none;
+}
+
+a {
+ border-radius: 0.125rem !important;
+ border: 1px solid transparent;
+}
+
+a.font-semibold {
+ background: var(--primary-light);
+ border: 1px solid var(--primary-dark);
+ font-weight: 500;
+ color: var(--primary-dark);
+}
+
+.rounded-md {
+ border-radius: 0.125rem !important;
+}
+
+.rounded-xl {
+ border-radius: 0.125rem !important;
+}
+
+.rounded-2xl,
+.rounded-search {
+ border-radius: 0.25rem !important;
+}
+
+#navbar-transition {
+ background: var(--gray);
+}
+
+#topbar-cta-button a span {
+ background: var(--primary-dark);
+ border-radius: 0.125rem;
+}
+
+#content-side-layout a {
+ border: none;
+}
+
+#content-side-layout a.font-medium {
+ color: var(--primary);
+ font-weight: 600;
+}
+
+a.card svg {
+ background: var(--primary-dark);
+ opacity: 0.8;
+}
+
+/* dark mode */
+html.dark > body {
+ background-color: #161616;
+}
+
+html.dark #navbar-transition {
+ background: var(--primary-dark);
+}
+
+html.dark a.font-semibold {
+ background: #000;
+ border: 1px solid var(--gray);
+ outline-color: var(--gray);
+ color: #fff;
+}
+
+html.dark a.font-semibold svg {
+ background: #fff;
+}
+
+html.dark #topbar-cta-button a span {
+ background: #fff;
+ color: #000;
+}
+
+html.dark #topbar-cta-button svg {
+ color: #000;
+}
+
+html.dark a.card svg {
+ background: var(--primary-light);
+}
+
+/* Banner styling for theme support */
+#banner {
+ /* background-color: var(--primary-light); # original */
+ background-color: #22c55e;
+ border-color: var(--primary-light);
+ color: var(--primary-dark);
+ border-bottom: 1px solid var(--primary-dark) !important;
+}
+
+#banner a:hover {
+ opacity: 0.8;
+}
+
+#banner p span {
+ color: black !important;
+ margin: 0;
+ font-size: medium;
+}
+
+#banner strong {
+ color: black !important;
+ font-size: medium;
+}
+
+#banner svg path {
+ color: #000 !important;
+}
diff --git a/utilsforecast/.nojekyll b/utilsforecast/.nojekyll
new file mode 100644
index 00000000..e69de29b
diff --git a/utilsforecast/compat.mdx b/utilsforecast/compat.mdx
new file mode 100644
index 00000000..8b137891
--- /dev/null
+++ b/utilsforecast/compat.mdx
@@ -0,0 +1 @@
+
diff --git a/utilsforecast/dark.png b/utilsforecast/dark.png
new file mode 100644
index 00000000..4142a0bb
Binary files /dev/null and b/utilsforecast/dark.png differ
diff --git a/utilsforecast/data.html.mdx b/utilsforecast/data.html.mdx
new file mode 100644
index 00000000..e77e014e
--- /dev/null
+++ b/utilsforecast/data.html.mdx
@@ -0,0 +1,59 @@
+---
+description: Utilies for generating time series datasets
+output-file: data.html
+title: Data
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### generate_series
+
+> ``` text
+> generate_series (n_series:int, freq:str='D', min_length:int=50,
+> max_length:int=500, n_static_features:int=0,
+> equal_ends:bool=False, with_trend:bool=False,
+> static_as_categorical:bool=True, n_models:int=0,
+> level:Optional[List[float]]=None,
+> engine:Literal['pandas','polars']='pandas', seed:int=0)
+> ```
+
+*Generate Synthetic Panel Series.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| n_series | int | | Number of series for synthetic panel. |
+| freq | str | D | Frequency of the data (pandas alias). Seasonalities are implemented for hourly, daily and monthly. |
+| min_length | int | 50 | Minimum length of synthetic panel’s series. |
+| max_length | int | 500 | Maximum length of synthetic panel’s series. |
+| n_static_features | int | 0 | Number of static exogenous variables for synthetic panel’s series. |
+| equal_ends | bool | False | Series should end in the same timestamp. |
+| with_trend | bool | False | Series should have a (positive) trend. |
+| static_as_categorical | bool | True | Static features should have a categorical data type. |
+| n_models | int | 0 | Number of models predictions to simulate. |
+| level | Optional | None | Confidence level for intervals to simulate for each model. |
+| engine | Literal | pandas | Output Dataframe type. |
+| seed | int | 0 | Random seed used for generating the data. |
+| **Returns** | **Union** | | **Synthetic panel with columns \[`unique_id`, `ds`, `y`\] and exogenous features.** |
+
+
+```python
+synthetic_panel = generate_series(n_series=2)
+synthetic_panel.groupby('unique_id', observed=True).head(4)
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|----------|
+| 0 | 0 | 2000-01-01 | 0.357595 |
+| 1 | 0 | 2000-01-02 | 1.301382 |
+| 2 | 0 | 2000-01-03 | 2.272442 |
+| 3 | 0 | 2000-01-04 | 3.211827 |
+| 222 | 1 | 2000-01-01 | 5.399023 |
+| 223 | 1 | 2000-01-02 | 6.092818 |
+| 224 | 1 | 2000-01-03 | 0.476396 |
+| 225 | 1 | 2000-01-04 | 1.343744 |
+
diff --git a/utilsforecast/evaluation.html.mdx b/utilsforecast/evaluation.html.mdx
new file mode 100644
index 00000000..4815bb9a
--- /dev/null
+++ b/utilsforecast/evaluation.html.mdx
@@ -0,0 +1,131 @@
+---
+description: Model performance evaluation
+output-file: evaluation.html
+title: Evaluation
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### evaluate
+
+> ``` text
+> evaluate (df:~AnyDFType, metrics:List[Callable],
+> models:Optional[List[str]]=None,
+> train_df:Optional[~AnyDFType]=None,
+> level:Optional[List[int]]=None, id_col:str='unique_id',
+> time_col:str='ds', target_col:str='y',
+> agg_fn:Optional[str]=None)
+> ```
+
+*Evaluate forecast using different metrics.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | AnyDFType | | Forecasts to evaluate. Must have `id_col`, `time_col`, `target_col` and models’ predictions. |
+| metrics | List | | Functions with arguments `df`, `models`, `id_col`, `target_col` and optionally `train_df`. |
+| models | Optional | None | Names of the models to evaluate. If `None` will use every column in the dataframe after removing id, time and target. |
+| train_df | Optional | None | Training set. Used to evaluate metrics such as [`mase`](https://Nixtla.github.io/utilsforecast/losses.html#mase). |
+| level | Optional | None | Prediction interval levels. Used to compute losses that rely on quantiles. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| agg_fn | Optional | None | Statistic to compute on the scores by id to reduce them to a single number. |
+| **Returns** | **AnyDFType** | | **Metrics with one row per (id, metric) combination and one column per model. If `agg_fn` is not `None`, there is only one row per metric.** |
+
+
+```python
+from functools import partial
+
+import numpy as np
+import pandas as pd
+
+from utilsforecast.losses import *
+from utilsforecast.data import generate_series
+```
+
+
+```python
+series = generate_series(10, n_models=2, level=[80, 95])
+```
+
+
+```python
+series['unique_id'] = series['unique_id'].astype('int')
+```
+
+
+```python
+models = ['model0', 'model1']
+metrics = [
+ mae,
+ mse,
+ rmse,
+ mape,
+ smape,
+ partial(mase, seasonality=7),
+ quantile_loss,
+ mqloss,
+ coverage,
+ calibration,
+ scaled_crps,
+]
+```
+
+
+```python
+evaluation = evaluate(
+ series,
+ metrics=metrics,
+ models=models,
+ train_df=series,
+ level=[80, 95],
+)
+evaluation
+```
+
+| | unique_id | metric | model0 | model1 |
+|-----|-----------|-------------|----------|----------|
+| 0 | 0 | mae | 0.158108 | 0.163246 |
+| 1 | 1 | mae | 0.160109 | 0.143805 |
+| 2 | 2 | mae | 0.159815 | 0.170510 |
+| 3 | 3 | mae | 0.168537 | 0.161595 |
+| 4 | 4 | mae | 0.170182 | 0.163329 |
+| ... | ... | ... | ... | ... |
+| 175 | 5 | scaled_crps | 0.034202 | 0.035472 |
+| 176 | 6 | scaled_crps | 0.034880 | 0.033610 |
+| 177 | 7 | scaled_crps | 0.034337 | 0.034745 |
+| 178 | 8 | scaled_crps | 0.033336 | 0.032459 |
+| 179 | 9 | scaled_crps | 0.034766 | 0.035243 |
+
+
+```python
+summary = evaluation.drop(columns='unique_id').groupby('metric').mean().reset_index()
+summary
+```
+
+| | metric | model0 | model1 |
+|-----|----------------------|----------|----------|
+| 0 | calibration_q0.025 | 0.000000 | 0.000000 |
+| 1 | calibration_q0.1 | 0.000000 | 0.000000 |
+| 2 | calibration_q0.9 | 0.833993 | 0.815833 |
+| 3 | calibration_q0.975 | 0.853991 | 0.836949 |
+| 4 | coverage_level80 | 0.833993 | 0.815833 |
+| 5 | coverage_level95 | 0.853991 | 0.836949 |
+| 6 | mae | 0.161286 | 0.162281 |
+| 7 | mape | 0.048894 | 0.049624 |
+| 8 | mase | 0.966846 | 0.975354 |
+| 9 | mqloss | 0.056904 | 0.056216 |
+| 10 | mse | 0.048653 | 0.049198 |
+| 11 | quantile_loss_q0.025 | 0.019990 | 0.019474 |
+| 12 | quantile_loss_q0.1 | 0.067315 | 0.065781 |
+| 13 | quantile_loss_q0.9 | 0.095510 | 0.093841 |
+| 14 | quantile_loss_q0.975 | 0.044803 | 0.045767 |
+| 15 | rmse | 0.220357 | 0.221543 |
+| 16 | scaled_crps | 0.035003 | 0.034576 |
+| 17 | smape | 0.024475 | 0.024902 |
+
diff --git a/utilsforecast/favicon.svg b/utilsforecast/favicon.svg
new file mode 100644
index 00000000..e5f33342
--- /dev/null
+++ b/utilsforecast/favicon.svg
@@ -0,0 +1,5 @@
+
diff --git a/utilsforecast/feature_engineering.html.mdx b/utilsforecast/feature_engineering.html.mdx
new file mode 100644
index 00000000..9334ee75
--- /dev/null
+++ b/utilsforecast/feature_engineering.html.mdx
@@ -0,0 +1,374 @@
+---
+description: Create exogenous regressors for your models
+output-file: feature_engineering.html
+title: Feature engineering
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### fourier
+
+> ``` text
+> fourier (df:~DFType, freq:Union[str,int], season_length:int, k:int,
+> h:int=0, id_col:str='unique_id', time_col:str='ds')
+> ```
+
+*Compute fourier seasonal terms for training and forecasting*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Dataframe with ids, times and values for the exogenous regressors. |
+| freq | Union | | Frequency of the data. Must be a valid pandas or polars offset alias, or an integer. |
+| season_length | int | | Number of observations per unit of time. Ex: 24 Hourly data. |
+| k | int | | Maximum order of the fourier terms |
+| h | int | 0 | Forecast horizon. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| **Returns** | **Tuple** | | **Original DataFrame with the computed features** |
+
+
+```python
+import pandas as pd
+
+from utilsforecast.data import generate_series
+```
+
+
+```python
+series = generate_series(5, equal_ends=True)
+transformed_df, future_df = fourier(series, freq='D', season_length=7, k=2, h=1)
+transformed_df
+```
+
+| | unique_id | ds | y | sin1_7 | sin2_7 | cos1_7 | cos2_7 |
+|------|-----------|------------|----------|-----------|-----------|-----------|-----------|
+| 0 | 0 | 2000-10-05 | 0.428973 | -0.974927 | 0.433894 | -0.222526 | -0.900964 |
+| 1 | 0 | 2000-10-06 | 1.423626 | -0.781835 | -0.974926 | 0.623486 | -0.222531 |
+| 2 | 0 | 2000-10-07 | 2.311782 | -0.000005 | -0.000009 | 1.000000 | 1.000000 |
+| 3 | 0 | 2000-10-08 | 3.192191 | 0.781829 | 0.974930 | 0.623493 | -0.222512 |
+| 4 | 0 | 2000-10-09 | 4.148767 | 0.974929 | -0.433877 | -0.222517 | -0.900972 |
+| ... | ... | ... | ... | ... | ... | ... | ... |
+| 1096 | 4 | 2001-05-10 | 4.058910 | -0.974927 | 0.433888 | -0.222523 | -0.900967 |
+| 1097 | 4 | 2001-05-11 | 5.178157 | -0.781823 | -0.974934 | 0.623500 | -0.222495 |
+| 1098 | 4 | 2001-05-12 | 6.133142 | -0.000002 | -0.000003 | 1.000000 | 1.000000 |
+| 1099 | 4 | 2001-05-13 | 0.403709 | 0.781840 | 0.974922 | 0.623479 | -0.222548 |
+| 1100 | 4 | 2001-05-14 | 1.081779 | 0.974928 | -0.433882 | -0.222520 | -0.900970 |
+
+
+```python
+future_df
+```
+
+| | unique_id | ds | sin1_7 | sin2_7 | cos1_7 | cos2_7 |
+|-----|-----------|------------|----------|-----------|-----------|----------|
+| 0 | 0 | 2001-05-15 | 0.433871 | -0.781813 | -0.900975 | 0.623513 |
+| 1 | 1 | 2001-05-15 | 0.433871 | -0.781813 | -0.900975 | 0.623513 |
+| 2 | 2 | 2001-05-15 | 0.433871 | -0.781813 | -0.900975 | 0.623513 |
+| 3 | 3 | 2001-05-15 | 0.433871 | -0.781813 | -0.900975 | 0.623513 |
+| 4 | 4 | 2001-05-15 | 0.433871 | -0.781813 | -0.900975 | 0.623513 |
+
+------------------------------------------------------------------------
+
+source
+
+### trend
+
+> ``` text
+> trend (df:~DFType, freq:Union[str,int], h:int=0, id_col:str='unique_id',
+> time_col:str='ds')
+> ```
+
+*Add a trend column with consecutive integers for training and
+forecasting*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Dataframe with ids, times and values for the exogenous regressors. |
+| freq | Union | | Frequency of the data. Must be a valid pandas or polars offset alias, or an integer. |
+| h | int | 0 | Forecast horizon. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| **Returns** | **Tuple** | | **Original DataFrame with the computed features** |
+
+
+```python
+series = generate_series(5, equal_ends=True)
+transformed_df, future_df = trend(series, freq='D', h=1)
+transformed_df
+```
+
+| | unique_id | ds | y | trend |
+|------|-----------|------------|----------|-------|
+| 0 | 0 | 2000-10-05 | 0.428973 | 152.0 |
+| 1 | 0 | 2000-10-06 | 1.423626 | 153.0 |
+| 2 | 0 | 2000-10-07 | 2.311782 | 154.0 |
+| 3 | 0 | 2000-10-08 | 3.192191 | 155.0 |
+| 4 | 0 | 2000-10-09 | 4.148767 | 156.0 |
+| ... | ... | ... | ... | ... |
+| 1096 | 4 | 2001-05-10 | 4.058910 | 369.0 |
+| 1097 | 4 | 2001-05-11 | 5.178157 | 370.0 |
+| 1098 | 4 | 2001-05-12 | 6.133142 | 371.0 |
+| 1099 | 4 | 2001-05-13 | 0.403709 | 372.0 |
+| 1100 | 4 | 2001-05-14 | 1.081779 | 373.0 |
+
+
+```python
+future_df
+```
+
+| | unique_id | ds | trend |
+|-----|-----------|------------|-------|
+| 0 | 0 | 2001-05-15 | 374.0 |
+| 1 | 1 | 2001-05-15 | 374.0 |
+| 2 | 2 | 2001-05-15 | 374.0 |
+| 3 | 3 | 2001-05-15 | 374.0 |
+| 4 | 4 | 2001-05-15 | 374.0 |
+
+------------------------------------------------------------------------
+
+source
+
+### time_features
+
+> ``` text
+> time_features (df:~DFType, freq:Union[str,int],
+> features:List[Union[str,Callable]], h:int=0,
+> id_col:str='unique_id', time_col:str='ds')
+> ```
+
+*Compute timestamp-based features for training and forecasting*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Dataframe with ids, times and values for the exogenous regressors. |
+| freq | Union | | Frequency of the data. Must be a valid pandas or polars offset alias, or an integer. |
+| features | List | | Features to compute. Can be string aliases of timestamp attributes or functions to apply to the times. |
+| h | int | 0 | Forecast horizon. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| **Returns** | **Tuple** | | **Original DataFrame with the computed features** |
+
+
+```python
+transformed_df, future_df = time_features(series, freq='D', features=['month', 'day', 'week'], h=1)
+transformed_df
+```
+
+| | unique_id | ds | y | month | day | week |
+|------|-----------|------------|----------|-------|-----|------|
+| 0 | 0 | 2000-10-05 | 0.428973 | 10 | 5 | 40 |
+| 1 | 0 | 2000-10-06 | 1.423626 | 10 | 6 | 40 |
+| 2 | 0 | 2000-10-07 | 2.311782 | 10 | 7 | 40 |
+| 3 | 0 | 2000-10-08 | 3.192191 | 10 | 8 | 40 |
+| 4 | 0 | 2000-10-09 | 4.148767 | 10 | 9 | 41 |
+| ... | ... | ... | ... | ... | ... | ... |
+| 1096 | 4 | 2001-05-10 | 4.058910 | 5 | 10 | 19 |
+| 1097 | 4 | 2001-05-11 | 5.178157 | 5 | 11 | 19 |
+| 1098 | 4 | 2001-05-12 | 6.133142 | 5 | 12 | 19 |
+| 1099 | 4 | 2001-05-13 | 0.403709 | 5 | 13 | 19 |
+| 1100 | 4 | 2001-05-14 | 1.081779 | 5 | 14 | 20 |
+
+
+```python
+future_df
+```
+
+| | unique_id | ds | month | day | week |
+|-----|-----------|------------|-------|-----|------|
+| 0 | 0 | 2001-05-15 | 5 | 15 | 20 |
+| 1 | 1 | 2001-05-15 | 5 | 15 | 20 |
+| 2 | 2 | 2001-05-15 | 5 | 15 | 20 |
+| 3 | 3 | 2001-05-15 | 5 | 15 | 20 |
+| 4 | 4 | 2001-05-15 | 5 | 15 | 20 |
+
+------------------------------------------------------------------------
+
+source
+
+### future_exog_to_historic
+
+> ``` text
+> future_exog_to_historic (df:~DFType, freq:Union[str,int],
+> features:List[str], h:int=0,
+> id_col:str='unique_id', time_col:str='ds')
+> ```
+
+*Turn future exogenous features into historic by shifting them `h`
+steps.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Dataframe with ids, times and values for the exogenous regressors. |
+| freq | Union | | Frequency of the data. Must be a valid pandas or polars offset alias, or an integer. |
+| features | List | | Features to be converted into historic. |
+| h | int | 0 | Forecast horizon. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| **Returns** | **Tuple** | | **Original DataFrame with the computed features** |
+
+
+```python
+series_with_prices = series.assign(price=np.random.rand(len(series))).sample(frac=1.0)
+series_with_prices
+```
+
+| | unique_id | ds | y | price |
+|-----|-----------|------------|----------|----------|
+| 436 | 2 | 2001-03-26 | 2.369113 | 0.774476 |
+| 312 | 1 | 2001-05-08 | 4.405212 | 0.557957 |
+| 536 | 3 | 2000-11-04 | 4.362074 | 0.745237 |
+| 34 | 0 | 2000-11-08 | 6.111161 | 0.809978 |
+| 652 | 3 | 2001-02-28 | 1.448291 | 0.685294 |
+| ... | ... | ... | ... | ... |
+| 609 | 3 | 2001-01-16 | 0.215892 | 0.699703 |
+| 873 | 4 | 2000-09-29 | 5.398198 | 0.677651 |
+| 268 | 1 | 2001-03-25 | 2.393771 | 0.735438 |
+| 171 | 0 | 2001-03-25 | 3.085493 | 0.463871 |
+| 931 | 4 | 2000-11-26 | 0.292296 | 0.691377 |
+
+
+```python
+transformed_df, future_df = future_exog_to_historic(
+ df=series_with_prices,
+ freq='D',
+ features=['price'],
+ h=2,
+)
+transformed_df
+```
+
+| | unique_id | ds | y | price |
+|------|-----------|------------|----------|----------|
+| 0 | 2 | 2001-03-26 | 2.369113 | 0.870133 |
+| 1 | 1 | 2001-05-08 | 4.405212 | 0.869751 |
+| 2 | 3 | 2000-11-04 | 4.362074 | 0.877901 |
+| 3 | 0 | 2000-11-08 | 6.111161 | 0.629413 |
+| 4 | 3 | 2001-02-28 | 1.448291 | 0.088073 |
+| ... | ... | ... | ... | ... |
+| 1096 | 3 | 2001-01-16 | 0.215892 | 0.472261 |
+| 1097 | 4 | 2000-09-29 | 5.398198 | 0.887531 |
+| 1098 | 1 | 2001-03-25 | 2.393771 | 0.481712 |
+| 1099 | 0 | 2001-03-25 | 3.085493 | 0.433153 |
+| 1100 | 4 | 2000-11-26 | 0.292296 | 0.620219 |
+
+
+```python
+future_df
+```
+
+| | unique_id | ds | price |
+|-----|-----------|------------|----------|
+| 0 | 0 | 2001-05-15 | 0.874328 |
+| 1 | 0 | 2001-05-16 | 0.481385 |
+| 2 | 1 | 2001-05-15 | 0.009058 |
+| 3 | 1 | 2001-05-16 | 0.083749 |
+| 4 | 2 | 2001-05-15 | 0.726212 |
+| 5 | 2 | 2001-05-16 | 0.052221 |
+| 6 | 3 | 2001-05-15 | 0.942335 |
+| 7 | 3 | 2001-05-16 | 0.274816 |
+| 8 | 4 | 2001-05-15 | 0.267545 |
+| 9 | 4 | 2001-05-16 | 0.112129 |
+
+------------------------------------------------------------------------
+
+source
+
+### pipeline
+
+> ``` text
+> pipeline (df:~DFType, features:List[Callable], freq:Union[str,int],
+> h:int=0, id_col:str='unique_id', time_col:str='ds')
+> ```
+
+*Compute several features for training and forecasting*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Dataframe with ids, times and values for the exogenous regressors. |
+| features | List | | List of features to compute. Must take only df, freq, h, id_col and time_col (other arguments must be fixed). |
+| freq | Union | | Frequency of the data. Must be a valid pandas or polars offset alias, or an integer. |
+| h | int | 0 | Forecast horizon. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| **Returns** | **Tuple** | | **Original DataFrame with the computed features** |
+
+
+```python
+def is_weekend(times):
+ if isinstance(times, pd.Index):
+ dow = times.weekday + 1 # monday=0 in pandas and 1 in polars
+ else:
+ dow = times.dt.weekday()
+ return dow >= 6
+
+def even_days_and_months(times):
+ if isinstance(times, pd.Index):
+ out = pd.DataFrame(
+ {
+ 'even_day': (times.weekday + 1) % 2 == 0,
+ 'even_month': times.month % 2 == 0,
+ }
+ )
+ else:
+ # for polars you can return a list of expressions
+ out = [
+ (times.dt.weekday() % 2 == 0).alias('even_day'),
+ (times.dt.month() % 2 == 0).alias('even_month'),
+ ]
+ return out
+
+features = [
+ trend,
+ partial(fourier, season_length=7, k=1),
+ partial(fourier, season_length=28, k=1),
+ partial(time_features, features=['day', is_weekend, even_days_and_months]),
+]
+transformed_df, future_df = pipeline(
+ series,
+ features=features,
+ freq='D',
+ h=1,
+)
+transformed_df
+```
+
+| | unique_id | ds | y | trend | sin1_7 | cos1_7 | sin1_28 | cos1_28 | day | is_weekend | even_day | even_month |
+|----|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | 0 | 2000-10-05 | 0.428973 | 152.0 | -0.974927 | -0.222526 | 0.433885 | -9.009683e-01 | 5 | False | True | True |
+| 1 | 0 | 2000-10-06 | 1.423626 | 153.0 | -0.781835 | 0.623486 | 0.222522 | -9.749276e-01 | 6 | False | False | True |
+| 2 | 0 | 2000-10-07 | 2.311782 | 154.0 | -0.000005 | 1.000000 | 0.000001 | -1.000000e+00 | 7 | True | True | True |
+| 3 | 0 | 2000-10-08 | 3.192191 | 155.0 | 0.781829 | 0.623493 | -0.222520 | -9.749281e-01 | 8 | True | False | True |
+| 4 | 0 | 2000-10-09 | 4.148767 | 156.0 | 0.974929 | -0.222517 | -0.433883 | -9.009693e-01 | 9 | False | False | True |
+| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
+| 1096 | 4 | 2001-05-10 | 4.058910 | 369.0 | -0.974927 | -0.222523 | 0.900969 | 4.338843e-01 | 10 | False | True | False |
+| 1097 | 4 | 2001-05-11 | 5.178157 | 370.0 | -0.781823 | 0.623500 | 0.974929 | 2.225177e-01 | 11 | False | False | False |
+| 1098 | 4 | 2001-05-12 | 6.133142 | 371.0 | -0.000002 | 1.000000 | 1.000000 | 4.251100e-07 | 12 | True | True | False |
+| 1099 | 4 | 2001-05-13 | 0.403709 | 372.0 | 0.781840 | 0.623479 | 0.974927 | -2.225243e-01 | 13 | True | False | False |
+| 1100 | 4 | 2001-05-14 | 1.081779 | 373.0 | 0.974928 | -0.222520 | 0.900969 | -4.338835e-01 | 14 | False | False | False |
+
+
+```python
+future_df
+```
+
+| | unique_id | ds | trend | sin1_7 | cos1_7 | sin1_28 | cos1_28 | day | is_weekend | even_day | even_month |
+|----|----|----|----|----|----|----|----|----|----|----|----|
+| 0 | 0 | 2001-05-15 | 374.0 | 0.433871 | -0.900975 | 0.781829 | -0.623493 | 15 | False | True | False |
+| 1 | 1 | 2001-05-15 | 374.0 | 0.433871 | -0.900975 | 0.781829 | -0.623493 | 15 | False | True | False |
+| 2 | 2 | 2001-05-15 | 374.0 | 0.433871 | -0.900975 | 0.781829 | -0.623493 | 15 | False | True | False |
+| 3 | 3 | 2001-05-15 | 374.0 | 0.433871 | -0.900975 | 0.781829 | -0.623493 | 15 | False | True | False |
+| 4 | 4 | 2001-05-15 | 374.0 | 0.433871 | -0.900975 | 0.781829 | -0.623493 | 15 | False | True | False |
+
diff --git a/utilsforecast/grouped_array.mdx b/utilsforecast/grouped_array.mdx
new file mode 100644
index 00000000..1f9221b0
--- /dev/null
+++ b/utilsforecast/grouped_array.mdx
@@ -0,0 +1,193 @@
+
+```python
+# test _append_one
+data = np.arange(5)
+indptr = np.array([0, 2, 5])
+new = np.array([7, 8])
+new_data, new_indptr = _append_one(data, indptr, new)
+np.testing.assert_equal(
+ new_data,
+ np.array([0, 1, 7, 2, 3, 4, 8])
+)
+np.testing.assert_equal(
+ new_indptr,
+ np.array([0, 3, 7]),
+)
+
+# 2d
+data = np.arange(5).reshape(-1, 1)
+new_data, new_indptr = _append_one(data, indptr, new)
+np.testing.assert_equal(
+ new_data,
+ np.array([0, 1, 7, 2, 3, 4, 8]).reshape(-1, 1)
+)
+np.testing.assert_equal(
+ new_indptr,
+ np.array([0, 3, 7]),
+)
+```
+
+
+```python
+# test append several
+data = np.arange(5)
+indptr = np.array([0, 2, 5])
+new_sizes = np.array([0, 2, 1])
+new_values = np.array([6, 7, 5])
+new_groups = np.array([False, True, False])
+new_data, new_indptr = _append_several(data, indptr, new_sizes, new_values, new_groups)
+np.testing.assert_equal(
+ new_data,
+ np.array([0, 1, 6, 7, 2, 3, 4, 5])
+)
+np.testing.assert_equal(
+ new_indptr,
+ np.array([0, 2, 4, 8]),
+)
+
+# 2d
+data = np.arange(5).reshape(-1, 1)
+indptr = np.array([0, 2, 5])
+new_sizes = np.array([0, 2, 1])
+new_values = np.array([6, 7, 5]).reshape(-1, 1)
+new_groups = np.array([False, True, False])
+new_data, new_indptr = _append_several(data, indptr, new_sizes, new_values, new_groups)
+np.testing.assert_equal(
+ new_data,
+ np.array([0, 1, 6, 7, 2, 3, 4, 5]).reshape(-1, 1)
+)
+np.testing.assert_equal(
+ new_indptr,
+ np.array([0, 2, 4, 8]),
+)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### GroupedArray
+
+> ``` text
+> GroupedArray (data:numpy.ndarray, indptr:numpy.ndarray)
+> ```
+
+*Initialize self. See help(type(self)) for accurate signature.*
+
+
+```python
+from fastcore.test import test_eq, test_fail
+
+from utilsforecast.data import generate_series
+```
+
+
+```python
+# The `GroupedArray` is used internally for storing the series values and performing transformations.
+data = np.arange(20, dtype=np.float32).reshape(-1, 2)
+indptr = np.array([0, 2, 10]) # group 1: [0, 1], group 2: [2..9]
+ga = GroupedArray(data, indptr)
+test_eq(len(ga), 2)
+```
+
+
+```python
+# Iterate through the groups
+ga_iter = iter(ga)
+np.testing.assert_equal(next(ga_iter), np.arange(4).reshape(-1, 2))
+np.testing.assert_equal(next(ga_iter), np.arange(4, 20).reshape(-1, 2))
+```
+
+
+```python
+# Take the last two observations from each group
+last2_data, last2_indptr = ga.take_from_groups(slice(-2, None))
+np.testing.assert_equal(
+ last2_data,
+ np.vstack([
+ np.arange(4).reshape(-1, 2),
+ np.arange(16, 20).reshape(-1, 2),
+ ]),
+)
+np.testing.assert_equal(last2_indptr, np.array([0, 2, 4]))
+
+# 1d
+ga1d = GroupedArray(np.arange(10), indptr)
+last2_data1d, last2_indptr1d = ga1d.take_from_groups(slice(-2, None))
+np.testing.assert_equal(
+ last2_data1d,
+ np.array([0, 1, 8, 9])
+)
+np.testing.assert_equal(last2_indptr1d, np.array([0, 2, 4]))
+```
+
+
+```python
+# Take the second observation from each group
+second_data, second_indptr = ga.take_from_groups(1)
+np.testing.assert_equal(second_data, np.array([[2, 3], [6, 7]]))
+np.testing.assert_equal(second_indptr, np.array([0, 1, 2]))
+
+# 1d
+second_data1d, second_indptr1d = ga1d.take_from_groups(1)
+np.testing.assert_equal(second_data1d, np.array([1, 3]))
+np.testing.assert_equal(second_indptr1d, np.array([0, 1, 2]))
+```
+
+
+```python
+# Take the last four observations from every group. Note that since group 1 only has two elements, only these are returned.
+last4_data, last4_indptr = ga.take_from_groups(slice(-4, None))
+np.testing.assert_equal(
+ last4_data,
+ np.vstack([
+ np.arange(4).reshape(-1, 2),
+ np.arange(12, 20).reshape(-1, 2),
+ ]),
+)
+np.testing.assert_equal(last4_indptr, np.array([0, 2, 6]))
+
+# 1d
+last4_data1d, last4_indptr1d = ga1d.take_from_groups(slice(-4, None))
+np.testing.assert_equal(
+ last4_data1d,
+ np.array([0, 1, 6, 7, 8, 9])
+)
+np.testing.assert_equal(last4_indptr1d, np.array([0, 2, 6]))
+```
+
+
+```python
+# Select a specific subset of groups
+indptr = np.array([0, 2, 4, 7, 10])
+ga2 = GroupedArray(data, indptr)
+subset = GroupedArray(*ga2.take([0, 2]))
+np.testing.assert_allclose(subset[0].data, ga2[0].data)
+np.testing.assert_allclose(subset[1].data, ga2[2].data)
+
+# 1d
+ga2_1d = GroupedArray(np.arange(10), indptr)
+subset1d = GroupedArray(*ga2_1d.take([0, 2]))
+np.testing.assert_allclose(subset1d[0].data, ga2_1d[0].data)
+np.testing.assert_allclose(subset1d[1].data, ga2_1d[2].data)
+```
+
+
+```python
+# try to append new values that don't match the number of groups
+test_fail(lambda: ga.append(np.array([1., 2., 3.])), contains='new must have 2 rows')
+```
+
+
+```python
+# build from df
+series_pd = generate_series(10, static_as_categorical=False, engine='pandas')
+ga_pd = GroupedArray.from_sorted_df(series_pd, 'unique_id', 'ds', 'y')
+series_pl = generate_series(10, static_as_categorical=False, engine='polars')
+ga_pl = GroupedArray.from_sorted_df(series_pl, 'unique_id', 'ds', 'y')
+np.testing.assert_allclose(ga_pd.data, ga_pl.data)
+np.testing.assert_equal(ga_pd.indptr, ga_pl.indptr)
+```
+
diff --git a/utilsforecast/imgs/losses/mae_loss.png b/utilsforecast/imgs/losses/mae_loss.png
new file mode 100644
index 00000000..c9d3b7fa
Binary files /dev/null and b/utilsforecast/imgs/losses/mae_loss.png differ
diff --git a/utilsforecast/imgs/losses/mape_loss.png b/utilsforecast/imgs/losses/mape_loss.png
new file mode 100644
index 00000000..d0f9a66a
Binary files /dev/null and b/utilsforecast/imgs/losses/mape_loss.png differ
diff --git a/utilsforecast/imgs/losses/mase_loss.png b/utilsforecast/imgs/losses/mase_loss.png
new file mode 100644
index 00000000..90db8c90
Binary files /dev/null and b/utilsforecast/imgs/losses/mase_loss.png differ
diff --git a/utilsforecast/imgs/losses/mq_loss.png b/utilsforecast/imgs/losses/mq_loss.png
new file mode 100644
index 00000000..7e3f6da3
Binary files /dev/null and b/utilsforecast/imgs/losses/mq_loss.png differ
diff --git a/utilsforecast/imgs/losses/mse_loss.png b/utilsforecast/imgs/losses/mse_loss.png
new file mode 100644
index 00000000..d175d5e0
Binary files /dev/null and b/utilsforecast/imgs/losses/mse_loss.png differ
diff --git a/utilsforecast/imgs/losses/q_loss.png b/utilsforecast/imgs/losses/q_loss.png
new file mode 100644
index 00000000..942dbc30
Binary files /dev/null and b/utilsforecast/imgs/losses/q_loss.png differ
diff --git a/utilsforecast/imgs/losses/rmae_loss.png b/utilsforecast/imgs/losses/rmae_loss.png
new file mode 100644
index 00000000..39a05b2e
Binary files /dev/null and b/utilsforecast/imgs/losses/rmae_loss.png differ
diff --git a/utilsforecast/imgs/losses/rmse_loss.png b/utilsforecast/imgs/losses/rmse_loss.png
new file mode 100644
index 00000000..0ceadef0
Binary files /dev/null and b/utilsforecast/imgs/losses/rmse_loss.png differ
diff --git a/utilsforecast/imgs/plotting.png b/utilsforecast/imgs/plotting.png
new file mode 100644
index 00000000..549fad86
Binary files /dev/null and b/utilsforecast/imgs/plotting.png differ
diff --git a/utilsforecast/index.html.mdx b/utilsforecast/index.html.mdx
new file mode 100644
index 00000000..09be4f2e
--- /dev/null
+++ b/utilsforecast/index.html.mdx
@@ -0,0 +1,139 @@
+---
+description: Forecasting utilities
+output-file: index.html
+title: utilsforecast
+---
+
+
+## Install
+
+### PyPI
+
+
+```sh
+pip install utilsforecast
+```
+
+### Conda
+
+
+```sh
+conda install -c conda-forge utilsforecast
+```
+
+## How to use
+
+### Generate synthetic data
+
+
+```python
+from utilsforecast.data import generate_series
+```
+
+
+```python
+series = generate_series(3, with_trend=True, static_as_categorical=False)
+series
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|------------|
+| 0 | 0 | 2000-01-01 | 0.422133 |
+| 1 | 0 | 2000-01-02 | 1.501407 |
+| 2 | 0 | 2000-01-03 | 2.568495 |
+| 3 | 0 | 2000-01-04 | 3.529085 |
+| 4 | 0 | 2000-01-05 | 4.481929 |
+| ... | ... | ... | ... |
+| 481 | 2 | 2000-06-11 | 163.914625 |
+| 482 | 2 | 2000-06-12 | 166.018479 |
+| 483 | 2 | 2000-06-13 | 160.839176 |
+| 484 | 2 | 2000-06-14 | 162.679603 |
+| 485 | 2 | 2000-06-15 | 165.089288 |
+
+### Plotting
+
+
+```python
+from utilsforecast.plotting import plot_series
+```
+
+
+```python
+fig = plot_series(series, plot_random=False, max_insample_length=50, engine='matplotlib')
+fig.savefig('imgs/index.png', bbox_inches='tight')
+```
+
+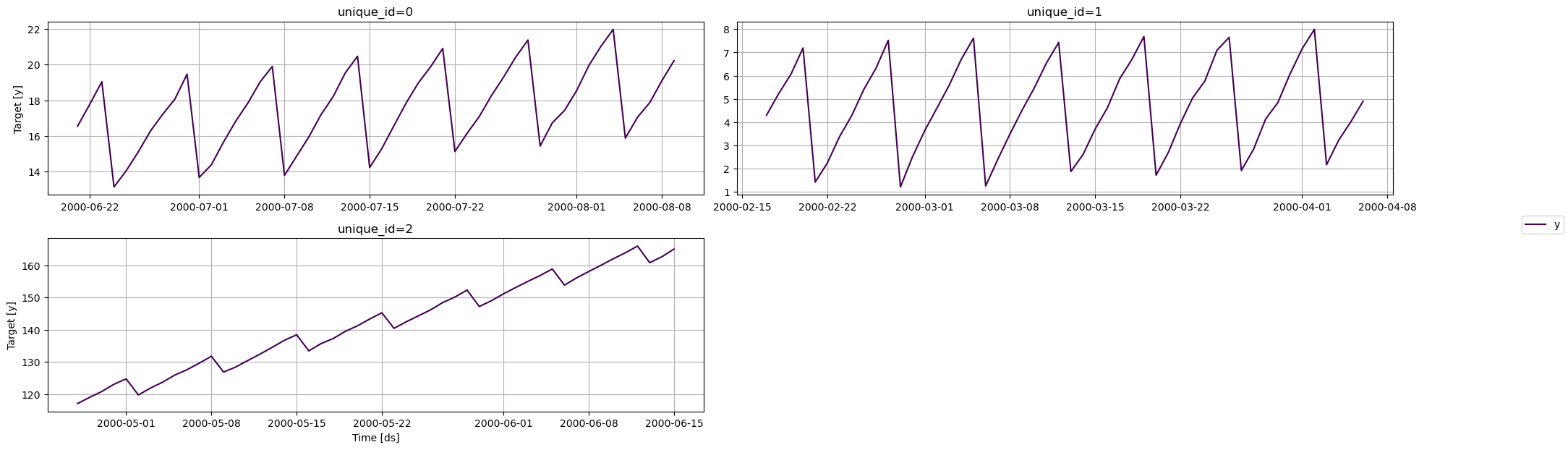
+
+### Preprocessing
+
+
+```python
+from utilsforecast.preprocessing import fill_gaps
+```
+
+
+```python
+serie = series[series['unique_id'].eq(0)].tail(10)
+# drop some points
+with_gaps = serie.sample(frac=0.5, random_state=0).sort_values('ds')
+with_gaps
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|-----------|
+| 213 | 0 | 2000-08-01 | 18.543147 |
+| 214 | 0 | 2000-08-02 | 19.941764 |
+| 216 | 0 | 2000-08-04 | 21.968733 |
+| 220 | 0 | 2000-08-08 | 19.091509 |
+| 221 | 0 | 2000-08-09 | 20.220739 |
+
+
+```python
+fill_gaps(with_gaps, freq='D')
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|-----------|
+| 0 | 0 | 2000-08-01 | 18.543147 |
+| 1 | 0 | 2000-08-02 | 19.941764 |
+| 2 | 0 | 2000-08-03 | NaN |
+| 3 | 0 | 2000-08-04 | 21.968733 |
+| 4 | 0 | 2000-08-05 | NaN |
+| 5 | 0 | 2000-08-06 | NaN |
+| 6 | 0 | 2000-08-07 | NaN |
+| 7 | 0 | 2000-08-08 | 19.091509 |
+| 8 | 0 | 2000-08-09 | 20.220739 |
+
+### Evaluating
+
+
+```python
+from functools import partial
+
+import numpy as np
+
+from utilsforecast.evaluation import evaluate
+from utilsforecast.losses import mape, mase
+```
+
+
+```python
+valid = series.groupby('unique_id').tail(7).copy()
+train = series.drop(valid.index)
+rng = np.random.RandomState(0)
+valid['seas_naive'] = train.groupby('unique_id')['y'].tail(7).values
+valid['rand_model'] = valid['y'] * rng.rand(valid['y'].shape[0])
+daily_mase = partial(mase, seasonality=7)
+evaluate(valid, metrics=[mape, daily_mase], train_df=train)
+```
+
+| | unique_id | metric | seas_naive | rand_model |
+|-----|-----------|--------|------------|------------|
+| 0 | 0 | mape | 0.024139 | 0.440173 |
+| 1 | 1 | mape | 0.054259 | 0.278123 |
+| 2 | 2 | mape | 0.042642 | 0.480316 |
+| 3 | 0 | mase | 0.907149 | 16.418014 |
+| 4 | 1 | mase | 0.991635 | 6.404254 |
+| 5 | 2 | mase | 1.013596 | 11.365040 |
+
diff --git a/utilsforecast/light.png b/utilsforecast/light.png
new file mode 100644
index 00000000..bbb99b54
Binary files /dev/null and b/utilsforecast/light.png differ
diff --git a/utilsforecast/losses.html.mdx b/utilsforecast/losses.html.mdx
new file mode 100644
index 00000000..7a201c3a
--- /dev/null
+++ b/utilsforecast/losses.html.mdx
@@ -0,0 +1,1232 @@
+---
+description: Loss functions for model evaluation.
+output-file: losses.html
+title: Losses
+---
+
+
+The most important train signal is the forecast error, which is the
+difference between the observed value $y_{\tau}$ and the prediction
+$\hat{y}_{\tau}$, at time $y_{\tau}$:
+
+$$
+
+e_{\tau} = y_{\tau}-\hat{y}_{\tau} \qquad \qquad \tau \in \{t+1,\dots,t+H \}
+
+$$
+
+The train loss summarizes the forecast errors in different evaluation
+metrics.
+
+
+```python
+from utilsforecast.data import generate_series
+```
+
+
+```python
+from polars.testing import assert_frame_equal as pl_assert_frame_equal
+models = ['model0', 'model1']
+series = generate_series(1000, n_models=2, level=[80])
+series_pl = generate_series(1000, n_models=2, level=[80], engine='polars')
+```
+
+## 1. Scale-dependent Errors
+
+### Mean Absolute Error (MAE)
+
+$$
+
+\mathrm{MAE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}) = \frac{1}{H} \sum^{t+H}_{\tau=t+1} |y_{\tau} - \hat{y}_{\tau}|
+
+$$
+
+
+
+------------------------------------------------------------------------
+
+source
+
+#### mae
+
+> ``` text
+> mae (df:~DFType, models:List[str], id_col:str='unique_id',
+> target_col:str='y')
+> ```
+
+\*Mean Absolute Error (MAE)
+
+MAE measures the relative prediction accuracy of a forecasting method by
+calculating the deviation of the prediction and the true value at a
+given time and averages these devations over the length of the series.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, actual values and predictions. |
+| models | List | | Columns that identify the models predictions. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+
+```python
+def pd_vs_pl(pd_df, pl_df, models):
+ pd.testing.assert_frame_equal(pd_df[models],
+ pl_df[models].to_pandas())
+```
+
+
+```python
+pd_vs_pl(
+ mae(series, models),
+ mae(series_pl, models),
+ models=models
+)
+```
+
+### Mean Squared Error
+
+$$
+
+\mathrm{MSE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}) = \frac{1}{H} \sum^{t+H}_{\tau=t+1} (y_{\tau} - \hat{y}_{\tau})^{2}
+
+$$
+
+
+
+------------------------------------------------------------------------
+
+source
+
+#### mse
+
+> ``` text
+> mse (df:~DFType, models:List[str], id_col:str='unique_id',
+> target_col:str='y')
+> ```
+
+\*Mean Squared Error (MSE)
+
+MSE measures the relative prediction accuracy of a forecasting method by
+calculating the squared deviation of the prediction and the true value
+at a given time, and averages these devations over the length of the
+series.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, actual values and predictions. |
+| models | List | | Columns that identify the models predictions. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+
+```python
+pd_vs_pl(
+ mse(series, models),
+ mse(series_pl, models),
+ models,
+)
+```
+
+### Root Mean Squared Error
+
+$$
+
+\mathrm{RMSE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}) = \sqrt{\frac{1}{H} \sum^{t+H}_{\tau=t+1} (y_{\tau} - \hat{y}_{\tau})^{2}}
+
+$$
+
+
+
+------------------------------------------------------------------------
+
+source
+
+#### rmse
+
+> ``` text
+> rmse (df:~DFType, models:List[str], id_col:str='unique_id',
+> target_col:str='y')
+> ```
+
+\*Root Mean Squared Error (RMSE)
+
+RMSE measures the relative prediction accuracy of a forecasting method
+by calculating the squared deviation of the prediction and the observed
+value at a given time and averages these devations over the length of
+the series. Finally the RMSE will be in the same scale as the original
+time series so its comparison with other series is possible only if they
+share a common scale. RMSE has a direct connection to the L2 norm.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, actual values and predictions. |
+| models | List | | Columns that identify the models predictions. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+
+```python
+pd_vs_pl(
+ rmse(series, models),
+ rmse(series_pl, models),
+ models,
+)
+```
+
+------------------------------------------------------------------------
+
+source
+
+#### bias
+
+> ``` text
+> bias (df:~DFType, models:List[str], id_col:str='unique_id',
+> target_col:str='y')
+> ```
+
+\*Forecast estimator bias.
+
+Defined as prediction - actual\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, actual values and predictions. |
+| models | List | | Columns that identify the models predictions. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+
+```python
+pd_vs_pl(
+ bias(series, models),
+ bias(series_pl, models),
+ models,
+)
+```
+
+------------------------------------------------------------------------
+
+source
+
+#### cfe
+
+> ``` text
+> cfe (df:~DFType, models:List[str], id_col:str='unique_id',
+> target_col:str='y')
+> ```
+
+\*Cumulative Forecast Error (CFE)
+
+Total signed forecast error per series. Positive values mean under
+forecast; negative mean over forecast.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, actual values and predictions. |
+| models | List | | Columns that identify the models predictions. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+
+```python
+pd_vs_pl(
+ cfe(series, models),
+ cfe(series_pl, models),
+ models,
+)
+```
+
+
+```python
+# case for cfe
+df = pd.DataFrame({
+ "unique_id": ["X","X","Y","Y"],
+ "y": [5, 10, 3, 7],
+ "y_hat": [7, 7, 1, 10]
+})
+# errors:
+# X: (7 - 5) + (7 - 10) = 2 + (-3) = -1
+# Y: (1 - 3) + (10 - 7) = -2 + 3 = 1
+expected = pd.DataFrame({
+ "unique_id": ["X", "Y"],
+ "y_hat": [-1, 1]
+ })
+
+# pandas
+out_pd = cfe(df, ["y_hat"])
+pd.testing.assert_frame_equal(
+ out_pd,
+ expected
+ )
+```
+
+
+```python
+df_pl = pl.from_pandas(df)
+out_pl = cfe(df_pl, ["y_hat"])
+pl_assert_frame_equal(
+ out_pl,
+ pl.from_pandas(expected)
+)
+```
+
+------------------------------------------------------------------------
+
+source
+
+#### pis
+
+> ``` text
+> pis (df:~DFType, models:List[str], id_col:str='unique_id',
+> target_col:str='y')
+> ```
+
+\*Compute the raw Absolute Periods In Stock (PIS) for one or multiple
+models.
+
+The PIS metric sums the absolute forecast errors per series without any
+scaling, yielding a scale-dependent measure of bias.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, actual values and predictions. |
+| models | List | | Columns that identify the models predictions. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+
+```python
+pd_vs_pl(
+ pis(series, models),
+ pis(series_pl, models),
+ models,
+)
+```
+
+
+```python
+# case for pis
+df = pd.DataFrame({
+ "unique_id": ["A","A","B","B"],
+ "y": [10, 15, 5, 7],
+ "y_hat": [12, 14, 4, 10]
+})
+# errors:
+# A: |12−10| + |14−15| = 2 + 1 = 3
+# B: |4−5| + |10−7| = 1 + 3 = 4
+expected = pd.DataFrame({
+ "unique_id": ["A", "B"],
+ "y_hat": [3, 4]
+ })
+
+# pandas branch
+out_pd = pis(df, ["y_hat"])
+pd.testing.assert_frame_equal(
+ out_pd,
+ expected
+)
+```
+
+
+```python
+df_pl = pl.from_pandas(df)
+out_pl = pis(df_pl, ["y_hat"])
+pl_assert_frame_equal(
+ out_pl,
+ pl.from_pandas(expected)
+)
+```
+
+------------------------------------------------------------------------
+
+source
+
+#### spis
+
+> ``` text
+> spis (df:~DFType, df_train:~DFType, models:List[str],
+> id_col:str='unique_id', target_col:str='y')
+> ```
+
+\*Compute the scaled Absolute Periods In Stock (sAPIS) for one or
+multiple models.
+
+The sPIS metric scales the sum of absolute forecast errors by the mean
+in-sample demand, yielding a scale-independent bias measure that can be
+aggregated across series.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, actual values and predictions. |
+| df_train | DFType | | |
+| models | List | | Columns that identify the models predictions. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+
+```python
+pd_vs_pl(
+ spis(series, series, models),
+ spis(series_pl, series_pl, models),
+ models,
+)
+```
+
+``` text
+/tmp/ipykernel_12558/2968355870.py:19: FutureWarning: The default of observed=False is deprecated and will be changed to True in a future version of pandas. Pass observed=False to retain current behavior or observed=True to adopt the future default and silence this warning.
+ .groupby(id_col)[target_col]
+```
+
+
+```python
+# case for scaled pis
+df_train = pd.DataFrame({
+ "unique_id": ["A","A","B","B"],
+ "y": [1, 3, 2, 6]
+})
+# Forecast data
+df = pd.DataFrame({
+ "unique_id": ["A","A","B","B"],
+ "y": [3, 3, 2, 8],
+ "y_hat": [6, 2, 3, 5]
+})
+# For A: errors = |3−6|+|3−2| = 3+1 = 4 ÷ mean(1,3)=2 → 2
+# For B: errors = |2−3|+|8−5| = 1+3 = 4 ÷ mean(2,6)=4 → 1
+expected = pd.DataFrame({
+ "unique_id": ["A", "B"],
+ "y_hat": [2.0, 1.0]
+ })
+
+# pandas branch
+out_pd = spis(
+ df = df,
+ df_train = df_train,
+ models = ["y_hat"],
+ id_col = "unique_id",
+ target_col = "y",
+)
+pd.testing.assert_frame_equal(
+ out_pd,
+ expected
+)
+```
+
+
+```python
+df_train_pl = pl.from_pandas(df_train)
+df_pl = pl.from_pandas(df)
+out_pl = spis(
+ df = df_pl,
+ df_train = df_train_pl,
+ models = ["y_hat"],
+ id_col = "unique_id",
+ target_col = "y",
+)
+pl.testing.assert_frame_equal(
+ out_pl,
+ pl.from_pandas(expected)
+)
+```
+
+## 2. Percentage Errors
+
+### Mean Absolute Percentage Error
+
+$$
+
+\mathrm{MAPE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}) = \frac{1}{H} \sum^{t+H}_{\tau=t+1} \frac{|y_{\tau}-\hat{y}_{\tau}|}{|y_{\tau}|}
+
+$$
+
+
+
+------------------------------------------------------------------------
+
+source
+
+#### mape
+
+> ``` text
+> mape (df:~DFType, models:List[str], id_col:str='unique_id',
+> target_col:str='y')
+> ```
+
+\*Mean Absolute Percentage Error (MAPE)
+
+MAPE measures the relative prediction accuracy of a forecasting method
+by calculating the percentual deviation of the prediction and the
+observed value at a given time and averages these devations over the
+length of the series. The closer to zero an observed value is, the
+higher penalty MAPE loss assigns to the corresponding error.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, actual values and predictions. |
+| models | List | | Columns that identify the models predictions. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+
+```python
+pd_vs_pl(
+ mape(series, models),
+ mape(series_pl, models),
+ models,
+)
+```
+
+### Symmetric Mean Absolute Percentage Error
+
+$$
+
+\mathrm{SMAPE}_{2}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}) = \frac{1}{H} \sum^{t+H}_{\tau=t+1} \frac{|y_{\tau}-\hat{y}_{\tau}|}{|y_{\tau}|+|\hat{y}_{\tau}|}
+
+$$
+
+------------------------------------------------------------------------
+
+source
+
+#### smape
+
+> ``` text
+> smape (df:~DFType, models:List[str], id_col:str='unique_id',
+> target_col:str='y')
+> ```
+
+\*Symmetric Mean Absolute Percentage Error (SMAPE)
+
+SMAPE measures the relative prediction accuracy of a forecasting method
+by calculating the relative deviation of the prediction and the observed
+value scaled by the sum of the absolute values for the prediction and
+observed value at a given time, then averages these devations over the
+length of the series. This allows the SMAPE to have bounds between 0%
+and 100% which is desirable compared to normal MAPE that may be
+undetermined when the target is zero.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, actual values and predictions. |
+| models | List | | Columns that identify the models predictions. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+
+```python
+pd_vs_pl(
+ smape(series, models),
+ smape(series_pl, models),
+ models,
+)
+```
+
+## 3. Scale-independent Errors
+
+### Mean Absolute Scaled Error
+
+$$
+
+\mathrm{MASE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}, \mathbf{\hat{y}}^{season}_{\tau}) =
+\frac{1}{H} \sum^{t+H}_{\tau=t+1} \frac{|y_{\tau}-\hat{y}_{\tau}|}{\mathrm{MAE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{season}_{\tau})}
+
+$$
+
+
+
+------------------------------------------------------------------------
+
+source
+
+#### mase
+
+> ``` text
+> mase (df:~DFType, models:List[str], seasonality:int, train_df:~DFType,
+> id_col:str='unique_id', target_col:str='y')
+> ```
+
+\*Mean Absolute Scaled Error (MASE)
+
+MASE measures the relative prediction accuracy of a forecasting method
+by comparinng the mean absolute errors of the prediction and the
+observed value against the mean absolute errors of the seasonal naive
+model. The MASE partially composed the Overall Weighted Average (OWA),
+used in the M4 Competition.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, actuals and predictions. |
+| models | List | | Columns that identify the models predictions. |
+| seasonality | int | | Main frequency of the time series; Hourly 24, Daily 7, Weekly 52, Monthly 12, Quarterly 4, Yearly 1. |
+| train_df | DFType | | Training dataframe with id and actual values. Must be sorted by time. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+
+```python
+pd_vs_pl(
+ mase(series, models, 7, series),
+ mase(series_pl, models, 7, series_pl),
+ models,
+)
+```
+
+### Relative Mean Absolute Error
+
+$$
+
+\mathrm{RMAE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}, \mathbf{\hat{y}}^{base}_{\tau}) = \frac{1}{H} \sum^{t+H}_{\tau=t+1} \frac{|y_{\tau}-\hat{y}_{\tau}|}{\mathrm{MAE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{base}_{\tau})}
+
+$$
+
+
+
+------------------------------------------------------------------------
+
+source
+
+#### rmae
+
+> ``` text
+> rmae (df:~DFType, models:List[str], baseline:str, id_col:str='unique_id',
+> target_col:str='y')
+> ```
+
+\*Relative Mean Absolute Error (RMAE)
+
+Calculates the RAME between two sets of forecasts (from two different
+forecasting methods). A number smaller than one implies that the
+forecast in the numerator is better than the forecast in the
+denominator.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, times, actuals and predictions. |
+| models | List | | Columns that identify the models predictions. |
+| baseline | str | | Column that identifies the baseline model predictions. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+
+```python
+pd_vs_pl(
+ rmae(series, models, models[0]),
+ rmae(series_pl, models, models[0]),
+ models,
+)
+```
+
+### Normalized Deviation
+
+$$
+
+\mathrm{ND}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}) = \frac{\sum^{t+H}_{\tau=t+1} |y_{\tau} - \hat{y}_{\tau}|}{\sum^{t+H}_{\tau=t+1} | y_{\tau} |}
+
+$$
+
+------------------------------------------------------------------------
+
+source
+
+#### nd
+
+> ``` text
+> nd (df:~DFType, models:List[str], id_col:str='unique_id',
+> target_col:str='y')
+> ```
+
+\*Normalized Deviation (ND)
+
+ND measures the relative prediction accuracy of a forecasting method by
+calculating the sum of the absolute deviation of the prediction and the
+true value at a given time and dividing it by the sum of the absolute
+value of the ground truth.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, times, actuals and predictions. |
+| models | List | | Columns that identify the models predictions. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+
+```python
+pd_vs_pl(
+ nd(series, models),
+ nd(series_pl, models),
+ models,
+)
+```
+
+### Mean Squared Scaled Error
+
+$$
+
+\mathrm{MSSE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}, \mathbf{\hat{y}}^{season}_{\tau}) =
+\frac{1}{H} \sum^{t+H}_{\tau=t+1} \frac{(y_{\tau}-\hat{y}_{\tau})^2}{\mathrm{MSE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{season}_{\tau})}
+
+$$
+
+------------------------------------------------------------------------
+
+source
+
+#### msse
+
+> ``` text
+> msse (df:~DFType, models:List[str], seasonality:int, train_df:~DFType,
+> id_col:str='unique_id', target_col:str='y')
+> ```
+
+\*Mean Squared Scaled Error (MSSE)
+
+MSSE measures the relative prediction accuracy of a forecasting method
+by comparinng the mean squared errors of the prediction and the observed
+value against the mean squared errors of the seasonal naive model.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, actuals and predictions. |
+| models | List | | Columns that identify the models predictions. |
+| seasonality | int | | Main frequency of the time series; Hourly 24, Daily 7, Weekly 52, Monthly 12, Quarterly 4, Yearly 1. |
+| train_df | DFType | | Training dataframe with id and actual values. Must be sorted by time. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+
+```python
+pd_vs_pl(
+ msse(series, models, 7, series),
+ msse(series_pl, models, 7, series_pl),
+ models,
+)
+```
+
+### Root Mean Squared Scaled Error
+
+$$
+
+\mathrm{RMSSE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}_{\tau}, \mathbf{\hat{y}}^{season}_{\tau}) =
+\sqrt{\frac{1}{H} \sum^{t+H}_{\tau=t+1} \frac{(y_{\tau}-\hat{y}_{\tau})^2}{\mathrm{MSE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{season}_{\tau})}}
+
+$$
+
+------------------------------------------------------------------------
+
+source
+
+#### rmsse
+
+> ``` text
+> rmsse (df:~DFType, models:List[str], seasonality:int, train_df:~DFType,
+> id_col:str='unique_id', target_col:str='y')
+> ```
+
+\*Root Mean Squared Scaled Error (RMSSE)
+
+MSSE measures the relative prediction accuracy of a forecasting method
+by comparinng the mean squared errors of the prediction and the observed
+value against the mean squared errors of the seasonal naive model.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, actuals and predictions. |
+| models | List | | Columns that identify the models predictions. |
+| seasonality | int | | Main frequency of the time series; Hourly 24, Daily 7, Weekly 52, Monthly 12, Quarterly 4, Yearly 1. |
+| train_df | DFType | | Training dataframe with id and actual values. Must be sorted by time. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+
+```python
+pd_vs_pl(
+ rmsse(series, models, 7, series),
+ rmsse(series_pl, models, 7, series_pl),
+ models,
+)
+```
+
+## 4. Probabilistic Errors
+
+### Quantile Loss
+
+$$
+
+\mathrm{QL}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{(q)}_{\tau}) =
+\frac{1}{H} \sum^{t+H}_{\tau=t+1}
+\Big( (1-q)\,( \hat{y}^{(q)}_{\tau} - y_{\tau} )_{+}
++ q\,( y_{\tau} - \hat{y}^{(q)}_{\tau} )_{+} \Big)
+
+$$
+
+
+
+------------------------------------------------------------------------
+
+source
+
+#### quantile_loss
+
+> ``` text
+> quantile_loss (df:~DFType, models:Dict[str,str], q:float=0.5,
+> id_col:str='unique_id', target_col:str='y')
+> ```
+
+\*Quantile Loss (QL)
+
+QL measures the deviation of a quantile forecast. By weighting the
+absolute deviation in a non symmetric way, the loss pays more attention
+to under or over estimation.
+A common value for q is 0.5 for the deviation from the median.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, times, actuals and predictions. |
+| models | Dict | | Mapping from model name to the model predictions for the specified quantile. |
+| q | float | 0.5 | Quantile for the predictions’ comparison. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+### Scaled Quantile Loss
+
+$$
+
+\mathrm{SQL}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{(q)}_{\tau}) =
+\frac{1}{H} \sum^{t+H}_{\tau=t+1}
+\frac{(1-q)\,( \hat{y}^{(q)}_{\tau} - y_{\tau} )_{+}
++ q\,( y_{\tau} - \hat{y}^{(q)}_{\tau} )_{+}}{\mathrm{MAE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{season}_{\tau})}
+
+$$
+
+------------------------------------------------------------------------
+
+source
+
+### scaled_quantile_loss
+
+> ``` text
+> scaled_quantile_loss (df:~DFType, models:Dict[str,str], seasonality:int,
+> train_df:~DFType, q:float=0.5,
+> id_col:str='unique_id', target_col:str='y')
+> ```
+
+\*Scaled Quantile Loss (SQL)
+
+SQL measures the deviation of a quantile forecast scaled by the mean
+absolute errors of the seasonal naive model. By weighting the absolute
+deviation in a non symmetric way, the loss pays more attention to under
+or over estimation. A common value for q is 0.5 for the deviation from
+the median. This was the official measure used in the M5 Uncertainty
+competition with seasonality = 1.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, times, actuals and predictions. |
+| models | Dict | | Mapping from model name to the model predictions for the specified quantile. |
+| seasonality | int | | Main frequency of the time series; Hourly 24, Daily 7, Weekly 52, Monthly 12, Quarterly 4, Yearly 1. |
+| train_df | DFType | | Training dataframe with id and actual values. Must be sorted by time. |
+| q | float | 0.5 | Quantile for the predictions’ comparison. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+### Multi-Quantile Loss
+
+$$
+
+\mathrm{MQL}(\mathbf{y}_{\tau},
+[\mathbf{\hat{y}}^{(q_{1})}_{\tau}, ... ,\hat{y}^{(q_{n})}_{\tau}]) =
+\frac{1}{n} \sum_{q_{i}} \mathrm{QL}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{(q_{i})}_{\tau})
+
+$$
+
+
+
+------------------------------------------------------------------------
+
+source
+
+#### mqloss
+
+> ``` text
+> mqloss (df:~DFType, models:Dict[str,List[str]], quantiles:numpy.ndarray,
+> id_col:str='unique_id', target_col:str='y')
+> ```
+
+\*Multi-Quantile loss (MQL)
+
+MQL calculates the average multi-quantile Loss for a given set of
+quantiles, based on the absolute difference between predicted quantiles
+and observed values.
+
+The limit behavior of MQL allows to measure the accuracy of a full
+predictive distribution with the continuous ranked probability score
+(CRPS). This can be achieved through a numerical integration technique,
+that discretizes the quantiles and treats the CRPS integral with a left
+Riemann approximation, averaging over uniformly distanced quantiles.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, times, actuals and predictions. |
+| models | Dict | | Mapping from model name to the model predictions for each quantile. |
+| quantiles | ndarray | | Quantiles to compare against. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+
+```python
+pd_vs_pl(
+ mqloss(series, mq_models, quantiles=quantiles),
+ mqloss(series_pl, mq_models, quantiles=quantiles),
+ models,
+)
+```
+
+### Scaled Multi-Quantile Loss
+
+$$
+
+\mathrm{MQL}(\mathbf{y}_{\tau},
+[\mathbf{\hat{y}}^{(q_{1})}_{\tau}, ... ,\hat{y}^{(q_{n})}_{\tau}]) =
+\frac{1}{n} \sum_{q_{i}} \frac{\mathrm{QL}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{(q_{i})}_{\tau})}{\mathrm{MAE}(\mathbf{y}_{\tau}, \mathbf{\hat{y}}^{season}_{\tau})}
+
+$$
+
+------------------------------------------------------------------------
+
+source
+
+### scaled_mqloss
+
+> ``` text
+> scaled_mqloss (df:~DFType, models:Dict[str,List[str]],
+> quantiles:numpy.ndarray, seasonality:int,
+> train_df:~DFType, id_col:str='unique_id',
+> target_col:str='y')
+> ```
+
+\*Scaled Multi-Quantile loss (SMQL)
+
+SMQL calculates the average multi-quantile Loss for a given set of
+quantiles, based on the absolute difference between predicted quantiles
+and observed values scaled by the mean absolute errors of the seasonal
+naive model. The limit behavior of MQL allows to measure the accuracy of
+a full predictive distribution with the continuous ranked probability
+score (CRPS). This can be achieved through a numerical integration
+technique, that discretizes the quantiles and treats the CRPS integral
+with a left Riemann approximation, averaging over uniformly distanced
+quantiles. This was the official measure used in the M5 Uncertainty
+competition with seasonality = 1.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, times, actuals and predictions. |
+| models | Dict | | Mapping from model name to the model predictions for each quantile. |
+| quantiles | ndarray | | Quantiles to compare against. |
+| seasonality | int | | Main frequency of the time series; Hourly 24, Daily 7, Weekly 52, Monthly 12, Quarterly 4, Yearly 1. |
+| train_df | DFType | | Training dataframe with id and actual values. Must be sorted by time. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+
+```python
+pd_vs_pl(
+ scaled_mqloss(series, mq_models, quantiles=quantiles, seasonality=1, train_df=series),
+ scaled_mqloss(series_pl, mq_models, quantiles=quantiles, seasonality=1, train_df=series_pl),
+ models,
+)
+```
+
+### Coverage
+
+------------------------------------------------------------------------
+
+source
+
+#### coverage
+
+> ``` text
+> coverage (df:~DFType, models:List[str], level:int,
+> id_col:str='unique_id', target_col:str='y')
+> ```
+
+*Coverage of y with y_hat_lo and y_hat_hi.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, times, actuals and predictions. |
+| models | List | | Columns that identify the models predictions. |
+| level | int | | Confidence level used for intervals. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+
+```python
+pd_vs_pl(
+ coverage(series, models, 80),
+ coverage(series_pl, models, 80),
+ models,
+)
+```
+
+### Calibration
+
+------------------------------------------------------------------------
+
+source
+
+#### calibration
+
+> ``` text
+> calibration (df:~DFType, models:Dict[str,str], id_col:str='unique_id',
+> target_col:str='y')
+> ```
+
+*Fraction of y that is lower than the model’s predictions.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, times, actuals and predictions. |
+| models | Dict | | Mapping from model name to the model predictions. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+
+```python
+pd_vs_pl(
+ calibration(series, q_models[0.1]),
+ calibration(series_pl, q_models[0.1]),
+ models,
+)
+```
+
+### CRPS
+
+$$
+
+\mathrm{sCRPS}(\hat{F}_{\tau}, \mathbf{y}_{\tau}) = \frac{2}{N} \sum_{i}
+\int^{1}_{0} \frac{\mathrm{QL}(\hat{F}_{i,\tau}, y_{i,\tau})_{q}}{\sum_{i} | y_{i,\tau} |} dq
+
+$$
+
+Where $\hat{F}_{\tau}$ is the an estimated multivariate distribution,
+and $y_{i,\tau}$ are its realizations.
+
+------------------------------------------------------------------------
+
+source
+
+#### scaled_crps
+
+> ``` text
+> scaled_crps (df:~DFType, models:Dict[str,List[str]],
+> quantiles:numpy.ndarray, id_col:str='unique_id',
+> target_col:str='y')
+> ```
+
+\*Scaled Continues Ranked Probability Score
+
+Calculates a scaled variation of the CRPS, as proposed by Rangapuram
+(2021), to measure the accuracy of predicted quantiles `y_hat` compared
+to the observation `y`. This metric averages percentual weighted
+absolute deviations as defined by the quantile losses.\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, times, actuals and predictions. |
+| models | Dict | | Mapping from model name to the model predictions for each quantile. |
+| quantiles | ndarray | | Quantiles to compare against. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **dataframe with one row per id and one column per model.** |
+
+
+```python
+pd_vs_pl(
+ scaled_crps(series, mq_models, quantiles),
+ scaled_crps(series_pl, mq_models, quantiles),
+ models,
+)
+```
+
+### Tweedie Deviance
+
+For a set of forecasts $\{\mu_i\}_{i=1}^N$ and observations
+$\{y_i\}_{i=1}^N$, the mean Tweedie deviance with power $p$ is
+
+$$
+
+\mathrm{TD}_{p}(\boldsymbol{\mu}, \mathbf{y})
+= \frac{1}{N} \sum_{i=1}^{N} d_{p}(y_i, \mu_i)
+
+$$
+
+where the unit-scaled deviance for each pair $(y,\mu)$ is
+
+$$
+
+d_{p}(y,\mu)
+=
+2
+\begin{cases}
+\displaystyle
+\frac{y^{2-p}}{(1-p)(2-p)}
+\;-\;
+\frac{y\,\mu^{1-p}}{1-p}
+\;+\;
+\frac{\mu^{2-p}}{2-p},
+& p \notin\{1,2\},\\[1em]
+\displaystyle
+y\,\ln\!\frac{y}{\mu}\;-\;(y-\mu),
+& p = 1\quad(\text{Poisson deviance}),\\[0.5em]
+\displaystyle
+-2\Bigl[\ln\!\frac{y}{\mu}\;-\;\frac{y-\mu}{\mu}\Bigr],
+& p = 2\quad(\text{Gamma deviance}).
+\end{cases}
+
+$$
+
+- $y_i$ are the true values, $\mu_i$ the predicted means.
+- $p$ controls the variance relationship
+ $\mathrm{Var}(Y)\propto\mu^{p}$.
+- When $1source
+
+#### tweedie_deviance
+
+> ``` text
+> tweedie_deviance (df:~DFType, models:List[str], power:float=1.5,
+> id_col:str='unique_id', target_col:str='y')
+> ```
+
+\*Compute the Tweedie deviance loss for one or multiple models, grouped
+by an identifier.
+
+Each group’s deviance is calculated using the mean_tweedie_deviance
+function, which measures the deviation between actual and predicted
+values under the Tweedie distribution.
+
+The `power` parameter defines the specific compound distribution: - 1:
+Poisson - (1, 2): Compound Poisson-Gamma - 2: Gamma - \>2: Inverse
+Gaussian\*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input dataframe with id, actuals and predictions. |
+| models | List | | Columns that identify the models predictions. |
+| power | float | 1.5 | Tweedie power parameter. Determines the compound distribution. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| target_col | str | y | Column that contains the target. |
+| **Returns** | **DFType** | | **DataFrame with one row per id and one column per model, containing the mean Tweedie deviance. ** |
+
+
+```python
+# Normal test
+for power in [0, 1, 1.5, 2, 3]:
+ # Test Pandas vs Polars
+ td_pd = tweedie_deviance(series, models, target_col="y", power=power)
+ td_pl = tweedie_deviance(series_pl, models, target_col="y", power=power)
+ pd_vs_pl(
+ td_pd,
+ td_pl,
+ models,
+ )
+ # Test for NaNs
+ assert not td_pd[models].isna().any().any(), f"NaNs found in pd DataFrame for power {power}"
+ assert not td_pl.select(pl.col(models).is_null().any()).sum_horizontal().item(), f"NaNs found in pl DataFrame for power {power}"
+ # Test for infinites
+ is_infinite = td_pd[models].isin([np.inf, -np.inf]).any().any()
+ assert not is_infinite, f"Infinities found in pd DataFrame for power {power}"
+ is_infinite_pl = td_pl.select(pl.col(models).is_infinite().any()).sum_horizontal().item()
+ assert not is_infinite_pl, f"Infinities found in pl DataFrame for power {power}"
+
+# Test zero handling (skip power >=2 since it requires all y > 0)
+series.loc[0, 'y'] = 0.0 # Set a zero value to test the zero handling
+series.loc[49, 'y'] = 0.0 # Set another zero value to test the zero handling
+series_pl[0, 'y'] = 0.0 # Set a zero value to test the zero handling
+series_pl[49, 'y'] = 0.0 # Set another zero value to test the zero handling
+for power in [0, 1, 1.5]:
+ # Test Pandas vs Polars
+ td_pd = tweedie_deviance(series, models, target_col="y", power=power)
+ td_pl = tweedie_deviance(series_pl, models, target_col="y", power=power)
+ pd_vs_pl(
+ td_pd,
+ td_pl,
+ models,
+ )
+ # Test for NaNs
+ assert not td_pd[models].isna().any().any(), f"NaNs found in pd DataFrame for power {power}"
+ assert not td_pl.select(pl.col(models).is_null().any()).sum_horizontal().item(), f"NaNs found in pl DataFrame for power {power}"
+ # Test for infinites
+ is_infinite = td_pd[models].isin([np.inf, -np.inf]).any().any()
+ assert not is_infinite, f"Infinities found in pd DataFrame for power {power}"
+ is_infinite_pl = td_pl.select(pl.col(models).is_infinite().any()).sum_horizontal().item()
+ assert not is_infinite_pl, f"Infinities found in pl DataFrame for power {power}"
+```
+
diff --git a/utilsforecast/mint.json b/utilsforecast/mint.json
new file mode 100644
index 00000000..6ef163e2
--- /dev/null
+++ b/utilsforecast/mint.json
@@ -0,0 +1,39 @@
+{
+ "$schema": "https://mintlify.com/schema.json",
+ "name": "Nixtla",
+ "logo": {
+ "light": "/light.png",
+ "dark": "/dark.png"
+ },
+ "favicon": "/favicon.svg",
+ "colors": {
+ "primary": "#0E0E0E",
+ "light": "#FAFAFA",
+ "dark": "#0E0E0E",
+ "anchors": {
+ "from": "#2AD0CA",
+ "to": "#0E00F8"
+ }
+ },
+ "topbarCtaButton": {
+ "type": "github",
+ "url": "https://github.com/Nixtla/utilsforecast"
+ },
+ "navigation": [
+ {
+ "group": "",
+ "pages": ["index.html"]
+ },
+ {
+ "group": "API Reference",
+ "pages": [
+ "preprocessing.html",
+ "feature_engineering.html",
+ "evaluation.html",
+ "losses.html",
+ "plotting.html",
+ "data.html"
+ ]
+ }
+ ]
+}
diff --git a/utilsforecast/plotting.html.mdx b/utilsforecast/plotting.html.mdx
new file mode 100644
index 00000000..59874d81
--- /dev/null
+++ b/utilsforecast/plotting.html.mdx
@@ -0,0 +1,84 @@
+---
+description: Time series visualizations
+output-file: plotting.html
+title: Plotting
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### plot_series
+
+> ``` text
+> plot_series (df:Optional[~DFType]=None,
+> forecasts_df:Optional[~DFType]=None,
+> ids:Optional[List[str]]=None, plot_random:bool=True,
+> max_ids:int=8, models:Optional[List[str]]=None,
+> level:Optional[List[float]]=None,
+> max_insample_length:Optional[int]=None,
+> plot_anomalies:bool=False, engine:str='matplotlib',
+> palette:Optional[str]=None, id_col:str='unique_id',
+> time_col:str='ds', target_col:str='y', seed:int=0,
+> resampler_kwargs:Optional[Dict]=None, ax:Union[matplotlib.ax
+> es._axes.Axes,numpy.ndarray,ForwardRef('plotly.graph_objects
+> .Figure'),NoneType]=None)
+> ```
+
+*Plot forecasts and insample values.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | Optional | None | DataFrame with columns \[`id_col`, `time_col`, `target_col`\]. |
+| forecasts_df | Optional | None | DataFrame with columns \[`id_col`, `time_col`\] and models. |
+| ids | Optional | None | Time Series to plot. If None, time series are selected randomly. |
+| plot_random | bool | True | Select time series to plot randomly. |
+| max_ids | int | 8 | Maximum number of ids to plot. |
+| models | Optional | None | Models to plot. |
+| level | Optional | None | Prediction intervals to plot. |
+| max_insample_length | Optional | None | Maximum number of train/insample observations to be plotted. |
+| plot_anomalies | bool | False | Plot anomalies for each prediction interval. |
+| engine | str | matplotlib | Library used to plot. ‘plotly’, ‘plotly-resampler’ or ‘matplotlib’. |
+| palette | Optional | None | Name of the matplotlib colormap to use for the plots. If None, uses the current style. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestep, its values can be timestamps or integers. |
+| target_col | str | y | Column that contains the target. |
+| seed | int | 0 | Seed used for the random number generator. Only used if plot_random is True. |
+| resampler_kwargs | Optional | None | Keyword arguments to be passed to plotly-resampler constructor. For further custumization (“show_dash”) call the method, store the plotting object and add the extra arguments to its `show_dash` method. |
+| ax | Union | None | Object where plots will be added. |
+| **Returns** | **matplotlib or plotly figure** | | **Plot’s figure** |
+
+
+```python
+from utilsforecast.data import generate_series
+```
+
+
+```python
+level = [80, 95]
+series = generate_series(4, freq='D', equal_ends=True, with_trend=True, n_models=2, level=level)
+test_pd = series.groupby('unique_id', observed=True).tail(10).copy()
+train_pd = series.drop(test_pd.index)
+```
+
+
+```python
+plt.style.use('ggplot')
+fig = plot_series(
+ train_pd,
+ forecasts_df=test_pd,
+ ids=[0, 3],
+ plot_random=False,
+ level=level,
+ max_insample_length=50,
+ engine='matplotlib',
+ plot_anomalies=True,
+)
+fig.savefig('imgs/plotting.png', bbox_inches='tight')
+```
+
+
+
diff --git a/utilsforecast/preprocessing.html.mdx b/utilsforecast/preprocessing.html.mdx
new file mode 100644
index 00000000..d00ba12e
--- /dev/null
+++ b/utilsforecast/preprocessing.html.mdx
@@ -0,0 +1,387 @@
+---
+description: Utilities for processing data before training/analysis
+output-file: preprocessing.html
+title: Preprocessing
+---
+
+
+------------------------------------------------------------------------
+
+source
+
+### id_time_grid
+
+> ``` text
+> id_time_grid (df:~DFType, freq:Union[str,int],
+> start:Union[str,int,datetime.date,datetime.datetime]='per_s
+> erie', end:Union[str,int,datetime.date,datetime.datetime]='
+> global', id_col:str='unique_id', time_col:str='ds')
+> ```
+
+*Generate all expected combiations of ids and times.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input data |
+| freq | Union | | Series’ frequency |
+| start | Union | per_serie | Initial timestamp for the series. \* ‘per_serie’ uses each serie’s first timestamp \* ‘global’ uses the first timestamp seen in the data \* Can also be a specific timestamp or integer, e.g. ‘2000-01-01’, 2000 or datetime(2000, 1, 1) |
+| end | Union | global | Initial timestamp for the series. \* ‘per_serie’ uses each serie’s last timestamp \* ‘global’ uses the last timestamp seen in the data \* Can also be a specific timestamp or integer, e.g. ‘2000-01-01’, 2000 or datetime(2000, 1, 1) |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestamp. |
+| **Returns** | **DFType** | | **Dataframe with expected ids and times.** |
+
+------------------------------------------------------------------------
+
+source
+
+### fill_gaps
+
+> ``` text
+> fill_gaps (df:~DFType, freq:Union[str,int],
+> start:Union[str,int,datetime.date,datetime.datetime]='per_seri
+> e',
+> end:Union[str,int,datetime.date,datetime.datetime]='global',
+> id_col:str='unique_id', time_col:str='ds')
+> ```
+
+*Enforce start and end datetimes for dataframe.*
+
+| | **Type** | **Default** | **Details** |
+|------|------------------|-------------------------|-------------------------|
+| df | DFType | | Input data |
+| freq | Union | | Series’ frequency |
+| start | Union | per_serie | Initial timestamp for the series. \* ‘per_serie’ uses each serie’s first timestamp \* ‘global’ uses the first timestamp seen in the data \* Can also be a specific timestamp or integer, e.g. ‘2000-01-01’, 2000 or datetime(2000, 1, 1) |
+| end | Union | global | Initial timestamp for the series. \* ‘per_serie’ uses each serie’s last timestamp \* ‘global’ uses the last timestamp seen in the data \* Can also be a specific timestamp or integer, e.g. ‘2000-01-01’, 2000 or datetime(2000, 1, 1) |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestamp. |
+| **Returns** | **DFType** | | **Dataframe with gaps filled.** |
+
+
+```python
+df = pd.DataFrame(
+ {
+ 'unique_id': [0, 0, 0, 1, 1],
+ 'ds': pd.to_datetime(['2020', '2021', '2023', '2021', '2022']),
+ 'y': np.arange(5),
+ }
+)
+df
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|-----|
+| 0 | 0 | 2020-01-01 | 0 |
+| 1 | 0 | 2021-01-01 | 1 |
+| 2 | 0 | 2023-01-01 | 2 |
+| 3 | 1 | 2021-01-01 | 3 |
+| 4 | 1 | 2022-01-01 | 4 |
+
+The default functionality is taking the current starts and only
+extending the end date to be the same for all series.
+
+
+```python
+fill_gaps(
+ df,
+ freq='YS',
+)
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|-----|
+| 0 | 0 | 2020-01-01 | 0.0 |
+| 1 | 0 | 2021-01-01 | 1.0 |
+| 2 | 0 | 2022-01-01 | NaN |
+| 3 | 0 | 2023-01-01 | 2.0 |
+| 4 | 1 | 2021-01-01 | 3.0 |
+| 5 | 1 | 2022-01-01 | 4.0 |
+| 6 | 1 | 2023-01-01 | NaN |
+
+We can also specify `end='per_serie'` to only fill possible gaps within
+each serie.
+
+
+```python
+fill_gaps(
+ df,
+ freq='YS',
+ end='per_serie',
+)
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|-----|
+| 0 | 0 | 2020-01-01 | 0.0 |
+| 1 | 0 | 2021-01-01 | 1.0 |
+| 2 | 0 | 2022-01-01 | NaN |
+| 3 | 0 | 2023-01-01 | 2.0 |
+| 4 | 1 | 2021-01-01 | 3.0 |
+| 5 | 1 | 2022-01-01 | 4.0 |
+
+We can also specify an end date in the future.
+
+
+```python
+fill_gaps(
+ df,
+ freq='YS',
+ end='2024',
+)
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|-----|
+| 0 | 0 | 2020-01-01 | 0.0 |
+| 1 | 0 | 2021-01-01 | 1.0 |
+| 2 | 0 | 2022-01-01 | NaN |
+| 3 | 0 | 2023-01-01 | 2.0 |
+| 4 | 0 | 2024-01-01 | NaN |
+| 5 | 1 | 2021-01-01 | 3.0 |
+| 6 | 1 | 2022-01-01 | 4.0 |
+| 7 | 1 | 2023-01-01 | NaN |
+| 8 | 1 | 2024-01-01 | NaN |
+
+We can set all series to start at the same time.
+
+
+```python
+fill_gaps(
+ df,
+ freq='YS',
+ start='global'
+)
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|-----|
+| 0 | 0 | 2020-01-01 | 0.0 |
+| 1 | 0 | 2021-01-01 | 1.0 |
+| 2 | 0 | 2022-01-01 | NaN |
+| 3 | 0 | 2023-01-01 | 2.0 |
+| 4 | 1 | 2020-01-01 | NaN |
+| 5 | 1 | 2021-01-01 | 3.0 |
+| 6 | 1 | 2022-01-01 | 4.0 |
+| 7 | 1 | 2023-01-01 | NaN |
+
+We can also set a common start date for all series (which can be earlier
+than their current starts).
+
+
+```python
+fill_gaps(
+ df,
+ freq='YS',
+ start='2019',
+)
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------------|-----|
+| 0 | 0 | 2019-01-01 | NaN |
+| 1 | 0 | 2020-01-01 | 0.0 |
+| 2 | 0 | 2021-01-01 | 1.0 |
+| 3 | 0 | 2022-01-01 | NaN |
+| 4 | 0 | 2023-01-01 | 2.0 |
+| 5 | 1 | 2019-01-01 | NaN |
+| 6 | 1 | 2020-01-01 | NaN |
+| 7 | 1 | 2021-01-01 | 3.0 |
+| 8 | 1 | 2022-01-01 | 4.0 |
+| 9 | 1 | 2023-01-01 | NaN |
+
+In case the times are integers the frequency, start and end must also be
+integers.
+
+
+```python
+df = pd.DataFrame(
+ {
+ 'unique_id': [0, 0, 0, 1, 1],
+ 'ds': [2020, 2021, 2023, 2021, 2022],
+ 'y': np.arange(5),
+ }
+)
+df
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------|-----|
+| 0 | 0 | 2020 | 0 |
+| 1 | 0 | 2021 | 1 |
+| 2 | 0 | 2023 | 2 |
+| 3 | 1 | 2021 | 3 |
+| 4 | 1 | 2022 | 4 |
+
+
+```python
+fill_gaps(
+ df,
+ freq=1,
+ start=2019,
+ end=2024,
+)
+```
+
+| | unique_id | ds | y |
+|-----|-----------|------|-----|
+| 0 | 0 | 2019 | NaN |
+| 1 | 0 | 2020 | 0.0 |
+| 2 | 0 | 2021 | 1.0 |
+| 3 | 0 | 2022 | NaN |
+| 4 | 0 | 2023 | 2.0 |
+| 5 | 0 | 2024 | NaN |
+| 6 | 1 | 2019 | NaN |
+| 7 | 1 | 2020 | NaN |
+| 8 | 1 | 2021 | 3.0 |
+| 9 | 1 | 2022 | 4.0 |
+| 10 | 1 | 2023 | NaN |
+| 11 | 1 | 2024 | NaN |
+
+The function also accepts polars dataframes
+
+
+```python
+df = pl.DataFrame(
+ {
+ 'unique_id': [0, 0, 0, 1, 1],
+ 'ds': [
+ datetime(2020, 1, 1), datetime(2022, 1, 1), datetime(2023, 1, 1),
+ datetime(2021, 1, 1), datetime(2022, 1, 1)],
+ 'y': np.arange(5),
+ }
+)
+df
+```
+
+| unique_id | ds | y |
+|-----------|---------------------|-----|
+| i64 | datetime\[μs\] | i64 |
+| 0 | 2020-01-01 00:00:00 | 0 |
+| 0 | 2022-01-01 00:00:00 | 1 |
+| 0 | 2023-01-01 00:00:00 | 2 |
+| 1 | 2021-01-01 00:00:00 | 3 |
+| 1 | 2022-01-01 00:00:00 | 4 |
+
+
+```python
+polars_ms = fill_gaps(
+ df.with_columns(pl.col('ds').cast(pl.Datetime(time_unit='ms'))),
+ freq='1y',
+ start=datetime(2019, 1, 1),
+ end=datetime(2024, 1, 1),
+)
+assert polars_ms.schema['ds'].time_unit == 'ms'
+polars_ms
+```
+
+| unique_id | ds | y |
+|-----------|---------------------|------|
+| i64 | datetime\[ms\] | i64 |
+| 0 | 2019-01-01 00:00:00 | null |
+| 0 | 2020-01-01 00:00:00 | 0 |
+| 0 | 2021-01-01 00:00:00 | null |
+| 0 | 2022-01-01 00:00:00 | 1 |
+| 0 | 2023-01-01 00:00:00 | 2 |
+| … | … | … |
+| 1 | 2020-01-01 00:00:00 | null |
+| 1 | 2021-01-01 00:00:00 | 3 |
+| 1 | 2022-01-01 00:00:00 | 4 |
+| 1 | 2023-01-01 00:00:00 | null |
+| 1 | 2024-01-01 00:00:00 | null |
+
+
+```python
+df = pl.DataFrame(
+ {
+ 'unique_id': [0, 0, 0, 1, 1],
+ 'ds': [
+ date(2020, 1, 1), date(2022, 1, 1), date(2023, 1, 1),
+ date(2021, 1, 1), date(2022, 1, 1)],
+ 'y': np.arange(5),
+ }
+)
+df
+```
+
+| unique_id | ds | y |
+|-----------|------------|-----|
+| i64 | date | i64 |
+| 0 | 2020-01-01 | 0 |
+| 0 | 2022-01-01 | 1 |
+| 0 | 2023-01-01 | 2 |
+| 1 | 2021-01-01 | 3 |
+| 1 | 2022-01-01 | 4 |
+
+
+```python
+fill_gaps(
+ df,
+ freq='1y',
+ start=date(2020, 1, 1),
+ end=date(2024, 1, 1),
+)
+```
+
+| unique_id | ds | y |
+|-----------|------------|------|
+| i64 | date | i64 |
+| 0 | 2020-01-01 | 0 |
+| 0 | 2021-01-01 | null |
+| 0 | 2022-01-01 | 1 |
+| 0 | 2023-01-01 | 2 |
+| 0 | 2024-01-01 | null |
+| 1 | 2020-01-01 | null |
+| 1 | 2021-01-01 | 3 |
+| 1 | 2022-01-01 | 4 |
+| 1 | 2023-01-01 | null |
+| 1 | 2024-01-01 | null |
+
+
+```python
+df = pl.DataFrame(
+ {
+ 'unique_id': [0, 0, 0, 1, 1],
+ 'ds': [2020, 2021, 2023, 2021, 2022],
+ 'y': np.arange(5),
+ }
+)
+df
+```
+
+| unique_id | ds | y |
+|-----------|------|-----|
+| i64 | i64 | i64 |
+| 0 | 2020 | 0 |
+| 0 | 2021 | 1 |
+| 0 | 2023 | 2 |
+| 1 | 2021 | 3 |
+| 1 | 2022 | 4 |
+
+
+```python
+fill_gaps(
+ df,
+ freq=1,
+ start=2019,
+ end=2024,
+)
+```
+
+| unique_id | ds | y |
+|-----------|------|------|
+| i64 | i64 | i64 |
+| 0 | 2019 | null |
+| 0 | 2020 | 0 |
+| 0 | 2021 | 1 |
+| 0 | 2022 | null |
+| 0 | 2023 | 2 |
+| … | … | … |
+| 1 | 2020 | null |
+| 1 | 2021 | 3 |
+| 1 | 2022 | 4 |
+| 1 | 2023 | null |
+| 1 | 2024 | null |
+
diff --git a/utilsforecast/processing.mdx b/utilsforecast/processing.mdx
new file mode 100644
index 00000000..05a98903
--- /dev/null
+++ b/utilsforecast/processing.mdx
@@ -0,0 +1,1146 @@
+
+```python
+import datetime
+from datetime import datetime as dt
+
+from fastcore.test import test_eq, test_fail
+from nbdev import show_doc
+
+from utilsforecast.compat import POLARS_INSTALLED
+from utilsforecast.data import generate_series
+```
+
+
+```python
+import polars.testing
+```
+
+------------------------------------------------------------------------
+
+source
+
+### to_numpy
+
+> ``` text
+> to_numpy
+> (df:Union[pandas.core.frame.DataFrame,polars.dataframe.frame.Da
+> taFrame])
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### counts_by_id
+
+> ``` text
+> counts_by_id
+> (df:Union[pandas.core.frame.DataFrame,polars.dataframe.fram
+> e.DataFrame], id_col:str)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### maybe_compute_sort_indices
+
+> ``` text
+> maybe_compute_sort_indices
+> (df:Union[pandas.core.frame.DataFrame,polars.
+> dataframe.frame.DataFrame], id_col:str,
+> time_col:str)
+> ```
+
+*Compute indices that would sort the dataframe*
+
+| | **Type** | **Details** |
+|--------|---------------------------|-------------------------------------|
+| df | Union | Input dataframe with id, times and target values. |
+| id_col | str | |
+| time_col | str | |
+| **Returns** | **Optional** | **Array with indices to sort the dataframe or None if it’s already sorted.** |
+
+------------------------------------------------------------------------
+
+source
+
+### assign_columns
+
+> ``` text
+> assign_columns
+> (df:Union[pandas.core.frame.DataFrame,polars.dataframe.fr
+> ame.DataFrame], names:Union[str,List[str]], values:Union[
+> numpy.ndarray,pandas.core.series.Series,polars.series.ser
+> ies.Series,List[float]])
+> ```
+
+
+```python
+engines = ['pandas']
+if POLARS_INSTALLED:
+ engines.append('polars')
+```
+
+
+```python
+for engine in engines:
+ series = generate_series(2, engine=engine)
+ x = np.random.rand(series.shape[0])
+ series = assign_columns(series, 'x', x)
+ series = assign_columns(series, ['y', 'z'], np.vstack([x, x]).T)
+ series = assign_columns(series, 'ones', 1)
+ series = assign_columns(series, 'zeros', np.zeros(series.shape[0]))
+ series = assign_columns(series, 'as', 'a')
+ series = assign_columns(series, 'bs', series.shape[0] * ['b'])
+ np.testing.assert_allclose(
+ series[['x', 'y', 'z']],
+ np.vstack([x, x, x]).T
+ )
+ np.testing.assert_equal(series['ones'], np.ones(series.shape[0]))
+ np.testing.assert_equal(series['as'], np.full(series.shape[0], 'a'))
+ np.testing.assert_equal(series['bs'], np.full(series.shape[0], 'b'))
+```
+
+------------------------------------------------------------------------
+
+source
+
+### drop_columns
+
+> ``` text
+> drop_columns
+> (df:Union[pandas.core.frame.DataFrame,polars.dataframe.fram
+> e.DataFrame], columns:Union[str,List[str]])
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### take_rows
+
+> ``` text
+> take_rows (df:Union[pandas.core.frame.DataFrame,polars.dataframe.frame.Da
+> taFrame,pandas.core.series.Series,polars.series.series.Series,
+> numpy.ndarray], idxs:numpy.ndarray)
+> ```
+
+
+```python
+for engine in engines:
+ series = generate_series(2, engine=engine)
+ subset = take_rows(series, np.array([0, 2]))
+ assert subset.shape[0] == 2
+```
+
+------------------------------------------------------------------------
+
+source
+
+### filter_with_mask
+
+> ``` text
+> filter_with_mask (df:Union[pandas.core.series.Series,polars.series.series
+> .Series,pandas.core.frame.DataFrame,polars.dataframe.fr
+> ame.DataFrame,pandas.core.indexes.base.Index,numpy.ndar
+> ray], mask:Union[numpy.ndarray,pandas.core.series.Serie
+> s,polars.series.series.Series])
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### is_nan
+
+> ``` text
+> is_nan (s:Union[pandas.core.series.Series,polars.series.series.Series])
+> ```
+
+
+```python
+np.testing.assert_equal(
+ is_nan(pd.Series([np.nan, 1.0, None])).to_numpy(),
+ np.array([True, False, True]),
+)
+if POLARS_INSTALLED:
+ np.testing.assert_equal(
+ is_nan(pl.Series([np.nan, 1.0, None])).to_numpy(),
+ np.array([True, False, None]),
+ )
+```
+
+------------------------------------------------------------------------
+
+source
+
+### is_none
+
+> ``` text
+> is_none (s:Union[pandas.core.series.Series,polars.series.series.Series])
+> ```
+
+
+```python
+np.testing.assert_equal(
+ is_none(pd.Series([np.nan, 1.0, None])).to_numpy(),
+ np.array([True, False, True]),
+)
+if POLARS_INSTALLED:
+ np.testing.assert_equal(
+ is_none(pl.Series([np.nan, 1.0, None])).to_numpy(),
+ np.array([False, False, True]),
+ )
+```
+
+------------------------------------------------------------------------
+
+source
+
+### is_nan_or_none
+
+> ``` text
+> is_nan_or_none
+> (s:Union[pandas.core.series.Series,polars.series.series.S
+> eries])
+> ```
+
+
+```python
+np.testing.assert_equal(
+ is_nan_or_none(pd.Series([np.nan, 1.0, None])).to_numpy(),
+ np.array([True, False, True]),
+)
+if POLARS_INSTALLED:
+ np.testing.assert_equal(
+ is_nan_or_none(pl.Series([np.nan, 1.0, None])).to_numpy(),
+ np.array([True, False, True]),
+ )
+```
+
+------------------------------------------------------------------------
+
+source
+
+### match_if_categorical
+
+> ``` text
+> match_if_categorical (s1:Union[pandas.core.series.Series,polars.series.se
+> ries.Series,pandas.core.indexes.base.Index], s2:Uni
+> on[pandas.core.series.Series,polars.series.series.S
+> eries])
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### vertical_concat
+
+> ``` text
+> vertical_concat (dfs:List[Union[pandas.core.frame.DataFrame,polars.datafr
+> ame.frame.DataFrame,pandas.core.series.Series,polars.ser
+> ies.series.Series]], match_categories:bool=True)
+> ```
+
+
+```python
+df1 = pd.DataFrame({'x': ['a', 'b', 'c']}, dtype='category')
+df2 = pd.DataFrame({'x': ['f', 'b', 'a']}, dtype='category')
+pd.testing.assert_series_equal(
+ vertical_concat([df1,df2])['x'],
+ pd.Series(['a', 'b', 'c', 'f', 'b', 'a'], name='x', dtype=pd.CategoricalDtype(categories=['a', 'b', 'c', 'f']))
+)
+```
+
+
+```python
+df1 = pl.DataFrame({'x': ['a', 'b', 'c']}, schema={'x': pl.Categorical})
+df2 = pl.DataFrame({'x': ['f', 'b', 'a']}, schema={'x': pl.Categorical})
+out = vertical_concat([df1,df2])['x']
+assert out.equals(pl.Series('x', ['a', 'b', 'c', 'f', 'b', 'a']))
+assert out.to_physical().equals(pl.Series('x', [0, 1, 2, 3, 1, 0]))
+assert out.cat.get_categories().equals(
+ pl.Series('x', ['a', 'b', 'c', 'f'])
+)
+```
+
+
+```python
+for engine in engines:
+ series = generate_series(2, engine=engine)
+ doubled = vertical_concat([series, series])
+ assert doubled.shape[0] == 2 * series.shape[0]
+```
+
+------------------------------------------------------------------------
+
+source
+
+### horizontal_concat
+
+> ``` text
+> horizontal_concat (dfs:List[Union[pandas.core.frame.DataFrame,polars.data
+> frame.frame.DataFrame]])
+> ```
+
+
+```python
+for engine in engines:
+ series = generate_series(2, engine=engine)
+ renamer = {c: f'{c}_2' for c in series.columns}
+ if engine == 'pandas':
+ series2 = series.rename(columns=renamer)
+ else:
+ series2 = series.rename(renamer)
+ doubled = horizontal_concat([series, series2])
+ assert doubled.shape[1] == 2 * series.shape[1]
+```
+
+------------------------------------------------------------------------
+
+source
+
+### copy_if_pandas
+
+> ``` text
+> copy_if_pandas
+> (df:Union[pandas.core.frame.DataFrame,polars.dataframe.fr
+> ame.DataFrame], deep:bool=False)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### join
+
+> ``` text
+> join (df1:Union[pandas.core.frame.DataFrame,polars.dataframe.frame.DataFr
+> ame,pandas.core.series.Series,polars.series.series.Series], df2:Uni
+> on[pandas.core.frame.DataFrame,polars.dataframe.frame.DataFrame,pan
+> das.core.series.Series,polars.series.series.Series],
+> on:Union[str,List[str]], how:str='inner')
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### drop_index_if_pandas
+
+> ``` text
+> drop_index_if_pandas
+> (df:Union[pandas.core.frame.DataFrame,polars.datafr
+> ame.frame.DataFrame])
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### rename
+
+> ``` text
+> rename
+> (df:Union[pandas.core.frame.DataFrame,polars.dataframe.frame.Data
+> Frame], mapping:Dict[str,str])
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### sort
+
+> ``` text
+> sort
+> (df:Union[pandas.core.frame.DataFrame,polars.dataframe.frame.DataFr
+> ame], by:Union[str,List[str],NoneType]=None)
+> ```
+
+
+```python
+pd.testing.assert_frame_equal(
+ sort(pd.DataFrame({'x': [3, 1, 2]}), 'x'),
+ pd.DataFrame({'x': [1, 2, 3]})
+)
+pd.testing.assert_frame_equal(
+ sort(pd.DataFrame({'x': [3, 1, 2]}), ['x']),
+ pd.DataFrame({'x': [1, 2, 3]})
+)
+pd.testing.assert_series_equal(
+ sort(pd.Series([3, 1, 2])),
+ pd.Series([1, 2, 3])
+)
+pd.testing.assert_index_equal(
+ sort(pd.Index([3, 1, 2])),
+ pd.Index([1, 2, 3])
+)
+```
+
+
+```python
+pl.testing.assert_frame_equal(
+ sort(pl.DataFrame({'x': [3, 1, 2]}), 'x'),
+ pl.DataFrame({'x': [1, 2, 3]}),
+)
+pl.testing.assert_frame_equal(
+ sort(pl.DataFrame({'x': [3, 1, 2]}), ['x']),
+ pl.DataFrame({'x': [1, 2, 3]}),
+)
+pl.testing.assert_series_equal(
+ sort(pl.Series('x', [3, 1, 2])),
+ pl.Series('x', [1, 2, 3])
+)
+```
+
+
+```python
+test_eq(_multiply_pl_freq('1d', 4), '4d')
+test_eq(_multiply_pl_freq('2d', 4), '8d')
+pl.testing.assert_series_equal(
+ _multiply_pl_freq('1d', pl_Series([1, 2])),
+ pl_Series(['1d', '2d']),
+)
+pl.testing.assert_series_equal(
+ _multiply_pl_freq('4m', pl_Series([2, 4])),
+ pl_Series(['8m', '16m']),
+)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### offset_times
+
+> ``` text
+> offset_times (times:Union[pandas.core.series.Series,polars.series.series.
+> Series,pandas.core.indexes.base.Index],
+> freq:Union[int,str,pandas._libs.tslibs.offsets.BaseOffset],
+> n:Union[int,numpy.ndarray])
+> ```
+
+
+```python
+pd.testing.assert_index_equal(
+ offset_times(pd.to_datetime(['2020-01-31', '2020-02-29', '2020-03-31']), pd.offsets.MonthEnd(), 1),
+ pd.Index(pd.to_datetime(['2020-02-29', '2020-03-31', '2020-04-30'])),
+)
+pd.testing.assert_index_equal(
+ offset_times(pd.to_datetime(['2020-01-01', '2020-02-01', '2020-03-01']), pd.offsets.MonthBegin(), 1),
+ pd.Index(pd.to_datetime(['2020-02-01', '2020-03-01', '2020-04-01'])),
+)
+```
+
+
+```python
+pl.testing.assert_series_equal(
+ offset_times(pl_Series([dt(2020, 1, 31), dt(2020, 2, 28), dt(2020, 3, 31)]), '1mo', 1),
+ pl_Series([dt(2020, 2, 29), dt(2020, 3, 28), dt(2020, 4, 30)]),
+)
+pl.testing.assert_series_equal(
+ offset_times(pl_Series([dt(2020, 1, 31), dt(2020, 2, 29), dt(2020, 3, 31)]), '1mo', 1),
+ pl_Series([dt(2020, 2, 29), dt(2020, 3, 31), dt(2020, 4, 30)]),
+)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### offset_dates
+
+> ``` text
+> offset_dates (dates:Union[pandas.core.series.Series,polars.series.series.
+> Series,pandas.core.indexes.base.Index],
+> freq:Union[int,str,pandas._libs.tslibs.offsets.BaseOffset],
+> n:Union[int,pandas.core.series.Series,polars.series.series.
+> Series])
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### time_ranges
+
+> ``` text
+> time_ranges (starts:Union[pandas.core.series.Series,polars.series.series.
+> Series,pandas.core.indexes.base.Index],
+> freq:Union[int,str,pandas._libs.tslibs.offsets.BaseOffset],
+> periods:int)
+> ```
+
+
+```python
+# datetimes
+dates = pd.to_datetime(['2000-01-01', '2010-10-10'])
+pd.testing.assert_series_equal(
+ time_ranges(dates, freq='D', periods=3),
+ pd.Series(pd.to_datetime(['2000-01-01', '2000-01-02', '2000-01-03', '2010-10-10', '2010-10-11', '2010-10-12']))
+)
+pd.testing.assert_series_equal(
+ time_ranges(dates, freq='2D', periods=3),
+ pd.Series(pd.to_datetime(['2000-01-01', '2000-01-03', '2000-01-05', '2010-10-10', '2010-10-12', '2010-10-14']))
+)
+pd.testing.assert_series_equal(
+ time_ranges(dates, freq='4D', periods=3),
+ pd.Series(pd.to_datetime(['2000-01-01', '2000-01-05', '2000-01-09', '2010-10-10', '2010-10-14', '2010-10-18']))
+)
+pd.testing.assert_series_equal(
+ time_ranges(pd.to_datetime(['2000-01-01', '2010-10-01']), freq=2 * pd.offsets.MonthBegin(), periods=2),
+ pd.Series(pd.to_datetime(['2000-01-01', '2000-03-01', '2010-10-01', '2010-12-01']))
+)
+pd.testing.assert_series_equal(
+ time_ranges(pd.to_datetime(['2000-01-01', '2010-01-01']).tz_localize('US/Eastern'), freq=2 * pd.offsets.YearBegin(), periods=2),
+ pd.Series(pd.to_datetime(['2000-01-01', '2002-01-01', '2010-01-01', '2012-01-01']).tz_localize('US/Eastern'))
+)
+pd.testing.assert_series_equal(
+ time_ranges(pd.to_datetime(['2000-12-31', '2010-12-31']), freq=2 * pd.offsets.YearEnd(), periods=2),
+ pd.Series(pd.to_datetime(['2000-12-31', '2002-12-31', '2010-12-31', '2012-12-31']))
+)
+# ints
+dates = pd.Series([1, 10])
+pd.testing.assert_series_equal(
+ time_ranges(dates, freq=1, periods=3),
+ pd.Series([1, 2, 3, 10, 11, 12])
+)
+pd.testing.assert_series_equal(
+ time_ranges(dates, freq=2, periods=3),
+ pd.Series([1, 3, 5, 10, 12, 14])
+)
+pd.testing.assert_series_equal(
+ time_ranges(dates, freq=4, periods=3),
+ pd.Series([1, 5, 9, 10, 14, 18])
+)
+```
+
+
+```python
+# datetimes
+dates = pl.Series([dt(2000, 1, 1), dt(2010, 10, 10)])
+pl.testing.assert_series_equal(
+ time_ranges(dates, freq='1d', periods=3),
+ pl.Series([dt(2000, 1, 1), dt(2000, 1, 2), dt(2000, 1, 3), dt(2010, 10, 10), dt(2010, 10, 11), dt(2010, 10, 12)])
+)
+pl.testing.assert_series_equal(
+ time_ranges(dates, freq='2d', periods=3),
+ pl.Series([dt(2000, 1, 1), dt(2000, 1, 3), dt(2000, 1, 5), dt(2010, 10, 10), dt(2010, 10, 12), dt(2010, 10, 14)])
+)
+pl.testing.assert_series_equal(
+ time_ranges(dates, freq='4d', periods=3),
+ pl.Series([dt(2000, 1, 1), dt(2000, 1, 5), dt(2000, 1, 9), dt(2010, 10, 10), dt(2010, 10, 14), dt(2010, 10, 18)])
+)
+pl.testing.assert_series_equal(
+ time_ranges(pl.Series([dt(2010, 2, 28), dt(2000, 1, 31)]), '1mo', 3),
+ pl.Series([dt(2010, 2, 28), dt(2010, 3, 31), dt(2010, 4, 30), dt(2000, 1, 31), dt(2000, 2, 29), dt(2000, 3, 31)])
+)
+# dates
+dates = pl.Series([datetime.date(2000, 1, 1), datetime.date(2010, 10, 10)])
+pl.testing.assert_series_equal(
+ time_ranges(dates, freq='1d', periods=2),
+ pl.Series([
+ datetime.date(2000, 1, 1), datetime.date(2000, 1, 2),
+ datetime.date(2010, 10, 10), datetime.date(2010, 10, 11),
+ ])
+)
+# ints
+dates = pl.Series([1, 10])
+pl.testing.assert_series_equal(
+ time_ranges(dates, freq=1, periods=3),
+ pl.Series([1, 2, 3, 10, 11, 12]),
+)
+pl.testing.assert_series_equal(
+ time_ranges(dates, freq=2, periods=3),
+ pl.Series([1, 3, 5, 10, 12, 14]),
+)
+pl.testing.assert_series_equal(
+ time_ranges(dates, freq=4, periods=3),
+ pl.Series([1, 5, 9, 10, 14, 18]),
+)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### repeat
+
+> ``` text
+> repeat (s:Union[pandas.core.series.Series,polars.series.series.Series,pan
+> das.core.indexes.base.Index,numpy.ndarray], n:Union[int,numpy.nda
+> rray,pandas.core.series.Series,polars.series.series.Series])
+> ```
+
+
+```python
+pd.testing.assert_index_equal(
+ repeat(pd.CategoricalIndex(['a', 'b', 'c'], categories=['a', 'b', 'c']), 2),
+ pd.CategoricalIndex(['a', 'a', 'b', 'b', 'c', 'c'], categories=['a', 'b', 'c'])
+)
+pd.testing.assert_series_equal(
+ repeat(pd.Series([1, 2]), 2),
+ pd.Series([1, 1, 2, 2])
+)
+pd.testing.assert_series_equal(
+ repeat(pd.Series([1, 2]), pd.Series([2, 3])),
+ pd.Series([1, 1, 2, 2, 2]),
+)
+np.testing.assert_array_equal(
+ repeat(np.array([np.datetime64('2000-01-01'), np.datetime64('2010-10-10')]), 2),
+ np.array([
+ np.datetime64('2000-01-01'), np.datetime64('2000-01-01'),
+ np.datetime64('2010-10-10'), np.datetime64('2010-10-10')
+ ])
+)
+np.testing.assert_array_equal(
+ repeat(np.array([1, 2]), np.array([2, 3])),
+ np.array([1, 1, 2, 2, 2]),
+)
+```
+
+
+```python
+s = pl.Series(['a', 'b', 'c'], dtype=pl.Categorical)
+pl.testing.assert_series_equal(
+ repeat(s, 2),
+ pl.concat([s, s]).sort()
+)
+pl.testing.assert_series_equal(
+ repeat(pl.Series([2, 4]), 2),
+ pl.Series([2, 2, 4, 4])
+)
+pl.testing.assert_series_equal(
+ repeat(pl.Series([1, 2]), np.array([2, 3])),
+ pl.Series([1, 1, 2, 2, 2]),
+)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### cv_times
+
+> ``` text
+> cv_times (times:numpy.ndarray, uids:Union[pandas.core.series.Series,polar
+> s.series.series.Series,pandas.core.indexes.base.Index],
+> indptr:numpy.ndarray, h:int, test_size:int, step_size:int,
+> id_col:str='unique_id', time_col:str='ds')
+> ```
+
+
+```python
+times = np.arange(51, dtype=np.int64)
+uids = pd.Series(['id_0'])
+indptr = np.array([0, 51])
+h = 3
+test_size = 5
+actual = cv_times(
+ times=times,
+ uids=uids,
+ indptr=indptr,
+ h=h,
+ test_size=test_size,
+ step_size=1,
+)
+expected = pd.DataFrame({
+ 'unique_id': 9 * ['id_0'],
+ 'ds': np.hstack([
+ [46, 47, 48],
+ [47, 48, 49],
+ [48, 49, 50]
+ ], dtype=np.int64),
+ 'cutoff': np.repeat(np.array([45, 46, 47], dtype=np.int64), h),
+})
+pd.testing.assert_frame_equal(actual, expected)
+
+# step_size=2
+actual = cv_times(
+ times=times,
+ uids=uids,
+ indptr=indptr,
+ h=h,
+ test_size=test_size,
+ step_size=2,
+)
+expected = pd.DataFrame({
+ 'unique_id': 6 * ['id_0'],
+ 'ds': np.hstack([
+ [46, 47, 48],
+ [48, 49, 50]
+ ], dtype=np.int64),
+ 'cutoff': np.repeat(np.array([45, 47], dtype=np.int64), h)
+})
+pd.testing.assert_frame_equal(actual, expected)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### group_by
+
+> ``` text
+> group_by (df:Union[pandas.core.series.Series,polars.series.series.Series,
+> pandas.core.frame.DataFrame,polars.dataframe.frame.DataFrame],
+> by, maintain_order=False)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### group_by_agg
+
+> ``` text
+> group_by_agg
+> (df:Union[pandas.core.frame.DataFrame,polars.dataframe.fram
+> e.DataFrame], by, aggs, maintain_order=False)
+> ```
+
+
+```python
+pd.testing.assert_frame_equal(
+ group_by_agg(pd.DataFrame({'x': [1, 1, 2], 'y': [1, 1, 1]}), 'x', {'y': 'sum'}),
+ pd.DataFrame({'x': [1, 2], 'y': [2, 1]})
+)
+```
+
+
+```python
+pd.testing.assert_frame_equal(
+ group_by_agg(pl.DataFrame({'x': [1, 1, 2], 'y': [1, 1, 1]}), 'x', {'y': 'sum'}, maintain_order=True).to_pandas(),
+ pd.DataFrame({'x': [1, 2], 'y': [2, 1]})
+)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### is_in
+
+> ``` text
+> is_in (s:Union[pandas.core.series.Series,polars.series.series.Series],
+> collection)
+> ```
+
+
+```python
+np.testing.assert_equal(is_in(pd.Series([1, 2, 3]), [1]), np.array([True, False, False]))
+```
+
+
+```python
+np.testing.assert_equal(is_in(pl.Series([1, 2, 3]), [1]), np.array([True, False, False]))
+```
+
+------------------------------------------------------------------------
+
+source
+
+### between
+
+> ``` text
+> between (s:Union[pandas.core.series.Series,polars.series.series.Series],
+> lower:Union[pandas.core.series.Series,polars.series.series.Serie
+> s], upper:Union[pandas.core.series.Series,polars.series.series.S
+> eries])
+> ```
+
+
+```python
+np.testing.assert_equal(
+ between(pd.Series([1, 2, 3]), pd.Series([0, 1, 4]), pd.Series([4, 1, 2])),
+ np.array([True, False, False]),
+)
+```
+
+
+```python
+np.testing.assert_equal(
+ between(pl.Series([1, 2, 3]), pl.Series([0, 1, 4]), pl.Series([4, 1, 2])),
+ np.array([True, False, False]),
+)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### fill_null
+
+> ``` text
+> fill_null
+> (df:Union[pandas.core.frame.DataFrame,polars.dataframe.frame.D
+> ataFrame], mapping:Dict[str,Any])
+> ```
+
+
+```python
+pd.testing.assert_frame_equal(
+ fill_null(pd.DataFrame({'x': [1, np.nan], 'y': [np.nan, 2]}), {'x': 2, 'y': 1}),
+ pd.DataFrame({'x': [1, 2], 'y': [1, 2]}, dtype='float64')
+)
+```
+
+
+```python
+pl.testing.assert_frame_equal(
+ fill_null(pl.DataFrame({'x': [1, None], 'y': [None, 2]}), {'x': 2, 'y': 1}),
+ pl.DataFrame({'x': [1, 2], 'y': [1, 2]})
+)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### cast
+
+> ``` text
+> cast (s:Union[pandas.core.series.Series,polars.series.series.Series],
+> dtype:type)
+> ```
+
+
+```python
+pd.testing.assert_series_equal(
+ cast(pd.Series([1, 2, 3]), 'int16'),
+ pd.Series([1, 2, 3], dtype='int16')
+)
+```
+
+
+```python
+pd.testing.assert_series_equal(
+ cast(pl.Series('x', [1, 2, 3]), pl.Int16).to_pandas(),
+ pd.Series([1, 2, 3], name='x', dtype='int16')
+)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### value_cols_to_numpy
+
+> ``` text
+> value_cols_to_numpy
+> (df:Union[pandas.core.frame.DataFrame,polars.datafra
+> me.frame.DataFrame], id_col:str, time_col:str,
+> target_col:Optional[str])
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### make_future_dataframe
+
+> ``` text
+> make_future_dataframe
+> (uids:Union[pandas.core.series.Series,polars.serie
+> s.series.Series], last_times:Union[pandas.core.ser
+> ies.Series,polars.series.series.Series,pandas.core
+> .indexes.base.Index], freq:Union[int,str,pandas._l
+> ibs.tslibs.offsets.BaseOffset], h:int,
+> id_col:str='unique_id', time_col:str='ds')
+> ```
+
+
+```python
+pd.testing.assert_frame_equal(
+ make_future_dataframe(
+ pd.Series([1, 2]), pd.to_datetime(['2000-01-01', '2010-10-10']), freq='D', h=2
+ ),
+ pd.DataFrame({
+ 'unique_id': [1, 1, 2, 2],
+ 'ds': pd.to_datetime(['2000-01-02', '2000-01-03', '2010-10-11', '2010-10-12'])
+ })
+)
+```
+
+
+```python
+pl.testing.assert_frame_equal(
+ make_future_dataframe(
+ pl.Series([1, 2]),
+ pl.Series([dt(2000, 1, 1), dt(2010, 10, 10)]),
+ freq='1d',
+ h=2,
+ id_col='uid',
+ time_col='dates',
+ ),
+ pl.DataFrame({
+ 'uid': [1, 1, 2, 2],
+ 'dates': [dt(2000, 1, 2), dt(2000, 1, 3), dt(2010, 10, 11), dt(2010, 10, 12)]
+ })
+)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### anti_join
+
+> ``` text
+> anti_join
+> (df1:Union[pandas.core.frame.DataFrame,polars.dataframe.frame.
+> DataFrame], df2:Union[pandas.core.frame.DataFrame,polars.dataf
+> rame.frame.DataFrame], on:Union[str,List[str]])
+> ```
+
+
+```python
+pd.testing.assert_frame_equal(
+ anti_join(pd.DataFrame({'x': [1, 2]}), pd.DataFrame({'x': [1]}), on='x'),
+ pd.DataFrame({'x': [2]})
+)
+test_eq(
+ anti_join(pd.DataFrame({'x': [1]}), pd.DataFrame({'x': [1]}), on='x').shape[0],
+ 0,
+)
+```
+
+
+```python
+pl.testing.assert_frame_equal(
+ anti_join(pl_DataFrame({'x': [1, 2]}), pl_DataFrame({'x': [1]}), on='x'),
+ pl_DataFrame({'x': [2]})
+)
+test_eq(
+ anti_join(pl_DataFrame({'x': [1]}), pl_DataFrame({'x': [1]}), on='x').shape[0],
+ 0,
+)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### ensure_sorted
+
+> ``` text
+> ensure_sorted
+> (df:Union[pandas.core.frame.DataFrame,polars.dataframe.fra
+> me.DataFrame], id_col:str, time_col:str)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### process_df
+
+> ``` text
+> process_df
+> (df:Union[pandas.core.frame.DataFrame,polars.dataframe.frame.
+> DataFrame], id_col:str, time_col:str,
+> target_col:Optional[str])
+> ```
+
+*Extract components from dataframe*
+
+| | **Type** | **Details** |
+|--------|---------------------------|-------------------------------------|
+| df | Union | Input dataframe with id, times and target values. |
+| id_col | str | |
+| time_col | str | |
+| target_col | Optional | |
+| **Returns** | **ProcessedDF** | **serie with the sorted unique ids present in the data.** |
+
+------------------------------------------------------------------------
+
+source
+
+### ProcessedDF
+
+> ``` text
+> ProcessedDF
+> (uids:Union[pandas.core.series.Series,polars.series.series.S
+> eries], last_times:numpy.ndarray, data:numpy.ndarray,
+> indptr:numpy.ndarray, sort_idxs:Optional[numpy.ndarray])
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### DataFrameProcessor
+
+> ``` text
+> DataFrameProcessor (id_col:str='unique_id', time_col:str='ds',
+> target_col:str='y')
+> ```
+
+*Initialize self. See help(type(self)) for accurate signature.*
+
+
+```python
+static_features = ['static_0', 'static_1']
+```
+
+
+```python
+for n_static_features in [0, 2]:
+ series_pd = generate_series(1_000, n_static_features=n_static_features, equal_ends=False, engine='pandas')
+ for i in range(n_static_features):
+ series_pd[f'static_{i}'] = series_pd[f'static_{i}'].map(lambda x: f'x_{x}').astype('category')
+ scrambled_series_pd = series_pd.sample(frac=1.0)
+ dfp = DataFrameProcessor('unique_id', 'ds', 'y')
+ uids, times, data, indptr, _ = dfp.process(scrambled_series_pd)
+ test_eq(times, series_pd.groupby('unique_id', observed=True)['ds'].max().values)
+ test_eq(uids, np.sort(series_pd['unique_id'].unique()))
+ for i in range(n_static_features):
+ series_pd[f'static_{i}'] = series_pd[f'static_{i}'].cat.codes
+ test_eq(data, series_pd[['y'] + static_features[:n_static_features]].to_numpy())
+ test_eq(np.diff(indptr), series_pd.groupby('unique_id', observed=True).size().values)
+```
+
+
+```python
+for n_static_features in [0, 2]:
+ series_pl = generate_series(1_000, n_static_features=n_static_features, equal_ends=False, engine='polars')
+ scrambled_series_pl = series_pl.sample(fraction=1.0, shuffle=True)
+ dfp = DataFrameProcessor('unique_id', 'ds', 'y')
+ uids, times, data, indptr, _ = dfp.process(scrambled_series_pl)
+ grouped = group_by(series_pl, 'unique_id')
+ test_eq(times, grouped.agg(pl.col('ds').max()).sort('unique_id')['ds'].to_numpy())
+ test_eq(uids, series_pl['unique_id'].unique().sort())
+ test_eq(data, series_pl.select(pl.col(c).map_batches(lambda s: s.to_physical()) for c in ['y'] + static_features[:n_static_features]).to_numpy())
+ test_eq(np.diff(indptr), grouped.count().sort('unique_id')['count'].to_numpy())
+```
+
+------------------------------------------------------------------------
+
+source
+
+### backtest_splits
+
+> ``` text
+> backtest_splits
+> (df:Union[pandas.core.frame.DataFrame,polars.dataframe.f
+> rame.DataFrame], n_windows:int, h:int, id_col:str,
+> time_col:str, freq:Union[int,str,pandas._libs.tslibs.off
+> sets.BaseOffset], step_size:Optional[int]=None,
+> input_size:Optional[int]=None,
+> allow_partial_horizons:bool=False)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### add_insample_levels
+
+> ``` text
+> add_insample_levels
+> (df:Union[pandas.core.frame.DataFrame,polars.datafra
+> me.frame.DataFrame], models:List[str],
+> level:List[Union[int,float]],
+> id_col:str='unique_id', target_col:str='y')
+> ```
+
+
+```python
+series = generate_series(100, n_models=2)
+models = ['model0', 'model1']
+levels = [80, 95]
+with_levels = add_insample_levels(series, models, levels)
+for model in models:
+ for lvl in levels:
+ assert with_levels[f'{model}-lo-{lvl}'].lt(with_levels[f'{model}-hi-{lvl}']).all()
+```
+
+
+```python
+series_pl = generate_series(100, n_models=2, engine='polars')
+with_levels_pl = add_insample_levels(series_pl, ['model0', 'model1'], [80, 95])
+pd.testing.assert_frame_equal(
+ with_levels.drop(columns='unique_id'),
+ with_levels_pl.to_pandas().drop(columns='unique_id')
+)
+```
+
diff --git a/utilsforecast/validation.html.mdx b/utilsforecast/validation.html.mdx
new file mode 100644
index 00000000..be619116
--- /dev/null
+++ b/utilsforecast/validation.html.mdx
@@ -0,0 +1,150 @@
+---
+description: Utilities to validate input data
+output-file: validation.html
+title: Validation
+---
+
+
+
+```python
+import datetime
+
+from fastcore.test import test_eq, test_fail
+```
+
+
+```python
+import polars.testing
+```
+
+------------------------------------------------------------------------
+
+source
+
+### ensure_shallow_copy
+
+> ``` text
+> ensure_shallow_copy (df:pandas.core.frame.DataFrame)
+> ```
+
+------------------------------------------------------------------------
+
+source
+
+### ensure_time_dtype
+
+> ``` text
+> ensure_time_dtype (df:~DFType, time_col:str='ds')
+> ```
+
+*Make sure that `time_col` contains timestamps or integers. If it
+contains strings, try to cast them as timestamps.*
+
+
+```python
+pd.testing.assert_frame_equal(
+ ensure_time_dtype(pd.DataFrame({'ds': ['2000-01-01']})),
+ pd.DataFrame({'ds': pd.to_datetime(['2000-01-01'])})
+)
+df = pd.DataFrame({'ds': [1, 2]})
+assert df is ensure_time_dtype(df)
+test_fail(
+ lambda: ensure_time_dtype(pd.DataFrame({'ds': ['2000-14-14']})),
+ contains='Please make sure that it contains valid timestamps',
+)
+```
+
+
+```python
+pl.testing.assert_frame_equal(
+ ensure_time_dtype(pl.DataFrame({'ds': ['2000-01-01']})),
+ pl.DataFrame().with_columns(ds=pl.datetime(2000, 1, 1))
+)
+df = pl.DataFrame({'ds': [1, 2]})
+assert df is ensure_time_dtype(df)
+test_fail(
+ lambda: ensure_time_dtype(pl.DataFrame({'ds': ['hello']})),
+ contains='Please make sure that it contains valid timestamps',
+)
+```
+
+------------------------------------------------------------------------
+
+source
+
+### validate_format
+
+> ``` text
+> validate_format
+> (df:Union[pandas.core.frame.DataFrame,polars.dataframe.f
+> rame.DataFrame], id_col:str='unique_id',
+> time_col:str='ds', target_col:Optional[str]='y')
+> ```
+
+*Ensure DataFrame has expected format.*
+
+| | **Type** | **Default** | **Details** |
+|-------------|----------|-------------|--------------------------------------------|
+| df | Union | | DataFrame with time series in long format. |
+| id_col | str | unique_id | Column that identifies each serie. |
+| time_col | str | ds | Column that identifies each timestamp. |
+| target_col | Optional | y | Column that contains the target. |
+| **Returns** | **None** | | |
+
+
+```python
+import datetime
+
+from utilsforecast.compat import POLARS_INSTALLED, pl
+from utilsforecast.data import generate_series
+```
+
+
+```python
+test_fail(lambda: validate_format(1), contains="got ")
+constructors = [pd.DataFrame]
+if POLARS_INSTALLED:
+ constructors.append(pl.DataFrame)
+for constructor in constructors:
+ df = constructor({'unique_id': [1]})
+ test_fail(lambda: validate_format(df), contains="missing: ['ds', 'y']")
+ df = constructor({'unique_id': [1], 'time': ['x'], 'y': [1]})
+ test_fail(lambda: validate_format(df, time_col='time'), contains="('time') should have either timestamps or integers")
+ for time in [1, datetime.datetime(2000, 1, 1)]:
+ df = constructor({'unique_id': [1], 'ds': [time], 'sales': ['x']})
+ test_fail(lambda: validate_format(df, target_col='sales'), contains="('sales') should have a numeric data type")
+```
+
+------------------------------------------------------------------------
+
+source
+
+### validate_freq
+
+> ``` text
+> validate_freq
+> (times:Union[pandas.core.series.Series,polars.series.serie
+> s.Series], freq:Union[str,int])
+> ```
+
+
+```python
+test_fail(lambda: validate_freq(pd.Series([1, 2]), 'D'), contains='provide a valid integer')
+test_fail(lambda: validate_freq(pd.to_datetime(['2000-01-01']).to_series(), 1), contains='provide a valid pandas or polars offset')
+```
+
+
+```python
+test_fail(lambda: validate_freq(pl.Series([1, 2]), '1d'), contains='provide a valid integer')
+test_fail(lambda: validate_freq(pl.Series([datetime.datetime(2000, 1, 1)]), 1), contains='provide a valid pandas or polars offset')
+test_fail(lambda: validate_freq(pl.Series([datetime.datetime(2000, 1, 1)]), 'D'), contains='valid polars offset')
+```
+